Abstract
Antibodies targeting the highly conserved prehairpin intermediate (PHI) of class I viral membrane-fusion proteins are generally weakly neutralizing and are not considered viable therapeutic agents. We previously demonstrated that antibodies targeting the gp41 N-heptad repeat (NHR), which is transiently exposed in the HIV-1 PHI, exhibit enhanced broad neutralization in cells expressing the Fc receptor, FcγRI. To enhance neutralization in cells lacking FcγRI, we here develop a bispecific antibody (bsAb) by fusing an NHR-targeting antibody to an antibody against CD4, the HIV-1 receptor on T cells. The bsAb provides a 5000-fold neutralization enhancement and shows unprecedented neutralization breadth compared to existing broadly neutralizing antibodies. Importantly, the bsAb reduces viral load in HIV-1-infected humanized male mice, and viral envelope sequencing under bsAb pressure revealed an NHR mutation that potentially impairs viral fitness. These findings validate the NHR as a potential HIV-1 therapeutic target, setting the stage for a new class of broadly neutralizing antibodies.
Similar content being viewed by others
Introduction
HIV-1 infection occurs through the fusion of viral and host membranes, a process initiated upon binding of viral glycoprotein envelope (Env) to host receptor CD4 and to coreceptors CCR5 or CXCR41,2. HIV-1 Env is a trimer composed of gp120 and gp41 subunits, with the trimer undergoing a series of conformational changes during the fusion process from the native, pre-fusion state to a transient prehairpin intermediate (PHI), followed by formation of a trimer-of-hairpins conformation (Fig. 1A)3,4,5,6.
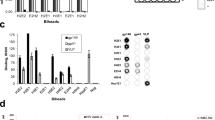
A Overview of HIV-1 infection. Infection by HIV-1 is initiated by binding of envelope (Env) protein, a trimer composed of gp120 and gp41 subunits, to CD4 on the surface CD4 + T cells. Subsequently, the Env trimer binds to host cell receptors including CCR5 and CXCR4, and triggers membrane fusion. During the fusion process, Env undergoes a conformational change, forming a prehairpin intermediate (PHI) followed by formation of a trimer-of-hairpins. gp41 is composed of N-heptad repeat (NHR), C-heptad repeat (CHR), and membrane-proximal external region (MPER). The N-heptad repeat (NHR) becomes transiently exposed in the PHI, which is normally not readily accessible in the native pre-fusion state. B Domain structure of HIV-1 Env protein, which is cleaved into gp120 and gp41. Numbers on top of each domain show sequence identity (percentage of amino acids that are identical). NHR sequence conservation in the logo plot (inset) is based on Los Alamos 2021 HIV database (>8000 HIV-1 Env proteins). The epitope of the anti-NHR mAb D5 is highlighted in orange. C A table showing sequence identity of NHR and the epitope of D5 among HIV-1 sequences. D Sequence conservation of NHR from 65 clustered groups in the Los Alamos 2021 HIV database (Table S1) is overlaid on the structure of gp41 inner-core mimetic 5-helix (PDB: 2CMR). The epitopes of D5 are shown in spheres, colored based on the extent of conservation indicated by the heatmap scale.
The N-heptad repeat (NHR) of gp41 is transiently exposed in the PHI (Fig. 1A, B) and is the site for binding by the FDA-approved peptide drug enfuvirtide (T-20). Enfuvirtide binds to a conserved, hydrophobic groove in the NHR7,8, while the NHR-binding antibody, D59, binds to a hydrophobic pocket immediately C-terminal to the enfuvirtide-binding site. The NHR is > 90% conserved among over 8000 Env sequences in the Los Alamos 2021 HIV database comprising most of the published Env sequences (Fig. 1B–D, Table S1). The hydrophobic pocket of NHR consists of seven discontinuous residues (L568, V570, W571, K574, Q575, Q577, and R579) that are 97–99% conserved, likely due to their importance in the fusion process9,10. Consistent with high conservation of the NHR, the mRNA sequences encoding these residues are found in stem V of the Rev response element, which is essential for viral replication11,12.
Viral neutralization by NHR-targeting antibodies has been modest and limited to tier-1 and tier-2 viruses (defined by their neutralization-sensitivity where tier-1 viruses are the most neutralization-sensitive)9,13,14,15. Recently, we showed that neutralization by NHR-targeting antibody D5 could be enhanced >5000-fold in cells expressing human FcγRI16. As well, the neutralization potency of an optimized D5 variant, D5_AR17 was also enhanced in FcγRI-expressing cells. This enhancement was independent of neutralization tiers, demonstrating broad neutralization across all tiers, including tier-3, the most difficult to neutralize18. We proposed that the enhancement of neutralization was due to binding of the Fc region of NHR-targeting antibodies to FcγRI, which prepositions the antibodies on host cells, thereby increasing their local concentration for binding to the transiently exposed NHR at the PHI. Although FcγRI is not expressed on CD4 + T cells, the major target cells for HIV-1 infection, it is expressed on macrophages and dendritic cells that are implicated in the early establishment of HIV-1 infection19,20.
Here, we investigate whether prepositioning at the site of viral fusion on T cells by a bispecific antibody (bsAb) can increase the local concentration of potency-optimized D5_AR to enhance its ability to prevent viral entry. Our approach, termed bispecific antibody-mediated prepositioning, uses an engineered bsAb to preposition D5_AR on CD4+ cells, achieving a 5000-fold enhancement in the neutralization potency. This results in the effective neutralization of a wide panel of pseudotyped and replication-competent viruses, including tier-3 strains—the most difficult strains to neutralize with patient antisera21. Using pseudovirus-based deep mutational scanning (DMS), we identified mutations in the NHR that decrease the neutralization potency of the bsAb, confirming that neutralization is dependent on bsAb binding to the NHR. We evaluated the in vivo efficacy of bsAb in HIV-1-infected humanized mice, revealing in vivo neutralization by the NHR-targeting antibody. Sequencing of viral Env under selective pressure in vivo suggests that a specific individual mutation in the NHR is deleterious to viral fitness. Overall, our findings validate the NHR as a druggable antibody target, introducing a new class of broadly neutralizing antibodies (bnAbs) against HIV-1.
Results
Generation and characterization of a bsAb for prepositioning on host cells
To develop a bsAb to preposition an NHR-targeting antibody at sites of viral infection, we generated hybrids between D5_AR and ibalizumab (iMab), an antibody against cellular CD4 (Fig. 2A, antibody sequences available in Table S2) that has previously been used in bispecific antibodies22,23,24,25,26. While iMab has previously been used in bsAbs targeting epitopes exposed in the prefusion state, our study uniquely employs iMab together with D5_AR, which specifically binds to the highly conserved NHR that is transiently and exclusively exposed in the PHI. iMab does not block the binding of gp120 to CD4, so it does not exert selective pressure against CD4 binding27. In order to promote correct light chain pairing of the two antibodies and avoid light chain pairing with the heavy chain of the other arm, we used the CrossMab approach, in which the domain structure of one IgG antibody arm remains unchanged, while in the second arm, the heavy chain and light chain domains are crossed-over in the Fab regions28. To promote dimerization of the heavy chains of the two antibodies, we used a knob-in-hole approach where a knob is inserted into one heavy-chain CH3 domain and a hole is inserted into the other (Fig. 2A). The knob incorporates S354C and T366W substitutions and the hole incorporates Y349C and T366S substitutions. The heavy chain with the hole has additional H435R and Y436F substitutions in the CH3 domain to abolish binding to protein A29. These modifications promote generation of highly pure heterodimeric bsAb, iMab/D5_AR (knob arm/hole arm). As controls (indicated as “c”), we inserted the Fab arm from mAb114, which targets unrelated Ebola glycoprotein, into either the knob arm or the hole arm to generate c/D5_AR and iMab/c, respectively. The control antibody (mAb114) showed no neutralization against HIV-1 as expected (Fig. S1).
