Abstract
Single-cell transcriptome profiling enables unparalleled characterization of the heterogeneous microenvironment of pediatric leukemias. To facilitate comparative analyses and generate pediatric leukemia signatures, we collect, process, and annotate single-cell data comprising over 540,000 cells from 159 different pediatric acute leukemia (myeloid, lymphoid, mixed phenotype lineages) and healthy bone marrow (BM) samples, profiled in our lab and curated from publicly available studies. The analysis identifies a leukemia-enriched signature of nine genes with over-expression in leukemic blast compared to healthy BM cells. This signature is also consistently over-expressed in leukemia samples compared to normal BM in bulk RNA-seq datasets (over 2000 samples). Outcome-based analysis on diagnosis samples using measurable residual disease (MRD) status depicts a significant association of oncogene-induced senescence and g-protein activation pathways with MRD positivity. MRD positivity across pediatric leukemias is also correlated with significant depletion of CD8+ and CD4+ naïve T-cells and M1-macrophages at diagnosis. To enable easy access to this comprehensive pediatric leukemia single-cell atlas, we develop the Pediatric Single-cell Cancer Atlas (PedSCAtlas, https://bhasinlab.bmi.emory.edu/PediatricSCAtlas/). The atlas allows for quick exploration of single-cell data based on genes, cell type composition, and clinical outcomes to understand the cellular landscape of pediatric leukemias.
Similar content being viewed by others
Introduction
Unraveling the heterogeneity of tumor landscapes is transforming the understanding of cancer progression, response to therapy, and the ability of tumors to adapt to novel conditions. A major limitation of bulk RNA-sequencing is that it represents only the average behavior of a sample, obscuring cell-to-cell variability in tumor ecosystems. Single-cell RNA-sequencing (scRNA-seq) overcomes these limitations by unmasking cellular heterogeneity1,2,3, and is being used for building atlases like the Human Cell Atlas (HCA) and the Human BioMolecular Atlas Program (HuBMAP), which examine various biological contexts and tissue makeup4,5,6,7,8,9,10 at a large scale.
Of the various single-cell “omics” approaches, scRNA-seq is the most utilized, enabling comprehensive characterization by measuring the expression levels of thousands of genes per cell11. ScRNA-seq is helping unravel the underlying mechanisms of tumor progression and relapse by enabling in-depth characterization of malignant, stromal, and immune cells in the tumor microenvironment and over the last decade, many cancer studies have been performed to identify potential diagnostic/therapeutic biomarkers using scRNA-seq12,13,14,15,16. Studies have also shed light on the clonal evolution of therapy-resistant tumor cells and the interaction of tumor cells with their environments17,18. Additionally, single-cell analysis enables identification of rare, clinically-relevant populations, as revealed by the identification of the presence of superior effector T-cells that are less prone to exhaustion, in a recent study demonstrating the enhanced therapeutic potential of PD1 blockade and IL-2 stimulation19. In the pediatric cancer field, there have been fewer single-cell studies than in adult cancer. A lower mutational burden in pediatric cancers is understood to make the identification of drug targets more challenging20, and given the evidence that pediatric and adult leukemias are molecularly distinct21, there is merit in pursuing pediatric leukemia focused single-cell studies.
Some of the recent single-cell studies in pediatric leukemia have investigated therapy resistance in early T-cell progenitor acute lymphoblastic leukemia (ETP-ALL)22, and characterized immune-suppressive environment association with minimal residual disease (MRD) in B-cell acute lymphoblastic leukemia (B-ALL)23. In our recent pediatric acute myeloid leukemia (AML) study24, we unraveled the heterogeneity of AML blasts and identified a blast-associated gene signature that could be used to discriminate AML blasts from non-blast cells. Similarly, our analysis of pediatric mixed phenotype acute leukemia (MPAL) samples demonstrated that B-cell/myeloid and T-cell/myeloid MPAL have unique profiles, which may account for the differences observed in disease development and/or response to current therapies25. In pediatric T-cell ALL (T-ALL), scRNA-seq analysis was utilized to identify a T-ALL blast-specific signature and MRD-associated immune environment changes26. Most single-cell studies involve small sample sizes and are limited to characterizing the single-cell landscape of a specific leukemia type. To explore potential candidates that may be commonly dysregulated across different leukemias, it is necessary to perform correlative studies after consolidating single cell data from emerging studies. Therefore, in this study, we have integrated pediatric leukemia datasets from AML, ALL, and MPAL to generate a large pediatric leukemia scRNA-seq atlas. Subsequently, we annotated the malignant and microenvironment cells, and performed a comparison of malignant and healthy bone marrow (BM) cells to identify genes commonly over-expressed in the leukemia cells. Further, to ensure leukemia specificity, genes expressed in BM cells from healthy donors were filtered out. The association of gene over-expression in leukemia was confirmed using public bulk RNA-seq datasets to generate a leukemia-enriched signature of nine genes that were over-expressed across different pediatric leukemias. The expression of the leukemia-enriched signature was further validated in malignant cells of scRNA-seq pediatric AML, adult AML, and adult MPAL publicly available datasets.
ScRNA-seq generates copious amounts of data and most single-cell analysis tools are both time- and effort-intensive and require prior knowledge of programming and access to costly computational resources. To overcome these limitations, we have developed an interactive online resource, the Pediatric Single-cell Cancer Atlas (PedSCAtlas). This unique, open-access resource facilitates the exploration and visualization of pediatric leukemias’ single-cell gene expression without requiring any analytical and bioinformatics expertise. It enables the exploration of gene expression profiles across leukemic and microenvironment cells of different major pediatric leukemias and their genetic subtypes. The PedSCAtlas also includes modules for data visualization and biomarker analysis to analyze and interpret single-cell data.
In this work, we integrate scRNA-seq datasets of pediatric AML, ALL, and MPAL samples along with healthy controls to construct a detailed, single-cell pediatric leukemia atlas. This enabled the identification of a leukemia-enriched signature of nine genes, which are overexpressed in leukemia cells, and validated on publicly available bulk and single-cell RNA-seq datasets. Additionally, based on this data, we have developed the Pediatric Single-cell Cancer Atlas (https://bhasinlab.bmi.emory.edu/PediatricSCAtlas/), an interactive resource that enables the exploration of single-cell gene expression and biomarker analysis in pediatric leukemias.
Results
Single-Cell RNA Sequencing Pediatric Leukemia Atlas
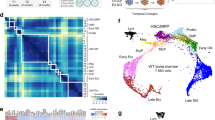
To generate the pediatric leukemia-enriched signature, we compiled a scRNA-seq dataset consisting of expression data for 231,883 cells extracted from 76 leukemic and 6 healthy BM samples. Table 1 describes the leukemia types, ages, and data sources of the samples used in the dataset. The expression of canonical immune markers is shown for each annotated cell type (Fig. 1a). The analysis of relative cell abundances across the leukemias revealed that samples are dominated by blasts, while normal immune cells were observed as minor populations, since most of the samples were collected at disease diagnosis (Fig. 1b). In contrast, healthy BM samples contained major immune lineages including T-cells, B-cells, and monocytes, with few immature cells (Fig. 1b). The clustering analysis and visualization on uniform manifold approximation and projection (UMAP) plots depict that non-blast immune cells (in shades of blue) from healthy and leukemic samples clustered together, while malignant blast cells (in shades of brown/red) show heterogeneity within and between different leukemia types (Fig. 1c). The cohort represents a diverse assortment of pediatric leukemia patients from various cytogenetic and immunophenotypic groups with different outcomes and therapeutic responses (Fig. 1d).
a Dot plot showing immune cell canonical marker expression in the leukemia dataset (n = 231,883 cells). For each annotated cell type, a corresponding dot color shows the level of expression (blue: low, red: high) and dot size indicating the percent of cells with expression of the target gene. b Proportion of cell types across different leukemias (AML, B-ALL, B/My MPAL, Healthy BM, T-ALL, and T/My MPAL). The different cell types are color coded as indicated in the legend below. c UMAP plots showing the split view of cell clusters from Healthy, AML, B-ALL, B/My MPAL, T-ALL, and T/My MPAL Bone marrow samples. The cell types are colored using the same colors as shown in panel b, with nonblast, and non-progenitor cell types lassoed in the Healthy BM group. The blast and healthy cells are represented with different shades of red and blue colors respectively. d Leukemia type (e.g. AML), cytogenetic category (e.g. inv(16)), mutational status (e.g. CEBPA), immunophenotype category (e.g. ETP-ALL), age, and number of cells for samples and patients in the pediatric leukemia cohort. All single-cell data were generated using 10x genomics technologies. Sorting methods include immune (sorted to enrich immune cells based on CD45), non-leukemia (CD45 + CD19-CD10- for B-ALL), and lymphoblast (CD19 + CD10+ for B-ALL). e Pediatric leukemia-enriched signature discovery workflow with each analytical and filtering step along with threshold and outputs. For each filtering step, the number of genes passing the threshold are shown. AML acute myeloid leukemia; B-ALL b-cell acute lymphoblastic leukemia; B/My MPAL b-cell/myeloid mixed phenotype acute leukemia; Healthy BM healthy bone marrow, T-ALL t-cell acute lymphoblastic leukemia, T/My MPAL t-cell/myeloid mixed phenotype acute leukemia, HSC hematopoietic stem cell, GMP granulocyte-monocyte progenitor, ETP-ALL early T-cell precursor acute lymphoblastic leukemia, UMAP uniform manifold approximation and projection, FC fold change, B-H Benjamini-Hochberg. Source data are provided as a Source Data file.
