Abstract
Oligonucleotide-based medicine faces challenges in efficiently crossing the blood-brain barrier and rapidly reducing toxic proteins. To address these challenges, here we establish an integrated modality, brain-penetrant DNA nanoflowers incorporated with oligonucleotide-based proteolysis targeting chimeras. Using FUS as a proof-of-concept, mutations of which cause frontotemporal dementia and amyotrophic lateral sclerosis, we demonstrate that a FUS-engaging RNA oligonucleotide crosslinked to a ligand for Cereblon efficiently degrade FUS and its cytoplasmic disease-causing mutants through a ubiquitin-proteasomal pathway. The DNA nanoflower contains hundreds of oligonucleotide binding sites and transferrin receptor-engaging aptamers, allowing efficient loading of the oligonucleotide-based degrader and engaging transferrin receptors for brain delivery. A single dose intravenous injection of this modality reaches brain parenchyma within 2 h and degrades 80% FUS protein there, sustained for two weeks without noticeable toxicity. DNA nanoflower oligonucleotide-based degrader is a therapeutic strategy for neurodegenerative diseases that leverages the advantages of designer oligonucleotides and targeted protein degradation.
Similar content being viewed by others
Introduction
Oligonucleotide (Oligo) therapies have achieved multiple successes in drug development in recent years. However, the delivery strategy remains challenging, especially the blood-brain barrier (BBB)-crossing delivery for neurodegenerative diseases and other diseases in the central nervous system. Oligos are negatively charged molecules and are cleared quickly in the cerebrospinal fluid1. Current published BBB-crossing strategies for Oligo delivery include metal-based or sugar-coated nanoparticles2,3, transferrin receptor (TfR)-binding peptides or antibodies4,5, and liposomes functionally coated with functional groups for transcytosis6,7, none of which have been clinically validated. Moreover, some face low delivery efficacy, and others show side effects such as anemia8,9. Although the TfR-targeting strategy is promising with mutations on the FC chain that may reduce anemia, protein-based TfR engager still requires sophisticated linker design for Oligo conjugation.
Frontotemporal dementia (FTD) and amyotrophic lateral sclerosis (ALS), two devastating progressive neurodegenerative diseases10, share clinical symptoms, pathological features, and genetic risks11,12. Emerging evidence reveals that abnormal protein aggregates formed by RNA-binding proteins, such as fused-in-sarcoma (FUS) and TAR DNA-binding protein 43 (TDP-43), are a pathological hallmark of FTD and ALS13,14. Under physiological conditions, FUS is located predominantly in the nucleus, while disease-causing mutations drive its mislocalization and aggregation in the cytoplasm12. Some de novo FUS mutants are associated with the most severe, early-onset ALS and FTD15. FUS mutations are classified into three categories: (1) point mutations in the IDR (intrinsically disordered region), RGG1 (arginine–glycine–glycine repeat 1) or RGG2 domains are likely to increase the aggregation propensity, (2) truncation mutants and mutations in the nuclear localization sequence (NLS) increase cytoplasmic accumulation of FUS15, and (3) mutations in the 3’ non-coding region of FUS mRNA may cause overexpression16,17. Despite the extensive efforts in drug development for FUS, only antisense Oligos (ASO)-mediated gene silencing shows some promise in a few rare cases18.
Proteolysis targeting chimeras (PROTACs) are heterobifunctional molecules19. By connecting one ligand for the protein of interest (POI) and the other for an E3 ubiquitin ligase, both of which do not naturally interact, with a well-designed linker, PROTACs engage the POI and E3 in a ternary complex20. This induced proximity allows ubiquitination of the POI by the E3, and subsequently proteasomal degradation of the POI. Because of this unique mechanism of action, PROTACs are attractive modalities for undruggable proteins, particularly in treating neurodegenerative diseases21. For example, PROTACs targeting pathogenic misfolded proteins, such as Tau protein and alpha synuclein, have been reported22,23. However, FUS-targeted PROTACs is not reported yet. Moreover, the BBB poses another challenge for effective delivery of PROTACs24,25. Besides peptides and small molecules, recent studies have demonstrated that Oligos are a promising ligand for undruggable proteins in PROTAC discovery26,27,28. For example, Oligo-PROTACs have been reported to degrade transcription factors28,29.
In this work, by taking advantage of Oligos in molecular recognition and programmability, we develop an RNA Oligo-PROTAC and a brain-penetrant DNA nanoflower containing hundreds to thousands of copies of TfR-engaging aptamers, which also contains numerous sequences for loading the Oligo-PROTAC. The nanoflower loaded with an RNA oligonucleotide-based PROTAC (FRONTAC) realizes BBB-crossing delivery and specific degradation of disease-causing proteins. We have proven our concept by designing a selective FRONTAC for FUS. The FRONTAC technology offers a general and promising strategy for targeting disease-causing proteins in the central nervous system.
Results
Engineering a selective FUS-engaging oligo for targeted protein degradation
Because RNA-binding protein FUS recognizes the GGUG motif30, we selected five RNA Oligos with high affinity to develop FUS degraders using the Oligo-PROTAC strategy (Fig. 1a). In addition, to increase the affinity between FUS and the Oligo-PROTAC, we also engineered the FUS-RNA#9, an Oligo with the highest affinity, by tandemly duplicating the GGUG-containing motif (Fig. 1a, b). The Oligo-PROTACs were generated via a click reaction between azide-modified RNA Oligos and two alkyne-modified ligands: VH032 for von Hippel-Lindau (VHL) or thalidomide for Cereblon (CRBN) (Fig. 1a and Supplementary Fig. 1a). Among the tested Oligo-PROTACs, dFUS-E9C, consisting of an improved FUS-RNA#E9 and CRBN ligand (CRBNL), is the most potent, whereas dFUS-E9V failed to degrade FUS, which consists of the FUS-RNA#E9 and VHL ligand (VHLL) (Fig. 1c, d). To investigate the differential degradation efficacy of dFUS-E9C and dFUS-E9V, we examined the formation of ternary complexes by bimolecular fluorescence complementation assays (Supplementary Fig. 1b). The fluorescence intensity was significantly boosted in the presence of dFUS-E9C or dFUS-E9V (Supplementary Fig. 1c, d), indicating the ternary complex formation in both cases. The gating strategy for cell sorting to detect BiFC signals is shown in Supplementary Fig. 1e. However, dFUS-E9C but not dFUS-E9V promoted FUS ubiquitination (Supplementary Fig. 1f), explaining why dFUS-E9V failed to degrade FUS. Flow cytometric analyses revealed that FITC-labeled dFUS-E9C was efficiently delivered into cells (Supplementary Fig. 1g, h). Immunofluorescence images confirmed that FUS was substantially destroyed by dFUS-E9C (Fig. 1e, f). These data demonstrate that dFUS-E9C is an efficient FUS degrader.
a Mechanism of action of Oligo-PROTAC-mediated FUS degradation. b The RNA Oligos selected for Oligo-PROTAC synthesis and profiling. The FUS-recognition GGUG motif is marked in red. c, d Immunoblots (c) and quantitative analysis (d) of FUS in PC12 cells transfected with CRBN-based degrader (dFUS-…C) or VHL-based degrader (dFUS-…V) for 24 h. The samples derive from the same experiment, and the blots were processed in parallel. e, f Images (e) and quantitative analysis (f) of PC12 cells treated with dFUS-E9C (125 nM), followed by immunostaining with anti-FUS antibodies and DAPI. Scale bar, 20 μm. g Streptavidin pulldown assays to examine the binding of dFUS-E9C with FUS and CRBN. h MST assays to determine the affinity of dFUS-E9C to GFP and GFP-FUS. i Quantitative PCR to determine the FUS mRNA levels in PC12 cells treated with dFUS-E9C (125 nM) for 24 h. j Immunoblot analysis of FUS and GAPDH in SH-SY5Y cells treated with dFUS-E9C (125 nM) for 24 h with or without MG132 treatment. k Immunoblots of SH-SY5Y cells treated with dFUS-E9C (125 nM) together with excessive CRBNL. l Immunoblot analyses of FUS, CRBN, and GAPDH in SH-SY5Y cells treated with dFUS-E9C (125 nM) and siRNAs targeting CRBN. m HEK-293T cells were transfected with His-Myc-Ub followed by treatment with dFUS-E9C (125 nM) for 6 h. Cell lysates were subjected to immunoprecipitation with the FUS antibody, followed by immunoblotting with the indicated antibodies. The averages of n = 3 (d, f, h, i) biologically independent samples are shown. Data are shown as the mean ± SEM. Statistical significance in (d, f, i) was assessed using one-way ANOVA with multiple comparisons and two-tailed t tests, respectively. The data presented in (c, e, g, j–m) are representative of three independent experiments. Source data are provided as a Source Data file.
