Abstract
Current strategies for active targeting in the brain are entirely based on the effective interaction of the ligand with the orthosteric sites of specific receptors on the blood-brain barrier (BBB), which is highly susceptible to various pathophysiological factors and limits the efficacy of drug delivery. Here, we propose an allosteric targeted drug delivery strategy that targets classical BBB transmembrane receptors by designing peptide ligands that specifically bind to their transmembrane domains. This strategy prevents competitive interference from endogenous ligands and antibodies by using the insulin receptor and integrin αv as model targets, respectively, and can effectively overcome pseudotargets or target loss caused by shedding or mutating the extracellular domain of target receptors. Moreover, these ligands can be spontaneously embedded in the phospholipid layer of lipid carriers using a plug-and-play approach without chemical modification, with excellent tunability and immunocompatibility. Overall, this allosteric targeted drug delivery strategy can be applied to multiple receptor targets and drug carriers and offers promising therapeutic benefits in brain diseases.
Similar content being viewed by others
Introduction
The blood-brain barrier (BBB) is a major obstacle in the treatment of a variety of brain diseases, and effective treatment strategies often involve active targeted drug delivery via interactions between ligands and corresponding receptors that are highly expressed on the BBB. Despite some progress in bringing this strategy to the clinical stage, to date most active targeting is performed by ligands or antibodies recognizing the extracellular orthosteric site of the target receptor. For example, the insulin receptor (IR) is a membrane protein composed of two α-subunit and two β-subunit, organized into an extracellular portion, a single transmembrane helix, and an intracellular tyrosine kinase domain. The α-subunits form the extracellular domain, while the β-subunits include the transmembrane and intracellular regions, with the transmembrane portion anchoring the receptor to the cell membrane. Insulin, the natural ligand of the IR, binds to its extracellular domain, triggering insulin signaling and activating downstream pathways. Ligand molecules that bind to the extracellular domain of the IR are often used to construct brain-targeted drug delivery systems1,2. However, endogenous insulin may competitively inhibit the binding of these ligand molecules to insulin receptors. Simultaneously, shedding or mutation of the extracellular region of some membrane proteins leads to pseudotarget or target loss, resulting in targeting moieties that do not effectively recognize the target receptor. For example, serum levels of the proteoglycan glypican-3 in some patients with hepatocellular carcinoma are elevated through the shedding of glypican-3 from cell surfaces, which interferes with the efficacy of glypican-3-targeted therapy3. Mutations in the extracellular region of the epidermal growth factor receptor have also been reported to prevent cetuximab from binding to it and causing drug resistance4. Therefore, drug delivery targeting the extracellular region of target receptors faces several insurmountable challenges, including (1) competitive interference from endogenous ligands or antibodies at the target and (2) pseudotarget or target loss caused by shedding or mutation of the extracellular region, which significantly inhibits drug efficacy and may pose safety concerns. Another concern is that exogenous ligand molecules exposed on the carrier surface may display varying degrees of immunogenicity. The c(RGDyK) peptide, for example, which is usually considered nonimmunogenic, can induce acute immune responses and even death in mice given several doses of polyethylene glycol (PEG)-ylated liposomes bearing this peptide5. CDX peptides (snake neurotoxin candoxin derived peptide) targeting nicotinic acetylcholine receptors on the BBB tend to adsorb natural immunoglobulin M (IgM) after they are modified into liposomes, leading to enhanced immunogenicity6. In recent years, emerging allosteric modulators have been developed that recognize allosteric sites other than orthosteric sites and modulate receptor activation by changing the conformation of the receptor7,8, which has the advantages of high selectivity, overcoming resistance to orthosteric modulators, and expanding the concept of “target sites”. The allosteric binding strategy offers promising avenues for drug development while providing insight into the challenges associated with identifying orthosteric sites for active targeted delivery strategies.
Membrane proteins account for more than 60% of the currently available drug targets9. The mining of membrane proteins as targets has long focused on extracellular and intracellular regions, and transmembrane domains (TMDs) have been deemed “undruggable” compared with their water-soluble counterparts. As recognition of their various functions has increased, TMDs have been found not only to support proteins across cell membranes but also to have important roles in molecular and cellular recognition processes and in protein-protein interactions; Certain peptides derived from natural toxins, exemplified by VSTx1, interact with lipid membranes through hydrophobic patches, thereby facilitating specific interaction with the voltage sensor domain, also TMDs, of ion channel proteins10; moreover, TMDs have emerged as potential targets for the development of diagnostics and therapeutics. The TMD of chimeric antigen receptor is important in the activation and function of cell therapy11. A close relationship between the intracellular localization of proteins and the length and amino acid composition of their TMDs has been reported, and alterations in the TMD also affect the rate of endocytosis12. The TMD of the IR not only anchors the receptor within the plasma membrane but also plays a critical role in transmitting the ligand-binding signal13. Peptides containing sequences from the TMD of the IR effectively induced IR phosphorylation, whereas peptides derived from the TMD of other receptors, such as IGF-I, PDGF, HER2/erbB2/Neu, and TrkA receptors, did not. This suggests that, despite their shared hydrophobic characteristics, peptides with appropriately tailored sequences can demonstrate specificity toward the TMD of a targeted receptor14. Overall, specific recognition of membrane proteins can be achieved by identifying the TMD15, which holds great promise as an alternative target site for strategies that target the extracellular region of receptors.
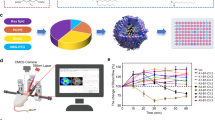
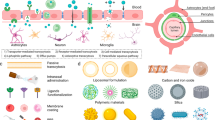
In this work, we construct a drug delivery platform based on the idea of allosteric recognition by using the TMD of membrane proteins (e.g., IR and integrin αvβ3) as a recognition site and explored potential applications for the treatment of brain diseases (Fig. 1). We design and screen peptide ligands that specifically interact with the TMD of the IR or integrin αv as target moieties and further evaluated the allosteric targeting strategy in several carrier types, including liposomes, lipid nanoparticles, and exosomes. Our results show that the designed lipophilic peptides can be spontaneously embedded in the lipid layer of the carrier, which recognizes the TMD of the target receptor in a manner similar to the interactions between TMDs. This drug delivery platform have the following features: (1) avoids competitive interference by endogenous ligands or antibodies; (2) effectively overcomes target-missing effects caused by shedding of extracellular regions; (3) contains “plug-and-play” targeting moieties without chemical modifications, and the constructed formulations have low immunogenicity and good stability; and (4) has excellent universality for various lipid carriers and can be extended to other forms of active targeting mediated by transmembrane receptors.
a Challenges to orthosteric targeting based on recognition of the ectodomain of membrane proteins, including endogenous ligands or antibodies competitively binding to ectodomain, pseudo-targets caused by shedding of the ectodomain, and target loss due to mutations in the ectodomain. b Lipid carriers (e.g., liposomes, lipid nanoparticles, or exosomes) are modified with allosteric modified peptides in a “plug-and-play” approach that recognizes the transmembrane domains of membrane proteins to mediate targeted delivery. c Allosteric targeting strategies mediate delivery to the brain, through the blood-brain barrier, via insulin receptors lacking ectodomain for the treatment of Alzheimer disease.
Results
Design and characterization of the allosteric peptide binder that binds to the insulin receptor transmembrane domain
Rational protein design was employed to develop peptide binders. Given the hydrophobic environment, a thorough evaluation of the peptides’ transmembrane characteristics and their interactions with the target region within the membrane was conducted. Based on the binding modes and interaction analyses of the protein complex, interaction and non-interaction interfaces were delineated. Custom ranges of amino acid mutations were defined for each position in the design chain to optimize the affinity and specificity of the peptides. Two detailed iterative rounds of design were performed, ultimately leading to the selection of the optimal sequence for further analysis, guided by Rosetta Design energy scores (Fig. 2a). The 3D structure of the peptide was predicted by using AlphaFold2 and aligned to the truncated chain B of the IR TMD to determine the binding mode. The 3D structure was optimized by 2000 cycles of steepest descent and 3000 cycles of conjugate gradient minimization by using HawkDock16. Then the free energies of chain B and IR transmembrane domain-binding peptide (ITP) binding to chain A were estimated with the MM/GBSA method17. The predicted MM/GBSA binding free energies of the native peptide and designed peptide to the TMD of the IR were −42.68 kcal/mol and −43.59 kcal/mol, respectively, suggesting that ITP binds more tightly to the IR than the native peptide. In addition, per-residue energy contribution analysis was used to identify “hot spots” (absolute energy contribution >2 kcal/mol) on the truncated chain B (Val27, Phe30, lle34, lle37, and Leu41) and the designed peptide (lle4, Leu8, Phe11, Leu15, lle18, and Lys22), which are important for recognition and binding with the TMD of the IR (Supplementary Fig. 1).
a Rational design flow chart of ITP and related sequence information. (1) Homodimer structure of the transmembrane domain (TMD) of insulin receptor (IR) (IR-TM chain A: blue ribbon, chain B: orange ribbon, PyMOL). (2) Sequence logo of top 5 designed unique sequences after the first round Rosetta Design. This image was created by weblogo (https://weblogo.berkeley.edu/logo.cgi). (3) 3D structures of the top 5 sequences, predicted by AlphaFold2, utilized for the second round of design. (4) The binding mode of transmembrane domain (TMD) of insulin receptor (IR) with ITP (IR-TM chain A: blue ribbon, ITP: cyan ribbon, PyMOL). (5) An overlay of chain B and ITP. The main chains are shown as tube representations, while side chains are displayed as sticks, with all amino acid side chains labeled in their respective colors (Discovery Studio Visualizer). (6) Sequences of the insulin transmembrane domain receptor, Chain B, and ITP. b Characterization of affinity between ITP and IR by surface plasmon resonance (SPR). c Quenching of coumarin-labeled insulin receptor protein fluorescence by binding to ITP or scrambled ITP. d–g Competitive binding assay demonstrates ITP binding to allosteric sites of IR by flow cytometry. Insulin was labeled with FITC. **p < 0.01, ***p < 0.001, ****p < 0.0001, NS not significant. h The effect of ITP on insulin activity in brain microvascular endothelial cells (BMECs); phosphorylation was characterized by western blotting. i Schematic representation of IR conformational changes monitored via time-resolved fluorescence resonance energy transfer (trFRET) approach (Created in BioRender. Tu, D. (2025) https://BioRender.com/n28f408). j Effect of increasing concentrations of insulin/ITP/Scrambled ITP on the trFRET signal measured with the SNAP-tagged IR. Statistical significance was tested with two-tailed unpaired Student’s t tests. Data in (b–g, j) are presented as the mean ± SD (n = 3 biologically independent experiments). Data in (h) are representative of three independent experiments with similar results. Source data are provided as a Source Data file.
