Abstract
The rhizosphere is a critical hotspot for metabolic activities involving arsenic (As). While recent studies indicate many functions for soil viruses, much remains overlooked regarding their quantitative impact on rhizosphere processes. Here, we analyze time-series metagenomes of rice (Oryza sativa L.)rhizosphere and bulk soil to explore how viruses mediate rhizosphere As biogeochemistry. We observe the rhizosphere favors lysogeny in viruses associated with As-oxidizing microbes, with a positive correlation between As oxidation and the prevalence of these microbial hosts. Moreover, results demonstrate these lysogenic viruses enrich both As oxidation and phosphorus co-metabolism genes and mediated horizontal gene transfers (HGTs) of As oxidases. In silico simulation with genome-scale metabolic models (GEMs) and in vitro validation with experiments estimate that rhizosphere lysogenic viruses contribute up to 25% of microbial As oxidation. These findings enhance our comprehension of the plant-microbiome-virome interplay and highlight the potential of rhizosphere viruses for improving soil health in sustainable agriculture.
Similar content being viewed by others
Introduction
The rhizosphere is a crucial zone for plant-soil interactions, where the release of labile nutrients by roots fosters rapid microbial proliferation and evolution1. This root-triggered activity not only broadens the genetic and functional repertoire of the soil ecosystem but also enhances plant adaptability2. Large-scale studies show that the impact of the rhizosphere on microbiome composition and function often surpasses influences from soil type, plant species, and climate3. Beyond these microbes, viruses are abundant biological entities in soils, with their particles outnumbering microbes tenfold4. Moreover, viruses have been reported to exhibit greater sensitivity to changes in soil factors compared to bacteria, potentially serving as crucial biological regulators of microbial survival in response to environmental stress and fluctuations5. However, the roles that soil viruses play in rhizosphere ecology remain a key unresolved question. A few insights from previous research suggest that the impact of viruses has been underestimated6. For instance, significantly increased viral activity observed in the wheat rhizosphere indicates that it functions beyond a mere microbial hotspot7. Additionally, the rhizosphere’s enhanced nutrient flow and microbial activity likely lead to more intricate interactions between viruses and their host cells, profoundly influencing the evolution and ecology of the microbiota8. Rhizosphere microbes serve as accelerators in soil biogeochemical cycles, while viruses can “reprogram” host metabolism by providing virus-encoded auxiliary metabolic genes (AMGs) through lysogenic conversion9. These dynamics may potentially drive rhizosphere processes, yet the extent and specific mechanisms of their influence remain unclear.
Arsenic (As) is a highly toxic metalloid and accumulates in soils worldwide, posing significant public health concerns for hundreds of millions globally through the food chain10. By 2040, 18.3% of rice-growing areas in China are predicted to face threats of As contamination exceeding safety limit11. Arsenite [As(III)] and arsenate [As(V)] are the most prevalent forms in soils, with As(V) being significantly less toxic and mobile than As(III)12. Rice (Oryza sativa L.) can selectively recruit a phylogenetically diverse microbiota involved in As oxidation from the bulk soil pool by releasing exudates and oxygen into the rhizosphere13. Consequently, the rice rhizosphere, a metabolic hotspot for As processes, provides an essential model for investigating strategies to mitigate As risk. Recent studies have shown that soil viruses can facilitate As biotransformation by mediating arsM gene transduction, thereby improving microbial adaptability14. This highlights the role of soil viruses as vectors in the co-evolutionary dynamics of As biotransformation. Due to the increased connectivity among soil microaggregates from flooding, the rice rhizosphere environment presents unique challenges and fosters tighter interactions with viruses15. This raises the question: could rhizosphere conditions further amplify the remodeling effect of soil viruses on host microbiota? Despite their potential, few studies have explored the role of soil viruses in regulating microbial As transformation in the rhizosphere, a critical aspect of As biogeochemical processes.
Here, we collected rhizosphere and bulk soils bi-daily time-series sampling over a 100-day rice growth season (Fig. 1a) to assess the viral impact on As biotransformation in both the rhizosphere and bulk soil under As contamination. The metagenomic analysis indicated a positive correlation between As-oxidizing microbes, for which viruses were identified, and As oxidation within the dynamic rhizosphere environment. By characterizing the profiles of viral lifestyle, AMGs and horizontal gene transfers (HGTs), we further investigated the mechanisms of viral influence on As oxidation. Moreover, we constructed genome-scale metabolic models (GEMs) for lysogens and host cells to in silico validate the metabolic reprogramming by rhizosphere viruses in As oxidation. Follow-up in vitro batch experiments using mitomycin-C (MC) induced virome quantified the contribution of rhizosphere lysogenic viruses to microbial As oxidation. By comprehensively understanding the interactions between viruses and As-transforming microbes in the rhizosphere, our study provides insights into the plant-microbiome-virome crosstalk-driven biogeochemical processes.
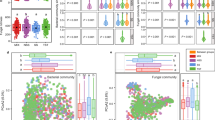
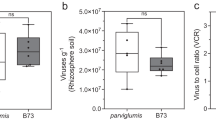
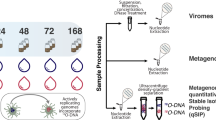
a Experimental design and bioinformatic workflow. The bioinformatic workflow is created with BioRender.com. b Relative change in As(III) proportion in the rhizosphere compared to bulk soil. c Enrichment ratio of As-transforming genes in the rhizosphere compared to bulk soil. The enrichment ratio was determined using the formula: Enrichment ratiocopies\(=\frac{{{Copies}}_{R}}{{{Copies}}_{B}}\). b, c The dashed line represents a value of 0. The regression line shown is a Loess estimate (n = 50). The shaded interval represents the 95% confidence interval, presenting the variability of the fit over time. d Abundance of As-oxidizing microbes for which viruses were identified (up) and As-reducing microbes for which viruses were identified (down) in the rhizosphere and bulk soil. The dots represent the microbial abundance at each time point. The dashed regression line shown is a Loess estimate (n = 50). The shaded interval represents the 95% confidence interval, presenting the variability of the fit over time. Statistical significance between the rhizosphere and bulk soil was determined using the two-sided Student’s t-test (up) and two-sided Welch’s t-test (down). e Distribution (up) and classification (down) of viruses associated with As oxidizing and As reducing microbes. The bipartite network (right) represents the correspondence between As-transforming genera (top 30) and vOTUs, where annotated classified vOTUs are highlighted in red circles and labeled with their corresponding family. f Correlation between the abundance of overall and virus-associated As-oxidizing microbes and the relative change in the proportion of As(III) in the rhizosphere compared to bulk soil. The yellow and red lines represent the linear regression models with 95% confidence interval (shaded band). Statistical significance for regression coefficients was assessed using a two-sided t-test (n = 50).
Results
Arsenic-transforming microbes with identified viruses
Figure 1a showed the whole experimental and analysis design, including time-series sampling and bioinformatic workflow. Based on our results, inorganic As species (As(V) and As(III)) dominate the As dynamics in the rhizosphere (accounting for at least 97%), as shown in Supplementary Fig. 1. The rhizosphere significantly promoted As oxidation, continuously reducing the proportion of As(III) throughout the rice vegetative stage (Supplementary Fig. 2a). Notably, during the elongation stage, As(III) proportion in the rhizosphere, relative to the bulk soil, decreased by up to 20% at most (Fig. 1b).
The abundances of As(III) oxidase genes (aioA, arxA, arxS, aioB, and arsH) in the rhizosphere were significantly higher than those in the bulk soil (Welch’s t-test, n = 50, p < 0.0001, Fig. 1c and Supplementary Fig. 2b). However, the abundance of As(V) reductase gene, arrB, in the rhizosphere was consistently lower than in the bulk soil (Fig. 1c). Previous studies have shown that nearly all microbes possess resistance pathways for inorganic arsenic12, including the reduction of As(V)16, as evidenced by the stable presence of arsC throughout the experimental period (Fig. 1c). Therefore, we define microbes in which inorganic arsenic resistance genes were detected but that lack As(III) oxidation function as As(V)-reducing microbes. Further analysis revealed that both the relative and absolute abundances of the As-oxidizing microbes in the rhizosphere were consistently higher than those in the bulk soil (Supplementary Figs. 3 and 4). Conversely, the relative and absolute abundances of As-reducing microbes in the rhizosphere remained similar to those in the bulk soil throughout the entire rice-growing season (Supplementary Figs. 3 and 4).
To ascertain the potential contribution of viruses in As redox processes in the rhizosphere, we further identified As-transforming microbes associated with identified viruses through host prediction (Supplementary Data 4). Although the abundance of As-transforming microbes for which viruses were identified in both the rhizosphere and bulk soil increased at the early stage, the abundance of As-oxidizing microbes for which viruses were identified in the rhizosphere was increased up to two times at the elongation stage than those in the bulk soil (Fig. 1d). Furthermore, the abundance of As-reducing microbes for which viruses were identified showed no significant difference between the rhizosphere and bulk soil during the growth season (Welch’s t-test, n = 50, p = 0.830, Fig. 1d).
We further identified the distinct taxonomic profiles of As-oxidizing microbes for which viruses were identified (Supplementary Fig. 5). Notably, taxa such as Terracidiphilus (7.61%), PCC-7407 (5.59%), and Gemmatirosa (4.94%), which represented the dominant among As-oxidizing microbial OTUs with identified viruses, were not the most abundant in the overall As-oxidizing OTUs. Furthermore, these dominant genera in As-oxidizing microbial OTUs with identified viruses were not the most abundant in the As-reducing microbial OTUs with identified viruses, emphasizing the distinct compositional differences between these As-transforming microbes with identified viruses (Supplementary Fig. 5).
Delving deeper into the viruses associated with the As-transforming microbes, we found that only 1.13% of these vOTUs had the association pattern with both As-oxidizing and As-reducing microbes (Supplementary Fig. 6). Then, we assigned the classification of As-transforming microbes with identified viruses at the family level, finding that only 5.38% of vOTUs were assigned to known phage families (Fig. 1e and Supplementary Data 6). Among identified viruses associated with As-oxidizing microbes, the majority of the vOTUs were identified from the Autographiviridae (50.00%) and Kyanoviridae (50.00%) families. In contrast, the majority of vOTUs associated with As reduction were assigned to the Straboviridae (30.19%), Peduoviridae (20.75%), Herelleviridae (16.98%), and Zierdtviridae (13.21%) families. Our virus-host bipartite network further highlighted that the viruses associated with As-oxidizing and As-reducing microbes (top 30) were distinct viral communities (Fig. 1e).
To explore the distribution of viruses associated with As-transforming microbes in soils across the globe, we analyzed the association between these viruses and the global soil virome dataset (GSV)17. Firstly, we found that 17.34% of the vOTUs matched species-level GSV (Supplementary Data 8). We then determined the abundance of these GSV-matched viruses in 1756 global soil metagenomes (Supplementary Data 9), finding that they were detected in soil metagenomes from diverse habitats worldwide (Supplementary Fig. 7a). Notably, their prevalence in paddy fields and wetlands was significantly higher than those in non-paddy fields, forests, and grassland habitats (Supplementary Fig. 7b). These results highlight the environmental ubiquity of viruses associated with As-transforming microbes, particularly in relatively anaerobic soils such as paddy soils, where they are likely to play a greater role in mediating microbial As transformation processes. Based on these findings, we further explored the broader ecological implications of virus-host interactions in the rhizosphere. Interestingly, we find that the relative decrease of As(III) proportion in the rhizosphere was significantly correlated with the abundance of As-oxidizing microbes for which viruses were identified (R = −0.65, p < 0.0001, n = 50), rather than with the abundance of entire As-oxidizing microbes (R = 0.11, p = 0.46, n = 50), suggesting the potential contribution of viruses in mediating As oxidation in the rhizosphere (Fig. 1f).
