Abstract
To what extent overdominance might contribute to the maintenance of genetic diversity within genomes is still an ongoing research question. Pseudo-overdominance created by the complementation of deleterious alleles in heterozygotes has recently become a subject of particular interest. Simulations and theory suggest that pseudo-overdominance may occur in low recombining regions. Here, we conduct a comprehensive investigation of large low-recombining (LLR) regions in cultivated populations of pearl millet, an outcrossing diploid African cereal. We examine seven large regions ranging from 5 to 88 Mb and six of them are pericentromeric. These LLR regions exhibit an excess of heterozygotes, a distinctive hallmark of overdominance. They display a tendency toward a higher diversity and a larger ratio of non-synonymous and deleterious variants. We conduct a more in-depth study of the largest 88 Mb region, identified on chromosome 3. Interestingly, haplotypes of this region have been introgressed from wild relatives. Using long read sequencing, we confirm their strong divergence and the presence of inversions across one of them. One of the haplotypes seems to be highly deleterious in the homozygous state. A total of 17% of the cultivated pearl millet genome exhibit a local population structure suggestive of overdominance or possibly pseudo-overdominance. Our empirical results contribute to the accumulation of knowledge, which will enhance our understanding of the potential role of overdominance or pseudo-overdominance in maintaining genetic diversity, particularly in low recombining regions.
Similar content being viewed by others
Introduction
Understanding how molecular polymorphism is maintained in populations is a longstanding question1,2 in evolutionary biology. Initially, selection was thought to be a major driving force1, with selective processes like overdominance favoring the maintenance of polymorphism in the population. Overdominance—a form of balancing selection3—is a selective mechanism whereby heterozygotes have higher fitness than both homozygotes at a given locus. Yet true overdominance is relatively uncommon4,5,6 and, as such, cannot therefore account alone for the genome-wide diversity patterns observed in populations. Although natural selection was initially considered to be a major driver in shaping diversity, theory and evidence suggest neutral processes have a considerable impact on the genetic diversity that prevails in any population2. Recently, there has been a renewed interest7,8,9,10 in another overdominance pattern, namely so-called pseudo-overdominance (POD)8.
POD is defined by Gilbert et al.10 as a ‘form of balancing selection that maintains complementary deleterious haplotypes, effectively masking the effects of recessive deleterious mutations’. Consequently, the complementation of deleterious alleles in repulsion results in a heterozygote advantage4,7,8. There is considerable confusion in the literature regarding the distinction between POD and associative overdominance11,12,13. True overdominant alleles and deleterious alleles involved in POD can lead through hitchhiking to associative overdominance (AOD)14,15, i.e., an apparent heterozygote advantage to nearby neutral loci10,12,16.
To take place, POD requires linkage disequilibrium (LD) between partially recessive deleterious alleles8,9. Strong LD can result from demographic factors, such as small population sizes, inbreeding17,18, as well as from genomic factors, such as low recombination rates. Low recombination rates can be expected in centromeric regions or when structural variants like inversions are present. Sturtevant and Mather proposed a mechanism by which POD might arise and posited the occurrence of an inversion, which allows accumulation of deleterious alleles in LD in two haplotypes19. Regions subject to POD should harbor an excess of heterozygotes compared to the rest of the genome8,9, and may contribute to higher-than-expected diversity across low-recombining regions10,20. This mechanism is particularly interesting, as genomic regions with a low recombination rate are generally expected to display lower neutral diversity due to linkage with loci under selection against deleterious (background selection21,22) or for beneficial (selective sweep23) variants10,20.
Recent simulations and theoretical studies have not allowed researchers to reach a consensus on the conditions required for the emergence of POD, its pervasiveness and the significance of its impact on variability8,9,10,12,24,25,26,27. Additionally, there is not yet sufficient empirical evidence to help answer these questions10,20,28. As a result, there is an urgent need to investigate POD patterns more frequently in genomic analyses, particularly since this phenomenon could be mistaken for other forms of balancing selection or could interfere with the detection of local adaptation10.
Here, we undertake a comprehensive study to identify large low-recombining (LLR) regions in a plant genome and to investigate whether true overdominance or POD processes might be at play in these regions. Pearl millet (Pennisetum glaucum (L.)) R. Br. syn. Cenchrus americanus (L.) Morrone, an allogamous diploid African cereal, has a genome size of 1.76 Gb with 7 metacentric chromosomes (2n = 14)29. We use local PCA30, a recent effective population genomics approach to detect LLR regions30,31,32,33 in Senegalese pearl millet landraces. We then characterize these regions’ diversity patterns, which include heterozygote excess, increased diversity and accumulation of deleterious mutations. Our findings demonstrate a potential interplay between overdominance (i.e., true and POD) and LLR regions. These regions cover at least 17% of the pearl millet genome. We conduct a thorough investigation of the largest region. By combining several datasets, we find that this region, which includes one apparently lethal haplotype, shows signatures of introgression from wild populations.
Results
Multiple divergent haplotypes in the Senegalese pearl millet population
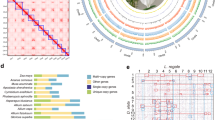
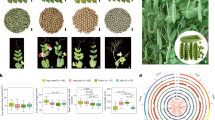
Pearl millet is a diploid allogamous cereal with high genetic diversity and a complex demographic history with gene flow from wild population34. We applied local PCA30, a population genomics approach that seeks outlier regions in terms of population structure compared to the rest of the genome30,32. We used a dataset, referred to as the Senegalese dataset, composed of 126 cultivated pearl millet accessions from Senegal genotyped by exome capture sequencing (n = 76,018 filtered SNPs). The population was structured into early- and late-flowering landraces, as previously reported35,36 (Supplementary Fig. 1). The local PCA analyses allowed us to identify seven regions ranging from 5.3 to 87.6 Mb (Table 1 and Supplementary Fig. 2). Six out of seven of the identified regions exhibit a population structure that deviated from the genome-wide pattern (early- vs late-flowering, Supplementary Fig. 1), where genetic diversity is organized into three to six distinct clusters. A population structure with three clusters distributed along the first PCA axis, with higher heterozygosity for the middle one, is expected when two non-recombining haplotypes are present in the population32 (Fig. 1a). We observed this pattern in the region identified on chromosomes 7 and 4 (Fig. 1b and Supplementary Fig. 2). When three non-recombining haplotypes: H1, H2 and the reference one, are present in the population, their combination results in six genotypic clusters37 (Fig. 1d: clusters RR, RH1, H1H1, RH2, H1H2 and H2H2). This pattern was observed in the candidate regions identified on chromosomes 2, 3 and 6 (Fig. 1e and Supplementary Fig. 2). Because no or low recombination is expected between haplotypes, the average heterozygosity within an individual is expected to be higher in heterokaryotic individuals32. We indeed observed significantly higher heterozygosity in heterokaryotic clusters (Fig. 1c, f, Supplementary Fig. 2 and Supplementary Data 1). The six regions with a clear local population structure deviating from that of the rest of the genome collectively spanned 307.9 Mb. All but one of these regions included the putative centromere (Table 1). These regions represented approximately 17% of the genome and encompassed 3163 genes (Supplementary Fig. 3). The number of genes across these regions ranged from 95 to 688 (Table 1, Supplementary Data 2, and Supplementary Fig. 3).
a Schematic representation of principal component analysis (PCA) expected pattern in the presence of two non-recombining haplotypes, the reference haplotype R and a single divergent alternate haplotype H1, in a population. The population structure is characterized by three clusters separated on the first PC axis. Each cluster aligns with one of the three possible genotypes: two homozygotes (RR and H1H1) and one heterozygote (RH1). b Results of a PCA analysis conducted in the LLR region identified on chromosome 7 from 129 to 178 Mb. Assignment of the accessions to the three clusters was performed with a kmean approach (red and blue circles represent likely homozygous genotypes, purple triangles likely heterozygous genotypes). c Observed heterozygosity pattern of the three clusters on chromosome 7. Each point displays mean individual heterozygosity with the confidence interval corresponding to ± 1.96 × the standard error of the mean (SEM). Sample sizes for clusters RR, RH1, H1H1: n = 59, 50, 17. d Schematic representation of PCA in the presence of a reference haplotype R and of two alternate haplotypes H1 and H2 in a population. The population structure is characterized by up to six clusters. Each cluster aligns with one of the six possible genotypes associated with the different haplotypes. Three clusters of homozygotes (clusters RR, H1H1 and H2H2) and three clusters of heterozygotes (RH1, RH2 and H1H2) are expected. e Results of a PCA analysis conducted on chromosome 2 from 131 to 187 Mb. The accessions were assigned to clusters using the kmean approach (dark, blue and red circles for the three likely homozygous genotypes, purple triangles for the three likely heterozygous genotypes). f Observed heterozygosity pattern of the six clusters on chromosome 2. Each point displays mean individual heterozygosity with the confidence interval corresponding to ± 1.96 × SEM. Sample sizes for clusters RR, RH1, H1H1, RH2, H1H2, H2H2: n = 17, 21, 10, 33, 24, 21.
