Abstract
Cubic phase lipid nanoparticles (cubosomes) show promise as carriers for nucleic acids due to their outstanding delivery performance and physical stability. However, both theoretical analysis and experimental results have shown that encapsulating long RNA is almost impossible while maintaining a cubic structure. Here we show successful encapsulation of long RNA (up to 4000 bases) in cubosomes by premixing RNA and lipids before self-assembly. Surprisingly, we discovered that long RNA within cubosomes folded into an isotropic phase, similar to other lipid self-assembly structures encapsulating nucleic acids. The cubosome–long RNA complex maintained excellent delivery efficiency even after 24 days at room temperature, underpinning its exceptional stability. Our premixing strategy not only demonstrates the feasibility of developing cold chain-free mRNA vaccines, but also offers insights for incorporating large cargo into other liquid crystals, such as peptides and/or polymer materials.
Similar content being viewed by others
Introduction
The 2023 Nobel Prize in Physiology or Medicine, awarded for the development of mRNA vaccines, has underscored the need for a safe and effective vector for long RNA1. Among many materials, lipids unequivocally stand out as the most promising due to their excellent biocompatibility and long-term safety2,3,4. Currently, the only viable lipidic vector for long RNA is lipid nanoparticles (LNPs)2. Despite their recent success, the amorphous structure of LNPs causes leakage and subsequent degradation of RNA at room temperature, essentially mandating cold chains for storage and distribution4,5. In addition, the fabrication hysteresis of LNPs leads to varied structures reported in the literature6,7, thereby impeding a fundamental understanding of the endosomal escape process. Therefore, in addition to the ongoing trial-and-error approach based on lipid synthesis and/or the optimization of LNP composition, strategies are needed to break through the current challenges posed by the materials themselves.
Cubosomes are thermodynamically stable LNPs with an inverse bicontinuous cubic structure. For approximately 40 years, lipid cubic phases have attracted considerable interest in the field of cellular delivery due to their excellent physical stability and endosomal escape performance8,9,10,11,12,13,14,15. In particular, the cubosome is known as topologically active, meaning that it can spontaneously fuse with the endosomal membrane due to its positive Gaussian modulus (\(\bar{\kappa }\) > 0)8,15. Recently, the intrinsic material benefits of cubosomes have led researchers to design lipidic vectors with cubic phases6,15,16,17,18.
Unfortunately, the small pore size of cubosome limits its use in the delivery of long RNA. Some studies, including ours, have shown that swelling the pore diameter is feasible19,20,21. However, the swollen cubic phase is either unstable or can only exist in the bulk phase. Therefore, loading long RNA inevitably shifts the phase to an inverted hexagonal or multilamellar phase which shows low stability or delivery efficiency. In general, liquid crystals are the result of equilibrium among fundamental interactions. This means that even a small inclusion of external molecules can lead to complete structural changes22. Therefore, the encapsulation of long RNA within cubosomes without sacrificing its cubic phase has been perceived as a formidable task.
In this work, we introduce a premixing strategy to encapsulate long RNA into cubosomes while maintaining the cubic phase. We prepared lipid–RNA precursors to localize long RNA in small lipid droplets, thereby minimizing the impact of the RNA on the overall structure of the cubosomes. The combination of in situ SAXS and Cryo-EM tomography studies unveils that long RNA within cubosomes folds into an isotropic phase. Furthermore, the cubosome–long RNA complex shows excellent and consistent delivery efficiency even after 24 days of storage at room temperature, highlighting its potential as a next-generation vector for long RNA.
Results and discussion
Premixing strategy for long RNA encapsulation in cubosomes
Encapsulating highly charged long RNA into cubosomes is a challenging task. This is because long RNA alters the equilibrium curvature of the lipid bilayer at both local and global levels, leading to structural changes. In fact, a complete structural transition or coexistence of biphasic structures has been reported in all previously studied cases (Supplementary Table 1)15,23. In biphasic cases, it is unknown whether long RNA exists within the cubic structure. Hence, the question remains regarding whether long RNA, including 1 kb RNA molecules (Rg ~ 10 nm), can be loaded into a cubosome without structural transformations (Fig. 1a).
a Schematic illustration of the challenge of long RNA encapsulation within the cubosomes. Whether long RNA larger than the unit cell of the cubosome can be loaded into the cubosome remains a materials challenge. b The premixing strategy was divided into three steps: microfluidic premixing of lipids and long RNA, droplet fusion, and self-assembly. Lipid droplets formed in the microfluidic mixer undergo fusion into intermediate structures. During the subsequent annealing process, lipids gradually reorganize, ultimately forming a highly ordered cubosome–long RNA complex (QllP, RNA).
We hypothesize that the formation of lipid–RNA precursors prior to self-assembly can confine the structural effects of the RNA on the final lipid assembly. To test our hypothesis, we utilized the droplet fusion method developed in our previous study10. Figure 1b illustrates the premixing strategy for encapsulating long RNA in cubosomes. Briefly, lipids in ethanol and RNA in DEPC water are introduced into each inlet of a staggered herringbone mixer (SHM). Within the microfluidic channel, cationic lipids and long RNA aggregate via electrostatic interactions to form lipid–RNA precursors. Because single-stranded RNA naturally exists in a folded state, the RNA solution was heated above the melting temperature to increase the flexibility of the RNA chain24. Flexible RNA chains can maximize interactions with lipids, which aids in the formation of uniform lipid–RNA precursors. Afterward, the precursor solution undergoes a mild thermal annealing process to evaporate the ethanol and promote self-assembly. It should be noted that the formation of cubosomes does not proceed via a simple stacking of lipid droplets. Instead, lipid molecules gradually reorganize through fusion and dynamic rearrangement, ultimately forming highly ordered cubosomes with embedded RNA.
Strikingly, the cubosome–long RNA complexes produced by the premixing strategy were able to retain a highly ordered cubic phase (Supplementary Fig. 2a). In contrast, the conventional postmixing method (Supplementary Fig. 1) shows a complete structural transition to the hexagonal phase as reported previously (Supplementary Fig. 2b)15,25. In fact, the influx of long RNA is known to disfavor highly curved structures and reduce higher-order reflections, resulting in a phase transition8,22. These findings clearly demonstrate that a premixing strategy is a simple and powerful approach for incorporating long RNA into cubosomes. In this study, we focused on understanding the structure of cubosome–long RNA complexes from a materials perspective.
