Abstract
Understanding the phylogenetic relationships among species is crucial for comprehending major evolutionary transitions. Despite the ever-growing volume of sequence data, constructing reliable phylogenetic trees effectively becomes more challenging for current analytical methods. In this study, we introduce a new solution to accelerate the integration of novel taxa into an existing phylogenetic tree using a pretrained DNA language model. Our approach identifies the taxonomic unit of a newly collected sequence using existing taxonomic classification systems and updates the corresponding subtree. Specifically, we leverage a pretrained BERT network to obtain high-dimensional sequence representations, which are used not only to determine the subtree to be updated, but also identify potentially valuable regions for subtree construction. We demonstrate the effectiveness of our method, named PhyloTune, through experiments on simulated datasets, as well as our curated Plant (focusing on Embryophyta) and microbial (focusing on Bordetella genus) datasets. Our findings provide evidence that phylogenetic trees can be constructed by automatically selecting the most informative regions of sequences, without manual selection of molecular markers. This discovery offers a guide for further research into the functional aspects of different regions of DNA sequences, enriching our understanding of biology.
Similar content being viewed by others
Introduction
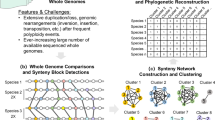
Phylogenetic trees serve as fundamental pillars in biological research, elucidating evolutionary relationships among organisms and offering profound insights into their shared history1,2,3. In addition, they play a pivotal role, for instance, in pinpointing distinct species communities to refine conservation strategies4,5,6, in deciphering virus origins7,8,9, or even in elucidating cancer progression and guiding therapies10,11,12,13. For nearly two decades, molecular phylogenies have primarily relied on data from a few genes obtained through PCR amplification and Sanger sequencing. However, advancements in sequencing technologies have resulted in large-scale datasets containing orders of magnitude more genes, posing challenges to accurately reconstructing the tree of life14,15. On the one hand, the exponential growth in the quantity of genetic data intensifies computational and storage burdens, leading to substantial time constraints and a super-exponential rise in the demand for computational and storage resources. On the other hand, the longer sequences may not only challenge the ability of computational resources but also contain inconsistencies or noise, leading to misleading or less precise results16,17.
Traditional phylogenetic tree construction methods are typically divided into two categories: distance-based and character-based15,18. In the context of species tree inference, distance methods involve calculating genetic distances between species pairs and using resulting matrices to build trees19,20,21. Character-based methods compare all DNA sequences in an alignment simultaneously, considering one site at a time to calculate scores for each tree. These methods include maximum parsimony, maximum likelihood, and Bayesian inference. However, identifying the tree with the highest score requires comparing all possible trees, which is computationally infeasible due to the NP-hard nature of tree construction22,23. To mitigate computational burdens, various heuristic tree search methods focusing on accelerating and parallelizing calculations have been developed, such as FastTree24, PhyloBayes MPI25, ExaBayes26, and RAxML-NG27. These heuristic tree search methods are not guaranteed to find the best tree, but make it feasible to analyze large data sets18. Nonetheless, there is considerable room for improving computational efficiency while maintaining accuracy, making the balance between the two a challenging and pressing issue.
Recent advances in deep learning offer promising opportunities for phylogenetic inference, which can be broadly categorized into classification-based and distance-based methods. Classification-based methods treat phylogenetic inference as a classification problem, aiming to train neural networks to predict the topology of sequences28,29. Distance-based methods utilize neural networks to improve distance representation, addressing challenges such as phylogenetic placement problems and data imputation problems in incomplete distance matrices30,31,32,33. However, applying deep learning to phylogenetic inference is still in its infancy34. Classification-based methods struggle with scalability and cannot infer branch lengths. Moreover, training deep learning models requires large datasets, but benchmark data with known true phylogenies are rare. As a result, almost all studies use simulated data for training, which may not accurately reflect the complexity and diversity of real biological systems, limiting the generalization performance of models on empirical data35. In addition, existing applications are predominantly based on convolutional neural network (CNN) structures, which may not always outperform traditional tree construction methods36. As large language models (LLMs) have revolutionized natural language understanding, their application to genome modeling problems has also increased37,38,39,40,41. Due to the similarities between DNA sequences and natural languages, genomic LLMs built on the Transformer architecture with a self-attention mechanism can skillfully model genomic information by capturing long-range dependencies. Such models have recently achieved unprecedented success across various downstream tasks, as exemplified by the generation of DNA sequences39, or the prediction of promoters37, and chromatin profiles41. Despite these advances, gaps remain in applying LLMs to phylogenetic inference.
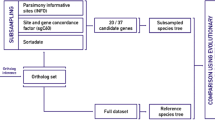
In this work, we propose PhyloTune, a method designed to accelerate phylogenetic updates by using pretrained DNA language models. In contrast to standard pipelines that align and analyze all sequences simultaneously (e.g., BuddySuite42 and MEGA43), our pipeline reduces the number and length of input sequences by identifying the smallest taxonomic unit of a new sequence within a given phylogenetic tree and extracting its high-attention regions (Fig. 1a). Specifically, we fine-tune the pretrained DNA large model (e.g., DNABERT37,44) using the taxonomic hierarchy information of the target phylogenetic tree to achieve precise precise taxonomic unit identification and high-attention region extraction (Fig. 1b). This targeted subtree construction obviates the need for reconstructing the entire tree from full-length sequences. We demonstrate the effectiveness of PhyloTune through experiments on simulated datasets and our curated datasets, including the Plant dataset focusing on Embryophyta and the microbial dataset from the Bordetella genus (see “Methods” for details). Results show that PhyloTune enables efficient phylogenetic updating through the smallest taxonomic unit identification and high-attention region extraction, with only a modest trade-off in accuracy. Moreover, attention-guided sequence regions may offer a scalable and interpretable alternative for phylogenetic analysis.
a Compared to the standard pipeline, PhyloTune introduces an innovative framework tailored to constrain updates within a specified subtree. By precisely identifying potentially informative regions within the subtree sequences, PhyloTune reduces the number and length of input sequences for alignment (e.g., MAFFT) and tree inference tools (e.g., RAxML), thereby improving tree update efficiency. b Overview of PhyloTune. For a given phylogenetic tree requiring updates, hierarchical linear probes (HLPs) are specifically designed to align with its taxonomic hierarchy. These probes are fine-tuned on a pre-trained DNA model to accurately classify query sequences at the smallest taxonomic unit within the specified tree while extracting high-attention regions from all sequences within the corresponding clade. c The PhyloTune model architecture tailored for the Plant dataset. It integrates a Transformer-based BERT module inherited from DNABERT and incorporates HLPs covering four taxonomic ranks: class, order, family, and genus.
Results
Overview of the PhyloTune method
PhyloTune enhances the efficiency of phylogenetic updates by identifying the taxonomic unit of a new sequence and extracting potentially valuable regions for the subtree sequences. Since taxonomic classifications reflect shared ancestry45,46, we leverage this principle to fine-tune a pretrained DNA language model based on the taxonomic hierarchy of the phylogenetic tree being updated, enabling precise inference of the smallest taxonomic unit for new sequences. Furthermore, given that transformer attention can capture biological signals37,38, we use attention scores to identify potentially valuable regions in sequences (hereafter referred to as high-attention regions). Subsequently, existing tools such as MAFFT for sequence alignment and RAxML for tree inference are then used to update the topology of the subtrees based on the extracted high-attention regions, allowing for more targeted and efficient updates of phylogenetic trees.
