Abstract
Pyrroloquinoline quinone (PQQ)-dependent methanol dehydrogenases (MDHs), the periplasmic metalloenzymes in Gram-negative methylotrophic bacteria, play a pivotal role in methane and methanol bio-utilization. Although the structures of many PQQ-dependent MDHs have been resolved, including the canonical heterotetrameric enzymes composed of two MxaF and two MxaI subunits with a molecule of PQQ and a calcium ion in the active site in MxaF, the biogenesis of these enzymes remains elusive. Here, we characterize a chaperone, MxaJ, responsible for PQQ incorporation by reconstructing a PQQ-dependent MDH assembly system in Escherichia coli. Using cryo-electron microscopy, we capture the structures of the intermediate complexes formed by the chaperone MxaJ and catalytic subunit MxaF during PQQ-dependent MDH maturation, revealing a chaperone-mediated molecular mechanism of cofactor incorporation. These findings not only advance our understanding on the biogenesis of PQQ-dependent MDH, but also provide an alternative engineering way for methane and methanol bioconversion.
Similar content being viewed by others
Introduction
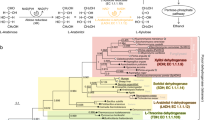
Annually, more than 40 million tons of methane, a potent greenhouse gas with 86 times stronger global warming potential (GWP) than CO2, are sunk by methanotrophic bacteria (methanotrophs), consuming methane as sole carbon source1,2. In methanotrophic metabolism, methane is initially oxidized by methane monooxygenases (MMOs) to methanol, which is further oxidized to formaldehyde and enters into the central metabolism mainly through the ribulose monophosphate pathway (RuMP) or the serine cycle (Fig. 1a)1,3,4. Notably, all methanotrophs utilize pyrroloquinoline quinone (PQQ)-dependent methanol dehydrogenases (MDHs) to oxidize methanol (Supplementary Fig. 1), enabling the electrons released during methanol oxidation to be transferred from reduced PQQ to the respiratory chain to drive ATP generation (Fig. 1a)5,6,7. This canonical methanol oxidation system represents a universal metabolic module in Gram-negative bacteria that possess methanol assimilation systems.
a Metabolic network for methane and methanol utilization in Gram-negative methylotrophs. Carbon metabolism pathways are indicated by black arrows, while the energy generation processes based on PQQ-dependent MDH are indicated by the red arrows. b The structural unit of a PQQ-dependent MDH from Methylobacterium extorquens (PDB ID: 1H4I) and its synthesis gene cluster (mxa). The cofactors, PQQ and Ca2+, in the enzyme are highlighted, and the genes in the cluster with known functions are shown in color. The two subunits of the enzyme are encoded by mxaF and mxaI, the four genes shown in yellow encode the proteins involved in Ca2+ incorporation, mxaG encodes the electron acceptor cytochrome cL, mxaB encodes the cluster-specific transcription factor. The genes with unknown functions are shown in white.
Two types of PQQ-dependent MDHs, MxaFI-type and XoxF-type, had been identified in nature8,9. XoxF-type MDHs are homodimeric proteins, employing lanthanide ion as cofactor assisting PQQ in catalysis8,10. In contrast, MxaFI-type MDHs are composed of two large (MxaF) and two small (MxaI) subunits. Each MxaF subunit houses one PQQ molecule and one calcium ion in its catalytic center and represents the catalytic component of the enzyme, while the MxaI subunits tightly wrap against the MxaF subunits, but the function is unclear (Fig. 1b)8,11,12. Numerous studies have shown that the synthesis of MxaFI-type MDH requires the involvement of a series of auxiliary proteins whose coding genes are typically clustered with the coding genes of the two functional subunits (Fig. 1b)13,14. Some of these proteins, including MxaA, MxaC, MxaK and MxaL, have been clarified to play roles in Ca²⁺ incorporation15,16,17. While others, such as MxaJ and MxaR, have only been linked to the activity of the enzyme, their specific functions remain unclear13,18. Nevertheless, the unusual crystal structure of MxaJ resembles some features found in periplasmic binding proteins, and it has been suggested that some aspects of its structure might provide a more favorable framework for interacting with proteins rather than small molecules19. However, given the close association between PQQ-dependent MDH availability and methylotrophic growth of the natural hosts, studying the auxiliary proteins using traditional reverse genetics has been challenging. As a result, identification of the auxiliary proteins involved in PQQ-dependent MDH maturation has yet to be accomplished, and the complete assembly process of these enzymes remains elusive.
In this study, we decouple the relationship between PQQ-dependent MDH biosynthesis and cellular growth by reconstructing the enzyme assembly system in Escherichia coli. This allows us to employ reverse genetics strategies to systematically identify the auxiliary proteins essential for the maturation of the enzyme. With the discovery of the key intermediates during PQQ-dependent MDH maturation, we resolved their structures using cryo-electron microscopy, thereby providing mechanistic insights into the biogenesis of the enzyme.
Results
Heterologous assembly of PQQ-dependent MDH
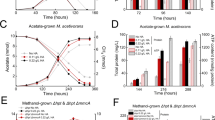
To comprehensively identify the auxiliary proteins involved in PQQ-dependent MDH maturation, we selected MxaFI-type MDH from the well-studied Methylobacterium extorquens AM1 as a model and reconstructed its assembly system in E. coli by heterologously expressing mxa cluster. Due to the unique arrangement of the natural operon, such as overlapping genes20,21, it was necessary to reorganize the expression operons in E. coli. Thus, 13 coding sequences from the mxa cluster, with the exception of the cluster-specific transcription factor coding gene mxaB22, were synthesized with codon usage optimization and recombined into three lac operons on two plasmids with designed ribosome binding sites (Fig. 2a). The resulting constructs were then transformed into E. coli to generate the recombinant strain (E.coli-13). After being cultured and induced in the medium containing PQQ, the heterologously expressed MDH (MDHEc13) was purified by Ni-affinity chromatography using a C-terminal His6-tag on the MxaF subunit. Similar to the endogenously synthesized MDH (MDHAM1) in M. extorquens, two functional proteins corresponding to MxaF and MxaI subunits were present in MDHEc13 (Fig. 2b). The comparable spectra (Fig. 2c) and the methanol oxidation activity (Fig. 2d, the first column) confirmed that the complete MDH was successfully assembled in E. coli-13, demonstrating that all auxiliary proteins required for the maturation of the enzyme were functionally expressed.
a Operon organization of the two plasmids expressing 13 genes of mxa cluster from M.extorquens AM1. b SDS-PAGE analysis of the purified MDHs assembled in M.extorquens AM1 and E. coli-13 (the strain containing 13 genes of mxa cluster) (One representative figure is shown in 3 independent experiments). c UV−vis spectra of the purified MDHs assembled in M.extorquens AM1 and E. coli-13. The absorbance at 345 nm reflects the semireduced state of PQQ and the enzyme is active. d Specific activity of the purified MDHs from E. coli-13 and its single gene deletion mutants. e Specific activity of the purified MDHs from E. coli−13 and its single-gene deletion mutants following 10 mM CaCl2 treatment in vitro at pH 9.5. The error bars represent the standard deviation from the mean, originating from three independent activity analyses.
