Abstract
Rubber tree (Hevea brasiliensis) is the primary source of natural rubber and economically important. We present the haplotype-resolved, telomere-to-telomere, gap-free genome assembly of the cultivar CATAS 7-33–97, with both haplotypes containing complete telomeric and centromeric regions. Structural variations, including a 32.71 Mb inversion on chromosome 8, are identified. The fully assembled 36 chromosomes enable comprehensive identification of rubber biosynthesis genes and their allele-specific expression. By integrating transcriptomic and metabolomic data, we reconstruct the rubber biosynthesis pathway and confirm the mevalonate (MVA) pathway as the major carbon source for rapid latex regeneration during tapping. Jasmonic acid (JA) plays a key role in promoting rubber yield by enhancing biosynthetic activity in response to mechanical wounding. We propose a model where JA-induced myelocytomatosis proteins 2 activate mevalonate kinase 1 expression, boosting MVA synthesis and rubber production. These findings provide insights into rubber tree genomics and its molecular response to tapping.
Similar content being viewed by others
Introduction
The rubber tree (Hevea brasiliensis) is the source of 99% of natural rubber (NR) globally, and its distinctive physical properties play pivotal roles across various industrial sectors1. Rubber plantation is the major income source for the farmers in many tropical countries2. It was first introduced to Southeast Asia in 1876 by Wickham, therefore, the descendants were called Wickham germplasm3. Rubber breeding was first started in 1915 when its vegetative budding propagation was achieved, through which the agronomic traits could be maintained4. After over one century of breeding, the rubber tree clones have undergone a lineage of four generations, resulting in a notable six-fold increase in yield5. However, the rubber yield seems hardly to be further elevated after rapid increasing at the early stage of domestication. Breeders attribute the stagnation in yield elevation to limited genetic diversity, as modern rubber clones predominantly descend from only nine ancestral trees6.
In addressing this challenge, researchers have turned to advanced genomic approaches. High-density genetic maps and genome-wide association studies were used to identify loci and genes related to rubber agronomic traits7,8,9,10. The application of marker-assisted selection and genomic selection (GS), based on the identification of key genes, represents an efficacious breeding strategy in plant breeding11,12,13,14,15. Recent genomic studies have revealed the process through which wild rubber trees have been artificially selected to evolve into high-yield clones9,16. These insights, derived from high-quality genomic data, are crucial for precise and efficient breeding. However, the presence of large gaps and unresolved complex regions in the existing genome assemblies limits the accuracy and effectiveness of these genomic tools.
To date, several rubber tree genomes have been published. The first published genome was RRIM 600, a widely planted clone around the world, which was assembled using Illumina data with a 43× coverage. However, this assembly had a contig N50 of only 2.9 Kb17. Subsequent assembly of RRIM 600, BMP24, and CATAS 7-33-97 was performed using Illumina, and BAC library data to obtain scaffold-level genomes18,19,20. In 2019, the first chromosome-level genome of the GT1 clone was published, which increased N50 to 152.7 Kb and reduced scaffolds to 60021. Further enhancement of the rubber genome quality was achieved by advances in long-read sequencing. The use of Oxford Nanopore Technologies (ONT), High throughput chromosome conformation capture (Hi-C), and BioNano facilitated the development of chromosome-level genomes for wild (MT/VB/25 A 57/8)9 and the cultivated clone (CATAS 8-79)16, which improved chromosomal continuity with contig N50 of 3.51 and 11.2 Mb, respectively. These significant advances offer crucial data resources for genetic study and clone improvements in rubber trees. Currently, all the published genome sequences are not haplotype-resolved. Besides, the large amount of gaps in these assemblies reduces the data integrity and hinders the investigation into the complex genomic regions. Notably, critical regions like telomeres and centromeres remain unresolved in these genomes.
In this study, a haplotype-resolved T2T gap-free reference genome assembly was successfully constructed in the widely cultivated rubber clone, CATAS 7-33-97. The comprehensive T2T genome marks a major step forward for revealing the complex genome’s recalcitrant structural features, and highlights previously overlooked structural variation among haplotypes. Besides, the T2T rubber genome enables the analysis of the evolution of genes related to rubber production and reveals the rubber biosynthesis regulon behind the tapping response.
Results
The haplotype-resolved T2T assembly for the rubber tree reference genome
We generated 193.19 Gb HiFi reads (~122× coverage), 654.04 Gb Hi-C reads (~414× coverage), and 199.24 Gb ultra-long ONT reads (~314× coverage) for assembling CATAS 7-33-97 genome (Fig. 1A, Table 1 and Supplementary Data 1). The initial assembly comprises 256 contigs with a size of 3.16 Gb (max length: 113.95 Mb) (Supplementary Data 2). Through contig ordering, orientation, filtering, chromosomal anchoring, and assembly optimization, we resolved two haplotypes collectively comprising 36 pseudochromosomes (Supplementary Fig. 1). Haplotype A (hapA) contained nine gaps (one gap on chromosomes 8, and eight on chromosome 11), while Haplotype B (hapB) displayed seven gaps (one gap on chromosome 4, and three gaps on chromosomes 11 and 18, respectively) (Supplementary Data 3). Then, haplotype-specific long reads were independently mapped to their corresponding haplotype assemblies. The existing gaps were manually filled by checking the existence of long reads spanning across the gap junctions (Supplementary Fig. 2). All gaps were successfully filled, resulting in two gap-free haplotypes for the rubber clone CATAS 7-33-97, each with a size of 1.56 Gb. The hapA contained 38,635 protein-coding genes alongside 4,664 snoRNAs, 1,343 rRNAs, 736 tRNAs, and 250 miRNAs, and hapB contained 38,618 protein-coding genes, 4694 snoRNAs, 1,416 rRNAs, 740 tRNAs, and 247 miRNAs (Supplementary Data 4). Both hapA and hapB genomes exhibited highly similar repetitive sequence compositions, with 84.63% and 84.53% identified as repeats. Long terminal repeats (LTRs) dominated these sequences (76.56% in hapA; 76.33% in hapB), primarily comprising Gypsy (48.96% hapA; 49.33% hapB) and Copia retroelements (13.44% hapA; 13.36% hapB). In terms of haplotype assembly contiguity, the contig N50 for hapA is 93.64 Mb, and for hapB, it is 94.38 Mb, with contig/chromosome (CC) ratios of 7.78 and 6.44, respectively. The high contiguity indicates the high quality of the haplotype assemblies. Regarding assembly completeness, we evaluated the overall completeness using a k-mer-based approach, which resulted in 97.5%. Benchmarking Universal Single-Copy Ortholog (BUSCO) analysis of gene space completeness showed values of 0.988 for hapA and 0.99 for hapB. We also assessed genome completeness based on LTR retrotransposons, with LTR assembly index values of 16.45 for hapA and 16.7 for hapB. Regarding assembly correctness, base error rates were 4.22e-7 for hapA and 3.69e-7 for hapB. The quality value for hapA and hapB was 65.52 and 65.72, respectively (Supplementary Data 5, Supplementary Fig. 3). These results demonstrate the high quality of our genome.
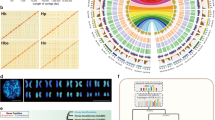
A The tree of cultivated rubber clone, CATAS 7-33-97. B The T2T genome of CATAS 7-33-97 is compared with the previously published genomes of wild rubber (MT/VB/25 A 57/8) and cultivated rubber (CATAS 8-79). Telomeric regions are marked with red dots, centromeric regions are marked with red blocks, and inverted regions are marked with yellow. C The circos plot of CATAS 7-33-97 genome characteristics, the center represents the syntenic regions between the hapA and hapB genomes. I–VIII represents the length of the 18 chromosome pairs in CATAS 7-33-97, telomere regions, GC content, gene density, Copia density, Gypsy density, tandem repeat density, and transposon element density, respectively. D Collinearity of chromosome 8 between hapA and hapB genomes. Source data are provided in Supplementary Data.
To identify the centromere regions in the rubber genome, ChIP-seq was performed using CENH3 antibodies, with two replicates for both immunoprecipitated samples and input control samples (Supplementary Data 6). Compared to the Input control group, distinct binding peaks were observed on each chromosome (Supplementary Fig. 4, Supplementary Data 7). By filtering the peaks on each chromosome, we identified the centromeric regions on each chromosome (Supplementary Data 8, Fig. 1B). Tandem Repeat (TR) sequences in the centromere regions were analyzed, revealing the presence of multiple TR sequences with varying periods (Supplementary Data 9). The rubber tree centromere is not composed of a single high-repeat TR sequence. Telomeres were identified by detecting a 7-base telomeric repeat sequence (CCCTAAA/TTTAGGG) on all chromosomes (Fig. 1B).
Variations were detected between the two haplotypes of CATAS 7-33-97 genome, including 28 inversions, ten transitions, 2,476,725 single nucleotide polymorphisms (SNPs), 169,365 insertions, and 170,240 deletions (Supplementary Data 10, Supplementary Fig. 5). 71.46% of the SNPs were located in the intergenic region, and only 3.5% were in the exons. To assess the impact of InDel between hapA and hapB on genomic heterozygosity, we screened for structural variants over 100 bp and identified 6330 InDel across 18 chromosome pairs. Annotation of the InDel indicated that most InDels were located in the non-coding region and only 114 in the exons. Strikingly, a 32.71 Mb multi-structured variant (sv33M) region (20,588,014 - 53,302,788) was found between chr8 chromosomal pairs, characterized by numerous structural variant events (Fig. 1B, D). It is noteworthy that a high degree of variations was observed in the chr8 corresponding regions in the wild (MT/VB/25 A 57/8) and the cultivated clone (CATAS 8-79) (Fig. 1B). To exclude any sequence misassembly, the long reads of CATAS 7-33-97, the wild, and CATAS 8-79 were mapped to the chr8A. The reads spanning the left and right boundaries of sv33M were examined. The read depth decreased rapidly within sv33M, with percentages of 82.35% (hapB), 92.31% (wild), and 92.86% (CATAS 8-79) compared to left region outside of sv33M. The percentages were 84.62% (hapB), 84.44% (wild), and 86.4% (CATAS 8-79) in comparison to the region outside of the right boundary (Supplementary Fig. 6). In particular, some long reads were found to span across both boundaries, demonstrating the real existing of this large inversion in all these genomes. This is demonstration of large structural variation between homologous chromosomes in the rubber tree. Due to its extremely large size, sv33M may have distinct genetic roles during evolution in Hevea brasiliensis.