Molecular docking revealed that dFUS-E9C engages FUS and CRBN in a ternary complex (Supplementary Fig. 1i). In addition, there are multiple interacting sites between FUS and CRBN in the presence of dFUS-E9C, indicating the compatibility of FUS-CRBN pair (Supplementary Fig. 2). To confirm that FUS–dFUS-E9C–CRBN ternary complex was indeed formed in cells, we performed streptavidin pulldown experiments in which biotin-dFUS-E9C successfully enriched FUS and CRBN (Fig. 1g). This ternary complex was partially inhibited by competitive addition of dFUS-E9C, indicating the binding specificity (Fig. 1g). We then measured the dissociation constant (Kd) of dFUS-E9C for FUS by microscale thermophoresis (MST) assays, and dFUS-E9C showed high affinity to FUS (Kd = 60.54 nM) (Fig. 1h). Moreover, dFUS-E9C did not affect the levels of FUS mRNA, ruling out its effect on transcriptional regulation or mRNA degradation (Fig. 1i).
We next investigated that whether dFUS-E9C elicited FUS degradation through the ubiquitin-proteasome system. As expected, the proteasome inhibitor MG132 treatment abolished dFUS-E9C-mediated FUS degradation (Fig. 1j). In addition, pretreatment with excessive CRBNL (thalidomide) or siRNA-mediated gene silencing of CRBN reversed the degradation efficacy of dFUS-E9C, indicative of a CRBN-dependent manner (Fig. 1k, l). Consistently, ubiquitinated FUS was elevated upon dFUS-E9C treatment (Fig. 1m). Taken together, we have demonstrated that dFUS-E9C degrades FUS through CRBN-mediated ubiquitination and subsequent proteasomal degradation.
dFUS-E9C attenuates mitochondrial damage induced by FUS overexpression
Next, we sought to determine whether FUS degrader can reverse cellular damages caused by the abnormal accumulation of FUS. Abnormal FUS accumulation results in mitochondrial dysfunctions and ultimately neuronal cell death31,32 (Supplementary Fig. 3a). We first confirmed that dFUS-E9C was able to degrade cytoplasmic aggregates caused by FUS overexpression in neuron-like SH-SY5Y cells (Supplementary Fig. 3b). FUS overexpression increased the intracellular levels of reactive oxygen species (ROS) by 192% (Supplementary Fig. 3c), consistent with previous studies33,34. Treating SH-SY5Y cells overexpressing FUS with dFUS-E9C significantly reduced the ROS levels, and MG132 treatment abolished the beneficial effect of dFUS-E9C (Supplementary Fig. 3c). Meanwhile, MG132 alone did not affect ROS production (Supplementary Fig. 3c). We next measured the levels of lactate dehydrogenase (LDH), an important indicator of cell damage and death. dFUS-E9C treatment abolished FUS overexpression-induced LDH release, and the protective effect of dFUS-E9C was inhibited by MG132 (Supplementary Fig. 3d). Consistently, dFUS-E9C rescued cell death caused by FUS overexpression, and MG132 blocked the protective role of dFUS-E9C (Supplementary Fig. 3e). Since mitochondrial dysfunctions could potentially underlies neuronal damages, we proceeded to investigate whether dFUS-E9C restores mitochondrial membrane potential (ΔΨm) using mitochondrial membrane potential probes JC-1 and TMRE. FUS overexpression caused a significant decrease in ΔΨm compared to control; however, dFUS-E9C treatment rescued the ΔΨm to normal levels, and consistently, this protective effect was again blocked by MG132 treatment (Supplementary Fig. 3f–i). Collectively, using an in vitro neurotoxicity model, we demonstrate the protective role for dFUS-E9C in rescuing mitochondrial function and cell viability by degrading overexpressed FUS.
The dFUS-E9C engages the conserved RGG2 and RRM domains of FUS
To explore the molecular details of the interaction between FUS and dFUS-E9C, we tested the binding between dFUS-E9C and FUS truncation mutants (Fig. 2a), and found that the RGG2 domain of FUS is essential for dFUS-E9C binding (Fig. 2b). In addition, the RRM (RNA recognition motif) domain also contributes to the binding, as expected (Fig. 2b). Molecular simulation results confirmed that more nucleotides engage the RGG2 domain than the RRM domain. The seven arginine residues in the RGG2 domain predominantly interact with FUS-RNA#E9, forming hydrogen bonds with the RNA structure (Fig. 2c and Supplementary Fig. 4). Furthermore, we also identified a network of hydrogen bonds between the main chain of several glycine residues (G389, G404, G408, and G418) and nucleotides of the FUS-RNA#E9 in our complex model, which further stabilizes the complex structure (Fig. 2c, Supplementary Fig. 4, and Supplementary Table 1). Several residues on the beta-sheet structure of RRM domain also contributes to the interaction by forming hydrogen bonds, including N284, Y325, K334, and T338 (Fig. 2d, Supplementary Fig. 4, and Supplementary Table 1), consistent with the binding between dFUS-E9C with FUS truncation mutants (Fig. 2b). To confirm the predicted interactions between FUS and dFUS-E9C, we mutated the critical bases in dFUS-E9C (Supplementary Fig. 4 and Fig. 2e). As anticipated, changing the predicted bases in dFUS-E9C attenuated or lost the binding (Fig. 2f, g). Importantly, these amino acid residues are highly conserved in human and popular model species, such as mouse, rat, and cynomolgus monkey (Supplementary Fig. 5a). This allows easy preclinical assessment for efficacy and toxicity. As expected, dFUS-E9C was productive for degradation in different model cell lines from human, mouse, and rat, including PC12 (rat), SH-SY5Y (human), and Neuro-2a (mouse) cells, with a similar DC50 (Fig. 2h, i). FUS was rapidly degraded: over 50% of the protein was degraded at 6 h post dFUS-E9C treatment (Supplementary Fig. 5b, c). Moreover, no disease-causing mutations has been reported in the dFUS-E9C engaging residues, making dFUS-E9C broadly applicable to all disease-causing FUS mutants (postnatal deletion of wild-type FUS has been demonstrated safe).
a A summary of tested FUS truncation mutants and their affinity to dFUS-E9C. b Streptavidin pulldown assays to determine the binding capability between Biotin-dFUS-E9C and FUS truncations. c, d Predicted interactions of RRG2 (c) or RRM (d) domain with dFUS-E9C by molecular simulation. Red bases are predicted to be critical for the interaction between the Oligo and FUS. e The critical bases of FUS-RNA#E9 responsible for FUS binding were mutated and marked in red. f MST assays to determine the binding affinity of GFP-FUS to dFUS-E9C and its mutants. g Streptavidin pulldown assays to assess the interaction between FUS and Biotin-dFUS-E9 and its mutants. h, i Immunoblots (h) and quantitative analysis (i) of PC12, SH-SY5Y, and Neuro-2a cells transfected with increasing concentrations of dFUS-E9C for 24 h. The averages of n = 3 (f, i) biologically independent samples are shown. Data are shown as the mean ± SEM. The data presented in (b, g, h) are representative of three independent experiments. Source data are provided as a Source Data file.