After establishing the recombinant E. coli strain capable of assembling complete MDH, the foreign genes, except for mxaF, mxaI, and mxaG (the coding gene of cytochrome cL, which only functions in electron transfer in vivo23), were deleted separately, and the MDHs synthesized by these gene-deletion strains were characterized. SDS-PAGE showed that MDHs synthesized in all gene-deletion strains still contained MxaF and MxaI subunits (Supplementary Fig. 2a), indicating the deleted genes are not related to subunit assembly. However, functional characterization revealed that MDHs produced in strains lacking mxaJ, mxaR, mxaS, mxaA, mxaC, mxaK, or mxaL were inactive, demonstrating the essential role of these seven genes in proper MDH assembly (Fig. 2d). Further analysis showed that the Ca2+ content of these enzymes was too low to produce the characteristic absorption of semireduced PQQ at 345 nm (Supplementary Fig. 2b and c)16,24, suggesting abnormal binding of cofactors.
After being incubated with Ca2+ in vitro at pH 9.5 (a condition that enable rescue Ca2+ incorporation25), both activity and the specific absorption at 345 nm were restored in the MDHs from strains lacking mxaR, mxaS, mxaA, mxaC, mxaK, or mxaL (Fig. 2e and Supplementary Fig. 2d). These results demonstrated that proteins MxaR and MxaS, along with the previously identified proteins MxaA, MxaC, MxaK, and MxaL15,16,17, collectively regulate Ca²⁺ incorporation into MDH. Unexpectedly, although in vitro Ca²⁺ treatment increased the Ca²⁺ content in MDH from the mxaJ-deletion strain to 73%, neither enzymatic activity nor the characteristic 345 nm absorption was recovered (Fig. 2e and Supplementary Fig. 2d), implying that the MDH produced by the mxaJ-deletion strain also lacks PQQ. Thus, protein MxaJ should be essential for PQQ incorporation. According to the above findings, by co-expressing only 9 essential genes (mxaF, mxaJ, mxaI, mxaR, mxaS, mxaA, mxaC, mxaK, and mxaL), we successfully assembled functional MDH in E. coli (Supplementary Fig. 3).
MxaJ: a chaperone for PQQ incorporation
Although we comprehensively identified the assembly system of a MxaFI-type MDH and successfully synthesized it in E. coli, the total protein of heterologously assembled enzymes consistently displayed lower activity compared to the enzyme extracted from the native host, even after being treated with excessive Ca²⁺ in vitro (Fig. 3a). Native-PAGE analysis revealed that, in addition to the complex of MxaF and MxaI (MxaF/MxaI), additional protein aggregates were also observed in MDHEc13 and MDHEc9 (MDH extracted from the E. coli strain containing only the 9 essential genes) (Fig. 3b), suggesting that the assembly efficiency of the complete MDH in the engineered strains is much lower than that in the native host. Mass spectrometry (MS) confirmed that the additional protein complex was mainly composed of MxaF and MxaJ (Supplementary Fig. 4 and Supplementary Table 1). Given the potential role of MxaJ in PQQ incorporation, we speculated that PQQ depletion is the primary cause of inactive complex accumulation, ultimately resulting in the reduced specific activity of heterologously assembled MDHs.
a Kinetic parameter of MDHs extracted from different strains. The preparations been treated with 10 mM Ca2+ in vitro at pH 9.5 were highlighted in bold and orange. Data are represented as mean ± SD. b Native-PAGE analysis of the aggregates in the purified MDH preparations from different strains (One representative figure is shown in 3 independent experiments). Protein MxaJ was set as a control. c Native-PAGE analysis of the aggregates in MDHs purified from E. coli strains containing 1 (E. coli-9) or 2 (E. coli-10) mxaI genes under the gradient (0, 10, 20 μM) PQQ supply. d Specific activities of the MDHs in c after Ca2+ treatment in vitro. e Native-PAGE analysis of the aggregates in different assembly reactions in vitro. f Specific activities of the mixtures in e after been treated with 10 mM CaCl2 in vitro at pH 9.5. The error bars represent the standard deviation from the mean, originating from four and three independent activity analyses, respectively, of the samples in c and e. P values were determined using a two-sided unpaired t-test.
To improve MDH assembly efficiency in E. coli, PQQ was supplied in the growth medium at different concentrations (ranging from 0 to 20 μM). However, increasing PQQ supply (from 10 μM to 20 μM) did not significantly enhance the specific activity of MDHEc9 (Fig. 3c, d, and Supplementary Fig. 5). Considering the functional MDH consists of MxaF and MxaI, we explored other potential reasons for the low assembly efficiency, such as limited MxaI or excess MxaJ. By co-expressing a second mxaI gene to boost MxaI production, the specific activity of the reconstructed MDHEc9 (MDHEc10) was significantly improved (Fig. 3c, d, and Supplementary Fig. 5). With sufficient Ca²⁺ (added in vitro), the specific activity of MDHEc10 reached approximately 1.2 μmol min⁻¹ mg⁻¹ (Fig. 3d), approaching the level of MDHAM1 in the native host. Surprisingly, in the absence of PQQ during growth, only the MxaF/MxaJ complex was observed, with no formation of the MxaF/MxaI complex or methanol oxidation activity (Fig. 3c, d and Supplementary Fig. 5). This indicates that PQQ incorporation is essential for assembling the MxaF/MxaI complex, while the MxaF/MxaJ complex appears to serve as a critical intermediate during MDH maturation.
Further in vitro studies demonstrated that both MxaJ and MxaI can bind to MxaF independently (Fig. 3e). However, the affinity of MxaF for MxaJ is approximately 10 times higher than for MxaI (Supplementary Fig. 6), causing MxaF to preferentially interact with MxaJ. Additionally, PQQ can only bind to the MxaF/MxaJ complex (with an equilibrium dissociation constant, KD, is about 77 nM), rather than to the MxaF/MxaI complex or MxaF alone (Supplementary Fig. 7). The incorporation of PQQ into the MxaF/MxaJ complex likely facilitates the interaction between MxaI and MxaF, accelerating the release of MxaJ from MxaF and forming the PQQ-loaded MxaF/MxaI complex (Fig. 3e, f, and Supplementary Fig. 8). Based on these findings, we propose that MxaJ functions as a chaperone, mediating the incorporation of PQQ into MxaF.
Mechanism of MxaJ-mediated PQQ incorporation
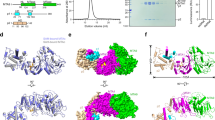
To elucidate the molecular mechanism of MxaJ-mediated PQQ incorporation in MDH, we resolved the structure of the MxaF/MxaJ complex using cryo-electron microscopy (cryo-EM) (Supplementary Fig. 9 and Table 2). The complex model comprises two MxaF/MxaJ units interacting over a large planar interface between two MxaF subunits (Fig. 4a), which is reminiscent of the interaction between the two MxaF subunits in complete MDH11. The loop region W418-Y439 (referred to as the roof sheet) of MxaF, which typically envelops the PQQ binding pocket of MxaF in complete MDH11, was captured by MxaJ and pulled away from the PQQ binding pocket of MxaF. Thus, the pocket is significantly exposed, enabling PQQ to pass through and bind to MxaF (Fig. 4b).
a Overall structure of the PQQ-unloaded MxaF/MxaJ complex. MxaJ (yellow) is shown in cartoon representation. MxaF is shown as a surface in green. b Close-up view of the PQQ binding pocket of MxaF opened by MxaJ. c Interactions between the roof sheet of MxaF and MxaJ. Critical amino acid residues of MxaF and MxaJ are shown in green and yellow stick representation, respectively. d Interactions between the L8 loop of MxaJ and MxaF. e Weblogo graphics of the multiple sequence alignments of the roof sheet of PQQ-dependent MDH and MxaJ L8 loop from PQQ-dependent dehydrogenase family (IPR017512) and methanol oxidation system protein MoxJ family (IPR022455), respectively. f Close-up view of the overlapped binding sites between MxaJ (yellow) and MxaI (purple) on MxaF (green).