Moreover, these regions harbored SNPs with higher linkage disequilibrium (LD) when studying the whole population, but a tendency to a lower LD when considering only the cluster of the most frequent homokaryotype (Supplementary Fig. 2). They also displayed lower estimated recombination rates compared to the rest of the genome (Supplementary Fig. 4).
Signatures of overdominance and pseudo-overdominance
We tested whether the candidate regions displayed genetic hallmarks that could be associated with true overdominance or POD8: heterozygote excess, accumulation of deleterious mutations, and maintenance of higher genetic diversity. We first assessed if those regions harbored higher heterozygote excess than expected from the average observed in the rest of the genome. The six regions with a clear local population structure pattern (i.e., all but the one on chromosome 1) displayed significantly lower FIS in comparison with the rest of the genome. Five of them were significantly negative, i.e., a signature of heterozygote excess compared to HWE expectations (Fig. 2, Table 1). Negative FIS is relatively uncommon and can notably be indicative of a selective heterozygote advantage38 (Supplementary Method 1, Supplementary Tables 1–3, and Supplementary Figs. 5, 6).
The mean FIS is represented by the dot, and the bar represents ± 1.96 × SEM confidence intervals. The dashed line indicates a zero FIS value. The FIS fixation coefficient is a metric of departure from Hardy-Weinberg equilibrium (HWE) departure. Values around zero meet a neutral expectation. Positive values indicate a heterozygote deficit, while negative values indicate a heterozygote excess compared to HWE expectations. Values were calculated for each region and for the whole genome, excluding these regions. The number of SNPs for each category is: all genome 76,018; Chr1 100–162 Mb 800, Chr2 131–187 Mb 800, Chr3 101–189 Mb 1000, Chr4 118–160 Mb 700, Chr6 133–201 Mb 800, Chr6 224–229 Mb 300, Chr7 129–178 Mb 700.
A second hallmark is an increase in diversity. However, five of the six candidate regions contain the centromeres, which are expected to exhibit low diversity39,40. Indeed, as in other species39,40,41,42,43,44,45, we observed a positive correlation between diversity (π, πS) and recombination (Supplementary Figs. 7c, 8c). Based on the diversity-recombination rate relationship, one would expect a significant decrease in diversity in the candidate regions. Contrary to this expectation, almost all regions showed no significant reduction neither in π or πS (Supplementary Table 4, Supplementary Figs. 7a, 8a), as if the negative effect of low recombination on diversity was compensated by overdominance or associative overdominance.
In an attempt to disentangle the effects of overdominance and low recombination, we also estimate π and πS for the most frequent homokaryotypes. For the six regions we could test, diversity values for π and πS for the most frequent homokaryotypes are reduced by almost 40% on average compared to when all karyotypes are considered (Supplementary Figs. 7b, 8b and Supplementary Table 5). Reductions were significant for four regions for π and three regions for πS. We verified that these lower diversity estimates were not simply a consequence of reduced sample size. Such an effect has been ruled out for almost all estimates, except for πS in candidate regions on chromosomes 2 and 6 (Supplementary Figs. 7e, 8e and Supplementary Table 6). Overall, these results suggested that π and πS were maintained in the regions despite low recombination39,40. Compared to the rest of the genome, we also found a tendency for higher values of πN that were significant for regions on chromosomes 1, 2 and 7 and almost significant on chromosome 3 (Supplementary Fig. 9a and Supplementary Table 4). The increase was not correlated with low recombination rates (Supplementary Fig. 9c, d). When only homokaryotes are considered, πN was reduced by 45% on average (Supplementary Fig. 9b and Supplementary Table 5). This reduction was significant for three regions, even though a small effect of sample size may occur as for πS (Supplementary Fig. 9e and Supplementary Table 6). A tendency for higher deleterious load in the LLR regions in terms of the πN /πS ratio was observed (Supplementary Fig. 10, Table 1 and Supplementary Table 7). When we used the pN/pS ratio, all six regions exhibited a higher ratio compared to the rest of the genome, with three of these regions showing statistically significant differences (Table 1). We further used SIFT46,47 and revealed a higher ratio of deleterious to tolerated mutations across the regions compared to that of the rest of the genome, with two of them being statistically significant (Table 1 and Supplementary Table 8). Overall, depending on the metric used, many of the regions showed the hallmark, suggesting possible true overdominance or POD in action.
In-depth study of the largest candidate region on chromosome 3
We delved deeper into the examination of the largest region on chromosome 3, spanning from 101 to 189 Mb (Table 1, Fig. 3 and Supplementary Fig. 2). Instead of the six expected clusters (Fig. 1d, and Supplementary Fig. 11c), this region was found to harbor only five distinct clusters (Fig. 3a and Supplementary Fig. 11a). The high heterozygosity noted for three of the clusters (Fig. 3b, and Supplementary Fig. 11b) suggested that one of the expected homozygous clusters (Fig. 1d and Supplementary Fig. 11c) was missing. We performed a PCAdapt analysis on the 1000 SNPs of the chromosome 3 candidate region on the Senegalese dataset to identify the SNPs that are responsible for the population structure. Visualizing genotypes for those 238 SNPs that had the most impact on the population structure enabled us to identify the haplotypes shaping the five clusters (Fig. 3c and Supplementary Fig. 11). The first divergent haplotype H1, spanned an area from around 120 to 175 Mb, and was found at both homozygous and heterozygous states in clusters (H1H1, RH1 and H1H2, Fig. 3c and Supplementary Fig. 11d). A second larger haplotype H2, spanning the whole 88 Mb region, was only found at heterozygous states (RH2 and H1H2, Fig. 3c and Supplementary Fig. 11d).
a Local PCA performed on 1000 SNPs from 101 to 189 Mb. The 126 accessions were assigned to five clusters using a kmeans approach: RR (blue), RH1 (purple), RH2 (brown), H1H2 (orange), and H1H1 (red). b Mean heterozygosity in the five clusters (RR, RH1, H1H1, RH2 and H1H2) and ± 1.96 × SEM. Sample sizes for clusters RR, RH1, H1H1, RH2, H1H2: n = 23, 53, 30, 8, 12. c Visualization of the genotypes for the 238 SNPs identified by PCAdapt analysis as the ones contributing the most to local population structure along the candidate region of chromosome 3. Positions of the 238 SNPs are displayed on the x-axis. On the y-axis, the 126 accessions are ordered in accordance with their respective cluster (RR, RH1, H1H1, RH2 and H1H2). The genotypes are colored in blue for the homozygous reference genotypes (0/0), red for the homozygous alternate genotypes (1/1) and purple for the heterozygous genotypes (0/1). Missing genotypes are indicated in gray. d Scaffolds of an individual from the cluster H1H2 (Autof-Pod103sr8) aligned to chromosome 3 of the reference genome. Three hybrid scaffolds (sorted as Super-Scaffold_100020, Super-Scaffold_100024, Super-Scaffold_100015) are aligned to the chromosome 3 region from 100 to 200 Mb. Green coloring corresponds to alignments with an identity above 50%, while orange corresponds to alignments with an identity of 25 to 50%. The two arrows indicate possible breakpoints of a main inversion from around 155 to 175 Mb.
We sequenced and assembled one heterozygous individual, Autof-Pod103sr8, belonging to cluster H1H2, using long-read sequencing and optical mapping (Supplementary Table 9). This assembled individual confirmed the presence of at least one large inversion on H1 with putative breakpoints at 155 and 175 Mb (Fig. 3d and Supplementary Fig. 12c). Other smaller inversions were present between 125 and 150 Mb (Fig. 3d and Supplementary Fig. 12c). The second haplotype H2 spans the entire candidate region from approximately 101 to 189 Mb and aligns with the reference genome with low sequence identity ranging from 25 to 50% across this region (orange dots in Supplementary Fig. 12c). The two largest scaffolds obtained from optical mapping showed good alignment to the reference genome at the beginning and end of the chromosome, but showed low congruence in the central region, approximately from 100 to 200 Mb (highlighted in yellow in Supplementary Figs. 12b, 13a), in contrast to an inbred line used as a control (Supplementary Fig. 13b). The low sequence identity observed in the central region of the chromosome, as indicated by both synteny plots and optical mapping, suggested substantial sequence divergence of H1 and H2 with the reference genome. This genetic divergence was also supported by FST estimates. All pairwise FST between clusters were significantly higher in the region than in the rest of the genome (Supplementary Fig. 14 and Supplementary Table 10). Notably, this was also the case between the two haplotypes H1 and H2 since FST between RH1-H1H2 and between H1H1-H1H2 were of the same order of magnitude. Although less pronounced, the same trend was observed when using DXY (Supplementary Fig. 15 and Supplementary Table 11).