The dFUS-E9C specifically degrades FUS and prefers cytoplasmic FUS aggregates
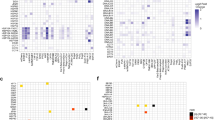
To determine the specificity of dFUS-E9C, we performed unbiased proteomic analyses. Only FUS was significantly degraded by dFUS-E9C (Fig. 3a). In contrast, the siRNA-mediated gene-silencing approach reduced FUS along with 7 other proteins (Fig. 3b). Six proteins were found to be up-regulated in both dFUS-E9C and siFUS treatment, suggesting the upregulation was caused by reduced FUS protein expression, not by dFUS-E9C or impaired CRBN function. Heatmap analysis and gene ontology revealed that dFUS-E9C-mediated changes of protein expression levels and biological processes are almost associated with FUS functions (Fig. 3c and Supplementary Fig. 6). Given that FUS mutations are the underlying cause of FTD/ALS and that they do not interfere dFUS-E9C binding15, we thus examined the degradation efficacy of dFUS-E9C on several common mutants, including P525L, R521C, R521G, and R521H. HEK-293T cells were transiently overexpressed with wild-type (WT) or FUS mutants, followed by treatment with different concentrations of dFUS-E9C. As expected, dFUS-E9C was able to degrade WT FUS and all the tested mutants, demonstrating the broad application of this degrader for FUS mutation carriers (Fig. 3d, e). We have noticed that the EC50 has shifted to ~0.5 μM, which is likely caused by the transient overexpression of FUS protein. Most of the disease-causing FUS mutations are enriched in the C-terminal NLS sequence12, suggesting that mislocalization underlies pathological protein aggregate formation. We tested whether cytoplasmic FUS was preferentially degraded. Strikingly, dFUS-E9C effectively degraded cytoplasmic GFP-FUSP525L, while nuclear GFP-FUSP525L was slightly degraded (Fig. 3f, g). Overexpression of WT and mutant FUS leads to protein aggregates15,16,35,36,37. Strikingly, the aggregated FUS was also remarkably inhibited by dFUS-E9C treatment (Fig. 3h, i). To decipher which form of FUS aggregates (amorphous or filamentous) are degraded by dFUS-E9C, we examined their assembly states using transmission electron microscopy (TEM) and immunostaining with the oligomer-specific antibody A11. Negative staining of A11 suggests that transiently overexpressed FUS and its mutants formed amorphous aggregates in cells, supported by the TEM results (Supplementary Fig. 7a–c). Streptavidin pulldown assays demonstrated that dFUS-E9C bound FUS variants as well as CRBN simultaneously (Fig. 3j) and promoted polyubiquitination of every single FUS variant (Fig. 3k). In conclusion, these findings demonstrate that dFUS-E9C preferentially degrades cytosolic FUS aggregates and can potentially be applied to all disease-causing FUS variants.
a, b Comparison of proteomic changes after treating neuron-like Neuro-2a cells with dFUS-E9C (a) or siFUS (b) for 24 h. Data are the mean of three biological replicates. c Heatmap showing proteins with significant changes between control and dFUS-E9C treatment. The overlapped proteins with significant changes in both dFUS-E9C and siFUS treatment are marked in red. d, e HEK-293T cells were transfected with GFP-FUS-WT, GFP-FUS-P525L, GFP-FUS-R521C, GFP-FUS-R521G, and GFP-FUS-R521H plasmids followed by transfecting with increasing concentrations of dFUS-E9C. Cell lysates were then subjected to immunoblotting (d), and the relative FUS levels were quantified by densitometry (e). f, g GFP-FUS-P525L-expressing HEK-293T cells were transfected with the indicated concentrations of dFUS-E9C, and the cytosol and nucleus compartments were isolated and subjected to immunoblotting (f). The relative FUS levels in the cytosol and nucleus were quantified by densitometry (g). h Immunofluorescence images of GFP-tagged wild-type FUS and disease-causing mutants in HEK-293T cells transfected with the indicated plasmids, followed by treating with dFUS-E9C (2 μM) for 24 h. Scale bar, 20 μm. i Quantification of the percentage of cells with GFP-FUS aggregates. j Streptavidin pulldown assays to determine the interactions between CRBN and FUS mutants in the presence or absence of dFUS-E9C. k HEK-293T cells were transfected with His-Myc-Ub plasmids together with GFP-FUS-WT, GFP-FUS-P525L, GFP-FUS-R521C, GFP-FUS-R521G, and GFP-FUS-R521H plasmids, followed by treatment with dFUS-E9C (2 μM) and MG132 (10 μM) for 24 h. Cell lysates were subjected to immunoprecipitation with the GFP antibody, followed by immunoblotting with the Myc and GFP antibodies. The averages of n = 3 (e, g, i) biologically independent samples are shown. Data are shown as the mean ± SEM. Statistical significance in (g, i) was assessed using two-tailed t tests. The data presented in (d, f, h, j, k) are representative of three independent experiments. Source data are provided as a Source Data file.
TfR-engaging DNA nanoflowers effectively deliver dFUS-PS9C across the BBB
Next, we examined whether dFUS-E9C could degrade FUS in vivo by infusing the indicated volume of dFUS-E9C (400 μM) into C57BL/6 mice through the lateral ventricle. The in vivo and ex vivo Cy5 signal was detected 12 h post injection (Supplementary Fig. 8a). As expected, dFUS-E9C reduced the FUS level in cortex and hippocampus in a dose-dependent manner (Supplementary Fig. 8b). To apply FUS degrader in vivo, we modified the nucleotide backbone of dFUS-E9C with phosphorothioate (hereafter referred to dFUS-PS9C) to increase its resistance to RNases. As expected, the serum stability of dFUS-PS9C was almost tripled compared to dFUS-E9C (Fig. 4a, b), while retaining strong efficacy on degrading FUS (Fig. 4c, d).
a, b Native PAGE analysis of the serum stability of FUS-RNA#E9, dFUS-E9C, and dFUS-PS9C (a), and the remaining Oligo levels at the indicated time points were quantified by densitometry (b). c, d Immunoblotting of FUS in SH-SY5Y cells transfected with increasing concentrations of dFUS-PS9C for 24 h (c), and the relative levels of FUS were quantified by densitometry (d). e Schematic showing the synthesis of DNA nanoflowers through rolling circle amplification and the generation of FRONTACFUS by functionalizing DNA nanoflowers with dFUS-PS9C. FRONTACFUS crosses the blood-brain barrier via TfR-mediated transcytosis. f Fluorescence spectral scanning of DNA nanoflowers (NF) and Cy5-labeled FRONTACFUS. g Dynamic light scattering to determine the particle size of NF and FRONTACFUS. h The loading efficiency of FRONTACFUS was determined by measuring the Cy5 signals. i Zeta potential analysis of dFUS-PS9C, NF, and FRONTACFUS. j Transmission electron microscopy (TEM) and scanning electron microscopy (SEM) images of NF and FRONTACFUS. Scale bars, 200 nm for TEM and 300 nm for SEM. k Immunofluorescence images of SH-SY5Y cells treated with PBS, Cy5-dFUS-PS9C, and Cy5-FRONTACFUS, followed by staining with LysoTracker (green) and DAPI (blue). Scale bar, 20 μm. l Immunoblotting analysis of FUS in SH-SY5Y cells treated with increasing concentrations of FRONTACFUS for 24 h, and the relative FUS levels were quantified by densitometry. m Serum stability determination of Cy5-dFUS-PS9C and Cy5-FRONTACFUS. n In vivo and ex vivo images of mice intravenously injected with PBS, sc-FRONTACFUS, and FRONTACFUS. The averages of n = 3 (b, d, h, i, m) biologically independent samples are shown. Data are shown as the mean ± SEM. The data presented in (a, c, f–n) are representative of three independent experiments. Source data are provided as a Source Data file.
Although Oligo-based medicines are popular in treating diseases in the central nervous system, a yet-to-be-addressed problem in Oligo-based therapy is efficient BBB-crossing delivery. Thus, patients have to suffer from invasive procedures, such as intrathecal injection, which are not widely available in common patient care practice. To overcome these issues, peripheral administration of BBB-crossing dFUS-PS9C would be ideal. To this end, we invented and established a strategy by using DNA nanoflower, a nucleic acid nanoparticle composed of hundreds of TfR-binding aptamers and dFUS-PS9C reverse complementary sequences, where dFUS-PS9C was loaded into the DNA nanoflower through Watson-Crick base pairing (Fig. 4e). We reasoned that the TfR-engaging DNA nanoflower (TfR-NF) can cross the BBB and deliver dFUS-PS9C to the brain via intravenous (i.v.) administration. Here we named DNA nanoflower incorporated with RNA Oligonucleotide-based PROTAC for FUS as FRONTACFUS.
