Abstract
Emerging evidence indicates that metabolic signals—including nutrient availability, biosynthetic intermediates, and energy balance—are linked to cell cycle progression. However, how these signals are sensed by the cell cycle machinery remains unclear. Citrate, a key intermediate in the TCA cycle, peaks during mitosis (M phase) and is detected by the glycolytic enzyme ATP-dependent 6-phosphofructokinase 1 muscle isoform (PFKM), accelerating mitotic progression. Mechanistically, citrate binds PFKM, disrupting its tetrameric structure into dimers. Dimeric PFKM interacts with nucleosomes and phosphorylates histone H3 at serine 10 (H3S10), functioning as a protein kinase to promote mitosis and cell proliferation. Structural simulations reveal that PFKM binds nucleosomes optimally when H3S10 aligns with its catalytic site. Disrupting citrate-PFKM or PFKM-H3 interactions reduces H3S10 phosphorylation, delays mitosis, and suppresses tumor growth and T-cell proliferation. Our findings demonstrate that PFKM acts as a citrate sensor, coupling metabolic signals to cell cycle regulation.
Similar content being viewed by others
Introduction
Cell proliferation is an important biological process that maintains the normal structure and function of organs and tissues. Cell cycle is a highly regulated process that enables cell growth, replication of genetic material, and cell division1. Cell cycle progression is finely orchestrated by the sequential activation of CDKs and cyclin partners2. Proper cell division is also controlled by cell cycle checkpoints that monitor the order, integrity, and fidelity of the main events of the cell cycle3. Deregulated cell cycle can lead to a variety of diseases, including cancer, aging and immune dysfunction.
It has been shown recently that certain metabolic pathways must be activated to meet fluctuating biosynthetic demands across cell cycle phases. In the S phase, nucleotide synthesis is activated to meet the needs of DNA replication4. In mitotic phase, fatty acid synthesis is upregulated to produce sufficient lipids to ensure membrane integrity of daughter cells during cell division5. In contrast, it has been rarely explored how these metabolic intermediates are accurately sensed by cell cycle machinery to ensure that there is enough material to complete the transition between cell cycle phases. A most recent study demonstrates that lactate stimulates efficient mitotic exit in proliferative human cells by remodeling the anaphase-promoting complex, which is initiated upon mitotic entry when lactate abundance reaches its apex6, implicating the importance of delineation of metabolite-sensing mechanism during cell cycle progression.
Citrate is an intermediate of the tricarboxylic acid (TCA) cycle, produced from oxaloacetate (OAA) and acetyl-CoA by citrate synthase (CS) in mitochondria. Citrate can be used for biosynthesis and the production of ATP via oxidative phosphorylation (OXPHOS)7. Although citrate is an important metabolic regulator involved in multiple pathological and physiological processes, including cancer and inflammation, whether and how the levels of citrate are sensed by cells to guarantee proper cell cycle progression remains unknown. In this study, we investigated the role of citrate in cell cycle progression and revealed PFKM as a citrate sensor. By sensing high levels of citrate in mitosis, PFKM interacts with and phosphorylates histone H3 to promote mitotic progression of the cells.
Results
High levels of citrate are sensed by PFKM to accelerate mitotic progression
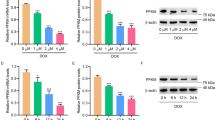
To determine whether the levels of citrate can be sensed by cell cycle machinery, we synchronized HeLa cells at mitotic (M) phase and then released the cells to G1, S or G2 phase, followed by liquid chromatograph mass spectrometer (LC-MS) analysis of citrate abundance at each phase, showing that the levels of citrate reached the peak at M phase compared to that at interphase (Fig. 1a), which is consistent with a previous metabolomics study8. Mitosis is a process by which cells divide and produce copies of the cells and is therefore the basis behind cell proliferation. In order to clearly analyze the stages of mitosis, we generated HeLa cells stably expressing EGFP-tagged histone H2B (HeLa/H2B-EGFP) to visualize the dynamics of chromosomal architecture in living cells (Supplementary Fig. 1a). To investigate the role of citrate in mitotic progression, we synchronized HeLa/H2B-EGFP cells in S phase with thymidine double block and incubated these cells with increasing doses of exogenous citrate. Mitotic index was calculated 14 h after the release from thymidine double block when most HeLa cells exited from mitosis (Supplementary Fig. 1b). The incubation of exogenous citrate, which simultaneously increased the levels of intracellular citrate (Supplementary Fig. 1c), reduced the number of the cells in M phase in a dose-dependent manner (Fig. 1b). Moreover, we measured mitotic duration from prophase to anaphase onset in synchronized HeLa/H2B-EGFP cells incubated with or without exogenous citrate basing on chromatin morphologies during the course of a video from time-lapse microscopy. As shown in Fig. 1c, exogenous citrate incubation markedly shortened mitotic duration in the cells. In contrast, depleting citrate synthase (CS) (Supplementary Fig. 1d), an enzyme in TCA cycle that catalyzes the production of citrate from acetyl-CoA and oxaloacetate, reduced intracellular citrate levels (Supplementary Fig. 1e), increased the number of mitotic cells (Fig. 1d) and prolonged the mitotic duration of the cells (Fig. 1e). Furthermore, we supplemented citrate to CS-depleted cells and observed that the mitotic index, which was elevated in CS-depleted cells, was reduced by the addition of citrate (Supplementary Fig. 1f). Consistently, the prolonged mitotic duration induced by CS depletion could be also rescued by citrate treatment (Supplementary Fig. 1g). These results reveal the important role of citrate in mitotic phase and implicate the potential association between the levels of citrate and the progression of mitosis.
a HeLa cells were synchronized in mitosis and subsequently released for the indicated time periods, followed by mass spectrometry to measure the levels of citrate in the cells. Rel: relative. b HeLa/H2B-EGFP cells were treated with different doses of citrate, mitotic index was calculated at 14 h after release from thymidine double block. c HeLa/H2B-EGFP cells were treated with different doses of citrate and synchronized for time-lapse microscopic imaging as described in Supplementary Fig. 1b. Representative images of cell cycle progression were shown and the transition from prophase to anaphase onset was highlighted in red rectangles. Mitotic duration from prophase to anaphase onset was measured according to the time points when corresponding chromatin morphologies occurred. 0 mM: n = 105; 0.5 mM: n = 109; 2.5 mM: n = 104; 5 mM: n = 112. d Citrate synthase (CS) was depleted in HeLa/H2B-EGFP cells with siRNA against CS, mitotic index was calculated at 14 h after release from thymidine double block. e HeLa/H2B-EGFP cells with or without CS depleted were synchronized for time-lapse microscopic imaging as described in Supplementary Fig. 1b. Representative images of cell cycle progression were shown and the transition from prophase to anaphase onset was highlighted in red rectangles. Mitotic duration from prophase to anaphase onset was measured. siNC: n = 100; siCS−1: n = 96; siCS−2: n = 99. f Flow chart depicting the LiP-SMap assay. Freshly prepared whole-cell lysates were treated with or without citrate followed by proteinase K (PK) digestion and analysis by mass spectrometry. The binding of citrate prevents PK digestion, leading to the differential MS peptide profiling. Compare peptide intensity change under citrate or control treatment, FC: fold change. g 17 proteins that bind three or more peptides were shown. h The binding mode of PFKM and citrate, the PFKM was shown in cartoon and colored in palegreen, and the dominate residues and citrate were shown in sticks and colored in palegreen and orange, respectively. i Microscale thermophoresis assay (MST) was used to detect the binding affinities of citrate with PFKM WT or K617A. Data represent the mean ± SD of three independent experiments. j PFKM-knockout HeLa/H2B-EGFP cells rescued with rPFKM WT or K617A were synchronized for time-lapse microscopic imaging as described in Supplementary Fig. 1b. Representative images of cell cycle progression were shown and the transition from prophase to anaphase onset was highlighted in red rectangles. Mitotic duration from prophase to anaphase onset was measured. PFKM KO + rPFKM WT: n = 88; PFKM KO + rPFKM K617A: n = 96. k PFKM-knockout HeLa cells were rescued with rPFKM WT or K617A. Colony formation assay was performed. (a, b, d, k) Data represent the mean ± SD of three independent experiments (two-tailed Student’s t-test), ns: no significance, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. (c, e, j) Each dot in the scatter plot represents a single cell. Data represent the mean ± SD of the two groups (two-tailed Student’s t-test). Source data are provided as a Source Data file.
To investigate how citrate levels are linked to mitosis, we carried out limited proteolysis-small molecule mapping (LiP-SMap) by incubating citrate and HeLa cell lysates and identified 468 citrate-interacting proteins (Fig. 1fand Supplementary Table. 1). Among them, 17 proteins contain three or more citrate-interacting peptides (Fig. 1g). To narrow down the candidates, molecular docking was used to determine the binding affinity of citrate with these proteins. Two of the proteins (PLEC, AHNAK) were not analyzed further because their structures were not available in current databases. The top 20 docking scores of the proteins with citrate are shown in Supplementary Fig. 1h. The protein of PFKM (citrate binding at D604-K617 peptide) or HSPB1 (citrate binding at F29-R37 peptide) with citrate obtained the lowest docking scores, suggesting that they had strong binding affinity with citrate.
PFKM is one of the isoforms of ATP-dependent 6-phosphofructokinase 1 (PFK1), the rate-limiting enzyme in glycolysis, catalyzing the transfer of a phosphate group from adenosine triphosphate (ATP) to fructose-6-phosphate (F-6-P), producing fructose-1,6-bisphosphate (F-1,6-BP)9. Our LiP-SMap result shows that PFKM contains 3 identified citrate-binding peptides ranging from E131 to K141, or from T175 to R184, or from D604 to K617 (Fig. 1g).
To map the interface where citrate interacts with PFKM, we performed the molecular dynamics (MD) simulation by adding citrate to PFKM tetramer. The conformations sampled from trajectories were used for the density analysis using VMD software10, in which a representative structure of citrate binding was selected from the density cluster. Based on the LiP-SMap data, citrate can bind three possible peptides. Further, MD simulations were used to determine the possible binding sites of citrate with PFKM based on these three citrate-binding possible peptides. The distribution of citrate around PFKM was observed by density analysis based on MD trajectories. From the density analysis, the only possible binding region of citrate is located at the peptide position from D604 to K617. The binding mode of citrate with PFKM is shown in Fig. 1h. To further evaluate the binding modes of the citrate with PFKM, the binding affinities of citrate with PFKM was estimated by using MM-GBSA calculations. The total binding free energies of citrate with PFKM is −8.46 kcal/mol. The dominate residues favoring for the interaction between citrate and PFKM are K617 (−3.69 kcal/mol), Q614 (−1.82 kcal/mol), and V613 (−1.39 kcal/mol) and the binding contribution of K617 is strongest in these residues (Fig. 1h). The interaction between citrate and PFKM was validated by microscale thermophoresis (MST) assay with purified recombinant PFKM and increasing doses of citrate. To specifically disrupt the association of citrate with PFKM, we mutated alkaline amino acid lysine (K) 617 in citrate-binding peptide to alanine (A). MST assay showed that K617A mutation dramatically abrogated the interaction between citrate and PFKM (Fig. 1i), further supporting that K617 is required for the interaction between citrate and PFKM, which is consistent with the previous study11. Of note, the in vitro PFKM glycolytic activity assay with immunoprecipitated Flag-rPFKM WT or K617A indicated that PFKM K617A processed comparable glycolytic activity to PFKM WT (Supplementary Fig. 1i), suggesting that K617A abrogates citrate-dependent regulation of PFKM activity without influencing its glycolytic activity.
To evaluate the contribution of the interaction between citrate and PFKM to mitotic progression, we generated PFKM-knockout HeLa/H2B-EGFP cells with gRNA and rescued the cells with gRNA-resistant (r) PFKM WT or K617A (Supplementary Fig. 1j). Mitotic duration was examined in these cells, showing that the expression of rescued rPFKM K617A prolonged mitotic duration of the cells compared to that of rescued rPFKM WT (Fig. 1j). Consequently, the proliferation of the cells was markedly suppressed by the expression of rescued rPFKM K617A (Fig. 1kand Supplementary Fig. 1k). Collectively, these results demonstrate that high levels of citrate promote the progression of mitosis, requiring its interaction with PFKM.
