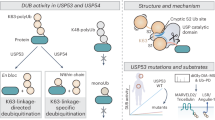
Abstract
DNA damage tolerance (DDT) is an important pathway that allows cells to bypass DNA lesions during replication. DDT is orchestrated by ubiquitination of PCNA, where monoubiquitinated PCNA (PCNA-Ub) initiates recruitment of TLS polymerases but also serves as a substrate for further ubiquitination, forming K63-polyubiquitinated PCNA that leads to HR-mediated bypass mechanisms. Recent work on USP1/UAF1 inhibition revealed that formation of K48-linked chains also occurs on PCNA, resulting in its proteasomal degradation. USP1/UAF1 is established as deubiquitinating enzyme (DUB) for PCNA-Ub, but little is known about removal of ubiquitin chains on PCNA. Here we show that USP1/UAF1 cleaves both K48 and K63-linked ubiquitin chains on PCNA efficiently, using an exo-cleavage mechanism. Kinetic analysis reveals that USP1/UAF1 prefers cleaving the ubiquitin-ubiquitin bond over cleavage of the ubiquitin-PCNA bond and therefore treats poly- and monoubiquitinated PCNA as different substrates. A cryo-EM structure of USP1/UAF1 with a K63-diubiquitin and structure-based mutagenesis suggests that this mechanistic preference is maintained in evolution. This unusual mechanism can cause temporal enrichment of monoubiquitinated PCNA during polyubiquitination. It will be interesting to see how this affects DDT pathway balance.
Similar content being viewed by others
Introduction
Ubiquitination plays a crucial role in regulation of the response to DNA damage1. There is extensive variety in ubiquitin signaling: Ubiquitin can be attached as a single entity or it can form different types of polyubiquitin chains by linking to any of the seven internal lysines or N-terminal methionine in ubiquitin2,3. Each of these different types of ubiquitination can lead to different outcomes on a protein substrate. Reversion of ubiquitination by deubiquitinating enzymes (DUBs) is therefore a crucial component in ubiquitin signaling4,5.
USP1 is a DUB with important roles in the response to DNA damage and is emerging as a promising clinical target6,7. USP1 functions as a heterodimer with its essential cofactor UAF1, which hyperactivates enzyme activity by relieving autoinhibition caused by insertions 1 and 3 in the USP1 catalytic domain8. USP1/UAF1 is a highly specific DUB that targets PCNA and Fanconi anemia proteins FANCI/FANCD29,10. This gives USP1/UAF1 important roles in two DNA damage response pathways: DNA damage tolerance (DDT) through action on PCNA, and interstrand crosslink (ICL) repair through action on FANCI/FANCD2.
During replication, PCNA and its interacting polymerase encounter numerous DNA lesions, which block the replicative polymerase from inserting the next base. This replication fork stalling can lead to fork collapse and overall genomic instability. To prevent this, DDT enables bypass of DNA lesions instead of waiting for DNA repair mechanisms, which allows for swift continuation of replication and maintenance of genome stability. Alternatively, repriming mediated by PrimPol can ensure that replication continues downstream of the lesion11, but this leaves a ssDNA gap behind the stalled PCNA, which subsequently requires gap filling by DDT12,13,14.
DDT is initiated by stalling of PCNA at the DNA lesion, which sets off a cascade of reactions that culminates in monoubiquitination of PCNA at its K164-residue (PCNA-Ub) by Rad6/Rad1815. PCNA-Ub exchanges its bound replication polymerase for a translesion synthesis (TLS) polymerase16,17,18. These polymerases can bypass DNA lesions due to their lack of proofreading activity and modified active sites that can accommodate different types of lesions19,20. While these features render the TLS polymerases more error-prone, TLS can be coupled with mismatch repair to correct for wrongly inserted bases21. PCNA-Ub is also a substrate for K63-linked polyubiquitination15, which is catalyzed by a different set of E2 and E3 enzymes. These have been identified as Ubc13/Mms2 and Rad5 in yeast15,22, while in humans multiple E3s have been identified23,24,25,26.
Polyubiquitinated PCNA (K63-PCNA-UbN) leads to two homologous recombination (HR)-mediated pathways, referred to as template switching (TS) and fork reversal (FR), that can also bypass DNA lesions15,27. Both pathways use the undamaged sister strand as a template and are considered to be error-free28. However, what exactly triggers formation of K63-PCNA-UbN and why HR-mediated bypass is required along with TLS is not clear. Additionally, it is unclear when fork reversal and template switching are preferred over translesion synthesis. It is possible that these HR-mediated bypass mechanisms are required for bulkier lesions which would not fit in the TLS polymerase active site29.
Timely removal of ubiquitin from PCNA is important since it allows for continuation of regular replication. Deubiquitination of PCNA-Ub is carried out by USP1/UAF19 and has been studied extensively. USP1/UAF1 can inhibit itself by autocleavage which happens upon exposure to UV-light9, which promotes DDT. It is thought that TLS polymerases are required to extend about 5-60 bp after the lesion30, after which a different USP1/UAF1 comes to deubiquitinate PCNA, allowing a replicative polymerase to take over again. Deubiquitination of K63-PCNA-UbN is less well established and it is still unclear what triggers formation of these chains on PCNA. While USP1/UAF1 activity on K63-linked chains on PCNA was reported once31, such a role has never been studied in detail.
Recent studies have shown that USP1 inhibition is effective in BRCA1/2 deficient tumors, as its inhibition is synthetically lethal with PARP inhibition in the absence of BRCA1/2. This can also help to prevent PARP resistance7,32,33. Interestingly, further analysis showed that absence of functional USP1 causes K48-linked chains to accumulate on PCNA that results in PCNA degradation by the proteasome. This K48 chain formation on PCNA is accomplished by UBE2K and RNF138 and requires monoubiquitination of PCNA by RAD6/RAD187. It is unclear whether other types of replication stress also trigger K48-linked ubiquitination of PCNA and under what circumstances K48-mediated degradation of PCNA occurs.
DUBs process ubiquitin chains using different cleavage patterns. They can cleave the entire chain off the substrate (base cleavage), cleave a ubiquitin-ubiquitin linkage in the middle of the chain (endo-cleavage), or start at the distal ubiquitin and processively cleave the chain one ubiquitin at a time (exo-cleavage). In context of PCNA ubiquitination, base cleavage immediately creates free PCNA, while endo- or exo-cleavage can leave either monoubiquitinated or polyubiquitinated PCNA, with potentially different effects on DDT.
Here we analyze the USP1/UAF1 cleavage mechanism on K63- and K48-linked polyubiquitinated PCNA to understand the cleavage preference and kinetics of this reaction. We find that USP1/UAF1 has a strong preference for exo-cleavage on polyubiquitinated PCNA. Kinetic modeling reveals that USP1/UAF1 is efficient at cleaving both K48- and K63-linked polyubiquitin chains on PCNA and prefers cleaving Ub-Ub bonds over the Ub-PCNA bond. By solving the cryo-EM structure of active full-length USP1/UAF1 covalently bound to a K63-linked diubiquitin chain, we show how USP1/UAF1 binds to K63-ubiquitin chains. Structure-function analysis reveals that UAF1 plays a minor role in USP1/UAF1’s exo-cleavage preference and that the preference for Ub-Ub is caused primarily by USP1. The result of this preference is relative enrichment of monoubiquitinated PCNA when polyubiquitination occurs.
Results
USP1/UAF1 prefers to cleave K63- and K48-PCNA-UbN using exo-cleavage
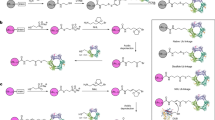
To study the cleavage mechanism of USP1/UAF1, we produced a variety of defined K63- or K48-ubiquitinated PCNA substrates (Fig. 1). We used different combinations of wildtype ubiquitin, ubiquitin K63R or K48R mutants, fluorescently labeled ubiquitin and activity-based ubiquitin probes. Together, these substrates help us to precisely follow deubiquitination of polyubiquitinated PCNA by USP1/UAF1. We used PCNA with defined monoubiquitination on K164, using established enzymatic modification34, and extended these to diubiquitin and/or longer ubiquitin chains with either K63 or K48 specific E2 enzymes and a chain-promoting E3 ligase, RNF8. For kinetic analysis we generated substrates using fluorescently labeled ubiquitin, where TAMRA (TMR) is covalently attached to an A28C-mutated ubiquitin. This allowed PCNA to be successfully modified with TMR-labeled UbA28C by the E1, E2s and E3 used in this study, which did not interfere with USP1/UAF1 cleavage (Supplementary Fig. 1).
Using different combinations of E2 and E3 enzymes, wildtype ubiquitin, ubiquitin mutants and the activity-based probe UbVAUb, we generated a toolbox that allowed us to study the cleavage mechanism of USP1/UAF1 on ubiquitin-modified PCNA at K164. UbA28C variants were fluorescently labeled with TAMRA (TMR) to allow quantitative analysis.
First, we aimed to determine whether USP1/UAF1 uses exo-, endo- or base-cleavage on PCNA, to characterize how USP1/UAF1 cleaves a ubiquitin chain. In this context, we define base-cleavage as cleaving a chain from PCNA through cleavage of the PCNA-Ub bond. We distinguish this from cleavage of a single ubiquitin from PCNA on monoubiquitinated PCNA (PCNA-Ub) and refer to the latter as mono-cleavage (Fig. 2A).
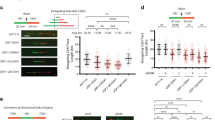
A Schematic of cleavage patterns on PCNA-UbN, PCNA-Ub2 and PCNA-Ub. Here we use terms as follows: Exo- and endo-cleavage both cleave a bond between two ubiquitins, whereas base-cleavage and mono-cleavage both cleave the bond between PCNA and ubiquitin. B−E SDS PAGE gels of USP1/UAF1 activity assays. B Conjugation of USP1/UAF1 to UbVAUb (complex of USP1 with a Ub2 chain). C Cleavage of K63-PCNA-UbVAUb (2 µM) results in conjugation of USP1/UAF1 (200 nM) to K63-PCNA-UbVAUb, seen as a large band shift. This indicates that USP1 performs exo-cleavage. D Fluorescence scan (left panel) and Coomassie-stained SDS PAGE gel (right panel) of K63-PCNA-UbVAUb-UbTMR (2 µM) and USP1/UAF1 (200 nM) reaction shows fast release of UbTMR. Subsequently, USP1 conjugates to PCNA-UbVAUb, only visible on the Coomassie stained gel. E DUB assay of USP1/UAF1 (200 nM) on K63-PCNA-UbN (2 µM, left panel) and K63-PCNA-Ub2 (2 µM, right panel). Assay shows rapid disappearance of chains, enrichment of PCNA-Ub and appearance of a free mono-ubiquitin, indicating that USP1/UAF1 prefers exo-cleavage on both longer K63-chains and diubiquitinated PCNA. F DUB assay of USP1/UAF1 (200 nM) on K48-PCNA-UbN (2 µM, left panel) and K48-PCNA-Ub2 (2 µM, right panel), shows that USP1 has an even stronger preference for exo-cleavage on K48-chains. BCDEF: Uncropped gels are provided as source data. All assays were repeated three times to similar results.