FRONTACFUS enters the brain through TfR-mediated transcytosis, as demonstrated that TfR-NF was accumulated in cells compared to the scrambled nanoflowers (sc-NF), and TfR depletion remarkably reduced the entry of Cy5-TfR-NF, suggesting that TfR is required for effective internalization (Supplementary Fig. 9a–d). In addition, pretreatment with chlorpromazine (CPZ), an inhibitor of clathrin-dependent endocytosis, completely blocked the uptake of Cy5-TfR-NF and Cy5-sc-NF (Supplementary Fig. 9a–d). The gating strategy for cell sorting to detect Cy5 signals is shown in Supplementary Fig. 9e. Fluorescence spectroscopy confirmed that Cy5-labeled dFUS-PS9C was successfully loaded into the DNA nanoflowers (Fig. 4f). The particle size was increased from 186 nm to 292 nm after loading (Fig. 4g). As the mass ratio of DNA nanoflowers to Cy5-labeled dFUS-PS9C increased, the loading efficiency also increased, and reached 79.4% with the mass ratio of 5:1 (Fig. 4h). In addition, Zeta potential analyses also indicated the successful loading of dFUS-PS9C (Fig. 4i). TEM and scanning electron microscopy (SEM) confirm the success of nanoflower generation and that dFUS-PS9C loading did not affect the structure of nanoflowers while slightly increase the particle size, suggesting a successful loading of the degrader (Fig. 4j). We next examined the delivery efficacy of DNA nanoflowers by directly adding FRONTACFUS into the culture medium of SH-SY5Y cells. Compared with Cy5-dFUS-PS9C, FRONTACFUS rapidly entered SH-SY5Y cells (Fig. 4k) and efficiently degraded FUS with DC50 = 28 nM (Fig. 4l). Meanwhile, direct addition of dFUS-PS9C only led to minimal degradation of FUS even at 125 nM (Supplementary Fig. 9f). Interestingly, DNA nanoflowers further doubled the half-life of dFUS-PS9C in the serum from 36 h to over 72 h (Fig. 4m).
Next, we examined whether FRONTACFUS can be delivered into the brain via i.v. administration. We scrambled the TfR aptamer sequence, and generated a dummy DNA nanoflowers that loads similar amounts of dFUS-PS9C (sc-FRONTACFUS) as the negative control. BALB/c mice were i.v. administered with Cy5-FRONTACFUS or Cy5-sc-FRONTACFUS via tail vain injection. As expected, Cy5 signals were observed in the brain of FRONTACFUS-treated mice but not in that of sc-FRONTACFUS (Fig. 4n). The distribution of both nanoparticles in peripheral organs remains the same, indicating that the periphery uptake of FRONTACFUS is not TfR-dependent. Taken together, we demonstrated that FRONTACFUS successfully cross the BBB via TfR-mediated transcytosis.
FRONTACFUS rapidly degrades FUS in the brain
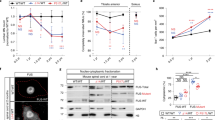
Next, we examined the degradation efficacy of FRONTACFUS in vivo. Mice were administered a single dose of FRONTACFUS via tail vein injection. Compared to Cy5-dFUS-PS9C (without a TfR engager), Cy5-labeled FRONTACFUS was enriched in the mouse brain starting at 1 h and peaked at 12 h post i.v. injection (Fig. 5a, b). Cy5 fluorescence in brain sections also confirmed the brain delivery of FRONTACFUS (Fig. 5c). Mice were treated (i.v. injection) with increasing concentrations of FRONTACFUS, and peripheral organs and brain tissues were harvested 24 h post injection. FUS in the hippocampus and cortex regions was significantly degraded at 10 mg/kg of FRONTACFUS (Fig. 5d–f). Immunohistochemistry and immunofluorescence images demonstrated that FRONTACFUS effectively degraded >50% ~ > 70% of total FUS protein in the hippocampus (CA1, CA2, and CA3), dentate gyrus (DG), and cortex regions (Fig. 5g, h and Supplementary Fig. 10a, b). Because FRONTACFUS were also enriched in the lung and liver, FUS in these organs was also degraded (Supplementary Fig. 10d, e). No obvious changes in neuron numbers in these brain regions from mice injected with Cy5-labeled FRONTACFUS compared to saline-injected controls (Fig. 5g and Supplementary Fig. 10a, c). The same was observed in peripheral organs by histological analyses (Supplementary Fig. 10f). In addition, because TfR was also ubiquitously expressed in immune cells and hemopoietic cells, we examined the toxicity of FRONTACFUS in these cells. FRONTACFUS did not affect the number of immune cells and hemopoietic cells (Fig. 5i, j and Supplementary Table 2). In conclusion, these results demonstrate that FRONTACFUS is a potent and safe degrader of FUS in the brain.
a In vivo and ex vivo images of mice intravenously injected with PBS, Cy5-dFUS-PS9C, and Cy5-FRONTACFUS. b Real-time monitoring of brain Cy5 signals in mice administered (i.v.) with Cy5-FRONTACFUS. c Fluorescence images of brain sections from mice receiving Cy5-dFUS-PS9C or Cy5-FRONTACFUS through i.v. injection. Scale bar, 50 μm. d Immunoblots of the hippocampus and cortex area in mice receiving i.v. injection of different doses of FRONTACFUS. e, f Immunoblots (e) and quantification (f) of the hippocampus and cortex area in mice receiving i.v. injection of NF control and FRONTACFUS (10 mg/kg). g, h Immunohistochemistry images (g) and quantification (h) of brain sections in mice receiving i.v. injection of NF control and FRONTACFUS. Scale bar, 50 μm. i, j The number of red blood cells (RBCs) and reticulocytes (RETs) in mice receiving i.v. injection of NF control and FRONTACFUS were determined. The averages of n = 3 (f, h–j) biologically independent samples are shown. Data are shown as the mean ± SEM. Statistical significance in (f, h–j) was assessed using two-tailed t tests. The data presented in (a–e, g) are representative of three independent experiments. Source data are provided as a Source Data file.
FRONTACFUS displays sustained efficacy with suboptimal neurotoxicity in vivo
To assess the sustained degradation effects and potential side effects of FRONTACFUS, mice were administered with a single dose of FRONTACFUS via tail vein injection, and hind-limb splay (a neurobehavioral indicator), body weight, and inflammatory effects were continuously monitored every other day for 2 weeks (Fig. 6a). Immunoblotting analysis showed that the levels of FUS in the hippocampus and cortex regions remained significantly reduced by approximately 60% 7 days post injection of FRONTACFUS, and the degradation persisted even on day 14, with a decrease of FUS around 20% to 40% (Fig. 6b, c), indicating the sustained degradation efficacy. Moreover, the FUS expression is restored between day 7 and day 14 (Fig. 6b, c), suggesting the FRONTACFUS-bearing neurons stayed healthy and functional. It is noted that none of the mice displayed hind-limb splay defects (Fig. 6d, e). There were no significant changes in body weights between the PBS control and the FRONTACFUS-treated groups (Fig. 6f). In addition, acute inflammatory response were only observed on day 2, which was quickly resolved at day 4 (Fig. 6g, h). Immunohistochemistry analyses confirmed that FRONTACFUS did not cause astrogliosis or microgliosis, signs of neuroinflammation (Fig. 6i, j), suggesting low immunogenic effects of FRONTACFUS in the brain. Next, we assessed neurotoxicity with a battery of histological staining (Supplementary Fig. 11a–f), and all of them revealed that FRONTACFUS did not affect the morphology and quantity of neurons in the cortex and hippocampus. Taken together, these results indicate that FRONTACFUS is a sustainable FUS degrader without noticeable neurological toxicity.
a Schematic of the administration of FRONTACFUS in BALB/c mice. Created in BioRender. Ge, R. (2025) https://BioRender.com/0t3564g. b, c The mice were intravenously administered with PBS control or FRONTACFUS (single dose, 10 mg/kg) for 7 days and 14 days, and the hippocampus and cortex were isolated for immunoblotting (b), and the levels of FUS were quantified by densitometry (c). d, e The representative hind-limb splay images (d) and statistical analysis (e) of mice after administration of PBS control or FRONTACFUS. f Measurement of body weight at the indicated intervals after administration of PBS or FRONTACFUS. g, h The serum was obtained at the indicated time points, and IL-6 (g) and TNF-α (h) concentrations were determined by ELISA. i, j Immunohistochemistry images (i) and quantification (j) of mouse brain sections stained with antibodies against GFAP and IBA1. Scale bar, 50 μm. The averages of n = 3 (c, e–h, j) biologically independent samples are shown. Data are shown as the mean ± SEM. Statistical significance in (c, f–h, j) was assessed using two-tailed t tests. The data presented in (b, d, i) are representative of three independent experiments. Source data are provided as a Source Data file.
