Abstract
Cellular Indexing of Transcriptomes and Epitopes by Sequencing (CITE-seq) enables paired measurement of surface protein and mRNA expression in single cells using antibodies conjugated to oligonucleotide tags. Due to the high copy number of surface protein molecules, sequencing antibody-derived tags (ADTs) allows for robust protein detection, improving cell-type identification. However, variability in antibody staining leads to batch effects in the ADT expression, obscuring biological variation, reducing interpretability, and obstructing cross-study analyses. Here, we present ADTnorm, a normalization and integration method designed explicitly for ADT abundance. Benchmarking against 14 existing scaling and normalization methods, we show that ADTnorm accurately aligns populations with negative- and positive-expression of surface protein markers across 13 public datasets, effectively removing technical variation across batches and improving cell-type separation. ADTnorm enables efficient integration of public CITE-seq datasets, each with unique experimental designs, paving the way for atlas-level analyses. Beyond normalization, ADTnorm includes built-in utilities to aid in automated threshold-gating as well as assessment of antibody staining quality for titration optimization and antibody panel selection. Applying ADTnorm to an antibody titration study, a published COVID-19 CITE-seq dataset, and a human hematopoietic progenitors study allowed for identifying previously undetected phenotype-associated markers, illustrating a broad utility in biological applications.
Similar content being viewed by others
Introduction
Recent advances in single-cell multimodal profiling, such as Cellular Indexing of Transcriptomes and Epitopes by sequencing (CITE-seq), have enabled the paired profiling of gene expression alongside surface protein expression1,2,3,4. This paired multimodal profiling of single cells has allowed researchers to achieve more precise cell-type annotation (e.g., of immune cells)5,6, study the relationship between transcriptomic state and surface phenotype7,8,9, and readily adapt results to flow cytometry for validation1,4. Given its extraordinary potential, there is increasing application of CITE-seq for atlas construction10,11,12 and in large cohort disease-related studies13,14,15. To effectively leverage the data being generated, there is a pressing need for computational tools for CITE-seq data integration across studies.
Surface proteome profiling by CITE-seq gives rise to specific data characteristics and sources of technical noise inherent to antibody staining. Owing to the high copy number of surface proteins and efficient molecular capture of antibody-derived tags (ADTs), protein expression is considerably less sparse than other single-cell modalities such as mRNA expression or genome-wide chromatin accessibility. Consequently, the protein expression captured by CITE-seq often closely matches the information-rich multi-peak density distributions observed in flow cytometry1 (Supplementary Fig. 1A). Density distributions of protein expression of CITE-seq data frequently exhibit a negative peak, representing background signal arising from non-specifically bound or unbound (free-floating) antibody16, and one or more positive peak(s) representing cells expressing the target protein. Similar to fluorescence-based techniques, the signal-to-noise ratio between the negative- and positive-expression peak(s) is highly sensitive to antibody staining conditions, including antibody concentrations17, staining volumes and time18, and antibody panel composition19. Because of these unique considerations, the normalization and integration approaches devised for other single-cell modalities may not be directly translatable, highlighting the need for methodologies tailored to the intricacies of protein data.
Recent normalization algorithms designed for CITE-seq data, similar to established scRNA-seq approaches20,21, have primarily focused on modeling sequencing bias and ambient expression to remove background signals. Centered log-ratio (CLR) normalization was initially proposed for CITE-seq1 and is also the Seurat default normalization method for the protein component of CITE-seq data, using library size to account for variable sequencing depth and cell size. However, unlike scRNA-seq, which offers relatively unbiased transcriptional profiling, CITE-seq protein panels target only a handful of manually selected proteins, typically between 10 and 300. Therefore, the overall ADT library size is highly sensitive to panel composition, can be easily skewed by high expression of a few subset-specific proteins, and unreliably reflects sequencing depth or cell size. More sophisticated algorithms, including totalVI22, DSB16, and DecontPro23, attempt to model ambient contamination and remove or re-center the background signal to zero. However, these negative-expression peaks in ADT abundance mirror expression distributions by conventional cytometry and are essential for reliable threshold-gating of cells for cell-type annotation24. Improper or incomplete removal of background ADT expression can make it difficult to distinguish between negative-, mid-, and high-expression peaks. For example, CD4 is crucial for T cell lineage identification and is a typical trimodal expression cell surface protein marker. Its positive population corresponds to CD4 T cells, the mid-peak represents monocytes, and the negative population corresponds to non-CD4 T cells, such as CD8 T cells. The negative peak provides essential background information that aids in accurately distinguishing the positive populations. Manipulating the data to eliminate all negative peaks is unnecessary and results in a loss of critical information for cell-type annotation. Consequently, normalization of the negative peak in CITE-seq should emphasize its essential role in cell-type identification rather than its artificial removal. Finally, although other methods have been developed for integration of lower-dimensional embeddings, we focus here on methods that output normalized ADT expression on a per-marker basis, enabling downstream analyses such as threshold-gating of cells for cell-type annotation or differential ADT abundance.
In this work, we introduce ADTnorm, a robust and scalable method tailored to normalize and integrate CITE-seq surface protein data across diverse experimental conditions. By aligning negative and positive expression peaks, ADTnorm effectively mitigates batch artifacts while preserving biologically informative features. We demonstrate that ADTnorm outperforms existing approaches on 13 publicly available CITE-seq datasets, consistently improving cell-type discrimination and enabling large-scale data integration. In addition, ADTnorm provides automated threshold-gating of cell types and a stain quality score to guide antibody titration and panel selection. These features streamline CITE-seq workflows, facilitating interpretable detection of phenotypic markers in complex studies such as those on COVID-19 and human hematopoietic progenitors. Together, ADTnorm expands the potential of multimodal single-cell analyses by delivering an accurate, fast, and user-friendly approach for normalizing and comparing surface protein expression across large and heterogeneous datasets.
Results
ADTnorm leverages a non-parametric strategy to integrate CITE-seq datasets while eliminating batch effects
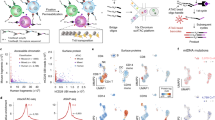
Instead of individually modeling each source of technical noise, we constructed a non-parametric strategy, ADTnorm, building on methods originally conceived for cytometry data25 to remove the batch effects through strategic peak identification and alignment. ADTnorm uses a curve registration algorithm26 to identify protein density landmarks, including the negative and positive peaks, and relies on local minima to detect the valleys separating adjacent peaks. Employing a functional data analysis approach27, ADTnorm normalizes protein expression by aligning the landmarks across datasets (Fig. 1A, Supplementary Fig. 1B and “Methods”), effectively simulating a scenario where all data are derived from the same experiment with equivalent background and antibody staining quality. ADTnorm is implemented as an R package (https://github.com/yezhengSTAT/ADTnorm)28 with an interactive graphical user interface to simplify landmark adjustments (Supplementary Fig. 2) and a Python wrapper (https://github.com/donnafarberlab/ADTnormPy) available to facilitate ADTnorm’s integration into existing CITE-seq analysis workflows (Supplementary Note).
