Abstract
Errors in chromosome segregation during gametogenesis, such as nondisjunction (NDJ) errors, have severe consequences in human reproduction, and a better understanding of their etiology is of fundamental interest in genetics. Mapping NDJ errors to meiotic/mitotic stages typically requires proband-parent comparison, limiting its applicability. Herein, we develop Mis-segregation Error Identification through Hidden Markov Models (MeiHMM), a method for inferring NDJ error stage and crossover events based on only genomic data of trisomic probands. Guided by triallelic genotype/haplotype configurations, MeiHMM discerns the allelic origin at each locus, which informs NDJ error during gamete formation, without identifying the parental origin of the trisomy. In 152 Down syndrome (DS) cases, MeiHMM achieved an accuracy of 96.1% in classifying NDJ errors, with a sensitivity of 91.6% in crossover identification, compared to proband-parents trio analysis. 17% of Meiosis II errors were misclassified as Meiosis I, mainly due to small proximal crossover events. Applying MeiHMM to 509 children with DS-associated childhood leukemia, we demonstrate that NDJ error is associated with the age of disease onset, somatic genomic abnormalities, and prognosis. Thus, MeiHMM is an effective method for trisomic NDJ error classification and crossover identification that can be applied broadly to study the etiology of congenital aneuploidy conditions.
Similar content being viewed by others
Introduction
Aneuploidy can occur during human meiosis due to nondisjunction (NDJ) errors, but it is challenging to study because most cannot give rise to viable embryos, with the exception of trisomies 13, 18 and especially 21—the most common aneuploid condition at birth. Therefore, Down syndrome (DS) offers a unique opportunity and is an ideal model for understanding the origin of these events. Individuals with DS are at increased risk of intellectual disability, congenital heart disease, and are predisposed to hematologic malignancies1. Trisomy 21 arises mostly from NDJ or mis-segregation of chromosome 21 (chr 21) during meiosis2,3, or relatively uncommonly, from postzygotic mitosis4. In more than 90% of cases, the extra copy of chr 21 originates from the maternal gamete5. This is because female gametogenesis is more prone than male gametogenesis to erroneous chromosomal segregation due to decades of arrest at prophase I6,7. Inaccurate chromosomal segregation during gametogenesis can be grossly classified into meiosis I (MI) or meiosis II (MII) errors using genotyping data8,9. MI errors are inferred when two homologous parental chromosomes are identified in the gamete and MII errors are inferred when two sister chromatids are identified. These observations can be the result of several mechanisms of mis-segregation, such as NDJ of the homologous chromosomes, precocious separation of sister chromatids, or reverse segregation. The stage at which the NDJ takes place is associated with the number and the position of the chromosomal crossovers. In maternal NDJ of chromosome 21, absence of crossover or crossover distally located from the centromere is associated with MI errors. In contrast, MII errors are associated with pericentromeric crossovers10.
The type/stage of NDJ error of trisomy 21 is usually inferred from proband-parent trios by comparing the heterozygosity patterns of genotypes at the pericentromeric region between the child with DS and the parents10,11,12,13. However, when the parents’ genetic data are unavailable, there is currently no generalizable method to infer the type of error, aside from recent work by refs. 14,15 analyzing NDJs based on low coverage whole genome sequencing (WGS) data in preimplantation genetic testing.
In this work, we develop a method, Mis-segregation Error Identification through Hidden Markov Models (MeiHMM), to infer the type of NDJ and locate the chromosomal crossovers using proband genotype alone, based on the frequencies of single nucleotide polymorphisms (SNPs) or haplotypes in the general population. Applying this to WGS data of 152 DS cases demonstrates high accuracy compared to trio analysis. We further investigate the association between the type of NDJ error and disease characteristics in a cohort of DS-associated acute lymphoblastic leukemia (ALL). MeiHMM does not infer the parental origin of trisomy 21, but it is applicable to NDJ errors of both maternal and paternal origins. For simplicity in the description, the following sections assume a maternal origin for trisomy 21, which is true in ~90% of DS cases5, though the framework is the same for NDJ of paternal origin as well.
Results
MeiHMM implements variant- and haplotype-based analyses to identify NDJ timing and cross over locations
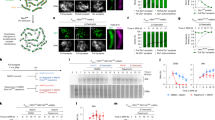
The types of chr 21 NDJ errors can be distinguished by the number of unique haplotypes across the q arm of this chromosome as follows (Fig. 1A): (1) Errors of MI origin are characterized by having three different haplotypes in the centromeric region, i.e., one allele with the paternal haplotype, two remaining alleles with two different maternal haplotypes due to the unsuccessful separation of the homologous chromosomes in the oocyte; (2) By contrast, errors of MII result when the duplicated sister chromatids fail to separate; such failure is inferred when only two haplotypes are represented in the centromeric region, i.e., one allele from the paternal haplotype and two identical alleles from the maternal haplotype; (3) Mitotic errors are inferred when there is complete duplication of maternal chr 21. (Alternatively, this could be inferred as a MII error with no recombination, a noted limitation of this gross classification10) Applying a Hidden Markov model, MeiHMM (1) segments chr 21 into blocks of two- or three haplotypes which informs the stage of NDJ, and (2) identifies the position of chromosome crossover (recombination during meiosis) on the basis of the boundaries of these haplotype blocks.