Mechanism of cubosome–long RNA complex formation
To understand the underlying mechanism of long RNA encapsulation in the premixing strategy, we conducted an in situ structural study of cubosome–long RNA complex formation. Figure 2a shows Cryo-TEM images obtained for each collected solution at various annealing time points. At the initial time point (t = 0 min), small droplets containing the lipid–RNA precursors were found. During the annealing process (t = 3 min), lipid droplets not only fused with each other but some particles had already self-assembled with periodic internal cubic structures. At full treatment (t = 6 min), highly ordered cubosome–long RNA complexes were found (additional Cryo-TEM image is available in Supplementary Fig. 3). Six minutes of annealing was unexpectedly fast, as it corresponds to almost half the time required for cubosome formation without long RNA. The shortened annealing time is thought to result from the negatively charged long RNA accelerated membrane fusion and self-assembly, as the driving force of higher-order self-assembly is the release of counterions26. This is an important finding that strongly supports the presence of long RNAs, as it was not observed in cubosomes without long RNA10.
a In situ Cryo-TEM images of the cubosome–long RNA complexes obtained at different annealing time points, demonstrating the mechanism of cubosome–long RNA complex formation (Scale bar, 100 nm). b, c Particle characterization. DLS results showing particle size before and after self-assembly (b) and Zeta potential of QllP, LIPID and QllP, RNA (c). Data are presented as mean ± SE. Data were obtained from three biological independent experiments (n = 3). d In situ SAXS profile at different annealing time points. Cubosome–long RNA complex exhibits rapid self-assembly as observed in Cryo-TEM study and invariant peak at q = 0.12 \({\text{\AA} }^{-1}\) (black arrow). e Super-resolution confocal microscope images showing the colocalization of cubosomes and long RNAs. Pearson’s coefficient was 0.981, indicating a high degree of colocalization. Similar results were obtained across three independent experiments. Scale bar is 10 µm. f Z-stack image of cubosome–long RNA complex showing yellow colors, indicating the long RNA is localized inside the cubosome. Texas Red for lipids (Red), FAM for long RNA (green).
Cubosome–long RNA complex formation was also monitored through dynamic light scattering, which provides information on the average particle size. The initial size of the droplet was about 39 nm, and the cubosome–long RNA complex (QllP, RNA) was about 140 nm. The increased cubosome size compared to that of RNA-free cubosomes (QllP, LIPID, about 100 nm) is thought to be the result of reduced charge repulsion between droplets mediated by the RNA, leading to an increased number of fused droplets per cubosome (Fig. 2b). Notably, the cubosomes exhibit a net positive surface potential, facilitating interaction with the negatively charged endosomal membrane (Fig. 2c).
In situ SAXS was also performed to explore the structural development during the formation of cubosome–long RNA complexes. At the 3 min annealing point, the SAXS scan revealed seven sharp Bragg peaks with relative q positions at ratios of \(\sqrt{2},\sqrt{4},\sqrt{6},\sqrt{10},\sqrt{12},\sqrt{14}\), and \(\sqrt{18}\), which is consistent with the well-ordered primitive cubic phase (Im3m). As expected, the crystallinity increased with annealing time (Fig. 2d). Notably, unlike other peaks, we observed a characteristic peak at q = 0.12 \({\text{\AA} }^{-1}\), the position of which was invariant during the annealing process (black arrow). Previous studies have reported similar peaks in other lipid mesophases, where the peak was assigned to the inter-nucleic acid spacing27,28. Therefore, the appearance of a characteristic peak at 0.12 \({\text{\AA} }^{-1}\) in the cubosome–long RNA complex is attributed to the periodic arrangement of the long RNA within the cubosome. Moreover, the increase in peak intensity during the annealing process suggested that the ordering of long RNA within the cubic structure increased as self-assembly progressed.
The presence of long RNA within cubosomes was studied using confocal microscopy. Cubosomes and RNAs were tagged with red and green fluorescent dyes, respectively. 2D and 3D images of the cubosome–long RNA complexes showed that most of the particles were yellow, indicating the colocalization of lipids and RNA (Fig. 2e, f). The Pearson correlation coefficient, which explains the correlation between lipids and RNA, indicated a very strong correlation with a value of 0.981. This finding implies that most long RNA is localized inside the cubosomes. In addition to the fluorescence images, the encapsulation efficiency was also confirmed by Ribogreen assay. The loading capacity was estimated to be approximately two mRNA molecules per cubosome (Supplementary Fig. 4a), which is comparable to that of conventional LNPs29. Furthermore, the integrity of the long RNA was confirmed by gel electrophoresis (Supplementary Fig. 4b, c), and the results demonstrated that encapsulation of long RNA within the cubosomes was almost 100%. The acceleration of cubic structure formation, the appearance of a characteristic peak, and the colocalization of lipids and RNA clearly indicate that the premixing strategy can preserve the cubic structure of the cubosome–long RNA complex.
Structure of cubosome–long RNA
Understanding the structure of RNA in the cubosome can provide valuable insights into how this structure can be maintained. The structural effect of RNA in cubosomes was analyzed via SAXS (red lines, Fig. 3a). The cubosome–long RNA complex revealed a shift in the diffraction peaks toward higher q-values (Fig. 3b). Compared to the cubosome-only sample, the lattice parameter decreased from 174\({{\text{\AA}}}\) to 151\({{\text{\AA}}}\). The radius of the water channel was also confirmed to be smaller in the cubosome–long RNA complex (\({r}_{w}^{{cubosome}-{RNA\; complex}}=28{{\text{\AA}}}\)) than in the cubosome-only sample (\({r}_{w}^{{cubosome}}=35{{\text{\AA} }}\))30. The decreases in the lattice parameter and the water channel radius appear to be caused by the electrostatic interaction between the cationic lipid (DOTAP) and the negatively charged long RNA8,9,19.
a 2D X-ray scattering pattern showing the presence of a characteristic peak in QllP, RNA marked with red dotted line (top). b Integrated SAXS scan data showing a highly ordered primitive cubic structures for both QllP, LIPID and QllP, RNA. The \({q}_{{RNA}}\) peak was only observed in QllP, RNA and assigned to a structure formed by the long RNAs. c Schematic illustration of the cubosome–long RNA complex representing the isotropic phase of RNA in a cubosome. Long RNA exhibits a d-spacing of 51 \({{\text{\AA}}}\) and domain size of 550 \({{\text{\AA} }}.\) d Reconstructed 3D structure of QllP, RNA showing long RNA encapsulated inside the cubosome in red (d, top). The FFT of sliced images at location 1 and 2 indicates the local structure near the long RNA (d, bottom). Scale bar, 100 nm. Cryo-TEM images showed comparable results from seven biologically independent experiments. Tomographic analysis was performed three times. e Iso-surface rendering of the reconstructed cubosome–long RNA complex showing distinct differences of internal structure induced by long RNA. f Electron density profiles across the regions containing RNA (top) and without RNA (bottom) indicating that the dimension affected by RNA is approximately 50 nm.