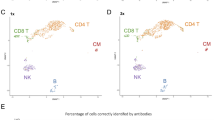
A. ADTnorm normalization procedure starts with the identification of landmarks and aligns peaks and valleys across batches through functional data analysis. The detected peaks and valleys can also be used for automatic threshold-gating (auto-gating), and antibody staining quality can guide the selection of antibodies and staining concentrations. B. Comparison of ADT expression distribution across studies of three T cell lineage markers (CD3, CD4 and CD8) after transformation by Arcsinh and CLR, or normalization by ADTnorm. UMAP embeddings colored by study or cell type were generated after the normalization of 9 ADT markers shared across all 13 studies. Cell type annotations were defined by manual threshold-gating by two immunologists on each sample separately, independent of the normalization work in this paper (“Methods”). The corresponding manual gating strategy is summarized in Supplementary Fig. 3 and Supplementary Table 2. C. Study-level batch correction and broad-level cell type separation quantified by Silhouette score and Adjusted Rand Index (ARI) across methods. ADTnorm was applied using default parameters or customized landmark alignment adjustments. Gray arrows indicate the direction of improved integration performance, i.e., minimized batch effect and maximized cell type separation. The measure center is the Silhouette or ARI metrics calculated on all the cells. The vertical and horizontal error bars represent each method’s standard deviations of 20 bootstrap samples. D. Violin plots displaying CD19 expression in the 10X_malt_10k dataset following normalization under the severe imbalanced setting (“Methods”). Abnormal artifacts introduced to specific cell types during the normalization are highlighted by red squares. The sample size for the box plots is the total cell number within each cell type. E, F. Average auto-gating accuracy (E) and stain quality score (F) across cell types (x-axis) and studies (colors). The sample size for the box plots is 13, representing 13 public datasets. Box boundaries of (D−F) represent the 25th and 75th percentiles. The center line indicates the median. Whiskers extend to the largest and smallest points within 1.5x the interquartile range from the 25th or 75th percentile. Data points beyond the whiskers are outliers. (B, E) and (F) share the same color legend for studies.
Benchmarking ADTnorm with 14 other existing methods on 13 public CITE-seq datasets
Leveraging 13 public CITE-seq datasets of immune cells (Supplementary Table 1)29, we benchmarked the integration performance of ADTnorm against 14 methods from three broad groups: (1) scaling methods commonly applied to cytometry and single-cell data, including Arcsinh transformation, CLR1, log-transformation of count per million (logCPM), and a hybrid approach combining Arcsinh and CLR transformations (Arcsinh + CLR); (2) popular single-cell batch effect removal tools, including Harmony30 implemented on the raw counts, Arcsinh-transformed, CLR-transformed or logCPM-transformed data, fastMNN31, and CytofRUV32; and (3) methods tailored to CITE-seq normalization, including DSB16, decontPro23, totalVI22, and sciPENN8. We obtained a ground-truth manual annotation of these CITE-seq datasets by independently annotating each sample using flow-cytometry gating principles (“Methods”, Supplementary Fig. 3 and Supplementary Table 2). Although the frequencies of these subsets varied, we identified similar immune cell subsets across all samples of the 13 public datasets (Supplementary Fig. 4A). Across the 13 datasets, ADTnorm effectively reduced batch variability, such that negative and positive populations for each surface protein marker could be consistently identified across studies (Fig. 1B, Supplementary Fig. 5, protein density distributions in Supplementary Note). UMAP embeddings of the normalized ADT expression revealed effective batch integration by ADTnorm while preserving cell type separation at both broad and refined annotation levels, treating either the study-level or individual samples as batches (Supplementary Figs. 5, 6). ADTnorm was applied using default parameter settings in the ADTnorm R function for landmark detection (default) or leveraged manually adjusted landmark locations (customized) using interactive R Shiny GUI (“Methods” - Default and customized ADTnorm normalization settings). Both ADTnorm settings, default and customized, outperformed other tools in balancing cell-type separation with cross-study batch effect removal as quantified by Silhouette scores, Adjust Rand Index (ARI), and the Local Inverse Simposon’s Index (LISI) (“Methods”, Fig. 1C and Supplementary Fig. 7A–C). Furthermore, ADTnorm can facilitate the seamless integration of new datasets without reprocessing existing ones by aligning landmarks to predetermined locations (Supplementary Note). It can also incorporate users’ prior knowledge about a batch’s cell type composition. For example, because the Buus 2021 T cell dataset is composed of only T cells, ADTnorm is adjusted to align the singular peak in CD3 as positive-expression (Fig. 1B and Supplementary Note). ADTnorm is also highly scalable, with a fast processing speed and low memory consumption compared to other methods (Supplementary Fig. 7D, E). Also, ADTnorm is designed to process protein markers independently, allowing adaption to parallel processing.
We next explored the downstream impact of protein normalization on joint embeddings of RNA and protein data. Following batch-correction of ADT expression by the above methods and batch-correction of the RNA expression using reciprocal PCA10, we computed the multimodal embedding using the weighted nearest neighbor (WNN) algorithm10 (Supplementary Fig. 8A and “Methods”). As totalVI and sciPENN already incorporate gene expression into their protein normalization process, we omitted them from the WNN integration comparison. As expected, methods with sub-optimal removal of ADT batch effects resulted in skewed WNN integration (Supplementary Fig. 9). ADTnorm markedly minimized batch influences and achieved superior accuracy in segregating cell types as quantified by ARI (Supplementary Fig. 8B), underscoring its utility in post-normalization multimodal integration.
Impact of imbalanced cell type composition on the normalized counts
As surface protein expression varies across cell types, batch correction may be sensitive to variable cell subset composition between batches. In the benchmarking analysis, Harmony exhibited a robust batch correction performance but did not achieve equally strong cell-type separation. This aligns with prior evaluations of scRNA-seq batch correction33 and prompted further investigation into potential over-corrections or artifact biases in the normalized counts across different methods. To evaluate the resilience of normalization methods, we subsampled specific cell subsets from a few batches, devising three scenarios featuring increasingly skewed cell-type compositions (“Methods”; Supplementary Fig. 4B). Careful examination revealed that Harmony, fastMNN, and CytofRUV were highly sensitive to compositional differences, producing unexpected and inaccurate results. For example, CD19 is a highly specific B cell-lineage marker. However, in some batches, Harmony- and fastMNN-normalized CD19 expression was significantly higher in CD4 T cells than in CD8 T cells, and CytofRUV-normalized CD19 expression in CD8 T cells was comparable to that in B cells, patterns not supported by biological expectations (Fig. 1D and Supplementary Fig. 10). Similar discrepancies were noted with DSB, totalVI, and sciPENN across other vital lineage markers (Supplementary Figs. 11, 12). ADTnorm distinguishes itself by meticulously preserving the ranking of protein expression across cells within each batch, thereby reducing the risk of biologically irrelevant anomalies.
ADTnorm provides automatic cell type gatings for annotation
Beyond its primary role in batch correction, ADTnorm leverages intermediate landmark detection results to perform automated threshold-gating (auto-gating) for cell type annotation and to assess staining quality to aid in the optimization of CITE-seq experiments (“Methods”). Valley landmarks identified during ADTnorm normalization can be used to perform automated cell type annotation using predefined gating rules (Supplementary Table 2; Supplementary Fig. 13A–C). While ADTnorm auto-gating showcased high accuracy for a majority of the studies, achieving between 80–100% for comprehensive and nuanced cell type distinctions, auto-gating was underperformed for dendritic cells, memory CD4 T and memory CD8 T cells for some samples in the Hao 2020, Kotliarov 2020, and Witkowski 2020 datasets (Fig. 1E).
