Abstract
Cytoplasmic pattern recognition receptors (PRR) for double-stranded RNA, such as RIG-I/MDA5, are key mediators of anti-viral responses. Here we screen for synergistic drug-virotherapy combinations and find that the reovirus type III Dearing strain (Rt3D)-palbociclib combination augments oncolytic virus-induced stress responses and increases interferon production and signaling. Data from RIG-I agonist and ER stress-inducing agents further confirms the crosstalk between RNA-sensing and ER stress in inducing cancer cell death and interferon production. Combined Rt3D-palbociclib also increases innate immune activation and IFN-induced HLA expression within tumor cells, with accompanying alterations in the epigenetic landscape and endogenous retroviral (ERV) elements. Analysis of the immunopeptidome in treated cells further reveals changes to HLA-captured peptides, including altered expression of peptides from cancer or testis antigens and ERVs. Our findings thus highlight the crosstalk between stress signaling and PRR activation for mediating enhanced anti-cancer efficacy.
Similar content being viewed by others
Introduction
The innate immune systems plays a crucial role in cancer therapy, detecting tumour-associated molecular patterns. However, cancers often evade immune surveillance by exploiting immune-evasion mechanisms which allow the cancer to grow and metastasize. Oncolytic viruses (OV) are a promising new field of cancer therapy that can potently activate PRRs for dsRNA (RIG-I/MDA-5) and dsDNA (STING) to induce anti-tumour innate and adaptive immune responses. These biological self-amplifying agents have been shown to be cancer cell tropic, kill through multiple mechanisms, and have the potential to promote anti-tumour responses1. Pelareorep is an oncolytic virus derived from a proprietary isolate of the reovirus type III Dearing strain (Rt3D). It has been granted orphan drug status by the FDA for malignant glioma, ovarian and pancreatic cancer and is actively under clinical investigation.
Rt3D is a non-enveloped dsRNA virus with a cytoplasmic life cycle. Despite this, Rt3D can induce endoplasmic reticulum (ER) stress2,3,4, due to remodelling of ER membranes as part of the formation of viral inclusions5,6. ER stress activates signalling via the unfolded protein response (UPR) via the three effector pathways IRE1α-XBP1, PERK-eIF2a and ATF67. Increasing the anti-cancer efficacy of Rt3D through manipulation of ER stress/UPR/ISR signalling has been established in a number of studies2,4,8,9.
We sought to establish synergistic combinations by screening Rt3D against 80 small molecules in melanoma. The CDK4/6 inhibitor palbociclib was a lead hit, with combination efficacy validated in vitro and in vivo. RNAseq and proteomics approaches indicated palbociclib in combination with Rt3D induced a pronounced ER stress / UPR activation signature.
Endogenous retroviral elements (ERVs) are remnants of ancient viral infections that have integrated into host germline DNA. In cancer, ERVs are often dysregulated and can be activated through treatments that modify the genome such as DNA methylation and histone modifications. The activation of ERVs in cancer may be therapeutic by triggering immune responses and generating neoantigens. Palbociclib potently increased interferon signalling and ERV transcripts in response to Rt3D. Knockdown studies indicated key UPR proteins and the RNA sensor RIG-I, were essential to the phenotype observed. Mechanistically independent experiments indicated ER stress boosted cancer cell death and interferon production in response to RNA sensor agonists.
Here we describe a therapeutically exploitable anti-cancer interaction between UPR activation and cytoplasmic RNA sensing, through the combination of palbociclib and Rt3D. This combination bolsters cell death, interferon production and immune activation, whilst also alters DNA modification and ERV elements. This study thus implicates palbociclib as a sensitiser for both ER stress and dsRNA-induced PRR signalling and offers new combinatorial opportunities for the development of cancer treatment.
Results
Palbociclib enhances anti-cancer efficacy of reovirus Type 3 Dearing (Rt3D) and the viral defence response
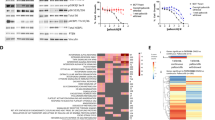
We performed a high-throughput small molecule screen using a non-biased approach to seek synergistic drug-virus interactions. The A375 BRAFV600E-mutant melanoma cell line was selected for screening due to the susceptibility of melanoma to Rt3D infection4. Compounds used are shown (Fig. S1A). The CDK4/6 inhibitor (CDK4/6i), palbociclib, enhanced cytotoxicity across a range of viral doses (Figs. 1A and S1B). Palbociclib-mediated CDK4/6 inhibition repressed phospho-retinoblastoma protein (Rb) in RB-wild-type cells, inducing G1/S cell-cycle arrest10. This was corroborated in A375 and other melanoma cell lines (Fig. S1C). To validate findings from the screen, the Rt3D-palbociclib combination was tested with orthogonal methods in A375 and other melanoma, the murine BRAF-mutant melanoma cell line 4434, head and neck and breast cancer cells, including those with acquired palbociclib resistance (Figs. 1B–D and S1D–I). The requirement for intact Rb signalling was shown in A2058 RB-null cells, in which palbociclib failed to mediate G1 arrest or augment Rt3D-induced cytotoxicity (Fig. S1J, K).
A A375 were treated with Rt3D (MOI 1) and 80 compounds at various doses. Low Z scores (below −2, dashed line) indicate sensitisers (palbociclib is shown in red, with relative Z scores that represent PD-0332991 doses 1 μM (−2.041) and PF-332991 doses at 1 μM (−5.076) and 0.5 μM (−5.570)). B Cell viability in A375 cells treated with Rt3D ± palbociclib (1 μM) by MTT (left) and crystal violet (right) 72 h after treatment (data are presented as mean values ± SEM, n = 3 biologically independent experiments). C Cell viability in melanoma cells MeWo (BRAF/RAS wild-type), and DO4 (Ras mutant) treated with Rt3D ± palbociclib (1 μM) by MTT (72 h). D Cell viability measured at 72 h by MTT assay in 4434 cells treated ∓ palbociclib (1 μM) (data are presented in (C, D) as mean values ± SEM, n = 3 biologically independent experiments). E Volcano plot of RNA sequencing showing upregulated and downregulated genes for A375 cells treated with palbociclib (1 μM), Rt3D (MOI 0.1) or the combination compared to basal at 48 h. Highlighted genes in red belong to the GSEA set ‘reactome interferon signalling’. F RT-qPCR of IFNa, IFNb, IL-28, IL-29, ISG15 and OASL in A375 cells treated with Rt3D ± palbociclib (1 μM) at 48 h. Data presented are mean values ± SEM, n = 3 biologically independent experiments. P values were determined by one-way ANOVA corrected for multiple comparisons. G RT-qPCR of mouse IFNα and IFNβ in murine BRAF-mutant melanoma 4434 cells treated with Rt3D ± palbociclib (1 μM) at 48 h. Data are presented as mean values ± SEM, n = 3 biologically independent experiments. P values were determined by one-way ANOVA corrected for multiple comparisons. H Proteomic analysis of A375 cells treated with Rt3D (0.1) in combination with palbociclib (1 μM) reveal upregulated (red) and downregulated (blue) proteins categorised under the GO term ‘defence response to virus’ (24 h, data are from 1 independent experiment). I IFNα and IFNβ, measured in cell-free supernatant from Rt3D-treated samples (MOI 0.05-0.5) ± palbociclib (1 μM) by ELISA. Results from 2 independent experiments. The lines joining the open and closed circles indicate the data points derived from the same experiment. J Tumour volumes for animals bearing A375 tumours treated with either a sham injection with vehicle (n = 13), an intra-tumoural injection of Rt3D (1 × 106 pfu, n = 11), palbociclib (n = 10), or the combination (n = 9). Tumour volumes for C57BL/6 mice bearing murine BRAF-mutant 4434 tumours were treated with a sham injection with vehicle (n = 6), an intra-tumoural injection of Rt3D (1 × 106 pfu, n = 9), palbociclib (n = 6), or the combination (n = 8). Data are presented as mean values ± SEM, and p values were derived from one-way ANOVA adjusted for multiple comparisons from area under curve values from individual mice, where p > 0.05 ns, *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001. Source data are provided as a Source Data file.
To probe mechanistic interactions between Rt3D and palbociclib, we analysed the transcriptome and the proteome in A375. At the transcriptomic level, data indicated a more pronounced interferon signature in tumour cells with combination treatment (Fig. 1E). These data were corroborated in separate RT-qPCR experiments for type I IFNα, IFNβ, type III IL-28 and IL-29 and IFN-stimulated genes ISG15 and OASL (Fig. 1F). Increased mouse Ifnα was observed for the mouse melanoma cell line (4434) with Rt3D-palbociclib combination therapy, despite the unremarkable effect on cell kill (Fig. 1G). At the proteomic level, proteins detected within the GO term ‘defence response to virus’ were upregulated in the Rt3D-palbociclib combination treatment in the human A375 melanoma line (Fig. 1H). To confirm that the observed IFN expression translated to IFN protein, an ELISA was performed for IFNα/β and IL-28/29 (Figs. 1I and S1L). Efficacy of Rt3D-palbociclib combination therapy was tested in vivo using human A375 tumour-bearing CD1 nude mice and in an immunocompetent model bearing the murine 4434 BRAF-mutant melanoma (Fig. 1J).
Rt3D-palbociclib-induced interferon signalling increases antigen-processing machinery and immunogenicity
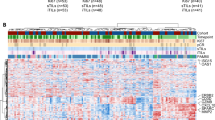
Proteomic analyses of proteins with GO terms ‘immune response’, ‘interferon regulation/response’ and ‘antigen-processing and presentation’ showed increases in the expression of class I and II HLA-related proteins with Rt3D-palbociclib, that were also evident in transcriptomic analysis (Figs. 2A and S2A–C). FACS analysis of non-permeabilized cells confirmed cell surface expression of HLA class I and class II (Figs. 2B and S2D), which was reduced by ruxolitinib-mediated inhibition of IFN signalling (Figs. 2C and S2E). Analysis of proteasome subunits showed that Rt3D induced a change from PSMB 1-6 subunits to incorporate low molecular weight proteins PSMB8 (LMP7), 9 (LMP2) and 10 (MECL-1). These subunit replacements convert the standard proteasome into an immunoproteasome. Additionally, changes were observed in transporter associated with antigen processing (TAP) and ERAP proteins (Figs. 2D and S2F). Given these changes, and the fact that Rt3D-palbocilib increases cell surface class I HLA expression, we performed immunopeptidomic analyses to look for evidence of treatment-related modulation of peptide presentation. We focused on potential tumour-associated antigens and showed changes in presentation of multiple cancer/testis antigens (CTA) with Rt3D-palbociclib treatment (Fig. 2E). A total of 63 CTA immunopeptides from 44 genes were identified, many showing increased expression upon combination treatment. In particular, Rt3D-palbociclib increased the expression of CTA derived from the protein MAGEA4. MAGEA4 mRNA expression levels were not altered by Rt3D-palbociclib treatment, nor by disruption to IFN signalling by the JAK/STAT inhibitor, ruxolitinib. However, protein levels dropped with treatment, suggesting increased turnover by the proteasome (Fig. 2F).