In order to study the USP1 cleavage mechanism on PCNA (Fig. 2A), we used a diubiquitin with a VME-like warhead that mimics the K63 linkage (K63-linked diubiquitin vinyl amide (VA), here termed UbVAUb)35, that conjugates to USP1 as it attempts to cleave the Ub-Ub bond (Fig. 2B, Fig. 1). We enzymatically attached this diubiquitin probe to PCNA’s K164 residue, aiming for one diubiquitin probe per PCNA trimer. This allows us to distinguish between cleavage of the PCNA-Ub bond (base-cleavage) or the Ub-Ub bond (exo-cleavage) by following the band shift after USP1 attempts to cleave K63-PCNA-UbVAUb. In these assays, we used a relatively high concentration of USP1/UAF1 (200 nM), to make sure the bands are clearly visible on gel. Addition of USP1/UAF1 results in conjugation to K63-PCNA-UbVAUb (Fig. 2C), revealing that USP1 performs exo-cleavage.
We then generated chains with 3 ubiquitins on PCNA by attaching a TMR-labeled UbK63R to K63-PCNA-UbVAUb. This yields K63-PCNA-UbVAUb-UbTMR, a substrate that allows distinction between exo-, endo- and base-cleavage by following the fluorescent signal. On this substrate, cleavage by USP1/UAF1 results in rapid formation of free UbTMR, consuming K63-PCNA-UbVAUb-UbTMR almost completely within 1 min (Fig. 2D), which can only happen if USP1/UAF1 uses an exo-cleavage mechanism. After UbTMR is cleaved off, USP1 continues to exo-cleave the chain, resulting in conjugation to K63-PCNA-UbVAUb as demonstrated by the same 40 kDa shift of USP1 as seen in Fig. 2C. We do not observe a band for fluorescent USP1 + PCNA-Ub3, which would indicate endo-cleavage, or a band expected for base- cleavage (either a free fluorescent Ub3 band, or a USP1+UbVaUb (as detected in 2B)). All labeled, cleaved ubiquitin appears as a free UbTMR, and USP1 conjugates to PCNA-Ub2, thus indicating exo-cleavage.
We then performed deubiquitination assays on native K63-linked polyubiquitinated PCNA (K63-PCNA-UbN) and K63-linked diubiquitinated PCNA (K63-PCNA-Ub2), using 200 nM of USP1/UAF1, to make sure the reaction runs to completion and follow USP1/UAF1 as it cleaves the entire substrate pool. On K63-PCNA-Ub2, USP1/UAF1 cleaves the ubiquitin-ubiquitin isopeptide-bond (exo-cleavage), as we can observe that PCNA-Ub and free monoubiquitin are being produced (Fig. 2E, right panel). Only when USP1/UAF1 has cleaved most PCNA-Ub2, it switches to mono-cleavage of PCNA-Ub, removing the last ubiquitin. USP1/UAF1 therefore does not cleave the PCNA-Ub bond when there are chains on PCNA, but does cleave it when chains have depleted and only PCNA-Ub is present.
On longer ubiquitin chains (K63-PCNA-UbN), USP1/UAF1 still appears to favor exo-cleavage over endo- and base-cleavage, as it primarily produces free monoubiquitin (Fig. 2E, left panel). There is a small degree of endo-cleavage since minor amounts of free ubiquitin chains of different lengths appear as well. Base-cleavage is unlikely, since PCNA-Ub accumulates at first and occurrence of base cleavage on PCNA-Ub2 is negligible (Fig. 2E, right panel). The free ubiquitin chains which are produced by endo-cleavage are cleaved by USP1/UAF1 as well and are completely consumed after 8 min, which is also when most K63-PCNA-UbN has been processed.
Interestingly, we also observed that USP1/UAF1 appears to preferentially cleave K63-PCNA-UbN over mono-cleavage of PCNA-Ub. This is demonstrated by the increase of PCNA-Ub until t = 4 min, while K63-PCNA-UbN gradually depletes. Only when most polyubiquitinated PCNA has depleted (+/− 4 min), USP1 starts to mono-cleave PCNA-Ub as seen by their decreased intensity from t = 8 min and onwards.
In the crosslinking experiment (Fig. 2D), we did not observe endo-cleavage as USP1 does not conjugate to PCNA-Ub3 (K63-PCNA-UbVAUb-UbTMR). Since crosslinking renders USP1 inactive immediately upon attempted endo-cleavage of this substrate, USP1/UAF1 seems to have a strong preference for exo-cleavage in the first turnover. To test this, we extended the longer polyubiquitin chains on PCNA (K63-PCNA-UbN) with a terminal fluorescently labeled TMR-UbK63R/A28C. This allows us to study the initial preference on longer ubiquitin chains on PCNA. Here, we also observe that after cleavage almost all fluorescence is found as a mono-ubiquitin, indicating that the UbTMR is cleaved off alone (Supplementary Fig. 2A). This verifies that USP1 has a strong preference for exo-cleavage when approaching K63-PCNA-UbN, especially in the first turnover. We observe the same effect when we add a TMR-UbK63R/A28C to a free K63-ubiquitin chain (K63-UbN), which shows that it is not PCNA, but the ubiquitin chain itself that requires USP1 to use exo-cleavage (Supplementary Fig. 2B).
When cleaving K48-linked chains on PCNA, USP1/UAF1 exclusively employs exo-cleavage (Fig. 2F) and even the minor amounts of endo-cleavage are missing. Chains shorten over time and only free ubiquitin is produced. As was observed for K63-PCNA-UbN USP1/UAF1 starts cleaving monoubiquitinated PCNA only after the majority of K48-linked chains has been processed, suggesting that USP1/UAF1 prefers cleaving Ub-Ub bonds over the Ub-PCNA bond in K48-linked chains as well. It is possible that the increased specificity for exo-cleavage is mediated by the more compact structure of K48-chains whereas the relative flexibility of K63-chains allows the occasional use of endo-cleavage36.
Designing a global kinetic model
In order to further characterize the cleavage mechanism of USP1/UAF1, and confirm our observations based on the DUB assays (Fig. 2, Supplementary Fig. 2), we set out to generate a global kinetic model. We used K63- and K48-PCNA-Ub2, where either the proximal or distal ubiquitin was fluorescently labeled, along with the monoubiquitinated variant (PCNA-UbTMR). These substrates allowed us to follow USP1/UAF1 activity in fluorescence polarization assays (Fig. 3A, Supplementary Fig. 3A, B). Please note that we used a lower concentration of USP1/UAF1 (10 nM) than in the assays shown in Fig. 2, to enable kinetic analysis. In addition, we studied binding affinities of USP1/UAF1 for PCNA-Ub, PCNA and free ubiquitin by MST analysis and determined the activity of USP1/UAF1 on a minimal substrate (Ub-Rho) (Supplementary Fig. 3C,D). All activity assays and the binding data were fitted simultaneously to a single global kinetic model using KinTek explorer (Fig. 3B, Table S1, Supplementary Fig. 4, Supplementary Fig. 5A, B).
A Fluorescence polarization assays of USP1/UAF1 (10 nM) activity comparing a single substrate concentration (4 µM) of PCNA-UbTMR, K63-PCNA-UbTMR-Ub and K63-PCNA-Ub-UbTMR and PCNA-UbTMR, K48-PCNA-UbTMR-Ub and K48-PCNA-Ub-UbTMR. The assays reveal that USP1/UAF1 prefers cleaving the ubiquitin-ubiquitin bond of both chain types. Full concentration range and repeats are shown in Supplementary Fig. 3A. B Schematic representation of the final kinetic model created with Kintek, describing deubiquitination of either K63 or K48 chains on PCNA. The model shows the potential paths taken by a single entity of USP1/UAF1 (shown as USP1). Written version of the kinetic model and accompanying calculated constants are shown in Supplementary Fig. 3. Since the off rate for PCNA-Ub2 (k1) is threefold lower than the off rate for PCNA-Ub (k5), USP1 prefers to proceed to another PCNA-Ub2 after cleaving and releasing the distal ubiquitin from PCNA-Ub2. USP1/UAF1 uses a distributive cleavage mechanism (bind, cleave, release) instead of a processive mechanism (bind, cleave, cleave again), as k2 is much higher than k4. Since the processive mechanism is barely used, it is visualized in the schematic using a blurry appearance. C Simulation generated by the kinetic model (B, D) shows cleavage of substrates and appearance of different products as USP1/UAF1 cleaves PCNA-Ub2. Its preferred cleavage mechanism and preference for PCNA-Ub2 leads to accumulation of PCNA-Ub. The simulation conditions mirror the fluorescence polarization assay shown in (A). D Constants describing the different reactions and equilibriums in the kinetic model. kon is diffusion limited and therefore the same for each reaction. k1, k2, k3, k4, k5, k6 are the off rates for each reaction and represent affinity in this model. Table S1 shows full table, including constants of the K48 model. Constants are derived from experimental data shown in Supplementary Fig. 3.E USP1/UAF1’s preference for exo-cleavage and cleaving Ub-Ub bonds gives it a preference for cleaving poly-ubiquitinated PCNA over monoubiquitinated PCNA and can result in temporal enrichment of PCNA-Ub.
To formulate an appropriate model (Fig. 3B, Table S1), we started by describing all options when cleaving PCNA-Ub2. In Supplementary Fig. 4, we show all the steps that were considered and tested to create the final model, in a process that is described in the methods and in the legend of Supplementary Fig. 5. For example, we started by keeping options open for exo- and base-cleavage. Endo-cleavage was excluded since PCNA-Ub2 does not contain endo-cleavable bonds. Only a model in which USP1/UAF1 was instructed to solely use exo-cleavage resulted in a good fit, which agrees with the exo-cleavage preference observed in our PCNA-Ub2 gel-based DUB assays (Fig. 2E, F).
To deconvolute the model, we separated dissociation constants (off-rates, the numbered constants in the model) from cleavage of the isopeptide bond (kcut) itself. As a result, kcut could be shared in all cleavage steps, as USP1 is always cleaving an isopeptide bond, regardless of substrate. On the other hand, the numbered constants (k) are unique, as they are product-dependent. In classical Michaelis-Menten kinetics, these two values together appear as kcat.
