Abstract
Grain size is a key agronomic traits that influence grain yield in crops. The transcription factor GS2/OsGRF4 can improve grain size and yield, but its underlying mechanism remains unclear. Here we report a suppressor of the gain-of-function allele GS2AA (SUG1) that encodes a plant-specific protein DEP2/SRS1/EP2/OsRELA and acts as a transcriptional regulator. The sug1 mutants form short grains, while overexpression of SUG1 results in long grains. GS2 directly activates the expression of SUG1. SUG1 associates with transcription factors OsBZR1, OsMADS56 and OsSPL13 to control grain size through GA and BR signaling as well as growth pathways. Natural variation in SUG1 contributes to grain size diversity, and the SUG1Hap2 allele from indica varieties can be used to improve grain size and yield of japonica varieties with the SUG1Hap3 allele. Thus, our findings uncover that the GS2-SUG1 module controls grain size by integrating multiple growth signals, providing the potential targets for crop improvement.
Similar content being viewed by others
Introduction
Rice is one of the most important food crops, and further increases in rice grain yield are necessary to ensure food security1. Rice yield is mainly determined by tiller numbers per plant, grain number per panicle, and grain weight. Rice grain size is one of the major factors determining grain weight, grain yield, and appearance quality2,3. In plants, several signaling pathways controlling seed size have been described, such as MAPK signaling, G protein signaling, the ubiquitin-related pathway, phytohormone signaling, and transcriptional regulators2. In rice, several important factors that improve grain size have been discovered, which enable approaches for breeding new rice varieties4. Thus, understanding the genetic and molecular mechanisms underlying grain size control is crucial for improving crop yield.
Transcriptional regulation plays a crucial role in rice grain size control. For example, several SQUAMOSA PROMOTER BINDING PROTEIN-LIKE (SPL) transcription factors are necessary for grain size regulation in rice. The OsSPL13 positively regulates grain size by influencing cell expansion in the grain hull5, while OsSPL16 and OsSPL18 control grain size by promoting cell proliferation in the grain hull6,7. By contrast, OsSPL4 and OsSPL12 negatively regulate grain size by influencing cell proliferation in the grain hull8,9. The MADS-box transcription factors OsMADS1 and OsMADS56 play vital roles in rice grain size control10,11. OsMADS1 regulates grain length by limiting cell proliferation in the grain hull. The transcriptional activator WG7 (WIDE GRAIN 7), which contains a cysteine-tryptophan (CW) domain, associates the promoter region of OsMADS1 and activates its expression10. By contrast, OsMADS56 regulates grain length by promoting cell expansion in the grain hull. OsMADS56 influences the expression of several Gibberellin (GA) biosynthetic genes and signaling transduction genes11, indicating that GA signaling is crucial for grain size control by influencing cell expansion. OsBZR1 is a central transcription factor that activates BR signaling. Overexpression of OsBZR1 produces large grains, and OsBZR1 RNAi transgenic plants form small grains12. There are four OsBZRs member family proteins in rice. Simultaneous disruptions of OsBZR1, OsBZR3, and OsBZR4 result in small grains, suggesting that OsBZR1, OsBZR3, and OsBZR4 function redundantly to control grain size13.
GS2/GL2/PT2/GLW2/LGS1 encodes the transcription factor OsGRF4 and is regulated by OsmiR396. The gain of function allele GS2AA with 2 bp substitution in the targeting site of OsmiR396 causes high expression of GS2, resulting in large grains and high yield14,15,16,17,18. GS2AA promotes grain growth mainly by promoting cell expansion in spikelet hulls in rice16. GS2 interacts with the transcriptional coactivators OsGIF1/2/315,17. Overexpression of OsGIF1 leads to large grains in rice15,17. OsGSK2, a homology of Arabidopsis BIN2 kinase, physically interacts with GS2 and represses its transcription activity14, suggesting that GS2 participates in the BR signaling pathway. GS2 also increases nitrogen utilization, cellulosic production and cold tolerance19,20,21. However, the downstream targets of GS2 in regulating grain size are still unclear.
To understand the genetic and molecular mechanisms of GS2 in grain size control, we have conducted a genetic screen to identify modifiers of GS2AA in grain size. Here, we report the suppressor of GS2AA (SUG1) that encodes a plant-specific protein with a BTB/POZ-like domain, three coiled-coil domains and a putative nuclear localization signal (NLS). The sug1 mutants form short grains mainly due to short cells, while overexpression of SUG1 increases grain length. GS2 can associate with the promoter of SUG1 and activate its expression. Further results show that SUG1 associates with multiple transcription factors, such as OsBZR1, OsMADS56 and OsSPL13, to control grain size through BR and GA signaling as well as growth pathways, indicating that SUG1 may act as a transcriptional regulator or cofactor. We also reveal that natural variation in SUG1 contributes to grain size diversity, and the natural allele SUG1Hap2 from indica varieties can be used to improve grain size and yield of japonica varieties with the SUG1Hap3 allele. Thus, our findings reveal a regulatory mechanism that GS2 activates the transcription of SUG1 that integrates with distinct transcription factors to control grain size through several phytohormone and growth pathways, suggesting they are important targets for improving grain size and yield in crops.
Results
The sug1-1 and sug1-2 suppress the grain length phenotype of GS2 AA
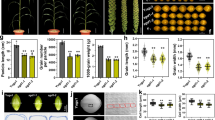
We previously reported that the transcription factor GS2 is a positive regulator of grain size and yield in rice15. The gain of function GS2AA allele in the japonica variety Zhonghua11 background (ZH11-GS2AA) and the indica variety Huazhan (HZ) background (HZ-GS2AA) formed large grains due to the disrupted cleavage by OsmiR396. GS2 mainly promotes cell expansion but also increases cell proliferation in spikelet hulls15,16. The GS2AA allele also produced long panicles compared with their parental lines14,15,16,18. To further understand the mechanism of GS2 in grain size control, we conducted a genetic screen to identify modifiers of GS2AA in grain size. Several suppressors of GS2AA (sug) were isolated from ethyl methanesulfonate (EMS) mutagenized M2 populations of ZH11-GS2AA and HZ-GS2AA, respectively. Among these suppressors, the sug1-1 ZH11-GS2AA plants produced shorter and wider grains than ZH11-GS2AA plants (Fig. 1a, g, h). The 1,000-grain weight of sug1-1 ZH11-GS2AA was slightly decreased compared with that of ZH11-GS2AA (Fig. 1i). The plant height and panicle length of sug1-1 ZH11-GS2AA were reduced in comparison to those of ZH11-GS2AA (Fig. 1b, c, j, k), while the tilling number, number of primary branches, number of secondary branches and grain number per panicle of sug1-1 ZH11-GS2AA were similar to those of ZH11-GS2AA (Supplementary Fig. 1). Similarly, sug1-2 HZ-GS2AA formed shorter, wider and thicker grains than HZ-GS2AA (Supplementary Fig. 2a, d-g). By contrast, the 1,000-grain weight of sug1-2 HZ-GS2AA was similar to HZ-GS2AA (Supplementary Fig. 2h), suggesting that genetic backgrounds may influence grain weight. The plant height and panicle length of sug1-2 HZ-GS2AA were decreased compared with HZ-GS2AA (Supplementary Fig. 2b-c, i, k). The tilling number, number of primary and secondary branches and grain number per panicle of sug1-2 HZ-GS2AA were comparable with those of HZ-GS2AA (Supplementary Fig. 2j, l-n). Thus, these results indicated that sug1-1 and sug1-2 mutations suppress the grain and panicle length phenotypes of ZH11-GS2AA and HZ-GS2AA, respectively.
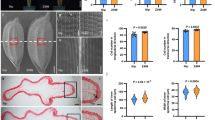
a–c Mature grains (a), plants (b) and panicles (c) of ZH11, ZH11-GS2AA, sug1-1 ZH11-GS2AA, sug1-1 ZH11-GS2AA;proSUG1:SUG1 #1, sug1-1 ZH11-GS2AA;proSUG1:SUG1 #2, and sug1-1 ZH11-GS2AA;proSUG1:SUG1 #3. d The SUG1 gene structure includes untranslated regions (white boxes), exons (black boxes), and introns (lines between boxes). The mutation sites of sug1-1 and sug1-2 are indicated in the diagram. e The schematic diagram of the SUG1 protein and the truncated proteins generated by sug1-1 and sug1-2 mutations. The SUG1 protein comprises a BTB/POZ-like domain, three coiled-coil domains, and a Nuclear localization signal. The mutation in sug1-1 and sug1-2 results in truncated proteins, which lack the C-terminal coiled-coil and nuclear localization signal. f The mutant sites of sug1-1 and sug1-2 were confirmed by sequencing. g–k Grain length (g), grain width (h), 1,000 grain weight (i), plant height (j), and panicle length (k) of ZH11, ZH11-GS2AA, sug1-1 ZH11-GS2AA, and genomic complementation lines sug1-1 ZH11-GS2AA;proSUG1:SUG1. n = 56 in (g, h) n = 3 in i, n = 13 in (j, k). Data in (g–k) are presented as mean ± SD. Different letters represent significant differences (P < 0.05) from ordinary one-way ANOVA multiple comparisons with Tukey’s multiple comparisons test. Bars: 3 mm (a), 10 cm (b), 5 cm (c). Source data are provided as a Source data file.
Identification of the SUG1 gene
A MutMap method was used to identify the SUG1 gene22. The sug1-1 ZH11-GS2AA and sug1-2 HZ-GS2AA were crossed with their parental lines ZH11-GS2AA and the HZ-GS2AA to generate F2 populations, respectively. In F2 populations, plants with short grains were selected to extract genomic DNA and pooled for whole genome resequencing. The genomic DNA of the ZH11-GS2AA and HZ-GS2AA were resequenced as controls. The SNP (Single Nucleotide Polymorphism) and INDEL (Insertion/deletion) analyses were conducted according to a previous report22. There were one candidate causal SNP and two candidate INDELs in chromosome 7 were identified in sug1-1 ZH11-GS2AA, while four candidate causal SNPs in chromosome 7 were identified in sug1-2 HZ-GS2AA (Fig. 1d, Supplementary Table 1 and Supplementary Table 2). Among these SNPs and INDELs, INDEL1 in sug1-1 ZH11-GS2AA and SNP4 in sug1-2 HZ-GS2AA occurred in the exons of the same LOC_Os07g42410 gene. The sug1-1 has a single base deletion (1233TG → T) at the seventh exon of the LOC_Os07g42410 gene, which produces a truncated protein of 438 amino acids. The sug1-2 has a C-to-T transition in the codon 701 (CAA/TAA) at the seventh exon of the LOC_Os07g42410 gene, resulting in a premature stop codon (Fig. 1d, e). We further confirmed these two mutations by sequencing LOC_Os07g42410 in the sug1-1 and sug1-2 mutants (Fig. 1f). These results suggested that the LOC_Os07g42410 is the candidate gene for SUG1.
