Abstract
Feeding is an innate behavior critical for survival but is also influenced by many non-nutritional factors such as emotion, social context and environmental conditions. Recently, tuberal nucleus somatostatin (TNSST) neurons have been identified as a key feeding regulation node. To gain a deeper understanding of the TNSST neural networks, we quantitatively characterised the brain-wide input-output configuration of mice TNSST neurons using the VITALISTIC method (Viral Tracing Assisted by Light-Sheet microscope and Tissue Clearing) and single-cell projectomes by fluorescence micro-optical sectioning tomography (fMOST). We found that TNSST neurons receive direct inputs from and send outputs to a broad range of brain regions, including many cortical and subcortical areas. Differently from AgRP neurons, the extensively studied ‘hunger’ neurons, TNSST neurons receive more diverse inputs from extra-hypothalamic regions and neuromodulatory centers. Using the projection-specific input tracing, we further revealed fine-tuning of the input-output configuration of TNSST neurons that align with specific functional needs.
Similar content being viewed by others
Introduction
Feeding regulation is critical for human health. While overeating results in obesity and other associated health problems, undereating causes metabolic derangements such as malnutrition or developmental delays. Large scale human genomic studies revealed that the expression of genomic loci associated with body weight is more enriched in the nervous system1, consistent with the idea that the nervous system plays an instrumental role in regulating feeding behavior.
It has been postulated that foraging, consumption and termination of feeding are regulated mainly by different neuronal circuits2, whereas different types of hypothalamic neurons may provide similar or redundant functional roles in feeding regulation3,4,5,6,7. While functional manipulation of specific types of neurons greatly facilitated our understanding of feeding regulation, how these feeding regulation neurons interact with other neural circuits, or how they integrate complex information, is still poorly understood. Several recent studies have started to uncover the complex and mutual influence of cortical or subcortical circuits on hypothalamic feeding-regulation neurons8,9,10,11,12. Earlier studies have reported broad afferent and efferent projections of AgRP, POMC and other hypothalamic neurons13,14,15, but a more quantitative and comprehensive analysis of the brain-wide network of these neurons is still missing.
The lateral tuberal nucleus (nucleus tuberalis lateralis, NTL) in the hypothalamus was previously considered an evolutionarily new brain structure that is recognized in primates and humans16. The presence of somatostatin (SST)-positive neurons in the tuberal nucleus (TN), the homologous brain region to human NTL, was recently confirmed in mice6. It was revealed that SST neurons in TN (TNSST neurons) are functionally important in the regulation of food intake in mice6. Further study found that TNSST neurons, together with their plastic change of subiculum inputs, are critical regulators of contextually conditioned feeding behavior8, a maladaptive feeding behavior that may contribute to the environmentally driven overconsumption of food and obesity. However, a comprehensive investigation of TNSST connectivity is lacking, which limits our understanding of how and what other neural circuits also participate in TNSST neurons-mediated feeding regulation.
The advances in viral tracing tools including modified rabies virus enable researchers to map monosynaptic inputs and outputs of genetically defined neuronal populations17,18. Recent developments in light-sheet microscopy and tissue clearing technologies have further enabled comprehensive whole-brain analysis of neural circuits without mechanical sectioning and manual counting19,20, which is time consuming and may introduce biases in quantification. In this study, we designed and implemented the VITALISTIC method (Viral Tracing Assisted by Light-Sheet microscope and Tissue Clearing), which provides an unbiased and quantitative description of the input configuration of TNSST neurons. Using VITALISTIC method, we compared and revealed brain-wide connectivity differences between TNSST and AgRP neurons, two important types of orexigenic neurons deep in the hypothalamus. Additionally, we also analysed the output of TNSST neurons revealed by sparse labelling and whole-brain reconstruction of single cell projectomes by fluorescence micro-optical sectioning tomography (fMOST), which provides high resolution reconstruction of single axons21,22. Finally, using a modified cTRIO (cell type specific tracing the relationship between input and output) method23, we revealed the functional fine-tuning of TNSST neurons projecting to different regions. Together, this study advanced our understanding of the TNSST circuits and provides an entry point for understanding complex yet integrated feeding regulation by cortical and subcortical inputs to hypothalamic orexigenic neurons.
Results
Mapping the brain-wide inputs to TNSST neurons
After viral infection and viral gene expression for tracing neural inputs, each mouse brain was harvested and cleared using the CUBIC method24,25,26. We then used our custom-built mesoSPIM light-sheet microscope19 to image cleared mouse brains and the whole-brain imaging data was analysed by the open-source MagellanMapper software27 (Fig. 1a). Because this pipeline involves multiple techniques, we termed this pipeline the VITALISTIC (Viral Tracing Assisted by Light-Sheet microscope and Tissue Clearing) method for simplicity and avoiding repetition of the combination of techniques. For studying mono-synaptic inputs to TNSST neurons, we used SST-cre mice and G-deleted rabies virus. This was done by first expressing in TNSST neurons the TVA receptor for EnvA fused to mCherry and the rabies glycoprotein (G), followed by injecting into TN a glycoprotein-deleted and GFP-expressing rabies virus (EnvA-RV-ΔG-GFP)17 (Fig. 1b). The max projection showed localised mCherry signal in TN and widely distributed GFP signal, representing the inputs to TNSST neurons (Fig. 1c).
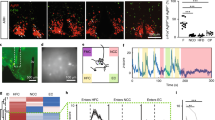
a Schematic diagram of the VITALISTIC method. Viral injection and gene expression followed by CUBIC tissue clearing for light sheet imaging. Automated whole-brain-analysis performed for the imaged individual neurons or individual synapse. b Representative diagram and images show rabies tracing for TNSST neurons. Expression of TVA-mCherry receptor for EnvA, rabies glycoprotein G and glycoprotein deleted GFP expressing rabies virus (RV-GFP). c Maximum projection of light sheet imaged whole mouse brain showing widely distributed RV-GFP signal representing the input neurons to TNSST and TVA-mCherry signal restricted in TN. d Circos map showing distributed inputs to TNSST neurons from ipsilateral and contralateral hemispheres. The brain region names are indicated along the circle with small fonts. e Electrophysiological recording of light-evoked response of TNSST neurons and ChR2 was expressed in AON. Panel showing representative trace together with number and percentage of light responsive recorded TNSTT neurons (n = 16 neurons, from 3 mice). PTX: picrotoxin, a GABAA receptor blocker; CNQX: 6-cyano-7-nitroquinoxaline-2,3-dione, a competitive AMPA/kainite receptor antagonist; APV: (2 R)-amino-5-phosphonovaleric acid, a NMDA receptor antagonist. f The proportion of the inputs to TNSST neurons in ipsilateral (blue) and contralateral (red) hemispheres. The values are the normalized ratio of the cell number in each area against the total number of input neurons. Error bars represent the SEM. n = 8 mice. g The cross-correlation of ipsilateral and contralateral inputs to TNSST neurons in major brain regions as shown in Fig. S1.
The Circos plot showed that TNSST neurons received inputs from both ipsilateral and contralateral hemispheres (Fig. 1d). Although deep in the bottom of hypothalamus, TNSST neurons received diverse inputs from many brain regions, including cortical regions such as the medial prefrontal cortex (mPFC) and the anterior olfactory nucleus (AON) (Fig. 1d and Supplementary Fig. 1). To confirm functional connectivity of such long range projections, we expressed channelrhodopsin (ChR2) in the AON and were able to record light-evoked response in TNSST neurons (Fig. 1e), demonstrating direct synaptic inputs from AON to TNSST neurons. Quantitative analysis showed that the ipsilateral hypothalamus was the biggest input source of TNSST neurons, followed by striatum and midbrain, with pons and medulla providing the least amount of inputs (Fig. 1f). Further breaking down the inputs to smaller brain regions revealed that the input pattern in the ipsilateral side correlated well with that in the contralateral side, evidenced by peak correlation at 0 lag (Fig. 1g and Supplementary Fig. 1), suggesting that the brain image registration and quantification pipeline was able to reliably capture small brain regions in both brain hemispheres.
In cortical regions, TNSST neurons receive considerable amount of inputs from mPFC regions, including anterior cingulate area, prelimbic area, infralimbic area and orbital area, suggesting strong influence of the valuation and decision-making system in modulating the neural activity of TNSST neurons (Supplementary Fig. 1). The cortical input analysis also revealed that the TNSST neurons may be more influenced by motor and somatosensory regions, while audio and visual inputs have lesser influence (Supplementary Fig. 1). In the hypothalamus, TNSST neurons received inputs from many hypothalamic areas that have been implicated in feeding regulation, among which lateral hypothalamus (LHA) and zona incerta (ZI) are the major ones (Supplementary Fig. 1). While ZI contains mainly GABAergic neurons28, LHA contains diverse neuronal types4,29. Therefore, we examined whether the TNSST neurons receive GABAergic or glutamatergic inputs from LHA. By non-selectively expressing ChR2 in the LHA and recording TNSST neurons, we found that 58% of recorded TNSST neurons received synaptic inputs from LHA neurons and the averaged response was about 200pA (Supplementary Fig. 2a, b). Among the TNSST neurons that received synaptic inputs from LHA, 41.7% of them received glutamatergic inputs, 16.6% received both GABAergic and glutamatergic inputs, and 41.7% received other uncharacterised types of inputs (Supplementary Fig. 2c–g). Immunostaining further revealed that about 10% of the LHA input neurons were orexinergic, suggesting diverse subtypes of LHA inputs to TNSST neurons (Supplementary Fig. 1h–k). Together, TNSST neurons mainly received glutamatergic and other non-GABAergic, potentially peptidergic, inputs from LHA neurons.