An additional important parameter was whether the reaction was processive (bind, cleave and immediately move to the next Ub on the chain), distributive (bind, cleave and release) or both. A model where USP1/UAF1 solely uses processive cleavage gave poor fit to the experimental data. We then wrote a model where USP1/UAF1 solely performs a distributive mechanism, with a bind, cleave and release for each cleavage event, which improved the fit. However, we obtained the best fit when USP1/UAF1 has the option to perform both distributive and processive cleavage. This was interesting because the constants calculated by the final model show that despite using the two mechanisms, USP1/UAF1 has a strong preference for the distributive cleavage mechanism (Table S1).
Important for this distributive cleavage model was that after cleavage of PCNA-Ub2, USP1/UAF1 immediately releases PCNA-Ub and remains bound to the cleaved-off distal ubiquitin, rather than to PCNA-Ub. Additionally, for this bind, cleave and release mechanism a proper fit could only be obtained if we explicitly instructed the model that once a substrate is released, USP1/UAF1 will not immediately bind and cleave that same substrate again. Released PCNA-Ub is absorbed in the general pool of available substrates and is treated equal to other ubiquitinated PCNAs present.
The full final model as written in KinTek is shown in Supplementary table 1 and a more schematic representation is shown in Fig. 3B. While our aim was to write separate kinetic models for K63- and K48-PCNA-Ub2, we found that the same final model could be applied to both. This means that USP1/UAF1 uses a similar approach cleaving these different chains on PCNA and as such, the following paragraphs regarding the kinetic model can be applied to either chain type.
USP1/UAF1 prefers to cleave ubiquitin-ubiquitin bond over PCNA-ubiquitin bond
In the schematic kinetic model (Fig. 3B, D) we show the path taken by a single entity of USP1/UAF1 as it cleaves PCNA-Ub2. We can follow this path by analyzing and comparing the reaction constants that are shown in the table in Fig. 3D. As mentioned previously, the model needed to accommodate the option for both distributive cleavage and processive cleavage mechanisms. However, comparison of the reaction constants for distributive (k2) and processive (k4) cleavage shows that USP1/UAF1 has a large (~ 170-fold) preference (k2: 932 ± 554 s-1, k4: 5.5 ± 3 s-1) for distributive cleavage, indicating that processive cleavage occurs rarely.
USP1/UAF1 starts by cleaving the distal ubiquitin from PCNA-Ub2, then first releases PCNA-Ub, followed by release of the distal ubiquitin. Subsequently, the same USP1/UAF1 will prefer to engage the next PCNA-Ub2 rather than PCNA-Ub, since the affinity derived from the off-rate shows that USP1/UAF1’s affinity for K63-PCNA-Ub2 (k1: 1244 s-1) is 2.5-fold higher than the affinity for PCNA-Ub (k5: 2812 s-1). Only when the pool of available PCNA-Ub2 depletes and the relative availability of PCNA-Ub increases, USP1/UAF1 will start cleaving PCNA-Ub as well. Also, after mono-cleavage of PCNA-Ub, USP1 first releases free PCNA before it releases the cleaved-off ubiquitin.
To generate a visual representation of this process, we used the kinetic model to simulate how the amounts of substrate and products vary over time (Fig. 3C). This simulation was run at concentrations that mimic the FP assay (Fig. 3C, Supplementary Fig. 6A, B), but could also be run at conditions that mimic the DUB assay (Fig. 2) to allow for visual comparison of the two (Supplementary Fig. 6C, D respectively). While chains on PCNA disappear rapidly, the amount of monoubiquitinated PCNA only declines when chains are depleted. This corresponds to our DUB assays (Fig. 2), where we observe how PCNA-Ub levels increase first and only decrease at a later stage. It demonstrates that the preference of USP1/UAF1 for cleaving PCNA-Ub2 can result in temporal enrichment of PCNA-Ub (Fig. 3E). Although the affinity for K48-PCNA-Ub2 is lower than for K63-PCNA-Ub2, as seen by its k1 value, the simulations show that the temporal enrichment of PCNA-Ub can also take place for K48-PCNA-Ub2.
Using the kinetic model, we can also calculate detailed reaction parameters (Supplementary Fig. 5C, D) and derive the catalytic efficiencies (kcat/KM) of the reaction (Table 1). We use Michaelis-Menten analysis for PCNA-Ub and distally labeled PCNA-Ub2. For the proximally labeled PCNA-Ub2 substrates however, Michaelis-Menten kinetics are not appropriate. USP1/UAF1’s preference for cleaving the distal ubiquitin causes a delay in cleavage of the proximal TMR-labeled ubiquitin, as shown in the kinetic simulation for PCNA-UbTMR-Ub (Supplementary Fig. 6A, B). We therefore use a substrate inhibition model to calculate the reaction rates (Supplementary Fig. 5C, D) for proximally labeled PCNA-Ub2 instead of the Michaelis-Menten model.
Similar to the kinetic model itself, the catalytic efficiencies (kcat/Km) of USP1/UAF1 for distally labeled K63- and K48-PCNA-Ub2 are comparable. These catalytic efficiencies show that USP1/UAF1 has a ~ 10-fold preference for cleaving Ub-Ub bonds (0.207 nM-1s-1 for K63-PCNA-Ub-UbTMR or 0.156 nM-1s-1 K48-PCNA-Ub-UbTMR) over the PCNA-Ub bond in PCNA-Ub2, (0.019 nM-1s-1 for K63-PCNA-UbTMR-Ub or 0.015 nM-1s-1 for K48-PCNA-UbTMR-Ub). The low catalytic efficiencies for proximally labeled PCNA reflect USP1/UAF1’s exo-cleavage and bind and release mechanism, as the enzyme would need to cleave the distal ubiquitin before we could detect the release of the proximal UbTMR (Fig. 3B, C, Supplementary Fig. 1E, F), resulting in these lower catalytic efficiencies. Cleavage of monoubiquitinated PCNA itself (PCNA-UbTMR: 0.090 nM-1s-1) is also less efficient compared with exo-cleavage of the distal ubiquitin.
Structure of USP1/UAF1 in complex with K63-linked diubiquitin
To gain structural insights into USP1/UAF1’s mechanism on ubiquitin chains and study why it prefers exo-cleavage, we performed cryo-EM single particle analysis on full length USP1/UAF1 crosslinked to the K63-linked diubiquitin VA probe warhead (UbVAUb). We obtained a reconstruction of the complex at 3.0 Å global resolution (Fig. 4, Fig. S8A).
A Domain architecture of USP1 and UAF1. Density was resolved for USP1’s insert 3 (dark gray), but not for insert 1 and insert 2 (light gray). B Final map of 3.0 Å final structure. DeepEMhancer sharpened map is colored as in A and shown at threshold 0.005. C DeepEMhancer sharpened map from (B), placed in the non-sharpened final density map (gray transparent map) shown at threshold 0.015 to accentuate density for flexible regions. D Atomic reconstruction of the structure of USP1/UAF1 bound to K63-linked UbVAUb. E Zoom showing the backbone of insert 3 of USP1 in the non-sharpened map shows how it anchors at the WD40 domain of UAF1.
The structure reveals how USP1/UAF1 binds the K63-linked diubiquitin chain (Fig. 3D). As expected for an exo-cleavage mechanism, the distal ubiquitin is placed in the USP1 hand, positioning its ubiquitin tail for catalysis in the active site (Supplementary Fig. 9A, B). The proximal ubiquitin is positioned at the base of the USP1 thumb in what looks like a binding pocket. They are linked by the VA-type warhead that is cross-linked to the catalytic cysteine (C90) of USP1 and links the distal and proximal ubiquitin (Supplementary Fig. 9A, B).
The overall structure of USP1/UAF1 is in concordance with existing USP1/UAF1 structures (7AY0, 7AY1, 7AY237 and USP1/UAF1 structures where USP1 is bound to an inhibitor (8A9K, 8A9J38, 9EBS))38,39. In Supplementary Fig. 8E, F, we show a superposition of our USP1/UAF1 structure to USP1/UAF1 structures that contain all UAF1 domains (7AY1, 8A9K, 8A9J, 9EBS)37,38,39 which shows that USP1/UAF1 adopts a highly similar conformation (Supplementary Fig. 8C). USP1 interacts with UAF1 primarily via the UAF1 β-propeller domain, through the tip of its fingers sub-domain, which is also seen for UAF1 that forms a complex with USP12 and USP4640,41,42,43.
Binding a diubiquitin chain induces the same minor conformational change in USP1’s fingers subdomain as was observed for USP1/UAF1 that binds a monoubiquitin37. Upon ubiquitin binding, the fingers close in on the distal ubiquitin to accommodate binding (Supplementary Fig. 8D). This movement also causes repositioning of UAF1, since it interacts with the tip of the USP1 fingers. Even between different USP1/UAF1 structures containing full length UAF1, there are minor differences in how UAF1 is positioned towards USP1. When we superpose UAF1 from these structures (7AY1, 9EBS, 8A9K, 8A9J) onto UAF1 from our structure we can observe that UAF1 maintains its internal structure with a rigid body movement that generates these minor differences (Supplementary Fig. 8E).
In this structure, USP1 D751 is facing the catalytic histidine (H593) (Supplementary Fig. 9C). We have previously shown that this aspartate is dispensable for catalysis and that D752 instead is critical for catalysis44. We built D752 to face the catalytic histidine, which follows the canonical positioning of this residue. However, there is also density to accommodate an alternative rotameric state, where D752 faces up towards N85 (Supplementary Fig. 9C). The alternative state of D752 was observed in one earlier structure (7AY137) but not in others37,38.
An unexpected feature in this structure is that insert 3 is partially ordered (residues 465 – 483), in contrast to previous structures of USP1/UAF1 (7AY0, 7AY1, 7AY237, 8A9K, 8A9J38, 9EBS39). It creates an additional interaction with the WD40 β-propeller domain of UAF1 (Fig. 4C, E). Since the resolution in this region was a bit lower than in the rest of the structure (Supplementary Fig. 8B), the actual positioning of the sidechains could differ. However, what does stand out is that the interface on UAF1 is made up of multiple lysines (K168, K203, K205, K207). These could potentially face and form interactions with multiple serines (S471, S472, S475) and a glutamate (E453) in USP1’s insert 3 to facilitate the interaction.
UAF1 plays a minor role in exo-cleavage preference for K63-PCNA-UbN
The structure suggests that positioning of UAF1 (Fig. 5A), may prevent cleavage via base- or endo-cleavage, especially the SLD and ancillary domain. These domains may need to move to accommodate distal ubiquitins if endo- or base-cleavage would occur. To test if these domains affect USP1/UAF1’s cleavage mechanism, we generated and purified deletion constructs and analyzed activity of USP1/UAF1ΔSLD (res. 9 – 563) and USP1/UAF1ΔAnc-SLD (res. 25-360) (Fig. 5B) on K63-PCNA-UbN and K48-PCNA-UbN. These deletions did not have a major effect on the thermal stability of the protein (Supplementary Fig. 10, supplementary Table S3).
