Abstract
To address the need for broadly protective SARS-CoV-2 vaccines, we developed an attenuated a SARS-CoV-2 vaccine virus that lacks the open reading frames of two viral structural proteins: the envelope (E) and membrane (M) proteins. This vaccine virus (ΔEM) replicates in a cell line stably expressing E and M but not in wild-type cells. Vaccination with ΔEM elicits a CD8 T-cell response against the viral spike and nucleocapsid proteins. Two vaccinations with ΔEM provide better protection of the lower respiratory tissues than a single dose against the Delta and Omicron XBB variants in hamsters. Moreover, ΔEM is effective as a booster in hamsters previously vaccinated with an mRNA-based vaccine, providing higher levels of protection in both respiratory tissues compared to the mRNA vaccine booster. Collectively, our data demonstrate the feasibility of a SARS-CoV-2 ΔEM vaccine candidate virus as a vaccine platform.
Similar content being viewed by others
Introduction
The SARS-CoV-2 virion consists of four structural proteins—the spike (S), envelope (E), membrane (M), and nucleoprotein (N) proteins—which are encoded in the last third of the viral RNA genome along with accessory proteins. The other two-thirds of the genome encodes non-structural proteins that function in viral RNA replication. Most vaccines against SARS-CoV-2 induce neutralizing antibodies against the S protein due to its key role in viral entry and high immunogenicity. Although these vaccines, based on mRNA, viral vectors, recombinant protein, and inactivated virus, induce robust neutralizing antibodies and systemic immune responses when administered intramuscularly, these responses are spike-specific and the induction of nasal mucosal immunity is inadequate1,2.
The frequent emergence of SARS-CoV-2 variants harboring multiple immune-evasive amino acid substitutions in the S protein has highlighted the need to explore approaches that target viral proteins other than the S protein3,4,5,6. Most mutations observed in the variants have accumulated in the S protein due to its higher mutation rate compared to other viral proteins, which is largely driven by selective pressure from the host immune response leading to the evolution of new variants7,8. While neutralizing antibodies mainly target the S protein, the epitopes targeted by T-cell responses are not limited to the S protein and can include the entire regions of both the surface and internal viral proteins9,10,11,12. Ideally, vaccines should induce both robust protective humoral and cellular immunity that can be reactive against several viral proteins at the site of initial infection in response to possible future SARS-CoV-2 variants.
To overcome the limitations of current vaccines, alternative strategies have been being implemented, such as an S protein booster to induce a mucosal immune response13, intranasal vaccines designed to induce S-independent CD8 T-cell immunity14, and dual antigen presenting vaccine platforms (e.g., self-amplifying RNA3,4, adenovirus vectors15,16 and modified vaccinia virus vectors17). In addition, live-attenuated vaccine (LAV) viruses that have limited replicative ability and reduced virulence are being developed. Strategies of SARS-CoV-2 attenuation have been achieved by genetic modification using codon-pair deoptimization18,19,20,21, accessory gene deletions22, and cold-adaptation23,24,25. Intranasal administration of LAV viruses would be expected to elicit local mucosal, humoral, and cellular immunity targeting multiple viral proteins, which would be beneficial in future pandemics caused by respiratory pathogens including SARS-CoV-226,27.
Here, we describe a gene-deletion attenuation approach that involves the generation of a SARS-CoV-2 isolate that lacks the open reading frames for two structural proteins, the E and M proteins, which we termed the ΔEM vaccine virus. Unlike current LAVs against SARS-CoV-2, the ΔEM vaccine candidate virus cannot undergo multiple rounds of virus replication in wild-type cells, can complete a single-round of replication allowing for expression of viral proteins without producing infectious progeny, thereby offering immune responses similar to those induced by natural infection but without any disease. In this study, we evaluated the protective efficacy of the ΔEM vaccine candidate virus in rodents and characterized their immune responses to ΔEM vaccination.
Results
Generation of an attenuated SARS-CoV-2 ΔEM virus
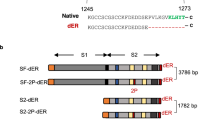
To generate a virus that is capable of only a single round of replication, we developed a SARS-CoV-2 ΔEM vaccine candidate virus by deleting the entire open reading frames (ORFs) for the E and M proteins from the genome of the SARS-CoV-2 isolate, Wuhan-Hu-1 (Fig. 1a). Using HEK293T cells that stably express the E and M proteins, we generated the ΔEM virus by reverse genetics using the circular polymerase extension reaction (CPER)28,29. The virus was then propagated on Vero TMPRSS2 cells stably expressing E and M proteins (Vero TMPRSS2/EM) that were generated by retrovirus transduction followed by transfection with a protein expression plasmid encoding human codon-optimized SARS-CoV-2 E and M, respectively, along with antibiotic-resistant genes. Single clones were selected based on the expression of both viral proteins. The clone that allowed ΔEM virus replication with the highest virus titer was used for these studies. In this stable cell line, the ΔEM virus replicated efficiently resulting in virus-induced cytopathic effects (CPE) (Fig. 1b, left panel). However, in wild-type Vero TMPRSS2 cells, there was limited cell-to-cell spread of the ΔEM virus, which resulted in the formation of foci, but no CPE (Fig. 1b, right panel; Supplementary Fig. 1).
a Schematic diagram of the viral genomes of the parental virus (top) and the ΔEM vaccine candidate virus (bottom) with the dashed lines around ORF E and ORF indicating the removal of the ORFs from the genome. b Infection of Vero TMPRSS2/EM cells (left panel) or Vero TMPRSS2 cells (right panel) with the ΔEM vaccine candidate virus. The SARS-CoV-2 N protein was visualized by DAB staining using an anti-N antibody. c Growth kinetics of the ΔEM vaccine candidate virus in Vero TMPRSS2/EM cells (MOI of 0.1 or 0.01) or Vero TMPRSS2 cells (MOI of 0.1). Virus titers in the cell supernatants were determined on Vero TMPRSS2/EM cells. The Day 0 samples were collected from media added immediately after washing unbound virus from the cells. Data represent the mean of experiments repeated three times (Vero TMPRSS2/EM cells) or twice (Vero TMPRSS2 cells), respectively. Error bars represent standard deviation (SD). ND; not detected. The limit of detection of infectious virus was 20 pfu/ml. d Body weight changes in hACE2 mice infected with the recombinant parental virus (Wuhan-Hu-1) or the ΔEM virus (mean with SD, n = 6 females/group). Statistical significance was determined by use of the unpaired two-tailed Student’s t-test (*P < 0.05, **P < 0.01, ****P < 0.0001). Exact Pvalues: day 4 (P = 0.013253), day 5 (P = 0.000006), day 6 (P < 0.000001), day 7 (P = 0.000001), day 8 (P = 0.000051), day 9 (P = 0.000042), day 10 (P = 0.000119), day 11 (P = 0.002217). e Survival of hACE2 mice infected with the recombinant parental virus (Wuhan-Hu-1) or the ΔEM virus (mean with SD, n = 6 females/group). Statistical significance was determined by use of the Log-rank Mantel-Cox test (**P = 0.004). f, g Virus titers of the recombinant parental virus (Wuhan-Hu-1) or the ΔEM vaccine candidate virus in lung, nasal turbinate (NT), and brain tissues of infected hACE2 mice (f) and hACE2 hamsters (g) (mean with SD, n = 4 females/virus). Tissues were collected 3 days after infection. Each dot in the bar graph indicates an individual animal in each group. ND; not detected. The dotted lines indicate the lower limit of detection (1.3 log10 pfu/g). Source data are provided as a Source Data file.
To examine the infectious progeny virus in the supernatant of infected cells, Vero TMPRSS2/EM cells were infected with the ΔEM virus at a multiplicity of infection (MOI) of 0.01. We observed an increase in virus titers over 3 days after infection, reaching a titer of more than 1 × 105 plaque-forming units (pfu)/ml (Fig. 1c). Infection of Vero TMPRSS2/EM cells at a higher MOI (0.1) resulted in less robust virus growth with CPE by day 2 after infection (Fig. 1c). The titer of ΔEM virus was lower than that of the parental recombinant virus, which can reach a titer of 107 pfu/ml; this may be due to differences in the expression kinetics of the viral E and M proteins between viral infection and cells stably expressing these proteins.
When Vero TMPRSS2 cells were infected with the ΔEM virus at an MOI of 0.1, there was no detectable infectious virus through three days after infection (Fig. 1c), confirming that the ΔEM virus is replication-deficient on wild-type cells.
To demonstrate the lack of replication of the ΔEM virus in vivo, we inoculated human (h)ACE2 transgenic (K18-hACE2) mice (females, n = 6/group) intranasally with 1 × 104 pfu of the ΔEM virus or the recombinant parental virus (Wuhan-Hu-1) as a positive control. K18-hACE2 mice infected with the parental virus showed a significant reduction in body weight starting at day 4 after infection with five of the six mice succumbing to infection within 10 days of inoculation (Fig. 1d, e). In contrast, K18-hACE2 mice infected with the ΔEM virus did not lose weight and all the mice survived (Fig. 1d, e).