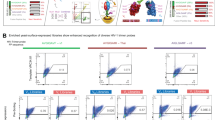
A Schematic of bispecific antibodies (bsAbs) generated by combining the antibody fragments of anti-NHR antibody D5_AR or anti-CD4 antibody iMab. mAb114 targeting unrelated Ebola glycoprotein is used as the control and is represented by the abbreviation ‘c’. A/B represents the antibody configuration, with antibody A contributing the knob arm (incorporating S354C and T366W substitutions) and antibody B contributing the hole arm (incorporating Y349C and T366S substitutions). B Flow cytometry to assess binding of bsAb to CD4-expressing TZM-bl cells and biotinylated NHR. Shown on the axes are the binding of the three antibodies to CD4 (probed with FITC-Fab fragment goat anti-human IgG, x-axis) and biotinylated NHR hydrophobic pocket mimetic IQN17 (probed with streptavidin-APC, y-axis). The configuration of the control Fab fragment is abbreviated as (c/) when on the knob arm and (/c) when on the hole arm. 10,000 cells were detected per measurement and results were plotted using FlowJo. The control shows binding to neither CCR5 nor NHR, while the bsAb binds both simultaneously. C Maximum % inhibition (left), IC50 (middle), and IC80 (right) for neutralization by iMab/D5_AR, its two control antibodies, and a mix of the two control antibodies, against a panel of 26 HIV-1 pseudotyped viruses representing diverse clades and tiers (see Table 1, source data are provided as a source data file). Each dot (n = 26) represents the average neutralization of biological duplicates of technical replicates for each pseudotyped virus. Maximum % inhibition values were calculated from the inhibition at the highest antibody concentration (50 μg/mL for iMab/D5_AR, and 50 μg/mL each for iMab/c and c/D5_AR in combination). IC50 and IC80 values were calculated using GraphPad Prism. Error bars indicate median +/- interquartile range. mAb114 Fab was used as a control Fab. The two-sided Wilcoxon-matched pairs signed test was used; **** indicates p < 0.0001. Black dotted lines indicate limit of quantification (50 μg/mL) of bsAb.
We next tested iMab/D5_AR alongside the controls c/D5_AR and iMab/c for simultaneous binding to CD4 and NHR on host cells using flow cytometry. We incubated TZM-bl cells which express CD430 with the bsAb or control antibodies, and after washing, with biotinylated NHR hydrophobic pocket mimetic IQN1712. We monitored binding at the cell surface with fluorescent probes streptavidin-APC and FITC-Fab fragment anti-human IgG. We found that only iMab/D5_AR, not the control antibodies, bound simultaneously to cells and to NHR mimetic (Fig. 2B). These results suggest that, consistent with its design, the bsAb effectively binds CD4 and NHR mimetic simultaneously.
NHR-targeting antibody exhibits broad and potent neutralization against HIV-1 through bispecific antibody-mediated prepositioning
To assess the neutralization breadth and potency of iMab/D5_AR, we tested it alongside control antibodies for neutralization in a luciferase-based reporter assay31 against a panel of 26 HIV-1 pseudotyped viruses representing diverse clades and neutralization tiers. The panel used in this study includes global HIV-1 Env reference strains32, highly neutralization-resistant tier-3 pseudoviruses, and transmitted/founder (T/F) pseudoviruses (RHPA4259 and WEAUd15.410), which are reported to be more infectious and have higher resistance to therapeutic fusion inhibitors and bnAbs33,34. This panel also includes three pseudoviruses (3103.v3.c10, WEAUd15.410.5017, and X2088_c9) reported to be resistant to 10E8/iMab24, which is in phase I clinical trials (NCT03875209 and NCT05890963) as an antibody-based prophylactic and therapeutic.
The two control antibodies containing one control Fab arm, c/D5_AR and iMab/c targeting either NHR only or CD4 only, as well as a combination of the two, showed limited to modest neutralization against all pseudoviruses, while iMab/D5_AR potently neutralized all of the pseudoviruses in the panel (Fig. 2C and Table 1). Notably, the neutralization IC80 values of iMab/D5_AR were in the range of efficacy for prevention of infection in humans (IC80 < 5 μg/mL)35 against all pseudoviruses in the panel (Table 1).
To further evaluate the neutralization potency of iMab/D5_AR, we expanded the virus panel to 119 pseudotyped viruses and compared it to the neutralization potencies of 12 previously identified bnAbs that bind to various epitopes of Env published in the CATNAP database which catalogs the efficacy of HIV-neutralizing antibodies. This panel of 12 bnAbs targets epitopes including CD4bs, V1/V2, and V3 of gp120, gp120/gp41 interface that are accessible at the native prefusion state36,37, and membrane-proximal external region (MPER) of gp41. iMab/D5_AR showed mean IC50 and IC80 values of 0.14 and 0.61 μg/mL, respectively, with an exceptional breadth of 95% (Fig. 3). Therefore, although the neutralization potency of iMab/D5_AR was comparable to that of other bnAbs (Fig. 3A), the breadth of protection was highest for the bsAb (Fig. 3B).
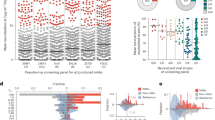
A Neutralization potencies of iMab/D5_AR and 12 bnAbs against a panel of 119 HIV-1 pseudotyped viruses. Antibodies are grouped by color based on which of six epitopes they target (CD4-binding site, CD4bs; V1/V2 loop of gp120, V1/V2 loop; high-mannose V3 loop, High-mannose V3 loop; gp120/gp41 interface; the membrane-proximal external region, MPER; and gp41 N-heptad repeat, NHR). The black dotted line indicates the limit of quantification (50 μg/mL). Error bars indicate median +/- interquartile range. Neutralization potencies of antibodies, apart from iMab/D5_AR and 10E8v4, were obtained from published studies curated in the Los Alamos HIV database (shown in gray background), CATNAP. Source data is provided as a source data file. Each dot represents the average neutralization of technical replicates for each pseudotyped virus. B Neutralization potency vs. breadth curve against 119 HIV-1 pseudotyped viruses. Antibodies were grouped by the epitope targeted (colors in key). Breadth was defined as non-protective when IC80 > 5 μg/mL. The blue dotted line indicates IC80 of 5 μg/mL. Source data ARE provided as a source data file.
