Abstract
Telomeres pose challenges during replication, with converging forks unlikely to resolve issues. Depleting TRF1 results in fragile telomeres, yet its exact role in telomere replication remains unclear. In our cellular model, insufficient TRF1 density at long telomeres leads to telomere fragility that is alleviated by restoring telomeric TRF1 levels. Our findings indicate that TRF1 mitigates lagging strand telomere fragility through fork reversal in a process involving telomerase activity, rather than merely alleviating fork barriers. Additionally, TFIIH, a crucial partner of TRF1, aids in restarting replication on the leading strand after fork reversal. When fork reversal is compromised, PrimPol-mediated repriming rescues fragility at leading strand telomeres, revealing a new role for this enzyme in human telomere replication. Lastly, our findings indicate that the TRF1-mediated decrease in telomere fragility is dependent on RNA:DNA hybrids, likely facilitating fork restart.
Similar content being viewed by others
Introduction
Human chromosome ends consist of approximately 10–15 kilobases of TTAGGG duplex repeats, ending with a 3′ overhang of about 100 to 300 nucleotides. The shelterin complex, composed of TRF1, TRF2, RAP1, TIN2, TPP1, and POT1, binds telomeres and fulfills important functions, such as inhibiting DNA repair pathways or facilitating telomeric DNA replication1. Telomeric DNA sequences are indeed hard-to-replicate regions prone to replication fork stalling due to their highly repetitive nature and various obstacles that impede fork progression, such as G-quadruplexes (G4s), persistent RNA:DNA hybrids, protein-DNA associations, i-loops or t-loops2. The replication-related defects observed in mammalian telomeres resemble those seen in aphidicolin-induced common fragile sites. These defects manifest as abnormal FISH signals on metaphase chromosomes, making them easily detectable using FISH approaches2,3. As telomere replication generally initiates from a single origin in the subtelomeric region and proceeds unidirectionally towards the chromosome end, careful regulation is required to manage potential replication fork stalling at telomeres, given that converging distal forks for rescue are unlikely4. Stalled replication forks can regress into four-way structures called reversed forks, which are thought to be a common response to replication stress, mediated by the coordinated action of various enzymes, including DNA translocases such as SMARCAL1, ZRANB3, or HLTF5. Fork reversal has been documented at both yeast and mammalian telomeres6,7,8,9, underscoring the essential role of SMARCAL1 in telomere maintenance—a role not shared by ZRANB3 or HLTF10.
In addition to fork reversal, other mechanisms, such as repriming and translesion DNA synthesis, also play a role in restarting compromised replication forks in the genome11. Aberrant processing of stalled replication forks can lead to one-ended double-strand breaks (DSBs) that might be repaired through the break-induced replication (BIR) mechanism12. All these processes have been observed at telomeres, particularly in ALT-positive (ALT+) cells, which maintain telomeres independently of telomerase through BIR-like recombination-dependent mechanisms13,14,15. Translesion DNA polymerases, such as Polη, also play crucial roles in managing replication stress at ALT telomeres16.
Telomeric DNA replication requires the action of several enzymes to prevent or recover from fork stalling, including shelterin proteins, nucleases, helicases, and DNA repair factors2. The telomere replication-promoting activity of the mammalian shelterin complex depends mainly on TRF1 and is proposed to rely on its ability to recruit DNA helicases, such as BLM, WRN, and RTEL13,17,18,19. In turn, RTEL1 helicase interacts with TRF2 to promote t-loop disassembly in S-phase20. Recently, TFIIH has been identified as a new factor recruited by TRF1 to promote both leading and lagging telomere replication, although the underlying mechanism remains unknown21.
Several studies have suggested that telomere replication stress correlates with telomere length22,23,24. Telomere elongation, through ectopic telomerase overexpression in human embryonic stem cells, induces fragile telomeres22. Although the underlying causes of this increased fragility were not investigated, this phenomenon could be due to an increased density of G4 structures at excessively long telomeres25. Alternatively, aligning with the idea that cellular levels of the fission yeast TRF1 homolog, Taz1, are likely insufficient to cover the entire length of the long telomeres in rap1Δ cells26, long mammalian telomeres might be less effectively covered by shelterin. Supporting this hypothesis, it has been suggested that the endogenous levels of TRF1 in mouse embryonic fibroblasts (MEFs) may be insufficient to fully prevent replication issues at telomeres, explaining why TRF1 overexpression in wild-type MEFs reduces the baseline level of fragile telomeres21.
In this study, we aim to further elucidate the mechanisms used by human cells to replicate long telomeres and the role of TRF1 in this process. To tackle this issue, we analyzed isogenic cell lines which exhibit different endogenous telomerase expression and vary in telomere length. We found increased levels of replication stress specifically at longer telomeres, which we linked to TRF1 undersaturation. Restoring TRF1 levels at telomeres alleviated fragility through mechanisms involving fork reversal and telomerase activity. Surprisingly, we additionally observed that TRF1’s ability to suppress telomere fragility depends on RNA:DNA hybrids, highlighting a beneficial role of RNA:DNA hybrids in maintaining telomere stability. These findings collectively indicate that TRF1 primarily facilitates fork repair mechanisms instead of removing replication barriers. Finally, our work revealed a role for PrimPol-mediated repriming in alleviating fragility at leading strand telomeres when fork reversal is compromised.
Results
Telomere replication defects are linked to telomere length
To explore whether telomere length affects replication stress at human telomeres without using the non-physiological approach of ectopic telomerase overexpression, we took advantage of isogenic post-crisis survivor clones with different telomere lengths. Briefly, eight post-crisis survivor clones (PCS-1 to PCS-8) were isolated after seeding pre-crisis IMR90 cells expressing the E6 and E7 oncogenes from human papillomavirus. These clones exhibited varying levels of telomerase activity (Fig. 1A) upon spontaneous reactivation of telomerase (Supplementary Fig. 1A). Clones PCS-1, PCS-2, and PCS-7 showed higher expression of hTERT mRNA and longer telomeres (Supplementary Fig. 1A, B). Whole genome sequencing analysis of PCS clones identified a translocation event between the hTERT gene promoter and the AHRR locus, located 815 kb from the hTERT gene, near to the 5p telomere (Supplementary Fig. 1C). The hTERT promoter rearrangement was detected in all PCS clones, alongside the presence of wild-type hTERT promoter sequences (Supplementary Fig. 1C). This suggests the emergence of a single survivor from IMR90E6/E7 cell crisis, followed by the isolation of eight colonies during the procedure.
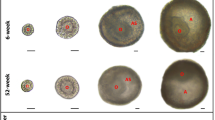
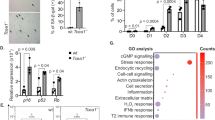
A Telomerase activity assessed by ddTRAP in PCS clones, normalized to PCS-8. SW39/TEL+ and IMRB/ALT+ cells (grey bars) were used as positive and negative controls, respectively. n = 2 independent biological replicates, except for PCS-7 and SW39 (n = 1). Mean ± SEM. Purple: clones with long telomeres; Cyan: clones with short telomeres. B TRF analysis (n = 1), with PCS-2 designated as PCSLT and PCS-3 designated as PCSST. C qRT-PCR analysis of hTERT expression, normalized to ACTB expression and to PCSST. n = 3 independent biological replicates, except for PCSLT cells (n = 4). Mean ± SEM. D Left: Representative FISH pictures on metaphase spreads from PCSST and PCSLT cells showing fragile telomeres (arrowheads). Right: Fragile telomeres with U2OS/ALT+ and IMRB/ALT+ cells as controls for ALT+ cells. A minimum of 5000 chromosome ends were analyzed per condition, based on n metaphase spreads collected from three independent biological replicates. Boxplot: Min, 1st quartile, Median, 3rd quartile, and Max. Kruskal–Wallis test. E Representative blot (above) and quantification (below) for C-circle assay (CCA) in PCSST and PCSLT cells, with SW39/TEL+ and IMRB/ALT+ cells as negative and positive controls, respectively. Signals were normalized to PCSST. n = 3 (SW39/TEL+ and IMRB/ALT+) or n = 4 biological replicates (PCSST and PCSLT). Mean ± SEM. Two-tailed ratio paired t-test. F TRF analysis (above) and CCA blot (below). PD population doubling. n = 1. G Fragile telomeres assessed by FISH on metaphase spreads. A minimum of 2200 chromosome ends were analyzed, based on n metaphase spreads collected from one experiment for PCSST-hTERT at PD9 and PD78, two independent biological replicates for PCSST-hTERT at PD18-21 and PD86-89, and three independent biological replicates for PCSST and PCSLT. Boxplot: Min, 1st quartile, Median, 3rd quartile, and Max. Kruskal–Wallis test. Source data are provided as a Source Data file.
For our study, we selected the PCS-2 clone with long telomeres (~ 11 kb), designated as PCSLT, and the PCS-3 clone with shorter telomeres (~ 4 kb), designated as PCSST (Fig. 1B and Supplementary Fig. 1B). Although the reasons for varying hTERT expression among PCS clones with the same promoter rearrangement are unclear, the PCSLT clone displayed more activation marks (H3Ac) on the rearranged hTERT promoter compared to the PCSST clone (Supplementary Fig. 1D). Importantly, the hTERT expression levels in PCSLT and PCSST mirror those observed in cancer cells that spontaneously reactivated telomerase, such as, respectively, HT1080 or HeLa cells (Fig. 1C).
The conventional approach to detect replication stress at telomeres involves observing abnormal telomeric structures in metaphase chromosomes, such as multiple signal foci, using a telomeric FISH probe2. Our FISH analysis indicated that longer telomeres in PCSLT show a higher incidence of fragile telomeres (Fig. 1D). However, the fragility observed was less pronounced compared to U2OS/ALT+ or IMRB/ALT+ cells (Fig. 1D) which, as previously described, exhibit heightened levels of replication stress and telomere fragility linked to a conservative BIR-like mechanism used to maintain telomeres in the absence of telomerase13,14,27,28. Over-elongated telomeres experiencing replication stress have been previously associated with the formation of C-circle (CC)-like molecules22,24. Using the CC assay (CCA)29, we observed that PCSST cells tested negative, whereas PCSLT cells exhibited detectable signals, albeit at lower levels compared to IMRB/ALT+ cells used as positive controls (Fig. 1E).
Next, to determine if the replication stress observed in PCSLT cells is related to the presence of long telomeres, we elongated the telomeres of PCSST cells by transducing them with a construct that enables hTERT overexpression (Supplementary Fig. 1E). As expected, telomerase overexpression gradually lengthened PCSST telomeres (Fig. 1F) and PCSST-hTERT cells became CCA-positive at PD40 (Fig. 1F). Additionally, these cells exhibited increased telomere fragility as telomere length increased (Fig. 1G), consistent with an impact of telomere length on fragility.