The molecular weight and purity of ITP, the transmembrane domain of IGF-IR (IGF-IR-TM), and the transmembrane domains of IR-related Receptor (IRR) (IRR-TM) were confirmed by electrospray ionization mass spectrometry and high-performance liquid chromatography (Supplementary Fig. 2). The α-helical structure of ITP was consistent with the prediction (Supplementary Fig. 3). Surface plasmon resonance (SPR) assay results showed that the dissociation constant (KD) between ITP and IR was 2.10 × 10−7 M (Fig. 2b), which was in sharp contrast to the KD for a scrambled peptide and IR (Supplementary Fig. 4a). Fluorescence resonance energy transfer (FRET) was used to further investigate the association of ITP with IR. Compared with that of the ITP condition, the scrambled ITP control strongly inhibited IR protein-coumarin fluorescence and weakened the FRET signal from fluorescein isothiocyanate (FITC) emission (Supplementary Fig. 4b, c). The calculated KD18 was consistent with the SPR results (Fig. 2c). In addition, ITP exhibited only mild interaction with itself, IGF-IR-TM, and IRR-TM, in contrast to its specific binding to IR-TM (Supplementary Fig. 4d–h). To demonstrate that ITP binds to IR by allosteric targeting, we evaluated the binding property via the competitive binding assay. Primary brain microvascular endothelial cells (BMECs) were isolated and preincubated with different concentrations of insulin or ITP, after which the uptake of FITC-labeled ITP or FITC-labeled insulin by the BMECs was measured by flow cytometry (Supplementary Fig. 5). The results showed that the uptake of FITC-insulin by BMECs was inhibited when they had been preincubated with insulin, because the previous insulin competitively bound to the ectodomain of the IR. In contrast, ITP pretreatment did not interfere with the uptake of FITC-insulin by the BMECs (Fig. 2d, e). Similar trends were observed in the uptake of FITC-labelled ITP by BMECs (Fig. 2f, g). The results above suggest that ITP and insulin bind to IR in a non-competitive manner. In addition, Western blots showed that ITP functions as a concentration-dependent agonist of IR. ITP induces the phosphorylation of IR as well as downstream signaling molecules AKT and ERK. Since ITP is an allosteric ligand, when insulin and ITP are treated simultaneously, their effects appear to be additive. Given that the IR internalization is dependent on IR phosphorylation, the results above suggest that ITP might independently induce IR internalization, thereby exerting an allosteric targeting effect19. Furthermore, the phosphorylation of AKT and ERK suggests that ITP may have an active role in regulating metabolism and promoting mitogenesis20 (Fig. 2h). We employed a method that has been reported to measure conformational changes in cell surface receptors, namely trFRET, to investigate the impact of ITP on the conformation of the N-terminal SNAP-tagged version of the IR (Fig. 2i)21. We observed that ITP exhibits a similar effect to insulin in inducing FRET signal changes associated with IR, suggesting that the conformational changes in IR induced by ITP may be analogous to those induced by insulin (Fig. 2j and Supplementary Fig. 6). Overall, these results suggest that ITP specifically recognizes the IR via allosteric binding.
Characterization of allosteric peptide-modified liposomes and their penetration of the blood-brain barrier
The mean diameter and ζ-potential of the PEGylated ITP-modified liposome (PEG-ITP-Lip) were 87.40 ± 9.81 nm and −24.57 ± 1.60 mV, respectively (Fig. 3a, b and Supplementary Table 1), and they were round and of uniform size (Fig. 3c and Supplementary Fig. 7). Given that ITP is embedded in the phospholipid layer of liposomes, this process may help to improve the stability of the liposomes. To evaluate the membrane fluidity, we labeled liposomes with NBD-PE (a lipid fluorescent probe) and measured the anisotropy values, with cholesterol-modified liposomes (CHL-Lips) used as controls. The results indicate that ITP reduced the mobility of phospholipids to a greater extent than did cholesterol (Fig. 3d and Supplementary Fig. 8). Moreover, the Young’s modulus of the ITP-modified liposomes (ITP-Lips) was approximately 33 times greater than that of the unmodified liposomes (Fig. 3e). The stability of the liposomes in mouse serum was evaluated with a FRET assay, which showed that the FRET efficacy of PEGylated ITP-modified liposomes (PEG-ITP-Lip) was 93.55% after 48 h of incubation in serum as compared with 69.50% for Lip and 82.51% for PEG-Lip (Fig. 3f and Supplementary Fig. 9). These FRET findings provide further evidence that ITP modification can improve the stability of liposomes.
a Size distribution and (b) transmission electron microscopy images of polyethyleneglycol ITP-modified liposomes (PEG-ITP-Lip) (scale bar = 50 nm). c Zeta potential of four liposomes. Data are presented as the mean ± SD (n = 3 biologically independent experiments). d–f Characterization of ITP modifications to improve the stability of liposomes. d The effect of ITP modification on membrane fluidity compared with cholesterol at different temperatures was characterized by measuring anisotropy values. Data are presented as the mean ± SD (n = 3 biologically independent experiments). e Young’s modulus of liposomes (Lip) and ITP-modified liposomes (ITP-Lip). Data are presented as the mean ± SD (n = 3 biologically independent experiments). *p < 0.05. f Fluorescence resonance energy transfer (FRET) efficiency of four liposomal formulations over 48 h of incubation with mouse serum. Liposomes were labeled with DiI and DiD. Data are presented as the mean ± SD (n = 3 biologically independent experiments). g–i In vitro uptake by bEnd.3 cells of liposomes modified with different ratios of ITP by (g) high content analysis system in confocal mode (Operetta CLS, PerkinElmer, USA) (scale bar = 10 μm) and (h, i) flow cytometry. Data are presented as the mean ± SD (n = 3 biologically independent experiments). ****p < 0.0001, **p < 0.01, NS not significant. j Schematic diagram of the in vitro BBB model (Created in BioRender. Tu, D. (2025) https://BioRender.com/h79d610). k Vertical 3D images of bEnd.3 monolayers (blue) interacting with liposomes (red). l Cumulative penetration of liposomes labeled with DiD (%) over 8 h. Data are presented as the mean ± SD (n = 3 biologically independent experiments). m Ex vivo and (n) in vivo fluorescence imaging of liposomes in normal mice with hair of brain and back removed. o The permeability of liposomes (green) in brain microvessels (red) as evaluated by immunofluorescence assay. The microvessels were stained with anti-CD34 antibody. Scale bar = 25 μm. Statistical significance was tested with two-tailed unpaired Student’s t tests. Data in (b, o) are representative of three independent experiments with similar results. Source data are provided as a Source Data file.
To verify the targeting of ITP, we examined the uptake of the ITP-modified liposomes (ITP-Lip) by bEnd.3 cells. Fluorescence imaging (Fig. 3g and Supplementary Fig. 10) and flow cytometry (Fig. 3h, i) showed that the 4% ITP modification (molar ratio) had the strongest uptake (p < 0.001). Moreover, 5% DSPE-PEG2000 (molar ratio) reduced the uptake of liposomes by RAW264.7 macrophage cells, potentially beneficial for their long-term circulation22 (Supplementary Fig. 11). The ability of ITP to cross the BBB was evaluated in vitro via a Transwell model (Fig. 3j). 3D imaging of bEnd.3 cell monolayers (blue) revealed that PEG-ITP-Lip/ITP-Lip (red) could effectively penetrate the BBB, but penetration of the PEG-Lip/Lip signal (red) was minimal (Fig. 3k). Further quantitative analysis was consistent with the fluorescence imaging results (Fig. 3l), which demonstrated that ITP modification could significantly promote liposomal penetration of the BBB in vitro. Ex vivo fluorescence imaging of the brain showed that the fluorescence intensity of PEG-ITP-Lip/ITP-Lip was significantly greater than that of the control group at all time points (Fig. 3m), and PEGylation effectively reduced the distribution of liposomes in the liver and extended their circulation time (Supplementary Fig. 12). The in vivo imaging results were consistent with the ex vivo imaging results, indicating that ITP still exhibits excellent brain-targeting effects in vivo (Fig. 3n). Immunofluorescence assays demonstrated that PEG-ITP-Lip/ITP-Lip could effectively penetrate brain microvessels and enter the brain parenchyma, which is essential for the subsequent treatment of brain diseases (Fig. 3o).
Evaluation of the allosteric mechanism in vitro and in vivo
To verify the mechanism by which ITP recognizes IR, the InsR gene was silenced in BMECs via siRNA transfection to suppress IR expression (Fig. 4a, b). The uptake of PEG-ITP-Lip/ITP-Lip was significantly inhibited in the BMECs with suppressed IR compared with their uptake by normal BMECs, indicating that ITP modifications were mediated through IR for active targeting (Fig. 4c, d). To validate the mechanism of ITP targeting in vivo, a brain-specific Insr-knockout mouse model was constructed with the Cre-loxP system (Insr CKO mice) (Supplementary Fig. 13). In vivo imaging of brains from Insr CKO mice revealed that the fluorescence intensity of PEG-ITP-Lip was significantly greater than that of PEG-Lip in flox/flox mice, but no difference was noted in fluorescence intensity between PEG-ITP-Lip and PEG-Lip in the Insr CKO mice (Fig. 4e and Supplementary Fig. 14). Ex vivo imaging showed that the fluorescence intensity of PEG-ITP-Lip in the brains of flox/flox mice was 2.06, 3.12, and 1.45 times greater than that of PEG-Lip at 1 h, 4 h, and 12 h (p < 0.05); moreover, no significant difference in fluorescence intensity was found in the Insr CKO mice (p > 0.05) (Fig. 4f–h). Moreover, immunofluorescence assays of brain sections were consistent with the in vivo and ex vivo fluorescence imaging results and revealed greater colocalization of PEG-ITP-Lip/ITP-Lip (green) with IR (red) in flox/flox mice than PEG-Lip/Lip (green) with IR (Fig. 4i). Thus, these in vitro and in vivo experiments demonstrated that the mechanism through which ITP modification achieves brain targeting is IR-mediated.