Differences in the brain-wide inputs between TNSST neurons and AgRP neurons
Next, we wanted to compare the input patterns between TNSST and AgRP neurons, two major deep hypothalamic orexigenic neurons. The input-output configuration of AgRP neurons has been reported previously13. Similar as mapping the inputs to TNSST neurons, we expressed TVA and G protein using cre-dependent AAV in the arcuate of AgRP-cre mice, followed by injecting ΔG-rabies virus in the arcuate. Using the VITALISTIC method, we analysed the input pattern of AgRP neurons, which was largely consistent with a previous report13, but with more details. The Circos plot clearly showed that the major input regions of AgRP neurons were the hypothalamus (Fig. 2a). Quantitative comparison between TNSST and AgRP neurons further revealed the differences of the input patterns of these two major orexigenic neurons. AgRP neurons received more inputs from the hypothalamus but far less inputs from almost all other major brain regions than TNSST neurons (Fig. 2b). Within the hypothalamus, AgRP neurons received very little inputs from zona incerta (ZI), but significant inputs from TN, which is opposite from that of TNSST neurons (hypothalamus panel, Supplementary Fig. 3 and Supplementary Fig. 1). Consistent with a previous report13, AgRP neurons received more inputs from DMH, but less from LHA (Supplementary Fig. 3) than TNSST neurons, suggesting that TNSST and AgRP neurons process distinct aspects of hypothalamic information for modulating feeding behavior. To examine the connectivity between lateral hypothalamus neurons and AgRP neurons, we injected AAV-hSyn-ChR2 in lateral hypothalamus of AgRP::Ai14 mice (Supplementary Fig. 4a). Electrophysiological recording of AgRP neurons showed that only 15.6% of recorded AgRP neurons responded to optogenetic stimulation of lateral hypothalamus neural terminals in arcuate region (Supplementary Fig. 4b), and the amplitude was one magnitude smaller than that of TNSST neurons (Supplementary Figs. 4b and 2b).
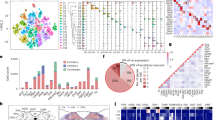
a Circos map showing distributed inputs to ARCAgRP neurons from ipsilateral and contralateral hemispheres. The brain region names are indicated along the circle with small fonts. (ARC: arcuate) b Graph comparing the proportion of the inputs to TNSST (red) and ARCAgRP neurons (blue) in ipsilateral (left) and contralateral (right) hemispheres. The values are the normalized ratio of the cell number in each area against the total number of input neurons. Error bars represent the SEM. n = 8 for SST-Cre mice and n = 7 for AgRP-Cre mice. *p < 0.05, **p < 0.01, ***p < 0.001, unpaired two-sided t test with Welch correction and Holm-Sidak correction. c Box plots showing comparison of the proportional inputs to TNSST neurons and AgRP neurons from major neuromodulatory centres. Error bars represent SEM. n = 8 for SST-Cre mice and n = 7 for AgRP-Cre mice. *p < 0.05, **p < 0.01, unpaired two sided t test with Welch correction and Holm-Sidak correction. Box plots showing median (yellow line), lower and upper edge of the box mean the first and third quartile respectively, and the whiskers extend to the last data points within 1.5 times of inter-quartile range (box height). d RNA Fluorescent In-Situ Hybridization (FISH) for dopaminergic receptors Drd1 and Drd2 in TNSST neurons. Representative images showing fluorescence labelling of Drd1, Drd2 receptors and SST in TN region. Pie graph showing the receptor expression percentage of TNSST neurons. n = 3 mice. e RNA FISH for Drd1 and Drd2 in arcuate AgRP neurons and quantification shown in pie chart. n = 3 mice. f RNA FISH for cholinergic receptors Chrm1 and Chrm2 in TNSST neurons. Representative images showing fluorescent labelling of Chrm1, Chrm2 receptors and SST in TN region. Pie graph showing the receptor expression percentage of TNSST neurons. n = 3 mice. g RNA FISH for Chrm1 and Chrm2 in arcuate AgRP neurons and quantification shown in pie chart. n = 3 mice.
Considering the important roles of neuromodulators in regulating feeding behavior, we examined the inputs from major neuromodulator centres. We found that TNSST neurons received significantly less inputs from the histamine centre, tuberal mammillary nucleus (TM), and AgRP neurons received significantly less inputs from other major neuromodulator centers such as ventral tegmental area (VTA) and pedunculopontine nucleus (PPN) (Fig. 2c), suggesting less influence of these neuromodulators on the function of AgRP neurons. Data-mining using publicly deposited single cell sequencing data revealed that AgRP neurons express lesser dopamine and serotonin receptors than TNSST neurons (Supplementary Fig. S4d). By performing RNA FISH to determine the mRNA expression levels, we found that more than half of TNSST neurons expressed Drd1 receptor, a much higher percentage than that of AgRP neurons30,31 (Fig. 2d, e), while a smaller fraction expressed Drd2 receptor. Moreover, about half of TNSST neurons expressed Chrm1 receptor and about a quarter of them expressed Chrm2 receptor, while a much smaller fraction of AgRP neurons expressed the Chrm1 cholinergic receptor (Fig. 2f, g). The differences of the expression levels of neuromodulator receptors in TNSST and AgRP matched their differences in inputs from major neuromodulator centers.
Mapping the brain-wide outputs of TNSST neurons
For mapping the outputs of single TNSST neurons with high resolution, we sparsely labelled TNSST neurons with virally expressed EYFP using AAV virus in the TN of SST-Cre mice. Brain samples were imaged using fMOST platform21,22 (Fig. 3a, b), which captures the whole-brain tissue at a high spatial resolution of 0.32 μm × 0.32 μm × 1 μm22. The projection of individually labelled axons were traced by a streamlined semi-automatic procedure that was previously described32. In total, 186 TNSST neurons were traced. By analysing this single neuron projectome results, we found that TNSST neurons projected mainly to ipsilateral side, but single axon projected broadly to many extra-hypothalamic regions including cortical regions (Fig. 3c, d). The cross-correlation of the outputs to ipsilateral and contralateral sides showed a peak at 0 lag for the majority of major brain regions, suggesting similarity of output targets in both hemispheres and supporting the precision of the output mapping pipeline (Fig. 3e and Supplementary Fig. 5). The previously reported projections to bed nucleus of the stria terminalis (BNST/BST), central amygdala (CeA), paraventricular nucleus of hypothalamus (PVH) and periaqueductal gray (PAG)6 were clearly an under-estimation of the complexity of the TNSST neurons’ projectome.
a Representative images showing the injection side and expression of EYFP in TN region of SST-Cre mouse. b Schematic showing the procedure of output mapping using fMOST platform. c Circos map showing distributed outputs of TNSST neurons in ipsilateral and contralateral hemispheres. The brain region names are indicated along the circle with small fonts. d Graph showing the proportion of the outputs of TNSST neurons in ipsilateral (blue) and contralateral (red) hemispheres. The values are normalized ratio of the axon length in each area against the total axon length of individual neuron. Error bars represent the SEM. n = 186 TNSST neurons. e The cross-correlation of ipsilateral and contralateral outputs of TNSST neurons in major brain regions as shown in Fig S5. f Location of all traced TNSST neurons, including some SST positive neurons at the boarder of TN. g Summary of the axon length (in μm) for all traced TNSST neurons. Each tick represents the value of the axon length of a given neuron in a brain region with color scaled to axon length. The values in these brain regions were used for clustering the projectome of TNSST neurons into 5 major groups. h Overview of whole-brain projectome of 5 major subtypes of TNSST neurons. Each color indicates one neuron. i and j, Scatter plot showing the input-output weights for ipsilateral (i) and contralateral (j) sides. Error bars represent the SEM. n = 186 neurons for output and n = 8 mice for input.
Based on the axon length of individual TNSST neurons in each brain region, which shows high correlation with synaptic terminal numbers, we clustered the axon projectome of the 186 TNSST neurons (Fig. 3f) into 5 major types (Fig. 3g, h), mainly separated by their projection patterns to regions in midbrain (i.e. PAG), pons (i.e. superior central nucleus raphe (CS)), pallidum (i.e. BST), striatum (i.e. lateral septum (LS)) and thalamus (i.e. paraventricular nucleus of the thalamus (PVT)). As a whole, it appeared that BST is the most commonly targeted extra-hypothalamic region by TNSST neurons, indicating that the functional output of TNSST neurons may critically depend on inhibiting downstream neurons in BST. Many BST-projecting TNSST neurons don’t project to PAG, which is consistent with a previous report that BST-projecting and PAG-projecting TNSST neurons have few overlap6. Notably, the projections to LS, PVT and lateral habenula (LH) were not appreciated in the previous report6, and such projection suggests potential link between feeding regulation, salience processing and mood regulation33,34. The projections to brain regions in pons and medulla (Supplementary Fig. 5) suggest potential interaction between TNSST neurons and interoception, including pain and nutrient sensing in parabrachial nucleus (PB) and nucleus of the solitary tract (NTS).
By analysing the percentage of input and output of TNSST neurons in major brain regions, we noted that most of the regions provided reciprocal connections with TNSST neurons with relatively similar weight (Fig. 3i, j), while cortical and hippocampal regions mainly provided input to TNSST neurons without receiving notable inputs from TNSST neurons (Supplementary Fig. 6). Within striatum, LS had reciprocal connections with TNSST neurons, but ACB (nucleus accumbens) and CP (caudoputamen) mainly provided inputs to TNSST neurons (Supplementary Fig. 6). Other regions such as SI (substantia innominata), BST and CEA had relatively balanced reciprocal connections with TNSST neurons (Supplementary Fig. 6). Most hypothalamic regions seemed to receive proportionally more input from TNSST neurons than providing input to TNSST neurons (Supplementary Fig. 6).
