Abstract
Wilms tumor (WT), the most common pediatric renal malignancy, exhibits a relatively low mutational burden compared to adult cancers, which hinders the development of targeted therapies. To elucidate the molecular landscape of WT, we perform integrative proteomic, phosphoproteomic, transcriptomic, and whole-exome sequencing analyses of WT and normal kidney tissue adjacent to tumor. Our multi-omics approach uncovers prognostic genetic alterations, distinct molecular subgroups, immune microenvironment features, and potential biomarkers and therapeutic targets. Proteome- and transcriptome-based stratification identifies three molecular subgroups with unique signatures, correlating with different histopathological subtypes and putative cellular origins at different stages of embryonic kidney development. Notably, we identify EHMT2 as a promising prognostic biomarker and therapeutic target associated with epigenetic regulation and Wnt/β-catenin pathway. In this work, we provide a comprehensive molecular characterization of WT, offering valuable insights into its pathogenesis and a foundational resource for future therapeutic development.
Similar content being viewed by others
Introduction
Wilms tumor (WT) is the most common malignant kidney tumor type in children, accounting for approximately 90% of all kidney tumors and 7% of all childhood malignancies1. The annual incidence of WT is 4.3 cases per million in East Asia, which is even lower than that in North America or Europe (8–9 cases per million)2. In children aged 0–4 in China, the incidence rate of this malignancy is higher than 7 cases per million3. With the implementation of standardized treatment protocols, the overall survival rate of WT has been significantly improved, but for children with unfavorable histological types, the 4-year survival rate varies from 30% to 85% depending on the tumor stage4. At present, approximately 40 somatic mutations or copy number (CN) variants are believed to be key cancer driver candidates in the oncogenesis and development of WT, including WT1, CTNNB1, AMER1, IGF2, TP53, and MYCN 5,6,7. Three of the most prominent genetic features and epigenetic alterations in WT are loss of function of WT1, activation of the Wnt signaling pathway, and overexpression of IGF2 8,9. However, the frequency of gene mutations and CN variants in WT is much lower than that in adult tumors, which has, to some extent, impeded the development of targeted therapies for children with WT10. Currently, the most promising molecular markers of high-risk WT, which are associated with poor prognosis, include loss of heterozygosity (LOH) of 1p and 16q11, increased 1q CN12, TP53 mutations and MYCN CN variants5,13. However, these chromosomal abnormalities and mutation-related risk stratification indicators still have some shortcomings in guiding treatment. This may lead to less accurate risk stratification or treatment regimen development in children, which can result in chemotherapeutic drug side effects of overtreatment or even tumor progression or recurrence due to insufficient treatment. Although genomics has played an important role in the exploration of the pathogenesis of WT and guiding individualized treatment14,15, somatic mutations do not always result in corresponding changes in protein expression or function in tumors. Therefore, combining the analysis of protein and mRNA expression in WTs using proteomic and transcriptomic techniques with genomic data is crucial to explore the possible pathogenesis, potential molecular markers, and therapeutic targets of WT15,16,17,18. Proteomic studies of WT have been limited to exploring molecular markers in urine and serum samples. These markers included prohibitin in urine and serum Apo C-1 and haptoglobin in blood19,20,21. However, proteogenomic analysis of WT samples and corresponding kidney samples with larger sample sizes has rarely been reported.
Proteomics is an efficient, high-throughput screening technique for protein expression profiling, protein interaction mapping, and quantitative analysis of protein modifications. By combining proteomics with whole exome sequencing and transcriptome analysis, proteogenomic analysis can provide multilevel information for mapping biological pathways associated with tumor development and metastasis, allowing for better staging of tumors, predicting treatment response, matching targeted therapies, and exploring therapeutic targets22,23,24.
In this work, we focused on performing in-depth bioinformatics analysis to interpret multi-omics of clinical samples from WT, which includes whole exome sequencing, quantitative proteomics, phosphorylated proteomics, and transcriptome analysis of both WT samples and normal adjacent to tumor (NAT) samples. Our aim was to discover potential molecular markers, therapeutic targets, and cancer driver candidates for tumorigenesis in WT and to provide a basis for clinical risk classification and treatment selection.