Auto-gating accuracy is likely influenced by the marker staining quality that leads to different separation power between negative and positive populations. Hence, we introduced a stain quality score, inspired by fluorescent stain index34, to detect protein markers with poor signal-to-noise separation (see “Methods” - Stain quality score for the full definition). CITE-seq is highly sensitive to antibody quality and concentration, requiring most laboratories to conduct multiple pilot studies for optimizing the antibody concentration cocktail to ensure reliable detection of positive peaks of the target proteins. The stain quality score is designed to assist researchers in refining antibody panels and concentrations during these pilot studies. The stain quality score integrates staining information from three key aspects: (1) peak number, (2) the degree of separation between positive and negative peaks, and (3) the concentration of the positive population. Specifically, a high stain quality score inherently favors markers with more peaks, a longer distance between negative and positive peak mode, and a sharply concentrated positive peak, as these markers demonstrate better antibody staining capacity (Supplementary Fig. 1C). Low-quality scores suggest suboptimal staining conditions, which require careful evaluation or potential exclusion from downstream analyses. For the same protein marker with a consistent peak number across samples, the stain quality score is an indicator of the separation power between negative and positive populations. Leveraging ADTnorm to assess staining quality of each marker to explain the variation in auto-gating accuracy, it revealed that CD56 and CD45RA, which are markers used for gating dendritic and memory T cells, featured less distinct peak separation in batches with poor auto-gating performance (Fig. 1F and Supplementary Fig. 13D; see “Methods” - Stain quality score for further case discussions).
ADTnorm provides a stain quality score to assist in optimizing antibody titration
Beyond the benchmarking analysis of batch correction and cell type separation (Fig. 1), we expanded ADTnorm’s utility to address the critical challenge regarding the antibody titration in the single-cell proteomics. To effectively stain for surface protein, antibody concentrations must be carefully tuned for each sample type. Sufficient antibodies are essential for positive-expression signal(s) to overcome background, but an overabundance of antibodies can obscure rare or low-expression markers by increasing background noise and can increase experimental costs. Although downstream analysis can often tolerate suboptimal staining conditions, variable staining quality is a major source of batch artifacts across samples and laboratories. To explore whether our stain quality score is sensitive enough for titration optimization35 and to evaluate ADTnorm’s ability to mitigate these batch effects, we utilized a titration CITE-seq study that analyzed 124 antibodies on human peripheral blood mononuclear cells (PBMCs)17. This study categorized antibody titration into four levels, including the manufacturer’s recommended concentration (1x) and adjustments to 1/25x, 1/5x, and double (2x) the recommended concentration. As anticipated, the higher concentrations (1x and 2x) typically yielded more distinct separation between negative and positive cell populations, whereas lower concentrations led to greater overlap between negative and positive populations or failed to identify any positive population (Fig. 2A and Supplementary Note). These trends were reflected in the stain quality scores, where markers with reduced separation at low antibody concentrations exhibited lower scores (Fig. 2B). Notably, conventional scaling transformations, Arcsinh and CLR, were unable to successfully integrate expression across titration batches (Supplementary Fig. 14 and Supplementary Note), but ADTnorm could effectively align negative and positive populations across concentrations, thus rescuing cell type discrimination for many protein markers profiled using suboptimal staining conditions (Supplementary Fig. 15) and minimizing batch effects (Fig. 2A). For markers at low titrations that exhibited no positive population, ADTnorm could only align the negative populations (Supplementary Fig. 16A). In these cases, excessively low stain quality scores could alert researchers of protein markers that consistently show poor discrimination between the positive population and the background, suggesting a potential need for revising antibody titration or antibody type (Fig. 2B and Supplementary Fig. 16B). For instance, the 0.04x panel of Fig. 2B indicates that CD36 can be efficiently profiled even after a 25-fold dilution of the antibody. For such markers, researchers can reduce antibody usage and sequencing costs by employing only 0.04x or 0.2x of the original titration. Conversely, some antibodies, such as CD26, CD127 and CD305, exhibit inadequate staining even at twice the commercially recommended concentration. In this case, a much higher concentration may be required, or alternative antibody clones should be tested. Notably, some markers show no detectable differences in staining quality across antibody titrations (right-most panel of Fig. 2B). If none of the tested concentrations effectively profile the positive population, the staining quality scores will consistently remain low, which suggests either that the target antigen is absent or that the antibody has poor binding specificity. For optimal CITE-seq staining, these antibodies are recommended to be reconfigured or excluded from the panel. We also assessed the influence of antibody titration on ADTnorm’s auto-gating accuracy, finding that auto-gating accuracy remains stable as long as lineage markers exhibited detectable positive staining (Supplementary Fig. 17).
A. ADT expression distributions of three T cell lineage markers (CD3, CD4 and CD8) across samples stained at 1/25, 1/5, 1 and 2 times the commercially recommended antibody concentration, following transformation by Arcsinh and CLR or normalization by ADTnorm. UMAP displays the batch correction across the four antibody concentrations and cell-type separation using 124 ADT markers from the original study17. The dashed squares highlight the CD4 T cell subtype clusters that were split due to titration variations and merged after batch correction by ADTnorm. B. The stain quality score evaluates antibody staining quality for target proteins, assessing their ability to successfully profile the positive population and achieve strong separation between positive and negative populations (see “Methods”). Four antibody concentration levels—1/25, 1/5, 1, and 2 times the commercially recommended concentration—are tested and represented by different colors. Recommended concentrations are indicated with solid-colored filling, while non-recommended concentrations are shown with transparent color filling. Protein markers are grouped based on their recommended antibody concentration within each panel and labeled accordingly. C. Data integration across three research institutes where CITE-seq was generated. UMAP shows the batch correction across three research institutes and cell type separation compared across Arcsinh, CLR, DSB and ADTnorm. DSB is the normalization method used in the original paper13. UMAPs were constructed on 192 ADT markers colored by research institute or cell type. D. Volcano plots displaying results of differential proportion of the positive cells for each protein marker between healthy donors and COVID-19 patients. The differential detection analysis was done for CD14+ Monocytes, CD16+ Monocytes and CD83+ CD14+ Monocytes, respectively. Cell type labels are from the original publication13 of the COVID-19 data. E. Dot plot displays consistently differentially expressed protein markers, i.e., CD38, CD64 and CD169, across three monocyte subsets. Points are colored by the average normalized ADT expression and the dot size is relative to the proportion of cells with positive-expression in healthy donors or COVID-19 patients.
ADTnorm integration across multi-center CITE-seq data enhances insights into COVID-19 study
We next investigated whether ADTnorm could facilitate the analysis of consortium efforts. Three UK medical centers profiled 192 protein markers using CITE-seq to study COVID-19 immune response across a diverse cohort of over 100 healthy donors and COVID-19 patients13. Staining quality was highly variable across the participating medical centers (Fig. 2C). Specifically, samples from Newcastle (Ncl) exhibited lower stain quality scores, characterized by a reduced separation between negative and positive peaks, whereas samples from Cambridge and Sanger displayed robust separations, higher stain quality scores, and greater frequencies of detectable positive signals (Supplementary Figs. 18, 19). These batch effects could not be effectively mitigated by Arcsinh and CLR scaling methods (Fig. 2C, Supplementary Fig. 20 and Supplementary Note). ADTnorm effectively reduced technical artifacts (Fig. 2C, ARI: 0.04), resulting in improved cell type separation (ARI: 0.312), both at the broad and refined annotation levels. Additionally, ADTnorm enhanced the joint RNA and ADT embedding compared to the RNA-only embedding reported in the original publication (Supplementary Fig. 21).