Smallest taxonomic unit
Identifying the smallest taxonomic unit of a new sequence involves two key tasks: novelty detection and taxonomic classification. Given a phylogenetic tree, a new sequence may belong to an unknown taxon at the genus rank but align with a known taxon at a higher rank. Therefore, identifying the smallest taxonomic unit requires first determining the lowest rank at which the sequence can be classified into a known taxon. Once this rank is established, taxonomic classification is performed to assign the sequence to the corresponding taxon. The two tasks are simultaneously addressed by utilizing pretrained DNA large language models (LLMs) to train a hierarchical linear probe (HLP) for each taxonomic rank of the phylogenetic tree to be updated (Fig. 1c). These probes learn classification boundaries specific to each rank to better identify out-of-distribution (OOD) sequences and classify in-distribution (ID) sequences. Notably, PhyloTune is the method to combine novelty detection and taxonomic classification for identifying the smallest taxonomic units. Traditional methods, such as identifying the most similar sequences in a reference database (BLAST47 and MMseqs248) or predicting taxonomic origins of k-mer queries (Kraken249), fail to ensure consistency across all taxonomic levels between the identified and query sequences.
High-attention regions
Recognizing high-attention regions of sequences in a subtree involves dividing all sequences equally into K regions and using the attention weight from the last layer of the transformer model to score these regions. The attention weight denotes how much focus each nucleotide should receive from other nucleotides within the same sequence. The weights at the last layer highlight the nucleotide most crucial for the downstream tasks, revealing key nucleotides or regions impacting functionality or classification tasks. These attention weights are iteratively optimized during training to generate gene embeddings that best provide the correct answer for the taxonomic unit of a sequence. Since the regions with high scores may differ across sequences, we use a voting method, i.e., minority-majority approach, to identify the top M (< K) regions with the highest scores as potentially valuable regions for tree construction. The settings of K and M can be set with reference to the attention scores across the sequence.
PhyloTune demonstrates the ability to rapidly update phylogenetic trees
To evaluate the feasibility of updating existing trees through targeted subtree reconstruction, we computed the normalized Robinson-Foulds (RF) distance and computational time between the updated trees obtained by reconstructing only subtrees and the complete trees built using all sequences (Fig. 2a for the schematic diagram). Through repeated experiments with five non-overlapping subtrees randomly selected from simulated datasets, our results demonstrate that for smaller datasets (with n representing the number of sequences used in ground-truth tree, n = 20, 40), the updated trees exhibit identical topologies to the complete trees (Fig. 2b). While minor discrepancies emerged with increasing sequence counts (n = 60, 80, 100), indicated by average RF distances for full-length trees of 0.007, 0.046, and 0.027, and for high-attention region trees of 0.021, 0.054, and 0.031. Notably, even complete trees reconstructed from complete sequences sets show non-trivial discrepancies from the ground truth in more complex topologies (e.g., RF = 0.038 and 0.020 for n = 60 and 80, respectively), which is consistent with known challenges in reconstructing complex topologies50,51.
a Schematic overview of tree reconstruction strategies. The original tree is built from sequences simulated on the ground-truth tree (gt) with one sequence removed as the new sequence. Updates are performed using: all sequences (complete tree), full-length sequences of a target subtree (full-length tree), or high-attention regions of the subtree (high-attention region tree). Using the example of the addition of species 4, the updated parts of the three trees using RAxML are highlighted in blue. b, c Robinson-Foulds (RF) distance and construction time compare updated trees to gt. Each box plot (n=5 independent experiments) shows the median, interquartile range (25th to 75th percentile), and whiskers to minima/maxima within 1.5 times IQR. d Example of updating the phylogenetic tree using PhyloTune. The original tree consisted of 677 species of 20 orders from Embryophyta. The tree was built using RAxML, with organisms colored based on order. The scale represents the normalized fraction of total branch length. The rugged bars at the outer circle represent the normalized length of input DNA sequences. (i) Update of out-of-distribution (OOD) sequences: the three newly added sequences belong to the order Fabales, but do not belong to any families or genera in the original tree, so only the subtree of Fabales is updated. (ii) Update of in-distribution (ID) sequences: the two newly added sequences belong to the genus Primulina, so the subtree of Primulina is updated. e Time comparison for the example tree. Blue and orange curves show subtree reconstruction times using full-length sequences and high-attention regions (one-third of the full length), respectively. The red dotted line indicates the time needed to update the tree using all sequences (about 20.1 h). Source data are provided as a Source Data file.
Importantly, our subtree update strategy significantly reduced computational cost: update time was relatively insensitive to total sequence numbers, in stark contrast to the exponential growth seen with complete tree reconstruction (Fig. 2c). Specifically, high-attention regions further reduced computational time by 14.3% to 30.3% compared to full-length sequences. While RF distances from high-attention regions were slightly higher (average differences of 0.004 to 0.014), indicating a modest trade-off in topological accuracy, this approach offers substantial efficiency gains. Despite potential limitations in capturing all global topological changes, subtree reconstruction remains a widely used strategy to balance computational efficiency and accuracy, as demonstrated by methods such as pplacerDC52 and SCAMPP53. Moreover, the well-known APG phylogeny of angiosperms54 is also constructed iteratively by connecting subtrees. Given that phylogenetic inference is inherently constrained by data availability and computational limitations, achieving an absolutely correct tree is challenging. Nonetheless, updating existing trees via subtree reconstruction offers a practical trade-off between efficiency and accuracy compared to de novo complete-tree reconstruction.
To further illustrate the process of updating a phylogenetic tree using PhyloTune, Fig. 2d presents a simplified example. The original tree comprises 677 species of 20 orders from Embryophyta. The sequences of these species were selected from the Plant dataset. We concatenated all molecular markers of each species for tree construction, with the total lengths ranging from 2, 493 bp to 7, 705 bp. When a new sequence is added, PhyloTune, fine-tuned on the Plant dataset, identifies its smallest taxonomic unit within the original tree and extracts high-attention regions for all sequences in the corresponding clade. We then utilized MAFFT55 and RAxML27 to perform alignment and subtree inference with only high-attention regions (Fig. 1a).
The smallest taxonomic unit for the new sequence is jointly determined by the novelty detection scores and taxonomic classification probabilities for four taxonomic ranks (class, order, family, and genus) as output by PhyloTune (Fig. 1c). If the sequence has a higher novelty detection score at any rank, it is classified as an out-of-distribution (OOD) sequence; otherwise, it is an in-distribution (ID) sequence. For OOD sequences (see example in Fig. 2di), subtrees are determined based on the maximum classification probability at the lowest rank among the ranks with a lower novelty detection score. Here, sequences receive a low novelty detection score at the order rank but a high score at the family rank, indicating that this sequence likely belongs to the known order (Fabales) while being classified into unknown families within the original tree. For ID sequences, the corresponding subtree is determined directly based on the genus with the highest probability (Fig. 2dii).