Next, we tested the neutralization efficacy of iMab/D5_AR against a panel of six replication-competent HIV-1 viruses, using the luciferase-based reporter assay. This panel includes T/F viruses (THRO, CH106, CH077, and CH058) and viruses with coreceptor tropism of CCR5 (Q23, THRO, CH106, CH077, and CH058) or CXCR4 (NL4-3). Compared to the two control antibodies, our NHR-targeting bsAb, iMab/D5_AR (Fig. 4A, B) and the FDA-approved NHR-targeting peptide enfuvirtide (Fig. S2A) consistently neutralized all of the replication-competent HIV-1 viruses in the panel, with iMab/D5_AR being particularly potent (Fig. S2B).
A A table showing neutralization potencies of iMab/D5_AR against six replication-competent HIV-1 viruses using the luciferase-based reporter assay. The IC80 values of bsAb at <5, 5–50, and > 50 μg/mL were assigned the colors of red, blue, and white, respectively. R5 and X4 tropisms indicate CCR5 and CXCR4, respectively. B Maximum % inhibition for neutralization by iMab/D5_AR, its control antibodies, and a mix of the two control antibodies against a panel of six replication-competent viruses on TZM-bl cells. Maximum % inhibition values were calculated from the inhibition at the highest antibody concentration (50 μg/mL for iMab/D5_AR, iMab/c, or c/D5_AR individually, and 50 μg/mL each for iMab/c and c/D5_AR in combination). Each dot represents an average of biological duplicates of technical replicates. Error bars indicate median +/− interquartile range. mAb114 Fab was used as a control Fab. Source data are provided as a source data file. C Neutralization of bsAb against NL4-3 replication-competent HIV-1 virus on human peripheral blood mononuclear cells (PBMCs). Each dot represents an average of technical replicates (n = 3 human donors). Error bars indicate mean +/- standard deviations (SD). Source data are provided as a source data file.
To test the neutralization efficacy of our bsAb on human primary cells, we performed neutralization assays in peripheral blood mononuclear cells (PBMCs) against the NL4-3 replication-competent HIV-1 virus. NL4-3 virus uses CXCR4 as a coreceptor. CXCR4 coreceptor usage has been associated with decreasing CD4 + T cell counts and accelerated progression to acquired immunodeficiency syndrome (AIDS)38. iMab/D5_AR showed potent neutralization even at 1 μg/mL while the two control antibodies showed weak or no neutralization (Fig. 4C). At a high concentration (100 μg/mL), iMab/c weakly neutralized the virus, and the combination of control antibodies neutralized the virus effectively.
Finally, we tested whether iMab/D5_AR could also inhibit HIV-1 infection against TZM-bl cells expressing FcγRI, since dendritic cells and macrophages expressing FcγRI are susceptible to HIV-1 infection. iMab/D5_AR showed potent neutralization against HXB2 (tier-1) and 25710 (tier-2) HIV-1 pseudoviruses in the presence of FcγRI (Fig. S3).
These results suggest that prepositioning of the NHR-targeting antibody on CD4-expressing cells via bispecific antibody-mediated prepositioning is an effective approach to neutralize HIV-1.
Deep mutational scanning confirms neutralization mechanism of iMab/D5_AR
To determine the neutralization mechanism of iMab/D5_AR, we used a pseudovirus-based DMS system39 with two mutant libraries each containing ~40,000 mutant Env sequences from the subtype A T/F virus BF520.W14M.C2 (called BF520 hereafter)40; these mutant libraries were produced for a previous study and described in detail previously39. The DMS system measures the impact of individual and multiple mutations within the Env protein on viral neutralization, which enables mapping of residues on the Env protein important for the neutralization by iMab/D5_AR. We first validated the DMS system using enfuvirtide, as clinical resistance to enfuvirtide has been well-characterized on the IAS-USA drug resistance mutations list41 and DMS of enfuvirtide has been characterized previously42 on subtype A BG505 Env. Consistent with these prior results, DMS using enfuvirtide against the BF520 Env mutant libraries identified residues 544-556 in the NHR as dictating sensitivity to enfuvirtide (Fig. 5A). While still within the NHR, the residues identified in the DMS map of iMab/D5_AR sensitivity (564, 568, 574, and 577; Fig. 5B) were different from those of enfuvirtide (Fig. 5C). These four residues are all in the hydrophobic pocket where D5_AR binds, consistent with the bsAb design.
DMS maps of all sites in the BF520 Env ectodomain where mutations lead to resistance from (A) enfuvirtide- and (B) iMab/D5_AR-mediated viral neutralization. Positive values and negative values represent residues where mutations cause resistance or sensitivity, respectively. Individual points on the line plots represent the mean effect of mutations on escape at that site. Interactive versions of the resistance maps for enfuvirtide and iMab/D5_AR are available https://dms-vep.org/HIV_Envelope_BF520_DMS_iMab.D5_AR_Enfuvirtide/htmls/Enfuvirtide_mut_effect.html and https://dms-vep.org/HIV_Envelope_BF520_DMS_iMab.D5_AR_Enfuvirtide/htmls/iMab.D5_AR_mut_effect.html, respectively. C DMS-identified resistance sites are shown for enfuvirtide (blue) and iMab/D5_AR (purple) in NHR, mapped onto a structure of the gp41 NHR (PDB: 2CMR). D Neutralization of BF520 Env pseudotyped viruses with individual mutations by enfuvirtide, and iMab/D5_AR shown as maximum % inhibition calculated from the inhibition at the highest antibody concentration (50 μg/mL). Each dot represents an average of biological duplicates of technical replicates. E Table showing neutralization IC50 (nM) values of enfuvirtide and iMab/D5_AR against each of the four point mutations in the NHR hydrophobic pocket. F Neutralization by enfuvirtide, iMab/D5_AR, and control antibodies of HIV-1 isolate MVP-5180 pseudovirus with deviations from the known D5 epitope sequences at residues 574 and 577. Results are shown as mean from duplicate experiments. Error bars indicate mean +/− standard deviations (SD).
Next, we inserted individual mutations of the four hydrophobic pocket residues into pseudoviruses and assessed their susceptibility to neutralization in the luciferase-based reporter assay (Fig. 5D, E). As expected, the individual mutations did not exhibit any resistance (decrease in neutralization compared to the wild-type (WT)) against enfuvirtide. The four mutations showed resistance to iMab/D5_AR, confirming that the binding of D5_AR to the hydrophobic pocket in the NHR is the mechanism by which iMab/D5_AR acts to neutralize. As the mutations K574N and Q577R exhibited the highest resistance to iMab/D5_AR, we searched the Los Alamos HIV database for HIV-1 Env sequences with deviations from the known D5 epitope residues (K574 and Q577). We found an HIV-1 group O (outlier) isolate, MVP-5180, with R574 and R577. MVP-5180 was effectively neutralized by enfuvirtide, but only weakly neutralized by iMab/D5_AR (Fig. 5F), again confirming that mutations in the hydrophobic pocket of the NHR alter neutralization mediated by iMab/D5_AR.