A Left: USP1/UAF1 showing the subdomains of UAF1 (WD40, Ancillary domain, SLD) in different tints of blue. Right: Theoretical depiction of USP1/UAF1 binding the chain in an endo-cleavage mechanism shows how UAF1 SLD and Ancillary domains could obstruct binding the ubiquitin chain in the middle. B DUB assays on K63-PCNA-UbN shows that deletion variants USP1/UAF1ΔSLD (200 nM) and USP1/UAF1ΔAnc-SLD (200 nM) use a more flexible mixture between exo- and endo- cleavage. Furthermore, the reaction in USP1/UAF1ΔAnc-SLD is slower. Constructs used for deletion mutants are shown in lower panel. Please note that UAF1ΔAnc-SLD migrates at the same height as PCNA-Ub. Uncropped gels are provided as source data. Assays were repeated three times independently to similar results. C DUB assays of USP1 alone (1 µM) on K63-PCNA-UbN and K63-PCNA-Ub2 (left) show that USP1 employs a mix of exo- and endo-cleavage, but does not seem to cleave at the base as indicated by cleavage of K63-PCNA-Ub2. On K48-PCNA-UbN and K48-PCNA-Ub2 (right panel) USP1’s activity is even more reduced compared to K63-PCNA-UbN, but still uses exo-cleavage. Uncropped gels are provided as source data. Assays were repeated three times independently to similar results.
Whereas USP1/UAF1ΔSLD behaves like wild-type USP1/UAF1 in terms of activity, it uses a slightly more flexible mixture between exo- and endo-cleavage. Deletion of both the SLD and Ancillary domain together allows for an even more flexible mixture of exo- and endo-cleavage, as relatively more free ubiquitin chains are generated compared to USP1/UAF1WT. In addition, this mutation makes USP1/UAF1ΔAnc-SLD overall slower, and this decreased activity can be seen through K63-PCNA-UbN chains disappearing at a slower rate, and prolonged PCNA-Ub accumulation. While USP1/UAF1WT and USP1/UAF1ΔSLD have consumed most PCNA-Ub after 16 min, USP1/UAF1ΔAnc-SLD has consumed only roughly 40 percent. Thus, despite USP1/UAF1ΔAnc-SLD using a more flexible cleavage mechanism between endo- and exo-cleavage, it still does not use base-cleavage and retains its preference for cleaving Ub-Ub bonds over mono-cleaving PCNA-Ub, resulting in accumulation PCNA-Ub. Thus, despite opening up more space to accommodate ubiquitin chain binding for endo- or base- cleavage, these USP1/UAF1 variants mostly retain their preference for cleaving Ub-Ub bonds over the PCNA-Ub bond.
We then tested activity of USP1 without UAF1, to understand their relative contribution to the mechanism. These experiments required higher concentrations of USP1, as the enzyme alone is less active45,46. When cleaving K63-PCNA-UbN, USP1WT generates an equal mixture of free chains and free ubiquitin at t = 5 min. Analysis of USP1WT on K63-PCNA-Ub2 confirms that USP1 still opts for the Ub-Ub bond rather than the PCNA-Ub bond. USP1WT therefore uses a mix of exo- and endo-cleavage on K63-PCNA-UbN (Fig. 5C), but still does not opt for base-cleavage. For K48 chains USP1WT is hardly active and can only be seen to generate free ubiquitin. In line with previous findings8, the USP1 inserts may obstruct binding and activity on K48-chains without UAF1.
These results show that UAF1 or its Ancillary and SLD domain together, may prevent some endo-cleavage, presumably by steric hindrance. However, the preference for the Ub-Ub bond over the PCNA-Ub bond is caused by USP1 alone. The results also show that the UAF1 WD40-domain is sufficient for most of the activation of USP1, as USP1/UAF1ΔAnc-SLD is still significantly faster than USP1 alone. The UAF1 WD40-domain is therefore sufficient to resolve the autoinhibition from insert 1 and insert 3. The observed interaction of UAF1WD40 with insert 3 (Fig. 4E), could contribute to this. UAF1’s SLD domain alone does not appear to play a role in PCNA-Ub or PCNA-UbN deubiquitination, in contrast to its role for FANCI/FANCD2 deubiquitination47.
Hyperactivation of USP1 by mutating a conserved negative patch
The structure shows how the proximal ubiquitin is positioned in a pocket next to the USP1 thumb, which consists mostly of negatively charged residues (Fig. 6A–C). Two of these residues (D751 and D752) are well conserved active site residues and E754, which faces the proximal ubiquitin, is also well conserved. The two remaining residues in this patch (D172 and E173) are positioned in a different loop. These residues are well conserved in USP1 homologs in evolution (Fig. 6D), but are not conserved in USP paralogs, except for USP30 (Supplementary Fig. 11A, B).
A Electrostatic surface of USP1 indicates negatively charged binding patch close to the proximal ubiquitin. B Electrostatic depiction of interface between USP1 and the proximal ubiquitin. C Details of the negatively charged patch in USP1: D751, D752 are active site residues, and E754 supports their positioning. D172, E173 are on a separate loop. D Sequence alignment of USP1 orthologs shows that D172 and E173 are well conserved in USP1 homologs. Residues are colored by charge. E Fluorescence polarization assays comparing activity of D172/E173 to lysine (DE/KK) mutant to wildtype USP1 on PCNA-UbTMR, K63-PCNA-UbTMR-Ub and K63-PCNA-Ub-UbTMR. USP1DE/KK/UAF1 displays a major increase in activity on PCNA-Ub bonds (read out in PCNA-UbTMR and K63-PCNA-UbTMR-Ub) but only a minor increase on Ub-Ub bonds (readout in K63-PCNA-Ub-UbTMR).
Only four USP structures complexed with Ub2 are available for comparison (USP21 + M1-Ub2; 2Y5B48, USP30 + K6-Ub2; 5OHP49, CYLD K63-Ub2; 3WXG, CYLD + M1-Ub2; 3WVE50). USP30 however, is bound to a K6-linked diubiquitin and as such, the proximal ubiquitin is positioned differently with regards to the catalytic domain and the residues corresponding to D172 and E173 are not involved (Supplementary Fig. 11A, B). The catalytic domain of CYLD adopts a fold slightly different from the canonical USP catalytic domain fold. The proximal ubiquitin of M1-and K63-Ub2 is twisted and positioned differently from the proximal ubiquitin bound to USP1, and CYLD has a protruding B-sheet which interacts with the proximal ubiquitin, which is not found in USP150. USP21 interacts with an M1-Ub2 chain with a preference for base-cleavage and has the C-terminal tail of the proximal subunit positioned for cleavage, placing the distal ubiquitin beyond that, on the tip of the fingers subdomain.
To study the role of D172 and E173 in catalysis we mutated the DE to AA or KK respectively and compared their activity to USP1WT/UAF1. Strikingly, both mutants show gain-of function, a small increase in activity for USP1DE/AA/UAF1, and a more substantial increase for USP1DE/KK/UAF1 for cleavage of K63-PCNA-UbN (Supplementary Fig. 11D). Both mutants still primarily use exo-cleavage as evidenced by the production of free ubiquitin and the decrease in chain length of PCNA-UbN, similar to USP1WT/UAF1.
We tested the mutants on fluorescently labeled K63-PCNA-Ub2 substrates to identify on which bond of the ubiquitin chains the reaction is enhanced. Hyper-activation of USP1DE/KK/UAF1 occurs on PCNA-UbTMR, K63-PCNA-UbTMR-Ub and only to a minor extent on K63-PCNA-Ub-UbTMR (Fig. 6E). The increase in activity is therefore much more pronounced on the PCNA-Ub bond than on the Ub-Ub bond. This also means that a positive charge into this negative pocket does not interfere with Ub-Ub cleavage.
We wondered why evolution maintains the slower D172/E173 in USP1WT, and not the increased activity on the PCNA-Ub bond of the positively charged K172/K173. We therefore tested thermal stability of these mutants and surprisingly, USP1DE/KK/UAF1 shows a 5˚C increase in thermal stability over USP1WT/UAF1 (Table S3, Supplementary Fig. 11E). As stability was evidently not the evolutionary push that favors USP1WT over USP1DE/KK, these results suggest that there is a different evolutionary advantage to retain a suboptimal USP1 with lower activity for mono-cleavage of the PCNA-Ub bond. It is possible that this lower activity of USP1WT creates a relative enrichment of PCNA-Ub, which gives more opportunities for fine tuning the DNA damage tolerance pathways.
Discussion
In this research, we reveal that USP1/UAF1 cleavage of polyubiquitinated PCNA has three unusual aspects. (1) USP1/UAF1 prefers exo-cleavage and therefore trims chains on PCNA down. (2) USP1/UAF1 has a kinetic preference for Ub-Ub bonds over Ub-PCNA bonds. (3) USP1/UAF1 uses a bind, cleave and release mechanism when acting on PCNA-UbN. These three aspects together result in a preference for USP1/UAF1 to act on chains on PCNA rather than on mono-ubiquitinated PCNA, which causes a temporal enrichment of PCNA-Ub when polyubiquitinated alternatives are present.
With this kinetic mechanism in mind, we can speculate about a potential role of USP1/UAF1 in directing DNA damage tolerance (Fig. 7), in anticipation of in vivo experiments that are required to verify this model. When there is PCNA ubiquitination, in the absence of damage, USP1/UAF1 will effectively remove monoubiquitin. This is important as it prevents unscheduled onset of DDT. Upon fork stalling, when PCNA is monoubiquitinated and translesion synthesis takes place, USP1/UAF1 will deubiquitinate PCNA-Ub after the lesion has been bypassed. However, when both mono- and polyubiquitination take place, or when there only is polyubiquitination, the kinetic data suggests that USP1/UAF1 will primarily trim chains, which could temporarily enrich for monoubiquitinated PCNA. Such activity would protect against formation of chains on PCNA, and this could prevent unscheduled HR-mediated bypass or PCNA degradation, and promote TLS instead. One can imagine that when a lesion cannot be resolved by TLS or if bypass takes too long, prolonged stalling would result in formation of sufficient K63-linked chains, resulting in HR-mediated DDT. This would be further enabled by the auto-catalytic degradation removal of USP1, which occurs e.g. upon exposure to UV-light9.