PhyloTune processes multiple sequences simultaneously and merges sequences from overlapping clades into a single sequence set for the largest clade. Taking Fig. 2d as an example, for five newly added sequences, PhyloTune outputs only two sequence sets: one for the order Fabales clade and one for the genus Primulina clade. These sequence sets include the high-attention regions of both the new sequences and all original sequences within each clade, used to reconstruct the subtrees for the Fabales and Primulina clades, respectively.
Figure 2e compares the time between our pipeline and standard phylogenetic reconstruction pipelines in the Plant dataset. By significantly reducing both sequence number and length for subsequent alignment and tree inference analyses, PhyloTune improves computational efficiency for phylogenetic tree updates. Specifically, our method reduces the time complexity from \(O(NL\log L)+O({N}^{2})\) to \(O(nl\log l)+O({n}^{2})\), where N and L represent the total number and maximum length of sequences in the original tree, while n and l denote the total number and maximum length of sequences in the subtree requiring update. For instance, when the updated sequence belongs to an unknown taxon of the order Malpighiales in the original tree, the time to align and update this subtree is only 4 min, which accounts for merely 0.33% of the time required by the standard pipelines. In addition, when using 8 A100 GPUs for inference, PhyloTune processes each sequence in approximately 0.01 seconds, a computation time significantly lower than that required for tree construction.
Performance evaluation of PhyloTune in identifying the smallest taxonomic units
PhyloTune identifies the smallest taxonomic unit of a new sequence in the original tree through two essential tasks: novelty detection and taxonomic classification. To assess its efficacy, we first evaluated the impact of training sample size on both classification and novelty detection performance using simulated datasets. Results indicate that performance improves consistently with increasing training samples (Fig. 3a). For instance, increasing the training set from 10 to 90 samples leads to a 35.5% improvement in Matthews Correlation Coefficient (MCC) and a 25.1% increase in AUROC. These findings underscore the challenges of fine-tuning pretrained large language models on limited data, where data scarcity can lead to overfitting.
a PhyloTune’s performance in identifying the smallest taxonomic unit on simulated datasets with varying training sequences. Top: Taxonomic classification metrics for known taxa. Bottom: Novelty detection metrics for unknown taxa. Line charts show the mean ± 95% confidence interval (CI, computed from SEM, n = 30 independent experiments). b Comparison of taxonomic classification between PhyloTune and MMseqs2 (using training data as a reference). Line charts show the mean ± 95% CI (n = 10 independent experiments). c Comparative analysis of novelty detection scores for PhyloTune and baseline, using in-distribution (ID) and out-of-distribution (OOD) test sequences from the Plant dataset (n = 15000 sequences). Source data are provided as a Source Data file.
Compared to MMseqs2, which relies on a reference database constructed from training samples, PhyloTune demonstrates a clear advantage in classifying sequences from known taxonomic units, particularly when training data is sparse (Fig. 3b). This suggests that AI-driven approaches are more adept at handling complex sequences with long-range homology, which are less likely to be well-represented in conventional reference databases56. Furthermore, traditional methods like MMseqs2 or BLAST lack a mechanism to assess consistency across taxonomic levels, limiting its ability to detect unknown taxonomic units.
To further evaluate PhyloTune, we compared it to a baseline on the Plant dataset. The baseline employs a fine-tuning strategy that freezes the backbone while training hierarchical linear probes (HLPs). PhyloTune significantly outperforms the baseline in both tasks (Table 1). In novelty detection, PhyloTune achieves 38.6% higher AUROC and 3.0% higher AUPR (Table 1a). The density distribution of novelty detection scores shows that PhyloTune generates more distinctive scores for both ID and OOD sequences, with most ID sequences receiving lower scores and OOD sequences receiving higher scores (Fig. 3c). In classification of known taxa, PhyloTune shows the largest improvements of 41.6, 23.5, 35.0, and 11.4% in precision, recall, F1 score, and MCC (Table 1b). Notably, traditional methods (e.g., BLAST, MMseqs2, and Kraken2) lack a mechanism to assess the consistency of identified taxa across hierarchical levels and exhibit a sharp decline in performance when classifying sequences containing unknown taxa (Supplementary Table 1).
To better assess performance across all abundance thresholds, we further presented a comparative analysis for PhyloTune and baseline on the Plant dataset in terms of macro-average Receiver Operating Characteristic (ROC) and Precision-Recall (PR) curves (Supplementary Fig. 1), as well as the ROC and PR curves for different categories (Supplementary Fig. 2). Although both methods employ weighted cross-entropy loss to address class imbalance during training, PhyloTune with a two-step training strategy significantly enhances the capability to capture distinctive class features. For example, at the class rank, PhyloTune outperforms the baseline by 5.1% in AUROC and 18.4% in AUPR. Despite these advances, challenges persist, particularly at taxonomic ranks like family and order, where category size and sample imbalance are more pronounced. For instance, the average precision (AveP) for Rubiaceae at the family rank and Cucurbitales at the order rank are only 57.58% and 53.98%, respectively. Hence, addressing class imbalance and the few-shot problem in DNA sequence data remains a critical frontier for enhancement through AI-driven methods.
Resolution of phylogenetic trees constructed with high-attention regions
To investigate whether the high-attention regions provided by PhyloTune encapsulate more sufficient informative sites for phylogenetic tree construction, we conducted a comparative analysis of phylogenetic trees constructed using high-attention and low-attention regions of sequences, respectively. We compared the normalized Robinson-Foulds (RF) distances between trees constructed using high- and low-attention regions and those built using the full-length sequence. Figure 4a illustrates the effect of varying the attention region length (i.e., 1/K of the full sequence length) on simulated datasets containing different numbers of sequences. In most cases, trees built from high-attention regions more closely resembled the ground truth trees than those built from low-attention regions. This trend was particularly pronounced when K = 3, where high-attention regions consistently outperformed low-attention regions across trees of all sizes (n = 20, 40, 60, 80, and 100). However, when only a quarter of the sequence length (K = 4) was extracted as an attention region, the accuracy of trees built from high-attention regions dropped significantly for datasets with 20 and 100 sequences. These findings highlight a trade-off between tree resolution and sequence length and underscore the need for further research to optimize fragment selection strategies, both in terms of region identification and fragment length.
a Difference in Robinson-Foulds (RF) distance between high-attention regions and full-length (RF(high, full)) versus low-attention regions and full-length (RF(low, full)) on the simulated datasets, across sequence counts and attention region lengths (1/K of the full length). Each box plot displays the median, interquartile range (25th to 75th percentile), and whiskers to minima/maxima within 1.5 times IQR (n = 10 independent experiments). b Difference in RF(high, full) versus RF(low, full) for order subtrees shown in Fig. 2d. c Phylogenetic trees for the angiosperm order Rosales constructed using high-attention (left) and low-attention (right) regions extracted by PhyloTune (same dataset as Fig. 2d). Source data are provided as a Source Data file.
