Abstract
During collagen biosynthesis, lysine residues undergo extensive post-translational modifications through the alternate action of two distinct metal ion-dependent enzyme families (i.e., LH/PLODs and GLT25D/COLGALT), ultimately producing the highly conserved α-(1,2)-glucosyl-β-(1,O)-galactosyl-5-hydroxylysine pattern. Malfunctions in these enzymes are linked to developmental pathologies and extracellular matrix alterations associated to enhanced aggressiveness of solid tumors. Here, we characterized human GLT25D1/COLGALT1, revealing an elongated head-to-head homodimeric assembly. Each monomer encompasses two domains (named GT1 and GT2), both unexpectedly capable of binding metal ion cofactors and UDP-α-galactose donor substrates, resulting in four candidate catalytic sites per dimer. We identify the catalytic site in GT2, featuring an unusual Glu-Asp-Asp motif critical for Mn2+ binding, ruling out direct catalytic roles for the GT1 domain, but showing that in this domain the unexpectedly bound Ca2+ and UDP-α-galactose cofactors are critical for folding stability. Dimerization, albeit not essential for GLT25D1/COLGALT1 activity, provides a critical molecular contact site for multi-enzyme assembly interactions with partner multifunctional LH/PLOD lysyl hydroxylase-glycosyltransferase enzymes.
Similar content being viewed by others
Introduction
Collagens are highly conserved proteins, from invertebrates to higher vertebrates. Their amino acid composition, post-translational modification patterns, and supramolecular assembly confer a versatile spectrum of physical and mechanical properties, resulting in the most abundant protein family in the animal kingdom constituting essential components of several animal tissues including bones, cartilage, tendons, and dermis1,2,3. During biosynthesis, collagens undergo extensive post-translational modifications to fold into right-handed triple helices. The repetitive unit consists of the Gly-Xaa-Yaa sequence motif, where positions Xaa and Yaa are frequently occupied by proline and hydroxyproline residues, respectively4,5,6,7. The role of proline hydroxylation in the global process of collagen biosynthesis is the most studied and understood to date, and it is essential to the winding process of the collagen right-handed triple helix8,9,10.
Lysine residues are less abundant than proline residues in collagen polypeptides, and are typically found at Yaa positions of the Gly-Xaa-Yaa motif11. The role of lysine residues and their post-translational modifications is mainly linked to the cross-linking processes that occur between collagen molecules once secreted into the extracellular space. In collagens and collagen-like protein segments, lysine residues are modified sequentially by specific biosynthetic enzymes which catalyze the formation of a simple but very conserved post-translational modification pattern: the α-(1,2)-glucosyl-β-(1,O)-galactosyl-5-hydroxylysine (Glc-Gal-Hyl)12,13,14,15,16. The path to the formation of Glc-Gal-Hyl starts with the formation of 5-hydroxylysine (Hyl) operated by Fe2+-dependent collagen lysyl hydroxylases (LH, also known as procollagen, lysyl 2-oxoglutarate dioxygenases (PLOD)). The LH/PLOD enzyme family is characterized in humans by three different isoforms (LH1/PLOD1, LH2/PLOD2, and LH3/PLOD3)17,18,19. The LH reaction occurs in the C-terminal dimeric lysyl hydroxylase domain of LH/PLODs. Hyl residues are then glycosylated by Mn2+-dependent galactosyltransferase enzymes, through an inverting reaction that adds the galactose moiety of the uridine diphosphate-α-galactose (UDP-α-Gal) donor substrate to the Hyl hydroxyl group, generating β-(1,O)-5-hydroxylysine (Gal-T activity). Gal-Hyl is further glycosylated by the multifunctional LH/PLOD enzymes through a retaining reaction, that requires Mn2+ and uridine diphosphate-α-glucose (UDP-α-Glc) donor substrate (Glc-T activity), ultimately generating the Glc-Gal-Hyl. Initially, the multifunctional LH/PLOD enzyme(s) were thought to be responsible for the entire Lys–to–Glc-Gal-Hyl biosynthetic pathway20,21. The subsequent identification of a specific family of collagen galactosyltranfserases (named GLT25D or COLGALT), underpinned their essential roles in collagen biosynthesis in vivo22,23,24,25,26,27. Additional biochemical studies on human LH/PLOD and GLT25D/COLGALT enzymes ultimately ruled out the single enzyme hypothesis. The resulting picture shows how the concerted action of two enzyme systems (i.e., LH/PLOD and GLT25D/COLGALT), alternatively acting on the same substrate and possibly directly interacting with each other28, leads to complete Glc-Gal-Hyl collagen post-translational modifications (Fig. 1a).
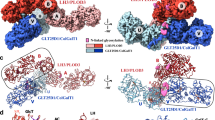
a Reaction scheme showing the collagen Lys–to–Glc-Gal-Hyl conversion post-translational modification process mediated by the alternate action of LH/PLOD and GLT25D1/COLGALT1, with color highlight of donor substrates and associated products. b Cartoon representation of the crystal structure of human GLT25D1/COLGALT1, showing the domain architecture of the enzyme with GT1 and GT2 domains connected via a rigid linker element, anchored to both domains via hydrophobic anchorages shown in details in (c).
While multifunctional lysyl hydroxylase and glucosyltransferase LH/PLOD enzymes have been the subject of extensive molecular and structural characterizations, much less is known regarding the GLT25D/COLGALT enzymes, which are responsible for the central galactosyltransferase reaction. Human GLT25D/COLGALT are soluble proteins anchored to the endoplasmic reticulum through a C-terminal Arg-Asp-Glu-Leu (RDEL) sequence, and have been found to co-localize with LH/PLOD enzymes28,29. Three human GLT25D/COLGALT isoforms have been identified: GLT25D1/COLGALT1, constitutively present in all human tissues, GLT25D2/COLGALT2, expressed mostly in nervous tissue, and GLT25D3/COLGALT3, found in all secretory tissues such as salivary glands, pancreas, placenta, as well as in the nervous system. However, only the first two isoforms are active, whereas isoform 3 (also called CERCAM) is inactive in vitro and potentially not related at all to collagen lysine modification.
Of considerable relevance is the involvement of the colgalt genes and their protein products GLT25D/COLGALT enzymes in numerous pathological conditions, including cerebral small vessel disease, liver pathologies, osteoarthritis, musculoskeletal defects, and cancer26,30,31,32. Liver fibrosis is a chronic pathological condition that sees an accumulation of proteins at the level of the extracellular matrix, whose evolution often leads affected subjects to develop cirrhosis and hepatocellular carcinoma. Recent work has observed high levels of GLT25D1/COLGALT1 in the plasma of cirrhosis patients compared to unaffected patients, and one of the proposed hypotheses is the stimulatory role of the enzyme against hepatic stellate cells33.
Efforts to decipher the structure-function relationships underlying GLT25D/COLGALT activity on collagen molecules have yielded interesting insights, but highlighted the presence of multiple Asp-x-Asp (DxD) motifs along the polypeptide sequence potentially relevant for enzymatic activity, thus demanding more in-depth structure-based investigations.
In this work, we merged computational and experimental structural biology methods with biochemistry and biophysics to reveal the quaternary assembly of GLT25D1/COLGALT1. The X-ray crystal structures of full-length human GLT25D1/COLGALT1 in its substrate-free state as well as in complex with the donor substrate UDP-α-Gal were determined using experimental phasing. Small-angle X-ray scattering, negative stain electron microscopy and mass photometry jointly revealed an elongated quaternary structure reminiscent of that of LH/PLOD enzymes, characterized by a head-to-head dimer with each monomer composed of two similar domains with Rossman fold architecture, both of which support binding of metal ions and the donor substrate. Structure-guided site-directed mutagenesis, native mass spectrometry and molecular dynamics simulations ultimately identified the C-terminal domain as the functional site for enzyme catalysis, while the N-terminal domain retains UDP-α-Gal unusually bound to Ca2+ exclusively for folding stability purposes. Cross-linking mass spectrometry analysis showed that the enzyme’s homodimerization is crucial for interaction with partner LH/PLOD enzymes. Collectively, our results provide insights into the molecular architecture and functions of a key human enzyme of collagen biosynthesis.
Results
GLT25D1/COLGALT1 is a Mn2+ dependent galactosyltransferase
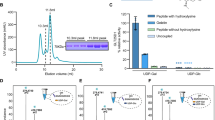
We established methods to recombinantly produce and purify human full-length GLT25D1/COLGALT1 in E. coli and obtained enzyme preparations suitable for evaluation of galactosyltransferase activity in vitro (Supplementary Fig. 1), through direct detection of Gal-T activity on synthetic collagen peptides using high-resolution mass spectrometry (HRMS), as well as by monitoring UDP generation from consumption of the UDP-α-Gal donor substrate in the presence of collagen hydroxylysines. Using luminescence-based detection of UDP as reaction product, we established a biochemical assay to evaluate steady-state enzyme kinetics (Supplementary Fig. 2). The results obtained from steady-state Michaelis-Menten analysis (Supplementary Table 5, Supplementary Fig. 2) were comparable to previously published investigations performed using radiolabeled UDP-α-Gal donor substrate34. Using these assays, we firstly evaluated the enzyme’s strong preference for the Mn2+ metal ion cofactor, highlighting complete absence of enzymatic activity in presence of other divalent metal ions, with the exception of Mg2+, which nevertheless yielded lower enzymatic activity compared to Mn2+ (Supplementary Fig. 3a, b). Given the ability of closely-related multifunctional LH/PLOD enzymes to process multiple donor substrates also in absence of acceptor substrates, we probed GLT25D1/COLGALT1 and found that differently from LH/PLOD, GLT25D1/COLGALT1 is specific for UDP-α-Gal and displays limited processing of UDP-α-Gal in absence of acceptor substrates (i.e., uncoupled activity, Supplementary Fig. 3c).
GLT25D1/COLGALT1 features an elongated multi-domain architecture
The 2.8 Å crystal structure of human wild-type GLT25D1/COLGALT1 was obtained by experimental phasing, exploiting single wavelength anomalous dispersion (SAD) of a Hg2+ derivative (Supplementary Tables 2 and 3). The quality of the electron density was excellent for the globular region of the two monomers of enzyme found in the asymmetric unit (Supplementary Fig. 4), whereas flexible regions located at the N- (residues 30–41) and C-termini (residues 572–618) could not be modeled.
Each GLT25D1/COLGALT1 polypeptide is characterized by two globular domains, which we named GT1 and GT2 (Fig. 1b), both characterized by similar glycosyltransferase architectures, but different topological organization (Supplementary Fig. 5a). In GLT25D1/COLGALT1, the two GT domains contact each other through electrostatics and hydrophobic contacts along helix α7 (GT1) and α8 (GT2). A 25-amino acid linker interconnects the two domains by shielding otherwise surface-exposed hydrophobic patches with two phenylalanine residues (Phe327 and Phe340, respectively, Fig. 1c).