Finally, we tested neutralization by D5_AR on FcγRI-expressing TZM-bl cells and confirmed that, while neutralization of the K574N mutation in pseudotyped virus was comparable to that of WT, the Q577R mutation decreased neutralization by D5_AR (Fig. S4).
iMab/D5_AR exhibits in vivo therapeutic efficacy
To determine in vivo efficacy of iMab/D5_AR, we used a humanized mouse model of HIV-1 infection43, for which human bone marrow-liver-thymus (BLT) is implanted into NOD/SCID/IL2rγnull (NSG) mice, and monitored by the presence of human CD45. The humanized mice were infected with tier-1B isolate NL4-3 virus, where the sensitivity of control antibodies was assessed on human primary cells in vitro (Fig. 4C). One week post-infection, we treated mice with weekly intraperitoneal injections of iMab/D5_AR, a combination of iMab/c and c/D5_AR, or PBS as a formulation control (Fig. 6A). Weekly injections were chosen to maintain sufficient antibody concentrations throughout the in vivo experiments (Fig. S5). We collected blood samples from the mice every 2 weeks to measure viral load (VL) titers. Mice treated with iMab/D5_AR exhibited a significant reduction in VL compared to the PBS formulation control, and the decrease in the VL persisted until the injections were discontinued at week 6 (Fig. 6B, C). The combination of control antibodies also reduced the VL, although not significantly compared to the control. We observed viral rebound at week 10, which correlated with decreased plasma concentrations of the bsAbs (Fig. 6D). Sequencing of the viruses from weeks 2 to 6 in iMab/D5_AR-treated mice revealed a mutation (30% Q575R) in the epitope of the NHR-targeting antibody (Fig. 6E), suggesting that the antiviral pressure we observed was primarily due to the binding of D5_AR to the NHR. Indeed, when we introduced Q575R into NL4-3 virus, the virus became resistant to iMab/D5_AR (Fig. S6). The mutation reverted (0.2% Q575R) by week 10 (Fig. 6E), at which point plasma concentrations of the bsAb were undetectable (Fig. 6D). Sequencing of the viruses in the mice treated with the combination of control antibodies showed no evidence of mutations in the NHR throughout the course of the experiment (Fig. S7A). However, mutations reported to confer resistance to iMab24 were detected at week 10 (Fig. S7B). These mutations were not found in the iMab/D5_AR-treated mice (Fig. S7C), indicating different antiviral pressures were imposed by the bsAb compared to the combination of control antibodies.
A Schematic representation of the experimental timeline for evaluating the efficacy of bsAbs in vivo. NOD/SCID/IL2rγnull (NSG) mice were implanted with human bone marrow-liver-thymus (BLT) tissues, followed by intravenous infection with 500 ng of NL4-3 replication-competent HIV-1 virus. Starting 1 week post-infection, mice were treated with weekly intraperitoneal injections of either iMab/D5_AR (2 mg), a combination of iMab/c (2 mg) and c/D5_AR (2 mg), or PBS as a control. Blood was drawn from the mice every two weeks for ten weeks for viral load (VL) titration. B Changes in plasma VL. Gray dashed lines indicate no change in VL from week 0. Black lines represent data from each mouse, and red lines represent the geometric mean of data from all mice (n = 8 for iMab/D5_AR, n = 4 for iMab/c + c/D5_AR, and n = 2 for PBS). C Geometric mean changes in plasma VL. Each line represents geometric changes in the VL on a log10 scale, and the error bars represent the standard deviation. iMab/D5_AR showed a statistical difference (p < 0.05 as determined by Mann-Whitney test) compared to the PBS group at weeks 2 and 4, whereas iMab/c + c/D5_AR did not show a significant difference from the PBS group (n = 8 for iMab/D5_AR, n = 4 for iMab/c + c/D5_AR, and n = 2 for PBS). D Plasma concentration of the bsAb in NSG mice. The gray dashed lines indicate the limit of quantification by ELISA at antibody concentration of 0.1 μg/mL. Black lines represent plasma concentration of the bsAbs from each mouse (n = 8 for iMab/D5_AR, n = 4 for iMab/c + c/D5_AR), with red lines representing the geometric mean. E The logo plots show the degree of conservation of the NHR from weeks 2–6 and week 10, with wild-type sequences shown in gray and mutation highlighted in blue.
Discussion
Here, we utilize bispecific antibody-mediated prepositioning to enhance binding to the transiently exposed NHR. Our bsAb targets the host cell receptor to preposition the NHR-targeting antibody on host cells, thereby enhancing its neutralizing potency at the HIV-1 PHI. This enhancement, which we observe in vitro in neutralization assays and in vivo in a humanized mouse model, is reminiscent of FcγRI-mediated potentiation at transiently exposed epitopes NHR and MPER16,17,24,44,45. Binding to viral lipids, a characteristic of gp41 MPER-targeting bnAbs that modulate neutralization46,47, has also been reported to contribute to enhancing neutralization in a similar manner to FcγRI by increasing the local antibody concentration48. We propose that our bsAb targeting CD4 prepositions the NHR-targeting antibody on the host cells in a manner similar to FcγRI-mediated or lipid-mediated potentiation.
The gp41 NHR is only transiently exposed in the PHI, preventing NHR-targeting antibodies from binding to intact virions49. In contrast, the gp41 MPER is accessible in the native pre-fusion state by MPER-targeting antibodies, which can bind to intact virions, likely due to tilting of the Env ectodomain, allowing access to the opposing MPER50,51,52. iMab/D5_AR neutralized all of the variants reported to be resistant to MPER-targeting antibodies (Fig. 2C), and it has been reported that MPER- and NHR-targeting antibodies are synergistic48. This suggests that antibodies targeting these two different regions could be used in combination.
We confirmed that binding to the NHR is responsible for the neutralizing activity of iMab/D5_AR. First, we used in vitro DMS-based mapping to identify mutations that led to resistance against the neutralizing activity of iMab/D5_AR, revealing residues in the hydrophobic pocket of the NHR-residues 564, 568, 574, and 577 (Fig. 5). Next, our in vivo efficacy study identified an additional resistance mutation, also in the NHR hydrophobic pocket, at residue 575 (Fig. 6E). Rapid reversion of the in vivo mutation (Q575R) in the absence of antiviral pressure suggests that it exerts a deleterious effect on viral fitness, consistent with the high conservation of the NHR (Fig. 1B). This finding further strengthens the rationale for developing NHR-targeting antibody therapeutics.
The passive infusion of bnAbs as a prophylactic approach to prevent HIV-1 acquisition has been shown to be effective in clinical trials and is currently under active investigation35,53,54. The potency of iMab/D5_AR is comparable to what was reported in the Antibody Mediated Prevention (AMP) clinical trials to assess the ability of the patient-derived gp120 CD4bs-targeting bnAb (VRC01) as preexposure prophylaxis35. In those studies, the authors found a correlation between HIV-1 prevention efficacy in humans and in vitro neutralization (IC80 < 5 μg/mL) among different HIV-1 strains. iMab/D5_AR showed IC80 < 5 μg/mL against 95% of viruses in a panel of 119 pseudoviruses (Fig. 3). As well, while its neutralization potency was comparable to those reported for other bnAbs, including those in clinical development, the breadth of protection was highest for iMab/D5_AR.