We further validated our approach using data from 19 orders depicted in Fig. 2d (excluding the order Porellales, which contains only a single individual). Based on the performance observed in the simulated datasets, we set K = 3 and M = 1. Similar trends were observed in the real-world data: nearly 90% of the trees constructed using high-attention regions achieved smaller or equal RF distances to the full-sequence trees than those constructed using low-attention regions (Fig. 4b). This suggests that when there is a need to improve tree-building efficiency due to the length of input sequences, using high-attention regions as a substitute for full sequences is an effective option. We then visualized the tree of orders encompassing a minimum of two genera to intuitively show the clustering of individuals of the same genus. By taking the angiosperm order Rosales as an example, the results indicate that high-attention regions facilitate a superior resolution of the phylogenetic topology, while in the topology constructed using low-attention regions, species from the genera Ficus and Artocarpus were interspersed (Fig. 4c). Similar scenarios were observed in orders Asparagales, Fabales, Gentianales and Lamiales (Supplementary Fig. 3). However, for Malpighiales and Poales, although the topologies of the trees were more clustered, the low-attention trees were closer to the full-length trees (Fig. 4b). Notably, regardless of whether high-attention, low-attention, or full-length regions were used, the resulting topologies of Malpighiales and Poales differed significantly from those reported in prior studies57,58. This highlights the inherent gap between gene trees constructed from different genes–or even different segments of the same gene–and the species tree59. It also suggests that comparison with full-length sequences is only a reference to assess whether high-attention regions can replace them, and does not fully reflect topology accuracy. For taxonomic groups with unstable topologies, careful consideration is needed in selecting appropriate markers or genetic regions to ensure reliable phylogenetic reconstruction60.
Differential performance of various markers in taxonomic classification and novelty detection
To further evaluate the effect of the model on different molecular markers, we analyzed the performance of nine molecular markers within the Plant dataset in terms of novelty detection and taxonomic classification. Among these nine markers, six are from the chloroplast genome, including both coding genes (atpB, matK, rbcL) and intergenic spacer regions (rpl32-trnL, trnL-trnF, trnH-psbA), while three others (5S rRNA, 28S rRNA, ITS) are from the nuclear genome. It is important to note that during the PhyloTune model training, all sequences are treated equally, without bias towards any specific molecular marker. Our findings indicate that performance varies across different markers, and performance for the same marker also varies across tasks. For example, the two nuclear markers (28S rRNA, ITS) as well as one IGS region from the chloroplast genome (trnL-trnF) demonstrate superior performance in novelty detection tasks, while other plastid markers (rpl32-trnL, trnH-psbA, and rbcL) perform relatively poorly (Fig. 5a). For taxonomic classification, trnL-trnF and rbcL excel in all assessed metrics, whereas atpB and trnH-psbA show comparatively weaker performance (Fig. 5b–d).
a Comparison of the average AUROC for four taxonomic ranks of different markers in novelty detection (n = 15000 sequences for each rank). b–d Comparison of the average macro precision, macro recall and macro F1-score for four taxonomic ranks of different markers in taxonomic classification (n = 14700, 14400, 13200, 10000 sequences for class, order, family and genus). e Pearson correlation coefficient between average attention, heterozygosity, the fixation index (FST), absolute divergence (DXY), and substitution rate based on nine molecular markers. f Attention heatmap of the chloroplast marker matK using PhyloTune (n = 1000 sequences). The red box highlights the attention peak region for the majority of sequences, with an example of the corresponding DNA sequences displayed above it. g Average attention, heterozygosity, substitution rate, FST, and DXY curves of matK. The blue shaded area denotes the peak region of attention. Source data are provided as a Source Data file.
The performance of molecular markers in taxonomy relies on the presence of informative variant sites that reflect the evolutionary relationships among species61. Here, the ability of these variant sites to be captured by the models is pivotal for taxonomic classification and novelty detection. To explain varying marker performance, we calculated the Pearson coefficients of average attention, heterozygosity, fixation index (FST), nucleotide substitution rate and absolute divergence (DXY) across different markers, where heterozygosity serves as a measure of genetic variation, FST is a statistical parameter used to quantify the degree of genetic differentiation among populations, substitution rate represents the evolutionary rate of different genetic regions, and DXY reflects the more ancient divergence between sequences. Overall, the model’s attention is positively correlated with most genetic parameters, particularly the substitution rate, suggesting that the rapid evolutionary rate may make these regions valuable for evolutionary studies (Fig. 5e). We also found a significant negative correlation between average attention and these genetic parameters for rbcL and atpB, while these markers also show relatively weak performance in identifying smallest taxonomic unit (Fig. 5a–d). These results revealed that for the molecular markers exhibiting inferior performance, the attention regions of our model were not focused on those exhibiting high genetic variation. In other words, our study demonstrates that the model autonomously selects markers conducive to classification, albeit the underlying mechanisms remain unclear; this nonetheless provides a valuable reference for future marker selection. Particularly in today’s era of explosive growth in sequencing data, the efficient screening of informative data for classification and phylogenetic tasks is of paramount importance. Detailed results of each molecular marker are shown in Supplementary Table 2, 3.
Attention heatmap of PhyloTune demonstrates its ability to capture genetic diversity
To gain deeper insights into the utility of attention weights provided by PhyloTune for phylogenetic tree construction, we plotted the attention heatmaps for nine molecular markers (Fig. 5f and Supplementary Figs. 4–6). We randomly sampled 1000 sequences for each molecular marker from the test set of the Plant dataset to plot their attention heatmaps. Our observation reveals a concentration of the model’s attention on molecular markers predominantly within the anterior and mid-portions, exhibiting distinct patterns across different markers (Supplementary Figs. 4–6). For example, for the marker matK, the attention is concentrated on the regions between 400 bp to 600 bp. By comparing the average attention of all sequences to heterozygosity, substitution rate, FST and DXY curves, we found that regions receiving increased attention often exhibit relatively high levels of these genetic parameters (Fig. 5g). However, it’s noteworthy that the model does not focus on all regions exhibiting high heterozygosity, substitution rate, FST and DXY. This finding concurs with the prior knowledge that a balanced mix of conserved and variable regions provides a comprehensive perspective on evolutionary relationships62; it suggests that the model might also rely on a certain ratio of variable and conserved sequence data to facilitate species classification, thereby echoing fundamental principles of phylogenetic inference.
Although AI models are often criticized for their lack of interpretability, the Transformer with a self-attention mechanism provides a solution to this problem37. Our analysis of attention also further suggests that the attention learned by LLMs may provide different perspectives and insights for molecular phylogenetic analysis.