Although both domains are structurally divergent from previously determined Rossman fold topologies, the GT1 domain appears more similar to known glycosyltransferases, yielding Z-scores higher than 15 with few bacterial and human GT-A glycosyltransferases (Supplementary Fig. 5b) according to DALI35, despite very limited sequence identity conservation. The closest structural homolog with an experimentally determined structure is human LH3/PLOD3, whose central accessory (AC) domain shares only 24% sequence identity, but shows very similar structural arrangement with RMSD of 1.2 Å (Supplementary Fig. 5c). This was further confirmed by extensive protein BLAST searches using the GLT25D1/COLGALT GT1 domain sequence as search query, which returned results including vertebrates, invertebrates, and giant viruses (Supplementary Fig. 6). The latter were of particular interest, as previous discoveries reported that Acanthamoeba polyphaga mimivirus is decorated with post-translationally modified collagens with high homology to mammalian orthologs36. Nevertheless, BLAST searches excluding LH/PLOD homologs identified almost exclusively the additional enzyme isoforms GLT25D2/COLGALT2 and CERCAM in vertebrates and GLT25D/COLGALT orthologs in insects (Supplementary Fig. 6). Collectively, these data highlight the conservation of the GT-A fold architecture of the GT1 domain, yielding closely matching folding topologies despite very limited amino acid sequence conservation (Supplementary Fig. 5b).
The GLT25D1/COLGALT1 GT2 domain shares only distant structural homology with known glycosyltransferases, and the resulting superpositions are less clear compared those obtained for GT1 (Supplementary Fig. 5d). Likewise, when superimposed to LH3/PLOD3, the GLT25D1/COLGALT1 GT2 domain shows only limited structural similarity with the N-terminal LH3/PLOD3 GT domain, with superposition RMSD values of 4.7 Å and Z-score lower than 10 according to DALI35 (Supplementary Fig. 5e). When subject to BLAST search, the sequence of this domain yielded more results compared to the GT1 domain, including vertebrates, invertebrates and giant viruses, but also bacteria (Supplementary Fig. 7). The presence of additional kingdoms in the BLAST alignment supports possible evolutionary ancestral gene fusion events ultimately leading to the two-domain GLT25D/COLGALT assembly.
GLT25D1/COLGALT1 is a dimeric enzyme
The asymmetric unit of the GLT25D1/COLGALT1 crystal structure is compatible with two enzyme molecules aligned in an antiparallel fashion through contacts mediated via their GT1 domain and wrapped around the unstructured N-terminal loop comprising residues 42–51 (Fig. 2a). This quaternary structure is characterized by a buried surface area of 1250 Å2, with 22 hydrogen bonds and 5 salt bridges out of 35 residues per monomer involved in the dimeric interface. Such assembly was supported by mass photometry (MP) data, showing that GLT25D1/COLGALT1 in solution is present with a molecular weight of 140 kDa, equivalent to the expected molecular weight for enzyme dimers (Fig. 2b), with minimal dispersion. Consistent results of 131 ± 10 kDa were obtained using size exclusion chromatography coupled to multi-angle light scattering (SEC-MALS) and small-angle X-ray scattering (SEC-SAXS) (Supplementary Table 6, Supplementary Fig. 8a, b), the latter producing ab initio sphere models of elongated molecular ensembles matching the size and overall envelope shape of crystallographic GLT25D1/COLGALT1 dimers with 95% correlation (Supplementary Fig. 8c). Orthogonal comparison of the unmodified experimental crystal structure with solution SAXS data yielded initial χ2 of 7.62, which further improved to 1.89 after modeling of flexible N- and C-termini (Fig. 2c). Furthermore, low-resolution 2D classes from a sample of GLT25D1/COLGALT1 analyzed using single particle negative staining electron microscopy revealed elongated objects highly comparable to the SEC-SAXS solution data and the crystallographic dimers (Fig. 2d).
a Structure of dimeric full-length GLT25D1/COLGALT1 enzyme, showing two molecules interconnected through their N-terminal domains. b Mass photometry analysis of recombinant wild-type GLT25D1/COLGALT1 samples. c Comparison of the GLT25D1/COLGALT1 dimer observed in the crystal structure with solution SAXS data (red circles) yields a χ2 of 7.62 (dashed line). Modeling of the N- and C-terminal flexible loops (shown as blue and spheres, respectively) yields a structural model of the dimeric enzyme (represented as a white cartoon) with improved fitting of the experimental solution SAXS data, resulting in χ2 of 1.89 (solid line). d Comparison of negative stain electron microscopy single particle 2D classes (gray background) with selected orientations of computed projections of the GLT25D1/COLGALT1 crystal structure (black background) shows matching molecular projections.
Probing the role of GLT25D1/COLGALT1 dimerization
To evaluate the importance of dimerization on GLT25D1/COLGALT1 function, we searched for amino acid residues located at the crystallographic dimer interface: Trp158, whose side chain is embedded into a cavity shaped by Arg53, Ala85, His113, Met157 of the other enzyme monomer; and Asp160, forming a hydrogen bonding network with Arg53 from both monomers (Fig. 3a). Mutation Trp158Arg was enough to disrupt the dimer interface of GLT25D1, resulting in monomeric species in solution as assessed by mass photometry and SEC-MALS (Fig. 3b, c). When prompted for enzymatic activity using either indirect luminescence-based detection of free UDP in the presence of gelatin, as well as direct evaluation of galactosyl-hydroxylysine modification of synthetic peptides using high-resolution mass spectrometry (HRMS), this mutant showed activities comparable to wild-type GLT25D1/COLGALT1 (Fig. 3d, Supplementary Table 5), supporting the hypothesis that enzyme dimerization does not constitute an essential requirement for the enzyme’s catalytic activity in vitro.
a Details of the dimeric interface, showing amino acid residues involved in the direct contacts between the two monomers. The bottom panel shows an open-book rendering of the two GLT25D1/COLGALT1 monomers, with the colored footprint of the dimerization partner on the molecular surface. b Results of the mass photometry comparison of wild-type GLT25D1/COLGALT1 and the Trp158Arg mutant, designed to disrupt the dimer interface. c Results of the SEC-MALS comparison of wild-type GLT25D1/COLGALT1 and the Trp158Arg mutant, showing alterations of the elution volume as well as molar mass. d Comparison of the Gal-T enzymatic activity of GLT25D1/COLGALT1 wild-type and Trp158Arg mutant using luminescence. The Michaelis-Menten curves show the steady-state kinetics characterization, described as apparent turnover number (v0/[enzyme]) as a function of gelatin substrate concentration. Quantitation of KM and kcat values for both wild-type and Trp158Arg mutant is provided in Supplementary Table 5. Error bars represent standard deviations from average of triplicate independent experiments. e Abundance ratios of wild-type and Trp158Arg GLT25D1/COLGALT1 GLT25D1Lys145-LH3Lys645 inter-protein cross-links (orange), GLT25D1Lys145 and LH3Lys645 mono-links (cyan), and total GLT25D1/COLGALT1 and LH3/PLOD3 peptides (green) detected in the XL-MS/MS experiments. The relative abundance of LH3/PLOD3 and GLT25D1/COLGALT1 proteins was calculated from the 10 most abundant peptides per protein. Data are reported as mean. Individual peptide and cross-link abundance ratios are shown as dots. (Mean ± SD: LH3/PLOD3 1.07 ± 0.11, GLT25D1/COLGALT1 1.02 ± 0.12); median values are 1.02 for LH3/PLOD3 and 1.00 for GLT25D1/COLGALT1, with interquartile ranges of 1.01–1.12 and 0.95–1.09, respectively. The minimum and maximum values observed were 0.96–1.30 for LH3/PLOD3 and 0.83–1.27 for GLT25D1/COLGALT1.
Intrigued by this observation, we used cross-linking mass spectrometry to explore whether GLT25D1/COLGALT1 dimerization could contribute to molecular interactions with partner multifunctional lysyl hydroxylase-glucosyltransferase LH3/PLOD3. We cross-linked a 1:1 stoichiometric mixture of wild-type and Trp158Arg GLT25D1/COLGALT1 with disuccinimidyl dibutyric urea (DSBU), followed by in-solution digestion with trypsin. The resulting peptide mixtures were analyzed using nano-liquid chromatography-tandem mass spectrometry (LC-MS/MS), revealing in both cross-linked samples a clear inter-protein cross-link between Lys145 of GLT25D1/COLGALT1 (GLT25D1Lys145) and Lys645 of LH3/PLOD3 (LH3Lys645) (Supplementary Fig. 9). However, the abundance of this GLT25D1Lys145-LH3Lys645 inter-protein cross-link was found to be approximately seven times higher using wild-type GLT25D1/COLGALT1 complex compared to the Trp158Arg mutant (Fig. 3e). Notably, this difference was not attributable to changes in protein concentration or lysine reactivity, as the overall abundance of both proteins, and the individual reactivity of GLT25D1Lys145 and LH3Lys645remained constant in both experiments (Fig. 3e). A nearly identical abundance of the GLT25D1Lys145 and LH3Lys645 mono-links, which occur when a lysine residue is covalently modified by DSBU without linkage to another residue, were measured in the two experiments (Fig. 3e). Collectively, these results show that GLT25D1/COLGALT1 dimerization critically contributes to macromolecular complex formation with multifunctional LH3/PLOD3 enzymes.
Both GT1 and GT2 domains bind donor substrates and metal ions
Analysis of the experimental electron density of GLT25D1/COLGALT1 revealed intense difference peaks in the GT1 domain of both enzyme monomers proximate to the DxD motif identified by residues Asp166 and Asp168, in a loop interconnecting strand β4 and helix α4, that could be modeled as UDP-α-Gal bound to a metal ion (Fig. 4a). The carboxylate group of Asp166 is not directly involved in the coordination of the metal ion. The donor substrate is surrounded by an extended amino acid network, characterized by a H-bonded interaction involving the side chains of Tyr126 with the C4 carbonyl of the uridine moiety, which is trapped by a pi-Sigma interaction with Arg61 and a pi-Alkyl interaction with Val143, in a hydrophobic cavity also shaped by residues Leu59, Asp91, Arg139. The ribose moiety is kept into a well-defined conformation by Trp135, and by hydrogen bonds with the main chain carbonyl and amino groups of residues Leu59, Arg61, and Ala167. Additional interactions involve the side chains of Arg147 and Asp265 with hydroxyl groups of the galactose moiety (Fig. 4a, b). Notably, Trp135 matches Trp130 in the GLT25D1/COLGALT1 mouse protein sequence, whose mutation (named fosse) has been associated to skeletal and muscular defects due to severely reduced expression of the mutated gene22. When co-crystallized with Mn2+ and UDP-α-Gal, the crystal structure of GLT25D1/COLGALT1 unexpectedly revealed electron density matching cofactors and donor substrates also in the GT2 domain of one of the two monomers in the asymmetric unit (Fig. 4c). In this domain the metal ion cofactor was found bound to a Glu-Asp-Asp motif adopting a conformation topologically equivalent to that found in Mn2+-dependent glycosyltransferases, with the carboxyl groups of Glu435 and Asp437 directly chelating the metal ion, while the side chain of Asp436 was interacting through H-bonded interactions with the hydroxyl groups of the ribose moiety of the cofactor (Fig. 4c, d). The uridine ring of the donor substrate was constrained by Gly408 and Gly411 of helix α10, in a hydrophobic groove defined by Cys412 of the same helix, plus Leu348 and Ala374 on the opposite side (Fig. 4c, d). No electron density could be observed for the galactose moiety, suggesting multiple conformations for the sugar possibly due to the absence of acceptor substrates in the wide crevice defined by residues Trp495, Leu497, Tyr567, and Pro558 (Fig. 4c, d).
a The GT1 domain is constitutively populated by a divalent metal ion (labeled as M2+, shown with a purple sphere) and UDP-α-Gal, as shown in the Fo–Fc omit electron density map (contour level 3.2 σ). Residues involved in coordination of the metal ion and interaction with the donor substrate are shown as sticks. b LIGPLOT+72 diagram illustrating the interaction network for donor substrates and cofactors observed in the GT1 domain. c When co-crystallized with excess Mn2+ and UDP-α-Gal, also the electron density of the GT2 domain (Fo–Fc omit map, contour level 3.2 σ) shows presence of donor substrates and cofactors. In this domain, the galactose moiety of the donor substrate is not visible, hence only UDP has been modeled (shown as sticks). Residues involved in coordination of the metal ion and interaction with the donor substrate are shown as sticks. d LIGPLOT+72 diagram illustrating the interaction network for donor substrates and cofactors observed in the GT2 domain. e Binding of Mn2+ and UDP-α-Gal induces conformational changes in the GT2 domain, by stabilizing the otherwise flexible C-terminal loop through direct interactions with the cofactor and the donor substrate. Shown is the superposition of GLT25D1 GT2 domains in substrate-free (light orange cartoon) versus substrate-bound (light blue cartoon) states, with highlight of the loop comprising residues 560–574 (brown), stabilized only when Mn2+ and UDP-α-Gal are present.