PFKM interacts with histone H3 at M phase
As PFKM is required for citrate-promoted mitotic progression, we next investigated how PFKM regulates mitotic progression. We depleted PFKM in HeLa/H2B-EGFP cells (Supplementary Fig. 2a) and synchronized the cells at S phase. 14 h after the release from thymidine double block, we observed that PFKM knockout caused more cells accumulated at M phase (Fig. 2a). Moreover, mitotic duration of the cells was also markedly prolonged by PFKM knockout (Fig. 2b), suggesting the important role of PFKM in mitosis.
a Mitotic index of HeLa/H2B-EGFP cells with or without PFKM knockout was calculated at 14 h after release from thymidine double block. b HeLa/H2B-EGFP cells with or without PFKM knockout were synchronized for time-lapse microscopic imaging as described in Supplementary Fig. 1b. Representative images of cell cycle progression were shown and the transition from prophase to anaphase onset was highlighted in red rectangles. Mitotic duration from prophase to anaphase onset was measured according to the time points when corresponding chromatin morphologies occurred. PFKM WT: n = 166; PFKM KO: n = 167. c Histone H3S10ph levels were examined by immunoblotting analysis in HeLa cells with or without PFKM knockout. d HEK293T cells with depletion of PFKM were transfected with empty vector or SFB-rPFKM, a co-immunoprecipitation (co-IP) assay was performed. e Co-IP assay was performed with anti-histone H3 antibody in HeLa cells. f HEK293T cells with depletion of PFKM were transfected with SFB-rPFKM, cells were synchronized as indicated in Supplementary Fig. 2g. I represents interphase; M represents mitosis; 4 h indicates interphase after removal of nocodazole for 4 h, immunoprecipitation was performed. g U87 cells were transfected with empty vector or HA-PFKM, immunoprecipitation was performed. h Bacterial purified His-PFKM were incubated with streptavidin agarose beads treated with biotinylated recombinant mononucleosome (Biotin-Nuc) or not. Pulldown (PD) assay was performed. i The binding mode of PFKM and nucleosomes, and binding free energy contributions are presented in units of kcal/mol. In this conformation, the PFKM is colored in slate blue, and the DNA, H2A, H2B, H3 and H4 in nucleosomes are colored in orange, magenta, green, cyan and wheat, respectively. While the dominate residues or bases in PFKM, DNA, and H3 are shown in sticks. The dominant residues of PFKM and their binding sites of nucleosomes are listed, and all energies are in units of kcal/mol. A: Alanine, K: Lysine, T: Threonine, P: Proline, R: Arginine, Q: Glutamine, V: Valine, H: Histidine, I: Isoleucine, L: Leucine. j PFKM-knockout U87 cells were transfected with rPFKM WT, K66A, or L79A, immunoprecipitation was performed. k Histone H3S10ph levels were examined in PFKM-knockout HeLa cells rescued with rPFKM WT or L79A, immunoblotting experiments were performed. l PFKM-knockout HeLa/H2B-EGFP cells rescued with rPFKM WT or L79A were synchronized for time-lapse microscopic imaging as described in Supplementary Fig. 1b. Representative images of cell cycle progression were shown and the transition from prophase to anaphase onset was highlighted in red rectangles. Mitotic duration from prophase to anaphase onset was measured. PFKM KO + rPFKM WT: n = 183; PFKM KO + rPFKM L79A: n = 157. m PFKM-knockout HeLa cells were rescued with rPFKM WT or L79A, a colony formation assay was performed using these cells. (a, m) Data represent the mean ± SD of three independent experiments (two-tailed Student’s t-test), *p < 0.05, ****p < 0.0001. (b, l) Each dot in the scatter plot represents a single cell. Data represent the mean ± SD of the two groups (two-tailed Student’s t-test). (c–h, j, k) Immunoprecipitation and immunoblotting experiments were performed with the indicated antibodies. Data are representative of three independent experiments. Source data are provided as a Source Data file.
Phosphorylation at a highly conserved residue serine (S) 10 in the histone H3 tail is considered to be a crucial event and also the well-established marker for mitosis12,13,14. Cells with abnormal histone H3S10 phosphorylation (histone H3S10ph) have difficulty in getting through mitosis and exiting mitosis12,14, underscoring the important role of histone H3S10ph in mitosis. We thus examined histone H3S10ph in HeLa cells with or without PFKM knockout (Supplementary Fig. 2b). Surprisingly, PFKM knockout decreased the levels of histone H3S10ph in cells (Fig. 2c), suggesting that PFKM might promote mitotic progression by regulating histone H3S10ph.
Aurora B kinase, a component of the chromosomal passenger complex, is responsible for histone H3S10ph during mitosis15. In order to know whether PFKM regulates histone H3S10ph through Aurora B, we examined the auto-phosphorylation of Aurora B at threonine (T) 232, which occurs once activated, in PFKM-knockout HeLa cells. Immunoblotting analysis showed that PFKM knockout did not influence the phosphorylation of Aurora B T232 (Supplementary Fig. 2c), suggesting that Aurora B activation is not involved in PFKM-promoted histone H3S10ph. To further investigate how PFKM regulates histone H3S10ph, we conducted mass spectrometry analysis of PFKM-associated proteins and identified a group of cell cycle regulators, including Kinesin-like protein KIF11 (KIF11), Histone 3 (H3), DNA-dependent protein kinase catalytic (PRKDC), Importin subunit beta-1 (IMB1), Cofilin-1 (COF1), GTP-binding nuclear protein Ran (RAN), Clathrin heavy chain 1 (CLTC), Tubulin beta-6 chain (TUBB6), Cytoskeleton-associated protein 5 (CKAP5), Serine/threonine-protein phosphatase 2 A 65 kDa regulatory subunit A alpha isoform (2AAA) (Supplementary Fig. 2d). The interaction between PFKM and histone H3 was validated by the Co-immunoprecipitation (Co-IP) experiment in HEK293T cells transfected with S-Flag-streptavidin-binding peptide (SFB) tagged PFKM (SFB-PFKM) (Fig. 2d) and was then confirmed by the pulldown experiment with His-tagged recombinant histone H3 (His-histone H3) in HeLa cells (Supplementary Fig. 2e). Additionally, recombinant PFKM could directly interact with recombinant histone H3 in vitro (Supplementary Fig. 2f). Endogenous PFKM interacted with endogenous histone H3 in HeLa cells (Fig. 2e). Taken together, these results indicate that PFKM interacts with histone H3 in vitro and within cells.
We next confirmed whether these two proteins interact with each other in M phase. HEK293T cells stably expressing SFB-PFKM were synchronized with thymidine double block, followed by the treatment of nocodazole, which arrests the cells in mitosis (Supplementary Fig. 2g). As shown in Fig. 2f, the two proteins’ interaction was markedly increased in M phase and then declined 4 h after M phase when the cells exited mitosis. We further conducted immunofluorescence assays, which also confirmed a significant increase in PFKM-H3 interaction during mitosis compared to interphase (Supplementary Fig. 2h). Of note, PFKM protein levels were not changed during the cell cycle (Supplementary Fig. 2i).
PFKM interacts with nucleosomes, which increases histone H3S10ph and promotes mitotic progression
As the majority of histone H3 is assembled in nucleosomes within cells, we thus tested whether PFKM interacts with histone H3 in the context of nucleosomes. Co-IP experiment showed that PFKM also interacted with other histone proteins, including histone H2A, H2B, and H4, in U87 cells (Fig. 2g), suggesting that PFKM interacts with nucleosomes within cells. Moreover, we performed the pulldown experiment by incubating commercial biotin-labeled recombinant nucleosomes with recombinant PFKM, indicating that PFKM directly interacts with nucleosomes (Fig. 2h).
To understand how PFKM interacts with nucleosomes, we conducted MD simulation with the crystal structures of PFKM (PDB: 4OMT) and nucleosomes (PDB: 3AV2). Based on the experimental result that PFKM interacts with both histone H3 and nucleosomes, three initial complexes molecular models of PFKM and nucleosomes were obtained from the results of ZDOCK16 and PFKM was located near the N-terminal of the H3 in these conformations. To further evaluate the binding modes of PFKM with nucleosomes, the binding affinities of PFKM with nucleosomes were estimated by using MM-GBSA calculations. The total binding free energies of three conformations of PFKM with nucleosomes are –97.76, −120.02, and −126.41, kcal/mol, respectively. Furthermore, the conformation with strongest total binding free energy was shown in Fig. 2i. The dominate residues contributing to the interaction between PFKM and nucleosomes are R622, K621, K564, K66, K557, R301, V83, H52, I84 and L79 of PFKM, and the A1, K4, T6, P16, R17, Q19 and R26 of histone H3 (Fig. 2iand Supplementary Fig. 2j). Among the dominate residues of PFKM, R622, K621, K564, K66, K557 and R301 mainly bind to DNA, V83 and L79 bind to both DNA and histone H3 at the same time, and H52 and I84 interact with histone H3 (Fig. 2i). Collectively, these results indicate that PFKM binds to DNA and histone H3 at the same time, the binding affinity contribution of DNA is stronger than that of histone H3, and the bases on DNA are the dominate part for PFKM binding to nucleosomes.
To identify the residues essential for the interaction between PFKM and nucleosomes, we generated a series of PFKM mutants, in which 10 residues in PFKM potentially contributing to its interaction with nucleosomes were mutated to alanine (A) individually. Mutated residues, including K66A and L79A, dramatically attenuated the interaction between PFKM and histone H3 in HEK293T cells (Supplementary Fig. 2k). Of note, PFKM K66A or L79A exhibited similar metabolic activity to PFKM WT (Supplementary Fig. 2l), suggesting that PFKM K66A or L79A only specifically disrupted the interaction between PFKM and histone H3. Furthermore, in U87 cells, either K66A or L79A mutation greatly suppressed the interaction between PFKM and nucleosome-associated histones, including histone H2B, histone H3, and histone H4 (Fig. 2j), suggesting that both K66 and L79 are required for the association of PFKM with nucleosomes.
To determine whether the interaction between PFKM and nucleosomes is required for histone H3S10ph, we first examined the role of PFKM L79, which is responsible for the interaction between PFKM and nucleosomes via binding DNA as well as histone H3, by rescuing PFKM-knockout HeLa cells with rPFKM WT or L79A (Supplementary Fig. 2m). As shown in Fig. 2k, tumor cells rescued with rPFKM L79A had much lower levels of histone H3S10ph than the cells rescued with rPFKM WT.
To investigate the role of PFKM-dependent histone H3S10ph in mitotic progression, we rescued PFKM-knockout HeLa/H2B-EGFP cells with rPFKM WT or L79A (Supplementary Fig. 2n). Time-lapse imaging showed that mitotic duration was much longer in rPFKM L79A-rescued cells than in rPFKM WT-rescued cells (Fig. 2l). Consequently, the colony formation of HeLa cells was greatly suppressed by rescued expression of rPFKM L79A (Fig. 2m).
PFKM phosphorylates histone H3S10
To further explore how PFKM increases histone H3S10ph by interacting with histone H3, we first tested whether the catalytic activity of PFKM is essential. We constructed the catalytic activity-dead PFKM D166A mutant, in which aspartic (D) 166 was mutated to A (Supplementary Fig. 3a). We rescued PFKM-knockout HeLa cells with rPFKM WT or D166A. PFKM knockout markedly decreased histone H3S10ph, while rescued expression of rPFKM WT, but not that of rPFKM D166A, recovered the levels of histone H3S10ph in PFKM-knockout cells (Supplementary Fig. 3b). Of note, D166A mutation did not influence the interaction between PFKM and histone H3 (Supplementary Fig. 3c).
Given the fact that the catalytic activity of PFKM and its interaction with histone H3, but not the activation of Aurora B, were required for PFKM-dependent histone H3S10ph (Fig. 2k, Supplementary Fig. 2c, and Supplementary Fig. 3b), we wondered whether PFKM phosphorylates histone H3S10 as a protein kinase. To test this hypothesis, we expressed and purified recombinant His-tagged PFKM (His-PFKM) from bacteria (Supplementary Fig. 3d), followed by an in vitro kinase assay by incubating His-PFKM with recombinant histone H3, which showed that PFKM indeed phosphorylated histone H3 (Fig. 3a). We also conducted an in vitro kinase assay using purified PFKM protein and histone H3 substrate, followed by mass spectrometry analysis, showing that three putative phosphorylation sites, including histone H3S10, H3S28, and H3 threonine (T) 45. Among them, histone H3S10 exhibited the highest phosphorylation abundance based on spectral peak intensity (Supplementary Fig. 3e). To test whether PFKM phosphorylates histone H3 at S10, we expressed and purified recombinant His-histone H3 S10A, in which S10 was mutated to A, from bacteria and performed the in vitro kinase assay by incubating histone H3 WT or S10A with recombinant PFKM, which showed that S10A mutation greatly abrogated PFKM-mediated histone H3 phosphorylation, suggesting that S10 is the major residue phosphorylated by PFKM (Fig. 3b and Supplementary Fig. 3f). Furthermore, immunoblotting analysis with anti-histone H3S10ph antibody showed that PFKM phosphorylated histone H3S10 in the in vitro kinase assay (Fig. 3c). Together, these results confirm that PFKM can function as a protein kinase to phosphorylate histone H3 at S10.
a The in vitro kinase assay was performed by incubating bacterial purified recombinant His-PFKM with commercial recombinant histone H3 in the presence of [32P] ATP. b The in vitro kinase assay was performed by incubating His-PFKM with bacterial purified recombinant His-histone H3 WT or S10A in the presence of [32P] ATP. c The in vitro kinase assay was performed by incubating His-PFKM with commercial recombinant histone H3. d, e PFKM-knockout HEK293T cells were transfected with Flag-rPFKM WT or indicated mutants. The in vitro kinase assays were performed by incubating immunoprecipitated Flag-rPFKM WT/ D166A (d) or Flag-rPFKM WT/ L79A (e) with commercial recombinant histone H3. f The in vitro kinase assay was performed by incubating His-PFKM with Biotin-Nucleosomes (Biotin-Nuc). g The catalytic center of PFKM and nucleosomes. In the conformation of catalytic center, H3S10 is shown in stick and colored in cyan, the ATP is shown in stick and colored in yellow, while the Mg2+ ions are shown in sphere and colored in pink, and the fructose-6-phosphate (F-6-P) is shown in line and colored in wheat. a–f Immunoprecipitation and immunoblotting experiments were performed with the indicated antibodies. a–f Data are representative of three independent experiments. Source data are provided as a Source Data file.
We next examined whether the PFKM activity is required for PFKM to phosphorylate histone H3S10. We immunoprecipitated Flag-rPFKM WT or D166A in HEK293T cells and performed the in vitro kinase assay with Flag-rPFKM proteins and recombinant histone H3. PFKM D166A lost the majority of its activity to phosphorylate histone H3S10 compared to PFKM WT (Fig. 3d). Similarly, PFKM L79A, which interacted with less histone H3, also had much weaker ability to phosphorylate histone H3S10 in the in vitro kinase assay (Fig. 3e). Of note, Aurora B was not detected in harshly washed Flag-rPFKM immunoprecipitants (Supplementary Fig. 3g), indicating that the phosphorylation of histone H3 by immunoprecipitated Flag-rPFKM proteins is not due to Aurora B contamination.
