Abstract
Gene expression during cell development and differentiation is orchestrated by distal regulatory elements that precisely modulate cell selective gene activity. Gene therapy vectors leverage these elements for precise spatiotemporal transgene expression. Here, we develop a one-shot approach to screen candidate regulatory sequences from large-scale epigenomics data for programmable transgene expression within gene therapy viral vectors. We assess a library of 15,000 short sequences derived from developmentally active elements during erythropoiesis using a clinically relevant reporter vector. These elements display a gradient of transcriptional enhancer activity in erythroid cells, with high cell type restriction and developmental stage specificity. Finally, replacing the canonical β-globin μLCR with a compact enhancer in a β-thalassemia lentiviral vector successfully corrects the thalassemic phenotype in patient-derived hematopoietic and stem and progenitor cells (HSPCs), while increasing viral titers and cell transducibility. Our approach provides further insights into enhancer biology with wider implications for human gene therapy.
Similar content being viewed by others
Introduction
Gene therapy has heralded a fundamental shift in modern medicine and has enabled curative single dose treatments for an array of diseases. The overall goal of this therapeutic modality is to achieve durable expression of a therapeutic gene that ameliorates or eliminates molecular disease symptoms. Conventional gene therapy viral vectors consist of two main components: (1) the therapeutic transgene and (2) a regulatory sequence driving the transgene expression. For several genetic or acquired diseases, gene therapy has successfully moved from proof-of–concept studies to clinical translation and commercially marketed clinical therapies1,2,3,4. Despite this promise, development of new gene therapy-based treatments has proven challenging due to transgene expression failing to reach therapeutic levels and substantial off-target effects (i.e. expression of the transgene in non-target cell types and/or ectopic activation of oncogenes near viral integration sites). Many of these limitations can be directly linked to the architecture of gene therapy vectors. A straight-forward strategy that can simultaneously amplify efficacy and safety of transgene vectors is the utilization of cell-type specific, compact regulatory sequences in the vector designs to precisely control expression of therapeutic transgenes in their proper context.
Recently, genome scale chromatin accessibility assays have identified millions of putative cis-regulatory elements5,6. The vast majority of these potential cis-regulatory elements are cell-selective and developmentally specific and, thus in principle, can be harnessed to control therapeutic transgene expression in a wide variety of contexts (i.e., location and time). Despite systematic efforts to biochemically delineate human regulatory DNA7,8,9, characterizing regulatory function (i.e., impact on gene expression) has remained challenging. While massively parallel reporter assays (MPRAs) can identify sequences with regulatory function, in many cases these assays have limited translatability because they are performed using non-integrating episomal vectors in highly proliferating malignant cell lines and not the therapeutically relevant cell type10,11,12. Moreover, when integrating vectors are employed (lentiviral), the readout of the assay is not a transgene, but the transcription of the regulatory sequence itself (located 3’ to the promoter). In this case, the architecture of the screening vector does not resemble a putative therapeutic (i.e., transgene control). Taken together, these limitations urge the development of alternative enhancer screening methods to identify and design control sequences for gene therapy vectors.
In this study, we describe a direct enhancer discovery pipeline to optimize gene therapy vectors including identification of cell type specific enhancer elements and design of a refined therapeutic vector. As a proof of concept, we apply this approach to β-hemoglobinopathies, the most common monogenic disorders worldwide and demonstrate that novel, cell type specific, compact, potent enhancers challenge the prototypic μLCR enhancer (a truncated version of the β-globin locus control region) of hemoglobinopathy vectors and enable efficient phenotypic correction.
Results
Large-scale evaluation of erythroid enhancers in a chromosomal and therapeutically relevant context
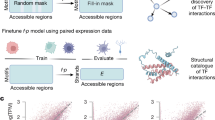
To identify enhancer elements for gene therapy applications we designed a 4-step discovery pipeline: (a) compilation of an test sequence library informed by functional genomics atlases, (b) addition of the library to a lentiviral vector relevant to clinical applications (including a cell type specific promoter and a chromatin insulator), (c) sequence selection based on direct protein readout, (d) downstream evaluation in a therapeutic vector and in patient-derived cells (Fig. 1).
A one-shot approach for screening and functionally validating erythroid specific enhancers for lineage-specific induction of transgene expression at therapeutic levels in gene therapy viral vectors. Briefly, 5393 DNase I hypersensitive sites (DHS) activated de novo during ex vivo human erythropoiesis were selected and broken down into ~3 198bp-long tiles each, comprising a library of total ~15 k elements. The tiles were then cloned into a clinically relevant GFP reporter lentiviral vector and HUDEP-2 cells were transduced at MOI (Multiplicity of infection) <1. The cells were then sorted by flow cytometry into 3 bins based on GFP expression. DNA libraries were constructed from each bin and read counts were assigned to each tile as function of GFP expression. Top performing tiles were then mapped to their full-size DHSs which were then in turn cloned into a therapeutic vector where the candidate elements were assessed based on their ability to achieve phenotypic correction of β-thalassemia patient donor derived erythroid cells. MFI: Mean Fluorescence Intensity.
Given that we aim for transcriptional enhancement in erythroid cells we sourced candidate sequences from a fine temporal accessibility mapping during adult ex vivo erythroid differentiation13. We focused on de novo accessible elements during erythropoiesis by excluding DNase I Hypersensitive Sites (DHSs) accessible at CD34+ hematopoietic stem and progenitor (HSPC) stages, in an attempt to eliminate elements potentially active in other lineages. Fully, this resulted in 5393 DHSs accessible at different stages of erythroid differentiation where 80% of the DHSs exhibit peak accessibility after day 7 of ex vivo erythroid differentiation (Supplementary Fig. 1a). These elements display strong lineage-specific accessibility as DNase I signal in non-erythroid hematopoietic cell types, including CD34+ HSPCs is markedly low (Fig. 2a). Furthermore, these elements are strongly enriched for transcription factor binding motifs of canonical drivers of erythropoiesis such as GATA1, TAL1, KLF1, and NFIX (Fig. 2b, Supplementary Data 1). In their majority, these DHSs represent distal non-coding elements either intronic or intergenic or (49.9% and 31.1%, respectively) whereas promoters are relatively under-represented (11.5%) (Fig. 2c).
a Normalized DNase I density over the 5393 DHS selected ( ± 5 kb from DHS center) across 8 human hematopoietic populations (b) Top 5 enriched motifs within the 5393 selected DHS, ranked by hypergeometric test q-value. c Frequency barchart of the genic features overlapped by the 5393 selected DHS (d) Histogram depicting the distribution of the maximum likelihood estimated effect of each tile on GPF expression. Gray dashed line represents the expected robust fitted gaussian (normal) distribution. Orange dashed lines annotate the 5 and 95 percentiles of the expected distribution. e Top enriched transcription factor binding site motifs within the potential enhancer and silencer tiles. Point size encodes the hypergeometric test q-value (-log10) and the color encodes the log2 ratio over all tiles. f Violins and boxplots depicting the distribution of chromatin accessibility by DNase I, and CUT&RUN density for GATA1, TAL1 as well as H3K27ac and H3K9me3 chromatin marks as measured in HUDEP-2 cells between potential enhancers (n = 897) and potential silencer (n = 481) tiles. Median is shown as a thick horizontal line, boxes extend to IQR (25th and 75th percentile) and whiskers extend to 1.5x IQR. Background thick gray line and gray band show median and IQR (25th and 75th percentile), respectively, of all tiles (n = 14,999). Benjamini-Hochberg corrected Wilcoxon rank sum test p-values are reported. g Enrichment of chromHMM overlapped by potential and enhancer and silencer tiles against the entire tile library. Point size encodes the binomial test q-value (-log10) and the color encodes the log2 ratio over all tiles.
To functionally screen our candidate regulatory elements (CREs), we designed a lentiviral vector closely matching those currently used in clinical therapeutics1,2,14. Our screening method deviated from traditional lentivrial MPRAs by employing a modified FACS based lenti-MPRA approach where the evaluation of the CREs was based on their effect on protein expression (Fig. 1). The enhancer screening library was generated by tiling each of the 5393 erythroid DHSs (median size 233 bp, Supplementary Fig. 1b) into overlapping oligos, resulting in a library of ~15 k (14,999) DHS fragments with a median of 3 tiles per DHS and a median size of 198 bp per tile (Supplementary Fig. 1c, d). The DHS-oligo library was cloned upstream of a minimal (169 bp) human β-globin promoter driving GFP expression (Fig. 1). To avoid positional effects driven by random vector integration we included a recently identified, dual functioning (enhancer-blocker and barrier) human chromatin insulator into our lentiviral vector which flanked our expression cassette15. To maintain the cell context as relevant to clinical translation as possible, we performed the screening in HUDEP-2 cells, a CD34+ cell derived human erythroid progenitor cell line. In contrast to another popular model of erythroid cells, the K562 erythroleukemia cell line, HUDEP-2 cells more faithfully recapitulate the chromatin accessibility profile of erythroid progenitors as they correlate better with day 7 ex vivo-derived adult erythroid progenitors, compared to K562 (Pearson’s r: 0.515 and 0.219, respectively) (Supplementary Fig. 2). Critically, we performed all HUDEP-2 transductions at multiplicity of infection (MOI) of 0.4 to achieve a theoretical maximum of one viral integration per cell.