Comparison of GT2 domains with and without donor substrates highlighted a conformational rearrangement of the otherwise flexible C-terminus of the domain for residues 562–580, wrapping around the donor substrate binding cavity and positioning the backbone carbonyl of residue Ser569 and Thr571 in a productive conformation for hydrogen bonding with both phosphates of the UDP donor substrate (Fig. 4e). The presence of donor substrates and cofactors only in one of the two crystallographic monomers could be explained with the different crystal packing network surrounding the GLT25D1/COLGALT1 molecules, which prevents the rearrangement of the GT2 C-terminal loop in one of the two molecules due to crystal contacts (Supplementary Fig. 10).
Mutations in the GT1 domain impact on enzyme folding stability
Structure-guided point mutants designed to interfere with metal ion or UDP-α-Gal binding in the GT1 domain (Fig. 4a, b) such as Asp166Ala (abolishing interaction with the bound metal ion), Asp265Ala (removing H-bonding contacts with the galactose moiety), Ile266Gln (preventing rigid positioning of the galactose moiety in the GT1 cavity), did not produce any recombinant enzyme samples suitable for biochemical studies, and were consistent with previously published attempts describing a Asp166Ala-Asp168Ala double mutant, designed to abolish the GT1 DxD motif34. We also attempted production of a Trp135Arg mutant, matching the homologous Trp130Arg fosse mutation in mouse22. The recombinant production yields for this mutant were significantly lower compared to wild-type GLT25D1/COLGALT1, consistent with the observed reduced expression levels reported for the fosse mutation22. SDS-PAGE analysis showed significant differences compared to wild-type GLT25D1/COLGALT1, suggesting severe folding stability issues (Supplementary Fig. 1). Control mutants such as Ile267Gln (in close proximity to Ile266, but not affecting UDP-α-Gal binding or positioning) instead resulted in well folded and active enzyme samples (Supplementary Fig. 1), matching the previously published Pro292Asn mutant, originally designed to generate a chimeric GLT25D1-CERCAM inactive galactosyltransferase, but still retaining enzymatic activity34.
The GT2 domain is responsible for the catalytic activity
Our experimental structure investigations revealed that the GT2 domain of GLT25D1/COLGALT1 binds metal ions and donor substrates transiently compared to GT1, inducing conformational changes of the extended C-terminus of the enzyme that could have an impact on catalysis (Fig. 4). The identification of an unusual EDD motif rather than the canonical DxD motif coordinating the metal ion cofactor within this domain, together with the presence of a UDP-α-Gal donor substrate with a flexible sugar moiety (Fig. 4c, d), may ultimately provide insights for the identification of the enzyme’s catalytic site. Conservation maps for the amino acid residues shaping the two putative ligand binding sites in homologous vertebrate enzymes highlighted that the amino acids involved in metal ion and UDP-α-Gal binding are highly conserved in both GT1 and GT2 domains (Fig. 5a). Prompted by this observation, and by our insights suggesting structural roles for the enzyme’s N-terminal GT1 domain, we generated a series of point mutations affecting metal ion and donor substrate binding in the GT2 domain, as well as residues potentially involved in acceptor substrate binding during transfer of the galactose moiety. All mutants generated could be purified and showed biophysical features compatible with well-folded proteins (Supplementary Fig. 1). Mutants Glu435Ala and Asp437Ala, within the EDD sequence responsible for metal ion coordination, were found completely inactive in both luminescence and HRMS assays (Fig. 5b, Supplementary Fig. 3, Supplementary Table 5). Similar results could be obtained with mutant Cys412Ser, affecting productive positioning of the uridine-ribose moieties in a hydrophobic cavity (Fig. 5b, Supplementary Fig. 3, Supplementary Table 5). Inspired by the potential roles found for aromatic and charged residues located at precise positions in the catalytic cavity of inverting glycosyltransferase enzymes37,38, we also generated mutants Trp495Ala and Asp522Ala, which both resulted in complete loss of GLT25D1/COLGALT1 Gal-T activity (Fig. 5b, Supplementary Fig. 3, Supplementary Table 5).
a Amino acid conservation diagram for different GLT25D/COLGALT homologs. The colors represent the different degree of amino acid conservation from dark blue (not conserved at all) to dark purple (fully conserved). The complete list used for the representation and the associated alignment is shown in Supplementary Figs. 6 and 7. b Luminescence-based comparison of the Gal-T enzymatic activity of wild-type GLT25D1/COLGALT1 and mutants designed to interfere with cofactor and substrate binding in the GT2 domain. The histograms show the average result of triplicate independent measurements (whose results are individually shown as overlayed dots). Error bars represent standard deviations from average of four independent experiments; Statistical evaluations based on pair sample comparisons between control and assay values using Student’s t-test. ****P-value < 0.0001. c A salt bridge involving Arg351 and Asp570 from the flexible C-terminal loop (blue) shapes the GLT25D1/COLGALT1 GT2 catalytic site when enzyme cofactors are bound. Mn2+ is shown as a purple sphere coordinated to Glu437. UDP as well as amino acid side chains directly involved in the described interactions are shown as sticks. d Mapping of the sites affected by known pathogenic mutations on the GLT25D1/COLGALT1 structure. Residues found mutated in the homologous fosse mouse phenotype (Trp135) and in cerebral small vessel disease (Leu151Arg, Ala154Pro, Gly377Arg) are shown in red and highlighted by arrows.
The conformational rearrangement of the GT2 C-terminus suggested a potential role also for Arg351, rearranging its side chain to form a hydrogen bond with Asp570 when cofactors and donor substrates are bound (Fig. 5c). We therefore generated mutants Arg351Ala and Asp570Ala. While production of mutant Arg351Ala resulted in poor quality enzyme preparations and was not suitable for subsequent biochemical studies, production of mutant GLT25D1/COLGALT1 mutant Asp570Ala was successful (Supplementary Fig. 1). Consistent with our hypothesis, this mutant was inactive (Fig. 5b, Supplementary Table 5).
The structural data obtained also provided a rationale for previously investigated mutations: Asp336, located at the C-terminus of the linker segment connecting GT1 and GT2 domains, whose hydrogen bond with GT2 Lys445 does not seem to be dramatically impacted when being mutated into a serine residue; and residues Asp461 and Asp433, the latter being involved in a hydrogen bonding network with the side chains of Lys508 and His547, likely critical for GT2 domain folding stability. Taken together, our molecular structures and site-directed mutagenesis results enabled the unambiguous identification of GLT25D1/COLGALT1 catalytic site and its unusual features.
Molecular significance of GLT25D1/COLGALT1 pathogenic mutations
The molecular structure and the biochemical characterizations of several GLT25D1/COLGALT1 mutants enable correlating the possible impact of disease-causing mutations associated with this enzyme. The fosse mutation Trp130Arg, associated to a mouse phenotype with developmental defects, localizes on Trp135 of the human ortholog. Trp135 shapes the UDP-α-Gal binding cavity in the GT1 domain of the enzyme by interlocking its side chain between the uridine and the galactose moieties of the cofactor (Fig. 4a). We evaluated the impact of this mutation experimentally and observed significantly reduced expression levels for the recombinant enzyme variant in vitro, as well as stability issues that hampered subsequent biochemical characterizations (Supplementary Fig. 1a). Mapping of pathogenic mutations associated to cerebral small vessel disease39,40 on the GLT25D1/COLGALT1 structure provided a rationale for the significance of these pathogenic enzyme variants (Fig. 5d). Mutations Leu151Arg and Ala154Pro localize at the C-terminus of the α3 helix in the GT1 domain. The hydrophobic side chains of Leu151 is buried into a hydrophobic pocket, therefore the introduction of a bulky Arg side chain at this position would likely induce severe perturbations to the stability of the GT1 domain. Likewise, replacing Ala154 with a Pro residue would likely impact on the structural stability of helix α3 near the homo-dimer interface. The possible effect of the Gly377Arg mutation39 could be inferred by Gly377 shaping an alpha turn segment connecting the strand β14 with helix α9 in close proximity to the binding site for the uridine ring of the catalytic UDP-α-Gal donor substrate. Introduction of an Arg side chain at this position may severely perturb GT2 domain conformational stability, resulting in loss of function.
Mn2+ binds transiently in the GT2 domain during catalysis
Intrigued by the observation of a dimeric ensemble potentially characterized by four aligned catalytic sites, we wondered whether metal ions and donor substrates could possibly have structural, rather than functional roles. We therefore purified recombinant GLT25D1/COLGALT1 without any cofactor supplementation and attempted to characterize the possible bound metal ions. DSF experiments performed by incubating the enzyme with metal ion solutions showed significant stabilization of GLT25D1/COLGALT1 only in the presence of Mn2+ (Fig. 6a), supporting the biochemical data indicating Mn2+ as the metal ion cofactor transiently bound during catalysis. Comparison with DSF data collected on mutants Glu435Ala and Asp437Ala incubated with Mn2+ did not result in stabilization (Fig. 6a), consistent with the structural data showing transient binding of this catalytic metal ion in the GT2 domain (Fig. 4).
a DSF analysis of wild-type GLT25D1 (left) incubated with Mn2+ (purple trace) shows significant stabilization induced by binding of the metal ion when compared to untreated samples (blue trace); removal of amino acid side chains critical for binding of metal ions in the GT2 domain such as Glu435 (center) and Asp437 (right) results in loss of the metal ion-induced stabilization effect. b DSF analysis of GLT25D1/COLGALT1 subject to EDTA (green trace) and EGTA (orange trace) treatment, showing the destabilization induced by metal ion chelation when compared to untreated samples (blue trace); Comparison between wild type (left) and mutants in the GT2 domain highlight very similar destabilization effects. c EDTA-treated samples show alterations in dispersity when tested using mass photometry. d EDTA-treated samples show altered elution profiles when tested using size-exclusion chromatography. e Native-MS spectra under partially-denaturing conditions of wild-type 30+ charge state (left) and Trp158Arg 31+ charge state (center), with the signals corresponding to the free protein and protein-ligand complex labeled; box plot (bounds: percentile 25–75%; mean value: black line; whiskers: maxima-minima) of ligand mass calculation obtained from the entire charge state distribution (n = 6 for wild-type, n = 7 for Trp158Arg) of Native-MS spectra (right); reference lines report the theoretical ligand mass as a function of distinct divalent cation cofactors.