A The types of chr 21 nondisjunction (NDJ) and their patterns of haplotypes along chr 21. Here, trisomy 21 is assumed to be of maternal origin. B Summary of the workflow to categorize chr 21 heterozygous SNPs. C Type 1 informative SNPs occur predominantly in two-haplotype blocks and type 2 informative SNPs occur predominantly in three-haplotype blocks. D Diagram of the hidden states and the observations of MeiHMM. E Distribution of the allele frequencies (AF) of two-copy alleles of chr 21 SNPs in simulated SNPs by sampling two (and duplicate one allele to make three alleles) or three alleles from gnomAD and AFLA references. F Distribution of the O/E ratio of hypothetical haplotypes. In panels E and F, yellow and blue colors indicate simulated values by assuming there are two or three haplotypes, respectively. G The results of MeiHMM of an example case with Meiosis I error. Red dashed lines indicate thresholds used to define type 1 (allele frequency <0.003) and type 2 (O/E ratio <0.1) informative SNPs. SNP Single-nucleotide polymorphism, ref alelle reference alelle, alt alelle alternative alelle, gnomAD Genome Aggregation Database, AFLA Allele Frequency Aggregator, AF allele frequency, O/E ratio ratio of observed vs expected frequency. Source data are provided as a Source Data file.
To implement MeiHMM, we categorize heterozygous SNPs on chr 21 into three groups (Fig. 1B). (1) Type 1 informative SNPs: for a given case, we first identify variants that are rare in the general population (alternative allele frequency <0.003 in gnomAD and ALlele Frequency Aggregator [ALFA]); if the genotype configuration at this SNP is alternative: alternative: reference alleles (Alt: Alt: Ref), this suggests a two-haplotype state because the probability of a rare allele occurring twice independently is exceedingly low; thus the configuration likely arises from the duplication of the rare allele, i.e., two-haplotype state (Fig. 1C; see “Methods” for details). (2) Type 2 informative SNPs: for variants that do not meet the criteria in #1 (common SNPs or rare variants without the defined genotype configuration), we utilize genotypes at adjacent variants to discern two- vs three-haplotype state: for a given variant and its two flanking SNPs, we generate hypothetical haplotypes for this three-variant block, assuming a two-haplotype state (Fig. 1C), i.e., one haplotype consists of the combination of the one-copy alleles across the three variants, whereas the other is defined by the combination of alleles with two copies. The likelihood of two- vs three-haplotype state for this region is then scored on the basis of the observed vs expected frequency (O/E) of the hypothetical haplotypes in the 1000 Genomes Project populations, with the expected frequency estimated as the product of the allele frequencies of incorporated alleles (see “Methods”). A lower O/E ratio suggests that the two-haplotype state is less likely, which informs the presence of the three-haplotype states. (3) The remainder of chr 21 SNPs are considered non-informative. Variants on chr 21 were classified into one of these three groups and these assignments were fed into a hidden Markov model to segment chr 21 into smoothened two-haplotype and three-haplotype blocks (Fig. 1D). The stage of NDJ error is then inferred from the segmentation results (Supplementary Fig. 1). Detailed definitions of the two types of informative SNPs and the hidden Markov model are available in “Methods”.
We performed simulations to evaluate the degree to which the aforementioned type 1 (rare variant-based) and type 2 (rare haplotype-based) informative SNPs inform the two- vs three-haplotype state on chr 21 (see “Methods” for details). First, for the rare variant-based inference, we generated triallelic genotypes for chr 21 SNPs according to the allele frequencies reported in gnomAD and ALFA, under the assumption of either a two- or three-haplotype state. Among 44,681 variants in gnomAD and ALFA for which (1) the alternative allele population frequency <0.003 and (2) the simulated genotype configuration is Alt: Alt: Ref, 99.7% were only seen under the two-haplotype hypothesis (Fig. 1E). For rare haplotype-based inference, we performed similar simulations but now by randomly sampling haplotypes from the 1000 Genomes project. In 1,653,980 hypothetical haplotypes with O/E ratio <0.1, 98.2% were only seen under the three-haplotype hypothesis (Fig. 1F).
Figure 1G shows an example of the segmentation results of MeiHMM, based on WGS data from Gabriella Miller Kids First Pediatric Research Program (see “Methods”). For this particular case, rare haplotype analysis of the pericentromeric region indicates a three-haplotype state, and therefore it is classified as MI error, with one crossover event identified in the q arm (Fig. 1G). Additional examples are included in Supplementary Fig. 2.
Performance metrics of MeiHMM
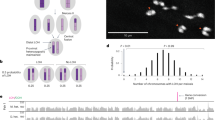
To test the accuracy of MeiHMM, we compared the NDJ error stage and crossover event calling results with those from 152 DS proband-parent trios with WGS data (see “Methods”). Overall, MeiHMM classified 76%, 19%, and 5% of the cases as MI, MII, and mitotic errors (Fig. 2A), respectively, of which 146 were consistent with trio analysis (accuracy = 96.1%; Fig. 2A). Misclassifications of the six cases were due to crossovers (Fig. 2B) near the centromere, resulting in exceedingly narrow two-haplotype blocks (Fig. 2C) in the pericentromeric region with a limited number of informative SNPs; consequently, MII errors were misclassified as MI errors (Fig. 2A).