In vectors containing only the promoter, GFP expression was low, and the overall population was largely overlapping with the untransduced/non-GFP expressing cells (Supplementary Fig. 3a). Five days post transduction we FACS-sorted the transduced cells selecting three equiproportional population bins (5% of total live singlets, each) across the GFP-positive fraction namely, GFP low, medium, and high (Supplementary Fig. 3b). We then sequenced the integrated lentiviral cassette and computed the relative enhancer tile frequencies in each GFP bin (Fig. 1, Methods). Because expression of integrated lentiviral cassettes is impacted by local genomic and epigenomic context (i.e. position-effect variegation)16, we assessed the number of unique integrations required for each element to yield reliable expression estimates. We found that for a library of our size (~15,000 sequences), ~800 integrations per element were need to achieve robust replicate concordance (r > 0.9), and that lower number of integrations compromised replicate concordance (100X r = 0.347; and 200X r = 0.423) (Supplementary Fig. 4), highlighting that lentiviral MPRA approaches are critically dependent on the number of unique insertions to overcome positional variegation effects16.
At a library coverage of 800X, we recovered the majority of the original enhancer tiles (14,668 fragments in total, 97.8% of the designed library). To estimate the effect of each fragment on altering GFP intensity we employed a statistical framework that estimates the latent effect of each sequence on expression by modeling the tile frequencies in each GFP bin through maximum likelihood (adapted from De Boer et al.17). Using this approach, we ranked each enhancer tile based on its estimated effect value, totally resulting in 6577 elements with positive and 8,091 elements with negative effect (Fig. 2d). The latter suggests that a large number of DHSs activating during erythroid differentiation may function as repressors.
Comparing the observed tile effect distribution to an expected normal distribution given the observed median and mean absolute deviation, we identified 481 elements that lie below the 5% (potential silencers) and 897 elements above the 95% (potential enhancers) (Fig. 2d, Supplementary Data 2). Comparing these two subsets we discovered that enhancer tiles are enriched for DNA binding motifs of key erythroid transcription factors such as GATA1 and TAL1 (hypergeometric test q-value < 1e-03), compared to the entire 15 K tile library. In contrast, silencing tiles are depleted of these motifs (hypergeometric test q-value < 1e-03) while they are strongly enriched for motifs of the SP transcription factor family (Fig. 2e, Supplementary Data 3). Moreover, we find enhancer tiles to exhibit significantly increased DNase I accessibility (Wilcoxon rank-sum test adjusted p-value 3.064e-4), binding of GATA1 and TAL1 (Wilcoxon rank-sum test p-value < 2.22e-16, each) and histone post-translational modifications canonically associated with enhancing cis-regulatory elements H3K27Ac and H3K4me1 (Wilcoxon rank-sum test adjusted p-value 6.851e-37 and 5.859e-25, respectively), over the silencing tiles (Fig. 2f). In line with these observations, characterization of chromatin states by composite mark profiles as discovered by chromHMM18 (Supplementary Fig. 5) revealed that enhancer tiles are enriched for regions with chromatin states characteristic of active erythroid enhancers (log2 ratio over tiles 0.443, binomial test adjusted p-value 2.515e-30) (Fig. 2g). Notably, we found silencing tiles to show strong enrichment for heterochromatin states, albeit with marginal statistical significance (log2 ratio over all tiles: 2.923, binomial test adjusted p-value: 0.085).
Given that each tile only represents a part of the DHS which they are derived from, we set to assess how the full-length DHSs, as the functional unit, correspond to tiles in terms of enhancer activity. To this end, we first selected a subset of 205 tiles, enriched in potential enhancers (117/205). Then, for each element in this mini-library we synthesized a new sequence element corresponding to the encompassing full-length DHS for each tile (Supplementary Fig. 6a–c, Supplementary Data 4). Following the same screening approach we applied to the original library, we observed a net positive shift in the GFP-MFI using the mini-library compared to the 15k-element library (Supplementary Fig. 6d). Finally, we found the effect on GFP expression of the strongest tile from each DHS to be well-correlated (r = 0.509) to the effect of the full-length DHS (Supplementary Fig. 6e), indicating that for many DHS elements, a compact sub-sequence encodes the majority of its regulatory function.
Identification of enhancers with lineage-selective and tuned developmental activity
Proper cell lineage and temporal activity of transgenes is paramount to safe and effective gene therapy vectors. To test for lineage specificity and developmental temporal activity of these sequences we individually characterized a subset of 40 candidate sequences from the full-length DHS mini-library (Supplementary Data 5). Each one of these sequences was individually cloned (in triplicate) in the same screening vector and finally tested their enhancer activity against the core elements of the β-globin LCR (HS2, HS3 and HS4), and a no-enhancer vector post transduction into HUDEP-2 cells. We found that 38 DHSs (95%) had a marked increase in GFP expression vs. the no enhancer control vector, with 27 of these exhibiting statistical significance (Fig. 3a). Critically, the expression intensity values from the individual vectors were well correlated with the activities measured from the mini-library (Pearson’s r = 0.72, Supplementary Fig. 7).
a GFP expression (log10 MFI) of the 40 DHSs individually cloned in the same backbone vector and transduced in HUDEP2 cells at a MOI < 1. Full length β-globin HS2, HS3 and HS4 were used as positive controls and a no-enhancer vector (gray band) as negative control. Mean ± SD is shown. Points are colored by T-test Benjamini-Hochberg adjusted p-value. Transductions were performed in three independent experiments and time points. b Correlation between the GFP intensity (log10 MFI) of the 40 selected elements after transduction in HUDEP-2 cells (x-axis) and in the K562 cell line (y-axis). Beta-globin HS2 and HS3 are annotated. Pearson’s r correlation coefficient is shown (c) Correlation between the GFP intensity (log10 MFI) of the 40 selected elements after transduction in HUDEP-2 cells (x-axis) and in CD34+ differentiated erythroid cells (y-axis). Pearson’s r correlation coefficient is shown. d Each of the 40 identified enhancers is individually transduced into CD34+ cells and each pool (n = 42) is subjected to ex vivo erythroid (Ery), megakaryocytic (Mk) and granulocytic-monocytic (GM) differentiation. After 7 days of differentiation the percentage of GFP+ cells are determined by flow cytometry. Median is shown as a thick horizontal line, boxes extend to IQR and whiskers extend to 1.5x IQR. e Mobilized peripheral blood CD34+ cells from healthy donors were transduced with a lentiviral library of the identified 40 vectors. The cells were transplanted into NBSGW mice and bone marrow was collected 16 weeks post transplantation. Percent of GFP+ cells was assessed by flow cytometry in all engrafted human hematopoietic lineages. Bars extend to mean of n = 3 ± SEM. MFI: Mean Fluorescence Intensity. Ery: Erythroid. Mk: Megakaryocytes. GM: Granulocyte-Monocyte. Source data for all relevant panels are provided within the Source Data file.
We then performed parallel experiments in both adult human mobilized CD34+ HSPC derived erythroid cells, which represent the target cell type for ex vivo human gene therapy, as well as the erythroleukemia K562 cell line, which has long served as a popular erythroid cell model. Notably, the activity of the enhancers in K562 cells was poorly correlated to HUDEP-2 cells (Pearson’s r = 0.17) (Fig. 3b). By contrast, the same enhancer constructs transduced into CD34+ HSPCs followed by in vitro erythroid differentiation (7 days, erythroblast stage) displayed activities that track well with that of HUDEP-2 (Pearson’s r = 0.76) (Fig. 3d). This result emphasizes the importance of cell type context in screen assays for the functional interrogation of regulatory DNA sequences.
As mentioned above, all tested DHSs were selected based on their increased accessibility in adult erythroid cells. However, this alone does not preclude any potential activity of these elements in other hematopoietic cell lineages. This is particularly relevant to the development of gene therapy products where transgenes integrating near proto-oncogenes have the potential to drive malignant transformations due to lineage-promiscuous enhancer elements. To assess to what degree the activity of these enhancers is restricted to the erythroid lineage, we individually transduced adult human mobilized CD34+ HSPCs with each of the aforementioned 40 vectors and allowed the cells to ex vivo differentiate towards the erythroid, myeloid (granulocytic/monocytic) and megakaryoctyic lineages (Fig. 3d and Methods). This revealed that these elements display robust erythroid GFP enhancing activity as measured by the fraction of GFP+ cells in the total population (median 11.7, IQR 7.658-19.925), while activity in the closely related megakaryocytic and the granulocytic/monocytic lineage remained minimal (median 1.53 and 1.645, respectively and IQR 0.803-2.82 and 1.085-2.608, respectively). Of all tested vectors only two were observed as outliers in the non-erythroid cultures. Expectedly, the control PGK-GFP vector displayed uniform activity across all lineages. Additionally, an element proximal to SKA1 gene displayed also activity across all lineages, despite transcription being driven by an erythroid promoter. To further explore the promoter-enhancer dynamics, we performed additional assays where three of the erythroid enhancers (PPARA, GATA1 and the positive control HS2) and the non-specific SKA1 enhancer were individually cloned upstream a non-erythroid promoter (minimal Thymidine kinase promoter). These vectors were tested alongside for their GFP expression in erythroid and non-erythroid cells (Supplementary fig. 8). Interestingly the erythroid enhancers showed a statistically significant enhanced GFP expression in erythroid cells compared to the non-erythroid population while the non-specific SKA1 enhancer performed similarly between the two cell lineages, irrespectively of the promoter in use emphasizing the pivotal function of regulatory sequences in determining cell specificity (Supplementary fig. 8).
To expand these findings across all hematopoietic lineages simultaneously and in vivo, we performed xenotransplantation experiments where mobilized adult human CD34+ cells were transduced with a pool of the described vectors. The transduced cells were transplanted in the NBSGW mouse model which efficiently supports multilineage human hematopoiesis19. Measurements at 16 weeks post-transplantation revealed robust GFP expression restricted to human hCD45+/CD235a+ erythroid progenitors (70% GFP+) and hCD45+/CD33+ myeloid progenitors (38% GFP+), with negligible levels of GFP detected within the human non-erythroid compartments (Fig. 3f). Cumulatively, individual evaluation of the candidate sequences revealed graded activity, enabling the precise modulation of transgene expression, in both amplitude and lineage specificity.