Origin of the divergent haplotypes of chromosome 3
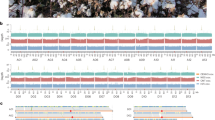
The divergence of the chromosome 3 haplotypes observed when comparing the optical maps and the scaffolds of Autof-Pod103sr8 with the reference genome (Supplementary Figs. 12, 13) as well as the genetic distances estimates (Supplementary Figs. 14, 15) led us to hypothesize that the variant may have been derived from introgression. Pearl millet domestication initially took place in the central Sahelian region and, following its geographical spreading, cultivated pearl millet underwent introgression from Western wild populations in Senegal34. We investigated the potential introgressed origin of the chromosome 3 haplotypes using an African dataset hosting data on 117 cultivated and 55 wild whole genomes distributed across Africa (Supplementary Fig. 16a). We first showed that the H1 haplotype was predominantly present, while the H2 haplotype was exclusively observed in cultivated pearl millet populations in Senegal (Supplementary Fig. 16b). Only Senegalese cultivated accessions from this African dataset were found within clusters RH2, H1H2 and H2H2 (Supplementary Fig. 16b: dark circles). Of the 238 SNPs identified by the PCAdapt analysis (Fig. 3), 158 SNPs were shared with the African dataset. Visualizing those SNP genotypes using accessions from Western and Central wild populations, we could see that H1 and H2 haplotypes in the cultivated Senegalese accessions were highly similar to those found in the wild Western population (Fig. 4a), suggesting that they might have been the result of introgression. We estimated fdM values48 to statistically assess this finding and found a signature supporting introgression between the wild Western population and the cultivated population (positive fdM values, Fig. 4b).
a Visualization of the SNP genotypes. Of the 238 SNPs depicted in Fig. 3, 158 SNPs were shared with the African dataset. Genotypes are shown in blue for the homozygous reference genotypes (0/0), red for the homozygous alternative genotypes (1/1) and purple for the heterozygous genotypes (0/1). Missing genotypes are shown in gray. Positions of the 158 SNPs are displayed on the x-axis. On the y-axis, 126 accessions are ordered as follows: clusters RR, RH1, H1H1, RH2 and H1H2 from top to bottom, as in Fig. 3, and the 47 wild accessions from Western Africa, then from Central Africa. b fdM values along chromosome 3. fdM, a statistic derived from ABBA-BABA tests, was calculated on non-overlapping 200 kb windows containing at least 200 SNPs. The fdM values ranged from −1 to 1. In a (((P1, P2) P3) Outgroup) topology, values around zero suggest no gene flow, positive values suggest gene flow between P3 and P2, while negative values indicate gene flow between P3 and P1. Here the analysis was performed using ((Cultivated_center, Cultivated_west), Wild_west, Outgroup), thus positive fdM values within the 101 to 189 Mb range indicate gene flow between wild and cultivated Western populations.
Segregation analysis of the divergent haplotypes of chromosome 3
A non-Mendelian segregation pattern was observed for two heterozygous SNPs in the progenies of three selfed individuals harboring the divergent chromosome 3 haplotype H2 found only in the heterozygous state (Supplementary Data 3, Fig. 5). The H2 homozygote genotypes expected at a frequency of 25%, were absent in two selfed progenies, and only found at 3% frequency in the third progeny. For these three populations, we also analyzed three other SNPs displaying strong linkage disequilibrium (r2 0.51 to 0.81, mean: 0.66, Supplementary Data 3) with the divergent haplotype outside the central region, and also observed the same non-Mendelian segregation pattern (Supplementary Data 3).
Segregation analysis of one SNP diagnostic marker of the chromosome 3 divergent haplotype H2 at the position 116,359,949 for three selfed-breeding parents (Pod 103-4, Pod 113-7 and Pod 74-3). Gray bars correspond to the expected frequencies of each genotype (Ref/Ref, Ref/Alt and Alt/Alt) under Mendelian segregation. Black bars are the observed genotype frequencies in the progeny. The homozygous alternate genotype was missing or only detected at very low frequency (<1%) in the progeny of each parent (p < 1.1.10−7 for Pod 103-4, p < 3.5.10−9 for Pod 113-7, p < 1.4.10−7 for Pod 74-3; these p-values are still significant with Bonferroni correction for three tests). Source data are provided as a Source Data file.
In contrast, we noted Mendelian segregation for four out of five SNPs in progenies from reference homozygote individuals, i.e., the control experiment (Supplementary Data 3). Experimental assessment of the recombination rate in progenies from heterozygote parents revealed lower recombination compared to that from homozygote parents (Supplementary Data 4). Overall, these results suggested that the H2 alternate haplotype behaved as highly deleterious at the homozygous state, which was in line with the absence of the H2H2 cluster observed earlier (Fig. 3a–c). With this extent of segregation bias, the H2 haplotype would likely disappear rapidly from the population.
Temporal evolution of the divergent haplotypes of chromosome 3
We used the temporal dataset of pearl millet landraces sampled 40 years apart, i.e., in 1976 and 2016 (Supplementary Fig. 17a) in Senegal, to assess how the frequencies of the different chromosome 3 haplotypes have evolved over time. A total of 377 accessions and 39,672 filtered SNPs were retained after filtering and used to perform a PCA analysis of the region from 101 to 189 Mb on chromosome 3. Over the 40-year sampling period, we found the same structure, with five clusters containing accessions from both 1976 and 2016 datasets (Supplementary Fig. 17a, b) and a sixth cluster represented by only three accessions (<1% of the population). This sixth cluster displayed high homozygosity and was only found in 2016 (Supplementary Fig. 17a, b). Visualization of the genotypes of SNPs confirmed that the sixth cluster contains the H2H2 homozygotes (Supplementary Fig. 17b, c). The number of accessions carrying the haplotype H2 (accessions of clusters RH2, H2H2 and H1H2) increased significantly between 1976 and 2016 (p-value of Fisher test = 0.003, Supplementary Table 12) whereas the number of accessions carrying the haplotype H1 (accessions of clusters RH1, H1H1 and H1H2) decreased (p-value of Fisher test = 0.038, Supplementary Table 12). At the cluster level, a significant decrease in the number of accessions was observed for cluster H1H1 (p-value: 0.032, Supplementary Table 12), whereas there was a trend toward an increase in cluster RH2 (p-value: 0.078, Supplementary Table 12). The maintenance of the haplotype H2 in heterozygotes, despite its very low frequency in the homozygous state, could be explained by a higher relative fitness of heterokaryotypes.
Association of the divergent haplotypes of chromosome 3 with phenotypic traits
We tested the association between the divergent haplotypes H1 and H2 of chromosome 3 and phenotypic traits. A total of 11 traits related to plant morphology and flowering time36 were assessed and obtained from three field experiments involving the 126 accessions of the Senegalese dataset. We found that the five clusters were significantly different with regard to fitness-related traits, including flowering time, total number of productive tillers, main panicle length, diameter and weight, and total seed weight (Supplementary Table 13). Heterozygote accessions with the 88 Mb divergent haplotype H2 displayed an earlier flowering time, and a greater number of productive tillers, but a smaller main panicle weight and diameter (Supplementary Table 13, clusters RH2 and H1H2). These phenotypes are characteristic of wild pearl millet and thus could be a phenotypic signature of introgression. However, GWAS analysis did not highlight any SNP associations with phenotypic traits across the chromosome 3 region (Supplementary Figs. 18, 19). Associations were only found between flowering time and SNPs at the beginning of chromosome 2 (Supplementary Figs. 18, 19), in accordance with the findings of a previous study36.
Discussion
With the increasing availability of high-quality genomes, there is mounting evidence suggesting that large structural variants such as inversions play a crucial role in the evolution of animals and plants49,50.
We employed a population genomics-based strategy, since approach of this type are known for their efficacy in detecting large low-recombining variants such as inversions30,32,33,49 or highly divergent sequences32,49. Among the seven regions flagged, six exhibited distinct patterns indicative of the presence of non-recombining haplotypes: a three to six clusters pattern, heightened heterozygosity, and increased LD across the entire set of individuals compared to a set of homozygotes only31,32,49. In our study, those genomic regions were large, ranging from 5 to 88 Mb, encompassing 17% of the genome. Considering the efficiency of genomics-based approaches and the growing availability of genomic data, we anticipate that an increasing number of studies will reveal the prevalence and diversity of LLR regions. We conducted a detailed analysis of the candidate region on chromosome 3. Our findings suggest that this region likely originated through introgression from a divergent wild relative population. The two haplotypes, H1 and H2, exhibit significant divergence from the reference genome and some evidence of a low recombination rate. The H1 haplotype carries an inversion and appears to have resulted from a double crossover event between 120 and 175 Mb, involving the reference haplotype and a divergent one. However, it remains unclear whether this latter haplotype corresponds to the H2 currently segregating in the population. FST values between H1 and H2 are high within the region, suggesting substantial genetic differentiation. Therefore, H1 and H2 might both originate from introgression events.
Low-recombining regions leading to a six clusters pattern have been observed across various taxa, including marine snails37,51, deer mice33, stick insects52 and zebra finches53. This pattern is anticipated when multiple rearrangements are present in the population. Our results highlight that haplotype diversity—featuring at least three non-recombining haplotypes—is more the rule than the exception, as evidenced by four out of the six genomic regions analyzed in this study. If the LLR involves an inversion, the emergence of multiple haplotypes may be further facilitated by the tendency of inversions to cluster in close proximity54. A notable example is observed in Heliconius, where consecutive nearby inversions resulted in the second inverted haplotype, responsible for a variety of wing morphotypes55,56.
