Abstract
The scarcity of cultivated gut bacteriophages hinders gut microbial research and application. Here we report the establishment and characterization of a Gut Phage Biobank (GPB) (https://db.cngb.org/genomics/datasets/GDS0000055) through a systematic isolation workflow and containing 104 isolates that target abundant or disease-associated gut bacteria. Genomic analysis reveals high diversity among these phages, and key genes of phage-bacteria interactions. The infection matrix demonstrates high host-specificity and varying infectivity of these phages under different conditions, unveiling phage-bacteria interaction mechanisms. In-depth characterization of the phages targeting obligate anaerobes uncovers a previously undescribed family and four previously undescribed genera, one of which is more prevalent than the well-known crAss-like phages globally except in Eurafrica. Cohort analysis reveals a higher prevalence of Mediterraneibacter and Dorea and a lower prevalence of Mediterraneibacter phages in Asian disease population. In vitro and in vivo evidence of phage inhibiting Dorea highlight the potential of phages in disease intervention. This biobank represents a valuable resource for advancing gut microbial research and holds promise for manipulating microbiomes.
Similar content being viewed by others
Introduction
The human gastrointestinal tract is a complex ecosystem harboring diverse microorganisms, with gut bacteria playing a central role in human diseases development1,2,3,4,5. Interventions such as probiotics, prebiotics, and fecal microbiota transplantation (FMT) have been explored to address chronic diseases associated with gut bacteria. Probiotics offer therapeutic benefits through beneficial microorganisms, while prebiotics selectively stimulate the growth of beneficial bacteria in the gut6,7. While FMT may restore gut microbiome homeostasis, it currently lacks precise regulation and control8,9. Bacteriophages, which specifically target and infect bacteria, have been successfully used in the treatment of multidrug-resistant infections10. In addition to probiotics, prebiotics and FMT, phages provide a targeted approach to address gut pathogens11,12, offering precise interventions for various gut bacteria-related diseases13,14,15,16,17. This approach complements these interventions in modulating the gut microbiome for therapeutic purposes.
To enhance the understanding and utilization of gut phages, several human gut virome databases have been established18,19,20,21. Although these virome databases collectively contain over one million viral contigs in total, the number of cultured phages is only 0.024% of that of viral contigs22,23,24,25,26,27,28. This highlights the necessity of isolating and identifying more gut phages to advance virome research and microbiome diversity. Increased phage isolates will broaden scope for research in taxonomy and biological traits. For example, crAss-like phages were fully characterized only after more than a decade of genome sequencing research, which became possible once these phages could be cultured in pure form24,29,30, leading to the formal recognition of the Crassvirales order by the International Committee on Taxonomy of Viruses (ICTV).
In addition to these efforts, researchers are exploring the limited availability of gut phages to investigate their interactions with gut bacteria, often revealing mutualistic, rather than antagonistic dynamics31,32. For instance, Bacteroides bacteria may use invertible promoters to mediate rapid phase variation of alternate capsular polysaccharides, balancing phage sensitivity and resistance29,33. However, understanding of the genetic elements and specific interactions between Bacteroides phages and bacteria is limited to a small subset of phages. Additionally, research investigating the interactions between phages and other obligate anaerobes, as well as between phages and facultative anaerobes under oxic and anoxic conditions, remains limited. To uncover mechanisms of phage-bacteria interactions under different conditions, it is crucial to isolate and characterize additional gut phage isolates.
Researchers utilized various methods for phage isolation. Briefly, these methods can be categorized into three types: (1) Direct plating to isolate phages from samples or concentrated samples33, identifying the desired phage among randomly distributed relatives. (2) Repetitive or concentration-based isolation through co-culturing with the candidate host25 to enhance the abundance of the target phage. (3) Screening with known genomic data, where qPCR assays are employed to identify samples with higher phage abundance, improving isolation efficiency22. Despite these approaches, developing systematic method for isolating previously undescribed phages remains a critical challenge.
In this study, we construct a gut phage biobank (GPB) through a comprehensive isolation of phages targeting abundant or disease-associated gut commensal bacteria. We analyze their genomic characteristics and the genes involved in phage-bacteria interactions. We conduct infection matrix analysis to assess the host specificity and infectivity of the phages under both oxic and anoxic conditions, while identifying potential factors contributing to these interactions. We investigate crAss-like phages by comparing their genomic and biological characteristics with previously reported crAss-like phages. Additionally, we conduct a detailed characterization of phages targeting obligate anaerobes, including those from orders Bacteroidales, order Lachnospirales and phylum Actinomycetota. We assess the prevalence and relative abundance of GPB phages across different continental and disease cohorts. We analyze the prevalence and relative abundance of Mediterraneibacter and Dorea bacterial strains, as previous studies have highlighted their association with obesity and T2D in European cohorts, along with the distribution of their corresponding phages, to verify whether similar association exist in Asian disease and health cohorts. Finally, we assess the ability of phage to inhibit the obesity-associated Dorea strain both in vitro and in vivo. This research provides valuable resources for manipulating the gut microbiome and offers insights for further research and potential therapeutic applications of gut phages and the virome.
Results
A comprehensive workflow for gut phage isolation
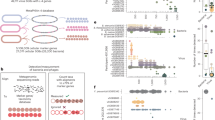
To address the challenges of gut phage isolation, we consolidated an integrated workflow by optimizing three complementary approaches (Fig. 1A). The classic plaque assay method involves directly co-culture followed by plaque screening. The repetitive enrichment method introduces a six-day, multi-cycle culture to enhance the recovery of slow-replicating or temperate phages. This approach may enhance the efficiency of isolating the phages with longer replication times and temperate phages. The third method leverages existing metagenomic data to selectively target phage sequences, improving efficiency for samples with prior sequencing.
A The comprehensive workflow for efficient gut phage isolation. The classic method involves a single co-culture of the bacterial host with the source sample, followed by concentration for plaque assays. The repetitive enrichment method enhances this approach by introducing multiple rounds of co-culture to increase the abundance of targeting phages. The metagenomics-guided method further refines the process by incorporating a metagenomic analysis step before selecting samples for repetitive co-culture and concentration. The numbers below each method indicate the number of phage isolates obtained and the corresponding number of infected bacterial hosts. B The three-column Sankey diagram visualizes the host species of GPB phages (middle column), whether these hosts are abundant within the human gut (left column), and the types of diseases associated with these host bacteria (right column). The Venn diagram illustrates the number of phages targeting highly abundant gut bacteria, disease-related gut bacteria, and their overlap.
We focused on 29 gut bacterial species (Supplementary Data 1) selected for either high abundance or disease association14,34,35,36,37,38. For the samples lacking metagenomic data, we applied the classic and repetitive enrichment method, yielding 81 phages for 56 bacterial strains, and 16 phages for 12 bacterial strains, respectively. To improve isolation of phages targeting obligate anaerobes, we employed metagenomics-guided screening on 12 preselected fecal/saliva samples (from 47 candidates, Figure S1), identifying7 additional phages targeting 6 bacterial strains.
Comparative analysis revealed key insights: (1) No phage overlap between isolation approaches, indicating that the three isolation approaches are complementary. (2) The classic method succeeded primarily with low-complexity samples, such as wastewater and lake water. In contrast, repetitive enrichment was essential for high-diversity samples, such as animal farm samples and feces, suggesting sample-dependent efficacy. (3) Classic isolation captured both facultative and obligate anaerobes, while repetitive enrichment yielded phage only against facultative anaerobes. Metagenomics-guided isolation enabled precise targeting of obligate anaerobes, by selecting 12 from a total of 47 fecal and saliva samples (Figure S1), significantly reducing both experimental time and sample requirements. Taken together, our findings demonstrate that employing multiple isolation strategies alongside diverse sample sources is essential for maximizing gut phage diversity.
In total, we isolated 104 phages targeting 4 bacterial phyla (51 Bacillota, 30 Pseudomonadota, 16 Bacteroidota, 7 Actinomycetota), 14 genera, and 17 species (Figure1B, Fig. 2A, Table 1 and Supplementary Data 1). Notably, 95% (99/104) of the isolated phages target disease-associated bacteria, 40% (42/104) infect highly abundant gut species, and 36% (37/104) exhibit both traits (Fig. 1B) - a deliberate contrast to broader isolation efforts like the Gut Phage Isolate Collection (GPIC)25.
A Host species distribution of the GPB phages (n = 104). Columns show the number of phages targeting each bacterial species, with host phyla indicated by the top legend. B Genome length of all sequenced GPB phages (n = 102), grouped by host species. Data are presented as box plots where the center line indicates the median, the box boundaries represent the 25th and 75th percentiles, and whiskers extend to 1.5 times the interquartile range (IQR). Dashed black lines at 20 kbp and 200 kbp highlight the broad genome size range. C Annotation rate of all sequenced GPB phages (n = 102), grouped by host species. The annotation rate is the ratio of functionally annotated CDSs to the total CDSs per phage. Data are presented as box plots where the center line indicates the median, the box boundaries represent the 25th and 75th percentiles, and whiskers extend to 1.5 times IQR. D Genome organization of all sequenced GPB phages (n = 102). Phages are arranged by genome length (pink lines) from largest to smallest. Box colors categorize genomes by size: >80 kbp (orange), 20−80 kbp (blue), and <20 kbp (yellow). The gene size distribution for each individual phage is presented as a box plot, colored according to the phage’s total genome size category: >80 kbp (orange), 20−80 kbp (blue), and <20 kbp (yellow). Each box plot displays the median (center line), 25th-75th percentiles (box boundaries), and whiskers extending to 1.5 times IQR. Genes larger than 5 kbp are labeled, and those with shared functional annotations are identically colored. E Heatmap of phage-bacteria interaction genes across GPB phage genomes. Rows represent functional gene categories, and columns represent individual phages (n = 102). The color intensity of blue indicates the number of genes in each category per phage. Host species for each phage are shown by the colored boxes at the top.