These findings highlight the value of the isogenic PCSLT/PCSST model cell lines for studying responses to telomeric replication stress.
PCSLT cells do not engage into BIR and ALT activities
Because PCSLT telomeres share features with ALT telomeres, such as telomere fragility and CCA positivity, both of which are outcomes of BIR activity at ALT telomeres28,30, we explored whether PCSLT cells undergo BIR to maintain their telomeres.
To assess the activation of BIR at PCSLT telomeres, we first compared fragility levels detected using FISH and CO-FISH techniques. Since BIR involves conservative DNA replication, fragility associated with BIR cannot be detected using CO-FISH, where newly replicated strands are degraded13. About 7–8% of telomeres showed fragility when analyzed using FISH in both U2OS/ALT+ and IMRB/ALT+ cells (Fig. 2A). In contrast, CO-FISH detected fragility in only 2–3% of telomeres in these cells (Fig. 2A). Conversely, telomere fragility in PCSLT cells was not reduced when evaluated by CO-FISH compared to FISH (Fig. 2A), indicating the absence of BIR-dependent replication at PCSLT telomeres. Additionally, CO-FISH revealed fragility in both leading and lagging strand telomeres of PCSLT cells (Fig. 2B).
A Fragile telomeres assessed by either FISH (blue) or CO-FISH (black) on metaphase spreads. FISH data are from Fig. 1D. For CO-FISH, fragility was the average fragility from both the leading and lagging strands. U2OS/ALT+ and IMRB/ ALT+ cells were used as controls for ALT+ cells. A minimum of 3800 chromosome ends were analyzed, based on n metaphase spreads collected from three independent biological replicates. Boxplot: Min, 1st quartile, Median, 3rd quartile, and Max. Two-tailed unpaired t-tests with Welch’s correction if needed. B Left: Representative CO-FISH pictures on metaphase spreads from PCSLT cells showing lagging (green) and leading (red) telomere fragility (arrowheads). Right: Lagging (green) and leading (black) telomere fragility. A minimum of 5000 chromosome ends were analyzed, based on n metaphase spreads collected from three independent biological replicates. Boxplot: Min, 1st quartile, Median, 3rd quartile, and Max. Two-tailed Mann–Whitney tests. C Representative blots (above) and quantification (below) for CCA in siPOLD3-treated PCSLT cells, using U2OS/ALT+ cells as control. Signals were normalized to siLuc. n = 5 (U2OS/ALT+) or n = 6 (PCSLT) independent biological replicates. Mean ± SEM, Two-tailed ratio paired t-tests. D Left: Representative images for Telo-FISH (red)/IF targeting BLM (cyan) and PML (green) in PCSLT cells, with U2OS/ALT+ cells as positive control. Arrowheads indicate PML-associated telomeres bound by BLM. Right: Percentage of PML-associated telomeres co-localizing with BLM per nucleus. n nuclei were analyzed from three independent biological replicates. Median in red, quartiles in orange. Two-tailed Mann–Whitney test. E Above: Representative CO-FISH pictures on metaphase spreads from IMRB/ALT+ cells showing T-SCE events (arrowheads). Below: T-SCEs in PCSST and PCSLT, with U2OS/ALT+ and IMRB/ALT+ as positive controls. n metaphases were analyzed from three independent biological replicates. Median in red, quartiles in orange. Kruskal–Wallis test. F Above: Representative images from native Telo-FISH targeting ssC-rich telomeric DNA (red) in U2OS/ALT+ and IMRB/ALT+ cells. Below: ssTeloC signals in PCSST and PCSLT, with U2OS/ALT+ and IMRB/ALT+ cells as positive controls. n nuclei were analyzed from three independent biological replicates. Median in red, quartiles in orange. Kruskal–Wallis test. Source data are provided as a Source Data file.
CC production in ALT+ cells is associated with POLD3-dependent telomere elongation during BIR30. Although, as expected, depletion of POLD3 reduced CC levels in U2OS/ALT+ cells, it had no impact on CC levels in PCSLT cells, providing additional evidence of the absence of BIR-related telomeric DNA synthesis in these cells (Fig. 2C and Supplementary Fig. 2A).
The co-localization of telomeres with PML bodies, known as ALT-associated PML bodies (APBs), serves as an additional marker of ALT+ cells15. In ALT+ cells, BIR takes place in PML bodies during the G2/M phase and is detected using the ALT telomere synthesis in APBs (ATSA) assay30. Although APB frequency was elevated in PCSLT cells compared to PCSST (Supplementary Fig. 2B), ATSA assay confirmed that PCSLT cells do not engage the BIR mechanism observed in ALT+ cells for telomere elongation as rare EdU+ telomeric signals were observed in PCSLT PML bodies (Supplementary Fig. 2C). In contrast, U2OS/ALT+ cells demonstrated active telomeric DNA synthesis in APBs during G2 phase (Supplementary Fig. 2C). In U2OS/ALT+ cells, consistent with the role of BLM helicase in telomere clustering and ALT telomere synthesis within APBs30,31,32, BLM co-localized with APBs (Fig. 2D). Conversely, BLM was not associated with PML body-localized telomeres in PCSLT cells (Fig. 2D). While PML is also required for CC production in ALT+ cells31, depletion of PML only slightly decreased C-circle levels in PCSLT cells (Supplementary Fig. 2D, E) suggesting those C-circle signals and APBs differ from the ones observed in ALT+ cells. Additionally, other ALT markers, such as telomeric sister chromatid exchanges (T-SCEs)15 and single-stranded C-rich telomeric DNA (ssTeloC)31,33, did not show increased levels in PCSLT cells compared to PCSST (Fig. 2E, F).
Overall, our results indicate that, although PCSLT cells exhibit some ALT-like characteristics, their telomeres do not undergo BIR. These findings align with previous studies demonstrating that telomerase-expressing cell lines with long telomeres exhibit a low frequency of APB-like foci and extra-chromosomal telomeric repeats, including CC22,23,24. However, PCSLT cells appear to be predisposed to ALT, as evidenced by increased PML and BLM recruitment to PCSLT telomeres upon ATRX depletion, and induction of ssTeloC signals (Supplementary Fig. 2F–I).
PCSLT telomeres exhibit replication-related damages
PCSLT telomeres do not undergo BIR but show fragility. Even if their molecular nature remains unknown, fragile telomeres are an indicator of replication defects at telomeres. In addition to BIR, single-strand DNA (ssDNA) gaps, that form uncondensed regions, or internal telomeric loops -both associated with replication stress- have been proposed to contribute to fragile telomeres28,34. To further investigate chronic telomeric replication stress in PCSLT cells, we therefore examined the presence of various markers linked to replication stress-induced DNA damage and repair responses. RPA32-S33 phosphorylation, a reliable marker of replication stress induced by ATR activation35, was significantly higher in the PCSLT cells compared to PCSST (Fig. 3A). The ssDNA binding protein RPA acts as an initial responder to stalled replication forks, and its phosphorylated form can regulate the formation of RAD51 filaments36. Supporting the elevated replication stress at PCSLT telomeres, we observed an increased recruitment of both RAD51 and RPA proteins to the telomeres (Fig. 3B). Moreover, RAD51 depletion in PCSLT cells led to an increase in their CC levels (Supplementary Fig. 3A, B) and reduced cell viability more significantly than in PCSST cells (Fig. 3C). Increased levels of RPA and pRPA, along with the telomeric recruitment of RPA and RAD51, have also been observed in PCSST-hTERT cells at later PD, further reinforcing the link between replication stress and long telomeres (Supplementary Fig. 3C, D). Finally, consistent with FANCM’s role as a key suppressor of telomeric DNA replication stress37,38, FANCM depletion significantly increased CC signal intensity and telomere fragility on both strands in these cells (Fig. 3D, E and Supplementary Fig. 3E).
A Above: Western blot of cell extracts from PCSST and PCSLT cells, probed for pRPA32-S33, RPA32, and Actin. Below: pRPA32-S33 (purple) and RPA32 (green) levels, normalized to Actin and to PCSST. Representative of n = 4 independent biological replicates. Mean ± SEM. Two-tailed ratio paired t-tests. B Left: Representative images for Telo-FISH (red)/IF experiments targeting RAD51 (cyan) and RPA (green) in PCSST and PCSLT cells. Right: Co-localization events between telomeres and RAD51 (cyan) or RPA (green) in PCSST and PCSLT. n nuclei were analyzed from three independent biological replicates. Median in red, quartiles in orange. Two-tailed Mann–Whitney tests. C Cellular viability of PCSST and PCSLT cells treated with siRAD51. 72 h after transfection in 6-wells plate, cells were counted using Burker slide. n measurements from three independent biological replicates and technical replicates for PCSST (n = 6) and PCSLT (n = 13). Mean ± SEM. Two-tailed unpaired t-test. D Representative blot (above) and quantification (below) of CCA in siFANCM-treated PCSLT cells. Signals were normalized to siLuc. n = 8 independent biological replicates. Mean ± SEM. Two-tailed ratio paired t-test. E Lagging (green) and leading (black) telomere fragility assessed by CO-FISH in siFANCM-treated PCSLT cells. A minimum of 8300 chromosome ends were analyzed, based on n metaphase spreads collected from three independent biological replicates. Boxplot: Min, 1st quartile, Median, 3rd quartile, and Max. Two-tailed unpaired t-test with Welch’s correction for the lagging strand comparison and two-tailed Mann–Whitney test for the leading strand comparison. F Left: Alkaline gel electrophoresis of ClaI-digested genomic DNA from PCSST and PCSLT cells. Ethidium bromide (EtBr) staining is shown as loading control. Right: Quantification of alkaline gel signals using the G-rich probe. The radioactive signal intensity below 6 kb was normalized to the total lane signal. The normalized signal from PCSST cells was set to 1 for comparison. n = 3 independent biological replicates. Mean ± SEM. Two-tailed ratio paired t-test. G 2D-gel analysis with telomeric probe showing i-loops (red arrows). Ethidium bromide staining is shown as loading control. U2OS/ALT+ cells were used as positive control. n = 1. Source data are provided as a Source Data file.
ssDNA nicks and gaps are intricately associated with replication stress, functioning as both triggers and manifestations of this stress. Accordingly, alkaline gel electrophoresis showed an increase in shorter telomeric DNA fragments when probing with either G-rich or C-rich probes in PCSLT cells (Fig. 3F and Supplementary Fig. 3F). This suggests the accumulation of SSBs at PCSLT telomeres and supports the fragility observed on both strands. Additionally, i-loops, linked to the formation of t-circles in response to nicks and ssDNA gaps34, were detected at PCSLT, but not PCSST, telomeres (Fig. 3G).
Collectively, these findings indicate that PCSLT telomeres undergo chronic replication stress, resulting in the accumulation of ssDNA gaps and increased fragility at the telomeres.