Enhancer-driven transgene activity parallels temporal dynamics of chromatin accessibility
The DHS elements included in the enhancer library were selected for their dynamic accessibility profile in the course of ex vivo erythroid differentiation13. We investigated whether these elements regulatory capacity (i.e., effect on transcription) paralleled the accessibility profile within their native genomic context. We transduced individual pools of CD34+ cells, each with one of the 40 selected enhancer elements and subjected them to ex vivo erythroid differentiation, and for each culture we measured GFP MFI as a proxy for enhancer function at regular intervals during the differentiation (Fig. 4a). Comparing the GFP MFI with the accessibility of their corresponding DHS during differentiation we found a strong correlation between the two across all elements tested similar to the HS2 and HS3 of the beta globin LCR (median r = 0.79) (Fig. 4b, Supplementary Fig. 9) strongly indicating that temporal enhancer functionality is maintained outside of their native genomic context and is solely encoded by the local sequence ( ~ 200 bp) of these elements.
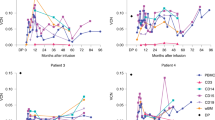
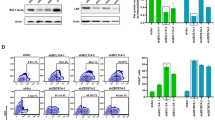
a Enhancer activity kinetics of the 42 elements as measured by GFP expression with flow cytometry during ex vivo erythroid differentiation from mobilized adult human CD34+ cells. Y-axis depicts MFI z-score across all elements within each time point measured. An early active (L196, teal) and a late active (L223, magenta) element were selected for further study. b Correlation (Pearson’s r) between the temporal profiles of enhancer activity (MFI) of the 42 elements and their in situ DNase I accessibility profile during ex vivo erythropoiesis. Median Pearson’s r is denoted with a dashed red line. c Comparison between the enhancer activity (MFI, light color) and in situ DNase I accessibility (dark color) for the L196 vector (left) and L223 (right). d The enhancer element in the L196 vector is derived from a DHS intronic to the PVT1 lncRNA and the element in L223 is an exonic DHS in the PPARA locus. Signal density tracks of the DNase I accessibility during erythroid differentiation, GATA1 CUT&RUN occupancy and H3K27Ac CUT&RUN enrichment in HUDEP-2 cells is shown for the PVT1 locus (left) and PPARA locus (right). Bottom track shows consensus footprinted motifs overlapping with each element. e Genetic deletion experiments of PVT1 DHS and PPARA DHS in HUDEP-2 cells result in significant repression of PVT1 and PPARA expression, respectively. Browser tracks of the DNase I accessibility in WT HUDEP-2 and mutant (ΔHS) are shown. Barplots show the mean ± SE of normalized gene counts from n = 4 experiments. Source data for all relevant panels are provided within the Source Data file.
Given that the enhancers display a precise temporal activity we interrogated the in situ functionality of strongest enhancers: L196 (early activation) and L223 (terminal activation) (Fig. 4c). Surprisingly, despite their strong enhancer function in both HUDEP-2 and ex vivo differentiated adult HSPCs, the two elements display weak H3K27ac levels (Fig. 4d), a histone modification typically associated with active enhancers. Despite the relatively low enrichment of H3K27ac, homozygous knockout these elements (Supplementary Fig. 10a) resulted in strong, reproducible transcriptional repression of their target genes (PVT1 for L196 and PPARA for L223) (Fig. 4e and Supplementary Fig. 10b). These results strongly indicate that these enhancers mirror their developmental activity in an engineered ectopically integrated vector and can therefore be used for temporal control of transgene expression in a gene therapy setting.
Direct preclinical translation of the novel enhancers
Because these enhancers drive robust gene expression, and their activity reflects observed developmental dynamics, we sought to apply them directly to a therapeutic vector for β-hemoglobinopathies as a translational paradigm. Here, we focused on the BCL11A-shmiRNA transgene approach. First, BCL11A knock down leads to HbF reactivation which is easily measurable in both healthy and diseased human erythroid cells. Secondly, BCL11A knock out in any other than the erythroid lineage would have an immediate negative effect in cells’ viability, proliferation, engraftment and differentiation capacity in vivo. Therefore, the shmiRNA vectors incorporating the optimal erythroid enhancers would permit us to simultaneously test their lineage-specificity and by extension, safety as well as their efficiency. A BCL11A-shmiRNA14,20,21 was cloned in the same backbone as the screening vector (Fig. 5a). Of note, both the PPARA–shmiRNA and PVT1–shmiRNA vectors showed a significant increase in viral production compared to the µLCR vector (Fig. 5b), likely due to the small size of the new enhancers (10% of the clinically utilized µLCR) (Fig. 5a). Subsequently, all shmiRNA vectors were transduced in HUDEP-2 cells, to evaluate HbF reactivation in a cell line with a low background of γ-globin expression. The small size of the new enhancers did not reduce their therapeutic effect as they were both able to achieve γ-globin reactivation in transduced cells at the same levels as the μLCR vector (Fig. 5c). HbF expression in untransduced cells (GFP–) was maintained at low levels, as expected. Notably, transduction with the new vectors did not significantly alter cell proliferation of neither HUDEP-2 cells (ANOVA F-value 2.135; p-value 0.104) nor primary CD34+ cells (ANOVA F-value 8.024e-3; p-value 0.999) transduced at a stable MOI, compared to untransduced cells or cells transduced with the μLCR-shmiRNA vector (Supplementary Fig. 11a, b). Subsequently, we transduced CD34+ cells from healthy donors with the above vectors with serial dilutions of the viruses which showed that the significantly larger μLCR-shmiRNA vector quickly reaches a plateau in HSPCs transduction ability (Supplementary Fig. 12a, b). HbF expression within the transduced populations was similar or higher using the new vectors compared to the μLCR-shmiRNA, with PVT1-shmiRNA achieving overall the highest percentage among the three groups (Supplementary Fig. 12c), while the differences in HbF expression were not dependent to the MOI used (Supplementary Fig. 12d).
a Schematic representation of the vector design showing the difference in size of the new enhancers compared to the μLCR. C1: chromatin insulator, βp: minimal β-globin promoter. b Titration of the PPARA, PVT1 and μLCR BCL11A-shmiRNA vectors in K562 cells based on qPCR. Bonferroni corrected Welch t-test values are shown. n = 10 virus productions in all groups. Median is shown as a thick horizontal line, boxes extend to IQR (25th and 75th percentile) and whiskers extend to 1.5x IQR. c HbF expression (%, mean ± SEM) within the GFP- and GFP+ populations assayed by flow cytometry in transduced HUDEP-2 cells with one of the PPARA (n = 5), PVT1 (n = 7), and μLCR (n = 7) BCL11A-shmiRNA vectors. Transductions in HUDEP-2 cells were performed in independent experiments. d Workflow of the in vivo experiments. Mobilized peripheral blood CD34+ cells from healthy donors were transduced at the same MOI with the μLCR and PVT1-shmiRNA. Mock-transduced cells from the same donor were used as control. Two days post transduction an equal number of cells was transplanted in NBSGW mice. Mice were sacrificed 16 weeks post transplantation. e Human chimerism in the bone marrow (BM) of the transplanted mice. hCD33, hCD19, hCD3, hCD34, hCD41 subpopulations were calculated within the hCD45 population (as % cells, mean ± SEM shown). f GFP expression (as %GFP+ cells, mean ± SEM shown) by flow cytometry in erythroid (hCD235a+) and non-erythroid (hCD45+) engrafted cells. Tukey HSD post-hoc test adjusted p-values are shown. g In vivo HbF expression (as % HbF+ cells, mean ± SEM) identified by flow cytometry in engrafted human erythroid (hCD235a+) cells at the time of sacrifice. In all in vivo experiments shown n = 5 mice per experimental group were used. Bonferroni corrected Welch t-test p-values shown. Non-significant (ns) ≥ 0.1. Source data for all relevant panels are provided within the Source Data file.
It is known that non-erythroid BCL11A suppression can be detrimental in cell engraftment, maintenance and multilineage differentiation22,23. To assess the compact vectors’ performance in vivo, both in terms of efficacy and safety, we transduced at the same MOI normal mobilized peripheral blood derived CD34+ cells with the PVT1-shmiRNA vector and compared its effect with that of the μLCR-shmiRNA vector in a xenotransplantation mouse model (Fig. 5d). Mock transduced cells were used as a control. The mice were sacrificed 16 weeks post transplantation and bone marrow was analyzed for multilineage reconstitution and HbF expression (within the erythroid lineage). Importantly, multilineage reconstitution was observed in all mice and was undistinguishable (ANOVA F-value 1.009; p-value 0.369) between the transduced and mock-transduced cells (Fig. 5e). In line with previous experiments, differences in transduction levels were lineage specific (population-vector interaction ANOVA F-value 431.352; p-value 5.34e–13) as GFP expression was restricted to the erythroid compartment. Within the hCD235a+ populations vector effects were pronounced in contrast to total white blood cells (hCD45 + ) where overall %GFP+ cells was non detectable. Despite the constant transduction MOI, cells transduced with the PVT1 vector achieved significantly higher percentage of GFP+ cells within the erythroid compartment (Fig. 5f) compared to the group transduced with the µLCR vector (Post-hoc Tukey HSD test p-value 9.06e–04). The superior transduction/transgene expression rate was also translated to a significantly higher overall percentage of HbF+ cells in the PVT1 group compared to the μLCR (Fig. 5g).