Leveraging ADTnorm’s integration and auto-gating, we next aimed to identify whether the expression of specific surface markers could be associated with COVID-19 disease (Supplementary Fig. 19E). Previous studies have identified compositional changes in the immune compartment associated with disease, including increases in the frequency of specific monocyte subsets in the PBMC compartment of mild, moderate, and severe COVID-19 patients (as noted in Fig. 1C of the original publication13). Other studies have identified biomarkers on blood monocytes associated with COVID-19 and type-I interferon signaling, including CD3836,37, CD6438,39 and CD16940,41. We sought to identify whether these trends could be attributed to changing subset frequencies within the monocyte compartment or to the upregulation of these markers across multiple subsets of monocytes. We analyzed the percent-positivity of these and other markers on CD14+, CD16+, and CD83+CD14+ monocytes, and observed upregulation of these markers among COVID-19 patients compared to healthy donors across multiple monocytes states (Fig. 2D, E and Supplementary Fig. 22A). Such upregulation mirrors the trends observed in scRNA-seq (Supplementary Fig. 22B). The normalization employed in the original publication, DSB, did not accurately represent these trends, masking positive expression of CD169 (Supplementary Fig. 23), failing to identify upregulation of CD169 with COVID-19 in any monocyte subset, and reducing signal of CD38 and CD64 in CD16 monocytes (Supplementary Fig. 22C). This demonstrates the utility of ADTnorm in isolating biologically relevant changes and uncovering previously concealed insights in surface protein expression.
Application of ADTnorm in the pilot exploration and final titrated analysis of human hematopoietic progenitor study
We applied ADTnorm to a recent human hematopoietic progenitor study12 to demonstrate its utility in protein data processing and its contribution to downstream biological discoveries. In the pilot study phase, 266 protein markers were examined to guide the selection of a final protein panel and optimize antibody titration. ADTnorm evaluated the stain quality of each protein marker across three cell isolation populations in the original publication: CD34high, CD34+CD271+, and bone marrow mononuclear cells (BMNC) (Fig. 3A). Among the protein markers with detectable positive peaks, stain quality generally was improved with increasing antibody concentration in the CD34high population (Fig. 3B). By leveraging stain quality scores from the titration pilot study, ADTnorm identified 82 protein markers with evidence of successful staining in at least two samples. Of these, 70 overlapped with the ultimate selection of 132 protein markers in the original study, which was determined using a semi-supervised classification model and expert domain knowledge (Fig. 3C). Density distributions of the remaining 12 protein markers indicated successful staining and suggested their potential inclusion in the final protein panel (Fig. 3D and Supplementary Fig. 24). Notably, the 70 protein markers recommended by ADTnorm generally exhibited higher stain quality compared to the remaining 62 markers in the final titration panel (Fig. 3E). Among the 62 markers, 24 markers demonstrated good stain quality scores in at least one donor sample (i.e., stain quality score > 2, Supplementary Fig. 25A).
A. Hematopoietic pilot study to determine the optimal antibody concentration in the titration cocktail. ADTnorm assigns stain quality scores to each protein marker per sample, allowing the identification of markers that exhibit successful staining and positive population signals. B. Among protein markers with detectable positive populations, stain qualities vary with antibody concentration, depending on cell sorting groups. The sample size for each box plot is the total number of protein markers with a staining quality score larger than two. C. Using stain quality scores, ADTnorm identified 82 protein markers with strong evidence of successful staining, 70 of which overlapped with the 132-plex protein panel in the original study. D. Density distribution of protein markers recommended by ADTnorm but not included in the original publication. The remaining 11 protein markers are presented in Supplementary Fig. 24. E. Comparison of stain quality scores for the 132-plex proteins panel in the original study. The sample size for the box plots is 70 for ADTnorm Recommended and 62 for Others in Original Paper. Box boundaries of (B) and (E) represent the 25th and 75th percentiles. The center line indicates the median. Whiskers extend to the largest and smallest points within 1.5x the interquartile range from the 25th or 75th percentile. Data points beyond the whiskers are outliers. F. UMAP visualization comparing protein data processed with Arcsinh transformation and ADTnorm normalization regarding batch correction and cell type separation. G. Heatmap depicting the percentage of positive populations across the hematopoietic cell type trajectory. The complete heatmap for all 132 proteins is in Supplementary Fig. 25B. H. Heatmap showing log2-fold-change of totalVI-normalized counts relative to the mean value per protein marker. G, H use the same hematopoietic cell type order as in the original study. The protein markers are ordered by hierarchical clustering based on ADTnorm positive population percentages. The complete heatmap ordered by hierarchical clustering of the totalVI is provided in Supplementary Fig. 25D. Red squares highlight the differences between the protein marker positive proportion obtained from ADTnorm and log2-fold-changes of the totalVI normalized counts used in the original study.
Using the full 132-plex protein panel in the original study, ADTnorm effectively mitigated batch effects across donor samples, achieving improved cell type separation (Fig. 3F and Supplementary Fig. 25B). Furthermore, the percentage of positive population obtained through ADTnorm provided a more interpretable metric for distinguishing the hematopoietic progenitor cell states and developmental trajectories(Fig. 3G, H), whereas log2-fold-change of normalized values (i.e., totalVI in the original study) remain invariant or zero across multiple cell types for nearly half of the proteins in the final titration panel (Supplementary Fig. 25C–E). Specifically, the log2-fold-changes of CD47 and CD52 across all the cell types were close to zero. However, ADTnorm revealed a high proportion of CD47 expression across hematopoietic and stromal cells, consistent with its role in protecting against macrophage-mediated phagocytosis42,43,44. Similarly, ADTnorm detected stable expression of CD52, a glycosylphosphatidylinositol-anchored protein with immunomodulatory function45, across various subsets of T cells. This mechanism allows CD52high T cells to suppress other T cells to maintain immune homeostasis, protecting against autoimmunity. The low CD52 signal indicated by totalVI could result in a misleading conclusion that this marker was not expressed on T cells. Moreover, log2-fold-change with respect to the mean across all cell types may indicate modest variations, but percentage-positivity reveals the key protein markers with widespread abundance. For instance, CD11a, a heterodimeric integrin critical for transendothelial migration, is highly expressed across many leukocytes46,47. CD162 with a high or moderate abundance on T cells, monocytes, and other leukocytes ensures the physiologic trafficking and homeostatic immune regulation48,49. Both markers’ signals were suppressed by the overall high mean in log2-fold-change quantification. In addition, among erythroblast subsets, over 95% of cells were positive for CD71, a transferrin receptor essential for iron uptake in hemoglobin synthesis, yet the log2-fold-change of totalVI-normalized counts was low, approximately 0.5. CD71 is an important marker, allowing for the identification of cells at various stages in erythropoiesis50.
Beyond improving cell state characterization, percent positivity analysis can also aid in assessing potentially misannotated populations. In the original study, a mixture of unsupervised and supervised methods was used to annotate T cell subsets. Taking the CD4-activated T cell population as an example, the totalVI log2-fold-change for CD8 was 1.8, suggesting a possible misclassification. ADTnorm revealed that 30% of these cells expressed CD8, quantifying this contamination. Additionally, percent positivity serves as a powerful metric for distinguishing subpopulations and refining cell type annotation. For instance, approximately 60% of CD8 effector memory T (TEM) cells expressed CD45RA, while 40% expressed CD45RO. This finding suggests that the majority of CD8 TEM cells are, in fact, terminally differentiated effector memory cells re-expressing CD45RA (TEMRA). In contrast, the totalVI log2-fold-change for CD45RA in these cells was only 0.7, failing to capture this critical biological distinction.
Overall, this case study highlights ADTnorm’s ability to evaluate the protein data from the early phase of protein panel design, improve batch correction, refine cell type annotation, and facilitate biological discoveries by providing interpretable and biologically meaningful metrics.