The diversity and genomic characteristics of the phages within GPB
We constructed a gut phage biobank (GPB) (https://db.cngb.org/genomics/datasets/GDS0000055) with isolated phages and deposited them in CNGBdb-EBB (E-BioBank) (https://db.cngb.org/ebb/bio_resources/) under accession numbers CNSebb2036927 - CNSebb2037028. The genomes of all the phage isolates were sequenced and submitted to the CNGB Sequence Archive (CNSA)39 of the China National GeneBank DataBase (CNGBdb)40 with accession number CNP0004979.
All GPB phages are double-stranded DNA (dsDNA) phages, with genome sizes ranging from 8,912 bp to 371,963 bp and GC content from 29 mol % and 59.25 mol % (Fig. 2B and Supplementary Data 1). Notably, phages targeting Cronobacter sakazakii, Escherichia coli, and Klebsiella pneumoniae have genomes exceeding 200 kbp, while four Eggerthella lenta phages and six Dorea longicatena phages are smaller than 20 kbp. All GPB phages belong to the Caudoviricetes class, which is predominant in the human gut virome. Gene-sharing network analysis of 102 GPB phages, 4,534 NCBI RefSeq phages, and a subcollection from four human gut virome databases (n = 13,476)18,19,20,21 identified 1,393 viral clusters (VCs) and 778 singletons (Figure S2 and Supplementary Data 2). 95 GPB phages were assigned to 38 VCs, corresponding to viral genus-level taxonomy, with three singletons and seven overlaps. Notably, 39 phage isolates across 21 VCs did not cluster with any RefSeq phages, suggesting that they represent isolates belonging to previously undescribed phage genera. These findings underscore the diversity in genome size, GC content, taxonomy, and gene-sharing network within the GPB phages.
The functional protein annotation rates for gut phages range from 21.88% to 91.49% (Fig. 2C), reflecting the high diversity of the GPB phages. Gene sizes range from 90 bp to 10 kbp, with 86% of genes smaller than 1 kbp and 97% smaller than 2.5 kbp (Fig. 2D). Notably, 21 genes exceed 5 kbp. Phages with genomes larger than 80 kbp tend to encode multiple genes over 5 kbp, while those between 20−80 kbp predominantly encode genes smaller than 5 kbp, except for four tail fiber proteins. Phages with genomes smaller than 20 kbp exclusively encode genes shorter than 2.5 kbp (Fig. 2D).
We focused on genes involved in phage-bacteria interaction, including those for adsorption, antimicrobial immunity, nucleic acid replication and transcription, progeny phage release, virulence factors, and antibiotic resistance (Fig. 2E and Supplementary Data 3). Three phages encode virulence factors that may enhance host bacterial virulence, but no drug-resistance genes were found. Except for four phages with incomplete genomes, all encode genes for adsorption and lysis, which are essential for bacterial infection. Additionally, 91 phages encode replication and transcription proteins, 69 encode antimicrobial immunity proteins, and 26 encode lysogenic proteins (Fig. 2E). Notably, 37 phages encode gene for the entire lytic cycle - adsorption, anti-immunity, replication and transcription, and lysis. This comprehensive gene set may provide these phages with greater autonomy in their reproduction compared to phages lacking genes for certain stages, which rely more heavily on host bacterial machinery.
Previous studies have shown that some phages carry auxiliary metabolic genes (AMGs), which can alter host metabolism and influence the entire microbial ecosystem41,42. We identified 56 AMGs in 26 gut phage genomes. The largest phage, CPB1014 ( ~ 371 kbp), contains six AMGs, while others carry fewer than four. Nine phages carry both AMGs and lysogenic genes (Fig. 2E and Supplementary Data 4). The 56 AMGs are involved in various metabolic processes, including cofactors and vitamin metabolism (26.8%), amino acid metabolism (21.4%), and other secondary metabolites biosynthesis (16.1%), terpenoids and polyketides metabolism (12.5%), energy metabolism (10.7%), folding, sorting, and degradation (8.9%), and glycan biosynthesis and metabolism (3.6%). These AMGs may modulate the phage-bacteria interaction by influencing the bacteria metabolism.
Phage-bacteria interactions under oxic and anoxic conditions
We challenged 40 bacterial strains (17 facultative anaerobes, 23 obligate anaerobes) from 17 species with 99 phage isolates under oxic/anoxic conditions (Fig. 3A). The results showed 17 distinct phage-bacteria interaction profiles, with each profile representing phages specific to particular bacterial species. These findings demonstrate that gut phages exhibit remarkable host specificity, minimal cross-species infectivity.
A Heatmap of host range assays for 99 GPB phages against 40 bacteria strains (17 species). Phages targeting facultative anaerobes were tested under both oxic and anoxic conditions. Phages targeting obligate anaerobes were tested only under anoxic condition. Purple blocks represent infections occurred under both anoxic and oxic conditions, blue under oxic only, and red under anoxic only. Phages are clustered by whole genome ANI within each host species. Colored sidebars represent bacteria species. B Heatmap of host range assays for E. faecalis phages. Columns represent bacterial strains clustered by ANI (98% cut-off). Rows represent phages clustered by ANI (99% cut-off). Red blocks indicate infections. Colors labeled aside and above represent bacterial and phages clusters. Fill colors indicate phage infectivity: blue for oxic-only, purple for both conditions. C Simulated linear comparison of PTS system CDSs among 4 E. faecalis strains. Arrows refer to annotated CDSs for three PTS system components. Same-colored arrows indicate CDSs with >95% identity and are linked with grey shading. Red-framed arrows indicate CDSs did not match any other CDS with the same functional annotations. D-E The killing curves of CPB1036 targeting E. faecalis JLDX0001 at MOI = 1 and 10 under oxic condition (D) and anoxic condition (E). Data are represented as mean ± standard deviation (SD) from three technical replicates. F Heatmap of host range assays for M. gnavus phages. Columns represent bacteria strain. Rows represent phages clustered by ANI with a cut-off of 99%. Red blocks indicate infections. Colored sidebars represent phages groups. G Multiple sequence alignment of two tail-associated proteins that exhibit inter-group differences. Red represents identical sequences across the inter-group, yellow indicates shared particle sequences, and unhighlighted regions show divergence. H Heatmap of host range assays for B. fragilis phages. Columns represent bacteria strains. Rows represent phages. Red blocks indicate infections. I Mimetic gene structure of B. fragilis RTGS0060 and four B. fragilis phages. Purple diamonds indicate CRISPR repeats, and same-colored rectangles represent CRISPR spacers that match phage sequences.
We first explored phage host range for facultative anaerobes under oxic and anoxic conditions. For Enterococcus faecalis phages under oxic condition, eight bacterial strains clustered by genomic similarity showed consistent phage sensitivity, linked to variations in phosphotransferase system (PTS) systems but not phage receptor binding proteins (Fig. 3B-C). For C. sakazakii under anoxic condition, all three strains were infected by five phages, with RTGS0181 showing broader susceptibility due to distinct peptidoglycan-associated lipoprotein (PAL) structures (Figure S3A-B). Oxygen conditions influenced infectivity: 25 of 26 E. faecalis phages formed plaques under both conditions, C. sakazakii phages infected fewer hosts under oxic condition, while Lacticaseibacillus paracasei phage, isolated under anoxic condition, only formed plaques under such condition (Fig. 3A). We further assessed phage-mediated bacterial inhibition in liquid culture under both oxic and anoxic conditions (Fig. 3D-E and Figure S4). Most phages exhibited similar bactericidal kinetics regardless of oxygen availability (Figure S4A-L). However, one E. faecalis phage and two B. cereus phages showed reduced inhibition of their specific host under anoxia compared to oxia (Fig. 3D-E and Figure S4M-N). Host range assays revealed strict species/strain specificity of gut phages, suggesting limited cross-species interactions.
We next investigated the host range of phages targeting obligate anaerobes under anoxic condition. For Mediterraneibacter gnavus (formerly Ruminococcus gnavus) phages, seven were classified into four genomic groups, with Group 2 and 4 showing variable host infectivity within groups (Fig. 3F). In Group 2, comparative genomic analysis revealed a 7-amino acids deletion in the tail protein of phage CPB0986 and a 162-amino acids deletion in CPB1098 relative to CPB1099 (Fig. 3G and Figure S5). In Group 4, the tail tape measure protein of CPB1094 showed a 21-amino acids deletion compared to CPB1088 (Fig. 3G and Figure S6), suggesting these structural variations may influence host range. In the host range assays of Bacteroides fragilis phages, phage CPB0999 showed distinct infectivity, while CPB0997, CPB0998, and CPB1129 shared two host, with CPB0997 additionally showing weak infectivity (turbid plaques) against RTGS0060 (Fig. 3H). Genomic comparison revealed >93% coverage and >99% identity among CPB1129, CPB0998 and CPB0997. We further explored whether RTGS0060 carries any phage defense systems43, and found that RTGS0060 carries CRISPR-Cas systems with spacers that match all four B. fragilis phages, explaining host resistance (Fig. 3I). Given that CPB0997 was isolated through co-culture with RTGS0060, its limited infectivity against this strain may result from adaptive evolution during repeated exposure.
Collectively, our dual oxygen condition analysis reveals: (1) Oxygen-dependent phage-host interaction dynamics affecting both host range and bactericidal efficacy; (2) Mechanistic roles of bacterial surface components (like PTS and PAL) and phage tail proteins; (3) CRISPR-Cas mediated resistance overcoming through iterative co-culture. These findings underscore the need for oxygen-matched isolation strategies when targeting facultative anaerobes.