Ex vivo therapeutic correction of β-thalassemia major in primary, patient-derived HSPCs
Given the results from transduced healthy HSPCs we next tested our enhancers in cells derived from patients with β-thalassemia major. Expanded HSPCs derived from two mobilization clinical trials24,25 were transduced with the PVT1 and μLCR-shmiRNA vectors (Methods) and cultured in erythroid medium for 18 days (terminal differentiation). Untransduced cells were used as control. Monitoring the growth of cells post transduction with the µLCR-shmiRNA and the PVT1-shmiRNA we observed significant effects over the untransduced cells (ANOVA F-value 5.423; p-value 5.54e–03). Specifically, transduction with PVT1-shmiRNA vector conferred a marked increase in proliferative ability over µLCR-shmiRNA (slope 1.001 and 0.841, respectively), where by the end of differentiation PVT1-shmiRNA transduced cells resulted in a > 3-fold net increase in total erythroid cells over untransduced (Bonferroni adjusted Wilcoxon rank sum test p-value 0.035) (Fig. 6a). HbF+ cell frequency in the untransduced population was surprisingly high, which is compatible with the increased fitness of cells with elevated HbF levels. However, despite the high background, HbF expression in both transduced groups was significantly higher in both the enucleated (ANOVA F-value 4.285; p-value 0.0394) and nucleated (ANOVA F-value 11.29; p-value 1.75e–03) populations compared to the untransduced cells. Use of PVT1 enhancer displayed consistently robust HbF and superior induction in both populations over µLCR, while HbF increase with µLCR was either insignificant (nucleated) or marginally significant (enucleated) over untransduced (Fig. 6b). Similarly, although transduction with vectors utilizing either enhancer increased production of both Aγ and Gγ chains, use of PVT1 successfully restores α to β-like chain imbalance to 1:1 ratio and consistently yields overall better results over untransduced (Fig. 6c, d).
Mobilized peripheral blood CD34+ cells from 5 patients with beta-thalassemia major were transduced with the PVT1 and μLCR-shmiRNA vectors in the presence of cytokines and small molecules. The cells were cultured under erythroid conditions for 18 days, 2 days post transduction. a Absolute cell numbers (mean ± SEM; n = 5) during erythroid differentiation of transduced and untransduced cells. Bonferroni corrected Wilcoxon rank sum test p-values are shown. b HbF expression (as %HbF+ cells) measured by flow cytometry in enucleated and nucleated cells on day 18 of ex vivo erythroid differentiation. Tukey HSD post-hoc test adjusted p-values shown. c Globin chain production displayed as γ to β-like, γ to α, and β-like to α ratio in transduced and untransduced cells measured by HPLC. Tukey adjusted post-hoc test with Welch correction p-values shown. d HPLC trace profiles of globin chains in transduced and untransduced β0/β+ patient CD34+ derived erythroid cells. e Percentage of enucleated cells on day 18 of the differentiation measured by flow cytometry. Bonferroni corrected Welch t-test p-values are shown. f Flow cytometry measured intensity of reactive oxygen species (ROS) post CFSE staining of erythroid cells on day 11 of the erythroid culture. Post-hoc Tukey HSD test adjusted p-values shown. g Characteristic morphology of transduced and untransduced erythroid cells in different erythroid differentiation and maturation stages. Non-significant (ns) ≥0.1. Boxplots extend to IQR (25th and 75th percentile). Median is shown as a thick horizontal line and whiskers extend to 1.5x IQR. Untr.: Untransduced control. Source data for all relevant panels are provided within the Source Data file.
This increase resulted in improved enucleation of the transduced over the untransduced cells (Fig. 6e), more pronounced in the PVT1-shmiRNA-group compared to μLCR-shmiRNA-group, (Post-hoc Tukey HSD test adjusted p-value 2.035e–04 and 3.713e–03, respectively) that was accompanied by a marked improvement of the thalassemic phenotype in transduced populations (Fig. 6e). We then analyzed reactive oxygen species (ROS) levels after transduction, a hallmark of oxidative stress in thalassemia caused by either increased autoxidation of α-globin chains or by increased iron levels. Overall, transduced cells achieved lower ROS levels compared to untransduced (One-way ANOVA F-value 6.316; p-value 0.0526). Although the difference in ROS levels between PVT1-shmiRNA and uLCR-shmiRNA vectors was not significant, transduction with the PVT1-shmiRNA-vector resulted in overall lower levels of reactive oxygen species (ROS) compared to uLCR (median 6.651e + 03 and 14.207e + 03, respectively) (Fig. 6f). Finally, transduced cells exhibit higher numbers of maturing erythroid cells at the end of EC, in contrast to an erythroid maturation blockade in the untransduced population (Fig. 6g).
Overall, our results demonstrate that vectors incorporating these compact, potent, and erythroid-specific enhancers can overcome long-standing limitations of viral vectors. Moreover, these enhancers effectively replace the β-globin μLCR, which has long been considered the gold standard in globin vector development. Specifically, results in both healthy donor and thalassemic donor derived CD34+ cells indicate that while there is no direct evidence that fetal hemoglobin induction is increased by a further enhancement of transgene expression, the differences in enhancer size lead to a higher number of transduced cells enhancing the overall therapeutic effect.
Discussion
Gene therapy has recently met great success for a variety of monogenic hereditary diseases, cancer, and neurodegenerative disorders and holds great promise in becoming a standard durable clinical option for many disorders of genetic or epigenetic background. Nonetheless, current vector designs suffer by suboptimal safety and efficacy in terms of tissue specificity, transgene expression levels, and viral titers26. In vivo gene therapy in particular, requires both tissue specificity and tightly controlled gene expression as the use of strong promoter-enhancer pairs can result in tissue toxicity due to transgene overexpression27,28,29. In the context of ex vivo gene therapy where high or even supranormal therapeutic protein levels are required, transgene expression is driven by strong promoter2,30 or enhancer sequences31,32. Unfortunately, a lentiviral vector equipped with the constitutive MNDU3 promoter developed for cerebral adrenoleukodystrophy (CALD) gene therapy lead to the first recorded genotoxic events as seven out of 67 treated patients developed myelodysplastic syndrome (MDS), AML33. In addition to adequate transgene expression levels, successful gene therapy protocols mandate high transducibility and efficient viral production. Hemoglobinopathies, a popular disease model for gene therapy has a history of vector development for more than 30 years and a recently approved product for β-thalassemia. The incorporation of the μLCR enhancer has enabled both therapeutic transgene expression levels and erythroid-specific expression patterns, representing the sine qua non in globin vector design1,4. However, its large size ( ~ 3 kb) yields low viral titers rendering vector production suboptimal and highly expensive in addition to impeding efficient cell transducibility34,35.
Here, we introduce a platform for large-scale discovery of short and potent enhancer elements with precise spatiotemporal activity, enabling the development of optimized viral vectors towards efficient, safe, and cost-effective gene therapy applications. Capitalizing on our previous efforts of systematic mapping and characterization of the cis regulatory DNA elements involved in human erythropoiesis13, we designed a flow cytometry-based screening strategy using a lentiviral vector that allowed us to assess the transcriptional enhancer activity of 15,000 short sequences.
Although large scale lentiviral screening strategies have also been developed by others36, these approaches largely rely on highly-proliferative cell lines36,37,38,39,40,41 and/or non-human models42,43,44,45. It is now well understood that the activity of cis-regulatory elements is determined by their sequence composition, genomic context, and the availability of trans-acting proteins (transcription factors) that actuate them. In contrast to the prevailing MPRA approaches, we evaluated potential cis-regulatory elements in a relevant cell type and therapeutic vector, enabling direct clinical translation. We demonstrate that enhancer function in HUDEP-2, but not in K562 cells, corresponded with high fidelity to their performance in adult CD34+-derived erythroid progenitors, highlighting the importance of using an appropriate cell context for evaluation.
We show that these validated erythroid-specific enhancer elements display precise spatiotemporal activity patterns both in vitro and in vivo as their activity is restricted to the erythroid lineage and further confined to specific stages of erythroid maturation. In the case of hemoglobinopathies current gene therapy paradigms entail the ex vivo transduction of a patient’s hematopoietic stem and progenitor cells which maintain multilineage differentiation capacity. Therefore, a constitutive promoter-enhancer pair would lead to ectopic pan-hematopoietic expression with unknown effects. Indeed, it was recently shown that the HS2 component of the μLCR enhancer has promiscuous cell type activity and thus only partial lineage specific LCR activity35. As such, we considered lineage specificity of these elements as an additional safety layer for these vectors.
In order to explore the potency of a μLCR–free vector design, we designed a clinically relevant vector featuring a therapeutic transgene, a truncated β-globin promoter, novel erythroid-specific enhancer elements and the C1 chromatin insulator15. We show that replacement of the μLCR by the PVT1 enhancer element resulted in overall comparable potency in terms of HbF + % cells, γ-globin output, oxidative stress reduction, erythroid cell enucleation and phenotypic correction. Importantly, owing to its small size ( ~ 9-fold difference compared to the μLCR), we were able to achieve up to 5-fold increase in lentiviral particle yield in addition to significantly increased transducibility of HSPCs. These features are key to enabling the clinical-scale, GMP-grade vector production at lower MOIs and the successful transgene delivery which in turn is expected to reduce the overall commercial cost and improve patient’s accessibility46. Moreover, the compact size of the new enhancers allows for the incorporation of additional features, supporting various facets of treatment, such as multiple therapeutic transgenes47, selection or expansion cassettes48, and safety elements.
In summary, we present an integrated large-scale FACS- and lentiviral-based screening assay informed by gene regulatory atlases for the discovery of short, cell-type selective transcriptional enhancers with a broad range of amplitude. These enhancer elements can be specifically optimized for the design of safer, more efficient and lower cost production gene therapy vectors.