Viral lifestyle in rhizosphere
To elucidate the mechanism of viruses in mediating microbial As transformation, we identified the lifestyle of these vOTUs (Supplementary Data 7). The rhizosphere significantly enhanced the abundance of lysogenic viruses (Welch’s t-test, n = 50, p < 0.0001, Fig. 2a) and decreased the prevalence of lytic viruses associated with As-oxidizing microbes (Welch’s t-test, n = 50, p = 0.00194, Fig. 2a). Additionally, the relative decrease in As(III) proportion in the rhizosphere correlated with the abundance of As-oxidizing microbes with identified lysogenic viruses (R = −0.6, p < 0.0001, n = 50, Supplementary Fig. 8), further emphasizing the role of viral lysogeny in facilitating rhizosphere As oxidation. Furthermore, the abundance of viruses associated with As-oxidizing microbes fluctuated with rice growth stages. In the vegetative stage, lysogenic virus abundance increased, while lytic virus levels stayed constant. In the reproductive stage, lysogenic virus levels decreased to the bulk soil levels, and lytic virus levels increased accordingly (Supplementary Fig. 9a). Conversely, for viruses associated with As-reducing microbes, the rhizosphere only decreased the abundance of lytic viruses associated with As-reducing microbes (Welch’s t-test, n = 50, p < 0.0001, Fig. 2b and Supplementary Fig. 9b).
Distribution of the abundance of lysogenic and lytic viruses involved in As oxidation (a) and As reduction (b), viral integrase genes (c), and virus to host abundance ratio (d) for both As oxidation and reduction. a–c (right), d (left) Statistical significances between the rhizosphere and bulk soil were determined using the two-sided Welch’s t-test (n = 50). c (left), d (right) Statistical significances between rhizosphere and bulk soils were determined using the two-sided Student’s t-test (n = 50).
We subsequently determined the abundance of integrase genes, a key indicator of viral lysogeny18 (Supplementary Data 10), which further confirmed the enrichment of lysogenic viruses associated with As-oxidizing microbes in the rhizosphere (Welch’s t-test, n = 50, p < 0.0001, Fig. 2c). Moreover, the trend of rhizosphere viral lysogeny associated with As-oxidizing microbes aligned with the aforementioned findings, peaking during the vegetative stage and declining to the bulk soil levels in reproduction (Supplementary Fig. 9c). However, the abundance of integrase genes in viruses associated with As-reducing microbes showed no significant difference between the rhizosphere and bulk soil (Student’s t-test, n = 50, p = 0.173, Fig. 2c and Supplementary Fig. 9c).
The virus-host ratio (VHR) was further examined to validate their interactions, with a lower value indicating a tendency towards lysogenic interactions19. The VHR for As-oxidizing microbes in the rhizosphere was significantly lower than in the bulk soil (Welch’s t-test, n = 50, p < 0.0001, Fig. 2d), while the VHR for As-reducing microbes was consistent with that in the bulk soil (Student’s t-test, n = 50, p = 0.525, Fig. 2d).
Enriched auxiliary metabolic genes
Given the vital roles of AMGs carried by viruses in lysogenic lifestyle in regulating microbial community functioning20, we identified rhizosphere enriched AMGs of viruses associated with As-oxidizing microbes (Fig. 3a). Those viruses significantly enriched genes involved in As oxidation and phosphorus (P) metabolism. In addition to inorganic P metabolism AMGs, the abundance of As-oxidizing, organic P synthesis and phosphate (Pi)-starvation regulation AMGs in the rhizosphere peaked at the elongation stage (Fig. 3b). Notably, lysogenic viruses in the rhizosphere were also enriched during the elongation stage (Supplementary Fig. 9a), providing cross-validation that lysogenic viruses, rather than lytic viruses, are more likely to carry and enrich As oxidation- and P-related AMGs.
a Enrichment analysis of viral genes associated with As-oxidizing microbes in the rhizosphere. Statistical significance was determined using the Wald test. Each circle represents a gene, with the color indicating the metabolic pathway it belongs to. The right figure displays the counts of enriched genes within these metabolic pathways. b Enrichment ratio of the AMGs pathway in the rhizosphere to the bulk soil. The dashed regression line shown is a Loess estimate (n = 50). The shaded interval represents the 95% confidence interval, presenting the variability of the fit over time. The dashed gray line represents a value of 0. c Protein structure of enriched AMGs associated with As-oxidizing microbes in the rhizosphere. The pTM score measures the accuracy of the entire protein structure. The pLDDT is shown as a per-atom confidence on a 0–100 scale where a higher value indicates higher confidence. d As-oxidizing AMGs abundance in the rhizosphere and bulk soil. e Abundance and composition of host microbes at the genus level with As-oxidizing AMGs in the rhizosphere and bulk soil. f P-metabolism AMGs abundance in the rhizosphere and bulk soil. g Abundance and composition of host microbes at the genus level with P-metabolism AMGs in the rhizosphere and bulk soil.
To confirm that these AMGs related to As oxidation and P metabolism can encode structurally complete proteins capable of performing functions, we predicted their corresponding protein structures using AlphaFold321 (one sequence was selected from each gene family). The results show that the pTM scores of these AMGs are all above 0.5, and most are greater than 0.8 (Fig. 3c), indicating that the predicted overall fold of the complex is similar to the true structure with high confidence21. These include the detection of As oxidase genes (arxS, arsH, and aioB) and certain P metabolism genes (phnK, phnL, phnM, pstS, ugpC, RegX3, and phoP) that can be encoded by viruses, catalyzing As oxidation and P co-metabolism.
Specifically, our time-series data revealed that enriched As oxidase AMGs peaked during the elongation stages of rice (Fig. 3d). We identified 40 microbes spanning 12 genera, dominated by PCC-7407, Gemmatirosa, MT45, Stieleria and Anaeromyxobacter associated with viruses carrying As oxidase genes (Fig. 3e). The abundance of the host microbes in the rhizosphere was also significantly higher than in the bulk soil (Fig. 3e), suggesting that these AMGs might confer adaptation advantages to their hosts in the rhizosphere.
Moreover, the rhizosphere also enriched AMGs associated with P metabolism, including inorganic P metabolism genes (phnK, phnL, phnM and pstS genes), organic phosphorus synthesis genes (ugpC gene) and Pi-starvation regulation genes (RegX3 and phoP genes) (Fig. 3a and Supplementary Fig. 4f). The As-oxidizing microbes with AMGs associated with P metabolism exhibited significantly adaptive advantages in the rhizosphere (Fig. 3g), particularly for Henriciella and Bradyrhizobium, which were associated with viruses carrying AMGs associated with inorganic P metabolism (Supplementary Fig. 10).
Virus-mediated horizontal gene transfers
To assess the potential contribution of virus-mediated HGTs in microbial As oxidation in rhizosphere, we pinpointed 40 instances of virus-to-host transfer of aioB, arsH, and arxS genes by a phylogenetic pipeline for HGT-indicative topologies22 (Fig. 4a and Supplementary Data 13). For arsH, the transfers were frequently detected from viruses to the Koribacter. The aioB gene was detected exclusively in transfers from viruses to Onthovivens. Further analysis revealed a trend in specific host lineages, particularly Kaistella, which frequently acquired the arxS gene from viruses.
a The phylogenetic origin of As-oxidizing genes from virus to host by HGTs. Phenotypic correlations of As-oxidizing genes are noted with colors and statistical support was assessed using standard bootstraps (SBS) (n = 1000). b Enrichment ratio of As-oxidizing genes by virus-mediated HGTs in the rhizosphere compared to bulk soil. The enrichment ratio was determined using the formula: Enrichment ratioread depths\(=\frac{{{Read\; depths}}_{R}}{{{Read\; depths}}_{B}}\). The solid regression line shown is a Loess estimate (n = 50). The shaded interval represents the 95% confidence interval, presenting the variability of the fit over time. Statistical significance between the rhizosphere and the bulk soil was determined using the two-sided Welch’s t-test (n = 44). c Enrichment ratiocopies of recipient microbes in the rhizosphere compared to bulk soil. The solid regression line shown is a Loess estimate (n = 50). The shaded interval represents the 95% confidence interval, presenting the variability of the fit over time. Statistical significance between the rhizosphere and the bulk soil was determined using the two-sided Welch’s t-test (n = 50). d Impact of virus-mediated HGT on the enrichment ratio of recipient microbe. The violin plot shows the enrichment ratio of recipient microbe in the pre-HGT and post-HGT periods, with the mean indicated by the solid point and the standard deviation represented by vertical line extending from the solid point. Statistical significance of Enrichment ratiocopies of recipient microbes between pre-HGT and post-HGT was determined using the two-sided Student’s t-test. Sample sizes for each microbe were as follows: Koribacter (pre-HGT n = 15, post-HGT n = 35), Kaistella_34dR (pre-HGT n = 17, post-HGT n = 33), Limnoglobus (pre-HGT n = 11, post-HGT n = 39), Kaistella_44dR (pre-HGT n = 22, post-HGT n = 28), Bacillus (pre-HGT n = 21, post-HGT n = 29), and Acidobacterium (pre-HGT n = 24, post-HGT n = 26). In the presented genomic map, genes colored in green represent the host’s inherent genes, while genes marked in other colors indicate the location of As-oxidizing gene introduced via virus-mediated HGT.
To explore the prevalence of virus-mediated HGT genes involved in As oxidation, we analyzed their gene enrichment ratios (Supplementary Data 14). Following the seeding stage of rice growth, a significant increase in the distribution of virus-mediated HGT genes involved in As oxidation was observed in the rhizosphere relative to bulk soil (Welch’s t-test, n = 44, p = 0.004, Fig. 4b). This increase was particularly pronounced during the elongation and grain filling stages.
To assess the influence of virus-mediated HGTs of As oxidase on the adaptation of these recipient microbes in the rhizosphere, we determined their enrichment ratios. These microbes showed a significantly higher abundance in the rhizosphere (Welch’s t-test, n = 50, p = 0.0007, Fig. 4c), especially during the elongation stage. To determine the timing of these transfers, we assessed the top 15 abundant recipient contigs retrieved from the time series samples. The earliest detection of As oxidase from virus-mediated HGTs was identified in six contigs, belonging to Koribacter, Kaistella, Limnoglobus, Bacillus, and Acidobacterium. These HGT occurrences were concentrated during the elongation stage (Supplementary Data 15), aligning with the trend of enrichment ratios observed for As oxidase and recipient microbes associated with virus-mediated HGTs (Fig. 4b, c). Subsequently, these recipient microbes demonstrated significant adaptability in the rhizosphere in the post-HGT period compared to the pre-HGT period (Fig. 4d). We also mapped the insertion points of these As-oxidizing genes associated with virus-mediated HGTs within the recipient microbial genomes.