Previously characterized phages targeting obligate anaerobes
Given the scarcity of phages infecting obligate anaerobes, our study concentrated on these specific phages. The GPB comprises 38 phage isolates infecting obligate anaerobic bacteria, including 8 Bacteroides phages, 7 M. gnavus phages, 7 D. longicatena phages, 6 E. lenta phages, 4 Parabacteroides distasonis phages, 4 Phocaeicola vulgatus phages, 1 Bifidobacterium longum phage, and 1 Clostridium perfringens phage. We selected several phage candidates displaying distinct genomic and phenotypic characteristics for comprehensive analysis to better understand the interactions between GPB phages and their respective bacterial hosts (Supplementary Results and Figure S7-11).
Gene-sharing network analysis revealed four GPB phages clustering with crAss-like phages, the most abundant phages in the human gut microbiome. These included two Bacteroides cellulosilyticus phages, one Bacteroides thetaiotaomicron phage, and one P. distasonis phage (Supplementary Results). Notably, the Parabacteroides-infecting phage is unique, as previously reported crAss-like phage isolates typically infect Bacteroides22,23,24,28. Phylogenetic analysis of their large terminase subunit (TerL) alongside 100 reference crAss-like genomes from ICTV, including 31 known strains22,23,26,28,44, placed them into four families: Crevaviridae (3 species), Intestiviridae (19 species), Steigviridae (22 species), and Suoliviridae (36 species) (Fig. 4A). Accordingly, CPB1101 and CPB1112 occupied distinct clades, while CPB1000 and CPB1023 clustered with phage Bc03. Notably, the Steigviridae family contained the highest proportion of isolated phages and had a significantly more tRNA genes, suggesting that the increased tRNA genes content may enhance isolation potential. In contrast, crAss-like phages from other families may possess distinct lifecycles or host interactions (e.g., non-lytic replication), limiting plaque assays detection.
A The phylogenic tree of large terminase subunits of GPB crAss-like phages and previously reported crAss-like phages. Phages labeled with circles represent crAss-like phages that can form plaques. previously reported ones are labeled with empty circles (○) while the ones isolated in this study are labeled with filled circles (●). The phage labeled with a cultural tube indicates that it can propagate but does not form plaques. The adjacently located strains boxed by the dotted squares represent them belonging to the same species at the ANI cutoff of 95%. Colored shades represent taxonomy as depicted in the legend. The outgroup represents Cellulophaga phage phi13:2. B The genomic similarity of isolated M. gnavus phage isolates. Phages isolated in this study are indicated by pink at the top, while previously isolated phages are indicated by blue. The heatmap is clustered based on the genomic similarity, with the intensity of the red color indicating the genomic similarity between each two phages. Clusters containing GPB phages are highlighted with boxes, where green represents a previously undescribed phage genus. C The genomic similarity of isolated P. vulgatus phage isolates. Phages isolated in this study are indicated by pink at the top, while previously isolated phages are indicated by blue. The heatmap is clustered based on the genomic similarity, with the intensity of the red color indicating the genomic similarity between each two phages. Clusters highlighted with green boxes represent previously undescribed phage genus or family composed of GPB phages. D Phylogenetic tree of D. longicatena phages. The phages with pink color represent previously undescribed phage genus discovered in this study, while the phages with blue color represent D. longicatena phage contigs from the GPD database. E The multiple sequence alignment results of D. longicatena phages. Phages in the red text represent GPB phages. The arrows indicate the direction of the CDSs, with different colors representing various functional categories. Gray regions represent sequence similarity greater than 70%. The two CDSs with opposite orientations (phage DNA encapsidation protein and holin family protein) are highlighted with blue-bordered arrows.
Beyond the four crAss-like phages, we examined M. gnavus, D. longicatena and P. vulgatus phages due to the scarcity of available isolates (Supplementary Results). For M. gnavus phages, Average Nucleotide Identity (ANI) analysis of our seven isolates and six previously reported ones revealed three distinct groups (Fig. 4B). Four of our isolates clustered with known ones (Group 1), while CPB1088 and CPB1094 formed Group 2, and CPB1090 comprised Group 3, indicating they may represent distinct lineages. Gene-sharing network analysis confirmed these three phages belong to clusters without RefSeq overlaps, representing two undescribed genera (Supplementary Data 2). For P. vulgatus phage, genomic comparison of our four isolates with 15 reported ones identified two uncharacterized groups (Fig. 4C). Gene-sharing network analysis revealed that CPB1107 and CPB1128 clustered with 278 unidentified genomes, indicating they are uncharacterized isolates from an undescribed genus, while CPB1111 and CPB1114, identified as singletons, likely represent distinct and undescribed genera (Supplementary Data 2).
Given the lack of previously reported isolates of the D. longicatena phages, we examined their evolutionary relationship with 834 D. longicatena phage contigs identified in the Gut Phage Database (GPD). This analysis revealed significant divergence, with our isolates formed a distinct clade, indicating their uniqueness (Fig. 4D). Gene-sharing network analysis showed that our 7 phages clustered with 9 host-unknown contigs, forming 16 phages with conserved genomic organization divided into two sub-clusters (Figure S11C). Notably, all coding sequences (CDSs) shared uniform orientation, except for a rare back-to-back arrangement between a DNA encapsidation protein and a holin gene (Fig. 4E). We hypothesized that these phages may employ a common promoter to optimize genome organization45,46, potentially enhancing early phage DNA encapsulation and late holin expression, indicating a previously uncharacterized phage family with unique genomic characteristics.
Geographical distribution of GPB phages
Since crAss-like phages are regarded as the most abundant bacteriophages in the gut23,24,47, we assessed the abundance and prevalence of our crAss-like phages alongside other GPB phages across ten metagenomic cohorts from five continents (Fig. 5A, S12A and Supplementary Data 5). We identified 32 GPB phages with a global prevalence equal to or exceeding that of ΦcrAss001, including 25 targeting obligate anaerobes - M. gnavus, P. vulgatus, P. distasonis, B. fragilis, B. cellulosilyticus, B. thetaiotaomicron E. lenta, and B. longum and 7 targeting facultative anaerobes - K. pneumoniae, E. faecalis, C. sakazakii, and E. coli. Our four crAss-like phages exhibited relative abundance similar to previously reported crAss-like phages (Figure S12A), although their prevalence varied from 0% to 18.85% (Fig. 5A). Additionally, four P. vulgatus phages and two P. distasonis phages (CPB1100 and CPB1133), representing isolates of undescribed genera, exhibited higher global prevalence (1.76% to 2.91%) than our crAss-like phages. Notably, two M. gnavus phages (CPB1094 and CPB1088), classified as representatives of an uncharacterized genus, showed higher global prevalence than p-crAssphage, despite similar abundance (Fig. 5A and S12A).
A The prevalence of GPB phages and crAss-like phages in ten cohorts from six continents. Each column represents a phage strain, the colored columns represent the host species of these phages depicted in the legend (legend ‘Control’ represents the crAss-like phages reported). Prevalence analysis was performed according to previous literature17 with a subtle refinement, specifically, a phage was considered present in a sample if the mapped reads covered ≥60% of the genome length. The GPB phages with very high prevalence are labeled in red fonts, the GPB crAss-phages are labeled in orange fonts, and the representative crAss-like phages reported before are labeled in green fonts. B The prevalence of GPB phages, p-crAssphage, and ΦcrAss001 in each continent. The base map was sourced from China’s Standard Map Service (Approval No. GS(2016)1666). C Left - Prevalence of M. gnavus strains in normal healthy cohort and obesity cohort from Asia. Right - Prevalence of M. gnavus strains in normal healthy cohort and T2D cohort from Asia. D Left - Prevalence of M. gnavus phages in normal healthy cohort and obesity cohort from Asia. Right - Prevalence of M. gnavus phages in normal healthy cohort and T2D cohort from Asia. E Left - Prevalence of D. longicatena strains in normal healthy cohort and obesity cohort from Asia. Right - Prevalence of D. longicatena strains in normal healthy cohort and T2D cohort from Asia.
We then analyzed the geographical distribution of these GPB phages, focusing on the 25 isolates targeting obligate anaerobes and two representative crAss-like phages (p-crAssphage and ΦcrAss001) (Fig. 5B). All 25 GPB phages were detected in the Asian cohorts, with the two M. gnavus phages exhibiting the highest prevalences, which may be because the isolated samples used in this study originated from Asia. In European and African cohorts, p-crAssphage was most prevalent, while the two M. gnavus phages showed a slightly lower prevalence. Despite regional prevalence, a similar distribution pattern was observed across Asia, Europe, and Africa, potentially due to the proximity of the Afro-Eurasian continent. In both North and South America, the two M. gnavus phages were the most prevalent, though other phages were below the detection limit. The distribution of GPB phages in Oceania showed a distinct pattern, with the fewest phages and significantly lower prevalence overall. Only the two M. gnavus phages and p-crAssphage were detected, likely due to the region’s geographical isolation. These findings emphasize the importance of considering geographic variation in studying the distribution of gut microbiome, especially phages.
Metabolic diseases-associated bacteria and phages in Asian cohorts
Obesity and T2D have been reported to be associated with an imbalance in gut microbiota, with high relative abundances of M. gnavus and D. longicatena suggesting a potential association with these diseases, particularly in European populations48,49,50,51. However, given the various geographic distributions of gut phages, we hypothesized that the distribution of these bacteria and their phages may differ in Asian disease cohorts. To investigate this, we utilized Asian disease and health cohorts to evaluate the distribution of these bacteria and their phages. This will enable us to further explore the interplay between phages, bacteria, and human health.