We also acknowledge potential limitations to our study. The resistance mutations identified in the in vitro DMS and in the in vivo study were different. One possible explanation is that the DMS used the BF520 envelope, while the in vivo study used the NL4-3 envelope, and previous studies have shown that mutations can have varying effects across different HIV-1 strains55. Despite these differences, all identified residues are contact residues of the NHR-targeting antibody, confirming that the binding of D5_AR to the hydrophobic pocket in the NHR is the mechanism by which iMab/D5_AR acts to neutralize. Another limitation is that the in vivo efficacy of iMab/D5_AR was not statistically more potent than the combination of control antibodies (iMab/c and c/D5_AR, Fig. 6). This outcome may be attributed to the high plasma concentrations of the antibodies (2 mg injected weekly). While iMab/D5_AR effectively neutralized NL4-3 replication-competent HIV-1 virus at 1 μg/mL, the combination of control antibodies also demonstrated effective neutralization at 100 μg/mL in the human PBMCs neutralization assay (Fig. 4C).
Overall, our results show that targeting the PHI with the NHR-targeting antibody iMab/D5_AR leads to extremely broad and effective neutralization. This further validates the NHR as a druggable target for antibodies (Fig. 7) and establishes a new class of broadly neutralizing antibodies against HIV-1.
Sites of vulnerability to broadly neutralizing antibodies on the HIV-1 Env trimer, shown from two perspectives (PDB: 7SKA). NHR is inaccessible in this state and is only transiently exposed in the prehairpin intermediate conformation.
Methods
Sequence conservation in HIV-1 envelope
8394 HIV-1 Env sequences were obtained from the Los Alamos 2021 HIV database (Table S1). Env with X, stop codons, or sequences with NHR frameshifts were filtered, resulting in 8155 unique Env sequences. These sequences were aligned using Geneious alignment (Geneious Prime) and used to calculate sequence identities for gp120 and gp41. The sequences were grouped or clustered into those with 90% homology within Env (129 sequences), and clusters with more than 1 Env (65 sequences) were then aligned to generate multiple sequence alignment (MSA) using Geneious Prime and uploaded onto the consurf server for overlay onto 2CMR structure on chain A.
Bispecific antibody expression and purification
Genes encoding the light and heavy chain of antibodies were synthesized by Integrated DNA Technologies (IDT). Sequences of heavy and light chain of antibodies can be found in the Supplementary Information. The genes were cloned into linearized bispecific IgG1 plasmids obtained as described56 using In-Fusion HD Cloning Kit Master Mix (Clontech). Heavy knob, light chain, CrossMab heavy chain with hole, and CrossMab light chain were co-transfected at a 1:1:1:1 ratio. The bsAb was produced in Expi293F cells (Thermo Fisher Scientific) using FectoPRO (Polyplus). The cells were incubated under 8% CO2 at 37 °C with shaking (120 rpm). The cells were harvested 4–5 days post-transfection by spinning at 4,000 x g for 10 min and filtered through a 0.22-μm filter. The bsAb was purified using a 5 mL MabSelect Sure PRISMTM column (Cytiva) on the AKTA pure FPLC followed by further purification using size-exclusion chromatography on AKTA FPLC using GE Superdex 200 increase 10/300 GL column (GE HealthCare).
HIV-1 virus production
HIV-1 pseudotyped lentiviruses were produced by co-transfection of HIV-1 Env plasmids (10 μg) and psg3Δ Env backbone plasmids (20 μg, catalog number 11051 from NIH AIDS Reagent program) to HEK293T cells using BioT (Bioland Scientific LLC) following the manufacturer’s instructions. The medium was replaced with fresh medium after 16 h post-transfection, and the supernatant was harvested 2 d post-media replacement. The supernatant was centrifuged at 300 × g for 5 min and filtered through a 0.45-μm filter, and stored at −80 °C until further use. Strains of pseudotyped HIV-1 are listed in the excel dataset file.
Replication-competent viruses were produced and titrated as described previously57. Briefly, HIV-1 full length plasmids were transfected to HEK293T cells using FuGENE (Promega) following the manufacturer’s instructions. Media replacement and virus harvest were performed as described above.
Cell culture
TZM-bl cells (obtained through the NIH AIDS Reagent program from John C. Kappes and Xiaoyun Wu) and HEK293T cells were maintained in Dulbecco’s modified eagle medium (DMEM) supplemented with 10% fetal bovine serum (FBS), 1% L-glutamine (Corning) and 1% penicillin-streptomycin (Corning) and incubated with 5% CO2 at 37 °C in a humidified atmosphere. TZM-bl cells transduced to stably express FcγRI were maintained in DMEM supplemented as above and blasticidin (Thermo Fisher Scientific) as described previously44. No commonly misidentified cell lines were used in the study.
Viral neutralization assay
TZM-bl cells with tat-regulated luciferase reporter gene expression were used for quantification of viral infection and antibody neutralization as described previously58,59. Briefly, 5000 TZM-bl cells were seeded overnight in white-walled 96-well plates with 5% CO2 at 37 °C in a humidified atmosphere. The medium was aspirated the next day, and bsAb (50 μL) in various concentrations was added to the cells for 1 h, and a mixture (50 μL) containing HIV-1 pseudotyped lentivirus and DEAE-dextran (10 μg/mL) was added to the cells. After two days, cells were lysed, and BriteLite Plus reagent (Perkin Elmer) was added for determination of luciferase activity. Relative luminesce unit (RLU) values were measured using a Synergy HTX multimode reader (BioTek) and % infection was calculated by the following formula: [% infection = (the luminescent signal of the test well—the luminescent signal of the background well) / (the luminescent signal of virus only well—the luminescent signal of background well) × 100]. Wells containing cells only were used for background measurement.
Human PBMCs neutralization assay
PBMCs were isolated from healthy donors (Stanford Blood Center) using Lymphoprep (Stemcell Technologies) following the manufacturer’s protocol. PBMCs (2 × 107) were stimulated with phytohemagglutinin (500X, Invitrogen) and interleukin-2 (100 U/mL, PeproTech) in complete RPMI 1640 medium containing 10% FBS and 1% penicillin-streptomycin, and incubated for 48 h at 37 °C in a humidified atmosphere with 5% CO2. After stimulation, PBMCs (2 × 105 cells in 100 μL) were seeded in white-walled 96-well plates. The bsAb (50 μL) at various concentrations was added to the cells for 1 h, followed by NL4-3 replication-competent HIV-1 virus (50 μL). After 24 h, the cells were centrifuged and washed 3 times with complete RPMI 1640 medium. Complete RPMI 1640 medium with bsAb (200 μL) was added to the cells. After 4 d incubation, the cells were centrifuged, and supernatants were collected. HIV-1 infection and replication were determined by measuring the p24 antigen in the culture supernatant using HIV-1 p24 antigen enzyme-linked immunosorbent assay (ELISA) kit (ZeptoMetrix RETROtek) following the manufacturer’s protocol.