To confirm that LOC_Os07g42410 is the SUG1 gene, we performed the genetic complementary test. A plasmid with the coding sequence of LOC_Os07g42410 driven by its native promoter (proSUG1:SUG1) was transferred into the sug1-1 ZH11-GS2AA mutant. The grain length, grain width, grain weight, plant height and panicle length of sug1-1 ZH11-GS2AA;proSUG1:SUG1 transgenic plants were similar to those of ZH11-GS2AA, indicating that the proSUG1:SUG1 construct complemented the phenotypes of sug1-1 ZH11-GS2AA (Fig. 1a–c, g–k). Thus, these results demonstrated that the SUG1 gene is LOC_Os07g42410.
SUG1 encodes a 1365 amino acid protein that contains a BTB/POZ-like domain, three coiled-coil domains and a putative nuclear localization signal (NLS) (Fig. 1e). The homolog of SUG1 in Arabidopsis is CIP7, which has been reported to promote the expression of light-regulated genes23. SUG1 is a plant specific protein DEP2/SRS1/EP2/OsRELA that has been reported to influence panicle architecture, plant height, grain shape and leaf angle in rice24,25,26,27, but its genetic and molecular mechanisms in grain size control are totally unclear.
Expression pattern and subcellular localization of SUG1
To examine the expression pattern of SUG1, we conducted qRT-PCR analysis and found that SUG1 was expressed in the roots, stems, leaves and panicles (Supplementary Fig. 3a). We further generated the transgenic plants containing the GUS reporter gene under control of the SUG1 promoter (proSUG1:GUS) and analyzed the expression patterns of SUG1 during the development of panicles. We observed GUS (β-glucuronidase) activity throughout panicle developmental stages, with a particularly strong GUS activity in younger panicles and grain hulls. The GUS activity was gradually declined during the panicle and spikelet growth (Supplementary Fig. 3b, c).
To explore the subcellular localization of SUG1, we produced pro35S:GFP-SUG1 transgenic lines in the ZH11 background. The pro35S:GFP-SUG1 transgenic plants exhibited a significant increase in grain length compared to ZH11, indicating that the GFP-SUG1 fusion protein possesses its function in grain size control (Supplementary Fig. 4a–c). Remarkably, GFP fluorescence was observed in the nuclei and cytoplasm of pro35S:GFP-SUG1 transgenic rice (Supplementary Fig. 4d), implying that SUG1 is localized in both nuclei and cytoplasm.
SUG1 regulates grain size by influencing cell expansion
To understand the effect of sug1 single mutant on grain size, we crossed sug1-1 ZH11-GS2AA with the parental line ZH11 to generate the F2 progenies. The sug1-1 single mutant was identified from the F2 segregated populations. The grain length of sug1-1 was significantly decreased compared with that of ZH11 grains (Supplementary Fig. 5a, d), while the grain width and grain thickness were significantly increased compared with those of ZH11, respectively (Supplementary Fig. 5e, f). The weight of sug1-1 grains was decreased in comparison to that of ZH11 grains (Supplementary Fig. 5g). The length-width ratio of sug1-1 grains was dramatically reduced compared with that of wild-type grains (Supplementary Fig. 5h), indicating that SUG1 is necessary for grain shape regulation. The sug1-1 also showed shorter plants and panicles than ZH11 (Supplementary Fig. 5b, c, i, j). By contrast, there were no significant differences in other agronomic traits, such as tiller number, primary panicle branches, secondary panicle branches and grain number per panicle (Supplementary Fig. 5k–n). Similarly, the sug1-2 single mutant exhibited shorter and wider grains and shorter plants and panicles than its parental line HZ (Supplementary Fig. 6).
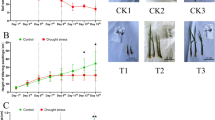
We further produced the loss-of-function mutant allele sug1-cr in the ZH11 background using CRISPR/Cas9 technology (Supplementary Fig. 7a). The sug1-cr contains a base T insertion in the seventh exon of SUG1, which results in a reading frame shift and early termination (Supplementary Fig. 7b, c). The grains of sug1-cr were shorter, wider and thicker than ZH11 (Fig. 2a, f–h), while the weight of sug1-cr grains was decreased compared with ZH11 (Fig. 2i). The sug1-cr exhibited shorter plants and panicles (Fig. 2b, c, j, k). The tilling number, number of primary branches, number of secondary branches and grain number per panicle of sug1-cr were comparable with those of ZH11 (Supplementary Fig. 7d–g). Thus, the sug1-cr has similar grain and panicle size phenotypes to sug1-1 and sug1-2.
a–c Mature rice grains (a), mature plants (b), and panicles (c) of ZH11 and sug1-cr. d-e The outer surface (d) and inner surface (e) of ZH11 and sug1-cr spikelet hulls. The white frames in e marks the complete cellular boundaries in the surface of spikelet hulls. f–k Grain length (f), grain width (g), grain thickness (h), 1,000-grain weight (i), plant height (j), and panicle length (k) of ZH11 and sug1-cr. n = 89 in f–g, n = 60 in h, n = 4 in (i) n = 14 in (j–k). l–o Outer epidermal cell length (l), longitudinal cell number (m), cell width (n), and transverse cell number (o) of ZH11 and sug1-cr lemmas. n = 21 in (l–n) and n = 20 in (o). p Mature rice grains of ZH11 and SUG1 over-expressed transgenic plants. q Relative transcript level of SUG1 in ZH11 and SUG1 over-expressed transgenic plants. n = 3 biological replicates. r–t Grain length (r), grain width (s), and 1,000 grain weight (t) of ZH11 and proACTIN:SUG1 transgenic plants. n = 50 in (r-s) n = 4 in (t). Data in (f–o, q–t) are presented as mean ± SD, relative to the parental line (ZH11). Two-tailed unpaired Student’s t-test was used for statistical analysis. Bars: 3 mm (a, p), 10 cm (b), 5 cm (c), 50 µm (d–e).
The final size of a spikelet hull is coordinately influenced by cell division and cell expansion2. To determine the cellular basis for SUG1 in regulating grain size, we performed morphological and cellular analysis for the outer surface of ZH11 and sug1-cr spikelet hulls. As shown in Fig. 2d, the average cell length of the sug1-cr was greatly decreased in the longitudinal direction compared with that of ZH11, while the cell number had no change between ZH11 and sug1-cr spikelet hulls in the longitudinal direction (Fig. 2l, m). The cell width of sug1-cr was increased in the grain-width direction compared with that of ZH11 (Fig. 2n), whereas the cell number of sug1-cr in the grain-width direction was similar to that of ZH11 (Fig. 2o). Similarly, the inner epidermal cells of sug1-cr grains were shorter and wider than those of ZH11 (Fig. 2e, Supplementary Fig. 7h, i). These results indicated that sug1-cr forms short and wide grains predominantly by influencing cell expansion in spikelet hulls.
Overexpression of SUG1 forms long grains
To further investigate the role of SUG1 in regulating grain size, we asked whether overexpression of SUG1 gene could result in large grains. To achieve this, we introduced the proACTIN:SUG1 construct into the ZH11 variety. The proACTIN:SUG1 transgenic plants exhibited longer grains than ZH11, while the grain width of proACTIN:SUG1 transgenic plants was similar to that of ZH11 (Fig. 2p–s). As a consequence, the grain weight of the proACTIN:SUG1 transgenic lines were remarkably increased compared with that of ZH11 (Fig. 2t). These results indicated that the SUG1 functions as a positive regulator in the regulation of grain length and weight in rice.
GS2 and OsGIF1 acts genetically with SUG1 to regulate grain length
Considering that sug1-1 and sug1-2 were identified as suppressors of GS2AA in grain size, we asked whether GS2 and SUG1 could act genetically to control grain size. To test this, we measured the size of ZH11, ZH11-GS2AA, sug1-1 and sug1-1 ZH11-GS2AA grains. The length of ZH11-GS2AA grains was increased by 27.22% compared with ZH11, while the length of sug1-1 ZH11-GS2AA grains was increased only by 16.92% in comparison to the sug1-1 single mutant (Fig. 3a, b), indicating that sug1-1 partially suppresses the grain length phenotype of ZH11-GS2AA. By contrast, the width of ZH11-GS2AA grains was increased by 14.88% compared with ZH11, while the width of sug1-1 ZH11-GS2AA grains was increased by 17.99% in comparison to the sug1-1 single mutant (Fig. 3c). The grain length-width ratio of sug1-1 ZH11-GS2AA double mutant was similar to that of the sug1-1 single mutant (Fig. 3d), suggesting that sug1-1 is epistatic to GS2AA with respect to grain shape. The loss-of-function of GS2/GRF4 does not obviously affect the grain size in rice, while knocked out of both GS2 and its closest homolog GRF3 leads to small grains28. We crossed sug1-1 with grf3-cri1 grf4-cri2 and isolated sug1-1 grf3-cri1 grf4-cri2 triple mutant (Supplementary Fig. 8a). The length of grf3-cri1 grf4-cri2 grains was decreased compared with ZH11, and sug1-1 enhanced the short grain phenotype of grf3-cri1 grf4-cri2 (Supplementary Fig. 8b), suggesting that SUG1 and OsGRF3/4 have partially overlapped function in grain length. Similar genetic phenomena have also been reported previously29,30,31. For example, the SGD1 and SiUBC32 have partial overlapping functions in regulating grain size, while the double mutant sgd1 siubc32 formed smaller grains than each single mutant29. Thus, these genetic analyses suggested that GS2 and SUG1 act, at least in part, in a common pathway to control grain length.
a Mature rice grains of ZH11, ZH11-GS2AA, sug1-1, and sug1-1 ZH11-GS2AA. b–d Grain length (b), grain width (c), and length-width ratio (d) of ZH11 (n = 62), ZH11-GS2AA (n = 61), sug1-1 (n = 60), and sug1-1 ZH11-GS2AA (n = 72). e Mature rice grains of ZH11, sug1-cr, OsGIF1-OE, and sug1-cr OsGIF1-OE. f–h Grain length (f), grain width (g), and length-width ratio (h) of ZH11 (n = 110), sug1-cr (n = 128), OsGIF1-OE (n = 132), and sug1-cr OsGIF1-OE (n = 128). i The relative expression levels of SUG1 in young panicles of ZH11-GS2AA and OsGIF1-OE compared with ZH11. n = 3 biological replicates. j The PF1 fragment in the 2 kb promoter region of SUG1 contains an ACAGTC sequence, but PF2 does not. A represents the probe derived from the PF1 fragment of the SUG1 promoter, which contains the ACAGTC motif, and A-m represents the mutation probe of A, where ACAGTC is mutated to TACGTT. k ChIP-qPCR analysis revealed that GS2 specifically binds to PF1 in the promoter of SUG1. The panicles of pro35S:MYC and pro35S:MYC-rGS2 transgenic plants were used to extract total chromatin and immunoprecipitated by anti-MYC antibodies. The fold enrichment was normalized to ACTIN, and the means were calculated based on three biological samples. Data are presented as mean ± SD, relative to the pro35S:MYC. Two-tailed unpaired Student’s t-test was used for statistical analysis. l The EMSA experiment determined a direct interaction between GS2 and the SUG1 promoter. m The dual-luciferase assays shown that GS2 activates the transcriptional activation from SUG1 promoter in rice protoplasts. The activity of the empty vector was set to one. Data are presented as mean ± SD. Two-tailed unpaired Student’s t-test was used for statistical analysis. n = 3 biological replicates. Data in (b–d, f–i) are presented as mean ± SD. Different letters represent significant differences (P < 0.05) from ordinary one-way ANOVA multiple comparisons with Tukey’s multiple comparisons test. Bars: 3 mm (a, e). Experiments in (k–m) were repeated independently at least twice with similar results. Source data are provided as a Source data file.