In silico simulation with genome-scale metabolic models
We further in silico quantified the contribution of viral lysogeny in microbial As oxidation using GEMs of 693 species-level As-oxidizing microbes with lysogenic viruses (lysogen) or without lysogenic viruses (host cell) (Fig. 5a and Supplementary Data 16). Using flux balance analysis (FBA) with the GEMs of lysogens and host cells, we investigated the number of MAGs with enhanced As oxidation due to lysogenic viruses and their corresponding phylum-level classifications. We identified 99 species-level lysogens that completely relied on lysogenic virus for As oxidation (virus-enabled), mainly classified in Proteobacteria, Acidobacteriota, Bacteroidota, Gemmatimonadota, Chloroflexota and Verrucomicrobiota phyla. We further observed that 59 species-level lysogens displayed reduced As oxidation flux without lysogenic viruses (viral-aided), mainly from Acidobacteriota, Proteobacteria, Gemmatimonadota, Chloroflexota and Bacteroidota phyla. Furthermore, we quantified that lysogenic viruses contribute approximately 21.0% to As oxidation in the rhizosphere (Fig. 5b). Specifically, among the microbial As-oxidizing flux regulated by viruses, 4.2% of virus-contribution is attributed to the viral-aided As oxidation efficiency, while 16.8% of virus-contribution is due to the virus-enabled As oxidation reaction.
a A phylogenetic tree was built for 693 species-level As-oxidizing MAGs based on the marker genes. The tree branches are colored according to phylum classification. The two inner rings are colored to indicate AMGs associated with As-oxidizing and P-metabolic pathways. The next outer ring shows the average relative abundance of MAGs in the rhizosphere. GEMs simulate the As-oxidizing flux for rhizosphere As-oxidizing MAGs in both the lysogen (with lysogenic viruses) and host cell (without lysogenic viruses) state. The difference in As-oxidizing flux between the two states, divided by the As oxidizing flux in the lysogen state, represents the proportion of viral contribution to the As oxidation for each MAG. The proportion is displayed in the outermost ring layer. The right bar plot shows the number of MAGs for which As-oxidizing flux is virus-aided or virus-enabled. b The schematic diagram for quantifying the virus contribution to microbial As oxidation, created with BioRender.com. The total virus-contribution is represented as \({C}_{{virus}}\). \({{Flux}}_{{host\; cell},i}\) and \({{Flux}}_{{lysogen},i}\) represents the As-oxidizing flux of the \(i \mathrm{th}\) host cell and lysogen, respectively. \({{{\rm{Growth}}}}_{i}\) denotes the maximum growth for the \(i \mathrm{th}\) MAG, and \(n\) indicates the number of MAGs. The symbol ∑ indicates the sum of all values for As-oxidizing MAGs. The circular diagram quantifies the virus contribution to microbial As oxidation flux through FBA simulation, showing the proportion of As-oxidizing flux that is virus-aided versus entirely virus-enabled. \({{{\boldsymbol{C}}}}_{{virus}-{aided}}\) represents \({{{\boldsymbol{C}}}}_{{virus}}\) considering only \({{Flux}}_{{host\; cell},i}\ne 0\); while \({{{\boldsymbol{C}}}}_{{virus}-{enabled}}\) represents \({{{\boldsymbol{C}}}}_{{virus}}\) considering only \({{Flux}}_{{host\; cell},i}=0\). c Impact of rhizosphere environmental variation on viral contribution to As oxidation in lysogens. The relationship between the upper bound flux of rhizosphere environmental factors and the ratio of total As-oxidizing flux in lysogens to host cells. The promotion of As oxidation by lysogenic viruses in these lysogens is enhanced by rhizosphere environmental factors such as oxygen, carbon sources, inorganic P, organic P, and H+. d The phylum-level taxa composition in which the promotion of As oxidation by lysogenic viruses is enhanced by the rhizosphere environmental factors.
Firstly, we observed significant increases in oxygen, carbon, and available P levels in the rhizosphere compared to bulk soil, as well as a significant decrease in pH (Supplementary Fig. 11). We then in silico estimated the influence of these rhizosphere environmental gradients on the virus-regulated As oxidation process by incorporating these factors as exchange boundaries into the GEMs. We found that the increased upper bounds for oxygen, H+, inorganic P, organic P and carbon source fluxes can amplify the role of lysogenic viruses in microbial As oxidation across multiple taxa (Fig. 5c, d). These lysogens affected by environmental factors are most sensitive to elevated oxygen levels, followed by the elevated H+, carbon source and P levels. Given that the oxygen concentration in the rhizosphere was up to 100 µmol L−1 higher than in the bulk soil (Supplementary Fig. 11), radial oxygen loss (ROL) caused by root activity may be the primary factor enhancing viral metabolic reprogramming in the microbial As oxidation process.
We then demonstrated the taxonomic composition of lysogens in which the promotion of As oxidation by lysogenic viruses is enhanced by rhizosphere environmental factors. These lysogens affected by oxygen that amplify the role of lysogenic viruses in As oxidation predominantly originated from the Proteobacteria (31.82%), Bacteroidota (29.55%), Gemmatimonadota (13.64%) and Verrucomicrobiota (11.36%) phyla (Fig. 5d and Supplementary Fig. 12). The lysogens regulated by H+, inorganic P, organic P, and carbon sources that enhance the role of lysogenic viruses in As oxidation mainly belong to the Acidobacteriota and Proteobacteria phyla. Additionally, the regulation of As oxidation by lysogenic viruses in lysogens from the Actinobacteriota, Myxococcota, and Planctomycetota phyla is promoted by the increase of H+, carbon source and P levels in the rhizosphere (Fig. 5d and Supplementary Fig. 11). Meanwhile, in lysogens from the Elusimicrobiota phylum, the viral contribution to As oxidation is solely facilitated by an increase in carbon sources and organic P caused by root exudates (Fig. 5d).
In vitro validation with batch experiments
To validate the contribution of viruses in the lysogenic lifestyle from the rhizosphere and the bulk soil to microbial As transformation, we conducted in vitro batch culture experiments with induced lysogenic viruses (Fig. 6a). The experimental design comprised two groups: a treatment group with bacteria and lysogenic viruses (VB) and a control group with only bacteria (B). TEM images displayed the typical morphologies of MC-induced lysogenic viruses (Fig. 6b). Notably, lysogenic viruses from the rhizosphere microbes significantly enhanced the As oxidation capacity of the microbes under As(III) environment when compared to those from the bulk soil (Welch’s t-test, n = 5, p = 0.0217, Fig. 6c). Specifically, rhizosphere lysogenic viruses contributed 25.63% to microbial As oxidation. In contrast, the viruses in lysogenic lifestyle from the bulk soil increased the As(V) reduction capacity of microbes by 32.88%.
a The workflow for inducing lysogenic viruses in bacterial cells from soils, and the design of in vitro culture experiments, created with BioRender.com. b TEM image of MC-induced lysogenic viruses. Each experiment was repeated independently three times, with similar results obtained across all replicates. Representative images from these experiments are shown. The scale bars shown in the figures are 100 nm. c Effects of lysogenic viruses from the rhizosphere and the bulk soil on As oxidation of bacterial microbes. Three biotic replicates were performed for VB and B treatments at each time point, and the points represent the relative change in As(V) proportion in VB calculated using the mean value. The solid lines represent regression lines (Logistic model, n = 5). Statistical significance between the rhizosphere and the bulk soil was determined using the two-sided Welch’s t-test.
Discussion
Here, we demonstrated that rice rhizosphere significantly altered viral roles in microbe-mediated As transformation, shifting from promoting As reduction in the bulk soil to enhancing As oxidation in the rhizosphere. Notably, the more prevalent arrB gene in bulk soil encodes an As-reducing enzyme in the microbial pericytoplasmic space, leading to the direct release of more toxic As(III) into the environment23. Conversely, rice plants release exudates and oxygen into the rhizosphere, encouraging As-oxidizing microbes to thrive13,24. According to the “piggyback the winner” concept25, this can induce positive feedback effects, in which viral infections of relatively dominant As-oxidizing microbes boost their activities, resulting in more efficient As oxidation in the rhizosphere compared to bulk soil. This loop is akin to the “Matthew effect” or “the strong get stronger” dynamic, a principle introduced in plant-soil systems26. Viruses have been overlooked in this theoretical perspective, but we demonstrate that those associated with As-transforming microbes are not only widely distributed in global soils but also serve as important factors in rhizosphere system-driven As transformation processes.
The As oxidation was significantly correlated with the abundance of As-oxidizing microbes for which viruses were identified, suggesting a potential contribution of rhizosphere viruses to As oxidation. Our findings further reveal a reproducible trend of viral lysogeny associated with As-oxidizing microbes in the rhizosphere. Lysogeny serves not only as an efficient defense mechanism against lytic viral attacks but also as a driving force for host dynamics and evolution27,28. Additionally, the abundance of these rhizosphere viruses fluctuates across different growth stages of rice, with populations of both lysogenic and lytic viruses progressively converging to those observed in bulk soil as root activity diminishes. This pattern suggests that root activity indirectly (via environmental changes or bacterial community modulation) or directly modulates the lysogeny of these viruses in the rhizosphere. The shared nutrients provided by the roots may lead to a “free rider” effect29, where rhizosphere microbes tend to compete for these resources. As a result, As-oxidizing microbes in the rhizosphere rely less on the nutrients released from viral lysis. Viral lysogeny is favored and selected in the nutrient- and host-rich environment of the rhizosphere. Furthermore, root-derived antimicrobial substances in the rhizosphere, such as alkaloids, terpenoids, and polyphenols30, may hypothetically stimulate lysogeny associations between microbes and viruses, potentially enhancing their adaptability in response to these compounds.
This cross-interaction between viral dynamics, microbial behavior, and plant-root influence provides a comprehensive understanding of the ecological processes occurring in the rhizosphere. By integrating these concepts, we suggest that the preference for lysogeny in the rhizosphere may not only be a viral response but could also represent an adaptive strategy, influenced directly or indirectly by plant root activity. Our study highlights the key role of viruses in rhizosphere As oxidation, broadening the traditional understanding that As dynamics are mainly influenced by physicochemical factors and microbes31. This nuanced perspective reveals the multi-layered interactions between viruses, microbes, and plants, shaping the overall As transformation in the rhizosphere.
Viruses in the rhizosphere have a greater degree of reprogramming on microbial As oxidation than those in the bulk soil by encoding for more As-oxidizing AMGs. For instance, virus-encoded enzymes like AioB catalyze aerobic As oxidation, and ArsH oxidizes toxic organic As(III), while arxS encodes a transcriptional regulator for As(III) anaerobic oxidation24,31. As a hub of plant-microbe signaling32, the cascade of intricate chemical signals may facilitate communication with the virome in the rhizomicrobiome, coordinating the enrichment of As oxidation-related genes for microbial thriving. Notably, virus-mediated HGT genes associated with As oxidation are more prevalent in the rhizosphere, aligning with the previous finding indicating higher gene abundance in rhizosphere soil compared to bulk soil33. Furthermore, this abundance advantage is particularly pronounced after the tillering stage, coinciding with the peak in rhizosphere As oxidation genes. This suggests that virus-mediated HGTs contribute to the functional stability of rhizosphere As oxidation in complex communities34. HGTs are thought to enhance microbial competency for rhizosphere colonization33,35. Our findings further validate this, showing a significant increase in the enrichment ratio of these host microbes in the rhizosphere following the acquisition of As oxidation-related HGT genes. Therefore, the rhizosphere, as a crucial hotspot for virus-mediated HGT genes, may serve as an ‘accelerator’ within ‘evolutionary incubator’ of soils.