Flow cytometry
TZM-bl cells expressing CD430 were incubated with bsAb (10 nM) in flow cytometry buffer (1% [w/v] BSA in PBS containing 0.05% [w/v] sodium azide) for 1 h at 4 °C. The cells were washed four times with flow cytometry buffer and probed with biotinylated NHR hydrophobic pocket mimetic IQN17 (10 nM)12 for 1 h at 4 °C. After washing with flow cytometry buffer, the cells were probed with streptavidin-APC (1:200, BioLegend) and Fluorescein (FITC)- Fab fragment goat anti-human IgG (1:50, Jackson ImmunoResearch) in flow cytometry buffer for 1 h at 4 °C. The cells were washed four times and sorted using AccuriTM C6 Plus (BD Biosciences). 10,000 cells were detected per measurement, and results were analyzed using FlowJo.
Deep mutational scanning
The previously described pseudovirus-based DMS platform39,60 was used to map HIV Env resistance from neutralization by enfuvirtide and iMab/D5_AR. Briefly, we used the two previously described biologically independently produced mutant libraries of HIV Env strain BF52039, which each contained ~40,000 mutants with an average of ~2.5 non-synonymous mutations per mutant. During antibody selections, VSV-G pseudotyped neutralization standard viruses were spiked-in to be 0.5-1% of the virus pool. For each antibody selection, we incubated 1 million TZM-bl cells in a single well of a six-well dish with the antibody of interest at 2x the target concentration in 1 mL. We then added 1 million infectious units of the pool of mutant viruses plus VSV-G standards in 1 mL to each condition, supplemented with 100 μg/mL DEAE dextran. 12 h after infection, each condition was mini-prepped to isolate the unintegrated lentivirus genomes of non-neutralized viruses, and then sequencing library preparation and sequencing were performed as described in Radford et al. 39.
Deep mutational scanning data analysis was performed using dms-vep-pipeline-3. See (https://github.com/dms-vep/HIV_Envelope_BF520_DMS_iMab.D5_AR_Enfuvirtide) for a repository containing the analysis performed for this work (version 3.4.5). See (https://dms-vep.org/HIV_Envelope_BF520_DMS_iMab.D5_AR_Enfuvirtide/) for HTML renderings of key analyses and results as well as interactive plots. Analysis of sequencing data and modeling of the effects of mutations on HIV Env function and antibody resistance were performed as previously described in Radford et al. 39 and Dadonaite et al. 60. Briefly, for antibody resistance modeling, we calculated the non-neutralized fraction of each mutant in each antibody selection using the counts of each mutant and the counts of the non-neutralized standard viruses in antibody incubation versus mock incubation conditions. We used the software package polyclonal61 version 6.7 (see https://jbloomlab.github.io/polyclonal/ for documentation) to model the effects of each individual mutation on resistance from each antibody. For Fig. 5, the line plots display the mean resistance score of mutations at each site.
Generation of humanized mice
Humanized mice were established as described previously43, in accordance with protocols (ARC-2010-038) approved by the UCLA Institutional Animal Care and Use Committees. Briefly, 7–10 weeks old male NSG mice (Strain #005557 from the Jackson Laboratory) were irradiated sublethally and implanted with BLT and CD34+ cells purified from fetal liver. Ten weeks after implantation, the engraftment frequency was determined by staining the blood lymphocytes and splenocytes for human CD45. One week later, the mice were then infected i.v. with 500 ng NL4-3 replication-competent HIV-1 virus. After confirmation of viral infection, the bsAbs were injected intraperitoneally (2 mg) weekly for 6 weeks. In the case of the combination of iMab/c and c/D5_AR, 2 mg of each antibody was injected. Mice were retro-orbital bled every 2 weeks for plasma VL titration.
Plasma viral load quantification
Mouse blood was clarified through centrifugation at 350 × g, and RNA extraction was performed using the QIAamp Viral RNA Mini Kit (Qiagen). Quantification of HIV-1 RNA was carried out using real-time RT-PCR with TaqMan RNA-To-Ct One-Step reagents (Thermo Fisher Scientific). The primers used were HIV-1_F: 5′-CAATGGCAGCAATTTCACCA-3′ and HIV-1_R: 5′-GAATGCCAAATTCCTGCTTGA-3′, along with a probe designed to hybridize to HIV-1 NL4-3 5′-[6-FAM]CCCACCAACAGGCGGCCTTAACTG [Tamra-Q]-3′.
Quantification of plasma concentration of bsAbs from humanized mice using ELISA
Nunc 96-well Maxisorp plates (Thermo Fisher Scientific) were coated overnight at 4 °C with 250 ng of streptavidin in coating buffer (0.1 M sodium bicarbonate, pH 8.6). Wells were blocked with 150 μL of ChonBlock buffer (Chondrex) for 1 h at 37 °C. Biotinylated IQN17 (20 nM) or biotinylated CD4 (200 ng/mL, ACROBiosystems) in ELISA buffer (3% BSA/PBS) was added to the wells. After incubation for 1 h at 37 °C, plates were washed with 0.05% (v/v) Tween 20 in PBS (PBST) three times, followed by incubation with serially diluted mice sera (1:100–1:7,812,500) or serially diluted bsAbs (5–0.000064 μg/mL) for standards. After washing three times with PBST, the plates were incubated for 1 h at 37 °C with goat anti-human IgG-horseradish peroxidase (SouthernBiotech) at a 1:8000 dilution. The plates were washed six times with PBST, followed by the addition of Turbo 3,3’,5,5’ tetramethylbenzidine (TMB)-ELISA substrate solution (Thermo Fisher Scientific) and 2 M sulfuric acid stop solution. Absorbance was measured at 450 nm using a BioTek Synergy HT Microplate Reader.
HIV-1 Env sequencing
Mouse plasma HIV-1 RNA extraction was performed using QIAamp MinElute Virus Spin Kit (Qiagen). Subsequently, cDNA was generated using SMARTScribe Reverse Transcriptase (Takara) following the manufacturer’s instructions, using an Oligo(dT) primer. The Env sequence was amplified by PCR using CloneAmp HiFi PCR Premix (Takara) with a forward primer (5’-GAGCAGAAGACAGTGGC-3’) and a reverse primer (5’-CCACTTGCCACCCATCTTATAG-3’). The cycling conditions were 2 min at 98 °C, followed by 45 cycles of 98 °C for 10 s, 58 °C for 15 s, and 72 °C for 45 s, concluding with 2 min at 72 °C. Next, nested PCR was performed using a forward primer (5’-GAGCAGAAGACAGTGGC-3’) and a reverse primer (5’-CTATTCTTAAACCTACCAAGCCTC-3’) to amplify the ectodomain of the HIV-1 env. The PCR product was sequenced using Oxford Nanopore sequencing (Plasmidsaurus).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data supporting the findings in this study are available within the paper and the Supplementary Information. The Source Data and Supplementary Information are included with this publication. The sequences of all the Env used in this study are available at CATNAP. Source data are provided with this paper.
Code availability
No new code was generated, and previously published code was used for DMS39.