Roles of Ca2+ and UDP-α-Gal donor substrate in the GT1 domain
Parallel to Mn2+ binding experiments, we tested the effect of removal of the tightly bound metal ion in the GT1 domain by incubating GLT25D1/COLGALT1 with different concentrations of ethylenediaminetetraacetic Acid (EDTA). We consistently observed destabilization, with a differential scanning fluorimetry (DSF) thermal shift of 3.0–3.5 °C compared to untreated samples (Fig. 6b). We observed nearly identical destabilization effects by treating GLT25D1/COLGALT1 mutants Glu435Ala and Asp437Ala with EDTA (Fig. 6b). As these mutants were designed to interfere with transiently bound metal ions in the GT2 domain, we concluded that EDTA treatment impacts on protein stability by stripping the metal ion bound the GT1 domain. Nevertheless, EDTA-treated samples still showed DSF profiles compatible with a partially folded protein sample, thus we wondered whether removal of metal ions through EDTA treatment could impact on the enzyme’s conformational stability. EDTA-treated wild-type GLT25D1/COLGALT1 samples were therefore subject to mass photometry analysis and size exclusion chromatography. While mass photometry did not show significant alterations in the oligomeric state of the enzyme upon EDTA treatment (Fig. 6c), stripping of the metal ion altered the elution profile of the enzyme, compatible with alterations of the enzyme’s hydrodynamic radius (Fig. 6d). Surprisingly, ICP-MS experiments aimed at identifying Mn bound to enzyme in the GT1 domain did not reveal the presence of this transition metal ion. We therefore decided to further investigate the ligands consistently bound in the GLT25D1/COLGALT1 GT1 domain. Native mass spectrometry (Native-MS) analysis of recombinant wild-type and Trp158Arg GLT25D1/COLGALT1 samples under controlled partially unfolding conditions revealed the coexistence of free- and ligand bound-monomers, leading to the identification of Ca2+ as the most probable bound metal ion in the GT1 domain (Fig. 6e); ICP-MS confirmed the identification, indicating a 1:1 Ca2+:GLT25D1/COLGALT1 stoichiometry. Prompted by this unexpected finding, we performed DSF analysis of GLT25D1/COLGALT1 samples in the presence of the highly selective Ca2+ chelating agent ethyleneglycol-bis(β-aminoethyl)-N,N,Nʹ,Nʹ-tetraacetic acid (EGTA), obtaining a folding destabilization profile matching that observed using EDTA (Fig. 6b).
Taken together, these data highlight the presence of Ca2+ bound in the GLT25D1/COLGALT1 GT1 domain, supporting structural roles for this metal ion cofactor and the UDP-α-Gal donor substrate in this domain. The absence of direct functional roles during catalysis for the GT1 domain is consistent with previous hypotheses derived from chimeric studies of GLT25D1/COLGALT1 and the CERCAM inactive paralog34, and is further supported by the absence of the essential negatively charged amino acid side chains near the galactose moiety of the UDP-α-Gal donor substrate (Fig. 5a, b) required for the enzymatic activity of inverting GT-A glycosyltransferases37,38.
MD simulations support distinct roles for GT1 and GT2 domains
To better characterize the molecular features observed in GLT25D1/COLGALT1 GT1 and GT2 domains, we carried out a comparative molecular dynamics study of the individual domains in the presence or in absence of metal ions and donor substrates. Since the focus of this study is to investigate how UDP-Gal ligand and metal ion cofactor binding influence the structural stabilization of these domains rather than to observe conformational transitions, conventional MD simulations are sufficient to capture the relevant dynamics. To this end, three independent 1-microsecond all-atom MD simulations per system in explicit solvent were perfomed, yielding sufficient accuracy to describe atomic interactions and solvation effects on nanosecond to sub-microsecond timescales, typical of domain stabilization and conformational flexibility. To assess the impact of cofactors binding on GT1 and GT2 stability and dynamics, analyses focused on conformational ensembles rather than time-dependent trajectories, to ensure a statistically robust characterization of the different simulated states. Equilibration and independence from the initial configurations were evaluated by analyzing the root mean square deviations of the protein backbone in each independent simulation replicate separately. After verifying the convergence, metatrajectories were built by concatenating the equilibrated part of the three replicates, on which ensemble-based analyses were performed. Further details on equilibration and the assessment of initial configurations independence are provided in the Methods section and in the Supplementary Methods. Simulations indicated that no large conformational changes occur in the GT1 and GT2 domains upon ligand binding. However, analyses of internal dynamics reveal that the binding of donor substrates and metal ions in the GT1 domain significantly stabilizes its structure, while a less pronounced effect is observed for the GT2 domain. Principal component analysis (PCA) was used to identify the dominant modes of motion in simulations by capturing directions of the largest variance in the data. More specifically, for each domain, GT1 and GT2, the bound and unbound trajectories were combined and subjected to PCA. The first two principal components (PC1 and PC2), representing the majority of conformational variance were calculated, and snapshots of the bound and unbound trajectories were projected onto these components. Histograms of normalized projection frequencies provided a visual representation of the conformational space explored by each system, highlighting their dynamic differences (Supplementary Fig. 12a–d). For the GT1 domain, projections of bound and unbound conformations onto PC1 exhibit overlapping distributions, indicating access to similar conformational spaces. However, the unbound state displays broader and less pronounced peaks, reflecting greater conformational diversity and flexibility. In contrast, the bound state shows sharper and higher peaks, suggesting that cofactor binding restricts the GT1 domain to a narrower set of conformations (Supplementary Fig. 12a). Similarly, along PC2, the bound state maintains a more confined distribution compared to the broader, multi-peaked profile of the unbound state (Supplementary Fig. 12b). For the GT2 domain, projections of bound and unbound conformations along PC1 demonstrate that the unbound state predominantly samples a single conformational state, indicated by a single peak centered at zero, reflecting minimal movement and inherent structural stability. In contrast, the bound state exhibits a bimodal distribution with one peak overlapping the unbound state and a second, sharper peak, suggesting that cofactors binding induces an additional conformation (Supplementary Fig. 12c). Along PC2, the unbound state shows a narrow distribution centered around zero, whereas the bound state shows a slight shift toward negative values, indicating subtle conformational adjustments upon cofactors binding (Supplementary Fig. 12d). These results suggest that binding of metal ions and donor substrates contribute to the overall GT1 domain stabilization by limiting its structural dynamics. In contrast, the GT2 domain remains structurally stable also in the unbound state, while cofactor binding induces an additional conformational state, resulting in minor alterations to its internal dynamics without causing large-scale structural changes.
Hydrogen bond persistence analysis along the simulation time corroborates the PCA results. In the GT1 domain, the bound state exhibits more persistent hydrogen bonds (60–90% of the simulation time) compared to the unbound state (<60%) (Supplementary Fig. 12e, f), indicating greater rigidity and stability. Conversely, the GT2 domain shows minimal differences in hydrogen bond persistence between bound and unbound states, maintaining a dynamic hydrogen bond network with only slight variations (Supplementary Fig. 12g, h). This result suggests that cofactor binding has limited impact on the structural dynamics of the GT2 domain.
Distance fluctuation (DF) analysis, which characterizes the coordinated internal dynamics of the GT1 and GT2 domains, further supports these findings. In the GT1 domain unbound state, the loop hosting residues 130–140, the C-terminal portion of helix a7 of the GT1 domain and the linker between GT1 and GT2 domains are highly dynamic and less restricted to a well-defined conformation, losing their coordination with the rest of the domain, (Supplementary Fig. 13a). Ligand binding enhances internal coordination of these regions, restricting their dynamics to more stable conformations. In contrast, the GT2 domain shows minimal differences in DF between bound and unbound states, maintaining similar dynamic coordination patterns with only slight variations in flexible regions (Supplementary Fig. 13b). These observations suggest that cofactor binding increases the internal coordination of the GT1 domain while exerting a less pronounced effect on the GT2 domain. In particular, without the stabilizing effect induced by donor substrate and metal ion, the loop hosting residues 130–140, the C-terminal portion of helix a7 of the GT1 domain and the linker between GT1 and GT2 domains become highly dynamic and less restricted to a well-defined conformation (Supplementary Fig. 13c). GT1 and GT2 domains respond differently to ligand and cofactor binding within the simulated timescales, confirming that the relevant structural stabilization processes are well-characterized by the simulations performed.
Overall results from MD simulations and analysis support the hypothesis that cofactor binding significantly enhances the structural stability of the GT1 domain while exerting a less pronounced effect on the structural dynamics of the GT2 domain.
Discussion
Collagen hydroxylysine galactosylation is a fundamental step in the complex post-translational modification pathway that leads to Glc-Gal-Hyl, the most abundant O-linked glycosylation found in the animal kingdom12,13,16. Following decades of intensive biochemical investigations addressing the enzymes involved in this process and their associated functions, recent years have witnessed increasing efforts to produce a comprehensive molecular description of these reactions and the actors involved, with particular attention to multifunctional LH/PLOD lysyl hydroxylases-glucosyltransferases17,41. However, the lack of experimental structure data about GLT25D/COLGALT galactosyltransferases has limited our understanding of the molecular mechanisms of these glycosyltransferases, despite previous significant efforts16,28,34. In this work, we have determined the experimental structure of full-length human GLT25D1/COLGALT1, and used a combination of biochemistry, biophysics and site-directed mutagenesis to characterize its functionality in vitro.
Our results reveal an elongated, multi-domain dimeric quaternary structure characterized by two Rossmann fold-type domains per monomer (named GT1 and GT2, respectively) interconnected through a rigid linker region, and with dimer contacts wrapping around the N-terminus of each monomer (Fig. 1b, c). Surprisingly, both GT1 and GT2 domains were found indispensable for catalysis and were capable of binding metal ions and UDP-α-Gal donor substrate (Fig. 4), with the ligand binding regions of both domains showing the highest degree of sequence conservation (Supplementary Figs. 6 and 7). The N-terminal domain GT1 is closely related to the inactive central accessory (AC) domain of LH/PLOD enzymes and shares features more similar those typical of inverting GT-A glycosyltransferases, whereas the GT2 domain features a distinctive topology not matched by well-characterized glycosyltransferase enzymes.
Previous work addressing the putative functional roles of GLT25D1/COLGALT1 DxD motifs showed that point mutations affecting residues involved in metal ion coordination Asp166 and Asp168 in the GT1 domain completely abolished the activity34. The same work also indicated that replacing the entire N-terminus of the enzyme with that of inactive CERCAM did not impact on the ability of the enzyme to glycosylate collagen substrates34. These results are consistent with the experimental identification of Ca2+ and UDP-α-Gal bound in the GT1 domain of the enzyme (Fig. 6b–e). Contrary to the frequent presence of Mn2+ and Mg2+ metal ion cofactors37,38, we could not find evidence of other Rossman fold GT-A glycosyltransferases with bound Ca2+ in their catalytic site. Combined with the absence of negatively charged amino acid residues at key positions essential for inverting glycosyltransferase catalysis37,38, the presence of a stably bound Ca2+ ion interacting with UDP-α-Gal rules out direct catalytic roles for the GLT25D1/COLGALT1 GT1 domain. In addition, MD simulations performed on ligand-free and ligand-bound GLT25D1/COLGALT1 GT1 domain further supported the structural roles for the bound metal ion and donor substrate, highlighting increased molecular flexibility around residues 130–140 and 310–330 of this domain in absence of metal ions and UDP-α-Gal (Supplementary Figs. 12 and 13). Notably, these regions encompass residues proximate to UDP-α-Gal binding such as Trp135, matching the site affected by the loss-of-function mouse fosse mutation22 and the main contact site of the GT1 domain with neighbor GT2, where mutations associated to cerebral small vessel disease can be mapped (Fig. 5d). Increased molecular flexibility in this portion of the GLT25D1/COLGALT1 polypeptide may significantly perturb the globular organization of this multi-domain enzyme. Such hypothesis is supported by our data showing that after EDTA/EGTA treatment, GLT25D1/COLGALT1 is characterized by folding instability and conformational alterations in solution without significant changes in the enzyme’s oligomeric state (Fig. 6b–d).