Development of Workflow to Identify Leukemia-Enriched Signature
The systematic approach for signature discovery is outlined in Fig. 1e. The signature represents genes over-expressed in major pediatric acute leukemia types compared to healthy BM, profiled using scRNA-seq and bulk-RNA-seq techniques. First, the gene-expression profile of blast cells of each leukemia type was compared with the young adult healthy BM cells using differential expression (DE) analysis. Additionally, gene set enrichment analysis (GSEA) was performed to identify significantly enriched gene sets across leukemias. Genes commonly over-expressed in all leukemia types with minimal expression in normal tissues were identified. Further, to reduce false positives, we verified the expression of blast over-expressed genes in bulk RNA-seq data of primary leukemia and normal BM cases. Subsequently, analysis was performed on genes in leukemia-enriched signatures to evaluate the correlation of expression with clinical blast percentage and prognostic outcomes.
Signature Identification
DE analysis comparing the profile of each leukemia type’s blast cells with young adult healthy BM cells identified 673 AML, 1162 B-ALL, 1416 T-ALL, 1133 B/My MPAL, and 956 T/My MPAL significantly over-expressed genes (Fig. 1e). The expression profile of the top 10 over-expressed genes in blast cells for different leukemias and healthy BM is shown in Fig. 2a. The top differentially expressed genes (DEGs) for AML include general myeloid progenitor markers (MPO, AZU1, ELANE)27. Further, T-ALL analysis identified cellular differentiation and chromatin modification-related genes (HIST1H1C, HES4). T/My MPAL top markers include GATA2, a transcription factor involved in hematopoiesis and critical for proliferation of hematopoietic stem cells (HSCs)28, and T-cell activation marker ITM2A29. The similarity in DEGs among leukemias was assessed by calculating Jaccard Index (JI) values (Fig. 2b). Interestingly, B-ALL and B/My MPAL exhibited a high overlap (JI = 0.433) between their DEGs, suggesting similar blast expression profiles. Common DEGs include SOCS2, a feedback inhibitor of JAK-STAT pathways highly expressed in HSCs30, and PMAIP1, a pro-apoptotic gene from the BCL-2 family31. Interestingly, the second highest similarity was observed between T/My MPAL and AML (JI = 0.329), possibly due to the myeloid lineage commonality; however, B/My MPAL and AML have a lower similarity (JI = 0.177). The lowest JI was observed between B-ALL and T/My MPAL (0.131), indicating the most diverging among the blast populations profiled. 98 genes were commonly over-expressed in all leukemia types studied (Supplementary Data 1), including SOX4, which has been associated with oncogenic survival signals in ALL and shown to drive AML32.
a Combined heatmap showing normalized, scaled expression of top significant (Bonferroni adj. p < 0.05) differentially expressed genes (DEGs) derived from comparing blast cells and healthy BM cells for each leukemia type (two-sided Wilcoxon rank sum test). Top 10 genes were identified based on log2FC values. b Heatmap showing similarity in DEGs for each leukemia’s blast cells as the Jaccard Similarity/Index, calculated as the number of intersecting genes divided by the size of the union of genes between two leukemia types. c Dot plot with gene sets which are common or specific to leukemia types that were identified by comparing enrichment scores (from gene set enrichment analysis) in blast and healthy BM cells (two-sided t-tests). The top significantly enriched (Benjamini-Hochberg adj. p < 0.05) gene sets were selected based on T-statistic. d Volcano plot showing differential expression analysis results from comparing different leukemias versus normal BM samples in bulk RNA-seq datasets (moderated two-sided t-test, limma). Genes significantly over-expressed (log2FC > 0.25, Benjamini-Hochberg adj. p < 0.05) in leukemia types are shown in red, genes significantly under-expressed (log2FC <−0.25, Benjamini-Hochberg adj. p < 0.05) are shown in blue, and non-significant genes are shown in grey. The shapes of the points represent the results of the differential expression analysis for different leukemia types. e A dot plot illustrating a summary of bulk RNA-seq DEG analysis (moderated two-sided t-test, limma) across different leukemias, with the color of the dots representing the log2FC, the shape representing whether the gene is over- (triangle) or under-expressed (circle) in leukemia compared to normal, and the Benjamini-Hochberg adjusted p-value represented as asterisks on the dots (*, < 0.05; **, <0.01; ***, <0.001; ****, <0.0001). AML acute myeloid leukemia, B-ALL b-cell acute lymphoblastic leukemia, B/My MPAL b-cell/myeloid mixed phenotype acute leukemia, T-ALL t-cell acute lymphoblastic leukemia, T/My MPAL t-cell/myeloid mixed phenotype acute leukemia, FC fold change. Source data are provided as a Source Data file.
GSEA was performed to assess dysregulated biological pathways among leukemia blast cells compared to healthy BM. A total of 658 gene sets were found to be over-enriched (T-statistic > 20) across different leukemias (Figs. 2c, S1). Pediatric leukemia-enriched gene sets include oncogene-induced senescence, G-protein activation, and EGFR signaling, which point to leukemogenesis mechanisms across leukemias. In addition, we identified varying numbers of leukemia-specific gene sets representing heterogeneity at pathway-level (Fig. 2c), including IL-1 processing in AML, KRAS signaling in B-ALL, TP53-regulation of G1 cell-cycle arrest in B/My MPAL, cholesterol biosynthesis in T-ALL, and loss of SMAD2 signaling in T/My MPAL (Fig. 2c). A healthy adult BM single-cell dataset4 was used to filter out leukemia-dysregulated genes with high expression (normalized expression across clusters =>0.5) in normal cells. The analysis identified 42 genes over-expressed in leukemic blast cells with minimal expression in the normal cells of the healthy BM (Table S1). To visualize the low expression of the candidate leukemia markers in healthy BM, we generated a heatmap (Fig. S2) for candidate leukemia over-expressed markers (n = 98), and their average expression across clusters in the healthy adult dataset; with the 42 genes forwarded to the bulk DE step having low expression across clusters.
To further validate leukemia overexpression of 42 candidate genes from single cell analysis, we utilized RNA-seq leukemia datasets from the Therapeutically Applicable Research to Generate Effective Treatments (TARGET) initiative. DE analysis was performed to compare the transcriptomes of normal BM and leukemia samples to identify over-expressed genes in leukemias. This analysis shows the majority of these 42 genes are significantly over-expressed in at least one leukemia type (Fig. 2d). Nine genes—TAF1D, SYNCRIP, PAN3, MYL6B, MYB, HACD1, GNA15, CCNB1IP1, and CCDC88A—were significantly over-expressed in all five leukemias. In addition, we identified signatures for major leukemia groups: AML (n = 21), ALL (n = 19), and MPAL (n = 17) (Table S2). Interestingly, most of the genes are significantly over-expressed in multiple leukemia types (e.g., MIR181A1HG, PMAIP1, ARMH1), while some have expression restrictive to one of the leukemias (NUFIP2 in T-ALL) (Fig. 2e). Multiple genes from the AML signature of 21 genes show a significant positive association between gene expression and blast percentage in the TARGET-AML dataset, supporting the blast-specific expression of these genes (Fig. S3). Additionally, while the expression of these genes varies across different leukemia types in the TARGET datasets (Fig. S4), they are all significantly over-expressed compared to normal BM samples, as determined during our differential expression testing.
To ensure the signature is robust to changes in filtering thresholds utilized in the discovery pipeline, we evaluated the impacts of altering the adjusted p-values and fold change (FC) thresholds on the differential expression of pediatric leukemia-enriched signature genes. For the single-cell analysis, lowering the adjusted p-value threshold did not impact the identification of signature genes, as the p-values for all genes were highly significant, near zero. Increasing the FC threshold affected the identification of over-expressed genes, as the majority have a FC below 2. Overall, GNA15 was the most robust gene in the single-cell threshold filtering (Fig. S5a). In the bulk differential expression analysis, increasing the FC threshold sequentially impacted genes from our signature, with all genes filtered out with FC cutoff >2.7. Overall, HACD1 was the most robust differentially expressed gene in the bulk RNA-seq data (Fig. S5b).
An ablation analysis was performed to further assess the robustness of the signature by randomly selecting the cells or samples utilized in the differential expression analysis. In the analysis, 5000 blast cells were randomly sampled for each leukemia type along with additional randomly selected 5000 cells from healthy BM samples for comparison. The analysis was permutated 100 times to determine the impact of randomly selected cells on the genes. Interestingly, 7 out of 9 genes were identified as differentially expressed in 100% of permutations. PAN3 and SYNCRIP genes were identified as differentially expressed in 98% and 82% of permutations, respectively (Fig. S5c). We performed additional permutation analysis on the bulk RNA-seq data to further test the robustness of the signature. The permutation analysis was repeated 100 times by randomly selecting 95% of samples from each leukemia type before differential expression analysis. Of the leukemia-enriched signature, eight genes were detected as differentially expressed in 100% of permutations, and GNA15 was detected in 80% of the runs (Fig. S5d).