Comparative genomics of rubber-producing genes in Euphorbiaceae
The T2T rubber genome enables the detailed evolution investigation of the genes related to rubber production. To identify orthologroups of rubber-producing related genes, we collected genomes from eight species (Arabidopsis thaliana22, Eucommia ulmoides23, Manihot esculenta24, Oryza sativa25, Populus trichocarpa26, Ricinus communis27, Taraxacum kok-saghyz28, and Vitis vinifera29) in addition to rubber tree. A total of 27,311 orthogroups were classified, occupying 89.7% of the overall genes, in which 8364 orthogroups were present in all these species (Supplementary Data 11, Supplementary Fig. 7). The rubber clone CATAS 7-33-97 (hapA) has 35,026 genes being divided into 16,584 orthogroups. In comparison to the low-yield wild rubber genome (MT/VB/25 A 57/8), the CATAS 7-33-97 exhibits 1297 expanded orthogroups and 662 contracted orthogroups (Fig. 2A). This represents 95.97% of the families exhibiting significant expansions and contractions among these species (P value ≤ 0.01). GO enrichment analysis revealed that the expanded gene family are involved in wounding response, terpene biosynthesis, hydrocarbon metabolism, jasmonic acid response, and lipid metabolism (Fig. 2B). This suggests that these gene family expansions may relate to the rubber yield, as latex is produced after the plant is injured and need to be replenished by synthesizing terpene from sucrose30. We also analyzed the GO enrichment results of the contraction gene. These genes were mainly enriched in the benzoate metabolic process, toxin catabolic process, response to reactive oxygen species, and sulfur compound metabolism. This suggests that this genotype may optimize resource allocation for latex biosynthesis by downregulating defense-related pathways (Supplementary Fig. 8).
A The Hevea gene cluster expansion and contraction: the pink, red and blue segments of the pie chart represent the maintained, expanded and contracted gene, respectively. B GO enrichment results for gene clusters with highly significant expansion of CATAS 7-33-97. P values were calculated by one-sided hypergeometric test. C The number of rubber-producing genes in Euphorbiaceae plants. D The number of genes involved in the mevalonate (MVA), methylerythritol 4-phosphate (MEP), initiator synthesis, and rubber elongation pathways in Euphorbiaceae genomes. E Phylogenetic tree of REF and SRPP genes in Euphorbiaceae plants. The REF and SRPP genes are labeled in green and blue, respectively.
To further study the evolution of rubber biosynthesis genes in Euphorbiaceae, we collected seven genome of Euphorbiaceae species, including two less rubber-producing plants, Ricinus communis (RC039)27 and Manihot esculenta (SC205)24, and five rubber tree genomes GT121, wild accession MT/VB/25 A 57/8, CATAS 8-79, CATAS 7-33-97 (scaffold level), and CATAS 7-33-97 (T2T). There are four classes of genes related to rubber biosynthesis in the plant, MVA pathway genes, Methylerythritol 4-phosphate (MEP) pathway genes, initiator synthesis genes, and rubber elongation genes (Fig. 2C, Supplementary Data 12). By comparing six genome assemblies with differences in the number of genes in four types, we found that the number of genes for rubber elongation type in rubber was significantly higher than those of Ricinus communis and Manihot esculenta (Fig. 2D). It could be inferred that the high rubber-producing ability of rubber trees was attributed to the expansion of genes in the rubber elongation type.
The rubber elongation factor (REF) and small rubber particle protein (SRPP) genes are recognized as the most important rubber-producing genes, which share high sequence similarity. The difference lies in the absence of a carboxy-terminus domain in REF protein. A total of 19 REF/SRPP genes were identified in this CATAS 7-33-97 genome, most of which exist in a cluster at edge of chr3, showing a state of dramatic expansion (Supplementary Fig. 9). Compared with the previously published assemblies, this T2T genome revealed an extra SRPP, HbSRPP11 in rubber tree. In the wild rubber tree (MT/VB/25 A 57/8), we identified 5 REF and 9 SRPP genes in the genome, which are significantly less than other cultivated clones. In Ricinus communis and Manihot esculenta, no REF but 3 and 5 SRPP genes were identified respectively. These results suggest REF/SRPP gene numbers were correlated to rubber-producing capability. With remarkably high homology between REF and SRPP genes, phylogenetic analysis was conducted on the identified 94 REF and SRPP genes. The SRPP genes were grouped into four clusters, while the REF genes were divided into three clusters (Fig. 2E), indicating the REF and SRPP subgroups may have originated through gene duplication events, followed by divergence and convergent evolution, leading to the formation of distinct subgroups for the REF and SRPP genes, respectively. Moreover, rubber trees have significantly more cis-prenyltransferase (CPT) genes than Ricinus communis and Manihot esculenta, suggesting its roles on contributing latex production (Fig. 2C).
Consistent allele-specific expression (ASE) in rubber tree
Using the collected transcriptome data of CATAS 7-33-97, we identified the differentially expressed bi-alleles in flower (FL), callus (CA), primary embryo (PE), cotyledonary embryo (CE), mature embryo (ME), and the latex in different tappings (T1, T7, TN)31,32 (Supplementary Data 13, Supplementary Fig. 10, 11). 10,136 alleles displayed specific expression in all these samples (Fig. 3A), among which 9546 (94.18%) showed consistent allele-specific expression (ASE), i.e., always dominantly expressed in a given haplotype. In these consistent ASE alleles, 4753 preferred to be highly expressed in hapA chromosomes, while 4793 alleles were always dominant in hapB (Fig. 3B). It was also surprising that only 590 showed dynamic ASE (dominant allele varied in different samples) pattern. The high percentage of consistent ASE pattern is quite different with that in cassava24, which is about 33%, indicating a distinguish ASE profiles in rubber tree.
A Differential bi-allelic (DBA) between CATAS 7-33-97 haplotypes (pink) and differentially expressed bi-allelic (DEBA) genes in various samples (green). B Consistent ASE pattern in rubber tree. The horizontal bar chart on the left represents the number of highly expressed bi-allelic genes in each haplotype. The vertical bar chart on the top shows the number of ASE genes in each haplotype in the samples connected by solid dots and lines. Note that the majority of ASE genes were highly expressed in a given haplotype in all samples. Cotyledonary embryo (CE), callus (CA), flower (FL), mature embryo (ME), primary embryo (PE). C The expression profile of the ASE genes in hapA and hapB, red dots represent genes involved rubber biosynthesis. P values are calculated using a two-sided quasi-likelihood F-test based on a negative binomial distribution. D Genes involved in the rubber biosynthesis pathways, i.e., Mevalonate (MVA), methylerythritol 4-phosphate (MEP), initiator synthesis (IS), and rubber elongation (RE) pathways. Genes highlight in orange are in ASE. Source data are provided in Supplementary Data.
Among these tissues and organs, TN is the latex sample collected from trees tapped for five years, representing the producing NR in mature rubber tree. A total of 4735 alleles showed showed ASE pattern, in which 2349 was highly expressed in hapA genome, while 2386 was dominantly expressed on hapB. These genes’ functions were primarily concentrated in biological processes, nucleic acid phosphodiester bond hydrolysis involved in DNA and RNA metabolism (Supplementary Figs. 10, 11). These processes represent crucial mechanisms by which cells regulate the stability of DNA and RNA. Interestingly, 17 out of 82 genes related to rubber biosynthesis (genes in MVA, MEP, initiator synthesis, and rubber elongation pathway) showed consistent ASE pattern (Fig. 3C, D), leading to expression imbalances between homologous chromosomes that can influence rubber production.
The mevalonate serves as a major carbon reservoir for rubber biosynthesis
Rubber yield is gained by tapping of the bark to collect latex produced in the laticifers (Fig. 1A). When a virgin tree is start to be tapped, the latex yield would increase gradually and reach a stable level after 7–10 tappings. The rubber-producing capability is enhanced during these consecutive taps (repeated mechanical wounding). To unravel the mechanism under this phenomenon, we collected latex samples (T1-T10) from unharvested trees (8 years old) from the first to the tenth tapping, as well as latex samples (TN) from trees that have been tapped for five years (13 years old). (Fig. 4A). As expected, the dry rubber yield gradually elevated during the first seven taps (T1-T7, 11.97 ± 1.01 g) and maintained at stable level afterward (T8-TN, Fig. 4B and Supplementary Data 14). Rubber particle sizes, which is negatively correlated with rubber biosynthesis activity33,34,35, reduced from 1.36 ± 1.11 μm to 0.96 ± 0.01 μm (Fig. 4C, Supplementary Data 15). Latex sucrose content, a negative indicator of latex metabolic activity36, was also dropped to lowest level of 5.08 ± 1.27 mM at T7 (Fig. 4D, Supplementary Data 16), demonstrating latex metabolism reached a very high level after 7 consecutive tappings. The rubber synthetic activity (atom percentage of 13C) measurement showed an increment from 1.09 ± 0.004 APC13C (T1) to 1.16 ± 0.015 APC13C (T7) and reached stable high activity afterward to TN (Fig. 4E, Supplementary Data 17). We therefore propose that the seventh tapping (T7) represents a critical time point for rubber synthetic activity assessment. Correlation analysis indicates a significant relationship among rubber particle size, sucrose content, dry rubber yield, and rubber synthesis rate. Specifically, there is a positive correlation between rubber particle size and sucrose content (r = 0.7), as well as between dry rubber yield and rubber synthesis rate (r = 0.69) (Supplementary Fig. 12).