Additional Experiments on the Bordetella Genus to Validate PhyloTune’s Generalizability
To explore the generalizability of PhyloTune beyond simulated and plant datasets, we extended our analysis to microbial data, focusing on the Bordetella genus (Supplementary Table 4). Our findings indicate that PhyloTune outperformed MMseqs in identifying the correct taxonomic unit, as shown in Supplementary Table 5. PhyloTune achieved AUPR values greater than 95% at both clade and species levels, demonstrating its superior ability to detect sequences from unknown taxa. Comparison of phylogenetic trees constructed from high- and low-attention sequence regions reveals that, in complex clade (clade1), trees built from high-attention regions exhibit greater similarity to the full-length tree than those constructed from low-attention regions (Fig. 6). For simpler clades (clade2,3), trees derived from both high- and low-attention regions had identical topologies. Further analysis of attention scores revealed that regions with higher attention scores were generally associated with higher levels of heterozygosity, substitution rate, FST, and DXY (Supplementary Fig. 7). This pattern was consistent with findings from the Plant dataset, further supporting the utility of attention in phylogenetic tree construction and their potential to capture evolutionary signals. However, some genes in the Bordetella dataset showed weaker correlations (p-value > 0.1, Supplementary Data 1), likely due to the larger number of genes with fewer sequences per gene, which limits the model’s generalization capacity. This highlights the challenge of applying pre-trained models to datasets with many genes but limited sequence data, an area for future exploration. More details on the experimental setup and results are provided in Supplementary Information.
a Difference in Robinson-Foulds (RF) distance between trees constructed from high-attention regions and full-length sequences (RF(high, full)) versus those constructed from low-attention regions and full-length sequences (RF(low, full)). High- and low-attention regions are defined by dividing the sequence from the Bordetella dataset into K = 12 segments and selecting the top M (1, 2, 3, and 4) regions with the highest and lowest attention scores, respectively. b Comparison of phylogenetic tree topologies of clade1 constructed from high- and low-attention regions at different M values. Source data are provided as a Source Data file.
Discussion
Although deep learning has made breakthrough progress in computational biology, its application in phylogenetic inference is still in its infancy. The continual growth in the number and length of DNA sequences, propelled by advancements in high-throughput technologies, presents formidable challenges for phylogenetic updates, necessitating a delicate balance between computational efficiency and accuracy. PhyloTune offers a pioneering solution by integrating deep learning with traditional methods. Leveraging the open-source genomic large language model, it efficiently identifies taxonomic units for new sequences and extracts potentially more valuable regions, thereby reducing the input for alignment and tree construction tools in terms of both sequence number and length.
PhyloTune attempts to harness genomic large language models for phylogenetic updates. Diverging from approaches that directly predict tree topology (e.g., FastTree, RAxML) or update tree nodes (e.g., DEPP31), PhyloTune focuses on refining tree update efficiency by narrowing the update scope and sequence processing length. Comprehensive experiments conducted on simulated, plant (Embryophyta) and microbial (Bordetella genus) datasets underscore the feasibility of our approach.
In addition, PhyloTune introduces the concept of the smallest taxonomic unit identification to refine the update scope, extending beyond standard taxonomic classification. Traditional classification methods, while effective in taxonomic assignments, are limited in detecting novel taxonomic units due to the lack of mechanisms for evaluating sequence consistency across taxonomic levels. Moreover, PhyloTune utilizes model attention mechanisms to assess the importance of individual sequence sites, offering an alternative approach to sequence analysis across different molecular markers.
While PhyloTune is trained within a taxonomic framework, its novelty detection and attention mechanisms provide potential to identify inconsistencies, detect novel lineages, and refine phylogenetic relationships by leveraging its novelty detection scores and attention mechanisms. Future advancements, such as incorporating unsupervised clustering, could further enhance its capability to discover entirely new evolutionary patterns, broadening its applicability across diverse datasets lacking predefined taxonomic structures. While our work presents a step toward bridging AI and phylogenetic inference, the journey toward seamlessly integrating the two, possibly achieving end-to-end phylogenetic tree construction and updates, remains a frontier for exploration. With the proliferation of foundational models, leveraging their modeling prowess to enhance phylogenetic tree construction holds promise and warrants further investigation.
Methods
Dataset
Simulated datasets
The simulated datasets were generated using Seq-Gen v1.3.563. First, a rooted binary tree with a specified number of species (n) was randomly generated using a custom Python script. Sequences of 3000 bp were then simulated along these trees using Seq-Gen. To evaluate the smallest taxonomic unit identification, species (excluding the outgroup) were divided into two taxonomic units based on the topology of a randomly generated tree (n = 100). The outgroup was expanded into a branch of five species, designated as an unknown taxonomic unit for novelty detection. For sequences belonging to known taxonomic units, we performed stratified 10-fold cross-validation to generate 10 sets of training and testing datasets. In addition to the original training set size, we subsampled 20, 30, 40, 50, ⋯ , 80 sequences to study the impact of varying training sample sizes. To minimize sampling bias, three independent sequence matrices were generated with Seq-Gen, resulting in a total of 30 experimental replicates for each training sample size.
Plant dataset
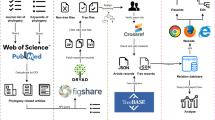
The Plant dataset comprises 157, 742 nucleotide sequences of Embryophyta (land plants), representing 19, 887 species from 150 genera. The original sequences were manually retrieved from the NCBI nucleotide database. We collected sequences from the nine most commonly used molecular markers: atpB, matK, rbcL, rpl32-trnL, trnL-trnF, trnH-psbA, 5S rRNA, 28S rRNA, and ITS, covering the coding genes (CDS) and intergenic spacer (IGS) regions of chloroplast DNA and nuclear DNA. Except for filtering sequences longer than 1530 bp to fit the input length limit of the used pre-trained model DNABERT and removing duplicate sequences, no additional processing was performed on the sequences.
For the identification of smallest taxonomic unit, we recorded taxonomic information for each sequence across four ranks: class, order, family, and genus (Fig. 7a). To ensure that each taxon has a sufficient number of sequences for training and evaluation, we selected the 150 genera with the largest number of sequences to generate the Plant dataset. For the top 100 most abundant genera, 50 and 100 samples were randomly selected to form the validation and test sets, respectively, with the remaining samples forming the training set. For the other 50 genera, 100 samples were downsampled to form an additional part of the test set (Fig. 7b). The top 100 genera and their corresponding taxa in class, order, and family are referred to as ID taxa, or known taxa, as they are observed and learned by the model during training. The remaining 50 genera are referred to as OOD taxa, or unknown taxa, as they are only observed by the model during testing. It is important to note that lineages of these unknown genus taxa may fall within the distribution. According to whether the taxon is observed during training, the test set samples of the Plant dataset can be divided into five types: known genus, unknown genus but known family, unknown genus and family but known order, unknown genus and family and order but known classes, and unknown taxa in all ranks (Fig. 7a). Intuitively, from genus to class, the number of samples belonging to unknown taxa gradually decreases. Table 2 shows the number of ID and OOD taxa in each taxonomic rank of the Plant dataset, as well as the corresponding number of samples. The ratio of ID and OOD test samples at the genus level is 2: 1, but this ratio at the class level increases sharply to 49: 1. In addition, the training set of Plant retains the imbalanced characteristics of DNA sequences in NCBI, with distributions at all ranks clearly exhibiting class imbalance (Fig. 7c). More than 95.5% of the sequences belong to the class Magnoliopsida, which consists of 29 orders with an exponentially decreasing distribution. The order with the largest number of sequences is Poales, which consists of two families and seven genera. Further details on the Plant dataset can be found in Supplementary Data 2.
a Taxonomic hierarchy and sample type. The Plant dataset, built on the Embryophyta, organizes samples across four taxonomic ranks: class, order, family, and genus. Samples are categorized into five types based on their known taxonomic resolution: ① known genus, ② unknown genus but known family, ③ unknown genus and family but known order, ④ unknown genus and family and order but known classes, and ⑤ unknown taxa in all taxonomic ranks. b Splitting of training, validation and test sets. Fifty and 100 samples of each known genus are randomly sampled to generate validation and test sets, and the remaining samples are used for the training set. 100 samples of an unknown genus are randomly sampled to generate another part of the test set. c Distribution of the Plant training set for each of the four taxonomic ranks, all of which exhibit significant class imbalance. Source data are provided as a Source Data file.