In addition, we assessed the signature coverage across patient samples in our pediatric leukemia single-cell atlas and determined our signature identification is not biased toward a particular leukemia type or cytogenetic subtype (Figs. 3a, S6a–c). As these genes are all significantly over-expressed in leukemic compared to healthy cells, we assessed the individual gene expression across leukemic and healthy cell types across our pediatric leukemia atlas (Fig. S6d). PAN3, SYNCRIP, and HACD1 show minimal to no expression in non-blast healthy cell types, while other genes exhibit low-level expression. Specifically, as shown in Fig. S6d, CCDC88A is expressed in mono/macrophage and progenitor cells, CCNBIP1 and MYB in progenitor cells, GNA15 in erythroid, mono/macrophages and progenitor cells, MYL6B in erythroid and progenitor cells, and TAF1D in most cell types. Overall, 91% of blast cells express at least one of the signature genes. Further, we confirm this in a larger TARGET cohort with major genetic subtypes, where the signature expression shows inter-patient heterogeneity. The heatmaps and unsupervised hierarchical clustering analysis reveal the signature is not specific to certain genetic and molecular subtypes across pediatric leukemias (Fig. S7a–c).
a Heatmap of proportion of cells expressing the leukemia-enriched signature at diagnosis, per patient in the pediatric leukemia atlas, calculated as the proportion of blast cells in a sample with counts above zero. Additional patient information is annotated on top of the heatmap. Hierarchical clustering analysis was performed on samples (columns) and genes (rows) using Euclidean distance. b Average log2 fold change of the signature genes’ expression compared to the average expression of CD34, CD45/PTPRC, and CD74 in the Roy et al.33 healthy hematopoiesis dataset, per tissue type. Adult bone marrow (ABM), early fetal liver (eFL), fetal bone marrow (FBM), fetal liver (FL), and pediatric bone marrow (PBM). c Boxplots depict normalized expression in AML cells (Lambo et al.34) labeled as malignant (red) or normal microenvironment (blue). Two-sided Wilcoxon rank sum test was used to compare the expression levels between malignant (n = 189,240) and normal (n = 119,292) cells. The p < 0.0001 represents where the p-value is estimated to be less than 2.2e-16. Middle bar represents the median, lower/upper hinges correspond to first/ third quartiles, the upper whiskers extend to the largest value no further than 1.5 times IQR. d Average expression of the signature genes across malignant AML cell types (Van Galen et al.27) in heatmap and overall in bar-plot. e Distributions of module scores of the signature across MPAL and healthy reference cells (Granja et al.35). HACD1 was not found in the count matrix for this dataset and therefore is not included in the score calculation. Categories include reference cells from healthy BM samples, healthy-like MPAL cells, and subtypes of malignant MPAL cells (Erythroid-like, Lymphoid-like, Myeloid-like, Progenitor-like, TNK-like). Bars in the violin plots represent the median. f Distribution of module score in malignant MPAL cells overall compared to healthy-like MPAL and healthy reference cells. AML acute myeloid leukemia, B-ALL b-cell acute lymphoblastic leukemia, B/My MPAL b-cell/myeloid mixed phenotype acute leukemia, T-ALL t-cell acute lymphoblastic leukemia, T/My MPAL t-cell/myeloid mixed phenotype acute leukemia, FC fold change. Source data are provided as a Source Data file.
Exploring Leukemia-Enriched Signature Profile in Additional Normal Datasets
To validate the malignancy association of our signature, we evaluated expression in two additional normal tissue scRNA-seq datasets: (i) two BM samples from healthy pediatric donors sequenced in our lab, and (ii) a previously published study with healthy hematopoietic tissues across different developmental stages.
The expression of the signature was assessed by calculating the fold change of the genes against a reference set of canonical hematopoietic cell lineage markers (CD74, CD45, CD34). The analysis depicted lower expression (i.e. negative FC) of our signature as compared to hematopoietic cell lineage markers across most cell types in the two pediatric BM samples sequenced by our lab (Fig. S8a). Most genes are expressed in only a small fraction of normal cells (averaging 18%) in pediatric bone marrow samples of similar age, highlighting their association with malignant phenotypes (Fig. S8a).
Next, we assessed the expression of our signature in the fetal liver, fetal BM, pediatric BM, and adult BM tissue types of the publicly available dataset from Roy et al. (Cell Reports 2021)33. Overall, the pediatric leukemia-enriched signature has heterogeneous but low average expression across the normal tissue developmental stages. When comparing the expression of genes to hematopoietic canonical markers (CD74, CD45, CD34) in the hematopoietic cells from fetal, pediatric, and adult samples, the signature genes have lower expression than the canonical markers across most tissue stages, providing further support for malignant phenotype association (Fig. 3b).
For validation using an alternative approach, we compared the aggregated expression of our signature to the signature utilized by Roy et al. for annotating the normal tissue types of lineages, such as ELANE for Myeloid, CD79A for Lymphoid, and KLF2 for Hematopoietic stem cells (Supplementary Data 2). Across all healthy tissue developmental stages (fetal, pediatric, adult) the signature has lower expression in comparison to the control signature (Fig. S8b). Further comparison by stratifying datasets into broad canonical cell types also showed significantly lower expression of signature genes as compared to the control gene signature, supporting the malignancy association of our signature (Fig. S8c).
Exploring Pediatric Signature Profile in External Leukemia Datasets
To validate the malignant cell type association of the pediatric leukemia-enriched signature in independent pediatric datasets, we performed a comparative analysis with a recent study by Lambo et al. with 308,000 cells (GSE235063) from diagnosis, relapse, and remission AML BM samples34. Following pre-processing, normalization, and cell type annotation (Fig. S9a), the analysis of the signature revealed significant overexpression in AML malignant cells compared to normal cells (Fig. 3c). The feature plots also depicted that the expression of signature genes is mostly restricted to UMAP regions enriched with malignant AML cells (Fig. S9b–d).
Similarly, to assess the expression of pediatric signature genes in adult acute leukemia datasets, we performed a comparative analysis with: (i) van Galen et al.27 adult AML study with over 37,000 cells27 and (ii) Granja et al.35 adult MPAL study with over 52,000 cells35. The expression of the signature genes is heterogenous across malignant subtypes in adult AML. On average, CCDC88A, SYNCRIP, and TAF1D have the highest expression in malignant adult AML cells (Figs. 3d, S10a). Signature expression based on module score is significantly higher in malignant than normal cells from adult AML BM (Fig. S10b). The adult AML dataset was further stratified into different lineage malignant cell types (as annotated in the Van Galen study) to evaluate our signature expression. The expression of the signature varies across subtypes (Fig. S10c), ranging from highest in stem-like populations GMP-like (granulocyte-monocyte progenitor), HSC-like (hematopoietic stem cell), and Prog-like (progenitor) and lower in more differentiated populations cDC-like, Mono-like, and ProMono-like (pro-monocyte).
The analysis of adult MPAL depicted the highest expression of TAF1D, GNA15, and SYNCRIP genes (Fig. S10d). The clustering of adult MPAL samples based on the expression of the pediatric signature genes depicted separate clustering for MPAL samples from the control PBMCs and BM cells. Interestingly, control CD34+ enriched samples clustered more with some T/My MPAL samples, depicting similarity in the profiles (Fig. S10d). This hints toward a similar expression profile of pediatric leukemia-enriched signature genes between T/My MPAL samples and normal HSCs. Further calculation of the signature enrichment based on the module score depicted higher expression of pediatric leukemia genes in lymphoid-like and progenitor-like blast cells as compared to erythroid-like, myeloid-like, and TNK-like blasts (Figs. 3e, S10e, f). This is further confirmed through the higher signature module scores in MPAL malignant cells overall as compared to healthy donor reference (CD34 + , BMMC, PBMC samples) and MPAL healthy-like cells (Figs. 3f, S10g).
Exploring Prognostic and Biological Trends of Genes in the Leukemia-Enriched Signature
We perform multivariate survival analysis to assess if the expression of our signature has a significant correlation with survival while adjusting for confounding factors. We observed heterogenous, leukemia-specific prognostic associations where some of the genes are associated with poor outcomes in specific types or subtypes of leukemia. After adjusting for age and cytogenetic groups available in the AML dataset, MYB, CCDC88A, PAN3, and SYNCRIP are significantly associated with poor survival (HR > 1, p < 0.05) in AML (Fig. 4a). MYB is a proto-oncogene transcription factor and has been associated with leukemogenesis, hematopoiesis36, and cell cycle via regulation of cyclin B37. CCNB1IP1 and MY6LB are associated with poor outcomes (HR > 1, p < 0.05) and PAN3, GNA15, SYNCRIP, and TAF1D are associated with better outcomes (HR < 1, p < 0.05) in the B-ALL dataset after adjusting for patient age (Fig. 4a). Interestingly, some genes—PAN3 and SYNCRIP—have opposite survival associations in the AML and B-ALL cohorts, which can be attributed to heterogeneous microenvironments and genetic differences. Further detailed analysis of different MYB expression levels and outcomes in the AML dataset depicted a significant correlation between expression levels and outcomes, with the highest expression associated with poor outcomes (HR = 1.5, p = 0.007, Fig. 4b). Further survival associations within each cytogenetic group depicted consensus association of higher MYB expression with poor outcome, independent of different baseline risks (Fig. 4c). Similarly, expression levels of PAN3 in the B-ALL dataset showed a significant correlation with outcomes, with the highest expression showing significantly better outcomes (HR = 0.56, p = 1.08e-08, Fig. 4d).
Multivariate survival analysis was performed to explore the independent associations of pediatric leukemia-enriched signature genes with outcomes. TARGET datasets from the Survival Genie tool were utilized for this analysis. In the TARGET-AML analysis, cytogenetic groups and age were included as covariates; for the TARGET-ALL-P2 and TARGET-ALL-P3 analyses, age was included as a covariate. When examining survival associations based on each gene’s expression, continuous log2(FPKM + 1) was used as a covariate. The Cox proportional hazards model was used to estimate hazard ratios (two-sided Wald test). a Scatter plot showing the hazard ratio (x-axis) and -log10 p-value (y-axis) of each gene’s association with survival across TARGET datasets. Each color corresponds to a different gene of the signature. b The survival area curve illustrating the estimated survival probability based on the discretized range of MYB expression values in the TARGET-AML-Legacy dataset (Cox HR = 1.50, p = 0.007). Expression ranges from low to high are shown with shades of blue and red colors, respectively. c Survival area curves showing the estimated survival probabilities based on MYB expression levels stratified based on cytogenetics categories in the AML cohort. d Survival area curve depicting the estimated survival probability based on the discretized range of PAN3 expression levels in TARGET-ALL-P2-B-cell dataset (Cox HR = 0.56, p = 1.08e-08). e Associated biological processes and interactions among genes from the signature from the literature or using Harmonizome 3.0 and KEGG databases. AML acute myeloid leukemia, B-ALL b-cell acute lymphoblastic leukemia, T-ALL t-cell acute lymphoblastic leukemia, HR hazard ratio, FPKM fragments per kilobase of transcript per million mapped reads. Source data are provided as a Source Data file.