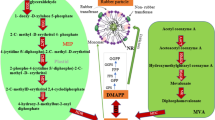
A Rubber tapping system diagram. Rubber trees that are 8 years old and have reached a circumference of 50 cm undergo the first tapping (T1) to collect latex, and the next tapping takes place after 3 days interval. The consecutive tappings were performed until T10. In addition, latex is collected from trees that have been tapped for 5 years (TN). B-E Measurement of dry rubber yield (B), rubber particle size (C), sucrose content (D) and synthetic activity of the latex (E) from each tapping. The center line represents the median; box bounds indicate the 25 and 75th percentiles; whiskers extend to the minimum and maximum values. For each time point, n = 16, biological replicates. P values were calculated using a two-sided Student’s t-test. F Top differentially accumulated metabolites (DAMs) in T1, T7 and TN latex, the color scale legend represents the standardized metabolite values to show the relative differences in the accumulated metabolite abundance at different periods. The classification of each metabolite is represented in parentheses. Metabolite abundance (MA) are normalized based on the z-score method (scale bar). G Schematic diagram of natural rubber biosynthesis pathway. The color scale legend of blue and yellow represent the expression levels of genes (log2(TMM)). REF and SRPP genes share one scale threshold because of excessively high expression levels, while the other genes share another identical scale. The color scale legend of red and green represent the metabolite abundance of DAM. Red font highlight differentially expressed genes and MVA. Source data are provided in Supplementary Data.
To elucidate how the rubber synthetic activity is elevated after consecutive tappings, comparative metabolomic and transcriptomic analyses were carried out among T1, T7, and TN (Supplementary Data 18). Total 390 differentially accumulated metabolites (DAMs) were identified among T1, T7, and TN taps. These DAMs were classified into 20 categories with 4 expression patterns (Supplementary Fig. 13). The category of amino acids and derivatives contains the largest number of DAMs (58), reflecting that the cytoplasm (latex) needs to be replenished after evacuation (tapping) in laticifers. Among the top 20 DAMs, the up-regulated metabolites mainly involve coumarins, amino acids and their derivatives, organic acids, saccharides, nucleotide derivatives, phenolic acids, and monoterpenoids, among others. Mevalonic acid, a precursor for isoprene pyrophosphate (IPP) involved in rubber biosynthesis, was also up-regulated. The down-regulated ones are primarily dominated by saccharides, which are considered as carbon source for rubber biosynthesis37 (Fig. 4F).
Rubber biosynthesis is a prolonged terpene biosynthesis pathway, mainly including precursor IPP synthesis and rubber polymer formation. Based on this high-quality genome, all the genes involved in rubber biosynthesis were identified and a pathway diagram was reconstructed according to their expression profiles responding to tapping (Fig. 4G, Supplementary Fig. 14). We found that transcript levels of MVA pathway genes are overall higher than those of MEP pathway genes. In the MVA pathway, each enzyme has at least one gene being significantly up-regulated by tapping. Among all genes in the MVA pathway, mevalonate kinase 1 (MVK1) is the most significantly up-regulated one, with over 10-fold transcript increment. Moreover, MVA, the substrate of MVK, was also found to increase by 14 and 8 folds at T7 and TN tappings (Fig. 4G). We conducted a correlation analysis between the rubber biosynthesis pathway gene and all DAMs. The results showed that a cluster containing 64 metabolites was significantly positively correlated with most of the rubber biosynthesis genes (Supplementary Fig. 15, Supplementary Data 19). These metabolites could be categorized into 18 groups, mainly including amino acids and derivatives, organic acids, nucleotides and derivatives, coumarins, etc. Notably, MVA, an organic acid, was the only differentially accumulated metabolite identified to directly participate in the rubber biosynthesis pathway. It exhibited significant correlations with the expression of 44 rubber synthesis-related genes and an extremely significant correlation with the MVK1 (p = 0.000418). Both the transcript and metabolite expression data strongly suggest that MVA accumulation is an indicator of rubber synthetic activity elevation, and this metabolite and its derivatives (mevalonate-5-phosphate and mevalonate-5-pyrophosphate) may serve as a major carbon reservoir for rubber biosynthesis in laticifers.
Consecutive tapping enhances rubber biosynthesis through JA signaling pathway
A total of 6466 differential expression genes (DEGs) were identified between three tappings (Supplementary Fig. 16A, B, E). The most significant differences were observed between T7 and T1 (4941 DEGs), in which 2328 genes were up-regulated and 2613 down-regulated (Supplementary Fig. 16C). These DEGs could be categorized into four clusters, with 1256 consecutively up-regulated genes in clusters1 (C1) and 1,476 down-regulated genes in the clusters4 (C4) (Fig. 5A, Supplementary Data 20). Gene ontology (GO) enrichment results indicated that terms related to the regulation of RNA biosynthetic processes (GO:2001141), heterocycle biosynthetic processes (GO:0018130), regulation of macromolecule biosynthetic process (GO:0010556) and nitrogen compound metabolic processes (GO:0051171) were significantly enriched in C1 (Fig. 5B), representing genes positively correlated with latex yield (r = 0.999) and synthetic activity (r = 0.951) in T1, T7 and TN tappings. In contrast, the C4 cluster observed genes with declined expression trends, positively correlating with negative indicators of latex metabolic activity, such as rubber particle size (r = 0.979) and sucrose content (r = 0.982). The Venn diagram comparison of these four enriched GO terms (GO: 2001141, 0018130, 0010556, and 0051171) revealed an overlap in 91 genes (Supplementary Fig. 16D).
A K-means clustering showed the expression changes over time, with T1, T7, and TN representing key tapping intervals. n = 1256, 2026, 1708, 1476, gene number. B Functional category of GO enrichment among the four major clusters. P values were calculated by one-sided hypergeometric test. C Regulatory network of key transcription factors involved in the macromolecular synthesis. D-G Jasmonic acid (JA), JA isoleucine conjugate (JA-ILE), JA valine (JA-Val) and indole-3-acetic acid (OxIAA) levels in the latex of T1, T7 and TN. n = 3, biological replicates. H The application of jasmonic acid (JA) promotes rubber biosynthesis activity. Rubber trees were treated with JA at the T1, T4, T7, and TN stages, followed by tapping of both the JA-treated and mock groups to assess rubber biosynthesis activity. n = 9, biological replicates. I The application of jasmonic acid (JA) promotes dry rubber yield during the early stages of tapping. Rubber trees were treated with JA at the T1, T4, T7, and TN stages, followed by tapping of both the JA-treated and mock groups to measure dry rubber yield. For T1, T4, and T7, n = 18; for TN, n = 12, biological replicates. J In vivo bioluminescence images of dual-luciferase activity. Each tobacco leaf was divided into four quadrants, including the following constructs: empty pGreenII 62-SK vectors (EV-SK), empty pGreenII 0800-LUC vectors (EV-LUC), HbMYC2 overexpression in pGreenII 62-SK (35S::HbMYC2-SK), and HbMVK1 promoter in pGreenII 0800-LUC (Pro-HbMVK1-LUC). K Statistical analysis of dual-luciferase reporter gene activity. n = 5, biological replicates. L Yeast one-hybrid assays showed the interaction between HbMYC2 and the promoter of HbMVK1. M Mechanism of rubber biosynthesis in response to tapping. Data were presented as mean values ± SD (*, p ≤ 0.05; **, p ≤ 0.01), p values in D–I, K are calculated with two-sided Student’s t-test. Source data are provided as a Source Data file.
The vast majority of these genes encode plant transcription factors (TF), in which MYC2 was predicted to be the hub gene in a String network regulating rubber biosynthesis (Fig. 5C, Supplementary Data 21). ERF1B and JAZ8 were JA signaling pathways TF, which may work together with MYC2 to regulate the increase of rubber synthetic activity during tapping (Fig. 5C). Furthermore, the hormone levels, JA, JA-ILE, and OxIAA, were significantly elevated with consecutive tappings, especially up-regulated by more than 4 folds from T1 to T7. (Fig. 5D–G, Supplementary Data 22). This may suggest JA’s role in regulating rubber synthetic activity. To investigate the role of JA in promoting rubber biosynthesis, untapped rubber trees were treated with exogenous JA (0.1 % w/v) 24 h prior to tapping at each of the T1, T4, and T7 timepoints. The atom percentage of 13C was significantly higher in JA-treated trees than the mock trees at T4 and T7 tappings (Fig. 5H, Supplementary Data 23), indicating exogenous JA application elevated rubber synthetic activity. At stages T1 and T4, the JA-treated group showed higher dry rubber yield than the control group, and at stage T7, the yield in the JA-treated group was significantly higher (Fig. 5I, Supplementary Data 24). And we collected latex produced by rubber trees treated with JA, and qRT-PCR results showed that JA upregulated the expression of MVK1 in latex (Supplementary Fig. 17). At the TN stage, the rubber biosynthesis activity and dry rubber yield in JA-treated trees were similar to those in the mock group, the differences were not statistically significant. We hypothesize that rubber trees at the early stage of tapping may exhibit greater sensitivity to JA. These results testified that consecutive tapping activated rubber biosynthesis by accumulating JA in latex.