Mapping inputs of TNSST neurons projecting to different brain regions
While stimulating TNSST neurons enhances feeding, only stimulating their projection terminals in selected brain regions also enhances feeding6. Considering the partial one-to-many projection pattern of TNSST neurons and their functional differences in different projection targets, we wanted to explore whether the TNSST neurons projecting to different brain regions receive different inputs. Utilising a combination of cre- and flp-dependent viral tools as well as cre- or flp-expressing mouse lines, cTRIO (cell-type-specific tracing the relationship between input and output) has been adopted for mapping the input to subtypes of neurons projecting to a defined brain region23. Using SST-flp mice, we injected a retrograde Cre-expressing virus in chosen projection target regions, and injected flp-dependent TVA-expressing virus with cre-dependent G-expressing virus in TN (Strategy A, Fig. 4a). The alternative strategy, in which cre-dependent TVA-expressing virus and flp-dependent G-expressing virus were injected in TN, might result in non-specific input mapping due to potential mutual connection among TNSST neurons (Strategy B, Fig. 4a). The flp-dependent TVA-expressing AAV virus resulted in negligible rabies virus infection in wild type mice and the EnvA-ΔG-rabies virus had negligible infection without TVA-expressing virus (Supplementary Fig. S7). We previously showed that the TNSST neural terminals in PVN and BST/BNST promote feeding, while terminals in CeA and PAG don’t; moreover, consistent with current projectome mapping, it was reported that the BST-projecting and PAG-projecting TNSST neurons show limited overlap, while PVN-projecting and BST-projecting ones show higher level of overlap6. Although CeA and PVN were not shown to be major projection targets in the current single neuron projectome mapping, it is noteworthy that the current mapping may biased toward more middle part of TN (Fig. 3f). Therefore, we injected retrogradely transported Cre-expressing viruses in PVN, BST/BNST, CeA and PAG, as four representative downstream targets6 (Fig. 4b). The mice were sacrificed 8 days after the rabies virus injection and the brains were further processed for analysis.
a Schematic of two viral injection strategies. Strategy A: injection of retrograde Cre-expressing virus in chosen projection target regions and injection of flp-dependent TVA-expressing virus with cre-dependent G-expressing virus in TN, followed by injection of RV-GFP in TN after incubation time; Strategy B: injection of retrograde Cre-expressing virus in chosen projection target regions and injection of cre-dependent TVA-expressing virus with flp-dependent G-expressing virus in TN, followed by injection of RV-GFP in TN after incubation time. b Diagram showing injection Strategy A in a mouse brain. c Correlation matrix comparing overall input patterns of PVN-projecting TNSST (TNSST→PVN) neurons, BNST-projecting TNSST (TNSST→BNST) neurons, CeA-projecting TNSST (TNSST→CeA) neurons and PAG-projecting TNSST (TNSST→PAG) neurons. PVN: paraventricular nucleus; BNST: bed nucleus of the stria terminalis; CeA: central amygdala; PAG: periaqueductal gray. d Principal Component Analysis (PCA) showing differences between inputs of TNSST→PVN, TNSST→BNST, TNSST→CeA and TNSST→PAG neurons. e Bar graph showing significantly different input percentages from various region for four projecting areas. ACB: nucleus accumbens; MS: medial septum; VMH: Ventromedial hypothalamic nucleus; ARC: arcuate nucleus; SO: Supraoptic nucleus; RCH: Retrochiasmatic area; STN: Subthalamic nucleus; SN: Substantia nigra; VTA: Ventral tegmental area. (Mean±SEM; n = 4, 3, 4, 3 for PVN, BNST, CeA and PAG-projecting TNSST neurons, respectively; *p < 0.05, **p < 0.01, **p < 0.001 for uncorrected two-sided t test; red p values and stars indicate significance for two-sided t test after Holm-Šídák correction) f Representative images showing different input neurons for TNSST neurons projecting to indicated brain regions. g A model of the input-output configuration of TNSST neurons. TNSST neurons with different shade indicating that the ones with more distinct output pattern receive more distinct inputs (light vs. dark).
The overall input patterns of PVN-projecting TNSST (TNSST→PVN) neurons, BNST-projecting TNSST (TNSST→BNST) neurons, CeA-projecting TNSST (TNSST→CeA) neurons and PAG-projecting TNSST (TNSST→PAG) neurons were similar to each other, with correlation value ranging from 0.7 to 0.95, and the inputs to TNSST→BNST and TNSST→PAG neurons showed the lowest correlation (Fig. 4c). Principal component analysis further revealed that the inputs to TNSST→PVN and TNSST→BNST neurons were more similar than that of TNSST→CeA and TNSST→PAG neurons (Fig. 4d), which is consistent with our previous findings that many TNSST neurons projecting to BNST also project to PVN, and the overlap between TNSST→PAG and TNSST→CeA neurons is lower6. Moreover, stimulating TNSST terminals in PVN or BNST leads to feeding while stimulating their terminals in CeA or PAG does not6, which matches the distance of their input patterns in PCA analysis, indicating that there is fine-tuning of the input pattern for matching the functional specification of the output circuits. Considering the potential importance of dopaminergic inputs revealed in Fig. 2, we first examined whether the TNSST neurons projecting to different brain regions are similarly innervated by VTA inputs. Interestingly, the PAG-projecting TNSST neurons received more VTA inputs than the TNSST neurons projecting to other brain regions (Fig. 4e, VTA group). Since the percentage of input neurons in majority of brain regions were similar, we only plotted the regions that showed significant differences between the four groups of TNSST neurons (Fig. 4e). More stringent multi-comparison correction removed some significant differences, revealing the importance of post-hoc validation of the identified differences using whole-brain mapping methods. Together, the anatomical mapping suggests that the TNSST neurons receive broadcast signals from many brain regions, but the input pattern is fine-tuned for some output targets (Fig. 4g).
PAG-projecting TNSST neurons receive VTA inputs and promotes locomotion
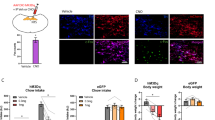
By immunostaining, we found that about 30% of VTA inputs neurons were dopaminergic (Fig. 5a–d). It was previously reported that about half of TNSST neurons respond to palatable food8, which is known to drive dopaminergic neurons35. We therefore examined whether the TN-projecting dopaminergic neurons also respond to palatable food. By injecting retrograde tracers in TN and cfos staining after letting mice consume high fat diet (HFD), we found that ~40% of TN-projecting dopaminergic neurons were cfos positive (Fig. 5e, g), supporting the link between the activity of VTA dopaminergic neurons and that of TNSST neurons in consuming HFD. Previous in vivo imaging work showed that the TNSST mainly responded to approaching palatable food, rather than consuming it8. Considering that PAG neurons could be involved in locomotion behavior36,37, we examined whether the projection of TNSST in PAG has any functional significance. By expressing ChR2 in TNSST neurons and optogenetically stimulating their terminals in PAG, we found that the locomotive activity of mice was increased without changing in maximum velocity and such increased locomotion was independent of the presence of food (Fig. 5h–j); while stimulating TNSST neural terminals in BNST didn’t significantly increase locomotion (Supplementary Fig. S8a).
a Schematic diagram of the imaged coronal brain section b Representative images of RV-GFP traced TNSST→PAG input neurons in VTA-SN region (left), immunohistochemical staining against TH-antibody (middle) and merged image (right). c Pie chart showing the number of TH + RV+ neurons among RV+ neurons within VTA-SN region. d Bar graph showing the percentage of TH + RV+ neurons over RV+ neurons within VTA-SN region. (N = 3 animals, Mean±SEM) e Experimental diagram showing CTB555 injected mice (in TN) were fasted overnight. Brains were collected after consuming HFD and were processed for FISH. f Representative coronal sections showing FISH for cfos (green), CTB555 (red), and DAT (blue). g Bar plot showing percentage of cfos + ,DAT + ,CTB+ neurons in DAT + ,CTB+ neurons within VTA-SN region. (N = 5 animals, Mean±SEM) h Representative coronal image showing ChR2-EYFP expression in TNSST neurons. i Total distance moved and j moving velocity when TNSST terminals in PAG were activated by LED with or without food presented. One-way ANOVA followed by Tukey’s multiple comparisons. Adjusted p value for comparing red (no-led, no-food) and yellow (with-led, with-food) groups is 0.0023; for comparing red (no-led, no-food) and blue (with-led, no-food) groups is 0.0045.
To confirm the functional connectivity of the VTA-TN-PAG circuit, we injected retrograde cre-dependent mCherry-expressing AAV virus in PAG to label the PAG-projecting TNSST neurons, and cre-dependent ChR2 in VTA of TH-cre::SST-cre mice (Fig. 6a). By patching red TN cells in acute brain slices, optogenetic stimulation of the VTA dopaminergic terminals in TN enhanced activity of PAG-projecting TNSST neurons (Fig. 6b–d). In vivo optogenetic stimulation of VTA dopaminergic neural terminals in TN resulted in more locomotion (Fig. 6e, f) without changing food intake (Supplementary Fig. S8b), similar to that of stimulating TNSST neural terminals in PAG (Fig. 5i). Further analysis of the movement trajectory showed that such increased locomotion resulted in more food approaching events, revealed by repeated approaching of the food port (Fig. 6g, h). Based on our tracing results, we hypothesize that silencing PAG-projecting TNSST neurons would block the effect of activating VTA dopaminergic terminals in TN. To examine this hypothesis, we injected retrograde cre-dependent GtACR1 in PAG and cre-dependent ChR2 in VTA of the TH-cre::SST-cre mice (Fig. 6i), so that blue light will activate VTA dopaminergic terminals and green light will inhibit PAG-projecting TNSST neurons. Whole-cell patch recording of red cells in acute slices showed that green light caused clear inward current and hyperpolarised the cells without notable rebound activity after turning off the light (Fig. 6j–l). In vivo optogenetic stimulation of VTA terminals and concurrent optogenetic silencing of PAG-projecting TNSST neurons failed to induce more locomotion and food approaching events (Fig. 6m–p). Together, these results suggest that the PAG-projecting TNSST neurons receive dopaminergic inputs, which may contribute to their response to palatable food during the approaching phase, and their output in PAG could facilitate food approaching behavior.