A Comparison of MeiHMM with trio analysis. B MeiHMM identified 87 of the 95 crossovers at similar positions and crossovers very close to the centromere leads to incorrect classifications. C Comparison of the minimum block size of correctly and incorrectly classified cases. D Frequencies of MI, MII, and mitotic errors identified by MeiHMM in 764 DS cases. E, F Distribution of the crossovers on chr 21 in (E) MI and (F) MII errors. MI meiosis I, MII meiosis II, DS Down syndrome, WGS whole genome sequencing. Source data are provided as a Source Data file.
MeiHMM also accurately identified crossover events (Fig. 2B). In the 95 crossover events identified by trio analysis, 87 were successfully discovered by MeiHMM (sensitivity = 91.6%) with no false positive crossover events (precision = 100%). Again, the discordance is largely due to the pericentromeric crossovers as described above, except for one case with two crossovers too close to each other (Fig. 2B). Two cases analyzed by both MeiHMM and trio are illustrated in Supplementary Fig. S3.
Next, we applied MeiHMM to a large dataset of 764 DS cases with proband-only data genotyped by using WGS, MeiHMM identified 75% (n = 571) of the errors as MI origin (no crossover, 47%; with crossover, 28%), 21% of the errors as MII, and 4% as mitotic errors (Fig. 2D). The frequencies of the error types were consistent with those in the cases by trio analysis (Fig. 2A), as well as those reported in a previous study using microsatellite markers10.
We identified an average of 0.47 crossover events per case in MI errors (range 0–3 per case, total 266 events; Fig. 2E), though this is likely an underestimation of the actual frequency of crossover events during gamete formation due to random selection of sister chromatids in MII followed by MI errors. In cases with MII errors (n = 165), an average of 1.19 crossover events were identified (range 1–3 per case, total 197 events; Fig. 2F). MI and MII errors were characterized by a distinct pattern of crossover position along chr 21. Crossovers in MI errors were more likely to be telomeric, especially in cases with only one crossover (Fig. 2E). In contrast, crossovers in MII errors were preferably pericentromeric, regardless of the number of crossovers, with a relatively heavier tail towards the telomeric region (Fig. 2F). These results were consistent with previous observations that the positioning of crossovers is associated with the type of error10.
Applying MeiHMM to SNP arrays
Because MeiHMM is based on genotype data, it can potentially be used for datasets generated by other genotyping platforms. To demonstrate this, we applied MeiHMM to a previously published SNP arrays dataset of 429 DS case-control series for congenital atrioventricular septal defects (AVSD)16. We elected to only use common variants and perform the haplotype-based analysis (Type 2 in Fig. 1B) because rare SNPs are poorly represented on SNP arrays. We observed MI, MII and mitotic errors in this DS-AVSD cohort with frequencies comparable to our DS-ALL dataset (Fig. 2D and Supplementary Fig. 4).
NDJ error is associated with disease phenotypes of DS-related ALL
The ability to infer chr 21 NDJ error using proband-only data enables the investigation of DS-associated health conditions when genotype data of the parents are not available. To demonstrate this, we applied MeiHMM to a cohort of children with DS-associated acute lymphoblastic leukemia (DS-ALL) (n = 509) genotyped using WGS17. Overall, MI, MII, and mitotic errors accounted for 73% (with crossover, 31%; no crossover, 43%), 22% and 5% of the dataset, respectively, similar to those observed in the general DS population without ALL (n = 764; P = 0.63).
However, there was significant heterogeneity in NDJ error within DS-ALL cases, especially when stratified by the age at ALL onset (P = 0.0026) and by ALL molecular subtype defined by leukemia genomic abnormalities (P = 0.017; Fig. 3A). MII or mitotic errors were over-represented in patients with ALL diagnosed at the age above 10 years (38%) compared to those diagnosed below 10 years (24%; P = 0.0024; Fig. 3A). Across different ALL subtypes, there seemed to be an over-representation of errors of MII origin in patients with ETV6::RUNX1-like, CRLF2 BCR::ABL1-like and C/EBPalt DS-ALL (Fig. 3A). Because MII errors are characterized by more crossover events, we further hypothesize that the specific location of chr 21 recombination (and consequently haplotype diversity of specific loci) may influence DS disease phenotypes. To test this, we compared the frequencies of two- and three-haplotype states in windows of 1 million bp along chr 21 between DS-ALL cases and non-ALL DS cases. Compared to non-ALL DS cases, DS-ALL patients older than 10 years were more likely to have 2 haplotypes (rather than 3 haplotypes) in the proximal q arm, especially at q21.1 (Fig. 3B). In contrast, DS-ALL patients younger than 10 years exhibited similar patterns of haplotype diversity relative to that of the general DS population without leukemia (Fig. 3B). In terms of molecular subtype, CRLF2 BCR::ABL1-like DS-ALL cases exhibited significantly more frequent two-haplotype blocks in the region spanning from q21.3 to q22.2, while C/EBPalt DS-ALL cases are more likely to have two-haplotype blocks at q22.12 when compared to non-ALL DS cases (Fig. 3C).