The interaction between MxaF and MxaJ is primarily mediated by the roof sheet of MxaF and the L8 loop of MxaJ (named based on previously reported MxaJ from Methylophaga aminisulfidivorans19). Specifically, roof sheet residues Q429 and A434 form hydrogen bonds with residues Y164 and N162 in the L8 loop, respectively. Additionally, two salt bridges are established between L8 loop residue R161 and MxaF residue D303, as well as between MxaJ residue Q225 and roof sheet residue R426 (Fig. 4c, d). Multiple sequence alignments revealed that these key residues in the roof sheet and the L8 loop are highly conserved across species (Fig. 4e), suggesting a conserved interaction pattern between MxaF and MxaJ. Additionally, comparing the structures of the MxaF/MxaJ complex and the native MDH tetramer, we observed that the binding sites of MxaJ and MxaI on MxaF partially overlap (Fig. 4f), indicating the competition between MxaJ and MxaI in binding to MxaF. However, due to the lower affinity to MxaF, MxaI cannot displace MxaJ in the MxaF/MxaJ complex unless PQQ binds, suggesting that the binding of PQQ changes the structure of MxaF.
Dissociation of MxaJ from the MxaF/MxaJ complex
To understand the molecular basis of MxaJ release from MxaF/MxaJ complex mediated by PQQ and MxaI, the cryo-EM structure of the PQQ-loaded MxaF/MxaJ complex was further solved (Supplementary Fig. 10 and Table 2). The overall structure of the PQQ-loaded MxaF/MxaJ complex is similar to the MxaF/MxaJ complex alone, with the PQQ binding pocket remaining open (Fig. 5a). However, two regions of MxaF, A100 to D105 and G175 to V180, undergo significant conformational changes upon PQQ incorporation (Fig. 5b, c). The region A100 to D105 is completely reformed by PQQ to construct the unique disulfide ring in the catalytic center of MDH, which is critical for the entire catalytic process24,26,27,28. Additionally, PQQ binding induces allosteric changes in region G175-V180, relieving spatial hindrance with MxaI residue Q40 and facilitating the formation of two hydrogen bonds (Fig. 5d, e). These changes will enhance the interaction between MxaF and MxaI, shifting the binding preference of MxaF toward MxaI. Due to competition between MxaI and MxaJ, the binding of MxaI triggers MxaJ release. By interacting with the main part of MxaF and the bound MxaI, the roof sheet is stably tied to the PQQ binding pocket, ultimately resulting in the closure of the pocket (Fig. 5f).
a Sectional view of the cryo-EM structure of PQQ-loaded MxaF/MxaJ complex. MxaJ is shown in pink, and MxaF is shown in blue. A PQQ molecule embedded in MxaF is shown in green. b The Cα distance between two MxaF subunits from different MxaF/MxaJ complexes (PQQ-unloaded and PQQ-loaded) after alignment. c Close-up view of regions A100-D105 and G175-V180 of MxaF subunit in PQQ-unloaded MxaF/MxaJ complex (green), PQQ-loaded MxaF/MxaJ complex (blue), and complete MDH (PDB: 1H4I, gray). d Close-up view of G175-V180 region of MxaF in MxaF/MxaJ complex before (green) and after (blue) PQQ combination. Complete MDH (gray) is superimposed structurally, and the Ca2+ in MDH is shown in green sphere representation. e Close-up view of the interactions between MxaI (purple) and region G175-V180 of MxaF in complete MDH. The PQQ-unloaded and the PQQ-loaded MxaF/MxaJ complexes were superimposed with complete MDH structurally. f Interaction of the roof sheet with the main part of MxaF (gray) and MxaI (purple) in complete MDH. The position of roof sheet in MxaF/MxaJ complex is shown in green. Critical amino acid residues are shown in stick representation. g Model of chaperone-assisted PQQ-dependent MDH assembly process.
Based on these discoveries, we propose a putative PQQ assembly process as illustrated in Fig. 5g. Initially, the folded MxaF is captured by the chaperone MxaJ to form the MxaF/MxaJ complex. Subsequently, a PQQ molecule passes through the channel between MxaF and MxaJ and binds to the catalytic center of MxaF. The small subunit, MxaI, then interacts with MxaF, triggering the release of chaperone MxaJ from the PQQ-loaded MxaF/MxaJ complex, forming the PQQ-loaded MxaF/MxaI complex. Finally, with the assistance of proteins MxaR, MxaS, MxaA, MxaC, MxaK, and MxaL, Ca²⁺ is incorporated into the catalytic center of MxaF to form a complete functional enzyme.
Discussion
Cofactors, including coenzymes, prosthetic groups, and metal ions, are embedded in the active sites of protein templates, where they support numerous critical functions that proteins cannot perform alone29. Despite most known protein complexes can incorporate cofactors independently through direct protein–cofactor interactions, the involvement of auxiliary proteins in cofactor incorporation has also been well documented for a wide range of systems, including hydrogenase, molybdoenzyme nitrate reductase, nitrogenase, cytochrome c, and complexes of respiratory chain30,31,32,33,34,35. Although heterologous expression of many such protein complexes has been accomplished36,37, the molecular mechanisms by which these auxiliary proteins operate remain unclear. In this study, using heterologous expression strategy, auxiliary proteins involved in MxaFI-type MDH assembly, including chaperone MxaJ, were comprehensively identified. By solving the structures of complexes MxaF/MxaJ and PQQ-loaded MxaF/MxaJ, the mechanism of PQQ incorporation mediated by MxaJ was revealed, providing a deeper insight into how chaperones induce conformational changes in client proteins for cofactor binding.
Although the mechanism of XoxF-type MDH maturation remains to be elucidated, the presence of xoxJ, encoding a MxaJ homolog, within the XoxF-type MDH synthesis cluster suggests a conserved PQQ incorporation mechanism among PQQ-dependent MDHs38. In contrast to XoxF-type MDH, the maturation of MxaFI-type MDH necessitates an additional small MxaI subunit and six auxiliary proteins for proper Ca²⁺ incorporation. MxaI mediates MxaJ dissociation from MxaF and stabilizes the roof sheet, thereby facilitating conformational rearrangement of E177, a residue that provides two coordination bonds to Ca²⁺, to establish the mature Ca²⁺-binding site (Fig. 5d). The remaining six auxiliary proteins, including the MoxR-class AAA+ ATPase (MxaR) and von Willebrand factor type A (VWA) domain-containing proteins (MxaS, MxaC, MxaL)8, likely assemble into a MoxR/VWA complex. This evolutionarily conserved prokaryotic machinery specializes in metal cofactor insertion and presumably facilitates Ca²⁺ transport and incorporation39,40, though its precise molecular mechanism requires further elucidation.