Additionally, we performed a correlative analysis of our signature genes with the previously published pediatric leukemic stem cell score signature (pLSC6) by Elsayed et al.38. The pediatric LSC6 signature consists of DNMT3B, ADGRG1/GPR56, CD34, SOCS2, SPINK2, and FAM30A genes and their study depicted a poor outcome association in the TARGET-AML cohort. The correlative analysis identified a significantly positive correlation (R = 0.48, p = 5e-04) between the average expression of pLSC6 and our pediatric leukemia-enriched signatures in the AML blast cells from our single-cell dataset (Fig. S11).
The leukemia-enriched signature genes depicted an association with the cell cycle and metabolism-related pathways (Fig. 4e). For example, CCNB1IP1 has a role in the progression of the cell cycle through G2/M by promoting cyclin-B degradation39. SYNCRIP is an RNA-binding protein involved in cell cycle progression and leukemogenesis40. Both CCDC88A and MYL6B are involved in cytoskeleton remodeling/binding and contribute to tumor cells’ ability to invade surrounding tissues and metastasize to distant sites41,42. TAF1D has a role in RNA polymerase I transcription43, and PAN3 is involved in RNA degradation44, which can lead to the accumulation of aberrant oncogenic proteins, resulting in cancer development. Finally, HACD1 is involved in fatty acid metabolism, which is significantly associated with cancer cells’ therapeutic resistance45.
Macrophages are Associated with Different MRD EOI Status
Assessment of minimal residual disease (MRD) at the end of induction (EOI) is a powerful tool for the prediction of relapse in pediatric leukemia46. In our study, 36 diagnosis samples are from patients who achieved MRD negativity (<0.01%), whereas 9 attained MRD positivity (>0.01%) at EOI (Supplementary Data 3). For the rest of the samples, either MRD information was not available, or they had induction failure (>5% blast cells at EOI); these samples were excluded from this analysis. The major immune (T/NK cells, monocytes/macrophages) and blast populations were assessed for dysregulation in enrichment and expression profiles between diagnosis samples of patients with MRD positive or negative outcomes. These analyses revealed T/NK cells with overexpression of cytotoxicity (GZMB, PRF1) and inflammation/interferon-gamma associated genes (NFKB2, CCL4, ISG15) in MRD+ as compared to MRD- (Fig. 5a) patients. We also tested the enrichment of T-cell type/phenotype signatures curated from the literature47. Comparative analysis of enrichment scores of these signatures revealed that diagnosis samples from MRD-negative patients displayed significantly higher enrichment (p < 0.0001) of CD8+ naïve, CD4+ naïve, and CD8+ cytokine/cytokine-receptor signatures (Fig. 5b). In contrast, T/NK cells from diagnosis samples of patients who experienced MRD positivity had higher expression (p < 0.001) of genes involved in CD4+ and CD8+ activation/effector function. Interestingly, the signature associated with CD8+ Cytotoxic T-cells had non-significant differences in expression between the MRD groups. To explore innate immune cells association with MRD outcomes, we performed DE on the monocytes/macrophages compartment. MRD negative patients showed significant up-regulation of M1-Macrophage genes (CD74, CSTA, S100A8, CD68, S100A12, TYROBP) (Fig. 5c). Further comparison of M1 and M2 signature enrichment depicted significant (p < 0.0001) over-representation of immunosuppressive M2 macrophages in patients that were MRD positive at EOI (Fig. 5d). On the other hand, myeloid cells in diagnosis samples from MRD negative patients depicted significant enrichment of the M1-macrophage signature (Fig. 5d). We further validate the MRD status association with cellular enrichment patterns at diagnosis through in-silico analysis using the TARGET datasets (Fig. S12). The analysis depicted significantly higher enrichment of effector CD4+ (p = 0.047) and CD8+ (p = 0.063) T-cells in the MRD positive B-ALL samples (TARGET-ALL-P2-B-ALL) as compared to MRD negative samples collected the time of disease diagnosis. Additionally, we validated the increased enrichment of M1-Macrophages (p = 0.023) in samples of MRD-negative T-ALL patients as compared to MRD-positive patients.
a Volcano plot showing differentially expressed genes (DEGs) in T/NK cells at diagnosis from minimal residual disease (MRD) negative versus positive patients. Two-sided Wilcoxon rank sum test used to identify genes significantly (Bonferroni adj. p < 0.05) up-regulated in MRD negative (log2FC < −0.5, blue) and MRD positive (log2FC > 0.5, red) groups. b Box plots with module scores of T-cell phenotype gene sets across cells at diagnosis from MRD negative (n = 11,285 cells) and positive (n = 2,301 cells) groups. Two-sided Wilcoxon rank sum tests were used to assess the difference in means. c Volcano plot showing DEGs in monocytes/macrophages from MRD negative versus positive patients at diagnosis. Two-sided Wilcoxon rank sum test used to identify genes significantly (Bonferroni adj. p < 0.05) up-regulated in MRD negative (log2FC < −0.5, blue) and MRD positive (log2FC > 0.5, red) groups. d Bar plots of M1-Macrophage and M2-Macrophage module scores for myeloid cells at diagnosis from MRD-positive (n = 904 cells, red) and -negative (n = 10,029 cells, blue) patients. Two-tailed Welch’s t-test with unequal variance was used for significance testing. Data are presented as mean values plus upper quartile. e Gene-set enrichment analysis (GSEA) on blast cells from MRD negative (purple) versus positive (gold) patients at diagnosis. Significantly dysregulated gene sets identified based on two-sided Wilcoxon rank sum tests (Benjamini-Hochberg adj. p < 0.01). The top 10 gene sets by T-statistic are shown for each group. f Violin plots showing enrichment scores (ES) for four of the top gene sets between blast cells at diagnosis from MRD negative (n = 49,542 cells, purple) and positive (n = 15,852 cells, gold) patients. Two-sided Wilcoxon rank sum tests are used to compare ES values between the groups. For boxplots in b and f, the middle bar represents the median, lower/upper hinges correspond to first/ third quartiles, the upper/lower whiskers extend to the largest/smaller value no further than 1.5 times IQR. In b, d, and f, p < 0.0001 represents where the p-value is estimated to be less than 2.2e-16. Dx, diagnosis; MRD, minimal residual disease; FC, fold change; GS, gene set. Source data are provided as a Source Data file.
Analysis of leukemia blasts based on MRD outcomes identified significant (adj. p < 0.01) pathway level enrichment of the deactivation of the β-catenin transactivating complex, regulation of PKL1 activity at G2/M transition, and NOTCH1-regulation transcription pathways for the diagnosis samples of MRD positive patients (Fig. 5e). On the other hand, gluconeogenesis, alternative complement system, and glucose metabolism pathways were enriched in the diagnosis samples of MRD negative patients (Fig. 5f). Interestingly, NOTCH1 is mutated in the majority of adult T-ALL patients, and NOTCH1 signaling is known to regulate leukemia cell growth48. Overall, GSEA results indicate that blast cells at diagnosis show patterns that may be predictive of future MRD status, with blasts from the MRD negative group enriched in glucose-related and metabolic pathways and blasts from the MRD positive group enriched in pathways related to cell growth and activation.
Pediatric Single-Cell Cancer Atlas - An Online Analytical Resource
We have developed a comprehensive resource, the pediatric single-cell cancer atlas (PedSCAtlas), to allow analysis, exploration, and visualization of expression signatures in different leukemia and healthy datasets that does not require prior bioinformatics training to use. The PedSCAtlas is an online web resource with over 540,000 cells from 159 different pediatric leukemia (AML, ALL, MPAL) and healthy bone marrow (BM) samples, profiled in our lab and curated from publicly available studies, along with multiple analytical tools (Fig. 6). Currently, four scRNA-seq datasets are available on the resource: (i) the pediatric leukemia single-cell atlas presented in this study, (ii), a publicly available dataset by Lambo et al. published in 202334, (iii) an integrated healthy pediatric dataset with BM samples from 2 pediatric healthy donors sequenced by our lab and 2 CD34+ sorted pediatric BM samples published in 2021 by Roy et al.33, and (iv) a healthy adult BM of ~400,000 cells from Human Cell Atlas initiative4. We have leveraged the published cell type annotations for the cells from the latter three datasets (ii-iv) to ensure consistency.
Overview of single-cell datasets in the Pediatric Single Cell Cancer Atlas (PedSCAtlas) and analysis modules. The single-cell RNA seq (scRNA-seq) pediatric leukemia atlas examined in this manuscript contains 82 BM samples (n = 76 leukemia, n = 6 healthy) retrieved from the Emory repository and public repositories. These 82 samples were integrated and uniformly processed to form an annotated and normalized pediatric leukemia scRNA-seq atlas. Users can access and analyze the pediatric leukemia dataset along with other leukemia, and healthy pediatric datasets through a common query page. The resource contains 540,000 cells from 159 different pediatric leukemia (AML, ALL, MPAL) collected at the time of disease diagnosis, EOI, and relapse, as well as healthy bone marrow (BM) samples. The “Analysis: Analysis of Pediatric Leukemia Datasets” module allows users to visualize cell metadata and assess gene expression of the scRNA-seq datasets available on the platform. The “DE: Exploration of DEGs” module allows users to access our DE results in both the SC and Bulk RNA-seq datasets. Finally, the “Testing: Leukemia Marker Testing” module allows users to investigate a gene of their choice and test if the gene would be a robust marker of a specific disease group. PedSCAtlas pediatric single-cell cancer atlas, DEGs differentially expressed genes, AML acute myeloid leukemia, UMAP uniform manifold approximation and projection, FC fold change. Created in BioRender. Bhasin, M. (2025) https://BioRender.com/k81i482. Source data are provided as a Source Data file.