In addition, we tested whether PFKM phosphorylates nucleosomes-packed histone H3. We performed the in vitro kinase assay with recombinant PFKM and nucleosomes, and observed that PFKM can phosphorylate histone H3S10 in the context of a DNA-bound nucleosomes (Fig. 3f). This notion was also supported by the results of MD simulations with PFKM and nucleosomes. The conformation of the catalytic reaction is shown in Fig. 3g, indicating that the most favorite binding mode of PFKM with nucleosomes is that S10 of histone H3 is located in the catalytic reaction center of PFKM, which is in agreement with the experimental results. The substrate F-6-P was shown in lines in Fig. 3g. And it shows that both histone H3S10 and F-6-P bind to PFKM at the similar site with the similar binding mode, suggesting that the histone H3S10 can be phosphorylated by PFKM. Collectively, these results demonstrate that PFKM interacts with nucleosomes to directly phosphorylate histone H3S10.
High levels of citrate enhance PFKM-dependent histone H3S10ph by disrupting tetrameric PFKM into dimer
As demonstrated earlier, high levels of citrate interact with PFKM to promote mitotic progression (Fig. 1). We then asked whether citrate regulates PFKM-mediated histone H3S10ph to promote mitosis. As shown in the in vitro kinase assay, citrate enhanced histone H3S10ph catalyzed by PFKM in a dose-dependent manner (Fig. 4a). Consistently, the interaction between PFKM and histone H3 was also enhanced by citrate (Fig. 4b). In addition to these in vitro experiments, we also treated the cells with increasing doses of citrate and observed that the supplementation of exogenous citrate enhanced the interaction between PFKM and histone H3 and therefore induced histone H3S10ph in a dose-dependent manner (Figs. 4c and 4d). Meanwhile, we depleted CS to reduce citrate levels in U87 cells (Supplementary Fig. 4a). The depletion of CS in U87 markedly decreased histone H3S10ph (Supplementary Fig. 4b), while the supplementation of exogenous citrate restored histone H3S10ph in CS-depleted cells (Fig. 4e). These results suggest that high levels of citrate may induce histone H3S10ph by enhancing the interaction between PFKM and histone H3.
a HEK293T cells were transfected with Flag-PFKM. The in vitro kinase assay was performed by incubating immunoprecipitated Flag-PFKM proteins with commercial recombinant histone H3 in the presence of increasing doses of citrate. b HEK293T cells were transfected with empty vector or Flag-histone H3. A co-IP assay was performed in the presence or absence of citrate (0.5 mM). c U87 cells were transfected Flag-PFKM and treated with different doses of citrate. A co-IP assay was performed. d U87 cells were treated with different doses of citrate. Histone H3S10ph levels were examined by immunoblotting analysis. e The U87 cells with CS depletion were supplemented with citrate (2.5 mM) for 24 h. Histone H3S10ph levels were examined by immunoblotting analysis. f The recombinant Sumo-His-PFKM treated with or without citrate was analyzed by gel filtration chromatography, followed by immunoblotting analysis. g PFKM-knockout HEK293T cells were transfected with Flag-rPFKM WT or F639L. The in vitro kinase assay was performed by incubating immunoprecipitated Flag-rPFKM WT or F639L with commercial recombinant histone H3. h PFKM-knockout HEK293T cells were transfected with HA-rPFKM WT or F639L and empty vector or Flag-histone H3. A co-IP assay was performed. i PFKM-knockout HeLa cells were transfected with HA-rPFKM WT or K617A, immunoprecipitation experiments were performed. j PFKM-knockout HeLa cells were rescued with rPFKM WT or K617A. Histone H3S10ph levels were detected. (a–j) Immunoprecipitation and immunoblotting experiments were performed with the indicated antibodies. Data are representative of three independent experiments. Source data are provided as a Source Data file.
Next, we explored how citrate enhances the interaction of these two proteins. PFKM can be regulated by multiple mechanisms, such as polymerization17,18,19,20. We thus examined the influence of citrate on PFKM polymerization. We constructed Sumo-His-tagged PFKM WT and tested whether citrate could disrupt the tetrameric form of PFKM. Gel filtration chromatography analysis confirmed that citrate disrupted tetrameric PFKM into dimeric form (Fig. 4f). Moreover, we performed MD simulations with PFKM tetramer in the absence or presence of citrate, which showed that in the absence of citrate, the R603 of one monomer and E647 from the other monomer, which were in the dimer interface, formed the salt-bridge, resulting in the stabilization of the tetramer. In the presence of citrate, the citrate binds to the dimer interface and breaks the salt-bridge between E647 and R603, thereby decreasing the stability between the dimers and attenuating the formation of tetrameric PFKM (Supplementary Fig. 4c). We next examined the interaction between dimeric PFKM and histone H3. The mutation of phenylalanine (F) 649 to L (F649L) has been shown to disrupt tetrameric PFKP into its dimeric form21. Sequence alignment shows that F639 residue of PFKM is conserved across the three isoforms of PFK1: PFKM, PFKP (platelet type), and PFKL (liver type) (Supplementary Fig. 4d). We constructed Sumo-His-tagged PFKM WT or PFKM F639L and tested whether F639L could also disrupt the tetrameric form of PFKM. Gel filtration chromatography analysis confirmed that PFKM F639L mainly existed as the dimeric form compared to PFKM WT (Supplementary Fig. 4e). And, compared to PFKM WT, PFKM F639L had much lower metabolic activity (Supplementary Fig. 4f), mainly due to dimeric PFKM has low affinity for its substrate fructose-6-phosphate22. In contrast, PFKM F639L catalyzed more histone H3S10ph than PFKM WT (Fig. 4g). Co-IP experiment showed that compared to PFKM WT, PFKM F639L interacted with more histone H3 (Fig. 4h). These results indicate that citrate enhances the interaction between PFKM and histone H3 by disrupting tetrameric PFKM into dimeric form, thereby increasing PFKM-dependent histone H3S10ph.
In addition, we performed MD simulations to analyze whether the dimeric forms of PFKP and PFKL have the kinase activity. The dimer of PFKL were obtained from the tetramer of PFKL bound ADP (PDB code 7LW1), which the Mg ions was constructed by align to the 4XYJ and the ADP was modified to ATP. PLIN2 was selected as the substrate of PFKL according to the Meng et al. results23. According to the results of Cao et al. 24, the dimer of PFKP from mouse was constructed based on the PFKP bound ATP and Mg ions from human (PDB code 4XYJ), and Lin41 was selected as the substrate of PFKP. Histone H3 was selected as the substrate of dimeric PFKM, which was used as the contrast. The initial binding mode of PFKM, PFKP, and PFKL with peptide substrate was constructed by ZDOCK, which docked the substrate to the PFKM, PFKP, and PFKL. The binding mode showed in Supplementary Fig. 4g. Our MD simulation results showed that the binding free energies of PFKL and PFKM with substrate peptide were −32.65, and −32.96 kcal/mol, respectively, and the binding free energy of PFKP was −50.57 kcal/mol, and the serine of the substrate all located in the catalytic activity site.
These results suggest that the PFKL and PFKP dimers can stably bind their substrates, which had the similar binding affinity as PFKM dimer. Notably, both dimeric forms maintained critical catalytic geometry with substrate peptide and with the strong binding free energies, and serine residues properly oriented within active sites, suggesting that the dimeric PFKP or PFKL might potentially function as protein kinases.
We also performed MD simulations to analyze whether the monomeric forms of PFKM and PFKL would similarly have kinase activity. Our MD simulations showed that monomeric PFKM could bind to histone H3 (−32.03 kcal/mol). The monomeric PFKL can bind PLIN2 and have the kinase activity, and the monomeric PFKM binds histone H3 and had the similar binding free energies, so we speculated that monomeric PFKM might have similar functions with monomeric PFKL. The binding mode of monomeric PFKL and PFKM was shown in Supplementary Fig. 4h. Notably, both monomeric forms maintained critical catalytic geometry with substrate and with the strong binding free energies, and serine residues properly oriented within active sites, it means that the both monomeric forms might have the similarly function as the protein kinase.
To further confirm that the interaction between citrate and PFKM is required for PFKM-dependent histone H3S10ph, we performed Co-IP experiment in PFKM-knockout HeLa cells rescued with rPFKM WT or K617A. The result showed that PFKM K617A, which disrupts citrate binding, had much weaker interaction with histone H3 than PFKM WT (Fig. 4i). Rescued expression of PFKM K617A markedly decreased histone H3S10ph levels in the cells compared to that of PFKM WT (Fig. 4j).
Collectively, these results demonstrate that citrate disrupts tetrameric PFKM into dimer and dimeric PFKM strongly interacts with histone H3, thereby increasing histone H3S10ph.
Citrate-regulated histone H3S10ph is required for tumorigenesis
To investigate the role of citrate-regulated histone H3S10ph in tumorigenesis, we first examined the interaction between PFKM and histone H3 in U87 glioma cells and RKO colorectal cancer cells. As shown in Supplementary Fig. 5a, the interaction between endogenous PFKM and histone H3 could be detected in these cancer cells. However, compared to PFKM WT, PFKM K617A exhibited much less interaction with histone H3 in these cells (Fig. 5a and Supplementary Fig. 5b). Rescued expression of PFKM K617A markedly decreased histone H3S10ph levels and consequently suppressed the proliferation of U87 and RKO cells, compared to that of PFKM WT (Fig. 5b,c). Moreover, we intracranially injected PFKM knockout U87/EGFRvIII cells (luciferase-expressing U87 cells stably expressing the active mutant of EGFR) rescued with rPFKM WT or K617A into randomized athymic nude mice. Bioluminescence imaging of mice indicated that tumor cells expressing rPFKM K617A developed much smaller tumor in mice than the cells expressing rPFKM WT (Fig. 5d). The dissected brain tissues were validated to contain tumors by hematoxylin and eosin (H&E) staining (Fig. 5e).
a PFKM-knockout U87 and RKO cells were transfected with HA-rPFKM WT or K617A, co-IP experiments were performed. b PFKM-knockout U87 and RKO cells were rescued with rPFKM WT or K617A. Histone H3S10ph levels were detected. c PFKM-knockout U87 and RKO cells were rescued with rPFKM WT or K617A. Colony formation assays were performed. d PFKM-knockout luciferase-expressing U87/EGFRvIII cells were rescued with SFB-rPFKM WT or K617A. The cells were intracranially injected into randomized nude mice (five mice per group), 12 and 15 days after inoculation, bioluminescence imaging was performed and representative images were shown (d, left panel). Relative luciferase intensities were normalized to those of mice injected with cells rescued with SFB-rPFKM WT at days 12 (d, right panel). Data represent the mean ± SD of luciferase intensities obtained from five mice (two-tailed Student’s t-test). e Representative images of H&E -stained coronal brain sections and tumor boundaries were shown (e left panel). Tumor areas in H&E-stained sections were calculated and normalized to those of the mice injected with cells rescued with SFB-rPFKM WT (e right panel). Data represent the mean ± SD (five mice). Student’s t-test (two-tailed) was used for two group comparisons. f PFKM-knockout U87 and RKO cells were rescued with rPFKM WT or L79A. Histone H3S10ph levels were detected. g PFKM-knockout U87 and RKO cells were rescued with rPFKM WT or L79A. Colony formation assays were performed. h PFKM-knockout luciferase-expressing U87/EGFRvIII cells were rescued with SFB-rPFKM WT or L79A. A colony formation assay was performed. i PFKM-knockout luciferase-expressing U87/EGFRvIII cells were rescued with SFB-rPFKM WT or L79A. The cells were intracranially injected into randomized nude mice (six mice per group). 10 days after inoculation, bioluminescence imaging was performed and representative images were shown (i, left panel). Relative luciferase intensities were normalized to those of mice injected with cells rescued with SFB-rPFKM WT at days 10 (i, right panel). Data represent the mean ± SD of luciferase intensities obtained from six mice (two-tailed Student’s t-test). j Representative images of H&E-stained coronal brain sections and tumor boundaries were shown (j, left panel). Tumor areas in H&E-stained sections were calculated and normalized to those of the mice injected with cells rescued with SFB-rPFKM WT (j, right panel). Data represent the mean ± SD (six mice). Student’s t-test (two-tailed) was used for two group comparisons. k, l RKO cells with PFKM knockout and rescued with rPFKM WT or L79A were injected subcutaneously in nude mice, n = 10. Tumor volume (mm3) at days 14 to 25 after subcutaneous injection were shown (k). Subcutaneous tumor growth (l, left panel) and tumor weight (l, right panel) at days 25 after subcutaneous injection were shown. Data represent the mean ± SD of the two groups (two-tailed Student’s t-test). a, b, f Immunoprecipitation and immunoblotting experiments were performed with the indicated antibodies. Data are representative of three independent experiments. c, g, h Data represent the mean ± SD of three independent experiments (two-tailed Student’s t-test), ns: no significance, **p < 0.01, ***p < 0.001, ****p < 0.0001. Source data are provided as a Source Data file.