Discussion
In summary, ADTnorm offers a fast, precise, and scalable solution for normalizing protein expression data, effectively minimizing batch artifacts within studies and enabling integration across studies. ADTnorm is designed for high adaptability, allowing for normalization at various batch levels, supporting missing data, and incorporating prior cell type knowledge. By addressing protein batch effects, ADTnorm also improves multimodal aggregation of RNA and protein modalities, enhancing cell type discrimination and improving interpretability. Unlike other normalization methods that may introduce abnormal expression artifacts, ADTnorm maintains the ranked order of cells within batches for each protein marker and delivers stable performance across datasets with uneven cell type compositions. Additionally, its auto-gating feature offers an expedited avenue for cell-type annotation. The integrated stain quality scoring system alerts researchers to suboptimal staining and assesses experiment quality, aiding in the calibration of antibody titration for pilot studies tailored to specific tissue systems. Especially in the integrative analysis of large datasets, the stain quality score enables rapid identification of problematic markers or batches, thereby enhancing reproducibility and reliability in downstream analyses. Among positive-expressing populations, ADTnorm’s landmark registration approach homogenizes variations in enrichment strength across samples. While it is possible that these variations represent biological differences, that interpretation is confounded by many sources of technical noise, including antibody concentrations, staining conditions, and sequencing artifacts. Notably, ADTnorm also preserves information about the proportion of positive-expressing events in each batch, offering valuable insights into disease status, as exemplified in the COVID-19 case study. This feature underscores the potential of ADTnorm to transcend mere normalization, contributing to the identification of disease-associated protein markers. In the hematopoietic progenitor case study, ADTnorm enhances protein data processing by improving stain quality assessment, mitigating batch effects, and refining cell type annotation through interpretable positive population percentages. Compared to traditional log-fold-change quantification, ADTnorm offers biologically meaningful insights that aid in identifying sub-cell types, correcting misannotations, and facilitating discoveries in hematopoietic progenitor cell characterization.
Due to ADTnorm’s high adaptability, we expect its utility may also extend beyond CITE-seq, allowing for the harmonization of protein expression across multiple technologies (e.g., flow cytometry, CyTOF, and CITE-seq together). Its application is also primed for expansion to multimodal assays by leveraging the normalized protein data as a bridge for cross-modality integration, such as scCUT&Tag-pro51, ASAP-seq52 and PHAGE-ATAC53, which profile surface proteins alongside epigenomic or chromatin accessibility features. ADTnorm stands as a pivotal tool in the evolving landscape of genomic research, facilitating comprehensive analyses across a broad spectrum of biological conditions and technological platforms.
Methods
Data source and processing
Public CITE-seq datasets were downloaded through URLs summarized in Supplementary Table 128. Datasets are identified by the first author’s last name or by “10X” for data obtained from the 10X Genomics websites. Empty droplets, cell aggregates, and apoptotic cells were removed from each dataset based on total UMI counts and the percentage of mitochondrial gene expression using the PerCellQCMetrics and isOutlier functions using default parameter values from the scuttle R package54. ADTnorm was then applied to the raw CITE-seq protein expression data after quality checks and cell filtering.
Cell type annotations used as the ground truth for benchmarking were determined independently by two immunologists through manual threshold-gating on each sample, separate from the normalization work in this manuscript. Manual threshold-gating in single-cell proteomics profiling is a standard approach for cell type annotation in cytometry data, where researchers define specific cell populations by visually gating cells on two-dimensional plots of paired protein markers. The protein markers used for manual gating and cell sub-population identification are summarized in Supplementary Table 2 and the gating logic is exemplified in Supplementary Fig. 2. Cells that could not be confidently assigned to a major cell type were labeled as “unknown", reflecting a consensus between both immunologists that no definitive classification could be made. These cells often exhibited high expression of mutually exclusive protein markers that define distinct major cell types (e.g., CD3+CD19+), suggesting that they could be doublets or rare bi-phenotypic lymphocytes. To ensure completeness for benchmarking, these “unknown" cells were retained.
The 13 public datasets were processed using ADTnorm, described below, along with 14 other existing normalization methods. Detailed implementation information for the comparative methods is provided in the Supplementary Table. 3. For consistency, when methods produced both integrated embeddings and normalized expression (e.g., totalVI), we evaluated performance using the normalized expression values.
ADTnorm normalization and integration pipeline
Landmark Detection. Supplementary Fig. 1B summarizes the model workflow and mathematical details. ADTnorm first applies arcsinh transformation to raw ADT counts with a co-factor 5 and a shift-factor 1 (i.e., arcsinh(1 + 1/5 x RawCount)), then identifies landmarks (peaks and valleys) in the density distribution of protein expression. The ADTnorm R function also allows users to specify their own preprocessing transformations, automatically disabling the default arcsinh transformation when alternative inputs are detected. Peaks are defined as local maxima within high-density regions (Supplementary Fig. 1A), and a curve registration algorithm26 is employed to identify all detectable peak locations. Between each adjacent pair of peaks, ADTnorm identifies valleys as local minima. In scenarios where only one peak is detected or in cases involving a shoulder peak (Supplementary Fig. 1C), valley detection depends on the density slope transitioning from the negative peak to the distribution’s right tail or shoulder peak. Peak and valley detection accuracy relies on precise kernel density estimation for each sample, making selecting a practical bandwidth crucial. The search for an appropriate bandwidth begins with a relatively large value. If no or only one peak is detected with this broader bandwidth, the search continues with narrower settings. For markers generally exhibiting multiple peaks, like CD4, an even narrower bandwidth is applied. Users can input prior information into the ADTnorm software to assist in selecting the optimal bandwidth for constructing the ADT density distribution.
CITE-seq ADT counts are discrete, unlike the continuous data from flow cytometry, with negative peaks often close to zero. Although the Arcsinh transformation effectively compresses large ADT counts into a more manageable range similar to log transformation, it remains nearly linear for counts near zero. Therefore, Arcsinh transformation potentially results in artificial peaks at this low range due to the discrete values. To eliminate suspicious negative peaks, ADTnorm merges peaks detected below a certain small threshold (neg_candidate_thres defined by users in ADTnorm function) near zero or applies a larger bandwidth to smooth these areas. Additionally, if the quality control and filtering steps are insufficiently rigorous, leaving empty droplets, a minor enriched peak might appear near zero before the true negative peak. ADTnorm is designed to recognize and disregard such spurious peaks. Conversely, doublets might create false-positive peak landmarks outside the typical range. ADTnorm uses the mean absolute deviation (MAD, mad function in the stats R package with default values) to assess whether a positive peak landmark is an outlier, excluding it from peak alignment procedures.
Landmark Alignment. ADTnorm leverages identified peaks and valleys in ADT density distributions to mitigate technical variations across batches, studies, platforms, and other experimental inconsistencies by aligning these landmarks across samples. This landmark alignment strategy is inspired by methodologies like guassNorm and fdaNorm25, initially developed for flow cytometry data. Specifically, ADTnorm utilizes functional data analysis, employing a warping function27 to perform a one-to-one transformation of ADT expression that uniformly adjusts the ADT density distribution in a monotone fashion. Mathematically, the kernel density estimate for each sample i is represented by a B-spline interpoland xi. The peak(s) and valley(s) detected for each sample serve as landmarks, and the landmark locations are denoted by tij where j = 1, . . , m. m is 2, meaning there is only one peak and one valley, and m is 3, indicating that this sample has two peaks and one valley. To align the peaks and valleys across sample, xi is transformed by a strictly monotone and invertible function hi known as a warping function for sample i, such that hi(Tstart) = Tstart where Tstart is the starting point of the ADT expression value range and hi(Tend) = Tend where Tend is the ending point of the ADT expression value range. Also, hi(t0j) = tij for j = 1, . . . , m, representing the transformation of the density curves xi so that the corresponding landmark j align to a fixed location t0j. By default, t0j is set to the mean value of tij across samples, but users can pre-define the target landmark alignment locations (target_landmark_location parameter in ADTnorm function). To obtain the optimal estimation of hi, the target function is set to minimizing ∫∣∣y(t) − xh(t)∣∣2dt + λ∫ω2(t)dt where y is a fixed function in the same class as xi and ω(t) measures the relative curvature of h. This penalty on the relative curvature ensures that the transformation function is both smooth and monotone.