A Type of chromosome 21 NDJ errors of DS-ALL patients in specific age groups and molecular subtypes. Associations were tested using Chi Squared test. Nominal p-values were reported without adjusting for multiple comparisons. B, C P-values (by Chi Squared test) of the frequency of two-haplotype blocks across chromosome 21 genomic positions in different (B) age groups and (C) molecular subtypes compared to non-ALL DS cases. For (B and C), dashed red line indicate Bonferroni corrected p-value of 0.05. Size of red circles indicate the odds ratio of two-haplotype state comparing DS-ALL and non-ALL DS cases, when adjusted p-value < 0.05. D, E Kaplan–Meier estimates of the probability of (D) event-free survival or (E) overall survival of DS-ALL patients with meiosis I, meiosis II, and mitotic errors. Differences in survival were tested by log-rank test. Source data are provided as a Source Data file.
We also observed that the type of trisomy 21 error was associated with treatment outcomes of DS-ALL in terms of both event-free survival (EFS; P = 0.016; Fig. 3D) and overall survival (OS; P = 0.0024; Fig. 3E). After adjusting for known prognostic factors such as age, initial white blood cell count, minimal residual disease at end-of-induction, and ALL molecular subtype, MII is independently associated with poorer event-free survival (P = 0.035; Supplementary Table 1). Because the ratio of two- vs three-haplotype blocks differ by NDJ error type, we examined the impact of the size of two-haplotype blocks on treatment outcomes. As shown in Supplementary Table 2, the proportion of two-haplotype blocks was linked to decreasing EFS (P = 0.012) and OS (P = 2.6 × 10−4), and it remained marginally significant after adjusting for NDJ error type (P = 0.038 for OS). These results imply that the proportion of chr 21 with two-haplotype state (thus the degree of homozygosity) may mediate the impact of NDJ error on leukemia treatment outcomes. Taken together, these results suggest that the NDJ error of trisomy 21 could modify specific phenotypes of DS-associated health conditions.
Discussion
The type of NDJ error and crossover events are central to understanding the etiology of DS (and trisomic chromosomal disorders in general). It has been found that the type of NDJ (MI, MII, or mitotic) is closely related to maternal age12,18, genetic polymorphisms11, lifestyle19,20, socioeconomic status21,22, and crossover events10. An altered crossover pattern is a strong risk factor for chromosomal NDJ2,8,23; and when NDJ happens, the crossover is associated with the type of error10. Because MeiHMM only requires data from the DS child, it can greatly expand the number of cases for which NDJ can be inferred and thus enable large association studies to explore the impact of NDJ on DS disease phenotypes. However, the exact biology underlying the association of NDJ error stage with DS disease phenotypes is unclear and warrants further investigation. We posit that duplication of the deleterious allele (in the two-haplotype state, MII) can result in increased severity of disease phenotypes, compared to cases with only one-copy of the risk allele (in the three-haplotype state, MI). In DS-ALL, we observe association of two-haplotype state with higher risk of unfavorable disease subtype CRLF2 BCR::ABL1-like, which is in line with this hypothesis. That said, this hypothesis should be experimentally tested in future studies.
We have demonstrated that MeiHMM can be applied to genotype data generated by WGS or SNP arrays. It should also be straightforward for MeiHMM to accommodate data generated from other sequencing or genotyping platforms, such as RNA sequencing, whole exome sequencing, or even a small, targeted assay designed specifically for NDJ error identification. In addition to its versatility in data types, MeiHMM can also be applied to study the NDJ of other trisomic chromosomal conditions (e.g., trisomy 13 or trisomy 18)24, either prenatally or postnatally. One limitation of MeiHMM is that it cannot detect crossover events involving small genomic regions with few informative SNPs. This is especially problematic for calling short two-haplotype blocks where rare variants are sparse. For example, as MII errors are associated with proximal crossovers, which results in short two-haplotype blocks at the pericentromeric region, leading to the misclassification of 17% (n = 6 of 35) of MII errors as MI errors (Fig. 2A). It should be noted that these misclassifications may introduce uncertainty in assessing the association of NDJ error type with DS phenotypes, such as treatment outcomes of ALL. Furthermore, because MeiHMM uses data of the proband-only, it does not infer the parental origin of trisomy 21.
The traditional method for NDJ staging relies on diads or trios, with restrictive utility because current DS genomic datasets are not trio-based. Prior efforts such as LD-PGTA15 and LD-CHASE14 are capable of inferring NDJ errors and crossover events in embryos generated through in vitro fertilization, using data from preimplantation genetic testing for aneuploidy. This elegant approach employs a haplotype-based framework, conceptually related to MeiHMM, even though it relies on sampling read pairs from low-coverage WGS data. In contrast, MeiHMM harnesses genotype information, making it more broadly generalizable across platforms; MeiHMM also makes use of rare variants (Type 1 informative SNP), which may be beneficial for regions with sparse linkage disequilibrium. Applying MeiHMM would greatly expand the ability to determine the origin of NDJ, essential in any DS cases genotyped for Chr 21 SNPs at a reasonable density, with potential value for studying the etiology of other congenital aneuploidies.