The GT2 domain displays amino acid sequence conservation for the residues surrounding the site transiently occupied by the Mn2+ cofactor and UDP-α-Gal during catalysis (Fig. 5a, Supplementary Fig. 7). Structural investigations proved essential to identify the GLT25D1/COLGALT1 catalytic site. Previous attempts to map the catalytic residues critical for GLT25D1/COLGALT1 enzymatic activity were not conclusive, likely due to the unusual EDD motif that we found responsible for binding Mn2+ in the GT2 domain, and the multiple DxD motifs within the enzyme polypeptide sequence (Supplementary Figs. 6 and 7). To ensure correct mapping of the GT2 catalytic network, we designed several point mutations surrounding the Mn2+ cofactor and the donor substrate observed in the crystal structures and found an extensive network of charged and hydrophobic side chains critically involved in UDP-α-Gal substrate recognition, activation, and catalysis (Fig. 5b, c).
Overall, substrate-free and substrate-bound structures for the GT2 catalytic cavity are almost identical (Fig. 4), except for the conformational rearrangement of the otherwise flexible C-terminal segment of the domain. This region is characterized by negatively charged amino acids, whose direct involvement in binding and processing of the donor substrate UDP-α-Gal has been demonstrated by our site-directed mutagenesis experiments. In this respect, our insights also support previous in silico hypotheses42.
The observed dimeric quaternary structure, confirmed by multiple orthogonal experiments in solution and by site-directed mutagenesis, does not appear to be a requirement for GLT25D1/COLGALT1 enzymatic activity in vitro (Fig. 3). Using cross-linking mass spectrometry we found that the dimeric architecture of GLT25D1/COLGALT1 shapes a direct molecular contact site for GLT25D1/COLGALT1-LH3/PLOD3 molecular interactions, providing experimental evidence to previous hypotheses regarding formation of multimeric biosynthetic hetero-complexes capable of fully converting collagen Lys residues into Glc-Gal-Hyl29,31. These results are consistent with the recently reported cryo-electron microscopy structure of a GLT25D1/COLGALT1-LH3/PLOD3 complex43. Notably, our biophysical data consistently indicate that a small fraction of our recombinant enzyme preparations is dispersed, possibly suggesting the need for further stabilizing homo- or hetero- protein-protein interactions with physiological relevance in vivo.
The elongated shape of the GLT25D1/COLGALT1 dimer resembles that of the tail-to-tail dimer observed for LH3/PLOD3, possibly facilitating extensive interactions among the two enzymes responsible for the subsequent processing of collagen lysines into Glc-Gal-Hyl and posing questions regarding the possible evolution of this two-enzyme system from a common ancestor. This is further supported by the structural similarity observed between the two non-functional domains of these enzymes (Supplementary Fig. 5c): the GLT25D1/COLGALT1 GT1 domain, unexpectedly found bound to Ca2+ and hosting UDP-α-Gal but only to preserve the conformational integrity of the domain, and the LH/PLOD AC domain, which cannot bind metal ions and/or co-substrates41. Such hypothesis prompts for extensive future work, also considering the unusual trafficking of multifunctional LH/PLOD enzymes through the secretory pathway due to absence of specific ER retention sequences44,45,46,47.
Methods
Chemicals
All chemicals were purchased from Sigma-Aldrich unless otherwise specified.
DNA constructs
The sequence encoding for human GLT25D1/COLGALT1, deprived of the N-terminal signal peptide and the C-terminal RDEL retention sequence (UniProt Q8NBJ5, residues 30 to 618), was synthesized by Genewiz and sub-cloned into a pCR8 cloning vector with in-frame 5’-BamHI and 3’-NotI restriction sites. GLT25D1/COLGALT1 mutants were generated using the Phusion Site Directed Mutagenesis Kit (Thermo Fisher Scientific). The entire plasmid was amplified using the primers listed in Supplementary Table 1. The linear mutagenized plasmids were then phosphorylated using T4 polynucleotide kinase (Invitrogen) prior to ligation. All plasmids were checked by Sanger sequencing (Microsynth) prior to cloning into a modified pET28b-SUMO vector (Novagen). The recombinant plasmids include an N-terminal 8xHis-tag followed by a small ubiquitin-like modifier (SUMO) tag preceding the in-frame 5’-BamHI restriction site, as well as an in-frame stop codon after the 3’-NotI restriction site.
Production of recombinant GLT25D1/COLGALT1
Chemically competent E. coli BL21 cells (Invitrogen) were transformed with the pET28b-SUMO-GLT25D1 recombinant plasmid using heat shock. A single colony was inoculated into Lysogeny broth (LB) medium supplemented with 100 μg ml−1 of kanamycin and grown overnight at 37 °C in a shaking incubator (New Brunswick). This pre-culture was then inoculated at a 1:50 ratio in 1 L of ZYP5052 autoinducing medium48 in a 5 L Erlenmeyer flask. The culture was kept at 37 °C for 3 h at 180 r.p.m. shaking speed. The temperature was then lowered to 17 °C, and further incubated overnight. Bacterial cells were harvested by centrifugation at 5000 × g, resuspended at 1:10 w/v ratio in a lysis buffer composed of 25 mM 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (HEPES)/NaOH, 500 mM NaCl, 10 μM leupeptin, 10 μM pepstatin, 0.3 mg ml−1 chicken egg white lysozyme, 500 μM MnSO4, pH 8.0. The suspension was subject to sonication (16 cycles, 9 s on, 6 s off pulses). Cell debris was removed by centrifuging the cell lysate at 60,000 × g for 45 min, at 4 °C in an Avanti J26 super centrifuge (Beckman Coulter). The supernatant containing soluble His-SUMO-GLT25D1/COLGALT1 protein was filtered through a MiniSart GF 0.8 μm filter (Sartorius), loaded onto a 5 mL Ni Sepharose Excel column (Cytiva) and eluted using a stepwise gradient of an elution buffer composed of 25 mM HEPES/NaOH, 500 mM NaCl, 500 mM imidazole, pH 8.0. The fractions of the eluate containing His-SUMO-GLT25D1/COLGALT1, as assessed by SDS-PAGE analysis, were pooled and dialyzed overnight against 2 L of 25 mM HEPES/NaOH, 500 mM NaCl, pH 8.0. The N-terminal 8xHis-SUMO tag was simultaneously cleaved by incubating the protein with 1 mg mL−1 His-tagged SUMO protease (1:300 v/v). Affinity-based removal of SUMO protease and 8xHis-SUMO-tag was achieved by passing the purified sample through a 5 mL Ni Sepharose Excel column (Cytiva) and collecting the flow-through fraction. The resulting sample was subject to buffer exchange in 30 kDa MWCO Vivaspin Turbo centrifugal filters (Sartorius) against 25 mM HEPES/NaOH, 100 mM NaCl, pH 8.0 and loaded into a HiScreen Capto Q column (Cytiva) pre-equilibrated with the same buffer. A linear NaCl gradient was then applied, resulting in GLT25D1/COLGALT1 elution with approximately 250 mM NaCl. The sample was concentrated using 30 kDa MWCO Vivaspin Turbo centrifugal filters (Sartorius) and subject to final polishing into a Superdex 200 Increase 10/300 GL (Cytiva), equilibrated with 25 mM HEPES/NaOH, 100 mM NaCl, pH 8.0. The final sample quality was assessed by reducing and non-reducing SDS-PAGE analysis. GLT25D1/COLGALT1-containing fractions were pooled, concentrated to 4 mg mL−1, and stored at −80 °C prior to usage. Results of SDS-PAGE sample quality controls for all proteins used in this study are shown in Supplementary Fig. 1.
Protein crystallization
GLT25D1/COLGALT1 spherulites were initially found in various conditions of nanoliter-dispensed droplets (0.1 μL protein at 4 mg mL−1 + 0.1 μL reservoirs), dispensed using a Gryphon crystallization robot (Art Robbins) using commercial crystallization screens in sitting-drop vapor diffusion drop plates (SwissSci). In particular, small microcrystals were initially identified after few days at 4 °C in conditions of the Morpheus crystallization screen49 (Molecular Dimensions), indicating a preference for poly-ethylene-glycol (PEG) as precipitant and pH between 6.0 and 7.0. Crystal optimization was carried out by systematically varying the precipitating agent, buffering agent, pH and additives composition of the initial hits from the commercial screen. Diffraction-quality crystals were eventually obtained by manually mixing 0.5 μL of protein concentrated at 3.5 mg mL−1 and 0.5 μL of an optimized reservoir solution composed of 8% PEG MW 4000, 100 mM 2-(N-morpholino)ethanesulfonic acid (MES)/NaOH, 20% glycerol, pH 6.5. Co-crystallization experiments were performed by setting up the same conditions and supplementing the protein solution with 500 μM MnCl2 and 1 mM UDP-α-Gal. Crystals were harvested using mounted Litholoops (Molecular Dimensions), flash-cooled and stored in liquid nitrogen prior to data acquisition. Heavy atom derivatives were prepared by soaking the GLT25D1/COLGALT1 crystals in mother liquor conditions containing 1 mM K2HgBr4. Crystals were incubated with the heavy atom solution for at least 5 h at 4 °C prior to cryoprotection, harvesting and flash-cooling in liquid nitrogen.
X-ray data collection, structure determination and refinement
X-ray diffraction data from single crystals were collected at 100 K at various beamlines of the European Synchrotron Radiation Facility (ESRF) in Grenoble, France and the Swiss Light Source (SLS) in Villigen, Switzerland. Experimental phasing was achieved by collecting X-ray diffraction data at the mercury inflection point at the ID23-1 beamline of the ESRF synchrotron equipped with an Eiger 2 16 M (Dectris) detector. Data were indexed and integrated with XDS50, followed by scaling and merging using AIMLESS51. Data collection statistics are given in Supplementary Table 2. Single wavelength anomalous dispersion (SAD) phasing was performed with the SHELXC/D/E pipeline and HKL2MAP52. A total of 13 heavy atom sites were identified at 4.0 Å resolution in SHELXD. Phasing with SHELXE resulted in non-ambiguous identification of the map handedness immediately after the initial rounds of density modification with phase extension to 2.80 Å, with 62% solvent content. The density modified experimental electron density map was used for tracing the initial model by combining multiple rounds of ARP/wARP and BUCCANEER53. The model was completed and the structure was refined by alternating steps of manual building in COOT and automated refinement with phenix.refine54 and REFMAC555. Crystal structures of GLT25D1/COLGALT1 in complex with donor substrates were subsequently determined using molecular replacement in PHASER, using the experimental structure of the enzyme as search model. Validation was carried out using MolProbity and the validation tools available on the Protein Data Bank server56. Final refinement statistics are summarized in Supplementary Table 3. Structural figures were prepared with PyMol (http://www.pymol.org). Superpositions were performed using the “super” command in PyMol. All-atom root mean square deviation (RMSD) values were computed accordingly.