The rhizosphere lysogenic viruses also enhance microbial As oxidation through enriching P-metabolism AMGs. Biogeochemical P cycling study of global soils indicates that the identified AMGs involved in P-acquisition have a greater influence on soil P availability than their corresponding bacterial genes under certain circumstances36. Due to plant uptake and a dense microbiome, the rhizosphere often faces P limitation37. Given the similar chemical properties of As and P, microbes commonly substitute As for P to alleviate P deficiency38. Pi-starvation signaling activates As(III) oxidation genes, promoting microbial oxidation of As(III) to As(V)39,40. Microbes then utilize As(V) to synthesize cell lipids (arsenolipids), allowing Pi for other critical cellular reactions such as nucleic acid synthesis24. One study indicated approximately half of the As(V) was incorporated into cellular lipids in the Agrobacterium tumefaciens under Pi-starvation and As-enriched conditions38. Our study extends this understanding to rhizosphere viruses, revealing their potential role in facilitating host As oxidation by enriching P-metabolism AMGs through synchronized dynamics and co-enrichment between the two pathways. Two-component systems like RegX3 and phoP AMGs were capable of transphosphorylation cross-talk in the Pi and As oxidation signal transduction systems, closely integrating As(III) oxidation with the Pi-starvation response in lysogens41,42. Furthermore, the enriched Pi uptake system AMGs (pstS), often co-exist with As oxidation gene clusters in bacterial genomic islands43. This can enhance uptake of As(V) and Pi of host microbes, transporting them into cells for participation in biosyntheses43.Additionally, the enriched AMGs of the Phn system, including phnK, phnL, and phnM, which facilitate phosphonate hydrolysis, are frequently found adjacent to aioBA genes in some As-oxidizing bacteria24,44. Meanwhile, the enriched ugpC AMG can facilitate the utilization of sn-glycerol-3P and synthesis of organic P compounds like pyruvate and pyrimidine45, promoting the proliferation of As-oxidizing microbes. Therefore, our research highlights the regulatory mechanism of rhizosphere viruses in As and P co-metabolism, enhancing host microbes' adaptability under Pi limitation and As(III) stress46.
We then determined that rhizosphere lysogenic viruses effectively reprogram the As oxidation by modulating the metabolic networks of hosts by cellular systems-level analysis. The in silico simulations using FBA indicated that the contribution of lysogenic viruses to the rhizosphere As oxidation was approximately 21%. Notably, about 16.8% of microbial As oxidation relied entirely on viral genomes to enable the reaction, highlighting that viruses regulate As oxidases and the production of key bioelectron carriers (such as cytochrome c)47. Additionally, approximately 4.2% of the As oxidation efficiency was enhanced through viral assistance. This suggests that rhizosphere lysogenic viruses can drive more efficient microbial As oxidation by influencing electron transfer intermediates (like NADPH)48 and terminal electron acceptors (including nitrate and iron)31,48 within host cells. We further validated the above conclusions, showing that rhizosphere lysogenic viruses contribute up to 25% of microbial As oxidation at the community level through in vitro virome experiments. By environmental gradient simulations with GEMs, we found that the ROL from rice roots not only, as previously discovered13, can mediate the enhancement of the As oxidation capacity of the rhizosphere microbiome but also significantly enhances the impact of rhizosphere lysogenic viruses on microbial As oxidation. Root activity, resulting in lower pH49 and higher levels of carbon sources50, elevates the proton gradient and energy state in lysogens, facilitating the viral regulation of microbial As oxidation. This aligns with previous evidence indicating that pH and dissolved organic carbon (DOC) are significant factors influencing the structure of soil viral communities both locally51 and globally17. Additionally, simulations confirmed that higher P-metabolism flux leads to greater contribution of viruses on host cell As oxidation flux, highlighting their facilitating role in rhizosphere microbial As oxidation by enhancing As and P co-metabolism. Our study reveals how rhizosphere ecological factors interact with viral reprogramming, essential for understanding virus-driven As oxidation mechanisms related to root activity. These findings also support the concept of utilizing rhizosphere viruses as a tool for improving soil health in sustainable agriculture. Targeted agricultural management practices, such as drainage and the supplementation of appropriate carbon and P sources, can influence virus-driven As oxidation, thereby offering a scientific foundation for sustainable development strategies.
By integrating time-series metagenomes, in silico simulations of lysogens, and in vitro batch experiments, our study reveals that lysogenic viruses enhanced a quarter of microbial As oxidation in Oryza sativa L. rhizosphere. Several sets of results converged to show that root activities enhance synergistic interactions between As-oxidizing microbes and corresponding viruses, promoting a “win-win” strategy. In the future, it is essential to continue exploring how these interactions affect broader ecological processes, including phosphate uptake, carbon metabolism, and energy production, through complex higher-order networks52. These investigations are crucial for effectively orchestrating rhizosphere interactions using synthetic biology technology53. Our research also establishes viruses as significant “ecological keystones”6 in the rhizosphere, shaping both current and future As biotransformation. Most importantly, our findings suggest that viruses could serve as powerful predictive tools for rhizosphere dynamics54, especially regarding rhizosphere biogeochemical cycles. Delving deeper into the viral ecological mechanisms related to these cycles will unlock greater potential for leveraging rhizosphere viruses to sustain agricultural soil health.
Methods
Experimental materials and rice cultivation
The experimental soil was collected from a rice field in Taizhou, Zhejiang province of China (28° 28′ N, 121°20′ E), and its physicochemical properties were measured (Supplementary Data 1). To prepare the As-spiked soils, an aqueous solution of Na₂HAsO₄ was added to a portion of the soils55. Due to the large amount of soil needed, a stepwise mixing method was used to ensure homogeneity in the soil As contamination56. The As-treated soil was dried for 48 h, then mixed with clean soil to achieve As contamination at 60 mg kg−1. The spiked soil moisture was adjusted to 60% of field capacity and pre-incubated in the dark at 25 °C for one month to equilibrate55. For our experiment, soil was sieved with a 2-mm mesh and used as a medium in plastic pots (diameter = 80 mm, height = 155 mm, 450 g dry soil per pot). Deionized water was added until the water depth reached 3 cm above the soil surface, ensuring stable redox conditions under greenhouse settings55. Seeds of rice (Oryza sativa cv. Guiyexiangzhan) were surface-sterilized, germinated and cultivated for 2 weeks as previously reported57, after which uniform seedlings were selected and transplanted into the potted soil at a density of one plant per pot for a total of 150 pots (50 time points × 3 replicates). Unplanted soil was used as a negative control for an additional 150 pots (50 time points × 3 replicates). Throughout the entire experimental period, all pots were kept flooded and under greenhouse conditions; greenhouse conditions were 12 h light (9 a.m. to 9 p.m.) and 12 h dark (9 p.m. to 9 a.m.) with constant temperature and relative humidity (30 °C; 70%).
Experimental design and sampling
Following transplantation, three plants were harvested every 48 h at 12 a.m. Rhizosphere soils were collected mainly based on the previously described protocols58. Briefly, the roots were collected first, and the attached loose soils were removed by shaking and kneading with sterilized gloves. Then the roots of each plant were transferred to a 50 mL tube containing 20 mL sterile phosphate-buffered saline (PBS) solution and sonicated 50–60 Hz for 20 s. The roots were transferred to a second tube and washing was repeated (second wash). The first and second washing solutions were combined and filtered through a 100 µm nylon mesh cell strainer to remove broken roots and large sediment. And then the rhizosphere solution was centrifuged at 10,000 × g and 4 °C for 5 min to get the rhizosphere soil pellet59. Bulk soil samples were collected from unplanted soils. For DNA extraction, replicate samples of rhizosphere and bulk soil were pooled separately at each time point and stored at −80 °C.
Chemical analyses
Soil physicochemical properties were determined according to Handbook of Soil Analysis60. Soil pH, Eh and oxygen were determined using oxygen and pH microelectrode sensors61 (Unisense OXY25, ROX-N, and pH-N; Science Park, Aarhus, Denmark) every 48 hours at 12 a.m. The fresh soil samples were homogenized and sieved through a 2-mm mesh sieve to remove visible residues (e.g., rocks, roots and organic debris). Then the samples (equivalent to 10 g oven-dried weight) were extracted using water (soil: water = 1:5) to determine soil DOC, following the procedures62. DOC concentration in the filtrates was measured using a Multi N/C TN/TC-analyzer (Analytik Jena AG, Jena, Germany). The soil samples were then divided into two parts, the first subsample was air-dried and further sieved through a 0.25-mm sieve for determination of AP63. The content of AP was determined through the sodium bicarbonate extraction molybdenum antimony anti-colorimetric method. The second subsample was freeze-dried, sieved (<60 mesh) and stored at −20 °C in the dark prior to analyses of the concentrations of As species64. Soil samples (0.2 g) were extracted with 10 mL of 2% phosphoric acid solution, and then sonicated for 60 min and centrifuged for 15 min at 3000 × g64. Then the soil extract was treated with 6 M HCl to prevent Fe oxidation and As species were immediately measured55. The species of As(III) and As(V) were quantified via high-performance liquid chromatography coupled with HPLC-ICP-MS (PerkinElmer Series 200 HPLC and NEXION300XX, ICP-MS) after chromatographic separation with 20 mM NH4H2PO4 (pH 6.5) as the mobile phase14. As transformation dynamics depend on the changing proportions of As species in soil65. The relative change in As(III) proportion was calculated as the percentage difference between the rhizosphere and bulk soil, normalized to bulk soil proportion:
where Relative changeAs(III) represents relative change in As(III) proportion in the rhizosphere; As(III) proportion is the average As(III) proportion of the three replicate samples in the rhizosphere and bulk soil at each time point.
DNA extraction, internal standard spike-in, and sequencing
DNA extraction from approximately 300 mg of each soil sample was performed using the E.Z.N.A. Soil DNA Kit (Omega Bio-tek, Norcross, GA). Extracted DNA concentration was quantified with a Qubit 4.0 fluorometer. The DNA Internal Standard (DIS) BMS5 was synthesized via plasmid linearization, following established protocols66,67. This DIS was amplified with NEB’s Q5 High-Fidelity 2X Master Mix and verified via gel electrophoresis. Corresponding bands were excised and purified using the Promega Wizard® SV Gel and PCR Clean-Up System. Purified DIS was then spiked into the DNA extracts, constituting 0.5% of the extracted DNA yield, as literature guidelines66,67. Sequencing libraries were generated with Illumina TruSeq Nano DNA LT Library Preparation Kit following the manufacturer’s recommendations. Libraries for DNA sequencing were prepared using 200 ng of input DNA for the Illumina NovaSeq 6000 (Illumina, San Diego, CA, USA) and produced a total of 65 - 108 million paired-end reads per sample (102 samples in total, see details in the Supplementary Data 2).
Metagenomic processing protocols
Raw reads were quality-controlled using Trimmomatic68 (v2.39) to trim adapters and primers, and to filter short (<50 bp) and low-quality reads (<20 bases). The number of DIS should be quantified by first using a BLASTn69 (v2.11.0) homology search against the DIS sequence to identify standard reads (bit-score >50) followed protocols66,67. Following quantification, the DIS sequences were removed from the dataset using SeqKit270 (v2.4.0). Clean data then were obtained by comparing the obtained clean sequences with the rice genome by the BWA-MEM71 (v1.0.6) algorithm to remove host genome contamination and low-quality data. Quality-controlled reads were categorized into 16 groups by sampling time and soil type (rhizosphere or bulk soil). Both the pooled metagenomic reads from each group and individual metagenomic reads per sample were de novo assembled using MEGAHIT72 (v1.2.9) with a minimum contig length of 500 bp (k-step = 10; k-min = 27). The metagenomic data, quality control statistics, and sequence information for the contigs generated from each sample are provided in Supplementary Data 2.