Six of the candidate regions include the pericentromeric regions. Comparable patterns were observed in plants57 and in birds58, with regions notably conserved within genera. Centromeres, which are recognized as flexible and unstable regions59, typically exhibit lower gene density and are characterized by high repeat content54. In addition, substantial impacts of rearrangements such as inversions on centromere drive have been observed54. Notably, the pivotal role of inversions in centromere evolution has been extensively demonstrated across various clades, including primates60 and Fabaceae species such as Vigna61 and soybean62. Large structural variants associated with the centromere are likely to evade more effectively any negative selection processes, with a reduced probability of breakpoints exerting a negative impact or of the elimination of recombinant gametes63. In addition, due to the near absence of recombination in centromeric regions, large structural variants such as inversion could also accumulate by a purely neutral process.
Another common hallmark of our candidate regions was their negative FIS values, which were significantly pronounced in five of them. In empirical studies, positive FIS values are observed more frequently than negative FIS values38. Negative FIS values can suggest overdominance. The selective advantage in heterozygotes was confirmed for chromosome 3 candidate region, where the H2 haplotype has been maintained for 40 years, despite its apparently strong deleterious effect in the homozygous state. However, negative FIS values can also be observed in cases of asexuality, disassortative mating, or very small populations38. Given the biology of pearl millet, these hypotheses to explain negative FIS values seem less likely.
True overdominance is generally regarded as rare4,5,6. The conditions for the occurrence of POD in natural populations have been theoretically studied26,64,65. Distinguishing between true overdominance and POD remains a significant challenge4. Conventional approaches, such as GWAS, are often confounded by strong linkage disequilibrium within POD zones or the presence of large structural variants, making it difficult to pinpoint the beneficial alleles associated with true overdominance. Consequently, the absence of significant associations between SNPs and fitness-related traits on chromosome 3 does not necessarily exclude true overdominance.
In contrast to true overdominance, POD requires regions of strong linkage between deleterious loci. Hence, the role of inversions in POD was theoretically proposed from the outset19, but inversions are unlikely to be maintained by POD alone in random mating populations12. Centromeres were also later identified as potential contributors to POD due to their low recombination rates9. All candidate regions exhibited low recombination rates, with six of them overlapping centromeric regions. POD is characterized by the presence of recessive or partially recessive deleterious mutations in repulsion with each other, resulting in haplotypes carrying distinct complements of mutations that enhance diversity and heterozygosity. We observed significantly higher πN values in four regions, suggesting that POD could emerge or is already taking place. Non-recombining haplotypes tend to accumulate deleterious mutations due to reduced recombination in heterozygotes9, unless they capture strongly selected beneficial alleles33. Hard selective sweep will facilitate their rapid rise to high frequencies, thereby limiting the accumulation of deleterious mutations33. In our case, interpreting πN/πS ratios is particularly challenging, as a reduction in effective population size due to any cause will lead to an increase in πN/πS12. For certain regions, metrics such as pN/pS ratios or the deleterious-to-tolerated mutation ratio indicate a tendency toward the expected accumulation of deleterious mutations in non-recombining haplotypes. Notably, one of the two haplotypes H2 appears to be strongly deleterious at homozygous state. The accumulation of deleterious mutations may partially account for this apparent lethality27, but other phenomena leading to half-lethal systems have been observed, such as gene disruption created by rearrangements66.
While some necessary conditions for POD to emerge—such as low recombination and deleterious mutations in repulsion—are now well established in simulation studies9,10, many uncertainties remain. Notably, questions persist regarding how much genetic load must accumulate or which demographic scenarios favor the emergence of POD. Most studies have focused on the emergence of structural variants and genetic load within a single population10,12,24,26. Recent simulation studies9,27 explored scenarios in which POD emerge through introgression. The heterozygote advantage was found to be positively associated with the divergence between populations and the deleterious load of the recipient population, provided that the deleterious loads of the two haplotypes are not too dissimilar27. Under very restricted conditions, the two haplotypes are maintained over an extended period27. In such cases, one of the probable fates is the evolution toward a half-lethal system27. Some of these favorable conditions align with the scenario observed for the region on chromosome 3, which likely originated through introgression from a divergent wild relative population.
What are the evolutionary consequences of maintaining polymorphism of non-recombining haplotypes when POD is involved? POD could be viewed as an ‘evolutionary trap’, where the relative advantage of heterozygotes perpetuates haplotypes within populations despite their deleterious effects at homozygous state. Conversely, POD might act as an ‘evolutionary escape’ from the steady accumulation of genetic load in regions of low recombination, such as those near centromeres, by partitioning the load between haplotypes, ultimately fixing half of the deleterious mutations in each haplotype. Simulations suggest that states where both homokaryotypes are maintained are typically transient but can persist for thousands of generations9,27. Evolutionary outcomes such as half-lethal systems (where one homokaryotype is lost) or balanced-lethal systems (where arrangements persist only at the heterozygous state) may persist longer but occur only under specific conditions for the latter one. In this latter case, population fitness decreases. Overall, simulation studies indicate that POD alone is unlikely to maintain polymorphism over long evolutionary timescales9,24,27. However, many parameters remain unexplored, such as the number of migrants or introgression events, whether involving the same rearrangement or a novel one. More generally, we would argue that cultivated plant species may be prone to POD because: (1) cultivated species, in particular annual species, often undergo severe population bottlenecks67,68 leading to a high cultivated genetic load69 and (2) cultivated species show frequent hybridization in secondary contact zones propelled by human migrations70,71,72.
Our empirical study revealed a significant excess of heterozygotes in a substantial portion of the pearl millet genome, coinciding with regions characterized by: (1) reduced recombination and (2) elevated diversity of deleterious mutations. While true overdominance cannot be ruled out, all conditions necessary for POD to play a role were observed in these regions. We postulate that POD might be one of the primary mechanisms for maintaining diversity across these expansive non-recombining genomic regions. In concordance with recent simulations and theoretical inquiries9,10, we suggest that this phenomenon could be more widespread than previously recognized, with gene flow and hybridization potentially contributing to its prevalence. We advocate for a reevaluation, through the lens of POD, of patterns identified in recent genomic studies, anticipating that such patterns will become increasingly apparent with the increase of genomic data availability and the advancement of long-read sequencing techniques. More generally, we believe that future theoretical studies should delve deeper into the role of gene flow and divergence in the emergence of POD. A comprehensive understanding of selection processes involving POD, background selection21,22 and positive selection will enrich our comprehension of how diversity in neutral and selected regions of the genome is shaped.
Methods
Sampling, sequencing and SNPs detection
SNPs detection is fully described in Supplementary Method 2. We used a previously published whole exome sequencing dataset36, which we then referred to as the Senegalese dataset (Supplementary Data 5). It consisted of 130 pearl millet (Cenchrus americanus glaucum) individuals from early- (87 accessions) and late-flowering Senegalese landraces (43 accessions).
We studied temporal variations using an exome sequencing dataset of 384 accessions. This dataset was referred to as the temporal dataset (Supplementary Data 5). Two early-flowering Senegalese landraces were sampled 40 years apart, i.e., in 1976 and 2016. A total of 96 plants were sampled per year and per landrace. We designed a specific capture approach73 targeting 1900 genes, including 1043 randomly picked genes, and 857 genes with a potential selection signature34.
We also generated a full genome high-depth sequencing dataset referred to as the African dataset, composed of 172 individuals, including 117 cultivated (Cenchrus americanus glaucum) and 55 wild (Cenchrus americanus monodii) plants distributed across the African continent (Supplementary Data 5, Supplementary Fig. 16a), with 31 being cultivated accessions from Senegal. One additional cultivated sample was generated as a replicate to assess the SNP calling accuracy. A mean read depth of 30X was targeted per sample. A replicate was used to estimate the SNP detection accuracy and indicated that approximately 93.4% of the SNPs had been correctly called. Moreover, optical mapping and long-read sequencing data were generated for one of the accessions, Autof-Pod103sr8 (accession SAMN26040898, see “Genome assembly” section).
Identification of large low-recombining regions
Putative chromosomal rearrangements were detected in the Senegalese dataset, using a population genomic approach named local PCA30,31,32. The method aims to detect regions with a population structure that differs from that of the rest of the genome and which could be candidates for the presence of non-recombining haplotypes. The method uses the R package lostruct30 (v. 0.0.0.9000) and is implemented in an R pipeline (https://forge.ird.fr/diade/dynadiv/inversions_detection_code/-/tree/main/localPCA). Briefly, the genome was divided into non-overlapping genomic windows composed of 100 SNPs, and a PCA was performed on each window. The first two principal components of the PCA were used to calculate the distance between each pair of windows and to compute the distance matrix. This distance matrix was used to perform a multidimensional scaling (MDS) analysis with 40 dimensions and to display the dissimilarity between windows across the genome. Outlier windows were identified using a 5% false discovery rate. For each dimension, we only considered chromosomes with significantly higher outlier windows compared to the rest of the genome (Gtest function from R library DescTools, v. 0.99.49, threshold p-value 0.05) and with a minimum of four outlier windows. We then clustered outlier windows to define candidate regions using a correlation value of 0.631.