To determine if whether infectious ΔEM virus was present in the tissues of the infected K18-hACE2 mice (n = 4 females/virus), lung, nasal turbinate, and brain tissues were collected 3 days after infection, and clarified tissue homogenates were used to determine virus titers on Vero TMPRSS2/EM cells. While high virus titers were detected in all tissues collected from animals infected with the wild-type virus, no infectious virus was detected in any of the tissues from the ΔEM virus-infected animals (Fig. 1f). We confirmed the lack of replication of the ΔEM virus in hACE2 transgenic hamsters (n = 4 females/virus). As expected, the parental virus replicated efficiently in the lung, nasal turbinate, and brain tissues, whereas there was no detectable infectious ΔEM virus in any of these tissues at 3 days after inoculation (Fig. 1g), confirming the lack of replication of the ΔEM virus in vivo.
Protective efficacy of ΔEM vaccination in K18-hACE2 mice
To assess the protective efficacy of the ΔEM vaccine candidate virus, we immunized groups of K18-hACE2 mice with 1 x 104 pfu of the ΔEM virus by intranasal inoculation.
Mice (females; n = 12 in each vaccinated group, n = 8 in the control group) were vaccinated with one or two doses of the ΔEM virus or were inoculated with PBS as a control. A second vaccination was given to one group four weeks after the initial vaccination. Four weeks after the last vaccination, mice were challenged with an ancestral SARS-CoV-2 isolate (1 x 105 pfu). After the challenge, neither of the ΔEM virus-vaccinated groups exhibited weight loss and all vaccinated mice survived the challenge (Fig. 2a, b). In contract, mice in the control group started to show significant weight loss on day 4 after challenge, with three of the eight mice succumbing to infection by day 11 after challenge (Fig. 2a, b).
a, b Body weight changes (a) and survival rate (b) (n = 12 females in each vaccinated group, n = 8 females in the control group) of hACE2 mice vaccinated once or twice with the ΔEM virus and then challenged with an ancestral SARS-CoV-2 isolate. Data represent the mean, and error bars indicate SD (a). Statistical significance was determined by use of the two-tailed Dunnett’s multiple comparisons test (a; *P < 0.05, **P < 0.01, ****P < 0.0001) and the Log-rank Mantel-Cox test (b; *P = 0.033). Exact P values: (a) control vs. ΔEM x2: day 4 (P = 0.0360), day 5 (P = 0.0012), day 6 (P < 0.0001), day 7 (P < 0.0001), day 8 (P = 0.0022), day 9 (P = 0.0348); control vs. ΔEM x1: day 5 (P = 0.0020), day 6 (P < 0.0001), day 7 (P = 0.0001), day 8 (P = 0.0064). c Efficacy of one or two vaccinations of the ΔEM vaccine candidate virus. Virus titers in the lung and nasal turbinate (NT) tissues of K18-hACE2 mice (mean with SD, n = 8 females/group) at 3 days after challenge with an ancestral SARS-CoV-2 isolate. The dotted line indicates the limit of detection (1.3 log10 pfu/g). Each dot in the bar graph indicates an individual mouse in each group. Statistical significance was determined by use of a one-way ANOVA with Dunnett’s multiple comparisons test (****P < 0.0001). Source data are provided as a Source Data file.
To evaluate the protective effect on respiratory tissues, lung and nasal turbinate were collected from K18-hACE2 mice (n = 8 females/group) three days after challenge. Mice in the control group had high virus titers in the lung and nasal turbinate tissues, close to 107 pfu/g and 106 pfu/g, respectively (Fig. 2c). A single vaccination resulted in a significant reduction of viral titers in both tissues by approximately 1000–10,000-fold compared to the control group. Two of the eight mice that received a single vaccination had no detectable infectious virus in their lung tissue. Two vaccinations provided better protection than the single vaccination, with no infectious virus detected in both respiratory tissues (Fig. 2c).
Immune responses in ΔEM-vaccinated K18-hACE2 mice
The anticipated benefit of intranasal administration of LAV vaccines is their potential to induce local mucosal, humoral, and cellular immunity targeting multiple viral proteins27. Therefore, we first analyzed the induction of S-specific IgA in the respiratory tract of ΔEM virus-vaccinated mice. K18-hACE2 mice (n = 7-8 females/group) were intranasally inoculated with ΔEM virus at 104 pfu. As a control, another group of mice (n = 4 females/group) was vaccinated with 1 μg of an mRNA vaccine (BNT162b2) by intramuscular injection. Four weeks after vaccination, mice were challenged with an ancestral SARS-CoV-2 isolate (1 x 105 pfu). Nasal wash and bronchoalveolar-lavage fluid (BALF) were collected on days 2 and 5 after challenge and S-specific IgA antibody was measured by an ELISA (Fig. 3a).
a Induction of spike-specific IgA in nasal wash and bronchoalveolar lavage fluid (BALF) of ΔEM virus-vaccinated mice (n = 4 females for the mock infection group; n = 6 females for the infection groups except that nasal wash samples on day 5 post-infection was collected from 7 females), mRNA-vaccinated mice (n = 6 females for both the mock and infection groups), or mock-vaccinated mice (n = 4 females) as measured by an ELISA. Samples were collected on days 2 or 5 following infection with an ancestral SARS-CoV-2 isolate. b, c Induction of an antigen-specific T cell population producing IFN-γ in the lungs of ΔEM virus- or mRNA-vaccinated mice (n = 6 females/group) as measured by use of an ELISpot assay. Representative ELISpot wells of cells isolated from lung tissue stimulated with S peptide pool (b; 2 x 105 cells/well) or N peptide pool (c; 4 x 105 cells/well) are shown. d, e The frequency of IFN-γ- and TNF-α-positive CD4+ and CD8+ T cells in the lungs of ΔEM virus- or mRNA-vaccinated mice as measured by flow cytometry. Cells collected from lung tissue were stimulated with S peptide pool (d, n = 6 females) or N peptide pool (e, n = 6 females). Box plots show the median center line and 10/90 percentiles. Whiskers show min and max values. Statistical significance was determined by use of a one-way ANOVA with Tukey’s multiple comparisons test (a, d, e) and a one-way ANOVA with Dunnett’s multiple comparisons test (b, c) (*P < 0.05, **P < 0.01, ****P < 0.0001; ns, not significant). Exact P values: (a) nasal wash samples on day 5 post-infection: ΔEM x1 vs. mock vaccination control (P = 0.0051), ΔEM x1 vs. mRNA x1 (P = 0.0944); BALF samples on day 5 post-infection: ΔEM x1 vs. mock vaccination control (P = 0.0492), ΔEM x1 vs. mRNA x1 (P = 0.0262). (b) ΔEM vs. mock (P = 0.0284), ΔEM vs. mRNA 0.1 μg (P = 0.0452), ΔEM vs. mRNA 1.0 μg (P = 0.7588). (c) ΔEM vs. mock (P < 0.0001), ΔEM vs. mRNA 0.1 μg (P < 0.0001), ΔEM vs. mRNA 1.0 μg (P < 0.0001). (d) S-reactive CD4+ T cells frequency: ΔEM vs. mock (P = 0.0109), ΔEM vs. mRNA 0.1 μg (P = 0.0494), ΔEM vs. mRNA 1.0 μg (P = 0.9897), mock vs. mRNA 1.0 μg (P = 0.0213); S-reactive CD8+ T cells frequency: ΔEM vs. mock (P = 0.0015), ΔEM vs. mRNA 0.1 μg (P = 0.0310), ΔEM vs. mRNA 1.0 μg (P = 0.9140), mock vs. mRNA 1.0 μg (P = 0.0066). (e) N-reactive CD4+ T cells frequency: ΔEM vs. mock (P = 0.9932), ΔEM vs. mRNA 0.1 μg (P = 0.9847), ΔEM vs. mRNA 1.0 μg (P = 0.5050), mock vs. mRNA 1.0 μg (P = 0.3592); N-reactive CD8+ T cells frequency: ΔEM vs. mock (P = 0.0076) ΔEM vs. mRNA 0.1 μg (P = 0.0076), ΔEM vs. mRNA 1.0 μg (P = 0.0144), mock vs. mRNA 1.0 μg (P = 0.9915). Source data are provided as a Source Data file.
In the nasal wash samples, there was no significant difference in the IgA antibody levels between the ΔEM- and mRNA-vaccinated groups at day 2 after challenge. However, by day 5 after challenge, the IgA antibody levels in the ΔEM-vaccinated mice were higher than those in the mRNA- and mock-vaccinated mice. Similarly, in BALF samples, IgA antibody levels in ΔEM-vaccinated mice were significantly higher at day 5 after challenge compared to mRNA- and mock-vaccinated mice.