Methods
Ethics
This study was conducted in compliance with all relevant ethical regulations and was approved by the ethics committees of the University of Washington in Seattle, the “George Papanikolaou” Hospital in Thessaloniki and Aristotle University of Thessaloniki.
Cell culture of immortalized cell lines
K562 cells were maintained at a density of 2.5 × 105 per mL in RPMI-1640 (Thermo Fisher Scientific) with 10% heat-inactivated Fetal Bovine Serum (Thermo Fisher Scientific) and 100 units/mL penicillin (Gibco) and 0.1 mg/mL Streptomycin (Gibco). Polybrene (8 ug/ml, Sigma) was added to the culture during lentiviral transduction.
HUDEP-2 cells were cultured in optimal expansion conditions49. During their expansion phase the cells were cultured in StemSpan H3000 medium (Stem Cell Technologies) supplemented with 100 ng/mL recombinant human stem-cell factor (rhSCF, Peprotech), 3 IU/mL recombinant human erythropoietin (rhEPO, Peprotech), 1 μM dexamethasone (Sigma), 1 μg/mL doxycycline (Sigma), and 100 units/mL penicillin (Gibco) and 0.1 mg/ml Streptomycin (Gibco). Polybrene (8 ug/ml, Sigma) was also added to the culture during lentiviral transduction. During differentiation, HUDEP-2 were maintained in Iscove’s modified Dulbecco’s medium (IMDM) supplemented with 330 μg/mL human holo-transferrin (Sigma), 10 μg/mL recombinant human insulin (Sigma), 2 IU/mL heparin (Sigma), 5% human AB plasma (obtained from Bloodworks Northwest, Seattle, WA), 3 IU/mL rhEPO (Peprotech), 100 ng/mL rhSCF (Peprotech), 1 μg/mL doxycycline (Sigma), and 100 units/mL penicillin (Gibco) and 0.1 mg/mL Streptomycin (Gibco).
Cell culture of primary CD34+ cells
Human CD34+ HSPCs from mobilized peripheral blood of deidentified healthy donors were obtained from Fred Hutchinson Cancer Research Center, Seattle, Washington, where all donors provided written informed consent under an IRB-approved protocol for the use of their samples in research. CD34+ cells from thalassemic patients were previously collected during two mobilization clinical trials (Thal-001, Thal-002) conducted at G.Papanikolaou Hospital, Thessaloniki, Greece24,25. All thalassemic donors provided written informed consent at the time of enrollment, and the use of these samples for research was approved by the relevant institutional review board. CD34+ HSPCs from mobilized donors were maintained in StemSpan H3000 with penicillin/streptomycin, the following cytokine cocktail: recombinant human Flt3 ligand (hFlt3, Peprotech), recombinant human thrombopoietin (rhTPO, Peprotech), and rhSCF (CC110, Stemcell Technologies) and in the presence of the small molecules StemRegenin1 (1 μΜ) (Cellagen Technology) and Ly2228820 (100 nM) (Selleckchem)50. For lentiviral transduction the cells were plated on retronectin coated plates (Takara Bio) and transduced in the presence of 8 μg/ml protamine sulfate (Sigma). Erythroid differentiation of CD34+ cells was carried out in 3 stages using an established protocol51. During Stage I the cells were expanded in the presence of 1μΜ Hydrocortisone, 5 ng/mL recombinant human IL-3 (rhIL-3, Peprotech), 100 ng/mL rhSCF (Peprotech) and 3IU/mL rhEPO (Peprotech), during stage II only in the presence of rhSCF (Peprotech) and rhEPO (Peprotech) and in the final stage III, rhSCF was omitted. Granulocytic/monocytic differentiation of CD34+ cells was achieved by culturing the cells in the same basal medium used for the ex vivo erythroid differentiation, with a different cytokine cocktail: 50 ng/mL rhSCF (Peprotech), 5 ng/mL rhIL-3 (Peprotech), 10 ng/mL rhGM-CSF (Peprotech), 20 ng/ mL rhG-CSF (Peprotech)52. For megakaryocytic differentiation CD34+ HSPCs were differentiated to the megakaryocyte lineage by culturing for 11 days in IMDM based media containing 30 ng/mL rhTPO (PeproTech), 1 ng/mL rhSCF (PeproTech), 7.5 ng/mL rhIL-6 (PeproTech), 13.5 ng/mL rhIL-9 (PeproTech), 20% BIT (StemCell Technologies), 40 µg/mL LDL (Millipore-Sigma), 0.05 mM beta mercaptoethanol (Millipore-Sigma), and 100 units/ml penicillin (Gibco) and 0.1 mg/ml Streptomycin (Gibco).
Selection of candidate DHS elements and DHS tiling strategy
The initial set of 11,805 DHS elements with dynamic accessibility during human ex vivo erythropoiesis as identified previously13 was filtered to exclude elements with >30 normalized DNase I counts per element at day 0 (undifferentiated CD34+ hematopoietic stem and progenitor cells). For each of the resulted 5393 DHS we designed multiple 198bp-long tiles that span the entire DHS with variable overlap between any two adjacent tiles (50 bp minimum) ensuring that one tile aligns with the center of the DHS. DHS smaller than 198 bp would be represented by a single fragment equal to the DHS size, while larger DHS were tiled by at least 3 fragments. This resulted in a set of 15,000 tile sequences.
Enhancer lentiviral-library construction and screening
A pool of 15,000 tiles was first chemically synthesized (Agilent Technologies) at 10 pmol each and then amplified in 9 cycles using Phusion DNA polymerase (New England Biolabs) from primers that added flanking overlaps to tiles allowing Infusion cloning in a screening vector. The screening vector was based on a lentiviral vector pRRLSIN.cPPT.PGK-GFP.WPRE (a gift from Didier Trono; Addgene plasmid # 12252), in which an enhancer blocker and barrier insulator element C115 was introduced 36 bp into the proximal portion of the 3′ LTR; from this position, the insulator element is copied into the 5′ LTR during proviral cDNA synthesis, thus flanking the expression cassette on both sides upon integration. In addition, the original PGK promoter was replaced with a minimal β-globin promoter (chr11:5227033-5227201, hg38) and EcoRI and KpnI sites in positions 3830 and 3846 in the original pRRLSIN vector were deleted and re-created in 5’ position of the β-globin promoter to allow insertion of DHS tiles. the titer of the lentiviral library was measured using qPCR in genomic DNA isolated from K562 cells that were infected with two known volumes of lentivirus prep. qPCR was carried out using SYBR-Green protocol with primers specific to RRE region in lentiviral genome (Forward, 5’-AACTCACAGTCTGGGGCATC-3’ and Reverse, 5’-TGGTGCAAATGAGTTTTCCA-3’) and to ALB region in human genome (Forward, 5’-GCTGTCATCTCTTGTGGGCTGT-3’ and Reverse, 5’-ACTCATGGGAGCTGCTGGTTCA-3’), with a known number of DNA molecules containing RRE and ALB amplicons as a standard curve. The vast majority of designed DHS fragments ( > 95%) were successfully synthesized and represented in the lentiviral library (Supplementary Fig. 3). A similar cloning strategy was used for the construction of the full-DHS library of 202 elements identified based on the 15.000 tile library. The elements were synthesized as 301-bp fragments (Twist Biosciences), pooled, and amplified using their flanking sequences with primers that added flanking overlaps to allow Infusion cloning in the screening vector. Cloning product for both libraries was introduced in chemically competent E. coli and resulting plasmid produced with redundancy of 144 clones per insert for the large tile library and 550 clones per insert for the small library. Forty candidate DHS fragments were also cloned individually in the screening vector, using an identical approach, as were enhancer controls consisting of the β-globin LCR HS2, HS3 and HS4 (all sequences are provided in Supplementary Data and β-globin µLCR (chr11:5280522-5281209; 5284350-5285412; 5287870 -5288668, hg38)), and a negative control consisting of sequence in ALB locus (chr4: 73,418,934-73,419,006, hg38). HUDEP-2 cells were transduced with either lentiviral enhancer screening library at MOI = 0.4. 5 days post transduction the cells were sorted based on their GFP expression on a MoFlo Astrios Cell Sorter. Specifically, 3 bins were selected represented the 5% of top, medium, and low GFP expressing cells with an average MFI of 15.000, 3.000 and 750 respectively (Supplementary Fig. 3b).
Sequencing of enhancer screening libraries and data preprocessing
Total DNA from enhancer screening library transduced HUDEP-2 cells was isolated using the DNeasy Blood & Tissue kit (Qiagen, 69504) and the enhancer inserts were PCR amplified using the following primers:
-
F: 5’-ACACGACGCTCTTCCGATCTNNNNGTCGAATTAAGGACCGGATCA-3’
-
R: 5’- GACGTGTGCTCTTCCGATCTNNNNAAGTGATGACAGCCGTACCA-3’
Samples were sequenced on HiSeq 4000 (Illumina) using a 2×76 paired-end sequencing. Cloning region flanking sequences were trimmed using seqtk (https://github.com/lh3/seqtk) using the trimfq command and removing 22 bases from read1 and 20 bases from read2. Illumina adapters were trimmed using Trimmomatic v0.36 setting the ILLUMINACLIP argument to 2:30:10:2:true. Sequences were aligned against the GRCh38 reference genome with BWA v0.7.1253 and sequences were filtered using samtools (http://www.htslib.org)54 with the following arguments: view -q 1 -f 64. Sequences were converted to genomic coordinates BED file using the bam2bed tool from the BEDOPS suite55. For every reference sequence (tile or full DHS depending on screening vector library) we counted the number of reads in each sample that overlap the reference sequence by ≥90% using the bedmap tool from BEDOPS.