The interactive resource offers a comprehensive overview of the currently available datasets, displaying clustering information, the proportion of different major cell types, and the expression of selected genes (Fig. 7a). Users have the flexibility to query all available datasets simultaneously or explore them individually through separate pages within the resource. The “Analysis” module allows visualization of expression profiles of leukemia and healthy datasets grouped based on clusters, cell types, or patient metadata such as genetics subtype, and outcome status. For example, Fig. 7a shows the three datasets grouped based on major leukemic and immune cell types. Additionally, Fig. 7b provides an insightful depiction of the expression profile of the PAN3 gene across leukemic blast and normal progenitor cells. PAN3 gene is part of our pediatric leukemia-enriched signature identified in this study and is implicated in miRNA-mediated repression49. Notably, while there is heterogeneity in the expression of PAN3 in blast cells from different leukemias, its expression is minimal in normal progenitor cells from young adult healthy bone marrow. Furthermore, Fig. 7c displays the expression of PAN3 in a large adult healthy bone marrow dataset, where little to no expression is observed across all major hematopoietic lineages. The “DE” module contains DEGs identified by comparing the transcriptomes of malignant blast cells to healthy BM (single cell) or comparing the transcriptomes of cancer samples to normal BM (bulk), for each leukemia. The third, “Marker Testing” module (available for the pediatric leukemia atlas), allows the user to evaluate whether a gene of interest would be a suitable marker for a leukemia type, based on expression in malignant cells compared to healthy BM, and the accuracy of the gene in differentiating leukemia blasts from healthy cells if used as a feature for developing a random forest classifier.
a Common query page showing available datasets (i.e., Pediatric Leukemia Atlas, AML Lambo, et al., Healthy Pediatric BM) along with description and overview by generating UMAP plots. Analysis page with all datasets currently available on the tool. Visualization of cell metadata in the three datasets. The cellular clusters from the datasets are colored based on cell types or other user-selected categories, including sample ID, Cluster ID, MRD status, etc. b Violin plot visualization of normalized PAN3 expression across different cell types in the pediatric leukemia atlas dataset. In the boxplots overlayed, the middle bar represents the median, lower/upper hinges correspond to first/third quartiles, the upper/lower whiskers extend to the largest/smaller value no further than 1.5 times IQR. The expression is shown for different leukemic blasts (AML: n = 106,070, B-ALL: n = 7502, B/My MPAL: n = 9085, T-ALL: n = 20,072, T/My MPAL: n = 15,671), and normal progenitor (n = 2465) cells. Users can visualize gene expression through various graphical outputs, including box plots and feature plots. These visualizations allow for the grouping of cells by multiple categories, such as cell type, leukemia type, time point, and patient. This flexibility enables detailed exploration of expression patterns across diverse subsets, facilitating insights into disease-specific trends and inter-patient variability. c Violin plot visualization of normalized PAN3 expression across different cell types within the healthy adult bone marrow dataset. This visualization highlights the distribution and variability of PAN3 expression among distinct cell normal populations. Additionally, users can explore expression using violin, box, and feature plots, with the flexibility to group cells by broad or fine-granular cell types. AML acute myeloid leukemia, B-ALL b-cell acute lymphoblastic leukemia, B/My MPAL b-cell/myeloid mixed phenotype acute leukemia, T-ALL t-cell acute lymphoblastic leukemia, T/My MPAL t-cell/myeloid mixed phenotype acute leukemia, UMAP uniform manifold approximation and projection, BM bone marrow. Source data are provided as a Source Data file.
Discussion
The past decade has seen a tremendous surge in utilization of single-cell assays, in part fueled by the development of high-throughput single-cell techniques combined with a decrease in cost and increase in efficiency50. ScRNA-seq has revealed cellular heterogeneity in complex tumor microenvironments as well as enabled deeper characterization of the complex molecular mechanisms of disease biology and clinical outcomes. The ever-growing publicly available datasets have resulted in the development of online resources that collect the datasets in one place like DISCO51, CellDepot52, and GEO53. Single-cell platforms that integrate multiple datasets, enable cell type identification, gene expression analysis, etc., are important resources for identifying biological changes associated with disease and response to therapy. In pediatric leukemia, the advancement of integrated resources and datasets is critical to provide the necessary data and tools to the community of researchers and clinicians who may not have access to samples and/or experience in deconvoluting large datasets. Therefore, we created a pediatric, acute leukemia atlas from the samples processed for the generation of single-cell data in our23,24,25,26 and other labs54,55, through uniform processing and annotation.
Our identification of a pediatric leukemia-enriched signature is key to exploring the mechanisms that may be commonly dysregulated across different pediatric leukemias. Of the nine leukemia-enriched genes identified using our integrated dataset, some e.g. MYB36, GNA1556, and SYNCRIP40, have previously been associated with ALL and/or AML, while genes like CCDC88A41, CCNB1IP157, TAF1D58, HACD159, have been studied in connection to other cancers. The signature provides a simplified view of the disease mechanisms across leukemias and comprises genes involved in the cell cycle, hematopoiesis, cytoskeleton remodeling, leukemogenesis, fatty acid metabolism, RNA polymerase-I transcription, and RNA degradation. Our pediatric leukemia-enriched signature development aims to identify a common set of genes representative of the malignant transformation of normal hematopoietic stem cells across major leukemias with different genetic alterations. This signature is consistently over-represented in pediatric acute leukemia cells and samples compared to normal cells and overall bone marrow. However, we observe significant inter-patient heterogeneity in gene expression, which does not appear to correlate with specific genetic or molecular subtypes of leukemia. After further validation, this leukemia-enriched signature may have potential use for distinguishing leukemic from non-leukemic cells. This signature can be further expanded by the inclusion of leukemia types and subtypes’ signatures identified by using PedSCAtlas resource to develop future assays for diagnosis of major leukemias (AML, ALL, MPAL) and their subtypes. Of the genes with significant leukemia-specific survival associations, these could be utilized as additional prognostic risk stratification strategies within these subtypes after further validation. PAN3, SYNCRIP, and HACD1 from the pediatric leukemia-enriched signature show minimal to no expression in non-blast healthy cell types, making them promising therapeutic targets pending further investigation and validation. In contrast, other genes in the signature exhibit low-level expression in normal cell types, which may pose potential toxic effects and limit their therapeutic applicability.
Further comparative analysis of samples based on MRD outcomes60 indicates expression profiles relevant to relapse or long-term remission outcomes. We observed that diagnostic samples from MRD-negative patients had higher enrichment of CD4+ and CD8+ naïve T-cells and M1-macrophages as compared to MRD positive patients. Diagnosis samples from MRD-positive patients had higher expression of inflammatory markers and enrichment of activated/effector function CD4+ and CD8 + T-cells and immunosuppressive M2-macrophages. These results support the hypothesis that the immune microenvironment at the time of disease diagnosis influences the depth and duration of the treatment response. This makes the assessment of immune microenvironment a potential prognostic and treatment modality for improving the outcomes of pediatric leukemias. Further studies are necessary to determine the predictive capability of the immune microenvironment of leukemia and outcomes in terms of MRD status at the end of treatment.
As pediatric leukemias harbor a significantly lower mutation burden compared to adult counterparts, there is a major focus on identifying common therapeutic targets across pediatric cancers to improve outcomes and reduce the toxic effects. Therefore, the approach adopted in this study to utilize the power of single-cell data for identifying genes that are over-expressed across different leukemias with minimal or no expression in normal stromal and immune cells might yield attractive next-generation, safer cancer treatments. To support this, our Pediatric Single-cell Cancer Atlas (PedSCAtlas) is a pediatric cancer research-focused resource, that provides immediate access to integrated datasets from multiple pediatric leukemia studies. We plan to improve the PedSCAtlas as follows: (1) addition of any new pediatric cancer datasets generated from our lab, and (2) periodic incorporation of publicly available single-cell pediatric leukemia studies from other labs. Each entry will be a new dataset for users to examine, along with the original pediatric leukemia single-cell atlas compiled for this study. Additionally, we will expand the PedSCAtlas by adding data from single-cell proteomics and epigenomics assays in the future to explore the protein and regulatory elements landscape in pediatric cancers. As we have seen some similar expression patterns of our pediatric leukemia-enriched signature in individual adult leukemia datasets, we plan to directly compare adult and pediatric leukemias in future studies to understand how their distinct genetic profiles influence their cellular makeup and expression in the leukemia transcriptomic environment.
Methods
The samples used in the development of the pediatric leukemia single-cell atlas were obtained from the published studies and our studies at Emory University. The samples from our studies were obtained from the Aflac Cancer and Blood Disorders Center Biorepository within the Children’s Healthcare of Atlanta (CHOA). Signed consent was provided by parents or legal guardians for pediatric patients that permitted the use of biological material in accordance with a protocol that was approved by the CHOA Institutional Review Board (IRB protocol #00034535). Sex and gender information was not available for all samples collected from public databases and therefore was not considered in the study design.