Results
Study design and multi-omics findings of the Wilms tumor cohort
This study recruited 96 patients who were diagnosed with WT before the age of 18. The cohort consisted of 91 WT samples and 74 NAT samples. To comprehensively profile the molecular features of WT, we performed whole-exome sequencing (WES), RNA sequencing (RNA-seq), quantitative proteomic and phosphoproteomic analysis on those samples (Fig. 1a, b and Supplementary Data 1a). The detailed clinical and pathological characteristics, as well as prognostic information were summarized in Fig. 1c and Supplementary Data 1b.
a The study design and overview of the multi-omics landscape of Wilms tumor. b Patient-centric Circos plot, with each circle representing one omics of WT (Wilms tumor)/NATs (normal adjacent to tumor). Gray gaps indicate unmeasured data. c Pie charts illustrate the patient distribution within the WT cohort based on key clinical and prognostic variables such as sex, tumor histology, risk group, tumor site, preoperative chemotherapy, distant metastasis, tumor stage, and prognostic outcomes in this study. d Boxplots showing the distribution of the number of RNAs, proteins, phosphorylated sites, and phosphorylated proteins identified by multi-omics profiling in WT and NATs. Blue: NATs; orange: WT. A two-sided Wilcoxon rank sum test was performed to determine the difference between WT and NATs. RNAs: NAT (n = 37), WT (n = 62); proteins: NAT (n = 71), WT (n = 88); phosphorylated sites: NAT (n = 23), WT (n = 23); phosphorylated proteins: NAT (n = 23), WT (n = 23). Data are presented as means \(\pm\) SEM. Boxplots show median value, box indicates 75 and 25th quartile and whiskers extend to the farthest value (largest and smallest). e Gene-wise mRNA‒protein correlations in NAT (left) and WT (right) samples (Spearman correlation). One-sided hyper-geometric test was performed to test the statistical significance of enrichment results by correlated mRNA-protein pairs in WT and NATs. Source data are provided as a Source Data file.
We identified a total of 369 non-silent mutations, including 317 substitutions (291 missense mutations, 18 nonsense mutations, 7 splicing mutations, and one translation start site mutation) and 52 indels (21 frameshift mutations, 29 in-frameshift mutations, 1 splicing mutation, and 1 nonsense mutation) in 36 WT samples, which resulted in a medium of tumor mutation burden (TMB) at 0.15 per million bases (Supplementary Fig. 1a, Supplementary Table 1, Supplementary Data 1c, d). We also identified a total of 16,835 RNAs encoding proteins from 62 WT and 37 NAT samples, 9956 proteins from 88 WT and 71 NAT samples, 9343 phosphorylation sites localized in 2918 proteins from 23 paired WT and NAT samples (Supplementary Fig. 1b, Supplementary Data 1e, f, g). Among the phosphorylation sites, 8703 (93.15%) were curated from Signor or PhosphoSite databases25,26 (Supplementary Data 1g), indicating high reliability of our phosphoproteome data. Notably, the number of identified genes, phosphorylation sites and phosphoproteins were significantly higher in WT compared to NATs (Wilcoxon test, p < 0.05), suggesting that tumor cells underwent complex alterations in gene expression and protein activity (Fig. 1d).
Additionally, we performed correlation analysis on the mRNA‒protein pairs identified by both transcriptomic and proteomic data. Specifically, we found that the WT samples (median = 0.27) had a higher Spearman correlation than the NAT samples (median = 0.2) (Fig. 1e, Supplementary Data 1h). The inconsistency between the transcriptome and proteome also implied the additional information that could not be observed by the transcriptome. In WT samples, the mRNAs/proteins involved in CDK regulation of DNA replication and genes controlling nephrogenesis were positively correlated (GSEA, adjusted p < 0.05, Fig. 1e, Supplementary Data 1i). The mRNA/protein levels in pathways such as respiratory electron transport, the IL-2 signaling pathway and fatty acid metabolism were positively correlated in NAT samples (GSEA, adjusted p < 0.05, Fig. 1e, Supplementary Data 1i). These results indicated that cell proliferation, DNA replication, and kidney development exhibited higher activities in WT samples than in NAT samples.
Tumor-NAT comparisons revealed tumorigenic genes and potential biomarkers
To identify the genes or proteins possibly associated with the tumorigenesis of WT, we conducted differential expression analysis at the mRNA, protein, and phosphoprotein levels by comparing WT samples to NAT samples. Through principal component analysis, tumors and NATs were efficiently distinguished based on their transcriptome, proteome, and phosphoproteome data (Supplementary Fig. 2a).