Note that ADTnorm also allows users to provide prior information regarding the cell type constitution to more properly align positive peaks across samples. For instance, in batches exclusively involving T cells (e.g., buus_2021_T), a single positive peak for CD3 protein markers is expected. By providing a list of such batches and markers where a uni-peak is expected as a positive peak, ADTnorm can precisely align the detected unique peak (e.g., the gray uni-peak of CD3 in buus_2021_T dataset of Fig. 1B) to the positive peaks in other samples, ensuring consistent and accurate peak alignment (Supplementary Note - Part 2 Protein Density Distribution). This functionality underscores ADTnorm’s adaptability and effectiveness in handling various experimental conditions and study designs. ADTnorm can be applied to integrate batch effects across studies (Supplementary Fig. 5) or batch effects between individual samples within studies, e.g., each donor is a batch (Supplementary Fig. 6 and Supplementary Note). Furthermore, by ignoring missing values, ADTnorm can be used to integrate ADT expression for markers profiled in some but not all batches, a capability not shared by all normalization methods (Supplementary Note).
Default and customized ADTnorm normalization settings
In the benchmark analysis with 14 existing methods, ADTnorm normalized the 13 public datasets using default parameter settings (default) or GUI-assisted, manually adjusted landmark detection (customized). The default setting (i.e., ADTnorm(default) in Fig. 1C) used the default parameter values of the ADTnorm function in the R package (https://github.com/yezhengSTAT/ADTnorm, Supplementary Note), which can handle general protein expression normalization scenarios. ADTnorm R function also offers adjustable parameters to refine landmark detection and provides intermediate density plot visualizations, allowing users to verify the reasonableness of detected peaks and valleys and the alignment of landmarks. A detailed tutorial (Supplementary Note and at https://yezhengstat.github.io/ADTnorm/articles/ADTnorm-tutorial.html) is available to facilitate ADTnorm’s usage, offering guidance on software utilization and parameter adjustment to accommodate different protein expression characteristics.
Additionally, a GUI implemented using the R shiny function (Supplementary Fig. 2) is provided to help users manually fine-tune landmark locations for tailored protein normalization. The customized setting (i.e., ADTnorm(customized) in Fig. 1C) in the benchmark analysis leveraged manual fine-tuning of peak and valley landmarks to ensure optimal landmark alignment. We recommend that users explore the bandwidth sliding bar at the top of the density distribution plot (Supplementary Fig. 2) to determine a proper density bandwidth for setting landmarks. This approach also helps verify whether the weak positive peaks are real or artifacts. At higher bandwidth levels, a true positive population tends to be more visible as one peak, whereas dubious peaks often display a zigzag pattern. Rarer positive populations that do not form visible peaks, even with higher bandwidths, can still be deemed as positive populations if the valley is positioned immediately to the right-hand side of the negative peaks. Similarly, when the negative population exhibits a zigzag pattern, increasing the bandwidth helps smooth out the discrete negative peak, facilitating the identification of an appropriate negative peak landmark.
Weighted nearest neighbor integration of the RNA and protein
Multimodal embeddings were evaluated to test the ADT integration performance of ADTnorm and existing methods. The RNA components are integrated using the Seurat reciprocal PCA (RPCA) strategy. Specifically, the raw gene expression data are first normalized by log-transformation of count per million (log CPM), and the top 5000 feature genes are selected by the “vst" method. Then, the normalized RNA data are scaled using the top features, followed by principal component analysis (PCA) for each study, respectively. Integration anchors are obtained by the FindIntegrationAnchors function of Seurat using the RPCA reduction method. We confirmed the RNA component integration performance by visualizing in UMAP and color-coded by batch and cell types in Supplementary Fig. 8A. The weighted nearest neighbor (WNN) strategy10 from Seurat is leveraged to further integrate the harmonized RNA and normalized protein components. Specifically, the FindMultiModalNeighbors function from Seurat is used to construct the WNN graph based on the top 30 PCs of the RNA component and the top 15 PCs of the protein component. We use default values for all other parameters in the above-mentioned across-modality integration pipeline.
Robustness evaluation on normalization methods by the imbalanced cell type constitution
To assess the robustness of normalization methods, we leveraged the 13 public datasets to create three imbalance scenarios with varying cell-type compositions.
-
Mild imbalance (default integration and benchmarking setting): In the standard setting used to illustrate the ADTnorm model and performance (Fig. 1A–C), one dataset out of 13 public datasets, i.e., buus_2021_T, contains only a single sample of 666 T cells. The remaining 12 datasets profile total PBMCs. This configuration introduces a mild imbalance for the data integration.
-
Moderate imbalance: To create a more challenging integration task with moderately imbalanced subset compositions across batches, we retained only T cells in the hao_2020 (24 samples) and triana_2021 (9 samples) studies, in addition to buus_2021_T. The remaining 10 public datasets were left intact, preserving their original PBMCs profiling.
-
Severe imbalance: For the most extreme scenario, we further filtered the triana_2021 study to include only CD8 T cells, while retaining all T cells from hao_2020 and buus_2021 studies. The rest of 10 public datasets were left unfiltered.
To access normalization performance, we evaluated the normalized expression of CD19 and CD4 across major cell types in the 10X_pbmc_10k and 10X_malt_10k datasets as examples. Since both datasets contain only one sample per study, they are free of within-study batch effect. The original cell type compositions were preserved for 10X_pbmc_10k and 10X_malt_10k.
Stain quality score
To determine the optimal concentration of antibodies to stain specific protein markers35, we proposed a stain quality score designed for ADT data. The stain quality score is inspired by the stain index widely used to optimize the quality and effectiveness of fluorescent staining of cells in flow cytometry55. The stain index is defined as the ratio of the separation between the positive and negative peaks divided by two times the standard deviation of the negative population.
To extend the stain index to capture separation in more diverse data distribution patterns beyond bimodal expression, such as multiple peaks, shoulder peaks or heavy right tail (Supplementary Fig. 1C), we designed the stain quality score as follows:
AUC(PosPeak) means the area under the curve of the positive peak in the corresponding density distribution. Therefore, the stain quality for protein markers with two peaks is positively correlated with the peak mode distance, the sharpness of the positive peak and the proportion of the positive population, and negatively correlated with the total standard deviation in the negative and positive populations.
For protein markers with three or more peaks, the stain quality score is positively correlated with the landmark distance between the right-most peak and the negative peak, the sharpness of the right-most peak and the proportion of non-negative populations. The score is negatively correlated with the sum of the standard deviation of each peak.
Due to the missing positive peak, for markers with one detected peak, we use the distance between peak and valley as the lower bound of the distance between any positive population and the negative peak mode. We continue to penalize the score for one peak by setting the PosPeakHeight to 0. The area under the curve of the right tail beyond the valley is used to distinguish markers that only have a negative population and markers with a heavy right tail or even a shoulder peak. In other words, although the independent positive peak failed to be detected, the positive population is still present.
Stain quality scores are comparable across markers with different peak numbers and generally give higher scores to markers with more peaks. For markers with the same number of identified peaks, better separation of positive and negative populations (longer distance between peak modes) and sharper peaks (lower standard deviation) leads to higher stain quality scores. Markers with two identified peaks score higher than those exhibiting only a shoulder peak. Distributions with only one identified peak and a heavy right tail will have a lower score, and distributions with only one peak and no right tail will be given the lowest score. Supplementary Fig. 1C provided a diagram illustrating the peak patterns and associated stain quality score order.