a Levels of insulin receptor (IR) protein in primary brain microvascular endothelial cells (BMECs) before and after siRNA transfection were evaluated by western blotting, with (b) corresponding semi-quantitative analysis. c, d The uptake of allosteric peptide-modified liposomes by BMECs was significantly decreased when IR expression was suppressed by siRNA transfection. Scale bar = 50 μm. e In vivo fluorescence imaging of PEG-Lip/PEG-ITP-Lip in Flox mice and brain-specific Insr-knockout mice (IR CKO) with hair of brain removed. f Ex vivo fluorescence imaging of PEG-Lip/PEG-ITP-Lip in Flox mice and Insr CKO mice, with (g, h) corresponding semi-quantitative analysis (normalized fluorescence intensity of (g) Flox or (h) CKO to PEG-Lip). i Distribution of liposomes in the brains of Flox mice (left) and CKO mice (right) and the co-localization of liposomes with IR, as characterized by immunofluorescence assay. White arrow indicates the co-localization of liposomes with IR. Scale bar = 50 μm. j Schematic diagram of using a peptide bound to the ectodomain of the IR (IEP) to block the IR (Created in BioRender. Ying, L. (2025) https://BioRender.com/ntzpswg). k Uptake of liposomes by BMECs with IEP blocking, as determined by flow cytometry. l Snapshot image of real-time single-cell multimodal analyzer, used to determine IR-α expression levels in BMECs treated or not treated with matric metalloproteinase 14 (MMP14). Scale bar = 20 μm. m Expression levels of IR-α on surfaces of BMECs treated or not treated with MMP14 as measured with a real-time single-cell multimodal analyzer. (F, Fluorescence intensity of IR-α on the cell membrane surface; F0: background fluorescence intensity of the culture dish). n Cellular uptake of liposomes by BMECs treated or not treated with MMP14. o Co-localization of liposomes with lysosomes, with (p) corresponding Pearson’s correlation coefficients. Scale bar = 5 μm. Statistical significance was tested with two-tailed unpaired Student’s t tests. *p < 0.05, **p < 0.01, ***p < 0.001, NS not significant. Data in (b, c, g, h, k, n, p) are presented as the mean ± SD (n = 3 biologically independent experiments). Data in (e, i) are representative of three independent experiments with similar results. Source data are provided as a Source Data file.
We further used IEP, a peptide bound to the insulin-binding site on the IR with a Kd value of 230 nM, to block the ability of BMECs to recognize ITP (Fig. 4j and Supplementary Fig. 15)23. Flow cytometry showed that IEP could not inhibit the uptake of PEG-ITP-Lip by the BMECs (p > 0.05), indicating that the site of action of ITP is different from that of IEP (Fig. 4k and Supplementary Fig. 16). We next used membrane-bound matrix metalloproteinase 14 (MT1-MMP/MMP14) to cleave the ectodomain of the IR in BMECs to construct a model of cells lacking the ectodomain of IR24. Expression of the α-subunit of IR (IR-α) on the surface of the cells was measured with a real-time single-cell multimodal analyzer, which showed that the expression of IR-α was 23.56 times lower in cells after MMP14 treatment, consistent with the western blotting results (Fig. 4l, m and Supplementary Fig. 17). Compared with uptake by normal BMECs, uptake of IEP-Lips in BMECs lacking IR-α was significantly inhibited (p < 0.05), but no significant difference was noted in the uptake of PEG-ITP-Lip (p > 0.05) (Fig. 4n). These findings further demonstrated that the allosteric targeting site of ITP is the IR TMD. In addition, we examined whether ITP-mediated BBB permeation depends on insulin-induced IR endocytosis. The results showed no significant difference in the penetration of PEG-ITP-Lip across the BBB model or its co-localization with IR within the cytoplasm, regardless of insulin concentration, whether low or high. This indicates that the transport of PEG-ITP-Lip is independent of insulin-induced IR endocytosis (Supplementary Figs. 18 and 19). In vivo imaging revealed no significant difference in barin fluorescence intensity between the fasting mice and normal mice, further confirming that ITP-mediated brain targeting is independent of insulin-induced IR endocytosis in vivo (Supplementary Fig. 20). Additionally, cellular uptake of PEG-ITP-Lip was significantly inhibited in vitro when pretreated with chlorpromazine, suggesting that the ITP-mediated endocytosis pathway is primarily clathrin-dependent (Supplementary Fig. 21). Given reports of a close relationship between the intracellular localization of proteins and the TMD12, we next examined the intracellular fate of PEG-ITP-Lip/ITP-Lip versus IEP-Lip. The results showed that IEP liposomes colocalized with lyosomes to a greater extent than did ITP liposomes; the Pearson correlation coefficients for that colocalization were PEG-ITP-Lip, 0.32; ITP-Lip, 0.31; and IEP, 0.67 (Fig. 4o, p), indicating that ITP modification can prevent rapid degradation by lysosomes after endocytosis.
Immunocompatibility evaluation of allosteric peptide modifications
Modification of functional molecules on the surface of PEGylated liposomes has been reported to have immunostimulatory toxic effects; for example, repeated administration of liposomes conjugated with the c(RGDyK) short peptide on the surface led to decreased body temperature and immune responses in mice. However, because ITP, as a targeting moiety, is embedded in the phospholipid layer by a “plug-and-play” approach, this strategy may help to improve the immunocompatibility of the formulation.
To test this hypothesis, we labeled the peptides with Cy5.5 fluorescein and prepared the corresponding liposomes. The fluorescence labeling percentage and fluorescence intensity of different liposomes were determined by Flow NanoAnalyzer (U30E, NanoFCM, China)25 (Supplementary Fig. 22). The results indicated that all liposomes and their associated peptides were loaded at comparable levels. Next, we used sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) to analyze the protein corona in serum from mice given the various liposome formulations and found that the concentration of PEG-ITP-Lip/ITP-Lip was significantly lower than that of IEP-Lip in terms of both the adsorbed protein species and the protein amount (Fig. 5a), and the adsorption of IgM by IEP-Lip was also greater than that by the other groups; the value was approximately 2.36 times greater than that of PEG-ITP-Lip (Fig. 5b, c). The reason for the lower ITP-modified adsorbed protein corona may be because ITP is embedded in the phospholipid layer, which would reduces its interaction with plasma proteins. A further series of in vivo experiments to evaluate immunocompatibility included a c(RGDyK)-modified liposome (RGD-Lip) as a positive control (Supplementary Fig. 23). Notably, mice in the RGD-Lip group developed hypothermia after the third dose, whereas mice in the other groups showed no changes in body temperature and otherwise appeared normal, suggesting that the readministration of RGD-Lip induces immune responses (Fig. 5d). We also evaluated the presence of IgM and IgG complexes in serum and in major tissues (liver, lung, and kidney) of mice after the third dose. Serum concentrations of IgM and IgG in the PEG-ITP-Lip/ITP-Lip -treated mice were significantly lower than those in the RGD-Lip/IEP-Lip -treated mice, and IgM and IgG immune complex despoits were the most prominent in liver, kidney, and lung in the RGD-Lip/IEP-Lip group compared with the other groups (Fig. 5e–h). Also, serum levels of IL-8 were comparable between the PEG-ITP-Lip/ITP-Lip group and the saline control group but were significantly lower than IL-8 levels in the RGD-Lip/IEP-Lip group (Fig. 5i, j).
a Separation of protein coronas on liposomes by SDS-PAGE. b IgM levels in protein coronas on liposomes, characterized by western blotting, with (c) corresponding semi-quantitative analysis. Data are presented as the mean ± SD (n = 3 biologically independent experiments). ***p < 0.001, **p < 0.01, *p < 0.05. d Body surface temperature of mice at 30 min after 3 doses of liposomes. The mice showed severe hypothermia when the RGD-Lip was given for the third time. e IgM and (f) IgG levels in the serum of mice with the corresponding liposomes as antigens at 1 h after the third dose were measured by indirect enzyme-linked immunosorbent assay. Data are presented as the mean ± SD (n = 3 biologically independent experiments). ***p < 0.001, **p < 0.01. g, h Immunohistochemical stains showing deposits of IgM and IgG immune complexes (red arrows) in liver, kidney, and lung at 1 h after the third dose. Scale bar = 50 μm. i IL-8 and (j) TNF-α levels in serum at 15 min after the third dose with liposomes. Data are presented as the mean ± SD (n = 3 biologically independent experiments). ***p < 0.001, **p < 0.01, NS not significant. Statistical significance was tested with two-tailed unpaired Student’s t tests. Data in (a) are representative of three independent experiments with similar results. Source data are provided as a Source Data file.
Overall, IEP-Lip, which targets the ectodomain of IR, could induce an immune response, but in contrast, the PEG-ITP-Lip/ITP-Lip formulations did not present a similar risk, which may be attributable to the specific modification of ITP that reduces its exposure in serum.
Therapeutic evaluation of ITP-LNP/siBACE1 in the treatment of Alzheimer disease
To validate the potential of allosteric targeting in conditions associated with ectodomain shedding of target receptors and the possibility of applying other vectors, we explored the effectiveness of lipid nanoparticles for delivering siRNA-BACE1 for the treatment of Alzheimer disease (AD), in which the ectodomain of the IR in the cerebrovasculature is reportedly cleaved26,27. For these experiments, we used APP/PS1 double transgenic mice, which show accelerated amyloid deposition and memory deficits, as a model of AD (Supplementary Fig. 24). Because the AD-like features of APP/PS1 mice can be improved by inhibiting BACE1, we evaluated the efficacy of these lipid nanoparticles in terms of both pathological and behavioral characteristics. After confirming that our siRNA sequences could effectively inhibit BACE1 mRNA (by 91%) and protein expression (by 76%) in mouse Neuro-2 cells, we tested the effects of siRNA-loaded ITP-modified lipid nanoparticles (ITP-LNPs/siRNAs) in APP/PS1 mice (Supplementary Figs. 25–27 and Supplementary Table 2). In vivo imaging demonstrated that ITP-modified lipid nanoparticles efficiently delivered Cy5-siBACE1 to the brain and enhanced the stability of Cy5-siBACE128 (Supplementary Fig. 28). Eight-month-old APP/PS1 mice were administered seven injections of 1 mg/kg siRNA in the form of ITP-LNP/siBACE1, IEP-LNP/siBACE1, LNP/siBACE1, ITP-LNP/siScr, LNP/siScr, or PBS (Fig. 6a), and the effects on spatial learning and memory, β-amyloid deposition, and BACE1 expression were tested. Spatial learning was evaluated with the novel object recognition test (NOR) and memory with the Morris water maze test (MWM)29. The NOR test findings showed improved spatial learning in APP/PS1 mice treated with ITP-LNP/siBACE1 (DI > 0), but the other treatment groups (including LNP/siBACE1) had no significant effects (Fig. 6b–d). Analysis of the swimming routes of the mice in the MWM test of memory (Fig. 6e) showed that APP/PS1 mice treated with ITP-LNP/siBACE1 and the WT (saline-treated) mice could repeatedly find the platform in the target quadrant, and the number of platform crossings, the length of the platform, and the length of the target quadrant were significantly greater than those of the mice given LNP/siBACE1, ITP-LNP/siScr, or LNP/siScr; no differences were noted in swimming speed (Fig. 6f–i). These findings indicate that treatment with ITP-LNP/siBACE1 led to the greatest improvement in the behavior of the APP/PS1 mice. We also found that treating APP/PS1 mice with ITP-LNP-siBACE1 led to greater deposition of β-amyloid (green) in the mouse brains, but no changes were found in the brains in the other treatment groups, including LNP/siBACE1 (Fig. 6j). Similarly, BACE1 expression in the hippocampus and cortex of APP/PS1 mice treated with ITP-LNP/siBACE1 was inhibited and similar to that in WT (saline control) mice compared with that in the other groups (Fig. 6k–m). Overall, ITP-LNP/siBACE1 was more effective at improving behavioral and AD-like pathological features in APP/PS1 mice than compared with the other treatment conditions, reflecting the potential of allosteric targeting for the treatment of disorders associated with deletion of the ectodomain of the target receptor.