In Asian cohorts, M. gnavus showed significantly higher prevalence and relative abundance in the obesity cohort compared to the healthy cohort, though no marked differences were observed between the T2D and healthy cohorts (Fig. 5C and S12B). Additionally, when assessing the prevalence and relative abundance of M. gnavus phages in the same cohorts, we found that five of the seven phages exhibited consistently low prevalence and relative abundance across all cohorts, whereas the globally prevalent M. gnavus phages CPB1094 and CPB1088 displayed higher prevalence in the healthy cohort, with comparable relative abundance (Fig. 5D and Figure S12C). D. longicatena exhibited higher prevalence in the Asian obesity cohort, despite similar relative abundance, while no notable differences were observed between the Asian T2D and healthy cohorts (Fig. 5E and S12D). Interestingly, no D. longicatena phages were detected in any cohort. In summary, M. gnavus and D. longicatena exhibited higher prevalence in the Asian obesity cohorts compared to healthy cohorts, consistent with results in European cohorts, whereas M. gnavus phages showed lower prevalence in the Asian obesity and T2D cohorts compared with healthy cohorts. These findings reveal the potential of M. gnavus phages, M. gnavus and D. longicatena as biomarkers for metabolic disease and as targets for our downstream intervention experiments.
Inhibition of D. longicatena by phage in vitro and in vivo
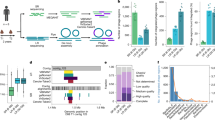
Gut phages offer a precise approach for manipulating the gut microbiome, with potential therapeutic applications for diseases linked to dysbiotic gut microbiota by selectively inhibiting the proliferation of specific bacterial hosts. In this study, we aimed to assess the inhibitory efficacy of phage targeting M. gnavus and D. longicatena strains, both implicated in obesity, through in vitro and in vivo experiments to evaluate their utility in obesity management. In vitro killing assays revealed that phages targeting M. gnavus exhibited limited inhibitory activity, whereas the D. longicatena phage CPB1092 displayed robust inhibition within 12 h (Supplementary Data 1 and Figure S13A-D). Genomic sequencing revealed the presence of lysogenic-related genes in the M. gnavus phages, likely attenuating their lytic activity, while no such genes were detected in the D. longicatena phage genomes. Consequently, we selected phage CPB1092, targeting D. longicatena strain RTGS0005, for in vivo studies, following established protocols with modifications52 (Fig. 6A). Although in vivo killing kinetics showed no significant differences between MOI of 1 and 10, we choose an MOI of 10 for in vivo experiments to maximize phage-host interactions and ensure effective infection.
A Murine phage therapy workflow. Six-week-old SPF WT C57BL/6 male mice (n = 15) were treated with four antibiotics by gavage for one week, washout for two days, then received D. longicatena RTGS0005 (107 CFU) at Day 0. The mice of phage-treated groups (n = 5) were given phages at MOI 10 from Day 1 to Day 4. Saline was used as a vehicle to prepare bacteria and phage solution. The phage-free groups (n = 5) were given vehicle (saline) instead of phage, and the blank group (n = 5) received neither bacteria nor phages except vehicle (saline). Stools were collected 12 hours after each gavage. Created in BioRender. Chen, Wan. (2025) https://BioRender.com/3oldu7j (B) Normalized CFU in murine phage therapy of phage CPB1092 targeting D. longicatena RTGS0005. Ten mice were orally administered D. longicatena RTGS0005 on Day 0 and then divided into a phage-free control group (n = 5) and a phage-treated group (n = 5). Data are presented as bar charts showing the mean ± SD of normalized CFU for the initial cohort at baseline (Day 0, gray bar, n = 10), the phage-free control group on Day 3 (red bar, n = 5), and the phage-treated group on Day 3 (blue bar, n = 5). Normalized CFU was calculated by dividing the CFU count of each mouse by a mean CFU value of a separate blank group (n = 5) at the corresponding time point. All individual data points are superimposed on the plot. Statistical significance between the phage-free and phage-treated groups on Day 3 was assessed using two-sided Mann-Whitney test, and the exact P value is shown. ns, not significant. C-D Relative abundance of (C) Dorea and (D) Dorea phages in murine samples during phage therapy of phage CPB1092 targeting D. longicatena RTGS0005. Mice were assessed at baseline (Day -1 and Day 0; gray, n = 10). From Day 1, mice were divided into a phage-free control group (red, n = 5) and a phage-treated group (blue, n = 5). Data are presented as box plots where the center line indicates the median, the box boundaries represent the 25th and 75th percentiles, and whiskers extend to 1.5 times the IQR. All individual data points are superimposed on the plot. Statistical significance between the phage-free and phage-treated groups at each time point (Day 1−4) was assessed using two-sided Mann-Whitney test, and the exact P value is shown. ns, not significant.
To investigate the effects of phage therapy on gut bacterial populations, we treated specific-pathogen-free (SPF) mice (n = 15) with a one-week regimen of four antibiotics, followed by a two-day washout period. The mice were then randomly allocated into three groups (n = 5 per group). The control group (n = 5) received a saline suspension throughout the experiment. The remaining two groups (n = 10), designated as the phage-treated and phage-free groups, were colonized with 107 CFU of a specific bacterial suspension one day prior to the initial phage administration (Day 0). Starting on Day 1, the phage-treated group (n = 5) was administered 108 PFU (MOI 10) of corresponding phages via oral gavage daily from Day 1 to Day 4, while the phage-free group (n = 5) received a saline vehicle at equivalent time points. We collected fecal samples 12 hours post-gavage to quantify residual host bacteria via CFU enumeration and to assess the relative abundance of the host bacteria and associated phages through metagenomic sequencing (see Methods).
CFU enumeration indicated low initial colonization of D. longicatena RTGS0005 on Day 1. By Day 3 of phage administration, CFU counts revealed robust bacteria proliferation in the phage-free group (Fig. 6B). In contrast, the phage-treated group exhibited a significant suppression of bacterial growth compared with the phage-free group (P = 0.029) (Fig. 6B). By Day 4, metagenomic sequencing demonstrated a significant reduction in the relative abundance of D. longicatena in the phage-treated group relative to the phage-free group (P = 0.029) (Fig. 6C). Phage presence was consistently detected in the phage-treated group from Day 1 until a marked decline on Day 4, with no phage was detected in the phage-free group (Fig. 6D).
Notably, the decline in phage relative abundance coincided with the reduction in host bacterial abundance, suggesting that phage persistence is dependent on host bacterial populations. Furthermore, metagenomic analysis revealed one-day delay in detecting significant differences in bacterial abundance compared to CFU enumeration, potentially attributable to persistence of bacterial DNA in the fecal samples. These findings underscore the therapeutic potential of GPB phages, such as D. longicatena phages, for modulating the gut microbiome to promote human health.
Discussion
In this study, we devised a systematic workflow for isolating gut phages, culminating in the creation of the Gut Phage Biobank (GPB), which contains phages targeting abundant or disease-relevant gut commensal bacteria. Within the GPB, we identified an undescribed phage family and several previously uncharacterized genera, suggesting that phage cultivation is essential for elucidating the dark matter of gut phages. The diversity and prevalence of phages within the GPB make them representatives of human gut phages, providing deeper insights into gut microbiome composition. Bioinformatic analyses of GPD revealed that 35.49% of phages infect multiple species, with some exhibiting extensive host range, even spanning across phyla (0.13%)18. In contrast, GPB phages showed high host species specificity, particularly those targeting Bacteroides and Parabacteroides, consistent with prior study on isolated gut phages25. Additionally, the species specificity and infectivity of GPB phages were influenced by cultivation conditions such as media compositions and oxygen availability, highlighting the complexity of phage-bacteria interactions. Notably, the inhibitory effects of Dorea-targeting phages in vitro and in vivo emphasize their potential as therapeutic tools for manipulating the gut microbiome.
Initially, our study aimed to investigate both lytic and lysogenic phages, focusing on their bacterial inhibition and bacterial hosts interactions. However, challenges in inducing sufficient prophage for comprehensive biological and genomic analyses led us to prioritize lytic phages, which offer immediate potential for bacterial inhibition and disease intervention. Consequently, we concentrated on phages capable of reliably forming plaques. Genomic analysis identified lysogenic genes in 26 GPB phages, with 20.99% (17/81) isolated using classic isolation and 28.13% (9/32) obtained via enrichment methods, including metagenomics-guided isolation. This suggests that enrichment methods may better capture non-traditional lytic phages or uncover diverse phage lifecycles present in gut samples. To improve the capture of temperate or carrier-state phages53, which are challenging to form plaques, optimizing plaque-free culturing methods and incorporating sequencing- guided detection will be essential.
The interactions between gut phages and their bacterial hosts are highly complex. Using metagenomics-guided isolation, we predicted the presence of phages targeting the bacteria of interest in 12 fecal samples, however, phages were isolated from only seven. This discrepancy may be attributed to several factors, including bacterial metabolic changes during culturing, adaptive immune mechanisms, and quorum-sensing mechanisms, which may influence phage-host dynamics and our ability to isolate phages33,45,54,55,56. Furthermore, killing kinetics showed that while some phages initially inhibited bacterial growth, 70% of experiments demonstrated an increase in OD at later stages. For instance, Dorea phages inhibited bacterial growth during the initial 8−12 hours, but growth resumed thereafter, suggesting the emergence of phage resistance populations (Fig. 3, Fig. 4 and Figure S8C-D). Different Dorea strains exhibited varying resistance patterns, with RTGS0004 developing resistance more rapidly than RTGS0005, indicating strain-specific resistance mechanisms. These findings emphasize the dynamic nature of phage-bacteria interactions and underscore the need to further explore resistance mechanisms to improve the effectiveness of phage-based interventions.