a Schematic diagram of the experimental strategy, indicating the viral injection and fiber implantation for optogenetic stimulation. b A representative image showing successful labelling of PAG-projecting TNSST neurons and projection of VTA dopaminergic neural terminals in TN. c A representative trace showing the effect of optogenetic stimulation of VTA dopaminergic terminals on TNSST neurons by whole-cell patch recording. d Quantification of the optogenetic stimulation effect by whole-cell patch recording of TNSST neurons. (Mean±SEM, two-tailed paired t-test, p = 0.0489, *p < 0.05) e Quantification of the effect of optogenetic stimulation of VTA dopaminergic terminals in TN on locomotion. n = 10, ***p < 0.001 with paired two-sided t-test. f Representative plots showing the trajectory of an animal in the behavioral chamber with or without optogenetic stimulation. The color indicates locomotion speed. g Quantification of the effect of optogenetic stimulation of VTA dopaminergic terminals in TN on food approaching behavior. n = 10, ***p < 0.001 with paired two-sided t-test. h Representative plots showing the distance to the food port during sessions with or without optogenetic stimulation. i Schematic diagram of the experimental strategy, indicating the viral injection and fiber implantation for optogenetic stimulation/inhibition. j A representative image showing patch clamp of a GtACR1-expressing PAG-projecting TNSST neuron. k Representative traces showing the effect of optogenetic inhibition on TNSST neurons by whole-cell patch recording. l Quantification of the optogenetic inhibition effect by whole-cell patch recording of TNSST neurons. (Mean±SEM, two-sided paired t-test, p = 0.0089, **p < 0.01) m Quantification of the effect of concurrent optogenetic stimulation of VTA dopaminergic terminals in TN and optogenetic inhibition of PAG-projecting TNSST neurons on locomotion. n = 11, paired two-sided t-test. n Representative plots showing the trajectory of an animal in the behavioral chamber with or without optogenetic manipulation. The color indicates locomotion speed. o Quantification of the effect of concurrent optogenetic stimulation of VTA dopaminergic terminals in TN and optogenetic inhibition of PAG-projecting TNSST neurons on food approaching behavior. n = 11, paired two-sided t-test. p Representative plots showing the distance to the food port during sessions with or without optogenetic manipulation.
Discussion
Many hypothalamic orexigenic neurons have been identified but how they interact with each other and how they together enable proper feeding behavior are still poorly understood2. In general, hypothalamic neurons, especially those close to the ventricles, have been considered mainly sensing peripheral nutritional or hormonal signals for exerting their functional output to regulate energy homeostasis. However, it has also been suggested that feeding behavior can be modulated by non-nutritional top-down information8,9. Relatively understudied, the TNSST neurons were recently reported as an important regulator of feeding6. To better understand the circuit mechanisms of TNSST neurons, we implemented the VITALISTIC method, combined with single cell projectome mapping by fMOST. Such analysis greatly facilitated our understanding of the modulation and functional impact of these neurons.
The VITALISTIC method
Mapping the input-output configuration of specific neurons has been very helpful in understanding their functional impact and neural mechanisms. It has been achieved by viral tracing and either mechanical sectioning of fixed brain tissue or optical sectioning of transparent brain tissue13,23,38,39. While mono-synaptic input tracing using modified rabies has been widely used18, its combination with tissue clearing and subsequent light sheet imaging is surprisingly rare40, probably because: 1) brain-wide input mapping is not a major focus of many studies utilising rabies tracing; 2) the limited availability of light sheet microscopes; and 3) challenging in analysing large scale whole brain imaging data. The recently developed mesoSPIM system provides the possibility for generating relatively smaller data size for whole brain imaging19, and thus makes brain-wide analysis more accessible for more neuroscience labs. Although we’re not the first one combining these different techniques40,41,42, we believe that it will become more widely used for understanding neural circuit configuration, and thus we coined the term VITALISTIC method so that it is easier to refer to the pipeline combining viral tracing, tissue clearing and light sheet imaging.
The advantage of tissue clearing and whole brain imaging has been reviewed extensively20. It appears to us that quantitative whole brain analysis could greatly reduce potential bias introduced by experimenters, either in the process of tissue sectioning or in the process of registering imaged data to the standard brain atlas. Taking input mapping for TNSST neurons as an example, although each cortical region contains a small number of input neurons, their combination actually accounted for almost 10% of all inputs to TNSST neurons. Manual analysis by experimenter might bias towards more ‘reasonable’ brain regions while neglecting sparse signal from more ‘unlikely’ brain regions based on an individual’s hypothesis. Therefore, an automatic imaging and analysis pipeline would provide an unbiased and quantitative result.
The differences between AgRP and TNSST neurons
A striking difference between the inputs of AgRP and TNSST neurons is the percentage of hypothalamic vs. extra-hypothalamic inputs. While AgRP neurons received mainly hypothalamic inputs, TNSST neurons received a considerable amount of inputs from many other major brain regions (Fig. 2b). Such disparity might reflect the potentially different roles of these two major orexigenic neurons. With more hypothalamic inputs, AgRP neurons are at a better position to integrate internal status such as temperature and other innate behaviors. Indeed, AgRP, but not TNSST, neurons were found to mediate cold-induced food consumption43. On the other hand, the more prominent inputs from extra-hypothalamic regions to TNSST neurons suggest that these neurons may be the main target of non-nutritional top-down regulation, and may suggest the involvement of TNSST neurons in mediating maladaptive eating behaviors. For example, TNSST, but not AgRP, neurons were found to mediate contextual conditioned feeding8, in which hippocampal inputs mediate a maladaptive behavior. Notably, human genetics and behavior analysis, combined with mouse hypothalamus single cell sequencing44, found that different clusters of SST+ neurons, but not AgRP neurons, enrich most genetic loci that are significantly associated with excess carbohydrate and fat intake45. Combined with another mouse single cell sequencing dataset46, the same report also found that SST_Pthlh neurons enrich more genes related with human dietary intake than AgRP neurons45. The SST_Pthlh neurons are indeed TNSST neurons according to the expression pattern of Pthlh in the hypothalamus (Allen Brain Atlas ISH experiment 73592531). The input pattern of TNSST neurons therefore supports the idea that TNSST neurons may be a key target of maladaptive feeding behavior that also have eminent impact for human behavior. Further study focusing on the plastic change of other extra-hypothalamic inputs, such as those from mPFC, may help to understand different forms of maladaptive feeding behavior.
Another notable difference between AgRP and TNSST neurons is their inputs from neuromodulator centers. Consistent with their difference in input weights, the expression level of major neuromodulator receptors also show difference between these two neuronal types. Interestingly, although AgRP neurons express relatively less Drd1 receptor and receive little input from VTA or SN, the Drd1 signalling in AgRP neurons has been reported to play a role in feeding regulation30,31, indicating other dopaminergic sources to AgRP neurons. The obvious input from the histamine center, tuberal mammillary nucleus, and the expression of histamine receptors on both AgRP and TNSST neurons indicate that histaminergic neurons may directly target the key orexigenic neurons to suppress feeding, a well-known effect of histamine47,48,49. While the histamine H1 receptor has gained attention in drug development, the prominent expression of H3 receptor in both AgRP and TNSST neurons warrants a further investigation of the their function50. The overall more neuromodulatory inputs and higher expression of neuromodulator receptors in TNSST neurons supports the idea that they could be a main target of top-down regulation of feeding behavior.
Potential hierarchical neural circuits for orchestrating feeding behavior
While several types of hypothalamic orexigenic neurons have been found to stimulate feeding3,4,5,6,7, whether and how they cooperate and contribute to the different aspects of feeding is still unclear. Our anatomical mapping revealed interesting connection between some of these hypothalamic orexigenic neurons. While AgRP neurons received more inputs from TN, they received less input from LHA and ZI compared with TNSST neurons (Supplementary Fig.1 and Supplementary Fig. 3), indicating that LHA and ZI are two major regulators of TN and TN is a major regulator of AgRP neurons. Because ZI predominantly has GABAergic neurons28, their input to TNSST neurons is highly likely inhibitory. Likewise, because TN mainly contains GABAergic neurons6, their connection with AgRP neurons is also most likely inhibitory. Because TNSST neurons also project to LHA (but less prominently to ZI) and receive inputs from the arcuate, it appears that there are reciprocal connections between TN and LHA/ZI neurons, as well as between TN and arcuate neurons. Such a reciprocal configuration would make sense if these different orexigenic neurons regulate different aspects or phases of feeding behavior.
Previous in vivo imaging work has found that the activity of AgRP neurons is increased in hunger state but is rapidly suppressed by the presence of food, without approaching or consuming food51,52. On the other hand, TNSST neurons respond when approaching food, especially palatable food, but not consuming food8, and some LHA/ZI GABAergic neurons show higher activity when consuming food4,28. Moreover, we found that the PAG-projecting TNSST neurons could promote locomotion, suggesting that the activity of TNSST neurons during food approach might contribute to approaching behavior. If we consider feeding as a serial action involving foraging, approaching, consuming and stopping, the activity profiles of AgRP, TNSST and LHA/ZI GABAergic neurons seem to match well with the sequential order of different phases of feeding behavior, and other anorexic neurons such as PBN neurons may function as a terminator of the consumption phase through projection to various brain regions53,54,55. Importantly, the reciprocal connections between AgRP, TNSST and LHA/ZI would provide a plausible neural mechanism underlying the sequential activity profile of them.