a Timeline for treatment of APP/PS1 or WT mice with siRNA-loaded lipid nanoparticles (ITP-LNP) or PBS by tail vein injection every 2 days. After treatment, the mice were given novel object recognition (NOR) and Morris water maze tests, after which serum and brain samples were obtained for subsequent evaluation. b Schematic diagram of the NOR test. The orange circle represents the familiar subject and the green triangle indicates the novel subject. c Discrimination index (DI) and (d) preference index (PI) findings from the NOR test. Data are presented as the mean ± SD (n = 6 biologically independent experiments). *p < 0.05, NS not significant. e Path diagram of APP/PS1 and WT mice performing the MWM test. The green dashed circles refer to the target platform in the MWM test. f Number of platform crossings, (g) time in the platform (s), (h) time in the target quadrant (%), and (i) swimming speed of mice in the MWM test. Data are presented as the mean ± SD (n = 6 biologically independent experiments). ***p < 0.001, **p < 0.01, *p < 0.05, NS not significant. j Immunofluorescence assays show deposition of β-amyloid (green) in the hippocampus and cortex of APP/PS1 and WT mice after treatment. Scale bar = 50 μm. k Expression levels of BACE1 protein in the hippocampus and cortex of APP/PS1 and WT mice after treatment, as characterized by western blotting, with (l, m) corresponding semi-quantitative analysis. Data are presented as the mean ± SD (n = 3 biologically independent experiments). ***p < 0.001, **p < 0.01, *p < 0.05. Statistical significance was tested with two-tailed unpaired Student’s t tests. Source data are provided as a Source Data file.
In addition, both hematoxylin-and-eosin staining of heart, liver, spleen, lung, and kidney and analysis of blood levels of alanine aminotransferase, aspartate aminotransferase, blood urea nitrogen, creatinine, alkaline phosphatase, and uric acid revealed no obvious pathological or functional changes among the various treatment groups (Supplementary Figs. 29 and 30). Analysis of several proinflammatory cytokines, including IL-1β, IL-6, TNF-α, and IFN-γ, also revealed no significant differences between treatment groups (Supplementary Fig. 31). Collectively, these results suggest that ITP-LNP/siBACE1 had good biocompatibility and safety in vivo.
Allosteric targeting as a platform strategy for active drug delivery
Research on mRNA delivery has received extensive attention, including the delivery of mRNA to specific tissues (e.g., liver, spleen, and lung) by selective organ targeting, but additional research is needed to overcome the BBB30. We extended our research on the IR transmembrane domain-binding peptide (ITP) to explore its potential for targeted delivery of mRNAs to the brain. We encapsulated eGFP-mRNA in lipid nanoparticles modified with the ITP (ITP-LNP/mRNA) or unmodified LNP (LNP/mRNA) and injected the particles intravenously in mice. Fluorescence imaging of the eGFP (green) in brain sections from those mice showed that ITP-LNP/mRNAs could effectively cross the BBB (Fig. 7a). Moreover, 3D fluorescence imaging of the whole brain confirmed that mRNA could be efficiently and consistently delivered into the brain parenchyma via allosteric targeting and expression of the eGFP (Fig. 7b, c).
a The expression of eGFP protein in the brains of mice was evaluated by immunofluorescence assay after intravenous injection of lipid nanoparticles modified with the ITP and containing mRNA (ITP-LNP/mRNA) versus LNP/mRNA. Scale bar = 50 μm. b, c 3D fluorescence images of eGFP (green) in the brains of mice obtained by light sheet fluorescence microscopy at 24 h after administration of (b) LNP/mRNA or (c) ITP-LNP/mRNA. d In vitro uptake of Cy5-siRNA-loaded exosomes (Exo/siRNA) and Cy5-siRNA-loaded ITP-Exosome (ITP-Exo/siRNA) by bEnd.3 cells, measured by confocal laser scanning microscopy (scale bar = 10 μm). e In vitro uptake of Exo and ITP-Exo by bEnd.3 cells measured by flow cytometry. Exosomes were labeled with DiO. f The encapsulation efficiency of siRNA in the exosomes/ITP-modified exosomes was determined with a RiboGreen assay. g Design of the ITP2 peptide bound to αv transmembrane domain (αv-TM). The first row illustrates the evolution of the sequence during the design process, with main chains represented by ribbons of different colors and side chains shown as sticks (PyMOL). The color descriptions are provided in the “All Sequences” section of the second row. In the second row, on the left, an overlay of anti-αv and ITP2 is represented in green and orange, respectively. The main chains are depicted using line ribbon representation, while side chains are displayed as sticks (Discovery Studio Visualizer). On the right, the sequences of all chains in this figure are provided. In the designed ITP2, the red font indicates introduced amino acids, while cyan highlights and underlines represent amino acids that have not been altered. h, i Characterization of affinity between ITP2 (left) / c(RGDyK) (right) and bEnd.3 vesicles by surface plasmon resonance. j In vitro uptake of ITP2, scrambled ITP2/c(RGDyK)-modified liposomes (ITP2-Lip, SP-Lip, and RGD-Lip) by bEnd.3 cells, measured by flow cytometry. Statistical significance was tested with two-tailed unpaired Student’s t tests. *p < 0.05, **p < 0.01, ***p < 0.001. Data in (e, f, j) are presented as the mean ± SD (n = 3 biologically independent experiments). Source data are provided as a Source Data file.
In addition to mRNA, we also examined the feasibility of this platform for delivering the small molecule amphotericin B (AmB) for the treatment of fungal meningitis31; for this purpose, we loaded PEGylated liposomes that were modified or not modified with ITP with AmB. The in vitro release profile showed slow release of the AmB-loaded PEG-ITP-Lips and significant growth inhibition of C. neoformans (minimum inhibitory concentration of 2 μg/mL) (Supplementary Figs. 32 and 33a). In vivo experiments with mice injected with AMB-PEG-ITP-Lips tested fungal burden and survival. Grocott methenamine silver staining of brain sections from the infected mice showed that PEG-ITP-Lip/AmB significantly reduced the amount of C. neoformans in the brain, and the survival rate of the mice was 42% at week 6; in contrast, mice treated with Lip/AmB had died by day 21 after infection, and those treated with PEG-Lip/AmB had died by day 24 (Supplementary Fig. 33b–d). Safety evaluation showed that the PEG-ITP-Lip/AmB particles exhibited no toxic effects on organ function, as demonstrated by hematoxylin and eosin staining and blood variable analyses. This lack of toxicity ensured no additional burden on the infected mice, which is crucial for significantly improving their survival rates. (Supplementary Figs. 34 and 35). Additionally, we conducted experiments to compare the uptake of ITP-Lip, IEP-Lip, and Lip by MCF-7 cells (high IR expression) and T-47D cells (low IR expression)32. The results revealed that ITP-Lip exhibited greater selectivity than IEP-Lip, highlighting that the ITP modification strategy effectively reduces potential impacts on non-targeted organs (Supplementary Fig. 36). Another particle type used as drug delivery vehicles is the exosome, a natural source of lipid vesicles33,34,35,36. We isolated exosomes derived from RAW264.7 macrophage cells and characterized their particle size, morphology, and common markers (CD63, CD81, and TSG101)37,38 (Supplementary Fig. 37). In vitro, the uptake of ITP-modified exosomes (ITP-Exos) encapsulated or not encapsulated with siRNA by bEnd.3 cells was significantly greater than the uptake of uncoated exosomes (Exos) (Fig. 7d, e). Moreover, compared with the control, the ITP-Exos could be loaded with siRNA more efficiently and leakage avoided (p < 0.05), which may be attributable to the improved stability of the exosome via ITP modification (Fig. 7f).
Given that more than 60% of targets in targeted therapy are membrane proteins39, we chose another membrane receptor, integrin alpha v (αv), to explore whether allosteric targeting could be expanded to a broader range of active targeted drug delivery. We initially constructed a complex of αv-TMD and its binder, anti-αv, based on the backbones of the A64–A86 helix and A121–A141 helix in the G3P transporter (PDB ID: 1PW4)40; sequence replacement of the anti-αv peptide was done with Discovery Studio visualizer 4.5 to improve the physicochemical properties, and the model of the αv-TMD/anti-αv complex was further relaxed by using Rosetta. After a reasonable starting all-atom model was retrieved, residue repacking at a specific position was done with the Rosetta platform to screen for peptides (ITP2) that specifically bind to αv-TMD40 (Fig. 7g and Supplementary Fig. 38). The HawkDock MM/GBSA module was then used to calculate the total binding free energy and the per-residue contributions to affinity (Supplementary Fig. 39). The hotspot residues in αV-TM include Val4, Leu7, Ala11, Leu15, and Leu19. In ITP2, the key amino acid residues contributing to its affinity are ILE5, Phe9, Thr13, Phe17, and Met20. Compared to the anti-αv peptide, the optimized ITP2 exhibits improved water solubility. The SPR results showed that the KD values of RGD, a classical peptide ligand that binds to the ectodomain of integrin, and ITP2, which binds to bEnd.3 vesicles expressing integrin αvβ3, were 17.10 μM and 9.66 μM, respectively; thus, ITP2 and RGD had similar affinities for integrin αvβ3 (Fig. 7h, i and Supplementary Fig. 40). Indeed, the uptake of ITP2-modified liposomes (ITP2-Lip) by bEnd.3 cells was significantly greater than the uptake of scrambled ITP2-modified liposomes (SP-Lip) (Fig. 7j and Supplementary Fig. 41). In vivo and ex vivo imaging further showed that mice given ITP2-Lip exhibited stronger brain fluorescence signals than did mice given SP-Lip group at all time points, demonstrating that ITP2 can effectively mediate delivery to the brain through BBB transport (Supplementary Figs. 42–44). Finally, delivery of the classical chemotherapy drug paclitaxel (PTX) encapsulated in liposomes (ITP2-Lip/PTX) by intravenous administration significantly extended the survival of mice bearing GL261 orthotopic gliomas (Supplementary Fig. 45).