We previously reported that OsGIF1 physically interacts with GS2 and influences grain size15. Considering that GS2 and SUG1 function partially in a common pathway to control grain length and shape, we asked whether SUG1 could act with OsGIF1 in a common pathway to control grain length and shape. To address this, we crossed sug1-cr with OsGIF1-OE plants overexpressing OsGIF1 driven by an ACTIN promoter and identified the sug1-cr OsGIF1-OE plants from the F2 segregated populations15 (Fig. 3e). We measured the grain size of ZH11, OsGIF1-OE, sug1-cr and sug1-cr OsGIF1-OE. The grain length of OsGIF1-OE was increased by 20.32% compared with that of ZH11, while the grain length of sug1-cr OsGIF1-OE was increased only by 13.57% in comparison to that of the sug1-cr single mutant (Fig. 3f), suggesting that GIF1 and SUG1 have overlapped function in grain length control. The grain width of OsGIF1-OE increased by 17.40% compared with that of ZH11, and the grain width of sug1-cr OsGIF1-OE was increased by 10.62% compared with that of sug1-cr single mutant (Fig. 3g). The grain length-width ratio of sug1-cr OsGIF1-OE was similar to that of the sug1-cr single mutant (Fig. 3h), suggesting that sug1-cr is epistatic to OsGIF1-OE with respect to grain shape. These results indicated that SUG1 and OsGIF1 have overlapped functions in the regulation of grain size and shape.
GS2 directly binds to the promoter region of SUG1 to activate its expression
Given that SUG1 acts genetically with GS2 to regulate grain size and shape, we asked whether the transcription factor GS2 could directly regulate the expression of SUG1. We investigated the expression levels of SUG1 in young panicles of ZH11-GS2AA and OsGIF1-OE. As shown in Fig. 3i, the transcriptional levels of SUG1 is significantly upregulated in ZH11-GS2AA and OsGIF1-OE compared with that in ZH11. Thus, these results showed that GS2 and OsGIF1 promote the expression of SUG1.
We then asked whether GS2 could associate with the promoter region of SUG1 in rice. In Arabidopsis, the core motif CTGACA can be recognized specifically by AtGRF932. In rice, GS2 can bind the core motif ACAGTA, which is similar to a reversed CTGACA sequence21. We next analyzed the 2 kb promoter sequence of SUG1, but no forward CTGACA sequence was found. However, we did find the ACAGTC motif, which is a reversed CTGACA sequence, within the promoter region of SUG1 (Fig. 3j). These analyses suggested that GS2 may bind the ACAGTC motif in the SUG1 promoter. To test this, we first fused MYC tag with rGS2 (a miR396-resistant variant of GS2 that disrupts the miR396 recognition without any alteration in amino acid) driven by a CaMV 35S promoter (pro35S:MYC-rGS2) and introduced in the ZH11 variety14. We then carried out a chromatin immunoprecipitation (ChIP) assay using pro35S:MYC and pro35S:MYC-rGS2 transgenic plants. We tested the enrichment of a SUG1 promoter fragment (PF1) that contains the ACAGTC sequence using ChIP-qPCR analysis. As shown in Fig. 3k, PF1 was significantly enriched, but there was no significant enrichment signal was observed in either the ACTIN promoter sequence or the SUG1 promoter fragment PF2, which lacks the ACAGTC sequence. These results indicated that GS2 associates with the promoter region of the SUG1 gene and activates its expression in rice.
To further test whether GS2 could directly bind to the promoter of SUG1, we conducted a DNA electrophoretic mobility shift assay (EMSA) using a GST-fused GS2 protein (GST-GS2). As shown in Fig. 3l, GST-GS2 exhibited the binding affinity towards the biotin-labeled probe A, which contains the ACAGTC sequence, while GST alone did not bind. When the concentrations of an unlabeled probe gradually increased, the binding signal was gradually diminished. Conversely, GST-GS2 failed to bind to the mutated biotin-labeled probe (A-m) that contains mutations in the ACAGTC sequence. These results reveal that GS2 can directly bind the ACAGTC sequence in vitro. We further conducted a transcriptional activation assay to evaluate the transcriptional regulatory relationship between GS2 and GIF1 on the promoter of SUG1. As shown in Fig. 3m, GS2 can activate the transcription of the SUG1 promoter, while GIF1 enhances GS2’s activation of the SUG1 promoter in rice protoplasts. Overall, these results demonstrated that GS2 directly binds to the promoter of SUG1 and promotes its expression.
SUG1 associates with multiple transcriptional factors
To further understand the biochemical function of SUG1, we identified the interacting proteins of SUG1 using the immunoprecipitation mass spectrometry (IP-MS) approach. Surprisingly, 33 transcriptional factors were identified as SUG1-associated proteins, such as MADS, SPL, MYB, WRKY and bZIP-type transcription factors (Supplementary Table 3). We further confirmed the association of SUG1 with transcription factors using Luciferase complementation assay, including OsMADS2, OsMADS4, OsMADS15, OsMADS22, OsMADS50, OsMADS51, OsMADS56, OsSPL10, OsSPL13, OsBZR3, OsBTF3, OsGBP1, LOC_Os01g64360 and LOC_Os06g05350 (Fig. 4a, Fig. 5a and g, Supplementary Fig. 9). Several transcription factors including OsSPL13 and OsMADS56 have been reported to regulate grain size by influencing cell expansion in the grain hull5,11. Considering that SUG1 lacks predicted DNA-binding domains, it is possible that SUG1 may act as a cofactor to associate with distinct transcription factors to regulate grain size as well as other growth and developmental processes.
a SUG1 associates with OsSPL13 in N. benthamiana leaves. b SUG1 associates with SUG1 in vivo. GFP-OsSPL13 and MYC-SUG1 were transiently expressed in N. benthamiana leaves. Total proteins from leaves were isolated and incubated with GFP-Trap-A agarose beads. The precipitated proteins were detected using anti‐GFP and anti‐MYC antibodies, separately. IB: immunoblot. IP: immunoprecipitation. c OsSPL13 interacts with SUG1 in vitro. The purified GST-OsSPL13 was incubated with MBP-SUG1 and pulled down by Glutathione‐Sepharose beads. The precipitates were detected by immunoblot with anti‐GST and anti‐MBP antibodies. d The dual-luciferase assays shown the transcriptional activation activities of SPL13, and the SUG1 could enhance the transcriptional activation activities of SPL13 in rice protoplasts. The luciferase (LUC)/renilla (REN) activity were measured by co-transformation of different effector construct and reporter constructs. The activity of the empty vector was set to one. Data are presented as mean ± SD. Two-tailed unpaired Student’s t-test was used for statistical analysis. n = 3 biological replicates. e Mature rice grains of ZH11, proACTIN:OsSPL13, sug1-cr, and sug1-cr proACTIN:OsSPL13. f Relative transcript level of OsSPL13 of ZH11, proACTIN:OsSPL13, sug1-cr, and sug1-cr proACTIN:OsSPL13. n = 3 biological replicates. g–h Grain length (g) and grain width (h) of ZH11 (n = 50), proACTIN:OsSPL13 (n = 56), sug1-cr (n = 60), and sug1-cr proACTIN:OsSPL13 (n = 51). Data in (f–h) are presented as mean ± SD. Different letters represent significant differences (P < 0.05) from ordinary one-way ANOVA multiple comparisons with Tukey’s multiple comparisons test. Bar = 3 mm (e). Experiments in (a-d) were repeated independently at least twice with similar results. Source data are provided as a Source data file.
a SUG1 interacts with OsBZR1, OsBZR2, OsBZR3 and OsBZR4 in N. benthamiana leaves. b–c SUG1 is associated with OsBZR1 in vivo (b) and in vitro (c). d The second lamina joints of ZH11 and sug1-cr seedlings were observed after being treated with various concentrations of BL for 2 days. e–f Quantification of the lamina joint angle (e) and the coleoptile length (f) of ZH11 and sug1-cr. n = 20. g SUG1 interacts with OsMADS56 in N. benthamiana leaves. h–i SUG1 is associated with OsMADS56 in vivo (h) and in vitro (i). j The seedlings of ZH11 and sug1-cr were treated with various concentrations of GA3 for 7 days. k The second leaf sheath length of ZH11 and sug1-cr after being treated with various concentrations of GA3 for 7 days. n = 19. Data in (e–f, k) are presented as mean ± SD. Two-tailed unpaired Student’s t-test was used for statistical analysis. Bars: 1 cm (d, j). Experiments in (a–k) were repeated independently at least twice with similar results. Source data are provided as a Source data file.
SUG1 interacts with OsSPL13 to control grain length
OsSPL13 was identified as a SUG1-interacting protein in IP-MS (Supplementary Fig. 10). Interestingly, both OsSPL13 and SUG1 regulate grain size by promoting cell elongation in the grain hull, and osspl13 and sug1 mutants show similar grain size and shape phenotypes5. We therefore asked whether SUG1 could interact with OsSPL13 to control grain size and shape. To test this, we first confirmed the interaction between SUG1 and OsSPL13. As shown in Fig. 4a, the coexpression of cLUC-OsSPL13 with SUG1-nLUC showed strong luciferase activity in N. benthamiana leaves, indicating that OsSPL13 associates with SUG1 in planta. To further confirm the association of SUG1 with OsSPL13, we performed a co-immunoprecipitation assay by transiently expressing MYC-SUG1 with GFP-OsSPL13 in N. benthamiana leaves. Total proteins were extracted and incubated with GFP-Trap-A agarose beads. The immunoprecipitated proteins were detected using anti-GFP and anti-MYC antibodies, respectively. The results showed that MYC-SUG1 was co-immunoprecipitated with GFP-OsSPL13, but not with the negative control (GFP) (Fig. 4b). We also performed co-immunoprecipitation assay in rice protoplasts, and the results shown that SUG1 associates with OsSPL13 in vivo (Supplementary Fig. 11). Additionally, an in vitro pull-down assay was carried out to investigate whether SUG1 could directly interact with OsSPL13. Maltose binding protein (MBP) fused SUG1 (MBP-SUG1) and glutathione S-transferase (GST) fused OsSPL13 (GST-OsSPL13) proteins were expressed in Escherichia coli cells and purified, respectively. The results showed that MBP-SUG1 physically interacted with GST-OsSPL13 in vitro, but not the negative control (GST) (Fig. 4c). Thus, these findings demonstrated the interaction between SUG1 and OsSPL13 in vitro and in vivo. Previous studies reported that OsSPL13 acts as a transcriptional activator5. Thus, we performed the dual-luciferase assays to test the transcriptional regulatory relationship between OsSPL13 and SUG1. As shown in Fig. 4d, OsSPL13 have transcriptional activation activity, and SUG1 can enhance the transcriptional activation activity of SPL13.