It is noteworthy that a MYC-responsive element was identified in the upstream region of the MVK1 (-456 bp), which the homologous gene MVK2 does not contain. To verify the direct regulatory effect of MYC2 on MVK1, we conducted interaction experiments. The Dual-Luciferase Reporter Assay revealed a significant luciferase signal in the experimental group (35S::HbMYC2-SK + Pro-HbMVK1-LUC) (Supplementary Fig. 18, Fig. 5J). The relative luciferase activity in the experimental group increased 2.1-fold compared to the control group (EV-SK + Pro-HbMVK1-LUC), demonstrating that MYC2 could activate the transcription of MVK1 (Fig. 5K). Yeast one-hybrid (Y1H) results further confirmed that MYC2 can interact with the G-box cis-element fragment on the MVK1 promoter (Fig. 5L). We therefore infered that consecutive tapping leads to an increase in endogenous JA levels, and the MYC2 in the activated JA pathway upregulates the expression of MVK1 gene in the mevalonate pathway, thereby promoting rubber biosynthesis (Fig. 5M).
Discussion
Our haplotype-resolved T2T genome facilitated the resolution of the intricate telomeric and centromeric regions of the rubber genome, in addition to structural variation and allelic expression imbalance among rubber haplotypes. Comparative genomic analyses of Euphorbiaceae plants revealed the expansion of the terpene synthesis gene family in cultivated rubber and copy number variations of the rubber elongation gene within the Rubber genus. Multi-omics data from consecutive tappings were employed to propose a model elucidating how tapping activates rubber biosynthesis and improves latex yield. While the detailed regulatory pathways remain to be experimentally validated, this gap-free reference genome is expected to elucidate detail mechanisms soon, and to help in breeding more high-yield clones.
Haplotype assembly improves understanding of homologous chromosome variation in rubber tree
Currently, most genomes of highly heterozygous diploid or polyploid species are chimeric, overlooking the differences between haplotypes. However, researchers have recently assembled haplotype genomes in many species, revealing the impact of chromosomal rearrangements, structural variations, and haplotype-specific allele expression on phenotypes38,39,40,41. In this study, we assembled the haplotype-resolved T2T genome in the rubber tree, which has significant advantages over previously published genomes in both structural studies and allele expression comparisons. In addition, haplotype-resolved genome assembly can facilitate allele studies. In rubber tree, new RNA need to be synthesized to assemble ribosomes for protein translation and to initiate latex replenishment after each tapping. During this process, bi-allelic expression differences were detected in rubber-producing genes, indicating rubber biosynthesis potential may be determined by the parental origin of the allele (Supplementary Fig. 11). The strong allele preferences possibly be attributed to the genetically narrow background with high heterozygosity in rubber trees. This finding may have great significance in rubber breeding by utilizing dominant alleles to accelerate the genome selection strategies.
In terms of genome structure, our T2T genome has identified all telomeres and the centromeric regions across chromosomes. In most plants, centromeres are composed of large arrays of species-specific satellite repeats interspersed with LTR retrotransposons. The majority of these centromeres are enriched with DNA repeats ranging in size from 100 to 1000 bp42,43. Additionally, there is a diversity of satellite repeats for different chromosomal centromeres, which suggests a rapid evolution of the centromeric sequence44,45,46,47. In the centromeric regions of Hevea brasiliensis chromosomes, TR units of varying lengths were identified. Among these, the largest satellite repeat unit spans 451 bp, accounting for 13.62% of the centromeric region in chromosome 11B. Additionally, satellite repeats of 403 bp, 146 bp, and 60 bp showed relatively high abundance in centromeric regions (Supplementary Data 9, Supplementary Fig. 19). Notably, satellite repeats of 21, 19, 146, 15, 13, 14, and 145 bp were widely distributed across multiple chromosomes (Supplementary Fig. 19). Similar observations have been reported in potato, where some chromosomes lack detectable satellite repeats in their centromere sequences48. Based on these findings, we hypothesize that the sequences in rubber tree centromeres may not be composed solely of a single LTRs unit, and their structural integrity likely relies on synergistic interactions among multiple repeat units.
The genetic basis and hormonal regulatory network of rubber tree yield traits
NR production is influenced by multiple factors, including sucrose transport, laticifer development, rubber biosynthesis, latex flow with laticifer plugging, and latex regeneration37,49,50,51,52,53. From a genetic perspective, multiple studies have shown that the amplification of the REF/SPRR gene family is a key factor enabling plants to possess latex production capability. Additionally, the presence of more latex-related genes may further enhance the rubber biosynthesis capacity of cultivated rubber trees. Our study found that cultivated rubber tree (CATAS 7-33-97) had three more REF genes (REF2, REF3, REF4) and two more SRPP genes (SRPP3 and SRPP10) compared to wild rubber tree (MT/VB/25 A 57/8). The REF2, REF3 and REF4 genes are tandemly arranged in the genome. Among this gene cluster, REF1 and SRPP1 had the highest expression levels in CATAS 7-33-97, especially during key stages of the tapping response, where they exhibited significant induced expression. The transcripts level of REF2, REF3 and REF4 were lower than REF1, they were still highly expressed in latex (Supplementary Data 25). In contrast, SRPP3 and SRPP10 genes showed negligible expression. Based on these results, we speculate that the gene dosage effect caused by multiple REF genes could be one of the reasons for the high yield of cultivated rubber.
The gene family expansion in cultivated rubber further reveals the genetic changes that occurred during the breeding/domestication process. We analyzed amplified genes in CATAS 7-33-97 (GO-enriched P value < 1e-5) across multiple latex developmental stages (Supplementary Data 26, Supplementary Data 27, Supplementary Fig. 20). Among these, we identified a significant upregulation of JAZ8. In the JA signaling pathway, JAZ proteins bind to TFs, such as MYC2 and are involved in stress responses and resistance mechanisms in plants. The correlation analysis between differentially expressed amplified genes and DAMs indicates that the function genes significantly positively correlated with MVA include those involved in the following processes: fatty acid biosynthesis, E3 ubiquitin-protein ligase, ADP-ribosylation factor GTPase-activating protein, calcium-dependent protein kinase, etc. Therefore, we speculate that the amplified genes in cultivated rubber trees promote rubber production.
To investigate additional causes of higher yield in cultivated rubber trees, we analyzed 326 marker-trait association candidate genes related to latex yield from our previous publication9, identifying 71 DEGs during tapping (Supplementary Data 28). These genes primarily associate with rubber biosynthesis and latex flow regulation. Notably, the methyl jasmonate (JA) esterase 1 (MJE1) gene exhibited a 4.8-fold upregulation from T1 to T7. The MJE1-encoded protein hydrolyzes methyl jasmonate (MeJA) into JA, further confirming JA’s crucial role in latex yield enhancement.
Hormones play a direct regulatory role in the growth and development of rubber trees as well as in rubber synthesis. When plants experience biotic stress or mechanical damage (tapping), JA activates downstream transcription factor networks and regulates key enzymatic pathways, thereby inducing the synthesis of antimicrobial peptides and phenolic compounds at wound sites. This dual mechanism effectively inhibits pathogen invasion while promoting tissue repair54,55.Our study observed progressive JA accumulation and concurrent IAA elevation during sustained rubber tree tapping. Although the precise molecular mechanisms remain uncharacterized, prior research confirms IAA positively regulates latex yield by promoting laticifer cell differentiation56. Notably, JA and auxin signaling pathways coordinate plant development through core functional modules involving COI1, MYC2, and JAZ57,58. JA and ethylene reportedly affect rubber production by acting on laticifer cell etc59,60,61,62,63, with JA and ethylene exhibiting context-dependent antagonistic/synergistic interactions in stress responses64,65. The AP2/ERF transcription factor ORA59 integrates JA and ethylene signaling to amplify antimicrobial capacity66,67. The observed overexpression of JAZ, MYC2, AP2, and ERF1B TFs in our experiments suggests a dual mechanistic role for MYC2 under mechanical wounding: direct regulation of mevalonate kinase MVK1 expression, coupled with coordinated mediation of IAA and ethylene biosynthesis to potentiate latex yield. These findings collectively indicate that JA interactions with multiple phytohormones likely orchestrate rubber tree responses to mechanical damage and productivity regulation. Future studies should employ hormone interaction network modeling to elucidate these regulatory mechanisms.
Methods
Plant materials and sequencing
The rubber tree clone, CATAS 7-33-97, is planted in the National Rubber Tree Germplasm Repository (Danzhou, Hainan, 19°52′N, 109°50′E, South China). Young bronze leaves were collected, immediately frozen in liquid nitrogen, and sent for library construction. For latex transcriptome and metabolome analysis, the trees were tapped half spiral tapping (S/2) with 3 days interval (1/2 S 3 d). Fresh latex samples were collected and immediately flash-frozen in liquid nitrogen, followed by transportation to Wuhan Metware Biotechnology Co., Ltd (Wuhan, China) for metabolite profiling, three biological replicates were collected at each time point. Detailed experimental procedures are provided in the supplementary methods. The CATAS 7-33-97 WGS data were generated from the PacBio Sequel II platform, producing high-fidelity reads (HIFI) data; the Oxford Nanopore PromethION 48 platform, producing ONT sequences; and the Illumina platform, producing Hi-C sequences.
Chromatin immunoprecipitation (ChIP-Seq) experiments were performed using formaldehyde-crosslinked nuclei from young leaves of CATAS 7-33-97, with IP and Input control samples each containing two replicates. Detailed protocols for HbCENH3 antibody production and the complete ChIP-Seq workflow are provided in the supplementary methods.
Genome assembly and annotation
We refer to the grape genome assembly process29,68,69. The HiFi data, ONT data, and Hi-C data underwent self-correction, trimming, assembly, and phasing using the default parameters of the hifiasm (v.0.19.4)70. The initial output of hifiasm resulted in two haplotype genome sequences. Ragtag with default parameters was applied to generate homology-based scaffolds by referring to the wild rubber genome71. Short contigs from the cytoplasmic genome and repetitive sequences were removed (Supplementary Fig. 21). For feature annotation, juicer (v1.6)72 was employed in conjunction with homology-based scaffolds and Hi-C data. Subsequently, 3D-DNA software was used for scaffolding73. The assembly results were manually corrected using Juicebox Assembly Tools (v2.20.00)74, resulting in a chromosome-level genome. However, gaps persisted in the genome. Finally, minimap2 (v2.24)75 was used to assign haplotype-specific long reads to haplotypic GAP regions. The BAM files were observed through IGV76 to fill the GAP and confirm the sequences in GAP positions. In the end, we obtained a gap-free genome for CATAS 7-33-97.
