Abstract
Stress stimulation-mediated liquid-liquid phase separation is a key activity in living organisms, but its biophysical characteristics are poorly understood. Here, we report a UV-light stress stimulation behaviour in a binary community of synthetic protocells of condensates and proteinosomes, showing that condensates could behave like Condensate Pumps to enable a stepwise controlled transmembrane mass transfer regardless of the permeability barrier of proteinosomes. The stimulation mechanism of interfacial tension-induced proteinosome deformation and transient high osmotic pressure arisen by the dissociation of condensate is proposed. Accordingly, under UV-light stress stimulation, unexpected characteristics could be triggered by transmembrane pumping oversized biomacromolecules into proteinosomes including liquid-liquid reentrant phase separation, DNA unwinding, and protein synthesis. Therefore, our results not only reveal unique physical principles and potential characteristics of macromolecular assemblies at droplet-membrane interface but also highlight a pathway for transmembrane transport of biomacromolecules which is anticipated to serve as a powerful technique to inducing higher-order behaviour in synthetic protocells community.
Similar content being viewed by others
Introduction
In eukaryotic cells, membrane-less organelles (MLOs) such as processing bodies (P-bodies, or PBs), stress granules, are liquid-like condensates formed through liquid-liquid phase separation (LLPS)1,2,3 under various cellular stress conditions, such as oxidative, osmotic or heat-shock stress, and play a crucial role in organizing cellular components4, regulating messenger RNA (mRNA) translation and degradation5. Recent recognition that these liquid-like biomolecular condensates are ubiquitous within living cells with specific geographical characteristics and form a plethora of new liquid-liquid and liquid-solid interfaces with membranes or cytoskeletal systems6,7,8,9. Interactions at these local contact interfaces do mechanical work, impacting the local mechanics, morphology, organization and function of condensates10,11. For example, the dynamics of PBs and stress granules, might be regulated by endoplasmic reticulum contact sites, and the number of PBs affects the translational capacity of the endoplasmic reticulum12.
In this regard, recently, the emerging of synthetic protobiology which is concerned with the construction, reconstitution, and functionalization of individual cell-like entities, is providing a pathway for unraveling certain unknown biophysical behaviors or the potential contributions of the confining membrane by creating various living cell/synthetic cellular bionics. Currently, studies focusing on the role of condensates as biomolecular compartments (protocell models) explore contact-dependent and through-space chemical interactions as steps toward the development of higher-order cell-like dynamic behaviors such as fusing13,14, phagocytosing15,16, preying17, signal processing18,19,20,21,22, self-reconfiguring23,24, etc. These pioneered studies, along with diverse investigations on alternative protocell models have successfully demonstrated primarily integration of functional components into certain dynamic micro-compartmentalized constructs25. In contrast, the stress stimulation triggered unraveled characteristic of synthetic protocell populations still receives relatively little attention.
Here, inspired by the interaction between condensates and membrane organelles in biological systems, we address an interaction mode between two different populations of artificial cells (Fig. 1). Among this work, condensates are designed to be dynamically photo-responsive26,27,28, and proteinosomes of which membrane was cross-linked could withstand strong external interferences, such as osmotic pressure. We refer to the co-assembly of condensate and proteinosome as the interactive protocells and reveal interacting relationship between the two communities in two aspects. On the one hand, condensates were stable on the proteinosome membrane for a longer period of time. The condensates acted as a condensate pump to facilitate an oversize biomacromolecule transmembrane transfer driven by the local osmotic pressure with the transient dissociation under external stimuli, and proteinosomes as a guest achieved temporary storage of cargos. After external stimuli disappearing, proteinosomes enriched reassembled condensates in specific regions. On the other hand, the deformation of proteinosomes occurred under the hypertonic stimulation of condensates. A series of biological processes inside proteinosomes were triggered and controlled by the formation-dissociation process of condensates, such as reentrant phase separation, DNA unwinding and protein synthesis, which were accompanied by regular fluctuations of components inside proteinosomes. Thus, it is anticipated that our work should highlight a way for oversized biomacromolecules to transmembrane transport in synthetic biomimetic systems, and provides opportunities for the development of reconfigurable cytomimetic materials with structural, functional, and signal transduction complexity.
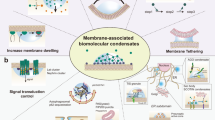
Azo Coas and proteinosomes are attracted by electrostatic interaction and embedded in the membrane due to interfacial tension (1). The dissociation and re-assembly of Azo Coas are reversible under UV (405 nm) and blue light (488 nm) (2). Azo Coas loaded with oversize biomolecules boosts transmembrane delivery (3, 4) accompanied with the emerging of reentrant phase separation inside proteinosomes encapsuling positive molecules (5). The reentrant condensates (Re Coas) have the opposite photo-responsive behavior to Azo Coas in proteinosomes (6, 7). Relying on the pumping capability of Azo Coas, the protocell system enables a range of biological processes, such as DNA unwinding and protein synthesis. Illustrations (8) of partial wetting with the contact angles θα + θβ + θγ = 360°, showing the force balance between the droplet interfacial tension Σαγ and the mechanical tensions Σαβ and Σβγ within the two membrane segments.
Results
Dynamic assembly of condensates around proteinosomes
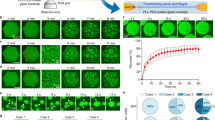
To achieve a type of stress stimulation responsive condensate, cationic azobenzene trimethylammonium bromide (AzoTAB) containing a quaternary ammonium group (Supplementary Fig. 2) and anionic succinyl dextran (Su-Dex) with different molecular weights (6 kDa, 70 kDa, 500 kDa, 1000 kDa, 2000 kDa) and degrees of substitution (DS) of ca.2 (Supplementary Fig. 3) at a AzoTAB/Su-Dex molar ratio of 3:1, were specially used to generate UV-light responsive condensates (Azo Coas). When there were more cis-AzoTAB, condensates dissociated; otherwise, condensates re-assembled26. And PEG-bis(N-succinimidyl succinate) (NHS-PEG16-NHS) cross-linked proteinosomes were prepared using bovine serum albumin (BSA)-polymer conjugates29 with molecular weight cut-off (MWCO) of the membrane approximately 70 kDa (Supplementary Fig. 5a–e). It maintained low permeability to macromolecules exceeding 150 kDa, of which the interception by proteinosomes could remain stable for 30 min (Supplementary Fig. 5f, g). Then the interactive protocells were prepared by adding condensates formed by Su-Dex500kDa and AzoTAB (Azo Coas500) into the proteinosome solution (Supplementary Fig. 1). Adhesion of condensates onto negatively charged proteinosome occurred at a wide range of pH, especially at pH 8.0, which was driven by electrostatic interaction (Fig. 2a1, a2, Supplementary Figs. 6, 7). Adhesion force at the condensate-membrane interface was much higher than 3.14 × 107 μN per unit area (μN/μm2) due to the liquid bridge broke at the far end of the interface (Fig. 2e, Supplementary Table 1. This value was calculated under the condition of a droplet interfacial tension γ of 1.6 × 10−5 N/m. See Methods for more details), when the greatest attraction (Fm) related to the fracture site of the liquid bridge could be calculated by the Drop Adhesion Force (DAF) module based on the structure and profile of the liquid bridge formed by condensed phase between the needle and membrane. The interactive protocell communities were further confirmed by flow cytometry (Fig. 2b–d). After mixing Azo Coas500 and proteinosomes, both forward scattered light (FSC-A) and side scattered light (SSC-A) were observed to increase compared with the single population, indicative of the increase in size and their complexity (Fig. 2b, d, Supplementary Fig. 8a–c). The red fluorescence representing Azo Coa500 population overlapped with the green fluorescence representing proteinosome population after co-incubation, demonstrating the formation of protocell communities (Fig. 2c, Supplementary Fig. 8d–f).
CLSM images of protocell communities consisting of Azo Coa500 and proteinosome populations (a1) and local magnification of their contact surface (a2). Red fluorescence, Azo Coas500 enriched with rhodamine B isothiocyanate labeled Su-Dextran500kDa (RBITC-Su-Dex500kDa); Green fluorescence: proteinosomes encapsulating fluorescein isothiocyanate isomer I labeled neutral dextran500kDa (FITC-Dex500kDa). 2D dot plots of FSC-A vs SSC-A (b) and FITC-H vs RBITC-H (c) collected from the mixed binary populations of Azo Coas500 and proteinosomes. d FSC-A histograms, determined from Supplementary Figs. 8a–c, of Azo Coa500 population, proteinosome popuplation and the mixed binary populations, respectively. e Drop adhesion force (DAF, Fa) between condensed phase and proteinosome membrane. θ (92.2°) represents the contact angle. CD represents the contact diameter. f In-situ CLSM 3D images of Azo Coa500 formation on the proteinosomes membrane after co-incubating for 0 min, 5 min, 10 min, 15 min, 20 min, and 25 min. Red fluorescence recorded the morphology, quantity, and localization of Azo Coas500. g In-situ CLSM 2D images of the formation and location of Azo Coas500 over time. g1 Preferential location of formed Azo Coas500 on proteinosomes. g2 Electrostatic adsorption between free condensates and proteinosomes. g3 Coalescence between membrane-fixed and free condensates. Scale bars in a1, a2, f, g1, g2 and g3, 10 μm. All relevant experiments in a1, a2, g1, g2 and g3 are performed independently at least three times with similar results. Source data are provided as a Source Data file.