In summary, allosteric targeting as a delivery platform is not only effective for delivering nucleic acids and small molecules but also is suitable for use with a wide range of lipid carriers; notably, this approach can also be extended to other membrane proteins. We conclude that allosteric targeting has great potential for further expansion in drug delivery research.
Discussion
Peptides have been widely used as ligands for targeted drug delivery and diagnostic imaging owing to their small size, ease of synthesis and functionalization, and high targeting affinity, thus combining the advantages of small molecules and biomacromolecules41. However, the complexities and barriers present in the in vivo environment still present ongoing challenges for the optimal design of peptide-based targeting moieties: (1) Conventional peptide ligands are susceptible to enzymatic barriers. The introduction of nonnatural amino acids can effectively improve the biostability of peptides but can sometimes increase their potential immunogenicity42. (2) Specific binding of low-affinity ligands to receptors usually requires high or even supersaturated concentrations to ensure effective recognition. However, high-affinity ligand conjugation easily traps nanoparticles in endosomes/lysosomes, preventing them from being effectively dissociated, resulting in decreased cellular translocation efficiency. Therefore, the design of ligands with moderate affinity is particularly important for efficient transport of the formulation43. (3) Although some peptide ligands can effectively recognize and bind to target receptors, they are unable to effectively penetrate cell barriers present because of poor membrane permeability. For example, integrin αvβ3 is highly expressed in the BBB, but its typical ligand, the RGD peptide, cannot effectively mediate BBB penetration for brain-targeted delivery. He et al. modified liposomes after coupling the RGD peptide to the cell-penetrating peptide R8, thereby facilitating penetration of the BBB by the formulation and targeting gliomas in the brain44. In the present study, our design of peptide ligands based on TMDs is beneficial for optimizing targeting moieties, as the resulting ligands have a natural affinity for lipid membranes and can be spontaneously embedded within them. Anchoring peptide ligands in the phospholipid layer could prevent their degradation by enzymes and suppress immunogenicity while facilitating membrane penetration. Moreover, because the TMD is less accessible than the extracellular region, this approach may facilitate the design of peptide ligands with moderate binding to better distinguish between target cells and nontarget cells.
It is noteworthy that current computational design methods for targeting protein interactions in transmembrane domains generally fall into two categories. The first is the rational design approach exemplified by the Computed Helical Antimembrane Protein (CHAMP) method, developed by William F. DeGrado’s team40, which achieved precise assembly by establishing the Crick parameter equation for transmembrane helices and using RosettaMP design to obtain de novo CHAMP sequences, combined with a more rational energy scoring mechanism (Rosetta’s ‘PackStat’ score)45 to continuously improve the design accuracy. Another class such as the multipass transmembrane protein design method developed by David Baker’s team46, which successfully constructed a complex topology containing 8 transmembrane domains by realising the connection of dispersed helices and sequence optimisation through Rosetta. Notably, these breakthroughs all used the Rosetta suites as core tool, confirming its versatility in the field of transmembrane design. In this study, transmembrane peptide design was achieved using the basic Rosetta module to validate the allo-targeted delivery strategy. From the perspective of transmembrane peptide design, there is still room for significant improvement in the future, including at least the following aspects: (1) integrating the ProteinMPNN deep learning model to expand the sequence space and achieve the synergistic optimisation of peptide length and sequence diversity; (2) adopting the combined generative model-MD strategy to dynamically evaluate the binding free energy; (3) combined with a virtual screening-conformational validation cycle to systematically improve the target affinity and transmembrane stability of peptides.
Lipid carriers such as liposomes, lipid nanoparticles, and lipid nanodiscs have all had important roles in the field of life sciences as tools for studying proteins, nucleic acids, and others, and have been useful in the development of drugsas well. Bionic carriers, such as extracellular vesicles, cell membrane vesicles, and cell membrane-encapsulated preparations, further enrich the variety of lipid carriers, and even cells themselves can be regarded as living lipid carriers. Such carriers often require further functional integration, though, and mild, efficient, and noncovalent modifications are most suitable for ligand-engineered carriers. This “plug-and-play”-style modification of targeting ligands that recognize the TMD of membrane proteins means that additional functions are tunable and modularized, and this modularity allows the efficient construction of personalized drug delivery systems by establishing a library of ligands in advance and combining them with appropriate vectors simply and efficiently.
Engineered bacteria have also shown potential for drug delivery and smart healthcare, especially as interactive cancer treatment tools47. Bacteria can be endowed with programmable properties to interact with specific targets through artificial, self-assembled receptors48. Thus, designing ligands and assembly strategies based on TMD interactions could assist with the intelligent delivery of various types of lipid-containing carriers.
In conclusion, we designed and screened peptide ligands that bind specifically to the TMD of the IR and constructed a series of allosteric brain-targeted lipid delivery systems. This allosteric targeting approach can effectively avoid pseudotargeting or loss of target, and the “plug-and-play” targeting moieties effectively improved the biocompatibility and stability of the formulation and showed good universality for various lipid-based carriers and membrane receptors. Therefore, this approach is expected to enrich the concept of active targeting and provide valuable insights for the development of smart healthcare.
Method
Experimental animals
BALB/c mice (6–8 weeks, 18–22 g) and Sprague-Dawley (SD) rats (8 weeks, 180–220 g) were purchased from the Chongqing Academy of Chinese Materia Medica (Chongqing, China). Brain-specific InsR-knockout mice (C57BL/6J-Insrem1cyagen) were obtained from Cyagen Biosciences, Inc. (Guangzhou, China). APP/PS1 mice (8 months) were purchased from Beijing Viewsolid Biotechnology Co., Ltd. (Beijing, China). Male and female mice were used in the experiment, and mice were randomly assigned to experimental groups. All animals were raised in a specific pathogen-free laboratory animal environment with a 12-h light-dark cycle at 22–24 °C and 30–50% relative humidity. All animal experiments were performed under the relevant guidelines and regulations of the Institutional Animal Care and Use Committee (IACUC) of Southwest University (IACUC Issue No. IACUC-20200525-02 & IACUC- 20240228-24).
Modeling the homodimer structure of the insulin receptor
The design of the peptide binder was inspired by the native structure of the transmembrane domain (TMD) of the IR. First, the monomer structure (chain A) of the TMD of the IR (aa 14–42) was retrieved from the PDB database49 (PDB ID: 2MFR). Then, the monomer structure (chain B) was created from chain A, and both structures were submitted to the ZDOCK server50,51,52 to obtain the homodimer structure. The homodimer complex with the best score from the 2000 ZDOCK predictions was selected for de novo protein design.
De novo design of peptides binders targeting the TMD of the insulin receptor
Starting from the predicted homodimer structure of the TMD of the IR, we used Rosetta Design, which includes the Rosetta fast relax53 and fixbb protocols54, to design potential peptide binders55. The aa 14–19 and aa 42 in chain B were truncated because they had little interaction with chain A. RosettaDesign was run with chain B (aa 20–41) having been set as the designed chain, and chain A set as the target protein. Based on the results of the protein-protein interaction analysis, the residues in chain B were categorized into interaction interface residues and non-interaction interface residues. In the resfile, different ranges of amino acid mutations were set for these two categories. For the non-interaction interface residues, a total of nine residues were identified: 21, 22, 25, 28, 29, 32, 33, 36, and 39 on chain B. Given that these positions are primarily located in the hydrophobic environment of the lipid bilayer, the mutation range for these amino acids was restricted to 11 non-polar amino acids: ACFGILMPVWY. For the near-interaction interface residues, a total of 13 residues were identified at positions 20, 23, 24, 26, 27, 30, 31, 34, 35, 37, 38, 40, and 41 on chain B. The mutation ranges for these positions were carefully tailored. Specifically, for Ser31 and Gly35, which form a common small-X3-small motif frequently observed in transmembrane helix pairs (PMID: 26725515), the mutation range was restricted to small amino acids (ASGT). Next, for the terminal amino acids Ile20, Phe40, and Leu41, we considered that, during transmembrane insertion, the peptide must first anchor to the hydrophilic groups of the lipid heads. Thus, the mutation range for these three residues was set to soluble groups (KHRDEST) (PMID: 17395823). Finally, for the remaining 9 amino acid residues at the interaction interface, we carefully balanced the hydrophilic and hydrophobic properties of the peptide. The mutation range was restricted to 8 hydrophobic amino acids and 2 small polar amino acids (GAVLIMFYST). For the receptor chain A, no new mutation design was performed; only repacking was conducted. The number of designed sequences was set to 10,000 and the other settings were set to the default parameters. The 3D structures of the top 5 designed unique sequences were predicted by using AlphaFold251,56 and aligned to the truncated chain B for redesign55. After two rounds of design, the most promising sequence (ELGILIFLYLFSLILGIIYWKK) was identified, but its efficacy required validation.
Surface plasmon resonance (SPR) measurements
The kinetics of binding between the peptides to IR or to bEnd.3 vesicles were determined by surface plasmon resonance (SPR) (Nicoya Lifescience, Waterloo, Canada). Briefly, the L1 sensor chip was installed, the system was filled with HEPES buffered saline (HBS), 80% iso-propanol was injected throughout the pipeline to remove air bubbles from the system, and HBS was used to stabilize the drift signal. Then, the L1 sensor chip was activated with 20 mM CHAPS three times, and the IR recombinant liposome or bEnd.3 vesicles were immobilized on an L1 sensor chip at a constant flow rate. Different concentrations of peptides were injected to measure the interaction between the peptides and the immobilized samples. All concentrations were tested in triplicate.