In literature, it has been established that USP1/UAF1 deubiquitinates PCNA when no lesions are present, to prevent a premature start of DNA damage tolerance (DDT). However, when PCNA stalls at a DNA lesion, monoubiquitination at lysine 164 is required to initiate DDT, which can be promoted by autocleavage of USP1. Monoubiquitinated PCNA (PCNA-Ub) promotes translesion synthesis (TLS), a mechanism that can rapidly bypass DNA lesions to prevent replication fork stalling. Following successful bypass of the DNA lesion, USP1/UAF1 deubiquitinates PCNA-Ub to allow PCNA to resume with replication. Alternatively PCNA-Ub can undergo K63-polyubiquitination, which promotes HR-mediated bypass of DNA. Recently it has been shown that PCNA-Ub can undergo K48-polyubiquitination as well, leading to degradation of PCNA. In this study, we have shown that USP1 is highly efficient at cleaving both K63- and K48-polyubiquitinated PCNA, with a strong preference for exo-cleavage; trimming the ubiquitin chain one ubiquitin at a time. USP1/UAF1 preferentially cleaves polyubiquitinated PCNA over monoubiquitinated PCNA, thereby allowing temporary enrichment of PCNA-Ub. This mechanism of USP1/UAF1 could have several implications: USP1/UAF1 could protect PCNA against undesired K63-and K48-polyubiquitination, thereby safeguarding against unscheduled HR-mediated bypass or premature degradation of PCNA respectively. Furthermore, USP1/UAF1’s mechanism can cause a temporary enrichment of mono-ubiquitinated PCNA that could affect the balance between TLS and HR.
When replication impediments cause severe replication stress and cannot be resolved by either TLS or HR-like bypass, K48-ubiquitination and subsequent removal of PCNA would offer a solution. Followed by upstream PrimPol mediated repriming11 replication would still be able to continue, but would leave behind a ssDNA gap. In turn, these ssDNA gaps can be filled post replicative in a PCNA ubiquitination dependent manner12.
This DNA damage tolerance pathway choice is affected by many factors, but it will certainly be regulated by local concentrations of E3 ligases. Although USP1/UAF1 is efficient at deubiquitinating PCNA, its affinity for PCNA is relatively low8. Competition with the different relevant E3 ligases could shift the balance in DDT. Alternatively, the type of DNA damage could decide which pathway is required. TLS may rapidly bypass smaller DNA lesions, whereas bulky or complicated lesions, that do not fit in the TLS polymerase active site19,20, might require HR-mediated bypass. Additionally, since specific DNA lesions require specific TLS polymerases for bypass51,52, it is possible that HR-mediated bypass is important when the relevant TLS polymerase is not available.
The low affinity of USP1/UAF1 for PCNA suggests that it might be assisted by a different protein. Recent work has shown that ATAD-5 (ELG1), a subunit of the RFC PCNA unloader complex, plays a scaffolding role, interacting with USP1/UAF1 to boost PCNA deubiquitination53. The interaction between USP1 and ATAD-5 has been shown previously54, along with the importance of ATAD-5 for PCNA deubiquitination55,56. There are other potential proteins that can be involved like for example, USP1/UAF1s interplay with the SPRTN pathway57. Additionally, USP1/UAF1’s role in BRCA1 resistance33 and USP1 inhibition in BRCA1/2 deficient tumor resistance7,33 suggests that there are possible interactors in this pathway, like for example RAD51AP158.
Our findings of USP1/UAF1 as a kinetic regulator of different ubiquitin signals on PCNA is supported by the evolutionary conservation of two negatively charged residues (D172, E173) in USP1. Mutating these residues to positively charged lysines, generates a hyperactive USP1 which is enhanced for cleavage of the PCNA-Ub bond. Conservation of the sub-optimal situation suggests that this lowered activity is vital to USP1s role in fine-tuning DDT pathways.
Our structure also revealed a partially ordered insert 3 (residues 465 – 483), which together with insert 1 (residues 227 – 423), auto-inhibits USP18. UAF1 binding resolves this autoinhibitory effect, anchoring insert 3 at its WD40-domain. This binding of insert 3 to UAF1 could prevent collapse of insert 3 on blocking loop 1 and 2 (BL1 residues 536 −551, BL2 residues 564 – 592) in the USP1 catalytic domain. The structure shows multiple serines in insert 3 near the interface and these contacts could be important for USP1 activation by UAF1.
We showed that D752 is important in catalysis44 but the cryo-EM density map is ambiguous on its rotameric state, making it hard to interpret its role. It is possible that D752 is indeed flexible and fulfills a dual role. Facing the active site, D752 could support the other catalytic residues whereas its alternative conformation could reinforce the oxyanion hole stabilizing role of N85. However, since the active site is currently engaged with the VA-warhead, it is possible that the active site is trapped in an intermediate state and that this affects positioning of D752.
Only a few DUBs are described with a strong preference for exo-cleavage. Two DUBs associated with the proteasome use exo-cleavage of K48-linked chains. By exo-cleaving, they can fine tune chain length to protect against degradation and recycle ubiquitin59, as only chains of Ub3 and longer trigger degradation60. These are USP1461 and UCHL5 (UCH37)62,63. Mechanistically this is achieved by a structural component that allows binding of chains to only promote exo-cleavage61,64. Most other DUBs use a flexible mixture between exo- and endo-cleavage65, due to a requirement for binding longer chains65,66,67. Other DUBs use a mixture of cleavage mechanisms that depends on the chain type, like USP9X68. In USP1/UAF1, the SLD and ANC domains partially promote exo-cleavage by preventing binding in the middle of the chain, but no further affinity for long chains is observed.
There is a difference in spatial organization for K63- and K48-linked ubiquitin chains. Nevertheless USP1/UAF1 uses the same cleavage mechanism on both chain types. While processing the cryo-EM dataset, resolving density of the proximal ubiquitin proved difficult at first and required extensive processing. This suggests that there is some flexibility in the position of the proximal ubiquitin. This minor flexibility could explain why USP1 is able to occasionally perform endo-cleavage on K63-PCNA-UbN. K48-chains on the other hand adopt a more rigid conformation than K63-chains, which could explain why USP1/UAF1 is only able to use exo-cleavage on K48 polyubiquitinated PCNA.
The findings in this study reveal detailed, mechanistic insights into how USP1/UAF1 operates on polyubiquitinated PCNA in vitro. It is possible that these kinetic mechanisms play a role in in regulation of PCNA in DNA damage tolerance. With USP1 inhibitors currently undergoing clinical trials, understanding the different functions of USP1 becomes increasingly relevant. This study yields interesting insights into the role of USP1 in regulating ubiquitinated PCNA, by protecting against K63- or K48-PCNA-UbN and promoting mono- over polyubiquitination.
Methods
Constructs and mutagenesis
USP1 (pFastbac-HTb, N-terminal His-tag, res. 21-785, G670A + G671A9 variants (USP1D172A/E173A, USP1D172K/E173K)) were generated using site-directed mutagenesis. UAF1 (pFastbac1, N-terminal strep-tag, res. 9-677) and its deletion variants (UAF1ΔSLD (res. 9 – 563), UAF1ΔAnc-SLD (res. 25-360) were generated using 5’-phosphorylated primers and site-directed mutagenesis). USP1 and UAF1 expressed and purified for cryo-EM had short C-terminal extensions as described previously44. Enzymes used for in vitro ubiquitination assays were expressed using the following constructs: hUBA1 (pET3a-UBA1), UbCH5CS22R (pET28a-UbCH5CS22R), Ubc13/Mms2 (pGEX-UBC13, pET16b-MMS2), UBE2K (pETM30-GST-UBE2K1–155) and RNF8RING (pGEX6p1-RNF8RING(351-485)). Ubiquitin (pET3a-Ub) variants (UbA28C, UbK63R, UbK48R, UbA28C/K63R, UbA28C/K48R, UbL73P) were generated using site-directed mutagenesis.
Protein purification
USP1/UAF1 was copurified by coexpression of USP1 and UAF1 in Sf9 cells, using a protocol adapted from earlier work8. After lysis and centrifugation, we first performed step-affinity purification using streptactin-XT beads (IBA), followed by nickel-affinity and size-exclusion chromatography, which improved our final yield of purified protein. PCNA, ubiquitin and enzymes for in vitro ubiquitination were expressed and purified following previously described methods: UBA169, UbCH5CS22R70 and PCNA were purified34,69. RNF8RING following71. UBC13/MMS2 following72, UBE2K1-155 following73. Ubiquitin variants (UbA28C, UbK63R, UbK48R, UbA28C/K63R, UbA28C/K48R, UbL73P) were purified following34.
Purification of PCNA-Ub, PCNA-Ub2, PCNA-Ub3 and PCNA-UbN variants
Different variants of mono- or diubiquitinated PCNA were made using either unlabeled ubiquitin (Ub) or TAMRA (TMR)-labeled ubiquitin (UbTMR). For fluorescent labeling we made use of UbA28C, to generate three different types of TMR-labeled ubiquitin variants (UbTMR, UbK63R-TMR, UbK48R-TMR). The labeling procedure followed previously described methods74.
Monoubiquitinated PCNA (PCNA-Ub, PCNA-UbL73P and PCNA-UbTMR) was made enzymatically using UBA1 and UbCH5CS22R (UBE2D3)34, aiming to monoubiquitinate every monomer of PCNA. To generate the K63-PCNA-UbVAUb activity-based probe, the protocol was adapted, substituting K63-linked UbVAUb for Ub (35 UbiQ-087 UbiQ Bio B.V.), and performing the reaction at 4 °C for 45 min, so that on average only one of the monomers is modified with UbVAUb. Note that in Mulder et al., 2014, UbVAUb is termed diUb VME.
For producing K63-linked PCNA-Ub2 variants (K63-PCNA-Ub-Ub, K63-PCNA-UbTMR-Ub, K63-PCNA-Ub-UbTMR), 12 μM PCNA-Ub or PCNA-UbTMR was mixed with 32 μM UbK63R or UbK63R-TMR, 200 nM UBA1, 4 μM UBC13/MMS2 and 2 μM RNF8RING in reaction buffer (50 mM HEPES pH 7.5, 150 mM NaCl, 1 mM DTT, 5 mM MgCl2, 5 mM ATP). Diubiquitination was performed for 60 min at 37 °C to ensure that every monomer was diubiquitinated, after which PCNA-Ub-UbK63R was purified using anion-exchange and size exclusion as described for PCNA-Ub34.
K63-PCNA-UbVAUb-UbTMR, was produced using the same protocol, adding UbK63R-TMR to K63-PCNA-UbVAUb.
K48-linked PCNA-Ub2 variants (K48-PCNA-Ub-Ub, K48-PCNA-UbTMR-Ub, K48-PCNA-Ub-UbTMR) were produced in an analogous manner: 16 μM PCNA-Ub or PCNA-UbTMR was mixed with 100 μM UbK48R or TMR-labeled UbK48R and 20 μM E2-25K1-155, 10 μM RNF8RING for 60 min while shaking at 500 rpm at 37 °C. The sample was purified following the steps described for other ubiquitinated PCNA variants.
K63-PCNA-UbN and K48-PCNA-UbN were generated using the same methods as their PCNA-Ub2 counterparts, using 100 μM WT ubiquitin. K63-PCNA-UbN-UbTMR was produced using this method as well, adding 100 μM UbK63R-TMR to 8 μM purified K63-PCNA-UbN.