A Proteomic analysis of A375 cells treated with Rt3D (0.1–1) ± palbociclib (1 μM) reveal upregulated (red) and downregulated (blue) proteins categorised under the GO term ‘antigen processing and presentation’ (48 h, data are from 1 independent experiment). B Expression of HLA-A,B,C (class I) and HLA-DR,DP,DQ (class II) in A375 cells treated with Rt3D ± palbociclib by FACS analysis. Data are representative of 3 independent biological experiments (shown in Fig. S2D). C Expression of HLA-A,B,C (class I) and HLA-DR,DP,DQ (class II) in A375 cells treated with Rt3D plus palbociclib ± ruxilitinib (RUX). Data are representative of 2 independent biological experiments (shown in Fig. S2E). D Proteomic analysis of A375 cells treated with Rt3D (0.1–1) ± palbociclib (1 μM) reveal upregulated (red) and downregulated (blue) proteosome subunits. PSMB8, 9 and 10 (highlighted) are incorporated into the proteosome to switch the standard proteasome into an immunoproteasome (24 h, above, 48 h, below, data are from 1 independent experiment for each time point). E Fold change in CTA petides from HLA-I with Rt3D-palbociclib versus basal conditions. Peptides belonging to the MAGEA4 protein are highlighted (1–8). Data represent 1 independent experiment. F Proteomic analysis of A375 cells treated with Rt3D (0.1–1) ± palbociclib (1 μM) reveal upregulated (red) and downregulated (blue) MAGEA4 protein (48 h, top). RT-qPCR of MAGEA4 in A375 cells treated with Rt3D ± palbociclib, ∓ruxilitinib (RUX) (1 μM) at (48 h). Proteomics data represent 1 independent experiment and PCR data are presented as mean values ± SEM, n = 3 independent biological experiments. G ATP release by cell titre glo assay in supernatants from A375 cells treated with indicated Rt3D doses ∓ palbociclib (1 μM). Data are presented as mean values ± SEM, n = 3 independent biological experiments. P values were determined by a two-way ANOVA with adjustments made for multiple comparisons. H HMGB1 release by western blot of supernatants of treated A375 cells. Data are representative of n = 2 independent biological experiments (available in source file). I Calreticulin (CRT, white arrows) cell surface expression by confocal microscopy of treated A375 cells, quantified for 3 fields of view per experiment, for 3 independent biological repeats. J Pictomicrographs of macrophages stained with CD11b-FITC co-cultured with treated, phrodo-stained tumour cells (red), showing engulfed (yellow arrows) and non-engulfed (white arrows) cells, enlarged examples are shown on the left. Percentages of macrophage population that dual-stain for phrodo are shown. Data are presented as mean values ± SEM, n = 3 independent biological experiments. P values were determined by ANOVA with adjustments made for multiple comparisons. P values were derived where p > 0.05 ns, *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001. Source data are provided as a Source Data file.
Next, we looked for Rt3D-palbociclib-induced tumour cell-intrinsic adjuvanticity through immunogenic cell death (ICD). ICD has been previously described for single-agent CDK4/6 inhibition11,12. Surrogate ICD markers (ATP and HMGB1 release) were increased following combined Rt3D-palbociclib treatment relative to either agent alone (Fig. 2G, H). The marked reduction in ATP in the supernatant at MOI 1 most likely reflected almost complete cell death at that viral dose. Surface expression of calreticulin (CRT) is also a surrogate marker of ICD. Unpermeabilised cells stained with CRT were imaged by confocal microscopy. Surface staining of unpermeabilised cells with anti-CRT showed areas of green fluorescence that were brighter and covered a larger area in the combination therapy. These levels of CRT were comparable with thapsigargin, which activates the UPR though inhibition of ER calcium pumps (Fig. 2I). Given these observations of increased antigen processing/presentation and ICD, which serve as ‘find me’ and ‘eat me’ signals13, we next tested if this resulted in elevated phagocytosis by cultured immune cells. CD11b-FITC-labelled PBMC-derived human macrophages were co-cultured with tumour cells that had prior staining with pHrodo, a dye that fluoresces in acidic environments, such as phagolysosomes14. Macrophage engulfment of Rt3D-infected tumour cells was significantly enhanced by palbociclib, measured by dual-staining (Figs. 2J and S2G). Taken together, these results support the notion that Rt3D-palbociclib treatment of tumour cells may increase activation of both innate and adaptive anti-tumour immunity.
Rt3D-palbociclib increases immune cell activation status
To develop this idea, we next investigated the infiltration of T cells and myeloid cells following Rt3D-palbociclib treatment in an immunocompetent model in vivo. Tumour-infiltrating CD4+, CD8+ and NK cells were analysed after a single intra-tumoural injection of Rt3D into 4434 tumours. Data revealed significantly increased frequencies of CD69 and/or Granzyme B-positive cells with combination therapy relative to their single-agent counterparts (Figs. 3A and S3A). It has been reported that the NK cell receptor, NKp46, recognises externalised calreticulin15. Given the effects on ICD markers, we next investigated the impact of this treatment on DC activation. We used a Kaede mouse model system to track immune cells in the tumour via their photoconversion from Kaede green to Kaede red16,17,18. Furthermore, we multiplexed this experiment with phagocytic assays in vivo using the mCherry-expressing 4434 tumour model. Our analysis segregated 4 populations: unlabelled, antigen-free DCs (negative staining for both Kaede red and mCherry); DCs from tumour origin (Kaede red+); tumour antigen-loaded DCs (mCherry+ DCs); and DCs from tumour origin, also loaded with tumour antigen (Kaede red+ and mCherry+) (Figs. 3B, and S3B, C). In tumour-draining lymph nodes, the activation status of type I and type II conventional DCs (cDC1, cDC2) was analysed for all 4 populations. Whilst no changes were observed for CD80, Rt3D-palbociclib combination therapy led to increased CD86 and CD40 co-stimulatory receptors, and PD-L1, in Kaede red populations relative to either single-agent, in cDC1 (Fig. 3C). In cDC2, again, no changes were observed for CD80, however, Rt3D-palboiclib combination therapy lead to increased CD86, in Kaede red and mCherry populations relative to either single-agents (Fig. 3C). In the tumour, both CD86 and PD-L1 were increased with Rt3D-palbociclib over their single-agent counterparts in mCherry + cDC1s (Fig. 3D). In tumour cDC2s, the effects were less marked. Taken together, we found clear evidence of immune response triggered by the combination therapy in tumours when compared with single-agent groups. This may warrant assessment of Rt3D-palbociclib in additional combination with anti-PD-1 therapy in future, more primarily immune-focused studies.
A Mice bearing 4434 tumours were treated with sham injection with vehicle (n = 6), palbociclib (100 mg/kg daily by oral gavage, n = 5), a single injection of 5 × 106 pfu Rt3D (n = 6), or the combination (n = 6). Tumours were harvested at day 6 after injection and TILs stained for activation markers. Data show proportions of CD69+ or GzmB+ cells in CD8, CD4 T or NK cells. A one-way ANOVA was used with the p value corrected for multiple comparisons. B Kaede mice bearing 4434-mcherry tumours were treated with palbociclib (100 mg/kg daily by oral gavage, blue arrows) followed by a single injection of 5 ×106 pfu Rt3D. The following day, tumours were photoconverted and, 48 h later, collected and stained for activation markers. Data are shown for 4 gated populations as shown: unlabelled, antigen-free DCs (negative staining for both kaede red and mCherry); DCs from tumour origin (kaede red+); tumour antigen-loaded DCs (mCherry+ DCs); and DCs from tumour origin, also loaded with tumour antigen (kaede red+ and mCherry+). Created in BioRender. Nenclares, P. (2025) https://BioRender.com/rwram82. C Data for experiment as describe in B, display type I conventional DCs, cDC1, and type II conventional DCs, cDC2 in lymph nodes. D. Data for type I conventional DCs, cDC1, and type II conventional DCs, cDC2 in tumours. For data in C and D, a one-way ANOVA was used with the p value corrected for multiple comparisons (PBS/vehicle, n = 7, PBS/pal n = 6, Rt3D/vehicle n = 7, Rt3D/pal n = 7). P values were derived where p > 0.05 ns, *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001. Source data are provided as a Source Data file.
Rt3D-palbociclib increases RNA sensor expression and JAK/STAT-dependent expression of endogenous retroviral (ERV) species
To understand more mechanistically the palbociclib-specific effects within combination therapy, transcriptomic analysis of Rt3D-palbociclib was compared with Rt3D. This revealed marked downregulation in genes sets associated with chromatin accessibility and nucleosome organisation. Histone mRNA expression was significantly downregulated, a result that was corroborated in all histones detected in proteomic analysis (Fig. 4A, B). Additionally, analysis of proteins with the GO term ‘DNA modification’ revealed decreases in DNA methyltransferases, including suppression of DNA methyltransferase-1 (DNMT1) and EZH2, by palbociclib alone (Fig. 4C). It has been reported that reduced DNMT function, either through CDK4/6 inhibition or direct DNMT-inhibition, can trigger release of ERV RNA species, which can act as tumour-associated antigens and stimulate interferon responses19,20. The RNA sensors RIG-I (DDX58), MDA-5 (IFIH1) and TLR3 were all significantly upregulated with Rt3D-palbociclib relative to single-agent counterparts (Fig. 4D). To determine the contribution of RNA sensors to combined therapeutic efficacy, knockdown of DDX58 (but not of IFIH1 or TLR3) rescued cells from the cytotoxicity of Rt3D-palbociclib combination (Figs. 4E and S4A). DDX58 knockdown also reduced the expression of virus-induced IFN response genes (Fig. S4B), implicating its importance both in cell death and the host response to viral infection.