Profiles of As-transforming microbes
All contigs > 1.5 kb were clustered into species-level contigs on the basis of 95% ANI of the shorter sequence73. As alignment fraction (AF) has been increasingly recognized for improving the robustness and reliability of species delineation74,75,76, AF was estimated using a pipeline77, and only contigs meeting the AF threshold were retained. Subsequently, species-level contigs were assigned using the Genome Taxonomy Database (GTDB) (release 214)78 by CLARK79 (v1.1). The protein-coding genes, or open reading frames (ORFs), identified in the contigs using Prodigal, were annotated through DIAMOND80 (v2.0.2) searching against AsgeneDB, a microbial As-transforming gene database81, using recommended parameters81 (e-value < 10−5, identity >30%). A total of 10,349 contigs encoding As oxidase genes and 17,752 contigs encoding As reductase genes were identified as putative arsenic transformation microbes (Supplementary Data 3). BWA-MEM71 (v1.0.6) was used to align the clean reads back to the representative contigs and protein-coding genes separately. To calculate the abundance of contigs or genes, CoverM82 (v0.7.0) was then used to calculate the number of reads aligned per contig/gene base (--methods reads_per_base) with strict parameters (--min-read-percent-identity 0.99 --min-read-aligned-percent 0.75) for accuracy83. Furthermore, the read number for each sample was determined using SeqKit2 (algorithm: stat) for normalizing the relative abundance by read per kilobase per million mapped reads (RPKM). Then the absolute abundance was calculated as follows66,67:
Abundance (copies ng−1) represents the number of contig or gene copies per nanogram of DNA in the sample. NDIS denotes the number of DIS copies added to the sample. \({m}_{{DNA}}\) indicates the mass of the extracted DNA from the sample, measured in nanograms (ng). \({N}_{{reads\; of\; target\; sequence}}\) refers to the count of reads that are mapped to the target sequence. \({L}_{{length\; of\; target\; sequence}}\), expressed in base pairs (bp), is the length of this sequence. \({N}_{{reads\; of\; DIS}}\) is the number of reads mapped to DIS. \({L}_{{length\; of\; DIS}}\), measured in base pairs (bp), indicates the length of the DIS sequence. Additionally, NDIS was calculated as:
Where C, V, NA, and MDIS denote the concentration of DIS added to the sample (ng μL−1), the volume of DIS added to the sample (μL), Avogadro’s constant and molar mass of DIS (double-stranded), respectively.
The detailed information of the identified As-transforming contigs are provided in the Supplementary Data 3 and 4. To quantify the enrichment of As-transforming contigs or genes in the rhizosphere relative to bulk soil, we calculated their enrichment ratios based on copy numbers and RPKM values. The enrichment ratio greater than 1 indicates a higher presence of these contigs or genes in the rhizosphere as compared to bulk soil. The enrichment ratios were determined using the following formulas:
-
(1)
Enrichment ratio of copy numbers
$${{\rm{Enrichment}}}\; {{{\rm{ratio}}}}_{{{copies}}}=\frac{{{Copies}}_{R}}{{{Copies}}_{B}}$$(4) -
(2)
Enrichment ratio of RPKM
Where \({{Copies}}_{R}\) and \({{RPKM}}_{R}\) represent the absolute copy numbers and RPKM observed in the rhizosphere samples; \({{Copies}}_{B}\) and \({{RPKM}}_{B}\) denote the corresponding measures in bulk soil samples.
Viral contig prediction
We used a viral detection pipeline that combined four viral signatures to identify viral contigs: (1) the presence of viral protein families; (2) the absence of microbial protein families; (3) the presence of viral nucleotide signatures; (4) multiple adjacent genes on the same strand. For detecting the presence of viral nucleotide signatures of ssDNA and dsDNA viruses, VirFinder84 (v1.1) was used to detect the presence of viral nucleotide signatures of ssDNA and dsDNA viruses, with cutoffs based on recommended guidelines. VirFinder has been shown to have better performance in evaluating viral contigs of ≥ 1.5kb85. All identification of viral signatures followed the previously established pipeline based on a combination of rules that are specific to different fragment lengths77, this viral detection pipeline established for genome fragments longer than 1 kb achieved false-positive rates of only 0.43%. To further minimize the potential for misidentification of short sequences (<5 kb), we only retained those viral sequences with a viral gene percentage >20%, as recommended by ref. 86. To control the quality of viral contigs, CheckV87 (v1.0) was utilized to refine viral contigs by identifying host-virus boundaries, removing contaminants and flanking host regions on assembled proviruses. Then quality-controlled viral contigs longer than 1.5 kb were further identified as representative vOTUs using the thresholds of 95% ANI and 85% AF, as recommended by Roux et al.88.
Profiles of viruses associated with As-transforming microbes
Three bioinformatic approaches were employed to predict virus-host associations. (1) Contigs of As-transforming microbes were used for searching CRISPR spacers by a combination of MinCED (v0.4.2) and PILER-CR89 with default parameters. To minimize spurious predictions, we dropped arrays with fewer than three spacers, those with nonconserved repeats (< 97% average identity to consensus repeat) or those in microbial contigs containing fewer than four CRISPR-associated proteins90. Spacers were searched against viral contigs using BLASTn, allowing a maximum of one mismatch or gap over ≥95% of the spacer length90. (2) Considering that host-range predictions based on sequence similarity are the most reliable88, integrated prophages in As-transforming microbes were identified using BLASTn with stringent parameters: alignment required > 95% identity and e-value < 10−5 over >1500 bp on microbial contigs > 1.5 times the length of the viral contig90,91. (3) Host and virus tRNA genes were predicted via tRNAscan-SE (v1.23) with default settings. The recovered tRNAs were compared by BLASTn and only perfect hits (100% coverage and 100% identity) were considered indicative of putative host-virus pairs92. In total, we identified 150 viruses associated with 343 As-oxidizing microbes from 112 genera and 299 viruses associated with 570 As reducing microbes from 225 genera (Supplementary Data 5). A bipartite network was constructed on the results generated from the methods described above using ggraph package in R studio, with the edges presenting virus-host associations. The association between vOTUs with As-transforming microbes and the species-level global soil virome dataset (GSV)17 was identified using BLASTn with the 95% ANI threshold88 (Supplementary Data 8). Then the normalized abundance and sample information of GSV in global soil metagenomes was downloaded from Zenodo at https://zenodo.org/records/10463783. The total abundance of GSV-matched viruses associated with As-transforming microbes was extracted, counted and visualized in RStudio (Supplementary Data 9). Lysogenic viruses were identified by combining the alignment results of the integrated prophage previously mentioned and predictions from Checkv and VIBRANT93 (v1.2.1). Classification for viral contigs was performed with a semi-supervised learning model PhaGCN94 (v2.0; default parameters) using a recommended cut-off score > 0.536 (Supplementary Data 6). The identified viral contigs were further analyzed using BWA-MEM to align clean reads, and the number of reads per contig base was calculated using CoverM (v0.7.0) with the strict parameters previously mentioned. The absolute abundance and lifestyle of viral contigs was determined using the previously mentioned formula and results were provided in Supplementary Data 7. VHR was determined by comparing the abundance of viruses associated with As-transforming microbes to the abundance of the corresponding As-transforming microbial hosts19.
Profiles of viral genes associated with As-transforming microbes
Prodigal95 (v2.6.3) was used for ORF prediction of viral contigs associated with As-transforming microbes using the meta option95. To determine the integrase gene in viral contigs, ORFs were compared to integrase proteins downloaded from Pfam (PF07508, PF00589, PF02899, PF09003, PF12482, PF12834, PF12835, PF13009, PF13102, PF13495, PF14659, PF14882, PF18644) using HMMER (v3.1b2) with an e-value threshold of 0.0000119. The 18 ORFs associated with As-oxidizing microbes and 24 ORFs associated with As-reducing microbes were identified as integrase genes (Supplementary Data 12). The annotations of gene function were searched against orthologous gene databases eggNOG96 (v5.0.0), AsgeneDB81, MCycDB97, NCycDB98, PCycDB45, and SCycDB99 using DIAMOND with recommended parameters of each database (e-value < 10−5, identity > 30%). The function of ORFs included categories such as metabolism, genetic information processing, environmental information processing, cellular processes, and organismal systems. Then the metabolic ORFs were further filtered based on DRAM-v annotation with default parameters, removing those with the auxiliary score greater than 3 and assigned an A, V, T or B flag100. Among them, 9 ORFs and 9 ORFs were identified as viral AMGs associated with As oxidation and P metabolism. The numbers of reads mapping to each viral gene in each sample were counted using the CoverM (--methods count). Differential expression (DE) in the rhizosphere compared to bulk soil was estimated using “DEseq” function in DESeq2 package in R. For all DESeq2 analyses, non-normalized count tables were used as input. The enriched genes in the rhizosphere compared to bulk soil were filtered based on Log2FoldChange > 0 and adjusted p value (p adj) < 0.01, using the Wald test (Supplementary Data 11). Then the number of reads per gene base was calculated using CoverM with the strict parameters previously mentioned. The absolute abundance of viral genes was determined using the aforementioned formula (Supplementary Data 12). The protein structure and main confidence to check the accuracy of structures metrics (pLDDT and pTM) of each gene was further supported by AlphaFold321. Then ChimeraX101 was used for the visualization of protein structure.
Profiles of virus-mediated horizontal gene transfers
To identify As-oxidizing microbes nested within viral clades (virus-to-host HGT), prophages previously identified were removed from microbial contigs to systematically detect viral-host gene exchange. Specifically, all ORFs annotated as As-oxidizing genes were classified as from either host or virus contigs. Host and viral proteins were then clustered into protein families using a similarity-based approach and the Markov clustering (MCL) algorithm (inflation = 2) after comparing sequences to one another using Diamond BLASTp (sensitive mode, E-value < 10−5, query coverage >50%). To detect instances of HGT, an automated and conservative algorithmic pipeline22 previously established was used to identify HGT-indicative topologies in the phylogenetic trees generated from each As-oxidizing protein family. In total, 40 instances of HGT were identified, characteristics including the recipient, donor, phylogenetic statistics and topology notes were recorded (Supplementary Data 13). Mapping information of recipient genomes were extracted from the quality-controlled BAM files. After identifying each genomic region of HGT within these recipient microbial genomes, samtools102 (v1.9 bedcov) was used to calculate the read depth of sequencing coverage of HGT genes in the BED file103. The obtained read depth of each HGT gene was normalized by the length of genomic region and total number of bp sequenced in the sample to allow for sample-to-sample comparison104 (Supplementary Data 14).
To compare the difference in HGT gene abundance between rhizosphere and bulk soil, the enrichment ratio was determined using the following formula:
Where \({{Read\; depths}}_{R}\) represent the read depths observed in the rhizosphere samples, and \({{Read\; depths}}_{B}\) denote the corresponding measures in bulk soil samples.
To potentially determine the timing of HGT events in the rhizosphere, all assembled contigs from time-point rhizosphere samples were mapped to recipient host contigs, using a 95% ANI and 85% AF. SeqKit2 was used to extract the assembled contigs of the top 15 abundant recipient hosts from samples at all time points. BLASTn was then used to ascertain whether these assembled contig genomes from specific time-point samples contained HGT genes (applying the identity threshold consistent with the parameter previously mentioned by CoverM), recording the appearance time of these HGT genes in the host contigs. We identified the recipient time of the six contigs (Supplementary Data 15). The recipient host contigs 42dR_1026030, 22dR_752297, and 44dR_1213478 were only assembled in specific time-point rhizosphere samples but were absent in bulk soil samples. Additionally, 10dN_812543, 30dR_950833, and 34dR_1015320 contigs were assembled multiple times in the rhizosphere time-series samples. Through alignment, we identified specific time points on their genomes when HGT genes began to be contained, and these HGT genes reappeared in subsequent samples with a frequency of ≥ 3 times after this time point. Then the sample corresponding to the earliest detection of HGT genes in each recipient host contig was defined as the HGT occurrence node. The time before this node is defined as pre-HGT, and the time after this node is defined as post-HGT, to determine the abundance changes of hosts after receiving virus-mediated HGT genes. Genomes of the recipient hosts were annotated by Prokka105 and genome maps were visualized using Geneious (v2023.2.1).