Repetitive sequences prediction
The TRs within the entire genome were initially annotated using Tandem Repeats Finder77. Subsequently, transposable elements (TEs) within the rubber tree genome were identified through a combination of ab initio prediction and homology-based methods. To begin, an ab initio repeat library specific to the rubber tree was constructed using default parameters with MITE-hunter78 and RepeatModeler79, which included LTR_FINDER, ltr_harverst, and LTR_retriver components tailored for plant genomes. This library was then compared against TEclass Repbase (http://www.girinst.org/repbase) to classify the various types of repeat families. For comprehensive genome-wide identification of repeats, RepeatMasker80 was employed to search for both known and novel TEs, utilizing both the de novo repeat library and the Repbase TE library. Overlapping TEs belonging to the same repeat class were subsequently merged and analyzed.
Non-coding RNA prediction
For the prediction of non-coding RNAs, a multi-step approach was employed. Initially, the cmscan program within the Infernal (v1.1.4)81 was utilized to align genome sequences against the Rfam database82, facilitating the detection of various RNA types including MicroRNA, rRNA, small nuclear RNA, and small nucleolar RNA (snoRNA). Subsequently, tRNAscan-SE (v2.0.9)83 was applied with eukaryote-specific parameters to identify transfer RNA (tRNA) sequences within the rubber tree genome. Lastly, the prediction of ribosomal RNAs (rRNAs) and their subunits was accomplished using models constructed by RNAmmer (v1.2)84. This comprehensive approach ensured a thorough exploration of diverse non-coding RNA species within the genome.
Functional gene annotation
The annotated genes were subjected to further annotation by aligning their corresponding translated protein sequences against various databases, including the Non-Redundant Protein Database (NR), Kyoto Encyclopedia of Genes and Genomes85, Eukaryotic Orthologous Groups of proteins86, GO87, and Swiss-Prot databases88, utilizing the blastp program with parameters set to -evalue 1e-5 and -max_target_seqs 1. For the genes related to rubber production, we performed a manual correction.
Annotation evaluation
To evaluate the genome annotation, BUSCO89 analysis was employed. The annotated protein sequences were aligned against the corresponding BUSCO databases using HMMER390, and the integrity of the alignment results was used to determine the presence of BUSCOs.
Chromatin immunoprecipitation (ChIP-seq) sequencing
Chromatin immunoprecipitation experiments were conducted using the previously described method91. The preparation of the HbCENH3 antibody was completed by WUHAN GENECREATE BIOLOGICAL ENGINEERING CO. LTD. Based on the homology comparison of AtCENH3 in Arabidopsis and the analysis of the antigenic epitopes of this protein, a 120 AA amino acid sequence specific to the HbCENH3 protein in rubber trees was screened out and inserted into the expression vector PETb2M (Supplementary Data 29, Supplementary Data 30). The protein was expressed and purified in Escherichia coli (Supplementary Fig. 22A). The polypeptide was used to immunize Japanese white rabbits (two biological replicates) to prepare antiserum and purify the HbCENH3 protein antibodies (G2825 and G2826) through antigen affinity purification, yielding 20 mg of each antibody. The purified antibodies were adjusted to a working concentration of 10 µg/mL (Supplementary Fig. 22B), ELISA and Western blot results showed that the antibody had good titer, sensitivity and specificity (Supplementary Data 31, Supplementary Fig. 22C). The ChIP-Seq experiment was conducted following published method92. Briefly, the purified antigen was mixed with Freund’s complete adjuvant (FCA), and the specific peptide fragment was injected into rabbits. Subsequently, antiserum was collected and the antibody was purified. Young leaves of CASTA 7-33-97 were used for nuclear DNA extraction, followed by cross-linking with formaldehyde and ultrasonic treatment to shear the chromatin into 300–500 base pair fragments. Immunoprecipitation was performed using the HbCENH3 fragment-specific antibody and incubated at 4 °C. The antibody-chromatin complex was isolated by binding to protein A/G Mag beads (GenScript, China). For sequencing, the immunoprecipitated DNA and input DNA were used to construct libraries. DNA sequencing libraries were generated using the Universal Plus DNA Library Preparation Kit (Vazyme). Sequencing was performed by Wuhan IGENEBOOK Biotechnology Co., Ltd. (http://www.igenebook.com) on the Illumina NovaSeq 6000 platform. The original antibody synthesis and validation reports have been uploaded to GitHub (https://github.com/Licc900/rubber_t2t).
Identification of telomeres and centromeres in the Rubber genome
To analyze the telomere repeat units using the TIDK (v0.2.0) (https://github.com/tolkit/telomeric-identifier). Then, we conducted a whole-genome search using the following parameters: tidk search -f hap.fa -s TTTAGGG -o tidk_search --dir telomere_find. Finally, we used TIDK plots and circos (v0.69.6)93 to visualize telomere peaks.
Raw data of ChIP-seq were filtered using Trimmomatic (v0.38)94. Then, clean reads were mapped to the CATAS 7-33-97 haplotype-resolved genome using BWA (v0.7.15)95. Potential PCR duplicates were removed using Samtools (v1.3.1)96. After merging the two technical replicates of the IP and Input control sample’s BAM files, peaks were called using MACS2 (v2.2.9.1)97 with the following parameters: (-g 3e9, -q 0.05, -bdg). Wig files generated by MACS software were used for data visualization through IGV76. By screening peaks on each chromosome (length, P value, proximity), we manually determined the centromeric regions on each chromosome.
Structural variation analysis
To investigate structural variations among rubber genomes, we compared the 4 haplotype genomes sequentially. First, use minimap2 to compare the two genomes to obtain the bam file, and then use SyRI (v0.23.4)98 to mine the generated VCF variation information of the bam file. Finally, structural variations were visualized using plotsr (v1.1.0) and NGenomeSyn (v1.41)99,100.
On chromosome 8, we observed a large structural variation region to verify the accuracy of the structural variation. We used minimap275 to map the SMRT long-reads of CATAS 7-33-97, MT/VB/25 A 57/8, and CATAS 8-79 to the hapA genome and then used IGV76 to observe the read characteristics of the 20 kb region upstream and downstream of this region and verify the structural variation.
Gene family expansion and contraction analysis
The orthogroups in the ten genomes were analyzed using OrthoFinder101 to infer their orthologous relationships. The species tree alignment sequences were then trimmed using Trimal102, and a species phylogenetic tree was constructed using raxmlHPC-PTHREADS with the PROTGAMMAJTT model. Finally, CAFE5103 was employed to analyze the expansion and contraction of orthogroups. Orthogroups exhibiting significant expansion or contraction (p ≤ 0.01) in cultivated rubber were selected for GO enrichment analysis.
Identification of rubber production-related genes
Firstly, we extracted the CDS sequences and converted them into protein sequences. Then, we used the published CDS sequences of rubber biosynthesis genes as a database20 and employed Blast+104 for conducting alignments and searches. Subsequently, we manually checked the alignment and identified the sequences that align with the genes. For SRPP and REF genes, we aligned the database sequences to the hapA genome and identified any unannotated ones. Additionally, we incorporated NCBI’s Conserved Domain Database (https://www.ncbi.nlm.nih.gov/Structure/bwrpsb/bwrpsb.cgi)105 and MEME106 as complementary tools for aiding in the identification process, thereby enhancing the overall accuracy.
Bi-allelic expression analysis
Firstly, syntenic blocks between hapA and hapB were identified utilizing MCScanX107. Subsequently, genes within these syntenic blocks were aligned using BLASTP (e value = 1e-10). Gene pairs with the highest bit score and greater than 50% identity for each gene in the block were selected. Ultimately, 30,586 pairs of bi-allelic were obtained.
For expression analysis, RNA-seq data were aligned with the hapA and hapB sequences using HISAT2 (v2.2.0)108. The alignment records with unique alignment positions were extracted based on the characteristics of HISAT2 alignment. Subsequently, quantification was performed using the Rsubread (v2.16.1)109 package. Finally, differential gene expression was determined by analyzing the identified gene pair sequences from previous steps.
Latex physiology diagnosis and rubber biosynthesis activity measurement
5 ml fresh latex from each tree was chilled on ice immediately for 5 min, and the rubber latex physiology diagnosis was carried out when the samples were carried back to the laboratory. Detailed measurements were according to those previously described110. For rubber molecular weight determination, 0.2 ml latex was dried and dissolved in 10 ml tetrahydrofuran (THF) for 3 days, then filtered with a 0.45 μm Minisart SRP 25 filter (Sartorius, Gottingen, Germany). The dissolved NR was passed through a Styragel HR5gel permeation chromatography (GPC) column (Waters Corporation) installed on a Waters 1515 isocratic HPLC pump system, and the signal was detected with a 2414 refractive index detector (RID) (Waters Corporation, Milford, MA, USA). GPC was carried out using 30 μl of solution at 40 °C, and the flow rate was set as 1.0 ml min-1. The latex rubber biosynthesis activity was determined by calculating the 13C content percentage of the total carbon according to Deng et al.111.
To study the correlation between rubber yield-related substances, ggstatsplot (v0.12.2)112 was used to conduct correlation analysis and statistical significance on the rubber particle size, dry rubber yield, sucrose content, and synthetic activity of the latex.
Latex metabolite profiling and identification of DAMs
Sample preparation for LC-MS/MS Analysis
The latex sample was thawed on ice after being taken out from the −80 °C freezer (three biological replicates were performed for each time point). A centrifuge tube and weighing spoon were precooled in liquid nitrogen, and then 50 mg of the powdered sample was weighed. Subsequently, 1.2 mL of 70% methanol internal standard solution was added to the sample, which was vortexed for 15 min to ensure complete mixing. The mixture was then centrifuged at 16,000 × g for 3 min at 4 °C. After centrifugation, the supernatant was carefully pipetted and filtered through a 0.22 μm microporous membrane. The filtered supernatant was stored in an injection vial for subsequent LC-MS/MS detection.