Recently, reducing CO₂ into methanol using solar energy via chemical catalysts has been one of the most efficient methods for carbon assimilation41,42,43,44. Thus, bioconversion of methanol offers a promising solution to global energy crises and climate challenge. Due to the excellent properties of PQQ-dependent MDHs in catalyzing methanol oxidation, they were considered the optimal biocatalysts for methanol bio-industrial conversion7. Addressing the mechanism of PQQ-dependent MDHs biogenesis not only advances our understanding on the function and evolution of their assembly system, but also provides a more efficient way for the application of these enzymes in methanol bioconversion. This advancement is expected to promote the development and revolution of the methanol-based carbon assimilation technologies, contributing to the achievement of carbon neutrality.
Methods
Strains and culture conditions
The strains used in this study are listed in Supplementary Table 3. M. extorquens AM1 and its transformants were cultured at 30 °C in “Hypho” minimal salts medium supplemented with 15 mM succinate (HS) or 0.5% (v/v) methanol (125 mM) (HM) as the sole carbon source45. To minimize the contamination, rifamycin (50 μg mL−1) was routinely used and the additional tetracycline (20 μg mL−1) was used for selection purposes. E. coli Trans5α and BL21 (DE3) were cultivated in LB medium with appropriate antibiotics at the concentrations of 50 μg mL−1, 30 μg mL−1, and 40 μg mL−1 for kanamycin, chloramphenicol, and tetracycline, respectively. For the preparation of solid media, 1.5% (w/v) agar was added.
Gene cloning and plasmid construction
The plasmids used in this study are listed in Supplementary Table 4. In order to obtain naturally synthesized MDH, the MxaF subunit containing the C-terminal His6-tag was expressed in M. extorquens AM1 by using pCM80 vector. The coding sequence of mxaF gene was amplified from the genomic DNA (gDNA) of M. extorquens AM1 by using the primer pair whose reverse primer contained a sequence encoding His6-tag. Then, the PCR products were cleaned up and integrated with the linearized pCM80 vector digested by XbaI and SacI using Gibson Assembly Master Mix (NEB, USA) to generate plasmid pCM80-mxaF-his.
For the expression of the 13 genes from mxa cluster in E. coli, two plasmids pACYCDuet-1-mxaFI-ACKL and pHZDNh-mxaJGRSDEH were constructed. The plasmid pACYCDuet-1-mxaFI-ACKL contains two subunits encoding genes mxaF and mxaI under the control of the first T7 promotor, as well as contains four Ca2+ incorporation related genes mxaA, mxaC, mxaK, and mxaL controlled by the second promoter. To construct this plasmid, the coding sequences of the six genes were optimized and linked into two fragments mxaFI and mxaACKL by four de novo designed ribosome binding site (RBS) sequences. Then, the two fragments were synthesized and integrated into vector pACYCDuet-1 at the positions between NcoI and SacI, as well as between NdeI and XhoI successively, to obtain plasmids pACYCDuet-1-mxaFI and pACYCDuet-1-mxaFI-ACKL, respectively. For the purpose of protein purification, the His6-tag was linked to the C-terminus of MxaF.
In order to construct plasmid pHZDNh-mxaJGRSDEH, a modified pET28a (+) vector, pHZDNh, that deleted the sequence between NcoI and NdeI was generated at first by RF cloning46. Then the DNA fragment containing the coding sequences of proteins MxaJ, MxaG, MxaR, MxaS, MxaD, MxaE, and MxaH was synthesized and integrated into vector pHZDNh at the position between NdeI and XhoI. Each coding sequence is preceded by an RBS sequence. For the construction of the single gene deletion plasmids, a series of corresponding restriction sites were added at both ends of genes mxaJ, mxaR, mxaS, mxaD, mxaE, and mxaH.
The single gene deletion plasmids were constructed based on the plasmids pACYCDuet-1-mxaFI-ACKL and pHZDNh-mxaJGRSDEH. The fragments mxaCKL, mxaAKL, mxaACL, and mxaACK were amplified from the plasmid pACYCDuet-1-mxaFI-ACKL and separately integrated with the linearized pACYCDuet-1-mxaFI digested by NdeI and XhoI to generate gene mxaA, mxaC, mxaK, or mxaL deletion plasmid, respectively. The plasmid pHZDNh-mxaJGRSDEH was digested by a single restriction enzyme whose cleavage sites had been added to both ends of the target gene and ligated by using T4 ligase to obtain the corresponding single gene deletion plasmid.
The plasmid pHZDNh-mxaJRS that contains the coding sequences of protein MxaJ, MxaR, and MxaS was constructed using Golden Gate Assembly47. Firstly, the coding sequence of MxaJ was amplified from plasmid pHZDNh-mxaJGRSDEH using the primer pair whose reverse primer contains two BsaI restriction sites. Then the fragment was inserted into the vector pHZDNh, digested by NdeI and XhoI to yield plasmid pHZDNh-mxaJ. Secondly, the fragment of mxaRS was also amplified from plasmid pHZDNh-mxaJGRSDEH. After being digested by BsaI, the fragment mxaRS and the linearized plasmid pHZDNh-mxaJ were ligated by using T4 ligase to obtain the final plasmid pHZDNh-mxaJRS.
For overexpressing mxaI, the plasmid pHZDNh-mxaJRIS was built. Briefly, the coding sequence of protein MxaI was amplified from plasmid pACYCDuet-1-mxaFI using the primers containing a designed RBS sequence, Then, the fragment was integrated with the linearized plasmid pHZDNh-mxaJRS digested by BamHI and EcoRI, whose restriction sites had been pre-added between the coding sequences of MxaR and MxaS.
The plasmid pACYCDuet-1-mxaF-6his used for the production of protein MxaF was constructed by integrating the coding sequence of his6-tag-containing MxaF amplified from plasmid pACYCDuet-1-mxaFI with the linearized vector pACYCDuet-1 digested by NcoI and SacI.
To construct the plasmid pACYCDuet-1-F-J-6his, the coding sequences of MxaF and MxaJ were amplified from plasmids pACYCDuet-1-mxaFI and pHZDNh-mxaJ, respectively. The his6-tag was fused to the C-terminus of the MxaJ rather than MxaF. The two fragments were successively integrated into vector pACYCDuet-1 at the positions between NcoI and SacI, as well as between NdeI and XhoI.
To obtain protein MxaI and MxaJ for MDH assembly in vitro, the coding sequences were amplified from plasmid pACYCDuet-1-mxaFI and pHZDNh-mxaJ, respectively, and then separately integrated with the linearized plasmid pHZDNh digested by NdeI and XhoI to yield plasmid pHZDNh -mxaI-6his and pHZDNh -mxaJ-6his. All the Primers used for gene cloning are listed in Supplementary Table 5.
Genetic manipulation
The traditional chemical transformation method was employed to introduce the plasmid into E. coli and the competent cells were purchased from TransGen Biotech, China. The transformation of M. extorquens AM1 was conducted by using electroporation at a voltage of 2 kV as previously described48, and the transformants bearing the corresponding plasmid were selected by using tetracycline resistance phenotype.