Bordetella dataset
The Bordetella dataset was created using a publicly available dataset on Bordetella genus64. We selected 140 core genes with high coverage in all Bordetella species for our experiments. The dataset included 39 species, with 13 designated as known species for training and testing, and the remaining species as unknown species for testing. Species were grouped into three clades and one outgroup, with sequences randomly allocated for training, validation, and testing. A more detailed taxonomic structure and dataset statistics are summarized in Supplementary Table 4.
Model architecture
The architecture of PhyloTune consists of the Bidirectional Encoder Representations from Transformers (BERT)65 and a set of hierarchical linear probes (HLPs). Figure 1c illustrates the adaptation of PhyloTune to the Plant dataset. The BERT module’s configuration depends on the selected pretrained model; for instance, we fine-tuned DNABERT for the Plant dataset, while for the simulated and Bordetella datasets, we used DNABERT-S. In principle, PhyloTune can be applied to any BERT-based pretrained DNA language model, leveraging the model’s robust representation capabilities to identify the smallest taxonomic units and high-attention regions.
The design of the HLPs is tailored to the taxonomic hierarchy of the phylogenetic tree being updated. For example, the Plant dataset encompasses four taxonomic ranks (class, order, family, and genus), so we implemented four HLP layers, each corresponding to novelty detection and classification at a specific rank. In contrast, the hierarchical structures of the simulated dataset and the Bordetella dataset consist of one and two levels, respectively, leading to the implementation of a single-layer HLP and a two-layer HLP for these datasets. Inspired by BERTax56, each HLP in PhyloTune receives input from the final hidden state of the [CLS] token, along with outputs from higher taxonomic levels. This hierarchical information flow captures inter-level dependencies, enhancing both classification accuracy and novelty detection.
Training strategy
PhyloTune employs a two-step training strategy. In the first step, the BERT module remains frozen while only the randomly initialized HLPs are trained. Subsequently, all layers are unfrozen, and the entire model is trained together. The number of epochs were adjusted based on the size of the training set. For the Plant dataset, we used 100 epochs for Step 1 and 30 epochs for Step 2, while for the Bordetella dataset, we used 10 and 5 epochs, respectively. Throughout training, learning rate reduction occurs when a plateau is reached, and early stopping based on the validation set is applied. Due to the limited number of training samples in the simulated dataset (fewer than 90), only the first step of fine-tuning is performed to prevent overfitting.
The training goal of fine-tuning was to enable the model to correctly classify each sequence at every taxonomic rank. In classification tasks, cross-entropy loss is commonly used to quantify the difference between the predicted and actual probability distributions. We fine-tuned the model by minimizing this difference during training. To prevent bias toward predicting the most common taxa, class weights were calculated for each taxonomic rank, balancing the taxa and allowing the model to focus more on samples from underrepresented taxa. The entire model was trained using a weighted cross-entropy loss (L), defined as follows:
where
Here \({y}_{s,r,i}^{{\prime} }\) and ys,r,i are the ground-truth and predicted values of sequence s for taxon i at rank r, respectively. R is the set of taxonomic ranks, Nr is the number of known taxa at rank r, nr,i is the number of training sequences for taxon i at rank r, and N is the total number of training sequences. It is important to note that R, Nr, nr,i, N are dependent on the statistics of the training set. For example, in the Plant dataset, R = {class, order, family, genus}, Nr = {5, 34, 65, 100} for r ∈ R, and N = 137, 742 (see Table 2). In contrast, for the Bordetella dataset, R = {clade, species}, Nr = {3, 13} for r ∈ R, and N = 1, 930 (see Supplementary Table 4). Since all simulated datasets have only one taxonomic level, we directly implemented it using the SVM function from scikit-learn v1.4.266.
Evaluation
For evaluation of smallest taxonomic unit identification, all experiments were conducted on the test set that was unseen during training. The baseline method used in Table 1 only utilized the first step of the two-step fine-tuning process, i.e., freezing the backbone and fine-tuning randomly initialized HLPs. For a fair comparison, traditional methods, including MMseqs2, BLAST, and Kraken2, were evaluated using the training set as the reference database.
For evaluation of phylogenetic trees constructed in high-attention regions, we utilized high- and low-attention regions extracted by PhyloTune to construct phylogenetic trees and compute their normalized Robinson-Foulds (RF) distance with the full-length trees using ETE367. Tree construction involved MAFFT v.7 for sequence alignment and RAxML v.8.2 for subtree inference. For the simulated dataset, we generated datasets with varying sequence numbers (n = 20, 40, 60, 80, 100), and for each size, 10 trees were randomly generated for validation. For the Plant dataset, species were selected from the training set to form the sequences used for tree construction evaluation, with each species containing at least five of the nine molecular markers. The examples of the five new sequences in Fig. 2d were randomly selected from the test set. For the Bordetella dataset, the entire training set was used to evaluate three clades.
For attention interpretability analysis, heterozygosity and substitution rate were calculated using the formulas from ref. 68, while FST and DXY were calculated based on ref. 69. For the Plant dataset, 1000 test sequences per molecular marker were randomly selected for analysis, while for the Bordetella dataset, the entire training set was used.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All datasets analyzed in this article are publicly available. The simulated datasets are available at https://github.com/danruod/PhyloTune/tree/main/datasets/simulated. The Plant dataset is available at https://github.com/danruod/PhyloTune/blob/main/datasets/Plant.zip. The Bordetella dataset is available at https://github.com/danruod/PhyloTune/blob/main/datasets/Bordetella.zip. Source data are provided in this paper.
Code availability
The alignment and tree inference tools use MAFFT v.7 (https://mafft.cbrc.jp) and RAxML v.8.2 (https://github.com/stamatak/standard-RAxML), respectively. The Robinson-Foulds distance are calculated using ETE3 (http://etetoolkit.org). Other data analyses including novelty detection and taxonomic classification used Python v3.11.9 (https://www.python.org/), PyTorch v2.0.1 (https://pytorch.org/), Transformers v4.42.3 (https://huggingface.co/docs/transformers), NumPy v1.24.3 (https://www.numpy.org/), pandas v2.0.2 (https://pandas.pydata.org/), scikit-learn v1.4.2 (https://scikit-learn.org/stable/), SciPy v1.10.1 (https://scipy.org/), seaborn v0.13.2 (https://seaborn.pydata.org/) and Matplotlib v3.7.1 (https://matplotlib.org/). The code developed in this manuscript and pretrained model weights are deposited in Github: https://github.com/danruod/PhyloTuneand https://doi.org/10.5281/zenodo.1553385370.