Furthermore, we rescued PFKM-knockout U87 and RKO cells with rPFKM WT or L79A (Supplementary Fig. 5c). As shown in Fig. 5f, tumor cells rescued with rPFKM L79A had much lower levels of histone H3S10ph than the cells rescued with rPFKM WT. The ability of colony formation of U87 and RKO cells was greatly suppressed by rescued expression of rPFKM L79A (Fig. 5g). To confirm the role of PFKM-dependent histone H3S10ph in tumor growth in vivo, we knocked out PFKM in U87/EGFRvIII cells, and rescued the cells with SFB-rPFKM WT or L79A (Supplementary Fig. 5d). Similarly, tumor cells rescued with rPFKM L79A had much lower levels of histone H3S10ph than the cells rescued with rPFKM WT (Supplementary Fig. 5e). Soft agar assay showed that rescued expression of rPFKM L79A dramatically decreased the number of colonies in U87/EGFRvIII cells (Fig. 5h). We next intracranially injected these cells into randomized athymic nude mice. Bioluminescence imaging of mice indicated that tumor cells expressing rPFKM L79A developed much smaller tumor in mice than the cells expressing rPFKM WT (Fig. 5i). The dissected brain tissues were validated to contain tumors by H&E staining (Fig. 5j). In addition, we also confirmed the role of PFKM-dependent histone H3S10ph in colorectal cancer (CRC) development by subcutaneously implanting athymic nude mice with PFKM-knockout RKO cells rescued with rPFKM WT or L79A. Tumor cells rescued with rPFKM WT elicited a rapid tumor growth in 9 out of 10 mice, while the cells rescued with rPFKM L79A completely failed to do so (Fig. 5k, l).
Collectively, these results clearly demonstrate that citrate-regulated PFKM-dependent histone H3S10ph is required for tumor cell proliferation and tumorigenesis.
Citrate-regulated histone H3S10ph is also required for T cell proliferation
To determine whether citrate-regulated histone H3S10ph is also required for the proliferation of normal cells, we treated human peripheral blood mononuclear cells (PBMCs) pre-activated by α-CD3/CD28 with increasing doses of citrate. 14 h after the release from thymidine double block, we observed that citrate decreased the number CD8+ T cells at M phase in a dose-dependent manner (Fig. 6a), suggesting the shortened mitotic duration. And citrate therefore increased their proliferation in a dose-dependent manner (Fig. 6b). Moreover, we depleted PFKM in human PBMCs (Supplementary Fig. 6a) and rescued the cells with rPFKM WT or K617A (Supplementary Fig. 6b), or rescued the cells with rPFKM WT or L79A (Supplementary Fig. 6c). These cells were then activated by α-CD3/CD28. Rescued expression of rPFKM K617A or L79A dramatically inhibits the proliferation of CD8+ T cells compared to that of rPFKM WT (Fig. 6c, d), underscoring the role of citrate-regulated mitosis in the proliferation of CD8+ T cells.
a CD8+ T cells were loaded on poly-Lysine coated dish before treated with thymidine and different doses of citrate. Mitotic index of these cells was calculated at 14 h after release from thymidine double block. b CD8+ T cells were treated with different doses of citrate for 48 h, and the proliferation of CD8+ T cells was examined. c PFKM-depleted PBMCs were transfected with rPFKM WT or K617A, cells were treated with α-CD3/CD28 for 48 h, and the proliferation of CD8+ T cells was examined. d PFKM-depleted PBMCs were transfected with rPFKM WT or L79A, cells were treated with α-CD3/CD28 for 48 h, and the proliferation of CD8+ T cells was examined. e Co-IP assays were performed with anti-histone H3 antibody in U87 and activated T cells. f Citrate levels were detected in U87 and activated T cells by mass spectrometry. g CD19 CAR-T cells were loaded on poly-Lysine coated dish before treated with thymidine and different doses of citrate. Mitotic index of these cells was calculated at 14 h after release from thymidine double block. h CD19 CAR-T cells were treated with different doses of citrate for 48 h, the proliferation of CD19 CAR-T cells was examined. i CD19 CAR-T cells were treated with different doses of citrate for 48 h, and CD19 CAR-T cells were co-cultured with CD19-expressing U87 cells for 24 h. U87 cells viability was measured by Cell Viability Assay. j CD19 CAR-T cells were treated with different doses of citrate for 48 h, and CD19 CAR-T cells were co-cultured with CD19-expressing RKO cells for 24 h. RKO cells viability was measured by Cell Viability Assay. a–d, f–j Data represent the mean ± SD of three independent experiments (two-tailed Student’s t-test), ns: no significance, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. (e) Immunoprecipitation and immunoblotting experiments were performed with the indicated antibodies. Data are representative of three independent experiments. Source data are provided as a Source Data file.
Through the analysis of glioma from the Chinese Glioma Genome Atlas (CGGA) database and colorectal cancer from The Cancer Genome Atlas (TCGA) database, we observed that PFKM expression was markedly increased in tumor patients (Supplementary Fig. 6d, e). Similarly, compared to the activated T cells, tumor cells had much higher expression of PFKM and meanwhile more interaction between PFKM and histone H3 was observed in tumor cells (Fig. 6e). Additionally, tumor cells had higher intracellular citrate levels than activated T cells (Fig. 6f). These results implicate the possible therapeutic window to treat tumor cells without obviously comprising T cell function.
In addition, we tested whether citrate incubation can boost the function of CAR-T. We incubated CD19 CAR-T cells with increasing doses of citrate and observed that citrate similarly shortened mitotic duration of CD19 CAR-T cells and promoted their proliferation in a dose-dependent manner (Fig. 6g, h). Furthermore, we pre-incubated CD19 CAR-T cells with increasing doses of citrate and then co-cultured these CAR-T cells with CD19-expressing tumor cells. Pre-incubation of citrate markedly improved the killing effect of CAR-T cells on tumor cells (Figs. 6i, j), suggesting that citrate incubation may be an effective strategy to boost the anti-tumor capability of CAR-T cells.
Discussion
Activity of metabolism is a critical factor which governs the fate of cells, such as the survival, proliferation and differentiation. Emerging evidence indicates that metabolic signals, including nutrient availability, biosynthetic intermediates and energetic balance, are coupled to cell cycle progression25,26. However, the mechanism by which these metabolic signals are integrated to the core machinery of cell cycle remains to be elucidated. In this study, we provide the evidence to support PFKM as a citrate sensor that links the levels of citrate to mitotic progression of the cells. In M phase, increased levels of citrate bind to and disrupt tetrameric PFKM to dimer. Dimeric PFKM interacts with and phosphorylates histone H3 at S10, thereby accelerating mitotic progression and the proliferation of tumor cells as well as T cells (Fig. 7).
High levels of citrate bind to PFKM and disrupt tetrameric PFKM into its dimeric form. Dimeric PFKM interacts with and phosphorylates histone H3 at S10, thereby promoting mitotic progression and cell proliferation. (Created in BioRender. Lin, P. (2025) https://BioRender.com/7z04413).
Emerging evidence shows that metabolism is tightly controlled by cell cycle regulator to meet the requirements for sufficient energy and biomass27. The inactivation of APC/C-Cdh1 in mid-G1 to early S phase activates PFKFB3 resulting in enhancement of glycolysis at the restriction period control, allowing further G1/S transition28,29. Mitochondria-targeted Cyclin B1/Cdk1 phosphorylates complex I of the respiratory chain with enhanced ATP generation, which provides cells with efficient bioenergy for G2/M transition30. However, it remains elusive how metabolic activity is sensed to coordinate with cell cycle progression. Here we uncover that the levels of citrate can be sensed by PFKM and subsequently translated into the levels of histone H3S10ph, thereby accelerating mitotic progression and promoting tumor cell proliferation as well as T cell proliferation.
Citrate is an important intermediate in the tricarboxylic acid (TCA) cycle that is required to support both mitochondrial bioenergetics and cytosolic macromolecular synthesis31,32. After its synthesis in mitochondria from acetyl-CoA and oxaloacetate (OAA) by citrate synthase, citrate becomes a substrate in the TCA cycle, and its subsequent complete oxidation provides the major source of cellular ATP production. On the other hand, citrate can also be transported out of the mitochondria and then broken down into acetyl-CoA for fatty acid and sterol biosynthesis in cytosol33,34. Intriguingly, when glucose contributes sufficiently to citrate synthesis, citrate also acts as a feedback inhibitor of PFK1, thereby slowing glycolysis and linking TCA cycle activity to glycolytic flux35,36. In contrast, our study reveals a distinct mechanism by which citrate functions through its interaction with PFKM. Specifically, citrate binding induces the transition of PFKM from a tetrameric to a dimeric state. This dimerized form of PFKM facilitates cell mitosis through non-metabolic functions, highlighting a novel regulatory role of citrate beyond its classical metabolic effects. Of note, during mitosis, glycolysis is suppressed, and PFK1 exhibits relatively low metabolic enzyme activity2. It has also been shown that tumor cells have much higher levels of citrate than normal cells, which may contribute to their enhanced proliferative capacity37,38.
PFKM has been recently shown to play the important role in many human cancers. Increased expression or glycolytic activity of PFKM promotes the development of hepatocellular carcinoma (HCC), ovarian cancer, and esophageal squamous cell carcinoma (ESCC)39,40,41,42. In addition, PFKM-depleted tumors display the decreased infiltration of PD-1+ CD8+ T cells and compromised tumor growth in a preclinical model of pancreatic ductal adenocarcinoma43. Consistent with these studies, we also observed the protumoral role of PFKM in both glioma and CRC. In contrast to its glycolytic function, however, we demonstrate here that PFKM functions as a protein kinase to phosphorylate histone H3S10, thereby promoting mitotic progression and tumor cell proliferation. Intriguingly, this PFKM-mediated histone H3 phosphorylation can be regulated by the levels of citrate, thereby linking cell cycle progression to metabolic state of the cell. Notably, 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 4, an isoform of PFK2 which synthesizes fructose-2,6-bisphosphate, a potent stimulator of glycolysis, has also been reported to phosphorylate SRC-3 as a protein kinase and enhance its transcriptional activity, thereby promoting aggressive metastatic breast cancer44.
Histone H3S10ph is a conserved epigenetic hallmark of chromatin condensation during mitosis across eukaryotes, serving as a canonical biomarker for these processes12,13,45. The Aurora B kinase is the predominant kinase that phosphorylates histone H3S10 in response to growth signals. This modification triggers HP1 dissociation from heterochromatin, thereby licensing chromosome segregation46,47. During the cell cycle, energy, nutrients, and growth signals work together to ensure that the cell cycle runs properly48. Our study unveils a novel metabolic checkpoint mechanism mediated by PFKM. We show that PFKM can sense citrate levels and phosphorylates histone H3 as a protein kinase, which promotes mitotic progression and cell proliferation, integrating metabolic signals into the cell cycle progression. Basing on the literature (Aurora B kinase) and the results in our study (Figs. 1j and 2l), we believe that the phosphorylation of histone H3S10 by these two proteins are both required for the progression of mitosis, coordinating growth signals and metabolic signals during cell cycle progression.
Emerging evidence highlights that metabolic enzymes can perform noncanonical functions critically involved in tumorigenesis49. As a representative example, PFK1 exists as three tissue-specific isoforms: PFKL, PFKP, and PFKM50. Intriguingly, these isoforms exhibit distinct moonlighting functions beyond their canonical catalytic roles. Under glucose deprivation, PFKL undergoes a functional switch through tetramer-to-monomer transition. While its enzymatic activity is inactivated in monomeric form, PFKL acquires a novel role in facilitating lipolysis by mediating lipid droplet-mitochondria tethering to promote fatty acid transfer and β-oxidation23. Similarly, PFKP demonstrates context-dependent functional plasticity during mouse embryonic stem cell (mESC) differentiation. It acts as a protein kinase to phosphorylate the developmental regulator Lin41 on serine residue. This post-translational modification stabilizes Lin41 by inhibiting its autoubiquitination and subsequent proteasomal degradation, ultimately promoting endodermal lineage specification through Lin41-mediated mRNA destabilization of ectodermal markers24. Notably, our study reveals a unique mechanism for PFKM during mitosis. Unlike PFKL’s monomer-dependent protein kinase activity, PFKM exerts its noncanonical function as a dimeric protein kinase independent of its metabolic enzymatic activity. This structural distinction and functional divergence among PFK isoforms underscore their context-dependent regulatory complexity.
Interestingly, we observed that compared to the activated T cells, tumor cells had much higher expression of PFKM, and meanwhile more interaction between PFKM and histone H3. Additionally, tumor cells had higher intracellular citrate levels than activated T cells. These results implicate the possible therapeutic window to treat tumor cells by targeting excess PFKM expression or high citrate levels without obviously comprising T cell function.
PFKM is regulated by multiple sets of mechanism during cancer progression. The transcription of PFKM is activated by ZEB1 or nuclear factor kappa B (NF-κB) to promote aerobic glycolysis, thereby facilitating the progression of HCC and ESCC41,42. In addition, the tetrameric form of PFKM is stabilized by NOS1-mediated S-nitrosylation of cystine 351, which abrogates the negative feedback regulation of its downstream metabolic intermediates and enhances aerobic glycolysis, thereby promoting tumor growth and metastasis of ovarian cancer cells39. In our study, we illustrate that the protein kinase activity of PFKM is regulated by the levels of citrate, in which citrate disrupts tetrameric PFKM into its dimeric form and increases PFKM’s interaction with histone H3, thereby facilitating PFKM-dependent histone H3S10ph and promoting tumor cell proliferation.
As shown in Supplementary Fig. 3e, in addition to histone H3S10, two more putative phosphorylation sites, including H3S28 and H3T45, could be phosphorylated by PFKM. Histone H3S28 phosphorylation has previously shown to function in DNA damage repair or chromatin compaction associated with cell mitosis and meiosis51,52. In contrast, AKT phosphorylates histone H3T45 upon DNA damage, and AKT-mediated phosphorylation of histone H3T45 regulates transcriptional termination53. Histone H3T45 phosphorylation was also associated with apoptosis in HL60 cells and purified human neutrophils54. These two putative phosphorylation sites (S28 and T45) on histone H3 by PFKM might also implicate the potential role of PFKM in DNA damage, transcriptional termination and apoptosis, which definitely requires further investigation.