A Volcano plot of RNA sequencing data showing upregulated and downregulated genes for A375 cells treated with Rt3D (MOI 0.1) plus palbociclib (1 μM), compared to Rt3D alone, at 48 h. Data are from n = 3 independent biological replicates. B Proteomic analysis of A375 cells treated with Rt3D (0.1–1) ± palbociclib (1 μM) reveal upregulated (red) and downregulated (blue) histone proteins (48 h, data are from 1 independent experiment). C Proteomic analysis of A375 cells treated with Rt3D (0.1–1) ± palbociclib (1 μM) reveal upregulated (red) and downregulated (blue) categorised under the GO term ‘DNA modification’ (48 h, data are from 1 independent experiment). D RT-qPCR of RIG-I (DDX58), MDA-5 (IFIH1) and TLR3 in A375 cells treated with Rt3D plus palbociclib (1 μM) at 48 h. Data presented are mean values ± SEM, n = 3 biologically independent experiments. P values were determined by one-way ANOVA corrected for multiple comparisons. E siRNA silencing of RNA sensors or scrambled control (SCR) in A375 cells treated with Rt3D (0.1) ∓ palbociclib (1 μM) by sulforhodamine B assay. Data are representative of 3 independent repeats. F RT-qPCR of Rt3D genome in A375-infected cells ± palbociclib (1 μM) at 48 h. Data presented are mean values ± SEM, n = 5 biologically independent experiments. G Replication of Rt3D by one-step virus growth assay for A375 ± palbociclib (1 μM). Data presented are mean values ± SEM, n = 3 biologically independent experiments. H RT-qPCR for the expression of a panel of 26 previously described ERVs in A375, treated with combination therapy, or Rt3D, respectively, from 1 independent biological experiment. I RT-qPCR of the ERVs MER21C, MLTA10, MLT1C627 or MER57B1, in A375 cells treated with Rt3D (MOI 0.1) ± palbociclib at 48 h. Data presented are mean values ± SEM, n = 3 biologically independent experiments. P values were determined by one-way ANOVA corrected for multiple comparisons. J MLT1C49, MER21C, MLTA10, MLT1C627 or MER57B1 expression in A375 cells treated with Rt3D (MOI 0.1) and/or palbociclib (1 μM) ± the JAK/STAT inhibitor, ruxolitinib (RUX, 1 μM) at 48 h. Data presented are mean values ± SEM, n = 3 biologically independent experiments. K Volcano plot of RNA sequencing data showing upregulated and downregulated ERVs in A375 cells treated with Rt3D ∓ palbociclib compared to basal at 48 h (using TETool kit). L Fold change in ERV peptides from captured HLA-I with Rt3D-palbociclib versus basal conditions. ERV Peptides Filtered: 5% Peptide FDR, Confirmed TMT Quant, Single Genomic Loci, Predicted as MHC Binding. P values were derived where p > 0.05 ns, *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001. Source data are provided as a Source Data file.
As Rt3D is a dsRNA virus, we surmised that enhanced interferon signalling RNA sensors could be driven by increased Rt3D genome levels. However, RT-qPCR analysis of Rt3D RNA showed that palbociclib did not alter viral RNA levels (Fig. 4F), nor did it increase viral replication (Fig. 4G). Therefore, we investigated the possibility that expression of ERV RNA species could be available for detection by the high levels of RNA sensors, contributing to the increased IFN response. A375 cells treated with Rt3D-palbociclib were screened for 26 previously described ERVs. Treatment induced expression of 5 ERVs - MLT1C49, MER21C, MLTA10, MLT1C627 and MER57B1 (Fig. 4H). Further analysis of these 5 upregulated ERVs showed that palbociclib increased the expression of 3 of 5 Rt3D-induced ERVs (MLT1C49, MER21C, MLTA10), over single-agent Rt3D in A375 (Fig. 4I). Similar testing in the RB-null cell line, A2058, revealed that Rt3D also induced expression of the same 5 ERVs, but additionally MER4A, MLT1B, ERV16A3 and THE1D (Fig. S4C). However, palbociclib had no additional effect on these Rt3D-induced ERVs in the RB-null A2058 cell line (Fig. S4D). Since the expression of ERVs was dependent on Rt3D, but not palbociclib, we considered that their expression was likely IFN-dependent, which was consistent with the location of the 5 ERVs expressed in both cell lines within STAT1-associated genes (Table S1). Abrogation of ERV expression with ruxolitinib confirmed their expression was dependent on interferon/JAK/STAT signalling (Figs. 4J and S4D). RNA sequencing corroborated that Rt3D and Rt3D-palbociclib induced ERV expression (Figs. 4K and S4E). Taken together, these data show that palbociclib drives marked alterations to chromatin, including reduced expression of histones and DNA methyltransferases. Rt3D treatment drives expression of ERVs, which are dependent on JAK/STAT signalling. Interestingly, we observed neither ERV transcription nor ISG induction due to single-agent palbociclib at this time point. Rather, palbociclib enhances RNA sensor expression and transcription of select ERVs induced by Rt3D. Finally, we analysed the immunopeptidome and found that ERV elements can be presented on HLA class I molecules, and that the expression of these is altered by Rt3D-palbociclib treatment (Fig. 4L). Analysis of the RNA expression levels of the ERV sequences encoding epitopes found on the immunopeptidome revealed no correlation with treatment, and so did not simply reflect generally increased expression of ERV elements (Fig. S4F, G), highlighting the likely complexity of the roles of differing ERVs in triggering an IFN response or being translated, processed and presented as potential tumour-associated antigens.
Rt3D-palbociclib augments ER stress
Since palbociclib exerts a G1 cell-cycle arrest, we also explored the cell-cycle effects with combination therapy. Palbociclib induced an increase in the G1 compartment, as expected. However, the combination of Rt3D-palbociclib led to a significantly increased sub-G1 population (Fig. 5A). Compared with single-agent therapies, combined Rt3D-palbociclib did not alter levels of phospho-Rb, CDK4, Cyclin D1, E1 or A2 at the protein level (Fig. S5A). Conversely, western analysis confirmed that Rt3D-induced caspase-3 and PARP cleavage was enhanced by palbociclib (Fig. 4B). Mechanistically, this suggested an apoptotic event, and we confirmed, using a panel of caspase inhibitors, that the caspase-4 inhibitor, Z-YVAD-FMK, or siRNA-mediated caspase-4 knockdown rescued the phenotype of the Rt3D-palbociclib combination treatment (Figs. 5C, D and S5B, C). Caspase-4 is localised to the endoplasmic reticulum (ER) and can be activated in response to ER stress-inducing agents21. Therefore, to investigate the possibility that Rt3D-palbociclib therapy induces ER stress, proteome data were analysed under the GO term ‘Response to ER stress’. This showed altered expression of proteins relating to ER stress with marked upregulation in UPR protein expression with Rt3D-palbociclib combination therapy (Fig. 5E). Interestingly, proteomic data at an early time point (24 h), before the effects of Rt3D become pronounced, also indicated palbociclib-associated elevation of ER chaperones, such as calreticulin, calnexin, GRP78/BiP (HSPA5) and protein disulphide isomerases (Fig. 4F).
A FACS analysis of A375 cells treated with Rt3D ± palbociclib (1 μM) stained for PI at 48 h. Data presented are mean values ± SEM, n = 3 biologically independent experiments. P values were determined by two-way ANOVA of the sub-G1 compartment corrected for multiple comparisons. B Western blots showing apoptotic markers in A375 cells treated with Rt3D (0.01–1) ± palbociclib (1 μM) at 48 h, representative for 3 biological replicates. Quantification of cleaved caspase-3 bands are shown for the mean values ± SEM, n = 3 for the biologically independent experiments. A one-way ANOVA was used with the p value corrected for multiple comparisons. C Cell viability in A375 cells treated with Rt3D ± palbociclib (1 μM) ± caspase-4 inhibitor Z-YVAD-FMK, by MTT (72 h). Data represented are mean values ± SEM, n = 2 independent biological experiments. D siRNA silencing of caspase-4 or scrambled control (SCR), in A375 cells treated with Rt3D (0.1) plus palbociclib (1 μM) by crystal violet. Quantification is shown for the mean values ± SEM, n = 3 biological replicates. P values were determined by a two-tailed t-test. E Proteomic analysis of A375 cells treated with Rt3D (0.1–1) in combination with palbociclib (1 μM) showing upregulated (red) and downregulated (blue) proteins by combination treatment categorised under the gene ontology (GO) term ‘response to endoplasmic reticulum stress’ (48 h, data are from 1 independent experiment). F Proteomic analysis of A375 cells treated with Rt3D (0.1) in combination with palbociclib (1 μM) showing upregulated (red) and downregulated (blue) ER chaperones (24 h, data are from 1 independent experiment). G siRNA knockdown of UPR components or scrambled control (SCR), in A375 cells treated with Rt3D (0.1) plus palbociclib (1 μM) by crystal violet. Data show mean values ± SEM, n = 3 for the biologically independent experiments. A one-way ANOVA was used with the p value corrected for multiple comparisons. H RT-qPCR of CHOP at 48 h following Rt3D + palbociclib (1 μM) in A375. Data presented are mean values ± SEM, n = 3 for the biologically independent experiments. A one-way ANOVA was used with the p value corrected for multiple comparisons. I Cell viability in A375 and Mel624 cells treated with thapsigargin (TG, 0.04 μM) ± palbociclib (1 μM) by MTT assay at 72 h. Data presented are mean values ± SEM, n = 3 biologically independent experiments. A one-way ANOVA was used with the p value corrected for multiple comparisons. J RT-qPCR of the spliced form of XBP1 at 48 h following Rt3D + palbociclib (1 μM) in A375. Data presented are mean values ± SEM, n = 3 biologically independent experiments. A two-way ANOVA was used with the p value corrected for multiple comparisons. P values were derived where p > 0.05 ns, *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001. Source data are provided as a Source Data file.
Rt3D is known to induce ER stress2,3,4,9, linked to remodelling of ER membranes6,22, activating unfolded protein response (UPR) signalling via three effector pathways: IRE1α-XBP1, PERK-eIF2α-ATF4 and ATF67. These lead either to restoration of normal protein homoeostasis or apoptosis7. Therefore, we investigated the importance of these UPR components in the efficacy of combined Rt3D-palbociclib by trying to rescue the cell death phenotype with siRNA. CCAAT-enhancer-binding protein homologous protein (CHOP) induces death due to ER stress23. In line with our data on caspase-4, DDIT3 (CHOP) knockdown also rescued death due to combination treatment, as did PERK and ATF6 knockdown (Figs. 5G and S5D). Rt3D-palbociclib led to increased upregulation of DDIT3 compared with either single-agent counterpart, confirmed by qPCR (Fig. 5H).