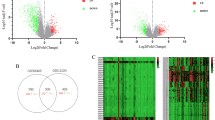
Specifically, we identified 6174 upregulated genes and 3714 downregulated genes in WT tissues at mRNA level (fold change >2 or <1/2, adjusted p-value < 0.05) (Supplementary Fig. 2b, Supplementary Data 2a). Additionally, we discovered 1827 upregulated and 2240 downregulated proteins (fold change > 1.2 or <5/6, adjusted p-value < 0.05) (Supplementary Fig. 2b, Supplementary Data 2a) in WT tissues. Regarding the phosphoproteome data, we retained 9343 phosphorylated sites for further analysis after quality control, leading to the identification of 1056 upregulated and 530 downregulated phosphoproteins in WT tissues (fold change >2 or <1/2, adjusted p-value < 0.05) (Supplementary Fig. 2b, Supplementary Data 2a). Furthermore, a total of 351 upregulated (4.78%) and 146 downregulated (2.79%) gene products were collectively identified through differential expression analysis using transcriptome, proteome, and phosphoproteome data across all three expression levels (Fig. 2a, b Supplementary Data 2a). Each omics dataset reveals a distinct set of differentially expressed gene products, highlighting the critical importance of multi-omics integration analysis in cancer research.
a Venn diagram depicting the overlap of genes upregulated in WT (Wilms tumor) for RNAs, proteins, and phosphorylated proteins. b Venn diagram depicting the overlap of genes downregulated in WT for RNAs, proteins, and phosphorylated proteins. c Boxplot showing the Z-score normalized RNA (upper panel) and protein (lower panel) abundance of the indicated genes in WT (RNA: n = 62, protein: n = 88) and NAT (normal adjacent to tumor) (RNA: n = 37, protein: n = 71). A two-sided Wilcoxon rank sum test was performed to determine the difference. Boxplots show median value, box indicates 75 and 25th quartile and whiskers extend to the farthest value (largest and smallest). d The pathways enriched by the upregulated (left) and downregulated (right) RNAs/proteins/phospho-proteins in WT compared to NAT. Bold texts represent categories of enriched pathways. Each circle represents one pathway, and the pathways are connected if they shared 20% genes. The colors within the nodes represent the omics data. e Representative genes involved in the pathways enriched by the differentially expressed RNAs/proteins between WT and NAT. f The kinases with differential activities between WT and NAT inferred by differently phosphorylated sites. Source data are provided as a Source Data file.
The differential expression analysis also revealed that some diagnostic biomarkers of WT were also found to be overexpressed in WT samples at both the mRNA and protein levels (Fig. 2c)27. WT samples with (chemo-treated samples) or without (treatment-naïve samples) chemotherapy were both included in the differential analysis between WT and NATs, and correlation analysis showed that WT samples with or without chemotherapy had high concordance in differentially expressed genes compared with NATs (Supplementary Fig. 2c, Supplementary Data 2b). In terms of the degree of differential gene expression, the upregulated proteins in the treatment-naïve group were decreased in chemo-treated group but were still overexpressed relative to NATs (Supplementary Figs. 2d, e Supplementary Data 2c).
The functional enrichment analysis of the differentially expressed mRNAs, proteins, and phosphoproteins revealed that the pathways-related to cell proliferation and cell cycle (cell cycle, mitotic metaphase and anaphase and retinoblastoma gene in cancer), RNA processing (mRNA splicing and RNA polymerase II transcription termination), epigenetic regulation (ERCC6 (CSB) and EHMT2 (G9a) positively regulate rRNA expression, epigenetic regulation of gene expression), renal development (development of ureteric collection system), NOTCH and Wnt signaling were highly enriched by upregulated mRNAs, proteins, and phosphoproteins in WT, suggesting their potential high activation in WT and their close association with the tumorigenesis of WT (Fig. 2d, Supplementary Data 2d). In contrast, the normal metabolism (amino acid metabolism, glycolysis/gluconeogenesis and fatty acid metabolism), renal function (proximal tubule transport) and stroma (cell‒cell communication, leukocyte trans-endothelial migration, focal adhesion) were enriched by downregulated mRNAs, proteins, and phosphoproteins, suggesting that the normal metabolic capability and renal function were impaired in WT (Fig. 2d, Supplementary Data 2d). Furthermore, we also found that pathways related to immune responses (complement cascade, FCGR activation, CD22-mediated BCR regulation and antigen processing and presentation) were specifically enriched by the downregulated mRNAs or proteins (Fig. 2d, Supplementary Data 2d), indicating a potential suppression or dysregulation of immune-related processes in WT. Specifically, the key genes involved in those pathways were differentially expressed between WT and NAT samples at both RNA and protein levels (Fig. 2e).
In addition, the analysis of differentially phosphorylated proteins and phosphorylation sites in WT unveiled several key kinases responsible for these phosphorylation changes. Notably, we observed increased activities of kinases in WT, such as ATM, CDK1, and CDK2, which might be regarded as potential therapeutic targets in WT (Fig. 2f, Supplementary Fig. 2f, Supplementary Data 2e).