References
Orloff, G. M., Orloff, S. L., Kennedy, M. S., Maddon, P. J. & McDougal, J. S. Penetration of CD4 T cells by HIV-1. The CD4 receptor does not internalize with HIV, and CD4-related signal transduction events are not required for entry. J. Immunol. 146, 2578–2587 (1991).
Moore, J. P., Trkola, A. & Dragic, T. Co-receptors for HIV-1 entry. Curr. Opin. Immunol. 9, 551–562 (1997).
Chan, D. C. & Kim, P. S. HIV entry and its inhibition. Cell 93, 681–684 (1998).
Eckert, D. M. & Kim, P. S. Mechanisms of viral membrane fusion and its inhibition. Annu. Rev. Biochem. 70, 777–810 (2001).
Harrison, S. C. Viral membrane fusion. Nat. Struct. Mol. Biol. 15, 690–698 (2008).
Ladinsky, M. S. et al. Electron tomography visualization of HIV-1 fusion with target cells using fusion inhibitors to trap the pre-hairpin intermediate. eLife 9, e58411 (2020).
Kilby, J. M. et al. Potent suppression of HIV-1 replication in humans by T-20, a peptide inhibitor of gp41-mediated virus entry. Nat. Med. 4, 1302–1307 (1998).
LaBonte, J., Lebbos, J. & Kirkpatrick, P. Enfuvirtide. Nat. Rev. Drug Discov. 2, 345–346 (2003).
Miller, M. D. et al. A human monoclonal antibody neutralizes diverse HIV-1 isolates by binding a critical gp41 epitope. Proc. Natl Acad. Sci. Usa. 102, 14759–14764 (2005).
Chan, D. C., Fass, D., Berger, J. M. & Kim, P. S. Core structure of gp41 from the HIV envelope glycoprotein. Cell 89, 263–273 (1997).
Fernandes, J., Jayaraman, B. & Frankel, A. The HIV-1 Rev response element: an RNA scaffold that directs the cooperative assembly of a homo-oligomeric ribonucleoprotein complex. RNA Biol. 9, 6–11 (2012).
Bruun, T. U. J. et al. Structure-guided stabilization improves the ability of the HIV-1 gp41 hydrophobic pocket to elicit neutralizing antibodies. J. Biol. Chem. 299, 103062 (2023).
Luftig, M. A. et al. Structural basis for HIV-1 neutralization by a gp41 fusion intermediate–directed antibody. Nat. Struct. Mol. Biol. 13, 740–747 (2006).
Gustchina, E. et al. Structural basis of HIV-1 neutralization by affinity matured Fabs directed against the internal trimeric coiled-coil of gp41. PLoS Pathog. 6, e1001182 (2010).
Sabin, C. et al. Crystal structure and size-dependent neutralization properties of HK20, a human monoclonal antibody binding to the highly conserved heptad repeat 1 of gp41. PLoS Pathog. 6, e1001195 (2010).
Montefiori, D. C. et al. The high-affinity immunoglobulin receptor FcγRI potentiates HIV-1 neutralization via antibodies against the gp41 N-heptad repeat. Proc. Natl. Acad. Sci. USA 118, e2018027118 (2021).
Rubio, A. A. et al. A derivative of the D5 monoclonal antibody that targets the gp41 n-heptad repeat of HIV-1 with broad tier-2-neutralizing activity. J. Virol. 95, e02350-20 (2021).
Bell, B. N., Bruun, T. U. J., Friedland, N. & Kim, P. S. HIV-1 prehairpin intermediate inhibitors show efficacy independent of neutralization tier. Proc. Natl. Acad. Sci. USA 120, e2215792120 (2023).
Groot, F., Welsch, S. & Sattentau, Q. J. Efficient HIV-1 transmission from macrophages to T cells across transient virological synapses. Blood 111, 4660–4663 (2008).
Pope, M. & Haase, A. T. Transmission, acute HIV-1 infection and the quest for strategies to prevent infection. Nat. Med 9, 847–852 (2003).
Seaman, M. S. et al. Tiered categorization of a diverse panel of HIV-1 Env pseudoviruses for assessment of neutralizing antibodies. J. Virol. 84, 1439–1452 (2010).
Pace, C. S. et al. Bispecific antibodies directed to CD4 domain 2 and HIV envelope exhibit exceptional breadth and picomolar potency against HIV-1. Proc. Natl. Acad. Sci. USA 110, 13540–13545 (2013).
Sun, M. et al. Rational design and characterization of the novel, broad and potent bispecific HIV-1 neutralizing antibody iMabm36. JAIDS J. Acquired Immune Defic. Syndromes 66, 473–483 (2014).
Huang, Y. et al. Engineered bispecific antibodies with exquisite HIV-1-neutralizing activity. Cell 165, 1621–1631 (2016).
Moshoette, T., Ali, S. A., Papathanasopoulos, M. A. & Killick, M. A. Engineering and characterising a novel, highly potent bispecific antibody iMab-CAP256 that targets HIV-1. Retrovirology 16, 31 (2019).
Moshoette, T., Papathanasopoulos, M. A. & Killick, M. A. HIV-1 bispecific antibody iMab-N6 exhibits enhanced breadth but not potency over its parental antibodies iMab and N6. Virol. J. 19, 143 (2022).
Jacobson, J. M. et al. Safety, pharmacokinetics, and antiretroviral activity of multiple doses of ibalizumab (formerly TNX-355), an anti-CD4 monoclonal antibody, in human immunodeficiency virus type 1-infected adults. Antimicrob. Agents Chemother. 53, 450–457 (2009).
Klein, C. et al. Progress in overcoming the chain association issue in bispecific heterodimeric IgG antibodies. mAbs 4, 653–663 (2012).
Tustian, A. D., Endicott, C., Adams, B., Mattila, J. & Bak, H. Development of purification processes for fully human bispecific antibodies based upon modification of protein A binding avidity. mAbs 8, 828–838 (2016).
Platt, E. J., Wehrly, K., Kuhmann, S. E., Chesebro, B. & Kabat, D. Effects of CCR5 and CD4 Cell Surface Concentrations on Infections by Macrophagetropic Isolates of Human Immunodeficiency Virus Type 1. J. Virol. 72, 2855–2864 (1998).
Montefiori, D. C. Evaluating neutralizing antibodies against HIV, SIV, and SHIV in luciferase reporter gene assays. CP in Immunology 64, (2004).
deCamp, A. et al. Global panel of HIV-1 Env reference strains for standardized assessments of vaccine-elicited neutralizing antibodies. J. Virol. 88, 2489–2507 (2014).
Keele, B. F. et al. Identification and characterization of transmitted and early founder virus envelopes in primary HIV-1 infection. Proc. Natl Acad. Sci. Usa. 105, 7552–7557 (2008).
Parrish, N. F. et al. Phenotypic properties of transmitted founder HIV-1. Proc. Natl. Acad. Sci. USA 110, 6626–6633 (2013).
Corey, L. et al. Two randomized trials of neutralizing antibodies to prevent HIV-1 acquisition. N. Engl. J. Med 384, 1003–1014 (2021).
Pancera, M. et al. Structure and immune recognition of trimeric pre-fusion HIV-1 Env. Nature 514, 455–461 (2014).