Aurora B-mediated histone H3S10 phosphorylation during mitosis has been well established47,55. Notably, histone H3S10 phosphorylation exhibits kinase promiscuity, with additional enzymes including ERK and p38 demonstrating catalytic capacity under specific cellular conditions56. Despite these advances, the potential crosstalk between metabolic states and histone phosphorylation has been largely unexplored. Our study directly addresses the frontier by identifying PFKM—conventionally known as a glycolytic enzyme—as a novel metabolic sensor-protein kinase capable of phosphorylating histone H3S10 to accelerate mitotic progression. This discovery establishes an unprecedented mechanistic link between cellular metabolism and mitotic histone modification.
Methods
Antibodies
The rabbit polyclonal anti-PFKM (55028-1-AP) and rabbit polyclonal anti-Aurora B (39262) were purchased from Proteintech. Rabbit polyclonal AurB (Thr232) (phosphor-specific) (TA325250) was purchased from Origene. Rabbit polyclonal anti-Histone H3 (ab1791) was purchased from Abcam. Rabbit polyclonal anti-phospho-Histone H3 (Ser10) (9701), rabbit monoclonal anti-HA (3724) and rabbit monoclonal anti-Histone H2B (12364S) were purchased from Cell Signaling Technology. Rabbit monoclonal anti-Flag (20543-1-AP) and mouse monoclonal anti-Actin (60008-1-lg) were purchased from Proteintech. Rabbit monoclonal anti-Histone H2A (A3692) was purchased from Abclonal. Mouse monoclonal antibody against Tubulin (T9026) and rabbit monoclonal anti-Histone H4 (05-858) were purchased from Sigma. FITC anti-human CD8 (980908) and Alexa Fluor® 700 anti-human CD8 Antibody (344724) were purchased from BioLegend.
Reagents
Puromycin (540411-100MGCN) and hygromycin (400053) were bought from EMD Biosciences. Cocktail (B14001) was bought from Selleck. Blasticidin S (60218ES10) was bought from Yeasen. ATP (tlrl-atp) was purchased from InvivoGen. [γ−32P] ATP (NEG502A) was purchased from PerkinElmer. Citrate (6132-04-3) was purchased from SHANGHAI EXPERIMENT REAGENT. Fructose-6-phoshphate (F3627) and anti-Flag M2 affinity gel (A2220) were purchased from Sigma. Recombinant histone H3 (M2507) was bought from New England Biolabs. Poly-Lysine (A3890401) was purchased from Invitrogen. Anti-HA agarose (AGM90054) was purchased from AOGMA. Ni-NTA resin (L00223I-50) and Glutathione resin (L00206) were purchased from GeneScript. Dynabeads Protein G (10003D) and Dynabeads Streptavidin Trial Kit (65801D) were purchased from Invitrogen. ImmunoCult™ Human CD3/CD28 T Cell Activator (10971) was purchased from STEMCELL. CellTrace CFSE Cell Proliferation Kit (65-0850-84) was purchased from Invitrogen.
Cell culture
HeLa cervical cancer cells, U87 glioma cells and RKO colorectal cancer cells, were maintained in Dulbecco’s modified Eagle’s medium (DMEM, Sigma-Aldrich, D5796) supplemented with 10% fetal bovine serum (FBS, ExCell Bio, FSD500) and antibiotics (Gibco, 15140–122). HEK293T kidney cells were maintained in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% bovine calf serum and antibiotics. Cells were authenticated using the short tandem repeat (STR) method and were incubated in 5% CO2 at 37 °C. Transfections were performed using PolyJet transfection reagent (SL100688), LipoFiterTM Liposomal Transfection Reagent (HB-LF-1000) and GeticoFectTM RNAiMax (130202).
Animal studies
5-week-old or 6-week-old female athymic nude mice were obtained from Lingchang Biotech (Shanghai, China). Littermates of the same sex were randomly assigned to experimental groups. The use of mice was in compliance with ethical regulations and was approved by the institutional review board at the Institute of Biochemistry and Cell Biology (SIBCB).
DNA constructs and mutagenesis
PCR-amplified human PFKM was cloned into pCDH-SFB, pCDH-HB, pCDH-Flag, pcDNA3-HA, pLVX-IRES-zsGreen, pLVX-IRES-mCherry, pCold I, pGEX-KG, and pET-28as vectors. PCR-amplified human histone H3 was cloned into pcDNA3-Flag and pCold I vectors. rPFKM K617A, L79A, H52A, K66A, V83A, I84A, R301A, K557A, K564A, K621A, R622A, D166A, F639L, and histone H3 S10A mutant were made using the QuickChange site-directed Mutagenesis kit (Stratagene, La Jolla, CA, USA). PFKM-F: ATGACCCATGAAGAGCACCA, PFKM-R: TTAGACGGCAGCTTCCCC, Histone H3-F: ATGGCCCGAACCAAGCAGAC, Histone H3-R: TTAAGCTCTCTCTCCCCGTATCCGG.
For gRNA cloning, the lentiCRISPRv2 vector was digested with BsmBI and ligated with BsmBI-compatible pre-annealed oligo-nucleotides. The following sequence was used for CRISPR-knockout strategy: PFKM gRNA: AATGAGGATCTTACCCACAG. The siRNA sequence used for non-targeting control (siNC): 5′-UUCUCCGAACGUGUCACGUTT-3′. The siRNA sequences used for targeting human CS (siCS−1 and siCS−2): 5′-GCCCUCAACAGUGAAAGUATT-3′ (siCS−1), 5′-GCAGGGUAAAGCCAAGAAUTT-3′ (siCS−2). The siRNA sequences used for targeting human PFKM: 5′-UCCAGAAAGCAGGUAAGAUTT-3′ (siPFKM).
Lentiviral transduction
HEK293T cells were co-transfected with plasmid together with lentivirus-packaging vectors pMD2.G and psPAX2. 24 and 48 (h) after transfection, the culture supernatants containing the released viral particles were collected and added with polybrene at a final concentration of 8 μg/mL. After 12–24 h, the media were replaced with fresh viral-free medium to allow further growth until use.
Quantitative real-time PCR
Total RNA was extracted with Trizol reagent (Invitrogen™, 15596018CN). cDNA was prepared by a ReverTra Ace qPCR RT Master Mix Kit (Toyobo, FSQ-301) using oligonucleotide (dT) and random primers. Quantitative Realtime PCR analyses were carried on Roche System (lightCycler® 96). Data were normalized to expression of a control gene (β-actin) for each experiment. Primer sequences of human CS were 5’-CACAGGGTATCAGCCGAACCAA-3’ (forward) and 5’-CCAATACCGCTGCCTTCTCTGT-3’ (reverse); β-actin were 5’-AGAGCTACGAGCTGCCTGAC-3’ (forward) and 5’-AGCACTGTGTTGGCGTACAG-3’ (reverse).
Purification of recombinant proteins and gel filtration
PFKM were purified from E. coli as followings: the vectors expressing PFKM were used to transform BL21. Protein expression was induced in the presence of 0.5 mM IPTG at 16 °C for 20 h (pCold I) or at 30 °C for 6 h (pGEX-KG and pET-28as). Cell pellets were gathered and sonicated in lysis buffer before centrifugation at 13400 g for 60 min (min) (4 °C). Supernatant were incubated with Ni-NTA resin or glutathione agarose for 2–3 h (4 °C). After that, beads were washed using lysis buffer containing 50 mM imidazole or not then stored in −80 °C or eluted with suitable buffer.
pET-28as-PFKM WT or K617A or F639L were purified using lysis buffer (20 mM Tris-HCl, pH 7.5, 50 mM potassium phosphate, 1 mM 2-mercaptoethanol, 10% glycerol, 10 mM imidazole and protease inhibitor cocktail) and elution buffer (lysis buffer with 200 mM imidazole). Protein was concentrated using an Amicon Ultracel-10K Centrifugal Filter Unit and buffer exchanged into FPLC buffer (20 mM HEPES, pH 7.5, 100 mM KCl, 1 mM TCEP, 1 mM ATP, 1 mM MgCl2, and 5% glycerol), followed by gel filtration experiment. PFKM was passed over a Superose 6 10/300 GL column or Superdex 200 Increase 10/300 GL, and fractions were collected, followed by western blot analysis.
His-histone H3 WT or S10A were expressed in E. coli and the proteins were purified on Ni-NTA resin under 8 M urea denaturing conditions, according to the manufacturer’s instructions (Qiagen). Renatured the proteins using dialysis buffer (2 mM EGTA, 20 mM 2-mercaptoethanol, 1 mM phenylmethylsulfonyl fluoride, 20 mM Tris-HCl, pH 8.0) at 4 °C overnight.
PFKM enzymatic activity assay
PFKM-knockout HEK293T cells were transfected with Flag-rPFKM WT or mutant. Flag-rPFKM WT and mutant proteins were immunoprecipitated for the measurement of enzymatic activity. The assays were performed in 50 mM Tris-HCl, pH 8.0, containing 10 mM MgCl2, 1 mM DTT, 200 μM Fructose-6-Phosphate and 20 μM ATP, at 30 °C for 30 min. PFKM activity was determined by monitoring the luminescence signal that accompanies the remaining ATP using the Kinase-Glo assay (Promega, V3771).
In vitro kinase assay
For phosphorylation reactions, recombinant His-PFKM from E. coli or immunoprecipitated Flag-PFKM from cell lysates of HEK293T cells were mixed with histone H3 or biotinylated recombinant mononucleosomes in kinase buffer (10 mM Tris-HCl, pH 7.5, 5 mM MgCl2, 20 mM NaCl, 1 mM EDTA, 2 mM DTT) in the presence of 0.5 mM ATP, or in kinase buffer (5 mM MOPS, pH 7.2, 5 mM MgCl2, 1 mM EGTA, 0.4 mM EDTA, 0.05 mM DTT, 2.5 mM β-glycerophosphate) with 50 μM ATP and 0.02 mCi/mL [γ−32P]ATP for 1 h at 30 °C. Terminated the reaction by adding SDS loading. Incorporation of γ−32P or P into histone H3 was visualized by SDS-PAGE and autoradiography. Besides, citrate (pH 7.4) was added to reaction 30 min prior to substrate H3.
Immunoprecipitation and immunoblotting analysis
Extraction of proteins with co-IP buffer (which is mild to maintain the interaction between proteins) from cultured cells were used for co-IP assay. Extraction of proteins with IP buffer (which is harsh and used for IP to get cleaner target proteins) from cultured cells were used for enzyme activity assay or in vitro kinase assay. Co-IP buffer: 50 mM Tris-HCl (pH 7.5), 1% Triton X-100, 0.01% SDS, 150 mM NaCl, 1 mM dithiothreitol, 0.5 mM EDTA, 0.1 mM sodium orthovanadate, 0.1 mM sodium pyrophosphate, 0.1 mM sodium fluoride, and protease inhibitor cocktail. IP buffer: 0.1% SDS and other compositions are the same as the co-IP buffer. The pH of used citrate was 7.4. Flag-tagged protein and HA-tagged protein were immunoprecipitated with anti-Flag M2 affinity gel and anti-HA agarose, respectively. For immunoblotting, cells were lysed in cold RIPA buffer for 30 min supplemented with 1% protease cocktail, pipetting up and down every ten minutes, following centrifugation at 12,000 g for 20 min. The pellets were discarded and supernatants were boiled in the loading buffer at 95 °C for 10 min. Protein samples were separated on an SDS-PAGE gel and transferred to a nitrocellulose membrane, followed by incubation with primary antibodies (dilution ratios depend on the specific antibody) overnight. Membrane was then washed with TBST three times, and incubated with secondary antibodies (1: 5000). After secondary antibody incubation, membranes were washed in TBST, then were added for visualization using enhanced chemiluminescence (ECL)57.
MicroScale Thermophoresis (MST) assay
Binding affinities of citrate with the purified proteins His-PFKM WT or K617A were measured by using the Monolith NT.115 (Nano Temper Technologies). The proteins were fluorescently labeled according to the manufacturer’s procedure and kept in the MST buffer (100 mM Tris-HCl, pH 6.0, 0.05% (v/v) Tween-20) at a concentration of 1 µM. For each assay, the labeled protein was mixed with the same volume of citrate at serially diluted concentrations at room temperature. The samples were then loaded into capillaries (Nano Temper Technologies) and measured at 25 °C by using 20% LED power and 40% MST power. Each assay was repeated three times. Data analyses were performed using MO. Affinity Analysis v.2.2.4 software. There is 95% confidence, that the Kd value is within the given range.
LiP-SMap analysis
HeLa cells in lysis buffer (100 mM HEPES pH 7.5, 150 mM KCl, 1 mM MgCl2) were mechanically ground and centrifugated 20,000 g for 5 min at 4 °C. The supernatant was retained and protein concentrations were measured with a BCA assay kit (Thermo Fisher Scientific, 23,228). Cell lysates containing about 100 μg of total protein sample were incubated with citrate (2.5 mM) or control for 10 min at 25 °C. Proteinase K was added simultaneously to each sample for the following 5 min. The sample was heated at 98 °C for 3 min, and then the digestion reaction was stopped with 5 % sodium deoxycholate. The sample was then reheated at 98 °C for 3 min.