Genomic alterations and their impact on the transcriptome, proteome, and phosphoproteome
The whole exome sequencing identified 7 significantly mutated genes in WT (MutsigCV, p-value < 0.05 or frequency ≥ 5%), including CTNNB1, WT1, AMER1, BRAF, TP53, CCNE1, and PIK3CB. Notably, mutations of TP53, WT1, CTNNB1, and AMER1 were also detected in the TARGET WT cohort (Fig. 3a, Supplementary Data 3a). Several genes were previously known oncogenes or tumor suppressor genes, with CTNNB1, AMER1 and WT1 being related to WT28. Compared to the TARGET database, our cohort had relatively higher mutation rates of WT1 and CTNNB1 but a significantly lower mutation rate of TP53 (Supplementary Fig. 3a).
a The recurrently mutated genes in 36 WT (Wilms tumor) samples. Top: mutation counts of the top seven mutated genes in each patient. Mutation types and their frequencies are demonstrated by a bar plot in the right panel. b The expression patterns of genes involved in the Wnt/β-catenin pathway and stroma in normal adjacent to tumor (NAT), wild-type WT samples (Wild), and WT samples carrying Wnt/β-catenin pathway-related mutations (Mutation). c The difference in stromal scores between WT samples with and without Wnt/β-catenin pathway-related mutations. A two-sided Wilcoxon rank sum test was performed to determine the difference. Boxplots show median value, box indicates 75 and 25th quartile and whiskers extend to the farthest value (largest and smallest). Wild type (n = 26), CTNNB1/AMER mutation (n = 9). Data are presented as means \(\pm\) SEM. d The frequently amplified and deleted genomic regions detected in the Xinhua-WT cohort by GISTIC2. e Genome-wide CNA-mRNA and CNA-protein correlations. Positive and negative correlations are indicated in red and green, respectively. Lower panel: the number of significant correlations. Blue bars stand for specific correlations with mRNA (left) or protein (right), and black bars stand for common correlation on both mRNA and protein levels. f Cis-regulatory CNAs aggregated at chromosomal cytobands. The statistical significance was determined by a two-sided hypergeometric test. g The amplified and deleted genes with potential cis-regulatory effects. Left panel: Gain of copy number located in 13q, 12q, 9q, 9p, 8p, 7p, 7q, 6q, and 1q. Right panel: Deletions located in 16q and 1p. The gray lines between genes represent correlations, and thicker lines indicate stronger correlations. Source data are provided as a Source Data file.
Mapping the mutations to cancer driver pathways revealed that the Wnt/β-catenin pathway was the most frequently mutated pathway in WT (Supplementary Fig. 3b, Supplementary Data 3b), and played key roles in cell proliferation as well as embryonic kidney development29,30,31,32. The expression profile of the Wnt/β-catenin pathway-related mutant samples (with CTNNB1, AMER1 mutations) showed activation of the Wnt/β-catenin pathway, including upregulation of MYC, WNT5A, FZD1, NKD2, and PLCB3 at the mRNA level (Fig. 3b, Supplementary Data 3c). The enrichment analysis of differentially expressed genes in Wnt/β-catenin pathway mutated samples revealed several stroma-related pathways such as muscle contraction pathways and focal adhesion, which were highly expressed in mutated samples compared to wild-type WT and NAT samples (Supplementary Fig. 3c, Supplementary Data 3d). Furthermore, downstream genes of the Wnt/β-catenin pathway related to muscle function, cell structure, and stromal components, including MYH3, MYL1, and VIM, were highly expressed in mutated samples compared to both wild-type WT samples and NAT samples (Fig. 3b, Supplementary Data 3c). Correspondingly, the stromal score was higher in Wnt/β-catenin mutant samples analyzed using transcriptomic data (Fig. 3c). These results indicated that the mesenchymal phenotype of tumor cells might be maintained by Wnt/β-catenin pathway activation.
The copy number alteration (CNA) analysis identified some well-known CNAs, such as LOH of 1p/16q and gain of 1q. Additionally, we identified several significantly CN gains/amplifications at 2p11.1, 2q11.1, 4p11, 6p12.1, 6q16.3, 9q21.11, 10q11.21, 12q12, 12q21.2, 18q11.1, and 19q11, and CN deletions/losses at 4q13.3 and 9q13 (Fig. 3d). Notably, the amplifications at 1q, 4p, 12q, and 19q, as well as the deletions at 1p, 9q, and 16q, were identified in the TARGET-WT cohort (Supplementary Fig. 3d)7.