UPLC conditions
The sample extracts were analyzed using an UPLC-ESI-MS/MS system (UPLC: ExionLCTM AD, https://sciex.com.cn/; MS: Applied Biosystems 6500 Q TRAP, https://sciex.com.cn/). The analytical conditions were as follows: UPLC was performed on an Agilent SB-C18 column (1.8 μm, 2.1 × 100 mm). The mobile phase consisted of solvent A (pure water with 0.1% formic acid) and solvent B (acetonitrile with 0.1% formic acid). The gradient elution program was as follows: initially 95% solvent A and 5% solvent B, followed by a linear gradient to 5% solvent A and 95% solvent B within 9 min, which was held for 1 min. Afterward, the mobile phase composition was adjusted to 95% solvent A and 5% solvent B within 1.1 min and maintained for 2.9 min. The flow rate was set at 0.35 mL/min, and the column oven temperature was maintained at 40 °C. The injection volume was 2 μL. The effluent was then directed to an ESI-triple quadrupole-linear ion trap (QTRAP)-MS for analysis.
ESI-Q TRAP-MS/MS parameters
The ESI source operation parameters were as follows: source temperature set to 500 °C; ion spray voltage (IS) of 5500 V (positive ion mode) and −4500 V (negative ion mode); ion source gas I (GS1), gas II (GS2), and curtain gas were set at 50, 60, and 25 psi, respectively. The collision-activated dissociation (CAD) was set to high. QQQ scans were acquired as multiple reaction monitoring (MRM) experiments, with nitrogen used as the collision gas at a medium setting. Declustering potential (DP) and collision energy (CE) for individual MRM transitions were optimized. Specific MRM transitions were monitored for each period based on the metabolites eluted during that time frame.
Metabolite identification and quantification
Metabolite identification and quantification were performed using a self-constructed database (MWDB, Metware Database) and a triple quadrupole mass spectrometer equipped with MRM. The MWDB is a proprietary local metabolite database developed by Metware Biotechnology Co., Ltd., containing high-quality reference MS/MS spectra, retention times (RT), and chemical information for known plant metabolites. The annotation was based on the combination of accurate precursor ion mass (Q1), product ion mass (Q3), RT, isotopic distribution, and MS/MS fragmentation patterns. An in-house intelligent MS2 spectral matching algorithm was employed to compare the MS2 spectra and RT of each sample against those in the database. The matching was performed with a mass tolerance of 20 ppm and a RT tolerance of 0.2 min.
Isotope signals, redundant signals corresponding to ions such as K⁺, Na⁺, and NH₄⁺, and fragment ions from larger molecules were excluded to minimize interference. Metabolite identification confidence was categorized into three levels: Level 1 – MS2 spectra and RT matched with a score ≥0.7; Level 2 – match score between 0.5 and 0.7; Level 3 – Q1, Q3, RT, DP, and CE consistent with the database but with limited spectral evidence. Standard compounds were used to validate metabolite identities when available. For metabolites lacking commercial standards, identification was supported by high-quality spectral matching and RT alignment.
In the MRM mode, the first quadrupole was used to select the precursor ions (parent ions) of target metabolites, while ions from other molecules were filtered out to reduce background noise. These precursor ions underwent collision-induced dissociation in the collision cell, forming a series of fragment ions. The third quadrupole then selected specific fragment ions as target ions, ensuring higher accuracy and reproducibility by eliminating non-target ions.
For metabolite quantification, chromatographic peaks were integrated, and the area under each peak was used to represent the relative content of each metabolite. All chromatographic peak areas were calibrated by comparing the RT and peak shapes of the same metabolite across different samples to ensure accurate and consistent quantification. Data analysis was conducted using Analyst 1.6.3 software, and chromatographic data were further processed using MultiQuant software to integrate and normalize the peak areas. The ion intensities of each metabolite were recorded as counts per second, and the extracted ion chromatograms (XICs) were generated to display the metabolite detection profile.
Selection of DAMs
Differential accumulation metabolites were initially screened based on the Variable Importance in Projection (VIP) values obtained from the Orthogonal Partial Least Squares Discriminant Analysis (OPLS-DA) model, with biological replicates ≥3. VIP values indicate the contribution of each metabolite to group differentiation in the model, with metabolites having VIP > 1 considered to exhibit significant differences. Subsequently, univariate analysis was performed to further refine the selection based on the fold change (FC) values. The criteria for selecting differential accumulation metabolites were as follows: 1. Metabolites with VIP > 1 were selected. VIP > 1 indicates that the metabolite has a significant impact on the differentiation between groups in the model; 2. Metabolites with FC ≥ 2 or FC ≤ 0.5 were selected. Metabolites showing a twofold or greater difference between the control and experimental groups (or a reduction by half) were considered significantly different.
Analyze differential expression, gene clustering, and functional enrichment
R package corrplot (v0.92)113 was used to present the correlation coefficient matrix and analyze feature samples. The vegan (v2.6.4)114 package for PCA analysis was evaluated sample similarity and dispersion. The edgeR (v3.38.1)115 was used for differential expression analysis. The statistical threshold of DEGs were the following parameters: FC > 2 and adjusted P ≤ 0.05. Tapping rubber displaying the temporal profiles of clusters was detected by k-means using R package Mfuzz (v2.62.0)116. For each clustering, genes were performed functional enrichment by TBtools (v2.041)117. GO terms of each gene are from genomic functional annotation. The top 5 with P-value representative GO terms were presented for each clustering. Protein interaction relationship was obtained using protein sequences in the STRING118 database, with Arabidopsis as a reference, and combined with expression differences between T7 and T1 visualized into a PPI network by cytoscape (v3.10.0)119.
Dual luciferase reporter assay
The dual luciferase reporter assay was performed as previously described9. Briefly, the promoter sequence of HbMVK1 was ligated to the reporter vector pGreen0800II-LUC, and the coding sequence (CDS) of HbMYC2 was cloned into the pGreenII 62-SK vector. The constructed vectors were individually transferred into Agrobacterium GV3101-p19 strain. Agrobacterium cultures containing the reporter and effector vectors were co-infiltrated into Nicotiana benthamiana leaves. Three days post-infiltration, in vivo imaging was performed using a Multifunctional Plant In Vivo Imaging System (ROCEL, Beijing, China). Simultaneously, the LUC/Renilla (Ren) activity ratio was measured according to the manufacturer’s instructions using the Dual Luciferase Reporter Gene Assay Kit (RG027, Beyotime Biotechnology Co., Ltd., Shanghai, China).
Y1H assay
The method of yeast monomerisation refers to previously published paper120. The CDS of HbMYC2 was cloned into the pGADT7 vector (Clontech, USA) as the prey, while a 60-bp fragment containing the G-box cis-element from the promoter region was inserted into the pAbAi vector (Clontech, USA) as the bait. The pAbAi-ProHbMVK1 construct was transformed into Y1HGold yeast cells (WEIDI BioTech, Shanghai, China) and selected on SD/-Leu medium. Successful transformants were verified by PCR. Subsequently, the pGADT7-HbMYC2 plasmid was introduced into Y1HGold cells harboring pAbAi-Pro-HbMVK1, and transformants were selected on SD/-Ura medium supplemented with 600 ng/mL Aureobasidin A (AbA). For controls: (1) The pAbAi-p53 and pGADT7-53 co-transformants served as the positive control; (2) Two negative controls were included: pAbAi-Pro-HbMVK1 with empty pGADT7 vector, and empty pAbAi vector with pGADT7-HbMYC2.
Statistics and reproducibility
The statistical significance was determined using a two-sided Student’s t-test. Samples used for multi-omics data collection, physiological parameter measurements, and JA treatments were not subjected to any statistical method for predetermining sample size; all data were included in the analyses without exclusions. The dual luciferase reporter assay was performed with three independent biological replicates, while the Y1H assay was conducted with three technical replicates.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The genome assembly, centromere identification (ChIP-seq), and latex transcriptome data used in this study have been deposited in the National Genomics Data Center, under accession number PRJCA022207. The assembled genome and annotation file of CATAS 7-33-97 are available at Figshare (https://doi.org/10.6084/m9.figshare.26644396.v1). The complete metabolomics dataset has been deposited in the National Genomics Data Center, under accession number OMIX008934. Source data are provided with this paper.
References
Arias, M. & Van Dijk, P. J. What is natural rubber and why are we searching for new sources. Front. Young-. Minds 7, 100 (2019).
FOX, J. M. et al. Rubber Rubber plantations expand in mountainous Southeast Asia: What are the consequences for the environment? Asia Pacific Issues 114, 1-8 (2014).
Hurley, P. E. History of natural rubber. J. Macromol. Science—Chemistry 15, 1279–1287 (1981).
Priyadarshan PM. Genesis and Development. In: Biology of Hevea Rubber). Springer International Publishing (2017).
Junquera, V., Meyfroidt, P., Sun, Z. & Latthachack, P. Grêt-Regamey A. From global drivers to local land-use change: understanding the northern Laos rubber boom. Environ. Sci. Policy 109, 103–115 (2020).
Cheng H. HeveaDB: A Hub for Rubber Tree Genetic and Genomic Resources. In: The Rubber Tree Genome (eds Matsui M, Chow K-S). (Springer, 2020).
Wu, W. et al. Ultrahigh-density genetic map construction and identification of quantitative trait loci for growth in rubber tree (Hevea brasiliensis). Ind. Crops Prod. 178, 114560 (2022).