Protein fragments from the limited proteolysis step were reduced with TCEP (final concentration is 5 mM) (Thermo Scientific) for 40 min at 37 °C and then alkylated by incubating 30 min at 25 °C with 20 mM iodoacetamide (Sigma Aldrich) in the dark. Samples were diluted with 0.1 M ammonium bicarbonate to a final concentration of 1% sodium deoxycholate. Digestions were completed by treatment with sequencing-grade trypsin (Promega) at an enzyme substrate ratio of 1:50 for 16 h at 37 °C. Trypsin was inactivated in order to stop peptide digestions by adding a volume of formic acid that lowered the pH to less than 2. Acidified peptide mixtures were loaded onto the monospin C18 column (SHIMADZU-GL) desalted, and eluted with 60% acetonitrile-0.1% formic acid. Samples were dried in a vacuum centrifuge, solubilized in 0.1% formic acid, and the same number of peptides were analyzed by mass spectrometry immediately.
The peptide mixture was loaded onto an Easy-nLC1200 system equipped with a home-made reverse phase C18 column (75 µm*30 cm, 1.9 µm) with a 200 min gradient from 1% to 90% of buffer B (buffer A: 0.1% formic acid in water; buffer B: 0.1% formic acid in 80% Acetonitrile) at 250 nL/min. The eluted peptides were ionized and directly introduced into a hybrid Orbitrap Eclipse mass spectrometer (Thermo Scientific, San Jose, CA) using a nano-spray source with the application of a distal 2.3-kV spray voltage.
Each of the samples was acquired in DIA mode. For DIA, the cycle time was 3 s, and each cycle consisted of a MS1 scan of 349.5–1,500.5 m/z with 120,000 resolution (Automatic Gain Control (AGC) target: 4e5 ions, maximum injection time: 10 ms, 349.5–1,500.5 m/z), and MS2 spectra at 30,000 resolution (AGC target: 2e5 ions, maximum injection time: 72 ms, Normalized Collision Energy (NCE): 32. The DIA isolation scheme consisted of 60 variable windows covering the 349.5–1500.5 m/z range with an overlap of 1 m/z per each window.
DIA data files were processed in Spectronaut v18 (Biognosys) using directDIA. Data were searched against the Human reference proteome with 20,387 entries (UniProtKB-TrEMBL), accessed on 03/08/2021. Data extraction parameters were set as dynamic, and non-linear iRT calibration with precision iRT was selected. Trypsin/P was set as the digestion enzyme, and two missed cleavages were allowed. Cysteine carbamidomethylation was set as a fixed modification, and methionine oxidation and protein N-terminus acetylation were set as dynamic modifications. For the protein level, identification was performed requiring a 1% q-value cutoff on the precursor ion and protein levels. Unique protein groups were reported with at least one peptide identifications. The protein level quantification was based on the peak areas of extracted ion chromatograms (XICs) of 3–6 MS2 fragment ions, specifically b- and y-ions, with local normalization and q-value sparse data filtering applied. In addition, iRT profiling was selected.
The investigation aims at identifying conformotypic peptides that significantly change their abundance across the different metabolite concentration points. Results acquired from at least three biological replicates are statistically tested for differential peptides abundances between conditions applying a fold change ≥ 1.5 or ≤ 0.67 and p-value < 0.01.
Colony formation assay
For colony formation assay, HeLa cells (4500), U87 cells (8000), RKO cells (4500) and U87/EGFRvIII cells (8000) were plated in 6-well plate in triplicates and cultured for 15-30 days before staining viable colonies with Nitro-blue tetrazolium (Sigma N5514). Data represent the mean ± SD of three independent experiments.
Flow sorting
For flow sorting experiments, HeLa/H2B-EGFP cells with PFKM knockout and rescued with rPFKM WT or K617A or L79A (pLVX-IRES-mCherry) were prepared as 7 × 106 cells in 0.5 mL PBS with 2% antibiotics. U87 and RKO cells expressing CD19 were prepared as 1 × 107 cells in 0.5 mL PBS with 2% antibiotics. Cells expressed mCherry were collected from flow sorting and plated on 35 mm dishes. The Sony MA900 instrument was used for flow sorting experiments, and mCherry channel was analyzed.
HeLa and RKO cells with PFKM knockout and rescued with rPFKM WT or K617A or L79A (pLVX-IRES-zsGreen), U87 cells with PFKM knockout and rescued with rPFKM WT or K617A (pLVX-IRES-zsGreen) were prepared as 7 × 106 cells in 0.5 mL PBS with 2% antibiotics. Cells expressed zsGreen were collected from flow sorting and plated on 35 mm dishes. The Sony MA900 instrument was used for flow sorting experiments, and EGFP channel was analyzed.
Thymidine double block and live cell time-lapse microscopic image
We used HeLa/H2B-EGFP cells, which allow high-resolution imaging of both mitotic chromosomes and interphase chromatin in live cells as described58. Briefly, HeLa/H2B-EGFP cells were treated with 2 mM thymidine for 18 h at 37 °C, washed twice with PBS, released for 9 h, and treated again with 2 mM thymidine for an additional 18 h, then washed twice with PBS and released for 9.5 h before imaging by time-lapse microscopy. Images were collected at 7 min or 10 min intervals. Mitotic index indicates the percentage of cells in mitosis according to corresponding chromatin morphologies, mitotic index be calculated as follows (every microscope field of view): \({{\rm{Mitotic}}}\; {{\rm{index}}}=\frac{{{\rm{Prophase}}}+{{\rm{Metaphase}}}+{{\rm{Anaphase}}}+{{\rm{Telophase}}}}{{{\rm{Total}}}\; {{\rm{number}}}\; {{\rm{of}}}\; {{\rm{cells}}}\; {{\rm{observed}}}}\times 100\). Perkin Elmer Operetta used for time-lapse experiments consisted of an Olympus microscope and a CO2-equilibrated chamber heated to 37 °C and equipped with 10 ×, 20 × and 40 × objectives.
Immunofluorescence staining
HeLa cells were transfected with Flag-histone H3 and HA-PFKM. Cells were synchronized by thymidine double block (2 mM) (cells in interphase were acquired), and then were released for 4 h, followed by nocodazole (50 ng/ml) treatment for 12 h (cells in mitosis were acquired). Then cells were fixed and incubated with primary antibodies (1:200), Alexa Fluor dye-conjugated secondary antibodies (1:500) and DAPI (1:2000) according to standard protocols. Cell imaging was performed using Olympus FV3000 confocal laser scanning microscope.
Protein sample preparation and mass spectrometry analysis
The gel band was excised into small pieces and washed in order with water, 50% acetonitrile. The protein was then reduced with 10 mM TCEP (Thermo Scientific) in 100 mM NH4HCO3 at room temperature for 30 min and alkylated with 55 mM iodoacetamide in 100 mM NH4HCO3 in the dark for 30 min. After that, the gel pieces were washed with 100 mM NH4HCO3 and 100% acetonitrile, and then dried using a SpeedVac. Finally, they were digested with 12.5 ng/μL chymotrypsin (Promega) in 50 mM NH4HCO3 for 16 h at 37 °C, and the tryptic peptides were extracted twice with 50% acetonitrile/5% formic acid and dried using a SpeedVac. The sample was reconstituted with 0.1% formic acid, desalted using a MonoSpinTM C18 column (GL Science, Tokyo, Japan), and then dried with a SpeedVac.
LC/tandem MS (MS/MS) analysis of peptides. The peptide mixture was analyzed by a home-made 30 cm-long pulled-tip analytical column (100 μm ID packed with ReproSil-Pur C18-AQ 1.9 μm resin, Dr. Maisch GmbH), the column was then placed in-line with an Easy-nLC 1200 nano HPLC (Thermo Scientific, San Jose, CA) for mass spectrometry analysis. Peptides eluted from the LC column were directly electro-sprayed into the mass spectrometer with the application of a distal 2.0-kV spray voltage. Data-dependent tandem mass spectrometry (MS/MS) analysis was performed with a Q Exactive Orbitrap mass spectrometer (Thermo Scientific, San Jose, CA). A cycle of one full-scan MS spectrum (m/z 300–1800) was acquired followed by top 20 MS/MS events, sequentially generated on the first to the twentieth most intense ions selected from the full MS spectrum at a 30% normalized collision energy. Full scan resolution was set to 70,000 with automated gain control (AGC) target of 3e6. MS/MS scan resolution was set to 17,500 with isolation window of 1.8 m/z and AGC target of 1e5. The number of microscans was one for both MS and MS/MS scans and the maximum ion injection time was 50 and 100 ms, respectively. The dynamic exclusion settings used were as follows: charge exclusion, 1 and >8; exclude isotopes, on; and exclusion duration, 30 s. MS scan functions and LC solvent gradients were controlled by the Xcalibur data system (Thermo Scientific). The analytical column temperature was set at 55 °C during the experiments. The mobile phase and elution gradient used for peptide separation were as follows: 0.1% formic acid in water as buffer A and 0.1% formic acid in 80% acetonitrile as buffer B, 0–3 min, 3%–5% B; 3–98 min, 5–45% B; 98–110 min, 45%–100% B, 110–120 min, 100% B. The flow rate was set as 300 nL/min.
Data Analysis. The acquired MS/MS data were analyzed against a homemade database (including the target proteins combined with E. coli database from Uniprot) using PEAKS Studio 8.5. Cysteine alkylation by iodoacetamide was specified as fixed modification with mass shift 57.02146 and methionine oxidation, protein n-terminal acetylation as variable. Additionally, phosphorylation on STY was set as dynamic modification with mass shift 79.9663. To accurately estimate peptide probabilities and false discovery rates, we used a decoy database containing the reversed sequences of all the proteins appended to the target database.
Analysis of CGGA and TCGA datasets
RNA data and clinical information for glioma were downloaded from CGGA database59(https://www.cgga.org.cn/index.jsp), and TCGA datasets of patients with Colorectal Adenocarcinoma dataset were downloaded from cBioPortal database (https://www.cbioportal.org/), respectively, and used for differential PFKM expression analyses.
Intracranial injection
We intracranially U87/EGFRvIII cells (2 × 105 in 5 µL PBS per mouse) with PFKM knockout and rescued with SFB-rPFKM WT or K617A or L79A into randomized 6-week-old female athymic nude mice. A small hand-controlled twist drill that is 1 mm in diameter is used to make a hole in the animal’s skull. The cell suspension is drawn up into the cuffed Hamilton syringe. The needle of the Hamilton syringe is slowly lowered into the central hole of the guide screw until the cuff rests on the screw surface. The cell suspension is slowly injected into the mouse’s brain60. Five or six mice per group in each experiment were included. At days 10, mice were anaesthetized with isoflurane inhalation and were subsequently intraperitoneally injected with 100 µL of 7.5 mg/mL D-luciferin (Xenogen). Bioluminescence imaging with a CCD camera (IVIS, Xenogen) was initiated 10 min after injection with 10 min exposure time. Bioluminescence from the region of interest was defined manually. All bioluminescent data were collected and analyzed using IVIS. After bioluminescence imaging, the brain of each mouse was collected, fixed in 4% formaldehyde, and embedded in paraffin. Tumor formation and phenotype were determined by histologic analysis of hematoxylin and eosin (H&E) -stained sections. The use of mice was in compliance with ethical regulations and was approved by the institutional review board at the Institute of Biochemistry and Cell Biology (SIBCB).
Subcutaneous injection
Two million of RKO cells with PFKM knockout and rescued with rPFKM WT or L79A were injected subcutaneously into the flank of female athymic nude mice (5 weeks old, ten mice per group). Tumor volumes were measured every two-three day with calipers (using the formula mm3 = (L × W2)/2) and begin at days 14. Tumors were harvested at days 25. The use of mice was in compliance with ethical regulations and was approved by the institutional review board at the Shanghai Institute of Biochemistry and Cell Biology (SIBCB).
Molecular dynamic simulations
Totally 61 peptides from 17 human proteins, which the 6 protein (PDB code 4J15 for DARS1, 6DV5 for HSPB1, 6XTX for MCM2, 8AS5 for NPM1, 6I2I for TUBA1B) were taken from the Protein Date Bank, the others 9 protein (EPRS1, HNRNPC, MYH9, EIF5A, DDX46, MIDN, HNRNPU, TRIM28, NAP1L4) were predicted by Alphafold261 A 30*30*30 Å box was set, the center of which was next to the possible binding peptide. Then the citrate was used as the ligand. The citrate model was assessed by the virtual screening with the possible protein structures by Autodock Vina62.
The initial conformation of PFKM (PDB code 4OMT)63 was taken from the Protein Data Bank (https://www.rcsb.org), and the dimer and tetramer conformation of PFKM were constructed based on the tetramer of PFKP bound with ATP and Mg ions (PDB code 4XYJ)21, which was in the R-state. And the tetramer conformation of PFKM bound with ATP and Mg ions were used to investigate the binding mode of PFKM with citrate. The initial complex model of PFKM with citrate was generated by adding 108 small molecules citric acid according to the experimental concentration using Packmol software64. The electronic potential of citrate was calculated using the Gaussian 16 program with the B3LYP functional under 6–311 G* basis set65. The partial charges of the substrate molecules were derived using the RESP charge fitted with the antechamber module in Amber 1666. The other parameters, including vdW, bond, angle and torsion terms, were obtained with the antechamber module66.
The initial structures of PFKM (PDB code 4OMT)63 and nucleosomes (PDB code 3AV2)67 were also taken from the Protein Data Bank (https://www.rcsb.org). The missing residues of nucleosomes were constructed using Modeller9.2068, and the N-terminals of the histones were taken from the AlphaFold model61.