Next, we conducted correlation analysis between CNAs and mRNA/protein expression to assess the impact of CNAs on mRNA and protein expression in WT. Our findings revealed that the correlation between CNA and mRNA is stronger than that between CNA and protein, likely due to the complex regulation of protein translation. The hotspot of CNAs that potentially affected both mRNA and protein expression levels were primarily located within chromosomal regions 6p, 12p, and 12q (Fig. 3e). The joint analysis of CNA-mRNA and CNA-protein correlations identified 225 CNA cis-regulated genes that were primarily located within cytobands, such as 1p, 16q, 12q, 9q, 6p, 7p, 7q, 1q, 13q, 8p, 6q and 9p (hypergeometric test, adjusted p < 0.05, Fig. 3f, Supplementary Data 3e), suggesting their potential regulatory roles in tumorigenesis or tumor progression. Furthermore, the amplified genes exhibited a high enrichment in pathways related to RNA splicing, regulation of G0 to G1 transition, DNA replication, mitotic sister chromatid segregation, chromatin remodeling and G1/S transition of the mitotic cell cycle. In contrast, the deleted genes, primarily located in 1p and 16q, were enriched in pathways, such as electron transport chain, carboxylic acid catabolic process, fatty acid oxidation, hexose metabolic process, and organic acid metabolic process (Fig. 3g, Supplementary Data 3f, hypergeometric test, adjusted p < 0.05). Particularly, among genes located in cis-regulatory CN amplifications (the top 2 CNAs: 6p and 12q), the expression levels of most genes were negatively correlated with EFS and OS. In contrast, genes in cis-regulatory CN deletions (1p and 16q) showed expression levels positively correlated with event-free survival (EFS) and overall survival (OS) (Supplementary Fig. 3e), further supporting a tumor-promoting role for amplified genes and a tumor-suppressing role for deleted genes.
Specifically, energetic metabolism enzymes, such as AKR7A2, COQ9, COX4I1, AKR1A1, and ALDH4A1, were located in the 1p or 16q region. Deletions in these regions significantly reduced both RNA and protein expression levels (Supplementary Fig. 3f), indicating a potential association between 1p and 16q deletions and metabolic reprogramming regarding reduced energy supply from the aerobic respiration process in WT. Collectively, these results indicated that CN gains and deletions might result in cell cycle progression and dysfunctional metabolism in WT, respectively.
Molecular stratification of Wilms tumor based on transcriptome and proteome
To elucidate the intertumoral heterogeneity among WT tumors, we employed an integrative approach using multi-omics data for tumor sample classification. Specifically, we classified 59 WT samples with both transcriptomic and proteomic data into three distinct subgroups exhibiting unique clinical, pathological, and molecular characteristics (Fig. 4a, Supplementary Figs. 4a,b,c,d, Supplementary Data 4a).
a The subgroups identified by spectral clustering and their clinical relevance. Heatmap depicting the RNA and protein expression patterns of the representative genes in the three subgroups, Subgroup 1 (S1), Subgroup 2 (S2) and Subgroup 3 (S3). b Bar chart of the proportions of blastemal, stromal and epithelial components in S1 (n = 24), S2 (n = 13), and S3 (n = 22). A one-sided Chi-squared test was performed to test the difference. c Box plot of the stromal scores (left) and immune scores (right) in S1 (n = 24), S2 (n = 13) and S3 (n = 22). Data are presented as means \(\pm\) SEM. Boxplots show median value, box indicates 75 and 25th quartile and whiskers extend to the farthest value (largest and smallest). A two-sided Wilcoxon rank sum test was performed to determine the difference between the subtypes. The false discovery rate adjustments were made for multiple comparisons. d Heatmap of alteration pathways in the proteomic subgroups at the RNA, protein and phospho-protein levels. e The mRNA and protein expression level of the signature genes of the alteration pathways highly expressed in S1 (left), S2 (middle), and S3 (right). f Classification of activated kinases in S1, S2 and S3. g Heatmap depicting the subgroup-specific phosphorylation sites and corresponding kinases. The labels on the right represent the phosphorylated sites. h Kaplan‒Meier plots of event-free survival (left) and overall survival (right) of WT subgroups in the TARGET cohort. The p-value was calculated by the log-rank test. i The proportions of tumors with stages I, II, III, IV, or V in subtypes S1, S2, and S3 in database GSE31403. A Chi-square test was performed to test the difference. Source data are provided as a Source Data file.
It has been well recognized that WT predominantly exhibits three main pathological types (blastemal, epithelial, and stromal histology), and blastemal histology is correlated with a worse prognosis. Notably, Subgroup1 (S1), Subgroup2 (S2), and Subgroup3 (S3) correlated with blastemal-dominant, stromal-dominant, and epithelial-dominant samples, respectively, based on HE staining, suggesting a high concordance between the pathological classification and molecular subgroups (Fig. 4b, Supplementary Fig. 4e). These results demonstrated that pathological cell types of WT were the key determinants of its molecular characteristics, while molecular subgroups revealed more complicated and accurate information than HE staining results. Notably, S1 had a higher proportion of high-risk patients and anaplastic pathology (Supplementary Fig. 4f). Additionally, S2 had the highest stromal score, while S3 exhibited the highest immune score (Fig. 4c), suggesting that a distinct tumor microenvironment in S2 and S3 might influence tumor growth dynamics and the interplay between the tumor and its surroundings.