Elicitation of resident memory T cells in the respiratory tract is crucial to rapidly limit viral spread and disease severity1,30,31. Th1-biased IgG subclasses, IgG2c and IgG3, were strongly induced in ΔEM-vaccinated mice (K18-hACE2 female mice, single vaccination, n = 8/group) compared to Th2-biased IgG subclasses (Supplementary Fig. 2), suggesting that cell-mediated immune responses are integral to the respiratory tract defense facilitated by ΔEM vaccination. Therefore, we analyzed the induction of antigen-specific T cells in the lungs of the ΔEM virus-vaccinated mice. Control groups of mice (K18-hACE2 female mice, single vaccination, n = 6/group) were vaccinated with two different amounts (1.0 and 0.1 μg) of an mRNA vaccine (BNT162b2) by intramuscular injection. Two weeks after vaccination, cells collected from the lung tissue were stimulated with either S or N peptide pools and the induction of antigen-specific T cells was qualified by using an IFN-γ enzyme-linked immunospot (ELISpot) assay. Vaccination with the ΔEM virus induced SARS-CoV-2 S- and N-reactive T cells secreting IFN-γ (Fig. 3b, c), whereas mRNA vaccination induced only S-reactive IFN-γ-secreting T cells in the lung, as expected2. The S-reactive T cells induced in the lungs of the ΔEM-vaccinated mice were comparable to those in those receiving high amount of mRNA vaccine. Subsequently, we used flow cytometry to analyze the frequencies of helper CD4+ T cells and cytotoxic CD8+ T cells secreting both IFN-γ and TNF-α32,33. The numbers of S-reactive CD4+ and CD8+ T cells were significantly increased in the lung tissue of the ΔEM-vaccinated animals compared to in the non-vaccinated control group and the animals immunized with the low dose of mRNA vaccine and comparable to those in animals immunized with the high dose of mRNA vaccine (Fig. 3d). Although induction of N-reactive CD4+ T cells by ΔEM vaccination was limited, the number of N-reactive CD8+ T cells was significantly higher in the lung tissue of the ΔEM-vaccinated animals compared to the non-vaccinated control and mRNA-vaccinated animals (Fig. 3e). Collectively, these results suggest that ΔEM vaccination induces lung resident T-cell responses comprising S-reactive CD4+ T cells and S/N-reactive CD8+ T cells.
Protective efficacy of ΔEM vaccination against SARS-CoV-2 variants in hamsters
We next explored the protective efficacy of the ΔEM vaccine candidate virus against a SARS-CoV-2 Delta variant34 and the Omicron variant XBB35. Wild-type Syrian hamsters (n = 4 females) were vaccinated intranasally with 2 × 104 pfu of the ΔEM virus. Six weeks later, the animals were challenged with the Delta variant (8 × 104 pfu). On day 3 after challenge, this single vaccination resulted in an approximately 10-fold reduction in virus titers in the lung tissue with no infectious virus detected in the lung tissue of one of the ΔEM-vaccinated hamsters; however, there was no reduction in virus titers in the nasal turbinate tissues of the same animals compared to the control group (n = 4 females; Supplementary Fig. 3a). In vaccinated hamsters (n = 4 females) challenged with Omicron XBB (1 × 105 pfu), there was a 10–100-fold reduction of the challenge virus titers in the lung and nasal turbinate tissues compared to animals in the control group (n = 4 females), with no infectious virus detected in the lung tissue of one of the vaccinated animals (Supplementary Fig. 3b).
Next, we examined the efficacy of two doses of ΔEM virus (2 × 104 pfu, intranasally) given four weeks apart. In addition, we evaluated vaccination in female and male hamsters. Six weeks after the second vaccination, one group of animals was challenged with the Delta variant (8 x 104 pfu). On day 3 after challenge, the Delta variant replicated efficiently in both female (n = 7) and male (n = 4) hamsters in the respiratory tissues of mock-vaccinated control animals (lung tissue: 108 pfu/g; nasal turbinate tissue: 107 pfu/g) (Fig. 4a). In contrast, no infectious virus was detected in the lungs of any of the vaccinated hamsters (n = 8 females, n = 4 males) (Fig. 4a). In the nasal turbinate, virus was detected in 3 of the 8 vaccinated female hamsters, but at significantly lower virus titers and not in any of the vaccinated male hamsters (Fig. 4a).
Efficacy of two vaccinations with the ΔEM vaccine candidate virus in hamsters. a Virus titers in the lung and nasal turbinate (NT) tissues on day 3 after challenge with a Delta variant (n = 7 female and n = 4 male control animals; n = 8 female and n = 4 male vaccinated animals) or an Omicron XBB variant (n = 6 female and n = 4 male control animals; n = 8 female and n = 4 male vaccinated animals). b Virus titers on day 6 after challenge with a Delta variant (n = 6 female and n = 4 male control animals; n = 6 female and n = 6 male vaccinated animals) or an Omicron XBB variant (n = 6 female and n = 6 male control animals; n = 6 female and n = 6 male vaccinated animals). Data represent the mean, and each dot in the bar graph indicates an individual hamster. Error bars represent SD. The dotted line indicates the limit of detection (1.3 log10 pfu/g). Statistical significance was determined by use of a two-way ANOVA with Šídák’s multiple comparisons test (*P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001). Exact P values: (a) Delta-challenged female lung (P < 0.0001), female NT (P < 0.0001), male lung (P < 0.0001), male NT (P < 0.0001); XBB-challenged female lung (P < 0.0001), female NT (P < 0.0001), male lung (P < 0.0001), male NT (P < 0.0001). (b) Delta-challenged female lung (P = 0.0036), female NT (P = 0.0171), male lung (P < 0.0001), male NT (P = 0.0128); XBB-challenged female lung (P = 0.0165), female NT (P = 0.0004), male lung (P = 0.0209), male NT (P = 0.0012). Source data are provided as a Source Data file.
Challenge with the Omicron XBB variant (1 × 105 pfu) after two vaccinations yielded similar results. On day 3 after challenge, no infectious virus was detected in the lungs of the vaccinated animals (n = 8 females, n = 4 males), and mean virus titers in the nasal turbinate of vaccinated hamsters were reduced by 10,000-fold compared to mock-vaccinated control animals (n = 6 females, n = 4 males) (Fig. 4a).
At six days after challenge, infectious virus was still detectable but at lower levels in the respiratory tissues of many control animals inoculated with either the Delta variant (n = 6 females, n = 4 males) or the Omicron XBB variant (n = 6 females, n = 6 males). No infectious virus was detected at this timepoint in the respiratory tissues of the vaccinated animals (Fig. 4b).
Protection against the Delta variant conferred by two doses of ΔEM vaccination was nearly as strong as that observed in hamsters that were initially infected with an ancestral SARS-CoV-2 isolate and re-infected with the Delta variant eight weeks later. No detectable infectious virus was found in the lung and nasal turbinate tissues of previously infected hamsters on day 3 after challenge with the Delta variant (Supplementary Fig. 4).
By using sera collected one day prior to a second vaccination (27 days after the first vaccination) or the challenge (41 days after the second vaccination), we determined neutralizing antibody titers against an ancestral SARS-CoV-2 isolate using a focus reduction neutralization test with titers reported as the reciprocal of the dilution at which the number of foci is reduced by 50% (FRNT50). No neutralizing antibodies were detected after the first vaccination with ΔEM virus (n = 4 females, n = 4 males; limit of detection 1:20 dilution of serum) while sera from hamsters vaccinated twice with the ΔEM virus (n = 6 females, n = 6 males) had FRNT50 values that ranged 51 to 209 (geometric mean of 131) (Supplementary Fig. 5a). In the sera collected from hamsters 55 days after a previously infection with an ancestral SARS-CoV-2 isolate (n = 3 females), neutralizing antibodies titers were higher, with FRNT50 values ranging from 767 to 1269 (geometric mean: 1096) (Supplementary Fig. 5a).
Next, we examined the neutralizing antibodies against the two challenge viruses. The sera from hamsters vaccinated twice with the ΔEM virus (n = 6 females, n = 6 males) had FRNT50 values that ranged from 42 to 229 (geometric mean of 119) against the Delta variant, which differs from the ancestral isolate by only two amino acids in the receptor-binding domain (Supplementary Fig. 5b). Conversely, there was no neutralizing activity against the Omicron XBB variant (Supplementary Fig. 5b), as previously reported by others35,36.
We also compared the pathological features in the lung tissue at day 6 after challenge between the control and vaccinated hamsters after challenge with the Delta variant. Histopathological analysis was conducted on hematoxylin and eosin (H&E)-stained lung sections from the control group (n = 4 females; n = 4 males) and the double-vaccinated group (n = 6 females; n = 6 males). Histopathological changes were assessed by a third-party pathologist using a semi-quantitative, 5-point grading scheme based on 4 different histopathological parameters: perivascular inflammation, bronchial or bronchiolar epithelial degeneration or necrosis, bronchial or bronchiolar inflammation. In 7 of 12 hamsters in the ΔEM vaccinated group, minor inflammation within the lungs was the only noted pathology (Fig. 5a) with individual total pathology scores of 0/16 to 1/16 (Fig. 5b). In contrast, there was more severe pathology noted for all four scored parameters (Fig. 5a) in the control hamsters with individual total pathology scores ranging from 9/16 to 13/16 (Fig. 5b).