Methods
Data and genotyping
This study complies with all relevant ethical regulations and was approved by the St. Jude Children’s Research Hospital Institutional Review Board and Baylor College of Medicine Institutional Review Board. All the WGS data used in this study were generated by the Gabriella Miller Kids First Pediatric Research Program. WGS and SNP array data were obtained for a total of 764 and 429 children with non-mosaic DS, respectively. WGS data of both parents were also obtained for a subset of DS children with WGS data, forming 152 trios. In addition, WGS data of DS-associated acute lymphoblastic leukemia cases (DS-ALL, n = 509) was obtained, and subtype classification was performed in conjunction with RNA-Seq data as described in our previous publication17. Briefly, subtypes including CRLF2, ETV6::RUNX1, IGH::IGF2BP1, TCF3::PBX1, BCR::ABL1, KMT2A, and DUX4 were classified by rearrangement identification (using fusionCatcher version 0.99.7d)25; high hyperdiploid subtype was identified by digital karyotyping (using RNASeqCNV version 1.2.2)26; C/EBPalt subtype was defined by overexpression and/or alterations of CEBPD, CEBPA or CEBPE genes17; BCR::ABL1-like and ETV6::RUNX1-like subtypes were called by gene expression analysis27; PAX5alt subtype was defined by unique gene expression pattern and presence of PAX5 alterations27; and PAX5 P80R and IKZF1 N159Y were called using variant identification27. DS-ALL Patients were enrolled in the following Children’s Oncology Group trials: 9904 (ClinicalTrials.gov identifier NCT00005585), 9905 (NCT00005596), 9906 (NCT00005603), AALL0232 (NCT00075725), AALL0331 (NCT00103285), AALL08B1 (NCT01142427), AALL0932 (NCT01190930), AALL1131 (NCT02883049), AALL1731 (NCT03914625), and APEC14B1 (NCT02402244). All samples were collected with informed consent obtained from patients, parents, and/or guardians, in written and signed form. WGS was performed using Kapa Hyperprep library preparation kit (Roche), sequenced on the NovaSeq platform (Illumina), and coverage of reference and alternative alleles were extracted using the GATK (version 3.5-0) germline short variant discovery pipeline28, based on which triallelic genotype calls were made. SNPs with variant allele frequencies between 0.13 and 0.40 or between 0.60 and 0.87 were deemed to have configuration of Alt: Ref: Ref or Alt: Alt: Ref, respectively. Only SNPs with coverage of at least 30 reads, and the frequencies of alternative allele carriers in the cases were within 10% of the expected proportion (estimated using overall population allele frequency from gnomAD29 and ALFA30) were kept for analysis. SNP array data was retrieved from a previously published dataset16 of SNP 6.0 arrays (Affymetrix). For each SNP on chromosome 21, genotyping was performed by unsupervised hierarchical clustering using average linkage and Euclidean distance based on three features: intensity A, intensity B and the ratio between B and A. Only SNPs with genotype counts broadly following Hardy-Weinberg equilibrium (Chi squared statistics <20) were kept for analysis.
Type 1 informative SNPs: variants with rare two-copy alleles
On chromosome 21, let \(M\) be the number of heterozygous SNPs in a DS proband that is properly genotyped, \({{\mathbf{\Omega }}}={({{{\bf{G}}}}_{i})}_{i=1}^{M}\) denote the genotypes of these SNPs where \({{{\bf{G}}}}_{i}\) is the genotype of the \({i}^{{th}}\) SNP. As there are three copies of chromosome 21, the genotype of a heterozygous SNP always consists of two copies of one allele and a single copy of another allele (hereafter referred to as two-copy allele and one-copy allele, respectively). \({{{\bf{G}}}}_{i}\) can be further denoted as
where \({a}_{i}\) is the two-copy allele and \({b}_{i}\) is the one-copy allele. As an example, for a heterozygous SNP with genotype \({TTG}\), we have \({a}_{i}=T\) and \({b}_{i}=G\). The position of the \({i}^{{th}}\) heterozygous SNP on chromosome 21 is denoted as \({l}_{i}.\) The genotypes \({({{{\bf{G}}}}_{i})}_{i=1}^{M}\) are ordered according to their positions, i.e., \({l}_{i} < {l}_{j},\forall {i} < j\).
Let \({{\rm{AF}}}(l,m,g)\) denotes the allele frequency of a SNP on chromosome 21, at position \(l\), with \(m\in \{{{\rm{eur}}},{{\rm{afr}}},{{\rm{amr}}},{{\rm{eas}}},{{\rm{sas}}}\}\), representing European, African, American, East Asian, and South Asian genetic ancestries, and \(g\in \left\{A,T,C,G\right\}\), representing the allele. We define a variant as having rare two-copy allele status if
where \({\{{m}_{k}\}}_{k}\) are the ancestries that account for more than 25% in the genetic ancestry admixture of the subject.
In inequality (2), the allele frequencies were obtained from gnomAD29, or from ALFA30 if the SNP is not reported in gnomAD. The genetic ancestry admixture fractions were estimated by using iAdmix31.
In genomic ranges with two haplotypes, as the third allele is a duplicate, rare alleles are as likely to be duplicated as the common alleles. In contrast, when there are three independent haplotypes, a rare allele is unlikely to appear twice, making rare two-copy alleles more likely to appear in two-haplotype blocks (Fig. 1B).