Through in-situ confocal laser scanning microscope (CLSM) 3D imaging, the assembling procedure of Azo Coa onto proteinosomes was revealed. Initially, fewer Azo Coa500 formed on the surface of proteinosomes with small diameters, and then Azo Coa500 microdroplet diameters gradually increased. Meanwhile, new Azo Coas500 continued to generate on other empty parts of the proteinosome membrane, resulting in a gradual increase in the number of Azo Coa500 (Fig. 1f). Moreover, in-situ 2D observations further indicated that formed Azo Coas500 preferentially located on the membrane and underwent coalescence due to the high density and proximity on the membrane of proteinosomes, which substantiated by the red fluorescence enhancement on the membrane (Fig. 2g1). Besides, electrostatic adsorption between free droplets (Fig. 2g2) and the membrane, along with coalescence between membrane-fixed and free condensates (Fig. 2g3) were considered to be the other two factors contributing to the increases on the condensate number and diameter on the membrane of proteinosomes. Nevertheless, the number of membrane-adhered condensates increased slightly with an increase of proteinosome diameter, it was ultimately in the range of 4~6 on the same equatorial plane (Supplementary Fig. 9a, b). The membrane network restricted droplet fusion on the membrane, thereby achieving size control and structural stabilization of the membrane-adhered condensates (Supplementary Fig. 9c, d).
Stress stimulation-induced large deformation of proteinosomes and oversize biomacromolecule transmembrane transfer
It is widely recognized that the close relationship between membrane organelles and membrane-less organelles enables various cellular functions in living cells, just like the regulation between PBs and endoplasmic reticulum at their contact sites in human cells12,30. Correspondingly, we also observed an obvious invagination but no endocytosis of Azo Coa on the proteinosomes membrane caused by interfacial tension at contact sites (Fig. 2a2). Especially, when exposing the formed interactive protocells to ultraviolet light (405 nm, UV), conformation transition of AzoTAB from trans- to cis- status resulted in the rapidly dissociation of Azo Coas, and a much more notable proteinosome deformation was observed (Supplementary Fig. 10). The whole deformation-recovery process of proteinosomes was delineated into four states (Fig. 3a). Initially, condensates were tethered to proteinosomes and invaginated at their contact sites (i). Subsequently, after Azo Coa dissociation, morphological changes of proteinosomes became more and more obvious until the maximum deformation occurred (ii, iii). The proteinosome deformation finally returned to the initial state within a certain period of time (iv).
a Schematic illustration of cargo pumping and proteinosome deformation processes resulting from condensate dissociation: (i) Azo Coa formation onto proteinosomes caused membrane invagination; (ii) Proteinosomes deformation; (iii) The maximum deformation of proteinosomes; (iv) Deformation recovery of proteinosomes. b Diffusion trajectory of Su-Dex500kDa following the first dissociation of Azo Coas on the proteinosome. The symbol t0 represented the starting time of UV light (405 nm) irradiation; RBITC-Su-Dex500kDa, red fluorescence. Scale bars, 10 μm. c In-situ CLSM images of the repetitive pumping of Azo Coas1000. UV-blue light irradiation cycles were repeated 8 times (UV1/Vis1, UV2/Vis2, ∙∙∙∙∙∙, UV8/Vis8). Contrast changes in bright-field images indicated the formation and dissociation of Azo Coas1000 on proteinosomes; green fluorescence, proteinosome; red fluorescence, Azo Coa1000. Scale bars, 20 μm. d Fluorescence intensity curve of Su-Dex1000kDa in proteinosomes with recyclable Azo Coa1000 pumps. Arrows represent UV (purple, 405 nm) and blue (blue, 488 nm) light irradiation, respectively. e CLSM 3D images of proteinosomes after 8 injections of Azo Coas1000. Scale bars, 10 μm. Spatially positioned pumping capacity of Azo Coas proved by the fluorescence intensity ratios [I after pumping/I before pumping] of Su-Dex1000kDa (f1) in target proteinosome 1 and non-target proteinosomes 2, 3 and the corresponding CLSM images (f2) before and after pumping. Scale bars, 100 μm. Green fluorescence in c, FITC-Dex500kDa; red fluorescence in c, e, f2, RBITC-Su-Dex1000kDa. Data in f1 are presented as mean ± SD (n = 3 examined over 3 independent experiments). All relevant experiments in b, c are performed independently at least three times with similar results. Source data are provided as a Source Data file.
To be of great interest, a phenomenon that noticeable red fluorescence within proteinosomes was observed after condensate dissociation, though the molecular weight of Su-Dex500kDa was significantly higher than the MWCO (ca. 70 kDa) of proteinosomes, indicating that Azo Coa acted as a high-pressure pump to squeeze Su-Dex into proteinosomes (Fig. 3b, Supplementary Fig. 11). To further verify the pump-like property of Azo Coas, we utilized Su-Dex with a higher molecular weight of 1000 kDa to prepare interactive protocells. Azo Coas1000 also acted like a pump, capable of pumping Su-Dex1000kDa into proteinosomes (Fig. 3c, Supplementary Fig. 12a, b). Specially, under blue light (488 nm), the conformation transition of AzoTAB from cis- to trans- status allowed re-assembled Azo Coas1000 to be positioned on the proteinosome membrane. The enhanced contrast of the proteinosome membrane in bright-field images accompanied with the red fluorescence enhancement showing the generation of new Azo Coa1000 on the membrane, albeit with smaller droplet diameters (Fig. 3c). Accordingly, by controlling the switch of 405 nm and 488 nm light, a directed accumulated delivery of Su-Dex1000kDa was achieved. After 8 cycles, the fluorescence intensity of Su-Dex1000kDa within proteinosomes increased to 250% (from ca. 37.93 mM to 95.56 mM of Su-Dex1000kDa) of the initial state with 35% ~ 47% of pumping fraction (Fig. 3d, e and Supplementary Fig. 12c). Similarly, by using Su-Dex with different molecular weights, such directed pumping capacity of the formed condensates still well maintained (Supplementary Figs. 13, 14), and even Su-Dex with a molecular weight of 2000 kDa. While, higher molecular weight of Su-Dex, more pumping cycles from 6 cycles (500 kDa) to 13 cycles (2000 kDa), to complete the total delivery of cargos (Supplementary Figs. 12–14). Successful pumping of Su-Dex70kDa was also observed, though outflow occurred at the 4th cycle due to its lower molecular weight than MWCO (Supplementary Fig. 15). More importantly, amphiphilic biomacromolecule proteins, such as human serum albumin (HSA, PI 4.7~4.9), bovine hemoglobin (Hb, PI 6.8~7.0), and lysozyme (LZ, PI 10.5~11.35) from chicken egg white, which have different charge properties under experimental conditions (pH 8.0), were pumped into proteinosomes, showing a sustained increase in the fluorescence intensity inside proteinosomes and a slow decrease outside (Supplementary Fig. 16). The distribution ratios (Iin/Iout) of HSA, Hb and LZ eventually reached 2.01 ± 0.12, 1.66 ± 0.83, and 1.81 ± 0.90, respectively. AzoTAB was uniformly dispersed after pumping. The result was obtained by in-situ observation of 5-amino-fluorescein (5-AF), a positive small molecule green fluorescent dye with similar properties, in Azo Coas (Supplementary Fig. 17). Particularly, it should be mentioned that by controlling the lighting site, we could selectively pump targeted Azo Coas for spatially positioned pumping (Fig. 3f, Supplementary Fig. 18). CLSM images revealed that, in the same field of view, the pumping efficiency of the target Azo Coa100-proteinosome assembly (proteinosome 1, 144% from ca. 12.87 mM to 17.65 mM of Su-Dex1000kDa) was superior to non-pumped assemblies located 200~300 μm apart (proteinosome 2, 103% from ca. 12.33 mM to 12.85 mM; proteinosome 3, 105% from ca. 11.91 mM to 12.53 mM of Su-Dex1000kDa).
Considering the effect of membrane pore difference on membrane permeability, glutaraldehyde, a crosslinker with a shorter chain, was used to verify the pumping capacity of Azo Coa1000 across the glutaraldehyde cross-linked proteinosome membrane, though it had similar MWCO to the NHS-PEG16-NHS cross-linked membrane (Supplementary Figs. 5e, 19). The condensates were pumped 8 times outside the membrane, and the fluorescence intensity of RBITC-Su-Dex1000kDa in proteinosomes at the 8th cycle was twice that of the initial (Supplementary Fig. 19b), which was slightly lower than that of NHS-PEG16-NHS cross-linked protinosomes (Icycle8/I initial ~250%). Our results demonstrated the excellent transmembrane delivery capability of the condensate as a pump to the cross-linked membrane.