Mass photometry
Mass photometry measurements were carried out on a Refeyn Two Mass Photometer (Refeyn) using 24 × 50 mm2 glass coverslips (Refeyn). Initial 200 nM stocks of purified human GLT25D1/COLGALT1 wild-type and mutants were diluted 1:10 using 25 mM HEPES/NaOH, 100 mM NaCl, pH 8.0, and analyzed under a field of view of 4 μm × 11 μm at a frame rate of 500 Hz. The data were collected using AcquireMP (Refeyn), then processed and plotted using DiscoverMP (Refeyn). Results of the data analysis are summarized in Supplementary Table 4.
Size exclusion chromatography coupled with multi-angle light scattering (SEC-MALS)
30 μl of 4 mg mL−1 recombinant GLT25D1/COLGALT1 were injected into a Protein KW-802.5 analytical size-exclusion column (Shodex) and separated with a flow rate of 1 mL min−1 in phosphate buffer saline using a Prominence high-pressure liquid chromatography (HPLC) system (Shimadzu). For molecular weight characterization, light scattering was measured with a miniDAWN multi-angle light scattering detector (Wyatt), connected to a RID-20A differential refractive index detector (Shimadzu) for quantitation of the total mass and to a SPD-20A UV detector (Shimadzu) for evaluation of the sole protein content. Chromatograms were collected and analyzed using the ASTRA7 software (Wyatt), using an estimated dn/dc value of 0.185 ml/g. The calibration of the instrument was verified by injection of 10 μL of 2.5 mg mL−1 monomeric BSA.
Differential scanning fluorimetry (DSF)
DSF assays on recombinant GLT25D1/COLGALT1 samples (wild-type and mutants) at a concentration of 1 mg mL−1 in 25 mM HEPES/NaOH, 100 mM NaCl, pH 8.0 were performed using a Tycho NT.6 instrument (NanoTemper Technologies GmbH). Data were analyzed and plotted using Graphpad Prism 757.
Evaluation of Gal-T activity through luminescence
Reaction mixtures (5 μL total volume in 25 mM HEPES/NaOH, 100 mM NaCl, pH 8.0) were prepared by initially incubating GLT25D1/COLGALT1 (final concentration 1 mg mL−1) at 37 °C for 3 h with variable amounts of bovine skin gelatin, previously solubilized through heating denaturation at 95 °C for 10 min. After incubation, 100 μM of UDP-α-sugar and 50 μM MnCl2 or MgCl2 were added to start the enzymatic reactions. After a precise timing between 1 and 120 min the reactions were stopped by heating the samples at 95 °C for 2 min, then the samples were transferred into Proxiplate white 384-well plates (Perkin-Elmer). 5 μL of the UDP-Glo luminescence detection reagent (Promega) were added and let incubate for 1 h at 25 °C. The plates were then transferred into a GloMax Discovery plate reader (Promega) configured according to manufacturer’s instructions for luminescence detection. Initial velocities were measured through the determination of slopes from linear fit interpolation of individual time points using GraphPad Prism 757. After visual inspection of double reciprocal plots, which were linear (Supplementary Fig. 2c), the initial velocity values, expressed as apparent mean turnover values and associated errors, were directly fitted to the Michaelis-Menten equation using GraphPad Prism 757, obtaining the values of apparent kcat and Km accounting for propagation of statistical errors58. All experiments were performed in triplicates. Control experiments were performed using identical conditions by selectively removing GLT25D1, donor or acceptor substrates. Results of the data analysis are summarized in Supplementary Table 5.
Evaluation of Gal-T activity through HR-LCMS
5 μM recombinant human LH3/PLOD3 (obtained in house as described in previous work) was incubated with 5 μM GLT25D1/COLGALT1 (wild-type or mutants), 50 μM FeCl2, 100 μM 2-oxoglutarate (2-OG), 500 μM sodium ascorbate, 50 μM MnCl2, 100 μM UDP-α-sugar, and 1 mM peptide (Ac-GIKGIKGIKGIK-COOH) substrate. The reactions were allowed to proceed for 3 h at 37 °C. 5 μL of each reaction were diluted with 43 μL of Milli-Q water and acidified by addition of 2 μL of formic acid, to reach a final volume of 50 μL, and then analyzed on an Exion LC AD UHPLC (AB Sciex) coupled to a X500B ESI-HRMS/MS system (AB Sciex) by a full scan acquisition. The column oven was kept at 40 °C, while the autosampler was cooled at 10 °C. Peptides were separated by reverse phase HPLC on a Hypersil Gold C18 column (150 × 2.1 mm, 3 μm particle size, 175 Å pore size, Thermo Fisher Scientific) using a linear gradient (2–50% solvent B in 15 min), with solvent A consisting of 0.1% aqueous formic acid and solvent B of acetonitrile (ACN) containing 0.1% formic acid. Flow rate was kept constant at 0.2 mL min−1. Mass spectra were generated in positive polarity under constant instrumental conditions: ion spray voltage 4500 V, declustering potential 100 V, curtain gas 30 psi, ion source gas 1 40 psi, ion source gas 2 45 psi, temperature 350 °C, collision energy 10 V. Spectra analyses were performed using the SCIEX OS 2.1 software (AB Sciex). Results of the data analysis are summarized in Supplementary Table 5.
Size exclusion chromatography coupled with small-angle X-ray scattering (SEC-SAXS)
Solution scattering data were collected at ESRF BM29 using a sec−1 frame rate on Pilatus 1 M detector located at a fixed distance of 2.87 m from the sample, allowing a global q range of 0.01–4.00 nm. SEC-SAXS experiments were carried out using Nexera High Pressure Liquid/Chromatography (Shimadzu) system connected online to SAXS sample capillary59. For these experiments, 50 μL of GTL25D1/COLGALT1 concentrated at 4 mg mL−1 were injected into a Superdex 200 PC 3.2/300 Increase column (GE Healthcare), pre-equilibrated with 25 mM HEPES/NaOH, 200 mM NaCl, pH 8.0. For SEC-SAXS data, frames corresponding to GTL25D1/COLGALT1 protein peak were identified, blank subtracted, and averaged using CHROMIXS60.
SEC-SAXS modeling and data analysis
Radii of gyration (Rg), molar mass estimates and distance distribution functions P(r) were computed using PRIMUS61 within the ATSAS package62. Comparison of experimental SAXS data and 3D models from crystal structures was performed using CRYSOL63. A summary of SAXS data collection and analysis results is reported in Supplementary Table 6. Ab initio sphere models were generated from the SAXS data using GASBOR, by providing the software with P(r) functions derived from experimental data and the number of residues defining GLT25D1/COLGALT1 monomers and without imposing any internal symmetry. 20 independent ab initio GASBOR runs were performed and subject to similarity analysis using SUPCOMB and DAMSEL from ATSAS, allowing selection of a representative individual model based on normalized spatial discrepancy. Superpositions were carried out using the SUPALM program available in the saspy ATSAS plugin in PyMol64. For SAXS rigid-body modeling, the model of a GLT25D1/COLGALT1 monomer from the experimental crystal structure was subject to loop modeling using CORAL65. Evaluation of the agreement between molecular models and experimental SAXS profiles was carried out using CRYSOL63.
Single particle negative staining electron microscopy
200 mesh, Cu grids (Ted Pella) grids were negatively charged using a PELCO EasiGlow (Ted Pella) with the following parameters: pressure 40 mBar, power 15 mA, time 30 s. 2 μL of 1 mg mL−1 GLT25D1/COLGALT1 were applied on each grid and left for 60 s. The excess sample was blotted using Whatman paper and then a 25 μL drop of 2% (w/v) uranyl acetate solution was applied, gently stirring for 1 min. The excess stain was blotted with filter paper and the grids were left dry overnight. 350 EM micrographs were acquired on a JEM 1200EXIII (JEOL) electron microscope equipped with a Mega View III CCD camera (Olympus). Data processing was carried out using RELION, including auto-picking and multiple rounds of 2D classification. Comparison of electron microscopy single particle analysis 2D classes and GLT25D1/COLGALT1 molecular models from crystal structures was carried out using AlignProjections within the COSMIC2 webserver66.
GLT25D1/COLGALT1-LH3/PLOD3 cross-linking and enzymatic proteolysis
5 μM GLT25D1/COLGALT1 (wild-type and Trp158Arg) were mixed with 5 μM human LH3/PLOD367 (total protein concentration 10 μM) in 50 mM HEPES/NaOH, 100 mM NaCl, pH 8.0. The samples were incubated at room temperature for 1 h with 0.5 mM DSBU. The cross-linker was dissolved in neat dimethyl sulfoxide (DMSO) immediately before adding it to the peptide solution. The reaction was quenched by adding 2 μL of 1 M Tris/HCl solution. Cross-linked samples were loaded onto S-Trap microcolumns (Protifi) according to the manufacturer’s instructions. In brief, after loading, samples were washed with 90:10% methanol/50 mM ammonium bicarbonate. The samples were then digested with trypsin (Promega) (1:20 trypsin/protein) for 1.5 h at 47 °C. The digested peptides were eluted using 50 mM ammonium bicarbonate. Two additional elutions were made using 0.2% formic acid and 0.2% formic acid in 50% ACN. The three fractions were pooled together and vacuum-centrifuged to dryness. The resulting peptides were resuspended in 5% ACN, 0.1% trifluoroacetic acid for subsequent LC-MS/MS analyses.
Liquid chromatography coupled to mass spectrometry for cross-linking peptide analysis
Proteolyzed cross-linked samples were analyzed via LC-MS/MS on an UltiMate 3000 RSLC nano-HPLC system (Thermo Fisher Scientific) coupled to a timsTOF Pro mass spectrometer equipped with CaptiveSpray source (Bruker Daltonics). Peptides were trapped on a C18 column (precolumn Acclaim PepMap 100, 300 μm × 5 mm, 5 μm, 100 Å, Thermo Fisher Scientific) at 50 °C and separated on a self-packed Picofrit (New Objective) nanospray emitter (360 μm o.d. × 75 μm i.d. × 400 mm L, 15 μm Tip i.d.) with C18-stationary phase (3.0 μm, 120 Å, Dr. Maisch GmbH). After trapping, peptides were eluted by a linear 90 min water−ACN gradient from 3% (v/v) to 50% (v/v) ACN. For the timsTOF Pro MS settings, the following parameters were adopted. The values for mobility-dependent collision energy ramping were set to 95 eV at an inverse reduced mobility (1/k0) of 1.6 V s/cm2 and 23 eV at 0.73 V s/cm2. Collision energies were linearly S-5 interpolated between these two 1/k0 values and kept constant above or below. No merging of TIMS scans was performed. Target intensity per individual PASEF precursor was set to 20,000. The scan range was defined between 0.6 and 1.6 V s/cm2 with a ramp time/accumulation time of 166 ms. 14 PASEF MS/MS scans were triggered per cycle (2.57 s) with a maximum of seven precursors per mobilogram. Precursor ions in an m/z range between 100 and 1700 with charge states ≥3+ and ≤8+ were selected for fragmentation. Active exclusion was enabled for 0.4 min (mass width 0.015 Th, 1/k0 width 0.015 V s/cm2).
XL-MS data analysis
Cross-linked peptides were identified using MeroX68 (version 2.0.1.7, https://stavrox.com). The search parameters were set as follows: tryptic cleavage C-terminal to Lys and Arg, allowing up to four missed cleavages; peptide length range 3–30 amino acids; fixed modification: Cys alkylation (iodoacetamide); variable modification: Met oxidation. The cross-linker was specified to react with Lys and N-termini. Cross-link identification was performed in RISEUP mode, allowing a maximum of three missing ions. The precursor and fragment mass tolerances were set to 10 ppm and 20 ppm, respectively. The signal-to-noise ratio was required to be >2.0. False discovery rate (FDR) filtering was applied at 1% at the peptide-spectrum match (PSM) level, with a minimum score cut-off of 100. To improve FDR estimation, a contaminant database (CRAP database) was included.