Structure and diversity analysis
PCAs were performed with SNPRelate74 (v. 1.28.0). Population structure analysis of the Senegalese cultivated dataset was performed with the R package LEA75 (v. 3.6.0) and the snmf() function. Cross-entropy was calculated with 1 to 10 K clusters. To obtain further evidence that the candidate regions might harbor non-recombining haplotypes, we first visualized the population structure by performing PCAs separately on each candidate region (SNPRelate74 v. 1.28.0). When the population was clearly structured, we assigned accessions to each cluster using a k-means clustering approach76 and the R function kmeans (). The number of k clusters in the population was chosen visually on the PCA plots. We conducted k-means clustering 100 times and selected the result with the lowest total within-cluster sum of squares. Then for each cluster we calculated the mean heterozygosity using vcftools (v. 0.1.17, --het parameter), by dividing the number of heterozygous sites by the total number of sites. We conducted Wilcoxon tests (wilcox.test () R function, v. 4.2.1) to statistically assess the difference in heterozygosity between clusters.
For each SNP, we computed the observed number of heterozygotes (Ho) and the corresponding expected number (He) under Hardy-Weinberg equilibrium using vcftools (v.0.1.17, --hardy option). We then calculated the FIS inbreeding coefficient for each SNP as follows: (He-Ho)/He. We calculated the mean FIS for each candidate region and compared it to the rest of the genome using Wilcoxon tests (wilcox.test () R function, v. 4.2.1). We also used t-test to compare the mean FIS for each candidate region with a value of 0, to assess departure from HWE expectation. We calculated the total number of genes within each candidate region using the annotation file of the new genome77 (ENA accession ERZ15184682). We plotted the number of genes along the chromosomes within non-overlapping genomic windows of 200 kb.
We assessed linkage disequilibrium between pairs of SNPs along each chromosome. The analysis was performed on the Senegalese dataset. We randomly picked SNPs using vcftools with the --thin parameter (v. 0.1.16, --thin 1000). We then used plink78 (v. 1.90, -- r2 parameter) to obtain intra-chromosomal r2 values for all pairs of SNPs and plotted them across the chromosomes. The analysis was performed for the whole population and for a cluster of the most frequent homokaryotypes.
We used the ReLERNN79 (v.1.0.0, default parameters) method to calculate the recombination parameter ρ = 4Nc, where N is the effective size and c is the recombination rate per nucleotide. ReLERNN79 generates simulations under the assumption of demographic equilibrium to train a recurrent neural network (RNN) that will later estimate the recombination landscape of the empirical dataset. This approach exhibits low sensitivity to demographic bias, and it demonstrates strong performance in fitting the esimates to inversions frequency. We assigned windows to each candidate region on the condition that at least half of the window covered the region. For each candidate region, we used Wilcoxon tests (wilcox.test () R function, v. 4.2.1) to assess if the mean recombination rate across the region was lower compared to the rest of the genome, excluding the candidate regions. The analysis was performed on the Senegalese dataset.
Study of deleterious mutations on the Senegalese dataset
SIFT46,47 (https://github.com/rvaser/sift4g) was used to predict the effect of the mutations on protein functions. We first created the SIFT database for the pearl millet reference genome (https://github.com/pauline-ng/SIFT4G_Create_Genomic_DB). SNPs were annotated using SIFT4G Annotator (https://github.com/pauline-ng/SIFT4G_Annotator) by assigning a SIFT score ranging from 0 (most deleterious) to 1 (most tolerated). We used a default threshold of 0.05, i.e., an SNP was annotated as deleterious if the SIFT score <0.05. We then calculated the ratio of deleterious to tolerated mutations for each candidate region and for the whole genome. We used G-tests (GTest() function from R library DescTools, v. 0.99.49) and a resampling procedure to assess if the ratio of deleterious to tolerated mutations of each candidate region was different from the rest of the whole genome. We computed the ratio for 1000 regions distributed randomly along the genome.
Diversity analysis of the candidate regions
Since diversity π is weighted by sequence length, we wanted to ensure that we were estimating the true sequenced length, i.e., the one for which we had sufficient sequencing depth and quality for polymorphic and non-polymorphic sites. Pixy handles missing data and uses invariant sites to calculate unbiased estimation of nucleotide diversity and divergence80. To obtain the invariant sites for the Senegalese dataset, we used the --include-non-variant-sites option of the module GenotypeGVCFs of gatk (v.4.2.0.0). We removed invariant sites with ‘LowQual’ annotation. We then applied similar filters as for the SNPs. We masked as missing data the genotypes with DP < 5 and DP > 100, and only kept the invariant sites with less than 50% of missing data. Diversity π was estimated using pixy80 (v. 1.2.10.beta2) within non-overlapping genomic windows of 10 Mb. The mean diversity π was then calculated for the whole genome and for each candidate region.
To estimate diversity at synonymous πS and non-synonymous πN sites, we used the script dNdSpiNpiS (v. 3 with the default parameters (https://kimura.univ-montp2.fr/PopPhyl/index.php?section=tools)). First, we generated a GFF file (ENA accession: ERZ24912374) using Liftoff 81 and the annotation files of the Tift 23D2B1-P1-P5 reference genome (https://doi.org/10.5524/100192). The -polish option was employed to re-align exons and ensure proper restoration of coding sequences. To extract phase information for the coding sequence (CDS) features, we used GenomeTools (gt gff3 -sort -tidy -retainids). Based on the VCF file, we then generate the diploid fasta sequence for each individual using the FastaAlternateReferenceMaker module from GATK82 (v. 4.2.3.0). We then intersect these fasta sequences with the GFF file generated to keep only CDS with SNPs, leading to a total of 6453 CDS. For each CDS, individuals with missing data were excluded when estimating πN and πS. As for π, these estimates should be corrected to account for truly invariant sites versus those resulting from insufficient sequencing depth. We used the invariant sites file generated in the previous section to estimate the expected percentage of bases called per CDS and applied this measure as a correction factor for the πN and πS estimates. The mean diversities πN and πS of the CDS were then calculated for the whole genome and for each candidate region. We used bootstraps to set 95% confidence intervals.
We used Wilcoxon tests or resampling procedures (1000 tests) to compare diversity estimates (π, πN, πS) between candidate regions and the rest of the genome. We additionally computed diversity estimates using only individuals from the most frequent homokaryotype for each candidate region identified with k-means clustering. The aim was to estimate the population diversity along pericentromeric and low-recombining regions without the effect of the presence of multiple haplotypes. For each candidate region, we used Wilcoxon tests to compare the mean estimate between the entire population and the most frequent homokaryotype. To evaluate the potential artifact of subsampling on the significance of these comparisons, we applied the same approach to the rest of the genome, excluding the candidate regions. We further assessed the correlation between the estimated recombination rate computed with ReLERNN79 and diversity estimates. To do so, we computed the nucleotide diversity estimates within genomic windows of the same size as imposed by ReLERNN79, from 15 Mb to 19 Mb, depending on the chromosomes.
In addition, πN/πS ratios were obtained by dividing the mean diversities πN and πS for the whole genome and for each candidate region. We compared πN/πS ratios between candidate regions and the rest of the genome excluding each candidate region using a resampling procedure with 1000 tests. We also computed pN and pS using the script dNdSpiNpiS (v. 3 with the default parameters, https://kimura.univ-montp2.fr/PopPhyl/index.php?section=tools). We computed the ratio pN/pS for the genome and for the candidate regions. We used a resampling procedure and 1000 tests to assess if the ratio pN/pS of each candidate region was different in comparison with the rest of the genome excluding the region tested.
In-depth study of the largest 88 Mb candidate region on chromosome 3
We conducted a PCAdapt83 analysis (pcadapt package, version 4.3.3) on SNPs within the candidate region on chromosome 3. We performed the analysis with the Senegalese dataset using k = 2. We applied a 5% false discovery rate (Benjamin–Hochberg method) and identified SNPs exhibiting the highest differentiation on the first two PCA axes. Genotypes for these SNPs were visualized across the candidate region with the accessions ordered according to clusters previously identified in the population.
We used pixy80 (v. 1.2.10.beta2) to estimate the statistics Dxy and FST84 between the clusters of accessions harboring the different haplotypes on chromosome 3. We computed the statistics Dxy and FST within non-overlapping genomic windows of 100 kb. We plotted Dxy and FST values of the windows along the chromosomes. We used Wilcoxon tests and a resampling procedure to compare the mean FST and the mean Dxy found within each haplotype with the rest of the genome and for all pairs of clusters of accessions. We used the region between 125 and 175 Mb for the clusters harboring only the first alternate haplotype, and the region from 101 to 189 Mb for the clusters harboring the second alternate haplotype.
Optical maps were generated for the Autof-Pod103sr8 heterozygote Senegalese cultivated sample from the E group (see “Results”). Optical maps were generated by the French Plant Genomic Resources Centre (CNRGV) of the French National Research Institute for Agriculture, Food and Environment (INRAE)77. De novo assembly was performed with the Bionano Solve pipeline85 (v. 3.5.1) using molecules longer than 150 kb and with more than nine labels. We compared the optical maps of Autof-Pod103sr8 to the pearl millet reference genome (GCA_947561735.1). The seven chromosomes were first converted into optical maps using the fa2cmap_multi_color.pl script of Bionano Solve85 (v3.3). We aligned the Autof-Pod103sr8 optical maps to the chromosomes using the runCharacterize.py script of Bionano Solve with RefAligner (v3.3, default parameters). Alignments were visualized with Bionano Access85,86 (v 3.7). As a control, we also aligned other optical maps of the PMiGAP257 (ENA accession ERZ14864807) Senegalese cultivated inbred accession.