Enhancer screening analysis
To estimate the effect of each tile from the 14,999-tile screening library (15 K library) on GFP expression we employed a maximum likelihood estimation framework adapted from MAUDE17 developed for sorting-based screenings. The log-likelihood of the observed read counts in each GFP bin were calculated under a negative binomial model based on the expected reads per bin following a normal distribution null model. The expected mean effect score was the basis for ranking tiles for enhancer activity. For the mini-library of 205 full-size DHS sequences the effect size of each sequence on GFP expression was estimated using linear regression of the per element counts against the MFI of each GPF sorting bin.
Lentivirus production
The VSV.G-pseudotyped Lentivirus vectors (LV) were produced in 293FT cells grown in DMEM and transiently transfected using calcium phosphate method with a plasmid mix containing packaging vector psPAX2, envelope pseudotyping vector pVSV-G, and library screening vector containing DHS tiles or elements, with or without shmiRNA cassette. Sixteen hours after transfection, new medium was added to the culture. The supernatant was collected twice: twenty-four and forty-eight hours after medium change. LV-containing supernatants were filtered through a 0.45 μm filter (Millipore). For the LV concentration we used Spin-XR UF concentrators (Corning). To estimate the viral titers, K562 cell were seeded in 48-well plate at 5 × 104 cells/well and infected with serial dilutions of the vector stocks in the presence of 8 µg/mL Polybrene, maintained in culture for at least five days, and genomic DNA from cells was isolated. To estimate the physical viral titer, real-time quantitative PCR was performed in ABI 7500 (Applied Biosystems) with the use of the following primers and probes:
-
Gag F: 5’-GGAGCTAGAACGATTCGCAGTTA-3’
-
Gag R: 5’-GGTTGTAGCTGTCCCAGTATTTGTC-3’
-
Gag Probe: 5’-FAM-ACAGCCTTCTGATGTTTCTAACAGGCCAGG-TAMRA-3’
-
hAlb F: 5’-TGAAACATACGTTCCCAAAGAGTTT-3’
-
hAlb R: 5’-CTCTCCTTCTCAGAAAGTGTGCATAT-3’
-
hAlb Probe: 5’-VIC-TGCTGAAACATTCACCTTCCATGCAGA-TAMRA-3’
Each DNA sample was run in 25 μL reaction volume using Taqman Universal PCR Master Mix (Applied Biosystems). Thermal cycling was started for 2 min at 50 °C, followed by 10 min at 95 °C and 40 thermal cycles of 15 s at 95 °C and 1 min at 60 °C. Vector copy number/cell was calculated by normalizing to the endogenous ALB gene.
Genome editing
To delete the two erythroid enhancers (coordinates) we designed and produced TALEN monomers, flanking in pairs the 5’ and 3’ ends of the PVT1 and PPARA enhancers using adaptations of previously described methods (Cermak et al., Sakuma et al.). TALEN mRNAs were prepared using a mMessageMachine T7 Ultra Kit (#AM1345, Ambion). Sense and anti-sense TALENs recognize the sequences are presented in Supplementary Data 6.
For all transfections, a BTX ECM830 device (BTX Harvard Apparatus) with a 2 mm gap cuvette was used. Two days post transfection we performed single cell sorting in 96 well plates. Single and double knock out clones (SKO and DKO) were evaluated by an in-out PCR of each clone with the following primers:
-
PVT1 In Rev: 5’-GCCCCAGCAAAGACGTTAAG-3’
-
PVT1 Out For: 5’-TGCGGAGTGAGCCTTATTCA-3’
-
PVT1 Out Rev: 5’GTTGTGGGGTACAAGCCAGA-3’
-
PPARA In Rev: 5’GGCCACACACATGGCTTTTT-3’
-
PPARA Out For: 5’GGCCTAGGTTTTTGCTGGGA-3’
-
PPARA Out Rev: 5’TTTGAGGTCATCAGCTGCGT-3’
RNA sequencing
For gene expression analysis, total RNA from expanded HUDEP-2 edited and wild-type clones was collected using the mirVana RNA isolation kit (ThermoFisher Scientific) or RNeasy Mini Kit (Qiagen) from sorted ( > 20,000 cells) and bulk cultures ( > 1,000,000 cells). Illumina libraries were constructed using the TruSeq Stranded Total RNA with Ribo-Zero Globin (Illumina). Finally, libraries were quantified using Fragment Analyzer (Advanced Analytical). RNA-seq libraries were sequenced with HiSeq 4000 using a 2 x 76 bp read length.
Differential gene expression analysis
RNA-seq reads were aligned against the GRCh38 reference genome using STAR56. Gene counts were obtained using featureCounts57. Differential expression analysis was performed using DESeq258. Differentially expressed genes were called at adjusted p-value < 0.05 and absolute log2 fold change > 0.58.
DNase I accessibility
DNase I-seq from HUDEP2 cells was performed according to an established protocol13. Specifically, 100,000-200,000 live cells were collected, and nuclei were extracted in the presence of 0.04% de-ionized IGEPAL CA-630 incubated at 4 °C for 10 min. Nuclei were treated with a gradient of DNase I solution (40 IU to 100 IU of DNase I) for 3 min at 37 °C. DNase I digestion was quenched by adding an equal volume of 5X Stop buffer and 20 μL RNase A (Sigma, R4642) followed by incubation at 37 °C for 60 min. After incubation, 1 µL of Proteinase K (Sigma, P4850) was added and the reactions were incubated at 50 °C for 60 min. Digested genomic DNA was visualized on 1.2% agarose gel and the fragment size profile was generated using the Fragment Analyzer (Advanced Analytical). Prior to genomic library generation, fragments were subjected to size selection with large fragment depletion by magnetic bead separation. Fragment size distribution and concentration of the fractionated sample was measured with Fragment Analyzer (Advanced Analytical). Illumina compatible, double-stranded DNA libraries from the size fractionated samples were constructed using the ThruPLEX DNA-seq Kit (Takara Bio) according to manufacturer’s instructions. DNase I-seq libraries were sequenced on NextSeq 500 with a 2x36bp read length. Adapter trimmed FASTQ files were aligned against GRCh38 using BWA53. Hotspots were detected using hotspot2 software (https://github.com/Altius/hotspot2). All downstream DNase I-seq analyses were performed on DNase I hotspots (genomic regions with statistically significant enrichment in DNase I cleavage).
CUT&RUN
HUDEP-2 CUT&RUN was performed as described59. Briefly, 2x10E5 HUDEP cells were washed twice with wash buffer (20 mM HEPES pH7.5, 150 mM NaCl, 0.5 mM spermidine, supplemented with proteinase inhibitor cocktail Roche 4693159001) and incubated with 15 μL Concannavalin A-coated beads (Bangs Laboratories BP-531). The beads were cleared and resuspended in 100 μl antibody binding buffer (20 mM HEPES pH7.5, 150 mM NaCl, 0.5 mM spermidine, 0.02% digitonin, 2 mM EDTA pH8) with the primary antibody and incubated overnight at 4 C. The beads were cleared and washed twice with digitonin-wash buffer (20 mM HEPES pH7.5, 150 mM NaCl, 0.5 mM spermidine, 0.02% digitonin) and were resuspended in 100 μL digitonin-wash buffer with 0.5 μL protein-MNase (14 μg/mL, kindly provided by Dr. Steven Henikoff from FredHutchinson Cancer Research Center) and incubated at 4 C for 1 hour. The beads were cleared and washed twice with digitonin-wash buffer and then resuspended in 100 μL digitonin-wash buffer. 2 μL of 0.1 M CaCl2 were added and the digestion went on for 30 min at 4 C. The reaction was stopped by adding 100 μL stop buffer (340 mM NaCl, 20 mM EDTA pH8, 4 mM EGTA, 0.02% digitonin, 20 μg/mL glycogen, 50 μg/mL RNase A) and the reaction mixture was incubated at 37 C for 10 min. The beads were cleared and the supernatant was collected for proteinase K digestion (55 C 30 min) and DNAs were cleaned by phenol-chloroform-iso amyl alcohol extraction and ethanol precipitation. Antibodies used were obtained from following suppliers: GATA1 (Acam ab11852), TAL1 (GeneTex GTX116020). Illumina compatible, double-stranded DNA libraries from the size fractionated samples were constructed using the ThruPLEX DNA-seq Kit (Takara Bio) according to manufacturer’s instructions. DNase I-seq libraries were sequenced on NextSeq 500 with a 2x36bp read length. Adapter trimmed FASTQ files were aligned against GRCh38 using BWA v0.7.1253.
ChromHMM
Analysis and annotation of functional chromatin states was performed using ChromHMM v1.2518. GRCh38 aligned BAM files from HUDEP-2 DNaseI-seq, and GATA1, TAL1, H3K27ac, H3K4me1, H3K4me3, H3K27me3 and H3K9me3 CUT&RUN experiments were used as inputs. A 15-state model was chosen, paired mode was enabled, and bin size was set to 100 bp.
Motif enrichment analysis
Transcription factor binding site (TFBS) motif over- and under-representation analysis by hypergeometric test was performed using hyperMotif (https://github.com/ggeorgol/hyperMotif/releases/tag/v1.0.0). A compilation of TFBS motif models from public databases was used, obtained from60.