Proteomic data analysis was conducted using MaxQuant69 (version 2.6.2.0) and Skyline70 (version 24.1.1.339). Bruker raw files were processed using data-dependent acquisition (DDA) searches against the UniProt Human Reference Proteome (UP000005640), which includes 20,406 canonical proteins. Contaminants were filtered using the default MaxQuant contaminant database. The search parameters included tryptic cleavage C-terminal to Lys and Arg, allowing up to two missed cleavages. The peptide length was set to 6–25 amino acids. Fixed modifications included Cys alkylation (iodoacetamide), while variable modifications included Met oxidation and N-terminal acetylation. For label-free quantification (LFQ), a minimum of three unique peptides per protein was required. The MS1 and MS2 tolerances were set to 10 ppm and 20 ppm, respectively. The Match Between Runs (MBR) feature was enabled, with a retention time alignment window of 0.4 min and an ion mobility window of 0.05. PSM and protein-level FDR were controlled at 1%. MaxQuant and MeroX results were imported into Skyline for MS/MS spectrum inspection and peptide quantification at MS1 level. Protein quantification was based on the ten most abundant validated peptides per protein. MS/MS spectra for the PLOD3-GLT25D1 inter-protein cross-link and relevant dead-end products were imported into Skyline, following previously established workflows71, and quantified.
ICP-MS measurements
To quantify the possible presence and concentration of metal ions in GLT25D1/COLGALT1 recombinant preparations, 900 μL of a sample containing 4 mg mL−1 GLT25D1/COLGALT1 (purified without usage of divalent ions) were treated with 300 μL 65% ultrapure HNO3 and 200 μL 30% w/w H2O2, made up to 5 mL with ultrapure water and analyzed on a single quadrupole inductively coupled plasma mass spectrometer (SQ-ICP-MS, iCAP RQ Thermo Fisher Scientific), equipped with a quartz cyclonic chamber cooled at 3 °C, a MicroMist nebulizer 179 (400 μL min−1), a quartz torch, a Ni sampler, skimmer cones, and a QCell pressurized with helium (3 V, KED mode). Quantitative determinations for manganese and calcium were obtained by the external standard calibration with four standards (0, 1, 5, 10 μg L−1) prepared daily in the same buffer used for sample preparation, at the same dilution and HNO3 concentration. While manganese could not be detected, calcium was detected with a 1:1 stoichiometry.
Native mass spectrometry (Native-MS)
Native-MS was employed to identify metal cofactors of GLT25D1/COLGALT1. Recombinant wild-type and Trp158Arg GLT25D1/COLGALT1 enzymes (20 μM) were analyzed under partially denaturing conditions (50 mM ammonium bicarbonate, 40% acetonitrile), in order to improve droplets desolvation and to minimize the adducts formation during electrospray. Samples were directly injected into an Orbitrap Fusion instrument (Thermo Fisher Scientific) using metal-coated borosilicate emitters (~1 μm internal diameter). The main instrumental parameters were set as follows: resolving power at m/z 200, 120,000; IRM pressure, 3 mTorr (intact protein mode); ion spray voltage, 1.1–1.2 kV; ion-transfer tube temperature, 275 °C; in-source fragmentation, 75 V; AGC target, 4 × 105; maximum injection time, 100 ms. Final spectra were obtained by averaging the signal over 60 s acquisition. Ligand mass was calculated as the difference between detected bound- and free-protein signals over the entire charge state distribution of the spectra (Supplementary Fig. 10).
Molecular dynamics simulations
Molecular dynamics simulations were carried out using the AMBER software package (version 24) with the ff19SB force field, using its GPU-accelerated pmemd.cuda utility during equilibration and production, and sander otherwise. Comprehensive descriptions of the preparation of the simulated systems and molecular dynamics simulation protocols can be found in the Supplementary Information. Analyses of MD trajectories were carried out with the cpptraj program distributed within the ambertools suite (version 24) and in-house scripts.
Principal component analysis (PCA) was performed to identify the major modes of motion in the molecular dynamics simulations of the GT1 and GT2 domains, combining the trajectories of the ligand-free and ligand-bound states of each domain. The covariance matrix of the backbone atoms of the simulated systems was calculated, and diagonalized to obtain the principal components (eigenvectors) and their corresponding eigenvalues. The eigenvalues corresponding to each principal components were used to evaluate the percentage of the variance in the motion of the protein represented by each component. Since the first two components, PC1 and PC2, account for most of the variance in all the simulated systems, the snapshots of the trajectories were projected only onto these principal components, and histograms of the normalized frequency of the conformational states along PC1 and PC2 were generated.
Hydrogen bond persistence analysis was performed using the hbond command in the cpptraj module of AMBER24. A hydrogen bond was considered present based on geometric criteria: a donor–acceptor distance cutoff of 3.5 Å and a hydrogen-donor-acceptor angle cutoff of 135°. The persistence of each hydrogen bond was monitored throughout the simulation time, and the hydrogen bond frequencies were normalized based on the total number of hydrogen bonds to represent the percentage of time each bond was present during the trajectory. These results were then grouped into 10% frequency intervals to generate histograms, illustrating the persistence of hydrogen bonds over the course of the simulation.
The distance fluctuation (DF) parameter for each pair of residues i and j in a protein system is defined as Dij = 〈(dij − 〈dij〉)2〉, where dij is the time-dependent distance between the Cα atoms of residues i and j, and the brackets indicates the time-averaged distance over the entire trajectory. This calculation results in a matrix that characterizes the extent of movement or flexibility between all residue pairs.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Unless otherwise stated, all data supporting the results of this study can be found in the article, supplementary, and source data files. Coordinates and structure factors have been deposited in the Protein Data Bank under accession codes 9EVJ, 9EVK, and 9EVL. In solution SEC-SAXS data have been deposited in the SASBDB under accession code SASDVZ2. XL-MS data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the project accession PXD061021. Native MS data have been deposited in the MassIVE database with dataset identifier MSV000097322 [https://doi.org/10.25345/C5TH8C06M]. MD simulation data, including initial coordinate files, simulation input files, and final output coordinate files have been deposited in Zenodo [https://doi.org/10.5281/zenodo.15020231]. The numerical source data for Supplementary Fig. 9 can be found in the raw data deposited on PRIDE under accession number PXD061021. The numerical source data for Supplementary Fig. 11 can be found in the raw data deposited on MassIVE under accession number MSV000097322 [https://doi.org/10.25345/C5TH8C06M]. Source data are provided with this paper.
References
Bourhis, J. M. et al. Structural basis of fibrillar collagen trimerization and related genetic disorders. Nat. Struct. Mol. Biol. 19, 1031–1036 (2012).
Fidler, A. L., Boudko, S. P., Rokas, A. & Hudson, B. G. The triple helix of collagens—an ancient protein structure that enabled animal multicellularity and tissue evolution. J. Cell Sci. 131, jcs203950 (2018).
Myllyharju, J. & Kivirikko, K. I. Collagens and collagen-related diseases. Ann. Med. 33, 7–21 (2001).
Ishikawa, Y. & Bachinger, H. P. A molecular ensemble in the rER for procollagen maturation. Biochim. Biophys. Acta 1833, 2479–2491 (2013).
Gelse, K., Poschl, E. & Aigner, T. Collagens-structure, function, and biosynthesis. Adv. Drug Deliv. Rev. 55, 1531–1546 (2003).
Kivirikko, K. I. & Myllyla, R. Posttranslational enzymes in the biosynthesis of collagen: intracellular enzymes. Methods Enzymol. 82, 245–304 (1982).
Kadler, K. E., Holmes, D. F., Trotter, J. A. & Chapman, J. A. Collagen fibril formation. Biochem. J. 316, 1–11 (1996).
Bella, J. Collagen structure: new tricks from a very old dog. Biochem. J. 473, 1001–1025 (2016).
Myllyharju, J. Prolyl 4-hydroxylases, the key enzymes of collagen biosynthesis. Matrix Biol. 22, 15–24 (2003).
Gorres, K. L. & Raines, R. T. Prolyl 4-hydroxylase. Crit. Rev. Biochem. Mol. Biol. 45, 106–124 (2010).
Yamauchi, M. & Sricholpech, M. Lysine post-translational modifications of collagen. Essays Biochem. 52, 113–133 (2012).
Cummings, R. D. The repertoire of glycan determinants in the human glycome. Mol. Biosyst. 5, 1087–1104 (2009).
Hennet, T. Collagen glycosylation. Curr. Opin. Struct. Biol. 56, 131–138 (2019).
Mori, K., Suzuki, T., Miura, K., Dohmae, N. & Simizu, S. Involvement of LH3 and GLT25D1 for glucosyl-galactosyl-hydroxylation on non-collagen-like domain of FGL1. Biochem. Biophys. Res. Commun. 560, 93–98 (2021).
Tvaroska, I. Glycosylation modulates the structure and functions of collagen: a review. Molecules https://doi.org/10.3390/molecules29071417 (2024).
De Giorgi, F., Fumagalli, M., Scietti, L. & Forneris, F. Collagen hydroxylysine glycosylation: non-conventional substrates for atypical glycosyltransferase enzymes. Biochem. Soc. Trans. 49, 855–866 (2021).
Scietti, L. & Forneris, F. Full-length human collagen lysyl hydroxylases. in Encyclopedia of Inorganic and Bioinorganic Chemistry (ed. Scott, R. A.) 1–12 (Wiley, 2020).
Salo, A. M. & Myllyharju, J. Prolyl and lysyl hydroxylases in collagen synthesis. Exp. Dermatol. https://doi.org/10.1111/exd.14197 (2020).
Yamauchi, M., Terajima, M. & Shiiba, M. Lysine hydroxylation and cross-linking of collagen. Methods Mol. Biol. 1934, 309–324 (2019).
Kivirikko, K. I. & Myllyla, R. Collagen glycosyltransferases. Int. Rev. Connect Tissue Res. 8, 23–72 (1979).
Wang, C. et al. The third activity for lysyl hydroxylase 3: galactosylation of hydroxylysyl residues in collagens in vitro. Matrix Biol. 21, 559–566 (2002).
Geister, K. A. et al. Loss of function of Colgalt1 disrupts collagen post-translational modification and causes musculoskeletal defects. Dis. Model Mech. https://doi.org/10.1242/dmm.037176 (2019).
Terajima, M. et al. Role of glycosyltransferase 25 domain 1 in type I collagen glycosylation and molecular phenotypes. Biochemistry 58, 5040–5051 (2019).
Webster, J. A. et al. Collagen beta (1-O) galactosyltransferase 1 (GLT25D1) is required for the secretion of high molecular weight adiponectin and affects lipid accumulation. Biosci. Rep. https://doi.org/10.1042/BSR20170105 (2017).
Baumann, S. & Hennet, T. Collagen accumulation in osteosarcoma cells lacking GLT25D1 collagen galactosyltransferase. J. Biol. Chem. 291, 18514–18524 (2016).
Yang, J. et al. Collagen beta(1-O) galactosyltransferase 2 deficiency contributes to lipodystrophy and aggravates NAFLD related to HMW adiponectin in mice. Metabolism 120, 154777 (2021).
Kehayova, Y. S., Wilkinson, J. M., Rice, S. J. & Loughlin, J. Osteoarthritis genetic risk acting on the galactosyltransferase gene COLGALT2 has opposing functional effects in articulating joint tissues. Arthritis Res. Ther. 25, 83 (2023).