Chanroj, V., Rattanawong, R., Phumichai, T., Tangphatsornruang, S. & Ukoskit, K. Genome-wide association mapping of latex yield and girth in Amazonian accessions of Hevea brasiliensis grown in a suboptimal climate zone. Genomics 109, 475–484 (2017).
Cheng, H. et al. Chromosome-level wild Hevea brasiliensis genome provides new tools for genomic-assisted breeding and valuable loci to elevate rubber yield. Plant Biotechnol. J. 21, 1058–1072 (2023).
Francisco, F. R. et al. Unravelling rubber tree growth by integrating GWAS and biological network-based approaches. Front. Plant Sci. 12, 768589 (2021).
Degen B, Müller NA. A simulation study comparing advanced marker-assisted selection with genomic selection in tree breeding programs. G3 Genes|Genomes|Genetics 13, jkad164 (2023).
Grattapaglia D. Status and Perspectives of Genomic Selection in Forest Tree Breeding. In: Proc. Genomic Selection for Crop Improvement: New Molecular Breeding Strategies for Crop Improvement (eds Varshney RK, Roorkiwal M, Sorrells ME). (Springer, 2017).
Isik F. Genomic Prediction of Complex Traits Complex traits in Perennial Plants: A Case for Forest Trees Forest trees. In: Proc. Genomic prediction of complex traits: Methods and Protocols). (Springer, 2022).
Souza, L. et al. Genomic selection in rubber tree breeding: a comparison of models and methods for dealing with G× E. bioRxiv, 603662 (2019).
Pootakham W, Shearman JR, Tangphatsornruang S. Development of molecular markers in Hevea brasiliensis for marker-assisted breeding. The Rubber Tree Genome, 67-79 (Springer, 2020).
Chao, J. et al. Genomic insight into domestication of rubber tree. Nat. Commun. 14, 4651 (2023).
Rahman, A. Y. A. et al. Draft genome sequence of the rubber tree Hevea brasiliensis. BMC Genomics 14, 1–15 (2013).
Lau, N.-S. et al. The rubber tree genome shows expansion of gene family associated with rubber biosynthesis. Sci. Rep. 6, 28594 (2016).
Pootakham, W. et al. De novo hybrid assembly of the rubber tree genome reveals evidence of paleotetraploidy in Hevea species. Sci. Rep. 7, 41457 (2017).
Tang, C. et al. The rubber tree genome reveals new insights into rubber production and species adaptation. Nat. Plants 2, 1–10 (2016).
Liu, J. et al. The chromosome-based rubber tree genome provides new insights into spurge genome evolution and rubber biosynthesis. Mol. Plant 13, 336–350 (2020).
Sloan, D. B., Wu, Z. & Sharbrough, J. Correction of persistent errors in Arabidopsis reference mitochondrial genomes. Plant Cell 30, 525–527 (2018).
Li, Y. et al. High-quality de novo assembly of the Eucommia ulmoides haploid genome provides new insights into evolution and rubber biosynthesis. Horticult. Res. 7, 183 (2020).
Hu, W. et al. Allele-defined genome reveals biallelic differentiation during cassava evolution. Mol. Plant 14, 851–854 (2021).
Kawahara, Y. et al. Improvement of the Oryza sativa Nipponbare reference genome using next generation sequence and optical map data. Rice 6, 1–10 (2013).
Tuskan, G. A. et al. The genome of black cottonwood, Populus trichocarpa (Torr. & Gray). Science 313, 1596–1604 (2006).
Lu, J. et al. A chromosome-level genome assembly of wild castor provides new insights into its adaptive evolution in tropical desert. Genomics Proteom. Bioinforma. 20, 42–59 (2022).
Lin, T. et al. Genome analysis of Taraxacum kok-saghyz Rodin provides new insights into rubber biosynthesis. Natl. Sci. Rev. 5, 78–87 (2018).
Shi, X. et al. The complete reference genome for grapevine (Vitis vinifera L.) genetics and breeding. Horticult. Res. 10, uhad061 (2023).
Tang, C. et al. The sucrose transporter HbSUT3 plays an active role in sucrose loading to laticifer and rubber productivity in exploited trees of Hevea brasiliensis (para rubber tree). Plant Cell Environ. 33, 1708–1720 (2010).
Li, F., Wang, Y., Gao, H., Zhang, X. & Zhuang, N. Comparative transcriptome analysis reveals differential gene expression in sterile and fertile rubber tree varieties during flower bud differentiation. J. Plant Physiol. 265, 153506 (2021).
Wang, Y. et al. Transcriptomes analysis reveals novel insight into the molecular mechanisms of somatic embryogenesis in Hevea brasiliensis. BMC Genomics 22, 1–18 (2021).
Ohya, N., Tanaka, Y. & Koyama, T. Activity of rubber transferase and rubber particle size in Hevea latex. J. Rubber Res. 3, 214–221 (2000).
Men, X., Wang, F., Chen, G.-Q., Zhang, H.-B. & Xian, M. Biosynthesis of natural rubber: current state and perspectives. Int. J. Mol. Sci. 20, 50 (2018).
Yamashita, S. et al. Purification and characterization of small and large rubber particles from Hevea brasiliensis. Biosci. Biotechnol. Biochem. 82, 1011–1020 (2018).
Jacob, J. L. et al. Yield-limiting Factors, Latex Physiological Parameters, Latex Diagnosis, And Clonal Topology (CRC Press, 2018).
Tupý, J. Some aspects of sucrose transport and utilization in latex producing bark of Hevea brasiliensis Muel. Arg. Biol. Plant. 27, 51–64 (1985).
Sun, H. et al. Chromosome-scale and haplotype-resolved genome assembly of a tetraploid potato cultivar. Nat. Genet. 54, 342–348 (2022).
Han, X. et al. Two haplotype-resolved, gap-free genome assemblies for Actinidia latifolia and Actinidia chinensis shed light on the regulatory mechanisms of vitamin C and sucrose metabolism in kiwifruit. Mol. Plant 16, 452–470 (2023).
Hu, W. et al. Resequencing of 388 cassava accessions identifies valuable loci and selection for variation in heterozygosity. Genome Biol. 22, 1–23 (2021).
Zhou, Q. et al. Haplotype-resolved genome analyses of a heterozygous diploid potato. Nat. Genet. 52, 1018–1023 (2020).
Comai, L., Maheshwari, S. & Marimuthu, M. P. Plant centromeres. Curr. Opin. Plant Biol. 36, 158–167 (2017).
Melters, D. P. et al. Comparative analysis of tandem repeats from hundreds of species reveals unique insights into centromere evolution. Genome Biol. 14, 1–20 (2013).
Su, H. et al. Centromere satellite repeats have undergone rapid changes in polyploid wheat subgenomes. Plant Cell 31, 2035–2051 (2019).
Ma, H. et al. Centromere plasticity with evolutionary conservation and divergence uncovered by wheat 10+ genomes. Mol. Biol. Evol. 40, msad176 (2023).
Liu, Y. et al. Pan-centromere reveals widespread centromere repositioning of soybean genomes. Proc. Natl. Acad. Sci. 120, e2310177120 (2023).
Lv, Y. et al. A centromere map based on super pan-genome highlights the structure and function of rice centromeres. J. Integr. Plant Biol. 66, 196–207 (2024).
Gong, Z. et al. Repeatless and repeat-based centromeres in potato: implications for centromere evolution. Plant Cell 24, 3559–3574 (2012).
Long, X. et al. Characterization of a vacuolar sucrose transporter, HbSUT5, from Hevea brasiliensis: involvement in latex production through regulation of intracellular sucrose transport in the bark and laticifers. BMC Plant Biol. 19, 591 (2019).
Auzac, J. D., Chrestin, H., Marin, B. & Lioret, C. A plant vacuolar system: the lutoïds from Hevea brasiliensis latex. Physiologie Vegetale 20, 311–331 (1982).
D’Auzac, J., Prevot, J. C. & Jacob, J. L. What’s new about lutoids? A vacuolar system model from Hevea latex. Plant Physiol. Biochem. 33, 765–777 (1995).
Puskas, J. E., Gautriaud, E., Deffieux, A. & Kennedy, J. P. Natural rubber biosynthesis - a living carbocationic polymerization? Prog. Polym. Sci. 31, 533–548 (2008).
Kongsawadworakul, P. et al. The leaf, inner bark and latex cyanide potential of Hevea brasiliensis: evidence for involvement of cyanogenic glucosides in rubber yield. Phytochemistry 70, 730–739 (2009).
Howe, G. A. Jasmonates as signals in the wound response. J. Plant Growth Regul. 23, 223–237 (2004).
León, J. & Sánchez-Serrano, J. J. Wound signalling in plants. Jexpbot 52, 1 (2001).
Koryati, T. Aplication Of PGR IAA And Kinetin On Rubber Clones To Accelarate Mature Tapping. Int. J. Sci. Technol. Res 6, 96–101 (2017).
Yang, J. et al. The crosstalks between jasmonic acid and other plant hormone signaling highlight the involvement of jasmonic acid as a core component in plant response to biotic and abiotic stresses. Front. Plant Sci. 10, 1349 (2019).
Ku YS, Sintaha M, Cheung MY, Lam HM. Plant hormone signaling crosstalks between biotic and abiotic stress responses. Int. J. Mol. Sci. 19, 3206 (2018).
Lacote, R. et al. Long-term effect of ethylene stimulation on the yield of rubber trees is linked to latex cell biochemistry. Field Crops Res. 115, 94–98 (2010).
Zhu, J. & Zhang, Z. Ethylene stimulation of latex production in Hevea brasiliensis. Plant Signal. Behav. 4, 1072–1074 (2009).
Tungngoen, K. et al. Involvement of HbPIP2;1 and HbTIP1;1 aquaporins in ethylene stimulation of latex yield through regulation of water exchanges between inner liber and latex cells in hevea brasiliensis. Plant Physiol. 151, 843–856 (2009).