Type 2 informative SNPs: variants in rare hypothetical haplotypes
When the three copies of chromosomes 21 originate from only two haplotypes (i.e., two of the three chromosomes are identical), then the two distinct haplotypes on heterozygous SNPs can be represented as \({({a}_{i})}_{i=1}^{M}\) and \({({b}_{i})}_{i=1}^{M}\), where former is formed by all the two-copy alleles, and the latter is formed by all the one-copy alleles. For the locus of the \({i}^{{th}}\) SNP, we can generate the hypothetical haplotypes formed by itself and its two immediately adjacent SNPs (a total of 3 loci), which can be represented as \(({a}_{i-1},{a}_{i},{a}_{i+1})\) and \(({b}_{i-1},{b}_{i},{b}_{i+1})\). For genomic ranges with three haplotypes, this procedure shuffles and recombines the alleles, creating arbitrarily rare haplotypes that do not appear in the general population (Fig. 1B). The frequency of the hypothetical haplotype \(({a}_{i-1},{a}_{i},{a}_{i+1})\) in the population can be calculated as \(F({a}_{i-1},{a}_{i},{a}_{i+1})/N\), where \(F({a}_{i-1},{a}_{i},{a}_{i+1})\) and \(N\) represent the number of occurrences of \(({a}_{i-1},{a}_{i},{a}_{i+1})\) and the total number of haplotypes reported in the 1000 Genomes project32, respectively. The expected frequency of the hypothetical haplotype \(({a}_{i-1},{a}_{i},{a}_{i+1})\), under the assumption of no linkage disequilibrium, can be calculated as \({{\rm{AF}}}({l}_{i-1},{a}_{i-1})\times {{\rm{AF}}}({l}_{i},{a}_{i})\times {{\rm{AF}}}({l}_{i+1},{a}_{i+1})\). We score \(({a}_{i-1},{a}_{i},{a}_{i+1})\) as the ratio between observed and expected frequencies (O/E ratio):
Note that differing with inequality (2), here the allele frequency \({{\rm{AF}}}({l}_{i},{a}_{i})\) was obtained from 1000 Genomes project (n = 3202 individuals, retrieved on April 4th, 2022). Because of the small number of available haplotypes in this database, the genetic ancestries were ignored when calculating the allele frequency for haplotype analysis. The hypothetical haplotype formed by single-copy alleles \(({b}_{i-1},{b}_{i},{b}_{i+1})\) can be scored similarly, and a SNP is defined to be in a rare hypothetical haplotype when
The steps involved in identifying rare two-copy alleles and rare hypothetical haplotypes are summarized in Fig. 1C.
Hidden Markov models
For each DS case, we constructed a hidden Markov model (Fig. 1F) on the set of heterozygous SNPs \(\Omega\), with the components defined as follows:
-
1.
vector of hidden states,
$${{\bf{Q}}}={\left({q}_{i}\right)}_{i=2}^{M-1},\,{q}_{i}\in \left\{2{{\rm{H}}},\, 3{{\rm{H}}}\right\},$$(5)where \(2{{\rm{H}}}\) and \(3{{\rm{H}}}\) indicate two-haplotype and three-haplotype states, respectively.
-
2.
vector of observations
$${{\bf{O}}}={\left({o}_{i}\right)}_{i=2}^{M-1},{o}_{i}=\left\{\begin{array}{cc}{{\rm{T}}}1,\hfill & {{\rm{if}}}\left(2\right)\,{{\rm{is}}}\; {{\rm{true}}},\hfill \\ {{\rm{T}}}2,\hfill & {{\rm{if}}}\left(4\right)\,{{\rm{is}}}\; {{\rm{true}}},\hfill \\ {{\rm{Others}}},& {{\rm{otherwise}}},\hfill \end{array}\right.$$(6)where \({{\rm{T}}}1\) and \({{\rm{T}}}2\) stand for type 1 and type 2 informative SNPs, respectively. For cases genotyped by SNP arrays, only rare hypothetical haplotypes were used as only common SNPs are covered by the platform.
-
3.
To define the initial, state transition, and emission probabilities, we first randomly selected 40 cases for manual inspection of two- vs three-haplotype state, most of which were visually apparent (e.g., based on AF of two-copy alleles and O/E ratio of hypothetical haplotype, Fig. 1G). For two-haplotype state, the initial probability was set as the proportion of samples with the first variant along the chromosome being assigned to the two-haplotype state; the transition probabilities were set as the proportion of variants followed by a state transition (a crossover event); For the emission probabilities, we used the proportion of each type of variants being assigned to the two-haplotype state. This was then repeated to set the parameters for the three-haplotype state.
The most likely vector of hidden states was estimated by using the Viterbi algorithm33.
Simulation
We estimated the distributions of (i) the allele frequency of two-copy allele and (ii) the score of the 3-SNP hypothetical haplotypes, under the assumptions of the triallelic chromosome 21 having two or three haplotypes, by sampling random genotypes/haplotypes from reference databases. The simulation was done by repeating the following procedure for 100 times:
-
i.
Allele frequencies of two-copy alleles
-
a.
Three haplotypes: for each SNP reported in reference database (gnomAD or ALFA), three independent alleles were sampled based on allele frequencies in the reference, and the allele frequency of the two-copy allele was recorded if the genotype was heterozygous.
-
b.
Two haplotypes: for each SNP, two independent alleles were sampled based on allele frequencies in the reference, and one was doubled at equal probability to make three alleles. If the final genotype was heterozygous, the allele frequency of the two-copy allele was recorded.
-
a.
-
ii.
O/E ratio of hypothetical haplotypes
-
a.
Three haplotypes: three complete haplotypes of chromosome 21 were sampled from the 1000 Genomes dataset, and the O/E ratios for all hypothetical haplotypes formed by three adjacent SNPs were recorded.
-
b.