Considering that SUG1 physically interacts with the OsSPL13 protein and promotes its transcriptional activation activity, we tried to explore the genetic relationship between SUG1 and OsSPL13 in grain size regulation. We constructed the CDS of OsSPL13 driven by the ACTIN promoter (proACTIN:OsSPL13) and introduced it into the ZH11 variety. The proACTIN:OsSPL13 transgenic plants produced longer grains than ZH11 (Supplementary Fig. 12). We then crossed sug1-cr with the proACTIN:OsSPL13 transgenic plants to generate the sug1-cr proACTIN:OsSPL13 lines (Fig. 4e). The expression level of OsSPL13 in sug1-cr proACTIN:OsSPL13 plants was similar to that in proACTIN:OsSPL13 (Fig. 4f). As shown in Fig. 4e, the sug1-cr mutation suppressed the long grain phenotype of proACTIN:OsSPL13, indicating that the long grain phenotype of proACTIN:OsSPL13 depends on the functional SUG1 (Fig. 4g-h).
SUG1 associates with OsBZR1 and is involved in BR responses
In IP-MS analysis, we found that SUG1 is associated with OsBZR3, which is a homologous protein of OsBZR1 (Supplementary Fig. 13). Overexpression of OsBZR1 resulted in increased grain size and weight, like those observed in proACTIN:SUG1 lines12,13,33. Considering that OsBZR1, OsBZR2, OsBZR3 and OsBZR4 may function redundantly to control grain size13, we asked whether SUG1 could associate with OsBZR1, OsBZR2, OsBZR3 and OsBZR4. As shown in Fig. 5a, SUG1 associates with OsBZR1/2/3/4 using luciferase complementation assay, respectively.
We then focused on testing the interaction between SUG1 and OsBZR1 because overexpression of OsBZR1 caused large and heavy grains12,13,33. We performed co-immunoprecipitation assay in N. benthamiana leaves and rice protoplasts, and the results showed that SUG1 associates with OsBZR1 in vivo (Fig. 5b and Supplementary Fig. 14). Furthermore, an in vitro pull-down assay was carried out to investigate whether SUG1 could directly interact with OsBZR1. The results shown that SUG1 can directly interact with OsBZR1 in vitro (Fig. 5c). These results demonstrated that SUG1 interacts with OsBZR1 in vitro and in vivo. Previous studies showed that OsBZR1 possesses transcriptional repression or activation activity, which depends on the target genes12,33,34,35,36,37,38,39. We performed the dual-luciferase experiments, and the result revealed that OsBZR1 has transcriptional repression activity at the overall level. However, SUG1 can not influence the transcriptional repression activity of OsBZR1 at the overall level (Supplementary Fig. 15). It is possible that the effect of SUG1 on the transcriptional activity of OsBZR1 depends on specific downstream genes or is context-dependent. Similar phenomena have reported in previous studies40,41.
Considering that SUG1 interacts with the OsBZR1, we tried to explore the genetic relationship between SUG1 and OsBZR1 in grain size regulation. The proACTIN:OsBZR1 transgenic plants produced longer grains than ZH11 (Supplementary Fig. 16a), consistent with previous studies33. The expression level of OsBZR1 in sug1-1 proActin:OsBZR1 plants was similar to that in proACTIN:OsBZR1 (Supplementary Fig. 16b). The sug1-1 mutation suppressed the long grain phenotype of proACTIN:OsBZR1, indicating that the long grain phenotype of proACTIN:OsBZR1 depends on the functional SUG1 (Supplementary Fig. 16c-e).
Considering that OsBZR1 is involved in BR signaling13,36,42, and SUG1 interacts with OsBZR1, we asked whether SUG1 influences BR responses. To test this possibility, we first performed a lamina joint bending experiment to test the BR sensitivity of sug1-cr. As shown in Fig. 5d-e, the sug1-cr mutant exhibited reduced bending compared to ZH11 in response to the active BR brassinolide (BL). In addition, the coleoptile elongation assay showed that sug1-cr displayed decreased sensitivity to BL treatment in comparison to the wild type (Fig. 5f). These findings indicated that sug1-cr is insensitive to BL compared with ZH11. We further analyzed the transcript levels of several genes involved in BR biosynthesis and signaling in young panicles of ZH11 and sug1-cr plants. The relative expression levels of BR biosynthetic genes such as BRD1 and DWARF4 were upregulated in sug1-cr panicles (Supplementary Fig. 17). This upregulation could be attributed to feedback regulation, as previously reported43,44,45. We also found that the expression levels of BR signal transduction genes like DLT, XIAO, XTR1, BU1, BRI1 and BZR1 were also significantly downregulated in sug1-cr panicles compared with those in wild-type panicles43,46,47,48,49 (Supplementary Fig. 17). Meanwhile, the expression level of the negative regulatory factor MDP1 in BR signaling pathway was increased in sug1-cr (Supplementary Fig. 17). Thus, these results showed that SUG1 is involved in BR responses.
SUG1 associates with OsMADS56 and is involved in GA responses
In IP-MS analysis, we found that SUG1 associates with OsMADS56 (Supplementary Fig. 18). OsMADS56 has been reported to regulate seed size by promoting cell expansion in the grain hulls50. Similarly, SUG1 promotes grain growth due to the increased cell expansion in the grain hull. As shown in Fig. 5g, SUG1 associates with OsMADS56 using luciferase complementation assay. We performed co-immunoprecipitation assay in N. benthamiana leaves and rice protoplasts, and the results showed that SUG1 associates with OsBZR1 in vivo (Fig. 5h and Supplementary Fig. 19). We further investigated the interaction between SUG1 and OsMADS56 using pull down assays, and the results showed that SUG1 can directly interact with OsMADS56 in vitro (Fig. 5i). These results demonstrated that SUG1 interacts with OsMADS56 in vitro and in vivo. We further examined the transcriptional activity of OsMADS56 in rice protoplasts using the dual-luciferase assay. The results showed that OsMADS56 has transcriptional activation activity, and SUG1 can enhance the transcriptional activation activity of OsMADS56 (Supplementary Fig. 20).
Considering that SUG1 interacts with the OsMADS56, we tried to explore the genetic relationship between SUG1 and OsMADS56 in grain size regulation. The proACTIN:OsMADS56 transgenic plants produced longer grains than ZH11 (Supplementary Fig. 21a), consistent with previous studies11,50. The expression level of OsMADS56 in sug1-1 proActin:OsMADS56 plants was similar to that in proACTIN:OsMADS56 (Supplementary Fig. 21b). The sug1-1 mutation suppressed the long grain phenotype of proACTIN:OsMADS56, indicating that the long grain phenotype of proACTIN:OsMADS56 depends on the functional SUG1 (Supplementary Fig. 21c–e).
OsMADS56 is an important regulator of GA homeostasis and signaling transduction50,51. Considering OsMADS56 positively controls grain size and physically interacts with SUG1, we wondered whether SUG1 participates in GA responses. We examined the sensitivity of ZH11 and sug1-cr seedlings to exogenous GA3. When treated with different concentrations of GA3, the second leaf sheath length of sug1-cr was significantly decreased compared with that of ZH11 (Fig. 5j, k). Moreover, we investigated the expression levels of genes involved in GA biosynthesis and signal transduction in young panicles of ZH11 and sug1-cr. Our results revealed that several GA biosynthetic genes, including CPS1, GA2OX4, GA2OX5, GA2OX6, were significantly increased in sug1-cr. Conversely, GA3OX2, GA20OX1, GA20OX2, GA20OX3 were found to be significantly reduced in sug1-cr (Supplementary Fig. 22a). Consistent with this, in GA-insensitive mutants, the expression levels of GA biosynthetic genes CPS1 and GA2OX genes were increased, while the GA3OX2 and GA20OX genes were reduced50,52. We also observed a significant increase in the expression of GA signal transduction genes, such as SLR1, WRKY36, and EUI1, and a notable decrease in GID2 expression in sug1-cr compared to the wild type (Supplementary Fig. 22b). The SLR1, WRKY36, and EUI1 are negative regulators of GA signaling, and the overexpression of WRKY36 resulted in a GA-insensitive phenotype, like those observed in sug1-cr53. Similarly, the expression of GID2, a positive regulator of GA signal transduction, was also reduced in the GA-insensitive mutant oscbl553. These results indicated that SUG1 is involved in GA responses.
Nucleotide polymorphism and selection signatures of SUG1
To investigate whether SUG1 is selected for rice domestication, we searched for the nucleotide polymorphism of the SUG1 genomic region (10.9 kb) in wild and cultivated rice using 446 O. rufipogon and 1083 O. sativa varieties54. The nucleotide diversity of the SUG1 genomic region in cultivated rice (π = 0.0011588) is lower than that in wild rice (π = 0.0024757) (Supplementary Fig. 23a), and the genetic diversity ratio of wild to cultivated rice varieties (πW: πC = 3.678) exceeded the 5% upper-quantile nucleotide polymorphism across the whole genome (πW: πC > 3)54 (Supplementary Fig. 23d). The 446 O.rufipogon samples were divided into three types based on genome sequence and structural variation, including Or-I, Or-II, and Or-III, among which indica and japonica are originated from Or-I and Or-III, respectively54. We next measured the nucleotide polymorphisms across SUG1 genomic region in Or-I, Or-III, indica and japonica varieties. The results showed that the nucleotide diversity of indica (π = 0.0004926) and japonica (π = 0.0000768) were much lower than that of Or-I (π = 0.0013209) and Or-III (π = 0.0020646), respectively (Supplementary Fig. 23b, c). The nucleotide diversity ratio of Or-III to japonica varieties (πOr-III: πjaponica = 17.416) was much higher than the genome-wide threshold of selection signals (πW: πC > 14)54 (Supplementary Fig. 23d). The genetic diversity ratio of Or-I to indica varieties (πOr-I: πindica = 4.247) also exceeded the genome-wide threshold of the selection signals (πW: πC > 3)54 (Supplementary Fig. 23d). These analysis results implied that the SUG1 genomic region could have selective signatures during rice domestication. To further investigate the evolutionary signature of SUG1, we calculated the fixation index (FST) of the SUG1 genomic region (10.9 kb). The results displayed that there was an obviously high degree of differentiation between the japonica and indica populations (Supplementary Fig. 24).