K63-UbN was produced by mixing 50 μM UbWT with 0.5 μM UBA1, 0.5 μM UBC13/MMS2 and 0.5 μM RNF8RING in reaction buffer (50 mM HEPES pH 7.5, 150 mM NaCl, 1 mM DTT, 5 mM MgCl2, 5 mM ATP). Reaction was performed overnight at 37 °C, after which K63-UbN was purified using cation-exchange on a POROS-XS column (Thermofisher). UbN-UbTMR was produced using the same method, adding 100 μM UbK63R-TMR to 10 μM purified K63-UbN.
K63-UbVAUb conjugation assays
Ubiquitin probe crosslinking analysis was performed by mixing enzyme and substrate to final concentrations of 200 nM USP1/UAF1 and 2 μM PCNA-UbVAUb-UbTMR (trimer), 2 μM PCNA-UbVAUb (trimer) or 6 µM UbVAUb. Enzyme and substrate were prepared at 2x concentration in reaction buffer (20 mM Hepes pH 7.5, 150 mM NaCl, and 2 mM DTT). Before mixing, 7.5 μl substrate and 7.5 μl enzyme were separately added to 5 μL 4x SDS-page loading buffer as t0. 15 μl samples were taken at the indicated time points. Samples were loaded on a NuPAGE 4-12 % Bis-Tris SDS gel, and were separated by running for 30 min at 160 V. The gel containing PCNA-UbVAUb-UbTMR was imaged for fluorescent TMR signal using a Typhoon FLA 9000 biomolecular imager (Cytiva), after which all gels were stained using Coomassie.
PCNA-Ub, PCNA-Ub2 and PCNA-UbN deubiquitination assays by SDS page analysis
DUB activity assays were performed at RT, unless stated otherwise, in reaction buffer consisting of 20 mM Hepes, pH 7.5, 150 mM NaCl, and 2 mM DTT. 200 nM USP1/UAF1 was prepared at a 2x concentration (400 nM USP1/UAF1). Different ubiquitinated PCNA variants (0.67 μM of PCNA-Ub, 0.67 μM PCNA-Ub2 or 1.33 μM PCNA-UbN (trimer concentration)) were prepared at 2x concentration as well (1.33 μM for PCNA-Ub/PCNA-Ub2 and 2.67 μM for PCNA-UbN). Before mixing, 7.5 μl substrate and 7.5 μl enzyme were separately added to 5 μl 4x SDS-page loading buffer as t0. Reaction was initiated by mixing remaining enzyme and substrate 1:1, after which 15 μl samples were taken at the indicated time points. Samples were loaded on a NuPAGE 4-12% Bis-Tris SDS gel and bands were separated by running for 30 min at 160 V. Whenever TMR-labeled substrates were used, gels were first imaged using a Typhoon FLA 9000 biomolecular gel scanner (Cytiva) to visualize the TMR signal, after which the gel was stained with Coomassie. Coomassie blue stained gels were imaged using a Gel Doc EZ imaging system (Bio-Rad Laboratories, Inc.). For assays testing USP1 without UAF1 the same protocol was followed, except the reaction was performed at 37 °C with final concentrations of 1 µM USP1, 6 µM PCNA-UbN or 4 µM PCNA-Ub2, and 5 µL timepoints were taken as indicated.
Fluorescence Polarization activity assays
For fluorescence polarization (FP) activity assays, TMR-labeled mono- or diubiquitinated variants of PCNA were used (PCNA-UbTMR, PCNA-UbTMR-UbK63R, PCNA-Ub-UbK63R-TMR, PCNA-UbTMR-UbK48R, PCNA-Ub-UbK48R -TMR). All reactions were performed in a 384-well plate (Corning, flat bottom, low flange) on a PHERASTAR plate reader (BMG Labtech), using a 540/590/590 nm filter block (BMG Labtech) at 25 °C. The FP-buffer consisted of 20 mM Hepes, pH 7.5, 150 mM NaCl, 5 mM DTT and 0.05% Tween-20.
Single-point assays were performed using 10 nM USP1/UAF1 and 1 μM of ubiquitinated PCNA, prepared separately at 2x concentration in FP-buffer. 10 μl of enzyme was pipetted to the plate in triplicates. To initiate the reaction, 10 μl of substrate was pipetted to the plate, after which measurement was started immediately. Release of UbTMR could be measured by the increase in fluorescence polarization.
For the global kinetic fit analysis, 10 nM USP1/UAF1 was prepared at 2x concentration in FP-buffer. USP1/UAF1 was tested against a substrate dilution series ranging from 4 μM to 0.5 μM, prepared at a 2x concentration. 10 µl of each substrate concentration from the dilution series was pipetted to a 384-well plate in triplicate. The experiment was started by injecting 10 µl of 2x USP1/UAF1 to each well using the built-in syringe of the instrument. The first datapoint was 26 s after injection, with subsequent datapoints taken every 14 s.
Microscale thermophoresis
For MST analysis, the USP1/UAF1 was fluorescently labeled at random sites. For this, all cysteines in USP1/UAF1 were first reduced by adding 5 mM DTT to 12 µL final volume containing 20 µM USP1WT/UAF1WT or USP1KK/UAF1, followed by incubation for 10 min at room temperature, after which the sample was buffer exchanged to 20 mM Hepes, 150 mM NaCl using a desalting column (Zeba Spin 40 K MWCO, 75 µl, Thermofisher). Then 12 µL USP1/UAF1 was mixed with 40 µM of fluorescent label (DY-547P1-maleimide, Dyomics) and incubated for 30 min at room temperature. 5 mM DTT was added to inactivate the non-ligated DY-547P1, after which the sample was buffer exchanged to 20 mM Hepes (pH 7.5), 150 mM NaCl, 2 mM DTT to get rid of non-ligated fluorescent label.
Binding analysis of monoubiquitinated PCNA made use of non-cleavable UbL73P75. 5 nM labeled USP1/UAF1 was tested against a dilution series of 250 µM to 1.91 nM of PCNA, 250 µM to 7.63 nM UbWT or 62.5 µM to 1.91 nM of PCNA-UbL73P75, each prepared at 2-fold higher concentration in 20 mM Hepes pH 7.5, 150 mM NaCl, 2 mM DTT, 0.05% Tween. 15 µl of enzyme was mixed with 15 µl of different substrate concentrations, loaded into microscale thermophoresis (MST) capillaries in duplicates and inserted into the MST system (NanoTemper). Two biological replicates were performed for each MST experiment. Data was analyzed and affinities were derived using NanoTemper’s internal software.
Kintek/Michaelis Menten analysis
The data from the FP activity assay and MST were analysed using global kinetic modeling using KinTek Explorer Software (KinTek Corporation). A comprehensive enzymatic kinetic model was constructed to study USP1/UAF1 activity across all tested substrates. These included anisotropy readouts from FP assays for substrates PCNA-UbTMR, K63-PCNA-UbTMR-Ub, K63-PCNA-Ub-UbTMR, K48-PCNA-UbTMR-Ub, K48-PCNA-Ub-UbTMR. The modeling software only requires three substrate concentrations. This allowed removal of concentrations that had less optimal repeats for kinetic modeling. Additionally, MST measurements of USP1/UAF1 binding to PCNA, UbWT and PCNA-UbL73P were used as input to the model as well. Previously published data on enzymatic cleavage of minimal substrate Ub-Rho by USP1/UAF144, were also used for fitting of the model.
The initial model incorporated all possible reaction pathways, including base-cleavage and exo-cleavage, as well as all potential intermediates in the enzymatic process (Supplementary Fig. 4). Following substrate cleavage, USP1 can first recognize PCNA and then the cleaved Ub, or vice versa. To connect the different sets of data in one model several assumptions were made. In fluorescence polarization (FP) activity assays, PCNA substrates were treated as monomers with a single ubiquitination site, as the readout was based on the release of free UbTMR. On-rate constants kon were fixed at diffusion-limited values (108 M-1s-1). The isopeptide bond cleavage rate (kcut) was set as uniform across all reaction steps. Off-rate constants (k2 to k6) were unified for structurally similar substrates and products, regardless of labeling. For example, koff was identical for K63-PCNA-Ub-UbTMR and K63-PCNA-UbTMR-Ub. All kinetic parameters were shared between K63- and K48-labeled substrates, except for k1 which are not shared between K63-PCNA-Ub2 and K48-PCNA-Ub2.
After constructing the initial model, an iterative fitting process was applied, attempting to simplify the model. In each step, kinetic constants that failed to converge or had a value of zero were removed, along with their associated reaction steps. If elimination worsened the fit, the parameter was reintroduced. This refinement continued until no further simplification was possible.
The final model describes two distinct pathways following distal Ub cleavage: distributive and processive (Table S1, Fig. 3B). It consists of seven kinetic parameters, which were tested for correlations using FitSpace (Supplementary Fig. 12). The cleavage rate (kcut) correlated with the dissociation constants of PCNA-Ub-Ub and PCNA-Ub (k1 and k5), suggesting that their relative ratio is more critical than their absolute values.
Using this model, we simulated the enzymatic cleavage of PCNA-Ub-Ub, tracking changes in substrate, product, and PCNA-Ub intermediate levels for FP assays (Fig. 3C) and also for DUB assays (Fig. 2E). Additionally, the model allows simulations under Michaelis-Menten conditions for each substrate, enabling the calculation of catalytic efficiency (kcat/KM) (Table 1).
Cryo-EM sample preparation
USP1/UAF1 (final 4 µM) was conjugated to K63-linked UbVAUb (UbiQ Bio B.V.) (final 16 µM) by incubating for 30 min at 37 °C in reaction buffer (20 mM HEPES pH 7.5, 150 mM NaCl, 2 mM TCEP) to generate 4 µM (0.75 mg/ml) of complex. Grids (Cu 300 Holey Carbon R1.2/1.3 (Quantifoil Micro Tools GmbH)) were glow-discharged for 1 min at 30 mA using a GloQube Plus (Quorum) just before use. 3 μL of sample was applied to the grid and vitrified using a Vitrobot (Mark IV chamber (Thermo Fisher Scientific)) maintained at 4 °C and 100% humidity.
Cryo-EM data collection
Data was collected at NeCEN using a 300 kV Titan Krios electron microscope (Thermo Fisher Scientific) with a K3 BioQuantum direct electron detector (Gatan). EPU-2.10.0 was used for automated data collection in super-resolution mode using AFIS centering. 5833 movies of 50 frames were acquired in super resolution mode at 0.418 Å/pixel size (Counted super-resolution) with a total dose of 50 e−/Å2 (14 e−/pixel/s). Defocus values ranged between −1.2 μm and −2.4 μm.