Removal of RNA:DNA hybrids increases PCSLT telomere fragility
Our next objective was to identify the source of fragility at PCSLT telomeres. Earlier research has documented that ectopic TERRA expression disrupts telomere replication and increases telomere fragility through RNA:DNA hybrid formation, a phenomenon that can be mitigated by RNaseH1 (RH1) overexpression39. Initially, we assessed the levels of telomeric (UUAGGG) RNA:DNA hybrids in PCSST and PCSLT cells using DNA:RNA Immunoprecipitation (DRIP)40. Our results showed that telomeric RNA:DNA hybrids tended to be more abundant at PCSLT telomeres (Fig. 4A). We therefore tested whether removing telomeric RNA:DNA hybrids through RH1 overexpression could reduce the observed telomere fragility in PCSLT cells. Unexpectedly, however, while RH1 overexpression in PCSLT cells effectively eliminated these hybrids (Fig. 4B and Supplementary Fig. 4A, B), it markedly augmented telomere fragility on the leading strand (Fig. 4C). In contrast, overexpression of a catalytically inactive (RH1-CD) enzyme showed no such effect (Fig. 4C). Consequently, the recruitment of both RAD51 and RPA at PCSLT telomeres was enhanced after RH1 overexpression, but not following RH1-CD overexpression (Fig. 4D).
A DNA:RNA immunoprecipitation (IP) analysis using S9.6 antibody in extracts from PCSST and PCSLT-EV cells. IP and input samples were analyzed by qPCR with primer sets amplifying telomeric DNA. Values in IP were normalized first to input and then to PCSLT-EV. Values obtained for control experiments with IgG were subtracted. n = 3 independent biological replicates. Mean ± SEM. Two-tailed ratio paired t-test. B Western blot of cell extracts from PCSLT cells transduced with pLHCX empty vector (EV), pLHCX-RNaseH1-Myc (RH1) or pLHCX-catalytically dead RNaseH1-Myc (RH1-CD), probed for Myc and Actin. Representative of n = 6 biological replicates. C Lagging (green) and leading (black) telomere fragility assessed by CO-FISH in PCSLT cells transduced with EV, RH1, or RH1-CD. A minimum of 11500 chromosome ends were analyzed, based on n metaphase spreads collected from three independent biological replicates. Boxplot: Min, 1st quartile, Median, 3rd quartile, and Max. Kruskal–Wallis tests. D Left: Representative images for Telo-FISH (red)/IF experiments targeting RAD51 (cyan) and RPA (green) in PCSLT cells transduced with EV, RH1, or RH1-CD. Right: Co-localization events between telomeres and RAD51 (cyan) or RPA (green). n nuclei were analyzed from three independent biological replicates. Median in red, quartiles in orange. Kruskal–Wallis tests. Source data are provided as a Source Data file.
These observations suggest that telomeric RNA:DNA hybrids are probably not the cause of PCSLT telomere fragility. Instead, they may arise as a consequence of telomeric replication stress and contribute to the cellular response to it.
Restored TRF1 levels at PCSLT telomeres mitigate telomere fragility
It has been proposed that TRF1 undersaturation at long telomeres could contribute to replication stress21,26. Consequently, we hypothesized that TRF1 levels may be insufficient at long telomeres of PCSLT cells. Supporting this hypothesis, chromatin immunoprecipitation experiments showed a decreased density of TRF1 at PCSLT telomeres compared to PCSST telomeres (Fig. 5A and Supplementary Fig. 5A).
A Above: Representative blot from ChIP analyses showing TRF1 binding at telomeres in PCSST and PCSLT cells and in PCSLT cells transduced with pLPC empty vector (EV) or pLPC-TRF1 (TRF1). Below: Quantifications. Signals were normalized first to telomeric input signals and then to either PCSLT (left part) or EV-transduced PCSLT cells (right part). n = 3 independent biological replicates for PCSLT and PCSST and n = 4 independent biological replicates for PCSLT cells transduced with either EV or TRF1. Mean ± SEM. Two-tailed ratio paired t-tests. B Western blot of cell extracts from PCSLT cells transduced with pLPC/pLHCX-empty vector (EV), pLPC-TRF1 (TRF1) or pLHCX-RNaseH1-Myc (RH1), probed for TRF1 and Myc. Representative of n = 3 biological replicates. C Above: Representative CO-FISH pictures on metaphase spreads from PCSLT cells transduced with either EV, TRF1, or both TRF1 and RH1 showing lagging (green) and leading (red) telomere fragility (arrowheads). Below: Lagging (green) and leading (black) telomere fragility. A minimum of 5200 chromosome ends were analyzed, based on n metaphase spreads collected from three independent biological replicates. Boxplot: Min, 1st quartile, Median, 3rd quartile, and Max. Kruskal–Wallis tests. Source data are provided as a Source Data file.
To investigate TRF1 undersaturation as a cause of PCSLT telomere fragility, we transduced PCSLT cells with a construct enabling TRF1 overexpression (Supplementary Fig. 5B, C). This overexpression effectively restored telomeric TRF1 levels in PCSLT cells to levels comparable to those observed at PCSST telomeres (Fig. 5A). Remarkably, achieving full TRF1 saturation at PCSLT telomeres reduced telomere fragility on both leading and lagging strands to levels comparable to those seen in PCSST cells. In contrast, TRF1 overexpression had no impact on telomere fragility in PCSST cells (Fig. 5B, C and Supplementary Fig. 5D).
In agreement with previous observations41, restoring TRF1 saturation at PCSLT telomeres tended to decrease telomeric RNA:DNA hybrids (Supplementary Fig. 5E). However, restoring TRF1 levels did not provide resistance to RH1 overexpression. Instead, we observed that RH1 overexpression negated the protective effect of TRF1 on telomere fragility on both strands (Fig. 5B, C and Supplementary Fig. 5D), suggesting that the TRF1-dependent repression of telomere fragility requires RNA:DNA hybrids.
Replication stress markers persist at PCSLT-TRF1 telomeres
While RNA:DNA hybrids are often considered harmful due to their interference with DNA replication promoting genome instability42, RNA:DNA hybrids and active transcription also play a positive role in HR-mediated repair of DSBs43,44, Ku-mediated fork protection45, replication (re)priming or restart46,47,48, and protection of reversed forks against excessive DNA resection49,50. At telomeres, telomeric RNA:DNA hybrids have been proposed to stimulate HR-dependent processes51. Since we found RNA:DNA hybrids to be essential for TRF1’s ability to reduce telomere fragility, we hypothesized that TRF1, instead of alleviating fork barriers, may slow down fork progression and facilitate the resumption of telomeric DNA replication at stalled forks in a process requiring RNA:DNA hybrids.
To test whether PCSLT-TRF1 telomeres still experience fork stalling, we first examined whether replication stress markers persisted in PCSLT cells overexpressing TRF1 (Supplementary Fig. 6A, B). Accordingly, we observed similar RPA and pRPA levels (Fig. 6A) and comparable recruitment of RAD51 and RPA at telomeres (Fig. 6B). Similarly, CCA signals (Fig. 6C) and APB-like structures (Fig. 6D) remained unchanged in PCSLT cells with restored telomeric TRF1 density. To further assess the impact of TRF1 on fork stalling, we examined the effect of TRF1 overexpression on the structure of replication forks traveling through the telomeric repeats of the SV40 mini-chromosome. As previously reported, the inclusion of telomeric DNA in an SV40 mini-chromosome amplifies the cone signal observed on two-dimensional gel electrophoresis, indicative of fork reversal9 (Fig. 6E and Supplementary Fig. 6C). Our results revealed that TRF1 overexpression slightly enhanced the cone signal in the SV40 mini-chromosome containing telomeric DNA, while no such increase was observed in the SV40 mini-chromosome lacking telomeric repeats (Fig. 6E). This suggests that TRF1 does not supress telomeric fork reversal under these experimental conditions.
A Above: Western blot of cell extracts from PCSLT cells transduced with pLPC empty vector (EV) or pLPC-TRF1 (TRF1), probed for pRPA32-S33, RPA32, and Actin. Below: pRPA32-S33 (purple) and RPA32 (green) levels, normalized to Actin and to PCSLT-EV. Representative of n = 6 independent biological replicates. Mean ± SEM. Two-tailed ratio paired t-tests. B Co-localization events between telomeres and RAD51 (cyan) or RPA32 (green) in PCSLT cells transduced with either EV or TRF1. n nuclei were analyzed from three independent biological replicates. Median in red, quartiles in orange. Two-tailed Mann–Whitney tests. C Representative blot and quantification for CCA in PCSLT cells transduced with either EV or TRF1. Signals were normalized to PCSLT-EV. n = 3 independent biological replicates. Mean ± SEM. Two-tailed ratio paired t-test. D Co-localization events between telomeres and PML in PCSLT cells transduced with either EV or TRF1. n nuclei were analyzed from three independent biological replicates. Median in red, quartiles in orange. Two-tailed Mann–Whitney test. E Above left: Schematic representation of the pTelN mini-chromosome containing 115 telomeric repeats (red arrow) and the SV40 replication origin (ORI). Above right: Schematic representation of the 2D-gel migration profile of the specified DNA replication intermediates. Below left: 2D-gel analysis of plasmid DNA using the 5.2 kb BamHI-SacI probe, in cells with and without ectopic overexpression of TRF1. Below right: Quantification of cone signal (reversed fork migration) as a ratio of cone signal to total replication intermediates (RI), normalized to EV. n = 3 independent biological replicates. Mean ± SEM. Two-tailed ratio paired t-test. F, G Lagging (green) and leading (black) telomere fragility assessed by CO-FISH in PCSLT-EV or PCSLT-TRF1 cells under the specified conditions. siSM1: siSMARCAL1. Cells were treated with either 15 µM BIBR (F) or 5 µM Olaparib (PARPi) (G), or DMSO as control, during 48 h before BrdU/BrdC incorporation. A minimum of 8600 (F) or 6200 (G) chromosome ends were analyzed, based on n metaphase spreads collected from three independent biological replicates. Boxplot: Min, 1st quartile, Median, 3rd quartile, and Max. Kruskal–Wallis tests. Source data are provided as a Source Data file.
TRF1 requires fork reversal to repress lagging telomere fragility
After demonstrating that TRF1 does not suppress fork reversal in the SV40 mini-chromosome assay, we next investigated whether fork reversal is essential for TRF1’s ability to safeguard telomeres from fragility. Although this may initially seem counterintuitive, it may be consistent with findings showing that fork slowing/reversal efficiently promotes replication fork stabilization and restart, contributing to the maintenance of genome stability52,53,54,55,.