Two haplotypes: similarly, two complete haplotypes of chromosome 21 were sampled. One of the haplotypes was doubled to make a total of three haplotypes, and the O/E ratios for all the hypothetical haplotypes formed by three adjacent SNPs were recorded.
Trio analysis and performance evaluation metrics
When genotyping data of the DS child and both parents were available, the stage of NDJ error and crossover event can be determined as described previously11. Briefly, SNPs that were homozygous non-reference in one parent but homozygous reference in the other were used as markers to infer the parent of trisomy 21 origin. In the DS child, these SNPs were heterozygous, and the two-copy alleles were from the parent of origin of the extra copy of chromosome 21. After the parent of origin was determined, SNPs that were heterozygous in the parent of origin but homozygous reference in the other parent were used to segment chromosome 21 into two-haplotype or three-haplotype blocks. For such SNPs, the non-reference allele could be doubled or absent when there were two haplotypes in the DS child. When there were three haplotypes, it was always the reference allele that was doubled. In these SNPs, whether the non-reference is doubled were used in HMMs as observations to segment chromosome 21 into blocks of two or three haplotypes.
Performances of MeiHMM in NDJ error and crossover event identification were evaluated by comparing to the results of trio analysis on the same set of DS cases (n = 152). For NDJ error classification, accuracy was used to measure the overall performance. It is defined as
For crossover event identification, let \({{\rm{TP}}}\), \({{\rm{FP}}}\) and \({{\rm{FN}}}\) denote the number of true positives, false positives, and false negatives, respectively, \({{\rm{precision}}}={{\rm{TP}}}/({{\rm{TP}}}+{{\rm{FP}}})\) and \({{\rm{sensitivity}}}={{\rm{TP}}}/({{\rm{TP}}}+{{\rm{FN}}})\), were used as the performance evaluation metrics.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The WGS data of the DS cases (both ALL and non-ALL) and the parents were obtained from the Gabriella Miller Kids First Pediatric Research Program and NIH INvestigation of Co-occurring conditions across the Lifespan to Understand Down syndromE (INCLUDE) Project, which are available through dbGaP with accession number phs002330.v2.p1 [https://www.ncbi.nlm.nih.gov/projects/gap/cgi-bin/study.cgi?study_id=phs002330.v2.p1]. Because the raw sequencing data contain personal genomic and genetic information, access to these data is restricted for the advancement of research only. Access to the data is facilitated through dbGaP website by making applications to the data access committee and the expected timeframe of response is dependent on the complexity of each application. Access is granted annually with the option of renewal. SNP array data were obtained from a previously published study, which is publicly available through Gene Expression Omnibus with accession number GSE60607. Source data are provided with this paper.
Code availability
The code for MeiHMM is available at https://github.com/jjyanglab/MeiHMM34.
References
Bull, M. J. Down syndrome. N. Engl. J. Med. 382, 2344–2352 (2020).
Sherman, S. L. et al. Trisomy-21 - association between reduced recombination and nondisjunction. Am. J. Hum. Genet. 49, 608–620 (1991).
Antonarakis, S. E. et al. The meiotic stage of nondisjunction in trisomy-21 - determination by using DNA polymorphisms. Am. J. Hum. Genet. 50, 544–550 (1992).
Antonarakis, S. E., Avramopoulos, D., Blouin, J. L., Talbot, C. C. & Schinzel, A. A. Mitotic errors in somatic-cells cause trisomy-21 in about 4.5-percent of cases and are not associated with advanced maternal age. Nat. Genet. 3, 146–150 (1993).
Antonarakis, S. E. Parental origin of the extra chromosome in trisomy 21 as indicated by analysis of DNA polymorphisms. Down Syndrome Collaborative Group. N. Engl. J. Med. 324, 872–876 (1991).
Garcia-Cruz, R. et al. Dynamics of cohesin proteins REC8, STAG3, SMC1 beta and SMC3 are consistent with a role in sister chromatid cohesion during meiosis in human oocytes. Hum. Reprod. 25, 2316–2327 (2010).
Wolstenholme, J. & Angell, R. R. Maternal age and trisomy-a unifying mechanism of formation. Chromosoma 109, 435–438 (2000).
Ottolini, C. S. et al. Genome-wide maps of recombination and chromosome segregation in human oocytes and embryos show selection for maternal recombination rates. Nat. Genet. 47, 727–735 (2015).
Bell, A. D. et al. Insights into variation in meiosis from 31,228 human sperm genomes. Nature 583, 259 (2020).
Lamb, N. E. et al. Susceptible chiasmate configurations of chromosome 21 predispose to non-disjunction in both maternal meiosis I and meiosis II. Nat. Genet. 14, 400–405 (1996).
Chernus, J. M. et al. A candidate gene analysis and GWAS for genes associated with maternal nondisjunction of chromosome 21. PLoS Genet. 15, https://doi.org/10.1371/journal.pgen.1008414 (2019).
Lamb, N. E., Yu, K., Shaffer, J., Feingold, E. & Sherman, S. L. Association between maternal age and meiotic recombination for trisomy 21. Am. J. Hum. Genet. 76, 91–99 (2005).
Oliver, T. R. et al. New insights into human nondisjunction of chromosome 21 in oocytes. PLoS Genet. 4, https://doi.org/10.1371/journal.pgen.1000033 (2008).