While lateral hypothalamus (LHA) GABAergic neurons also play important roles in feeding4, they don’t seem to be well connected with either AgRP or TNSST neurons. In our study, we found that the major connection between LHA and TNSST neurons is glutamatergic, while even less connection between LHA and AgRP neurons were found using electrophysiological recording, consistent with our current and others’ previous anatomical mapping13. Therefore, AgRP and TNSST neurons may be modulated by glutamatergic or neuromodulatory inputs from LHA, and the LHA GABAergic neurons may be downstream targets of TNSST neurons. It remains possible that ZI GABAergic neurons interact with LHA GABAergic neurons for orchestrating consumption behavior or prey hunting behavior36,56.
Outputs of TNSST neurons
While the VITALISTIC method performed reasonably well for mapping input, mapping output by imaging synaptic terminal is still challenging using mesoSPIM system. Using the highest magnification of the mesoSPIM system, the synaptic terminal is at its detection limit. Therefore, we adopted results from fMOST for more precise mapping of the output configuration with higher image resolution. Such output mapping provided unprecedented details of the projection pattern of TNSST neurons, which will greatly advance our understanding of their neural network in feeding regulation and beyond. Our previous report suggested a partial one-to-many projection pattern of TNSST neurons6, which is further supported by single neuron projectome mapping in the current study. The projection to BNST/BST turned out to be one of the most prominent outputs of TNSST neurons, strongly indicating the critical role of BNST in feeding regulation. It is possible that the BNST is a master hub in facilitating feeding behavior, so that many TNSST neurons project to BNST while different subpopulations project to different other brain regions for fine-tuning feeding behavior (i.e. PAG for promoting locomotion) or concurrently modulating other physiological functions. It should be noted that both AgRP and TNSST promote feeding through their terminals in BNST57; the exact role of BNST in feeding regulation worth a deeper investigation58,59. Moreover, the distinct projection to LS was underappreciated before. The LS neurotensin-positive GABAergic neurons project to TN for regulating hedonic feeding60, supporting a potential reciprocal inhibitory connection between TN and LS for delicately modulating hedonic feeding behavior.
It is important to keep in mind that current output mapping is based on measuring the length of EYFP-labelled axon segments in various brain regions. While these measurements correlate with EYFP-labelled synaptic terminal counts, they should not be interpreted as actual synaptic outputs. The distribution of presynaptic terminals along axon is unlikely to be uniform through the whole axon, and the proximal axon segments in hypothalamus are unlikely to harbour similar density of presynaptic terminals as distal axon segments in other regions. As a result, we have to interpret the output weights measured by axon length cautiously. For example, the output in hypothalamus, especially within TN, may be over-estimated. Therefore, future output mapping using more definitive presynaptic marker such as synaptophysin-GFP should be done for a more accurate description of the output pattern. Nevertheless, the current mapping based on axonal signal provided a great starting point and an initial framework for future investigations.
Together, using the VITALISTIC and fMOST methods, we revealed the input output configuration of TNSST neurons, and contrasted important differences between that of AgRP neurons. The input of TNSST neurons that project to various brain areas is tailored to correspond to different functional outputs. Anatomical analysis combined with functional assays leads to a better understanding of the configuration of the hypothalamic orexigenic neural networks, and we propose a hierarchical neural network in orchestrating different phases of feeding behavior.
Methods
Mice
All experimental procedures involving use of animals were approved by the Institutional Animal Care and Use Committee of the Agency for Science, Technology and Research (A*STAR) of Singapore. Animals were housed in a standard 12 h light-dark cycle with access ad libitum to water and standard mouse chow (Harlan 2018 Teclab Global 18% protein rodent diet). 2–5 months old female and male mice were used for all experiments. SST-Cre (stock # 013044; RRID: IMSR_JAX:013044), AgRP-Cre (stock # 012899; RRID: IMSR_JAX: 012899) and Ai14 (stock # 007914; RRID: IMSR_JAX:007914) were purchased from Jackson Laboratories. SST-Flp mice were kindly provided by Dr. Josh Huang at Cold Spring Harbor Laboratory. TH-Cre mice were kindly provided by Dr. Luis de Lecea at Stanford University.
Viral vectors and stereotaxic surgery
We purchased AAVrg-Cre from University of North Carolina (UNC) vector core and CAV2-Cre from Plateforme de Vectorologie de Montpellier (PVM). The viruses used in monosynaptic retrograde tracing AAV8/Dj-EF1a-DIO-oG-WPRE-hGH, AAV8-CAG-FLEx (FRT)-TC (TVA-mCherry) and EnvA pseudotyped G-deleted rabies-GFP were purchased from the viral vector core of Salk Institute for Biological Studies. AAV-hSyn-ChR2-EYFP and AAVrg-DIO-stGtACR1-fusionred viruses were purchased from BrainVTA. Stereotaxic surgeries performed as following as previous studies6. Briefly, animals were anesthetized with Ketamine/Xylazine mixture (0.1 µl/10 g), put artificial tears ointments, and kept at 33 °C on the heating pad. Atipamezole (rehearsal) and buprenorphine (analgesic) were administered to animals.
For monosynaptic retrograde input tracing of TNSST and AgRP neurons, animals were injected with 0.3 µl equal volume mix of AAV8/Dj-EF1a-DIO-oG-WPRE-hGH and AAV8-CAG-FLEX-TC (TVA-mCherry) by 10 µl NANOFIL syringes with 33 Gauge needles (NF33BV-2) and Micro4 syringe pump (World Precision Instruments SYS-Micro4) at 2 nl/sec into tuberal nucleus (TN) (AP: −1.6, ML: ±0.9, DV: −5.5) and arcuate nucleus (ARC) (AP:−1.5, ML: ±0.2, DV:−5.80) in SST-Cre and AgRP-Cre mice respectively. 3 weeks later 0.35 µl EnvA G- deleted rabies GFP (RV-GFP) was injected into same region. 8 days post-rabies injection mice were sacrificed for CUBIC clearing.
For projecting specific TNSST retrograde tracing either 0.2 µl AAVrg-Cre or CAV2-Cre were injected in following coordinates unilaterally: bed nuclei of the stria terminalis (BNST) (AP: +0.6, ML: ±1.0, DV: −3.7), central amygdalar nucleus (CEA) (AP: −1.0, ML: ±2.4, DV: −4.2), periaqueductal gray (PAG) (AP: −4.3, ML: ±0.4, DV: −2.2), paraventricular hypothalamic nucleus (PVH) (AP: −0.5, ML: ±0.2, DV: −4.6). 2-3 weeks after recovery and proper virus expression, 0.3 µl equal volume mix of AAV8/Dj-EF1a-DIO-oG-WPRE-hGH and AAV8-CAG-FLEx(FRT)-TC (TVA-mCherry) injected into TN. Following injection and timeline is similar with the monosynaptic retrograde input tracing, brains were processed for histology and imaging.
To labelling inputs to TN in VTA-SN region for cFos experiment, WT mice were injected with 0.2 µl of 0.25% retrograde tracer Cholera Toxin subunit B (CTB555).
For electrophysiology experiments, SST-Cre:Ai14 mice were injected with 0.2 µl AAV5-hSyn-hChR2(H134R)-EYFP in anterior olfactory nucleus (AON) (AP: +3.00, ±0.3, DV: −2.70), lateral hypothalamus (LH) (AP:−1.0, ML: ±1.10, DV: −5.10).
For optogenetic experiments, SST-Cre mice were injected into TN with 0.15 µl AAV9-Ef1α-DIO-hChR2(H134R)-EYFP. Fiber optics (core diameter: 200 μm, numerical aperture: 0.37, Inper, China) were implanted into the PAG (AP: −4.35, ML: ±0.0, DV: −1.85) immediately after viral injections and were secured to the skull with bone screws and dental cement.
Histology and clearing
Animals were transcardially perfused with phosphate buffered saline (PBS) and 4% paraformaldehyde (PFA) in PBS. Collected brains post-fixed in 4% PFA overnight. Then, transferred into 15% sucrose (4 h) and 30% sucrose (overnight) respectively. Brains were prepared for cryostat sectioning by freezing in OCT compound. 40 µm brain sections were collected in PBS. For monosynaptic input tracing of TNSST neurons, brains were sectioned at 50 µm in the coronal plane with Leica Cryostat (CM1950) and were stored in 1X PBS at 4 °C until further processing. For the quantifications, we used Allen Institute’s reference atlas to assign regions and manually quantified the cells in every region. Further data analysis was carried out using ImageJ software and GraphPad Prism8. All values were presented as the Mean ± SEM. For relevant immunohistochemistry experiment, free-floating coronal sections were prepared. The sections were washed with PBS 3 × 5 min followed by incubation in the blocking buffer (5% serum, 3%BSA and 0.4% Triton X-100 in PBS) for 2 h at room temperature. Sections were incubated in Rabbit anti-TH (1:1000, Millipore AB152) in blocking solution at 4 °C overnight. Sections were washed in 0.1% Triton X-100 in PBS for 3 × 5 min, followed by Alexa Flour coupled donkey anti-rabbit 488 (1:500; Invitrogen A21206) incubation in blocking solution for 2 h at room temperature. Sections were washed and incubated with PBS for 3 × 5 min, followed by Hoescht stain in 1 X PBS (1:1000) for 10 minutes at room temperature. At the end sections were mounted onto SuperFrost (Fisherbrand, #12-550-15) microscope slides, Fluoromount-G (Southern Biotech, 0100-01) was applied and coverslips were sealed with nail polish. Sections were imaged using A1R+ si confocal microscope in Nikon Imaging Centre in Singapore Bioimaging Consortium.