To further characterize the three subgroups of WT, we analyzed the differentially expressed genes and pathways in each subgroup. S1 was characterized by elevated mRNA and protein expression involved in the pathways of cell cycle and epigenetic regulation, and DNA damage response (DDR), encompassing genes such as CDK1/2/4, EHMT1/2, and HDAC2. S2 displayed the highest expression of proteins enriched in the Wnt signaling, myogenesis, collagen, and focal adhesion pathways, such as COL1A1, COL3A1, and MYH2. S3 demonstrated elevated expression of immune cell markers, major histocompatibility complex and interferon-induced proteins, with significant proteins such as CD19, HLA-E, and IFIT1 (Fig. 4d, e, Supplementary Data 4b, c).
Integration of the phosphoproteome revealed differential kinase activity and substrates among subgroups, with CDK1, CDK2, CDK5, CSNK1A1, MAPK8, GSK3A, PDPK1 and GSK3B showing higher activities in S1, and AKT1, RPS6KA1, CDK6, and MAPK1 in S3, underscoring the heterogeneity of signaling pathways within each subgroup. Notably, S1 displayed distinct kinase activity hotspots, potentially warranting kinase-targeted therapy (Fig. 4f, g).
Interestingly, S2 was characterized by prominent CTNNB1 mutations, along with stromal features and ECM pathway activation, suggesting that Wnt signaling activation might promote the maintenance of the mesenchymal phenotype in WT. Key CNAs, including deletions at 1p, 16q, and gains at 1q, 6p, and 12q, were more prevalent in S1, aligning with S1 signatures such as cell cycle, epigenetic regulation, and tRNA regulation-related pathway activation (Supplementary Fig. 4g). Furthermore, we used genes that were specifically highly expressed in each subgroup as signature genes. Using the nearest template prediction (NTP) model33, we classified the WT samples from the TARGET database into 3 subgroups. Survival analysis showed that S1 had lower EFS rate (p = 0.035) and OS rate (p = 0.38) (Fig. 4h). This result corresponded to the high-risk histology and activated cell cycle and DNA damage response (DDR) pathways observed in S1. Due to differences in the distribution of pathology types between TARGET and our cohort, we separated diffuse anaplastic WT (DAWT) and relapsed favorable histology WT (FHWT) in TARGET cohort. As shown in Supplementary Fig. 4h, S1 still correlated with a worse prognosis compared to S2 and S3 in both relapsed FHWT and DAWT. Additionally, we included another public database containing primary FHWT samples (GSE31403), and result showed that S1 has significantly higher proportion of stages III–V samples, further suggesting that S1 was associated with poor prognosis (Fig. 4i).
Kidney developmental perspective of WT tumorigenesis
In addition to common tumorigenic molecular features, we identified the activation of pathways associated with kidney development in WT, particularly those involved in embryonic development and the Wnt signaling pathway (Fig. 2c). Given that WT is classified as an embryonal tumor, our findings prompted a deeper investigation into the intricate relationship between kidney development and WT onset. Early kidney development progresses through four stages: metanephric mesenchyme (MM), ureteric bud (UB), cap mesenchyme (CM), and renal vesicle (RV). Several studies have demonstrated that WT exhibits a gene profile resembling early kidney development. For instance, critical regulators of kidney development9,34,35,36,37, such as SIX2, SALL1, WT1, and EYA1, were upregulated in WT at both the mRNA and protein levels (Supplementary Table 2). Moreover, we also collected 103 signature genes representing 12 renal developmental phases and assessed their expression in each sample using the gene set variation analysis (GSVA) algorithm (Supplementary Data 5a). As illustrated in Fig. 5a, S1 showed a strong association with MM and CM, S2 with mesangial cells and renal interstitium (RI), and S3 with the S-shaped body (Fig. 5a). These findings aligned with the predominant pathological types within each molecular subgroup, suggesting potential origins of distinct WT tumor cell populations. WT displayed distinct gene signatures reflective of various developmental stages, including MM, CM, RV, mesangial cells and RI. Notably, signatures corresponding to UB, ureteric tip, and relatively mature cell types such as podocytes, proximal tubules, and distal tubules were absent, implying a specific developmental trajectory in WT tumorigenesis (Supplementary Fig. 5a). The TARGET-WT cohort demonstrated a similar expression pattern of these developmental stages (Supplementary Fig. 5b), further supporting the strong association between WT and early embryonic kidney development, particularly stages preceding the mesenchymal-epithelial transition (MET), such as MM and CM. MET is a critical process for early kidney development and governed by several genes with pivotal roles38,39,40. In our cohort, we observed downregulation of key MET-promoting genes (CDH6, CDH4, and FGF1) at the protein level, suggesting potential dysregulation of the MET process in WT tumorigenesis (Fig. 5b).