Previous studies have also suggested linkages between CDK4/6 inhibition and ER stress24,25. Therefore, we next tested if palbociclib could sensitise cells to ER stress-induced cytotoxicity. Palbociclib enhanced cell kill due to the UPR activator, thapsigargin (TG), confirming that it can behave as an ER stress sensitiser (Figs. 5I and S5E). ER stress has been demonstrated to induce recruitment of XBP1 and transcriptional machinery to an enhancer element in the Ifnb1 promoter during PRR activation, resulting in synergistic Ifnβ expression in a study using mouse macrophages26. Palbociclib increased the spliced form of XBP1, and an IRE1α endonuclease reporter signal, supporting the involvement of the IRE1α-XBP1 branch in the palbociclib-associated ER stress signature (Figs. 5J and S5F–H).
ER stress enhances RNA sensor-driven interferon responses
Due to the ER stress signatures and dsRNA response induced by Rt3D-palbociclib, and the importance of both these pathways in cell death, we suspected that the combination was exploiting crosstalk between UPR signalling and cytoplasmic dsRNA sensing to mediate increased IFN response and cytotoxicity. Such UPR-RNA crosstalk has been previously described in immune cells26,27,28. We next tested non-viral tool compounds specific to ER stress/UPR activation (thapsigargin) and RNA sensor activation (poly I:C, 3p-hpRNA), to further investigate UPR-RNA crosstalk in tumour cells under alternative treatment conditions. Co-treatment with thapsigargin and poly I:C, either transfected (activating the cytoplasmic dsRNA sensor) or untransfected (acting via endosomal TLR3), enhanced cell kill relative to single-agents (Figs. 6A, B and S6A). In response to escalating doses of poly I:C, transcriptional expression of cytoplasmic RNA sensors and type I/III interferons plateaued, suggesting saturation of signalling capacity. However, combining poly I:C with thapsigargin facilitated increased expression of DDX58 (RIG-I), IFIH1 (MDA-5), IFNα, IFNβ, IL-28, IL-29 and DDIT3 (CHOP), mediating increased downstream signalling from escalating levels of input RNA agonist (Figs. 6C and S6B). The RIG-I agonist, 3p-hpRNA, combined with thapsigargin, induced similar transcriptional changes to those observed for poly I:C (Figs. 6D and S6C, D), as well as increased levels of expression of the ERV MLT1C49 (Fig. 6E). Taken together, these data demonstrate generalisable crosstalk between ER stress and RNA agonism in tumour cells resulting in augmented transcription of RNA sensors, amplification of IFN expression and enhanced cell death.
A A375 cells treated with transfected or B untransfected poly I:C (10 μg/mL), combined with thapsigargin (TG, 0.01, 0.05 μM) and cell viability shown by crystal violet assay. C RT-qPCR of RIG-I (DDX58), MDA-5 (IFIH1), IFNα IFNβ in A375 cells treated with transfected poly I:C (0.1, 2, 10 μg/mL), ±thapsigargin (TG, 0.05 μM) at 48 h. Data presented are mean values ± SEM, n = 3 biologically independent experiments. A two-way ANOVA was used with the p value corrected for multiple comparisons. D RT-qPCR of indicated gene expression in A375 cells treated with RIG-I agonist 3p-hpRNA (1 μg/mL) ± thapsigargin (0.05 μM) at 48 h. Data presented are mean values ± SEM, n = 3 for the biologically independent experiments. A one-way ANOVA was used with the p value corrected for multiple comparisons. E RT-qPCR of the ERV MLT1C49 in A375 cells treated with 3p-hpRNA ± thapsigargin at 48 h. Data presented are mean values ± SEM, n = 3 for the biologically independent experiments. A one-way ANOVA was used with the p value corrected for multiple comparisons. F IFNβ measured by ELISA in cell-free supernatant from A375 cells treated with Rt3D (0.1 MOI) ∓ palbociclib (1 μM) ± IRE1a inhibitor (STF-083010). Data shown are from 1 independent biological experiment, but is representative of 2 independent experiments. G RT-qPCR of RIG-I (DDX58) and IFNb in A375 cells treated with 1 μM palbociclib (+) ± poly I:C (2, 10 μg/mL) at 48 h. Data presented are mean values ± SEM, n = 3 for the biologically independent experiments. A two-way ANOVA was used with the p value corrected for multiple comparisons. H Tumour volumes for C57BL/6 mice bearing 4434 tumours treated with either sham injection with vehicle (n = 6), a single intra-tumoural injection of poly I:C (50 μg, n = 12), palbociclib (90 mg/kg, n = 6), or the combination (n = 11). Data are presented as mean values ± SEM, and p values were derived from one-way ANOVA adjusted for multiple comparisons from area under curve values from individual mice, where p > 0.05 ns, *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001. Source data are provided as a Source Data file.
To further test the linkage between UPR signalling and IFN response in Rt3D-palbociclib-treated cells, we evaluated effects of drugs that modulate the UPR on Rt3D-palbociclib-induced IFNβ production9. IRE1α inhibition reduced the elevated IFNβ seen with Rt3D-palbociclib therapy (Fig. 6F), as did an inhibitor of eIF2α, salubrinal. However, the PERK inhibitor GSK2656157 upregulated IFNβ protein expression, whilst the ATF6 inhibitor, ceapin, had no effect on IFNβ (Fig. S6E, F). Together with knockdown data (Figs. 4E and 5G), these experiments support ER stress/UPR activation plus RNA sensor agonism through combined Rt3D-palbociclib as a key modulator of interferon signalling and enhanced anti-cancer efficacy in response to dsRNA sensor activation.
Finally, we asked whether palbociclib sensitises cells to RNA in a non-viral context. Indeed, poly I:C/palbociclib significantly amplified DDX58 (RIG-I) and IFNβ expression above levels seen with palbociclib alone (Fig. 6G). Returning to tumour therapy, this combination reduced tumour burden in vivo, demonstrating that palbociclib treatment can more widely sensitise tumours to RNA agonism, with therapeutic benefit (Fig. 6H).
Discussion
Boosting innate immune effector function by targeting nucleic acid sensors, linked to subsequent priming of antigen-specific T cell-mediated anti-tumour immunity, is currently an exciting area of anti-cancer research, with STING, TLR3 and RIG-I agonists currently under therapeutic investigation29,30,31. Oncolytic viruses can also be classified as nucleic acid sensor agonists, but they have more pleiotropic effects, including inducing ER stress and UPR activation. ER stress due to Rt3D is well described3,8,32, and we have previously exploited this therapeutically4,9. In this study, ER stress/UPR signalling plus RNA agonism potentiated IFN signalling and enhanced cell death and immunogenicity in tumour cells, heralding significant combinatorial opportunities.
We are unaware of any previous description of CDK4/6 inhibition as an ER stress sensitiser. Palbociclib-treated young rats and CDK4-knockout mice display insulin-deficiency, defective pancreatic beta-cell function and increases in ER stress markers24,25. Beta-cells are sensitive to perturbation of ER stress, with these studies phenocopying observations in PERK-knockout mice33. Consistent with these findings, we confirm that single-agent palbociclib increased ER stress-associated proteins and the IRE1α endonuclease reporter signal in treated tumour cells.
In addition to the ER stress signature seen with Rt3D-palbociclib, we observed a potent increase in the IFN response within tumour cells, both at the transcriptional and protein levels, with an escalation in the expression of RNA sensors. Hence, induction of the UPR goes beyond protein homoeostasis, interacting with PRR signalling and enhancing inflammatory responses; this is the first demonstration, to our knowledge, of such crosstalk between these pathways in tumour cells and its immunological consequences. Importantly, this crosstalk extends beyond the use of oncolytic viruses and a targeted small molecule as a therapy combination, since enhanced UPR signalling using thapsigargin also liberated interferon gene expression to poly I:C dsRNA in melanoma cells. Furthermore, IRE1α inhibition was able to reduce Rt3D-palbociclib-induced IFNβ protein, further demonstrating the link between UPR and IFN responses. Research investigating UPR and PRRs crosstalk has not previously been described in the context of cancer therapeutics, which has broad implications for the clinical development of agents, including nucleic acid sensor agonists and oncolytic viruses. From data in this study and others, it is apparent that ER stress in and of itself can induce an increase in type I IFNs, but these changes are modest in comparison with PRR agonists and viruses28,34,35,36. Spliced-XBP1 (by the endonuclease IRE1α) has, however, been shown to bind to an enhancer site of the IfnB1 gene26, with numerous studies confirming spliced-XBP1 boosts IFNβ only in combination with certain PRR activators26,27,32,36,37.
As with the tool compound thapsigargin, palbociclib sensitised cancer cells to ER stress, which, in combination with PRR activation, also led to increased IFN expression. This combination translated to therapy, as shown by the combination with poly I:C in vivo, illustrating the capacity of palbociclib to amplify the IFN response in a non-viral context.
In this study, we addressed the consequences of the observed increased IFN on tumour cell-intrinsic immunogenicity. Increased HLA expression by Rt3D-palbociclib was observed both in RNA sequencing and proteomic data and confirmed on the cell surface by FACS analysis. MHC II expression on tumour cells has been previously reported and associated with a therapeutic response to PD-138. Expression of both HLA class I and II was reduced with inhibition of JAK/STAT signalling, demonstrating a link between increased IFN signalling and antigen presentation. CDK4/6i has previously been reported to increase antigen presentation39, consistent with the findings in this study. In vivo, we observed activation of tumour cDC1s with Rt3D-palbociclib therapy (over Rt3D alone), that was not seen in cDC2s. Since cDC1s have been shown to be key in cross-presenting antigens to CD8+ T cells and the generation of cytotoxic T cell responses, this was supported by our data, in a separate in vivo experiment, that demonstrated increased expression of CD69+ CD8 T cells with Rt3D-palbociclib compared with Rt3D alone. In tumour-draining lymph nodes, the activation status of both cDC1 and cDC2 was increased with Rt3D-palboiclib combination therapy in cells that had migrated from the tumour and/or were associated with tumour material via tumour mCherry expression.