Pumping mechanism of condensates during stress stimulation
Amphiphilic small molecule AzoTAB usually binds proteins, which has the potential to change proteinosome membrane permeability. We found that AzoTAB slightly enhanced the membrane permeability of proteinosomes (Supplementary Fig. 5g, i). Nevertheless, the interception of macromolecules by proteinosomes mixed with AzoTAB remained stable for 30 min without external interference (Supplementary Fig. 5h, i). What′s more, AzoTAB with a high concentration of 20 mM did not cause proteinosome deformation (Supplementary Fig. 20). The transmembrane pumping of cargos required overcoming the membrane resistance, which caused proteinosome deformation and might be driven by the osmotic pressure difference caused by the transient dissociation of condensates releasing a high concentration of components. The micropipette aspiration technique31,32,33 was specially used to verify whether the pressure difference on both sides of the proteinosome membrane can drive the transmembrane movement of oversize biomolecules (Fig. 4a1), and the bending modulus (κ) and the area expansion modulus (K) were calculated. In detail, a micropipette with an inner diameter of 5 μm (Dp) was placed on the surface of FITC-Dex500kDa-encapsuled proteinosomes and the suction was gradually increased. During the aspiration process, the membrane segment inside micropipette experienced a suction of approximately 0 to 22 Pa per unit area (Pa/μm²), and the aspiration length (∆Lp) increased with increasing suction pressure (∆P) due to the membrane’s elastic (Fig. 4b, Supplementary Fig. 21a). After slowly reducing the suction pressure, a significant leakage of FITC-Dex500kDa was observed outside the proteinosome (Fig. 4c), demonstrating that applying a certain external force to the membrane could facilitate the transmembrane transport of oversize biomolecules exceeding the MWCO (70 kDa) of proteinosome membrane. In addition, it was observed that κ and K of the proteinosome membrane were independent of the contents within proteinosomes (Fig. 4d, Supplementary Fig. 21b). And the measured values of κ and K were significantly lower than those of the known giant unilamellar vesicles (GUVs)31, suggesting that the proteinosomes membrane was more flexible than lipid bilayers and was more prone to deformation in response to external environmental changes (Supplementary Table 2).
a1 Schematic illustration of proteinosome membrane properties determined by micropipette aspiration technique. The proteinosome was aspirated into a micropipette with increasing suction pressure (∆P). RP and RV0 denoted the micropipette and the proteinosome radii. As ∆P was gradually increased, the aspiration length (∆Lp) increased due to the membrane’s elastic response to the mechanical stress. L0 denoted the aspiration length when the minimum suction pressure was initially applied. ∆Lp at each ∆P were recorded. The elastic coefficient (EC), apparent area strain (αapp), the bending (κ) and apparent area expansion (Kapp) moduli of the proteinosome membrane were calculated. See Methods for more details. a2 Schematic illustration of micropipette-like properties of Azo Coas. Osmotic pressure difference (∆∏) generated by the transient dissociation of micropipette-like Azo Coas acted as the suction pressure (∆P). Invagination depth (∆Lc) was related to Azo Coa diameters (DAzo Coa). See Methods for more details. b Fluorescence images of the proteinosome encapsulating 10 mM FITC-Dex500kDa aspirated into a micropipette (2Rp = 5 μm) with increasing suction pressure. c Fluorescence images of the proteinosome with decreasing suction pressure. Scale bars in b, c, 50 μm. d Elastic coefficient-strain measurement for proteinosomes at 25 °C, corresponding the data in (b). The blue dots denoted the natural ln (EC) against αapp. A linear fit was made. The red dots denoted EC against αapp, which is nearly linear. κ and Kapp of the proteinosome were measured to be 3.7303 × 10−20 J and 4.2950 mN/m, respectively. e Schematic illustration of directional transmembrane transport of oversize biomolecules driven by transient osmotic pressure differences at membrane invagination sites. Invagination width (Dp) was defined as x∙DAzo Coa, where x (taken as 1.05) represented the expansion coefficient of Azo Coas upon dissociation. ∆Lc of Azo Coas did not exceed DAzo Coa, and ∆Lc = x ∙ 1/2 DAzo Coa was used for data analysis in our work. See Methods for more details. All relevant experiments in (c) are performed independently at least three times with similar results. Source data is provided as a Source Data file.
Driven by the interfacial tension, which did mechanical work such as partial to complete wetting transitions11, budding34, and membrane tube formation35, condensates ultimately adhered to the cross-linked proteinosome membrane with the invagination depth (∆Lc) not exceeding the diameter of condensed microdroplets (DAzo Coa) (Supplementary Fig. 22a). The complete, transient dissociation by an individual Azo Coa under UV light (405 nm) resulted in the osmotic pressure differences of approximately 854 to 1616 Pa, which was obtained by formula (12) (Supplementary Tables 3, 4, Supplementary Figs. 23, 24). The osmotic pressure differences per unit area were about 28.26 to 61.27 Pa/μm2 at the contact surface when the invagination depth (∆Lc) was x · 1/2 DAzo Coa (Fig. 4a2, Supplementary Figs. 22b, 25, Supplementary Table 5), where x is the expansion coefficient (taken as 1.05, x represents the volume expansion degree of condensates before the transient dissociation). This result was consistent with the conclusion of the micropipette aspiration experiment shown above. The osmotic pressure difference generated by the dissociation of an individual condensate far exceeded the suction pressure provided during our micropipette aspiration, thereby offering a sufficient force for the directional transmembrane transport of cargos loaded within condensates. Furthermore, membrane invaginations (∆Lc = x · 1/2 DAzo Coa) under osmotic pressure produced varying degrees of apparent area strains (αapp) ranging from 5.51 × 10−3 to 7.86 × 10−3 J (Supplementary Table 5), which was attributed to the reduction in membrane density, subsequently facilitating oversize biomolecules enter proteinosomes at the moment of membrane strain at invagination regions. It was also confirmed by the failed influx of Su-Dex500kDa in the proteinosomes without Azo Coas (Supplementary Fig. 26).
Therefore, our results confirmed that the pumping of condensates occurred through invaginations at contact sites due to interfacial tension difference. High osmotic pressure differences resulting from the dissociation of Azo Coas drove cargos to overcome the membrane resistance and transmembrane transport into proteinosomes, as well as induce proteinosome deformation. The concentration of dextran encapsuled in proteinosomes and the diffusion of cargos into the surrounding environment gradually reduced pressure differences on both sides of the membrane, until proteinosomes were at the maximum deformation state. In addition, transmembrane transport required that the osmotic pressure outside proteinosomes was greater than that inside, so it was reasonable to speculate that the transmembrane transport only occurred between the initial and the maximum deformation of proteinosomes (Fig. 3a ii, iii). The deformation recovery was due to the hyperosmotic conditions inside proteinosmes caused by the concentrated dextran until reaching the initial state.
Stress stimulation induced reentrant phase separation
Intracellular MLOs rapidly exchange components with the surrounding environment by virtue of their fluid properties and display dynamic patterns of assembly and dissolution in a multi-component mixed system, ultimately enabling precise regulation of various biological processes such as enzyme reaction regulation36,37,38,39 and translation inhibition40. Here, we created a micron-scale, tractable, minimalistic ternary mixing system constructed utilizing the pumping of Azo Coas, consisting of the pumped AzoTAB, Su-Dex, and cationic quaternized dextran with a molecular weight of 500 kDa (Q-Dex500kDa, DS of 0.25) encapsulated in proteinosomes (Fig. 5a, Supplementary Figs. 4, 27). Trans-AzoTAB and Su-Dex are known to undergo LLPS via multivalent interactions at low cis-AzoTAB concentrations under blue light (488 nm), reentering into a phase-separated regime consisting of Q-Dex and Su-Dex from a well-mixed state at high cis-AzoTAB concentrations under UV light (405 nm) (Fig. 5b, c). The composition and spatial organization of biomolecular condensates are ultimately controlled by the nature of intermolecular interactions between trans-AzoTAB and Su-Dex as well as their interactions with Q-Dex41.