Ariad, D. et al. Aberrant landscapes of maternal meiotic crossovers contribute to aneuploidies in human embryos. Genome Res. 34, 70–84 (2024).
Ariad, D. et al. Haplotype-aware inference of human chromosome abnormalities. Proc. Natl. Acad. Sci. USA 118, https://doi.org/10.1073/pnas.2109307118 (2021).
Ramachandran, D. et al. Contribution of copy-number variation to Down syndrome-associated atrioventricular septal defects. Genet. Med. 17, 554–560 (2015).
Li, Z. et al. Genomic landscape of Down syndrome-associated acute lymphoblastic leukemia. Blood https://doi.org/10.1182/blood.2023019765 (2023).
Allen, E. G. et al. Maternal age and risk for trisomy 21 assessed by the origin of chromosome nondisjunction: a report from the Atlanta and National Down Syndrome Projects. Hum. Genet. 125, 41–52 (2009).
Yang, Q. H. et al. Risk factors for trisomy 21: maternal cigarette smoking and oral contraceptive use in a population based case control study. Genet. Med. 1, 80–88 (1999).
Keen, C. et al. The association between maternal occupation and Down syndrome: a report from the National Down Syndrome Project. Int. J. Hyg. Envir. Heal 223, 207–213 (2020).
Christianson, R. E., Sherman, S. L. & Torfs, C. P. Maternal meiosis II nondisjunction in trisomy 21 is associated with maternal low socioeconomic status. Genet. Med. 6, 487–494 (2004).
Hunter, J. E. et al. The association of low socioeconomic status and the risk of having a child with Down syndrome: a report from the National Down Syndrome Project. Genet. Med. 15, 698–705 (2013).
Warren, A. C. et al. Evidence for reduced recombination on the nondisjoined chromosomes-21 in Down syndrome. Science 237, 652–654 (1987).
Carey, J. C. Trisomy 18 and trisomy 13 syndromes. In Cassidy and Allanson's Management of Genetic Syndromes (eds Carey, J. C. et al.) 737–956 (Wiley-Blackwell, 2020).
Nicorici, D. et al. FusionCatcher – a tool for finding somatic fusion genes in paired-end RNA-sequencing data. Preprint at bioRxiv https://doi.org/10.1101/011650 (2014).
Barinka, J. et al. RNAseqCNV: analysis of large-scale copy number variations from RNA-seq data. Leukemia 36, 1492–1498 (2022).
Gu, Z. et al. PAX5-driven subtypes of B-progenitor acute lymphoblastic leukemia. Nat. Genet. 51, 296–307 (2019).
Poplin, R. et al. Scaling accurate genetic variant discovery to tens of thousands of samples. Preprint at bioRxiv https://doi.org/10.1101/201178 (2018).
Karczewski, K. J. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581, 434–443 (2020).
Phan, L. et al. ALFA: allele frequency aggregator. www.ncbi.nlm.nih.gov/snp/docs/gsr/alfa/ (2020).
Bansal, V. & Libiger, O. Fast individual ancestry inference from DNA sequence data leveraging allele frequencies for multiple populations. BMC Bioinforma. 16, 4 (2015).
Genomes Project, C. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Rabiner, L. R. A tutorial on hidden Markov-models and selected applications in speech recognition. Proc. IEEE 77, 257–286 (1989).
Li, Z. et al. Inferring chromosome segregation error stage and crossover in trisomic disorders with application to Down syndrome. Github https://doi.org/10.5281/zenodo.15632851 (2025).
Acknowledgements
We would like to thank the patients and families who participated in this study for donating specimens for research, and the clinicians and research staff for assistance in sample collection, processing, and curation. We also thank Jeremy Hunt, Ting-Nien Lin, and Pam McGill from the Department of Pharmacy and Pharmaceutical Sciences at St. Jude for technical support and Dr. Cherise Guess from the Department of Scientific Editing at St. Jude for proofreading. This work was in part supported by the National Institutes of Health (X01HL145686-01 [P.J.L., K.R.R.], R01CA249867 [K.R.R., P.J.L.], P30CA125123-14S4 [K.R.R.], 1R03HD103908-01 [J.J.Y.], U10CA98543 [M.L.L.], U10CA180886 [M.L.L.], U10CA98413 [M.L.L.], U10CA180899 [M.L.L.], U24CA114766 [M.L.L.] and U24CA196173 [M.L.L.]), Department of Defense (W81XWH-20-1-0567 [P.J.L.]), the Lynch family (K.R.R.), and the American Lebanese Syrian Associated Charities (J.J.Y.).
Author information
Authors and Affiliations
Contributions
G.W., S.L.S., K.R.R., P.J.L. and J.J.Y. conceived the study. Z.L., W.Y., G.W., T.-C.C., Z.C., M.D., L.S., E.F., T.C.R., E.G.A., S.L.S. and J.J.Y. performed data analysis. M.S., A.J.C., N.A.H., J.M.G.-F., B.L.W., E.A.R., S.P.H., M.L.L., E.F., T.C.R., E.G.A., S.L.S., K.R.R. and P.J.L. provisioned study materials. All authors wrote, reviewed, and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, Z., Yang, W., Wu, G. et al. Inferring chromosome segregation error stage and crossover in trisomic disorders with application to Down syndrome. Nat Commun 16, 6316 (2025). https://doi.org/10.1038/s41467-025-61413-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-61413-w