In the cubosome–long RNA complex sample, another notable difference was the presence of a characteristic peak at q = 0.12 \({\text{\AA} }^{-1}\) (Fig. 3b arrow, Fig. 3a red dashed line). Similar to our SAXS data, previous studies demonstrated that a \({q}_{{DNA}}\) peak was observed near 0.12 \({\text{\AA} }^{-1}\) due to the intercalation of double-stranded nucleic acid in lamellar26,28,31,32 and inverted hexagonal lipid mesophases27,32,33. Based on our SAXS results, we interpret the local conformation of the long RNA within the cubosome as an isotropic phase with short-range order (Fig. 3c). Notably, the characteristic peak we discovered, \({q}_{{RNA}}\), likely arises from the folded structure of RNA instead of inter-RNA arrangements. This interpretation is supported by the relatively small persistence length of single-stranded RNAs, which allows for flexible conformations and enables interaction with both the hydrophilic and hydrophobic moieties of lipids34. Consequently, RNA molecules are presumed to be spatially confined within the water channels of the cubosome, exhibiting a d-spacing of 51 \({{\text{\AA} }}\). The domain size was calculated to be 550 Å. Importantly, this d-spacing was much smaller than the inter-water channel distance of the cubic phase, but close to the persistence length of RNA. This similar length scale supports the interpretation of an isotropic RNA arrangement with localized order, rather than a well-defined secondary structure such as inverted hexagonal packing.
Next, we investigated the local structure of the cubosome influenced by the long RNAs. We performed cryo-electron tomography to directly visualize the local structure. Through 3D rendering of the cubosome–long RNA complex, we determined the position of the long RNA within the cubosome (Supplementary Fig. 5 and Supplementary video 1). A 3D image was reconstructed showing the volumes of the cubosome–long RNA complex (blue) and long RNA (red) (Fig. 3d top). In the reconstructed volume, two slices at different heights were prepared that represent the regions with (slice 1) and without (slice 2) long RNA (Fig. 3d bottom). The red boxes shown in the slice images indicate X–Y regions containing long RNA. The Fast Fourier transformation (FFT) of the red box in slice 2 showed a primitive cubic phase (yellow circles), whereas that of slice 1 showed a disordered topology without a periodic lattice structure. The contrast in the internal crystallinity of the region where the long RNA is located is more clearly displayed by the iso-surface rendering of the reconstructed cubosome–long RNA complex (Fig. 3e and Supplementary videos 2 and 3). These results demonstrate that long RNA is confined to a small region, rather than affecting the overall topology of the cubosome–long RNA complex.
To understand detailed structural effects of long RNA on cubic phase, local electron density profiles from two slice images were studied. In slice 1, water channels (yellow square) were distributed between lipid bilayers (blue) with high electron density, showing a repetitive and regular order with an average spacing of 14.3 nm. Interestingly, a region showing a distorted structure was found presumably due to RNA inclusion (Fig. 3f, top red panel). In contrast, slice 2 (the region without long RNA) showed high periodicity throughout the region (Fig. 3f, bottom), indicating a clear difference in ordering with and without long RNAs. The lattice parameter and domain size obtained from tomography closely matched those from SAXS (15 and 55 nm, respectively). More data from other directions are shown in Supplementary Fig. 6.
Effect of RNA length on the structure of the cubosome
The sizes of the mRNAs used for the CRISPR/Cas9 system and COVID-19 vaccines range from 2000 to 4000 bp. To expand the application spectrum of cubosome–long RNA complexes, it is necessary to explore the effect of RNA length on the formation of cubosome–long RNA complexes. We varied the RNA length from 1 kb to 4 kb. Different lengths of RNA were loaded into cubosomes through the premixing method, and SAXS scans were performed. All the samples prepared for SAXS had the same nucleotide-to-lipid (NP) ratio. Impressively, the cubosomes maintained the primitive cubic phase even with RNA lengths up to 4 kb, as shown in Fig. 4a. As seen by others, RNA encapsulation screens charge repulsion between positively charged lipids, leading to a decrease in the lattice parameter (Fig. 4b)8,10. Encapsulation of longer RNA further decreases the lattice parameter, but the change is not significant and plateaus at 15 nm.
a SAXS scan results of the cubosome–long RNA complexes with different RNA lengths. b Lattice parameters of cubosomes with various RNA lengths. c The characteristic peaks from different RNA lengths after polynomial background subtraction. d Average domain size derived from the FWHM of the \({q}_{{RNA}}\) peaks shown in (c).
The presence of \({q}_{{RNA}\,}\) peak was consistent regardless of RNA length (Fig. 4a, red box). Encapsulation of a 4 kb RNA into cubosomes led to broadening of the peak with a slight increase in the peak position (Fig. 4c). This indicates a decrease in ordering with smaller domain size32. However, the retention of the 4 kb \({q}_{{RNA}}\) peak demonstrates that long RNAs remained aligned within the cubosome. Compared with shorter RNAs, the 4 kb RNA had a smaller domain size, indicating that increased RNA length reduced long-range ordering (Fig. 4d). The d-spacing of 1 kb RNA was 51 \({{\text{\AA} }}\), while that of 4 kb RNA was 48 \({{\text{\AA} }}\), indicating a slight decrease in inter-RNA spacing. Our findings highlight that the premixing strategy allows encapsulation of up to 4 kb of RNA and 80 μg of RNA content (Supplementary note 1 and Supplementary Fig. 7) in the cubosome, suggesting that cubosomes are a promising vector for mRNA therapy.
Delivery performance of the cubosome–long RNA complex
The long RNA delivery performance of cubosomes was investigated under various storage conditions. Physical stability of cubosome–long RNA complex was first evaluated. As expected from crystallin nature, no structural changes in \({q}_{{RNA}}\) positions were observed in the samples refrigerated for 24 days ensuring the long-term stability of the complex (Fig. 5a). As shown in Fig. 5b, the cubosome–long RNA complexes still retained the primitive cubic phase even after storage for 24 days at various temperatures. The colloidal stability of the cubosome–long RNA complex was also demonstrated by measuring the particle size every 7 days after storage at room temperature, and no changes were observed in terms of size after 24 days (Fig. 5c).