a Schematic illustration of reentrant LLPS mediated by Azo Coa pumps. b In-situ CLSM images of Azo Coa500 pump-induced recyclable reentrant LLPS in proteinosomes. The entire UV-blue light irradiation cycles were repeated 6 times (UV1/Vis1, UV2/Vis2, ∙∙∙∙∙∙, UV6/Vis6). Azo Coa500 dissociation and reentrant condensate (Re Coa) formation under UV light (405 nm) occurred, while Azo Coa500 regeneration and Re Coa dissociation occurred under blue light (488 nm). Scale bars, 20 μm. c Fluorescence colocalization of Q-Dex500kDa and Su-Dex500kDa inside proteinosomes, indicating Re Coas were formed by Q-Dex500kDa and Su-Dex500kDa. d Binding affinity between Su-Dex500kDa and trans-AzoTAB (left), Q-Dex500kDa (right), respectively. See Methods and Supplementary Table 6 for specific parameters and results. e CLSM images of fluorescence localization of Re Coas on the proteinosome membrane at different profiles after reducing Azo Coa500 to the total concentration of AzoTAB and Su-Dex mixture of 6 mM. Scale bars in (c, e), 10 μm. f Fluorescence intensity curve of Su-Dex500kDa in proteinosomes during the process of Azo Coa pumping-proteinosome membrane deformation-internal reentrant LLPS. Arrows represent UV (purple, 405 nm) and blue (blue, 488 nm) light irradiation, respectively. Segment ab, compression and decompression of the contents resulting from proteinosome deformation (enhanced fluorescence) and deformation recovery (decreased fluorescence); segment bc, the release of Su-Dex (enhanced fluorescence) caused by Re Coa dissociation and Azo Coa500 regeneration on the membrane (decreased fluorescence) under blue light; segment cd, the enrichment of Su-Dex (enhanced fluorescence) caused by Azo Coa500 dissociation and Re Coa regeneration. Fluorescence accumulated continuously within proteinosomes due to the repetitive pumping. Green fluorescence in (b, c, e), FITC-Q-Dex500kDa; red fluorescence in (b, c), RBITC-Su-Dex500kDa. All relevant experiments in (e) are performed independently at least three times with similar results. Source data are provided as a Source Data file.
Earlier studies suggested that a system is said to be undergoing a reentrant phase transition (RPT) if a monotonic variation of any thermodynamic field results in two (or more) phase transitions, and it finally attains a state which is macroscopically similar to the initial state or the system reenters the original state42. Most of the studies on reentrant phase separation have been regulated through a monotonic variation of ions43, RNA44, and salt45 concentration. In contrast, reentrant phase behaviors in this work were achieved by transiently changing AzoTAB concentration through light stimulation.
When the interactive protocell was first exposed to UV light (405 nm), negative RBITC-Su-Dex could be pumped into proteinosomes, where positive FITC-Q-Dex encapsulated in the proteinosome exhibited fluorescence localization on the membrane due to electrostatic interaction (Fig. 5b). Then, upon switching between UV light (405 nm) and blue light (488 nm), Azo Coas continuously pumped Su-Dex into proteinosomes, leading to a gradual accumulation of Su-Dex. Compared with proteinosomes encapsuling neutral Dex, Q-Dex might accelerate the enrichment of Azo Coa on the membrane due to the electrostatic interaction with Su-Dex (Supplementary Fig. 17b). Finally, with the dissociation of Azo Coas, reentrant phase separation between accumulated Su-Dex and Q-Dex within proteinosomes could occur under UV light (405 nm) (Fig. 5c). These new, small reentrant condensates (Re Coas) gradually increased in size over time and sedimented at the bottom of the proteinosome cavity due to gravitational forces. Surprisingly, Re Coas showed an opposite photo-responsive behavior to Azo Coas. Under UV light (405 nm), due to the competition affinity between AzoTAB and Q-Dex towards Su-Dex, Azo Coas dissociated and Re Coas formed, while Azo Coas regenerated and Re Coas underwent dissociation under blue light (488 nm) (Fig. 5b, Supplementary Fig. 28). Isothermal titration calorimetry (ITC) was employed to reveal a stronger affinity of trans-AzoTAB/Su-Dex than Q-Dex/Su-Dex (Fig. 5d, Supplementary Table 6). However, no formation of Re Coas was observed under UV light (405 nm), and Q-Dex was excluded from Azo Coas under blue light (488 nm) after mixing Azo Coas with Q-Dex without proteinosomes under the same experimental conditions (Supplementary Fig. 29). Only when the concentration of Su-Dex was doubled, the opposite photo-responsive behavior between Azo Coa and Re Coa occurred. The affinity competition between trans-AzoTAB and Q-Dex was the essence of the opposite photo-responsive behavior, and the continuous pumping of Azo Coas to reach the concentration threshold of Su-Dex for triggering reentrant phase separation was the other prerequisite.
To be of great interest, the position of formed Re Coas within proteinosomes could be regulated by adjusting the amount of Su-Dex pumped by Azo Coas (Fig. 5b, Supplementary Figs. 30, 31). As the total concentration of Azo Coa decreased from 16 mM to 6 mM, Re Coas experienced two position states of stable distribution within the proteinosome cavity (Fig. 5b) and transfer to the inner membrane (Fig. 5e). The transfer time was reduced from 3 min to about 0.5 min at the 3rd cycle (Supplementary Figs. 30, 31) due to the positive charges provided by excess Q-Dex in Re Coas. Reentrant droplets induced by the lower Azo Coa concentrations were visually smaller, and more pumping times were required to achieve size growth of small Re Coas. A series of biological processes, such as transmembrane signal transduction induced by condensate dissociation, the formation, growth, relocation, and dissociation-regeneration of reentrant droplets, were composition-dependent, which was consistent with a periodic fluctuation of Su-Dex within the proteinosome (Fig. 5f, Supplementary Fig. 30d).
Stress stimulation-mediated biological processes
Stress stimulation is a normal phenomenon that could disrupt homeostasis in living cells accompanied with physiological and behavioral change. Alternatively, to further demonstrate such studied UV light-based stress stimulation could also be utilized as an effective strategy for regulating certain biological reactions in the constructed artificial cell communities, DNA unwinding and controlled synthesis of green fluorescent protein (GFP) were specially shown (Fig. 6a). In detail, a small quantity of negatively charged DNA labeled by SYBR Green (SYBR Green-DNA) and Q-Dex500kDa were co-encapsulated in proteinosomes. DNA unwinding process can be monitored by the fluorescence intensity emitted by SYBR Green. Due to its electrostatic attraction with Q-Dex500kDa, DNA was specifically enriched into the reentrant condensates (Supplementary Fig. 32). RBITC labeled DNase (RBITC-DNase, 31 kDa) capable of uncoiling double-stranded DNA was loaded into Azo Coas which then could be sequestered into Re Coas after each pumping (Fig. 6b, c, Supplementary Fig. 33). After pumping DNase once and thrice by Azo Coas, DNA within proteinosomes was uncoiled to different degrees within 15 min, with the latter uncoiling 2.14 times as fast as the former (Fig. 6a, d). Despite the accumulation of DNase within proteinosomes being 140% and 154% of the initial state during the 1st and 3rd cycles (Supplementary Fig. 33), respectively, it was evident that a greater number and size of reentrant droplets were formed after the 3rd cycle, accelerating the reaction rate of DNA unwinding (Fig. 6c). These results demonstrated the capability of Azo Coas to control the target components and its regulation ability over biological processes within the assemblies.
a Schematic illustration of DNA unwinding and green fluorescent protein (GFP) synthesis controlled by condensate pumps (Azo Coas500) for transmembrane transporting DNase and plasmid GFP DNA. In the DNA unwinding process, the amount of DNase31kDa pumped into proteinosome was adjusted by the number of pumping cycles (1 or 3 times). During the GFP synthesis, proteinosomes encapsulating E. coli lysate containing a large amount of transcription and translation-related factors and Q-Dex500kDa were co-incubated with Azo Coas500 loaded with plasmid GFP DNA. GFP DNA was pumped into proteinosomes, thereby constructing a cell-free protein expression system within protocell communities. In- situ CLSM images of DNA unwinding after once pumping (b), thrice pumping (c) of Azo Coas500. DNA is labeled with SYBR Green I (SYBR Green-DNA, green fluorescence); RBITC-DNase, red fluorescence. Scale bars, 25 μm. d Fluorescence intensity changes of DNA unwinding induced by DNase for once pumping (circles, blue line) and thrice pumping (squares, green line) of Azo Coas500. Control group (rhombuses, gray line) shows fluorescence quenching of SYBR Green-DNA without DNase. e Fluorescence intensity at different positions of proteinosomes after GFP synthesis. Box plots show the mean line (midline), 25/75 percentiles (boxes), and SD (whiskers). Data are represented as mean ± SD (n = 5). Positions 1~4 were shown in (f) and 5 was shown in Supplementary Fig. 35. f CLSM image of GFP synthesis within proteinosomes. Scale bar, 3 μm. All relevant experiments in b, c are performed independently at least three times with similar results. Source data is provided as a Source Data file.