Protein expression and purification
For the homologous expression of MDH in M. extorquens AM1, the positive colonies were initially grown in 10 mL HS medium at 30 °C with shaking (220 rpm) until the mid-exponential phase. Then 2 mL of a sub-culture was transferred into 400 mL of HM medium for protein expression under the same growth conditions. The expression continued for 96 h, and 100% methanol was added every 24 h to a final concentration of 0.5% (v/v) to maintain the cell growth and protein expression.
For the protein expression in E. coli, the recombinants with corresponding expression plasmids were cultivated in LB medium at 37 °C by shaking up to an OD600 of 0.6. IPTG was added routinely to the final concentration of 0.2 mM to induce the protein expression at 16 °C overnight. When expressing MDH, the additional PQQ and CaCl2 were added to the final concentration of 10 μM and 100 μM, respectively, while adding IPTG.
The general protein purification method was described below. The cells were harvested by centrifugation at 5500 × g for 20 min and resuspended in lysis buffer (20 mM Tirs-HCl pH 8.0, 150 mM NaCl, 5 mM imidazole). Then the cells were cracked by the high-pressure homogenizer and centrifuged at 11,000 × g for 30 min. The supernatant was loaded onto nickel affinity columns (GE Healthcare) pre-equilibrated using the lysis buffer. After washed out the unspecific bound proteins by washing buffer (20 mM Tris-HCl pH 8.0, 150 mM NaCl, and 50 mM imidazole), the target protein was eluted by elution buffer (20 mM Tris-HCl pH 8.0, 150 mM NaCl, 400 mM imidazole) and changed into storage buffer (20 mM Tris-HCl, pH 8.0, 150 mM NaCl, 10% (v/v) glycerol) by using desalting column (GE Healthcare). The protein was further concentrated by ultrafiltration as required and the concentration was determined by Bradford kit (Bio-red) with bovine serum albumin as a standard.
For the purification of MxaF/MxaJ complex for structural studies, the strain containing plasmid pACYCDuet-1-mxaF-J-his, was cultured and induced in 5 L LB medium under the conditions as mentioned above. Cells were harvested by centrifugation at 5500 × g for 20 min and resuspended in 25 mM Tris-HCl buffer (pH 8.0) containing 500 mM NaCl, 20 mM imidazole. Then the cells were disrupted by the high-pressure homogenizer and the cell debris was removed by centrifugation at 17,000 × g for 1 hour. The supernatant was applied onto a Ni-NTA column FPLC system (Cytiva) pre-equilibrated with buffer (25 mM Tris-HCl, pH 8.0, 500 mM NaCl, 20 mM imidazole). The target protein was dialyzed against buffer containing 25 mM Tris-HCl, pH 8.0, overnight, and applied to a DEAE column FPLC system (Cytiva). Target proteins were eluted at ~125 mM NaCl, and then dialyzed against buffer containing 25 mM Tris-HCl, 150 mM NaCl, pH 8.0, overnight. After being concentrated, the sample was applied to a pre-equilibrated size exclusion column (HiLoad Superdex 200 10/30, 24 mL, Cytiva) at a flow rate of 0.5 mL min−1. MxaF/MxaJ complexes showed a single peak on the elution profiles, and the purity of the samples was checked by SDS-PAGE. The purified proteins were concentrated to 10 mg mL−1 in 25 mM Tris-HCl, 150 mM NaCl, pH 8.0, buffer for cryo-EM assays. For the preparation of the PQQ-loaded MxaF/MxaJ complex, the buffer used for size-exclusion chromatography was added additional 250 μM PQQ.
Activity assays
The activity of MDH was determined by a traditional dye-linked assay as previously described49,50. Briefly, the standard reaction mixture contains 100 mM Tris-HCl (pH 9.0), 15 mM NH4Cl, 200 μM 2,6-dichlorophenol-indophenol (DCPIP), 1 mM phenazine methosulfate (PMS), 500 μM Methanol, and 10 to 50 mg L-1 of enzyme. By tracking the decrease of the absorbance at OD600, the PMS-mediated DCPIP reduction was monitored, and the reduction rate was calculated based on the extinction coefficient of DCPIP at pH 9.0 (ε600 = 21 mM−1 cm−1)51,52. To minimize the impact of the substrate-independent reduction of DCPIP on the enzyme activity measurement, each reaction was set with a control that without substrate addition, and the background activity was subtracted. The kinetic parameters for MDHs were calculated according to the specific activities of the enzyme towards diverse concentrations (0, 10, 30, 50, 100, 250, and 500 μM) of methanol based on the Michaelis-Menten equation.
Spectrophotometric measurement
200 μL protein preparations were added into the 96-well plate to record the absorption spectra from 250 nm to 600 nm at room temperature by using Spectra Max M2e spectrophotometer (Molecular Devices) with 5 nm spectral band width.
Ca2+ incorporation in vitro
For the incorporation of Ca2+ in vitro, the reactions took place in 20 mM Tris-HCl (pH 9.5) buffer based on the previous studies16,53. The protein preparations were incubated with 10 mM CaCl2 at room temperature overnight, and then the mixtures were desalted by using a desalting column and 20 mM Tris-HCl buffer (pH 8.0).
Metal analysis by inductively coupled plasma optical emission spectrometry
The purified MDH with or without Ca2+ incorporation in vitro was changed into 20 mM Tris-HCl buffer (pH 8.0) by using a desalting column. The calcium contents of the enzymes were determined by using an Inductively Coupled Plasma Optical Emission Spectrometry (ICP-OES; PerkinElmer 7300DV) at the wavelength of 317.9 nm.
Identification of MxaF/MxaJ complex
The target bands in the Native-PAGE gel were collected, and the in-gel digestion was conducted as described previously with minor modifications54. Proteins were reduced by 10 mM dithiothreitol (DTT) at 97 °C for 10 min and alkylated by 40 mM 2-Chloroacetamide (CAA) at room temperature for 30 min. Gel pieces were dehydrated with 100% acetonitrile. Proteolytic digestion was performed with 30 ng μL−1 trypsin (Sequencing grade, Promega) dissolved in 50 mM ammonium bicarbonate and was incubated at 37 °C overnight. The peptides were extracted with 100% acetonitrile and were dried under vacuum.
Peptide separation was carried out on a nanoElute liquid chromatography system (Bruker Daltonics). Digested peptides were separated within 60 min at a flow rate of 350 nL min−1 on a homemade column 25 cm × 75 μm, 1.9 μm C18-AQ particles (Dr. Maisch). Mobile phases A and B were water and ACN with 0.1% formic acid, respectively. The %B was linearly increased from 5 to 32% within 50 min, then increased to 95% within 6.9 min, and finally maintained at 95% for the last 3.1 min.
All samples were analyzed on a hybrid TIMS quadrupole time-of-flight mass spectrometer (Bruker timsTOF Pro2) via a CaptiveSpray nanoelectrospray ion source. The mass spectrometer was operated in data-dependent mode for sample analysis. The accumulation and ramp times were set at 100 ms each, and the mass spectra were recorded in the range from m/z100 to 1700 in positive electrospray mode. The ion mobility was scanned from 0.6 to 1.5 V s/cm2. We acquired dda-PASEF with four PASEF scans per top-N acquisition cycle, an intensity threshold of 1000 arbitrary units (a.u.) as indicated by the Bruker acquisition software, and a target value of 6000 a.u. for precursor selection. Precursors that reached this target value were excluded for 0.4 min. Singly charged precursors were filtered out based on their m/z-ion mobility position, and precursors with a mass below 700 Da were isolated with a quadrupole selection window of 2 Th and otherwise with 3 Th.