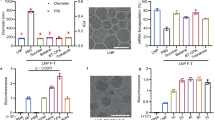
a–d Structural stability of the cubosome–long RNA complex. The SAXS study at various storage period (a) and temperature (b) shows no structural changes. In addition, no changes in the size of QllP, RNA (c) or leakage of long RNA from QllP, RNA (d) were found, indicating the excellent physical stability of the cubosome–long RNA complex. e Cell viability test of cubosome–long RNA complexes at 0.5 µg/mL concentration. The average value of the control group is set to 100%. Data are shown as mean values with SE and all data were obtained from three biologically independent experiments (n = 3). f Evaluation of the cellular delivery efficiency of the cubosome–long RNA complexes. Cubosome–long RNA complexes were more effective than LNPs in delivering 1 kb. Data are shown as mean value (mean ± SE, n = 3; biological independent experiments). g The delivery performance was consistent even after storage at RT for 24 days (mean ± SE, n = 3; biological independent experiments). Statistical analysis was performed using ordinary one-way ANOVA followed by Tukey’s multiple comparisons test; no significant differences were observed (p = 0.9798). h Visualization of the cubosome–long RNA complex and LNP in transfected HepG2 cells after 24 h. Lipids and mRNAs are shown in green (NBD-PE) and red (Cy5), respectively. Scale bar 25 µm. i, j Fluorescence intensites of cells containing lipids (i) and mRNAs ( j ) were quantified from (h) (n = 3; individual cells pooled from independent replicates). k Protein expression image of cubosome-eGFP RNA complex transfection at 24 h post-treatment in HeLa cells. Green fluorescent image and merged with the bright field image are shown respectively. Scale bar 200 µm. l, m, Total fluorescence intensity of cells (l) and percentage of GFP expressing cells (m) were quantified from (k). All data are presented as mean ± SE, n = 3. Statistical analysis was performed using ordinary one-way ANOVA and Tukey’s multiple comparisons test (**p = 0.0041).
RNA can easily leak or degrade after being stored at room temperature for a long period of time5. Therefore, gel electrophoresis was performed after the cubosome–long RNA complexes were stored for 24 days at room temperature. Remarkably, no sub-bands were observed in the gel, implying that RNA leakage did not occur even after storage at room temperature for 24 days (Fig. 5d, Lane 6). These results demonstrate that the cubosome–long RNA complexes effectively protect the long RNAs from degradation without the need for polymeric additives. These findings suggest that cubosomes are a promising candidate to address the physical stability issues of LNP–mRNA vaccines.
To determine whether cubosome–long RNA complex is capable of delivering long RNA, we treated various cell lines with the cubosome–long RNA complex and mRNA loaded LNP. After 24 h of incubation, both the cubosome–long RNA complex and the LNP showed excellent cellular viability (Fig. 5e). Dose-dependent evaluation showed that cubosome–long RNA complexes had no significant effect on the cellular viability up to a concentration of 0.5 µg/mL mRNA (Supplementary Fig. 8). Therefore, all subsequent studies were conducted at an mRNA concentration of 0.5 µg/mL. Quantitative reverse transcription polymerase chain reaction (qRT‒PCR) revealed that the level of eGFP mRNA (1 kb) was significantly greater in the cubosome–long RNA complex than in the LNP in all cell lines assessed (Fig. 5f).
Transfection efficiency varies among different cell types, and cells that are generally refractory to transfection, such as those with low proliferation rates or high sensitivity to exogenous materials, pose significant challenges for nucleic acid delivery. Hepatic cells, in particular, play a central role in detoxifying foreign substances, and hepatocellular carcinoma (HCC) cells often exhibit resistance to conventional therapies. This characteristic extends to in vitro models such as HepG2 and LX2 cells35,36, which are commonly used to study HCC-related pathologies and therapeutic strategies37,38. Despite these challenges, the cubosome–long RNA complexes exhibited relatively high transfection efficiency in hepatic cell lines, including HepG2 (a hepatoma cell line) and LX2 (a hepatic stellate cell line), compared to LNPs. Given that these cell types are typically difficult to transfect39, our results underscore the potential of cubosome–long RNA complexes as promising delivery vehicles.
Notably, in the case of longer RNA molecules such as Cas9 mRNA (4.4 kb), the intracellular RNA levels were approximately four-fold higher in the cubosome–long RNA complexes–treated group compared to the lipoplex group (Supplementary Fig. 9), further supporting the effective delivery capability of cubosome–long RNA complexes for large RNA cargos. Moreover, to evaluate the storage stability of the cubosome–long RNA complexes, we stored the cubosome–long RNA complex at room temperature for 4 weeks and treated the cells. Remarkably, mRNA delivery efficiency remained stable for up to 24 days (Fig. 5g), suggesting that cubosome–long RNA complexes can preserve mRNA integrity and function even after being stored at room temperature for an extended period of time. Collectively, these findings demonstrate the potential of cubosome–long RNA complexes for robust and stable RNA delivery, particularly in challenging cell types such as hepatic cells.
Next, we fabricated a cubosome–long RNA complex and LNP using fluorescently labeled mRNA (Cy5) and lipid (NBD-PE) to measure the amount of each LNP inside the cells. Twenty-four hours post-treatment, stronger fluorescent signals from both lipids and mRNA were observed in cells treated with the cubosome–long RNA complex than in those treated with the LNP (Fig. 5h–j and Supplementary Fig. 10). Taken together, these results show that the cubosome–long RNA complex can deliver long RNA and protect the RNA from degradation even under room temperature storage conditions, indicating that the cubosome–long RNA complex has the potential to be a vector for long RNA.
To establish cubosome as effective mRNA delivery systems, delivered mRNA needs to undergo translation into functional protein. Accordingly, we examined whether the improved mRNA delivery efficiency of cubosomes is directly associated with elevated levels of protein expression. At 24 h after transfection, cubosome–long RNA complex treated cells exhibited elevated total fluorescence intensity and a higher percentage of GFP-expressing cells compared to other groups (Fig. 5k–m). These findings suggest that cubosome–long RNA complexes can be used as a more uniform and efficient means of mRNA delivery.