To optimize brain clearing method CUBIC for whole brain imaging with light sheet microscope, protocol has adopted from Susaki et al.24. Briefly, deeply anesthetized animals, overdose Ketamine/Xylazine mixture, transcardially perfused with 20 ml cold PBS and 30 ml cold 4%PFA. Brains were collected and further incubated in 4% PFA on shaker at 4 °C for 24 h. Brains were washed with PBS for 2 h 3 times to remove PFA before clearing. Then, they were immersed in 50% CUBIC-L (1:1 mixed with ddH2O) solution for 6 h. Further, transferred into CUBIC-L with gentle shaking at 37 °C for 5 days. Solution has been refreshed every day. After delipidation, brains were washed 2 h 3 times with PBS at room temperature. Then, brains transferred into the 50% CUBIC-R (1:1 mixed with ddH2O) solution for 6 h. Further, incubated in CUBIC-R at room temperature with shaking for Reflactive Index (RI) matching.
Light sheet imaging and data analysis
Cleared brain tissues were imaged using custom built mesoSPIM microscope19, equipped with 488 nm, 561 nm and 647 nm Toptica lasers. Images acquired by 647 nm illumination were used for Allen Brain Atlas registration using CCFv3 atlas. Images were analysed by Magellanmapper27. We included all level 8 brain regions and excluded fiber tracts and cerebellum in final results.
(The following is paraphrased from previously published method paper27,61): We detected nuclei in cleared mouse brains using a 3D Laplacian of Gaussian blob detection technique throughout each whole brain. To perform detections in large images several hundred gigabytes (GB) to over a terabyte (TB) in size, we subdivided the image into many smaller chunks to reduce RAM requirements and maximize parallel processing. We loaded images through the Numpy library’s memory mapped method (load with the mmap_mode option) to load only the necessary parts of the image on-the-fly, allowing us to load small images a chunk at a time without reading the entire volumetric image into memory. We divided the image shape into overlapping chunks to ensure that nuclei at borders would not be missed, with overlap size of approximately the nucleus diameter. After determining the offset and shape of each chunk, we set the image array as a class attribute, initiated multiprocessing (the multiprocessing. Pool in the standard Python library), and accessed each chunk as a view in a separate process via class methods to avoid duplicating arrays in memory. Thus, we could control total memory usage by the size of chunks and the number of CPU (central processing unit) cores available for separate processes.
3D cell detection poses a number of challenges including adapting to local variation such as staining inhomogeneity, background variation, and autofluorescence, in addition to overlapping cells in dense tissue62. To address these issues, we analyzed images in a local manner by further subdividing each chunk for preprocessing based on its immediate surroundings. We split each chunk into sub-chunks using the same approach as above but ran each sub-chunk serially within each CPU process. In each sub-chunk, we first clipped intensity values at the 5th and 98.5th percentiles (percentile in Numpy) to remove extreme outliers, rescaled the intensities from 0 to 1, and further saturated signal by clipping the rescaled intensities at the 50th percentile. We next enhanced edges by using unsharp masking with a Gaussian sigma of 8 (filters.gaussian in scikit-image) to identify sharp details as the difference between an image and its blurred version (which we amplified by a factor of 0.3) and adding back those details to the original image. We mildly eroded the resulting signal with an erosion filter using an octahedron structuring element of size 1 (morphology.erosion with morphology.octahedron in scikit-image) to separate out blobs.
To detect blobs, we implemented the 3D Laplacian of Gaussian blob detector from the scikit-image library (feature.blob_log) as a multi-scale interest point operator. We set the minimum and maximum sigma based on the microscopy resolution, with 10 intermediate values, detection threshold of 0.1, and overlap fraction threshold of 0.55 below which duplicated blobs are eliminated. Initially we missed many nuclei positioned above one another in the z-direction, likely because the anisotropy necessitated by the relatively thick lightsheet at 5x in our setup limited resolution in the z-direction. To improve detection along the z-axis, we interpolated the images in this direction to near isotropy before detection (transform.resize in scikit-image). The blob detector had a tendency to cluster detections in the bottom and topmost z-planes in each ROI from nuclei visible within the ROI but whose centroids are outside. To avoid this clustering and minimize duplication with adjacent ROIs, we cropped nuclei from these planes on the assumption that they would be captured better in the adjacent, overlapping ROIs.
Overlapping chunks minimized missing nuclei at edges but also necessitated pruning blobs duplicately detected in adjacent chunks. Pruning involves checking for duplicates within all potentially overlapping regions. Since the overlapping portions of the regularly spaced chunks collectively form a grid pattern throughout the full volumetric image, we could limit our search to these grid planes along each axis. After completing detection on the whole image, we first pooled all detected blobs into a single array. Along a given axis of the full image, we determined the boundaries for each overlapping region and all of its blobs. Within each overlapping region, we found all blobs close to another blob by taking the absolute value of the difference between all blobs with one another and finding blobs within a given tolerance in all dimensions. The tolerance was titrated so that the ratio of final blobs in overlapping regions to the next adjacent regions of same volume was about 1:1. For each close pair of blobs found, we replaced both blobs with a new blob that took the mean of their coordinates. To minimize memory usage, we checked smaller groups of blobs against one another until completing all comparisons. We checked overlapping regions simultaneously in multiprocessing along a given axis for efficiency, re-pooled all blobs, and pruned along the next axis to account for blobs that may have been duplicately detected in overlapping chunks along multiple axes, at grid intersections.
Parameters for preprocessing, detections, and pruning steps were optimized through a Grid Search approach, a type of hyperparameter tuning, to check combinations of parameters systematically. To evaluate the accuracy of each set of parameters, two students in our lab generated truth sets of nuclei locations and radii using our serial 2D nuclei annotation tool, taking ROIs of size 42 × 42 × 32 pixels (x, y, z) from representative ROIs of all major brain structures (n = 15 forebrain, eight midbrain, and seven hindbrain; n = 2766 nuclei). They separately generated additional truth sets at a slightly lower magnification (4x), size 60 × 60 × 14 pixels (n = 40 ROIs, 1116 nuclei) to increase representation. After detecting nuclei on these images with a given set of parameters, matches between detections and ground truth was determined using the Hungarian algorithm, a combinatorial optimization method to determine optimal assignments between two sets, as implemented in optimize.linear_sum_assignment in Scipy. After scaling nuclei coordinates for isotropy, we found the Euclidean distances between detected and truth nuclei points through distance.cdist, which serves as the cost matrix input to optimize.linear_sum_assignment to find optimal pairings between points based on closest distance. We took correctly identified detections, or true positives (TP), as pairings within a given tolerance distance. Unpaired detections or those in pairs exceeding this threshold were considered false positives (FP), and the same for ground truth were false negatives (FN). Since a match for a given nucleus within the ROI may lie outside of it and thus go unseen, we first searched for pairings only within an inner sub-ROI, followed by a secondary search for pairings between only unmatched inner sub-ROI objects and the rest of the ROI. This approach reduced the total number of nuclei available but avoided missed border matches. As measures of performance of our detection compared with ground truth, we used the following standard equations:
Image downsampling and registration
To assign nuclei to the proper brain label, we employed automated label propagation by registering the E18.5 atlas to each of our imaged mouse brains using SimpleElastix, a toolkit that combines the programmatic access of SimpleITK to the Insight Segmentation and Registration Toolkit (ITK) with the Elastix image registration framework. Elastix has been recently validated as a computationally efficient and accurate tool for registration of mouse brains cleared by CUBIC63.
Registration involved rigid followed by non-rigid alignment. For rigid registration, we employed a translation (translation parameter map in SimpleElastix) with default settings except increasing to 2048 iterations (MaximumNumberOfIterations setting) followed by an affine (affine parameter map) with 1024 iterations, applied with an ElastixImageFilter, to shift, resize, and shear the atlas microscopy image to the same space as that of the sample brain. For non-rigid alignment, we employed a b-spline strategy (bspline parameter map) guided by the AdvancedNormalizedCorrelation similarity metric with grid spacing of size 60, measured in voxels rather than physical units (FinalGridSpacingInVoxels setting in place of FinalGridSpacingInPhysicalUnits), over 512 iterations. The TransformixImageFilter in SimpleElastix allowed us to apply the identical registration transformation to the atlas labels image, except that we set the final b-spline interpolation order (FinalBSplineInterpolationOrder) to 0 to avoid interpolating any new values, preserving the labels’ specific set of integer values. We applied this identical transformation to both the mirrored and edge-refined atlas labels.
To evaluate the level of alignment from registration, we measured the similarity between each registered atlas histology and its corresponding sample image using a Dice Similarity Coefficient (DSC) as implemented by the GetDiceCoefficient function in SimpleElastix/ SimpleITK, given by the equation:
where S and T are two different sets of voxels. We took the foreground of each atlas and sample microscopy image to be its mean threshold (filters.threshold_mean function in scikit-image) and input them to a LabelOverlapMeasuresImageFilter to take the DSC.
Circos plots were generated as described in http://circos.ca/64.
Sparse labelling and fMOST mapping
AAV-hSyn-Con/Fon-EYFP and AAV-EF1a-FlpO were generated during the virus package step (1:8000 mixture ratio, 1.41 x 10^13 genomic copies/mL) by the Gene Editing Core Facility of the Institute of Neuroscience in Shanghai. 50 nl of the virus was injected into the TU of SST-Cre mice. 29 days after viral injection, the brains were harvested and post-fixed in 4% paraformaldehyde, followed by being embedded in Lowicryl HM20 resin (Electron Microscopy Sciences, 14340). The resin-embedded brains were imaged in a water bath containing propidium iodide (PI) under an fMOST microscope at a voxel resolution of 0.32 μm x 0.32 μm x 1 μm32. The sample was imaged for YFP (for tracing) and PI channels (for brain registration) at 1 μm step by a fixed diamond knife. The imaging-sectioning cycle was continued until the entire brain sample was fully imaged. The imaged neurons were traced using Fast Neurite Tracer (FNT), a software developed in previous publication32. Each traced neuron was independently checked by another person to ensure accuracy and each brain sample was subjected to random post-tracing quality check. The reconstructed neuron images were registered into the standard Allen CCFv3 as previously described32. The soma location, axon length and terminal number for each neuron was analysed using NeuroView (https://gitee.com/bigduduwxf/NeuronView), and visualised using neuron-vis (https://gitee.com/bigduduwxf/neuron-vis). A modified Hausdorff match distance65 was calculated for neuron pairs. After generating the similarity index of all neuronal pairs, we performed hierarchical clustering of the matrix using Ward’s linkage to identify projectome-defined neuron subtypes. More details and all dataset can be found in a published paper66.