Datasets
The pediatric leukemia single-cell atlas contains scRNA-seq data from five types of leukemia (acute myeloid leukemia, AML; B-cell acute lymphoid leukemia, B-ALL; T-cell acute lymphoid leukemia, T-ALL; B-cell/myeloid mixed phenotype acute leukemia, B/My MPAL; and T-cell/myeloid mixed phenotype acute leukemia, T/My MPAL) (Table 1). Viably frozen BM aspirates were obtained from the Aflac Cancer and Blood Disorders Biorepository (39 samples), and these samples were processed and sequenced, to generate data for our Emory repository: AML (n = 20), B-ALL (n = 6 patients multiplexed: n = 2 nonleukemic and n = 2 leukemic samples), B/My MPAL (n = 3), T-ALL (n = 10), and T/My MPAL (n = 2) patients; including data from previously published leukemia studies from our lab and collaborators23,24,25,26. A public dataset available on gene expression omnibus (GEO), accessible through the accession GSE154109 (n = 19)54, containing scRNA-seq data of BM samples from pediatric AML patients (n = 8, 4 of these patients overlap with 4 AML patients in the Emory repository, but were sequenced separately), pediatric B-ALL patients (n = 7), and young adult healthy donors (n = 4) have been included. Additionally, a scRNA-seq dataset of 24 samples from the Single-cell Pediatric Cancer Atlas Portal (ScPCA, project SCPCP000007)55 has been integrated, with samples from pediatric AML patients (n = 20), T/My MPAL patients (n = 2), and healthy young adult donors (n = 2). Overall, our integrated dataset contains 82 samples (n = 76 acute leukemia samples, n = 6 healthy samples). Most of the acute leukemia samples were obtained at the time of disease diagnosis (n = 74), one sample (PM8r) was taken at post-treatment relapse, and one sample (PT9s) was taken at the second malignancy. In addition, one sample (PA15d) is a peripheral blood mononuclear cell (PBMC) sample, while the rest are BM samples. Detailed sample and patient information have been included in Supplementary Data 3.
We have also processed and integrated bulk RNA-seq data from over 2000 samples from the TARGET datasets. We compiled the TARGET- AML,- ALL-P1, -ALL-P2, and -ALL-P3 datasets for our analyses. These datasets contain both primary leukemic BM (n = 2092) and normal BM (n = 324) samples. Leukemic BM samples correspond to multiple leukemias: AML (n = 1661), B- and Pre-B ALL (n = 115), T-ALL (n = 245), B/My MPAL (n = 32), and T/My MPAL (n = 39). In addition, we have included scRNA-seq data from almost 400,000 cells from healthy BM of adults generated under the Human Cell Atlas (HCA) initiative4 as an additional control group.
For further validation, we generated and processed scRNA-seq data for two pediatric BM samples from healthy donors (ages 11.4 and 11.9 years), aligned with 10x Genomics Cell Ranger v7.0.0. The single cell RNA-seq libraries were prepared using the Chromium single cell 3’v2 reagent kits (10x Genomics), with CMO (cell multiplexing oligo) tagging. The libraries were sequenced on an Illumina NovaSeq 6000. The processed and normalized data from these samples is also available on the PedSCAtlas resource.
Data Pre-Processing, Normalization, and Integration
For the Emory repository samples (Table 1), scRNA-seq datasets were aligned using Cell Ranger50 to generate count matrices and processed using Seurat61. For the publicly available datasets, the cell count matrices were obtained. All cells underwent quality control based on mitochondrial transcript content and feature count filtering. The high-quality cells from each dataset underwent log-normalization, scaling, dimensionality reduction, and clustering. Batch correction was performed with Harmony62 to minimize the technical effects from different sources. Dimensionality reduction was applied, and cells were clustered using a K-nearest neighbor graph-based approach.
For the bulk RNA-seq dataset, raw counts along with clinical metadata for each sample were downloaded from the genomic data commons (GDC) portal (https://portal.gdc.cancer.gov/) using the GenomicDataCommons package63. Matrices were concatenated based on their intersecting genes, forming a gene-by-sample count matrix. The raw counts were normalized using the Voom algorithm (limma) which estimates the mean-variance relationship to compute appropriate gene expression level weights for normalizing the counts data64. Specific algorithm parameters and version details have been provided in Table S3.
Cell Type Annotation
Due to the patient heterogeneity of pediatric acute leukemia, cell type annotation is inherently more complicated in scRNA-seq analysis of these samples. Therefore, multiple metrics were used to objectively annotate clusters of cells as malignant and non-malignant for each leukemia type. Each type (AML, B-ALL, T-ALL, B/My MPAL, T/My MPAL) was annotated separately, with the first step combining the type’s samples with our young adult healthy control single-cell samples. Generally, assessment from three metrics was used to annotate each cluster: the proportion of cells from healthy samples assigned to the cluster, the sample-specificity of the cluster, and canonical marker expression. These assessments, along with the final annotation assigned for each cluster, are shown in Supplementary Fig. 13. For the B-ALL analysis, samples had relatively lower proportions of B-ALL blast clusters compared to other leukemia types, due to the way in which these samples were processed. PB1d-PB7d sample data was retrieved from a public repository, GSE154109, which reports in their methods that tumor samples were sorted to decrease tumor cell abundance54. In addition, the four B-ALL samples from the Emory repository were also sorted. Two samples were sorted to have high leukemic fractions (PB8d and PB9d), whereas the other two have low leukemic fractions (PB10d and PB11d). Due to these preprocessing alterations, only 5 clusters were assigned as blasts (3, 6, 7, 9, 11). For some clusters with high healthy proportions, a single sample had most of its cells located in the otherwise normal cluster; in these cases, that sample’s cells were assigned as blasts, others as the corresponding non-malignant cell type. These clusters are noted with red lassoes in Supplementary Fig. 13. Any cells from healthy samples assigned to blast clusters are labeled as “progenitors”. Non-blast clusters are labeled based on canonical marker expression. For healthy samples, the majority label assigned from each leukemia analysis (Fig. S13) is used in the combined analysis. Copy number variation (CNV) prediction analysis was performed to validate our blast annotations using the inferCNV tool from the Trinity CTAT project (https://github.com/broadinstitute/infercnv). The four healthy young adult BM samples from GSE154109 were used as normal reference cells, and a sample’s blast and non-blast (microenvironment) cells were utilized as observation cells. The algorithm outputs a heatmap of predicted CNVs across chromosome regions, along with cell-wise CNV predictions. Samples from patients with known chromosomal abnormalities were used for this analysis, and regions with known abnormalities (e.g. chromosome 11 for KMT2A + B/My MPAL sample PM1d) were compared across leukemia sample blast and microenvironment cells, to confirm our malignant non-malignant cells annotation (Figs. S14–18). Additionally, we also predicted CNVs in the malignant blast cells using normal progenitor cells from healthy samples as a reference to further evaluate the accuracy of CNV prediction and malignant cell annotation (Fig. S14e, f).
Leukemia-Enriched Signature Identification Approach
The analysis for leukemia-enriched signature identification was performed through systematic analysis using multiple R language packages (Table S3). First, differential expression (DE) was performed using Seurat’s FindMarkers function to identify significantly Differentially Expressed Genes (DEGs) (FC (log2)>0.25, Bonferroni adjusted P < 0.05, non-mitochondrial, non-ribosomal genes) by comparing leukemic blast cells to healthy BM cells. Further comparative analysis of the significant DEGs was performed to determine the overlap of genes among different leukemias. The comparative analysis allowed the identification of genes that are common across all leukemias. Further, to select the genes that are over-expressed in the leukemic blast cells and minimally expressed in normal cells, we performed a filtering analysis with the adult healthy BM dataset4. Genes with normalized expression <0.5 across all clusters in the healthy dataset are considered to have a minimal expression in normal cells and therefore leukemogenesis-associated. Jaccard Index (JI), the number of intersecting genes divided by the union of genes in the two sets, was calculated to compare the similarity of DEGs between leukemia types.
Using the bulk RNA-seq TARGET dataset of leukemia and normal BM samples, we performed DE analysis (edgeR lmFit method) to identify genes over-expressed in each leukemia compared to normal BM samples. Candidate genes that were also found in the bulk DEGs, formed the leukemia-specific (i.e. AML, ALL, and MPAL) and leukemia-enriched (common among all leukemias) signatures.
Coverage and Ablation Analysis
We assess the transcriptomic heterogeneity of the signature in our pediatric leukemia single-cell atlas by calculating percent expression across blast versus non-blast normal cell types, samples, leukemia types, and cytogenetic subtypes. Percent expression is calculated as the proportion of cells in a target group with counts above zero for a gene. To examine the robustness of our pediatric leukemia-enriched signature genes, we systematically evaluate the signature by (i) altering filtering thresholds and (ii) removing a random subset of cells and samples before performing differential expression. First, we evaluated the impacts of altering the adjusted p-values and fold change thresholds on the resulting signature genes by testing combinations of these filters. Second, we performed an ablation analysis and repeated the differential expression steps with different subsets of our leukemia cohorts. In the analysis, 5000 blast cells were randomly sampled for each leukemia type along with additional randomly selected 5000 cells from healthy BM samples for comparison. The analysis was permutated 100 times to determine the impact of randomly selected cells on the resulting signature. Additionally, we performed permutation analysis on the bulk RNA-seq data to test the robustness of the signature. The analysis was performed 100 times by randomly selecting 95% of samples from each leukemia type and healthy control group for differential expression analysis.
Processing of Additional External Single-cell Datasets
We obtained and processed additional external healthy and leukemia scRNA-seq datasets from published studies to validate our leukemia-enriched signature and added these datasets to the PedSCAtlas for interactive analysis. Each dataset underwent the following pre-processing steps using Seurat functions. Cells were filtered based on mitochondrial transcript content and feature counts, with specific thresholds for each dataset shown in the supplementary information (Table S3). The high-quality cells underwent log-normalization, scaling, dimensionality reduction, and clustering. Batch correction with Harmony62 was applied for the Roy et al. dataset33 to adjust for tissue types, and for the Lambo et al dataset34 to adjust for patient-specific effects. The malignant and normal cell types were labeled using the annotation provided by the authors of each dataset.
The healthy hematopoiesis scRNA-seq dataset from Roy et al.33 of healthy adult, pediatric, fetal BM, and liver tissues were analyzed after downloading cell count matrices and metadata from GSE155259. To illustrate the leukemia-specific expression of the signature, we calculate the log2 fold change between the expression of each leukemia-enriched gene and the average expression of canonical hematopoietic genes (CD74, CD45/PTPRC, CD34). In addition, we computed the module score for a control signature of genes utilized in the annotation of canonical cell types (Supplementary Data 2) and compared this to the module score of the signature to evaluate overall expression during different developmental stages.