Since various studies on LNP have reported that LNP release RNA into the cytosol via endo/lysosomal pathways, we determined whether cubosomal mRNA delivery occurs via endo/lysosomal pathways similar to those of LNPs. To test this possibility, we performed live-cell imaging using PE-tagged cubosome and lysotracker (Supplementary Fig. 11a). The fluorescence intensity of cubosome increased rapidly during the first hour; however, they gradually decreased over time after initial increase (Supplementary Fig. 11b). Concurrently, co-localized cubosome–long RNA complexes with lysosome increased up to 4 h (Supplementary Fig. 11c). Taken together, these data suggest that cubosome–long RNA complexes enter cells and travel through the lysosomal pathway, releasing their encapsulated mRNA.
In summary, we reported a premixing strategy that enables the encapsulation of long RNA within a structured lipid nanoparticle. The premixing of lipids and RNA localizes long RNA within the confined volume of lipid droplets and facilitates their assembly into cubosome–long RNA complexes. Interestingly, long RNA dramatically accelerates self-assembly kinetics. In the cubosome–long RNA complex, in-depth structural studies discovered that RNA adopts an isotropic phase within the cubosome. Remarkably, our cubosome–long RNA complex preserved its structure without degradation of long RNA even after 24 days of storage at room temperature. The successful delivery of long RNA, including 4 kb RNA, and its high transfection efficiency into cancer cells and hepatic stellate cells highlight the potential of using cubosome–long RNA complexes as next-generation vectors for long RNA.
Methods
materials
1,2-dioleoyl-3-trimethylammonium-propane (chloride salt); DOTAP, 1,2-dioleoyl-sn-glycero-3-phosphocholine; DOPC, 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine-N-[methoxy(polyethylene glycol)−2000] (ammonium salt); DOPE-PEG2000, cholesterol(ovine), 1,2-dihexadecanoyl-sn-glycero-3-phosphocholine; DPPC, 1,2-dipalmitoyl-sn-glycero-3-phosphoethanolamine-N-(7-nitro-2-1,3-benzoxadiazol-4-yl) (ammonium salt): NBD-PE were obtained from Avanti Polar Lipids. Glycerol monooleate (GMO) was purchased from Nu-chekprep (USA). GMO and cholesterol were dissolved in fresh chloroform solvent (Sigma-Aldrich) at the desired concentration. Heptadecan-9-yl 8-((2-hydroxyethyl)(6-oxo-6-(undecyloxy)hexyl)amino)-octanoate; SM-102 was acquired from Echelon Biosciences. Texas Red-DHPE, RNA markers, Formaldehyde Load dye, and DEPC-treated water were bought from Invitrogen. GelRed Nucleic Acid Gel stain was obtained from biotium. CleanCap® EGFP mRNA (5moU) was customized from TriLink, and other kinds of RNA were purchased from Miltenyi Biotec and Sigma-Aldrich.
For in vitro assays, Dulbecco’s modified Eagle’s medium (DMEM) and phosphate-buffered saline (PBS) were purchased from Welgene, fetal bovine serum (FBS) from Cytiva, ZellShield from Minerva biolabs. Trizol was obtained from Ambion. LIVE/DEAD™ Viability/Cytotoxicity Kit, SYBR Green Real-Time PCR Master Mix, Opti-MEM, and NucBlue™ Live ReadyProbes™ Reagent were obtained from Thermo Fisher Scientific and ReverTra Ace™ qPCR RT Master Mix was from TOYOBO. N¹-methylpseudouridine-5′-triphosphate was customized from TriLink. T7 RNA polymerase mix, ATP/CTP/GTP, 3′-O-Me-m7G(5’)ppp(5’)G RNA Cap Structure Analog, DNase I, RNA Cleanup Kit, and E. coli poly(A) Polymerse were bought from New England Biolabs.
Synthesis of cubosome–long RNA complex
SHM device was used for synthesis of cubosome–long RNA complex in this study. The lipid feeding solution with GMO: DOTAP: PEG2000 = 95: 4: 1 mol% was prepared in ethanol (0.15 M total lipid concentration). The molar ratio of each lipids was kept constant in all formulations. 20 µg of RNA was dissolved in DEPC water and heated before use (65 °C/5 min). Cubosome–long RNA complex was formulated using a syringe pump (NE-1000, New Era Pump Systems, NY, USA) with flow condition 0.05 mL/min and flow rate ratio 3. Collected droplet were transferred into the round-bottom flask and then self-assembled while a rotary evaporator operated at P = 75 mbar and T = 57 °C. For structural analysis, cubosome–long RNA complexes were stored at different temperatures (−20/4/23 °C) and periods (0/7/15/24 days). For cubosomes, the lipid composition and synthesizing conditions were identical to those of cubosome–long RNA complexes, and the water feeding solution was prepared with absence of RNA.
For in vitro experiments, LNPs were prepared with different lipid compositions (SM-102: DPPC: PEG2000: cholesterol = 50: 10: 1.5: 38.5 mol%), flow condition 1 mL/min and chaotic microfludic mixer. liposomes were prepared with DOPC instead of GMO (DOPC: DOTAP: PEG2000 = 95: 4: 1 mol%). Except for the lipids used, all conditions for lipid nanoparticle formulation are the same as those for cubosomes/cubosome–long RNA complexes.
Cryogenic transmission electron microscopy (Cryo-TEM) and 3D tomography
The cubosome–long RNA complex morphology was observed by using Cryo-TEM (FEI Tecnai F20 G2 at 200 kV). To prepare samples for Cryo-TEM study Lacey carbon grids were glow-discharged (15 s, 25 mA) in Cressington 108 (Cressington Scientific Inc., USA). A volume of 2.3 µl of the sample was applied on the grid, which was then blotted for 3.0 s and plunge-frozen in liquid ethane to rapidly freeze the sample using a Vitrobot FP5350/60 (FEI, USA). All experimental procedures were conducted under cryogenic conditions (<−178 °C). Images were taken under cryogenic conditions with Cryo-electron microscopy Tecnai F20 G2 (FEI, USA) at accelerating voltage of 200 kV and RIO 16 camera (GATAN, USA). For 3D tomography, single-axis tilt series were acquired at each position of interest at 14,000 × g magnification from −42° to +40° tilt angles with a step size of 1°. Tomography acquisitions were performed using the Latitude T software. Tomographic reconstruction was performed using the IMOD software package v. 4.11.24. The thickness of slice was 1.17 nm. The Gatan program was used for the Fast Fourier Transform (FFT).