Moreover, by taking advantage of precise transmembrane transport control and sequestration capability of Azo Coas and Re Coas, the ability to trigger protein synthesis within protocell communities was also developed in vitro transcription and translation machinery along with plasmid DNA encoding GFP (GFP DNA) using in vitro cell free protein expression Kit of Escherichia coli (E. coli) system (Fig. 6a). We experimentally demonstrated that the plasmid GFP DNA labeled by PI (PI-GFP DNA) could be sequestered by Azo Coas then being pumped into proteinosomes (Supplementary Fig. 34a). A large number of formed Re Coas within proteinosomes could sequester the genetic material after two pumping cycles of Azo Coa, and finally located on the inner membrane (Supplementary Fig. 34b). Therefore, proteinosomes encapsulating E. coli lysate containing a large amount of transcription and translation-related factors and Q-Dex500kDa were co-incubated with Azo Coas500 loaded with plasmid GFP DNA in a 1.5 mL of centrifuge tube. Two rounds of UV treatment (405 nm) were performed to induce Azo Coas to pump DNA twice into the proteinosome, thereby triggering GFP expression in proteinosomes. After a transcription and translation reaction at room temperature (25 °C) for 2 h, green fluorescence emitted by the synthesized GFP was observed (Fig. 6f, Supplementary Fig. 35a). The distribution of synthesized GFP exhibited a significant bias though there was a certain amount of GFP DNA leakage (Fig. 6e). Fluorescence intensity on the proteinosome membrane and within proteinosomes was significantly stronger than that of the solution outside proteinosomes, by factors of 6.07 and 2.58, respectively, due to the preferential spatial distribution of GFP DNA-enriched Re Coas on the membrane. No significant green fluorescence was observed after 2 h when GFP DNA was not pumped, suggesting that the pumping of condensates was the key to protein expression in the interactive protocell communities (Supplementary Fig. 35b, c).
Discussion
Inspired by biological systems, building biomimetic platforms to exploit the potential functions of interactive protocell communities using the physical principles and potential characteristics of macromolecular assemblies at droplet-membrane interface has become a central focus of the field. In this work, we proposed a binary community of synthetic protocells of photo-responsive Azo Coa microdroplets and proteinosomes with a cross-linked, flexible membrane. The high-density distribution of condensates on the proteinosomes membrane led to in-situ coalescence, and the spatial localization of the membrane network further realized the size control and structural stability of the condensates, which thus highlighted the potential contribution of membranes to the assembly dynamics of condensates. In addition, we have developed the function of Azo Coas as a high-pressure pump, using invagination caused by interfacial tension and hyperosmotic conditions resulting from the dissociation of Azo Coas to open up a way for oversize biomolecule transmembrane transduction. The osmotic pressure stimulation, generated by Azo Coa dissociation, drove the directed transport of skeleton molecules involved in the formation of condensates and functional molecules from the outer to the inner of the proteinosome according to the concentration gradient, to form reentrant LLPS, bringing about periodic fluctuations of those components, which was consistent with the directional exchange of the C-terminal domain of RNA polymerase II from condensates involved in transcription initiation to condensates involved in RNA processing after oxidative stress simulation46. The composition-dependent, controllable phase behavior between extracellular Azo Coas and intracellular reentrant Re Coas acting as the similar process reflected the fine-tune biomolecular interactions and revealed the assembly regulation in non-equilibrium state characteristic of the multi-component phase behaviors. Functional molecules such as enzymes and factors associated with transcription and translation processes were ultimately selectively localized into Re Coas, enabling directed and controlled assembly and processing of biochemically active substances, such as DNA unwinding reaction and protein synthesis.
Overall, our work introduced a strategy for the influx of biomacromolecules beyond membrane permeability into membranes in synthetic biomimetic systems, which could be used as an effective technique for boosting the development of reconfigurable cytomimetic materials with structural, functional, and signal transduction complexity.
Methods
Materials
Albumin from bovine serum (BSA, 66 kDa, Sigma-Aldrich, 98%), 1,6-Diaminohexane (Kermel, 99.7%), N-(3-Dimethylaminopropyl)-N′- ethylcarbodiimide hydrochloride (EDAC, Sigma-Aldrich, 98%), Dextran (MW 6 kDa, 70 kDa, 500 kDa from Leuconostoc spp. Sigma-Aldrich), Dextran (MW 1000 kDa, 2000 kDa, Jinpin, Shanghai, 95%), Deoxyribonuelease I (DNase, 31 kDa, Solarbio), DNA (~100 kDa, Thermo Fisher Scientific), Human serum albumin (HSA, 66 kDa, Sigma-Aldrich, 96%), Lysozyme from chicken egg white (LZ, 14.4 kDa, Sigma-Aldrich, 95%), Hemoglobin from bovine blood (Hb, 64.5 kDa, Sigma-Aldrich, 95%), Dimethyl sulfoxide (DMSO, Energy Chemical, 99.9%), 4-(Dimethylamino)pyridine (DMAP, Sigma-Aldrich, 99%), (3-Chloro-2-hydroxypropyl)trimethylammonium chloride (Aladdin, 65 wt% in H2O), Succinic anhydride (Aladdin, 99%), 4-Ethoxyaniline (Aladdin, 98%), 1,2-Dibromoethane (Aladdin, 99%), Trimethylamine (Aladdin, 30 wt% in H2O), Phenol (Aladdin, 99.5%), Fluorescein isothiocyanate (FITC, Sigma-Aldrich, 90%), Rhodamine B isothiocyanate (RBITC, Sigma-Aldrich), Fluorescein isothiocyanate labeled dextran(FITC-Dextran, MW 4 kDa, 10 kDa, 20 kDa, 70 kDa, 150 kDa, 500 kDa, Sigma-Aldrich, 98%). Cell-free protein expression (CFPE) Kit of GFP was purchased from Mich Scientific. Milli-Q water was used to prepare all the solutions in this study.
Characterization methods
Optical images were performed on a Leica DMi8 manual inverted fluorescence microscope at 40 × magnifications. Confocal fluorescence images were performed on a Leica SP8 confocal laser scanning microscope (CLSM). 1H-NMR spectra of chemical compounds were measured on a Bruker Avance-400 MHz NMR spectrometer using D2O and DMSO-d6 as the solvent. The test result was analyzed using MestReNova. Protocell co-assembly analysis was undertaken on a Flow cytometer (CytoFLEX, Beckman Coulter, Inc.). UV-vis spectrum measurements were performed on a PerkinElmer spectrophotometer (Lambda 750S, USA). Zeta potential measurements were performed using Malvern Zetasizer Nano-ZSP at 25 °C.
Drop adhesion force (DAF)/interfacial tension (γ) was performed on an Optical Surface Analyzer (OSA 100S-M, NBSI). Isothermal titration calorimetry (ITC) experiments were performed using Nano-ITC (TA, USA).
Synthesis of quaternized dextran
Dissolve 150 mg of dextran and 280 mg of NaOH in 14.25 mL of Milli-Q water at 35 °C to prepare quaternized dextran9. After dextran completely dissolved, 1.1 mL of (3-Chloro-2-hydroxypropyl)trimethylammonium chloride (65 wt% in water) was added dropwise to the stirring reaction mixture, which was then allowed to react overnight. The final mixture was neutralized with acetic acid and extensively dialyzed against water using a dialysis tube with 3.5 kDa (80% yield).
Synthesis of succinyl dextran
Dissolve 100 mg of dextran and 185 mg of succinic anhydride in 15 mL of DMSO to prepare succinyl dextran9 at 60 °C. Once dextran was fully dissolved, add 5 mg of DMAP and stir the reaction mixture for 16 h. After the reaction, 30 mL of water was added and the solution was extensively dialyzed against water using dialysis tube with 3.5 kDa (80% yield).
Synthesis of AzoTAB
AzoTAB was synthesized using a three-step reaction26,27. Personal protective equipment was used during the synthesis and purification steps.
Add concentrated HCl (17 mL) and ice (80 g) to a 1:1 v/v ethanol-water solution (160 mL) containing 4-ethoxyaniline (10.3 mL, 80 mmol, 1 equiv.) and sodium nitrite (5.5 g, 80 mmol, 1 equiv.), while maintaining the mixture in an ice bath (T = 0 °C) under a fume hood. Stir the mixture for 1 h, then carefully add cold water (42 mL) containing phenol (7.5 g, 80 mmol, 1 equivalent) and NaOH (6.4 g, 160 mmol, 2 equiv.), continuing to stir for an additional 90 min while keeping the temperature below 5 °C. Subsequently, adjust the pH of the solution to 1 using concentrated HCl and let it stand for 30 min. Filter the resulting precipitate, wash thoroughly with water, and vacuum-dry overnight to obtain 4-ethoxy-4’-hydroxyazobenzene (azoH) as a dark brown powder.
Mix azoH (2.4 g, 10 mmol, 1 equiv.), 1,2-dibromoethane (5.6 g, 3 equiv.), potassium carbonate (2.07 g, 1.5 equiv.), and potassium iodide (0.083 g, 0.05 equiv.) in 50 mL of butanone, and reflux the mixture in the dark under a fume hood for 48 h. The reaction mixture was then hot filtered to remove solid impurities and salt, with the residue being washed with butanone. The filtrate was collected, and the solvent was removed under reduced pressure. The resulting solid was dissolved in dichloromethane (20 mL) and extracted with NaOH solution (1 M, 2 × 8 mL) and pure water (2 × 8 mL). The organic phase was dried over MgSO4 and concentrated. The crude product was purified by recrystallization from ethanol, yielding 4-ethoxy-(4′-(2-bromoethyloxy)phenyl) azobenzene (azoBr) as an orange powder upon vacuum drying.