To investigate whether SUG1 has been utilized during rice genetic improvement, we analyzed the haplotypes of the SUG1 genomic region using a dataset of 1,641 rice cultivars obtained from the 3 K Rice Genomes Project55. In cultivated rice, a total of 28 variants were identified, including 13 SNPs in the 2,330 bp promoter region, 4 SNPs in the coding region, and 10 SNPs and 1 indel in intron, which classified SUG1 into four major haplotypes Hap1-Hap4 (Fig. 6a). Despite the existence of the diverse haplotypes in the cultivated rice, these variants did not lead to frame-shifts or changes to conserved sites within the functional domains (Fig. 6a). As shown in Fig. 6b, the SUG1 gene in the indica group mainly consists of two haplotypes, Hap1 ((448/1016, 44%) and Hap2 (498/1016, 49%)). In Japonica accessions, the SUG1 gene mainly contains Hap3 (466/625, 75%) and Hap4 (131/625, 21%). To determine whether the major four SUG1 haplotypes have been utilized in rice genetic breeding, we further analyzed the distribution of four haplotypes using landrace and improved accessions56. We found that SUG1Hap2 accounts for 25.5% of rice landrace, while it increases to 50.3% in improved varieties. The similar trend was observed in indica varieties, with SUG1Hap2 accounting for 48.6% in landrace and 72.3% in improved varieties (Supplementary Fig. 25). It is possible that SUG1Hap2 has been selected during the deliberate breeding selection of rice indica varieties for long grain alleles, thereby increasing its proportion in the indica subspecies. Interestingly, japonica landrance with the SUG1Hap2 allele only accounts for 1.45%, and improved japonica varieties with the SUG1Hap2 allele is about 3.3%. These results suggested that there is a great potential to improve grain size in japonica varieties using the SUG1Hap2 allele.
a Haplotype analysis of SUG1 from 1,641 Asian cultivated rice accessions. b The distribution frequency of the four SUG1 haplotypes in diverse Asian cultivated rice varieties (n = 462/512/492/175, 448/498/26/44, 14/14/466/131). Source data are provided as a Source data file. c The grain length was analyzed in 1253 Asian cultivated rice varieties (with phenotype data) by SUG1 haplotypes (n = 408/387/307/151). Data are presented as mean ± SD. Different letters represent significant differences (P < 0.05) from ordinary one-way ANOVA multiple comparisons with Tukey’s multiple comparisons test. Source data are provided as a Source data file. d Mature grains of ZH11 and transgenic lines ZH11;gSUG19311 (three independent transgenic lines). Bar = 3 mm. e–i The grain length (e), grain width (f), 1,000 grain weight (g), and grain yield per plant (h) and grain yield per plot (i) of ZH11 and transgenic lines ZH11;gSUG19311. The field trials were conducted in Lingshui, Hainan in 2024. Data are presented as mean ± SD. Two-tailed unpaired Student’s t-test was used for statistical analysis. n = 193/169/216/104 in (e-f) n = 3 in (g) n = 24/20/20/20 in (h) and n = 5 in (i). Source data are provided as a Source data file. j A proposed molecular framework for the GS2-SUG1 module-mediated grain size control. SUG1 positively regulates grain size. GS2 directly binds to the promoter region of SUG1, and OsGIF1 interacts with GS2 to promotes the expression of SUG1. SUG1 interacts with a range of transcription factors, such as OsSPL13, OsBZR1, and OsMADS56, to regulate grain size through BR and GA signaling as well as growth pathways.
The SUG1 Hap2 allele from 9311 improves grain size and grain yield of ZH11
Considering that SUG1 predominantly influences grain length, we therefore compared the grain length distribution of the Hap1-Hap4 representative varieties using the phenotypic data provided by the 3 K Rice Genomes Project55. The results showed that the grain length of the representative varieties with Hap2 was obviously higher than that of Hap3 (Fig. 6c). The indica variety 9311 belongs to Hap2, and the Zhonghua 11 (ZH11) and Nipponbare (NIP) are Hap3. To further assess the functional difference between SUG1Hap2 and SUG1Hap3, we amplified the genomic fragment that consisted of the 2,330 bp of 5’ flanking sequence, the SUG1 gene, and 639 bp of 3’ flanking sequence from the 9311 (Hap2) and ZH11 (Hap3) variety. This fragment was then introduced into ZH11 (Hap3) to generate the gSUG19311 and gSUG1ZH11 transgenic plants, respectively. We measured grain size of 12 independent ZH11;gSUG1ZH11 lines and 15 independent ZH11;gSUG19311 lines. The results showed that the ZH11;gSUG19311 transgenic plants exhibited longer grains than ZH11 and ZH11;gSUG1ZH11 transgenic plants, while the grain width of the ZH11;gSUG19311 transgenic plants was similar to that of the ZH11 and ZH11;gSUG1ZH11 transgenic plants (Supplementary Fig. 26 and Fig. 6d-g). Therefore, we further used three independent ZH11;gSUG1ZH11 lines and three independent ZH11;gSUG1ZH11 lines for subsequent field yield test. The grain yield per plant and grain yield per plot of gSUG19311 showed a significant increase compared to that of gSUG1ZH11 and ZH11(Fig. 6h-i and Supplementary Fig. 27). These findings indicated that the SUG1Hap2 allele from indica varieties can be used to improve grain size and yield of the japonica varieties with the SUG1Hap3 allele.
Discussion
Grain size is one of the key components of grain yield in rice. Understanding the molecular mechanisms of grain size control will be vital for improving grain yield. We have previously reported that the transcription factor GS2 regulates grain size and yield15. GS2 has been reported to increase nitrogen utilization and yield and is proposed to be a new green revolution gene20. However, how GS2 regulates grain size remains unclear. Here, we identified the suppressor of GS2AA (SUG1) that functions as a transcription regulator. The sug1 can partially suppress the grain length phenotype of the GS2AA allele (Fig. 1a–c and g–k). SUG1 encodes plant specific protein DEP2/EP2/SRS1/OsRELA, which has been described to influence panicle shape, grain shape and leaf angle24,25,26,27. However, the genetic and molecular mechanisms of SUG1 in grain size regulation are still unclear. Our results show that sug1 mutants produce short and wide grains, while plants overexpressing SUG1 form long grains, indicating that SUG1 is a positive regulator of grain length (Fig. 2a, f–i, p–t). Cellular observation indicates that SUG1 regulates grain size by affecting cell expansion in the grain hull (Fig. 2d-e, l-o, Supplementary Fig. 7h, i). The genetic analyses show that sug1-1 is partially epistatic to GS2AA in terms of grain length (Fig. 3a, b). We further demonstrated that GS2 binds to the promoter of SUG1 and promotes its expression (Fig. 3i-l). Supporting this, GS2 and SUG1 have similar expression patterns during panicle development (Supplementary Fig. 3b, c)16,19. The transcription coactivator OsGIF1 physically interacts with GS2 to influence grain size15. Consistent with this, our genetic analyses indicated that SUG1 and OsGIF1 have overlapped functions in grain length and grain shape control (Fig. 3e–h). Furthermore, the transcription activation experiment indicates that GS2 can activate the transcription of the SUG1 promoter, and OsGIF1 enhances GS2’s activation of the SUG1 promoter (Fig. 3m). Thus, our findings revealed that SUG1 acts downstream of GS2 and OsGIF1 and is a direct target of GS2 in grain size control (Fig. 6j).
SUG1 is a plant specific protein, and its molecular and biochemical functions are totally unknown. In this study, we identified SUG1-interacting proteins using IP-MS. Interestingly, 33 transcription regulators were identified in this IP-MS assay, including MADS, SPL, MYB, WRKY and bZIP-type transcription factors, suggesting that SUG1 may function as a transcriptional regulator (Supplementary Fig. 9, Supplementary Table 3). Among SUG1-associated proteins, several transcriptional factors (e.g. OsSPL13, OsBZR1/2/3/4, and OsMADS56) have been shown to regulate grain size5,13,50. The transcription factor OsSPL13 has been reported to control grain size by regulating cell expansion in the grain hull5. The spl13 mutant showed similar phenotypes to sug1 mutants5. In this study, we demonstrate that SUG1 physically interacts with OsSPL13 in vivo and in vitro (Fig. 4a–c). Genetic analyses support that SUG1 and OsSPL13 have overlapped function in grain length control (Fig. 4e–g). OsSPL13 has been proposed to regulate cell expansion possibly by influencing microtubule and cell wall pathways5. It will be a valuable challenge to investigate if SUG1 regulates microtubule and cell wall pathways in the future. OsBZRs are the central transcriptional factors in the BR signaling pathway. OsBZR1 acts redundantly with other family members to regulate grain size13. Here we uncover that SUG1 can interact with OsBZR1 (Fig. 5a–c). The sug1-cr exhibited decreased BR sensitivity compared with the wild type (Fig. 5d–f). Consistent with this, the expression of BR biosynthetic and signaling-related genes were changed in sug1-cr (Supplementary Fig. 17). It is plausible that SUG1 regulates grain size and BR response, at least in part, by interacting with OsBZR1 and family members. OsMADS56 has been described to control grain size by regulating GA signaling50. Our biochemical data supports that SUG1 physically interacts with OsMADS56 in vitro and in vivo (Fig. 5g–i). The sug1-cr mutant is insensitive to exogenous GA3 and changes the expression of several GA-related genes (Fig. 5j–k). It is plausible that the SUG1 influences grain size and GA response partially by interacting with OsMADS56. We also found that SUG1 enhances the transcriptional activation activities of OsSPL13 and OsMADS56 (Fig. 4d and Supplementary Fig. 20). However, the SUG1 had no effect on the transcriptional repression activity of OsBZR1 at the overall level (Supplementary Fig. 15). Its possible that the effect of SUG1 on the transcriptional activity of OsBZR1 was context dependent, tissue specific or depended on specific downstream genes12,35,36,38,39,57,58,59. Thus, our results reveal a previously unknown mechanism by which SUG1 acts as a hub to integrate multiple transcription factors, which are involved in several phytohormone signaling and growth and development processes, to control grain growth in rice (Fig. 6j).
As two major subspecies of Asian cultivated rice, indica and japonica have produced significant differences in grain size and shape due to long-term natural and artificial selection54,60. indica varieties usually produce long grains, while japonica varieties form short grains. Recent studies reveal that several genes, such as GSE5/GW5 and GS3, contribute to grain size and shape domestication in indica and japonica populations60,61,62. In this study, we showed that the nucleotide polymorphisms of the SUG1 locus decreased significantly compared with those of the wild rice Or-III, suggesting that SUG1 locus has been subjected to artificial selection during rice domestication. Furthermore, haplotype analysis revealed that SUG1Hap2 may have been widely utilized by deliberate breeding selection of rice indica varieties for larger grain alleles (Supplementary Fig. 25), while SUG1Hap2 has not yet been widely used for breeding selection of japonica varieties to obtain larger grain alleles. We further demonstrate that the natural allele (SUG1Hap2) from indica variety 9311 can be used to improve grain size and yield of japonica variety ZH11 (Fig. 6d–i and Supplementary Figs. 26 and 27). Plants carrying gSUG19311 in ZH11 background exhibited increased grain length and grain yield compared with ZH11 (Fig. 6d–i and Supplementary Figs. 26 and 27). It will be promising to utilize the SUG1Hap2 allele from indica varieties to improve grain size and yield of japonica varieties in the future. Considering that homologs of SUG1 exist in other crops, it is a worthwhile challenge to identify their beneficial alleles and improve grain yield in key crops.