Metagenome binning, refinement and annotation (functional, phylogenetic, and taxonomic)
MAGs were recovered for metagenomic assemblies using metaWRAP106 (v1.3) on the basis of tetranucleotide frequencies (TNF) and coverage information, contigs shorter than 1500 bp were discarded. The resulting MAGs were refined using the module ‘bin_refinement’ from metaWRAP to combine and improve the results generated by the three binners. During refinement, the completeness and contamination of all MAGs were estimated using CheckM82 (v1.1.6) via the lineage-specific workflow with the options ‘-c 50 -x 10’ to filter MAGs to be at least 50% complete, with < 10% contamination. The MAGs were further quality-filtered with the function ‘--checkM_method (lineage_wf)’ to avoid low-quality genomes, and then the filtered 1956 MAGs were dereplicated and clustered into 1356 species-level MAGs using dRep107 (v3.0.9) with the default parameters. A total of 693 species-level MAGs putatively encode As oxidases with oxygen-independent and oxygen-dependent pathways. We extracted these As-oxidizing MAGs for the construction of a maximum-likelihood phylogenetic tree. A multiple sequencing alignment (MSA) of the core genes generated by GTDB-tk108 (v2.1.1) were subjected to iqtree2109 with the following options ‘-m MFP -bb 1000’ and the constructed phylogenetic tree was visualized with ggtree (v3.2.1). These MAGs were classified with GTDB-TK using the ‘classify_wf’ function and default parameters according to the GTDB78 (release 214). To confirm the presence of these As-oxidizing MAGs in the rhizosphere, metagenomic reads from rhizosphere samples were aligned to each MAG. This was executed using the CoverM command line tool in combination with the BWA aligner, employing previously mentioned parameters, and calculating their relative abundance using the “genome” setting in CoverM110. The average relative abundance of these MAGs in the rhizosphere samples was calculated to assess their presence in the rhizosphere using the R package dplyr111 (Supplementary Data 16).
Connecting MAGs to viruses and masking lysogenic virus sequence
MAGs were aligned to viral contigs using BLASTn to identify integrated viral sequences. A viral contig was determined to be integrated in a MAG if it aligned by e-value < 10−5 and > 95% identity over > 1000 bp on a microbial contig that was >1.5 times the length of the viral contig20,77,91. To exclude possible decayed prophages, that is, integrated virus genomes which are now inactive and progressively removed from the host genome, all predictions for which 30% or more of the genes displaying a best hit to Pfam (35.0)112 were excluded using the thresholds (hmmsearchscore ≥ 50 and E ≤ 0.001) recommended by the ref. 17,90. Then the base positions of the aligned target regions in the MAGs were replaced with ‘N’ to mask the integrated viral sequences. MAGs are termed ‘lysogens’ before masking and ‘host cells’ after masking. Then the ORFs in lysogen and host cell MAGs were predicted using Prodigal with the ‘-p single’ parameter. Then the predicted ORFs were annotated with orthologous gene databases as previously mentioned (Supplementary Data 16).
Reference database
The reference used to link genome annotation to metabolism was the MetaCyc database113, a comprehensive reference database of metabolic pathways and enzymes from all domains of life. The selection of the MetaCyc database was predicated on two principal criteria. First, MetaCyc is meticulously curated by hand, and it has far wider coverage of metabolic pathways and reactions compared to other databases such as KEGG114, with particular emphasis on the inclusion of numerous critical secondary metabolic reactions within host cells. Second, the As-related metabolic reactions are only covered within MetaCyc.
Reconstruction of genome-scale metabolic models (GEMs)
All functional annotation results of MAGs were compiled into the Eggnog-mapper annotation file format in R studio. The Python package Emapper2gbk115 (v0.3.1) was utilized to connect these between the annotation file and GEM reconstruction tool in order to generate genome annotated inputs. To compile genome-annotated results, we organized functional annotations derived from the ORFs of MAGs using R studio. Subsequently, we transformed these annotations into a format compatible with GenBank using the Python tool Emapper2gbk. This conversion facilitated the utilization of the PathoLogic module within PathwayTools116 (v23.0), enabling the direct mapping of metabolic pathways from genome annotations, guided by the comprehensive MetaCyc database. Consequently, each GEM was reconstructed with an automated command-line version of PathwayTools using the mpwt program115, then the model was converted into sbml format with padmet115. All GEMs were constructed without gap-filling or manual curation to avoid the risk of introducing false positives, particularly in the case of poorly known organisms, which could mask potential metabolic dependencies117,118. Thus, we chose to rely on drafts of GEMs, i.e., we chose false negatives (due to flaws in genome annotation) over false positives.
Flux balance analysis (FBA)
The GEMs simulations of lysogen and host cell were handled by the package COBRApy119 (v0.29.0) for FBA to assess the impact of viruses. The media compositions were defined as LB medium for organism to grow, setting the upper flux of exchange boundary for reactions at 10 mmol gDW−1 h−1120. The oxygen flux boundaries of GEMs were determined by the As oxidation pathway, setting the boundary at 50 mmol gDW-1 h-1 for aerobic oxidation and 0 mmol gDW−1 h−1 for anaerobic oxidation. By setting the corresponding As oxidation reaction as objective in GEMs, we compared the objective value between lysogen and host cell of each MAG to identify the change in their As-oxidizing capability (Supplementary Data 17). Further, we integrated a bacterial biomass equation into GEM based on universal bacterial model120. Model growth was simulated with biomass as the objective function to calculate the maximum growth rate of each MAG (Supplementary Data 17). To simulate environmental conditions within the rhizosphere, components such as oxygen, carbon sources (the representative organic acids, polysaccharides, lipids, and amino acids detected in rhizosphere exudates, as detailed in Supplementary Data 18), inorganic P (common forms in soil, Supplementary Data 19), organic P (substrates catalyzed by the enriched organic P AMGs, Supplementary Data 19), and H+ were respectively provided to the models. Exchange fluxes for the target simulation compounds were set in a gradient within [0–1000 mmol gDW−1 h−1], simulating the objective value of As oxidation reaction in both lysogens and their corresponding host cells (Supplementary Data 20). This method aims to assess the influence of the rhizosphere environment on the viral-regulated As oxidation network. We then fitted the rhizosphere environmental gradients and the ratio of As-oxidizing flux in lysogen to host cell. Lysogens with a slope greater than 0 were defined as those in which the promotion of As oxidation by lysogenic viruses is enhanced by rhizosphere environmental factors.
Preparation of soil viral and bacterial suspensions
The induction of lysogenic viruses was performed as previously described by tangential flow filtration (TFF) and MC induction assays14,18. After 50 days, 25 g of rhizosphere and bulk soil samples were collected from each time point and pooled together. Approximately 600 g of these collected samples were then washed using sterile saline magnesium buffer (SM; per L containing 5.8 g of NaCl, 2 g of MgSO4, and 50 mL of Tris-HCl, adjusted to pH 7.5). This washing process involved vortexing at a maximum speed for 3 min followed by shaking for 2 h at 4 °C and 250 rpm. Subsequently, the soil slurry was centrifuged at 2500 × g at 4 °C for 20 minutes. The resulting supernatant was filtered through a 2 μm cellulose acetate filter, and further filtered and concentrated using the TFF system (Cogent μScale, Merck, America) with pore sizes of 0.22 μm and 100 kDa. The filtrate (from 100 kDa to 0.22 μm) was removed as it contained the extracellular free virus suspension. Approximately 500 mL of the cell suspension containing particles larger than 0.22 μm was retained as the bacterial community121,122.
To provide carbon, energy, and other nutrient resources for the following lysogenic virus-induced processes and activities of the virome and host microbiome in culture experiments, sterile water-soluble organic carbon (WSOC) solutions were prepared according to the previously reported method18. In detail, soil from the rhizosphere and bulk soil was suspended in sterile deionized water and homogenized by vortexing at maximum speed for 3 min. Then, the soil slurry was centrifuged at 5000 × g for 20 minutes, and the supernatant was filtered through 0.02 μm PVDF filter and then autoclaved. These bacterial suspensions were then inoculated into sterile WSOC solutions, both with and without the addition of MC (at a final concentration of 1 μg mL−1)18. The suspensions were incubated with shaking at 200 rpm for 24 h in the dark at room temperature123. Subsequently, MC-induced lysogenic viruses were collected after passing through a 0.22 μm sterile filter (Thermo Scientific). Following filtration, Amicon Ultra filters (Millipore) were employed to concentrate and collect the lysogenic viruses124. Both viral and bacterial suspensions were observed, verified, and counted with epifluorescence microscopy (EFM) as previously described123. The virus-like particle (VLP) concentrations in the rhizosphere and bulk soil virus suspensions were 3.37 × 10⁹ and 2.91 × 10⁹ VLPs per gram of soil, respectively (see details in the Supplementary Methods and Supplementary Fig. 13). In addition, the morphologies of lysogenic viruses were observed in suspension using transmission electron microscopy (TEM).
In vitro culture experiments
In order to simulate the in-situ environment, we added 1 mL of virus suspensions or sterile WSOC solution along with 3 mL of bacterial suspensions into each sterile 6 mL cell culture plate, maintaining a volume ratio of 1:3 (virus to bacteria)121. The experiment was structured into four distinct treatments: (1) B in the rhizosphere (B R): 1 mL sterile WSOC solution from rhizosphere + 3 mL bacteria suspension from rhizosphere; (2) VB in the rhizosphere (VB R): 1 mL MC-induced lysogenic virus suspension from rhizosphere + 3 mL bacteria suspension from rhizosphere; (3) B in bulk soil (B B): 1 mL sterile WSOC solution from bulk soil + 3 mL bacteria suspension from bulk soil; and (4) VB in bulk soil (VB B): 1 mL MC-induced lysogenic virus suspension from bulk soil + 3 mL bacteria suspension from bulk soil. All treatments were exposed to As(III) environments at 20 mg kg−1 and replicated in triplicate. Subsequently, each sample was sealed with a sterilized gas-permeable sealing membrane and incubated at 25 °C in the dark within a controlled environment room. As species from the culture were determined at 0, 3, 6, 12 and 24 h using HPLC-ICP-MS.
Relative change in As(V) proportion in VB at each time was calculated as:
where Relative changeAs(V) represents relative change in As(V) proportion in VB; As(V) proportion are the average As(V) proportion of the three biotic replicates in VB and B treatments at each time point.
Statistics and reproducibility
Analysis in this project was performed using R version 4.1.1125. A two-sided p value of < 0.05 was considered significant for all analyses. The statistical significance between the two groups was determined using either the two-sided Welch’s two-sample t-test (for unequal variances) or two-sided Student’s two-sample t-test (for equal variances). No data were excluded from the analyses. The investigators were not blinded to allocation during experiments and outcome assessment.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All raw sequence data generated in this study have been deposited in the National Genomics Data Center under project accession code PRJCA026734. All data are publicly accessible at https://ngdc.cncb.ac.cn/gsa/browse/CRA018565 (GSA: CRA018565). Detailed information on the sample sequence data can be found in Supplementary Data 2 [https://doi.org/10.6084/m9.figshare.28517573]. The supplementary and source data for this paper are available as Supplementary Data and Source Data files, which have been deposited in the Figshare database [https://doi.org/10.6084/m9.figshare.28517573]. The databases used in this study include the GTDB database Release 214 [https://data.ace.uq.edu.au/public/gtdb/data/releases/release214/214.0/], the MetaCyc database (https://metacyc.org/), the AsgeneDB database [https://data.cyverse.org/dav-anon/iplant/home/xinwei/AsgeneDB], the MCycDB database [https://github.com/qichao1984/MCycDB/blob/main/MCycDB_2021.zip], the NcycDB database [https://github.com/qichao1984/NCyc/tree/master/data], the PcycDB database [https://github.com/ZengJiaxiong/Phosphorus-cycling-database], and the SCycDB database [https://github.com/qichao1984/SCycDB/blob/master/SCycDB_2020Mar.zip]. Source data are provided with this paper.