Pancera, M., Changela, A. & Kwong, P. D. How HIV-1 entry mechanism and broadly neutralizing antibodies guide structure-based vaccine design. Curr. Opin. HIV AIDS 12, 229–240 (2017).
Connor, R. I., Sheridan, K. E., Ceradini, D., Choe, S. & Landau, N. R. Change in coreceptor use correlates with disease progression in HIV-1–infected individuals. J. Exp. Med. 185, 621–628 (1997).
Radford, C. E. et al. Mapping the neutralizing specificity of human anti-HIV serum by deep mutational scanning. Cell Host Microbe 31, 1200–1215.e9 (2023).
Simonich, C. A. et al. HIV-1 neutralizing antibodies with limited hypermutation from an infant. Cell 166, 77–87 (2016).
Wensing, A. M. et al. 2017 update of the drug resistance mutations in HIV-1.Top Antivir. Med. 24, 132–133 (2017).
Dingens, A. S., Arenz, D., Overbaugh, J. & Bloom, J. D. Massively parallel profiling of HIV-1 resistance to the fusion inhibitor enfuvirtide. Viruses 11, 439 (2019).
Kitchen, S. G. et al. In vivo suppression of HIV by antigen specific T cells derived from engineered hematopoietic stem cells. PLoS Pathog. 8, e1002649 (2012).
Perez, L. G., Costa, M. R., Todd, C. A., Haynes, B. F. & Montefiori, D. C. Utilization of immunoglobulin G Fc receptors by human immunodeficiency virus type 1: a specific role for antibodies against the membrane-proximal external region of gp41. J. Virol. 83, 7397–7410 (2009).
Perez, L. G., Zolla-Pazner, S. & Montefiori, D. C. Antibody-dependent, FcγRI-mediated neutralization of HIV-1 in TZM-bl cells occurs independently of phagocytosis. J. Virol. 87, 5287–5290 (2013).
Rujas, E. et al. Functional optimization of broadly neutralizing HIV-1 antibody 10E8 by promotion of membrane interactions. J. Virol. 92, e02249-17 (2018).
Irimia, A. et al. Lipid interactions and angle of approach to the HIV-1 viral membrane of broadly neutralizing antibody 10E8: Insights for vaccine and therapeutic design. PLoS Pathog. 13, e1006212 (2017).
Kim, S., Filsinger Interrante, M. V. & Kim, P. S. Enhancing HIV-1 neutralization by increasing the local concentration of membrane-proximal external region-directed broadly neutralizing antibodies. J. Virol. 97, e01647–22 (2023).
Nelson, J. D. et al. Antibody elicited against the gp41 N-heptad repeat (NHR) coiled-coil can neutralize HIV-1 with modest potency but non-neutralizing antibodies also bind to NHR mimetics. Virology 377, 170–183 (2008).
Lee, J. H., Ozorowski, G. & Ward, A. B. Cryo-EM structure of a native, fully glycosylated, cleaved HIV-1 envelope trimer. Science 351, 1043–1048 (2016).
Carravilla, P. et al. Molecular recognition of the native HIV-1 MPER revealed by STED microscopy of single virions. Nat. Commun. 10, 78 (2019).
Yang, S. et al. Dynamic HIV-1 spike motion creates vulnerability for its membrane-bound tripod to antibody attack. Nat. Commun. 13, 6393 (2022).
Mendoza, P. et al. Combination therapy with anti-HIV-1 antibodies maintains viral suppression. Nature 561, 479–484 (2018).
Sobieszczyk, M. E. et al. Safety, tolerability, pharmacokinetics, and immunological activity of dual-combinations and triple-combinations of anti-HIV monoclonal antibodies PGT121, PGDM1400, 10-1074, and VRC07-523LS administered intravenously to HIV-uninfected adults: a phase 1 randomised trial. Lancet HIV 10, e653–e662 (2023).
Haddox, H. K., Dingens, A. S., Hilton, S. K., Overbaugh, J. & Bloom, J. D. Mapping mutational effects along the evolutionary landscape of HIV envelope. eLife 7, e34420 (2018).
Weidenbacher, P. A.-B. et al. Converting non-neutralizing SARS-CoV-2 antibodies into broad-spectrum inhibitors. Nat. Chem. Biol. 18, 1270–1276 (2022).
Provine, N. M., Puryear, W. B., Wu, X., Overbaugh, J. & Haigwood, N. L. The infectious molecular clone and pseudotyped virus models of human immunodeficiency virus type 1 exhibit significant differences in virion composition with only moderate differences in infectivity and inhibition sensitivity. J. Virol. 83, 9002–9007 (2009).
Montefiori, D. C. Measuring HIV Neutralization in a Luciferase Reporter Gene Assay. in HIV Protocols (eds. Prasad, V. R. & Kalpana, G. V.) vol. 485 395–405 (Humana Press, 2009).
Sarzotti-Kelsoe, M. et al. Optimization and validation of the TZM-bl assay for standardized assessments of neutralizing antibodies against HIV-1. J. Immunol. Methods 409, 131–146 (2014).
Dadonaite, B. et al. A pseudovirus system enables deep mutational scanning of the full SARS-CoV-2 spike. Cell 186, 1263–1278.e20 (2023).
Yu, T. C. et al. A biophysical model of viral escape from polyclonal antibodies. Virus Evolut. 8, veac110 (2022).
Acknowledgements
We thank members of the Peter Kim laboratory for fruitful discussion and useful comments on the manuscript. TZM-bl and TZM-bl/FcγRI cells were obtained from the NIH AIDS Reagent Program, contributed by Drs. John C. Kappes and Xiaoyun Wu. The replication-competent viruses were gifted by Dr. Catherline Blish (Stanford University). This work was supported by the National Institutes of Health (NIH) award number 5DP1AI158125 (P.S.K.), the Virginia & D.K. Ludwig Fund for Cancer Research (P.S.K.), and the Chan Zuckerberg Biohub (P.S.K.).
Author information
Authors and Affiliations
Contributions
Conceptualization, S.K. and P.S.K.; investigation, S.K. and P.S.K.; validation and analysis, S.K., M.S.S., B.W., M.P.; methodology, S.K., C.E.R., J.D.B. and P.S.K.; software, C.E.R., J.D.B.; assistance for in vitro validation, D.X., J.Z., D.M.P., K.A.T., M.V.F.I., T.U.J.B.; assistance for in vivo pharmacokinetics, J.D.; in vivo neutralization, V.Z., S.G.K.; writing-original draft, S.K., C.E.R., J.D.B., P.S.K.; writing, review and editing, S.K., C.E.R., D.X., J.Z., D.M.P., K.A.T., J.D., M.V.F.I., T.U.J.B., V.Z., B.W., M.P., S.G.K., J.D.B., M.S.S., P.S.K.; supervision, P.S.K.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Victoria Walker-Sperling and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kim, S., Radford, C.E., Xu, D. et al. A broad antibody with enhanced HIV-1 neutralization via bispecific antibody-mediated prepositioning. Nat Commun 16, 4617 (2025). https://doi.org/10.1038/s41467-025-60035-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-60035-6