Methods
Plant materials and growth conditions
ZH11 (a japonica variety) and HZ (an indica variety) were used as the wild type. ZH11-GS2AA is the near-isogenic line of GS2AA in ZH11 background16 and HZ-GS2AA is the near-isogenic line of GS2AA in HZ background15. The mutant sug1-1 was isolated from an M2 population of ZH11-GS2AA (EMS-treated), and sug1-2 was isolated from an M2 population of HZ-GS2AA (EMS-treated). Rice (Oryza sativa) plants were cultivated in open fields at Lingshui, Hangzhou, and Beijing during the natural growing season. N. benthamiana plants were grown in a greenhouse with 16 h light and 8 h dark.
Morphological analysis
The plants and grains were taken photographs and measured after maturity. Grains from the main panicles were scanned using a scanner (MICROTEK, Scan Marker i560), and then grain length and grain width were measured by the Rice Test System (WSEEN). To calculate the grain weight, one hundred dry grains were weighed, and the average value was determined based on three biological replicates. Grain thickness was evaluated using a vernier caliper, with at least 50 seeds being counted for each sample.
Cellular Analysis
Scanning electron microscopy (SEM) was used for cellular analysis by scanning the matured grains. Outer epidermal cell size and inner epidermal cell size of the central part of the lemmas were measured using Image J software. The widest part of the grain was counted to obtain the cell number in the grain-width direction and the longest part of the grain was used to count the cell number in the grain-length direction.
Identification of the SUG1 gene
To generate F2 populations, sug1-1 ZH11-GS2AA and sug1-2 HZ-GS2AA were crossed with ZH11-GS2AA and HZ-GS2AA, respectively. Subsequently, we proceeded to clone the SUG1 gene using these F2 populations. We used the NextSeq 500 system (Illumina) to resequence the genomes of ZH11-GS2AA and plants displaying the sug1-1 ZH11-GS2AA mutant phenotypes (50 individuals mixture for each sample). Similarly, we resequenced the genomes of HZ-GS2AA and plants displaying sug1-2 HZ-GS2AA mutant phenotypes (50 individuals mixture for each sample). To identify potential causal genes, we analyzed the SNP/INDEL index by the MutMap methods, as previous study22,63. By aligning the reads to the Nipponbare and 9311 reference sequence, we obtained the special SNPs and INDELs and calculated the corresponding SNP/INDEL-index. Then, the SNPs or INDELs with a SNP/INDEL-index of 1 were selected for subsequent sequence analyses.
Vector construction and transformation
We constructed plasmids using Uniclone One Step Seamless Cloning Kit (Genesand, SC612). The primers gSUG1-F and gSUG1-R were used to amplify genomic sequence of SUG1. Then we inserted the SUG1 genomic sequence into the pMDC99 vector to construct the gSUG1 plasmid. We amplified coding sequence of the SUG1 using primer pair ACTIN-SUG1-F and ACTIN-SUG1-R and inserted it into the pIPKb003 vector to generate the proACTIN:SUG1 construct. We amplified the coding sequence (CDS) of the SUG1 by the primers GFP-SUG1-F and GFP-SUG1-R and inserted it into the pMDC43 vector for constructing the pro35S:GFP-SUG1 plasmid. The promoter of SUG1 (2330 bp) was amplified and inserted into the pMDC164 vector to generate the proSUG1:GUS plasmid by primer pair pMDC164-SUG1-F and pMDC164-SUG1-R. For pro35S:MYC-rGS2 construction, we adopted an overlapping PCR approach by using the primers rGS2-MYC-F, rGS2-MYC-R, rGS2-F and rGS2-R to produce a miR396-resistant variant of GS2 that does not cause amino acids change. The primers SUG1-CRI-F and SUG1-CRI-R were used to generate the sug1-cr plasmid, and the 20 bp target site sequence was AGCTGGAGGGGGTTCAAGAC. The CDS of the OsSPL13 was amplified using primers ACTIN-SPL13-F and ACTIN-SPL13-R, and inserted into the pIPKb003 vector to generate the proACTIN:OsSPL13 construct. All the constructs were transformed into the japonica variety ZH11 using Agrobacterium Tumefaciens GV3101 to obtain the transgenic plants. Primer sequences used for PCR amplification were listed in the Supplementary Data 1.
RNA extraction and quantitative Real-time PCR (qRT-PCR)
The total RNA was extracted from rice young panicles using the RNAprep pure kit (TIANGEN, DP439). The first strand cDNA was produced using cDNA Synthesis Kit (Vazyme, R211). Quantitative real-time PCR was completed using SYBR qPCR Mix (Genstar, A301-10) by a Lightcycler 480 (Roche,Switzerland). Rice ACTIN1 was used as an internal control. Primers used were listed in Supplementary Data 1.
GUS staining
Different lengths of developing panicles and spikelet hulls of proSUG1:GUS plants were stained in the GUS staining buffer solution (0.1% Triton X-100, 50 mM NaPO4, 0.4 mM each K3Fe(CN)6/K4Fe(CN)6 and 1 mM X-Gluc). Samples were incubated at 37°C for 6 h after being vacuumed for 30 mins. After that, 70% ethanol was used to remove chlorophyll. Then the samples were photographed with a Nikon D7100.
Subcellular localization of OsSUG1
To investigate the subcellular localization of OsSUG1 in rice, we analyzed the GFP fluorescence from the roots of 7 day-old pro35S:GFP-SUG1 transgenic seedlings. Laser scanning confocal microscopy (Zeiss LSM710) was used to visualize the GFP fluorescence after treating the samples with the nuclear staining buffer DAPI for 20 min.
Firefly luciferase complementation assay
The coding region sequence of SUG1 was amplified and inserted into the pCAMBIA-split_nLUC vector (digested with BamHI and SalI) to generate SUG1-nLUC. The cDNA sequences of OsBZR1, OsBZR2, OsBZR3, OsBZR4, OsSPL10, OsSPL13, OsMADS2, OsMADS4, OsMADS15, OsMADS22, OsMADS50, OsMADS51, OsMADS56, OsBTF3, OsGBP1, LOC_Os01g64360, LOC_Os06g05350 were amplified and fused to the pCAMBIA-split_cLUC vector (digested with BamHI and SalI) to generate plasmids including cLUC-OsBZR1, cLUC-OsBZR2, cLUC-OsBZR3, cLUC-OsBZR4, cLUC-OsSPL10, cLUC-OsSPL13, cLUC-OsMADS2, cLUC-OsMADS4, cLUC-OsMADS15, cLUC-OsMADS22, cLUC-OsMADS50, cLUC-OsMADS51, cLUC-OsMADS56, cLUC-OsBTF3, cLUC-OsGBP1, cLUC-LOC_Os01g64360, cLUC-LOC_Os06g05350, respectively. N. benthamiana leaves were co-transformed with various combinations of plasmids as indicated by Agrobacterium tumefaciens strain GV3101. Following a 48-h incubation, NightSHADE LB 983 imaging apparatus was used to measure luciferase intensity64. According to previous studies65,66, we selected GUS as negative control. All primers required for these constructs are listed in Supplementary Data 1.
Pull-Down assay
The CDS of SUG1 was amplified and inserted into the pMAL-C2 to generate MBP-SUG1 construct. The CDSs of OsBZR1, OsSPL13 and OsMADS56 were amplified and cloned into the pGEX4T-1 to generate GST-OsBZR1, GST-OsSPL13 and GST-OsMADS56 plasmid, respectively. To test the interactions, the purified recombinant proteins as indicated were incubated in TGH buffer for a duration of 30 min at 4 °C with Glutathione Sepharose 4B beads (GE Healthcare, USA). After washed with TGH buffer for 5 times, the beads were added SDS-loading buffer and denatured at 98 °C for 10 min. Then, the corresponding samples were separated and analyzed by anti-MBP (NEB, E8032, dilution, 1:10000) and anti-GST (Abmart, M20007, dilution, 1:5000), respectively. The results were photoed using the Tanon-4500 gel imaging system.
Co-immunoprecipitation assay
We amplified the coding region of SUG1 and inserted it into pCambial1300-221-MYC vector to generate pro35S:MYC-SUG1 construct. The CDSs of OsMADS56, OsBZR1 and OsSPL13 were cloned into the pMDC43 vector to produce pro35S:GFP-OsMADS56, pro35S:GFP-OsBZR1 and pro35S:GFP-OsSPL13 plasmids, respectively. Primers used for these constructs were listed in Supplementary Data 1. The plasmids pMDC43, pro35S:GFP-OsMADS56, pro35S:GFP-OsBZR1, pro35S:GFP-OsSPL13 and pro35S:MYC-SUG1 were individually introduced into the GV3101. Different combinations of activated GV3101 with plasmids were injected into N. benthaminana leaves for incubation about 48 h. After that, the leaves were collected to extract total proteins with extraction buffer (NaCl (150 mM), Tris-HCl pH 7.5 (50 mM), EDTA (1 mM), Triton X-100 (1%), glycerol (5%) and 1× Roche protease inhibitor cocktail). The proteins were then immunoprecipitated using 20 µL GFP-Trap-Agrose (Chromotek, gta-20) at 4°C for 30 mins, and washed at least five times with the Wash buffer (NaCl (150 mM), Tris-HCl pH 7.5 (50 mM), EDTA (1 mM), glycerol (5%), Triton X-100 (0.2%) and 1× Roche protease inhibitor cocktail). Then, the beads were added 1× SDS loading buffer and boiled at 95 °C for 10 min. The eluted proteins were separated in the SDS-polyacrylamide gel and detected by immunoblotting using an anti-GFP antibody (Abmart, Cat.No: M20004, dilution, 1:5000) and anti-MYC antibody (Abmart, Cat. No: M20002, dilution, 1:5000), respectively.
Electrophoretic mobility shift assay (EMSA)
In order to construct the GST-GS2 plasmid, we amplified the CDS of GS2 using the GST-GS2-F and GST-GS2-R primers, and inserted it into the pGEX4T-1 vector. The GST and GST-GS2 recombinant proteins were expressed by Escherichia coli BL21 (DE3) strain, and purified using Glutathione Sepharose 4B beads (GE Healthcare, 17-0756-01). For the generation of double-stranded biotinlabeled DNA probes and non-biotinlabeled DNA probes, the sense and antisense oligonucleotides were annealed. The annealing reactions were carried out using PCR thermocycler with an initial temperature of 95 °C for 10 min, followed by a gradual decrease of 0.5 °C per minute until reaching 25 °C. EMSA was accomplished using the Light Shift Chemiluminescent EMSA Kit (Thermo Fisher Scientific, 20148) according to the instructions. The primers were listed in Supplementary Data 1.
Chromatin immunoprecipitation and quantitative real-time PCR analysis
The ChIP assay was conducted on the basis of the previous study with some modifications67. Approximately 1.5 g young panicles from transgenic plants pro35S:MYC or pro35S:MYC-rGS2 were fixed with 1% formaldehyde for 15 mins under vacuum at room temperature. The termination of the crosslinking process was achieved by adding 0.125 M glycine and incubating for 5 mins. The panicles were grounded in the liquid nitrogen for isolating nuclei. Upon lysing of the nuclei, the chromatin complexes were extracted and fragmented into pieces of ~500 bp through sonication (Bioruptor pico, Diagenode). 10 ug anti-MYC antibody (Abcam, Cat. No:ab32) and 20 ul protein A + G beads (Millipore, 16-663) were used for immunoprecipitations. The QIAGEN DNA purification kit (QIAGEN, 28104) was used to recover the precipitated DNA for the following qRT-PCR. The primers were listed in Supplementary Data 1.