Small-Angle X-ray Scattering (SAXS)
Synchrotron SAXS was performed at Beamline 4 C of Pohang Accelerator Laboratory. The sample-to-detector distance was 1 m which the accessible q-range from 0.03 to 0.7 Å−1. All the samples were measured at 25 °C after equilibrium with exposure time of 30 s. The synchrotron X-ray source had an average photon energy of 16.9 keV with an average beam size of 100 μm × 300 μm (V × H). The scattered photons were recorded by using a Rayonix 2D SX 165 CCD detector (Rayonix, USA) and 2D X-ray diffraction images were integrated using the custom software provided by the beamline 4C40. For in vitro mRNA expression experiments, custom-built lab SAXS was used. The X-ray source had an average photon energy of 8.0478 keV with a average beam size of 1000 µm × 1000 µm (V × H).
Super-resolution microscopy
For super-resolution microscopy (Carl Zeiss, Elyra7) imaging, we designed primer that perfectly matched the 1 kb RNA used in the experiment (seq. [FAM] TGG GGG TGT TCT GCT GGT AGT GG). The prepared primer was annealing with the 1 kb RNA and dialysis was performed for 24 h to remove primer that did not pair with the 1 kb RNA. The primer was conjugated with FAM dye, and Texas Red-DHPE lipid was used to distinguish cubosomes from RNA. The excitation/emission spectra of the primer and Texas Red-DHPE are 498 nm/517 nm and 595 nm/613 nm, respectively. The cubosome–long RNA complexes (QllP, RNA) were prepared by mixing GMO: DOTAP: PEG2000: Texas Red-DHPE at a molar ratio of 94.5: 4: 1: 0.5, respectively.
DLS
The size, polydispersity index (PDI), and zeta potential of the samples were determined by using a dynamic light scattering instrument (Zetasizer Nano ZSP, Malvern, UK). The presented data corresponded to the mean of three successive measurements. The PDI was taken as the width of the particle size distribution and was calculated by using Origin program.
Agarose gel electrophoresis
1% agarose gels containing 10 µl of GelRed Nucleic Acid stain was prepared. For RNA markers and RNA, formaldehyde loading dye was essential and the procedure suggested in the protocol was followed. Samples included RNA markers were loaded into each well and electrophoresed with autoclaved 1× TAE running buffer at 100 V for 40 min. To identify the long RNA integrity, Trizol mixture (Trizol: chloroform: isopropanol = 25: 24: 1 v/v%) and sample were used in 1:1 ratio to crush the cubosome–long RNA complex. Subsequently, centrifuge (14,000 × g, 4 °C, 15 min) was conducted and then supernatant was collected for electrophoresis. Supernatant solutions have to be mixed with loading dye before loaded into the well. The stained RNA bands were visualized and photographed using Shenhua SH-523 multifunctional imager. The RNA markers are a set of 10 transcripts with lengths of 0.5, 1, 1.5. 2, 2.5, 3, 4, 5, 6, and 9 kb.
Cell culture
HEK293T, HeLa, and HepG2 cells were maintained in DMEM (Welgene, Gyeongsan, Korea) containing 10% (v/v) FBS (Cytiva, Incheon, Korea) and 1% ZellShield (Minerva biolabs, Berlin, Deutschland). LX2 cells were cultured in DMEM with 2% FBS and 1% ZellShield. HCT116 cells were cultured in Rosewell Park Memorial Institute (Welgene) containing 10% (v/v) FBS and 1% ZellShield. Cells were grown at 37 °C in the presence of 5% CO2 according to suppliers’ instructions.
Transfection
Before 24 h of transfection, cells were plated in clear-bottom 6-well plates or glass bottom 35 mm 4-well confocal dish. The number of cells seeded per well varied by plate type: 3.5 × 105 cells in 6-well plate and 3.5 × 10⁴ cells in confocal dish. Before 30 min of transfection, old media was replaced with fresh media, and transfection mixture was prepared by mixing cubosome–long RNA complex or lipoplex with Opti-MEM(ThermoFisher) to a concentration of 7 µg/ml. Transfection mixture was added to cultured cells drop by drop to a final concentration of 0.25 to 1 µg/ml(0.5 to 2 µg of mRNA in 2 ml of 6-well plate and 0.25 µg of mRNA in 500 µl of 4-well confocal dish). Positive control for GFP expression test, Polyetherimide (PEI) was pre-mixed with mRNA at a ratio of 1.5:1. PEI mixture was treated to cells in the same amount of mRNA as in other lipid-mRNA complex. After 4 h, cells were washed with PBS (Welgene) 3 times and fresh media was added.
Cell viability assay
Cell viability was determined by LIVE/DEAD™ Viability/Cytotoxicity Kit (Thermo Fisher Scientific, Seoul, Korea). Before cell viability assay, calcein/EthD-1 mixture was prepared by adding them into PBS, to a concentration of 1 µM. After being treated with cubosome–long RNA complex or LNP-RNA complex (0.25–1 µg/ml) and incubated for 24 h, medium was removed and cells were washed with PBS. Calcein/EthD-1 mixture was added to each well and after 30 min of incubation at room temperature.
RNA isolation and qRT–PCR
Transfected cells were resuspended in 500 µl of Trizol (Ambion, Carlsbad, CA), and incubated for 5 min at 4 °C for lysation. 100 µl of chloroform were added to the mixture, and centrifuged at 15,000 × g for 15 min at 4 °C. Transparent supernatant was collected as RNA-dissolved solution. RNA was precipitated by adding 250 µl of absolute isopropanol. After centrifugation at 15,000 × g for 10 min at 4 °C, the supernatants were discarded, and the pellet were washed in 75% (v/v) ethanol with several times of inverting. RNA pellet were centrifuged at 10,000 × g for 10 min at 4 °C, and supernatants were discarded. Remaining supernatants were dried in room temperature for 10 min, and pellet were diluted in RNase-free water.
cDNA was synthesized from 1 µg of total RNA using ReverTra Ace™ qPCR RT Master Mix(TOYOBO, Seoul, Korea). qRT–PCR was performed using SYBR Green Real-Time PCR Master Mix(Thermo Fisher Scientific) in qTOWER³. The human GAPDH gene was used as the internal control for normalization. Sequence of qRT–PCR primer are shown in Supplementary Table 2.
In vitro transcription
20 µl transcription reactions containing T7 RNA polymerase mix (New England Biolabs, Ipswich, MA) and 120 ng DNA template, supplemented with 7.5 mM ATP/CTP (New England Biolabs), 7.5 mM N¹-methylpseudouridine-5′-triphosphate (TriLink, San Diego, CA), 2 mM GTP (New England Biolabs), and 8 mM 3′-O-Me-m7G(5′)ppp(5′)G RNA Cap Structure Analog (New England Biolabs) were incubated for 24 h at 37 °C. To remove DNA template, 2 µl of DNase I (New England Biolabs) was added to transcription reactions. After incubation for 15 min, RNA was purified using Monarch® RNA Cleanup Kit (New England Biolabs).