Semi-quantitative pathological scores of individual hamsters analyzed using the indicated four parameters (a). A 5-point scoring system (0-within normal limits, 1-mild, 2-moderate, 3-marked, 4-severe) was used. A total pathology score was calculated for each hamster by adding the individual histopathological feature scores (b). A maximum total pathology score of 16 is possible for an individual hamster (n = 4 females, n = 4 males for non-vaccinated controls, n = 6 females, n = 6 males for vaccinated hamsters). Statistical significance was determined by use of a two-way ANOVA with Šídák’s multiple comparisons test (****P < 0.0001). Data represent the mean, and error bars indicate SD (a and b). Each symbol indicates an individual hamster. Source data are provided as a Source Data file.
Collectively, these results suggest that two vaccinations with the ΔEM virus could confer promising protection in the lower respiratory tract against possible future variants.
The ΔEM vaccine candidate virus as a booster vaccine in hamsters
Lastly, given the certain levels of immunity against SARS-CoV-2 either by vaccination or natural infection in human populations, we determined the efficacy of the ΔEM vaccine candidate virus as a booster vaccination.
We first assessed the systemic IgG responses against the spike proteins of an ancestral SARS-CoV-2 isolate with a D614G spike and the Omicron XBB variant. Groups of hamsters received the mRNA vaccine (group #1), the mRNA vaccine followed by a booster vaccination with the ΔEM vaccine candidate virus (group #2), or the ΔEM virus. Serum samples after the first vaccination but prior to the booster showed similar ELISA endpoint dilution titers of binding IgG antibodies against both spikes (the average titers ranged from 1:1360 to 1:3200); there were no significant differences between any of the vaccinated groups (Fig. 6a). After the boost vaccination with the ΔEM virus, the IgG antibody endpoint titers significantly increased by about 5–11-fold compared to those prior to the boost vaccination, and average titers ranged from 1:15360 to 1:23040, but with no significant difference between the vaccinated groups (Fig. 6a). Both vaccines were based on the original Wuhan-Hu-1 isolate, but the induced IgG antibodies in the sera reacted against both the ancestral spike protein and the Omicron XBB spike protein.
a Total IgG antibody endpoint titers against the spike proteins of an ancestral isolate (spike D614G) or Omicron XBB using hamster sera after one or two vaccinations with the ΔEM virus. Each dot in the bar graph indicates an individual hamster in each group (mean with SD, n = 4 females/group). Statistical significance was determined by using a two-way ANOVA with Šídák’s multiple comparisons test (*P < 0.05, **P < 0.01, ***P < 0.001). b Efficacy of the ΔEM vaccine virus as a booster vaccine. Virus titers of the Omicron XBB variant in lung and nasal turbinate (NT) tissues at day 3 after challenge. Each dot in the bar graph indicates an individual hamster in each group (mean with SD, n = 4 females/group). Statistical significance was determined by using the unpaired two-tailed Student’s t-test. (*P < 0.05, ***P < 0.001, ****P < 0.0001). c Induction of a spike-specific T cell population producing IFN-γ in the lungs of ΔEM virus- or mRNA-boosted hamsters that had previously received a prime mRNA vaccination as measured by use of an ELISpot assay (n = 6 females/group). Box plots show the median center line and 10/90 percentiles. Whiskers show min and max values. Statistical significance was determined by using a one-way ANOVA with Tukey’s multiple comparison test (*P < 0.05, **P < 0.01; ns, not significant). Exact P values: (a) Ancestral D614G spike: mRNA x2 (P = 0.0001), mRNA + ΔEM (P = 0.0018), ΔEM x2 (P = 0.0004); XBB spike: mRNA x2 (P = 0.0011), mRNA + ΔEM (P = 0.0003), ΔEM x2 (P = 0.0348). (b) lung: control vs. mRNA x2 (P = 0.0002), mRNA x2 vs. mRNA + ΔEM (P < 0.0001); NT: control vs. mRNA x2 (P = 0.0122), mRNA x2 vs. mRNA + ΔEM (P = 0.0129). (c) control vs. mRNA x2 (P = 0.2058), control vs. mRNA + ΔEM (P = 0.0011), mRNA x2 vs. mRNA + ΔEM (P = 0.0381). Source data are provided as a Source Data file.
We next examined protection against the Omicron XBB variant in hamsters vaccinated with the mRNA vaccine (1 μg/animal) and boosted with either the mRNA vaccine (n = 4 females) or the ΔEM virus (n = 4 females). Three days after challenge with the Omicron XBB variant, virus titers in the lung and nasal turbinate tissues of the hamsters boosted with the mRNA vaccine were reduced by about 10-fold compared to the control (Fig. 6b). In contrast, in the lung tissue of the ΔEM-boosted hamsters, there was no detectable infectious virus in three of four animals and more than a 10,000-fold reduction in the fourth animal (Fig. 6b), which is comparable to the protective efficacy observed in animals vaccinated twice with the ΔEM virus (Fig. 4a). In the nasal turbinate tissue of the mRNA-vaccinated, ΔEM-boosted hamsters, there was a 500- and 50-fold reduction in virus titers compared to the non-vaccinated and mRNA-boosted hamsters, respectively (Fig. 6b). These results demonstrate that although the levels of systemic ELISA antibody titers were similar between animals vaccinated with the mRNA vaccine and boosted with the mRNA vaccine or the ΔEM vaccine candidate virus, the levels of protection in both the lungs and nasal turbinate conferred by ΔEM were higher than those of the mRNA vaccine against both homologous and antigenically advanced challenge viruses.
To understand the difference in protective levels between boosting with the mRNA vaccine compared to the ΔEM virus (Fig. 6b), we examined the induction of S-reactive T cells in the lungs of the mRNA-boosted (n = 6 females) and ΔEM-boosted (n = 6 females) animals by using the ELISpot assay. The number of S-reactive IFN-γ-secreting T cells was higher in the ΔEM-boosted hamsters than in the control and mRNA-boosted hamsters (Fig. 6c). There was no significant difference in the number of S-reactive IFN-γ-secreting T cells between the control and mRNA-boosted hamsters. Collectively, these data provide a possible rationale for the protection conferred by booster vaccination with the ΔEM virus. Our data thus demonstrate the potential of the ΔEM vaccine candidate virus as a booster that can induce protective immunity against newer variants to which the animals have not been previously exposed.
Discussion
Intranasal LAVs can induce cellular immunity against exterior and interior viral proteins in addition to neutralizing antibody responses targeting the S protein at the site of infection, mimicking natural infection. Here, we developed one such LAV candidate, the SARS-CoV-2 ΔEM vaccine virus. To restrict productive viral infection and attempt to achieve a single round of replication for our vaccine candidate, the E and M proteins were deleted since these proteins are structural components of the virus necessary for assembly but are not needed for viral genome replication, which is facilitated by the viral nucleoprotein (N) and polymerase complex proteins. Previous groups have taken similar approaches to attenuate SARS-CoV-2. One group engineered a recombinant virus lacking the accessory proteins encoded by ORF3a, 6, 7, and 8 that can still replicate without compensating for the missing proteins22. In a similar approach, a trans-complementation system using SARS-CoV-2 virus lacking ORF3a and E has been established for safe use in BSL-2 containment37. Like other LAVs, vaccination with the ΔEM virus conferred promising respiratory protection in vaccinated animals (mice and hamsters), without requiring an adjuvant. The sCPD9 vaccine virus, which is a codon-pair deoptimized LAV demonstrated broad and complete protection against the Alpha, Beta, and Delta variants in the lung tissue of vaccinated animals18,19. Here, we demonstrated that our ΔEM vaccine candidate virus also offers broad and efficient protection against the Delta and Omicron XBB variants in the lungs of vaccinated female and male hamsters. Although protection in the nasal turbinate tissues was less pronounced compared to that in the lower respiratory tract, all but a few animals had a greater than 1,000-fold reduction in virus titers on day 3 after challenge, which lead to prompt virus elimination within 6 days of challenge.
Like other LAVs, there are potential concerns about recombination of the ΔEM vaccine candidate virus with circulating SARS-CoV-2 variants in the environment. Although this safety concern will be assessed in a future study, we would expect the ΔEM virus to be quickly eliminated. Indeed, infectious ΔEM virus was no longer detected in the respiratory tissues and brain tissue of transgenic hACE2 hamsters and mice three days after inoculation. Furthermore, since the ΔEM virus is a vaccine platform, updated versions of the vaccine virus can be generated by reverse genetics to replace the S protein with that of a circulating virus to accommodate the continued antigenic changes of SARS-CoV-2.