CDK4/6i has been shown to induce dsRNA-IFN responses through downregulation of the E2F target DNMT1 and expression of ERV elements20,39. Indeed, we observed marked modification to DNA, including downregulation of DNMT as a result of palbociclib treatment, and a marked downregulation in histones. Analysis of chromatin accessibility, by methods such as ATAC sequencing, may provide more insight into the palbociclib-driven effects on histone loss, and the consequences on the transcriptome (including ERV expression). By screening a selection of ERVs, we observed enhanced STAT1-driven transcription of select ERVs. This was not specifically a virus-mediated effect, because ERV expression was also accomplished by the treatment of RIG-I stimulation and amplified in combination with ER stress (thapsigargin).
Upregulation of shared tumour-associated antigens such as MAGEA4, as well as enhanced STAT1-driven ERV transcription, is likely to be beneficial in promoting anti-tumour-immune responses40,41,42. Herein, we demonstrate that peptides originating from CTA and ERV elements can be presented by HLA class I molecules on tumour cells. Whether these peptides and their expression reflect alterations in functionally relevant tumour-associated antigens, which contribute to adaptive immune therapy, is currently not known. Despite the presence of ERV peptides on the immunopeptidome, no changes were seen in the RNA expression of these ERVs with Rt3D-palbociclib treatment. This phenomenon was also observed for MAGEA4 expression, and suggests that post-transcriptional processing events (such as proteasome/immunoproteasome switching) may be responsible for the appearance of these peptides associated with HLA-I. Overall, the wider functional immune consequences of the tumour cell-intrinsic immune activation by CDK4/6 inhibition and double-stranded RNA sensor agonism described in this study require further investigation. Key questions, including i) the extent to which the immune system plays a role in Rt3D-palbociclib combination therapy in vivo, ii) the effects of Rt3D-palbociclib on the tumour-immune microenvironment, and iii) whether the potentially antigenic peptides we have found presented by Rt3D-palbociclib-treated cells (e.g. derived from ERV/CTA) are important for in vivo therapy, are currently being addressed in our laboratory. Interestingly, therapeutic effects were observed in vivo in the 4434 murine melanoma model, despite no increased killing of this cell line in vitro with combination therapy compared with virus alone. This would imply other important effects of combining palbociclib with Rt3D beyond superior direct tumour cell killing, perhaps through the increased IFN signalling and innate response that was observed with the combination therapy.
This study demonstrates that crosstalk between RNA sensing and ER stress pathways can increase tumour cell death, particularly in the application of the clinical therapies CDK4/6 inhibition and RNA agonism/oncolytic virotherapy. These interventions also increase the immunogenicity of tumour cell death, both in terms of innate interferon production and adaptive presentation by HLA class I peptides of potential TAA.
Our data support work towards clinical translational development of combined RNA sensing and ER stress pathway engagement. Clinically, both palbociclib and Rt3D are excellent candidates for combination, given the fact that the former is an approved anti-cancer agent and the latter has been the subject of multiple clinical trials demonstrating safety and tolerability. Of course, inevitably, there are limitations to this study that will require further investigation before we can consider translating our findings to the clinic. For instance, expanding the selection of animal models to those beyond BRAF-mutant melanoma would address the heterogeneity of the various histological subtypes of cancer and their respective tumour-immune microenvironments (TIME). Indeed, a more in-depth analysis of how the combination of Rt3D and palbociclib performs together within the complexities of different TIMEs might point the way towards potential triple therapy combinations with greater therapeutic efficacy. For example, an increase in antigen presentation may fail to reach its full therapeutic potential without the addition of immune checkpoint blockade. Further work is currently underway in our lab to fully define the functional immune consequences of such novel combinatorial approaches. Nevertheless, inter- and intra-tumoural variability, heterogeneity of the TIME, differences in immune function and the make-up of the microbiome between and within patients will undoubtedly impact the outcome of translating our current findings to the clinic. In fact, many of these factors will probably be most meaningfully studied within the context of early-phase trials that focus on detailed collection of biological samples.
Methods
Cell lines and therapeutic agents
Neat stocks of 1 × 109 TCID50 Rt3D were diluted 1:10 in DMEM for single-use aliquots and stored at −80 °C. Palbociclib was kindly provided by Pfizer.
A375, Mel624, MeWo, A2058, Fadu, HN5 and Cal27 were obtained from stocks within Kevin Harrington’s team at the ICR. D04 was obtained by generous donation from R. Marais, CRUK Manchester Institute. T47D, T47D-pR, MCF-7, MCF-7-pR were obtained by generous donation from N. Turner, ICR London and cultured in RPMI (no phenol red) plus oestradiol (1 nM) with the addition of 0.5 µM palbociclib for resistant (pR) cell lines. All other cell lines were obtained from stocks within Kevin Harrington’s team at the ICR. DMEM or RPMI was supplemented with 5% or 10% (v/v) FCS, 1% (v/v) glutamine, and 0.5% (v/v) penicillin/streptomycin (ICR, CSSD, UK).
Fucci and reporter cells
Cells were infected with Fluorescent Ubiquitination-based Cell-Cycle Indicator (FUCCI) cell-cycle vectors. G1 red expresses mCherry hCdt1(30-120aa); S–G2–M cyan expresses AmCyan hGeminin(1-110aa). Sequences derived from pRetroX-G1-Red and pRetroX-SG2M (Clontech) were cloned into pHR lentiviral vectors under puromycin or blasticidin selection. Lentiviral packaging was performed in 293T cells using MD2.g and psPAX2. A375 cells containing an IRE1α endonuclease reporter construct were generated by cloning the reporter construct into the lentivirus pHRSIN (kindly provided by Prof. Greg Towers, UCL, London, UK). The region encoding the 26 nt-nt intron from XBP1 excised by IRE1a riboendonuclease activity was incorporated by PCR in front of EGFP-FLAG and inserted between SFFV and WPRE elements (primers used were forward: GTTGGATCCGCCACCATGCCAGTGGCCGGGTCTGCTGAGTCCGCAGCACTCAGACTACGTGCACCTCTGCAGCAGGTGCAGGCCCAGTTGTCGTGAGCAAGGGCGAGGAG, and reverse: AATCAGGCGGCCGCTTACTTGTCGTCATCGTCTTTGTAGTCC).
Library screen and cell titre Glo assays
Cells were plated at 500 cells per well in 20 µL, in 384-well plates using a Thermo Scientific multidrop combi and incubated overnight. Readily available small molecule screen plates 1 (11) and 2 (12) were used compromising a range of different concentrations of 80 drugs (Prof. Chris Lord, ICR). Medium containing 20 µL of the drug ranging from 1 to 1000 nM was added to the cells using a Hamilton microlab star. Cells were infected with Rt3D in 5 µL per well, 2 h after the addition of the compounds. 72 h after treatment, cell viability was measured by CellTiter-Glo Luminescent Cell Viability Assay (G9681, CTG, Promega, UK). 25 µL of CTG was added per well and incubated at 37 °C for 10 min. Luminescence was measured on a Wallac Victor 2 V plate reader.
Screen analysis
In each plate, 8 dose points were used per compound. Combined with 4 doses of Rt3D, this was a comprehensive 8 × 5 matrix screen. Rt3D cytotoxicity levels between plates were compared for consistency. The effect of the drug-virotherapy combination was ranked as ‘therapy effect’ Z scores. In brief, Z scores are a normalised value between untreated (library alone) versus treated (virus plus library) for each dose used. A threshold value <−2 was used to define highly significant sensitisation (values of >2 would indicate combination therapy resistance). The data have been normalised to untreated or virus alone in each case.
MTT assays
Cells were seeded between 5 × 103 and 1 × 104 cells per well in 96-well plates overnight prior to treatment with palbociclib at indicated doses (Pfizer), thapsigargin (T9033, Sigma), 5-Azacytidine (A2385, Sigma), poly I:C (P1530, Sigma) and/or infected with Rt3D at indicated MOI. At the desired time point, 20 µL of 5 mg/mL 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (M6494, MTT) was added per well and incubated for 2–4 h. The supernatant was removed, and the formazan was solubilised in DMSO. Absorbance was read in a plate reader at 550 nm.
Crystal violet and sulforhodamine B (SRB) assays
Cells were seeded in 6-well plates at 3 × 105 cells per well overnight and treated as described. For crystal violet assays, medium was removed, and cells were washed in PBS. 2 mL of crystal violet (HT90132, scientific laboratory supplies), was added per well and incubated for 20 min, after which time it was removed, and the plates washed. For SRB assays, cells were fixed by adding an equal volume of 10% TCA for 1 h. Plates were then washed in tap water and left to dry prior to staining with 0.057% SRB (wt/vol, 341738, Sigma) in 1% acetic acid overnight. Plates were then washed in 1% acetic acid and left to dry.
Clonogenic assays
Cells were seeded in 6-well plates at 7.5 × 103 cells per well overnight. After treatment with palbociclib or Rt3D, cells were incubated for 2 weeks with renewal of cell medium every 4 days (replacing drug but not virus). Colonies were measured by crystal violet assay.
Cell-cycle assays
Cells were seeded in 6-well plates at 3 × 105 cells per well overnight prior to treatments. Post-treatment, cells were harvested, washed and then fixed with ethanol overnight. The following day, cells were pelleted out of ethanol, and stained with propidium iodide (P4864, 40 µg/mL) in the presence of 10 μg/mL RNase A (R4875, Sigma) prior to analysis on the BD LSR II analyser.
FACS analysis
Cells were seeded in 6-well plates at 3 × 105 cells per well overnight prior to treatments. At the desired time point, cells were harvested and spun at 300 × g, 5 min. Cells were resuspended in antibodies anti-human HLA-A,B,C (311403, biolegend), HLA-DR,DP,DQ (361709, biolegend) with viability dye (65-0865-14, eBioscience) for 30 mins. Cells were washed in PBS + 5% FBS and fixed in FluoroFix buffer (422101, BioLegend) prior to analysis on the BD LSR II analyser or the BD Symphony A5. Data was collected using DIVA software, version 9.3.1 (build 2021_11_30_11_38), and analysed using Flojo version 10.10.0.