Fluorescence resonance energy transfer (FRET) assay
FRET experiments were conducted at room temperature in a 1 × 10 mm quartz cell on a fluorescence spectrometer (F-7000, HITACHI, JPN) as described elsewhere18. In brief, 400 nM (7-hydroxy-3-carboxyamide coumarin)-labeled IR protein (ab70687; abcam) in 20 mM HEPES (pH 7.4) and 1.0 mM C14 betaine micelle buffer were titrated with increasing concentrations of FITC-tagged IR domain-binding peptide (ITP) or FITC-tagged scrambled ITP. The excitation wavelength was set at 390 nm. Emission at 410–590 nm was recorded. Samples containing the same amount of fluorescence acceptor (FITC-tagged ITP/FITC-tagged scrambled ITP) served as a reference. Quenching of the fluorescence signal of the coumarin-labeled IR protein was analyzed with the equation:
Allosteric binding of ITP and IR
Competitive binding assay57
Primary BMECs were pre-seeded in 12-well plates (2 × 105 per well) and cultured at 37 °C for 24 h. The medium was then discarded and replaced with fresh medium containing insulin at concentrations of 2, 50, and 1250 nM or ITP at concentrations of 1.25, 2.5, 5, and 10 μM. Cells were incubated at 37 °C for 1 h, after which the medium was replaced with fresh medium, 50 nM FITC-insulin was added, and the mixture was incubated at 37 °C for 1 h. Then, the BMECs were collected and washed three times with PBS, and quantitative uptake was tested with a FACS Calibur flow cytometer (BD Biosciences, USA). In addition, we carried out similar experiments in which cells were pretreated with ITP at concentrations of 5, 10, and 20 μM or insulin at concentrations of 2, 10, 50, 250, and 1250 nM, and then 5 μM FITC-ITP was added, and other steps were performed as above.
The effect of ITP on insulin activity57
BMECs were pre-seeded in 6-well plates (4 × 105 per well) and cultured at 37 °C for 24 h. The medium was discarded, replaced with fresh medium containing 1, 5, 10 μM ITP, or 2 nM insulin, or a mixture of ITP (10 μM) and insulin (2, 10, 50 nM), and the cells were incubated at 37 °C for 10 min. Then, the BMECs were collected, and phosphorylation of the IR-β and protein kinases (AKT (Thr308), AKT (Ser473), and ERK1/2 (T202/Y204)) were detected via western blotting.
Uptake by bEnd.3 cells in vitro
Uptake of ITP or coumarin (C6) -labeled liposomes by bEnd.3 cells in vitro was evaluated by fluorescence imaging and fluorescence-activated cell sorting. Briefly, bEnd.3 cells were preseeded in 96-well plates (Cell Carrier Ultra, PerkinElmer, 1 × 104 per well) and cultured at 37 °C for 24 h. Different molar ratios of ITP- or DSPE-PEG2000-modified C6-labeled liposomes were added to the plates at a final concentration of 1 mg/mL lecithin and incubated at 37 °C for 2 h. The bEnd.3 cells were then fixed with 4% paraformaldehyde (PFA) at room temperature for 1 h and stained with 4′,6-diamidino-2-phenylindole (DAPI) at room temperature for 5 minutes. Finally, the bEnd.3 cells were washed three times with PBS, and the samples were imaged with a high-content screening and analysis system (Operetta CLS, PerkinElmer, USA). For further quantitative analysis, bEnd.3 cells were preseeded in 12-well plates (2 × 105 per well) and cultured at 37 °C for 24 h. C6-labeled liposomes were added to the plates at a final concentration of 1 mg/mL lecithin and incubated at 37 °C for 2 h. Then, the bEnd.3 cells were collected and washed three times with PBS, and uptake of the liposomes was quantified with a FACS Calibur flow cytometer (BD Biosciences, USA).
Transcytosis assay in an in vitro model of the BBB
bEnd.3 cells were inoculated into 24-well Transwell plates and incubated; the transendothelial electric resistance of the cell monolayers was checked continuously until it reached 150 Ω·cm58. Then liposomes labeled with the lipophilic fluorescent dye DiD were added to the apical chamber of the Transwell plates and incubated at 37 °C for 2 h, after which the bEnd.3 cells were fixed with 4% PFA at room temperature for 1 h and stained with DAPI at room temperature for 5 min. The Transwell membranes were subsequently transferred to microscope slides and scanned on the z-axis by high-content screening to obtain images of fluorescence penetration.
For further quantitative analysis, the medium in the basolateral chamber was collected at 0.5, 1, 2, 4, and 8 h after the addition of liposomes, and the permeated DiD fluorescence intensity was measured with an Infinite F200 pro multimode reader (Tecan, Switzerland).
Construction and evaluation of brain-specific Insr-knockout mice (C57BL/6J-Insrem1cyagen)
Brain-specific Insr-knockout mice were generated by using Cre-loxP system59. In brief, gRNA to the mouse Insr gene, a donor vector containing loxP sites, and Cas9 mRNA were coinjected into fertilized mouse eggs to generate targeted conditional knockout offspring. F0 founder animals were identified by polymerase chain reaction (PCR) followed by sequence analysis and were subsequently bred to wild-type mice to test germline transmission and F1 animal generation. An F1-targeted mouse was bred with a tissue-specific Camk2a-Cre deletion mouse to generate F2.
Mice were genotyped via PCR as follows. Genomic DNA samples were prepared by lysing a 2- to 5-mm segment of the mouse tail. The sequences of primer 1 used were as follows: 5′-GAGTTTCTCAACCTTCCTAGCACG-3′ (sense) and 5′-GAAACAGAGTCAACTAGCATCACT-3′ (antisense) (homozygotes: one band of 291 bp; heterozygotes: two bands of 291 bp and 231 bp; wild-type allele: one band of 231 bp). The sequences of primer pair 2 used were as follows: 5′-GTTCTCCGTTTGCACTCAGG-3′ (sense) and 5′-CAGGTTCTTGCGAACCTCAT-3′ (antisense) (Cre amplicon: ~500 bp). The expression of IR in the brain was analyzed semiquantitatively by western blotting.
In vivo and ex vivo fluorescence imaging
For in vivo fluorescence imaging, mice were randomly assigned to different treatment groups, and DiR-labeled formulations were injected via the tail vein (100 μg DiR/kg). The mice were anesthetized at 0.5, 1, 2, 4, 8, and 12 h after injection, and the fluorescence distribution was observed and recorded with an in vivo imaging system (Vieworks, Smart-LF, Korean)60.
For ex vivo fluorescence imaging, DiR-labeled formulations were injected as described above, and the mice were euthanized at 1, 4, 8, and 12 h afterward. The brain, heart, lung, liver, spleen, and kidney were carefully collected and washed, and images of the fluorescence distribution in each isolated tissue sample were obtained.
Evaluation of the permeability of the brain microvasculature to the liposomes
The permeability of the brain microvasculature to the liposome formulations was assessed by immunofluorescence analysis6,61. Briefly, the mice were randomly assigned to one of four treatment groups, and C6-labeled liposomes were injected via the tail vein. The mice were euthanized at 2 h after administration, and brain tissues were carefully collected, dehydrated, embedded, sectioned, and transferred to microscope slides. Samples were fixed with 4% PFA at room temperature for 30 min, permeabilized with 0.1% Triton-100 at room temperature for 15 min, and blocked with 5% bovine serum albumin (BSA) solution at 37 °C for 1 h. Then, an anti-CD34 antibody was added and incubated at 4 °C overnight, after which a goat anti-rabbit IgG/Alexa Fluor 594 secondary antibody was added and incubated at 37 °C for 1 h. Samples were then stained with DAPI at room temperature for 5 min and imaged with high-content screening.
Silencing the Insr gene in primary brain microvascular endothelial cells (BMECs)
The sequences of the siRNA-Insr used were as follows: 5′-GCACAGACCAUUUGAGAAATT-3′ (sense) and 5′-UUUCUCAAAUGGUCUGUGCTT-3′ (antisense). BMECs were isolated from the brains of SD rats and cultured in Dulbecco’s modified Eagle’s medium supplemented with 20% fetal bovine serum (Gibco, USA)62. The medium was replaced with fresh medium without antibiotics or serum when the BMEC density reached 50%. A mixture of siRNA and Lipofectamine 2000 was added to 6-well plates, and incubated for 6 h, and then the medium was replaced with fresh medium containing 10% serum and antibiotics, and the cells were incubated for 48 h.
After Insr gene expression was silenced by siRNA, the protein content of the IR-treated BMECs was quantified via western blotting. In brief, the BMECs were lysed, and the protein concentration was measured via a bicinchoninic acid protein assay kit. Proteins were separated by sodium dodecyl sulfate polyacrylamide gel electroporesis (SDS-PAGE), and the separated proteins were transferred to polyvinylidene fluoride membranes. Then, the proteins were blocked with 5% nonfat milk at 37 °C for 1 h and incubated with an anti-insulin receptor-β antibody (ab278100) overnight at 4 °C, followed by another incubation with a secondary antibody, goat anti-rabbit IgG/HRP (bs-0295G-HRP), at 37 °C for 1 h, and the protein bands were visualized by BeyoECL Star. The intensity of the protein bands was analyzed semiquantitatively by using ImageJ software. Uptake of the various liposomes by the transfected BMECs was analyzed by fluorescence imaging and flow cytometry.
Distribution of liposomes in the brains of brain-specific Insr-knockout mice
The distribution of liposomes in the brain tissue was evaluated by immunofluorescence imaging. The CKO and Flox mice were randomly assigned to one of two groups, and liposomes were injected into the mice via the tail vein. The mice were euthanized at 2 h after administration, and the brains were carefully collected, dehydrated, embedded, sectioned, and transferred to microscope slides. The samples were fixed with 4% PFA at room temperature for 30 min, permeabilized with 0.1% Triton-100 at room temperature for 15 min, and blocked with 5% BSA solution at 37 °C for 1 h. Then, rabbit anti-insulin receptor beta polyclonal antibody (bs-0290R) was added and incubated at 4 °C overnight, after which goat anti-rabbit IgG/Alexa Fluor 594 secondary antibody was added and incubated at 37 °C for 1 h. The samples were then stained with DAPI at room temperature for 5 min and imaged with high-content screening.