References
Woese, C. R. & Fox, G. E. Phylogenetic structure of the prokaryotic domain: the primary kingdoms. Proc. Natl. Acad. Sci. USA 74, 5088–5090 (1977).
Ciccarelli, F. D. et al. Toward automatic reconstruction of a highly resolved tree of life. Science 311, 1283–1287 (2006).
Hug, L. A. et al. A new view of the tree of life. Nat. Microbiol. 1, 1–6 (2016).
Winter, M., Devictor, V. & Schweiger, O. Phylogenetic diversity and nature conservation: where are we? Trends Ecol. Evol. 28, 199–204 (2013).
Stiller, J. et al. Complexity of avian evolution revealed by family-level genomes. Nature 629, 851–860 (2024).
Zuntini, A. R. et al. Phylogenomics and the rise of the angiosperms. Nature 629, 843–850 (2024).
Du Plessis, L. et al. Establishment and lineage dynamics of the SARS-CoV-2 epidemic in the UK. Science 371, 708–712 (2021).
Lemieux, J. E. et al. Phylogenetic analysis of SARS-CoV-2 in Boston highlights the impact of superspreading events. Science 371, eabe3261 (2021).
Yu, J. et al. Phylogeny and molecular evolution of the first local monkeypox virus cluster in Guangdong Province, China. Nat. Commun. 14, 8241 (2023).
Naxerova, K. & Jain, R. K. Using tumour phylogenetics to identify the roots of metastasis in humans. Nat. Rev. Clin. Oncol. 12, 258–272 (2015).
Enriquez-Navas, P. M. et al. Exploiting evolutionary principles to prolong tumor control in preclinical models of breast cancer. Sci. Transl. Med. 8, 327ra24–327ra24 (2016).
Schwartz, R. & Schäffer, A. A. The evolution of tumour phylogenetics: principles and practice. Nat. Rev. Genet. 18, 213–229 (2017).
Zepeda-Rivera, M. et al. A distinct Fusobacterium nucleatum clade dominates the colorectal cancer niche. Nature 628, 424–432 (2024).
Reuter, J. A., Spacek, D. V. & Snyder, M. P. High-throughput sequencing technologies. Mol. Cell 58, 586–597 (2015).
Kapli, P., Yang, Z. & Telford, M. J. Phylogenetic tree building in the genomic age. Nat. Rev. Genet. 21, 428–444 (2020).
Brocchieri, L. Phylogenetic inferences from molecular sequences: review and critique. Theor. Popul. Biol. 59, 27–40 (2001).
Gilbert, P. S., Wu, J., Simon, M. W., Sinsheimer, J. S. & Alfaro, M. E. Filtering nucleotide sites by phylogenetic signal to noise ratio increases confidence in the Neoaves phylogeny generated from ultraconserved elements. Mol. Phylogenet. Evol. 126, 116–128 (2018).
Yang, Z. & Rannala, B. Molecular phylogenetics: principles and practice. Nat. Rev. Genet. 13, 303–314 (2012).
Gascuel, O. BIONJ: an improved version of the NJ algorithm based on a simple model of sequence data. Mol. Biol. Evol. 14, 685–695 (1997).
Rempel, A. & Wittler, R. SANS serif: alignment-free, whole-genome-based phylogenetic reconstruction. Bioinformatics 37, 4868–4870 (2021).
Wang, F. et al. MIKE: an ultrafast, assembly-, and alignment-free approach for phylogenetic tree construction. Bioinformatics 40, btae154 (2024).
Ng, M. P., Steel, M. & Wormald, N. C. The difficulty of constructing a leaf-labelled tree including or avoiding given subtrees. Discrete Appl. Math. 98, 227–235 (2000).
Berry, V., Scornavacca, C. & Weller, M. Scanning Phylogenetic Networks is NP-Hard. (2020).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2–approximately maximum-likelihood trees for large alignments. PLoS ONE 5, e9490 (2010).
Lartillot, N., Rodrigue, N., Stubbs, D. & Richer, J. PhyloBayes MPI: phylogenetic reconstruction with infinite mixtures of profiles in a parallel environment. Syst. Biol. 62, 611–615 (2013).
Aberer, A. J., Kobert, K. & Stamatakis, A. ExaBayes: massively parallel Bayesian tree inference for the whole-genome era. Mol. Biol. Evol. 31, 2553–2556 (2014).
Kozlov, A. M., Darriba, D., Flouri, T., Morel, B. & Stamatakis, A. RAxML-NG: a fast, scalable and user-friendly tool for maximum likelihood phylogenetic inference. Bioinformatics 35, 4453–4455 (2019).
Suvorov, A., Hochuli, J. & Schrider, D. R. Accurate inference of tree topologies from multiple sequence alignments using deep learning. Syst. Biol. 69, 221–233 (2020).
Zou, Z., Zhang, H., Guan, Y. & Zhang, J. Deep residual neural networks resolve quartet molecular phylogenies. Mol. Biol. Evol. 37, 1495–1507 (2020).
Bhattacharjee, A. & Bayzid, M. S. Machine learning based imputation techniques for estimating phylogenetic trees from incomplete distance matrices. BMC Genom. 21, 497 (2020).
Jiang, Y., Balaban, M., Zhu, Q. & Mirarab, S. DEPP: deep learning enables extending species trees using single genes. Syst. Biol. 72, 17–34 (2023).
Jiang, Y., Tabaghi, P. & Mirarab, S. Learning hyperbolic embedding for phylogenetic tree placement and updates. Biology 11, 1256 (2022).
Nesterenko, L., Blassel, L., Veber, P., Boussau, B. & Jacob, L. Phyloformer: Fast, accurate and versatile phylogenetic reconstruction with deep neural networks. Mol. Biol. Evol. 42, msaf051 (2025).
Sapoval, N. et al. Current progress and open challenges for applying deep learning across the biosciences. Nat. Commun. 13, 1728 (2022).
Al-Zubaidi, M. R., Thwiny, H. T. & Al-Biati, M. N. Modulation of chitosan nanoparticles properties for sheep pox mucosal vaccine delivery with cytotoxicity and release studies-in vitro. Iraqi J. Vet. Sci. 37, 111–119 (2023).
Zaharias, P., Grosshauser, M. & Warnow, T. Re-evaluating deep neural networks for phylogeny estimation: the issue of taxon sampling. J. Comput. Biol. 29, 74–89 (2022).
Ji, Y., Zhou, Z., Liu, H. & Davuluri, R. V. DNABERT: pre-trained Bidirectional Encoder Representations from Transformers model for DNA-language in genome. Bioinformatics 37, 2112–2120 (2021).
Avsec, Ž. et al. Effective gene expression prediction from sequence by integrating long-range interactions. Nat. Methods 18, 1196–1203 (2021).
Zhang, D. et al. DNAGPT: a generalized pretrained tool for multiple DNA sequence analysis tasks Preprint at https://arxiv.org/abs/2307.05628 (2023).