In both in vitro and in vivo experiments targeting D. longicatena associated with obesity, significant bacterial inhibition was observed, although complete eradication was not achieved. This aligns with previous findings showing that a single phage can only “knock-down” rather than “knock-out” target bacteria14. Consequently, expanding the gut phage collection is crucial to design more effective phage cocktails. Furthermore, in the animal experiments, incomplete bacterial colonization of D. longicatena was noted, likely due to the single bacterial inoculation. This suggests that phage therapy studies in the gut may require continuous bacterial inoculation to ensure stable colonization before phage intervention, allowing for more accurate evaluation of phage efficacy. Extending the duration of animal experiments would also facilitate monitoring phage resistance development, microbial community shifts, and potential immune system responses. While we focused on phages with strong antimicrobial activity for functional validation, phages with lower inhibitory efficiency may also be valuable. These phages could contribute to understanding phage-bacteria interactions, be incorporated into phage cocktails, or serve as derivatives in functional applications. Before applying phage therapy in human, it is essential to determine whether the observed phage differences are directly related to health status or indirectly influenced through their host bacteria. Further studies are needed to explore the stimulation of human immunity by phages and the localized inflammation caused by bacterial lysis57,58.
This research advances methods for efficiently isolating gut phages and facilitates the discovery of previously uncharacterized phages. To expand the gut phage biobank, it is crucial to isolated phages under diverse culture conditions and convert complex isolation processes into high-throughput, automated procedures. As the gut phage collection accumulates and more data on genomes and phage-bacteria interactions are integrated, our understanding of phage-mediated impacts on microbial diversity, ecosystem dynamics, evolution, and human health will significantly improve. Before using phages to modulate the gut microbiome for disease intervention, large-scale cohort studies and comprehensive analysis are needed to define and validate the causal relationships between the gut microbiome and human health. This relationship can then be applied clinically by combining metagenomic analysis of the gut microbiome with patient phenotypes to assess health status and risks. Based on the degree of gut dysbiosis, tailored intervention strategies can be proposed: for mild dysbiosis, single intervention such as dietary changes, probiotic, phage therapy, and/or FMT may suffice; for severe dysbiosis, a combinatory approach incorporating artificial microbial communities (gut commensal bacteria/ probiotics, specific phages, and/or phage derivatives) may be more effective. Ultimately, the goal is to establish comprehensive solutions that encompass diagnosis, precision treatment, and health monitoring and management.
Methods
Ethics statement
This study was conducted in accordance with the principles of the Declaration of Helsinki and was approved by the BGI Institutional Review Board on Bioethics and Biosafety (Approval No. BGI-IRB 17084-T4). All participants provided written informed consent after receiving a detailed explanation of the research procedures. Fecal samples were provided voluntarily and without compensation.
The animal experiments were conducted following guidelines from the Association for Assessment and Accreditation of Laboratory Animal Care, International (AAALAC international) and approved by the Institutional Animal Care and Use Committee (IACUC) of Gempharmatech Co., Ltd (AP#: GPTAP20230821-2).
Bacterial strains and phage resource
Supplementary Data 1 lists all the strains used for phage isolation in this study. For phage fecal resource collecting, 1.5-2.0 g fecal and saliva samples were picked into 20 mL SM buffer, vortex for 5 min, and stood at 4 °C overnight. For the collection of other phage resources, such as wastewater, sea water, and lake water, approximately 5 L of each liquid sample was collected in bottles and stored at a temperature of 4 °C temporarily for subsequent processing. To process the phage resources samples, we centrifuged the samples at 5000 × g for 15 minutes and sterilized the supernatant with a 0.45 μm filter to remove any endogenous bacteria. Store at 4 °C for further phage isolation.
Bacterial culture conditions
Bacteria were all cultured in BHI-YH medium (BHI medium supplemented with 5 g/L yeast extract, 5 mg/L hemin). The anaerobic bacteria were cultured in an anaerobic chamber (filled with 5% H2, 5% CO2, and 90% N2) at 37 °C. The BHI-YH for anaerobic bacteria was supplemented with 0.5 g/L L-cysteine hydrochloride and 0.25 g/L anhydrous sodium sulfide.
To culture facultative anaerobic bacteria, 10 μL of glycerol bacterial solution was inoculated onto BHI-YH solid plates (1.5 g/L agar added to BHI-YH) and incubated for 1 day. For anaerobic bacteria, inoculate 10 μL of glycerol solution onto reduced BHI-YH solid plates in an anaerobic chamber (MAWORDE) and incubate anaerobically for 1-2 days. Transferred single colony into 1 mL BHI-YH liquid broth and incubated overnight to provide starting bacterial cultures for the experiments.
Phage isolation
For the source samples without existing metagenomic data, we employed both the classical method and the repetitive enrichment method to isolate phages for all bacterial strains. For source samples that already had metagenomic data available, we specifically selected samples predicted to contain viral contigs linking to our candidate anaerobic bacteria for subsequent isolation.
For the classical enrichment method, we mixed 40 mL filtered samples with 10 mL quintuple-strength BHI-YH broth supplemented with 1 mM CaCl2 and MgCl2. We then inoculated the mixture with late log-phase candidate bacterial strains at a 1:100 ratio and cultured it at 37 °C overnight. After co-culturing, we centrifuged the culture at 5000 × g for 15 minutes and sterilized the supernatant using a 0.45 μm filter. Next, we added a precipitant solution to the filtered culture to obtain a concentrated phage solution. The precipitant solution consisted of 1 M NaCl and 10% (w/v) PEG-8000, and we allowed the mixture to settle at 4 °C overnight. The settled mixture was then centrifuged at 4 °C, 12,000 × g for 20 minutes, and the supernatant was discarded. Finally, we resuspended the pellet in 1 mL SM buffer to obtain the concentrated phage solution.
For the repetitive enrichment method, we took 1 mL filtered samples and added them to 9 mL broth supplemented with 1 mM CaCl2 and MgCl2. We then inoculated the mixture with late log-phase candidate bacterial host strains at a 1:100 ratio and incubated it at 37 °C overnight. After co-culturing, we centrifuged the culture at 5000 × g for 15 minutes and sterilized the supernatant using a 0.45 μm filter. We then took 1 mL filtered culture from the previous step and added it to 9 mL fresh broth. We inoculated this mixture with late log-phase single strains of bacteria at a 1:100 ratio and repeated the incubation and harvest steps three times. After the triple enrichments, we mixed all the remaining enriched cultures and added a precipitant solution to obtain a concentrated phage solution, as described above.
For the metagenomics-guided method, we initially select samples with available metagenomic data to predict candidate viral sequences that link to our candidate anaerobic bacteria host (see Fecal metagenomic analysis in method) for the subsequent repetitive enrichment method.
To identify isolated phages, we performed spot assays using the concentrated phage solution and picked the plaques that appeared. We then purified these plaques and stored the isolated phages at 4 °C.
Fecal metagenomic analysis
Raw data were cleaned by filtering out poor-quality reads using fastp59. Reads that mapped to the human genome were removed using BWA60 and SAMtools61. The taxonomy classification was performed using Kraken62. Clean reads were assembled using SPAdes with the ‘-meta’ option. Viral sequences were identified using DeepVirFinder63 (score ≥ 0.9 and P < 0.05) and VirSorter264 (max_score ≥ 0.8). We used a CRISPR spacer-based approach to assign host for viral sequences. CRISPR spacers were predicted from gut bacterial genomes using MinCED65 with the ‘-minNR 2’ option. Matches between the CRISPR spacers and identified viral sequences were identified using BLASTn66.
To select samples for phage isolation, we established the following criteria based on metagenomic sequencing (Figure S1): a minimum of 0.1% of total reads classified as viral reads according to Kraken; identification of more than 500 candidate viral sequences using DeepVirFinder and VirSorter2; predicted viral sequences were linked to the bacteria utilized in our study, evidenced by a complete alignment with five or more CRISPR spacers within one bacterial genome. The selected samples should meet both criteria 1 and criteria 2 or satisfy criterion 3.
Phage purification
To purify the isolated phages, we dropped 10 μL phage solution (SM buffer with a plaque) onto a double-layer plate with host bacteria, streaked to serial dilute with sterilized paper strips 3-4 times, and incubated at 37 °C overnight. When a single plaque appeared on the bacterial lawn, we picked the single plaque (chipped out the plaque) into 1 mL SM buffer and repeated the purification for 3−5 rounds to obtain a pure phage solution.
Phage enrichment and titration
To enrich purified phages, we performed spot assays by incubating 10−20 μL of pure phage solution overnight to form a large plaque. We then added the plaque to a 50 mL culture of early log-phase host bacteria and incubated it overnight. After centrifugation at 5000 × g for 15 minutes and filtration with a 0.45 μm filter, we obtained the phage lysate, which was stored at 4 °C. To determine the phage concentration, we diluted the lysate 1:10 with SM buffer and conducted spot assays.
DNA extraction and sequencing
DNase I and RNase A were added to 50 mL of phage lysate as the phage enrichment step described, at a final concentration of 10 μg/mL each, and the mixture was incubated at 37 °C for 1 hour. Another round of DNase I and RNase A were added, and the mixture was incubated at 37 °C for 30 minutes. The nucleases were then inactivated in a 65 °C water bath for 20 minutes.