In vivo experiments
All procedures were approved by the Animal Welfare and Ethical Review Board at the Institute of Cancer Research in accordance with Home Office Regulations under the Animals (Scientific Procedures) Act 1986. All animals were handled according to the Institute and U.K. Home Office guidelines and kept in a specific pathogen-free facility and exposed to a 12 h light/dark cycle at a constant temperature (22 ± 2 °C), with access to unrestricted food and water supply. CD1 nude mice or C57BL/6 female mice aged 6–8 weeks (Charles Rivers, Kent, UK) were subcutaneously injected with 3 × 106 A375 cells or 4 × 106 4434 cells, respectively, suspended in PBS in the right flank. Once tumours were established to ~6 mm in diameter, mice were allocated treatment groups stratified by tumour size, with n = 6–10 animals per group. Mice bearing tumours were treated daily with 60–90 mg/kg palbociclib (Pfizer) or vehicle (0.05 N sodium lactate buffer, pH 4.0) by oral gavage. Drug administration continued daily for 2 weeks. A Rt3D injection of 1 × 106 pfu dissolved in PBS (or a PBS sham) was administered as an intra-tumoral injection 3 days after drug administration commenced. Poly I:C was given as a single intra-tumoural injection of 50 μg (tlrl-pic, Invivogen). Kaede mice [https://knowledge.brc.riken.jp/resource/animal/card?__lang__=en&brc_no=RBRC05737] were kindly provided by Dr Y. Miwa (Tsukuba University) and Dr O. Kanagawa (RCAI, RIKEN) and Dr. M. Tomura (Osaka Ohtani University), and bred at Charles River. In these experiments, tumour was exposed to a 405-nm wavelength LED light from a fixed distance of 1 cm using a Dymax BlueWave QX4 system (DYM41572; Intertronics) outfitted with an 8 mm focusing lens for a total of 3 min at 50% of full power with a 5-s break for every 20 s. We used a 0.3 mm thick piece of black acrylic with a 12 mm diameter hole to shield the surrounding area, ensuring only the tumour would be exposed to near-UV light and photoconverted. During the 5-s 0% intensity steps, H2O was carefully applied to the tumour to prevent the skin drying out16,17,18. Established tumour volumes were measured at least twice weekly using Vernier calipers and the tumour volume was estimated from the formula: V = 0.5 × (length × width2). Permission was granted to perform animal experiments according to Home Office, with animals judged to have failed treatment if tumour diameter approached the size limit of 15 mm in one of three dimensions. These limits were adhered to in all experiments. The humane end-point was euthanasia by neck dislocation due to moribund status as determined by weight loss of 18% or more compared with maximum weight measured, inability to reach food or water, breathing difficulties, hunched intermittently, anorexia, dehydration, marked piloerection, lethargy and subdued. Animals weights were monitored twice weekly. Stock numbers for purchased animals are 000664.
Immune profiling of tumours
C57BL/6 mice (Charles Rivers, Kent, UK) were subcutaneously implanted with 4 × 106 4434 murine cells suspended in 0.1 mL PBS per flank. Tumours were allowed to grow to 6–8 mm before mice were allocated treatment groups stratified by tumour size. Mice bearing tumours were treated daily with 100 mg/kg palbociclib (kindly provided by Pfizer) or vehicle (0.05 N sodium lactate buffer, pH 4.0, refer to palbociclib stocks) by oral gavage. A Rt3D injection of 5 × 106 pfu dissolved in PBS (or a PBS sham) was administered as an intra-tumoural injection 3 days after drug administration commenced. Tumours were harvested and analysed for tumour-infiltrating lymphocytes by FACS. Tumours were harvested and minced with scissors in digestion mix (0.01% trypsin, 2.5 mg/mL collagenase, 2 mg/mL dispase and 1 mg/mL DNAse in RPMI) and incubated at 37 °C for 30 min. Thereafter, samples were kept on ice. Suspensions were passed through a 70 μm strainer using a 2.5 mL syringe plunger and washed through with RPMI + 5 mM EDTA until only connective tissue remained. Samples were centrifuged at 400 × g, for 5 min at 4 °C, and transferred into a V-well 96-well plate. Samples were stained in FACS buffer (PBS + 5% FCS) for 30 min on ice and protected from light, with extracellular antibodies. Cells were then washed in FACS buffer and permeabilized and stained with intracellular antibodies. Samples were then washed and fixed (1–2% PFA) prior to analysis of tumour-infiltrating lymphocytes by flow cytometry. Tumours were weighed on collection and counting beads were added when running the analysis to calculate cells per mg of tumour. All antibodies used are displayed in Table S4.
Western blotting
Samples harvested for western blotting were stained with antibodies as shown in Table S5. Antibodies were used at a 1:1000 dilution. Uncropped and unprocessed scans for all western blots are provided in the source file.
ZVAD experiments
Cells were seeded in 96-well plates at 5 × 103 cells per well overnight. An inhibitor sampler pack which contains caspase inhibitors of 1, 2, 3, 4, 6, 8, 9, 10, 13 and a pan-caspase inhibitor (FMKSP01, R&D systems), was used on cells treated with combination therapy to observe rescue from cells treated without inhibitors. The caspase-4 inhibitor (Z-YVAD-FMK, FMK005, R&D systems) was used in further experiments at the dose of 50 µM. Cell survival was measured by MTT assay 72 h later.
siRNA experiments
Cells were seeded in 12- or 6-well plates at 5 × 105 or 3 × 105 cells per well, respectively, overnight. Cells were transfected with siRNA (Qiagen) using Lipofectamine RNAiMAX transfection reagent (13778100, Thermo Fisher) as per manufacturer's instructions. 24 h later media was replaced containing therapy and cell survival was measured 48–96 h later by crystal violet assay.
Viral replication assays
Cells were plated at 1 × 105 in 24-well plates and treated the following day with palbociclib and/or Rt3D at a dose of MOI 5 (to ensure infection of all cells). After another 2 h, cells were washed twice with media and the inhibitors replaced. At 4, 24, and 48 h after infection, cells were harvested into the media and stored at −80 °C prior to processing. The lysate was subjected to 3× freeze/thawing between −80 and 37 °C, centrifuged at 8000 × g for 5 min and the supernatant was used to titre for Rt3D by TCID50 assay on L929 cells.
RT-qPCR
RNA was extracted from samples using RNeasy kit (74134, Qiagen, UK), and cDNA was synthesised using SensiFAST cDNA synthesis kit (BIO-65053, Bioline), before amplification against transcripts by qRT-PCR kit with SYBR green (BIO-92005, Bioline). Primers used against DDIT3 (CHOP) were commercially available QuantiTect primer assays (249900, Qiagen, UK). Please refer to Tables S2 and S3 in the supplementary material for the sequences of other primers used. Relative gene expression was calculated with the 2-ddCT method using 18S or beta actin as a housekeeping gene.
ATP release
Cells were plated at 3 × 105 cells per well in 6-well plates and incubated overnight prior to treatment the following day. Cell supernatants were collected at the desired time points and centrifuged at 800 × g for 4 min. 200 μL of cell-free supernatants were mixed with 50 μL CellTiter-Glo Luminescent Cell Viability Assay (Promega, G7571) in white 96-well plates (Greiner, 655075). After 10 min incubation at room temperature, luminescence was measured on a Victor 2V plate reader.
Confocal microscopy
Cells were seeded in glass-bottom dishes at 3 × 105 cells per well overnight prior to treatments. After treatment cell medium was removed, and cells were fixed in 10% formalin for 5–10 min. Samples were then washed and then blocked (with PBS + 1 % BSA, 2 % FBS and 0.05% Sodium Azide). Samples were stained with calreticulin primary antibody (PA3-900, Thermofisher, UK), secondary antibody conjugated to FITC (A-11070 anti-rabbit Alexafluor 488, Invitrogen, UK), Hoechst 33342, and imaged by confocal microscopy by Z stacking. 3 fields of view were taken for 3 independent repeats and CRT expression was quantified relative to cell number using cell profiler.
Phagocytosis assays
PBMCs were harvested from K3EDTA anti-coagulated whole blood by density gradient centrifugation with Lymphoprep and plated in 12-well plates at 2–4 × 106 cells per well in 10% RPMI + 10 ng/mL human CSF (300-25, Peprotech, UK). Medium was replaced every 2–3 days until adherent populations of macrophages had established, and these were used in experiments 8–10 days later. Tumour cells were stained with 20 ng/mL of pHrodo-SE (P36600, Thermofisher) for 30 min then washed and co-cultured with the macrophages14. 1–2 h after co-culture, samples were washed and stained for CD11b conjugated to FITC (#11-0118-42, Thermofisher), then fixed and analysed on the BD LSR II. Separately stained macrophages or treated cells labelled with pHrodo-SE served to determine cut-offs for phagocytosis. In 1 × 104 collected events, phagocytosis percentage was determined by the double-stained CD11b+ pHrodo-SE+ cells from the CD11b+ population above the threshold set in the controls.
Whole proteome LC-MS/MS analysis
Cells were seeded in 6 cm dishes at 9 × 105 cells per well overnight prior to treatments and collected at indicated timepoints by trypsinising cells and washing 3 times in PBS. One techinal replicate was collected for two time points, 24 and 48 h. At the 24-h time point, 4 samples were collected, consisting of mock, Rt3D at an MOI of 0.1, palbociclib at a dose of 1 μM, or the combination. At the 48-h time point, 6 samples were collected, consisting of mock, Rt3D at an MOI of 0.1 and 1, palbociclib at a dose of 1 μM, or the combination of palbociclib with each dose of Rt3D. This totalled 10 samples collected and analysed for mass spectrometry analysis and as such no statistical tests were performed on the analysis. Pellets were stored at −80 °C prior to processing. Cell pellets were lysed in 5% SDS/100 mM TEAB (tetraethylammonium bromide, Sigma) by probe sonication and heating at 90 °C for 10 min. Proteins were reduced by TCEP (Tris(2-carboxyethyl) phosphine, Sigma), alkylated by iodoacetamide (Sigma), and purified by trichloroacetic acid (Sigma) precipitation. The protein pellet was resuspended in 100 mM TEAB buffer and digested by trypsin (Thermo). 40 µg of protein digest was labelled by TMT11plex (Thermo Fisher), and 11 samples were pooled, dried in SpeedVac. The mixture was fractionated on an XBridge BEH C18 column (2.1 mm i.d. × 150 mm, Waters). Fractions were collected at every 30 s and then concatenated to 28 fractions and dried in SpeedVac. 1/3 of peptides were injected for online LC-MS/MS analysis on the Orbitrap Fusion Lumos hybrid mass spectrometer coupled with an Ultimate 3000 RSLCnano UPLC system. Peptides were first loaded on a PepMap C18 nano trap (100 µm i.d. × 20 mm, 100 Å, 5 µ), and then separated on a PepMap C18 column (75 µm i.d. x 500 mm, 2 µm) over a linear gradient of 6.4–30.5% CH3CN/0.1% formic acid in 90 min/cycle time at 120 min at a flow rate at 300 nL/min. All instruments and columns were from Thermo Fisher. The MS acquisition used MS3 level quantification with Synchronous Precursor Selection (SPS5) with the Top Speed 3 s cycle time. Briefly, the full MS survey scan in Orbitrap was m/z 375–1500. Multiply charged ions (2–5) with intensity threshold above 7 × 103 were fragmented in ion trap at 35% collision energy with isolation width at 0.7 Da in quadruple. The dynamic range was 40 s at ±10 ppm. The top 5 MS2 fragment ions were SPS selected with the isolation width at 0.7 Da and fragmented in HCD at 65% NCE and detected in Orbitrap.