The lack of data on the effectiveness of ΔEM vaccination in preventing transmission of SARS-CoV-2 variants and the durability of the vaccine-induced immune response are limitations of this study that require further evaluation. Given the low titer of the ΔEM virus, we plan to investigate whether a lower dose can provide adequate protection and explore methods for generating ΔEM vaccine virus with higher virus titers.
The current mRNA-based vaccines induce minimal mucosal immunity in the respiratory tract2, which may allow breakthrough infections. LAVs are highly protective against intranasal virus challenge and transmission because they mimic the multi-faceted immunity elicited by natural infection38. Resident memory T cells in the airway and lung tissue play vital roles in preventing respiratory virus infections39. A previous study found that although the population of respiratory memory T cells induced by LAVs was not significantly increased even after boosting, IFN-γ expression levels of memory T cells in the lung tissue were relatively high compared to those in blood18. Here, ΔEM vaccination significantly induced IFN-γ-producing T cells in the lung tissue of vaccinated animals, which was comparable to those induced by a high dose of mRNA vaccine. These ΔEM-induced T cells respond not only to the S protein but also the N protein, and possibly to other viral proteins. Among the ΔEM-induced lung T cells, the S-reactive T cells included CD4+ and CD8+ T cells, but the N-reactive T cells were mainly CD8+ T cells. This suggests that ΔEM vaccination preferentially and strongly induces virus-reactive CD8+ T cells rather than CD4+ T cells. Disease severity and viral load in COVID-19 patients are strongly associated with T cell response40 and in mild cases, high levels of virus-reactive CD8+ T cells are induced31,41. Virus-reactive CD8+ T cells in the respiratory tract are thought to play an important role in viral clearance14,42. Moreover, memory CD8+ T cells in convalescents have the potential to broadly respond to variant S proteins43,44.
In addition to CD8+ T cells, memory CD4+ T cells have been seen in convalescents45,46. While CD8+ T cells preferentially recognize the N protein, CD4+ T cells can recognize multiple viral proteins47. In our experimental setting, N-reactive, IFN-γ/TNF-α-producing CD4+ T cells were not induced in the lung tissue of ΔEM-vaccinated animals. This might be due to the single vaccination, the low dose of the vaccine, or the missing ORFs. Boosting might increase the N-reactive CD4+ population as the prime-boost vaccination regimen improved the protective efficacy of the ΔEM vaccine. Further investigation is needed to better understand the immune responses induced by ΔEM vaccination, including an assessment of the cell-mediated immunity induced by ΔEM vaccination compared with that induced by natural infection which was not investigated in these studies.
In wild-type hamsters, one vaccination of the ΔEM virus did not elicit detectable neutralizing antibodies, and there was little to no reduction in virus titers in the respiratory tissues. However, this lack of detectable neutralizing antibodies did not blunt the immune response produced by a second vaccination, which did induce neutralizing antibodies and a greater reduction in virus titers in the tissues of the vaccinated animals challenged with a Delta variant. With the Omicron XBB variant, IgG antibodies in the sera of ΔEM virus-vaccinated animals could bind to the spike but did not exhibit neutralizing activity against this variant. This finding suggests that, in addition to T cell immunity, the ΔEM virus vaccine may confer another layer of antibody-mediated immunity, such as ADCC48,49, which will be investigated in future studies.
In summary, here, we generated a single cycle-replicating ΔEM vaccine candidate virus based on the ancestral isolate and demonstrated its protective effectiveness against the homologous ancestral virus and its variants in small animal models and the induction of resident memory T cells in the vaccinated animals. Our data show that the ΔEM vaccine candidate virus is a promising platform for the development of an effective COVID-19 LAV candidate.
Methods
Cells
Vero TMPRSS2 cells50 and HEK293T cells (laboratory stock) were cultured in Dulbecco’s modified Eagle medium (DMEM) supplemented with 10% fetal bovine serum (FBS), and antibiotics. G418 (1 mg/ml) in DMEM was added to Vero TMPRSS2.
HEK293T EM cells (HEK293T cells stably expressing SARS-CoV-2 E and M) were generated as follows: a cDNA fragment encoding the human codon-optimized SARS-CoV-2 E (#141273, Addgene, a gift from Fritz Roth51) and M (#141274, Addgene, a gift from Fritz Roth51) genes was cloned into the retroviral vector pMXs-IRES-puromycin (pMXs-IP) (RTV-014, Cell Biolabs). To generate the retrovirus, Plat-GP cells (RV-103, Cell Biolabs) were co-transfected with pMXs-IP vectors encoding E and M along with an expression vector for VSV G (PT3343-5, Clontech) by using Lipofectamine 2000 (Invitrogen). Two days later, the culture supernatants containing the retroviruses were collected and used to transduce HEK293T cells. Stable cells were selected with 2 μg/ml puromycin (InvivoGen).
Vero TMPRSS2/EM cells (Vero TMPRSS2 cells stably expressing SARS-CoV-2 E and M) were generated in a similar manner as HEK293T EM cells, with slight modification. Vero TMPRSS2 cells were first transduced with a retrovirus carrying the E gene and selected with 7 μg/ml puromycin. E protein expression was confirmed by use of an immunofluorescent assay using an anti-SARS-CoV-2 E antibody (SARS-COV2-E-101AP, Thermo Fisher Scientific). Single clone cells with high E protein expression levels (Vero TMPRSS2/E) were obtained by cell cloning. Then, the Vero TMPRSS2/E cells were transfected with pIREShyg3 vector (#631620, Clontech) expressing SARS-CoV-2 M (#141274, Addgene, a gift from Fritz Roth51) and selected with 400 μg/ml Hygromycin B (InvivoGen). Single clone cells with high M protein expression levels were obtained by cell cloning. Among the selected cell clones, the Vero TMPRSS2/EM cell line that showed CPE after infection and supported ΔEM virus replication most efficiently was used for this study (Supplementary Table 1). All cells were incubated at 37 °C and 5% CO2 and were routinely tested for mycoplasma by using a PCR-based assay.
Viruses
An ancestral SARS-CoV-2 isolate (SARS-CoV-2/UT-HP095-1N/Human/2020/Tokyo)52, Delta variant (hCoV-19/USA/WI-UW-5250/2021 [B.1.617.2; UW-5250])53, and Omicron XBB subvariant (hCoV-19/USA/NY-MSHSPSP-PV73997/2022)54 were used as authentic challenge viruses in this study. All viruses were sequenced on the Illumina NGS platform to confirm the stability of substitutions and the absence of adventitious alterations. All virus experiments were performed under biosafety level 3 containment with an approved biosafety protocol (B00000263).
Generation of recombinant SARS-CoV-2 wild-type and ΔEM viruses
Fragments for circular polymerase extension reaction (CPER)
Six fragments with overlapping 20 nucleotide sequences for the CPER reaction28,29 were amplified from the full-length cDNA of SARS-CoV-2 (Wuhan-Hu-1 isolate) (NC_045512) cloned into the pBeloBAC11 vector by using PrimeSTAR GXL DNA polymerase (TaKaRa Bio) and the corresponding primer pairs with overlapping sequences at the 5’ end, which enables sequence-specific assembly.
Fragment F6 for the ΔEM virus lacks both the entire E and M ORF regions (nucleotides 26,245 to 27,191) including the intergenic region between them (Supplementary Fig. 6a). The linker fragment used to connect fragments F1 and F6 contains a polyA tail (30 adenines) and the hepatitis delta virus ribozyme for generating the authentic 3’ end of the viral RNA, a simian virus 40 polyA signal for efficient termination of transcription, and a spacer sequence followed by the cytomegalovirus promoter for viral RNA transcription. Each PCR product was purified with a QIAquick Gel Extraction Kit (Qiagen) after separation by agarose gel electrophoresis, and then used for the CPER to generate a circular infectious clone (Supplementary Fig. 6b).
CPER reaction
To generate an infectious cDNA clone, six fragments and a linker fragment were mixed at 0.1 pmol each in a 50 µl reaction volume and used for the following PCR reaction with PrimeSTAR GXL DNA polymerase: initial denaturation at 98 °C for 1 min; 15 cycles of denaturation at 98 °C for 10 s, annealing at 55 °C for 20 s, extension at 68 °C for 15 min, and a final extension at 68 °C for 15 min.
CPER transfection and virus rescue
The CPER product (30 µl of a 50-µl reaction volume) was directly transfected into wild-type HEK293T or HEK293T EM cells seeded in a 6-well plate (8.0 × 105 cells/well) by using TransIT-LT1 transfection reagent (Mirus Bio). The next day, the culture supernatant was replaced with fresh culture medium containing 5% FBS. On the fourth day after transfection, the supernatant was collected, and 1 ml of it was added to a T-25 flask of confluent wild-type Vero TMPRSS2 or Vero TMPRSS2/EM cells. Supernatants containing viruses were harvested when cytopathic effect appeared (4–7 days post-infection). To obtain high-titer virus stocks, the supernatant was passaged in fresh cells. The whole genome sequences of the recombinant parental virus and vaccine virus candidate were confirmed to be identical to that of the Wuhan-Hu-1 isolate by next-generation sequencing on the Illumina MiSeq platform55.