Statistical analyses
All statistical analyses were performed using R 4.0.5 and Python 3.8. Specific statistical tests, along with corresponding test statistic values, are detailed in figure legends and the main text. P-values, along with adjustments for multiple comparisons, are reported as applicable. Results are expressed as mean ± standard deviation (SD), mean ± standard error (SE) or median ± interquartile range (IQR), depending on data distribution. For boxplots, center lines represent the median, box limits represent the first and third quartiles, and whiskers extend to 1.5 times the IQR. Outliers are shown as individual points. Sample sizes of 10 or fewer are shown as individual points. Otherwise, sample sizes are denoted as appropriate in the main text and figure legends.
shmiRNA vector design and production
Lentiviral vectors were based on the pRRLSIN.cPPT.PGK-GFP.WPRE (kindly gifted from Didier Trono; Addgene plasmid # 12252; http://n2t.net/addgene:12252; RRID:Addgene 12252) in which the original human PGK promoter was exchanged for a truncated human beta-globin promoter (human genome hg38 coordinates: chr11:5227033-5227201). In addition, the vector contained a C1 insulator/barrier element15,61 inserted in the proximal portion of the 3′ LTR. From this position, the element is copied into the 5′ LTR during proviral cDNA synthesis, thus flanking the entire proviral expression cassette. EcoRI and KpnI sites in the original vector were deleted and recreated in the 5’ position proximal of beta globin promoter, serving as cloning sites for insertion of enhancer elements in this basal screening vector. Subsequently, positive control vector derivatives containing truncated HS2 element originating from human beta globin locus control region and a second one with expanded region composed of truncated HS2, HS3, and HS4 elements62 were created. Human genome hg38 coordinates for HS2 are chr11: 5,280,523-5,281,381, hg38, coordinates for HS2-HS3-HS4 are chr11: 5,280,522-5,281,209, 5,284,350-5,285,412, 5,287,870-5,288,668, respectively. Negative control vector was prepared by inserting a sequence from human albumin locus (human hg38 coordinates chr4: 73,418,934-73,419,006). Lentiviral vectors expressing miRNA-embedded shmiRNA targeting BCL11A were based on control vectors described above in which the GFP ORF was appended by a cassette containing BCL11A-targeting shmiRNA embedded in miR223 as described previously20,21.
FACS
To assess the multilineage engraftment of CD34+ cells in the bone marrow of NBSGW mice post transplantation, the following antibodies were used: CD45-APC (BD Biosciences), CD19-PE (BD Biosciences), CD3-PE (ExBIO), CD33-PerCp (ExBIO), CD235a-PE (ExBIO), CD34 (BD Biosciences), CD41 (BD Biosciences). After wash, cells were resuspended in FACS buffer and analyzed using a FACS-Calibur (BD Biosciences, San Jose, CA). To evaluate the differentiation stages during ex vivo erythroid differentiation culture cells were stained with the following antibodies: CD235a-PE (ExBIO) and CD36-APC (ExBIO). For the intracellular detection of human γ-globin expression, erythroid cells were fixed in 4% paraformaldehyde followed by permeabilization in 1:1 acetone/H2O, 100% acetone, 1:1 acetone/H2O and stained with anti-human gamma globin (51.7) from Santa Cruz. The fraction of enucleated cells was measured with the use of nuclear red (Thermo Fisher Scientific). was excluded using a forward scatter-area and sideward scatter-area gate. Flow cytometry data were then analyzed using FlowJo (version 10.0.8, FlowJo, LLC).
Globin HPLC
Individual globin chain levels from transduced ex vivo differentiated normal CD34+ cells or NBSGW bone marrow-engrafted human cells were quantified on a Shimadzu Prominence instrument with an SPD-10AV diode array detector and an LC-10AT binary pump (Shimadzu, Kyoto, Japan). Vydac 214TP™ C4 Reversed-Phase columns for polypeptides (214TP54 Column, C4, 300 Å, 5 µm, 4.6 mm i.d. x 250 mm) (Hichrom, UK) were used. Globin chains from ex vivo differentiated thalassemic cells were quantified on a Shimadzu LC-2060C 3D Liquid chromatography with a GmbH MultoHigh Bio 300, 250 × 3 mm column. A 38%–60% gradient mixture of 0.1% trifluoroacetic acid in water/acetonitrile was applied at a rate of 1 mL/min.
Xenotransplantation
Immunodeficient NOD.Cg-KitW-41J Tyr+ Prkdcscid Il2rgtm1Wjl/ThomJ (NBSGW) mice were obtained from the Jackson Laboratory (Bar Harbor, ME). Mice used in all experiments were housed in a pathogen-free animal facility at the University of Washington or the Gene and Cell Therapy Center, G.Papanikolaou Hospital (EL54BIOExp14) and were free of infection or disease. Housing rooms were maintained at a 12 h light/12 h dark cycle and controlled temperature of approximately 23 °C and 40–60% humidity. To evaluate the enhancers’ cell specificity, CD34+ cells were transduced with a pool of the same MOI of 40 different vectors. 2 days post transduction, ex vivo transduced 1 × 106 CD34+ cells from healthy donors were injected intravenously into NBSGW recipient mice, without any conditioning. Non-transduced CD34+ cells were used as controls. Five non irradiated 8–12 weeks female mice were transplanted per group in all experiments. Male mice were not included in the studies due to their well-documented low bone marrow engraftment capacity of human cells63,64. Sixteen weeks post transplantation, NBSGW mice were sacrificed and bone marrow cells were collected, for assessment of multilineage engraftment. To evaluate specificity, efficiency and in vivo safety of the BCL11A-shmiRNA vectors, CD34+ cells from healthy donors were transduced with the same MOI (40) with the two shmiRNA vectors. Mock transduced cells were transplanted in the control group. Transplantation of the cells was performed in the same manner as described above. Sixteen weeks post transplantation the recipient mice were sacrificed, and bone marrow was collected from 4 long bones. Multilineage engraftment, GFP and HbF expression were evaluated by flow cytometry. All procedures were approved by the institutional and/or regional animal care and use committee (University of Washington IACUC #2437-05, Directorate of Veterinary Medicine (PKM), Central Macedonia, Greece #658637(2586)).
Cytospin slide preparation
Cytospins of 0.3-0.5 × 105 thalassemic transduced and untransduced cells were prepared during erythroid differentiation, by cytocentrifugation (ROTOFIX 32, Hettich Zentrifugen) at 100 g for 5 minutes. Cytospins were air dried and then stained with Giemsa/May-Grünwald (Merck, Darmstadt, Germany) for 15 and 5 minutes, respectively.
Analysis of Reactive Oxygen Species (ROS) levels
Intracellular ROS levels from erythroid cells were determined using the CellROX Deep Red Flow Cytometry Assay kit (Invitrogen by Thermo Scientific), according to the manufacturer’s instructions. Briefly, 1 × 106 thalassemic ex vivo differentiated cells were incubated for 30 minutes at 37oC, protected from exposure to light with the CellROX reagent at a final concentrate on of 500 nM. Oxidation of the probe can be detected by the increase of fluorescence (APC) with flow cytometry.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Ex vivo CD34+ erythroid differentiation DNase I data generated previously is available from GEO GSE183268. DNase I data from CD4+ activated T cells, CD8+ activated T cells, macrophages, CD14+ monocytes, NK cells, CD19+ B cells K562 were obtained from ENCODE (accession numbers: ENCSR438USP, ENCSR294YEJ, ENCSR721XAP, ENCSR407WGG, ENCSR241BNZ, ENCSR381PXW, and ENCSR000EKS). DNase I data from HUDEP-2 cells is deposited in GEO with accession number: GSE252160. CUT&RUN data from HUDEP-2 cells is deposited in GEO with accession number: GSE252157. MPRA DNAseq data in HUDEP-2 cells is deposited in GEO with accession number GSE252159. Source data are provided with this paper.
References
Locatelli, F. et al. Betibeglogene autotemcel gene therapy for non–β0/β0 genotype β-thalassemia. N. Engl. J. Med. 386, 415–427 (2021).
Fumagalli, F. et al. Lentiviral haematopoietic stem-cell gene therapy for early-onset metachromatic leukodystrophy: long-term results from a non-randomised, open-label, phase 1/2 trial and expanded access. Lancet 399, 372–383 (2022).
Florian, E. et al. Hematopoietic stem-cell gene therapy for cerebral adrenoleukodystrophy. N. Engl. J. Med. 377, 1630–1638 (2017).
Kwiatkowski, J. L. et al. Betibeglogene autotemcel gene therapy in patients with transfusion-dependent, severe genotype β-thalassaemia (HGB-212): a non-randomised, multicentre, single-arm, open-label, single-dose, phase 3 trial. Lancet 404, 2175–2186 (2024).
Consortium, T. E. P. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2011).
Meuleman, W. et al. Index and biological spectrum of human DNase I hypersensitive sites. Nature 584, 244–251 (2020).
Visel, A. et al. ChIP-seq accurately predicts tissue-specific activity of enhancers. Nature 457, 854–858 (2009).
Heintzman, N. D. et al. Histone modifications at human enhancers reflect global cell-type-specific gene expression. Nature 459, 108–112 (2009).
Andersson, R. et al. An atlas of active enhancers across human cell types and tissues. Nature 507, 455–461 (2014).
Inoue, F. & Ahituv, N. Decoding enhancers using massively parallel reporter assays. Genomics 106, 159–164 (2015).
Melnikov, A. et al. Systematic dissection and optimization of inducible enhancers in human cells using a massively parallel reporter assay. Nat. Biotechnol. 30, 271–277 (2012).
Arnold, C. D. et al. Genome-wide quantitative enhancer activity maps identified by STARR-seq. Science 339, 1074–1077 (2013).
Georgolopoulos, G. et al. Discrete regulatory modules instruct hematopoietic lineage commitment and differentiation. Nat. Commun. 12, 6790 (2020).
Esrick, E. B. et al. Post-transcriptional genetic silencing of BCL11A to treat sickle cell disease. N. Engl. J. Med. 384, 205–215 (2020).
Papayanni, P.-G. et al. Investigating the barrier activity of novel, human enhancer-blocking chromatin insulators for hematopoietic stem cell gene therapy. Hum. Gene Ther. 32, 1186–1199 (2021).