Discussion
The BBB is a hurdle for drug delivery to the central nervous system. Although brain-shuttle and other BBB-crossing strategies leveraging antibodies or antibody fragments to target TfR or CD98 heavy chain (CD98hc) have achieved good BBB-crossing efficacy38,39, it remains not yet clinically available. Crosslinking Oligos to antibodies to achieve BBB-crossing delivery are compromised by the loading efficiency and downstream CMC (chemistry, manufacturing, and controls) problems. Some TfR-engaging antibodies cause severe anemia in animals and in clinical trials because reticulocytes express a high level of transferrin receptor9. Nucleic acid nanomaterials have good biocompatibility, programmability, and biodegradability and have important applications in biomedical fields. Nanoflowers synthesized by rolling-cycle amplification reaction have been widely used to deliver ASOs and interfering RNAs experimentally40,41. In this scenario, we designed a brain-penetrant FRONTACFUS which achieved efficient brain FUS degradation in vivo. FRONTACFUS not only solve the brain delivery issue but also improve the stability of the FUS-targeted Oligo, offering a general, modular, and straightforward approach to treating neurodegenerative diseases.
ALS and FTD are two deadly neurodegenerative diseases sharing many common risk factors and pathology. More than 50 genes have been identified as the cause of ALS and FTD diseases, among which mutations in C9orf72, superoxide dismutase 1 (SOD1), TDP-43, and FUS genes are the four most common types, accounting for over 70% of familial cases12. Using gene-silencing to suppress their toxicity has been regarded as a safe and effective strategy for treating ALS/FTD18,42. However, only intrathecal-administered tofersen, an ASO agent, has been recently approved for the treatment of SOD1 mutation-caused ALS, which only accounts for a small percentage of familial ALS cases43,44. Intrathecal injection also leads to poor patient compliance. PROTACs have shown prospects in treating neurodegenerative diseases because of its ability in handling “undruggable” targets. However, obvious challenges have halted the drug development, such as the targetability of the IDR of FUS and TDP-43 and the poor BBB-crossing property of PROTACs45,46,47. Herein, we established the FRONTAC strategy to address these challenges by successfully developing a selective Oligo-based FUS degrader and using a BBB-crossing nanoflower to achieve strong degradation efficacy in the mouse brain, suggesting great therapeutic potential of FRONTAC in treating neurodegenerative diseases caused by RNA-binding proteins.
Both experimental results and molecular docking data revealed that the engineered FUS-RNA#E9 primarily binds to FUS RGG2 domain and partially to the RRM domain. In line with this finding, it is reported that the RGG and RRM domains determine the binding specificity of FUS to FUS-RNA#930. The simultaneous engagement of two adjacent domains suggests that dFUS-E9C may function as an intramolecular bivalent degrader. Indeed, dFUS-E9C efficiently degraded FUS with a DC50 of ~25 nM, although this value was right-shifted in cells transiently expressing GFP-FUS, likely caused by FUS overexpression. In addition, we observed multiple interacting sites between the CRBN and FUS interaction surface in the presence of dFUS-E9C, which may further explain why CRBN-based but not VHL-based degrader effectively destroyed FUS, given that the proper pairing of E3 ligase with the POI is essential for degradation selectivity and efficiency48,49.
Given that the gene-silencing approach has been applied to patients carrying FUS mutations, we compare the off-target effect of dFUS-E9C with siRNAs. Unbiased mass spectrometry revealed that only FUS was significantly down-regulated by dFUS-E9C compared to eight proteins by siFUS, suggesting the superior selectivity of dFUS-E9C. The majority of disease-causing FUS mutants abolish nuclear localization and form aggregates in the cytoplasm37,50,51. Importantly, dFUS-E9C preferentially degrade cytoplasmic FUS aggregates and reduced cellular toxicity in an in vitro neurotoxicity model, indicating the therapeutic potential of dFUS-E9C. Although the formed cytoplasmic FUS aggregates in the cell model are in amorphous other than fibril-like state, FRONTACFUS has the potential to benefit patients with FUS-ALS by clearing pathological aggregates in neurons, due to the fact that ASO-mediated FUS mRNA silencing is effective in treating FUS mutation in ALS and reducing soluble protein supply may help the neurons to remove fibril-like aggregates18,52.
Methods
Cell culture and transfection
PC12 (#CRL-1721), Neuro-2a (#CCL-131), SH-SY5Y (#CRL-2266), and HEK-293T (#CRL-11268) cells were obtained from American Type Culture Collection (ATCC) and cultured in the indicated medium supplemented with 10% fetal bovine serum at 37 °C in 5% CO2, according to ATCC protocols. All cells were authenticated using short tandem repeat DNA fingerprinting and regularly examined for mycoplasma contamination. Oligo-PROTACs, siRNAs, and plasmids were transfected into cells using Lipofectamine 3000 (Thermo Fisher Scientific, Waltham, MA, USA) according to the manufacturer’s instructions. The CRBN-targeted siRNAs (#1: 5’-GUAGCUGCUUGUCUUCCUA-3’; #2: 5’-GAAGCACAGUUUGGAACAA-3’), FUS-targeted siRNA (5’-GTCCUAAUCCUACAUGUGAGA-3’), and TfR-targeted siRNA (5’-AACUUCAAGGUUUCUGCCAGC-3’) were obtained from the RiboBio Inc. (GuangZhou, China). Mammalian plasmid expressing FUS was obtained from the Public Protein/Plasmid Library (Nanjing, China), and its truncations were constructed via seamless cloning.
Oligo-PROTAC synthesis
Azide was modified on the 5’ end of RNAs and were synthesized from Beijing Genomics Institute (Beijing, China), Thalidomide-O-amido-propargyl was synthesized and characterized as previously described26. To generate Oligo-PROTACs, thalidomide-O-amido-propargyl was incubated with N3-RNA at a ratio of 10:1 in click reaction buffer (10 mM Tris-HCl, 10 mM CuSO4, 10 mM sodium erythorbate) at 37 °C for 4 h. The reaction product was then purified using a 3 K ultrafiltration membrane (Millipore, Burlington, MA, USA). The generated Oligo-PROTACs were separated by 15% native polyacrylamide gel electrophoresis (PAGE), stained with GelRed solution (UE Everbright, Suzhou, China), and imaged under UV illumination using an Amersham Imager 600 System (GE Healthcare, Pittsburgh, PA, USA).
Flow cytometry analysis
The uptake efficiency of Oligo-PROTACs was measured by flow cytometry. Different concentrations of FITC-dFUS-E9C were transfected into PC12 cells for the indicated time. Cells were then collected and washed twice with PBS, and resuspended in 1 ml of PBS. The FITC-positive cells were examined by a LSR Fortessa flow cytometer (Becton Dickinson, Mountain View, CA, USA), and the data were analyzed with FlowJo software.
Serum stability determination
FUS-RNA#E9, dFUS-E9C, and dFUS-PS9C (10 μM) were incubated in DMEM medium containing 10% fetal bovine serum at 37 °C for the indicated time, followed by separation with 15% natural PAGE, stained with a GelRed solution, and imaged under UV illumination using an Amersham Imager 600 System. For FRONTACFUS stability determination, Cy5-dFUS-PS9C or Cy5-FRONTACFUS (10 μM) were incubated in DMEM containing 10% fetal bovine serum for the indicated time points at 37 °C. The Cy5-FRONTACFUS was centrifuged at 10,625×g for 10 min and resuspended in the complete medium. The fluorescence intensity of Cy5-dFUS-PS9C and Cy5-FRONTACFUS was detected by a microplate reader (Biotek, Porsgrunn, Norway).