Schegg, B., Hulsmeier, A. J., Rutschmann, C., Maag, C. & Hennet, T. Core glycosylation of collagen is initiated by two beta(1-O)galactosyltransferases. Mol. Cell Biol. 29, 943–952 (2009).
Liefhebber, J. M., Punt, S., Spaan, W. J. & van Leeuwen, H. C. The human collagen beta(1-O)galactosyltransferase, GLT25D1, is a soluble endoplasmic reticulum localized protein. BMC Cell Biol. 11, 33 (2010).
Kehayova, Y. S., Watson, E., Wilkinson, J. M., Loughlin, J. & Rice, S. J. Genetic and epigenetic interplay within a COLGALT2 enhancer associated with osteoarthritis. Arthritis Rheumatol. 73, 1856–1865 (2021).
Guo, T. et al. COLGALT2 is overexpressed in ovarian cancer and interacts with PLOD3. Clin. Transl. Med. 11, e370 (2021).
Wang, Y. et al. Exosomes secreted by adipose-derived mesenchymal stem cells foster metastasis and osteosarcoma proliferation by increasing COLGALT2 expression. Front Cell Dev. Biol. 8, 353 (2020).
Wang, S. et al. Upregulation of GLT25D1 in hepatic stellate cells promotes liver fibrosis via the TGF-beta1/SMAD3 pathway in vivo and in vitro. J. Clin. Transl. Hepatol. 11, 1–14 (2023).
Perrin-Tricaud, C., Rutschmann, C. & Hennet, T. Identification of domains and amino acids essential to the collagen galactosyltransferase activity of GLT25D1. PLoS ONE 6, e29390 (2011).
Holm, L. & Rosenstrom, P. Dali server: conservation mapping in 3D. Nucleic Acids Res. 38, W545–W549 (2010).
Luther, K. B. et al. Mimivirus collagen is modified by bifunctional lysyl hydroxylase and glycosyltransferase enzyme. J. Biol. Chem. 286, 43701–43709 (2011).
Lairson, L. L., Henrissat, B., Davies, G. J. & Withers, S. G. Glycosyltransferases: structures, functions, and mechanisms. Annu. Rev. Biochem 77, 521–555 (2008).
Taujale, R. et al. Deep evolutionary analysis reveals the design principles of fold A glycosyltransferases. Elife https://doi.org/10.7554/eLife.54532 (2020).
Miyatake, S. et al. Biallelic COLGALT1 variants are associated with cerebral small vessel disease. Ann. Neurol. 84, 843–853 (2018).
Teunissen, M. W. A. et al. Biallelic variants in the COLGALT1 gene causes severe congenital porencephaly: a case report. Neurol. Genet. 7, e564 (2021).
Scietti, L. et al. Molecular architecture of the multifunctional collagen lysyl hydroxylase and glycosyltransferase LH3. Nat. Commun. 9, 3163 (2018).
Sadakierska-Chudy, A., Szymanowski, P., Lebioda, A. & Ploski, R. Identification and in silico characterization of a novel COLGALT2 gene variant in a child with mucosal rectal prolapse. Int. J. Mol. Sci. https://doi.org/10.3390/ijms23073670 (2022).
Peng, J. et al. The structural basis for the human procollagen lysine hydroxylation and dual-glycosylation. Nat. Commun. 16, 2436 (2025).
Salo, A. M. et al. Lysyl hydroxylase 3 (LH3) modifies proteins in the extracellular space, a novel mechanism for matrix remodeling. J. Cell Physiol. 207, 644–653 (2006).
Wang, C., Ristiluoma, M. M., Salo, A. M., Eskelinen, S. & Myllyla, R. Lysyl hydroxylase 3 is secreted from cells by two pathways. J. Cell Physiol. 227, 668–675 (2012).
Chen, Y. L. et al. Lysyl hydroxylase 2 is secreted by tumor cells and can modify collagen in the extracellular space. J. Biol. Chem. 291, 25799–25808 (2016).
Banushi, B. et al. Regulation of post-Golgi LH3 trafficking is essential for collagen homeostasis. Nat. Commun. 7, 12111 (2016).
Studier, F. W. Protein production by auto-induction in high density shaking cultures. Protein Expr. Purif. 41, 207–234 (2005).
Gorrec, F. The MORPHEUS protein crystallization screen. J. Appl. Crystallogr. 42, 1035–1042 (2009).
Kabsch, W. Xds. Acta Crystallogr D. Biol. Crystallogr 66, 125–132 (2010).
Evans, P. R. & Murshudov, G. N. How good are my data and what is the resolution? Acta Crystallogr D. Biol. Crystallogr 69, 1204–1214 (2013).
Pape, T. & Schneider, T. R. HKL2MAP: a graphical user interface for macromolecular phasing with SHELX programs. J. Appl. Crystallogr. 37, 843–844 (2004).
Cowtan, K. The Buccaneer software for automated model building. 1. Tracing protein chains. Acta Crystallogr D. Biol. Crystallogr 62, 1002–1011 (2006).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr D. Biol. Crystallogr 66, 213–221 (2010).
Murshudov, G. N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr D. Biol. Crystallogr 67, 355–367 (2011).
Gore, S. et al. Validation of structures in the protein data bank. Structure 25, 1916–1927 (2017).
Graphpad Software, L.J. Graphpad Prism 7 (Graphpad Software, USA).
Bevington, P. H. Data Reduction and Error Analysis for the Physical Sciences, 56–62 (McGraw-Hill, 1969).
Brennich, M. E., Round, A. R. & Hutin, S. Online size-exclusion and ion-exchange chromatography on a SAXS beamline. J. Vis. Exp. https://doi.org/10.3791/54861 (2017).
Panjkovich, A. & Svergun, D. I. CHROMIXS: automatic and interactive analysis of chromatography-coupled small angle X-ray scattering data. Bioinformatics https://doi.org/10.1093/bioinformatics/btx846 (2017).
Konarev, P. V., Volkov, V. V., Sokolova, A. V., Koch, M. H. J. & Svergun, D. I. PRIMUS: a Windows PC-based system for small-angle scattering data analysis. J. Appl. Crystallogr. 36, 1277–1282 (2003).
Franke, D. et al. ATSAS 2.8: a comprehensive data analysis suite for small-angle scattering from macromolecular solutions. J. Appl. Crystallogr 50, 1212–1225 (2017).
Svergun, D., Barberato, C. & Koch, M. H. J. CRYSOL—a program to evaluate X-ray solution scattering of biological macromolecules from atomic coordinates. J. Appl. Crystallogr. 28, 768–773 (1995).
Panjkovich, A. & Svergun, D. I. SASpy: a PyMOL plugin for manipulation and refinement of hybrid models against small angle X-ray scattering data. Bioinformatics 32, 2062–2064 (2016).
Petoukhov, M. V. et al. New developments in the ATSAS program package for small-angle scattering data analysis. J. Appl. Crystallogr. 45, 342–350 (2012).
Cianfrocco, M. A., Wong-Barnum, M., Youn, C., Wagner, R. & Leschziner, A. COSMIC2: a science gateway for cryo-electron microscopy structure determination. in Proceedings of the Practice and Experience in Advanced Research Computing 2017 on Sustainability, Success and Impact Article 22 (Association for Computing Machinery, 2017).
Mattoteia, D. et al. Identification of regulatory molecular “hot spots” for LH/PLOD collagen glycosyltransferase activity. Int. J. Mol. Sci. https://doi.org/10.3390/ijms241311213 (2023).
Iacobucci, C. et al. A cross-linking/mass spectrometry workflow based on MS-cleavable cross-linkers and the MeroX software for studying protein structures and protein-protein interactions. Nat. Protoc. 13, 2864–2889 (2018).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
MacLean, B. et al. Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 26, 966–968 (2010).
Rojas Echeverri, J. C. et al. A workflow for improved analysis of cross-linking mass spectrometry data integrating parallel accumulation-serial fragmentation with MeroX and Skyline. Anal. Chem. 96, 7373–7379 (2024).
Laskowski, R. A. & Swindells, M. B. LigPlot+: multiple ligand-protein interaction diagrams for drug discovery. J. Chem. Inf. Model 51, 2778–2786 (2011).
Acknowledgements
We thank Ms. Lisa Negro and Ms. Martina Soffientini for assistance with production of recombinant GLT25D1/COLGALT1, Dr. Marco Fumagalli for support with HRMS assays, and Dr. Gianluca Santoni for support with SAD X-ray data collection setup. We thank Dr. Federica Marasca and Prof. Antonella Profumo for support with ICP-MS analysis. We also would like to thank Prof. Giulia F. Mancini and Prof. M. Freccero for inspiring discussions during manuscript revision. We thank Prof. Andrea Sinz for providing access to the Center for Structural Mass Spectrometry at Martin Luther University Halle-Wittenberg, Halle (Saale), Germany. We thank the European Synchrotron Radiation Facility (ESRF) and the Swiss Light Source (SLS) for the provision of synchrotron radiation facilities. We thank Centro Grandi Strumenti (University of Pavia) for the provision of HRMS and NS EM instrumentation. This work was supported by the Italian Association for Cancer Research (AIRC, Grants MFAG 20075 and Bridge 27004 to F.F.), the Mizutani Foundation for Glycoscience (Grant 200039 to F.F.), the Ehlers-Danlos Society (Rarer Types EDS Grant 2022 to F.F.), the Giovanni Armenise-Harvard Foundation (CDA 2013 to F.F.), the NextGenerationEU-MUR PNRR program through Italian Ministry for University and Research (grant PRIN PNRR 2022 P20224WAME to C.I. and F.F.), the University of Pavia (InROAD fellowship to L.S.). The Forneris Lab also acknowledges support from EU funding within the NextGenerationEU-MUR PNRR Extended Partnership initiative on Emerging Infectious Diseases (Project no. PE00000007, INF-ACT), and by the Italian Ministry of Health (Piano Operativo Salute, IMMUNO-HUB). Some of the instrumentation used for this research was acquired through funding by Regione Lombardia, regional law n° 9/2020, resolution n° 3776/2020. None of the funding sources had roles in study design, collection, analysis, and interpretation of data, in the writing of the report and in the decision to submit this article for publication.
Author information
Authors and Affiliations
Contributions
F.F. conceived the project and designed research. M.D.M., L.S. and S.R.R. produced recombinant GLT25D1/COLGALT1 and performed biochemical characterizations, with support from D.M. A.P. generated all mutants. L.S. crystallized GLT25D1/COLGALT1. F.F. and L.S. solved the structure and carried out structural refinements, with support from M.D.M. D.M. and M.D.M. performed HRMS assays. S.L. carried out mass photometry experiments and associated data analysis. A.V. and C.I. carried out XL-MS experiments and associated data analysis. E.M. and G.C. carried out molecular dynamics simulations and associated analyses. C.S. performed Native-MS experiments for the identification of cofactors bound in the GT1 domain. M.D.M., S.R.R., L.S. and F.F. analyzed the data, prepared the figures and wrote the paper, with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
De Marco, M., Rai, S.R., Scietti, L. et al. Molecular structure and enzymatic mechanism of the human collagen hydroxylysine galactosyltransferase GLT25D1/COLGALT1. Nat Commun 16, 3624 (2025). https://doi.org/10.1038/s41467-025-59017-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-59017-5
This article is cited by
-
Molecular basis of collagen galactosylation by GLT25D1
Nature Communications (2026)
-
Enzymatic craftsmanship in collagen glycosylation
Nature Communications (2025)