SMARCAL1 is the primary DNA translocase recruited to stalled replication forks at telomeres to aid in their repair and resumption via fork reversal10. Consistent with our hypothesis, TRF1’s capacity to prevent fragility on the lagging strand telomere was completely eliminated when SMARCAL1 was depleted, whereas fragility at the leading strand telomeres remained unchanged (Fig. 6F and Supplementary Fig. 6D). Previous studies in yeast and mammals have reported that telomerase activity contributes to the management of reversed telomeric forks6,7,8,9. Further supporting the role of fork reversal in ensuring TRF1’s function in preventing fragility on the lagging strand telomere, we observed that treatment with the BIBR telomerase inhibitor produced effects similar to SMARCAL1 depletion in PCSLT-TRF1 cells (Fig. 6F). Importantly, we found that telomerase activity inhibition and repression of fork reversal act through the same pathway, as indicated by their epistatic relationship (Fig. 6F).
Finally, given the role of PARP1 in promoting fork reversal56, we tested whether inhibiting PARP1 with olaparib could mimic the effects of SMARCAL1 depletion or telomerase activity inhibition on PCSLT-TRF1 telomere fragility. In contrast to the results observed in SMARCAL1-depleted PCSLT-TRF1 cells, where increased fragility was specifically observed on the lagging strand and not the leading strand, olaparib treatment reversed the protective effect of TRF1 on telomere fragility, impacting both the lagging and leading strands (Fig. 6G). These results suggest that PARP1’s role extends beyond promoting fork reversal and align with the demonstration that PARP1 inhibitors accelerate fork elongation speed57. In this context, the effect of PARP1 mirrors the role we propose for TRF1 at telomeres, where it likely slows fork progression to help reduce genomic instability.
In summary, these findings highlight TRF1’s role in mitigating lagging telomere fragility through fork reversal.
PrimPol rescues leading telomere fragility in PCSLT-TRF1 cells lacking SMARCAL1
When fork reversal is impaired, repriming of DNA synthesis using the highly error-prone primase-polymerase PrimPol serves as an alternative pathway for restarting replication forks58. Although PrimPol has been reported to enable DNA replication repriming to bypass G4s55, its role in the replication of human telomeres has not yet been established.
To test whether PrimPol-dependent repriming was used by SMARCAL1-depleted PCSLT-TRF1 cells, we examined the impact of PrimPol depletion and observed increased leading telomere fragility (Fig. 7A and Supplementary Fig. 7A–C). This suggests that, in the absence of fork reversal, PrimPol takes over in dealing with replication stress at leading telomeres. Notably, PrimPol knockdown alone did not significantly affect telomere fragility in PCSLT-TRF1 cells (Fig. 7A), indicating that fork reversal, considered a higher fidelity DNA damage tolerance pathway59, remains the primary pathway for managing fork blockage. Interestingly, in PCSLT-TRF1 cells depleted of both SMARCAL1 and PrimPol, fragility assessed by CO-FISH was high on both strands (Fig. 7A and Supplementary Fig. 7A–C), suggesting that BIR was still not induced under these conditions.
A Lagging (green) and leading (black) telomere fragility assessed by CO-FISH in PCSLT-EV, PCSLT-TRF1 or PCSLT-TRF1-RH1 cells under the specified conditions. siSM1: siSMARCAL1. Cells were treated with 60 µM spironolactone (SP), or DMSO as control, during 25 h before BrdU/BrdC incorporation. A minimum of 5900 chromosome ends were analyzed, based on n metaphase spreads collected from three independent biological replicates. Boxplot: Min, 1st quartile, Median, 3rd quartile, and Max. Kruskal–Wallis tests. Source data are provided as a Source Data file. B Model. Stalled replication forks at PCSLT telomeres undergo SMARCAL1-dependent reversal, generating new 3′ telomeric ends. The reversed forks are bound by telomerase. Fork replication can resume through a homologous recombination-dependent restart process that requires RAD51 and might be facilitated by long 3′ overhangs generated by the telomerase and the formation of RNA:DNA hybrid formation, possibly involving TFIIH. If reversal is suppressed, PrimPol-dependent repriming can promote fork restart. This process may also be dependent on RNA:DNA hybrids. Optimal levels of TRF1 are required to suppress telomere fragility, by facilitating the reversal/restart processing of stalled forks. Protein illustrations were generated using BioRender (Vaurs, M. (2025) https://BioRender.com/ke04c93).
These observations provide evidence for PrimPol’s role in promoting an alternative damage tolerance pathway at SMARCAL1-depleted leading strand telomeres experiencing replication stress.
TFIIH is required for TRF1 to repress leading telomere fragility
Recently, TFIIH has been identified as a critical effector of TRF1, through non-canonical functions of the transcription and nucleotide excision repair factor21. Considering TFIIH’s role in facilitating the assembly of the RNA PolII preinitiation complex, we hypothesized that its transcription-related activity might contribute to its reported function in supporting TRF1’s role in telomere replication21 by promoting the formation of RNA:DNA hybrids.
To investigate whether inhibiting TFIIH mimics RNAseH1 overexpression, we treated PSCLT-TRF1 cells with spironolactone (SP), an inhibitor of TFIIH. However, while we observed a significant increase in fragility of the leading strand telomere in SP-treated PCSLT-TRF1 cells, lagging strand telomere was not affected (Fig. 7A). Building on our findings highlighting the role of fork reversal in mitigating fragility of the lagging strand in PCSLT-TRF1 cells, we propose that telomeres in SP-treated PCSLT-TRF1 cells still undergo fork reversal and that TFIIH may be crucial for homology-dependent restart to alleviate leading strand fragility (Fig. 7B).
Therefore, the RNA:DNA hybrids that are required to enable TRF1’s function in mitigating lagging strand telomere fragility do not depend on TFIIH and rely on still unknown mechanisms (Fig. 7B).
Discussion
The increased structural complexity of long telomeres and the higher incidence of replication barriers, such as G4s25, led to suggest that long telomeres may be more susceptible to replication stress. Previous studies have supported this hypothesis by using ectopic overexpression of telomerase22,23. Additionally, the detrimental effect of telomere length on stability is evidenced by the observation that long telomeres undergo trimming23. In this study, we found that PCSLT cells, which have long telomeres and spontaneous reactivation of telomerase, exhibit more replication stress markers at telomeres than isogenic PCSST cells with shorter telomeres, supporting the negative impact of telomere length on telomere replication. Replication stress markers also characterize the telomeres of ALT+ cells, which maintain telomeres independently of telomerase activity through a BIR-like mechanism13,14,15,27. Some markers, such as C-circles29, are used to indicate ALT activity. CC however, are also detected in PCSLT cells, consistent with findings on TEL+ cells with over-lengthened telomeres and high levels of CC22,24. PCSLT cells exhibit additional ALT+ markers, such as the co-localization of telomeres with PML bodies (APBs), also reported in some TEL+ cells with elongated telomeres23. However, BIR is not activated at PCSLT telomeres, and no T-SCEs were detected, indicating the absence of ALT activation. Thus, caution should be exercised when using ALT markers like CC or APBs to screen for ALT activity60, as they indicate telomeric replication stress rather than true ALT activity.
Our work indicated that telomere fragility in PCSLT cells was due to TRF1 undersaturation at telomeres. The mechanism by which TRF1 facilitates telomeric DNA replication is still unclear. Initially, it was suggested that TRF1 aids telomeric replication by resolving fork barriers through the recruitment of DNA helicases like BLM3,17,18,19. However, the role of TRF1 in telomere replication is certainly more complex, especially given that BLM deficiency in MEFs does not induce the leading strand telomere fragility observed in the absence of TRF13,17. TRF1 is now proposed to engage with additional partners, such as TFIIH, to mitigate replication challenges21.
We made the unexpected observation that TRF1 may play a role in fork restart/recovery through fork reversal, rather than removing upstream fork barriers. This is consistent with in vitro evidence showing that exogenous TRF1 and TRF2 hinder DNA replication progression through telomeric repeats by acting as fork barriers61,62. Our finding that overexpressing TRF1 slightly increased the cone signals detected in 2D gels, indicative of fork reversal at telomeric repeats on the SV40 mini-chromosome, further supports this notion. Along this line, replication fork slowing has recently been proposed as a mechanism to preserve genomic integrity at genomic loci prone to G4 formation55. This concept does not inherently contradict the idea that TRF1 may assist in fork barrier removal at some stage. It is plausible that fork reversal, by slowing down replication forks, provides an opportunity for TRF1 or other factors to remove subsequent barriers encountered during replication of the lagging strand telomere, such as G4s.
The TRF1-dependent rescue of lagging strand telomere fragility was completely lost upon SMARCAL1 depletion, while leading strand telomere fragility remained unaffected. This indicates a crucial role for fork reversal in repressing lagging strand telomere fragility. One of the main functions of fork reversal is to allow RAD51-dependent shielding of arrested forks against resection63,64,65,66. At telomeres, extensive resection of the lagging strand leads to the formation of ss G-rich DNA which, through the formation of G4 structures, can increase telomere fragility2,17,18. Stretches of ssDNA might also lead to aberrant chromatinization/condensation resulting in a fragile telomere phenotype3. We therefore propose that reversed forks help mitigate lagging strand telomere fragility by shielding arrested forks from nuclease activity5. RAD51 plays a key role in fork protection in multiple ways: it not only catalyzes fork reversal but also prevents the degradation of the regressed nascent DNA arm by MRE11 and EXO1/DNA263,64,65,66. Additionally, RAD51 may promote strand invasion reaction for HR-dependent replication resumption67,68. Crucially, RAD51 ensures that the replicative helicase remains properly poised to the stalled fork, promoting DNA replication restart69. We hypothesize that telomerase activity aids in the formation of the necessary 3′ overhang at reversed forks (Fig. 7B). Although telomerase inhibition with BIBR similarly increased the fragility of mouse RTEL1−/− telomeres undergoing fork reversal, telomerase depletion had the opposite effect6. This raises the possibility that BIBR traps TERT at reversed forks, interfering with processes required for efficient fork restart. Thus, telomerase-independent mechanisms might be more efficient at generating the 3′ overhang at reversed telomeric forks but must compete with telomerase for binding.
Our data indicated that fork reversal is not an absolute requirement for TRF1’s ability to repress leading strand telomere fragility, as PrimPol appears to assume responsibility for leading strand telomere repriming in SMARCAL1-depleted cells. PrimPol depletion alone did not increase leading strand telomere fragility, indicating that fork reversal is the primary pathway used by PCSLT-TRF1 cells to recover from fork stalling. This provides evidence for a role of PrimPol in telomeric leading strand synthesis and supports the notion that fork reversal and fork repriming operate independently58. Given that restoration of TRF1 telomeric levels reduces fragility on both strands, and that both fork reversal and fork repriming are used to reduce telomere fragility, TRF1 might play a role in both pathways (Fig. 7B).