The ZDOCK program16 was used to dock the PFKM to nucleosome. And three different initial conformations were selected for molecular dynamic simulations. The binding positions of the ATP and Mg ions with PFKM were obtained from the structure of PFKP with ATP and Mg ions (PDB code 4XYJ) by aligning the two structures.
MD simulations were performed on the dimeric forms of PFKP, PFKL and PFKM, and the monomeric forms of PFKL and PFKM with their peptide substrates. The dimeric form and monomeric form of PFKL were obtained from the tetramer of PFKL bound ADP (PDB code 7LW1), which the Mg ions was constructed by align to the 4XYJ and the ADP was modified to ATP. The substrate of PFKL was selected PLIN2, which was predicted by Alphafold261. The dimer of PFKP from mouse was constructed based on the PFKP bound ATP and Mg ions from human (PDB code 4XYJ), and the substrate of PFKP selected Lin41, which was also predicted by Alphafold261. ZDOCK program16 was used to docked the substrate to the PFKM, PFKP and PFKL.
Atomistic molecular dynamic simulations of initial models were carried out in the AMBER18 program using AMBER14SB force field for protein69,70 and Parmbsc1 force field for DNA71. Each system was neutralized with a number of neutralized ions and then immersed in a solvent box filled with TIP3P water molecules72, to warrant a distance of at least 10 Å between the surface of each protein models and the water box edge. The entire systems were subject to the energy minimization in three stages to remove the bad contacts. Firstly, the solvent and the neutralized ions were minimized by holding the protein and ligand using a restraint with strength of 100 kcal/(mol Å2), and then the minimization was performed by holding the protein and ligand using a constraint of 10 kcal/(mol Å2). Finally, the whole systems were minimized by removing any constraint. Each stage was performed using the steepest descent minimization of 1000 steps followed by 9000 steps conjugate gradient minimization. NVT (constant Number of atoms, Volume and Temperature) simulations were carried out by heating the whole system linearly with time gradually from 100 to 300 K in the first 300 ps, and the Berendsen thermostat73 was used to maintain the temperature of the whole system. Subsequently, the system was equilibrated under the temperature of 300 K for 1 ns was followed by an NPT (constant Number of atoms, Pressure, and Temperature) production run. During the heating stage, all the protein and ligands were restrained by a restrained of 100 kcal/ (mol Å2), and under equilibration stage the restraint strength was decreased to 10 kcal/ (mol Å2). During the NPT production run, the Berendsen barostat74 was used to control the pressure at 1 atm, and the Langevin thermostat was employed to control the temperature of the systems at 300 K. All bonds associated with hydrogen atoms were constrained by employing the SHAKE algorithm75, and the Hydrogen Mass Repartitioning (HMR) method was adopted, such that the integration time step of 4 fs could be used. A cutoff value of 12 Å was set for nonbonded interactions, and the Particle Mesh Ewald method76 was employed for treating electrostatic interactions. For each system, five independent molecular dynamic simulations were carried out using different velocities that were randomly generated at the beginning of the simulations and run for 1 μs. The analysis of each molecular dynamic trajectory was performed with the cpptraj module in Amber 1870.
MM-GBSA calculation
To understand the interaction between PFKM with citrate and PFKM with nucleosomes, and the dimeric forms of PFKP, PFKL and PFKM, and the monomeric forms of PFKL and PFKM with their peptide substrates, the binding free energies were calculated using the MM-GBSA method. For each complex, 500 snapshots were extracted from the last 100 ns along the molecular dynamic trajectory at an interval of 200 ps. The MM-GBSA method77 was performed to compute the binding free energies. The binding free energy (\(\Delta G\)) can be represented as:
where \(\Delta {E}_{{MM}}\) is the difference of molecular mechanic energy between the complex and each binding partner in the gas phase, \(\Delta {G}_{{sol}}\) is the solvation free energy contribution to binding. \(\Delta {E}_{{MM}}\) is further divided into two parts:
where \(\Delta {E}_{{ele}}\) and \(\Delta {E}_{{vdW}}\) are described as the electrostatic interaction and van der Waals energy in the gas phase, respectively. The solvation free energy is expressed as:
where \(\Delta {G}_{{gb}}\) and \(\Delta {G}_{{np}}\) are the polar and non-polar contributions to the solvation free energy, respectively.
Pulldown and mass spectrometry analysis
HEK293T cells were transfected with empty vector or SFB-PFKM. Vector or SFB-PFKM protein was pulled down with streptavidin agarose beads in co-IP buffer, and PFKM-associated proteins were detected by mass spectrometry analysis. The protein complexes were boiled at 95 °C for 10 min. PFKM-associated proteins were separated from the complexes using SDS-PAGE. The protein samples were digested by Filter Aided Sample Preparation (FASP) Method78, except that chloroacetamide was used in place of iodoacetamide.
The tryptic peptides were separated through a nano-emitter column (15-cm length, 75-μM inner diameter) packed in-house with 3-μM C18 ReproSil particles (Dr. Maisch GmbH) and introduced into MS via a nanoelectrospray ion source (source voltage, 2 kV). A linear gradient from 4% to 30% of acetonitrile with 0.1% (v/v) formic acid was used for peptide separation. Top 10 ions were selected at an isolation window of 2.0 m/z units. Dynamic exclusion was enabled for 90 sec to avoid choosing former target ions, and lockmass was enabled using 445.120025. All mass spectral data were acquired on LTQ-Orbitrap (Thermo Electron Finnigan). Data were acquired using Xcalibur software.
Raw Orbitrap full-scan MS and ion trap MS2 spectra were processed by MaxQuant79. Database searching was performed using the Andromeda search engine80 against UniProt human sequence database, concatenated with known contaminants and reversed sequences of all entries. An FDR < 0.01 for proteins and peptides and a minimum peptide length of 6 amino acids were required. His or GST pulldown assays were performed in co-IP buffer, and streptavidin pulldown assay was performed in buffer: 25 mM HEPES, pH 7.6, 0.37 mM EDTA, 0.35 mM EGTA, 70 mM KCl, 5 mM MgCl2, 1 mM DTT, 0.02% NP-40, 10% glycerol, and 0.1 mg/mL BSA.
For metabolite extraction
Cells were collected (~5 × 106 cells per sample) in 6 cm dish. Tilt plate, aspirate of all media, immediately add 1 mL 40:40:20 methanol: acetonitrile: water with 0.5% formic acid (ice cold). Incubate plates on ice for 5 min, add 50 µL 15% NH4HCO3, scrap cells from the plates with a cell lifter. Transfer cell lysate/methanol mixture to 1.5 mL tubes on ice. Centrifuge at 15,000 g for 10 min in cold room to pellet cell debris and proteins. Transfer 700 µL supernatant to 1.5 mL tubes on ice. Fifty microliters were transferred to LC vials containing glass inserts for analysis.
Metabolites were extracted for LC-MS/MS analysis to measure citrate
LC-MS/MS analysis was conducted using an Agilent 1290 Infinity II UHPLC system tandem with Agilent 6495 mass spectrometer. All data were collected and processed using the QQQ Quantitative Analysis Software. Chromatographic separation was achieved on Waters ACQUITY UPLC HSS T3 (100 mm × 2.1 mm, 1.8 μm) column. The mobile phase consisted of 0.1%(v/v) formic acid in water as solvent A and 0.1%(v/v) formic acid in methanol as solvent B at a flow rate of 0.4 mL/min. The gradient of mobile phase B was 2% in 1 min, 2% to 15% in 2 min, 15% to 50% in 1 min, 50% to 95% in 1 min, held at 95% for 3 min, then 95% to 2% in 0.1 min, held at 2% for 3.9 min. The flow rate of mobile phase was kept at 0.4 mL/min, column temperature was set at 40 °C and the injection volume was 5 μL. Mass spectrometer operating in negative ion mode using the following settings: gas temperature 200 °C, gas flow 14 l/min, Nebulizer 20 psi, Sheath gas temperature 250 °C, Sheath gas flow 11 l/min, Capillary 3000 V, Nozzle Voltage 1500 V. The quantitative analysis of citrate was measured by multiple reaction monitoring (MRM) mode. The MRM transitions used was m/z 191 → 110.9, and the dwell time was set at 200 ms.
Isolation and culture of T cells
Peripheral blood mononuclear cells (PBMCs) were isolated from healthy donors by gradient density centrifugation and then prepared at 3 × 106 cells/mL in T cell medium (X-VIVOTM 15, 10% fetal bovine serum, glutamax, human IL-2, human IL-7, and human IL-15). PBMCs were rested in X-vivo medium for one day before stimulated by α-CD3/CD28 activator for subsequent experiments. Human CD8+ T lymphocytes were selected from PBMCs using anti-human CD8 (Biolegend,1:200) staining and flow cytometry was carried out.
Flow cytometry
Cells were harvested and washed by PBS for 2 times, then stained in PBS containing 2% FBS with antibodies at the recommended dilution on ice for 30 min in the dark and then washed twice with PBS prior to Flow Cytometry analysis.
T Cell proliferation
T Cell proliferation was assayed by analysis of the Carboxy fluorescein succinimidyl ester (CFSE) staining. CellTrace CFSE Cell Proliferation Kit (Invitrogen) was used to form a stock concentration of 5 mM CFSE, which was diluted to 1 mM using PBS. Cells were labeled as recommended by the manufacturer. The cells were incubated for 15 min at 37 °C to label the cells. The labeled cells were then centrifuged at 500 g for 5 min at room temperature and the pellets were re-suspended in culture medium. Surface antibody (anti-human CD8, 1:200) staining and flow cytometry was carried out. Proliferation-mediated CFSE decay was analyzed by flow cytometry.
In vitro cytotoxicity assay
Tumor cells stably expressing CD19 were plated onto 96-well plates. CD19 CAR-T cells were pretreated with increasing doses of citrate, and then co-cultured with tumor cells for 24 h in the complete T cell medium. Tumor cells post co-culture were analyzed by CellCounting-Lite 2.0 Luminescent Cell Viability Assay (Vazyme) for cell viability.
Statistical analysis
Statistics were performed using GraphPad Prism software (v.8.0.2). Student’s t-test (two-tailed) was used for two-group comparisons other the situation mentioned above. p < 0.05 was considered to be significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The mass spectrometry data generated in this study have been deposited in the ProteomeXchange under accession code PXD065595. All data supporting the conclusions are available from the authors on request. Source data are provided with this paper.
References
Suski, J. M., Braun, M., Strmiska, V. & Sicinski, P. Targeting cell-cycle machinery in cancer. Cancer Cell 39, 759–778 (2021).
Icard, P. & Simula, L. Metabolic oscillations during cell-cycle progression. Trends Endocrinol. Metab. 33, 447–450 (2022).
Bury, M., Le Calve, B., Ferbeyre, G., Blank, V. & Lessard, F. New insights into CDK regulators: novel opportunities for cancer therapy. Trends Cell Biol. 31, 331–344 (2021).
Angus, S. P. et al. Retinoblastoma tumor suppressor targets dNTP metabolism to regulate DNA replication. J. Biol. Chem. 277, 44376–44384 (2002).
Scaglia, N., Tyekucheva, S., Zadra, G., Photopoulos, C. & Loda, M. De novo fatty acid synthesis at the mitotic exit is required to complete cellular division. Cell Cycle 13, 859–868 (2014).
Liu, W. et al. Lactate regulates cell cycle by remodelling the anaphase promoting complex. Nature 616, 790–797 (2023).
Icard, P., Poulain, L. & Lincet, H. Understanding the central role of citrate in the metabolism of cancer cells. Biochim. Biophys. Acta 1825, 111–116 (2012).
Lee, H. J. et al. Proteomic and metabolomic characterization of a mammalian cellular transition from quiescence to proliferation. Cell Rep. 20, 721–736 (2017).
Moreno-Sanchez, R., Rodriguez-Enriquez, S., Marin-Hernandez, A. & Saavedra, E. Energy metabolism in tumor cells. FEBS J. 274, 1393–1418 (2007).
Humphrey, W., Dalke, A. & Schulten, K. VMD: Visual molecular dynamics. J. Mol. Graph. Model. 14, 33–38 (1996).
Usenik, A. & Legisa, M. Evolution of allosteric citrate binding sites on 6-phosphofructo-1-kinase. PLoS One 5, e15447 (2010).
Wei, Y., Yu, L., Bowen, J., Gorovsky, M. A. & Allis, C. D. Phosphorylation of histone H3 is required for proper chromosome condensation and segregation. Cell 97, 99–109 (1999).
Wei, Y., Mizzen, C. A., Cook, R. G., Gorovsky, M. A. & Allis, C. D. Phosphorylation of histone H3 at serine 10 is correlated with chromosome condensation during mitosis and meiosis in Tetrahymena. Proc. Natl Acad. Sci. USA 95, 7480–7484 (1998).
Wilkins, B. J. et al. A cascade of histone modifications induces chromatin condensation in mitosis. Science 343, 77–80 (2014).
Vader, G. The chromosomal passenger complex: guiding Aurora-B through mitosis. J. Cell Biol. 173, 833–837 (2006).
Pierce, B. G. et al. ZDOCK server: interactive docking prediction of protein-protein complexes and symmetric multimers. Bioinformatics 30, 1771–1773 (2014).
Aaronson, R. P. & Frieden, C. Rabbit muscle phosphofructokinase: studies on the polymerization. The behavior of the enzyme at pH 8, pH 6, and intermediate pH values. J. Biol. Chem. 247, 7502–7509 (1972).
Fernandes, P. M., Kinkead, J., McNae, I., Michels, P. A. M. & Walkinshaw, M. D. Biochemical and transcript level differences between the three human phosphofructokinases show optimisation of each isoform for specific metabolic niches. Biochem J. 477, 4425–4441 (2020).