Immunoprecipitation in combination with mass spectrometry (IP-MS)
The SUG1 interacting protein screen was conducted using immunoprecipitation combined with mass spectrometry (IP-MS). We obtained total proteins from the mixed 3-5 cm young panicles of pro35S:MYC-SUG1 transgenic plants, and the young panicles of pro35S:12×MYC transgenic plants were act as control. The extracted proteins were then precipitated by anti-MYC-Tag mAb Agarose Conjugated beads (Abmart, M20012) and washed with Buffer (NaCl (150 mM), Tris-HCl (10 mM), NP-40 (0.5%), EDTA (0.5 mM), 1 × protease inhibitor cocktail and PMSF (1 mM)) for five times. Subsequently, the beads were added SDS buffer (4% SDS, 100 mM Tris-HCl pH7.5) and boiled for 20 mins. The proteins were then digested using the filter-aided sample preparation (FASP) method68, and underwent analysis as previously showed69. For MS analyses, peptides were analyzed by LTQ Orbitrap Elite mass spectrometer (Thermo Fisher Scientific) coupled online to an Easy-nLC 1000 (Thermo Fisher Scientific) in the data-dependent mode. The MS data was analyzed using a pre-release version of Thermo Scientific Proteome DiscovererTM software version 1.4, The proteome sequences for Oryza sativa from IRGSP (http://rice.plantbiology.msu.edu/) were used for the database searching. Information about interacting transcription factors of SUG1 by IP-MS was summarized in Supplementary Table 3.
Brassinosteroid treatment
The seeds of ZH11 and sug1-cr were surface sterilized with NaClO (30%) and washed with sterilized water for five times before they were germinated at 28°C. For the lamina inclination test, the seedlings were cultured with the 96-well PCR plates that the bottoms were cut off. When the second leaf grew out with the blade slightly spreading out but still vertical, different concentrations of 0, 0.01, 0.1 and 1 μM BL (Yuanye, S18015) were added to the adaxial side of the leaf blade tip. After 3 days of normal growth, the plants were photographed, and the leaf angles were measured using ImageJ. For the coleoptile elongation assay, seeds were grown on 0.3% agar medium with various concentrations of 0, 0.01, 0.1, 1 and 10 μM BL for 1 week under continuous darkness at 28°C. Then, the coleoptile length were measured.
Gibberellin treatment
Germinated seeds were pretreated with 10 μM uniconazole (Yuanye, S18186) for 1 days to inhibit endogenous GA biosynthesis, then grown on 96-well plates in distilled, deionized H2O with different concentrations of GA3 (Yuanye, S28506) for 7 days. The seedlings were photographed, and the second leaf sheath length were measured using Image J.
Rice protoplast preparation and dual-luciferase assays
We conducted rice protoplast preparation and transformation as described in a previous study70. Rice protoplasts were prepared using ZH11 seedlings grown for 10–14 days at 28°C. Briefly, the stems of well-grown ZH11 were thinly sliced and incubated with enzyme solution (Cellulose RS (1.5%), Macerozyme R-10 (0.75%), MES pH 5.7 (10 mM), mannitol (0.6 M), BSA (0.1%), and CaCl2 (10 mM)) for about 4 h in the dark. Then an equal volume of W5 solution (KCl (5 mM), NaCl (154 mM), MES pH 5.7 (2 mM) and CaCl2 (125 mM)) was added and gently shaken to fully release the protoplasts. The protoplasts were collected using a miracloth (AMRESCO, 1363488), washed 2 times with W5 solution and then suspended in MMG solution (MES pH5.7 (4 mM), mannitol (0.4 M), MgCl2 (15 mM)). Transient transfection of protoplasts was performed by a PEG-mediated method. We used a DLR assay to evaluate the effect of SUG1 on the transcriptional activity of OsSPL13, OsBZR1 and OsMADS56 in rice protoplasts, as previously study71. We used pPTRL plasmid as an internal control, which contains Renilla LUC driven by 35S promoter. The protoplast transformation for each combination contained 6 μg of effectors, 6 μg of reporter, and 1 μg of pPTRL. The firefly luciferase activity and Renilla luciferase activity were measured using a DLR assay kit in a luminometer device (Promega) after the transformed protoplasts were incubated for 16 h at 28°C.
Haplotype analysis of SUG1
To investigate natural variation of SUG1, a dataset of 1,641 cultivated rice accessions consisting of 1016 indica varieties and 625 japonica varieties was used for haplotype analysis55. A dataset of the 332 Oryza sativa accessions (141 landraces and 191 improved varieties) was used for genetic improvement analysis56. The SNPs and Indels in the genomic regions (10.9 kb including the 2330 bp promoter and 3’UTR sequence) of SUG1 from 1,641 cultivar accessions were acquired from 3000 Rice Genome Project (https://snp-seek.irri.org/) for haplotype classification using DnaSP 5.10.01 software. By classification, four major haplotypes (Hap1 to Hap4) were identified in 1,641 cultivated rice. The rest were rare types, which were not considered.
Nucleotide diversity and fixation index calculation analysis
To examine whether SUG1 is selected during rice domestication, we used the published data for evolutionary analysis, including the following six groups: 446 wild rice species (155 wild_Or-I, 121 wild_Or-II and 170 wild_Or-III), 1083 cultivated rice species (484 japonica, 520 indica, 30 aus, 5 aromotic and 44 intermedia), 155 wild_Or-I, 170 wild_Or-III, 484 japonica species and 520 indica species54. SNPs for the nucleotide polymorphism analysis were filtered with a minor allele frequency (MAF) of >5% according to the published paper55. Missing genotypes were imputed using Beagle 5.4 software (https://faculty.washington.edu/browning/ beagle/beagle.html). VCFtools software (https://vcftools.sourceforge.net/) was used to calculate the nucleotide diversity (π), and fixation index (FST) for each subgroup with a window size of 50 bp and a step of 25 bp across the SUG1 locus (Chr. 7: 26,039,855 bp to 26,050,689 bp). Based on the published articles, the sequence diversity (π) of O. rufipogon was estimated at ∼0.003, and the sequence diversity is 0.0024 for O. sativa, and 0.0016 and 0.0006 for indica and japonica, respectively54. FST > 0.3, which covered ~3% of the complete rice genome, represents highly differentiated loci.
Statistics & Reproducibility
All data shown as the mean ± SD (standard deviation), unless indicated otherwise. Statistical analysis was performed using GraphPad Prism 8 software (GraphPad Software, Inc. San Diego, CA, USA). All details on statistics have been indicated in figure legends. The P-value was calculated using the two-tailed unpaired Student’s t-test. No statistical method was used to predetermine sample size. Images were analyzed with ImageJ.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data and materials are available from the corresponding authors upon request. The authors declare that all data supporting the findings of this study are available within the article and Supplementary Information files. The raw sequence data for MutMap generated in this study have been deposited in the Genome Sequence Archive database (accession number, GSA: CRA016489) [https://ngdc.cncb.ac.cn/gsa/browse/CRA016489] in National Genomics Data Center, China National Center for Bioinformation. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium (accession number: PXD052270). Sequence details from this study can be found in China Rice Data Center (https://www.ricedata.cn/gene/) with the following accession numbers: SUG1 (LOC_Os07g42410), GS2 (LOC_Os02g47280), OsGIF1 (LOC_Os03g52320), OsSPL13 (LOC_Os07g32170), OsBZR1 (LOC_Os07g39220), OsBZR2 (LOC_Os01g10610), OsBZR3 (LOC_Os06g35900), OsBZR4 (LOC_Os02g13900), OsMADS56 (LOC_Os10g39130). Source data are provided with this paper.
References
Zhang, Q. Strategies for developing green super rice. Proc. Natl Acad. Sci. USA 104, 16402–16409 (2007).
Li, N., Xu, R. & Li, Y. Molecular networks of seed size control in plants. Annu. Rev. Plant. Biol. 70, 435–463 (2019).
Zuo, J. & Li, J. Molecular genetic dissection of quantitative trait loci regulating rice grain size. Annu. Rev. Genet. 48, 99–118 (2014).
Guo, T. et al. Advances in rice genetics and breeding by molecular design in China. Sci. Sin. Viate 49, 1185–1212 (2019).
Si, L. et al. OsSPL13 controls grain size in cultivated rice. Nat. Genet. 48, 447–456 (2016).
Wang, S. et al. Control of grain size, shape and quality by OsSPL16 in rice. Nat. Genet. 44, 950–954 (2012).
Yuan, H. et al. OsSPL18 controls grain weight and grain number in rice. J. Genet. Genomics 46, 41–51 (2019).
Hu, J. et al. The elite alleles of OsSPL4 regulate grain size and increase grain yield in rice. Rice 14, 90 (2021).
Zhang, X. F., Yang, C. Y., Lin, H. X., Wang, J. W. & Xue, H. W. Rice SPL12 coevolved with GW5 to determine grain shape. Sci. Bull. 66, 2353–2357 (2021).
Huang, Y. et al. Wide Grain 7 increases grain width by enhancing H3K4me3 enrichment in the OsMADS1 promoter in rice (Oryza sativa L). Plant J. 102, 517–528 (2020).
Zuo, Z. W. et al. Control of thousand-grain weight by OsMADS56 in rice. Int. J. Mol. Sci. 23, 125 (2021).
Zhu, X. et al. Brassinosteroids promote development of rice pollen grains and seeds by triggering expression of Carbon Starved Anther, a MYB domain protein. Plant J. 82, 570–581 (2015).
Liu, D. et al. Diversification of plant agronomic traits by genome editing of brassinosteroid signaling family genes in rice. Plant Physiol. 187, 2563–2576 (2021).
Che, R. et al. Control of grain size and rice yield by GL2-mediated brassinosteroid responses. Nat. Plants 2, 15195 (2015).
Duan, P. et al. Regulation of OsGRF4 by OsmiR396 controls grain size and yield in rice. Nat. Plants 2, 15203 (2015).
Hu, J. et al. A rare allele of GS2 enhances grain size and grain yield in rice. Mol. Plant 8, 1455–1465 (2015).
Li, S. et al. The OsmiR396c-OsGRF4-OsGIF1 regulatory module determines grain size and yield in rice. Plant Biotechnol. J. 14, 2134–2146 (2016).
Sun, P. et al. OsGRF4 controls grain shape, panicle length and seed shattering in rice. J. Integr. Plant Biol. 58, 836–847 (2016).
Chen, X. et al. A missense mutation in large grain size 1 increases grain size and enhances cold tolerance in rice. J. Exp. Bot. 70, 3851–3866 (2019).
Li, S. et al. Modulating plant growth-metabolism coordination for sustainable agriculture. Nature 560, 595–600 (2018).
Gao, Y. et al. MYB61 is regulated by GRF4 and promotes nitrogen utilization and biomass production in rice. Nat. Commun. 11, 5219 (2020).