Dissolve 1 g of azoBr (4.4 mmol, 1 equiv.) in 80 mL of dry tetrahydrofuran (THF), then add a 33% solution of trimethylamine in ethanol (4.2 mL, 11.5 mmol, 4 equiv.). The mixture was stirred in the dark in a fume hood for 6 days. The resulting precipitate was filtered, washed with THF, and dried under vacuum. The crude product was recrystallized twice from ethanol and dried overnight under vacuum, yielding azobenzene trimethylammonium bromide (azoTAB) as an orange powder (36% yield). 1H-NMR (400 MHz, DMSO): δ = 7.86 (d3,J(H-H) = 8 Hz, 2H; Ar-H), 7.82 (d3,J(H-H) = 8 Hz, 2H; Ar-H), 7.17 (d3,J(HH) = 8 Hz, 2H; Ar-H), 7.08 (d3,J(H-H) = 8 Hz, 2H; Ar-H), 4.56 (m, 2H; CH2O), 4.11 (q3,J(H-H) = 6 Hz, 2H; CH2O), 3.82 (m, 2H; CH2N), 3.18 (s, 9H; CH3N), 1.35 ppm (t3,J(H-H) = 4 Hz, 3H; CH3)13;C NMR (400 MHz, DMSO): δ = 161.3 (Ar-C), 159.9 (Ar-C), 147.1 (Ar-C), 146.4 (Ar-C), 124.7 (Ar-C), 124.5 (Ar-C), 115.8 (Ar-C), 115.4 (Ar-C), 64.5 (CH2O), 64.1 (CH2N), 62.5 (CH2O), 53.6 (CH3N), 15.0 ppm (CH3); MS (ESI): m/z: calcd for C19H26N3O2: 328.4 [M–Br–]+; found: 328.2.
Conversion efficiency of AzoTAB under light simulation was not 100%26. Under UV light, AzoTAB existed in almost all-cis-configuration (94.65%), while more than half of AzoTAB exists in trans-configuration (50.43%) under blue light.
Preparation of stock solutions of AzoTAB and Su-Dextran
A trans-azoTAB stock solution (5 mL, 30 mM) was prepared by dissolving an appropriate amount of the azobenzene derivative in PBS buffer (50 mM, pH 8.0). The stock solution was stored for 30 min in the blue light (488 nm) irradiation to ensure complete isomerization to the trans-. As the same, Su-Dex (5 mL, 30 mM repeating unit concentration) with different molecular weights (70 kDa, 500 kDa, 1000 kDa, 2000 kDa) stock solutions were prepared in PBS buffer (50 mM, pH 8.0). The stock solutions are diluted according to the experimental requirements.
Preparation of Azo condensates, proteinosomes and their interactive protocell communities
Azo Condensates (Azo Coas) were produced by mixing Su-Dex (15 μL, 16 mM repeating unit concentration) and trans-azoTAB (45 μL, 30 mM) to reach final concentrations of 4 mM Su-Dex and 12 mM trans-azoTAB at a charge molar ratio of 3:1 (+/−).
Proteinosomes were prepared by mixing an aqueous BSA-NH2/PNIPAAm solution with 2-ethyl-1-hexanol followed by shaking the mixture by hand for 10 s. Typically, 0.06 ml of aqueous BSA-NH2/PNIPAAm (4.0 mg/ml, pH 8.0, 50 mM PBS buffer) was mixed with 1.0 ml of the oil. The proteinosomes were then cross-linked in the continuous oil phase by the addition of PEG-bis(N-succinimidyl succinate) (0.5 mg), which reacted with free primary amine groups of BSA-NH229.
Interactive protocells were prepared by mixing the Azo condensate solution (20 μL) and the proteinosome solution (10 μL) for 10 min.
Proteins labeled by fluorescence dyes
Dissolve proteins (50 mg) in 2.0 mL of Na2CO3/NaHCO3 buffer (100 mM), and then add FITC or RBITC (50 μL, 1 mg/mL in DMSO) dropwise. Stir the solution at room temperature for 6 h at 200 rpm. Subsequently, perform extensive dialysis against milliQ water (dialysis tube ~3 kDa MWCO) to remove any unreacted dye. After freeze-drying, the protein labeled with fluorescence dyes were obtained.
Optical switching in-situ CLSM experiments of Azo Coa-proteinosome assemblies
The interactive protocell solution (10 μL) was then placed on a confocal dish, and the target assembly was selected under the microscope. The target interactive protocell was subjected to alternating UV (405 nm)-blue (488 nm) light irradiation using Leica SP8 Laser (405 nm ≥ 50 mW, 488 nm ≥ 20 mW; 552 nm ≥ 20 mW), causing the dissociation and regeneration of Azo Coas.
Drop adhesion force (DAF) measurement
DAF was used to assess the interaction between condensed phase and proteinosomes performed on an Optical Surface Analyzer (OSA 100S-M, NBSI). Generally, the substrate was first prepared by depositing the corresponding proteinosome solution onto a glass slide and dried naturally at room temperature. The condensed phase was obtained by centrifuging the condensate solution at 2800 × g for 30 min. The corresponding condensed phase was inhaled into a syringe pump (1 mL). The general process was as follows: the droplet was produced through compression, hanging at the tip of the needle, and moving the substrate began to contact with the droplet. Then, moving the substrate towards the opposite direction induced the droplet elongation and deformation until the droplet breakup. Contact angle (θ) and DAF (Fa) between the droplets and the corresponding surface can be directly calculated. More precisely, the greatest attraction/tension (Fm) is related to the fracture site of the liquid bridge. If the fracture occurs at the droplet-substrate interface (lower contact position), then Fm is approximately equal to Fa (Fm ≈ Fa); If the fracture occurs at a different location, Fa is greater than Fm. The closer the fracture position of the liquid bridge is to the droplet-substrate interface, the closer the two forces are.
In our experiment, the fracture of the liquid bridge occurred at the upper midsection, indicating that Fa between condensates and proteinosome membrane was much greater than the measured attraction of 3.1413 × 107 μN/μm2.
Flow cytometry
The Azo Coa dispersions were prepared with final concentrations of 12 mM trans-azoTAB and 4 mM Su-Dex500kDa at a molar ratio of 3:1 (+/-). The samples were incubated for 10 min to ensure that the incubation duration was consistent with the generation time of the interactive protocells. The flow speed was set to be 10 μL/min, and the 2D dot plots were determined for a total volume of 10 μL.
Proteinosomes were prepared by the above methods. Gently shake before the test to ensure that the proteinosomes are evenly dispersed in the centrifuge tube. The flow speed was set to be 10 μL/min, and the 2D dot plots were determined for a total volume of 10 μL.
The interactive protocell solution was prepared by mixing Azo Coa and proteinosome solutions at a volume ratio of 2:1 and was incubated for 10 min to ensure the adhesion of the two communities. The flow speed was set to be 10 μL/min, and the 2D dot plots were determined for a total volume of 10 μL. All samples were characterized with a flow cytometer (Beckman Coulter, Inc.). Data was analyzed with FlowJo 10.8 software.
Pumping fraction and diffusion fraction
We randomly selected several interactive protocells labeled with RBITC-Su-Dex from independent samples, and recorded the diameters of proteinosomes at the maximum equatorial plane, the diameters and red fluorescence intensity of Azo Coas, as well as the fluorescence intensity inside proteinosomes after pumping. The fluorescence intensity was converted into Su-Dex concentration, and the total amount of Su-Dex in Azo Coas before pumping and the total amount of Su-Dex in proteinosomes after pumping could be calculated. The diffusion fraction was calculated by 1 - pumping fraction of Su-Dex.
Binding affinity determined by isothermal titration calorimetry (ITC)
ITC Experiments were performed with a Nano ITC from TA Instruments. All experiments were performed at 25 °C. The following conditions were used: Su-Dex500kDa (0.1 mM repeated units in 50 mM PBS, pH 8.0) was titrated with AzoTAB (20 mM in 50 mM PBS, pH 8.0): 25 injections, 2 uL/inj. Su-Dex500kDa (1 mM repeated units in 50 mM PBS, pH 8.0) was titrated with Q-Dex500kDa (20 mM in 50 mM PBS, pH 8.0): 25 injections, 2 uL/inj.
For each experiment, a control was performed by titrating the corresponding amount of AzoTAB, Q-Dex500kDa in 50 mM PBS (pH 8.0) in the absence of Su-Dex500kDa. Data were fitted using a non-linear least squares algorithm. All titrations were performed in a regime of concentrations where no phase-separation behavior could be observed.