3, 3’-Diaminobenzidine (DAB) staining
Wild-type Vero TMPRSS2 cells and Vero TMPRSS2/EM cells were infected with the ΔEM virus at a multiplicity of infection (MOI) of 0.01. Three days later, the cells were fixed with 4% paraformaldehyde and permeabilized with 0.05% Triton X-100. After being blocked with 1% FBS in PBS, the cells were incubated with a rabbit anti-SARS-CoV-2 nucleocapsid antibody (1:200, Rockland, #200-401-MS4) followed by an HRP-conjugated goat anti-rabbit IgG (1:3000, Life Technologies, G21234). The infected cells were then washed with PBS and visualized by using DAB (MP Biomedicals). The reaction was stopped by rinsing with water.
Animal experiments and approval
Animal studies were performed under a protocol approved by the Institutional Animal Care and Use Committee at the University of Wisconsin, Madison (protocol number V006426). Virus infections were performed under isoflurane, and all efforts were made to minimize pain. In vivo studies were not blinded. Group sizes were determined based on prior virus challenge studies, and no sample-size calculations were performed to determine the power of each study.
Animal studies analyzing T-cell responses were carried out in accordance with the recommendations in the Guide for the Care and Use of Laboratory Animals of the National Institutes of Health. The protocol (P19-72) was approved by the Institutional Animal Care and Use Committee at the Animal Experiment Committee of the Institute of Medical Science, the University of Tokyo.
Mouse vaccination and challenge experiments
Heterozygous K18-hACE2 C57BL/6 J mice (strain 2B6.Cg-Tg[K18-ACE2]2Prlmn/J) were obtained from The Jackson Laboratory and Charles River Laboratories. Animals were housed in groups under controlled conditions (22 ± 2 °C temperature, 55–65% humidity) with a 12 h light/dark cycle and fed standard chow diets. In the first set of challenge experiments, K18-hACE2 mice (females, 5–month-old) were anesthetized with isoflurane and inoculated intranasally with 1 × 104 pfu of the recombinant parental virus or ΔEM virus in a total volume of 50 μl of DMEM. In the second set of experiments, K18-hACE2 mice (females, 5–month-old) were anesthetized with isoflurane and vaccinated intranasally with 1 x 104 pfu of the ΔEM virus in a total volume of 50 μl of DMEM. Boost vaccination was performed four weeks after the prime vaccination. Challenge was performed four weeks after the last vaccination. Mice were anesthetized with isoflurane and inoculated intranasally with 1 x 105 pfu of an ancestral SARS-CoV-2 isolate (50 μl per mouse). Animal health was monitored daily for changes in body weight and signs of illness including inability to remain upright; no other scoring was performed. Three days after challenge, the mice were humanely sacrificed by isoflurane overdose and lung tissue was collected to measure virus titers, which were determined by performing the standard plaque assays on Vero TMRPSS2 cells or Vero TMRPSS2/EM cells56.
Hamster vaccination and challenge experiments
Syrian golden hamsters were obtained from Charles River Laboratories, Envigo. K18-hACE2 homozygous transgenic hamsters57,58 were from an established colony at UW-Madison. Animals were housed in groups and fed standard chow diets. In the first set of challenge experiments, hACE2 hamsters (females, 5–6-weeks-old) were anesthetized with isoflurane and inoculated intranasally with 4 x 104 pfu of the ancestral virus or ΔEM virus in a total volume of 100 μl of DMEM. In the second set of experiments, wild-type Syrian hamsters (females and males, 5–6-weeks-old) were anesthetized with isoflurane and vaccinated intranasally with 2 × 104 pfu of the ΔEM virus in a total volume of 100 μl of DMEM. Boost vaccination was performed four weeks after the prime vaccination. Challenge was performed six weeks after the last vaccination. Hamsters were anesthetized with isoflurane and inoculated intranasally with a Delta variant (8 × 104 pfu; determined on Vero ACE2/TMPRSS2 cells) or Omicron XBB (1 x 105 pfu; determined on Vero ACE2/TMPRSS2 cells) in a total volume of 100 μl of DMEM. In the third set of experiments, wild-type Syrian hamsters (females and males, 5–6-weeks-old) were anesthetized with isoflurane and vaccinated either intramuscularly with 10 μg of an mRNA vaccine (BNT162b2) or intranasally with 2 × 104 pfu of the ΔEM vaccine candidate virus. Boost vaccination with the ΔEM vaccine candidate virus was performed four weeks after the prime vaccination. Challenge was performed six weeks after the last vaccination. Hamsters were anesthetized with isoflurane and inoculated intranasally with Omicron XBB (105 pfu; determined on Vero ACE2/TMPRSS2 cells). Animal health was monitored daily. Three days after challenge, the hamsters were humanely sacrificed by isoflurane overdose. Nasal turbinates and lungs were collected to measure virus titers on Vero ACE2/TMPRSS2 cells, which were determined by performing the standard plaque assays56. Blood was collected from the sublingual vein under isoflurane anesthesia the day before boost vaccination and challenge, and the serum was used in the ELISA assay.
IgG ELISA assay
Ninety-six-well plates were coated overnight at 4 °C with 50 μL of recombinant ancestral (Hexa pro-His, produced in-house59) and Omicron XBB (Sino Biological, 40589-V08H40) spike antigens at 2 μg/ml in PBS. After being blocked with PBS containing 0.1% Tween 20 (PBS-T) and 3% non-fat milk, the plates were incubated in duplicate with heat-inactivated serum diluted in PBS-T with 1% milk. A hamster IgG secondary antibody conjugated with horseradish peroxidase (1:5000, Thermo Fisher Scientific), and mouse IgG1 (115-035-205), IgG2b (115-035-207), and IgG3 (115-035-209) subclass secondary antibodies conjugated with horseradish peroxidase (1:2000, Jackson ImmunoResearch) were used for detection. Plates were developed with SigmaFast o-phenylenediamine dihydrochloride solution (Sigma), and the reaction was stopped by adding 3 M hydrochloric acid. The absorbance was measured at a wavelength of 490 nm (OD490). Background absorbance measurements from pooled naïve hamster sera were subtracted from measurements from sera collected after immunization for each dilution. IgG antibody endpoint titers were defined as the highest serum dilution with an OD490 cut-off value of 0.15 which has been used as a cut-off value in our other vaccine studies for consistency54,59,60.
IgA ELISA assay
Ninety-six-well plates coated with the recombinant Wuhan-Hu-1 spike protein (Hexa pro-His, produced in-house59) were prepared as described above. After blocking, the plates were incubated in duplicate with heat-inactivated nasal wash or BALF samples diluted at 1:1 with PBS. IgA was detected by using an IgA secretory component ELISA kit (ABIN6962949, Antibodies-online). The absorbance was measured at a wavelength of 450 nm (OD450).
Intracellular cytokine staining of lung cells from ΔEM virus-vaccinated mice
Hemizygous K18-hACE2 C57BL/6 J mice (strain B6.Cg-Tg[K18-ACE2]2Prlmn/J) were obtained from the Jackson Laboratory. For the immunization, eight-week-old female K18-hACE2 mice were anesthetized with isoflurane and intranasally inoculated with VP-SFM medium (control) or the ΔEM virus (104 pfu). For comparison, mice were intramuscularly injected with mRNA vaccine (0.1 or 1.0 μg). Two weeks after the immunization, mice were euthanized, and their lungs were collected after perfusion with PBS. To harvest single cells, lungs were minced to yield 1–2 mm pieces and incubated with RPMI 1640 containing collagenase D for at least 30 min at 37 °C. Then, the cells were purified in 33% Percoll by centrifugation. The purified cells were resuspended in RPMI 1640 with 10% FCS at a concentration of 4 × 105 cells/100 μl/well. These cells were then stimulated for 6 h with or without 1 μg/mL SARS-CoV-2 peptide designed from the S protein (#LB01792) or N protein (#LB01786) of the ancestral strain (peptides and elephants) in the presence of GolgiStop (BD Biosciences) in a U-bottom plate at 37 °C under 5% CO2 in RPMI 1640 with 10% heat-inactivated FCS. After incubation, the cells were incubated with Live/dead fixable aqua (Thermo Fisher Scientific) (1:200), anti-CD16/32 (93) Ab (1:200), and antibodies specific to CD45 (30-F11) (1:100), CD4 (RM4-5) (1:100), and CD8a (53–6.7) (1:100). Following fixation and permeabilization with Cytofix/Cytoperm from the Fixation/Permeabilization Solution Kit (BD Biosciences), the cells were stained with antibodies specific to TNF-α (MP6-XT22) (1:100) and IFN-γ (XMG1.2) (1:50) (Biolegend). Data were acquired with CytoFLEX S (Beckman Coulter Inc.) and data analysis was performed using FlowJo software (Supplementary Fig. 7).