References
Korenblum, E. et al. Rhizosphere microbiome mediates systemic root metabolite exudation by root-to-root signaling. Proc. Natl. Acad. Sci. USA 18, 3874–3883 (2020).
Trivedi, P., Leach, J. E., Tringe, S. G., Sa, T. & Singh, B. K. Plant–microbiome interactions: from community assembly to plant health. Nat. Rev. Microbiol. 18, 607–621 (2020).
Ling, N., Wang, T. & Kuzyakov, Y. Rhizosphere bacteriome structure and functions. Nat. Commun. 13, 1–13 (2022).
Jansson, J. K. & Wu, R. Soil viral diversity, ecology and climate change. Nat. Rev. Microbiol. 21, 296–311 (2023).
Santos-Medellín, C., Blazewicz, S. J., Pett-Ridge, J., Firestone, M. K. & Emerson, J. B. Viral but not bacterial community successional patterns reflect extreme turnover shortly after rewetting dry soils. Nat. Ecol. Evol. 7, 1809–1822 (2023).
Braga, L. P. P. & Schumacher, R. I. Awaking the dormant virome in the rhizosphere. Mol. Ecol. 32, 2985–2999 (2023).
Muscatt, G. et al. Crop management shapes the diversity and activity of DNA and RNA viruses in the rhizosphere. Microbiome 10, 181 (2022).
Cheng, Z. et al. Interactive dynamics between rhizosphere bacterial and viral communities facilitate soybean fitness to cadmium stress revealed by time-series metagenomics. Soil Biol. Biochem. 190, 109313 (2024).
Pratama, A. A., Terpstra, J., de Oliveria, A. L. M. & Salles, J. F. The role of rhizosphere bacteriophages in plant health. Trends Microbiol. 28, 709–718 (2020).
Chen, Q. Y. & Costa, M. Arsenic: a global environmental challenge. Annu. Rev. Pharmacol. Toxicol. 61, 47–63 (2021).
Zhang, S. et al. Escalating arsenic contamination throughout Chinese soils. Nat. Sustain. 1–10 https://doi.org/10.1038/s41893-024-01341-7 (2024).
Yang, H.-C. & Rosen, B. P. New mechanisms of bacterial arsenic resistance. Biomed. J. 39, 5–13 (2016).
Jia, Y., Huang, H., Chen, Z. & Zhu, Y.-G. Arsenic uptake by rice is influenced by microbe-mediated arsenic redox changes in the rhizosphere. Environ. Sci. Technol. 48, 1001–1007 (2014).
Tang, X. et al. Lysogenic bacteriophages encoding arsenic resistance determinants promote bacterial community adaptation to arsenic toxicity. ISME J. 17, 1104–1115 (2023).
Pratama, A. A. & van Elsas, J. D. The ‘neglected’ soil virome—potential role and impact. Trends Microbiol. 26, 649–662 (2018).
Chen, S.-C. et al. The Great Oxidation Event expanded the genetic repertoire of arsenic metabolism and cycling. Proc. Natl. Acad. Sci. USA 117, 10414–10421 (2020).
Ma, B. et al. Biogeographic patterns and drivers of soil viromes. Nat. Ecol. Evol. 8, 717–728 (2024).
Huang, D. et al. Enhanced mutualistic symbiosis between soil phages and bacteria with elevated chromium-induced environmental stress. Microbiome 9, 150 (2021).
Xia, R. et al. Benzo[a]pyrene stress impacts adaptive strategies and ecological functions of earthworm intestinal viromes. ISME J. 17, 1004–1014 (2023).
Du, S. et al. Highly host-linked viromes in the built environment possess habitat-dependent diversity and functions for potential virus-host coevolution. Nat. Commun. 14, 2676 (2023).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Irwin, N. A. T., Pittis, A. A., Richards, T. A. & Keeling, P. J. Systematic evaluation of horizontal gene transfer between eukaryotes and viruses. Nat. Microbiol. 7, 327–336 (2022).
Glasser, N. R., Oyala, P. H., Osborne, T. H., Santini, J. M. & Newman, D. K. Structural and mechanistic analysis of the arsenate respiratory reductase provides insight into environmental arsenic transformations. Proc. Natl. Acad. Sci. USA 115, E8614–E8623 (2018).
Shi, K., Wang, Q. & Wang, G. Microbial oxidation of arsenite: regulation, chemotaxis, phosphate metabolism and energy generation. Front. Microbiol. 11, 569282 (2020).
Silveira, C. B. & Rohwer, F. L. Piggyback-the-Winner in host-associated microbial communities. npj Biofilms Microbiomes 2, 1–5 (2016).
Dan, X. et al. Strong rhizosphere priming effects on N dynamics in soils with higher soil N supply capacity: the ‘Matthew effect’ in plant-soil systems. Soil Biol. Biochem. 178, 108949 (2023).
Bondy-Denomy, J. et al. Prophages mediate defense against phage infection through diverse mechanisms. ISME J. 10, 2854–2866 (2016).
Doud, M. B. et al. Competition-driven eco-evolutionary feedback reshapes bacteriophage lambda’s fitness landscape and enables speciation. Nat. Commun. 15, 863 (2024).
Denison, R. F. et al. Cooperation in the rhizosphere and the “free rider” problem. Ecology 84, 838–845 (2003).
Wang, W. et al. Rice secondary metabolites: structures, roles, biosynthesis, and metabolic regulation. Molecules 23, 3098 (2018).
Zhu, Y. G., Xue, X. M., Kappler, A., Rosen, B. P. & Meharg, A. A. Linking genes to microbial biogeochemical cycling: lessons from arsenic. Environ. Sci. Technol. 51, 7326–7339 (2017).
Jamil, F., Mukhtar, H., Fouillaud, M. & Dufossé, L. Rhizosphere signaling: insights into plant–rhizomicrobiome interactions for sustainable agronomy. Microorganisms 10, 899 (2022).
Lopes, L. D., Pereira E Silva, M. D. C. & Andreote, F. D. Bacterial abilities and adaptation toward the rhizosphere colonization. Front. Microbiol. 7, 1341 (2016).
Wang, T. et al. Horizontal gene transfer enables programmable gene stability in synthetic microbiota. Nat. Chem. Biol. 18, 1245–1252 (2022).
Alzubaidy, H. et al. Rhizosphere microbiome metagenomics of gray mangroves (Avicennia marina) in the Red Sea. Gene 576, 626–636 (2016).
Liang, J.-L. et al. Hidden diversity and potential ecological function of phosphorus acquisition genes in widespread terrestrial bacteriophages. Nat. Commun. 15, 2827 (2024).
Langhans, C., Beusen, A. H. W., Mogollón, J. M. & Bouwman, A. F. Phosphorus for sustainable development goal target of doubling smallholder productivity. Nat. Sustain. 5, 57–63 (2022).
Wang, Q. et al. Phosphate starvation response controls genes required to synthesize the phosphate analog arsenate. Environ. Microbiol. 20, 1782–1793 (2018).
Qiao, Z., Huang, J., Cao, Y., Shi, K. & Wang, G. Genetics and proteomics analyses reveal the roles of PhoB1 and PhoB2 regulators in bacterial responses to arsenite and phosphate. Res. Microbiol. 170, 263–271 (2019).
Rawle, R. A., Kang, Y.-S., Bothner, B., Wang, G. & McDermott, T. R. Transcriptomics analysis defines global cellular response of Agrobacterium tumefaciens 5A to arsenite exposure regulated through the histidine kinases PhoR and AioS. Environ. Microbiol. 21, 2659–2676 (2019).
Rang, J. et al. SenX3-RegX3, an important two-component system, regulates strain growth and butenyl-spinosyn biosynthesis in Saccharopolyspora pogona. iScience 23, 101398 (2020).
Martín-Martín, S. et al. Self-control of the PHO regulon: the PhoP-dependent protein PhoU controls negatively expression of genes of PHO regulon in Streptomyces coelicolor. J. Antibiot. 71, 113–122 (2018).
Li, H., Li, M., Huang, Y., Rensing, C. & Wang, G. In silico analysis of bacterial arsenic islands reveals remarkable synteny and functional relatedness between arsenate and phosphate. Front. Microbiol. 4, 347 (2013).
Manav, M. C., Sofos, N., Hove-Jensen, B. & Brodersen, D. E. The abc of phosphonate breakdown: a mechanism for bacterial survival. BioEssays 40, 1800091 (2018).
Zeng, J. et al. PCycDB: a comprehensive and accurate database for fast analysis of phosphorus cycling genes. Microbiome 10, 101 (2022).
Wang, X. et al. Heavy metal contamination collapses trophic interactions in the soil microbial food web via bottom-up regulation. Soil Biol. Biochem. 184, 109058 (2023).
Watson, C. et al. Electron transfer through arsenite oxidase: insights into Rieske interaction with cytochrome c. Biochim. Biophys. Acta Bioenerg. 1858, 865–872 (2017).
Xu, R. et al. Diversity and metabolic potentials of As(III)-oxidizing bacteria in activated sludge. Appl. Environ. Microbiol. 87, e01769-21 (2021).
Zhang, K. et al. Interactive effects of soil pH and substrate quality on microbial utilization. Eur. J. Soil Biol. 96, 103151 (2020).
Luo, Y. et al. Rice rhizodeposition promotes the build-up of organic carbon in soil via fungal necromass. Soil Biol. Biochem. 160, 108345 (2021).
Huang, X. et al. Soil nutrient conditions alter viral lifestyle strategy and potential function in phosphorous and nitrogen metabolisms. Soil Biol. Biochem. 189, 109279 (2024).
Marschmann, G. L. et al. Predictions of rhizosphere microbiome dynamics with a genome-informed and trait-based energy budget model. Nat. Microbiol. 9, 421–433 (2024).
Ragland, C. J., Shih, K. Y. & Dinneny, J. R. Choreographing root architecture and rhizosphere interactions through synthetic biology. Nat. Commun. 15, 1370 (2024).
Varanda, C., Félix, M., do, R., Campos, M. D. & Materatski, P. An overview of the application of viruses to biotechnology. Viruses 13, 2073 (2021).
Shen, Y. et al. Influence of tetracycline on arsenic mobilization and biotransformation in flooded soils. Environ. Pollut. 292, 118416 (2022).
He, Y. et al. Dissipation of phenanthrene and pyrene at the aerobic–anaerobic soil interface: differentiation induced by the rhizosphere of PAH-tolerant and PAH-sensitive rice (Oryza sativaL.) cultivars. Environ. Sci. Pollut. Res. 22, 3908–3919 (2015).
Huang, Y., Dai, Z., Tang, C. & Xu, J. Labile and recalcitrant carbon inputs differ in their effects on microbial phosphorus transformation in a flooded paddy soil with rice (Oryza sativa L.). Appl. Soil Ecol. 198, 105372 (2024).
Santos-Medellín, C., Edwards, J., Nguyen, B. & Sundaresan, V. Acquisition of a complex root microbiome reshapes the transcriptomes of rice plants. New Phytol. 235, 2008–2021 (2022).
Boubsi, F. et al. Pectic homogalacturonan sensed by Bacillus acts as host associated cue to promote establishment and persistence in the rhizosphere. iScience 26, 107925 (2023).