DNA unwinding experiment
SYBR Green specifically labeled DNA (SYBR Green-DNA) at a final concentration of 0.4 mg/mL and Q-Dex500kDa at a final concentration of 16 mM were co-encapsulated into the proteinosomes. Add 1 μL of DNase (10 mg/mL) to 12 μL of Azo Coa500 solution, so that DNase was sequestered into the Azo Coas500. Then the Azo Coa loaded with DNase was mixed with the above-prepared proteinosome solution at a volume ratio of 2:1 and incubated for 10 min. A small amount of the mixture was then placed on a confocal dish, and the target assembly was selected under confocal fluorescence microscopy. The target Azo Coa-proteinosome assembly was subjected to alternating UV (405 nm)-blue (488 nm) light irradiation, causing the dissociation and regeneration of Azo Coas. The average fluorescence intensity of SYBR Green-DNA within proteinosomes was then monitored to evaluate the DNA uncoiling rate. Quantitative of RBITC-labeled DNase pumped by Azo Coas at each cycle was obtained by monitoring the average fluorescence intensity of RBITC-DNases within proteinosomes under blue light (488 nm).
Cell free protein expression in vitro of green fluorescent protein (GFP)
Green fluorescent protein (GFP) expression with a molecular weight of 26 kDa was performed using the in vitro cell-free protein expression (CFPE) Kit of reconstituted Escherichia coli (E. coli) system purchased from MICH Scientific. A total 33 μl of aqueous phase containing 8 μL of E. coli lysate rich in a large amount of transcription and translation-related factors, 3 μL of Q-Dex500kDa (176 mM), 5 μL of crosslinking agent (0.1 mg/mL), and 20 μL of BSA-PNIPAAm (10 mg/mL) was prepared proteinosomes for in vitro cell-free protein expression. Proteinosomes encapsulating reconstituted E. coli lysate and Q-Dex500kDa were co-incubated with Azo Coas500 loaded with plasmid DNA encoding GFP (6034 bp, 100 ng/mL) in a 1.5 mL centrifuge tube. UV light (405 nm) was irradiated twice, the time interval was 10 min, then centrifuged, remove the supernatant, and 5 μL of E. coli lysate to the centrifuge tube again. After the mixture was incubated at room temperature (25 °C) for 2 h, protein expression was observed by CLSM.
Micropipette aspiration technique for determination of proteinosome membrane properties (bending modulus κ and area expansion modulus K)
This method is based on the existing micropipette aspiration technique31.
The relationship between the induced elastic coefficient (EC) of the membrane deformation and the applied suction pressure (∆P) was determined upon aspiration of a portion of the proteinosome membrane:
Where RP and RV denoted the micropipette and the proteinosome radii, respectively. In our experiment, the inner diameter of the prepared micropipette (2Rp) was 5 μm, and the initial diameter of proteinosomes (2RV) was 25 μm. The proteinosome was aspirated into the micropipette with increasing suction pressure (∆P). As ∆P was gradually increased, the aspiration length (∆Lp) increased due to the membrane’s elastic.
Research has shown that the product of the bending modulus (κ) and αapp is directly proportional to the natural logarithm of membrane elastic deformation:
Where kB is the Boltzmann constant and T is the absolute temperature (T = 298.15 K).
The bending modulus, κ, was determined by plotting ln(EC) vs αapp and taking the product of the slope with kBT/8π. The plot of ln (EC) vs αapp for a proteinosome was shown in Fig. 4d (blue line). αapp was governed by the membrane area expansion and increases linearly with membrane elastic deformation:
By convention, the slope was known as the apparent area expansion modulus (Kapp).
The relationship between ∆Lp and the observed apparent area strain (αapp) was as follows:
Where A0 was the membrane area of the proteinosome measured at the initial state and A was the membrane area after pressurization.
We hypothesize that, similarly, Azo Coa localizes to the proteinosome membrane, and its dissociation causes an increase in αapp of the membrane. As shown in Fig. 4e, we defined the invagination depth (∆L) of Azo Coas on the proteinosome membrane, ranging from half to the entire Azo Coa diameter. The diameter of the invagination site corresponded to the inner diameter of the micropipette, where Azo Coa functioned as the micropipette in our Azo Coa-proteinosome assembly model. Therefore, Rp was related to Azo Coas. Assuming an expansion coefficient x of 1.05, where x represents the volume expansion degree of condensates before the transient dissociation. Rp could be expressed as x · 1/2 DAzo Coa. The force causing proteinosome membrane deformation was the osmotic pressure difference generated upon transient dissociation of Azo Coas, denoted as \({\Delta {\Pi }}\), which was related to the concentration of the released components after Azo Coa dissociation. Buffer salt concentration concentrated in Azo Coas was ignored.
Considering all factors above, we ultimately defined the Azo Coa-induced elastic deformation of the proteinosome membrane (αapp) as follows:
Changes in proteinosome membrane area were calculated from displacements in ∆Lp inside the micropipette using the geometric relations for total area of an aspirated proteinosome. Because the pressurized shape of the vesicle is almost a perfect sphere, the relations depend only on the diameter 2RV of proteinosome outside the micropipette, the internal diameter of the micropipette 2Rp, and the aspiration length ∆Lp: i.e.,
Hemispherical cap and cylindrical portion of the projection inside the pipette:
Spherical segment of the vesicle region outside the micropipette:
combine to specify the total area:
The suction on the membrane segment per unit area (μm2) within the micropipette was calculated as follows:
Osmotic pressure different measurements
Transient osmotic pressure difference, ∆∏ (Pa), resulting from Azo Coa dissociation was calculated from osmolality measurements using the Van’t Hoff equation:
Where R is the gas constant, T is the absolute temperature, ∆c is the charge molar concentration of the Azo Coa components released after the dissociation of Azo Coas (in mol kg−1) and RT = 2.479 kg MPa mol−1 at 25 °C.
The osmotic pressure difference on membrane invaginations per unit area (μm2), \({\Delta {\Pi }}_{{unit}}\), was calculated as follows:
Where Scontact area (μm2) is the contact area between Azo Coas and the membrane of the proteinosome.
Quantitative calculation of AzoTAB in Azo Coas
The standard curve method was used to quantify AzoTAB in Azo Coas by using the characteristic that the cis-AzoTAB has the maximum absorption peak near 450 nm. Firstly, solutions of AzoTAB with concentrations ranging from 0.1 mM to 0.7 mM were prepared. The solutions were then subjected to 1 h of UV irradiation (405 nm) to ensure complete conversion of trans-AzoTAB to cis-AzoTAB. The UV-Vis spectra of the solutions were measured, and a linear fit was performed for the absorbance at 450 nm as a function of AzoTAB concentration, yielding a fitting equation.
A solution of Azo Coas with a molar ratio of AzoTAB to Su-Dex of 3:1 was prepared, with a total molar concentration of 16 mM and a final volume of 20 μL. After centrifugation at 1800 × g for 5 min to separate the condensed phase from the supernatant, the supernatant was collected. It was then diluted 10-fold with water and subjected to 1 h of UV irradiation (405 nm). The UV-Vis spectra were measured, and the absorbance at 450 nm was determined. Thus, AzoTAB concentration in Azo Coas (Ccoa) could be obtained:
Where n0 represented the added amount of AzoTAB in the entire system. Cs and Vs represented the volume of the supernatant and the concentration of AzoTAB after centrifugation, respectively, with a dilution factor of 10.
Quantitative calculation of Su-Dex in Azo Coas
Su-Dex in Azo Coas was quantified by standard curve method of CLSM. Firstly, solutions of RBITC-Su-Dex with concentrations ranging from 0 mM to 100 mM were prepared. The fluorescence intensity of RBITC-Su-Dex was quantified at different concentrations using CLSM under the same parameters, resulting in a concentration-fluorescence intensity fitting curve. Fluorescence signal in this article was calibrated against concentration, and the intensity-concentration relationship was linear within the tested range.
Subsequently, a solution of Azo Coas with a molar ratio of AzoTAB to Su-Dex of 3:1 was prepared, with a total molar concentration of 16 mM and a final volume of 20 μL. The fluorescence intensity of RBITC-Su-Dex in the condensed phase and the surrounding environment solution was quantified under the same parameters. The distribution coefficient (K) of RBITC-Su-Dex was calculated based on the obtained data. Using the standard curve, the amount of RBITC-Su-Dex in the surrounding environment solution could be calculated, and by applying the distribution coefficient, the concentration of RBITC-Su-Dex in condensates could be determined.
Where Ccoa and Cs represented the concentration of RBITC-Su-Dex in the condensed phase and the surrounding environment solution, Icoa and Is represented the fluorescence intensity of RBITC-Su-Dex in the condensed phase and the surrounding environment solution, respectively. K was the distribution coefficient.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All relevant data supporting the findings of this study are available within the paper, Supplementary Information or from corresponding authors upon request. Source data are provided with this paper.