For RNA poly A tailing, 10 ng of in vitro transcribed-RNA was added to 20 µl of E. coli poly(A) Polymerse (New England Biolabs) reactions and incubated for 30 min at 37 °C. Final RNA production was purified using Monarch® RNA Cleanup Kit.
Image acquisition and quantification
To quantify the extent of intracellular cubosome–long RNA complex, cells were imaged using Leica DMi8 fluorescence microscope (Leica Microsystem, Wetzlar, Germany). Before imaging, NucBlue™ Live ReadyProbes™ Reagent (Thermofisher) was treated to the cell. Laser and detector settings were kept the same while imaging. Images were randomly acquired from 3 to 5 different fields of each samples. For cell counting and diameter measurement, LasX software and Photoshop CS6 (Adobe) were used. ROIs were manually selected and intensity of each fluorescent channel was quantified in Image J software (National Institutes of Health, Bethesda, MD, USA).
To quantify GFP expression cell, Incucyte® S3 live-cell analysis system (Satorius, Goettingen, Germany) was used. Cells were imaged in phase contrast channel and green channel using 4× and 10× objective in Incucyte S3 software (IncuCyte 2019B Rev2), after 24 h after transfection. Cell number and GFP intensity were quantified using Aivia software (DRVision Technologies). Each images were taught in pixel classifier and identically applied.
For live cell imaging, 1× of NucSpot® Live 488(Biotium, Fremont, CA) and 50 nM of Lysotracker™ Deep Red(ThermoFisher) were added to cells before 10 min of transfection. Incubator T Series(LCI, Namyangju, Korea) were used for maintain cells at 37 °C in a humidified atmosphere of 5% CO2 for 24 h. Live cell images were acquired using a Leica STELLARIS Confocal Microscope (63× oil-immersion objective; 1024 × 1024 pixels; 2D acquisition). Fluorescence intensity and area for each channel were quantified using Fiji software, based on Image J.
Statistics and reproducibility
Unless otherwise specified, all statistical analyses were conducted using data collected from at least three biological independent replicates. Information on biological replicates and statistical analyses is provided in the method and figure legends.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data in this study are included in the main manuscript and the Supplementary Information. Furthermore, except for image data, the source data for Figs. 1–5 are available in the Source Data file. Source data are provided with this paper.
References
Callaway, E. & Naddaf, M. Pioneers of mRNA COVID vaccines win medicine Nobel. Nature 622, 228–229 (2023).
Verma, M. et al. The landscape for lipid-nanoparticle-based genomic medicines. Nat. Rev. Drug Discov. 22, 349–350 (2023).
Yin, H. et al. Non-viral vectors for gene-based therapy. Nat. Rev. Genet. 15, 541–555 (2014).
Hou, X., Zaks, T., Langer, R. & Dong, Y. Lipid nanoparticles for mRNA delivery. Nat. Rev. Mater. 6, 1078–1094 (2021).
Schoenmaker, L. et al. mRNA-lipid nanoparticle COVID-19 vaccines: structure and stability. Int. J. Pharm. 601, 120586 (2021).
Cárdenas, M., Campbell, R. A., Yanez Arteta, M., Lawrence, M. J. & Sebastiani, F. Review of structural design guiding the development of lipid nanoparticles for nucleic acid delivery. Curr. Opin. Colloid Interface Sci. 66, 101705 (2023).
Viger-Gravel, J. et al. Structure of lipid nanoparticles containing siRNA or mRNA by dynamic nuclear polarization-enhanced NMR spectroscopy. J. Phys. Chem. B 122, 2073–2081 (2018).
Leal, C., Bouxsein, N. F., Ewert, K. K. & Safinya, C. R. Highly efficient gene silencing activity of siRNA embedded in a nanostructured gyroid cubic lipid matrix. J. Am. Chem. Soc. 132, 16841–16847 (2010).
Kim, H. & Leal, C. Cuboplexes: topologically active siRNA delivery. ACS Nano 9, 10214–10226 (2015).
Kim, H., Sung, J., Chang, Y., Alfeche, A. & Leal, C. Microfluidics synthesis of gene silencing cubosomes. ACS Nano 12, 9196–9205 (2018).
Abourehab, M. A. S. et al. Cubosomes as an emerging platform for drug delivery: a review of the state of the art. J. Mater. Chem. B 10, 2781–2819 (2022).
Varghese, R., Salvi, S., Sood, P., Kulkarni, B. & Kumar, D. Cubosomes in cancer drug delivery: a review. Colloids Interface Sci. Commun. 46, 100561 (2022).
Karami, Z. & Hamidi, M. Cubosomes: remarkable drug delivery potential. Drug Discov. Today 21, 789–801 (2016).
Dyett, B. P., Yu, H., Strachan, J., Drummond, C. J. & Conn, C. E. Fusion dynamics of cubosome nanocarriers with model cell membranes. Nat. Commun. 10, 4492 (2019).
Zheng, L. I., Bandara, S. R., Tan, Z. I. & Leal, C. Lipid nanoparticle topology regulates endosomal escape and delivery of RNA to the cytoplasm. Proc. Natl. Acad. Sci. USA 120, e2301067120 (2023).
Yu, H. et al. Formation of particulate lipid lyotropic liquid crystalline nanocarriers using a microfluidic platform. J. Colloid Interface Sci. 634, 279–289 (2023).
El Mohamad, M. et al. Regulating the structural polymorphism and protein corona composition of phytantriol-based lipid nanoparticles using choline ionic liquids. J. Colloid Interface Sci. 657, 841–852 (2024).
Yu, H. et al. Inverse cubic and hexagonal mesophase evolution within ionizable lipid nanoparticles correlates with mRNA transfection in macrophages. J. Am. Chem. Soc. 145, 24765–24774 (2023).
Tyler, A. I. I. et al. Electrostatic swelling of bicontinuous cubic lipid phases. Soft Matter 11, 3279–3286 (2015).
Zabara, A. et al. Design of ultra-swollen lipidic mesophases for the crystallization of membrane proteins with large extracellular domains. Nat. Commun. 9, 544 (2018).
Kim, H., Song, Z. & Leal, C. Super-swelled lyotropic single crystals. Proc. Natl. Acad. Sci. USA 114, 10834–10839 (2017).