Loh, S. C., Othman, A. S. & Veera Singham, G. Identification and characterization of jasmonic acid- and linolenic acid-mediated transcriptional regulation of secondary laticifer differentiation in Hevea brasiliensis. Sci. Rep. 9, 14296 (2019).
Hao, B.-Z. Wu J-L. laticifer differentiation in hevea brasiliensis: induction by exogenous jasmonic acid and linolenic acid. Ann. Bot. 85, 37–43 (2000).
Zhu, JK. Abiotic stress signaling and responses in plants. Cell 167, 313–324 (2016).
Zhu, Z. & Lee, B. Friends or foes: new insights in jasmonate and ethylene co-actions. Plant Cell Physiol. 56, 414 (2015).
Pré, M., Atallah, M., Champion, A., Vos, M. D. & Memelink, J. The AP2/ERF domain transcription factor ORA59 integrates jasmonic acid and ethylene signals in plant defense. Plant Physiol. 147, 1347–1357 (2008).
Huang, L.J. et al. The AP2/ERF transcription factor ORA59 regulates ethylene-induced phytoalexin synthesis through modulation of an acyltransferase gene expression. J. Cell. Physiol. 239, e30935 (2024).
Wang, X. et al. Integrative genomics reveals the polygenic basis of seedlessness in grapevine. Curr. Biol. 34, 3763–3777 (2024).
Liu, Z. et al. Grapevine pangenome facilitates trait genetics and genomic breeding. Nat. Genet. 56, 2804–2814 (2024).
Cheng, H., Concepcion, G. T., Feng, X., Zhang, H. & Li, H. Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nat. Methods 18, 170–175 (2021).
Alonge, M. et al. Automated assembly scaffolding using RagTag elevates a new tomato system for high-throughput genome editing. Genome Biol 23, 258 (2022).
Durand, N. C. et al. Juicer provides a one-click system for analyzing loop-resolution Hi-C experiments. Cell Syst. 3, 95–98 (2016).
Dudchenko, O. et al. De novo assembly of the Aedes aegypti genome using Hi-C yields chromosome-length scaffolds. Science 356, 92–95 (2017).
Durand, N. C. et al. Juicebox provides a visualization system for Hi-C contact maps with unlimited zoom. Cell Syst. 3, 99–101 (2016).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Thorvaldsdóttir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinforma. 14, 178–192 (2013).
Benson, G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res. 27, 573–580 (1999).
Han, Y. & Wessler, S. R. MITE-Hunter: a program for discovering miniature inverted-repeat transposable elements from genomic sequences. Nucleic Acids Res. 38, e199–e199 (2010).
Flynn, J. M. et al. RepeatModeler2 for automated genomic discovery of transposable element families. Proc. Natl. Acad. Sci. USA 117, 9451–9457 (2020).
Chen N. Using repeat masker to identify repetitive elements in genomic sequences. Curr. Protoc. Bioinf. 5, 4.10. 11-14.10. 14 (2004).
Nawrocki, E. P. & Eddy, S. R. Infernal 1.1: 100-fold faster RNA homology searches. Bioinformatics 29, 2933–2935 (2013).
Kalvari, I. et al. Rfam 14: expanded coverage of metagenomic, viral and microRNA families. Nucleic Acids Res. 49, D192–D200 (2021).
Chan, P. P., Lin, B. Y., Mak, A. J. & Lowe, T. M. tRNAscan-SE 2.0: improved detection and functional classification of transfer RNA genes. Nucleic Acids Res. 49, 9077–9096 (2021).
Lagesen, K. et al. RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res. 35, 3100–3108 (2007).
Kanehisa, M. Goto S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Galperin, M. Y., Makarova, K. S., Wolf, Y. I. & Koonin, E. V. Expanded microbial genome coverage and improved protein family annotation in the COG database. Nucleic Acids Res. 43, D261–D269 (2015).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Boeckmann, B. et al. The SWISS-PROT protein knowledgebase and its supplement TrEMBL in 2003. Nucleic Acids Res. 31, 365–370 (2003).
Manni, M., Berkeley, M. R., Seppey, M., Simão, F. A. & Zdobnov, E. M. BUSCO update: novel and streamlined workflows along with broader and deeper phylogenetic coverage for scoring of eukaryotic, prokaryotic, and viral genomes. Mol. Biol. Evol. 38, 4647–4654 (2021).
Potter, S. C. et al. HMMER web server: 2018 update. Nucleic Acids Res. 46, W200–W204 (2018).
Sani, E., Herzyk, P., Perrella, G. & Colot, V. Hyperosmotic priming of Arabidopsis seedlings establishes a long-term somatic memory accompanied by specific changes of the epigenome. Genome Biol. 14, R59 (2013).
Hofstatter, P. G. et al. Repeat-based holocentromeres influence genome architecture and karyotype evolution. Cell 185, 3153–3168 (2022).
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009).
Bolger AM, Marc L, Bjoern U. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Li H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv e-prints (2013).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Zhang, Y., Liu, T., Meyer, C. A., Eeckhoute, J. & Johnson, D. S. Model-based analysis of ChIP-Seq (MACS). Methods Mol. Biol.1150, 81–95 (2008).
Goel, M., Sun, H., Jiao, W.-B. & Schneeberger, K. SyRI: finding genomic rearrangements and local sequence differences from whole-genome assemblies. Genome Biol. 20, 1–13 (2019).
Goel, M. & Schneeberger, K. plotsr: visualizing structural similarities and rearrangements between multiple genomes. Bioinformatics 38, 2922–2926 (2022).
He, W. et al. NGenomeSyn: an easy-to-use and flexible tool for publication-ready visualization of syntenic relationships across multiple genomes. Bioinformatics 39, btad121 (2023).
Emms, D. M. & Kelly, S. OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol. 20, 1–14 (2019).
Capella-Gutiérrez, S., Silla-Martínez, J. M. & Gabaldón, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Mendes, F. K., Vanderpool, D., Fulton, B. & Hahn, M. W. CAFE 5 models variation in evolutionary rates among gene families. Bioinformatics 36, 5516–5518 (2020).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinforma. 10, 1–9 (2009).
Marchler-Bauer, A. et al. CDD/SPARCLE: functional classification of proteins via subfamily domain architectures. Nucleic Acids Res. 45, D200–D203 (2017).
Bailey, T. L., Johnson, J., Grant, C. E. & Noble, W. S. The MEME suite. Nucleic Acids Res. 43, W39–W49 (2015).
Wang, Y. et al. MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic acids Res. 40, e49–e49 (2012).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Liao, Y., Smyth, G. K. & Shi, W. The R package Rsubread is easier, faster, cheaper and better for alignment and quantification of RNA sequencing reads. Nucleic Acids Res. 47, e47–e47 (2019).
Nie, Z. et al. Profiling ethylene-responsive genes expressed in the latex of the mature virgin rubber trees using cDNA microarray. PLoS One 11, e0152039 (2016).
Deng, X. et al. Jasmonate signalling in the regulation of rubber biosynthesis in laticifer cells of rubber tree, Hevea brasiliensis. J. Exp. Bot. 69, 3559–3571 (2018).
Patil, I. Visualizations with statistical details: The’ggstatsplot’approach. J. Open Source Softw. 6, 3167 (2021).
Wei, T. et al. Package ‘corrplot’. Statistician 56, e24 (2017).
Oksanen, J. et al. The vegan package. Community Ecol. Package 10, 719 (2007).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Kumar, L. & Futschik, M. E. Mfuzz: a software package for soft clustering of microarray data. Bioinformation 2, 5 (2007).
Chen, C. et al. TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 13, 1194–1202 (2020).
Szklarczyk, D. et al. The STRING database in 2017: quality-controlled protein–protein association networks, made broadly accessible. Nucleic Acids Res. 10, kw937 (2016).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Duan, Y. et al. Ustilaginoidea virens secreted effector UvSec117 hijacks OsWRKY31-OsAOC module to suppress jasmonic acid-mediated immunity in rice. Plant Biotechnol. J. 22, 3342–3344 (2024).
Acknowledgements
This research was financially supported by National Scientific Fund of China (No.32471914 to.H.C.), the Central Public-interest Scientific Institution Basal Research Fund for Chinese Academy of Tropical Agricultural Sciences (1630022023010 to H.C.), Chinese Academy of Tropical Agricultural Sciences for Science and Technology Innovation Team of National Tropical Agricultural Science Center (NO. CATASCXTD202401 to H.C.), Hainan Province Science and Technology Talent Innovation Project (KJRC2023C18 to H.C.), Hainan Province Science and Technology Special Fund (ZDYF2024XDNY233 to Z.Y.N.), the Project of National Key Laboratory for Tropical Crop Breeding (No. NKLTCBCXTD01 to H.C., NKLTCB202321 to Z.D.). We thank Dr. Huang Sanwen for his advice on the project design.
Author information
Authors and Affiliations
Contributions
H.C., and Y.F.Z. designed the project. C.C.L. performed a comparative genomic and evolutionary analysis of rubber-producing genes. Z.Y.W., J.X.Q., X.W., and S.C. assembled and annotated the genome. T.K.W has completed relevant interaction experiments. Z.Y.N., T.K.W., W.G.W., H.S.H., Z.W.A., Z.D., and D.X. collected the sample in this study. Y.Y., Z.X.J., X.B.W., and W.H were involved in OMICS analyses. B.Q.L participated in image layout and schematic drawing. C.C.L. and Y.Y. wrote the manuscript with contributions from H.C. and Y.F.Z.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Wen-Biao Jiao, Lingling Chen, Stephen Ryu and Massimiliano Corso for their contribution to the peer review of this work. [A peer review file is available].
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, C., Yuan, Y., Nie, Z. et al. The haplotype-resolved telomere-to-telomere genome and OMICS analyses reveal genetic responses to tapping in rubber tree. Nat Commun 16, 6255 (2025). https://doi.org/10.1038/s41467-025-61527-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-61527-1