MST assays
The affinity of Oligo-PROTACs to GFP-FUS or GFP were determined by MST. The dFUS-E9C was serially diluted and mixed with GFP-FUS or GFP protein. After incubation for 10 min, the mixture was loaded into Monolith NTTM hydrophobic capillaries for measurement using a Monolith NT.115 instrument (NanoTemper Technologies, Munich, Germany). The Kd values were calculated using MO Affinity Analysis software (NanoTemper Technologies).
MD simulations
The modeling structures of the RRM and RGG2 domains of FUS were obtained from the AlphaFold protein structure database (UNP ID: P35637; amino acids: 283-432). The RNAComposer web server was utilized to build the structure of FUS-RNA#E9. Additionally, the structure of CRBN with CRBNL was obtained from the Protein Data Bank (PDB ID: 6RBN, Chain B). These structures were collectively used for subsequent simulations.
To obtain a rational structure and dynamic trajectories of the FUS–Oligo-PROTAC–CRBN ternary complex, multiple simulation methods were employed sequentially, ranging from local to global and from molecular docking to MD simulation. (1) Zdock software was utilized to predict the complex structure of FUS and FUS-RNA#E9, using the NMR structure of the FUS RRM domain and snRNA (PDB ID: 6SNJ) as a reference for RRM-RNA binding. The disordered region of the FUS protein, RGG2, and the remaining portion of the RNA were left free in the docking results to facilitate binding in subsequent MD simulations. A total of 1 μs MD simulation was conducted using Amber23 to optimize the interaction between RRM and the FUS-RNA#E9 5’-terminal structure, and to sample the stable complex state of the RGG2 domain and the 3’-terminal region of the FUS-RNA#E9. (2) Subsequently, with the stable complex structure of FUS/FUS-RNA#E9 obtained from step 1 and the CRBNL complex structure from the PDB, the triple structure for FUS–dFUS-E9C–CRBN was simulated using Zdock. Several triple complex conformations were selected to run 500 ns MD simulations to optimize the interactions of FUS with FUS-RNA#E9 and CRBNL. Subsequently, various conformations of the linker between FUS-RNA#E9 and CRBNL were constructed through MD simulation. The sampled linker conformations were matched to the triple complex conformations, with the best conformation selected by a custom Python ternary model prediction script. (3) Finally, 500 ns MD simulations were performed to validate the ternary interaction of the triple complex system.
Immunoblotting
Proteins were separated by 10% SDS-PAGE and transferred to polyvinylidene difluoride membranes (Millipore). The membrane was blocked with 5% fat-free milk in Tris-buffered saline supplemented with 0.1% Tween 20 (TBST) for 2 h, probed with primary antibody at 4 °C overnight, washed five times with TBST, and then probed with the horseradish peroxidase (HRP)-conjugated secondary antibody (Abways, Shanghai, China) at room temperature for 1 h. After washing five times with TBST, the protein on the membrane was visualized with an enhanced chemiluminescence substrate kit (Millipore) using an Amersham Imager 600 System (GE Healthcare). Primary antibodies used for immunoblotting were as follows (see Supplementary Data 1): anti-FUS (#11570-1-AP, Proteintech, Wuhan, China), anti-GAPDH (#AB0037, Abway), anti-Flag (#AB0008, Abways), anti-Myc (#AB0001, Abways), anti-GFP (#AB0045, Abways), anti-TfR (#CY6618, Abways), anti-PCNA (#AB0051, Abways), and anti-CRBN (#28494-2-AP, Proteintech).
Coimmunoprecipitation and streptavidin−biotin pulldown
Cells were lysed with RIPA cell lysis buffer supplemented with phenylmethanesulfonylfluoride (PMSF) and protease inhibitor cocktail. For streptavidin−biotin pulldown, biotin-modified dFUS-E9C was added to the cell lysates and incubated at 4 °C for 6 h. Subsequently, streptavidin agarose beads (#C006390, Sangon Biotech, Shanghai, China) were added and incubated at 4 °C overnight. The beads were washed six times with RIPA cell lysis buffer, boiled in SDS loading buffer for 10 min, and subjected to immunoblotting with the indicated antibodies. For coimmunoprecipitation, agarose beads (Santa Cruz Biotechnology, Dallas, TX, USA) conjugated with GFP or FUS antibodies were added to the cell lysates and incubated at 4 °C overnight. The beads were washed six times with the RIPA cell lysis buffer and boiled in SDS loading buffer for 10 min before immunoblotting with the indicated antibodies.
Immunofluorescence microscopy
Cells grown on coverslips were fixed with 4% paraformaldehyde, permeabilized with 0.5% Triton X-100, blocked with 4% bovine serum albumin, followed by probing with the primary antibodies and Alexa Fluor 488- or Alexa Fluor 568-conjugated secondary antibodies (Abcam, Cambridge, UK) and DAPI. Coverslips were mounted with 90% glycerol on a slide and imaged with a TCS SP8 confocal microscope (Leica Microsystems, Wetzlar, Germany). Primary antibodies used for immunostaining were as follows: anti-FUS (#sc-47711, Santa Cruz Biotechnology), anti-α Synuclein (#66412-1-Ig, Proteintech), and anti-oligomer A11 (#AHB0052, Invitrogen).
Intracellular ROS determination
The fluorescent probe 2’,7’-dichlorodihydrofluorescein diacetate (DCFH-DA) (#S0033, Beyotime Biotech, Shanghai, China) was used to measure cellular ROS. SH-SY5Y cells were transfected with FUS plasmids for 24 h followed by treating with PBS or Oligo-PROTACs in the presence or absence of MG132 for 24 h, followed by incubating with DCFH-DA for 20 min. After washing with PBS three times, the fluorescence signal in cells was detected using a microplate reader (Biotek) with an excitation/emission at 488/525 nm.
Mass spectrometry (MS) proteomics
Cells were treated as specified and lysed with RIPA lysis buffer. Proteins were digested in solution by trypsin (Promega, Madison, WI, USA) overnight at 37 °C. The resulting peptides were analyzed by LC-MS/MS using a nano-LC system (EASY-nLC 1200, Thermo Fisher Scientific) and an Orbitrap Eclipse Tribrid mass spectrometer (Thermo Fisher Scientific). The Swiss-Prot human proteome database in fasta format excluding protein isoforms (downloaded on 11.06.2023, 20,462 sequences) was used for direct data-independent acquisition (DIA) analysis with default settings. Standard settings in Spectronaut were used for quantification at the MS2 level. False discovery rate (FDR) thresholds were specified at 1% for PSMs, peptides, and protein groups. For quantitative comparisons, protein intensity values were log2-transformed.
Cell viability determination
SH-SY5Y cells were transfected with FUS plasmids for 24 h followed by treating with PBS or Oligo-PROTACs in the presence or absence of MG132 for the indicated time points. The viable cells were measured by a CCK-8 solution kit (Beyotime Biotech, #C0038) according to the manufacturer’s instructions. The absorbance value at 450 nm was detected by a microplate reader (Biotek).
LDH release assay
SH-SY5Y cells were transfected with FUS plasmids for 24 h followed by treating with PBS or Oligo-PROTACs in the presence or absence of MG132 for 24 h. Then, cells were incubated with LDH detection reagents (Beyotime, #C0016) for 30 min according to the manufacturer’s manual, and the absorbance value at 490 nm was detected by a microplate reader (Biotek).
JC-1 and TMRE staining
SH-SY5Y cells transiently overexpressing FUS were seeded into 24-well plates, followed by treatment with PBS or Oligo-PROTACs in the presence or absence of MG132 for 24 h. Cells were stained with JC-1 (Beyotime, #C2006) and TMRE (Beyotime, #C2001) kits according to the steps in the manual, and then subjected to DAPI staining, followed by examination with a TCS SP8 confocal microscope (Leica).