Our surprising finding that overexpression of RNaseH1 in PCSLT-TRF1 cells completely abolishes TRF1’s ability to reduce both leading and lagging strand fragility—mimicking the effect of combined SMARCAL1 and PrimPol depletion—suggests a role for RNA:DNA hybrids in both fork reversal and PrimPol-dependent repriming. We propose that telomeric RNA:DNA hybrids may both help protect reversed telomeric forks and support HR-dependent replication restart by facilitating DNA invasion. These findings are consistent with previous studies showing that RNA:DNA hybrids promote HR via an R-to-D-loop transition70 and help protect stalled replication forks49,50. They also align with recent evidence that telomeric RNA:DNA hybrids are induced in response to oxidative stress at telomeres—a condition that also leads to the formation of i-loops and single-strand breaks—potentially to enhance DNA damage signaling and repair71. Their findings further suggest that telomeric RNA:DNA hybrids may contribute to the dissociation of TRF1 from telomeres71. Thus, it is possible that TRF1 slows replication fork progression, triggering fork reversal, and that the resulting telomeric RNA:DNA hybrids facilitate TRF1 removal, enabling replication to continue. It remains to be determined whether the telomeric RNA species correspond to canonical TERRA molecules transcribed from subtelomeric promoters or are generated locally, potentially via the transcription-promoting activity of TFIIH, and whether they form in cis or in trans (Fig. 7B). Alternatively, the ATPase/translocase activity of TFIIH XPB subunit might contribute to fork restart by removing nucleosomes72, or unwinding DNA to facilitate fork recovery.
In summary, our study reveals an unexpected function for TRF1 in reducing telomere fragility via fork reversal and highlights the roles of RNA:DNA hybrids in this process. Future research is needed to clarify the roles of TRF1 and RNA:DNA hybrids in stalled fork processing, and to understand how TFIIH facilitates leading strand telomere restart.
Methods
Post-crisis survivors
Forty million puromycin-resistant IMR90E6/E7 cells, which had grown to crisis at PD105, were seeded across twenty 15 cm dishes (2 million cells per dish). After eight weeks, cloning cylinders were used to isolate the eight individual clones, named PCS-1 through PCS-8.
Cell culture procedures
U2OS, HeLa, HeLa 1.3 (kindly provided by T. de Lange, Rockefeller University, NY, USA) and HEK293T were grown in DMEM (Gibco) supplemented with 10% Fetal Bovine Serum (FBS) (Cytiva) and 1% penicillin/streptomycin (P/S) (Capricorn). HT1080 and HT1080-ST (kindly provided by J. Lingner, EPFL, Lausanne, Switzerland), IMRB (Coriell Institute for Medical Research, New Jersey, USA) and SW39 (kindly provided by W. Wright, UT Southwestern, Dallas, USA) SV40T-immortalized human fetal lung fibroblasts were grown in EMEM (Gibco) supplemented with 10% FBS, 1% non-essential amino acids (Gibco) and 1% P/S. The post-crisis survivor (PCS) and the parental IMR90E6/E7 cell lines were grown in DMEM supplemented with 15% FBS and 1% P/S. Cells were cultured at 37 °C under a humidified atmosphere of 5% CO2. Mycoplasma contaminations were tested by qPCR in culture medium using mycoA/B-F/R primers listed in Supplementary Data 1.
Transfections with siRNAs
siRNAs (Supplementary Data 2) were transfected at a final concentration of 125 nM, using Lipofectamine 2000 (ThermoFisher Scientific) as per manufacturer’s instructions. Briefly, 170000 to 200000 cells (for 6-wells plates), or 42500 to 50000 cells (for poly-lysine-coated coverslips), resuspended in Opti-MEM (Gibco) medium, were used for transfections. Six hours after transfection, medium was changed. Unless otherwise stated, cells were collected three days post-transfection.
Retrovirus generation and transduction
One day after plating into T25 flasks, HEK293T cells were transfected with 1 µg of pVPack-VSV-G vector, 2 µg of pVPack-GP, and 6 µg of the plasmid of interest using Polyethylenimine (Polysciences Inc.). Six hours after transfection, medium was changed. Two days after transfection, the retroviral particle-containing medium was filtered through 0.45 µm filters and incubated with 8 µg/ml Polybrene (Sigma) for 20 min. Cells seeded in T25 flasks the day before transduction were then exposed to the retrovirus-containing medium for 24 h before three washes with 1× PBS. Unless otherwise stated, cells were collected and plated for experiments three days post-transduction. All plasmids used in this study are listed in Supplementary Data 3.
ddTRAP
Briefly, 500000 cells were lysed on ice in NP-40 buffer (10 mM Tris-HCl pH 8.0, 0.1 mM MgCl2, 1 mM EDTA, 10% (v/v) glycerol, 1% (v/v) NP-40, 150 mM NaCl, 0.25 mM Sodium Deoxycholate, 5 mM β-mercaptoethanol, 0.1 mM AEBSF (ThermoFischer Scientific)) for 30 min. 1 µl of the lysate at 2 µg/µl was added to 50 μl of 1× TRAP buffer (200 mM Tris-HCl, pH 8.3, 15 mM MgCl2, 2.5 mM each dNTPs, 200 nM TS primer (Supplementary Data 1), and 0.4 mg/ml BSA), and extension reaction was performed by incubating the mix at 25 °C for 40 min, followed by inactivation at 95 °C for 5 min. For the droplet digital PCR reaction, 3 µl of the reaction mix were added to 1× EvaGreen Supermix, 50 nM TS primer, 50 nM ACX primer (Supplementary Data 1) in a 20 μl reaction. PCR program was as follows: initial denaturation for 5 min at 95 °C followed by 40 cycles of 95 °C/30 s, 54 °C/30 s, and 72 °C/30 s.
Genomic DNA extraction
Cells were lysed in TNES buffer (10 mM Tris-HCl pH 8.0, 1 mM EDTA, 0.5% SDS, 10 mM NaCl), incubated for at least 1 h with 50 µg/ml of RNase A (Thermo Scientific) at 37 °C before overnight incubation in the presence of 100 μg/ml proteinase K at 37 °C (Merck Life Science BV). Genomic DNA was extracted with phenol-chloroform-isoamyl alcohol 25:24:1, precipitated with isopropanol and resuspended in TE (10 mM Tris-HCl pH 7.5, 1 mM EDTA).
Telomere restriction fragment (TRF) analysis
10 μg of RNA-free genomic DNA were digested overnight with 20 U HinfI and RsaI and directly loaded on a 0.6% agarose gel. After 6 h of migration at 75 V, the gel was first subjected to a depurination step in 0.25 M HCl for 10 min and then incubated two times for 15 min in denaturation buffer (0.5 M NaOH, 1.5 M NaCl). Next, neutralization was achieved by incubating the gel twice during 15 min in 0.5 M Tris-HCl, 3 M NaCl, 1 mM EDTA, pH 7.5. Finally, the gel was incubated in 20× SSC solution (3 M NaCl, 0.3 M sodium citrate, pH 7.0) and left overnight to transfer by capillarity on a Hybond N+ nylon membrane (Cytiva). After transfer, the membrane was pre-hybridized for 1 h at 42 °C in ULTRA hyb-Oligo hybridization buffer (Ambion) prior overnight incubation with the radioactive telomeric probe in the same buffer at 42 °C. The C-rich telomeric probe was prepared as follows: 10 μM (CCCTAA)4 telomeric sequence was denatured for 5 min at 68 °C and 1 μl was used for radioactive labeling with 6 μl of [γ-32P] ATP (10 mCi/ml) (PerkinElmer) in the presence of 10 U of T4 polynucleotide kinase (Roche) in a 20 μl-reaction mix containing 1× PNK buffer; this mix was incubated for 20 min at 37 °C before inactivation of the enzyme with 2 μl of 0.5 M EDTA, pH 8.0. Forty microliters of 1× PNK buffer were then added to the radioactive probe before purification on a G-25 column (Cytiva). After hybridization, the membrane was washed twice with Stringent wash buffer I (2× SSC, 0.1% SDS) for 5 min followed by two washes of 5 min with Stringent wash buffer II (0.2× SSC, 0.1% SDS). Signal was analyzed using Phosphorimager.
C-circle assay (CCA)
2–5 μg of RNA-free genomic DNA were digested with 20 U of HinfI and RsaI before purification using phenol-chloroform extraction. 30 ng of digested genomic DNA were resuspended in 10 μl of 10 mM Tris pH 8.0 and added to 10 μl of reaction mix (4 μg/ml BSA, 4 mM DTT, 0.1% Tween-20, 1 mM each dATP, dCTP, dGTP and dTTP, 1× Phi29 Buffer) with or without 7.5 U of Phi29 DNA polymerase (NEB). Samples were incubated at 30 °C for 8 h and then at 65 °C for 20 min. Amplification products were slot-blotted on a Hybond N+ nylon membrane (Cytiva). The membrane was probed as described for TRF analysis and washed once for 10 min at room temperature (RT) with Stringent wash buffer I, once for 3 min at 42 °C with Stringent wash buffer II and twice for 5 min at RT with Stringent wash buffer I. Signal was analyzed as described above for TRF.
RNA extraction and qRT-PCR
RNA was extracted using TriPure Isolation Reagent (Sigma-Aldrich), followed by chloroform extraction and precipitation with isopropanol. RNA was treated with 2 U of DNase I (ThermoFisher Scientific) for 30 min at 37 °C followed by a second phenol-chloroform extraction and isopropanol precipitation. One μg of the purified RNA was reverse transcribed using MMLV-RT (ThermoFisher Scientific) and random hexamers (ThermoFisher Scientific). RNA was incubated at 42 °C for 1 h followed by 10 min at 95 °C. cDNA was then analyzed by qPCR with the primers listed in Supplementary Data 1 using KAPA SYBR FAST (Sigma-Aldrich) with the following program: initial denaturation for 10 min at 95 °C followed by 40 cycles of 95 °C/15 s and 60 °C/30 s, and a last step at 95 °C for 15 s. The results were calculated according to the 2−ΔΔCt methodology and are shown as relative expression to the corresponding reference sample.
Chromatin Immunoprecipitation (ChIP)
Chromatin immunoprecipitation of H3Ac and H3 was carried out using OneDay ChIP kit (Diagenode) as per manufacturer’s instruction using rabbit polyclonal anti-H3 (Abcam, 1:100) and rabbit polyclonal anti-H3Ac (Millipore, 1:100). Immunoprecipitated DNA was analyzed by qPCR using primers listed in Supplementary Data 1.