Enzyme-linked immunospot (ELISpot) assay
ELISpot was performed using the mouse IFN-γ ELISpot Kit (55188, BD Biosciences) and the ELISpot Flex: Hamster IFN‑γ kit (3102-2 A, MABTECH) according to the manufacturers’ instructions. Briefly, for mice, cells from the lungs were isolated as described above and resuspended in RPMI 1640 with 10% FCS at a concentration of 2 × 105 cells/100 μl/well for stimulation with S peptide or 4 × 105 cells/100 μl/well for stimulation with N peptide. For hamsters, wild-type Syrian hamsters (females, 5–6-weeks-old) were anesthetized with isoflurane and vaccinated intramuscularly with 10 μg of mRNA vaccine (BNT162b2). Four weeks after this vaccination, the hamsters were boost vaccinated either intramuscularly with the mRNA vaccine or intranasally with 2 × 104 pfu of the ΔEM vaccine candidate virus. Boost vaccination with the ΔEM virus was performed four weeks after the prime vaccination. Six weeks after the last vaccination, lungs were minced to yield 1–2 mm pieces and incubated with RPMI 1640 containing collagenase D for at least 30 min at 37 °C. Then, the cells were purified in 33% Percoll by centrifugation. The purified cells were resuspended in RPMI 1640 with 10% FCS at a concentration of 2 x 105 cells/100 μl/well for stimulation with S peptide. Then, the cells were incubated with SARS-CoV-2 peptides at a final concentration of 1 μg/mL in 96-well ELISpot plates at 37 °C. Sixteen hours later, the cells and supernatants were removed, and the ELISpot membranes were stained for IFN-γ. The spot numbers were quantified by using an ImmunoSpot S6 Analyzer, ImmunoCapture software, and BioSpot software (Cellular Technology).
Histopathology
Left lung lobes of the hamsters were fixed in 10% neutral buffered formalin and processed for paraffin embedding. The paraffin blocks were sectioned and placed on glass slides, followed by hematoxylin and eosin (H&E) staining for histopathological examination. Whole slide images were created by digital scanning on the Aperio AT2 scanner (Leica Biosystems, Inc.) and were evaluated by a semi-quantitative analysis using the following four parameters: perivascular inflammation, bronchial/bronchiolar epithelial degeneration or necrosis, bronchial/bronchiolar inflammation, and alveolar inflammation. A 5-point scoring system was utilized [0-within normal limits, 1-mild (<25%), 2-moderate (25%–50%), 3-marked (50%–75%), or 4-severe (>75%)]. A total pathology score was calculated for each hamster by adding the individual histopathological feature scores.
Focus reduction neutralization test (FRNT)
A 2-fold dilution series of serum starting at a dilution of 1:20 was mixed with approximately 800 focus-forming units of an ancestral virus, a Delta variant, or an Omicron XBB variant/well and incubated for 1 h at 37 °C. The antibody-virus mixture was inoculated onto Vero E6/TMPRSS2 cells in 96-well plates and incubated for 1 h at 37 °C. An equal volume of methylcellulose solution was added to each well. The cells were incubated for 16 h at 37 °C and then fixed with formalin. After the formalin was removed, the cells were immunostained with a mouse monoclonal antibody against SARS-CoV-1/2 nucleoprotein (clone 1C7C7, Sigma-Aldrich, catalog #MA5-29982, 1:10,000 dilution), followed by a horseradish peroxidase-labeled goat anti-mouse immunoglobulin (ThermoFisher, catalog #31430, 1:2000 dilution). The infected cells were stained with TrueBlue Substrate (SeraCare Life Sciences) and then washed with distilled water. After cell drying, the focus numbers were quantified by using an ImmunoSpot S6 Analyzer, ImmunoCapture software, and BioSpot software (Cellular Technology). The FRNT50 value was then calculated using a four-parameter nonlinear regression in Graphpad Prism.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data underlying graphs generated in this study are provided in the Source Data file. There are no restrictions in obtaining access to primary data. All virus sequences in this study were previously deposited and are available from the GISAID database (https://gisaid.org/), Genbank database (https://www.ncbi.nlm.nih.gov/genbank/), or UniprotKB database (https://www.uniprot.org/).
References
Wherry, E. J. & Barouch, D. H. T cell immunity to COVID-19 vaccines. Science 377, 821–822 (2022).
Tang, J. et al. Respiratory mucosal immunity against SARS-CoV-2 after mRNA vaccination. Sci. Immunol. 7, eadd4853 (2022).
Rice, A. et al. Heterologous saRNA Prime, DNA Dual-Antigen Boost SARS-CoV-2 Vaccination Elicits Robust Cellular Immunogenicity and Cross-Variant Neutralizing Antibodies. Front Immunol. 13, 910136 (2022).
McCafferty, S. et al. A dual-antigen self-amplifying RNA SARS-CoV-2 vaccine induces potent humoral and cellular immune responses and protects against SARS-CoV-2 variants through T cell-mediated immunity. Mol. Ther. 30, 2968–2983 (2022).
Afkhami, S. et al. Respiratory mucosal delivery of next-generation COVID-19 vaccine provides robust protection against both ancestral and variant strains of SARS-CoV-2. Cell 185, 896–915.e819 (2022).
Hajnik, R. L. et al. Dual spike and nucleocapsid mRNA vaccination confer protection against SARS-CoV-2 Omicron and Delta variants in preclinical models. Sci. Transl. Med. 14, eabq1945 (2022).
Thakur, S. et al. SARS-CoV-2 Mutations and Their Impact on Diagnostics, Therapeutics and Vaccines. Front. Med. (Lausanne) 9, 815389 (2022).
Rak, A., Isakova-Sivak, I. & Rudenko, L. Overview of Nucleocapsid-Targeting Vaccines against COVID-19. Vaccines (Basel) 11, 1810 (2023).
Tai, W. et al. An mRNA-based T-cell-inducing antigen strengthens COVID-19 vaccine against SARS-CoV-2 variants. Nat. Commun. 14, 2962 (2023).
Tarke, A. et al. SARS-CoV-2 vaccination induces immunological T cell memory able to cross-recognize variants from Alpha to Omicron. Cell 185, 847–859 e811 (2022).
Liu, J. et al. Vaccines elicit highly conserved cellular immunity to SARS-CoV-2 Omicron. Nature 603, 493–496 (2022).
Grifoni, A. et al. Targets of T Cell Responses to SARS-CoV-2 Coronavirus in Humans with COVID-19 Disease and Unexposed Individuals. Cell 181, 1489–1501.e1415 (2020).
Mao T. et al. Unadjuvanted intranasal spike vaccine booster elicits robust protective mucosal immunity against sarbecoviruses. bioRxiv, (2022).
Ishii, H. et al. Neutralizing-antibody-independent SARS-CoV-2 control correlated with intranasal-vaccine-induced CD8(+) T cell responses. Cell Rep. Med. 3, 100520 (2022).
Dangi, T., Class, J., Palacio, N., Richner, J. M. & Penaloza MacMaster, P. Combining spike- and nucleocapsid-based vaccines improves distal control of SARS-CoV-2. Cell Rep. 36, 109664 (2021).
Gabitzsch, E. et al. Dual-Antigen COVID-19 Vaccine Subcutaneous Prime Delivery With Oral Boosts Protects NHP Against SARS-CoV-2 Challenge. Front. Immunol. 12, 729837 (2021).
Chiuppesi, F. et al. Development of a multi-antigenic SARS-CoV-2 vaccine candidate using a synthetic poxvirus platform. Nat. Commun. 11, 6121 (2020).
Nouailles, G. et al. Live-attenuated vaccine sCPD9 elicits superior mucosal and systemic immunity to SARS-CoV-2 variants in hamsters. Nat. Microbiol. 8, 860–874 (2023).
Trimpert, J. et al. Live attenuated virus vaccine protects against SARS-CoV-2 variants of concern B.1.1.7 (Alpha) and B.1.351 (Beta). Sci. Adv. 7, eabk0172 (2021).
Wang, Y. et al. Scalable live-attenuated SARS-CoV-2 vaccine candidate demonstrates preclinical safety and efficacy. Proc. Natl. Acad. Sci. USA 118, e2102775118 (2021).
Trimpert, J. et al. Development of safe and highly protective live-attenuated SARS-CoV-2 vaccine candidates by genome recoding. Cell Rep. 36, 109493 (2021).
Liu, Y. et al. A live-attenuated SARS-CoV-2 vaccine candidate with accessory protein deletions. Nat. Commun. 13, 4337 (2022).
Seo, S. H. & Jang, Y. Cold-Adapted Live Attenuated SARS-Cov-2 Vaccine Completely Protects Human ACE2 Transgenic Mice from SARS-Cov-2 Infection. Vaccines (Basel) 8, 584 (2020).
Xu, J. et al. The Cold-Adapted, Temperature-Sensitive SARS-CoV-2 Strain TS11 Is Attenuated in Syrian Hamsters and a Candidate Attenuated Vaccine. Viruses 15, 95 (2022).