DNA nanoflower synthesis
All DNA sequences were synthesized by Beijing Genomics Institute (Beijing, China). The 5’ phosphate modified linear DNA template was annealed with the primer (see Supplementary Table 3) and then cyclized by T4 ligase (Vazyme, #C301) for 16 h at 16 °C, followed by heating at 65 °C for 10 min to inactivate the T4 ligase. The cyclized template was subjected to rolling circle amplification with Phi29 DNA polymerase (#E013, Novoprotein, Suzhou, China) at 30 °C for 3 h, followed by heating at 65 °C for 10 min to inactivate the DNA polymerase. The synthesized DNA nanoflower was centrifuged at 10,625 × g for 10 min, washed twice with ddH2O, and resuspended in ddH2O. The concentration was measured using NanoDrop (Thermo Fisher Scientific). To load Oligo-PROTACs, DNA nanoflower was mixed with Oligo-PROTACs in TM buffer (10 mM Tris-HCl, 12.5 mM MgCl2, pH 8), heated at 95 °C for 10 min, cooled down naturally to room temperature, centrifuged at 10,625×g for 10 min, and resuspended in ddH2O. For loading efficiency determination, DNA nanoflowers and Cy5-labeled Oligo-PROTACs were mixed in TM buffer at different mass ratios, centrifugated at 10,625×g for 10 min, and the fluorescence intensity was then measured with a microplate reader (Biotek).
DNA nanoflower characterization
The sizes of DNA nanoflowers with or without Oligo-PROTACs were measured by dynamic light scattering (DLS) using 12 mm square glass cuvette. The zeta potentials of DNA nanoflower with or without Oligo-PROTACs were measured by zeta potential analyzer (Malvern Panalytical Ltd, UK) using a disposable folding capillary cell. DNA nanoflower droplets were placed in a copper mesh and dried, the morphology of DNA nanoflower was visualized by a GeminiSEM transmission electron microscope (Carl Zeiss, Germany) and a HT-7800 scanning electron microscope (Hitachi Ltd, Japan).
Immunohistochemistry (IHC)
The 5-μm-thick paraffin tissue sections were deparaffinized with xylene and rehydrated through a graded alcohol series to distilled water, and the sections were subjected to HE staining or IHC staining. For IHC staining, the sections were probed in sequence with the primary antibody and HRP-conjugated anti-mouse secondary antibodies (Servicebio, Wuhan, China), followed by staining with 3,3’-diaminobenzidine tetrahydrochloride (DAB) (Servicebio). The coverslip was mounted and examined under a fluorescence microscope. Primary antibodies used for IHC were as follows: anti-FUS (#sc-47711, Santa Cruz Biotechnology), anti-GFAP (#GB12100-100, Servicebio), and anti-IBA1 (#GB15105-100, Servicebio).
ELISA
Serum samples obtained in animal experiments were subjected to ELISA assays to assess the pro-inflammatory factors using ELISA kits for mouse IL-6 (#PI326, Beyotime Biotech) and TNF-α (#PT512, Beyotime Biotech). The measurements were conducted in accordance with the manufacturer’s guidelines.
Real-time quantitative PCR (qPCR)
Total RNA was extracted using Trizol (#R401, Vazyme Inc., Nanjing, China). The 1 μg of purified RNA was reverse-transcribed using a HiScript III 1st Strand cDNA Synthesis Kit (Vazyme Inc.). Real-time qPCR was then performed by using SYBR kit (Vazyme Inc.) with the following primer sequences: GAPDH forward primer: 5’-GCAAGTTCAACGGCACAG-3’; GAPDH reverse primer: 5’-GCCAGTAGACTCCACGACAT-3’; FUS forward primer: 5’-GTAAGGCACCTAAGCCAGAT-3’; FUS reverse primer: 5’-CCCGATCATATCCTCCTCT-3’.
Animal experiments
Male 8-week-old BALB/C mice and C57BL/6 mice (Beijing Vital River Laboratory Animal Technology Co., Ltd, Beijing, China) were housed at 22 °C in air-conditioned rooms with a standard rodent and water feeding and controlled humidity (60% ± 10%). The mice were randomly allocated by blinded investigators to each group. For in vivo imaging, PBS, Cy5-sc-FRONTACFUS, and Cy5-TfR-FRONTACFUS were injected into BALB/C mice via the tail vein. The fluorescence images were monitored at the indicated time with a Syngene PXi imaging system (Synoptics Technologies Ltd., Cambridgeshire, UK). At the end of the time points, the mice were sacrificed, and the brains and various major organs (heart, liver, spleen, lung, and kidney) were isolated for ex vivo imaging.
For the pharmacodynamics study, NF control or FRONTACFUS (10 mg/kg) was injected into male 8-week-old BALB/C mice via the tail vein. Peripheral blood was harvested in an EDTA-anticoagulation tube for routine blood test with a hemocytometer. The mice were then sacrificed, and their brains and various major organs were isolated for further analysis. All procedures involving mice were performed in accordance with the relevant regulatory guidelines and were approved by the Animal Care and Use Committee of Shandong Normal University (ID: AEECSDNU2156).
Statistical analysis
All experiments were repeated independently at least three times. Data are expressed as mean ± standard error of the mean (SEM). GraphPad Prism v8.0 (GraphPad Software, La Jolla, CA, USA) was used for statistical analyses. Significant differences were identified using Student’s t test for pairwise comparisons and analysis of variance (ANOVA) for comparisons among multiple groups, with P values less than 0.05 considered statistically significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data are included in the main text and Supplementary Information or are available from the authors. Source data is available for Figs. 1–6 and Supplementary Figs. 1, 3, 5, 10, and 11 in the associated source data file. The FACS data have been deposited in FLOW Repository under the accession codes FR-FCM-Z9D8, FR-FCM-Z9D9, and FR-FCM-Z9DE. The mass spectrometry proteomics data have been deposited in ProteomeXchange Consortium via the PRIDE partner repository under the codes of PXD062670 and PXD062671. Source data are provided with this paper.
References
Luo, M. et al. Delivering the promise of gene therapy with nanomedicines in treating central nervous system diseases. Adv. Sci. 9, e2201740 (2022).
Kong, L., Wu, Y., Alves, C. S. & Shi, X. Efficient delivery of therapeutic siRNA into glioblastoma cells using multifunctional dendrimer-entrapped gold nanoparticles. Nanomedicine 11, 3103–3115 (2016).
Grafals-Ruiz, N. et al. Brain targeted gold liposomes improve RNAi delivery for glioblastoma. Int J. Nanomed. 15, 2809–2828 (2020).
Rofo, F. et al. A brain-targeting bispecific-multivalent antibody clears soluble Amyloid-beta aggregates in Alzheimer’s disease mice. Neurotherapeutics 19, 1588–1602 (2022).
Jagadeesan, N. et al. Modulation of hippocampal protein expression by a brain penetrant biologic TNF-α inhibitor in the 3xTg Alzheimer’s disease mice. J. Transl. Med. 22, 291 (2024).
Yu, D. et al. Multiplexed RNAi therapy against brain tumor-initiating cells via lipopolymeric nanoparticle infusion delays glioblastoma progression. Proc. Natl. Acad. Sci. USA 114, E6147–e6156 (2017).
Cohen, Z. R. et al. Localized RNAi therapeutics of chemoresistant grade IV glioma using hyaluronan-grafted lipid-based nanoparticles. ACS Nano 9, 1581–1591 (2015).
Tian, G. et al. Engineered design of theranostic upconversion nanoparticles for tri-modal upconversion luminescence/magnetic resonance/X-ray computed tomography imaging and targeted delivery of combined anticancer drugs. J. Mater. Chem. B 2, 1379–1389 (2014).
Gadkar, K. et al. Mathematical PKPD and safety model of bispecific TfR/BACE1 antibodies for the optimization of antibody uptake in brain. Eur. J. Pharm. Biopharm. 101, 53–61 (2016).
Van Langenhove, T. et al. Genetic contribution of FUS to frontotemporal lobar degeneration. Neurology 74, 366–371 (2010).
Sengupta, U. & Kayed, R. Amyloid β, Tau, and α-Synuclein aggregates in the pathogenesis, prognosis, and therapeutics for neurodegenerative diseases. Prog. Neurobiol. 214, 102270 (2022).
Mackenzie, I. R., Rademakers, R. & Neumann, M. TDP-43 and FUS in amyotrophic lateral sclerosis and frontotemporal dementia. Lancet Neurol. 9, 995–1007 (2010).
Seelaar, H. et al. Frequency of ubiquitin and FUS-positive, TDP-43-negative frontotemporal lobar degeneration. J. Neurol. 257, 747–753 (2010).