We generated long reads based on Oxford Nanopore for the same individual Autof-Pod103sr8, which carried the variant on chromosome 3. High molecular weight DNA was extracted from isolated plant nuclei and used to prepare RAD004 and LSK109 libraries for Nanopore sequencing (https://www.protocols.io/view/high-molecular-weight-dna-extraction-from-plant-nu-83shyne)87. DNA was quantified by fluorometry (Qubit) and qualitatively assessed using pulsed field electrophoresis to ensure that the fragment sizes ranged from 40 to 150 kb. Libraries were constructed using the SQK-RAD004 Rapid Sequencing or SQK-LSK109 1D ligation genomic DNA kits (Oxford Nanopore Technologies) according to the supplier’s recommendations, with minor modifications: 4 µg of initial DNA without shearing, 7 min incubation for DNA repair and 13 min incubation for ligation for SQK-LSK109 library preparation, while only 1.5 µL of fragmentase was used versus 2.5 µL for RAD004 library preparation.
Base calling on ONT reads was performed with guppy (v. 6.0.6, dna_r9.4.1_450bps_hac.cfg model). SMARTdenovo88 (v. 1.0) and reads longer than 1 kb were used for the assembly. Three rounds of Medaka (v. 1.7, https://github.com/nanoporetech/medaka) were performed to polish and correct contigs with high-quality long reads (longer than 2 kb and with a quality score Q above 10, filtered with NanoFilt89 v. 1.0). Polishing with high-quality Illumina short reads (21X of coverage, accession SAMN26040898, bioproject PRJNA805042) was also performed using Hapo-G90 (v 1.3 default parameters). We used BUSCO91 (Benchmarking Universal Single-Copy Orthologs, v. 5.4.3) and the Poales dataset (odb10, 4896 genes) to assess the contig quality. We ordered and oriented the ONT contigs using the assembled optical maps and the hybrid scaffolding pipeline of Bionano Solve85 (v. 3.3, hybridScaffold.pl script with --B2 --N2 parameters). Scaffold alignments to chromosome 3 of the reference genome were visualized with D-genies92 (v. 1.4, enabling the ‘hide noises’ and ‘sorted contigs’ options and displaying alignments with more than 25% of identity). We performed annotation transfer to the scaffolds using Liftoff 81 (v 1.6.3, – copies – sc 0.50) using the annotation files of the previous Tift 23 D2B1-P1-P5 reference genome (https://doi.org/10.5524/100192)29. We calculated the total number of genes spanning the candidate region on chromosome 3.
To study changes in the frequency of the chromosome 3 haplotypes, the temporal dataset was used over a span of 40 generations. We first confirmed the characteristic clustering of the population on chromosome 3 in both 1976 and 2016 using PCAs (SNPRelate74 v. 1.28.0). We then identified SNPs with the highest differentiation using PCAdapt (k = 2), and visualized their genotypes. We applied a 5% false discovery rate (Benjamin–Hochberg method) and identified SNPs exhibiting the highest differentiation on the first two PCA axes. We examined whether the frequencies of the different haplotypes or clusters changed between 1976 and 2016 using a Fisher’s exact test with the R function fisher.test ().
We assessed segregation of individuals carrying the chromosome 3 haplotypes by selecting 10 SNPs distributed along the chromosome. These SNPs were chosen from the 1500 SNPs with the highest contribution to the second component axis according to Faye et al.36 and are referred to here as diagnostic SNPs of the chromosome 3 haplotypes. We selfed 130 accessions in a phenotyping experiment carried out in 2016. From this set, a random sample of 95 parental plants was genotyped using the 10 diagnostic SNPs. We only retained five SNPs that met two criteria, namely: (1) they had fewer than 20% missing data for genotyping; and (2) they were polymorphic within at least one of the progenies. Subsequently, we selected three parents among those heterozygous for the five diagnostic SNPs. The progeny of each of these three parents—totaling 94 individuals per progeny—was genotyped using the five diagnostic SNPs. In addition, we chose two homozygous-reference progenies for the diagnostic SNPs as a control. For these segregations, we designed a new set selected from among all SNPs, excluding the one identified by Faye et al.36 (see above). Given the expectation of greater homozygosity than observed in the previous set, the decision was taken to select 24 SNPs. They were distributed in regions and at intervals similar to those of the diagnostic SNPs. We applied the same filters as above, and one of the SNPs was disregarded due to its presence only in the heterozygous state, indicating a potential duplicated region. After filtering, the final SNP control set consisted of five SNPs. We conducted a Chi-square test to assess Mendelian segregation and estimated recombination rates based on offspring genotyping. We computed pairwise r2 values between each of the five filtered SNPs using plink78 (v. 1.90, -- r2 parameter) to obtain intra-chromosomal r2 values for all SNP pairs all along the chromosomes.
To investigate associations between the chromosome 3 haplotypes and phenotypic traits, we used 11 phenotypic trait measurements described in Faye et al.36 were used for the Senegalese cultivated dataset. Briefly, three field experiments were performed: one in Senegal in 2016 and two in Niger and Senegal in 2017. Three fully randomized repetitions were performed per trial, while 8 and 10 individuals per accession for each repetition were phenotyped in Niger and Senegal, respectively. A total of 11 traits associated with fitness were measured: date when half of flowers were in bloom (later referred to as flowering time); main panicle length; main panicle diameter; main panicle weight; main stem length; main stem diameter; total seed weight and 1000 seed weight of the main panicle; total number of tillers; and total number of productive and non-productive tillers. For each trait, the minimal and maximal measures were eliminated, and the mean trait value was calculated.
We tested the association of chromosome 3 haplotypes with each phenotypic trait. We first tested the relationship between each trait and the sample clusters identified across the candidate region. We used the R function lm () to fit a linear model and test the effects of the nine repetitions, the two flowering types and the clusters on each trait (formula: trait ~ repetitions + types + clusters). We then tested the association between the genotypes of each SNP and the phenotypic traits. Genotypes with missing values were imputed using the impute () function of the LEA R package75 (v. 3.8). We tested the effects of the nine repetitions using the lm () function (formula: trait ~ repetitions) based on the mean residual values for each trait and each sample. We used emma93 (v.1.1.2) and LFMM94 (v. 1.0) for association mapping between the residual values and genotypes. LFMM corrects for population structure while emma corrects for the genetic relatedness between the accessions. We used k = 2 for the population structure correction. We applied a Bonferroni correction to the p-value significance threshold to account for multiple testing.
To study the geographic distribution of the chromosome 3 haplotypes, we used the African dataset, which included accessions distributed across Africa, to study the geographic distribution of the chromosome 3 haplotypes. PCA (SNPRelate74, v. 1.28.0) was first performed with the full dataset. We intersected SNPs from the Senegalese and the African datasets. A local PCA was performed (SNPRelate74, v. 1.28.0) with SNPs shared between the two datasets across the candidate region on chromosome 3 for cultivated accessions.
We further investigate the introgressed origin of the chromosome 3 haplotypes. We used Illumina short reads of two Cenchrus pedicellatus accessions (accessions: srx4310736 and srx4310737) as an outgroup. The methods of identification of SNPs and genotypes are fully described in Supplementary Method 2. We then tested for introgression between wild and cultivated pearl millet populations using the fdM statistic based on the ABBA-BABA test48. For the ((P1, (P2, P3)), O) genealogy, where O is the outgroup species, fdM uses frequencies of the derived allele to estimate gene flow from P3 to P2 (positive values) or from P3 to P1 (negative values). We used sNMF95 (v 2.0) software implemented in the R package LEA75 (v 3.14) to verify the assignment to wild vs cultivated and geographical subgroups. We first ran the analyses on all accessions to distinguish wild vs cultivated and then only on the cultivated accessions. The procedure was as follows: 10 analyses with K ranging from 1 to 10, with 5 repetitions for each K and 5 million SNPs randomly sampled along the genome for each of the 10 analyses. We only used accessions that could be assigned to the following respective groups with a threshold above 0.7 and in line with the sampling geographic coordinates: WILD_West (27 accessions from Senegal-Mauritania); WILD_Center (22 accessions from Niger, Mali and Chad); CULT_West (27 accessions from Senegal); CULT_Center (20 accessions from Mali, Burkina Faso and Niger); and CULT_East_South (29 accessions from the rest of Africa). Then we calculated fdM values using the following topology ((CULT_Center, (CULT_West, WILD_West)), Out) along 200 kb non-overlapping windows using scripts from Martin et al.96.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The fastq files for the Senegalese, temporal and WGS datasets were deposited in GenBank under Bioproject PRJNA805042 and PRJNA771656. The VCF files and the scaffolds of Autof-Pod103sr8 that support the findings of this study are openly available in the DataSuds repository (IRD, France) [https://doi.org/10.23708/SN2T4A]. Long reads and optical mapping data of the Autof-Pod103sr8 genotype can be found under ENA accession study PRJEB57746. ONT long reads are accessible with ENA accession ERR12178246. Assembly of Autof-Pod103sr8 at the contig-level was deposited on ENA under accession CAVLDM02. The GFF file of the annotation transfer to the scaffolds of Autof-Pod103sr8 is accessible with ENA accession ERZ22147468. Bionano raw data and assembled optical maps for Autof-Pod103sr8 can be found under ENA accession ERZ21830311. Optical maps for the PMiGAP257 genotype are available under ENA accession ERZ14864807. Source data are provided with this paper and GitHub repository [https://github.com/msalson/low-recombining_regions_study]. Source data are provided with this paper.