References
Banani, S. F., Lee, H. O., Hyman, A. A. & Rosen, M. K. Biomolecular condensates: organizers of cellular biochemistry. Nat. Rev. Mol. Cell Biol. 18, 285–298 (2017).
Shin, Y. & Brangwynne, C. P. Liquid phase condensation in cell physiology and disease. Science 357, eaaf4382 (2017).
Mehta, S. & Zhang, J. Liquid-liquid phase separation drives cellular function and dysfunction in cancer. Nat. Rev. Cancer 22, 239–252 (2022).
Riback, J. A. et al. Composition-dependent thermodynamics of intracellular phase separation. Nature 581, 209–214 (2020).
Fu, Y. & Zhuang, X. m6A-binding YTHDF proteins promote stress granule formation. Nat. Chem. Biol. 16, 955–963 (2020).
Chen, Y. et al. Construction of coacervate-in-coacervate multi-compartment protocells for spatial organization of enzymatic reactions. Chem. Sci. 11, 8617–8625 (2020).
Lu, T. & Spruijt, E. Multiphase complex coacervate droplets. J. Am. Chem. Soc. 142, 2905–2914 (2020).
Mountain, G. A. & Keating, C. D. Formation of multiphase complex coacervates and partitioning of biomolecules within them. Biomacromolecules 21, 630–640 (2020).
Li, J. et al. Programmable spatial organization of liquid-phase condensations. Chem 8, 784–800 (2022).
Gouveia, B. et al. Capillary forces generated by biomolecular condensates. Nature 609, 255–264 (2022).
Lu, T. et al. Endocytosis of coacervates into liposomes. J. Am. Chem. Soc. 144, 13451–13455 (2022).
Lee, J. E., Cathey, P. I., Wu, H., Parker, R. & Voeltz, G. K. Endoplasmic reticulum contact sites regulate the dynamics of membraneless organelles. Science 367, eaay7108 (2020).
Mason, A. F., Buddingh, B. C., Williams, D. S. & van Hest, J. C. M. Hierarchical self-assembly of a copolymer-stabilized coacervate protocell. J. Am. Chem. Soc. 139, 17309–17312 (2017).
Li, J., Liu, X., Abdelmohsen, L., Williams, D. S. & Huang, X. Spatial organization in proteinaceous membrane-stabilized coacervate protocells. Small 15, e1902893 (2019).
Martin, N. et al. Antagonistic chemical coupling in self-reconfigurable host-guest protocells. Nat. Commun. 9, 3652 (2018).
Rodriguez-Arco, L., Li, M. & Mann, S. Phagocytosis-inspired behaviour in synthetic protocell communities of compartmentalized colloidal objects. Nat. Mater. 16, 857–863 (2017).
Zhao, C. et al. Membranization of coacervates into artificial phagocytes with predation toward bacteria. ACS Nano 15, 10048–10057 (2021).
Adamala, K. P., Martin-Alarcon, D. A., Guthrie-Honea, K. R. & Boyden, E. S. Engineering genetic circuit interactions within and between synthetic minimal cells. Nat. Chem. 9, 431–439 (2017).
Buddingh, B. C., Elzinga, J. & van Hest, J. C. M. Intercellular communication between artificial cells by allosteric amplification of a molecular signal. Nat. Commun. 11, 1652 (2020).
Yang, Q. et al. A cascade signaling network between artificial cells switching activity of synthetic transmembrane channels. J. Am. Chem. Soc. 143, 232–240 (2021).
Thamboo, S. et al. Mimicking cellular signaling pathways within synthetic multicompartment vesicles with triggered enzyme activity and induced ion channel recruitment. Adv. Funct. Mater. 29, 1904267 (2019).
Chen, H. et al. DNA-based artificial receptors as transmembrane signal transduction systems for protocellular communication. Angew. Chem. Int. Ed. 62, e202301559 (2023).
Mu, W. et al. Superstructural ordering in self-sorting coacervate-based protocell networks. Nat. Chem. 16, 158–167 (2024).
Ji, Y., Lin, Y. & Qiao, Y. Plant cell-inspired membranization of coacervate protocells with a structured polysaccharide layer. J. Am. Chem. Soc. 145, 12576–12585 (2023).
Zhang, Y. et al. Osmotic-induced reconfiguration and activation in membranized coacervate-based protocells. J. Am. Chem. Soc. 145, 10396–10403 (2023).
Huang, Y. et al. Reversible light-responsive coacervate microdroplets with rapid regulation of enzymatic reaction rate. ChemSystemsChem 3, e2100006 (2021).
Martin, N. et al. Photoswitchable phase separation and oligonucleotide trafficking in DNA coacervate microdroplets. Angew. Chem. Int. Ed. 58, 14594–14598 (2019).
Fraccia, T. P. & Martin, N. Non-enzymatic oligonucleotide ligation in photoswitchable coacervate protocells sustains compartment-content coupling. Nat. Commun. 14, 2606 (2023).
Huang, X. et al. Interfacial assembly of protein-polymer nano-conjugates into stimulus-responsive biomimetic protocells. Nat. Commun. 4, 2239 (2013).
Dimova, R. How membranes influence intracellular phase separation. Nat. Chem. 16, 10–11 (2024).
Lu, L., Doak, W. J., Schertzer, J. W. & Chiarot, P. R. Membrane mechanical properties of synthetic asymmetric phospholipid vesicles. Soft Matter 12, 7521–7528 (2016).
Christian, D. A. et al. Spotted vesicles, striped micelles and Janus assemblies induced by ligand binding. Nat. Mater. 8, 843–849 (2009).
Olbrich, K., Rawicz, W., Needham, D. & Evans, E. Water permeability and mechanical strength of polyunsaturated lipid bilayers. Biophys. J. 79, 321–327 (2000).
Kusumaatmaja, H. & Lipowsky, R. Droplet-induced budding transitions of membranes. Soft Matter 7, 6914–6919 (2011).
Yuan, F. et al. Membrane bending by protein phase separation. Proc. Natl. Acad. Sci. USA. 118, e2017435118 (2021).
Zhang, L. et al. 53BP1 regulates heterochromatin through liquid phase separation. Nat. Commun. 13, 360 (2022).
Cai, D. et al. Phase separation of YAP reorganizes genome topology for long-term YAP target gene expression. Nat. Cell Biol. 21, 1578–1589 (2019).
Strom, A. R. et al. Phase separation drives heterochromatin domain formation. Nature 547, 241–245 (2017).
Larson, A. G. et al. Liquid droplet formation by HP1α suggests a role for phase separation in heterochromatin. Nature 547, 236–240 (2017).
Simon, J. R., Eghtesadi, S. A., Dzuricky, M., You, L. & Chilkoti, A. Engineered ribonucleoprotein granules inhibit translation in protocells. Mol. Cell 75, 66–75 (2019).
Booth, R., Qiao, Y., Li, M. & Mann, S. Spatial positioning and chemical coupling in coacervate-in-proteinosome protocells. Angew. Chem. Int. Ed. 58, 9120–9124 (2019).
Narayanan, T. & Kumar, A. Reentrant phase transitions in multicomponent liquid mixtures. Phys. Rep. 249, 135–218 (1994).
Onuchic, P. L., Minn, A. N., Alshareedah, I., Deniz, A. A. & Banerjee, P. R. Divalent cations can control a switch-like behavior in heterotypic and homotypic RNA coacervates. Sci. Rep. 9, 12161–12170 (2019).
Kaur, T. et al. Sequence-encoded and composition-dependent protein-RNA interactions control multiphasic condensate morphologies. Nat. Commun. 12, 872 (2021).
Banerjee, P. R., Milin, A. N., Moosa, M. M., Onuchic, P. L. & Deniz, A. A. Reentrant phase transition drives dynamic substructure formation in ribonucleoprotein droplets. Angew. Chem. Int. Ed. 56, 11354–11359 (2017).
Guo, Y. E. et al. Pol II phosphorylation regulates a switch between transcriptional and splicing condensates. Nature 572, 543–548 (2019).
Acknowledgements
We thank the National Natural Science Foundation of China (22171058, 22475056, X. H.; 22307030, H. C.), the Fundamental Research Funds for the Central Universities (HIT.OCEF.2023040), China-German Mobility Program (M-0470) to X. H., and the China Postdoctoral Science Foundation (2023M730844, X. W.) for financial support.
Author information
Authors and Affiliations
Contributions
Y. H. and H. C. performed the experiments and undertook data analysis. X. Q. and S. L. carried out the fluorescence microscopy measurements and isothermal titration calorimetry experiment. X. H. led the project. X. H., X. L. and X. W. conceived, designed and supervised the study, as well as revised the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Lingxiang Jiang, Jianbo Liu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Huang, Y., Chen, H., Qiao, X. et al. Liquid-liquid phase separation-boosted transmembrane delivery in interactive protocell communities. Nat Commun 16, 5231 (2025). https://doi.org/10.1038/s41467-025-60541-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-60541-7