Förster, S. et al. Order causes secondary Bragg peaks in soft materials. Nat. Mater. 6, 888–893 (2007).
Angelov, B. et al. DNA/fusogenic lipid nanocarrier assembly: millisecond structural dynamics. J. Phys. Chem. Lett. 4, 1959–1964 (2013).
Becskei, A. & Rahaman, S. The life and death of RNA across temperatures. Comput. Struct. Biotechnol. J. 20, 4325–4336 (2022).
Leal, C. et al. Stacking of short DNA induces the gyroid cubic-to-inverted hexagonal phase transition in lipid-DNA complexes. Soft Matter 9, 795–804 (2013).
Rädlr, J. O., Koltover, I., Salditt, T. & Safinya, C. R. Structure of DNA-cationic liposome complexes: DNA interaction in multilamellar membranes in distinct interhelical packing regimes. Science 275, 810–814 (1997).
Arteta, M. Y. et al. Successful reprogramming of cellular protein production through mRNA delivered by functionalized lipid nanoparticles. Proc. Natl. Acad. Sci. USA 115, E3351–E3360 (2018).
Patel, S. et al. Naturally-occurring cholesterol analogues in lipid nanoparticles induce polymorphic shape and enhance intracellular delivery of mRNA. Nat. Commun. 11, 983 (2020).
Li, S. et al. Payload distribution and capacity of mRNA lipid nanoparticles. Nat. Commun. 13, 5561 (2022).
Pilkington, C. P. et al. A microfluidic platform for the controlled synthesis of architecturally complex liquid crystalline nanoparticles. Sci. Rep. 13, 12684 (2023).
Kulkarni, J. A. et al. On the formation and morphology of lipid nanoparticles containing ionizable cationic lipids and siRNA. ACS Nano 12, 4787–4795 (2018).
Pattipeiluhu, R. et al. Liquid crystalline inverted lipid phases encapsulating siRNA enhance lipid nanoparticle mediated transfection. Nat. Commun. 15, 1303 (2024).
Aburai, K., Hatanaka, K., Takano, S., Fujii, S. & Sakurai, K. Characterizing an siRNA-containing lipid-nanoparticle prepared by a microfluidic reactor: small-angle x-ray scattering and cryotransmission electron microscopic studies. Langmuir 36, 12545–12554 (2020).
Giraud, L., Viricel, W., Leblond, J. & Giasson, S. Single stranded siRNA complexation through non-electrostatic interactions. Biomaterials 113, 230–242 (2017).
Xu, L. et al. Human hepatic stellate cell lines, LX-1 and LX-2: new tools for analysis of hepatic fibrosis. Gut 54, 142–151 (2005).
Song, Y. et al. Activated hepatic stellate cells play pivotal roles in hepatocellular carcinoma cell chemoresistance and migration in multicellular tumor spheroids. Sci. Rep. 6, 36750 (2016).
Gao, S. et al. Ex vivo gene delivery to hepatocytes: techniques, challenges, and underlying mechanisms. Ann. Biomed. Eng. 40, 1851–1861 (2012).
Zhang, B. et al. TLR3 activation of hepatic stellate cell line suppresses HBV replication in HepG2 cells. Front. Immunol. 9, 2921 (2018).
Yaghmur, A., Østergaard, J. & Mu, H. Lipid nanoparticles for targeted delivery of anticancer therapeutics: recent advances in development of siRNA and lipoprotein-mimicking nanocarriers. Adv. Drug Deliv. Rev. 203, 115136 (2023).
Kim, K. W. et al. Small-angle X-ray scattering beamline BL4C SAXS at Pohang Light Source II. Biodesign 5, 24–29 (2017).
Acknowledgements
This work was supported by the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT) (No. RS-2023-00209955, participants: H.J., J.P., J.C.P., S.B., and H.K.). This research was also supported by the Basic Science Research Program through the NRF, funded by the Ministry of Education (NRF-2022R1C1C1006015, participants: I.S., J.R., and J.H.K.), This research was partially supported by the Korea Institute of Science and Technology (KIST) (Institutional 2E32345 (participants: H.J., I.S., J.P., J.H.K., and H.K.), 2E33561 (participants: I.S., J.R., and J.H.K.), and 2E33111 (participants: H.J., J.P., J.C.P., S.B., K.H.L., Y.J., and H.K.). This research was supported by the Bio & Medical Technology Development Program of the National Research Foundation (NRF) funded by the Korean government (MSIT) (No. RS-2025-02309992, participants: I.S., J.R., and J.H.K.). This research was also partially supported by Korea Health Industry Development Institute (KHIDI) (Grant No. RS-2022-KH125686, participants: Jahyun Koo). This work was also partially supported by Korea Institute of Planning and Evaluation for Technology in Food, Agriculture and Forestry (IPET) through High-Risk Animal infectious Disease Control Technology Development Program, funded by Ministry of Agriculture, Food and Rural Affairs (MAFRA)(RS-2024-00396818, participants: H.J., I.S., J.P., J.C.P., S.B., J.R., J.H.K., and H.K.). This work was supported by UST Young Scientist+ Research Program 2023 through the University of Science and Technology (No. 2023YS23, participant: H.J.). We thank Dr. Yoon-ha Jang for technical assistance with the in vitro mRNA synthesis. We also thank Dr. Youli Li and Prof. Safinya for their insightful comments on the description of mRNA folding state.
Author information
Authors and Affiliations
Contributions
H.K. and J.H.K proposed the idea. H.J. designed and performed the overall experiments, analyzed the data under the supervision of H.K.; I.S. performed in vitro experiments and analysis under the supervision of J.H.K.; J.P. prepared cubosome samples and performed analysis for in vitro experiments. Cryo-TEM and tomography study were conducted by Y.S.; J.C.P. and S.B. contributed to the experiment of synchrotron SAXS and confocal microscopy imaging; J.R. contributed to the in vitro expression experiments. Y.J., J.K., and K.H.L. provided experimental advice. J.H.K. and H.K. provided funding support. H.J., I.S., J.H.K., and H.K. wrote the manuscript. All authors commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
Y.J., K.H.L., J.H.K., and H.K. are in the process of applying for Korean invention patents (10-2023-0015749 and 10-2023-0015776) related to the subject matter of this manuscript. Remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Jin, H., Seo, I., Park, J. et al. Premixing enables loading of long RNA in cubic phase lipid nanoparticles. Nat Commun 16, 5054 (2025). https://doi.org/10.1038/s41467-025-60380-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-60380-6