Stain quality scores can also provide insights into auto-gating performance. Auto-gating accuracy is primarily influenced by the ease of distinguishing between positive and negative peaks. For example, in the case of CD45RA, high-quality antibodies enable successful profiling of the positive population, ensuring that peak number and positive population concentration remain consistent across samples. This allows the stain quality score to effectively reflect the separation power between negative and positive populations. Conversely, a low stain quality score for CD45RA suggests a significant overlap between negative and positive populations, making it difficult to confidently separate the positive peak from the negative peak and reducing auto-gating accuracy, particularly in memory-related cell types (Fig. 1E). However, when comparing protein markers, the stain quality score can be influenced by both peak number and peak separation power. The score inherently favors markers with multiple peaks, as these tend to demonstrate better antibody staining capacity (Supplementary Fig. 1C). For instance, CD45RA typically exhibits 2-3 peaks, whereas CD25 is a single-peak marker with a heavy right tail (Supplementary Note - Part 2, Protein Density Distribution). As a result, CD25 generally receives a lower stain quality score, as its positive population is not as enriched as CD45RA’s to form distinct positive peaks. Beyond antibody staining ability, the absence of a detectable positive peak for CD25 may also stem from the lower proportion of Treg cells compared to memory-like immune cells. Thus, while CD25 may have a lower stain quality score overall, this does not necessarily indicate poor separation between negative and positive populations but rather reflects differences in marker biology and population abundance. CD25 can still effectively distinguish between the negative peak and the heavy right tail (i.e., positive population), allowing for the highly accurate identification of the Treg population (Fig. 1E, F). In summary, stain quality scores provide a valuable metric for understanding auto-gating performance across samples for the same markers.
Computational environment for evaluating runtime and memory
Software performance assessments (Supplementary Fig. 7D, E) were conducted on a dedicated server at Fred Hutchinson Cancer Center in terms of running time and memory consumption. The server was equipped with an Intel(R) Xeon(R) Gold 6254 CPU @3.10GHz, featuring 18 cores, 36 threads, and 754GB RAM. For GPU-accelerated tasks, an NVIDIA-SMI GPU with 12GB of VRAM was utilized. The computational environment was hosted on Ubuntu 18.04.6 LTS, with kernel version 4.15.0-213-generic. The software was compiled and run using GCC version 8.3.0 and CUDA toolkit 12.2. Evaluations were performed under minimal system load to ensure consistent and reproducible results.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw data used in the paper were downloaded from multiple sources, depending on the original studies. Data generated in the studies used in this paper have been deposited in multiple databases summarized in Supplementary Table 1. The corresponding organized public data for the 13 public studies were uploaded as demo data to be part of the ADTnorm software repository (https://github.com/yezhengSTAT/ADTnorm/tree/main/data). The raw and processed 13 public datasets, COVID-19 study and the human hematopoietic progenitor study are available at Zenodo (https://zenodo.org/records/15477967)29.
Code availability
ADTnorm package is implemented in R and is accompanied by a Python wrapper of the R function. The source codes and detailed instructions for running ADTnorm are publicly available at https://github.com/yezhengSTAT/ADTnormfor the R package and https://github.com/donnafarberlab/ADTnormPyfor the Python wrapper. The software is also available through Zenodo at https://zenodo.org/records/1547805528 and Code Ocean at https://doi.org/10.24433/CO.5673910.v156.
References
Stoeckius, M. et al. Simultaneous epitope and transcriptome measurement in single cells. Nat. Methods 14, 865–868 (2017).
Shahi, P., Kim, S. C., Haliburton, J. R., Gartner, Z. J. & Abate, A. R. Abseq: Ultrahigh-throughput single cell protein profiling with droplet microfluidic barcoding. Sci. Rep. 7, 1–12 (2017).
Peterson, V. M. et al. Multiplexed quantification of proteins and transcripts in single cells. Nat. Biotechnol. 35, 936–939 (2017).
Mimitou, E. P. et al. Multiplexed detection of proteins, transcriptomes, clonotypes and crispr perturbations in single cells. Nat. Methods 16, 409–412 (2019).
Caron, D. P. et al. Multimodal hierarchical classification of cite-seq data delineates immune cell states across lineages and tissues. Cell Rep. Methods 5, 1 (2025).
Wells, S. B. et al. Multimodal profiling reveals tissue-directed signatures of human immune cells altered with age. Nat. Immunol. https://doi.org/10.1101/2024.01.03.573877 (2025).
Zhou, Z., Ye, C., Wang, J. & Zhang, N. R. Surface protein imputation from single cell transcriptomes by deep neural networks. Nat. Commun. 11, 651 (2020).
Lakkis, J. et al. A multi-use deep learning method for cite-seq and single-cell RNA-seq data integration with cell surface protein prediction and imputation. Nat. Mach. Intell. 4, 940–952 (2022).
Zhou, S., Li, Y., Wu, W. & Li, L. scmmt: A multi-use deep learning approach for cell annotation, protein prediction and embedding in single-cell RNA-seq data. Brief. Bioinforma. 25, bbad523 (2024).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Guilliams, M. et al. Spatial proteogenomics reveals distinct and evolutionarily conserved hepatic macrophage niches. Cell 185, 379–396 (2022).
Zhang, X. et al. An immunophenotype-coupled transcriptomic atlas of human hematopoietic progenitors. Nat. Immunol. 25, 703–715 (2024).
Stephenson, E. et al. Single-cell multi-omics analysis of the immune response in covid-19. Nat. Med. 27, 904–916 (2021).
Baysoy, A., Bai, Z., Satija, R. & Fan, R. The technological landscape and applications of single-cell multi-omics. Nat. Rev. Mol. Cell Biol. 24, 695–713 (2023).
Anderson, N. D. et al. Transcriptional signatures associated with persisting cd19 car-t cells in children with leukemia. Nat. Med. 29, 1700–1709 (2023).
Mulè, M. P., Martins, A. J. & Tsang, J. S. Normalizing and denoising protein expression data from droplet-based single cell profiling. Nat. Commun. 13, 1–12 (2022).
Nettersheim, F. S. et al. Titration of 124 antibodies using CITE-seq on human PBMCs. Sci. Rep. 12, 20817 (2022).
Buus, T. B. et al. Improving oligo-conjugated antibody signal in multimodal single-cell analysis. eLife 10, e61973 (2021).
Colpitts, S. J. et al. Strategies for optimizing CITE-seq for human islets and other tissues. Front. Immunol. 14, 1107582 (2023).
Lopez, R., Regier, J., Cole, M. B., Jordan, M. I. & Yosef, N. Deep generative modeling for single-cell transcriptomics. Nat. Methods 15, 1053–1058 (2018).
Yang, S. et al. Decontamination of ambient RNA in single-cell RNA-seq with DecontX. Genome Biol. 21, 1–15 (2020).
Gayoso, A. et al. Joint probabilistic modeling of single-cell multi-omic data with totalvi. Nat. Methods 18, 272–282 (2021).
Yin, Y., Yajima, M. & Campbell, J. D. Characterization and decontamination of background noise in droplet-based single-cell protein expression data with decontpro. Nucleic Acids Res. 52, e4–e4 (2024).
Staats, J., Divekar, A., McCoy, J. P. & Maecker, H. T. Guidelines for gating flow cytometry data for immunological assays.Methods Mol. Biol. 2032, 81–104 (2019).
Hahne, F. et al. Per-channel basis normalization methods for flow cytometry data. Cytom. A. 77, 121–131 (2010).
Ramsay, J. O. & Li, X. Curve registration. J. R. Stat. Soc.: Ser. B 60, 351–363 (1998).