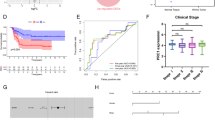
a The scores for 12 developmental phases of the kidney by the ssGSEA algorithm. The mean scores were calculated for each phase in normal tissue as well as in S1 (Subgroup 1), S2 (Subgroup 2), and S3 (Subgroup 3) WT tissues. A two-sided student t test was used to determine the difference between NAT (normal adjacent to tumor), S1, S2 and S3. b Expression of mesenchymal-epithelial transition signature genes in NAT, S1, S2 and S3 WT samples at the mRNA and protein levels. NS p > 0.05, * p < 0.05, ** p < 0.01, *** p < 0.001. A two-sided Wilcoxon rank sum test was performed to test the difference. c Normalized enrichment score (NES) of the kidney development phase-related gene sets in NATs, S1, S2 and S3. The normalized enrichment score was calculated using GSEA function from R clusterProfiler package. A two-sided Kolmogorov–Smirnov (K–S) test was performed to test the statistical significance. The false discovery rate adjustments were made for multiple comparisons. d Normalized enrichment score for TF target genes in WT samples. Bottom: highly expressed TFs in WT. The normalized enrichment score was calculated using GSEA function from R clusterProfiler package. A two-sided Kolmogorov–Smirnov (K–S) test was performed to test the statistical significance. The false discovery rate adjustments were made for multiple comparisons. Source data are provided as a Source Data file.
An analysis of key transcription factors (TFs) across distinct subgroups revealed that TFs associated with CM and RV were highly expressed in WT compared to NATs, particularly in S1 (Fig. 5c, Supplementary Fig. 5c). Moreover, TFs critical for early embryonic kidney development, such as SOX11, SALL1, WT1, and MAZ, were significantly overexpressed in WT samples (Supplementary Data 5b). These TFs exhibited strong correlations with their respective target gene expression within WT, highlighting their potential regulatory roles in WT development (Fig. 5d)41,42,43,44.
Immuno-landscape of WT based on integrated proteogenomic data
To characterize the immune microenvironment of WT, we employed the CIBERSORTx method45 to estimate the relative abundance of immune cells within the tumor tissue based on gene expression data. By comparing our cohort with adult renal carcinoma samples from the TCGA database and WT from the TARGET database, we found that the immune cell proportion of WT samples was significantly lower than that of adult renal carcinoma, like kidney renal clear cell carcinoma (KIRC) and kidney renal papillary cell carcinoma (KIRP) (Fig. 6a). Additionally, we observed no correlation between the immune cell proportion and TMB (Fig. 6b).
a Ridge plot of the immune signature score of WT (Wilms tumor) from our cohort and the TARGET database and 3 adult kinds of kidney tumors from the TCGA database, TCGA-KICH (Kidney Chromophobe), TCGA-KIRC (Kidney Renal Clear Cell Carcinoma), and TCGA-KIRP (Kidney Renal Papillary Cell Carcinoma). b Correlation analysis between tumor mutation burden (TMB) and immune signature score. c Cell abundance of listed immune cells in S1, S2 and S3. Kruskal test was used to determine the difference between subgroups. d RNA (upper) and protein (lower) abundance of chemokines, interferons, interleukins and other cytokines in S1, S2 and S3. e RNA (left) and protein (right) abundance of immune checkpoint genes/proteins in S1, S2 and S3. A two-sided student’s t test was used to determine the difference between subgroups. NS p > 0.05, * p < 0.05, ** p < 0.01, *** p < 0.001. Source data are provided as a Source Data file.
To further dissect the immune microenvironment of WT, we utilized the CIBERSORTx method to estimate the relative abundances of 22 distinct immune cell types. Consistently, WT had relatively lower abundances of most immune cells, including B cells, cytotoxic cells, macrophages and antigen presenting machinery (Supplementary Fig. 6a), suggesting that WT has a more immunosuppressive phenotype compared to adult tumors, similar to other childhood embryonal tumors46,47. Notably, M2 macrophages, gamma-delta T cells, activated mast cells, CD8 T cells, and resting CD4 memory T cells were significantly enriched in S3 subgroup (Fig. 6c, Kruskal-Wallis test, p < 0.05). These findings indicated that the S3 had higher immune cell infiltration and might represent an immune-enriched subtype, which was consistent with immunohistochemistry (IHC) results of CD4, CD8, CD45 and PD-L1 in different subgroups (Supplementary Fig. 6b). Despite the overall low immune infiltration observed in WT compared to adult renal tumors, S3 exhibited a higher immune cell proportion and increased abundance of immune cells, particularly antigen presenting machinery and cytotoxic cells. Moreover, S3 displayed relatively higher expression levels of chemokines, cytokines and interferons than subgroups S1 and S2 (Fig. 6d). Beyond immune infiltration, immunoinhibitory molecules played a crucial role in shaping the immune phenotype of tumors and influencing their responsiveness to immunotherapy. Our findings suggested a potential immune evasion mechanism in S3, as evidenced by significantly elevated expression of immune checkpoint genes, including CD274 and BTN3A1, at both mRNA and protein levels (Fig. 6e). These results highlighted the immunomodulatory characteristics of S3, suggesting that it may be a promising candidate for immunotherapy.