Next, a precipitant solution was added to the lysate, resulting in a final concentration of 1 M NaCl and 10% (w/v) PEG-8000. The mixture was thoroughly mixed and left to settle at 4 °C overnight. Subsequently, the mixture was centrifuged at 4 °C, 12,000 × g for 20 minutes. The supernatant was discarded, and the pellet was resuspended with 200 μL of SM buffer. Then, 0.5% SDS and 5 μL of proteinase K were added to the resuspension, and the mixture was incubated at 56 °C for 1 hour and after gently mixing. Another round of proteinase K was added, and the mixture was incubated for an additional hour. The proteinase K was then inactivated at 65 °C for 20 minutes.
The phage DNA was extracted using the GeneClean® Turbo Kit (MP Biomedicals, #111102400), following the protocol provided by the kit. Subsequently, the single-end library was prepared using the MGlEasy Universal DNA Library Prep Kit and then sequenced on the DNBSEQ-E5 platform (MGI, BGI-Shenzhen, China).
Phage genome assembly and annotation
The sequencing depth of each phage was about 1000×. Raw data were filtered with fastp59 to remove poor-quality reads. Clean reads were then assembled using SPAdes67. The completeness of sequence assembly was evaluated using CheckV68.
For functional gene annotation, gene prediction was performed using Prodigal69. BLASTp66 searches against the NCBI nr database70,71 and hmmscan72 searches against the UniProt/Swiss-Prot database73 were conducted. The protein structure-based prediction tool Phold (https://github.com/gbouras13/phold) was also applied. Final annotations were obtained through manual integration, prioritizing results from BLASTp first, followed by HMMscan, and lastly Phold. Lysogenic genes were examined for integrases, Cro/CI repressor proteins, immunity repressors, DNA partitioning protein A (ParA), and anti-repressor proteins. Auxiliary metabolic genes (AMGs) and anti-resistant genes were analyzed using VIBRANT74 and DefenseFinder70,71. Virulence factors and drug resistance genes were identified by comparing against databases such as the Virulence Factors Database (VFDB, http://www.mgc.ac.cn/VFs/) and the Comprehensive Antibiotic Resistance Database (CARD, https://card.mcmaster.ca/). Mobile genetic elements (MGEs) were identified by comparing them against existing MGEs sequences75.
Gene-sharing network construction
We compared viral sequences from four human gut virome databases - the Gut Phage Database (GPD), human Gut Virome Database (GVD), the Metagenomic Gut Virus (MGV) catalog, and the Cenote Human Virome Database (CHVD)18,19,20,21 - against GPB phage genomes using BLASTp66, selecting aligned sequences to form a subcollection (n = 13,476). Following this, Prodigal69 was utilized to predict CDSs in gut phage genomes (n = 102), NCBI RefSeq phage genomes (ProkaryoticViralRefSeq211-Merged, n = 4534), and the sequence subcollection. These protein sequences served as input for vConTACT276, which assesses homologous protein sequence identity across phage genomes, generating probabilistic viral clusters (VCs) indicative of high genomic similarity. The obtained VCs were equivalent to a genus level in accordance with the International Committee on Taxonomy of Viruses (ICTV)77. Phages were singletons or only clustered with our phages and were considered unclassifiable. The gene-sharing network was visualized using Cytoscape78.
Transmission electron microscopy
First, we carefully injected 10 mL of a 20% sucrose solution beneath the lysate. Next, we proceeded to centrifuge the mixture at 59,057 × g for 3 hours at 4 °C. This process resulted in the formation of pellets, which were then resuspended in 1 mL of SM buffer. Subsequently, we performed dialysis using a 100 kD MWCO dialysis in 1 L of SM buffer for 4 hours. We replaced the buffer with another 1 L of fresh SM buffer and continued dialysis overnight. Finally, we sent the purified samples to Beijing GeoAnalysis Technology Co., Ltd. for TEM picture-taking.
Phylogenetic analysis
For phylogenetic analysis, we aligned the sequences of phage large terminase subunits from the four GPB crAss-like phages, 28 reported cultured crAss-like strains, and 72 reference genomes from ICTV using mafft79. Phylogenetic trees were constructed via the maximum-likelihood method using IQ-TREE80 and further visualized by iTOL81. All Maximum Likelihood (ML) phylogenies were constructed with 1,000 ultrafast bootstrap pseudo-replications.
Prevalence and abundance analysis
To estimate the prevalence and relative abundance of each gut phage, we mapped reads from 1,690 human gut metagenomic samples to the genomes of GPB phages and crAss-like phages using Bowtie282. SAMtools61 was employed to filter the mapped reads and calculate the coverage of each phage genome. Prevalence and abundance analysis draws upon previous literature with a subtle refinement18. Specifically, a phage was considered present in a sample if the mapped reads covered ≥60% of the genome length. The prevalence of a phage was calculated by dividing the number of samples containing that phage by the total number of samples. The relative abundances were determined by calculating the ratio of mapped reads to the total reads count for each sample. The source of these 1690 human gut metagenomic samples used in our study is listed in Supplementary Data 5.
Host range assay
We added 1 mL of log-phase bacterial culture to 14 mL of top agar containing 5 mM CaCl2 and MgCl2. This mixture was then poured onto a plate and allowed to dry completely, which typically took around 20 minutes. Next, we carefully dripped 1 μL of a 106 PFU/mL phage solution onto the plate in triplicate. The plate was subsequently cultured at 37 °C for 24 hours.
Sequence comparison and structural prediction
For sequence comparison, CDSs within the genomes of gut bacteria strains and phage isolates were predicted by Prodigal69. Pairwise genomic sequence comparisons were conducted using BLASTn66, while protein sequence comparisons were performed with BLASTp. Multiple sequence alignments were performed and visualized using ESPript 3.0 (https://espript.ibcp.fr/ESPript/cgi-bin/ESPript.cgi). The average nucleotide identity (ANI) between the genome sequences was calculated using fastANI83.
For structural prediction, Alphafold284,85 algorithm was employed to identify amino acid sequences that bear resemblance to the input sequence through sequence alignment, extract relevant sequence features, train a neural network model, and subsequently generate the corresponding three-dimensional structure of the protein. Subsequently, protein superposition was conducted using Pymol86.
Killing curve
To assess the killing curves, the bacteria at log-phase and combined with phages at MOI 1 and 10, then we diluted to OD600nm 0.1 for the following killing curve observation. Subsequently, 200 μL of this mixture was added to each well of a 96-well plate and incubated at 37 °C with orbital shaking. Measurements of OD600nm were taken every 10 minutes to monitor bacterial growth. As a control, bacterial hosts were mixed with BHI-YH broth to serve as the free-phage control, while only BHI-YH liquid acted as the blank control. Each group was repeated three times.
Mouse Models
Six-week-old SPF WT C57BL/6 male mice (GemPharmatech) were housed in Tecniplast isocages (temperature 20-26 °C, relative humidity 40−70%) under a 12 h light/12 h dark cycle (illuminated, 08:00-20:00), fed with sterilized corn cobs, and a week was given to acclimate in the animal facility. The animal experiments were conducted following guidelines from the Association for Assessment and Accreditation of Laboratory Animal Care, International (AAALAC International)
Murine phage therapy and bacteria quantity
The mouse experiment was performed according to previous literatures52 with slight modifications. Six-week-old SPF WT C57BL/6 male mice (GemPharmatech) were housed in Tecniplast isocages (temperature 20−26°C, relative humidity 40−70%) under a 12 h light/12 h dark cycle (illuminated, 08:00-20:00), fed with sterilized corn cobs, and a week was given to acclimate in the animal facility. They were then administered a daily dose of 60 mg ampicillin, 60 mg vancomycin, 60 mg neomycin, and 48 mg metronidazole via gavage for seven days. After a two-day washout with sterile water, 10 mice were given 200 μL saline suspension of D. longicatena RTGS0005 (108 CFU/mL) after 5 minutes of oral gavage of 100 μL 3% NaHCO3 on Day 0.
On Day 0, the mice given D. longicatena RTGS0005 were separated into phage-treated groups and phage-free groups (n = 5 for each group). Then the phage-treated groups accepted 200 μL of 109 PFU/mL phages CPB1092 from Day 1 to Day 4. The phage-free control group received saline instead of phages. A blank group (n = 5) accepted neither bacteria nor phage but saline instead.
Stools were collected 12 hours after each gavage. For CFU counting, the fecal samples for D. longicatena counting were cultured on BHI-YH plates. To calculate the normalized CFU was calculated by dividing the average stool concentrations of CFU in the phage-treated or phage-free groups by the CFU concentrations of the blank group. For relative abundance calculation, the clean reads from the metagenomic sequencing of stools were mapped to the genome of bacteria or phages using Bowtie282. The relative abundance was calculated by dividing the number of mapped reads by the total number of reads.
The experiment aimed to determine whether phages inhibit host bacteria in vivo, rather than evaluating the effect of mouse sex on phage antibacterial activity. Thus, mouse sex was excluded from the study design and statistical analyses. Consequently, mouse sex was not considered as a variable in the study design, no sex-disaggregated data were collected during experimentation, and it was excluded from the statistical analyses.
Statistics & Reproducibility
Statistical analysis was performed using R (3.6.3), OriginPro 2024 (Learning Edition). To calculate the normalized CFU during the murine phage therapy experiments, we divided the average stool concentrations of CFU into the phage-treated or phage-free groups by the CFU concentrations of the blank group. The two-sided Mann-Whitney test was used to assess the statistical significance of the relative abundance and prevalence of gut phages in human cohorts, CFU concentration, and the abundance of bacteria and phages in murine experiments.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The genome sequencing data reported in this study have been deposited in CNGB Sequence Archive (CNSA)39 of China National GeneBank DataBase (CNGBdb)40 under the accession number CNP0004979. All the phages isolated in this study have been deposited in CNGBdb-EBB with accession number CNSebb2036927 - CNSebb2037028. All these phage isolates can be requested through CNGBdb-EBB or by direct correspondence with the lead contact, Dr. Minfeng Xiao (xiaominfeng@genomics.cn). These data and resources are strictly limited to academic research use, and this publication must be cited accordingly.