Code availability
The code used for the analyses performed in the study can be found on GitHub [https://github.com/msalson/low-recombining_regions_study and https://github.com/stella-huynh/localPCA].
References
Fisher, R. A. The Genetical Theory of Natural Selection (Clarendon Press, 1930).
Kimura, M. The Neutral Theory of Molecular Evolution (Cambridge University Press, 1983).
Charlesworth, D. Balancing selection and its effects on sequences in nearby genome regions. PLoS Genet. 2, e64 (2006).
Charlesworth, D. & Willis, J. H. The genetics of inbreeding depression. Nat. Rev. Genet. 10, 783–796 (2009).
Hedrick, P. W. What is the evidence for heterozygote advantage selection? Trends Ecol. Evol. 27, 698–704 (2012).
Ruzicka, F. et al. A century of theories of balancing selection. Preprint at bioRxiv https://doi.org/10.1101/2025.02.12.637871 (2025).
Glemin, S. Pseudo-overdominance: how linkage and selection can interact and oppose to purging of deleterious mutations. PCI Evol. Biol. https://doi.org/10.1101/2021.12.16.473022 (2022).
Waller, D. M. Addressing Darwin’s dilemma: can pseudo-overdominance explain persistent inbreeding depression and load? Evolution 75, 779–793 (2021).
Abu-Awad, D. & Waller, D. Conditions for maintaining and eroding pseudo-overdominance and its contribution to inbreeding depression. Peer Commun. J. 3, e8 (2023).
Gilbert, K. J., Pouyet, F., Excoffier, L. & Peischl, S. Transition from background selection to associative overdominance promotes diversity in regions of low recombination. Curr. Biol. 30, 101–107.e3 (2020).
Durmaz, E., Kerdaffrec, E., Katsianis, G., Kapun, M. & Flatt, T. How selection acts on chromosomal inversions. eLS 1, 307–315 (2020).
Charlesworth, B. The fitness consequences of genetic divergence between polymorphic gene arrangements. Genetics 226, iyad218 (2024).
Ohta, T. & Kimura, M. Linkage disequilibrium at steady state determined by random genetic drift and recurrent mutation. Genetics 63, 229–238 (1969).
Frydenberg, O. Population studies of a lethal mutant in Drosophila melanogaster: I. Behaviour in populations with discrete generations. Hereditas 50, 89–116 (1963).
Ohta, T. Associative overdominance caused by linked detrimental mutations. Genet. Res. 18, 277–286 (1971).
Zhao, L. & Charlesworth, B. Resolving the conflict between associative overdominance and background selection. Genetics 203, 1315–1334 (2016).
Slatkin, M. Linkage disequilibrium—understanding the evolutionary past and mapping the medical future. Nat. Rev. Genet. 9, 477–485 (2008).
Golding, G. B. & Strobeck, C. Linkage disequilibrium in a finite population that is partially selfing. Genetics 94, 777–789 (1980).
Sturtevant, A. H. & Mather, K. The interrelations of inversions, heterosis and recombination. Am. Nat. 72, 447–452 (1938).
Becher, H., Jackson, B. C. & Charlesworth, B. Patterns of genetic variability in genomic regions with low rates of recombination. Curr. Biol. 30, 94–100.e3 (2020).
Charlesworth, B., Morgan, M. T. & Charlesworth, D. The effect of deleterious mutations on neutral molecular variation. Genetics 134, 1289–1303 (1993).
Charlesworth, D., Charlesworth, B. & Morgan, M. T. The pattern of neutral molecular variation under the background selection model. Genetics 141, 1619–1632 (1995).
Smith, J. M. & Haigh, J. The hitch-hiking effect of a favourable gene. Genet. Res. 23, 23–35 (1974).
Connallon, T. & Olito, C. Natural selection and the distribution of chromosomal inversion lengths. Mol. Ecol. 31, 3627–3641 (2022).
Sianta, S. A., Peischl, S., Moeller, D. A. & Brandvain, Y. The efficacy of selection may increase or decrease with selfing depending upon the recombination environment. Evolution 77, 394–408 (2023).
Berdan, E. L., Blanckaert, A., Butlin, R. K. & Bank, C. Deleterious mutation accumulation and the long-term fate of chromosomal inversions. PLoS Genet. 17, e1009411 (2021).
Berdan, E. L. et al. Mutation accumulation opposes polymorphism: supergenes and the curious case of balanced lethals. Philos. Trans. R. Soc. B 377, 20210199 (2022).
McMullen, M. D. et al. Genetic properties of the maize nested association mapping population. Science 325, 737–740 (2009).
Varshney, R. K. et al. Pearl millet genome sequence provides a resource to improve agronomic traits in arid environments. Nat. Biotechnol. 35, 969–976 (2017).
Li, H. & Ralph, P. Local PCA shows how the effect of population structure differs along the genome. Genetics 211, 289–304 (2019).
Huang, K., Andrew, R. L., Owens, G. L., Ostevik, K. L. & Rieseberg, L. H. Multiple chromosomal inversions contribute to adaptive divergence of a dune sunflower ecotype. Mol. Ecol. 29, 2535–2549 (2020).
Mérot, C. Making the most of population genomic data to understand the importance of chromosomal inversions for adaptation and speciation. Mol. Ecol. 29, 2513–2516 (2020).
Harringmeyer, O. S. & Hoekstra, H. E. Chromosomal inversion polymorphisms shape the genomic landscape of deer mice. Nat. Ecol. Evol.6, 1965–1979 (2022).
Burgarella, C. et al. A western Sahara centre of domestication inferred from pearl millet genomes. Nat. Ecol. Evol. 2, 1377–1380 (2018).
Oumar, I., Mariac, C., Pham, J.-L. & Vigouroux, Y. Phylogeny and origin of pearl millet (Pennisetum glaucum [L.] R. Br) as revealed by microsatellite loci. Theor. Appl. Genet. 117, 489–497 (2008).
Faye, A. et al. Genomic footprints of selection in early-and late-flowering pearl millet landraces. Front. Plant Sci. 13, 880631 (2022).
Faria, R. et al. Multiple chromosomal rearrangements in a hybrid zone between Littorina saxatilis ecotypes. Mol. Ecol. 28, 1375–1393 (2019).
Waples, R. S. Testing for Hardy–Weinberg proportions: have we lost the plot? J. Heredity 106, 1–19 (2015).
Burgarella, C. et al. Mating systems and recombination landscape strongly shape genetic diversity and selection in wheat relatives. Evol. Lett. 8, 866–880 (2024).
Chen, J., Glémin, S. & Lascoux, M. Genetic diversity and the efficacy of purifying selection across plant and animal species. Mol. Biol. Evol. 34, 1417–1428 (2017).
Hwang, H.-Y. & Wang, J. Effect of recombination on genetic diversity of Caenorhabditis elegans. Sci. Rep. 13, 16425 (2023).
Begun, D. J. & Aquadro, C. F. Levels of naturally occurring DNA polymorphism correlate with recombination rates in D. melanogaster. Nature 356, 519–520 (1992).
Roselius, K., Stephan, W. & Städler, T. The relationship of nucleotide polymorphism, recombination rate and selection in wild tomato species. Genetics 171, 753–763 (2005).
Nachman, M. W. Patterns of DNA variability at X-linked loci in Mus domesticus. Genetics 147, 1303–1316 (1997).
Lercher, M. J. & Hurst, L. D. Human SNP variability and mutation rate are higher in regions of high recombination. Trends Genet. 18, 337–340 (2002).
Ng, P. C. SIFT: predicting amino acid changes that affect protein function. Nucleic Acids Res. 31, 3812–3814 (2003).
Vaser, R., Adusumalli, S., Leng, S. N., Sikic, M. & Ng, P. C. SIFT missense predictions for genomes. Nat. Protoc. 11, 1–9 (2016).
Malinsky, M. et al. Genomic islands of speciation separate cichlid ecomorphs in an East African crater lake. Science 350, 1493–1498 (2015).
Huang, K. & Rieseberg, L. H. Frequency, origins, and evolutionary role of chromosomal inversions in plants. Front. Plant Sci. 11, 296 (2020).
Wellenreuther, M. & Bernatchez, L. Eco-evolutionary genomics of chromosomal inversions. Trends Ecol. Evol. 33, 427–440 (2018).
Le Moan, A. et al. An allozyme polymorphism is associated with a large chromosomal inversion in the marine snail Littorina fabalis. Evol. Appl. 16, 279–292 (2023).
Lindtke, D. et al. Long-term balancing selection on chromosomal variants associated with crypsis in a stick insect. Mol. Ecol. 26, 6189–6205 (2017).