Ramsay, J. O., Wickham, H., Graves, S. & Hooker, G. fda: Functional data analysis. R package https://cran.r-project.org/web/packages/fda/index.html (2014).
Zheng, Y. et al. R package for manuscript ADTnorm: Robust integration of single-cell protein measurement across cite-seq datasets. Zenodo https://doi.org/10.5281/zenodo.15478055 (2025).
Zheng, Y. et al. Processed data for manuscript ADTnorm: Robust integration of single-cell protein measurement across cite-seq datasets. Zenodo https://doi.org/10.5281/zenodo.15477967 (2025).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with harmony. Nat. Methods 16, 1289–1296 (2019).
Haghverdi, L., Lun, A. T., Morgan, M. D. & Marioni, J. C. Batch effects in single-cell rna-sequencing data are corrected by matching mutual nearest neighbors. Nat. Biotechnol. 36, 421–427 (2018).
Trussart, M. et al. Removing unwanted variation with cytofruv to integrate multiple cytof datasets. eLife 9, e59630 (2020).
Chu, S.-K., Zhao, S., Shyr, Y. & Liu, Q. Comprehensive evaluation of noise reduction methods for single-cell rna sequencing data. Brief. Bioinforma. 23, bbab565 (2022).
Siddiqui, S. & Livák, F. Principles of advanced flow cytometry: a practical guide. T-cell development: methods and protocols. 89–114 (Springer, New York, 2022).
Kwon, D. The antibodies don’t work! the race to rid labs of molecules that ruin experiments. Nature 635, 26–28 (2024).
Qin, S. et al. Dynamic changes in monocytes subsets in covid-19 patients. Hum. Immunol. 82, 170–176 (2021).
Horenstein, A. L., Faini, A. C. & Malavasi, F. Cd38 in the age of covid-19: a medical perspective. Physiol. Rev. 101, 1457–1486 (2021).
Karawajczyk, M. et al. High expression of neutrophil and monocyte cd64 with simultaneous lack of upregulation of adhesion receptors cd11b, cd162, cd15, cd65 on neutrophils in severe covid-19. Ther. Adv. Infect. Dis. 8, 20499361211034065 (2021).
Li, Y. et al. Monocyte surface expression of fcγ receptor ri (cd64), a biomarker reflecting type-i interferon levels in systemic lupus erythematosus. Arthritis Res. Ther. 12, 1–12 (2010).
Doehn, J.-M. et al. Cd169/siglec1 is expressed on circulating monocytes in covid-19 and expression levels are associated with disease severity. Infection 49, 757–762 (2021).
Ortillon, M. et al. Monocyte cd169 expression in covid-19 patients upon intensive care unit admission. Cytom. A 99, 466–471 (2021).
Blazar, B. R. et al. Cd47 (integrin-associated protein) engagement of dendritic cell and macrophage counterreceptors is required to prevent the clearance of donor lymphohematopoietic cells. J. Exp. Med. 194, 541–550 (2001).
Matozaki, T., Murata, Y., Okazawa, H. & Ohnishi, H. Functions and molecular mechanisms of the cd47–sirpα signalling pathway. Trends Cell Biol. 19, 72–80 (2009).
Khalaji, A. et al. Don’t eat me/eat me signals as a novel strategy in cancer immunotherapy.Heliyon 9, e20507 (2023).
Bandala-Sanchez, E. et al. T cell regulation mediated by interaction of soluble cd52 with the inhibitory receptor siglec-10. Nat. Immunol. 14, 741–748 (2013).
Wojcikiewicz, E. P. et al. Lfa-1 binding destabilizes the jam-a homophilic interaction during leukocyte transmigration. Biophys. J. 96, 285–293 (2009).
Fagerholm, S. C., Guenther, C., Llort Asens, M., Savinko, T. & Uotila, L. M. Beta2-integrins and interacting proteins in leukocyte trafficking, immune suppression, and immunodeficiency disease. Front. Immunol. 10, 254 (2019).
Hirata, T. et al. P-selectin glycoprotein ligand 1 (psgl-1) is a physiological ligand for e-selectin in mediating t helper 1 lymphocyte migration. J. Exp. Med. 192, 1669–1676 (2000).
Hope, J. L. et al. Psgl-1 attenuates early tcr signaling to suppress cd8+ t cell progenitor differentiation and elicit terminal cd8+ t cell exhaustion.Cell Rep. 42, 112436 (2023).
Zhang, J., Socolovsky, M., Gross, A. W. & Lodish, H. F. Role of ras signaling in erythroid differentiation of mouse fetal liver cells: functional analysis by a flow cytometry–based novel culture system. Blood 102, 3938–3946 (2003).
Zhang, B. et al. Characterizing cellular heterogeneity in chromatin state with sccut&tag-pro. Nat. Biotechnol. 40, 1220–1230 (2022).
Mimitou, E. P. et al. Scalable, multimodal profiling of chromatin accessibility, gene expression and protein levels in single cells. Nat. Biotechnol. 39, 1246–1258 (2021).
Fiskin, E. et al. Single-cell profiling of proteins and chromatin accessibility using phage-atac. Nat. Biotechnol. 40, 374–381 (2022).
McCarthy, D. J., Campbell, K. R., Lun, A. T. & Wills, Q. F. Scater: pre-processing, quality control, normalization and visualization of single-cell rna-seq data in r. Bioinformatics 33, 1179–1186 (2017).
Shapiro, H. M. Practical flow cytometry. (John Wiley & Sons, New Jersey 2005).
Zheng, Y. et al. R scripts for manuscript “ADTnorm: Robust integration of single-cell protein measurement across cite-seq datasets”. Code Ocean https://doi.org/10.24433/CO.5673910.v1 (2025).
Acknowledgments
This work was supported by the National Institutes of Health grant, HG012797, to Y.Z. and Chan Zuckerberg Initiative award, DI-0000000345, to R.G. D.P.C. was supported by the Columbia University Graduate Training Program in Microbiology and Immunology (T32AI106711). P.A.S. was supported by U19AI128949. We also acknowledge the Scientific Computing Infrastructure at Fred Hutchinson Cancer Center, funded by ORIP grant S10OD028685, the J. Orin Edson Foundation, the Translational Data Science Integrated Research Center of the Fred Hutchinson Cancer Center and NIH U19AI128914. We also appreciate the timely and in-depth discussion with Drs. Helen Lindsay, Bernat Bramon Mora and Antonin Thiebaut from the University of Lausanne.
Author information
Authors and Affiliations
Contributions
R.G. and Y.Z. conceived the project. Y.Z., D.P.C. and R.G. designed the research and developed the method. Y.Z., J.Y.K. and D.P.C. developed the software and organized the usage manual and tutorial. Y.Z. and S.H.J. designed the auto-gating strategy. Y.T. and F.M. manually gated the protein data to provide a gold standard for the cell type annotation. R.G., P.A.S. and K.D.S. provided feedback and suggestions as the project progressed. All authors contributed to the preparation of the manuscript.
Corresponding author
Ethics declarations
Competing interests
R.G. has received consulting income from Takeda and Sanofi and discloses ownership in Ozette Technologies. Additionally, R.G. declares research collaborations with Owkin and 10X Genomics. Other authors declare no competing financial interests.
Peer review
Peer review information
Nature Communications thanks Ivan Costa, Sofie Van Gassen, and Xiang Lin for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zheng, Y., Caron, D.P., Kim, J.Y. et al. ADTnorm: robust integration of single-cell protein measurement across CITE-seq datasets. Nat Commun 16, 5852 (2025). https://doi.org/10.1038/s41467-025-61023-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-61023-6