Whole proteome data analysis
Raw spectra were processed using ProteomeDiscoverer v2.4(Thermo Scientific) and searched against FASTA sequence databases containing GENCODE v3243 protein sequences, UniProt (2019_05) Reovirus Proteins, translated gEVE database44 sequences and cRap contaminates using both Mascot server v2.4 (Matrix Science) and SequestHT with target-decoy scoring evaluated using Percolator45. The precursor tolerance was set at 20 ppm, fragment tolerance set at 0.5 Da and spectra were matched with fully tryptic peptides with a maximum of 2 missed cleavages. Fixed modifications included:carbamidomethyl [C] and TMT6plex [N-Term]. Variable modifications included: TMT6plex [K], oxidation [M], and deamidation [NQ]. Peptide results were initially filtered to a 1% FDR(0.01 q-value). The reporter ion quantifier node included a TMT-11-plex quantification method with an integration window tolerance of 15 ppm and integration method based on the most confident centroid peak at MS3 level. Protein quantification was performed using unique peptides only, with protein groups considered for peptide uniqueness. Log2 fold change ratios were calculated for each sample vs time point Basal sample using normalised protein abundances. Ratios and abundance were loaded into Perseus46 or further downstream analysis and plotting. Z-score scaling was used to generate heatmaps, proteins were clustered using k-means method and GO enrichment was performed using fisher exact test. Results and RAW spectral files have been uploaded to PRIDE repository47 under project accession PXD036540.
HLA-I peptide capture and TMT labelling
To capture HLA peptides, HLA class I molecules were purified from 100 × 106 cells by immunoaffinity precipitation (IP). Snap frozen cell pellets were lysed in 0.5% CHAPS (Abcam) supplemented with protease and phosphatase inhibitors (Thermo Scientific) for 1 h at 4 °C. The lysate was centrifuged (30 min, 10,000 × g at 4 °C) and supernatants used for HLA-I IP in which purified anti-human HLA-A,B,C antibody (W6/32, BioLegend) was crosslinked to Protein G Dynabeads (Thermo Scientific) with Dimethyl pimelimidate dihydrochloride (DMP, Sigma Aldrich). Beads were washed with phosphate buffered saline (PBS) 0.01% Tween 20 (Merck Millipore), incubated with antibody and PBS 0.1% Tween (15 min at RT), washed twice with PBS 0.01% Tween and twice with 0.2 M Triethanolamine (pH 8.2), prior to crosslinking. Beads and antibodies were crosslinked using 20 mM DMP in Triethanolamine (pH 8.2) (30 min at RT), and quenched with 50 mM Tris-HCl (15 min at RT). Beads were washed with PBS 0.1% Tween before incubation with cell lysate supernatants (3 h, at 4 °C). Beads were washed eight times with cold PBS followed by two washes with cold water prior to peptide elution, in 1 ml 0.2 % trifluoroacetic acid on ice. Four rounds of elution were performed on each sample. The beads were reused for a second round of IP using the lysate flowthrough from the first IP. Eluted peptides were pooled, dried, reduced with 5 mM TCEP (15 min at RT) and alkylated with 10 mM iodoacetamide (30 min at RT). Purified HLA class 1 peptides were labelled by 1 mg TMTpro, followed by desalting and fractionation on the high pH spin column (Thermo) as the step elution method by these solvent composition: 10%, 12.5%, 15%, 17.5%, 20%, 25% and 30%, then concatenated to 3 fractions. Samples were dried down by SpeedVac, and resuspended in 0.5% FA before LC-MS analysis.
Immunopeptidome MS analysis
Samples were analysed by LC-MS/MS on the Orbitrap Fusion Lumos mass spectrometer coupled to a U3000 RSLCnano UHPLC system. All instruments and columns used were from Thermo Fisher. The peptides were first loaded to a PepMap C18 trap (100 µm i.d. × 20 mm, 100 Å, 5 µm) at 10 µl/min with 0.1% FA/H2O, and then separated on a PepMap C18 column (75 µm i.d. × 500 mm, 100 Å, 2 µm) at 300 nl/min and a linear gradient of 4-32% ACN/0.1% FA in 60 min with the total cycle at 90 min. The Orbitrap full MS survey scan was m/z 375–2000 with the resolution 120,000 at m/z 200, with AGC (Automatic Gain Control) set at 40,000 and maximum injection time at 50 ms. Ions with charge at 1–4 with intensity above 50,000 counts were fragmented in HCD (higher collision dissociation) cell at 35% collision energy, and the isolation window at 1 Da. The fragment ions were detected in Orbitrap with AGC at 50,000 and 86 ms maximum injection time. The dynamic exclusion time was set at 30 s with ±10 ppm. The vials were then resuspended in 20% ACN/0.5% FA, diluted to 5% ACN, and followed by another LC-MS analysis with a shorter gradient.
Immunopeptidome data analysis
Raw MS spectra were converted to mzML format using ThermoRawFileParser48. Spectra were searched in a custom NextFlow and OpenMS (v2.6)49 pipeline using Comet (2019.01 rev. 5)50 and MSGFplus (v2020.07.02)51. Results from both search engines were processed by Percolator and merged using the OpenMS tool ConsensusID. The searches were performed against GENCODE v38 Protein Sequences, UniProt (June 2021) ReoVirus Proteins, cRap contaminates, and ERV proteins translated from gEVE database44 and hERVd database52. Search parameters included minimum peptide length of 8 amino acids and maximum of 15 amino acids with unspecified peptide cleavage. A maximum peptide charge of 7 was allowed. Precursor tolerance was set to 20 ppm. MSGFplus instrument type was set to ‘high_res’ and Comet fragment tolerance was set to 0.01 Da with a bin offset of 0, mass range of 600–2500 Da and maximum fragment charge of 5. Fixed modifications included TMTpro (N-term), TMTpro (K), and Carbamidomethyl (C). Variable modifications included Oxidation (M). Results were filtered at a 5% Peptide FDR, and A375 HLA binding was predicted using NetMHC (4.1)53. Immunopeptidome data files have been uploaded to PRIDE repository under project accession PXD036559.
RNA sequencing
Cells were seeded in 6-well plates at 3 × 105 overnight prior to treatments. 48 h after treatment cells were harvested using the RNeasy kit (74104, Qiagen) with on-column DNase digestion (79254, Qiagen). Samples were sequenced in-house (at the ICR, London). The STAR alignment software (v.2.5.1b) was used to align reads to the soft masked human reference genome containing all the repeat and low complexity sequences (GRCH38_sm). Custom generated GTF files for TE annotations were downloaded from the TETool kit website (http://labshare.cshl.edu/shares/mhammelllab/www-data/TEtranscripts/TE_GTF/). Downstream ERV expression analysis was carried out using TETranscript from TEToolkit suite (v2.03), or the SalmonTE (v0.2) pipeline.
ELISA
A375 cells were seeded at 2 × 105/mL and treated with 1 µM palbociclib then infected with Rt3D for 48 h. The production of human IFNα (Mabtech), IFNβ (PBL Interferon Source), IL-28 and IL-29, (R&D Systems) in cell-free supernatant was determined using matched-pairs antibodies according to the manufacturers’ instructions. Optical density absorbance readings were determined at 405 nm absorbance. For UPR inhibitor experiments the compounds STF-083010 (IRE1α inhibitor) was used at 20 µM and IFNβ was measured using the DuoSet ELISA (DY814-05, R&D Systems) as per manufacturer’s instructions.
Statistical analysis
Statistical analysis was performed in GraphPad Prism version 10.4.2. T-tests or ANOVA tests were used to make comparisons between groups. For in vivo analyses, the area under curve (AUC) was calculated for individual mice. A normality test was performed to verify the data follow a Gaussian distribution. If they do, a T-test or ANOVA test was used followed by a multiple comparisons test, if not, a non-parametric Mann-Whitney test was performed followed by a Dunn’s multiple comparison test. P values were derived where p > 0.05 ns, *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data are included in the Supplementary Information or available from the authors, as are unique reagents used in this Article. The raw numbers for charts and graphs are available in the Source Data file whenever possible. The RNAseq data for this study have been deposited in the European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB75360. Whole proteome results and RAW spectral files have been uploaded to the PRIDE repository under project accession PXD036540, and immunopeptidome data files have been uploaded to the PRIDE repository under project accession PXD036559. Source data are provided with this paper.
Change history
15 July 2025
A Correction to this paper has been published: https://doi.org/10.1038/s41467-025-61212-3
References
Kaufman, H. L., Kohlhapp, F. J. & Zloza, A. Oncolytic viruses: a new class of immunotherapy drugs. Nat. Rev. Drug Discov. 14, 642–662 (2015).
Smith, J. A. et al. Reovirus induces and benefits from an integrated cellular stress response. J. Virol. 80, 2019–2033 (2006).
Carew, J. S. et al. Reolysin is a novel reovirus-based agent that induces endoplasmic reticular stress-mediated apoptosis in pancreatic cancer. Cell Death Dis. 4, e728 (2013).
Roulstone, V. et al. BRAF-and MEK-targeted small molecule inhibitors exert enhanced antimelanoma effects in combination with oncolytic reovirus through ER stress. Mol. Ther. 23, 931–942 (2015).
de Castro, I. F. et al. Reovirus forms neo-organelles for progeny particle assembly within reorganized cell membranes. mBio 5, e00931-13 (2014).