Next, we analyzed the pediatric AML single-cell dataset from Lambo et al.34 after downloading cell count matrices and metadata from GSE235063. Cells without cell type labels were filtered out of the dataset. Expression of the signature genes was compared in malignant and normal cells from AML samples to support malignancy associations.
To assess the expression of pediatric leukemia-enriched genes in adult leukemias, we acquired and analyzed adult AML (van Galen et al.27) and adult MPAL (Granja et al.35) scRNA-seq publicly available datasets27,35. Single-cell expression data and cell metadata were obtained from GSE116256 and https://github.com/GreenleafLab/MPAL-Single-Cell-2019. The signature expression was examined in adult AML malignant blast subtypes (e.g. GMP-like) and compared with normal AML microenvironment cells. Similarly, expression was evaluated in adult MPAL malignant cells compared to healthy-like MPAL cells and normal reference cells (from healthy donors), and across MPAL malignant subtypes (e.g. myeloid-like, lymphoid-like).
Survival Analysis
The survival association of the 9 genes in the leukemia-enriched signature was assessed across the primary TARGET Bulk RNA-seq bone marrow from SurvivalGenie (https://bhasinlab.bmi.emory.edu/SurvivalGenie2/home)65 – TARGET-AML-Legacy, TARGET-ALL-P3, TARGET-ALL-P2-B-ALL, and TARGET-ALL-P2-T-ALL. Instead of stratifying samples into high and low-expression groups based on arbitrary thresholds, we assess the survival associations based on the continuous expression of each gene. To identify the independent survival associations of the pediatric leukemia-enriched signature genes, we include age and cytogenetic group (for AML only) in multivariate survival analysis. The survival association of each gene was determined based on log2(FPKM + 1) normalized data. Cox proportional hazard regression was performed using the survival R package66 to examine the relationship with overall survival, and hazard ratios and p-values were recorded. Survival area plots, which estimate the effect of a continuous variable on the outcome, are generated with the contsurvplot package67.
Gene Set Enrichment Analysis and Module Score
Gene set enrichment analysis (GSEA) was performed using escape Bioconductor package, which provides a framework for testing the enrichment of gene set in the scRNA-seq data68. Hallmark and Reactome gene sets were obtained from the Molecular Signatures Database, and enrichment of these gene sets was calculated in all cells. Differences in enrichment among populations of cells were tested using a student’s t-test. Gene sets with Benjamini-Hochberg corrected p-value of <.05 and a test statistic > 20 were considered significantly over-enriched in leukemia blasts versus healthy cells. In addition, we performed a deconvolution analysis using GSEA to quantify the proportions of immune cell types/subtypes in AML and ALL datasets from the TARGET initiative. The deconvolution analysis was performed using macrophage and T-cell phenotype gene sets compiled from the literature47 (Table S4). The predicted cell type enrichments were explored for correlation with MRD status. To eliminate bias due to blast cell expression in measuring enrichment scores, we only evaluated enrichment for cell types that do not belong to the tumor cell lineage being tested. For example, in AML, we calculated the enrichment of immune cell types, excluding those from the myeloid lineage.
Module scores are calculated to reflect the expression of a group of genes. This is calculated using the AddModuleScore function from the Seurat package61. The module score represents the average expression of a gene set in a cell, subtracted by the aggregated expression of random background genes.
Development of Pediatric Single-Cell Cancer Atlas Web Tool
The PedSCAtlas web resource source code is written in Python and utilizes the Plotly Dash library69. The tool is currently hosted on a Rocky Linux 8.6 (Green Obsidian) server using the Dash Apps program. The user guide and documentation of the tool are located on the GitHub repository for the PedSCAtlas70.
Four scRNA-seq datasets are available for analysis in the PedSCAtlas: (i) the pediatric leukemia single-cell atlas of 82 samples, (ii) a pediatric AML dataset from Lambo et al.34 with 75 samples, (iii) four healthy pediatric BM samples from our lab and Roy et al.33, and (iv) healthy adult BM of ~400,000 cells from Human Cell Atlas initiative4.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The pediatric leukemia single-cell atlas, pediatric AML external dataset (Lambo et al.34), healthy pediatric bone marrow, and healthy adult bone marrow datasets are accessible through the PedSCAtlas [https://bhasinlab.bmi.emory.edu/PediatricSCAtlas/] online resource. Cell count matrices that were generated by our lab during other studies are available on GEO at GSE236351, GSE227122, and GSE235923. Cell count matrices that were obtained from external published studies are available on the ScPCA portal at SCPCP000007 and on GEO at GSE154109, GSE132509, GSE155259, GSE235063, GSE139369, and GSE116256. Processed sequencing data - cell-gene count matrices - from the two healthy pediatric bone marrow samples generated for this study are available at GSE291349; the raw sequencing data in Fastq format are not available due to IRB resection on patient privacy concerns as consent to upload raw sequencing data was not obtained from the parents/legal guardians. The combined cell count matrix and cell type labels for the adult healthy bone marrow dataset were obtained from the Human Cell Atlas data explorer [https://explore.data.humancellatlas.org/projects/cc95ff89-2e68-4a08-a234-480eca21ce79]. The corresponding repository IDs and PedSCAtlas sample IDs are available in Supplementary Data 3. Source data are provided with this paper in the attached Source Data file and athttps://doi.org/10.5281/zenodo.15000525. Source data are provided with this paper.
Code availability
The dash app code for the PedSCAtlas is available on our GitHub repository70 at https://doi.org/10.5281/zenodo.14974956 and https://github.com/bhasin-lab/PedSCAtlas. The code is available via an MIT License.
References
Shalek, A. K. et al. Single-cell RNA-seq reveals dynamic paracrine control of cellular variation. Nature 510, 363–369 (2014).
Villani, A. C. et al. Single-cell RNA-seq reveals new types of human blood dendritic cells, monocytes, and progenitors. Science. https://doi.org/10.1126/science.aah4573 (2017).
Zeisel, A. et al. Cell types in the mouse cortex and hippocampus revealed by single-cell RNA-seq. Science 347, 1138–1142 (2015).
Regev, A. et al. The Human Cell Atlas. Elife 6, e27041 (2017).
Rozenblatt-Rosen, O., Stubbington, M. J. T., Regev, A. & Teichmann, S. A. The Human Cell Atlas: from vision to reality. Nature 550, 451–453 (2017).
Pullen, L. C. Human Cell Atlas Poised to Transform Our Understanding of Organs. Am. J. Transpl. 18, 1–2 (2018).
Hon, C. C., Shin, J. W., Carninci, P. & Stubbington, M. J. T. The Human Cell Atlas: Technical approaches and challenges. Brief. Funct. Genom. 17, 283–294 (2018).
Guo, J. et al. The adult human testis transcriptional cell atlas. Cell Res 28, 1141–1157 (2018).
Hay, S. B., Ferchen, K., Chetal, K., Grimes, H. L. & Salomonis, N. The Human Cell Atlas bone marrow single-cell interactive web portal. Exp. Hematol. 68, 51–61 (2018).
Cool, J., Conroy, R. S., Hanlon, S. E., Hughes, S. K. & Roy, A. L. Spatial and temporal tools for building a human cell atlas. Mol. Biol. Cell 30, 2435–2438 (2019).
Townes, F. W., Hicks, S. C., Aryee, M. J. & Irizarry, R. A. Feature selection and dimension reduction for single-cell RNA-Seq based on a multinomial model. Genome Biol. 20, 295 (2019).
Jia, Q., Chu, H., Jin, Z., Long, H. & Zhu, B. High-throughput single-сell sequencing in cancer research. Signal Transduct. Target. Ther. 7, 145 (2022).
Sun, Y.-F. et al. Dissecting spatial heterogeneity and the immune-evasion mechanism of CTCs by single-cell RNA-seq in hepatocellular carcinoma. Nat. Commun. 12, 4091 (2021).
Nofech-Mozes, I. et al. Pan-cancer classification of single cells in the tumour microenvironment. Nat. Commun. 14,1 (2023).
Zheng, L. et al. Pan-cancer single-cell landscape of tumor-infiltrating T cells. Science 374, https://doi.org/10.1126/science.abe6474 (2021).
Zhang, Z. et al. Integrated analysis of single-cell and bulk RNA sequencing data reveals a pan-cancer stemness signature predicting immunotherapy response. Genome Med. https://doi.org/10.1186/s13073-022-01050-w (2022).
Su, X. et al. Clonal evolution in liver cancer at single-cell and single-variant resolution. J. Hematol. Oncol. https://doi.org/10.1186/s13045-021-01036-y (2021).
Kumar, M. P. et al. Analysis of Single-Cell RNA-Seq Identifies Cell-Cell Communication Associated with Tumor Characteristics. Cell Rep. 25, 1458–1468.e1454 (2018).
Codarri Deak, L. et al. PD-1-cis IL-2R agonism yields better effectors from stem-like CD8(+) T cells. Nature 610, 161–172 (2022).
Faulk, K. E., Anderson-Mellies, A., Cockburn, M. & Green, A. L. Assessment of enrollment characteristics for Children’s Oncology Group (COG) upfront therapeutic clinical trials 2004-2015. PLoS One 15, e0230824 (2020).
Chaudhury, S. et al. Age-specific biological and molecular profiling distinguishes paediatric from adult acute myeloid leukaemias. Nat. Commun. https://doi.org/10.1038/s41467-018-07584-1 (2018).
Anand, P. et al. Single-cell RNA-seq reveals developmental plasticity with coexisting oncogenic states and immune evasion programs in ETP-ALL. Blood 137, 2463–2480 (2021).
Hunter, R. et al. B-cell acute lymphoblastic leukemia promotes an immune suppressive microenvironment that can be overcome by IL-12. Sci. Rep. https://doi.org/10.1038/s41598-022-16152-z (2022).
Mumme, H. et al. Single-cell analysis reveals altered tumor microenvironments of relapse- and remission-associated pediatric acute myeloid leukemia. Nat. Commun. 14, 6209 (2023).