Data was analyzed by MaxQuant Software (Version2.1.4.0)55. We used the reviewed E. coli (BL21-DE3) proteome (UniProt,4172 entries without isoforms) and default settings for the analyses. This comprised a false discovery rate (FDR) of 1%, two missed cleavages, the cleavage pattern for trypsin, cysteine carbamidomethylation as fixed modification, and methionine oxidation and protein N-terminal acetylation as variable modifications. For the analysis of proteomic data, iBAQ, and LFQ were used. The mass tolerance was set to 10 ppm for the main search, and known contaminants were excluded from the dataset56.
Isothermal titration calorimetry
Isothermal titration calorimetry (ITC) assays were conducted by using the MicroCal PEAQ-ITC system (Malvern Panalytical Ltd). All experiments were performed at 25 °C, in 20 mM HEPES buffer (pH 7.5) containing 150 mM NaCl. The ITC cell contained ~20 μM proteins (MxaF, MxaF/MxaI complex, or MxaF/MxaJ complex as required). The titrant syringe contained 200 µM PQQ, 80 µM MxaJ, or 150 µM MxaI as required in the same buffer. Titration settings were the following: 750 rpm stirring speed, 60 s initial delay, 8 µcal/s reference power, and 120 s spacing between each injection. Titrations consisted of a first 0.2 µL injection followed by 18 × 2 µL injections. Titrations of ligand solutions into the cell containing buffer without protein showed negligible heats of dilution. The resulting data were fitted using the ITC evaluation software to one set of sites model, provided by the manufacturer, to obtain the number of sites (n), association constant (K), binding enthalpy (ΔH), and entropy change (ΔS), from which the free energy change (ΔG) was calculated.
PQQ incorporation in vitro
The reactions were carried out in 1 mL 20 mM HEPES (pH 7.5) buffer, and the standard mixture contained 10 μM MxaF, 10 μM MxaJ, 20 μM MxaI, and 20 μM PQQ. After being incubated at room temperature for 1 h, 20 μL of the mixture was taken for Native-PAGE detection, and the remaining mixture was changed into 20 mM Tris-HCl (pH 8.0) buffer by using a desalting column (GE Healthcare) to remove the unbound PQQ and for further assays.
Surface plasmon resonance assays
Surface plasmon resonance (SPR) experiments were performed on a BIAcore 8 K instrument (Cytiva) at 25 °C. Protein MxaJ was immobilized to a level of 400 RU on the sample flow cell of S-series CM5 sensor chip using amine-coupling chemistry at pH 5.0, according to the manufacturer’s instructions. The reference flow cell was treated identically, with activation and deactivation, but without immobilization of protein MxaJ. Then, a dual injection experiment was performed. First, 10 µM MxaF in a running buffer consisting of 20 mM HEPES, 150 mM NaCl, 0.075% (v/v) Tween20, pH 7.5, was injected over the protein MxaJ surface for 120 s, followed by the injection of 20 µM PQQ or a mixture of 20 µM PQQ and 20 µM MxaI. Blank injection of buffer was also carried out and subtracted for each injection.
Determination of the structures of MxaF/MxaJ complexes
To prepare the cryo sample, the MxaF/MxaJ complex and the PQQ-loaded MxaF/MxaJ (MxaF/MxaJ/PQQ) complex samples were both vitrified using a Vitrobot Mark IV (ThermoFisher Scientific) plunge freezing device. The MxaF/MxaJ complex sample (4.0 μL, 1.0 mg mL−1) was applied to a Cu Quantifoil 1.2/1.3 holey carbon grid with glow discharged for 40 seconds. The PQQ-loaded MxaF/MxaJ complex (MxaF/MxaJ/PQQ complex) sample (4.0 μL, 1.0 mg mL−1) was applied to a 35 seconds glow-discharged Au Quantifoil 1.2/1.3 holey carbon grid. Grids above were then blotted using different conditions (blot time 5 s and blot force −6 for MxaF/MxaJ complex; blot time 6 s and blot force −6 for the MxaF/MxaJ/PQQ complex) at a temperature of 4 °C and a humidity level of >99% and plunge frozen into liquid ethane.
The prepared grids were transferred to a 300 kV Titan Krios transmission electron microscope equipped with Gatan K3 detector and GIF Quantum energy filter. Movies were collected at 105,000x magnification with a calibrated pixel size of 0.69 Å over a defocus range of −1.0 μm to −2.0 μm in super resolution counting mode with a total dose of 60 e−/Å2 using EPU (ThermoFisher Scientific) automated acquisition software.
The detailed data processing workflow is summarized in Supplementary Fig. 9 and 10. All the raw dose-fractionated images stacks were 2× binned, aligned, dose-weighted, and summed using MotionCor257. The contrast transfer function (CTF) estimation, particle picking and extraction, 2D classification, ab initio model generation, 3D refinements were performed in cryoSPARC v.4.2.158.
For the MxaF/MxaJ complex, a total of 5126 micrographs were collected for this dataset, from which 742,831 initial particles were picked and extracted with a box size of 400 pixels. After three rounds of iterative 2D classification, a clean set of 603,925 particles was used to generate five ab-initio reconstructions. To perform further discarding of bad particles, the heterogeneous refinement was processed by using the one dominant class with the associated particles of the five initial volumes. Among the five 3D classes, one volume containing ~42.71% of total particles was subjected to the homogeneous refinement, and iterative global CTF refinement was performed, yielding a final density map at 2.78 Å resolution estimated by the gold-standard Fourier shell correlation (FSC) 0.143 criterion. The final map was sharpened by DeepEMhancer.
The prototype MxaF/MxaJ/PQQ complex dataset was processed similarly. In another initially collected dataset, we picked out particles using blob-pick procedure of cryoSPARC from 1000 micrographs, and then these particles were subjected to 2D classification. After three rounds of 2D classification, we selected good particles in different views for Topaz training and then generated the Topaz model. Then we applied the Topaz59 procedure to select particles against entire micrographs. The 510,868 initial particles were picked and extracted from 5001 micrographs. After the extensive 2D classification, approximately 259,610 good particles were selected to generate the initial models and 3D classification and resulting to five distinct volumes. The one dominant class, containing 46.30% of total particles, was identified, which displayed clear features of secondary structural elements, especially in the area of the binding interface of the MxaF and MxaJ. These particles were subjected to homogeneous, non-uniform, and CTF refinements in cryoSPARC v.4.2.1, which yielded a final density map at 2.81 Å resolution estimated by the gold-standard Fourier shell correlation cut-off value of 0.143. The final map was sharpened by DeepEMhancer.