Maricque, B. B., Chaudhari, H. G. & Cohen, B. A. A massively parallel reporter assay dissects the influence of chromatin structure on cis-regulatory activity. Nat. Biotechnol. 37, 90–95 (2019).
Boer, C. G. de., Ray, J. P., Hacohen, N. & Regev, A. MAUDE: inferring expression changes in sorting-based CRISPR screens. Genome Biol. 21, 134 (2020).
Ernst, J. & Kellis, M. ChromHMM: automating chromatin-state discovery and characterization. Nat. Methods 9, 215–216 (2012).
McIntosh, B. E. et al. Nonirradiated NOD,B6.SCID Il2rγ −/− Kit W41/W41 (NBSGW) mice support multilineage engraftment of human hematopoietic cells. Stem Cell Rep. 4, 171–180 (2015).
Brendel, C. et al. Lineage-specific BCL11A knockdown circumvents toxicities and reverses sickle phenotype. J. Clin. Invest. 126, 3868–3878 (2016).
Guda, S. et al. miRNA-embedded shRNAs for lineage-specific BCL11A knockdown and hemoglobin F induction. Mol. Ther. 23, 1465–1474 (2015).
Luc, S. et al. Bcl11a deficiency leads to hematopoietic stem cell defects with an aging-like phenotype. Cell Rep. 16, 3181–3194 (2016).
Psatha, N. et al. Disruption of the BCL11A erythroid enhancer reactivates fetal hemoglobin in erythroid cells of patients with β-thalassemia major. Mol. Ther. Methods Clin. Dev. 10, 313–326 (2018).
Yannaki, E. et al. Hematopoietic stem cell mobilization for gene therapy of adult patients with severe β-thalassemia: results of clinical trials using g-csf or plerixafor in splenectomized and nonsplenectomized subjects. Mol. Ther. 20, 230–238 (2012).
Yannaki, E. et al. Hematopoietic stem cell mobilization for gene therapy: superior mobilization by the combination of granulocyte–colony stimulating factor plus plerixafor in patients with β-thalassemia major. Hum. Gene Ther. 24, 852–860 (2013).
Montini, E. et al. The genotoxic potential of retroviral vectors is strongly modulated by vector design and integration site selection in a mouse model of HSC gene therapy. J. Clin. Invest. 119, 964–975 (2009).
Belbellaa, B., Reutenauer, L., Messaddeq, N., Monassier, L. & Puccio, H. High levels of frataxin overexpression lead to mitochondrial and cardiac toxicity in mouse models. Mol. Ther. Methods Clin. Dev. 19, 120–138 (2020).
Huichalaf, C. et al. In vivo overexpression of frataxin causes toxicity mediated by iron-sulfur cluster deficiency. Mol. Ther. Methods Clin. Dev. 24, 367–378 (2022).
Alstyne, M. V. et al. Gain of toxic function by long-term AAV9-mediated SMN overexpression in the sensorimotor circuit. Nat. Neurosci. 24, 930–940 (2021).
Sessa, M. et al. Lentiviral haemopoietic stem-cell gene therapy in early-onset metachromatic leukodystrophy: an ad-hoc analysis of a non-randomised, open-label, phase 1/2 trial. Lancet 388, 476–487 (2016).
Psatha, N., Papayanni, P.-G. & Yannaki, E. A new era for hemoglobinopathies: more than one curative option. Curr. Gene Ther. 17, 364–378 (2018).
Yannaki, E. et al. Success stories and challenges ahead in hematopoietic stem cell gene therapy: hemoglobinopathies as disease models. Hum. Gene Ther. 32, 1120–1137 (2021).
Duncan, C. N. et al. Hematologic cancer after gene therapy for cerebral adrenoleukodystrophy. N. Engl. J. Med. 391, 1287–1301 (2024).
Han, J. et al. β-Globin lentiviral vectors have reduced titers due to incomplete vector RNA genomes and lowered virion production. Stem Cell Rep. 16, 198–211 (2021).
Cabriolu, A. et al. Globin vector regulatory elements are active in early hematopoietic progenitor cells. Mol. Ther. 30, 2199–2209 (2022).
Gordon, M. G. et al. lentiMPRA and MPRAflow for high-throughput functional characterization of gene regulatory elements. Nat. Protoc. 15, 2387–2412 (2020).
Kwasnieski, J. C., Fiore, C., Chaudhari, H. G. & Cohen, B. A. High-throughput functional testing of ENCODE segmentation predictions. Genome Res. 24, 1595–1602 (2014).
Gasperini, M. et al. A genome-wide framework for mapping gene regulation via cellular genetic screens. Cell 176, 377–390.e19 (2019).
Inoue, F. et al. A systematic comparison reveals substantial differences in chromosomal versus episomal encoding of enhancer activity. Genome Res. 27, 38–52 (2017).
Klein, J. C. et al. A systematic evaluation of the design and context dependencies of massively parallel reporter assays. Nat. Methods 17, 1083–1091 (2020).
Mauduit, D. et al. Analysis of long and short enhancers in melanoma cell states. Elife 10, e71735 (2021).
Patwardhan, R. P. et al. Massively parallel functional dissection of mammalian enhancers in vivo. Nat. Biotechnol. 30, 265–270 (2012).
Lambert, J. T. et al. Parallel functional testing identifies enhancers active in early postnatal mouse brain. Elife 10, e69479 (2021).
Kvon, E. Z. et al. Genome-scale functional characterization of Drosophila developmental enhancers in vivo. Nature 512, 91–95 (2014).
Dickel, D. E. et al. Genome-wide compendium and functional assessment of in vivo heart enhancers. Nat. Commun. 7, 12923 (2016).
Morgan, R. A., Gray, D., Lomova, A. & Kohn, D. B. Hematopoietic stem cell gene therapy: progress and lessons learned. Cell Stem Cell 21, 574–590 (2017).
Brusson, M. et al. Novel lentiviral vectors for gene therapy of sickle cell disease combining gene addition and gene silencing strategies. Mol. Ther. Nucleic Acids 32, 229–246 (2023).
Uchida, N. et al. Sustained fetal hemoglobin induction in vivo is achieved by BCL11A interference and coexpressed truncated erythropoietin receptor. Sci. Transl. Med. 13, eabb0411 (2021).
Psatha, N. et al. Enhanced HbF reactivation by multiplex mutagenesis of thalassemic CD34+ cells in vitro and in vivo. Blood 138, 1540–1553 (2021).
Psatha, N., Georgolopoulos, G., Phelps, S. & Papayannopoulou, T. Brief report: a differential transcriptomic profile of ex vivo expanded adult human hematopoietic stem cells empowers them for engraftment better than their surface phenotype. Stem Cell Transl. Med. 6, 1852–1858 (2017).
Giarratana, M.-C. et al. Proof of principle for transfusion of in vitro–generated red blood cells. Blood 118, 5071–5079 (2011).
Egeland, T. et al. Myeloid differentiation of purified CD34+ cells after stimulation with recombinant human granulocyte-monocyte colony-stimulating factor (CSF), granulocyte-CSF, monocyte-CSF, and interleukin-3. Blood 78, 3192–3199 (1991).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. GigaScience 10, giab008 (2021).
Neph, S. et al. BEDOPS: high-performance genomic feature operations. Bioinformatics 28, 1919–1920 (2012).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Skene, P. J., Henikoff, J. G. & Henikoff, S. Targeted in situ genome-wide profiling with high efficiency for low cell numbers. Nat. Protoc. 13, 1006–1019 (2018).
Vierstra, J. et al. Global reference mapping of human transcription factor footprints. Nature 583, 729–736 (2020).
Liu, M. et al. Genomic discovery of potent chromatin insulators for human gene therapy. Nat. Biotechnol. 33, 198–203 (2015).
Li, Q., Peterson, K. R., Fang, X. & Stamatoyannopoulos, G. Locus control regions. Blood 100, 3077–3086 (2002).
Notta, F., Doulatov, S. & Dick, J. E. Engraftment of human hematopoietic stem cells is more efficient in female NOD/SCID/IL-2Rgc-null recipients. Blood 115, 3704–3707 (2010).
Ng, E. S. et al. Long-term engrafting multilineage hematopoietic cells differentiated from human induced pluripotent stem cells. Nat. Biotechnol. https://doi.org/10.1038/s41587-024-02360-7 (2024).
Acknowledgements
The authors want to acknowledge Paige Ver Burg and Etaane Neumann for their help with plasmid construction and virus production. This work was supported by the NIH grant 5R01HL136375-02 (to G.S. and T.P.) and by a Topic-in-Focus Advanced Research Grant awarded by the European Hematology Association (to N.P.). This project is carried out within the framework of the National Recovery and Resilience Plan Greece 2.0, funded by the European Union – NextGenerationEU (Implementation body: HFRI-OGETHERA-15358) (to N.P.). This work is dedicated to the memory of Prof. George Stamatoyannopoulos who conceptualized and supervised the project until his death in June 2018.
Author information
Authors and Affiliations
Contributions
N.P., J.V. and G.S. designed and conceptualized the study; N.P., G.G., P.S., K.P., M.I., J.B., T.U., H.W., A.K., NI.V., MS.W. performed experiments. N.P., G.G., and J.V. analyzed data. N.P., G.S. and J.V. supervised the study. J.A.S, E.Y. and T.P. provided consultation. N.P., G.G., P.S., and J.V. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
N.P., G.G., J.S. and J.V. have filed a patent application (US PPA No. 63224537 and PCT No. US202114752) covering various aspects of this work. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Yongchun Zuo and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Psatha, N., Sova, P., Georgolopoulos, G. et al. Large-scale discovery of potent, compact and erythroid specific enhancers for gene therapy vectors. Nat Commun 16, 4325 (2025). https://doi.org/10.1038/s41467-025-59235-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-59235-x