Abe, A. et al. Genome sequencing reveals agronomically important loci in rice using MutMap. Nat. Biotechnol. 30, 174–178 (2012).
Yamamoto, Y. Y., Matsui, M., Ang, L. H. & Deng, X. W. Role of a COP1 interactive protein in mediating light-regulated gene expression in arabidopsis. Plant Cell 10, 1083–1094 (1998).
Abe, Y. et al. The small and round seed1 (SRS1/DEP2) gene is involved in the regulation of seed size in rice. Genes Genet. Syst. 85, 327–339 (2010).
Li, F. et al. Rice DENSE AND ERECT PANICLE 2 is essential for determining panicle outgrowth and elongation. Cell Res. 20, 838–849 (2010).
Zhu, K. et al. Erect panicle2 encodes a novel protein that regulates panicle erectness in indica rice. Genetics 184, 343–350 (2010).
Zhu, C. L. et al. OsRELA Regulates leaf inclination by repressing the transcriptional activity of OsLIC in rice. Front. Plant Sci. 12, 760041 (2021).
Huang, K. et al. Modulation of histone acetylation enables fully mechanized hybrid rice breeding. Nat. Plants 10, 954–970 (2024).
Tang, S. et al. An E2-E3 pair contributes to seed size control in grain crops. Nat. Commun. 14, 3091 (2023).
Bai, C. et al. OsMAPK6 phosphorylation and CLG1 ubiquitylation of GW6a non-additively enhance rice grain size through stabilization of the substrate. Nat. Commun. 15, 4300 (2024).
Duan, E. et al. The transcriptional hub SHORT INTERNODES1 integrates hormone signals to orchestrate rice growth and development. Plant Cell 35, 2871–2886 (2023).
Omidbakhshfard, M. A. et al. GROWTH-REGULATING FACTOR 9 negatively regulates arabidopsis leaf growth by controlling ORG3 and restricting cell proliferation in leaf primordia. PLoS Genet 14, e1007484 (2018).
Du, H. et al. OsBAK2/OsSERK2 expression is repressed by OsBZR1 to modulate brassinosteroid response and grain length in rice. J. Exp. Bot. 74, 4978–4993 (2023).
Ke, Y. et al. The versatile functions of OsALDH2B1 provide a genic basis for growth-defense trade-offs in rice. Proc. Natl. Acad. Sci. USA 117, 3867–3873 (2020).
Fang, Z. et al. Strigolactones and brassinosteroids antagonistically regulate the stability of the D53-OsBZR1 complex to determine FC1 expression in rice tillering. Mol. Plant 13, 586–597 (2020).
Qiao, S. et al. The RLA1/SMOS1 transcription factor functions with OSBZR1 to regulate brassinosteroid signaling and rice architecture. Plant Cell 29, 292–309 (2017).
Xiao, Y. et al. GSK2 stabilizes OFP3 to suppress brassinosteroid responses in rice. Plant J. 102, 1187–1201 (2020).
Ren, Y. et al. Oryza sativa mediator subunit OsMED25 interacts with OsBZR1 to regulate brassinosteroid signaling and plant architecture in rice. J. Integr. Plant Biol. 62, 793–811 (2020).
Wang, H. et al. The histone deacetylase HDA703 interacts with OsBZR1 to regulate rice brassinosteroid signaling, growth and heading date through repression of Ghd7 expression. Plant J. 104, 447–459 (2020).
Pei, Y. et al. Bifunctional transcription factors SlERF.H5 and H7 activate cell wall and repress gibberellin biosynthesis genes in tomato via a conserved motif. Dev. Cell 59, 1345–1359 e1346 (2024).
Zou, T. et al. DWARF AND LOW-TILLERING 2 functions in brassinosteroid signaling and controls plant architecture and grain size in rice. Plant J. 116, 1766–1783 (2023).
Bai, M. Y. et al. Functions of OsBZR1 and 14-3-3 proteins in brassinosteroid signaling in rice. Proc. Natl. Acad. Sci. USA 104, 13839–13844 (2007).
Tong, H. N. et al. DWARF AND LOW-TILLERING acts as a direct downstream target of a GSK3/SHAGGY-like kinase to mediate brassinosteroid responses in rice. Plant Cell 24, 2562–2577 (2012).
Tian, X. et al. WRKY53 Integrates classic brassinosteroid signaling and the mitogen-activated protein kinase pathway to regulate rice architecture and seed size. Plant Cell. 33, 2753–2775 (2021).
Sakamoto, T. et al. Erect leaves caused by brassinosteroid deficiency increase biomass production and grain yield in rice. Nat. Biotechnol. 24, 105–109 (2006).
Min, H. J. et al. OsBZR1 turnover mediated by OsSK22-regulated U-box E3 ligase OsPUB24 in rice BR response. Plant J 99, 426–438 (2019).
Jiang, Y. et al. XIAO is involved in the control of organ size by contributing to the regulation of signaling and homeostasis of brassinosteroids and cell cycling in rice. Plant J. 70, 398–408 (2012).
Tong, H. et al. DWARF AND LOW-TILLERING, a new member of the GRAS family, plays positive roles in brassinosteroid signaling in rice. Plant J. 58, 803–816 (2009).
Tanaka, A. et al. BRASSINOSTEROID UPREGULATED1, encoding a helix-loop-helix protein, is a novel gene involved in brassinosteroid signaling and controls bending of the lamina joint in rice. Plant Physiol. 151, 669–680 (2009).
Zhan, P. et al. Natural variations in grain length 10 (GL10) regulate rice grain size. J. Genet. Genom. 49, 405–413 (2022).
Hong, J. et al. Combined genome-wide association study and epistasis analysis reveal multifaceted genetic architectures of plant height in Asian cultivated rice. Plant Cell Environ. 46, 1295–1311 (2023).
Zhang, Y. et al. OsCBL5–CIPK1–PP23 module enhances rice grain size and weight through the gibberellin pathway. Plant J. 115, 895–909 (2023).
Lan, J. et al. Small grain and semi-dwarf 3, a WRKY transcription factor, negatively regulates plant height and grain size by stabilizing SLR1 expression in rice. Plant Mol. Biol. 104, 429–450 (2020).
Huang, X. et al. A map of rice genome variation reveals the origin of cultivated rice. Nature 490, 497–501 (2012).
Wang, W. et al. Genomic variation in 3,010 diverse accessions of Asian cultivated rice. Nature 557, 43–49 (2018).
Guo, H. et al. Differentiation, evolution and utilization of natural alleles for cold adaptability at the reproductive stage in rice. Plant Biotechnol. J. 18, 2491–2503 (2020).
Katan, Y., Agami, R. & Shaul, Y. The transcriptional activation and repression domains of RFX1, a context-dependent regulator, can mutually neutralize their activities. Nucleic Acids. Res. 25, 3621–3628 (1997).
Fry, C. J. & Farnham, P. J. Context-dependent transcriptional regulation. J. Biol. Chem. 274, 29583–29586 (1999).
Gao, X. et al. Rice qGL3/OsPPKL1 functions with the GSK3/SHAGGY-like kinase OsGSK3 to modulate brassinosteroid signaling. Plant Cell 31, 1077–1093 (2019).
Takano-Kai, N. et al. Evolutionary history of GS3, a gene conferring grain length in rice. Genetics 182, 1323–1334 (2009).
Duan, P. et al. natural variation in the promoter of GSE5 contributes to grain size diversity in rice. Mol. Plant 10, 685–694 (2017).
Mao, H. et al. Linking differential domain functions of the GS3 protein to natural variation of grain size in rice. Proc. Natl. Acad. Sci. USA 107, 19579–19584 (2010).
Huang, K. et al. WIDE AND THICK GRAIN 1, which encodes an otubain-like protease with deubiquitination activity, influences grain size and shape in rice. Plant J. 91, 849–860 (2017).
Chen, H. et al. Firefly luciferase complementation imaging assay for protein-protein interactions in plants. Plant Physiol. 146, 368–376 (2008).
Tian, J. et al. Maize smart-canopy architecture enhances yield at high densities. Nature 632, 576–584 (2024).
Hao, R. et al. On salt stress, PLETHORA signaling maintains root meristems. Dev. Cell 58, 1657–1669 e1655 (2023).
Zhu, J. Y., Sun, Y. & Wang, Z. Y. Genome-wide identification of transcription factor-binding sites in plants using chromatin immunoprecipitation followed by microarray (ChIP-chip) or sequencing (ChIP-seq). Methods Mol. Biol. 876, 173–188 (2012).
Wisniewski, J. R., Zougman, A., Nagaraj, N. & Mann, M. Universal sample preparation method for proteome analysis. Nat. Methods 6, 359–362 (2009).
Lyu, J. et al. Control of grain size and weight by the GSK2-LARGE1/OML4 pathway in rice. Plant Cell 32, 1905–1918 (2020).
Zhang, Y. et al. A highly efficient rice green tissue protoplast system for transient gene expression and studying light/chloroplast-related processes. Plant Methods 7, 30 (2011).
Hao, Y.-J. et al. Plant NAC-type transcription factor proteins contain a NARD domain for repression of transcriptional activation. Planta 232, 1033–1043 (2010).
Acknowledgements
We are grateful for Prof.Qian Qian for providing the plant material ZH11-GS2AA. We would like to thank Prof. Shouyi Chen and Prof. Jinsong Zhang for the dual-luciferase reporter (DLR) assay system. We would like to thank Prof. Jianmin Zhou for providing the c/n-LUC vectors and Prof. Kejian Wang for the Cas9 vector. We would like to thank Prof. Ran Xu for analysing the MutMap data. We would like to thank Mr. Yongqing Pan’s help with haplotype analysis. This work was supported by grants from the National Key Research and Development Program of China (2022YFF1002903, 2021YFF1000202), the strategic priority research program of the Chinese Academy of Sciences (XDB1090000), the National Natural Science Foundation of China (32270349, 31871219, 31771340, 31960159, 32072046), the China Postdoctoral Science Foundation (2022M723368), Key Research and Development Program of Hainan (ZDYF2021XDNY165), Hainan Provincial Natural Science Foundation of China (325RC840), and Hainan Seed Industry Laboratory Project (B21HJ0220).
Author information
Authors and Affiliations
Contributions
Y.H.L. conceived and designed this project. Y.J.L., K.H. and Y.H.L. designed experiments. Y.J.L. and K.H. performed most of the experiments. B.L.Z. performed rice genetic transformation. P.G.D. helped to analyse the MutMap data and identify the SUG1 gene. L.M.Z. helped to analyses the haplotype data. G.Z.Z. helped to perform the sug1-cr construct. C.Z. and N.N.H. helped to perform field yield tests. X.H.H., and Y.C.W. helped to perform the mass spectrometry analyses. Y.J.L., K.H., and Y.H.L. analyzed data, prepared figures and wrote the article. L.Y.Z. revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Xiaojin Luo and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, Y., Huang, K., Zhang, L. et al. A molecular framework for the GS2-SUG1 module-mediated control of grain size and weight in rice. Nat Commun 16, 3944 (2025). https://doi.org/10.1038/s41467-025-59236-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-59236-w