Uptake by BMECs after IR-EM-binding peptide (IEP) blockade
BMECs were preseeded in 12-well plates (2 × 105 per well) and cultured at 37 °C for 24 h, after which the medium was discarded, and fresh medium added that contained IEP at final IEP concentrations of 0.8%, 0.4%, and 0.2% (molar ratio) of the phospholipids, and the cells were then incubated for 1 h. The medium was then replaced with fresh medium, and the C6-labeled liposomes were added to the corresponding 12-well plates, which were incubated at 37 °C for 2 h. The BMECs were subsequently collected, washed three times with PBS, and their uptake of liposomes was quantified via FACS Calibur flow cytometry (BD Biosciences, USA)63.
Cleavage of the extracellular domain of IR by membrane-bound matrix metalloproteinase 14 (MT1-MMP/MMP14)
Membrane type 1 matrix metalloproteinase (MT1-MMP/MMP14), a membrane-bound Zn-containing endopeptidase, cleaves a wide variety of substrates ranging from the extracellular matrix to growth factor receptors, e.g., IR and lymphatic vessel endothelial hyaluronan receptor-1 (LYVE-1), and these experiments were done as described elsewhere with some modifications24,64,65. Briefly, BMECs were pre-seeded in 12-well plates (2 × 105 per well) and cultured at 37 °C for 24 h. The medium was then discarded and replaced with fresh medium containing 100 ng/mL recombinant human MMP14, and the cells were incubated at 37 °C for 12 h. The BMECs were subsequently collected, proteins were isolated, and cleavage of the extracellular domain of the IR was verified via western blotting.
Real-time single-cell quantitative analysis
BMECs (1 × 104) were seeded on poly-L-lysine-coated culture dishes and cultured overnight, after which recombinant human MMP14 was added to the culture dish and incubated for 12 h. The BMECs were then washed three times with PBS and fixed with 4% PFA for 0.5 h. Recombinant anti-insulin receptor alpha antibody [EPR23962−157] (ab283689) was added to the culture dish for 2 h at room temperature, followed by incubation with a FITC-labeled secondary antibody for 2 h at room temperature. The nanoprobe, a nanopipette with the ability to excite fluorescence and collect fluorescent signals, was slowly moved and positioned on the surface of BMECs with a triaxial microsystem. The FITC-labelled IR on the cell membrane of the BMECs was excited by the excitation light emitted from the nanoprobe, and the fluorescence intensity of individual cells was measured and recorded in real-time by using a single-cell multimodal analyzer (Jiangsu Rayme Biotechnology Co., Ltd.)66.
Lysosomal colocalization analysis
bEnd.3 cells were preseeded in 96-well plates (Cell Carrier Ultra, PerkinElmer, 1 × 104 per well) and cultured at 37 °C for 24 h. The various C6-labeled liposomes were added to the plates and incubated at 37 °C for 2 h, after which the medium was then discarded and replaced with fresh medium containing the LysoTracker at a final concentration of 1 μg/mL, and the cells were incubated for 1.5 h. The cells were then fixed with 4% PFA at room temperature for 0.5 h and stained with Hoechst 33342 at room temperature for 10 min. Finally, lysosomal and liposomal colocalization was imaged with a high-content screening system in confocal mode, and Pearson correlation coefficients were calculated by using GraphPad Prism 8.3.0 software67.
Characterization of the protein corona
Whole blood was collected from healthy BALB/c mice, and the serum was separated via centrifugation. Three hundred microlitres of liposomes were mixed with an equal volume of serum, and the mixtures were incubated at 37 °C for 1 h. Pellets were by centrifugation at 14,000 × g for 40 min and washed twice with cold PBS. The same procedure was performed with blank serum as the control. The collected pellets were resuspended in 60 μL of PBS and 15 μL of 5× SDS-PAGE sample buffer, and the mixtures were boiled for 10 min to denature the proteins. SDS-PAGE was used to separate the proteins, which were stained with a Fast Silver Stain Kit. In addition, adsorbed IgM was semiquantitatively analyzed via western blotting6.
Preparation and characterization of ITP-modified lipid nanoparticles (ITP-LNPs)
ITP-LNPs were prepared by using a microfluidic setup68. Briefly, Dlin-MC3-DMA, DSPC, Chol, DMG-PEG2000, and ITP were dissolved in ethanol at molar ratios of 50:10:34.5:1.5:4. eGFP mRNA was diluted in 10 mM citrate buffer (pH 3) to obtain a mRNA: lipid weight ratio of 1:5. The aqueous and ethanol solutions were mixed at a 3:l volume ratio at a mixing rate of 12 mL/min with a microfluidic apparatus. ITP-LNP were dialyzed against PBS (pH 7.4) by using a dialysis bag with a molecular weight of 8 ~ 14 kDa. Blank lipid nanoparticles (LNP) were prepared by the same procedure but without peptide. The ITP-LNPs were measured for size with a Zeta Sizer Nano Series (ZEN3600, Malvern, UK) and transmission electron microscopy images were obtained with a Hitachi Ht 7800 TEM instrument (Hitachi, Tokyo, Japan).
3D fluorescence imaging of the whole brain
These experiments were done as described elsewhere69. Briefly, mice were injected with ITP-LNP/mRNA or LNP/mRNA via the tail vein, and 24 h later, the mice were anesthetized with an overdose of isoflurane. To clarify brain tissue samples, they were fixed with 4% PFA at 4 °C for 24 h and then treated with Quadrol destaining solution at 37 °C for 2 days. The samples were then soaked in gradient dehydration solution on a shaker at 37 °C for 2 days, followed by treatment with dehydration solution for 2 days and BB-PEG transparent medium for 5–7 days until transparency was achieved. The samples were then stored in clarified media at room temperature. Bright images were obtained with a digital camera. The purified brains were imaged with a LiTonexL thin-section microscope (LiTonexL, Light Innovation Technology, China) with a 4× objective (numerical aperture [NA] 0.28, working distance [WD] 20 mm). The sheets were illuminated from all sides of the sample, and the merged images were saved.
Decolorization solutions
Quadrol was diluted with H2O to a final concentration of 25% v/v. Ammonium was diluted with H2O to a final concentration of 5% v/v.
Gradient tB delipidation solution
Pure tert-butanol (tB) was diluted with distilled water to prepare alcohol solutions with concentrations of 30% v/v, 50% v/v, and 50% v/v. Then, 70% (v/v) Quadrol was added to a final concentration of 3% (v/v). The pH of the solution was adjusted to above 9.5.
tB-PEG dehydration solution
The composition of the dehydration solution was 70% v/v tert-butanol, 27% v/v polyethylene glycol methacrylate Mn 500 (PEGMMA500), and 3% w/v Quadrol.
BB-PEG clearing medium (refractive index R.I. 1.543). BB-PEG was prepared by mixing 75% v/v benzyl benzoate (BB) and 25% v/v PEGMMA500 with 3% w/v Quadrol. The fresh medium was a thin, colorless liquid that turned pale yellow after 1 week.
Statistical analysis
All measurements were taken from distinct samples. Data are presented as means ± SDs, and statistical significance was determined by two-tailed unpaired Student’s t tests with GraphPad Prism software 8.3.0. *p < 0.05, **p < 0.01, and ***p < 0.001 were considered to indicate statistical significance. NS was used to indicate nonsignificance.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data supporting the findings of this study are available within the paper and its Supplementary Information. Source data are provided for Figs. 2b, c, e, g, h, j, 3a, c–f, i, l, 4a–c, g, h, k, m, n, p, 5a–f, i, j, 6c–i, k, l, m, 7e, f, h–j and Supplemental Figs. S1a, b, s2a–f, 3, s5a–h, s6c–d, s8a–c, s9a, b, s11c, s12b, s13a–d, s16b, s17a–c in the associated source data file.
References
Ulbrich, K., Knobloch, T. & Kreuter, J. Targeting the insulin receptor: nanoparticles for drug delivery across the blood-brain barrier (BBB). J. Drug Target. 19, 125–132 (2011).
Scapin, G. et al. Structure of the insulin receptor–insulin complex by single-particle cryo-EM analysis. Nature 556, 122–125 (2018).
Sun L. et al. Shed antigen-induced blocking effect on CAR-T cells targeting Glypican-3 in hepatocellular carcinoma. J. Immunother. Cancer 9, e001875 (2021).
Montagut, C. et al. Identification of a mutation in the extracellular domain of the Epidermal Growth Factor Receptor conferring cetuximab resistance in colorectal cancer. Nat. Med. 18, 221–223 (2012).
Wang, X. et al. Liposomes with cyclic RGD peptide motif triggers acute immune response in mice. J. Control. Release 293, 201–214 (2019).
Guan, J. et al. Enhanced immunocompatibility of ligand-targeted liposomes by attenuating natural IgM absorption. Nat. Commun. 9, 2982 (2018).
Conn, P. J., Lindsley, C. W., Meiler, J. & Niswender, C. M. Opportunities and challenges in the discovery of allosteric modulators of GPCRs for treating CNS disorders. Nat. Rev. Drug Discov. 13, 692–708 (2014).
Song, G. et al. Human GLP-1 receptor transmembrane domain structure in complex with allosteric modulators. Nature 546, 312–315 (2017).
Overington, J. P., Al-Lazikani, B. & Hopkins, A. L. How many drug targets are there? Nat. Rev. Drug Discov. 5, 993–996 (2006).
Mihailescu, M. et al. Structural interactions of a voltage sensor toxin with lipid membranes. Proc. Natl. Acad. Sci. USA 111, E5463–E5470 (2014).
Ying, Z. et al. A safe and potent anti-CD19 CAR T cell therapy. Nat. Med. 25, 947–953 (2019).
Cosson, P., Perrin, J. & Bonifacino, J. S. Anchors aweigh: protein localization and transport mediated by transmembrane domains. Trends Cell Biol. 23, 511–517 (2013).
Bershatsky, Y. V. et al. Diversity of structural, dynamic, and environmental effects explain a distinctive functional role of transmembrane domains in the insulin receptor subfamily. Int. J. Mol. Sci. 24, 3906 (2023).
Lee, J., Miyazaki, M., Romeo, G. R. & Shoelson, S. E. GMS activation with transmembrane domain ligands *. J. Biol. Chem. 289, 19769–19777 (2014).
Albrecht, C. et al. Transmembrane peptides as inhibitors of protein-protein interactions: an efficient strategy to target cancer cells? Front. Oncol. 10, 519 (2020).
Weng, G. et al. HawkDock: a web server to predict and analyze the protein-protein complex based on computational docking and MM/GBSA. Nucleic Acids Res. 47, W322–w330 (2019).