An, H. et al. ALS-linked FUS mutations confer loss and gain of function in the nucleus by promoting excessive formation of dysfunctional paraspeckles. Acta Neuropathol. Commun. 7, 7 (2019).
Nolan, M., Talbot, K. & Ansorge, O. Pathogenesis of FUS-associated ALS and FTD: insights from rodent models. Acta Neuropathol. Commun. 4, 99 (2016).
Sabatelli, M. et al. Mutations in the 3’ untranslated region of FUS causing FUS overexpression are associated with amyotrophic lateral sclerosis. Hum. Mol. Genet. 22, 4748–4755 (2013).
Mitchell, J. C. et al. Overexpression of human wild-type FUS causes progressive motor neuron degeneration in an age- and dose-dependent fashion. Acta Neuropathol. 125, 273–288 (2013).
Korobeynikov, V. A., Lyashchenko, A. K., Blanco-Redondo, B., Jafar-Nejad, P. & Shneider, N. A. Antisense oligonucleotide silencing of FUS expression as a therapeutic approach in amyotrophic lateral sclerosis. Nat. Med. 28, 104–116 (2022).
Li, K. & Crews, C. M. PROTACs: past, present and future. Chem. Soc. Rev. 51, 5214–5236 (2022).
Yang, Z., Pang, Q., Zhou, J., Xuan, C. & Xie, S. Leveraging aptamers for targeted protein degradation. Trends Pharm. Sci. 44, 776–785 (2023).
Tsai, J. M., Nowak, R. P., Ebert, B. L. & Fischer, E. S. Targeted protein degradation: from mechanisms to clinic. Nat. Rev. Mol. Cell Biol. 25, 740–757 (2024).
Qu, J. et al. Specific knockdown of α-Synuclein by peptide-directed proteasome degradation rescued its associated neurotoxicity. Cell Chem. Biol. 27, 751–762.e754 (2020).
Ge R. X. et al. Targeting neurodegenerative disease-associated protein aggregation with proximity-inducing modalities. Acta Pharmacol. Sin. https://doi.org/10.1038/s41401-025-01538-2 (2025).
Pandit, R., Chen, L. & Götz, J. The blood-brain barrier: physiology and strategies for drug delivery. Adv. Drug Deliv. Rev. 165-166, 1–14 (2020).
Yi, X., Manickam, D. S., Brynskikh, A. & Kabanov, A. V. Agile delivery of protein therapeutics to CNS. J. Control Release 190, 637–663 (2014).
Kong, L. et al. An engineered DNA aptamer-based PROTAC for precise therapy of p53-R175H hotspot mutant-driven cancer. Sci. Bull. 69, 2122–2135 (2024).
Chen, M. et al. Inducible degradation of oncogenic nucleolin using an aptamer-based PROTAC. J. Med. Chem. 66, 1339–1348 (2023).
Liu, J. et al. TF-PROTACs enable targeted degradation of transcription factors. J. Am. Chem. Soc. 143, 8902–8910 (2021).
Shao, J. et al. Destruction of DNA-binding proteins by programmable oligonucleotide PROTAC (O’PROTAC): Effective targeting of LEF1 and ERG. Adv. Sci. 8, e2102555 (2021).
Lerga, A. et al. Identification of an RNA binding specificity for the potential splicing factor TLS. J. Biol. Chem. 276, 6807–6816 (2001).
Tsai, Y. L. et al. ALS/FTD-associated protein FUS induces mitochondrial dysfunction by preferentially sequestering respiratory chain complex mRNAs. Genes Dev. 34, 785–805 (2020).
Deng, J. et al. FUS interacts with HSP60 to promote mitochondrial famage. PLoS Genet. 11, e1005357 (2015).
Kumar, R. et al. Amyotrophic lateral sclerosis risk genes and suppressor. Curr. Gene Ther. 23, 148–162 (2023).
Wang, Y. W. et al. HIF-1α-regulated lncRNA-TUG1 promotes mitochondrial dysfunction and pyroptosis by directly binding to FUS in myocardial infarction. Cell Death Discov. 8, 178 (2022).
Mackenzie, I. R. et al. Distinct pathological subtypes of FTLD-FUS. Acta Neuropathol. 121, 207–218 (2011).
Deutschländer, A. B., Ross, O. A., Dickson, D. W. & Wszolek, Z. K. Atypical parkinsonian syndromes: a general neurologist’s perspective. Eur. J. Neurol. 25, 41–58 (2018).
Dormann, D. et al. Arginine methylation next to the PY-NLS modulates transportin binding and nuclear import of FUS. EMBO J. 31, 4258–4275 (2012).
Niewoehner, J. et al. Increased brain penetration and potency of a therapeutic antibody using a monovalent molecular shuttle. Neuron 81, 49–60 (2014).
Chew, K. S. et al. CD98hc is a target for brain delivery of biotherapeutics. Nat. Commun. 14, 5053 (2023).
Kim, M. G., Park, J. Y., Shim, G., Choi, H. G. & Oh, Y. K. Biomimetic DNA nanoballs for oligonucleotide delivery. Biomaterials 62, 155–163 (2015).
Esteban-Fernández de Ávila, B. et al. Acoustically propelled nanomotors for intracellular siRNA delivery. ACS Nano 10, 4997–5005 (2016).
McCampbell, A. et al. Antisense oligonucleotides extend survival and reverse decrement in muscle response in ALS models. J. Clin. Invest 128, 3558–3567 (2018).
Miller, T. M. et al. Trial of antisense oligonucleotide Tofersen for SOD1 ALS. N. Engl. J. Med. 387, 1099–1110 (2022).
Benatar, M. et al. Design of a randomized, placebo-controlled, Phase 3 Trial of Tofersen initiated in clinically presymptomatic SOD1 variant carriers: the ATLAS Study. Neurotherapeutics 19, 1248–1258 (2022).
Tashima, T. Proteolysis-targeting chimera (PROTAC) delivery into the brain across the blood-brain barrier. Antibodies 12, 43 (2023).
Mohanty, P. et al. A synergy between site-specific and transient interactions drives the phase separation of a disordered, low-complexity domain. Proc. Natl. Acad. Sci. USA 120, e2305625120 (2023).
Patel, A. et al. A liquid-to-solid phase transition of the ALS protein FUS accelerated by disease mutation. Cell 162, 1066–1077 (2015).
Liu, Y. et al. Expanding PROTACtable genome universe of E3 ligases. Nat. Commun. 14, 6509 (2023).
Yang, Z. et al. Identification of a non-inhibitory aptameric ligand to CRL2(ZYG11B) E3 ligase for targeted protein degradation. Nat. Commun. 16, 2494 (2025).
Neumann, M. et al. A new subtype of frontotemporal lobar degeneration with FUS pathology. Brain 132, 2922–2931 (2009).
Munoz, D. G. et al. FUS pathology in basophilic inclusion body disease. Acta Neuropathol. 118, 617–627 (2009).
DeVos, S. L. et al. Tau reduction prevents neuronal loss and reverses pathological tau deposition and seeding in mice with tauopathy. Sci Transl Med. 9, eaag0481 (2017).
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (32270892 to S.X., 32200613 to M.C., and 32470811 to J.L.).
Author information
Authors and Affiliations
Contributions
H.Y., J.Z. and S.X. supervised the project and designed the experiments. R.G., M.C., S.W., P.Z., M.H.C., F.Z., J.Z.Z., Y.Z., J.L., G.N., Z.Y. and L.Z. performed the experiments. S.H., Q.L., W.P., M.L., C.X., H.Y., J.Z. and S.X. analyzed the data. H.Y., J.Z. and S.X. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
J.Z. and S.X. declare the filing of a patent application for the FRONTACFUS technology. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Zhuo Tang, Guizhi Zhu, and the other, anonymous, reviewer for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ge, R., Chen, M., Wu, S. et al. DNA nanoflower Oligo-PROTAC for targeted degradation of FUS to treat neurodegenerative diseases. Nat Commun 16, 4683 (2025). https://doi.org/10.1038/s41467-025-60039-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-60039-2
This article is cited by
-
Nuclear ribonucleoprotein condensates as platforms for gene expression regulation
Genes & Genomics (2025)