The metagenomic samples analyzed in this study are publicly available in NCBl Sequence Read Archive (SRA) or the European Nucleotide Archive (ENA), with all accession codes provided in Supplementary Data 5. Source data are provided with this paper.
Code availability
The present study did not generate code, and the mentioned tools used for the data analysis were applied with default parameters unless specified otherwise in the Methods section.
References
Forslund, K. et al. Disentangling type 2 diabetes and metformin treatment signatures in the human gut microbiota. Nature 528, 262–266 (2015).
Harsch, I. A. & Konturek, P. C. The role of gut microbiota in obesity and type 2 and type 1 diabetes mellitus: new insights into “old” diseases. Med. Sci. (Basel) 6, 32 (2018).
Britton, G. J. et al. Microbiotas from humans with inflammatory bowel disease alter the balance of gut Th17 and RORgammat(+) regulatory T cells and exacerbate colitis in mice. Immunity 50, 212–224.e214 (2019).
Liang, J. Q. et al. A novel faecal Lachnoclostridium marker for the non-invasive diagnosis of colorectal adenoma and cancer. Gut 69, 1248–1257 (2020).
Needham, B. D., Kaddurah-Daouk, R. & Mazmanian, S. K. Gut microbial molecules in behavioural and neurodegenerative conditions. Nat. Rev. Neurosci. 21, 717–731 (2020).
Oniszczuk, A., Oniszczuk, T., Gancarz, M. & Szymanska, J. Role of gut microbiota, probiotics and prebiotics in the cardiovascular diseases. Molecules 26, 1172 (2021).
Markowiak, P. & Śliżewska, K. Effects of probiotics, prebiotics, and synbiotics on human health. Nutrients 9, 1021 (2017).
Antushevich, H. Fecal microbiota transplantation in disease therapy. Clin. Chim. Acta 503, 90–98 (2020).
Wang, J.-W. et al. Fecal microbiota transplantation: review and update. J. Formos. Med. Assoc. 118, S23–S31 (2019).
Pirnay, J. A.-O. et al. Personalized bacteriophage therapy outcomes for 100 consecutive cases: a multicentre, multinational, retrospective observational study. Nat. Microbiol. 9, 1434–1453 (2024).
Duan, Y., Young, R. & Schnabl, B. A. O. X. Bacteriophages and their potential for treatment of gastrointestinal diseases. Alimentary Pharmacol. Therap. 51, 53–63 (2022).
Khan Mirzaei, M. & Deng, L. New technologies for developing phage-based tools to manipulate the human microbiome. Trend Microbiol. 30, 131−142 (2021).
Galtier, M. et al. Bacteriophages targeting adherent invasive escherichia coli strains as a promising new treatment for crohn’s disease. J. Crohns Colitis 11, 840–847 (2017).
Federici, S. et al. Targeted suppression of human IBD-associated gut microbiota commensals by phage consortia for treatment of intestinal inflammation. Cell 185, 2879–2898.e2824 (2022).
Duan, Y. et al. Bacteriophage targeting of gut bacterium attenuates alcoholic liver disease. Nature 575, 505–511 (2019).
Zheng, D. W. et al. Phage-guided modulation of the gut microbiota of mouse models of colorectal cancer augments their responses to chemotherapy. Nat. Biomed. Eng. 3, 717–728 (2019).
Kabwe, M., Dashper, S., Bachrach, G. & Tucci, J. Bacteriophage manipulation of the microbiome associated with tumour microenvironments-can this improve cancer therapeutic response? FEMS Microbiol. Rev. 45, fuab017 (2021).
Camarillo-Guerrero, L. F., Almeida, A., Rangel-Pineros, G., Finn, R. D. & Lawley, T. D. Massive expansion of human gut bacteriophage diversity. Cell 184, 1098–1109.e1099 (2021).
Nayfach, S. et al. Metagenomic compendium of 189,680 DNA viruses from the human gut microbiome. Nat. Microbiol. 6, 960–970 (2021).
Tisza, M. J. & Buck, C. B. A catalog of tens of thousands of viruses from human metagenomes reveals hidden associations with chronic diseases. Proc. Natl. Acad. Sci. USA 118, e2023202118 (2021).
Gregory, A. C. et al. The gut virome database reveals age-dependent patterns of virome diversity in the human gut. Cell Host Microbe 28, 724–740.e728 (2020).
Ramos-Barbero, M. D. et al. Characterization of crAss-like phage isolates highlights Crassvirales genetic heterogeneity and worldwide distribution. Nat. Commun. 14, 4295 (2023).
Guerin, E. et al. Isolation and characterisation of phi crAss002, a crAss-like phage from the human gut that infects Bacteroides xylanisolvens. Microbiome 9, 89 (2021).
Shkoporov, A. N. et al. PhiCrAss001 represents the most abundant bacteriophage family in the human gut and infects Bacteroides intestinalis. Nat. Commun. 9, 4781 (2018).
Shen, J. et al. Large-scale phage cultivation for commensal human gut bacteria. Cell Host Microbe 31, 665–677.e667 (2023).
Hryckowian, A. J. et al. Bacteroides thetaiotaomicron-infecting bacteriophage isolates inform sequence-based host range predictions. Cell Host Microbe. 28, 371–379.e375 (2020).
Bonilla-Rosso, G., Steiner, T., Wichmann, F., Bexkens, E. & Engel, P. Honey bees harbor a diverse gut virome engaging in nested strain-level interactions with the microbiota. Proc. Natl. Acad. Sci. USA 117, 7355–7362 (2020).
Papudeshi, B. et al. Host interactions of novel Crassvirales species belonging to multiple families infecting bacterial host, Bacteroides cellulosilyticus WH2. Microb. Genomics 9, 001100 (2022).
Shkoporov, A. N. et al. Long-term persistence of crAss-like phage crAss001 is associated with phase variation in Bacteroides intestinalis. BMC Biol. 19, 163 (2021).
Guerin, E. et al. Isolation and characterisation of PhicrAss002, a crAss-like phage from the human gut that infects Bacteroides xylanisolvens. Microbiome 9, 89 (2021).
Chevallereau, A., Pons, B. J., van Houte, S. & Westra, E. R. Interactions between bacterial and phage communities in natural environments. Nat. Rev. Microbiol. 20, 49–62 (2022).
Shkoporov, A. N., Turkington, C. J. & Hill, C. Mutualistic interplay between bacteriophages and bacteria in the human gut. Nat. Rev. Microbiol. 20, 737–749 (2022).
Porter, N. T. et al. Phase-variable capsular polysaccharides and lipoproteins modify bacteriophage susceptibility in Bacteroides thetaiotaomicron. Nat. Microbiol. 5, 1170–1181 (2020).
Yeoh, Y. K. et al. Prevotella species in the human gut is primarily comprised of Prevotella copri, Prevotella stercorea and related lineages. Sci. Rep. 90, 9055 (2022).
Yang, J. et al. Species-level analysis of human gut microbiota with metataxonomics. Front. Microbiol. 11, 2029 (2020).
Arumugam, M. et al. Enterotypes of the human gut microbiome. Nature 473, 174–180 (2011).
Underwood, M. A., German, J. B., Lebrilla, C. B. & Mills, D. A. Bifidobacterium longum subspecies infantis: champion colonizer of the infant gut. Pediatr. Res. 77, 229–235 (2015).
Xu, R. et al. Altered gut microbiota and mucosal immunity in patients with schizophrenia. Brain, Behav., Immun. 85, 120–127 (2020).
Guo, X. et al. CNSA: a data repository for archiving omics data. Database 2020, baaa055 (2020).
Chen, F. et al. CNGBdb: China National genebank database. Hereditas (Beijing 42, 799–809 (2020).
Chen, L. A.-O. et al. Large freshwater phages with the potential to augment aerobic methane oxidation. Nat. Microbiol. 5, 1504–1515 (2020).
Yi, Y. et al. A systematic analysis of marine lysogens and proviruses. Nat. Commun. 14, 6013 (2023).
Watson, B. N. J. et al. Different genetic and morphological outcomes for phages targeted by single or multiple CRISPR-Cas spacers. Philos. Trans. R. Soc. Lond. Ser. B, Biol. Sci. 374, 20180090 (2019).
Shkoporov, A. A.-O. et al. CrAss001 represents the most abundant bacteriophage family in the human gut and infects Bacteroides intestinalis. ss001 Represents Most Abund. Bacteriophage Fam. Hum. Gut. Infects. Bacter. 9, 4781 (2018).
Jiang, X. F. et al. Invertible promoters mediate bacterial phase variation, antibiotic resistance, and host adaptation in the gut. Science 363, 181–187 (2019).
Warman, E. A. et al. Widespread divergent transcription from bacterial and archaeal promoters is a consequence of DNA-sequence symmetry. Nat. Microbiol. 6, 746–756 (2021).
Siranosian, B. A., Tamburini, F. B., Sherlock, G. & Bhatt, A. S. Acquisition, transmission and strain diversity of human gut-colonizing crAss-like phages. Nat. Commun. 11, 280 (2020).
Palmas, V. et al. Gut microbiota markers associated with obesity and overweight in Italian adults. Sci. Rep. 11, 5532 (2021).
Gravdal, K. et al. Exploring the gut microbiota in patients with pre-diabetes and treatment naïve diabetes type 2 - a pilot study. BMC Endocr. Disord. 23, 179 (2023).