Mumme, H. L. et al. Single-cell RNA sequencing distinctly characterizes the wide heterogeneity in pediatric mixed phenotype acute leukemia. Genome Med. 15, 83 (2023).
Bhasin, S. S. et al. Pediatric T-cell acute lymphoblastic leukemia blast signature and MRD associated immune environment changes defined by single cell transcriptomics analysis. Sci. Rep. 13, 12556 (2023).
Van Galen, P. et al. Single-Cell RNA-Seq Reveals AML Hierarchies Relevant to Disease Progression and Immunity. Cell 176, 1265–1281.e1224 (2019).
Wlodarski, M. W., Collin, M. & Horwitz, M. S. GATA2 deficiency and related myeloid neoplasms. Semin Hematol. 54, 81–86 (2017).
Kirchner, J. & Bevan, M. J. ITM2A is induced during thymocyte selection and T cell activation and causes downregulation of CD8 when overexpressed in CD4(+)CD8(+) double positive thymocytes. J. Exp. Med 190, 217–228 (1999).
Vitali, C. et al. SOCS2 Controls Proliferation and Stemness of Hematopoietic Cells under Stress Conditions and Its Deregulation Marks Unfavorable Acute Leukemias. Cancer Res 75, 2387–2399 (2015).
Sayers, E. W. et al. Database resources of the national center for biotechnology information. Nucleic Acids Res 50, D20–d26 (2022).
Ramezani-Rad, P. et al. SOX4 enables oncogenic survival signals in acute lymphoblastic leukemia. Blood 121, 148–155 (2013).
Roy, A. et al. Transitions in lineage specification and gene regulatory networks in hematopoietic stem/progenitor cells over human development. Cell Rep. https://doi.org/10.1016/j.celrep.2021.109698 (2021).
Lambo, S. et al. A longitudinal single-cell atlas of treatment response in pediatric AML. Cancer Cell. https://doi.org/10.1016/j.ccell.2023.10.008 (2023).
Granja, J. M. et al. Single-cell multiomic analysis identifies regulatory programs in mixed-phenotype acute leukemia. Nat. Biotechnol. 37, 1458–1465 (2019).
Pattabiraman, D. R. & Gonda, T. J. Role and potential for therapeutic targeting of MYB in leukemia. Leukemia 27, 269–277 (2013).
Nakata, Y. et al. c-Myb contributes to G2/M cell cycle transition in human hematopoietic cells by direct regulation of cyclin B1 expression. Mol. Cell Biol. 27, 2048–2058 (2007).
Elsayed, A. H. et al. A 6-gene leukemic stem cell score identifies high risk pediatric acute myeloid leukemia. Leukemia 34, https://doi.org/10.1038/s41375-019-0604-8 (2020).
Singh, M. K. et al. HEI10 negatively regulates cell invasion by inhibiting cyclin B/Cdk1 and other promotility proteins. Oncogene 26, 4825–4832 (2007).
Vu, L. P. et al. Functional screen of MSI2 interactors identifies an essential role for SYNCRIP in myeloid leukemia stem cells. Nat. Genet. 49, 866–875 (2017).
Tanouchi, A. et al. CCDC88A, a prognostic factor for human pancreatic cancers, promotes the motility and invasiveness of pancreatic cancer cells. J. Exp. Clin. Cancer Res. 35, 190 (2016).
Belinky, F. et al. PathCards: multi-source consolidation of human biological pathways. Database (Oxford) https://doi.org/10.1093/database/bav006 (2015).
Gorski, J. J. et al. A novel TBP-associated factor of SL1 functions in RNA polymerase I transcription. EMBO J. 26, 1560–1568 (2007).
Wolf, J. & Passmore, L. A. mRNA deadenylation by Pan2-Pan3. Biochem Soc. Trans. 42, 184–187 (2014).
Du, A. et al. Fatty acids in cancer: Metabolic functions and potential treatment. MedComm – Oncol. 2, e25 (2023).
Jovanovska, A. et al. Clinical Significance of Minimal Residual Disease at the End of Remission Induction Therapy in Childhood Acute Lymphoblastic Leukemia. Open Access Maced. J. Med Sci. 7, 2818–2823 (2019).
Chu, Y. et al. Pan-cancer T cell atlas links a cellular stress response state to immunotherapy resistance. Nat. Med. 29, 1550–1562 (2023).
Sanchez-Martin, M. & Ferrando, A. The NOTCH1-MYC highway toward T-cell acute lymphoblastic leukemia. Blood 129, 1124–1133 (2017).
Wilczynska, A. & Bushell, M. The complexity of miRNA-mediated repression. Cell Death Differ. 22, 22–33 (2015).
Zheng, G. X. Y. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Li, M. et al. DISCO: a database of Deeply Integrated human Single-Cell Omics data. Nucleic Acids Res. 50, D596–D602 (2021).
Lin, D. et al. CellDepot: A Unified Repository for scRNA-seq Data and Visual Exploration. J. Mol. Biol. 434, 167425 (2022).
Barrett, T. et al. NCBI GEO: archive for functional genomics data sets—update. Nucleic Acids Res. 41, D991–D995 (2012).
Bailur, J. K. et al. Risk-associated alterations in marrow T cells in pediatric leukemia. JCI Insight 5, https://doi.org/10.1172/jci.insight.140179 (2020).
Hawkins, A. G. et al. The Single-cell Pediatric Cancer Atlas: Data portal and open-source tools for single-cell transcriptomics of pediatric tumors. bioRxiv, 2024.2004.2019.590243, https://doi.org/10.1101/2024.04.19.590243 (2024).
Li, M. et al. Downregulation of GNA15 Inhibits Cell Proliferation via P38 MAPK Pathway and Correlates with Prognosis of Adult Acute Myeloid Leukemia With Normal Karyotype. Front Oncol. 11, 724435 (2021).
Fang, X. et al. SNORD126 promotes HCC and CRC cell growth by activating the PI3K-AKT pathway through FGFR2. J. Mol. Cell Biol. 9, 243–255 (2017).
Man, Y. N., Sun, Y., Chen, P. J., Wu, H. & He, M. L. TAF1D Functions as a Novel Biomarker in Osteosarcoma. J. Cancer 14, 2051–2065 (2023).
Zhang, K., Qian, Y., Quan, X., Zhu, T. & Qian, B. A Novel Signature of Lipid Metabolism-Related Gene Predicts Prognosis and Response to Immunotherapy in Lung Adenocarcinoma. Front. Cell Dev. Biol. https://doi.org/10.3389/fcell.2022.730132 (2022).
Abou Dalle, I., Jabbour, E. & Short, N. J. Evaluation and management of measurable residual disease in acute lymphoblastic leukemia. Ther. Adv. Hematol. 11, 2040620720910023 (2020).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e3529 (2021).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
GenomicDataCommons: NIH / NCI Genomic Data Commons Access (Bioconductor 3.19, 2024).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47–e47 (2015).
Dwivedi, B., Mumme, H., Satpathy, S., Bhasin, S. S. & Bhasin, M. Survival Genie, a web platform for survival analysis across pediatric and adult cancers. Sci. Rep. https://doi.org/10.1038/s41598-022-06841-0 (2022).
Therneau, T. M. & Grambsch, P. M. Modeling Survival Data: Extending the Cox Model. (Springer, 2000).
Denz, R. & Timmesfeld, N. Visualizing the (Causal) Effect of a Continuous Variable on a Time-To-Event Outcome. Epidemiology (Cambridge, Mass.). https://doi.org/10.1097/EDE.0000000000001630 (2023).
Borcherding, N. et al. Mapping the immune environment in clear cell renal carcinoma by single-cell genomics. Commun. Biol. 4, 122 (2021).
Hossain, S. Visualization of Bioinformatics Data with Dash Bio. In Proc Python in Science Conference, 126–133, https://doi.org/10.25080/majora-7ddc1dd1-012 (2019).
Mumme, H. L. et al. Identification of Leukemia-Enriched Signature Through the Development of a Comprehensive Pediatric Single-Cell Atlas. Zenodo, https://doi.org/10.5281/zenodo.14974956 (2025).
Acknowledgements
Thanks to Taylor Lawrence and the Aflac Biorepository and Children’s Healthcare of Atlanta staff for providing samples along with de-identified patient information. Thanks to Beena Thomas for single cell assays. The study is supported through funding from the CURE Childhood Cancer Foundation (DKG) and Emory startup funds (MBh). Next generation sequencing services were provided by the Emory NPRC Genomics Core which is supported in part by NIH P51 OD011132. Sequencing data was acquired on an Illumina NovaSeq 6000 funded by NIH S10 OD026799.
Author information
Authors and Affiliations
Contributions
S.S.R., M.Q., D.D., S.M.C., D.S.W., C.C.P., and D.K.G. collected bone marrow samples and clinical data. MBa performed tissue extraction and single-cell assays. H.L.M., S.S.B., and M.B.h designed the study. H.L.M., D.O., and C.H. analyzed the data. H.L.M. and C.H. developed the web tool. H.L.M., D.O., C.H., S.S.B., D.S.W., C.C.P., D.K.G., and M.Bh. interpreted the results and wrote the manuscript. M.B.h supervised the study.
Corresponding author
Ethics declarations
Competing interests
MBh serves on the board of Canomiks Inc. as chief scientific advisor and has equity in it. DKG and DD hold equity in Meryx Inc. SSB serves as CEO of Anxomics LLC and has equity in it. The remaining authors declare no other competing interests.
Peer review
Peer review information
Nature Communications thanks Karen Keeshan, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Mumme, H.L., Huang, C., Ohlstrom, D. et al. Identification of leukemia-enriched signature through the development of a comprehensive pediatric single-cell atlas. Nat Commun 16, 4114 (2025). https://doi.org/10.1038/s41467-025-59362-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-59362-5