Gao, W. et al. The role of S-nitrosylation of PFKM in regulation of glycolysis in ovarian cancer cells. Cell Death Dis. 12, 408 (2021).
Park, J. S. et al. Mechanical regulation of glycolysis via cytoskeleton architecture. Nature 578, 621–626 (2020).
Webb, B. A. et al. Structures of human phosphofructokinase-1 and atomic basis of cancer-associated mutations. Nature 523, 111–114 (2015).
Reinhart, G. D. & Lardy, H. A. Rat liver phosphofructokinase: use of fluorescence polarization to study aggregation at low protein concentration. Biochemistry 19, 1484 (1980).
Meng, Y. et al. Glycolytic enzyme PFKL governs lipolysis by promoting lipid droplet-mitochondria tethering to enhance beta-oxidation and tumor cell proliferation. Nat. Metab. 6, 1092–1107 (2024).
Cao, L. et al. Glycolytic Pfkp acts as a Lin41 protein kinase to promote endodermal differentiation of embryonic stem cells. EMBO Rep. 24, e55683 (2023).
Aguilar, V. & Fajas, L. Cycling through metabolism. Embo Mol. Med. 2, 338–348 (2010).
Buchakjian, M. R. & Kornbluth, S. The engine driving the ship: metabolic steering of cell proliferation and death. Nat. Rev. Mol. Cell Biol. 11, 715–727 (2010).
Coller, H. A. Cell biology. The essence of quiescence. Science 334, 1074–1075 (2011).
Moncada, S., Higgs, E. A. & Colombo, S. L. Fulfilling the metabolic requirements for cell proliferation. Biochem. J. 446, 1–7 (2016).
Tudzarova, S., Colombo, S. L., Carcamo, K. aiS. & Moncada, S. S. Two ubiquitin ligases, APC/C-Cdh1 and SKP1-CUL1-F (SCF)-beta-TrCP, sequentially regulate glycolysis during the cell cycle. Proc. Natl Acad. Sci. USA 108, 5278–5283 (2011).
Wang, Z. et al. Cyclin B1/Cdk1 coordinates mitochondrial respiration for cell-cycle G2/M progression. Dev. Cell 29, 217–232 (2014).
Arnold, P. K. & Finley, L. W. S. Regulation and function of the mammalian tricarboxylic acid cycle. J. Biol. Chem. 299, 102838 (2023).
Li, Y. et al. Blockage of citrate export prevents TCA cycle fragmentation via Irg1 inactivation. Cell Rep. 38, 110391 (2022).
Icard, P. et al. Understanding the central role of citrate in the metabolism of cancer cells and tumors: an update. Int. J. Mol. Sci. 22, 6587 (2021).
Currie, E., Schulze, A., Zechner, R., Walther, T. C. & Farese, R. V. Jr. Cellular fatty acid metabolism and cancer. Cell Metab. 18, 153–161 (2013).
Poorman, R. A., Randolph, A., Kemp, R. G. & Heinrikson, R. L. Evolution of phosphofructokinase-gene duplication and creation of new effector sites. Nature 309, 467–469 (1984).
Mlakar, T. & Legisa, M. citrate inhibition-resistant form of 6-phosphofructo-1-kinase from Aspergillus niger. Appl Environ. Microbiol 72, 4515–4521 (2006).
Canelas, A. B., Pierick, A. T., Ras, C., Seifar, R. M. & Heijnen, J. J. Quantitative evaluation of intracellular metabolite extraction techniques for yeast metabolomics. Anal. Chem. 81, 7379–7389 (2009).
Parkinson, E. K. et al. Extracellular citrate and metabolic adaptations of cancer cells. Cancer Metastasis Rev. 40, 1073–1091 (2021).
Gao, W., Huang, M., Chen, X., Chen, J. & Liu, Q. The role of S-nitrosylation of PFKM in regulation of glycolysis in ovarian cancer cells. Cell Death Dis. 12, 408 (2021).
Tang, H., Lee, M., Sharpe, O., Salamone, L. & Shrager, J. B. Oxidative stress-responsive microRNA-320 regulates glycolysis in diverse biological systems. Faseb J.26, 4710–4721 (2012).
Xiong, X., Ke, X., Wang, L., Yao, Z. & Zhang, H. Splice variant of growth hormone-releasing hormone receptor drives esophageal squamous cell carcinoma conferring a therapeutic target. Proc. Natl Acad. Sci. USA 117, 201913433 (2020).
Zhou, Y., Lin, F., Wan, T., Chen, A. & Li, Q. ZEB1 enhances Warburg effect to facilitate tumorigenesis and metastasis of HCC by transcriptionally activating PFKM. Theranostics 11, 5926–5938 (2021).
Cortese, N. et al. Metabolome of pancreatic juice delineates distinct clinical profiles of pancreatic cancer and reveals a link between glucose metabolism and PD-1(+) cells. Cancer Immunol. Res 8, 493–505 (2020).
Dasgupta, S. et al. Metabolic enzyme PFKFB4 activates transcriptional coactivator SRC-3 to drive breast cancer. Nature 556, 249–254 (2018).
Sauve, D. M., Anderson, H. J., Ray, J. M., James, W. M. & Roberge, M. Phosphorylation-induced rearrangement of the histone H3 NH2-terminal domain during mitotic chromosome condensation. J. Cell Biol. 145, 225–235 (1999).
Carmena, M., Wheelock, M., Funabiki, H. & Earnshaw, W. C. The chromosomal passenger complex (CPC): from easy rider to the godfather of mitosis. Nat. Rev. Mol. Cell Biol. 13, 789–803 (2012).
Hirota, T., Lipp, J. J., Toh, B. H. & Peters, J. M. Histone H3 serine 10 phosphorylation by Aurora B causes HP1 dissociation from heterochromatin. Nature 438, 1176–1180 (2005).
Diehl, F. F., Sapp, K. M. & Vander Heiden, M. G. The bidirectional relationship between metabolism and cell cycle control. Trends Cell Biol. 34, 136–149 (2024).
Xu, D. et al. The evolving landscape of noncanonical functions of metabolic enzymes in cancer and other pathologies. Cell Metab. 33, 33–50 (2021).
Webb, B. A., Dosey, A. M., Wittmann, T., Kollman, J. M. & Barber, D. L. The glycolytic enzyme phosphofructokinase-1 assembles into filaments. J. Cell Biol. 216, 2305–2313 (2017).
Hans, F. & Dimitrov, S. Histone H3 phosphorylation and cell division. Oncogene 20, 3021–3027 (2001).
Sharma, K. L. et al. Mitogen-induced transcriptional programming in human fibroblasts. Gene 800, 145842 (2021).
Lee, J. H. et al. AKT phosphorylates H3-threonine 45 to facilitate termination of gene transcription in response to DNA damage. Nucleic Acids Res. 43, 4505–4516 (2015).
Hurd, P. J. et al. Phosphorylation of histone H3 Thr-45 is linked to apoptosis. J. Biol. Chem. 284, 16575–16583 (2009).
Nowak, S. J. & Corces, V. G. Phosphorylation of histone H3: a balancing act between chromosome condensation and transcriptional activation. Trends Genet 20, 214–220 (2004).
Rossetto, D., Avvakumov, N. & Cote, J. Histone phosphorylation: a chromatin modification involved in diverse nuclear events. Epigenetics 7, 1098–1108 (2012).
Lu, Z. et al. Activation of protein kinase C triggers its ubiquitination and degradation. Mol. Cell Biol. 18, 839–845 (1998).
Kanda, T., Sullivan, K. F. & Wahl, G. M. Histone-GFP fusion protein enables sensitive analysis of chromosome dynamics in living mammalian cells. Curr. Biol. 8, 377–385 (1998).
Zhao, Z. et al. Chinese Glioma Genome Atlas (CGGA): A Comprehensive Resource with Functional Genomic Data from Chinese Glioma Patients. Genomics Proteom. Bioinforma. 19, 1–12 (2021).
Liang, J. et al. Mitochondrial PKM2 regulates oxidative stress-induced apoptosis by stabilizing Bcl2. Cell Res. 27, 329–351 (2017).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Trott, O. & Olson, A. J. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. comput. Chem. 31, 455–461 (2010).
Kloos, M., Brüser, A., Kirchberger, J., Schöneberg, T. & Sträter, N. Crystallization and preliminary crystallographic analysis of human muscle phosphofructokinase, the main regulator of glycolysis. Acta Crystallogr. Sect. F. Struct. Biol. Commun. 70, 578–582 (2014).
Martínez, L., Andrade, R., Birgin, E. G. & Martínez, J. M. PACKMOL: a package for building initial configurations for molecular dynamics simulations. J. Comput Chem. 30, 2157–2164 (2009).
Frisch, M. J. et al. Fox DJ Gaussian 09. Revision D.1 edn. (Gaussian, Inc., Wallingford CT., 2009).
Case, D. et al. Amber 2016. (University of California, San Francisco, 2016).
Tachiwana, H. et al. Structures of human nucleosomes containing major histone H3 variants. Acta Crystallogr. Sect. D., Biol. Crystallogr. 67, 578–583 (2011).
Webb, B. & Sali, A. Comparative protein structure modeling using modeller. Curr. Protoc. Protein Sci. 86, 2 9 1–2 9 37 (2016).
Hornak, V. et al. Comparison of multiple Amber force fields and development of improved protein backbone parameters. Proteins 65, 712–725 (2006).
Roe, D. R. & Cheatham, T. E. PTRAJ and CPPTRAJ: software for processing and analysis of molecular dynamics trajectory data. J. Chem. Theory Comput. 9, 3084–3095 (2013).
Ivani, I. et al. Parmbsc1: a refined force field for DNA simulations. Nat. methods 13, 55–58 (2016).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Berendsen, H. J. C., Postma, J. P. M., Vangunsteren, W. F., Dinola, A. & Haak, J. R. Molecular-dynamics with coupling to an external bath. J. Chem. Phys. 81, 3684–3690 (1984).
Braun, E., Moosavi, S. M. & Smit, B. Anomalous effects of velocity rescaling algorithms: the flying ice cube effect revisited. J. Chem. theory Comput. 14, 5262–5272 (2018).
Ryckaert, J. P., Ciccotti, G. & Berendsen, H. J. C. Numerical-integration of cartesian equations of motion of a system with constraints - molecular-dynamics of N-alkanes. J. Comput Phys. 23, 327–341 (1977).
Darden, T., York, D. & Pedersen, L. Particle mesh ewald - an N.Log(N) method for ewald sums in large systems. J. Chem. Phys. 98, 10089–10092 (1993).
Kollman, P. A. et al. Calculating structures and free energies of complex molecules: combining molecular mechanics and continuum models. Acc. Chem. Res. 33, 889–897 (2000).
Winiewski, J. R., Zougman, A., Nagaraj, N. & Mann, M. Universal sample preparation method for proteome analysis. Nat. Methods 6, 359–362 (2009).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Cox, Jr. et al. Andromeda: a peptide search engine integrated into the MaxQuant environment. J Proteome Res. 10, 1794–1805 (2011).
Acknowledgments
This work was supported by the National Natural Science Foundation of China (grant no. 92357301, 92253305, and 32025013) to W. Y.; National Key Research and Development Program of China (grant no. 2024YFA1306003 and 2022YFA0806200) to W. Y.; The Strategic Priority Research Program of the Chinese Academy of Sciences (XDB0990000) to W.Y.; CAS Project for Young Scientists in Basic Research (grant no. YSBR-014) to W. Y.; Science and Technology Commission of Shanghai Municipality (24J12800600); the Innovative Research Team of High-level Local Universities in Shanghai (grant no. SHSMU-ZLCX20212302) to W. Y.; Shanghai Municipal Science and Technology Major Project; W.Y. is a SANS Exploration Scholar. National Key R&D Program of China (grant no. 2023YFF1204903) to H.C.; We thank the Genome Tagging Project (GTP) Center and the Core Facilities of SIBCB for technical support and all the Core Facilities of SIBCB for technical support. We thank Chemical Biology Core Facility in CEMCS for technical support. We thank the Mass Spectrometry & Metabolomics Core Facility at the Center for Biomedical Research Core Facilities of Westlake University for sample analysis. We thank Xueliang Zhu (SIBCB, CAS, Shanghai, China.) and Yunqi Tang (Shanghai Institute of Nutrition and Health, CAS, Shanghai, China.) for valuable advice. We also thank Chen Su and Qingrun Li of the Mass Spectrometry System (https://cstr.cn/31129.02.NFPS.MSAS) at the National Facility for Protein Science in Shanghai (https://cstr.cn/31129.02.NFPS), Shanghai Advanced Research Institute, Chinese Academy of Sciences, China for MS sample preparation, data collection and data analysis.
Author information
Authors and Affiliations
Contributions
W.Y. conceived the concept of the study. W.Y., P.L. and Y.Q. designed the study. P.L. and Y.Q. performed most of the experiments and contributed to data analyses and figure editing. G.L. designed the molecular dynamics strategy. G.L. and H.C. performed the simulations and data analysis. H.W., Y.Z., X.H., X.X., C.L. and R.Z. provided experimental assistance. H.G. assisted in reviewing and editing the manuscript. W.Y., P.L. and Y.Q. wrote the manuscript with comments from all authors. All authors contributed to the article and approved the submitted version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interest.
Peer review
Peer review information
Nature Communications thanks Shanshan Li who co-reviewed with Xinlin Xu; Lauren Albrecht and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lin, P., Qi, Y., Chu, H. et al. PFKM phosphorylates histone H3 and promotes mitotic progression by sensing the levels of citrate. Nat Commun 16, 6736 (2025). https://doi.org/10.1038/s41467-025-62111-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-62111-3