For chromatin immunoprecipitation of TRF1, 10 million cells were harvested by trypsinization, centrifuged at 805 × g (2000 rpm) at 4 °C for 3 min, and resuspended in 1% formaldehyde for 30 min at room temperature, followed by quenching with 125 mM glycine for 5 min. Cross-linked cells were centrifuged as above and pellets were resuspended in 500 µl of ChIP lysis buffer (1% SDS, 10 mM EDTA pH 8.0, 50 mM Tris-HCl pH 8.0), sonicated using a Bioruptor apparatus (Diagenode) and centrifuged at maximum speed for 10 min at 4 °C. 1 mg of lysate was diluted in ChIP dilution buffer (150 mM NaCl, 20 mM Tris-HCl pH 8.0, 1% Triton X-100, 2 mM EDTA pH 8.0) to reach a total of 1 ml. One hundred microliters of lysate were kept as input and 900 µl were incubated with 1 μg of anti-TRF1 antibody (St John’s laboratory) for 4 h at 4 °C. Immunoprecipitated complexes were isolated by incubation with 25 µl of Protein A/G magnetic beads (Thermo Scientific) at 4 °C overnight on a rotating wheel. Beads were washed four times with ChIP wash buffer (150 mM NaCl, 20 mM Tris-HCl pH 8.0, 1% Triton X-100, 0.1% SDS, 2 mM EDTA pH 8.0) and once with ChIP final wash buffer (500 mM NaCl, 20 mM Tris-HCl pH 8.0, 1% Triton X-100, 0.1% SDS, 2 mM EDTA pH 8.0). Beads were then incubated in 100 µl of ChIP elution buffer (1% SDS, 100 mM NaHCO3) containing 40 μg/ml RNase A (Thermo Scientific) for 15 min at RT followed by 1 h at 37 °C before overnight incubation at 65 °C for reverse cross-link. Input was treated as beads. DNA was purified using PCR purification kit (Qiagen), denatured with 0.2 M NaOH at 100 °C for 10 min, cooled down directly on ice, and neutralized (6× SSC, 0.2 M Tris pH 7.4) prior slot-blotting onto a Hybond N+ nylon membrane (Cytiva) that was processed as described above for TRF.
CO-FISH
Growing cells were incubated with 10 μM BrdU:BrdC (3:1) (MP Biomedicals, Sigma-Aldrich) for 16 to 20 h before metaphase arrest with 0.2 μg/ml colcemid (Roche) for 3 to 4 h. For experiments on transduced cells, cells were plated three days after transduction. For experiments on transfected cells, BrdU/BrdC was added 52 h after transfection. For experiments with BIBR, PARP inhibitor or spironolactone (SP), 15 µM of BIBR (Sigma), 5 µM of olaparib (Axon Medchem) or 60 µM of SP (Sigma) were added, respectively, 48 h, 48 h or 25 h, prior to BrdU/BrdC addition. Cells were then subjected to hypotonic shock with 75 mM KCl at 37 °C for 30 min before fixation with methanol:acetic acid solution (3:1, v/v). Metaphase spreads were prepared by dropping the cell suspensions onto Superfrost slides (VWR) before overnight drying in the dark. The slides were then rehydrated in 1× PBS and treated with 100 µg/ml RNase A (Merck Life Science BV) in 1× PBS for 10 min at 37 °C before incubation with 0.05 μg/ml Hoechst 33258 (Sigma) for 15 min at RT and exposition to 365-nm UV light (Stratalinker 1800UV irradiator; crosslink at energy = 5.4 × 103 J/m2). The BrdU/BrdC-substituted DNA strand was digested with 150 U of Exonuclease III (NEB) for 5 min at RT, followed by 20 min at 37 °C and 5 min at RT. After dehydration in ethanol series (70, 90, and 100%), the slides were first hybridized with a TeloC PNA probe in Hybridization solution I (0.5 µg/ml TelC-FITC probe (Panagene- FITC-OO-CCCTAACCCTAACCCTAA), 70% deionized formamide (Bio Science), 20 mM Tris pH 7.4, 1× Blocking agent (Sigma-Aldrich)) for 3 min at 83 °C, followed by 1.5 h at RT. After a first wash for 5 min in PNA-Wash buffer (70% formamide (Bio Science), 20 mM Tris-HCl pH 7.4), and a second one, for 5 min, in FISH-Wash II buffer (150 mM NaCl, 0.05% Tween-20, 50 mM Tris-HCl pH 7.4), the slides were dehydrated again using the ethanol series (70, 90, and 100%). Subsequently, the slides were incubated at RT for 1.5 h with the GGGTtAGGGttAGgGTTAGGGttAGGGttAGGGtTA (TAMRA) TeloG LNATM probe (with small letters indicating LNATM modified bases) in Hybridization solution II (50 nM TelG probe (Qiagen), 50% deionized formamide (Bio Science), 2× SSC, 1× Blocking agent (Sigma-Aldrich)). The unbound probe was washed off successively with two washes for 15 min each in LNA-Wash buffer (50% formamide (Bio Science), 2× SSC, 20 mM Tris-HCl pH 7.4) and three times, for 5 min each, in FISH-Wash II buffer. Finally, slides were serially dehydrated again, air dried and mounted with mounting medium (DAKO) containing 0.5 μg/ml DAPI (Sigma). Images were acquired with the Cell Observer Spinning Disk confocal microscope (Zeiss) with 100× objective. Pictures were analyzed using ImageJ.
FISH on metaphase spreads
After colcemid (Roche) treatment as described above, metaphase spreads on slides were rehydrated in 1× PBS before incubation for 10 min in the fixation buffer (3.7% formaldehyde, 1× PBS). Then, samples were incubated in 100 µg/ml RNase A (Merck Life Science BV) in 1× PBS for 20 min at 37 °C, before fixation again for 10 min. The slides were subsequently dehydrated using the ethanol series (70, 90, and 100%), hybridized at 83 °C for 3 min with 50 nM TeloG LNATM probe in Hybridization solution II and then incubated at RT for 1.5 h. The unbound probe was washed away and slides were mounted as described above for the CO-FISH experiments. Images were acquired and analyzed as described for CO-FISH experiments.
Immunofluorescence (IF), native or denaturing telomeric FISH and ATSA
For staining experiments on cultured cells, 15,000 to 20,000 cells were seeded the day before onto 4-well slides (Thermo Scientific). For staining experiments on transduced cells, cells were detached and seeded onto 4-well slides three days after transduction and staining was thus performed four days after transduction. Cells were treated with permeabilization buffer (20 mM Tris-HCl pH 8.0, 50 mM NaCl, 3 mM MgCl2, 300 mM Sucrose, 0.5% Triton X-100) for 5 min at RT, before fixation (3% formaldehyde, 2% sucrose, 1× PBS) for 15 min at RT and a second permeabilization step for 10 min at RT. For staining experiments on cells transfected with siRNAs, 42500 to 50000 cells were transfected on coverslips (Knittel Glass) as described above. Three days later, cells on coverslips were treated as described above for slides. Cells on slides/coverslips were incubated for 45 min at 37 °C in Blocking solution (1% BSA, 10% normal goat serum (Cell Signaling Technology), 0.1% Triton X-100, 1× PBS) before overnight incubation at 4 °C with primary antibody diluted in the Blocking solution. The next day, slides/coverslips were incubated for 40 min at 37 °C with secondary antibodies diluted in Blocking solution before fixation (3.7% formaldehyde, 1× PBS) for 2 min and treatment with 100 μg/ml RNase A (Merck Life Science BV) for 1 h at 37 °C. Cells on slides/coverslips were permeabilized and fixed again for, respectively, 10 min and 2 min, serially dehydrated with the ethanol series (70%, 90%, and 100%) and then air dried. Next, slides were hybridized at 83 °C for 3 min with 160 nM TeloG LNATM probe in Hybridization solution II and then incubated at RT for 2 h. For native FISH, slides were not heat-denatured. Unbound probe was washed off successively and slides were mounted as described for the CO-FISH experiments. For native FISH, an additional wash of 4 min with LNA-Wash buffer was performed.
For ATSA experiments, cells were first synchronized with 2 mM thymidine for 21 h followed by release for 4 h in fresh medium, incubation with 15 µM CDK1 inhibitor RO-3306 (Selleckchem) for 12 h and, finally, released for 1 h in fresh medium containing 10 µM EdU (ThermoFischer Scientific). Cells were then fixed and permeabilized as described above and EdU staining was performed using the Click-IT EdU Alexa Fluor 488 Imaging kit following manufacturer’s instructions (ThermoFischer Scientific). PML IF-FISH was subsequently performed as described above.
Images were acquired and analyzed as described for CO-FISH experiments.
All antibodies used in this study are listed in Supplementary Data 4.
Two-dimensional agarose gel electrophoresis
For Fig. 3G, 10 μg of genomic DNA, were digested overnight with HinfI and RsaI, precipitated with isopropanol and resuspended in 10 mM Tris-HCl pH 8.0. The first dimension was run on 0.4% agarose (US Biological) in 0.5× TBE without ethidium bromide (EtBr) for 16 h at 0.15 V/cm. The gel was then stained in 0.5× TBE with 0.3 µg/ml EtBr for 45 min. A 5.5 cm slice (from the well down) was cut for each lane. The second dimension was run on 0.8% agarose gel with 0.3 μg/ml EtBr for 3 h at 5 V/cm at 4 °C. After imaging, the DNA was blotted onto a membrane. For Fig. 6E, plasmid DNA was extracted using a modified Hirt protocol. Briefly, cells were washed in 1× PBS and resuspended in 1× TE. Lysis was performed by adding SDS to a final concentration of 0.6% and inverting the tubes gently. Chromosomal DNA was precipitated by adding NaCl to a final concentration of 1 M, followed by overnight incubation at 4 °C. Samples were centrifuged in a SW41-Ti rotor (Beckman) at 31611 × g (16000 rpm) for 3 h at 4 °C. The supernatant was collected and supplemented with 100 μg/ml proteinase K (Roche) before incubation at 37 °C for 1 h. The DNA, was precipitated in 100% ethanol, resuspended in Tris-HCl pH 8.0, treated with 100 μg/ml RNase A at 37 °C for 1 h and then extracted using phenol/chloroform/isoamyl alcohol, followed by chloroform extraction and precipitation with isopropanol. Plasmid DNA (3–4 μg) was then digested overnight with 90 U of SacI and BamHI. Subsequently, 40 U of DpnI was added and incubated at 37 °C for 1 h. The digestion product was precipitated with isopropanol and resuspended in 10 mM Tris-HCl (pH 8.0). The first dimension was run on 0.4% agarose in 0.5× TBE without EtBr for 42 h at 0.65 V/cm, then stained in 0.5× TBE with EtBr for 45 min. A 6.5 cm slice from the 5-kb size band and up was cleaved for each lane. The second dimension was run on 0.7% agarose gel with EtBr for 23 h at 3 V/cm at 4 °C. Prior to blotting, psoralen cross-linking was reversed by UV irradiation (254 nm) for 10 min. After migration, gels were processed by incubating for 30 min in depurination solution (0.25 M HCl), followed by 2 × 40 min in denaturing solution (1.5 M NaCl, 0.5 M NaOH), and 2 × 40 min in neutralizing solution (1.5 M NaCl, 0.5 M Tris-HCl pH 7.0). DNA was transferred by capillarity in 10× SSC onto an Amersham Hybond-X membrane (Cytiva). For Fig. 3G, probe was prepared as follows. After digestion of the pSty11 plasmid (Supplementary Data 3) with EcoRI, the 800-bp telomeric insert was purified, diluted to 20 ng/μl and 10 μl were mixed with 5 ng of (CCCTAA)4 telomeric primer and water before heat-denaturation and labeling using the Prime-a-Gene Labeling system (Promega) with [α-32P]dCTP (3000 Ci/mmol; PerkinElmer). For Fig. 6E, the 5.2-kb fragment obtained from the digestion of the pTelN plasmid (Supplementary Data 3) with BamHI and SacI was used for the probe. 30 ng of the purified fragment were heat-denatured and labeled as described for pSty11-derived fragment but without adding the telomeric primer. The labeled probes were purified, denatured, and added to the hybridization mix. Hybridization was performed at 65 °C for at least 4 h. Membranes were washed three times for 15 min with 1× Church Wash (25 mM NaPi pH 7.2, 0.1 M EDTA pH 8.0, 1% SDS) at 65 °C and radioactive signal was captured on phosphor-screens (FUJIFILM Storage Phosphor screen) before reading on Typhoon Trio Imager (GE).