Theodoris, C. V. et al. Transfer learning enables predictions in network biology. Nature 618, 616–624 (2023).
Nguyen, E. et al. HyenaDNA: long-range genomic sequence modeling at single nucleotide resolution. Adv. Neural. Inf. Process. Syst. 36, 43177–43201 (2023).
Bond, S. R., Keat, K. E., Barreira, S. N. & Baxevanis, A. D. BuddySuite: command-line toolkits for manipulating sequences, alignments, and phylogenetic trees. Mol. Biol. Evol. 34, 1543–1546 (2017).
Kumar, S., Nei, M., Dudley, J. & Tamura, K. MEGA: a biologist-centric software for evolutionary analysis of DNA and protein sequences. Brief. Bioinform. 9, 299–306 (2008).
Zhou, Z. et al. DNABERT-S: Pioneering species differentiation with species-aware DNA embeddings. Preprint at https://doi.org/10.48550/arXiv.2402.08777 (2024).
Parr, C. S., Lee, B., Campbell, D. & Bederson, B. B. Visualizations for taxonomic and phylogenetic trees. Bioinformatics 20, 2997–3004 (2004).
Schoch, C. L. et al. NCBI Taxonomy: a comprehensive update on curation, resources and tools. Database 2020, baaa062 (2020).
Ye, J., McGinnis, S. & Madden, T. L. BLAST: improvements for better sequence analysis. Nucleic Acids Res. 34, W6–W9 (2006).
Steinegger, M. & Söding, J. MMseqs2 enables sensitive protein sequence searching for the analysis of massive data sets. Nat. Biotechnol. 35, 1026–1028 (2017).
Wood, D. E., Lu, J. & Langmead, B. Improved metagenomic analysis with Kraken 2. Genome Biol. 20, 1–13 (2019).
Nabhan, A. R. & Sarkar, I. N. The impact of taxon sampling on phylogenetic inference: a review of two decades of controversy. Brief. Bioinform. 13, 122–134 (2012).
Bernot, J. P. et al. Major revisions in pancrustacean phylogeny and evidence of sensitivity to taxon sampling. Mol. Biol. Evol. 40, msad175 (2023).
Koning, E., Phillips, M. & Warnow, T. pplacerDC: a new scalable phylogenetic placement method. In Proc. ACM International Conference on Bioinformatics, Computational Biology, and Health Informatics. 1–9 (2021).
Wedell, E., Cai, Y. & Warnow, T. SCAMPP: scaling alignment-based phylogenetic placement to large trees. IEEE/ACM Trans. Comput. Biol. Bioinform. 20, 1417–1430 (2022).
Group, A. P. et al. An update of the Angiosperm Phylogeny Group classification for the orders and families of flowering plants: APG IV. Bot. J. Linn. Soc. 181, 1–20 (2016).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Mock, F., Kretschmer, F., Kriese, A., Böcker, S. & Marz, M. Taxonomic classification of DNA sequences beyond sequence similarity using deep neural networks. Proc. Natl. Acad. Sci. USA 119, e2122636119 (2022).
Xi, Z. et al. Phylogenomics and a posteriori data partitioning resolve the Cretaceous angiosperm radiation Malpighiales. Proc. Natl. Acad. Sci. USA 109, 17519–17524 (2012).
Wang, H., Wu, Z., Li, T. & Zhao, J. Phylogenomics resolves the backbone of Poales and identifies signals of hybridization and polyploidy. Mol. Phylogenet. Evol. 200, 108184 (2024).
Szöllősi, G. J., Tannier, E., Daubin, V. & Boussau, B. The inference of gene trees with species trees. Syst. Biol. 64, e42–e62 (2015).
Li, Y. et al. Phylogenomics of Bivalvia using ultraconserved elements reveal new topologies for Pteriomorphia and Imparidentia. Syst. Biol. 74, 16–33 (2024).
Patwardhan, A., Ray, S. & Roy, A. Molecular markers in phylogenetic studies-a review. J. Phylogenet. Evol. Biol. 2, 131 (2014).
Bose, N. & Moore, S. D. Variable region sequences influence 16S rRNA performance. Microbiol. Spectr. 11, e01252–23 (2023).
Rambaut, A. & Grass, N. C. Seq-Gen: an application for the Monte Carlo simulation of DNA sequence evolution along phylogenetic trees. Bioinformatics 13, 235–238 (1997).
Bridel, S. et al. A comprehensive resource for Bordetella genomic epidemiology and biodiversity studies. Nat. Commun. 13, 3807 (2022).
Devlin, J., Chang, M.-W., Lee, K. & Toutanova, K. BERT: pre-training of deep bidirectional transformers for language understanding. In Proc. 2022 Conference of the North American Chapter of the Association For Computational Linguistics Human Language Technologies (2019).
Pedregosa, F. et al. Scikit-learn: Machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Huerta-Cepas, J., Serra, F. & Bork, P. ETE 3: reconstruction, analysis, and visualization of phylogenomic data. Mol. Biol. Evol. 33, 1635–1638 (2016).
Nei, M. Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89, 583–590 (1978).
Nagylaki, T. Fixation indices in subdivided populations. Genetics 148, 1325–1332 (1998).
Deng, D. et al. PhyloTune: an efficient method to accelerate phylogenetic updates using a pretrained DNA language model. https://doi.org/10.5281/zenodo.15533853 (2025).
Acknowledgements
W.X., J.Z., and B.W. acknowledge support from the “Pioneer” and “Leading Goose” R&D Program of Zhejiang (2024SSYS0007). P.L. acknowledges support from the Key Technology Research and Development Program of Zhejiang Province (No. 2023C03138). G.C. acknowledges support from the Zhejiang Province Vanguard Goose-Leading Initiative (No. 2025C01114) and National Natural Science Foundation of China (Project No. 62376254, 32341017, 32341018). P.A.H. acknowledges support from the Research Grants Council of the Hong Kong Special Administrative Region, China, under Project T45-401/22-N.
Author information
Authors and Affiliations
Contributions
P.A.H., G.C., J.Z., P.L. and W.X. conceived and supervised the study. D.D. developed the algorithm, implemented the code, and conducted the experiments, including dataset preparation, model training, and statistical analysis. W.X. generated the simulated datasets, curated the plant and microbial datasets, reconstructed and analyzed phylogenetic trees, and performed traditional classification experiments. G.C. advised on the algorithm design. J.Z. and P.L. guided the experimental design. G.C., J.Z., P.L. and B.W. contributed to the analysis of experimental results. D.D. and W.X. drafted the manuscript and prepared the supplementary materials. H.P.C., P.L., Y.F., J.Z., B.W., G.C. and P.A.H. revised the manuscripts. All authors participated in the discussions and agreed with the contents of this work. P.A.H., G.C., W.X., J.Z. and B.W. provided research environment and funding support.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Deng, D., Xu, W., Wu, B. et al. PhyloTune: An efficient method to accelerate phylogenetic updates using a pretrained DNA language model. Nat Commun 16, 6905 (2025). https://doi.org/10.1038/s41467-025-61684-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-61684-3