For the initial model building of the MxaF/MxaJ and MxaF/MxaJ/PQQ complex, the PDB code 1H4I and 5SV6 were used as the starting model and fitted into the corresponding cryo-EM maps using UCSF Chimera v.1.1560. Mutation and manual adjustment were carried out with COOT v.0.9.361. Each residue was manually checked with the chemical properties taken into consideration during model building. Several rounds of the real-space refinement in Phenix-1.20.162 and manually building in Coot were performed until the final reliable models were obtained. Molprobity63 was used to validate geometry and check structure quality. Statistics associated with data collection, 3D reconstruction, and model building were summarized in Supplementary Table 2. Figures were generated using Chimera60 and PyMol v.2.0 (http://www.pymol.org).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data that support the findings of this study are presented in the main text or Supplementary Information. And the source data underlying Figs. 2b–e, 3a–f, 5b, and Supplementary Figs. 2a–d, 3b, c, 4, 5 are provided in the Source Data file with this paper or available from the corresponding author on request. Source data are provided with this paper. The cryo-EM maps have been deposited in the Electron Microscopy Data Bank (EMDB) under accession codes EMD-60838 (MxaF/MxaJ complex) and EMD-60840 (PQQ-loaded MxaF/MxaJ complex). The atomic coordinates have been deposited in the Protein Data Bank (PDB) under accession codes 9ISM (MxaF/MxaJ complex); and 9ISO (PQQ-loaded MxaF/MxaJ complex). Source data are provided with this paper.
References
Hanson, R. S. & Hanson, T. E. Methanotrophic bacteria. Microbiol. Rev. 60, 439–471 (1996).
Jackson, R. B. et al. Increasing anthropogenic methane emissions arise equally from agricultural and fossil fuel sources. Environ. Res. Lett. 15, 071002 (2020).
Fei, Q. et al. Bioconversion of natural gas to liquid fuel: Opportunities and challenges. Biotechnol. Adv. 32, 596–614 (2014).
Tucci, F. J. & Rosenzweig, A. C. Direct methane oxidation by copper- and iron-dependent methane monooxygenases. Chem. Rev. 124, 1288–1320 (2024).
Anthony, C. The c-type cytochromes of methylotrophic bacteria. Biochim Biophys. Acta 1099, 1–15 (1992).
Anthony, C. The pyrroloquinoline quinone (PQQ)-containing dehydrogenases. Respiration in archaea and bacteria: diversity of prokaryotic electron transport. Carriers 15, 203–225 (2004).
Krusemann, J. L., Rainaldi, V., Cotton, C. A., Claassens, N. J. & Lindner, S. N. The cofactor challenge in synthetic methylotrophy: bioengineering and industrial applications. Curr. Opin. Biotechnol. 82, 102953 (2023).
Keltjens, J. T., Pol, A., Reimann, J. & Op den Camp, H. J. PQQ-dependent methanol dehydrogenases: rare-earth elements make a difference. Appl. Microbiol. Biotechnol. 98, 6163–6183 (2014).
Chistoserdova, L. & Kalyuzhnaya, M. G. Current trends in methylotrophy. Trends Microbiol. 26, 703–714 (2018).
Pol, A. et al. Rare earth metals are essential for methanotrophic life in volcanic mudpots. Environ. Microbiol. 16, 255–264 (2014).
Ghoshl, M., Anthony, C., Harlos, K., Goodwin, M. G. & Blake, C. The refined structure of the quinoprotein methanol dehydrogenase from Methylobacterium extorquens at 1.94 Å. Structure 3, 177–187 (1995).
Anthony, C. & Williams, P. The structure and mechanism of methanol dehydrogenase. Biochim. Biophys. Acta 1647, 18–23 (2003).
Amaratunga, K., Goodwin, P. M., O’Connor, C. D. & Anthony, C. The methanol oxidation genes mxaFJGIR (S) ACKLD in Methylobacterium extorquens. FEMS Microbiol. Lett. 146, 31–38 (1997).
Tanaka, Y., Yoshida, T., Watanabe, K., Izumi, Y. & Mitsunaga, T. Cloning and analysis of methanol oxidation genes in the methylotroph Hyphomicrobium methylovorum GM2. FEMS Microbiol. Lett. 154, 397–401 (1997).
Morris, C. J., Kim, Y. M., Perkins, K. E. & Lidstrom, M. E. Identification and nucleotide sequences of mxaA, mxaC, mxaK, mxaL, and mxaD genes from Methylobacterium extorquens AM1. J. Bacteriol. 177, 6825–6831 (1995).
Richardson, I. W. & Anthony, C. Characterization of mutant forms of the quinoprotein methanol dehydrogenase lacking an essential calcium ion. Biochem. J. 287, 709–715 (1992).
Chistoserdova, L., Chen, S. W., Lapidus, A. & Lidstrom, M. E. Methylotrophy in Methylobacterium extorquens AM1 from a genomic point of view. J. Bacteriol. 185, 2980–2987 (2003).
Van Spanning, R. J. M. et al. Isolation and characterization of the moxJ, moxG, moxI, and moxR genes of Paracoccus denitrificans: inactivation of moxJ, moxG, and moxR and the resultant effect on methylotrophic growth. J. Bacteriol. 173, 6948–6961 (1991).
Choi, J. M., Cao, T.-P., Kim, S. W., Lee, K. H. & Lee, S. H. MxaJ structure reveals a periplasmic binding protein-like architecture with unique secondary structural elements. Proteins 85, 1379–1386 (2017).
Zhang, M. & Lidstrom, M. E. Promoters and transcripts for genes involved in methanol oxidation in Methylobacterium extorquens AM1. Microbiology 149, 1033–1040 (2003).
Vuilleumier, S. et al. Methylobacterium genome sequences: a reference blueprint to investigate microbial metabolism of C1 compounds from natural and industrial sources. PLoS One 4, e5584 (2009).
Springer, A. L., Auman, A. J. & Lidstrom, M. E. Sequence and characterization of mxaB, a response regulator involved in regulation of methanol oxidation, and of mxaW, a methanol-regulated gene in Methylobacterium extorquens AM1. FEMS Microbiol. Lett. 160, 119–124 (1998).
Anderson, D. J. & Lidstrom, M. E. The moxFG region encodes four polypeptides in the methanol-oxidizing bacterium Methylobacterium sp. strain AM1. J. Bacteriol. 170, 2254–2262 (1988).
Sarmiento-Pavia, P. D. & Sosa-Torres, M. E. Bioinorganic insights of the PQQ-dependent alcohol dehydrogenases. J. Biol. Inorg. Chem. 26, 177–203 (2021).
Goodwin, M. G. & Anthony, C. Characterization of a novel methanol dehydrogenase containing a Ba2+ ion at the active site. Biochem. J. 318, 673–679 (1996).
Chan, S. I. et al. Mechanism of pyrroloquinoline quinone-dependent hydride transfer chemistry from spectroscopic and high-resolution x-ray structural studies of the methanol dehydrogenase from Methylococcus capsulatus (Bath). J. Am. Chem. Soc. 143, 3359–3372 (2021).
Blake, C. C., Ghosh, M., Harlos, K., Avezoux, A. & Anthony, C. The active site of methanol dehydrogenase contains a disulphide bridge between adjacent cysteine residues. Nat. Struct. Biol. 1, 102–105 (1994).
Avezoux, A., Goodwin, M. G. & Anthony, C. The role of the novel disulphide ring in the active site of the quinoprotein methanol dehydrogenase from Methylobacterium extorquens. Biochem. J. 307, 735–741 (1995).