Alkaline gel analysis
Genomic DNA (10 μg) was digested overnight with 50 U ClaI and loaded onto an agarose gel prepared with 1× alkaline buffer (0.5 M NaOH, 0.1 M EDTA). Electrophoresis was performed at 2 V/cm for 10 min, then continued for 16 h and 20 min with buffer circulation at 12 ml/min. The gel was neutralized, stained with 0.3 µM EtBr, and visualized. Southern blotting was then performed as described in the above section using strand-specific single-stranded probes prepared as follows. (CCCTAA)4 or (TTAGGG)5 primers were end-labeled with T4-PNK (NEB) and [γ-32P] ATP (6000 Ci/mmol; PerkinElmer). Hybridization was carried out overnight at 50 °C. The membranes were washed three times with 4× SSC for 30 min each and with 4× SSC, 0.1% SDS for another 30 min at 50 °C. Radioactive signal was captured on phosphor screens (FUJIFILM Storage Phosphor screen), read on Typhoon Trio (GE), and analyzed on ImageJ.
Protein extraction and Western Blot
Proteins were extracted using RIPA buffer (150 mM NaCl, 50 mM Tris-HCl pH 7.4, 0, 1% NP-40, 0.25% sodium deoxycholate) supplemented with proteases inhibitors (Roche) and phosphatases inhibitors (Roche) for pRPA analyses. Western blotting was performed according to standard procedures. Blots were revealed with SuperSignal West Pico Chemiluminescent Substrate (Thermo Scientific) using Amersham Image Quant 800 and signal quantification was done using ImageJ.
All antibodies used in this study are listed in Supplementary Data 4.
DNA:RNA Immunoprecipitation (DRIP)
Cells grown in 15 cm dishes until 70% confluent were scraped, centrifuged at 500 × g at 4 °C for 5 min, and resuspended by vortexing in 1 ml of RA1 lysis buffer (Macherey-Nagel) containing 10 µl of β-Mercaptoethanol and 100 mM NaCl. Nucleic acids were purified with Phenol:Chloroform:Isoamyl Alcohol (25:24:1) followed by isopropanol precipitation and resuspended in TE buffer contaning 300 mM NaCl. Nucleic acids were sonicated twice using a Bioruptor apparatus (Diagenode) at 4 °C (30 s “ON”/30 s “OFF”; power: “High”; time: 7.5 min). 8 μg of nucleic acids were brought to a final volume of 500 μl with IP buffer (0.1% SDS, 1% Triton X-100, 10 mM HEPES pH 7.2, 0.1% sodium deoxycholate, 275 mM NaCl) and incubated with 4 μg of mouse monoclonal S9.6 antibody (Kerafast) at RT for 1h30min on a rotating wheel. Immunocomplexes were isolated by incubation with 25 μl of Dynabeads™ Protein G (Invitrogen) pre-blocked with 5 μg of sheared Escherichia coli DNA and 30 μg of BSA for 1 h at RT. Beads were washed four times in IP buffer at RT and incubated in elution buffer (50 mM Tris-Cl pH 8, 10 mM EDTA, 0.5% SDS) containing 10 μg/ml proteinase K (Sigma-Aldrich) and 40 μg/ml RNase A (NZYtech) at 50 °C for 30 min. Supernatants were recovered and purified with the Wizard® SV Gel and PCR Clean-Up System (Promega). Quantitative PCRs were performed using the iTaq Universal SYBR Green Supermix (Bio-Rad) on an Applied Biosystems 7500 Fast instrument with a 2-step program (45 cycles of denaturation at 95 °C for 15 s, annealing and extension at 60 °C for 30 s) with the oligonucleotides listed in Supplementary Data 1.
Reagent catalog numbers are listed in Supplementary Data 4.
Quantification, statistical analyses, and manuscript preparation
ImageJ (Version 1.54p, Java1.8.0_322 (64-bit)) was used to analyze confocal microscopy pictures and quantify C-circles, slot blots, and Western blots. GraphPad Prism (Version 8.02) was used to generate graphs and for statistical analyses. Statistical tests used in this study were indicated in Figure legends. Normality was assessed using the Shapiro–Wilk test. If the data did not follow a normal distribution, a two-tailed Mann–Whitney test was performed. If normality was confirmed, a two-tailed unpaired t-test was used, with Welch’s correction applied in cases of significantly different variances. Comparisons involving a normalized dataset were analyzed using a two-tailed ratio paired t-test. For multiple comparisons, Kruskal–Wallis test was performed. For CO-FISH analyses, statistic analysis on leading and lagging strand fragilities were performed separately. Violin plots were used to represent immunofluorescence combined with FISH and T-SCE quantifications, while box plots illustrated telomere fragility detected by FISH or CO-FISH, showing the median, as well as the first and third quartiles. Protein illustrations in the model of Fig. 7B were generated using BioRender (Vaurs, M. (2025) https://BioRender.com/ke04c93). All remaining illustrations were created by the authors using Inkscape (Version 1.3.2).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data supporting the findings of this study are available within the paper and its Supplementary Information. Source data are provided with this paper.
References
de Lange, T. Shelterin-mediated telomere protection. Annu. Rev. Genet. 52, 223–247 (2018).
Glousker, G. & Lingner, J. Challenging endings: how telomeres prevent fragility. BioEssays 43, e2100157 (2021).
Sfeir, A. et al. Mammalian telomeres resemble fragile sites and require TRF1 for efficient replication. Cell 138, 90–103 (2009).
Drosopoulos, W. C., Kosiyatrakul, S. T., Yan, Z., Calderano, S. G. & Schildkraut, C. L. Human telomeres replicate using chromosome-specific, rather than universal, replication programs. J. Cell Biol. 197, 253–266 (2012).
Adolph, M. B. & Cortez, D. Mechanisms and regulation of replication fork reversal. DNA Repair (Amst.) 141, 103731 (2024).
Margalef, P. et al. Stabilization of reversed replication forks by telomerase drives telomere catastrophe. Cell 172, 439–453.e14 (2018).
Matmati, S., Lambert, S., Géli, V. & Coulon, S. Telomerase repairs collapsed replication forks at telomeres. Cell Rep. 30, 3312–3322.e3 (2020).
Dehé, P. M., Rog, O., Ferreira, M. G., Greenwood, J. & Cooper, J. P. Taz1 enforces cell-cycle regulation of telomere synthesis. Mol. Cell 46, 797–808 (2012).
Huda, A. et al. The telomerase reverse transcriptase elongates reversed replication forks at telomeric repeats. Sci. Adv. 9, eadf2011 (2023).
Poole, L. A. et al. SMARCAL1 maintains telomere integrity during DNA replication. Proc. Natl. Acad. Sci. USA 112, 14864–14869 (2015).
Quinet, A., Tirman, S., Cybulla, E., Meroni, A. & Vindigni, A. To skip or not to skip: choosing repriming to tolerate DNA damage. Mol. Cell 81, 649–658 (2021).
Costantino, L. et al. Break-induced replication repair of damaged forks induces genomic duplications in human cells. Science 343, 88–91 (2014).
Roumelioti, F. et al. Alternative lengthening of human telomeres is a conservative DNA replication process with features of break-induced replication. EMBO Rep. 17, 1731–1737 (2016).
Dilley, R. L. et al. Break-induced telomere synthesis underlies alternative telomere maintenance. Nature 539, 54–58 (2016).
O’Sullivan, R. J. & Greenberg, R. A. Mechanisms of alternative lengthening of telomeres. Cold Spring Harb. Perspect. Biol. 17, a041690 (2025).
Garcia-Exposito, L. et al. Proteomic profiling reveals a specific role for translesion DNA polymerase η in the alternative lengthening of telomeres. Cell Rep. 17, 1858–1871 (2016).
Zimmermann, M., Kibe, T., Kabir, S. & de Lange, T. TRF1 negotiates TTAGGG repeat-associated replication problems by recruiting the BLM helicase and the TPP1/POT1 repressor of ATR signaling. Genes Dev. 28, 2477–2491 (2014).
Vannier, J. B., Pavicic-Kaltenbrunner, V., Petalcorin, M. I. R., Ding, H. & Boulton, S. J. RTEL1 dismantles T loops and counteracts telomeric G4-DNA to maintain telomere integrity. Cell 149, 795–806 (2012).
Maresca, C. et al. PARP1 allows proper telomere replication through TRF1 poly (ADP-ribosyl)ation and helicase recruitment. Commun. Biol. 6, 234 (2023).
Sarek, G., Vannier, J. B., Panier, S., Petrini, H. J. & Boulton, S. J. TRF2 recruits RTEL1 to telomeres in S phase to promote t-loop unwinding. Mol. Cell 57, 622–635 (2015).
Yang, Z., Sharma, K. & de Lange, T. TRF1 uses a noncanonical function of TFIIH to promote telomere replication. Genes Dev. 36, 956–969 (2022).
Rivera, T., Haggblom, C., Cosconati, S. & Karlseder, J. A balance between elongation and trimming regulates telomere stability in stem cells. Nat. Struct. Mol. Biol. 24, 30–39 (2017).
Pickett, H. A., Cesare, A. J., Johnston, R. L., Neumann, A. A. & Reddel, R. R. Control of telomere length by a trimming mechanism that involves generation of t-circles. EMBO J. 28, 799–809 (2009).
Jones, C. Y. et al. Hyperextended telomeres promote formation of C-circle DNA in telomerase positive human cells. J. Biol. Chem. 299, 104665 (2023).
Yang, S. Y. et al. G-quadruplexes mark alternative lengthening of telomeres. NAR Cancer 3, zcab031 (2021).