Pansu, M. & Gautheyrou, J. Handbook of Soil Analysis: Mineralogical Organic and Inorganic Methods. (Springer, Berlin; New York, 2006).
Zhao, K. et al. The only constant is change: endogenous circadian rhythms of soil microbial activities. Soil Biol. Biochem. 173, 108805 (2022).
Huang, Y., Lin, J., Tang, C. & Xu, J. Organic carbon inputs shift the profiles of phosphorus cycling-related genes in maize rhizosphere. Plant Soil 503, 595–609 (2024).
Li, X. et al. Acidification suppresses the natural capacity of soil microbiome to fight pathogenic Fusarium infections. Nat. Commun. 14, 5090 (2023).
Guo, T. et al. The influence of periphyton on the migration and transformation of arsenic in the paddy soil: rules and mechanisms. Environ. Pollut. 263, 114624 (2020).
Wang, H. T. et al. Effects of arsenic on gut microbiota and its biotransformation genes in earthworm Metaphire sieboldi. Environ. Sci. Technol. 53, 3841–3849 (2019).
Ju, F. et al. Wastewater treatment plant resistomes are shaped by bacterial composition, genetic exchange, and upregulated expression in the effluent microbiomes. ISME J. 13, 346–360 (2019).
Satinsky, B. M., Gifford, S. M., Crump, B. C. & Moran, M. A. Chapter twelve—Use of internal standards for quantitative metatranscriptome and metagenome analysis. in Methods in Enzymology, Vol. 531 (ed. DeLong, E. F.) 237–250 (Academic Press, 2013).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Shen, W., Sipos, B. & Zhao, L. SeqKit2: a swiss army knife for sequence and alignment processing. iMeta 3, e191 (2024).
Jung, Y. & Han, D. BWA-MEME: BWA-MEM emulated with a machine learning approach. Bioinformatics 38, 2404–2413 (2022).
Li, D., Liu, C.-M., Luo, R., Sadakane, K. & Lam, T.-W. MEGAHIT: an ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics 31, 1674–1676 (2015).
Jain, C., Rodriguez-R, L. M., Phillippy, A. M., Konstantinidis, K. T. & Aluru, S. High throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries. Nat. Commun. 9, 5114 (2018).
Varghese, N. J. et al. Microbial species delineation using whole genome sequences. Nucleic Acids Res. 43, 6761–6771 (2015).
Riesco, R. & Trujillo, M. E. Update on the proposed minimal standards for the use of genome data for the taxonomy of prokaryotes. Int. J. Syst. Evol. Microbiol. 74, 006300 (2024).
Gosselin, S., Fullmer, M. S., Feng, Y. & Gogarten, J. P. Improving phylogenies based on average nucleotide identity, incorporating saturation correction and nonparametric bootstrap support. Syst. Biol. 71, 396–409 (2022).
Nayfach, S. et al. Metagenomic compendium of 189,680 DNA viruses from the human gut microbiome. Nat. Microbiol. 6, 960–970 (2021).
Parks, D. H. et al. GTDB: an ongoing census of bacterial and archaeal diversity through a phylogenetically consistent, rank normalized and complete genome-based taxonomy. Nucleic Acids Res. 50, D785–D794 (2022).
Ounit, R., Wanamaker, S., Close, T. J. & Lonardi, S. CLARK: fast and accurate classification of metagenomic and genomic sequences using discriminative k-mers. BMC Genomics 16, 236 (2015).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 12, 59–60 (2015).
Song, X. et al. AsgeneDB: a curated orthology arsenic metabolism gene database and computational tool for metagenome annotation. NAR Genomics Bioinf. 4, lqac080 (2022).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015).
Woodcroft, B. J. et al. Genome-centric view of carbon processing in thawing permafrost. Nature 560, 49–54 (2018).
Ren, J., Ahlgren, N. A., Lu, Y. Y., Fuhrman, J. A. & Sun, F. VirFinder: a novel k-mer based tool for identifying viral sequences from assembled metagenomic data. Microbiome 5, 69 (2017).
Guo, J. et al. VirSorter2: a multi-classifier, expert-guided approach to detect diverse DNA and RNA viruses. Microbiome 9, 37 (2021).
Li, R. et al. Viral metagenome reveals microbial hosts and the associated antibiotic resistome on microplastics. Nat. Water 2, 553–565 (2024).
Nayfach, S. et al. CheckV assesses the quality and completeness of metagenome-assembled viral genomes. Nat. Biotechnol. 39, 578–585 (2021).
Roux, S. et al. Minimum information about an uncultivated virus genome (MIUViG). Nat. Biotechnol. 37, 29–37 (2019).
Edgar, R. C. PILER-CR: fast and accurate identification of CRISPR repeats. BMC Bioinformatics 8, 18 (2007).
Nayfach, S. et al. A genomic catalog of Earth’s microbiomes. Nat. Biotechnol. 39, 499–509 (2021).
Nishijima, S. et al. Extensive gut virome variation and its associations with host and environmental factors in a population-level cohort. Nat. Commun. 13, 5252 (2022).
Paez-Espino, D. et al. Uncovering Earth’s virome. Nature 536, 425–430 (2016).
Kieft, K., Zhou, Z. & Anantharaman, K. VIBRANT: automated recovery, annotation and curation of microbial viruses, and evaluation of viral community function from genomic sequences. Microbiome 8, 90 (2020).
Jiang, J. Z. et al. Virus classification for viral genomic fragments using PhaGCN2. Briefings Bioinf. 24, bbac505 (2023).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 11, 119 (2010).
Huerta-Cepas, J. et al. eggNOG 5.0: a hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Res. 47, D309–D314 (2019).
Qian, L. et al. MCycDB: a curated database for comprehensively profiling methane cycling processes of environmental microbiomes. Mol. Ecol. Resour. 22, 1803–1823 (2022).
Tu, Q., Lin, L., Cheng, L., Deng, Y. & He, Z. NCycDB: a curated integrative database for fast and accurate metagenomic profiling of nitrogen cycling genes. Bioinformatics 35, 1040–1048 (2019).
Yu, X. et al. SCycDB: a curated functional gene database for metagenomic profiling of sulphur cycling pathways. Mol. Ecol. Resour. 21, 924–940 (2021).
Shaffer, M. et al. DRAM for distilling microbial metabolism to automate the curation of microbiome function. Nucleic Acids Res. 48, 8883–8900 (2020).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. GigaScience 10, giab008 (2021).
Retel, C., Kowallik, V., Becks, L. & Feulner, P. G. D. Strong selection and high mutation supply characterize experimental Chlorovirus evolution. Virus Evol. 8, veac003 (2022).
Roux, S. et al. Ecogenomics of virophages and their giant virus hosts assessed through time series metagenomics. Nat. Commun. 8, 858 (2017).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Uritskiy, G. V., DiRuggiero, J. & Taylor, J. MetaWRAP—a flexible pipeline for genome-resolved metagenomic data analysis. Microbiome 6, 158 (2018).
Olm, M. R., Brown, C. T., Brooks, B. & Banfield, J. F. dRep: a tool for fast and accurate genomic comparisons that enables improved genome recovery from metagenomes through de-replication. ISME J. 11, 2864–2868 (2017).
Chaumeil, P.-A., Mussig, A. J., Hugenholtz, P. & Parks, D. H. GTDB-Tk: a toolkit to classify genomes with the Genome Taxonomy Database. Bioinformatics 36, 1925–1927 (2020).
Minh, B. Q. et al. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the enomic era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Mara, P. et al. Metagenomic profiles of archaea and bacteria within thermal and geochemical gradients of the Guaymas Basin deep subsurface. Nat. Commun. 14, 7768 (2023).
Wickham, H., François, R., Henry, L., Müller, K. & Vaughan, D. dplyr: a grammar of data manipulation. R version 1.1.2. RStudio https://dplyr.tidyverse.org/ (2023).
Mistry, J. et al. Pfam: the protein families database in 2021. Nucleic Acids Res. 49, D412–D419 (2021).
Caspi, R. et al. The MetaCyc database of metabolic pathways and enzymes—a 2019 update. Nucleic Acids Res. 48, D445–D453 (2020).
Kanehisa, M., Sato, Y., Kawashima, M., Furumichi, M. & Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 44, D457–D462 (2016).
Belcour, A. et al. Metage2Metabo, microbiota-scale metabolic complementarity for the identification of key species. eLife 9, e61968 (2020).
Karp, P. D. et al. Pathway Tools version 23.0 update: software for pathway/genome informatics and systems biology. Brief Bioinf. 22, 109–126 (2019).
Mataigne, V., Vannier, N., Vandenkoornhuyse, P. & Hacquard, S. Multi-genome metabolic modeling predicts functional inter-dependencies in the Arabidopsis root microbiome. Microbiome 10, 217 (2022).
Giordano, N. et al. Genome-scale community modelling reveals conserved metabolic cross-feedings in epipelagic bacterioplankton communities. Nat. Commun. 15, 2721 (2024).
Ebrahim, A., Lerman, J. A., Palsson, B. O. & Hyduke, D. R. COBRApy: constraints-based reconstruction and analysis for python. BMC Syst. Biol. 7, 74 (2013).
Machado, D., Andrejev, S., Tramontano, M. & Patil, K. R. Fast automated reconstruction of genome-scale metabolic models for microbial species and communities. Nucleic Acids Res. 46, 7542–7553 (2018).
Tong, D. et al. Viral lysing can alleviate microbial nutrient limitations and accumulate recalcitrant dissolved organic matter components in soil. ISME J. 17, 1247–1256 (2023).
Braga, L. P. P. et al. Impact of phages on soil bacterial communities and nitrogen availability under different assembly scenarios. Microbiome 8, 52 (2020).
Liang, X. et al. Lysogenic reproductive strategies of viral communities vary with soil depth and are correlated with bacterial diversity. Soil Biol. Biochem. 144, 107767 (2020).
Albright, M. B. N. et al. Experimental evidence for the impact of soil viruses on carbon cycling during surface plant litter decomposition. ISME Commun. 2, 1–8 (2022).
Allaire, J. J. RStudio: Integrated Development Environment for R. (2023).
Acknowledgements
This study was supported by the National Natural Science Foundation of China (42277283 to B.M., 42090060 to B.M., 41991334 to J.X.), National Key R&D Program of China (2024YFD1501800 to B.M.), National Key Science and Technology Major Project (Deep Earth Research, 2024ZD1000603 to B.M.), Zhejiang Science Foundation for Distinguished Young Scholars (LR22C190001 to Y.R.), Key R&D Program of Zhejiang Province (2023C02015 to B.M., 2023C02004 to B.M., 2024C03131 to B.M.), and Key R&D Program of Ningbo (2024Z267 to B.M.).
Author information
Authors and Affiliations
Contributions
B.M. and X.S. created the study design. X.S. performed the experiments. X.S., Y.L.W. and K.Z. performed the data analysis. X.S. and B.M. performed the data visualization. X.S., B.M., Y.L.W., D.T., R.G., X.L., D.K., Y.R., M.W., X.T., F.L., Y.L., Y.Z. and J.X. contributed to scientific discussion and wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Ruonan Wu, Kaixiang Shi and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Song, X., Wang, Y., Wang, Y. et al. Rhizosphere-triggered viral lysogeny mediates microbial metabolic reprogramming to enhance arsenic oxidation. Nat Commun 16, 4048 (2025). https://doi.org/10.1038/s41467-025-58695-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-58695-5
This article is cited by
-
Microbially-mediated halogenation and dehalogenation cycling of organohalides in the ocean
Nature Communications (2025)