Cryo-EM data processing
Data were imported and processed using CryoSparc76. Movies were binned 2x and motion corrected using patch motion correction (default settings), after which they were further processed using patch CTF estimation (default settings). An initial Topaz77 deep learning particle picking model was trained using manually picked particles, picked on a 100-micrograph subset. Particles were cleaned up using 2D classification and ab initio reconstruction and were used to train a new Topaz model. This Topaz model was then applied to the full dataset. The resulting particles were cleaned up using 2D classification, ab initio reconstruction and 3D classification, followed by NU-refinement. At this point, the particles were subjected to two different methods in parallel: (1): Iterative training of a new Topaz model and (2): Template picking using 2D projections of the NU-refined model.
-
1)
The NU-refined particles were used to train a new Topaz model, after which particles were cleaned up again. This process was repeated until eventually an improved Topaz model was generated. These were extracted with 4x binning (original box size: 360 pix). Particles were cleaned up using 2D classification and ab initio reconstruction, after which they were re-extracted using a 280-pixel box size. A 1 class 3D classification was run to generate a single 3D volume from all particles. This was followed by NU-refinement, global and local CTF refinement and second NU-refinement. This resulted in a 3D density map with improved density for insert 3, which was not present in the 3D volume generated using approach (2).
-
2)
2D projections of the NU-refined model were used for Cryosparc’s template picker performed on the entire dataset and were extracted using a 512-pixel box size (binned to 256 pix). 2D classification was used to remove non-particles. Particles were then cleaned up using subsequent rounds of ab initio reconstruction with three classes, each time picking the best class. The resulting class was used to train a new Topaz model, and particles were extracted with 352-pixel box size (binned to 176 pix). Ab initio classes were generated, whose particles and 3D density maps were used for heterogeneous refinement, which was repeated iteratively until an improved 3D density map was generated. These particles were re-extracted in a 352-pixel box size without binning, followed by NU-refinement. This resulted in a final model with a better resolved SLD compared to approach (1).
Particles of approach (1) and (2) were extracted at 360 pix and combined, removing only 2812 particles and therefore greatly increasing the number of particles. Since a 10-class 3D classification resulted in highly similar 3D volumes, the particles were combined into a single model using 3D classification and a mask covering the entire complex. The resulting 3D volume was NU-refined using the same mask, followed by local and global CTF refinement and another NU-refinement. This final 3D volume of combined particle stacks (1) and (2) had better SLD, insert 3 and proximal Ub density and improved local resolution. Until this point the density for the proximal ubiquitin was ambiguous and made it difficult to precisely place the proximal ubiquitin, suggesting some minor flexibility. We therefore used this refinement for 3D flex78. This map was used for local refinement, with a mask covering the entire complex and the fulcrum coordinates set on the proximal ubiquitin (xyz: −33.8, −2.9, 57.4). We generated a tight mask on the entire complex and ran a final NU refinement, which improved the density for the proximal ubiquitin even more. This yielded the final model, which was sharpened using DeepEMhancer79.
Model building
The final cryo-EM density and DeepEMhancer sharpened map were used to build the structure using real-space refinement in Phenix, PDB-Redo and manual jiggle fitting in Coot80,81,82. We started by rigid body fitting the full length AlphaFold-2 models83 of USP1 (uniprot ID: O94782) and UAF1 (uniprot: Q8TAF3), from the EBI database, and two copies of the experimental structure of ubiquitin (PDB ID: 1UBQ) into our final density map using ChimeraX84. The fitted model was real-space refined using Phenix80, after which the fit was adjusted in Coot by jiggle fitting and manual rebuilding. We used Coot to generate and fit the 4-aminobutyric acid linker (termed ABU in structure) in the active site with restraints according to ABU. The model was submitted to PDB-REDO82, together with the restraints file for ABU, which improved the overall model. This model was examined using Coot, after which we manually corrected remaining model geometry issues, class score and rotamer outliers. This model was submitted for one more round of Phenix real-space refinement. The output model was checked a final time in Coot, correcting for the few remaining geometry and rotamer outliers. This final model was validated using the PDB-validation tool, after which it was submitted to the PDB and EMDB. Figures including the cryo-EM model were made using USCF ChimeraX.
Structure based sequence alignment
To compare predicted structures from all human USPs, predictions from the EBI Alphafold-2 database83 were used. All predicted structures were superimposed onto USP1’s Alphafold-2 structure, using the matchmaker function from UCSF Chimera85. In turn, this superposition was used for structure-based sequence alignment, using Match -> align function in Chimera. Since the output showed that except for USP1 and USP30, this loop is not found in any of the other USPs and could therefore not be shown.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Cryo-EM reconstruction and structure coordinates are available on the Electron Microscopy Database (EMDB) and to the PDB under the following accession code: PDB: 9HNW, EMD: 52316. Source data are provided with this paper.
References
Al-Hakim, A. et al. The ubiquitous role of ubiquitin in the DNA damage response. DNA Repair 9, 1229 (2010).
Hershko, A. & Ciechanover, A. The ubiquitin system. Annu. Rev. Biochem. 67, 425–479 (1998).
Komander, D. & Rape, M. The ubiquitin code. Annu. Rev. Biochem. 81, 203–229 (2012).
Nijman, S. M. B. et al. A genomic and functional inventory of deubiquitinating enzymes. Cell 123, 773–786 (2005).
Komander, D., Clague, M. J. & Urbé, S. Breaking the chains: structure and function of the deubiquitinases. Nat. Rev. Mol. Cell Biol. 10, 550–563 (2009).
Cadzow, L. et al. Development of KSQ-4279 as a first-in-class USP1 inhibitor for the treatment of BRCA-deficient cancers. Eur. J. Cancer 138, S52 (2020).
Simoneau, A. et al. Ubiquitinated PCNA drives USP1 synthetic lethality in cancer. Mol. Cancer Ther. 22, 215–226 (2023).
Dharadhar, S., van Dijk, W. J., Scheffers, S., Fish, A., Sixma, T. K. Insert L1 is a central hub for allosteric regulation of USP1 activity. EMBO Rep. 22, e51749 (2021).
Huang, T. T. et al. Regulation of monoubiquitinated PCNA by DUB autocleavage. Nat. Cell. Biol. 8, 341–347 (2006).
Nijman, S. M. B. et al. The deubiquitinating enzyme USP1 regulates the fanconi anemia pathway. Mol. Cell. 17, 331–339 (2005).
Bainbridge, L. J., Teague, R. & Doherty, A. J. Repriming DNA synthesis: an intrinsic restart pathway that maintains efficient genome replication. Nucleic Acids Res. 49, 4831–4847 (2021).
Nusawardhana, A., Pale, L. M., Nicolae, C. M. & Moldovan, G. L. USP1-dependent nucleolytic expansion of PRIMPOL-generated nascent DNA strand discontinuities during replication stress. Nucleic Acids Res. 52, 2340–2354 (2024).
Piberger, A. L. et al. PrimPol-dependent single-stranded gap formation mediates homologous recombination at bulky DNA adducts. Nat. Commun. 11, 1–14 (2020).
Tirman, S. et al. Temporally distinct post-replicative repair mechanisms fill PRIMPOL-dependent ssDNA gaps in human cells. Mol. Cell. 81, 4026–4040.e8 (2021).
Hoege, C., Pfander, B., Moldovan, G. L., Pyrowolakis, G. & Jentsch, S. RAD6-dependent DNA repair is linked to modification of PCNA by ubiquitin and SUMO. Nature 419, 135–141 (2002).
Bienko, M. et al. Ubiquitin-binding domains in y-family polymerases regulate translesion synthesis. Science (1979) 310, 1821–1824 (2005).
Kannouche, P. L., Wing, J. & Lehmann, A. R. Interaction of human DNA polymerase η with monoubiquitinated PCNA: a possible mechanism for the polymerase switch in response to DNA damage. Mol. Cell. 14, 491–500 (2004).
Lehmann, A. R. et al. Translesion synthesis: Y-family polymerases and the polymerase switch. DNA Repair 6, 891–899 (2007).
Biertümpfel, C. et al. Structure and mechanism of human DNA polymerase η. Nature 465, 1044 (2010).
Kirouac, K. N. & Ling, H. Unique active site promotes error-free replication opposite an 8-oxo-guanine lesion by human DNA polymerase iota. Proc. Natl. Acad. Sci. USA. 108, 3210–3215 (2011).
Tsaalbi-Shtylik, A. et al. DNA mismatch repair controls the mutagenicity of Polymerase ζ-dependent translesion synthesis at methylated guanines. DNA Repair 142, 103755 (2024).
Parker, J. L. & Ulrich, H. D. Mechanistic analysis of PCNA poly-ubiquitylation by the ubiquitin protein ligases Rad18 and Rad5. EMBO J. 28, 3657–3666 (2009).
Unk, I. et al. Human SHPRH is a ubiquitin ligase for Mms2-Ubc13-dependent polyubiquitylation of proliferating cell nuclear antigen. Proc. Natl. Acad. Sci. USA. 103, 18107–18112 (2006).
Unk, I. et al. Human HLTF functions as a ubiquitin ligase for proliferating cell nuclear antigen polyubiquitination. Proc. Natl. Acad. Sci. USA. 105, 3768–3773 (2008).
Krijger, P. H. L. et al. HLTF and SHPRH are not essential for PCNA polyubiquitination, survival and somatic hypermutation: existence of an alternative E3 ligase. DNA Repair 10, 438–444 (2011).
Gallina, I. et al. The ubiquitin ligase RFWD3 is required for translesion DNA synthesis. Mol. Cell. 81, 442 (2021).
Stelter, P. & Ulrich, H. D. Control of spontaneous and damage-induced mutagenesis by SUMO and ubiquitin conjugation. Nature 425, 188–191 (2003).
Chiu, R. K. et al. Lysine 63-polyubiquitination guards against translesion synthesis–induced mutations. PLoS Genet 2, e116 (2006).
Sale, J. E., Lehmann, A. R. & Woodgate, R. Y-family DNA polymerases and their role in tolerance of cellular DNA damage. Nat. Rev. Mol. Cell Biol. 13, 141–152 (2012).
Fujii, S. & Fuchs, R. P. Defining the position of the switches between replicative and bypass DNA polymerases. EMBO J. 23, 4342–4352 (2004).
Yang, X. H. & Zou, L. Dual functions of DNA replication forks in checkpoint signaling and PCNA ubiquitination. Cell Cycle 8, 191–194 (2009).
Cadzow, L. et al. The USP1 inhibitor KSQ-4279 overcomes PARP inhibitor resistance in homologous recombination–deficient tumors. Cancer Res. 84, 3419–3434 (2024).
Lim, K. S. et al. USP1 Is required for replication fork protection in BRAC1-deficient tumors. Mol. Cell. 72, 925–941.e4 (2018).
Hibbert, R. G. & Sixma, T. K. Intrinsic flexibility of ubiquitin on proliferating cell nuclear antigen (PCNA) in translesion synthesis. J. Biol. Chem. 287, 39216–39223 (2012).