Faizuloev, E. et al. Cold-adapted SARS-CoV-2 variants with different temperature sensitivity exhibit an attenuated phenotype and confer protective immunity. Vaccine 41, 892–902 (2023).
Becerra, X. & Jha, A. Project NextGen - Defeating SARS-CoV-2 and Preparing for the Next Pandemic. N. Engl. J. Med. 389, 773–775 (2023).
Pilapitiya, D., Wheatley, A. K. & Tan, H. X. Mucosal vaccines for SARS-CoV-2: triumph of hope over experience. EBioMedicine 92, 104585 (2023).
Amarilla, A. A. et al. A versatile reverse genetics platform for SARS-CoV-2 and other positive-strand RNA viruses. Nat. Commun. 12, 3431 (2021).
Torii, S. et al. Establishment of a reverse genetics system for SARS-CoV-2 using circular polymerase extension reaction. Cell Rep. 35, 109014 (2021).
Tun, Z. H., Htike, N. T. T., Kyi-Tha-Thu, C. & Lee, W. H. Employing T-Cell Memory to Effectively Target SARS-CoV-2. Pathogens 12, 301 (2023).
Peng, Y. et al. Broad and strong memory CD4(+) and CD8(+) T cells induced by SARS-CoV-2 in UK convalescent individuals following COVID-19. Nat. Immunol. 21, 1336–1345 (2020).
Payne, R. P. et al. Immunogenicity of standard and extended dosing intervals of BNT162b2 mRNA vaccine. Cell 184, 5699–5714 e5611 (2021).
Mateus, J. et al. Low-dose mRNA-1273 COVID-19 vaccine generates durable memory enhanced by cross-reactive T cells. Science 374, eabj9853 (2021).
Singanayagam, A. et al. Community transmission and viral load kinetics of the SARS-CoV-2 delta (B.1.617.2) variant in vaccinated and unvaccinated individuals in the UK: a prospective, longitudinal, cohort study. Lancet Infect. Dis. 22, 183–195 (2022).
Wang, Q. et al. Alarming antibody evasion properties of rising SARS-CoV-2 BQ and XBB subvariants. Cell 186, 279–286.e278 (2023).
Wang, W. et al. Human and hamster sera correlate well in identifying antigenic drift among SARS-CoV-2 variants, including JN.1. J. Virol. 98, e0094824 (2024).
Zhang, X. et al. A trans-complementation system for SARS-CoV-2 recapitulates authentic viral replication without virulence. Cell 184, 2229–2238.e2213 (2021).
Alu, A. et al. Intranasal COVID-19 vaccines: From bench to bed. EBioMedicine 76, 103841 (2022).
Zhang, M., Li, N., He, Y., Shi, T. & Jie, Z. Pulmonary resident memory T cells in respiratory virus infection and their inspiration on therapeutic strategies. Front. Immunol. 13, 943331 (2022).
Tan, A. T. et al. Early induction of functional SARS-CoV-2-specific T cells associates with rapid viral clearance and mild disease in COVID-19 patients. Cell Rep. 34, 108728 (2021).
Schulien, I. et al. Characterization of pre-existing and induced SARS-CoV-2-specific CD8(+) T cells. Nat. Med. 27, 78–85 (2021).
Channappanavar, R., Fett, C., Zhao, J., Meyerholz, D. K. & Perlman, S. Virus-specific memory CD8 T cells provide substantial protection from lethal severe acute respiratory syndrome coronavirus infection. J. Virol. 88, 11034–11044 (2014).
Redd, A. D. et al. CD8+ T-Cell Responses in COVID-19 Convalescent Individuals Target Conserved Epitopes From Multiple Prominent SARS-CoV-2 Circulating Variants. Open Forum Infect. Dis. 8, ofab143 (2021).
Tarke, A. et al. Impact of SARS-CoV-2 variants on the total CD4(+) and CD8(+) T cell reactivity in infected or vaccinated individuals. Cell Rep. Med. 2, 100355 (2021).
Sekine, T. et al. Robust T Cell Immunity in Convalescent Individuals with Asymptomatic or Mild COVID-19. Cell 183, 158–168.e114 (2020).
Zuo, J. et al. Robust SARS-CoV-2-specific T cell immunity is maintained at 6 months following primary infection. Nat. Immunol. 22, 620–626 (2021).
Cohen, K. W. et al. Longitudinal analysis shows durable and broad immune memory after SARS-CoV-2 infection with persisting antibody responses and memory B and T cells. Cell Rep. Med. 2, 100354 (2021).
Izadi, A. & Nordenfelt, P. Protective non-neutralizing SARS-CoV-2 monoclonal antibodies. Trends Immunol. 45, 609–624 (2024).
Zedan, H. T. et al. SARS-CoV-2 infection triggers more potent antibody-dependent cellular cytotoxicity (ADCC) responses than mRNA-, vector-, and inactivated virus-based COVID-19 vaccines. J. Med. Virol. 96, e29527 (2024).
Imai, M. et al. Syrian hamsters as a small animal model for SARS-CoV-2 infection and countermeasure development. Proc. Natl. Acad. Sci. USA 117, 16587–16595 (2020).
Kim D.K., et al. A Comprehensive, Flexible Collection of SARS-CoV-2 Coding Regions. G3 (Bethesda) 10, 3399–3402 (2020).
Imai, M. et al. Characterization of a new SARS-CoV-2 variant that emerged in Brazil. Proc. Natl Acad. Sci. USA 118, e210653511 (2021).
Halfmann, P. J. et al. Long-term, infection-acquired immunity against the SARS-CoV-2 Delta variant in a hamster model. Cell Rep. 38, 110394 (2022).
Halfmann, P. J. et al. Broad protection against clade 1 sarbecoviruses after a single immunization with cocktail spike-protein-nanoparticle vaccine. Nat. Commun. 15, 1284 (2024).
Uraki, R. et al. Characterization of SARS-CoV-2 Omicron BA.4 and BA.5 isolates in rodents. Nature 612, 540–545 (2022).
Halfmann, P. J. et al. SARS-CoV-2 Interference of Influenza Virus Replication in Syrian Hamsters. J. Infect. Dis. 225, 282–286 (2022).
Gibson, S. A. et al. Differences in Susceptibility to SARS-CoV-2 Infection Among Transgenic hACE2-Hamster Founder Lines. Viruses-Basel 16, 1625 (2024).
Gilliland, T. et al. Transchromosomic bovine-derived anti-SARS-CoV-2 polyclonal human antibodies protects hACE2 transgenic hamsters against multiple variants. Iscience 26, 107764 (2023).
Chiba, S. et al. Multivalent nanoparticle-based vaccines protect hamsters against SARS-CoV-2 after a single immunization. Commun. Biol. 4, 597 (2021).
Halfmann, P. J. et al. Multivalent S2-based vaccines provide broad protection against SARS-CoV-2 variants of concern and pangolin coronaviruses. EBioMedicine 86, 104341 (2022).
Acknowledgements
We thank Addgene for providing the plasmids used in this study, and Susan Watson for scientific editing. Y.K. acknowledges support from the National Institute of Allergy and Infectious Diseases of the National Institutes of Health under Award Number P01AI165077, the Japan Program for Infectious Diseases Research and Infrastructure (JP23wm0125002), and the Japan Initiative for World-leading Vaccine Research and Development Centers (JP233fa627001), and Program on R&D of new generation vaccine including new modality application (JP233fa827005) from the Japan Agency for Medical Research and Development.
Author information
Authors and Affiliations
Contributions
Conceptualization: M.K., P.H., Y.K. Methodology: M.K., P.H., R.U. Investigation: M.K., P.H., R.U., K.T., T.A.A., S.S., R.D., L.B. Visualization: M.K., P.H., R.U. Funding acquisition: Y.K. Project administration: P.H., Y.K. Supervision: P.H., Y.K. Writing – original draft: M.K. Writing – review & editing: P.H., R.U., K.T., T.A.A., S.S., R.D., L.B., S.Y., Y.K.
Corresponding authors
Ethics declarations
Competing interests
Y.K. has received unrelated funding support from Daiichi Sankyo Pharmaceutical, Toyama Chemical, Tauns Laboratories, Inc., Shionogi & Co. LTD, Otsuka Pharmaceutical, KM Biologics, Kyoritsu Seiyaku, Shinya Corporation, and Fuji Rebio. The other authors declare that they have no competing interests.
Peer review
Peer review information
Nature Communications thanks Kai Dallmeier, and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kuroda, M., Halfmann, P.J., Uraki, R. et al. SARS-CoV-2 virus lacking the envelope and membrane open-reading frames as a vaccine platform. Nat Commun 16, 4453 (2025). https://doi.org/10.1038/s41467-025-59533-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-59533-4
This article is cited by
-
SARS-CoV-2 variants: biology, pathogenicity, immunity and control
Nature Reviews Microbiology (2025)