Companys, J. A.-O. et al. Gut microbiota profile and its association with clinical variables and dietary intake in overweight/obese and lean subjects: a cross-sectional study. Nutrients 13, 2032 (2021).
Ruuskanen, M. A.-O. et al. Gut microbiome composition is predictive of incident type 2 diabetes in a population cohort of 5,572 finnish adults. Diabetes Care 45, 811–818 (2022).
Wang, Y. et al. Phage cocktail targeting STEC O157:H7 has comparable efficacy and superior recovery compared with enrofloxacin in an enteric murine model. Microbiol. Spectr. 10, e0023222 (2022).
Schmidtke, D. T., Hickey, A. S., Liachko, I., Sherlock, G., & Bhatt, A. S. Analysis and culturing of the prototypic crAssphage reveals a phage-plasmid lifestyle. bioRxiv 20, 2024.03.20.585998 (2024).
Silpe, J. E. & Bassler, B. L. A host-produced quorum-sensing autoinducer controls a phage lysis-lysogeny decision. Cell 176, 268 (2019).
Shkoporov, A. N. et al. The human gut virome is highly diverse, stable, and individual specific. Cell Host Microbe. 26, 527–541.e525 (2019).
Rotman, E. et al. Rapid design of bacteriophage cocktails to suppress the burden and virulence of gut-resident carbapenem-resistant Klebsiella pneumoniae. Cell Host Microbe. 32, 1988–2003.e1988 (2024).
Gogokhia, L. et al. Expansion of bacteriophages is linked to aggravated intestinal inflammation and colitis. Cell Host microbe 25, 285–299.e288 (2019).
Norman, J. M. et al. Disease-specific alterations in the enteric virome in inflammatory bowel disease. Cell 160, 447–460 (2015).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Wood, D. E., Lu, J. & Langmead, B. Improved metagenomic analysis with Kraken 2. Genome Biol. 20, 257 (2019).
Ren, J. et al. Identifying viruses from metagenomic data using deep learning. Quant. Biol. 8, 64–77 (2020).
Guo, J. et al. VirSorter2: a multi-classifier, expert-guided approach to detect diverse DNA and RNA viruses. Microbiome 9, 37 (2021).
Bland, C. et al. CRISPR recognition tool (CRT): a tool for automatic detection of clustered regularly interspaced palindromic repeats. BMC Bioinforma. 8, 209 (2007).
Mount, D. W. Using the basic local alignment search tool (BLAST). CSH Protoc. 2007, pdb top17 (2007).
Prjibelski, A., Antipov, D., Meleshko, D., Lapidus, A. & Korobeynikov, A. Using SPAdes de novo assembler. Curr. Protoc. Bioinforma. 70, e102 (2020).
Nayfach, S. et al. CheckV assesses the quality and completeness of metagenome-assembled viral genomes. Nat. Biotechnol. 39, 578–585 (2021).
Hyatt, D. et al. Prodigal: Prokaryotic gene recognition and translation initiation site identification. Bmc Bioinform. 11, 119 (2010).
Tesson, F. et al. Systematic and quantitative view of the antiviral arsenal of prokaryotes. Nat. Commun. 13, 2561 (2022).
Abby, S., Néron, B., Ménager, H., Touchon, M. & Rocha, E. MacSyFinder: A Program to mine genomes for molecular systems with an application to CRISPR-Cas systems. PloS one 9, e110726 (2014).
Johnson, L. S., Eddy, S. R. & Portugaly, E. Hidden Markov model speed heuristic and iterative HMM search procedure. Bmc Bioinform. 11, 431 (2010).
Suzek, B. E., Huang, H., McGarvey, P., Mazumder, R. & Wu, C. H. UniRef: comprehensive and non-redundant UniProt reference clusters. Bioinformatics 23, 1282–1288 (2007).
Kieft, K., Zhou, Z. & Anantharaman, K. VIBRANT: automated recovery, annotation and curation of microbial viruses, and evaluation of viral community function from genomic sequences. Microbiome 8, 90 (2020).
Pärnänen, K. et al. Maternal gut and breast milk microbiota affect infant gut antibiotic resistome and mobile genetic elements. Nat. Commun. 9, 3891 (2018).
Bin Jang, H. et al. Taxonomic assignment of uncultivated prokaryotic virus genomes is enabled by gene-sharing networks. Nat. Biotechnol. 37, 632–639 (2019).
Lefkowitz, E. J. et al. Virus taxonomy: the database of the international committee on Taxonomy of viruses (ICTV). Nucleic Acids Res. 46, D708–D717 (2018).
Otasek, D., Morris, J. H., Boucas, J., Pico, A. R. & Demchak, B. Cytoscape automation: empowering workflow-based network analysis. Genome Biol. 20, 185 (2019).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Nguyen, L. T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 49, W293–W296 (2021).
Langmead, B., Wilks, C., Antonescu, V. & Charles, R. Scaling read aligners to hundreds of threads on general-purpose processors. Bioinformatics 35, 421–432 (2019).
Jain, C., Rodriguez-R, L. M., Phillippy, A. M., Konstantinidis, K. T. & Aluru, S. High throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries. Nat. Commun. 9, 5114 (2018).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Varadi, M. A.-O. et al. AlphaFold protein structure database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic acids Res. 50, D439–D444 (2022).
Schrodinger, L. L. C. The PyMOL molecular graphics system, version 1.8 4. Am. J. Infect. Dis. Microbiol. 4, 61−71 (2015).
Henke, M. A.-O. et al. Ruminococcus gnavus, a member of the human gut microbiome associated with Crohn’s disease, produces an inflammatory polysaccharide. Proc. Natl. Acad. Sci. USA 116, 12672–12677 (2019).
Yoo do, Y. et al. Bacteroides fragilis enterotoxin upregulates lipocalin-2 expression in intestinal epithelial cells. Lab. Investig. A. J. Tech. Methods Pathol. 93, 384–396 (2013).
Maini Rekdal, V. A.-O., Bess, E. A.-O., Bisanz, J. A.-O., Turnbaugh, P. A.-O. & Balskus, E. A.-O. Discovery and inhibition of an interspecies gut bacterial pathway for Levodopa metabolism. Science 364, eaau6323 (2019).
Bamba, T. et al. The pathogenic role of Bacteroides vulgatus in patients with ulcerative colitis. J. Gastroenterol. 30, 45–47 (1995).
Viladomiu, M. et al. Adherent-invasive E. coli metabolism of propanediol in Crohn’s disease regulates phagocytes to drive intestinal inflammation. Cell Host Microbe 29, 607–619.e608 (2021).
Duport, C., Jobin, M. & Schmitt, P. Adaptation in Bacillus cereus: from stress to disease. Front. Microbiol. 7, 1550 (2016).
Jiao, L. F. et al. Effects of zinc-exchanged montmorillonite with different zinc loading capacities on growth performance, intestinal microbiota, morphology and permeability in weaned piglets. Appl. Clay Sci. 112, 40–43 (2015).
Golińska, E. et al. Virulence factors of Enterococcus strains isolated from patients with inflammatory bowel disease. World J. Gastroenterol. 19, 3562–3572 (2013).
Yuan, J. et al. Fatty liver disease caused by high-alcohol-producing Klebsiella pneumoniae. Cell Metab. 30, 675–688.e677 (2019).
Banaszkiewicz, A. et al. Enterotoxigenic Clostridium perfringens infection and pediatric patients with inflammatory bowel disease. J. Crohn’s colitis 8, 276–281 (2014).
Mendes-Frias, A. et al. Akkermansia muciniphila and Parabacteroides distasonis as prognostic markers for relapse in ulcerative colitis patients. Front. Cell. Infect. Microbiol. 14, 1367998 (2024).
Curtis, M. M. et al. The gut commensal Bacteroides thetaiotaomicron exacerbates enteric infection through modification of the metabolic landscape. Cell Host Microbe. 16, 759–769 (2014).
Almeida, A. et al. A unified catalog of 204,938 reference genomes from the human gut microbiome. Nat. Biotechnol. 39, 105–114 (2021).
Acknowledgements
This work is supported by the National Natural Science Foundation of China (NSFC) (82250901 to R.L. and C.L., 82050007 to R.L. and C.L.). Genome sequencing is supported by the Global Phage Hub project (GPH) initiated by BGI-Shenzhen. Bacterial strains are provided by China National GeneBank (CNGB). We would like to thank Professor Jingmin Gu for helping to isolate several E. faecalis phages.
Author information
Authors and Affiliations
Contributions
M.X. conceived the study. B.X., C.L., and W.C. designed, conducted the experiments, and drafted the manuscript. B.X. and Z.L. performed bioinformatics and computational analyses. X.J. provided bacterial resources. C.W. and R.L. supported the design of animal experiments. X.X. and X.Z. provided the equipment and materials. J.L. and M.X. offered critical insights and revised the manuscript. All authors reviewed the manuscript before submission.
Corresponding authors
Ethics declarations
Competing interests
Authors affiliated with BGI Research (C.L., B.X., M.X., J.L and Z.L.) have filed a patent application titled “Isolation and Characterization of Bacteriophages Targeting Dorea longicatena and their Potential Applications” (Application No. 202510946726.2). It describes phage isolated in this study that target an obesity-associated bacteria, including genomic characterization, safety screening, and therapeutic cocktail development validated in in vitro and murine models. Other authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Corinne Maurice, and the other, anonymous, reviewer for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xing, B., Liu, C., Chen, W. et al. Gut Phage Biobank: a collection of bacteriophages targeting human commensal bacteria. Nat Commun 16, 11050 (2025). https://doi.org/10.1038/s41467-025-61946-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-61946-0