Gohlke, H. & Case, D. A. Converging free energy estimates: MM-PB(GB)SA studies on the protein-protein complex Ras-Raf. J. Comput. Chem. 25, 238–250 (2004).
Wang, Y. et al. Targeting trimeric transmembrane domain 5 of oncogenic latent membrane protein 1 using a computationally designed peptide. Chem. Sci. 10, 7584–7590 (2019).
Liao, Z. et al. Comprehensive insulin receptor phosphorylation dynamics profiled by mass spectrometry. FEBS J. 289, 2657–2671 (2022).
Leto, D. & Saltiel, A. R. Regulation of glucose transport by insulin: traffic control of GLUT4. Nat. Rev. Mol. Cell Biol. 13, 383–396 (2012).
Scholler, P. et al. HTS-compatible FRET-based conformational sensors clarify membrane receptor activation. Nat. Chem. Biol. 13, 372–380 (2017).
Zhou, H. et al. Dense and dynamic polyethylene glycol shells cloak nanoparticles from uptake by liver endothelial cells for long blood circulation. ACS Nano 12, 10130–10141 (2018).
Pillutla, R. C. et al. Peptides identify the critical hotspots involved in the biological activation of the insulin receptor. J. Biol. Chem. 277, 22590–22594 (2002).
Guo, X. et al. Regulation of age-associated insulin resistance by MT1-MMP-mediated cleavage of insulin receptor. Nat. Commun. 13, 3749 (2022).
Ji, Y. et al. The ELAVL3/MYCN positive feedback loop provides a therapeutic target for neuroendocrine prostate cancer. Nat. Commun. 14, 7794 (2023).
Leclerc, M. et al. Cerebrovascular insulin receptors are defective in Alzheimer’s disease. Brain 146, 75–90 (2023).
Xu, Q. et al. Animal models of Alzheimer’s disease: preclinical insights and challenges. Acta Mater. Med. 2, 192–215 (2023).
Liu H.-M. et al. Hypoxia-responsive ionizable liposome delivery siRNA for glioma therapy. Int. J. Nanomedicine. 12, 1065–1083 (2017).
Zhou, Y. et al. Blood-brain barrier–penetrating siRNA nanomedicine for Alzheimer’s disease therapy. Sci. Adv. 6, eabc7031 (2020).
Cheng, Q. et al. Selective organ targeting (SORT) nanoparticles for tissue-specific mRNA delivery and CRISPR–Cas gene editing. Nat. Nanotechnol. 15, 313–320 (2020).
Dammert, P. et al. Treatment of cryptococcal meningitis in Peruvian AIDS Patients using amphotericin B and fluconazole. J. Infect. 57, 260–265 (2008).
Sciacca, L. et al. Biological effects of insulin and its analogs on cancer cells with different insulin family receptor expression. J. Cell. Physiol. 229, 1817–1821 (2014).
van den Boorn, J. G., Schlee, M., Coch, C. & Hartmann, G. SiRNA delivery with exosome nanoparticles. Nat. Biotechnol. 29, 325–326 (2011).
Zhao, L. et al. Exosome-mediated siRNA delivery to suppress postoperative breast cancer metastasis. J. Control. Release 318, 1–15 (2020).
Alvarez-Erviti, L. et al. Delivery of siRNA to the mouse brain by systemic injection of targeted exosomes. Nat. Biotechnol. 29, 341–345 (2011).
Zhu, Z., Liao, L. & Qiao, H. Extracellular vesicle–based drug delivery system boosts phytochemicals’ therapeutic effect for neurodegenerative diseases. Acupunct. Herb. Med. 2, 229–239 (2022).
Yong, T. et al. Tumor exosome-based nanoparticles are efficient drug carriers for chemotherapy. Nat. Commun. 10, 3838 (2019).
Zhong, J. et al. High-quality milk exosomes as oral drug delivery system. Biomaterials 277, 121126 (2021).
Rask-Andersen, M., Almén, M. S. & Schiöth, H. B. Trends in the exploitation of novel drug targets. Nat. Rev. Drug Discov. 10, 579–590 (2011).
Yin, H. et al. Computational design of peptides that target transmembrane helices. Science 315, 1817–1822 (2007).
Jiang, Z., Guan, J., Qian, J. & Zhan, C. Peptide ligand-mediated targeted drug delivery of nanomedicines. Biomater. Sci. 7, 461–471 (2019).
Haro, K. J. et al. Photo-reactive and non-natural amino acid epitopes of human WT1 enhance immunogenicity and allow kinetic study of antigen processing. Blood 110, 2311–2311 (2007).
Yu, Y. J. et al. Boosting brain uptake of a therapeutic antibody by reducing its affinity for a transcytosis target. Sci. Transl. Med. 3, 84ra44–84ra44 (2011).
Liu, Y. et al. Paclitaxel loaded liposomes decorated with a multifunctional tandem peptide for glioma targeting. Biomaterials 35, 4835–4847 (2014).
Mravic, M. et al. De novo-designed transmembrane proteins bind and regulate a cytokine receptor. Nat. Chem. Biol. 20, 751–760 (2024).
Lu, P. et al. Accurate computational design of multipass transmembrane proteins. Science 359, 1042–1046 (2018).
Gurbatri, C. R., Arpaia, N. & Danino, T. Engineering bacteria as interactive cancer therapies. Science 378, 858–864 (2022).
Lahav-Mankovski, N. et al. Decorating bacteria with self-assembled synthetic receptors. Nat. Commun. 11, 1299 (2020).
Li, Q., Wong, Y. L. & Kang, C. Solution structure of the transmembrane domain of the insulin receptor in detergent micelles. Biochim. Biophys. Acta 1838, 1313–1321 (2014).
Pierce, B. G. et al. ZDOCK server: interactive docking prediction of protein-protein complexes and symmetric multimers. Bioinformatics 30, 1771–1773 (2014).
Mijit, A. et al. Mapping synthetic binding proteins epitopes on diverse protein targets by protein structure prediction and protein-protein docking. Comput. Biol. Med. 163, 107183 (2023).
Zhao, L. et al. Identification of anti-TNFα VNAR single domain antibodies from whitespotted bambooshark (Chiloscyllium plagiosum). Mar. Drugs 20, 307 (2022).
Nivón, L. G., Moretti, R. & Baker, D. A Pareto-optimal refinement method for protein design scaffolds. PLoS ONE 8, e59004 (2013).
Huang, P. S. et al. RosettaRemodel: a generalized framework for flexible backbone protein design. PLoS ONE 6, e24109 (2011).
Yang, J. et al. Computational design and modeling of nanobodies toward SARS-CoV-2 receptor binding domain. Chem. Biol. Drug Des. 98, 1–18 (2021).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Yunn, N.-O. et al. Agonistic aptamer to the insulin receptor leads to biased signaling and functional selectivity through allosteric modulation. Nucleic Acids Res. 43, 7688–7701 (2015).
Zou, Y. et al. Single siRNA nanocapsules for effective siRNA brain delivery and glioblastoma treatment. Adv. Mater. 32, 2000416 (2020).
Mori, H. et al. Socs3 deficiency in the brain elevates leptin sensitivity and confers resistance to diet-induced obesity. Nat. Med. 10, 739–743 (2004).
Xue, J. et al. Neutrophil-mediated anticancer drug delivery for suppression of postoperative malignant glioma recurrence. Nat. Nanotechnol. 12, 692–700 (2017).
Rafii, S. et al. Isolation and characterization of human bone marrow microvascular endothelial cells: hematopoietic progenitor cell adhesion. Blood 84, 10–19 (1994).
Navone, S. E. et al. Isolation and expansion of human and mouse brain microvascular endothelial cells. Nat. Protoc. 8, 1680–1693 (2013).
Xie, J. et al. Oriented assembly of cell-mimicking nanoparticles via a molecular affinity strategy for targeted drug delivery. ACS Nano 13, 5268–5277 (2019).
Ghiarone, T. et al. ADAM17 cleaves the insulin receptor ectodomain on endothelial cells and causes vascular insulin resistance. Am. J. Physiol.-Heart Circ. Physiol. 323, H688–H701 (2022).
Wong, H. L. X. et al. MT1-MMP sheds LYVE-1 on lymphatic endothelial cells and suppresses VEGF-C production to inhibit lymphangiogenesis. Nat. Commun. 7, 10824 (2016).
Huang, F. et al. Photoactivated specific mRNA detection in single living cells by coupling “Signal-on” fluorescence and “Signal-off” electrochemical signals. Nano Lett. 18, 5116–5123 (2018).
Li, W. et al. BBB pathophysiology–independent delivery of siRNA in traumatic brain injury. Sci. Adv. 7, eabd6889 (2021).
Rhym, L. H., Manan, R. S., Koller, A., Stephanie, G. & Anderson, D. G. Peptide-encoding mRNA barcodes for the high-throughput in vivo screening of libraries of lipid nanoparticles for mRNA delivery. Nat. Biomed. Eng. 7, 901–910 (2023).
Jing, D. et al. Tissue clearing of both hard and soft tissue organs with the PEGASOS method. Cell Res. 28, 803–818 (2018).
Acknowledgements
The authors would like to thank Christine Wogan, MS, ELS, of MD Anderson Cancer Center’s Division of Radiation Oncology for editorial assistance. This work was supported by the National Natural Science Foundation of China (NSFC No. 82373808, 82073789), and the Chongqing Science Fund for Distinguished Young Scholars (CSTB2023NSCQ-JQX0021) awarded to C.L. The authors wish to acknowledge Dr. Huan Zhao (Revvity) and Xiaogang Wang (Revvity) for their advice on the experimental design and helpful discussions. The authors would also like to thank Rayme Biotechnology and Revvity for equipment support.
Author information
Authors and Affiliations
Contributions
C.L., K.T., Z.T., Z.Y., B.K., and W.J. conceived the project, designed the experiments and analyzed the data. M.N., Z.K., W.X., X.W., X.L., Y.Y., S.J., and A.W. conducted the experiments and analyzed the data. A.W. and Y.M. helped interpret the results. All authors contributed to the writing of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Zhenpeng Qin, Na-Oh Yunn, and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Tang, K., Tang, Z., Niu, M. et al. Allosteric targeted drug delivery for enhanced blood-brain barrier penetration via mimicking transmembrane domain interactions. Nat Commun 16, 3410 (2025). https://doi.org/10.1038/s41467-025-58746-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-58746-x