Mulder, M. P. C., El Oualid, F., Ter Beek, J. & Ovaa, H. A native chemical ligation handle that enables the synthesis of advanced activity-based probes: diubiquitin as a case study. Chembiochem 15, 946 (2014).
Ye, Y. et al. Ubiquitin chain conformation regulates recognition and activity of interacting proteins. Nature 492, 266–270 (2012).
Rennie, M. L., Arkinson, C., Chaugule, V. K., Toth, R. & Walden, H. Structural basis of FANCD2 deubiquitination by USP1−UAF1. Nat. Struct. Mol. Biol. 28, 356–364 (2021).
Rennie, M. L., Arkinson, C., Chaugule, V. K. & Walden, H. Cryo-EM reveals a mechanism of USP1 inhibition through a cryptic binding site. Sci. Adv. 8, 6353 (2022).
Simoneau, A. et al. Characterization of TNG348: a selective, allosteric USP1 inhibitor that synergizes with PARP inhibitors in tumors with homologous recombination deficiency. Mol. Cancer Ther. 24, 678−691 (2025).
Dharadhar, S., Clerici, M., van Dijk, W. J., Fish, A. & Sixma, T. K. A conserved two-step binding for the UAF1 regulator to the USP12 deubiquitinating enzyme. J. Struct. Biol. 196, 437–447 (2016).
Li, H. et al. Allosteric activation of ubiquitin-specific proteases by β-propeller proteins UAF1 and WDR20. Mol. Cell. 63, 249–260 (2016).
Yin, J. et al. Structural insights into WD-repeat 48 activation of Ubiquitin-specific protease 46. Structure 23, 2043–2054 (2015).
Zhu, H., Zhang, T., Wang, F., Yang, J. & Ding, J. Structural insights into the activation of USP46 by WDR48 and WDR20. Cell Discov. 5, 1–4 (2019).
Keijzer, N. et al. Variety in the USP deubiquitinase catalytic mechanism. Life Sci. Alliance. 7, e202302533 (2024).
Cohn, M. A. et al. A UAF1-Containing multisubunit protein complex regulates the fanconi anemia pathway. Mol. Cell. 28, 786–797 (2007).
Cohn, M. A., Kee, Y., Haas, W., Gygi, S. P. & D’Andrea, A. D. UAF1 is a subunit of multiple deubiquitinating enzyme complexes. J. Biol. Chem. 284, 5343–5351 (2009).
Yang, K. et al. Regulation of the Fanconi anemia pathway by a SUMO-like delivery network. Genes Dev. 25, 1847–1858 (2011).
Ye, Y. et al. Polyubiquitin binding and cross-reactivity in the USP domain deubiquitinase USP21. EMBO Rep. 12, 350–357 (2011).
Gersch, M. et al. Mechanism and regulation of the Lys6-selective deubiquitinase USP30. Nat. Struct. Mol. Biol. 24, 920 (2017).
Sato, Y. et al. Structures of CYLD USP with Met1- or Lys63-linked diubiquitin reveal mechanisms for dual specificity. Nat. Struct. Mol. Biol. 22, 222–229 (2015).
Friedberg, E. C., Lehmann, A. R. & Fuchs, R. P. P. Trading places: how do DNA polymerases switch during translesion DNA synthesis?. Mol. Cell. 18, 499–505 (2005).
Paniagua, I. & Jacobs, J. J. L. Freedom to err: the expanding cellular functions of translesion DNA polymerases. Mol. Cell. 83, 3608–3621 (2023).
Ryu, E. et al. ATAD5 functions as a regulatory platform for Ub–PCNA deubiquitination. Proc. Natl. Acad. Sci. USA. 121, e2315759121 (2024).
Lee, K. Y. et al. Human ELG1 regulates the level of ubiquitinated proliferating cell nuclear antigen (PCNA) through its interactions with PCNA and USP1. J. Biol. Chem. 285, 10362–10369 (2010).
Bell, D. W. et al. Predisposition to cancer caused by genetic and functional defects of mammalian Atad5. PLoS Genet 7, e1002245 (2011).
Kim, S. et al. Polyubiquitinated PCNA triggers SLX4-mediated break-induced replication in alternative lengthening of telomeres (ALT) cancer cells. Nucleic Acids Res 52, 11785–11805 (2024).
Coleman, K. E. et al. USP1-trapping lesions as a source of DNA replication stress and genomic instability. Nat. Commun. 13, 1–19 (2022).
Cukras, S. et al. The USP1-UAF1 complex interacts with RAD51AP1 to promote homologous recombination repair. Cell Cycle 15, 2636 (2016).
Lee, M. J., Lee, B. H., Hanna, J., King, R. W. & Finley, D. Trimming of ubiquitin chains by proteasome-associated deubiquitinating enzymes. Mol. Cell. Proteom. 10, R110.003871 (2011).
Kiss, L., James, L. C. & Schulman, B. A. UbiREAD deciphers proteasomal degradation code of homotypic and branched K48 and K63 ubiquitin chains. Mol. Cell. 85, 1467–1476.e6 (2025).
Hu, M. et al. Structure and mechanisms of the proteasome-associated deubiquitinating enzyme USP14. EMBO J. 24, 3747 (2005).
Lam, Y. A., Xu, W., DeMartino, G. N. & Cohen, R. E. Editing of ubiquitin conjugates by an isopeptidase in the 26S proteasome. Nature 385, 737–740 (1997).
Yao, T. et al. Proteasome recruitment and activation of the Uch37 deubiquitinating enzyme by Adrm1. Nat. Cell Biol. 8, 994–1002 (2006).
Burgie, S. E., Bingman, C. A., Soni, A. B. & Phillips, G. N. Structural characterization of human Uch37. Proteins 80, 649 (2011).
Arif, S. et al. Mechanism of activation and regulation of deubiquitinase activity in MINDY1 and MINDY2. Mol. Cell. 81, 4176–4190.e6 (2021).
Wendrich, K. et al. Discovery and mechanism of K63-linkage-directed deubiquitinase activity in USP53. Nat. Chem. Biol. 21, 746–757 (2024).
Hermanns, T. et al. A family of unconventional deubiquitinases with modular chain specificity determinants. Nat. Commun. 9, 799 (2018).
Paudel, P. et al. Crystal structure and activity-based labeling reveal the mechanisms for linkage-specific substrate recognition by deubiquitinase USP9X. Proc. Natl. Acad. Sci. USA. 116, 7288–7297 (2019).
Hibbert, R. G., Huang, A., Boelens, R. & Sixma, T. K. E3 ligase Rad18 promotes monoubiquitination rather than ubiquitin chain formation by E2 enzyme Rad6. Proc. Natl. Acad. Sci. USA. 108, 5590–5595 (2011).
Brzovic, P. S., Lissounov, A., Christensen, D. E., Hoyt, D. W. & Klevit, R. E. A UbcH5/ubiquitin noncovalent complex is required for processive BRCA1-directed ubiquitination. Mol. Cell. 21, 873–880 (2006).
Mattiroli, F. et al. RNF168 ubiquitinates K13-15 on H2A/H2AX to drive DNA damage signaling. Cell 150, 1182–1195 (2012).
Pickart, C. M. & Raasi, S. Controlled synthesis of polyubiquitin chains. Methods Enzymol. 399, 21–36 (2005).
Pichler, A. et al. SUMO modification of the ubiquitin-conjugating enzyme E2-25K. Nat. Struct. Mol. Biol. 12, 264–269 (2005).
Dharadhar, S., Kim, R. Q., Uckelmann, M. & Sixma, T. K. Quantitative analysis of USP activity in vitro. Methods Enzymol. 618, 281–319 (2019).
Békés, M. et al. DUB-resistant ubiquitin to survey ubiquitination switches in mammalian cells. Cell Rep. 5, 826 (2013).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Bepler, T. et al. Positive-unlabeled convolutional neural networks for particle picking in cryo-electron micrographs. Nat. Methods 16, 1153–1160 (2019).
Punjani, A. & Fleet, D. J. 3DFlex: determining structure and motion of flexible proteins from cryo-EM. Nat. Methods 20, 860–870 (2023).
Sanchez-Garcia, R. et al. DeepEMhancer: a deep learning solution for cryo-EM volume post-processing. Commun. Biol. 4, 1–8 (2021).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D. Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. Biol. Crystallogr. 66, 486–501 (2010).
Joosten, R. P., Long, F., Murshudov, G. N. & Perrakis, A. The PDB_REDO server for macromolecular structure model optimization. IUCrJ 1, 213–220 (2014).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Pettersen, E. F. et al. UCSF ChimeraX: Structure visualization for researchers, educators, and developers. Protein Sci. 30, 70 (2021).
Meng, E. C., Pettersen, E. F., Couch, G. S., Huang, C. C. & Ferrin, T. E. Tools for integrated sequence-structure analysis with UCSF chimera. BMC Bioinforma. 7, 1–10 (2006).
Acknowledgements
We thank personnel of NeCEN and Xiaohu Guo from the NKI cryo-EM facility for assistance in data collection. We thank Andrea Murachelli and Shun Hsiao Lee for help with cryo-EM data processing and Robbie Joosten for helping with model building. The authors would like to acknowledge the Research High Performance Computing (RHPC) facility of the Netherlands Cancer Institute (NKI) which was used for cryo-EM data processing. This work is part of the Oncode Institute, which is partly financed by the Dutch Cancer Society. Research at the Netherlands Cancer Institute is supported by institutional grants of the Dutch Cancer Society and the Dutch Ministry of Health, Welfare and Sport. This study was supported by NWO grants OCENW.KLEIN.131 (T.K.S), TOP714.016.002 (T.K.S.) and Health Holland grant LSHM21048-H045 (T.K.S. & F.E.O.), and by the European Union’s Horizon Europe research and innovation programme under the Marie Skłodowska-Curie grant agreement No 101072903 (T.K.S).
Author information
Authors and Affiliations
Contributions
Conceptualization: NK, TS, FEO. Design: NK, FEO, AF. Execution: NK, NvL, JS, KM. Analysis: NK, JS, AF. Writing: NK, TS, with input from JS, KM, NvL, FEO.
Corresponding author
Ethics declarations
Competing interests
F.E.O. declares competing financial interests as co-founder and shareholder of UbiQ Bio BV. NK, JS, KM, NvL, AF and TS declare no competing interest.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Keijzer, N., Sakoltchik, J., Majumder, K. et al. USP1/UAF1 targets polyubiquitinated PCNA with an exo-cleavage mechanism that can temporarily enrich for monoubiquitinated PCNA. Nat Commun 16, 6991 (2025). https://doi.org/10.1038/s41467-025-61768-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-61768-0
This article is cited by
-
Bruceine A protects nuclear receptor 4A1 from ubiquitin-degradation to alleviate mesangial proliferative glomerulonephritis
Signal Transduction and Targeted Therapy (2025)