Multiflex fluorescence in-situ hybridization (FISH) using RNAScope®
Fresh Frozen Sample Prep and RNAScope® Fluorescent Multiplex Assay kit (ACDBio®) have been used for in-situ hybridization. Animals were deeply anesthetized with overdose Ketamine/Xylazine mixture and cervical dislocated. Brains were collected, immediately frost in OCT on dry ice and stored at −80 °C. Fresh frozen brains were sectioned at 20 µm on the slides, post-fixed in cold 4% PFA at 4 °C for 1 h followed, quickly washed with PBS by dehydration in 50%, 70% and 100% ethanol, respectively. Slides were dried at 40 °C for 20 min and permeabilized at room temperature with Protease IV provided with the kit. Probe hybridization was done at 40 °C for 2 h with HybEZ™ Oven. After hybridization, slides were washed in the Wash Buffer provided in the kit, then incubated in order of amplification reagents (amp 1–4). After incubating with DAPI for 3 min slides were mounted and coverslipped. RNAScope® probes used in study: cfos, Chrm1, Chrm2, DAT, Drd1, Drd2, AgRP and SST.
For cFos staining in TN inputs in VTA-SN region
CTB555 injected animals were habituated with the high fat prior to experiment. Overnight fasted animals received high fat and let it consume for 1 h. Then, animals were sacrificed for DAT and cfos double fluorescence in-situ hybridization.
Whole-cell patch recording in acute slices
SST-Cre::Ai14 or AgRP-Cre::Ai14 mice were injected 0.8 µl AAV5-hSyn-ChR2-YFP in AON or 0.5 µl AAV5-hSyn-Chr2-YFP in LHA for allowing ChR2-expression in both glutamatergic and GABAergic neurons. Experiments were performed on acute brain slices. Mouse brains were rapidly removed after decapitation and placed in high sucrose ice-cold oxygenated artificial cerebrospinal fluid (ACSF) containing the following (in mM): 230 sucrose, 2.5 KCl, 10 MgSO4, 0.5 CaCl2, 26 NaHCO3, 11 glucose, 1 kyneurenic acid, pH 7.3, 95% O2 /5% CO2. Coronal brain slices were cut at a thickness of 250 um using a vibratome (VT1200S; Leica Biosystems) and immediately transferred to an incubation chamber filled with ACSF containing the following (in mM): 119 NaCl, 2.5 KCl, 1.3 MgCl2, 2.5 CaCl2, 1.2 NaH2PO4, 26 NaHCO3, and 11 glucose, pH 7.3, equilibrated with 95% O2 and 5% CO2. Slices were allowed to recover at 32 °C for half an hour and then maintained at room temperature. Recordings were performed with pCLAMP11.
Whole-cell patch-clamp recordings were performed on TdTomato expressing neurons in TN of SST-Cre::Ai14 mice, or AgRP neurons in arcuate of AgRP::Ai14 mice. The cells were visualized using a CCD camera and monitor. Pipettes used for recording were pulled from borosilicate glass capillary tubes with filament (length 100 mm, outer diameter 1.5 mm, inner diameter 0.84 mm, WPI) using a P-1000 Flaming/Brown Micropipette Puller (Sutter Instruments). Patch pipettes (3–5 MΩ) were filled with internal solution containing (in mM): 105 K-gluconate, 30 KCl, 4 MgCl2, 10 HEPES, 0.3 EGTA, 4 Na-ATP, 0.3 Na-GTP, and 10 Na2-phosphocreatine (pH 7.3 with CsOH; 295 mOsm). The access resistance, membrane resistance, and membrane capacitance were monitored during the experiment to ensure the stability and the health of cells. To evoke synaptic transmission using ChR2, photostimulation (470 nm) was delivered by LED illumination system (pE-4000) and synaptic responses were recorded at holding potentials of −70 mV. A 525 nm light was delivered for checking response of GtACR1-expression cells.
Optogenetic stimulation for analysing locomotion
The open field test was used to measure locomotor activity of mice. Mice were handled and underwent tethering habituation processes prior to the test. The open field test consisted of a 30 min session in which mice were placed into the centre of an arena (40 cm × 40 cm × 40 cm) and allowed to move freely. A video camera positioned directly above the arena was used to track the movement of each mouse by a tracking software (EthoVision XT, Noldus). For optogenetic activation of the projections of TNSST neurons in PAG, a 473 nm laser (MBL589, CNI Lasers, China) was delivered through a rotary joint (FRJ_1X2i_FC-2FC, Doric) and related patch cords connected to the fiber optic implanted in mouse heads. A 20 Hz, 5 ms pulses stimulation paradigm was applied with a 3 s on 1 s off for 30 min, triggered by a stimulator (Model 2100, Isolated pulse stimulator, A-M systems) that was controlled by the EthoVision XT software. The laser power was adjusted to 3–5 mW/mm2 at the end of the optic fiber. For optogenetic activation with food, standard normal chow was presented. The total locomotor distance travelled and velocity were analysed. The apparatus was cleaned with 75% ethanol between tests to remove any scent from the previous mouse.
For optogenetic stimulation in TN, light from a blue LED (Prizmatix, 460 nm) was delivered through a rotary joint (FRJ_1X2i_FC-2FC, Doric) and related patch cords connected to the fiber optic ( ~ 2 mW/mm2) implanted in mouse heads. A 5s-on, 5s-off, continuous light stimulation was generated by a stimulator (Model 2100, Isolated pulse stimulator, A-M systems) that was controlled by the EthoVision XT software. For concurrent optogenetic inhibition, light from a green LED (Doric, Ce:YAG) was delivered into the same fiber ( ~ 2 mW/mm2) but controlled with a different stimulator for delivering a 7 s on, 3 s off, continuous green light with a 3 s delay to the start of blue light. Animal behavior was recorded by PhenoTyper and was analysed using DeepLabCut67. The coordinates of the animal head were used for calculating movement speed and trajectory.
Statistics
Statistical analyses were done using GraphPad Prism (GraphPad Software V8, Inc, La Jolla, CA), Python3.7.6.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Source data are provided with this paper. Any additional requests for information can be directed to, and will be fulfilled by, the corresponding authors. Source data are provided with this paper.
Code availability
Light sheet imaging data was analysed using an open source software Magellanmapper (https://github.com/sanderslab/magellanmapper). The analysed results were used to generate Circos plot using https://circos.ca/software/ and our custom codes that are available from https://github.com/yufu1981/plotting_results. Single cell projectome 3D visualization images were generated using codes from https://gitee.com/bigduduwxf/neuron-vis.
References
Locke, A. E. et al. Genetic studies of body mass index yield new insights for obesity biology. Nature 518, 197–206 (2015).
Sternson, S. M. & Eiselt, A. K. Three Pillars for the Neural Control of Appetite. Annu Rev. Physiol. 79, 401–423 (2017).
Aponte, Y., Atasoy, D. & Sternson, S. M. AGRP neurons are sufficient to orchestrate feeding behavior rapidly and without training. Nat. Neurosci. 14, 351–355 (2011).
Jennings, J. H. et al. Visualizing hypothalamic network dynamics for appetitive and consummatory behaviors. Cell 160, 516–527 (2015).
Krashes, M. J. et al. Rapid, reversible activation of AgRP neurons drives feeding behavior in mice. J. Clin. Invest 121, 1424–1428 (2011).
Luo, S. X. et al. Regulation of feeding by somatostatin neurons in the tuberal nucleus. Science 361, 76–81 (2018).
Zhang, X. & van den Pol, A. N. Rapid binge-like eating and body weight gain driven by zona incerta GABA neuron activation. Science 356, 853–859 (2017).
Mohammad, H. et al. A neural circuit for excessive feeding driven by environmental context in mice. Nat. Neurosci. 24, 1132–1141 (2021).
Stern, S. A. et al. Top-down control of conditioned overconsumption is mediated by insular cortex Nos1 neurons. Cell Metab. 33, 1418–1432 e1416 (2021).
Livneh, Y. et al. Homeostatic circuits selectively gate food cue responses in insular cortex. Nature 546, 611–616 (2017).
Horio, N. & Liberles, S. D. Hunger enhances food-odour attraction through a neuropeptide Y spotlight. Nature 592, 262–266 (2021).
Stutz, B. et al. AgRP neurons control structure and function of the medial prefrontal cortex. Mol. Psychiatry 27, 3951–3960 (2022).
Wang, D. et al. Whole-brain mapping of the direct inputs and axonal projections of POMC and AgRP neurons. Front Neuroanat. 9, 40 (2015).
Goto, M., Canteras, N. S., Burns, G. & Swanson, L. W. Projections from the subfornical region of the lateral hypothalamic area. J. Comp. Neurol. 493, 412–438 (2005).
Ugur, M., Doridot, S., la Fleur, S. E., Veinante, P. & Massotte, D. Connections of the mouse subfornical region of the lateral hypothalamus (LHsf). Brain Struct. Funct. 226, 2431–2458 (2021).
Kremer, H. P. H. the hypothalamic lateral tuberal nucleus: normal anatomy and changes in neurological diseases. Prog. Brain Res. 93, 13 (1992).
Callaway, E. M. & Luo, L. Monosynaptic Circuit Tracing with Glycoprotein-Deleted Rabies Viruses. J. Neurosci. 35, 8979–8985 (2015).
Luo, L., Callaway, E. M. & Svoboda, K. Genetic Dissection of Neural Circuits: A Decade of Progress. Neuron 98, 256–281 (2018).