Miller, K. M., Rog, O. & Cooper, J. P. Semi-conservative DNA replication through telomeres requires Taz1. Nature 440, 824–828 (2006).
Lu, R. & Pickett, H. A. Telomeric replication stress: the beginning and the end for alternative lengthening of telomeres cancers. Open Biol. 12, 220011 (2022).
Yang, Z., Takai, K. K., Lovejoy, C. A. & de Lange, T. Break-induced replication promotes fragile telomere formation. Genes Dev. 34, 1392–1405 (2020).
Henson, J. D. et al. DNA C-circles are specific and quantifiable markers of alternative- lengthening-of-telomeres activity. Nat. Biotechnol. 27, 1181–1185 (2009).
Zhang, J. M., Yadav, T., Ouyang, J., Lan, L. & Zou, L. Alternative lengthening of telomeres through two distinct break-induced replication pathways. Cell Rep. 26, 955–968.e3 (2019).
Loe, T. K. et al. Telomere length heterogeneity in ALT cells is maintained by PML-dependent localization of the BTR complex to telomeres. Genes Dev. 34, 650–662 (2020).
Sobinoff, A. P. et al. BLM and SLX4 play opposing roles in recombination-dependent replication at human telomeres. EMBO J. 36, 2907–2919 (2017).
Claude, E. et al. Detection of alternative lengthening of telomeres mechanism on tumor sections. Mol. Biomed. 2, 32 (2021).
Mazzucco, G. et al. Telomere damage induces internal loops that generate telomeric circles. Nat. Commun. 11, 5297 (2020).
Vassin, V. M., Anantha, R. W., Sokolova, E., Kanner, S. & Borowiec, J. A. Human RPA phosphorylation by ATR stimulates DNA synthesis and prevents ssDNA accumulation during DNA-replication stress. J. Cell Sci. 122, 4070–4080 (2009).
Shi, W. et al. The role of RPA2 phosphorylation in homologous recombination in response to replication arrest. Carcinogenesis 31, 994–1002 (2010).
Pan, X. et al. FANCM suppresses DNA replication stress at ALT telomeres by disrupting TERRA R-loops. Sci. Rep. 9, 19110 (2019).
Silva, B. et al. FANCM limits ALT activity by restricting telomeric replication stress induced by deregulated BLM and R-loops. Nat. Commun. 10, 2253 (2019).
Feretzaki, M. et al. RAD51-dependent recruitment of TERRA lncRNA to telomeres through R-loops. Nature 587, 303–308 (2020).
Arora, R. et al. RNaseH1 regulates TERRA-telomeric DNA hybrids and telomere maintenance in ALT tumour cells. Nat. Commun. 5, 5220 (2014).
Lee, Y. W., Arora, R., Wischnewski, H. & Azzalin, C. M. TRF1 participates in chromosome end protection by averting TRF2-dependent telomeric R loops. Nat. Struct. Mol. Biol. 25, 147–153 (2018).
Brambati, A., Zardoni, L., Nardini, E., Pellicioli, A. & Liberi, G. The dark side of RNA:DNA hybrids. Mutat. Res. Rev. Mutat. Res. 784, 108300 (2020).
Ohle, C. et al. Transient RNA-DNA hybrids are required for efficient double-strand break repair. Cell 167, 1001–1013.e7 (2016).
Ngo, G. H. P., Grimstead, J. W. & Baird, D. M. UPF1 promotes the formation of R loops to stimulate DNA double-strand break repair. Nat. Commun. 12, 3849 (2021).
Audoynaud, C. et al. RNA:DNA hybrids from Okazaki fragments contribute to establish the Ku-mediated barrier to replication-fork degradation. Mol. Cell 83, 1061–1074.e6 (2023).
Kumar, C., Batra, S., Griffith, J. D. & Remus, D. The interplay of RNA:DNA hybrid structure and G-quadruplexes determines the outcome of R-loop-replisome collisions. Elife 10, e72286 (2021).
Stuckey, R., García-Rodríguez, N., Aguilera, A. & Wellinger, R. E. Role for RNA:DNA hybrids in origin-independent replication priming in a eukaryotic system. Proc. Natl Acad. Sci. USA 112, 5779–5784 (2015).
Chappidi, N. et al. Fork cleavage-religation cycle and active transcription mediate replication restart after fork stalling at co-transcriptional R-loops. Mol. Cell 77, 528–541.e8 (2020).
Song, L. et al. Dynamic control of RNA-DNA hybrid formation orchestrates DNA2 activation at stalled forks by RNAPII and DDX39A. Mol. Cell 85, 506–522.e7 (2025).
Xu, Z. et al. DDX39A resolves replication fork-associated RNA-DNA hybrids to balance fork protection and cleavage for genomic stability maintenance. Mol. Cell 85, 490–505.e11 (2025).
Fernandes, R. V., Feretzaki, M. & Lingner, J. The makings of TERRA R-loops at chromosome ends. Cell Cycle 20, 1745–1759 (2021).
Vujanovic, M. et al. Replication fork slowing and reversal upon DNA damage require PCNA polyubiquitination and ZRANB3 DNA translocase activity. Mol. Cell 67, 882–890.e5 (2017).
Mutreja, K. et al. ATR-mediated global fork slowing and reversal assist fork traverse and prevent chromosomal breakage at DNA interstrand cross-links. Cell Rep. 24, 2629–2642.e5 (2018).
Neelsen, K. J. & Lopes, M. Replication fork reversal in eukaryotes: from dead end to dynamic response. Nat. Rev. Mol. Cell Biol. 16, 207–220 (2015).
Bai, G. et al. HLTF resolves G4s and promotes G4-induced replication fork slowing to maintain genome stability. Mol. Cell 84, 3044–3060.e11 (2024).
Ray Chaudhuri, A. et al. Topoisomerase I poisoning results in PARP-mediated replication fork reversal. Nat. Struct. Mol. Biol. 19, 417–423 (2012).
Maya-Mendoza, A. et al. High speed of fork progression induces DNA replication stress and genomic instability. Nature 559, 279–284 (2018).
Quinet, A. et al. PRIMPOL-mediated adaptive response suppresses replication fork reversal in BRCA-deficient cells. Mol. Cell 77, 461–474.e9 (2020).
Sale, J. E. Translesion DNA synthesis and mutagenesis in eukaryotes. Cold Spring Harb. Perspect. Biol. 5, a012708 (2013).
Udroiu, I., Marinaccio, J., Goffi, R. S., Micheli, E. & Sgura, A. Specificity and sensitivity of ALT-associated markers in cancer cells. FEBS Lett. 599, 989–1005 (2025).
Ohki, R. & Ishikawa, F. Telomere-bound TRF1 and TRF2 stall the replication fork at telomeric repeats. Nucleic Acids Res. 32, 1627–1637 (2004).
Leman, A. R. et al. Timeless preserves telomere length by promoting efficient DNA replication through human telomeres. Cell Cycle 11, 2337–2347 (2012).
Lemaçon, D. et al. MRE11 and EXO1 nucleases degrade reversed forks and elicit MUS81-dependent fork rescue in BRCA2-deficient cells. Nat. Commun. 8, 860 (2017).
Kolinjivadi, A. M. et al. Smarcal1-mediated fork reversal triggers Mre11-dependent degradation of nascent DNA in the absence of Brca2 and stable Rad51 nucleofilaments. Mol. Cell 67, 867–881.e7 (2017).
Mijic, S. et al. Replication fork reversal triggers fork degradation in BRCA2-defective cells. Nat. Commun. 8, 859 (2017).
Somyajit, K., Saxena, S., Babu, S., Mishra, A. & Nagaraju, G. Mammalian RAD51 paralogs protect nascent DNA at stalled forks and mediate replication restart. Nucleic Acids Res. 43, 9835–9855 (2015).
Ait Saada, A., Lambert, S. A. E. & Carr, A. M. Preserving replication fork integrity and competence via the homologous recombination pathway. DNA Repair (Amst.) 71, 135–147 (2018).
Thangavel, S. et al. DNA2 drives processing and restart of reversed replication forks in human cells. J. Cell Biol. 208, 545–562 (2015).
Liu, W. et al. RAD51 bypasses the CMG helicase to promote replication fork reversal. Science 380, 382–387 (2023).
Decottignies, A. TERRA and RAD51AP1 at the R&D-loop department of ALT telomeres. Mol. Cell 82, 3963–3965 (2022).
Nguyen, T. T. et al. Oxidative stress at telomeres triggers internal DNA loops, TRF1 dissociation, and TRF2-dependent R-loops. Nucleic Acids Res. 53, gkaf285 (2025).
Haase, J. et al. The TFIIH complex is required to establish and maintain mitotic chromosome structure. Elife 11, e75475 (2022).
Acknowledgements
We thank T. Michiels and T. de Lange for plasmids; T. de Lange, J. Lingner, and W. Wright for cell lines. We are grateful to S. Coulon, E. Lazzerini Denchi, and B. Azeroglu for their stimulating discussions and assistance in designing the experiments. We are grateful to M. Mahieu, S. de Tocqueville, and P. Revy for help with the experiments. We thank N. Bettin for helpful comments on the manuscript. M.V. is supported by the King Baudouin Foundation (Funds Maurange) and the FNRS; E.C. was supported by the Télévie/FNRS. A.D is a recipient from the FNRS. AD’s lab is supported by the FNRS, the Télévie/FNRS, the King Baudouin Foundation, and UCLouvain. Y.D.’s lab is supported by the Associazione Italiana per la Ricerca sul Cancro, AIRC IG 28954, and Worldwide Cancer Research, WWCR 24-0166. J.K. is supported by NIH grants AG0773424, CA234047, P30CA014195, and GM142173. J.N. was supported by NCI-NIH grant K99-R00CA252447. C.M.A.’s lab is supported by Portuguese national funds through FCT–Fundação para a Ciência e a Tecnologia, I.P. PTDC/MED-ONC/7864/2020 and 2021.00143.CEECIND, by LaCaixa Foundation LCF/PR/HP21/52310016, and by TessellateBIO.
Author information
Authors and Affiliations
Contributions
M.V., E.C. and A.D. designed the project, analyzed the data, and wrote the paper. M.V. and E.C. performed most experiments. J.N., under the supervision of J.K., generated the PCS survivor cell lines. E.Z., under the supervision of Y.D., conducted 2D-gel and alkaline gel experiments (Figs. 3F, G, S3F, 6E, and S6C). J.R. and C.M.A. conducted DRIP experiments (Figs. 4A, S4B, and S5E). A.D. supervised the work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Vaurs, M., Claude, E., Zanella, E. et al. TRF1 relies on fork reversal to prevent fragility at human telomeres. Nat Commun 16, 6439 (2025). https://doi.org/10.1038/s41467-025-61828-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-61828-5