Tenorio, R. et al. Reovirus σNS and μNS proteins remodel the endoplasmic reticulum to build replication neo-organelles. mBio 9, 1–15 (2018).
Hetz, C., Chevet, E. & Harding, H. P. Targeting the unfolded protein response in disease. Nat. Rev. Drug Discov. 12, 703–719 (2013).
Kelly, K. R. et al. Reovirus therapy stimulates endoplasmic reticular stress, NOXA induction, and augments bortezomib-mediated apoptosis in multiple myeloma. Oncogene 31, 3023–3038 (2012).
McLaughlin, M. et al. The PERK inhibitor GSK2606414 enhances reovirus infection in head and neck squamous cell carcinoma via an ATF4-dependent mechanism. Mol. Ther. Oncolytics 16, 238–249 (2020).
Teh, J. L. F. & Aplin, A. E. Arrested developments: CDK4/6 inhibitor resistance and alterations in the tumor immune microenvironment. Clin. Cancer Res. 25, 921–927 (2019).
Md Sakib Hossain, D. et al. Dinaciclib induces immunogenic cell death and enhances anti- PD1–mediated tumor suppression. J. Clin. Investig. 128, 644–654 (2018).
Kepp, O. et al. Consensus guidelines for the detection of immunogenic cell death. Oncoimmunology 3, e955691 (2014).
Gardai, S. J. et al. Cell-surface calreticulin initiates clearance of viable or apoptotic cells through trans-activation of LRP on the phagocyte. Cell 123, 321–334 (2005).
Miksa, M., Komura, H., Wu, R., Shah, K. G. & Wang, P. A novel method to determine the engulfment of apoptotic cells by macrophages using pHrodo succinimidyl ester. J. Immunol. Methods 342, 71–77 (2009).
Sen Santara, S. et al. The NK cell receptor NKp46 recognizes ecto-calreticulin on ER-stressed cells. Nature 616, 348–356 (2023).
Li, Z. et al. In vivo labeling reveals continuous trafficking of TCF-1+ T cells between tumor and lymphoid tissue. J. Exp. Med. 219, e20210749 (2022).
Dean, I., Kennedy, B. C., Li, Z., Berditchevski, F. & Withers, D. R. Protocol for transcutaneous tumor photolabeling to track immune cells in vivo using Kaede mice. STAR Protoc. 5, 102956 (2024).
Tomura, M. et al. Monitoring cellular movement in vivo with photoconvertible fluorescence protein ‘“Kaede”’ transgenic mice. www.pnas.org/cgi/content/full/ (2008).
Chiappinelli, K. B. et al. Inhibiting DNA methylation causes an interferon response in cancer via dsRNA including endogenous retroviruses. Cell 162, 974–986 (2015).
Goel, S. et al. CDK4/6 inhibition triggers anti-tumour immunity. Nature 548, 471–475 (2017).
Hitomi, J. et al. Involvement of caspase-4 in endoplasmic reticulum stress-induced apoptosis and Aβ-induced cell death. J. Cell Biol. 165, 347–356 (2004).
de Castro, I. F. Reovirus forms neo-organelles for progeny particle assembly within reorganized cell membranes. mBio 5, e00931-13 (2014).
Marciniak, S. J. et al. CHOP induces death by promoting protein synthesis and oxidation in the stressed endoplasmic reticulum. Genes Dev. 18, 3066–3077 (2004).
Sacaan, A. I. et al. CDK4/6 inhibition on glucose and pancreatic beta cell homeostasis in young and aged rats. Mol. Cancer Res. 15, 1531–1541 (2017).
Rane, S. G. et al. Loss of Cdk4 expression causes insulin-deficient diabetes and Cdk4 activation results in β-islet cell hyperplasia. 22, http://genetics.nature.com (1999).
Zeng, L. et al. XBP-1 couples endoplasmic reticulum stress to augmented IFN-β induction via a cis-acting enhancer in macrophages. J. Immunol. 185, 2324–2330 (2010).
Martinon, F., Chen, X., Lee, A.-H. & Glimcher, L. H. Toll-like receptor activation of XBP1 regulates innate immune responses in macrophages. Nat. Immunol. 11, 411–418 (2010).
Eckard, S. C. et al. The SKIV2L RNA exosome limits activation of the RIG-I-like receptors. Nat. Immunol. 15, 839–845 (2014).
Barber, G. N. STING: infection, inflammation and cancer. Nat. Rev. Immunol. 15, 760–770 (2015).
Elion, D. L. et al. Therapeutically active RIG-I agonist induces immunogenic tumor cell killing in breast cancers. Cancer Res. 78, 6183–6195 (2018).
Jiang, X. et al. Intratumoral delivery of RIG-I agonist SLR14 induces robust antitumor responses. J. Exp. Med. 216, 2854–2868 (2019).
Smith, J. A. et al. Endoplasmic reticulum stress and the unfolded protein response are linked to synergistic IFN-β induction via XBP-1. Eur. J. Immunol. 38, 1194–1203 (2008).
Harding, H. P. et al. Diabetes mellitus and exocrine pancreatic dysfunction in perk/ mice reveals a role for translational control in secretory cell survival. Mol. Cell 7, 1153–1163 (2001).
Woo, C. W. et al. Adaptive suppression of the ATF4-CHOP branch of the unfolded protein response by toll-like receptor signalling. Nat. Cell Biol. 11, 1473–1480 (2009).
Liu, Y.-P. et al. Endoplasmic reticulum stress regulates the innate immunity critical transcription factor IRF3. J. Immunol. 189, 4630–4639 (2012).
Carletti, T. et al. Viral priming of cell intrinsic innate antiviral signaling by the unfolded protein response. Nat. Commun. 10, 3889 (2019).
Hu, F. et al. ER stress and its regulator X-box-binding protein-1 enhance polyIC-induced innate immune response in dendritic cells. Eur. J. Immunol. 41, 1086–1097 (2011).
Johnson, D. B. et al. Melanoma-specific MHC-II expression represents a tumour-autonomous phenotype and predicts response to anti-PD-1/PD-L1 therapy. Nat. Commun. 7, 10582 (2016).
Dowless, M. et al. Abemaciclib is active in preclinical models of Ewing sarcoma via multipronged regulation of cell cycle, DNA methylation, and interferon pathway signaling. Clin. Cancer Res. 24, 6028–6039 (2018).
Smith, C. C. et al. Endogenous retroviral signatures predict immunotherapy response in clear cell renal cell carcinoma. J. Clin. Investig. 128, 4804–4820 (2018).
Smith, C. C. et al. Alternative tumour-specific antigens. Nat. Rev. Cancer 19, 465–478 (2019).
Cañadas, I. et al. Tumor innate immunity primed by specific interferon-stimulated endogenous retroviruses. Nat. Med. 24, 1143–1150 (2018).
Frankish, A. et al. GENCODE reference annotation for the human and mouse genomes. Nucleic Acids Res. 47, D766–D773 (2019).
Nakagawa, S. & Takahashi, M. U. gEVE: a genome-based endogenous viral element database provides comprehensive viral protein-coding sequences in mammalian genomes. Database 2016, baw087 (2016).
Käll, L., Canterbury, J. D., Weston, J., Noble, W. S. & MacCoss, M. J. Semi-supervised learning for peptide identification from shotgun proteomics datasets. Nat. Methods 4, 923–925 (2007).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 13, 731–740 (2016).
Perez-Riverol, Y. et al. The PRIDE database resources in 2022: a hub for mass spectrometry-based proteomics evidences. Nucleic Acids Res. 50, D543–D552 (2022).
Hulstaert, N. et al. ThermoRawFileParser: modular, scalable, and cross-platform RAW file conversion. J. Proteome Res. 19, 537–542 (2020).
Röst, H. L. et al. OpenMS: a flexible open-source software platform for mass spectrometry data analysis. Nat. Methods 13, 741–748 (2016).
Eng, J. K., Jahan, T. A. & Hoopmann, M. R. Comet: an open-source MS/MS sequence database search tool. Proteomics 13, 22–24 (2013).
Kim, S. & Pevzner, P. A. MS-GF+ makes progress towards a universal database search tool for proteomics. Nat. Commun. 5, 5277 (2014).
Paqes, J., Pavlíqek, A. & Paqes, V. HERVd: database of human endogenous retroviruses. Nucleic Acids Res. 30, http://www.girinst.org (2002).
Andreatta, M. & Nielsen, M. Gapped sequence alignment using artificial neural networks: application to the MHC class i system. Bioinformatics 32, 511–517 (2016).
Acknowledgements
We thank Oncolytics Biotech for providing the reovirus. Research described in this manuscript is funded by the CRUK programme (grant number CRM183X, A.M.). V.R. is supported by the Mark Donegan Foundation. M.McL. received funding from the Oracle Cancer Trust. K.J.H. is supported by the RM/ICR NIHR Biomedical Research Centre. We thank Dr Y. Miwa (Tsukuba University) and Dr O. Kanagawa (RCAI, RIKEN) and Dr. M. Tomura (Osaka Ohtani University) for the Kaede mice, and Harriet Whittock and Jacob Van Vloten for help with in vivo work produced for this study, and Mark Allen and Peter John-Baptiste for monitoring mice.
Author information
Authors and Affiliations
Contributions
V.R., J.K.-C., A.R., J.C., C.J.L., A.M., M.McL., K.J.H. contributed to the conception and design of the research as well as in writing the manuscript; V.R., J.C.-K., L.Y., A.B.-A., M.M., R.E., D.M., V.J., F.B. and E.A. performed the experiments and acquired data; J.W., M.M., J.C., R.E., C.J.L., D.M., R.C., A.R., T.T., P.M., P.N., R.V., H.P., A.M., M.McL. and K.J.H. contributed to the analysis and interpretation of the data; J.W., R.E., C.J.L. and R.C. provided support for bioinformatics analysis; C.C., H.B., M.P., S.F., I.D., E.C.P., A.W. and R.V. performed and supported in vivo experiments; N.M. contributed to the design and processing of the RNA sequencing experiments; K.J.H. is the guarantor, senior author and provided funding for the project. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Judith Smith, Samuel Workenhe and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Roulstone, V., Kyula-Currie, J., Wright, J. et al. Palbociclib and dsRNA sensor co-operate to enhance anti-cancer effects through ER stress and modulation of immune evasion. Nat Commun 16, 4855 (2025). https://doi.org/10.1038/s41467-025-60133-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-60133-5