Voigt, F. F. et al. The mesoSPIM initiative: open-source light-sheet microscopes for imaging cleared tissue. Nat. Methods 16, 1105–1108 (2019).
Ueda, H. R. et al. Whole-Brain Profiling of Cells and Circuits in Mammals by Tissue Clearing and Light-Sheet Microscopy. Neuron 106, 369–387 (2020).
Li, A. et al. Micro-optical sectioning tomography to obtain a high-resolution atlas of the mouse brain. Science 330, 1404–1408 (2010).
Gong, H. et al. Continuously tracing brain-wide long-distance axonal projections in mice at a one-micron voxel resolution. Neuroimage 74, 87–98 (2013).
Schwarz, L. A. et al. Viral-genetic tracing of the input-output organization of a central noradrenaline circuit. Nature 524, 88–92 (2015).
Susaki, E. A. et al. Advanced CUBIC protocols for whole-brain and whole-body clearing and imaging. Nat. Protoc. 10, 1709–1727 (2015).
Tainaka, K. et al. Chemical Landscape for Tissue Clearing Based on Hydrophilic Reagents. Cell Rep. 24, 2196–2210 e2199 (2018).
Matsumoto, K. et al. Advanced CUBIC tissue clearing for whole-organ cell profiling. Nat. Protoc. 14, 3506–3537 (2019).
Young, D. M. et al. Whole-Brain Image Analysis and Anatomical Atlas 3D Generation Using MagellanMapper. Curr. Protoc. Neurosci. 94, e104 (2020).
Zhao, Z. D. et al. Zona incerta GABAergic neurons integrate prey-related sensory signals and induce an appetitive drive to promote hunting. Nat. Neurosci. 22, 921–932 (2019).
Stuber, G. D. & Wise, R. A. Lateral hypothalamic circuits for feeding and reward. Nat. Neurosci. 19, 198–205 (2016).
Zhang, Q. et al. Food-induced dopamine signaling in AgRP neurons promotes feeding. Cell Rep. 41, 111718 (2022).
Gaziano, I. et al. Dopamine-inhibited POMCDrd2+ neurons in the ARC acutely regulate feeding and body temperature. JCI Insight 7, e162753 (2022).
Gao, L. et al. Single-neuron projectome of mouse prefrontal cortex. Nat. Neurosci. 25, 515–529 (2022).
Zhu, Y. et al. Dynamic salience processing in paraventricular thalamus gates associative learning. Science 362, 423–429 (2018).
Hu, H., Cui, Y. & Yang, Y. Circuits and functions of the lateral habenula in health and in disease. Nat. Rev. Neurosci. 21, 277–295 (2020).
Mazzone, C. M. et al. High-fat food biases hypothalamic and mesolimbic expression of consummatory drives. Nat. Neurosci. 23, 1253–1266 (2020).
Li, Y. et al. Hypothalamic Circuits for Predation and Evasion. Neuron 97, 911–924 e915 (2018).
Han, W. et al. Integrated Control of Predatory Hunting by the Central Nucleus of the Amygdala. Cell 168, 311–324 e318 (2017).
Beier, K. T. et al. Circuit Architecture of VTA Dopamine Neurons Revealed by Systematic Input-Output Mapping. Cell 162, 622–634 (2015).
Hunnicutt, B. J. et al. A comprehensive excitatory input map of the striatum reveals novel functional organization. Elife 5, e19103 (2016).
Menegas, W. et al. Dopamine neurons projecting to the posterior striatum form an anatomically distinct subclass. Elife 4, e10032 (2015).
Niedworok, C. J. et al. Charting monosynaptic connectivity maps by two-color light-sheet fluorescence microscopy. Cell Rep. 2, 1375–1386 (2012).
Mano, T. et al. CUBIC-Cloud provides an integrative computational framework toward community-driven whole-mouse-brain mapping. Cell Rep. Methods 1, 100038 (2021).
Yang, S. et al. An mPOA-ARC(AgRP) pathway modulates cold-evoked eating behavior. Cell Rep. 36, 109502 (2021).
Mickelsen, L. E. et al. Single-cell transcriptomic analysis of the lateral hypothalamic area reveals molecularly distinct populations of inhibitory and excitatory neurons. Nat. Neurosci. 22, 642–656 (2019).
Merino, J. et al. Genetic analysis of dietary intake identifies new loci and functional links with metabolic traits. Nat. Hum. Behav. 6, 155–163 (2022).
Campbell, J. N. et al. A molecular census of arcuate hypothalamus and median eminence cell types. Nat. Neurosci. 20, 484–496 (2017).
Clineschmidt, B. V. & Lotti, V. J. Histamine: intraventricular injection suppresses ingestive behavior of the cat. Arch. Int Pharmacodyn. Ther. 206, 288–298 (1973).
Itowi, N., Nagai, K., Nakagawa, H., Watanabe, T. & Wada, H. Changes in the feeding behavior of rats elicited by histamine infusion. Physiol. Behav. 44, 221–226 (1988).
Lecklin, A., Etu-Seppala, P., Stark, H. & Tuomisto, L. Effects of intracerebroventricularly infused histamine and selective H1, H2 and H3 agonists on food and water intake and urine flow in Wistar rats. Brain Res. 793, 279–288 (1998).
Ishizuka, T. & Yamatodani, A. Integrative role of the histaminergic system in feeding and taste perception. Front Syst. Neurosci. 6, 44 (2012).
Chen, Y., Lin, Y. C., Kuo, T. W. & Knight, Z. A. Sensory detection of food rapidly modulates arcuate feeding circuits. Cell 160, 829–841 (2015).
Betley, J. N. et al. Neurons for hunger and thirst transmit a negative-valence teaching signal. Nature 521, 180–185 (2015).
Campos, C. A., Bowen, A. J., Schwartz, M. W. & Palmiter, R. D. Parabrachial CGRP Neurons Control Meal Termination. Cell Metab. 23, 811–820 (2016).
Kim, D. Y. et al. A neural circuit mechanism for mechanosensory feedback control of ingestion. Nature 580, 376–380 (2020).
Phua, S. C. et al. A distinct parabrachial-to-lateral hypothalamus circuit for motivational suppression of feeding by nociception. Sci. Adv. 7, eabe4323 (2021).
Shang, C. et al. A subcortical excitatory circuit for sensory-triggered predatory hunting in mice. Nat. Neurosci. 22, 909–920 (2019).
Betley, J. N., Cao, Z. F., Ritola, K. D. & Sternson, S. M. Parallel, redundant circuit organization for homeostatic control of feeding behavior. Cell 155, 1337–1350 (2013).
Wang, Y. et al. A bed nucleus of stria terminalis microcircuit regulating inflammation-associated modulation of feeding. Nat. Commun. 10, 2769 (2019).
Zhang, J. et al. Dissection of the bed nucleus of the stria terminalis neuronal subtypes in feeding regulation. Physiol. Behav. 271, 114333 (2023).
Chen, Z. et al. A circuit from lateral septum neurotensin neurons to tuberal nucleus controls hedonic feeding. Mol. Psychiatry 27, 4843–4860 (2022).
Young, D. M. et al. Constructing and optimizing 3D atlases from 2D data with application to the developing mouse brain. Elife 10, e61408 (2021).
Shuvaev, S. A. et al. DALMATIAN: An Algorithm for Automatic Cell Detection and Counting in 3D. Front Neuroanat. 11, 117 (2017).
Nazib, A., Galloway, J., Fookes, C. & Perrin, D. Performance of Registration Tools on High-Resolution 3D Brain Images. Annu Int Conf. IEEE Eng. Med Biol. Soc. 2018, 566–569 (2018).
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009).
Mizrahi, A. et al. Comparative analysis of dendritic architecture of identified neurons using the Hausdorff distance metric. J. Comp. Neurol. 422, 415–428 (2000).
Jiao, Z. et al. Projectome-based characterization of hypothalamic peptidergic neurons in male mice. Nat. Neurosci. 28, 1073–1088 (2025).
Mathis, A. et al. DeepLabCut: markerless pose estimation of user-defined body parts with deep learning. Nat. Neurosci. 21, 1281–1289 (2018).
Acknowledgements
We thank Christine Chiam and Olivia Khoo for their excellent technical support; Dr. Josh Huang for providing SST-flp mice and Dr. Luis de Lecea for providing TH-ires-Cre mice. This work was supported by Biomedical Research Council (BMRC) of A*STAR (1530700142 to Y.F. and UIBR fund to Y.F.); the National Medical Research Council (NMRC) of Singapore (OFIRG20nov-0030 to Y.F.); Ministry of Science and Technology of China STI 2030-Major Projects (2021ZD0203203 and 2021ZD0204400 to X.X.); and the National Natural Science Foundation of China (31900708 to Q.L.).
Author information
Authors and Affiliations
Contributions
Y.F. conceived the project and wrote the manuscript. E.S. designed, implemented and analysed input and cTRIO mapping of the study. M.W. built mesoSPIM microscope. Y.X. and P.Z. contributed key behavioral experiments. Z.J. and X.X. contributed fMOST imaging and results analysis for output mapping. E.S., H.M. and Y.W. contributed rabies tracing and staining results. X.Y.Y., H.H. and S.Y.J. contributed electrophysiological recordings. T.S. and Q.L. contributed optogenetic and locomotion analysis. D.M.Y. implemented MagellanMapper in the lab. All authors discussed the results, commented on and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. [A peer review file is available].
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Senol, E., Wang, M., Xin, Y. et al. Brain-wide input-output analysis of tuberal nucleus somatostatin neurons reveals hierarchical circuits for orchestrating feeding behavior. Nat Commun 16, 5627 (2025). https://doi.org/10.1038/s41467-025-60585-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-60585-9
This article is cited by
-
High-fat diet disrupts a septal control on feeding to promote obesity in male mice
Nature Communications (2025)