Kim, K.-W. et al. A sex-linked supergene controls sperm morphology and swimming speed in a songbird. Nat. Ecol. Evol. 1, 1168–1176 (2017).
Ma, J., Wing, R. A., Bennetzen, J. L. & Jackson, S. A. Plant centromere organization: a dynamic structure with conserved functions. Trends Genet. 23, 134–139 (2007).
Jay, P. et al. Supergene evolution triggered by the introgression of a chromosomal inversion. Curr. Biol. 28, 1839–1845.e3 (2018).
Jay, P. & Joron, M. The double game of chromosomal inversions in a neotropical butterfly. C. R. Biol. 345, 57–73 (2022).
Wilkinson, M. J. et al. Centromeres are hotspots for chromosomal inversions and breeding traits in mango. N. Phytol. 245, 899–913 (2025).
Ishigohoka, J. et al. Distinct patterns of genetic variation at low-recombining genomic regions represent haplotype structure. Evolution 78, 1916–1935 (2024).
Barra, V. & Fachinetti, D. The dark side of centromeres: types, causes and consequences of structural abnormalities implicating centromeric DNA. Nat. Commun. 9, 4340 (2018).
Montefalcone, G., Tempesta, S., Rocchi, M. & Archidiacono, N. Centromere repositioning. Genome Res. 9, 1184–1188 (1999).
Dias, S. et al. Translocations and inversions: major chromosomal rearrangements during Vigna (Leguminosae) evolution. Theor. Appl. Genet. 137, 29 (2024).
Liu, Y. et al. Pan-centromere reveals widespread centromere repositioning of soybean genomes. Proc. Natl. Acad. Sci. USA 120, e2310177120 (2023).
Kirkpatrick, M. How and why chromosome inversions evolve. PLoS Biol. 8, e1000501 (2010).
Charlesworth, B. & Charlesworth, D. Rapid fixation of deleterious alleles can be caused by Muller’s ratchet. Genet. Res. 70, 63–73 (1997).
Pálsson, S. & Pamilo, P. The effects of deleterious mutations on linked, neutral variation in small populations. Genetics 153, 475–483 (1999).
Lamichhaney, S. et al. Structural genomic changes underlie alternative reproductive strategies in the ruff (Philomachus pugnax). Nat. Genet. 48, 84–88 (2016).
Glémin, S. & Bataillon, T. A comparative view of the evolution of grasses under domestication. N. Phytol. 183, 273–290 (2009).
Miller, A. J. & Gross, B. L. From forest to field: perennial fruit crop domestication. Am. J. Bot. 98, 1389–1414 (2011).
Liu, Q., Zhou, Y., Morrell, P. L. & Gaut, B. S. Deleterious variants in Asian rice and the potential cost of domestication. Mol. Biol. Evol. 34, 908–924 (2017).
Hufford, M. B. et al. The genomic signature of crop-wild introgression in maize. PLoS Genet. 9, e1003477 (2013).
Gonzalez-Segovia, E. et al. Characterization of introgression from the teosinte Zea mays ssp. mexicana to Mexican highland maize. PeerJ 7, e6815 (2019).
Janzen, G. M., Wang, L. & Hufford, M. B. The extent of adaptive wild introgression in crops. N. Phytol. 221, 1279–1288 (2019).
Mariac, C. et al. Optimization of capture protocols across species targeting up to 32000 genes and their extension to pooled DNA. Preprint at bioRxiv https://doi.org/10.1101/2022.01.10.474775 (2022).
Zheng, X. et al. A high-performance computing toolset for relatedness and principal component analysis of SNP data. Bioinformatics 28, 3326–3328 (2012).
Frichot, E. & François, O. LEA: an R package for landscape and ecological association studies. Methods Ecol. Evol.6, 925–929 (2015).
Hartigan, J. A. & Wong, M. A. Algorithm AS 136: a k-means clustering algorithm. Appl. Stat. 28, 100 (1979).
Salson, M. et al. An improved assembly of the pearl millet reference genome using Oxford Nanopore long reads and optical mapping. G3 13, jkad051 (2023).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. GigaScience 4, 7 (2015).
Adrion, J. R., Galloway, J. G. & Kern, A. D. Predicting the landscape of recombination using deep learning. Mol. Biol. Evol. 37, 1790–1808 (2020).
Korunes, K. L. & Samuk, K. pixy: unbiased estimation of nucleotide diversity and divergence in the presence of missing data. Mol. Ecol. Resour. 21, 1359–1368 (2021).
Shumate, A. & Salzberg, S. L. Liftoff: accurate mapping of gene annotations. Bioinformatics 37, 1639–1643 (2021).
McKenna, A. et al. The Genome Analysis Toolkit: a mapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Luu, K., Bazin, E. & Blum, M. G. B. pcadapt: an R package to perform genome scans for selection based on principal component analysis. Mol. Ecol. Resour. 17, 67–77 (2017).
Weir, B. S. & Cockerham, C. C. Estimating F-statistics for the analysis of population structure. Evolution 38, 1358 (1984).
Shelton, J. M. et al. Tools and pipelines for BioNano data: molecule assembly pipeline and FASTA super scaffolding tool. BMC Genomics 16, 734 (2015).
Yuan, Y., Chung, C. Y.-L. & Chan, T.-F. Advances in optical mapping for genomic research. Comput. Struct. Biotechnol. J. 18, 2051–2062 (2020).
Mariac, C. High molecular weight DNA extraction from plant nuclei isolation optimised for long-read sequencing V1. protocols.io https://www.protocols.io/view/high-molecular-weight-dna-extraction-from-plant-nu-83shyne (2019).
Liu, H., Wu, S., Li, A. & Ruan, J. SMARTdenovo: a de novo assembler using long noisy reads. Gigabyte 2021, 1–9 (2021).
De Coster, W., D’Hert, S., Schultz, D. T., Cruts, M. & Van Broeckhoven, C. NanoPack: visualizing and processing long-read sequencing data. Bioinformatics 34, 2666–2669 (2018).
Aury, J.-M. & Istace, B. Hapo-G, haplotype-aware polishing of genome assemblies with accurate reads. NAR Genomics Bioinform. 3, lqab034 (2021).
Manni, M., Berkeley, M. R., Seppey, M. & Zdobnov, E. M. BUSCO: assessing genomic data quality and beyond. Curr. Protoc. 1, e323 (2021).
Cabanettes, F. & Klopp, C. D-GENIES: dot plot large genomes in an interactive, efficient and simple way. PeerJ 6, e4958 (2018).
Kang, H. M. et al. Efficient control of population structure in model organism association mapping. Genetics 178, 1709–1723 (2008).
Frichot, E., Schoville, S. D., Bouchard, G. & François, O. Testing for associations between Loci and environmental gradients using latent factor mixed models. Mol. Biol. Evol. 30, 1687–1699 (2013).
Frichot, E., Mathieu, F., Trouillon, T., Bouchard, G. & François, O. Fast and efficient estimation of individual ancestry coefficients. Genetics 196, 973–983 (2014).
Martin, S. H., Davey, J. W. & Jiggins, C. D. Evaluating the use of ABBA–BABA statistics to locate introgressed loci. Mol. Biol. Evol. 32, 244–257 (2015).
Acknowledgements
We are grateful to all farmers and local fieldwork teams who helped with sample collection in this study. We would also like to thank D. Manley and J. Tregear for English proofreading. We acknowledge the ISO 9001-certified IRD i-Trop HPC (South Green Platform) at IRD Montpellier for providing HPC resources that have contributed to the research results reported within this paper. We are also greatly thankful for reviewers’ insights, which helped to improve the manuscript. This study was supported by a French Agence National de la Recherche (ANR) grant to C.B.-S. (ANR-19-CE02-00006-1; PEMILADAPT project), as well as an ANR grant to Y.V. (ANR-19-CE32-0009; DOMISOL project). Grants from the Marcus Borgströms Foundation and Nilsson–Ehle Foundation were awarded to C.B. M.S. received a PhD scholarship from the French government. C.B. received funding from the European Union Horizon 2020 research and innovation funding program under the Marie Skłodowska-Curie Fellowship Programme (grant no. 839643).
Author information
Authors and Affiliations
Contributions
C.B.-S. and Y.V. designed the study. M.S., M.D., S.H., C.M., J.O., S.A., N.R., P.C., C.B.-S. and Y.V. carried out the analyses. M.D. and S.H. contributed equally and shared second place in the co-authorship rankings. C.B., C.M., L.Z., M.C., S.A., N.R., A.B., A.F., N.K.; C.B.-S. and Y.V. contributed to sample collection and preparation. M.S., C.M., C.T.-D., J.O., P.C., A.-C.T., C.B., M.N., Y.V., C.B.-S. contributed to the interpretation of the results. M.S., C.B.-S. and Y.V. took the lead in writing the manuscript. All authors provided critical feedback, discussed the results and contributed to the final manuscript. C.B.-S. and Y.V. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Brian Charlesworth, Donald Waller and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s Note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Salson, M., Duranton, M., Huynh, S. et al. Interplay between large low-recombining regions and pseudo-overdominance in a plant genome. Nat Commun 16, 6458 (2025). https://doi.org/10.1038/s41467-025-61529-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-61529-z