Kirschning, A. The coenzyme/protein pair and the molecular evolution of life. Nat. Prod. Rep. 38, 993–1010 (2021).
Lubitz, W., Ogata, H., Rudiger, O. & Reijerse, E. Hydrogenases. Chem. Rev. 114, 4081–4148 (2014).
Leimkuhler, S. & Iobbi-Nivol, C. Bacterial molybdoenzymes: old enzymes for new purposes. FEMS Microbiol. Rev. 40, 1–18 (2016).
Jasniewski, A. J., Lee, C. C., Ribbe, M. W. & Hu, Y. Reactivity, mechanism, and assembly of the alternative nitrogenases. Chem. Rev. 120, 5107–5157 (2020).
Sanders, C., Turkarslan, S., Lee, D. W. & Daldal, F. Cytochrome c biogenesis: the Ccm system. Trends Microbiol. 18, 266–274 (2010).
McNeil, M. B. & Fineran, P. C. Prokaryotic assembly factors for the attachment of flavin to complex II. Biochim. Biophys. Acta 1827, 637–647 (2013).
Watson, S. A. & McStay, G. P. Functions of cytochrome c oxidase assembly factors. Int. J. Mol. Sci. 21, 7254 (2020).
Schiffels, J. et al. An innovative cloning platform enables large-scale production and maturation of an oxygen-tolerant [NiFe]-hydrogenase from Cupriavidus necator in Escherichia coli. PLoS One 8, e68812 (2013).
Solomon, J. B. et al. Ammonia synthesis via an engineered nitrogenase assembly pathway in Escherichia coli. Nat. Catal. 7, 1130–1141 (2024).
Featherston, E. R. et al. Biochemical and structural characterization of XoxG and XoxJ and their roles in Lanthanide-dependent methanol dehydrogenase activity. Chembiochem 20, 2360–2372 (2019).
Bhandari, V., Van Ommen, D. A. J., Wong, K. S. & Houry, W. A. Analysis of the evolution of the MoxR ATPases. J. Phys. Chem. A 126, 4734–4746 (2022).
Kahle, M., Appelgren, S., Elofsson, A., Carroni, M. & Adelroth, P. Insights into the structure-function relationship of the NorQ/NorD chaperones from Paracoccus denitrificans reveal shared principles of interacting MoxR AAA+/VWA domain proteins. BMC Biol. 21, 47 (2023).
Shih, C. F., Zhang, T., Li, J. & Bai, C. Powering the future with liquid sunshine. Joule 2, 1925–1949 (2018).
Jiang, X., Nie, X., Guo, X., Song, C. & Chen, J. G. Recent advances in carbon dioxide hydrogenation to methanol via heterogeneous catalysis. Chem. Rev. 120, 7984–8034 (2020).
Zhong, J. et al. State of the art and perspectives in heterogeneous catalysis of CO(2) hydrogenation to methanol. Chem. Soc. Rev. 49, 1385–1413 (2020).
Sha, F., Han, Z., Tang, S., Wang, J. & Li, C. Hydrogenation of carbon dioxide to methanol over non-Cu-based heterogeneous catalysts. ChemSusChem 13, 6160–6181 (2020).
Okubo, Y., Skovran, E., Guo, X., Sivam, D. & Lidstrom, M. E. Implementation of microarrays for Methylobacterium extorquens AM1. OMICS 11, 325–340 (2007).
van den Ent, F. & Lowe, J. RF cloning: a restriction-free method for inserting target genes into plasmids. J. Biochem. Biophys. Methods 67, 67–74 (2006).
Engler, C. & Marillonnet, S. Generation of families of construct variants using golden gate shuffling. Methods Mol. Biol. 729, 167–181 (2011).
Toyama, H., Anthony, C. & Lidstrom, M. E. Construction of insertion and deletion mxa mutants of Methylobacterium extorquens AM1 by electroporation. FEMS Microbiol. Lett. 166, 1–7 (1998).
Jahn, B. et al. Understanding the chemistry of the artificial electron acceptors PES, PMS, DCPIP and Wurster’s Blue in methanol dehydrogenase assays. J. Biol. Inorg. Chem. 25, 199–212 (2020).
Gutenthaler, S. M., Phi, M. T., Singer, H. & Daumann, L. J. Activity assays of methanol dehydrogenases. Methods Enzymol. 650, 57–79 (2021).
Good, N. M. et al. Pyrroloquinoline quinone ethanol dehydrogenase in Methylobacterium extorquens AM1 extends lanthanide-dependent metabolism to multicarbon substrates. J. Bacteriol. 198, 3109–3118 (2016).
Vu, H. N. et al. Lanthanide-dependent regulation of methanol oxidation systems in Methylobacterium extorquens AM1 and their contribution to methanol growth. J. Bacteriol. 198, 1250–1259 (2016).
Goodwin, M. G., Avezoux, A., Dales, S. L. & Anthony, C. Reconstitution of the quinoprotein methanol dehydrogenase from inactive Ca2+-free enzyme with Ca2+, Sr2+ or Ba2+. Biochem. J. 319, 839–842 (1996).
Shevchenko, A., Tomas, H., Havlis, J., Olsen, J. V. & Mann, M. In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat. Protoc. 1, 2856–2860 (2006).
Schwanhäusser, B. et al. Global quantification of mammalian gene expression control. Nature 473, 337–342 (2011).
Chen, Y. et al. Proteomic analysis identifies prolonged disturbances in pathways related to cholesterol metabolism and myocardium function in the COVID-19 recovery stage. J. Proteome Res. 20, 3463–3474 (2021).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Bepler, T. et al. Positive-unlabeled convolutional neural networks for particle picking in cryo-electron micrographs. Nat. Methods 16, 1153–1160 (2019).
Pettersen, E. F. et al. UCSF Chimera-a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. Biol. Crystallogr. 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D. Biol. Crystallogr. 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D. Biol. Crystallogr. 66, 12–21 (2010).
Acknowledgements
This work was supported by the Strategic Priority Research Program of the Chinese Academy of Sciences (XDC0120302 to H.F.J.) and Tianjin Synthetic Biotechnology Innovation Capacity Improvement Project (TSBICIP-KJGG-009-02 to H.F.J. and TSBICIP-CXRC-030 to H.C.Z.). We thank Professor Song Yang from Qingdao Agricultural University for providing the plasmid, strain, and assistance in strain cultivation; Shixin Zhang from Dalian Institute of Chemical Physics, Chinese Academy of Sciences, for technical support.
Author information
Authors and Affiliations
Contributions
H.F.J., F.G., H.J.D. and H.C.Z. conceived the project. H.C.Z., M.W., Q.L., C.Y., L.P.L., Y.W.L., and W.D.L. conducted biochemical experiments. J.Q.S., H.X.X., and F.G. obtained all cryo-EM structures. J.B. and J.C. conducted molecular dynamics simulations and evolutionary analysis. M.M.H. and Q.C.C. conducted mass spectrometry. J.S. conducted SPR experiments. H.F.J. and H.C.Z. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Chris Anthony and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhou, H., Sun, J., Cheng, J. et al. Deciphering the assembly process of PQQ dependent methanol dehydrogenase. Nat Commun 16, 6672 (2025). https://doi.org/10.1038/s41467-025-61958-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-61958-w