Identification of therapeutic strategies from proteogenomic analyses
Precision medicine plays a crucial role in cancer treatment by selectively targeting oncogenic pathways, including mutations, CNAs, differentially expressed proteins and kinases. Through a comprehensive analysis, we identified a total of 39 potential therapeutic targets (Fig. 7a). Remarkably, several of these candidates were significantly correlated with poor prognosis in WT and could be targeted by FDA-approved drugs (Supplementary Fig. 7a). These genes were enriched in tumor-promoting pathways, such as cell cycle, regulation of TP53 activity, and Wnt signaling pathway (Fig. 7b). Among these potential drug targets, the CDK2-RB1-E2F axis has emerged as a well-studied pathway implicated in driving cell proliferation across various malignancies48. In WT, the activation of this axis was supported by the elevated expression and kinase activity of CDK2, hyperphosphorylation of RB1, and upregulation of E2F target genes, particularly in S1 (Fig. 7c). Moreover, the inhibition of CDK2 with a small molecular inhibitor, BIX-01294 (S8006, Selleck), in a WT cell line led to a reduction in phosphorylated RB1 levels, confirming the regulatory effect of CDK2 on RB1 phosphorylation in WT cells (Supplementary Fig. 7b).
a Potential drug targets with FDA-approved drugs (left) by multi-omics data analysis in the Xinhua-WT cohort and their hazard ratios (right), as well as 95% confidence intervals (CIs), of EFS and OS using the TARGET-WT cohort (n = 125). The forest plots show the 95% confidence interval of hazard ratio with median value, 97.5 and 2.5th quartiles. A two-sided log-rank test was performed to test the difference. b The pathways enriched by the potential drug targets. A one-sided hypergeometric test was used to determine the statistical significance. c Heatmap depicting the kinase activities and RNA/protein expression levels of CDK1 and CDK2, phosphorylation levels of RB1 (pS249 and pS37), and RNA and protein expression levels of E2F target genes. d Western blot of EHMT2 and GADPH expression in WT (Wilms tumor) and NAT (normal adjacent to tumor) samples. N: NATs (n = 18); T: Tumors (n = 18). Western blot was repeated three times independently with similar results. e Western blot of the expression of dimethylated H3K9 and H3 in SK-NEP-1 cells after EHMT2 knockdown. EHMT2 knockdown and Western blot was repeated three times independently with similar results. f mRNA expression of pre-rRNAs after EHMT2 knockdown in SK-NEP-1 cells. A two-sided student t test was performed to test the difference. Data are presented as means \(\pm\) SEM. EHMT2: **** p = 8.1E-05 (NC and siEHMT2-1), **** p = 2.9E-05 (NC and siEHMT2-2). Pre-rRNA_1: * p = 1.2E-02 (NC and siEHMT2-1), ** p = 2.7E-03 (NC and siEHMT2-2). Pre-rRNA_2: ** p = 6.9E-03 (NC and siEHMT2-1), ** p = 2.9E-03 (NC and siEHMT2-2). The false discovery rate adjustments were made for multiple comparisons. Each group has three biological replicates. g Pathway enrichment results of downregulated and upregulated genes after EHMT2 knockdown. h Heatmap of the RNA abundance of differentially expressed genes related to genes controlling nephrogenesis, the cell cycle and the Wnt signaling pathway. i. Western blot of the expression of dimethylated H3K9 and H3 in SK-NEP-1 cells after treatment with the EHMT2 inhibitor at the indicated concentrations. Inhibitor treatment and Western blot was repeated three times independently with similar results. Source data are provided as a Source Data file.
EHMT2, a histone-lysine N-methyltransferase, modulates histone H3 through mono-methylation and di-methylation, functioning as a transcriptional repressor, with implications in hepatocellular carcinoma49 and melanoma50. Interestingly, EHMT2 was highly expressed in S1 at both the mRNA and protein levels (Fig. 7a), and its elevated expression was associated with shortened EFS and OS in the TARGET cohort (Supplementary Fig. 7c). Additionally, EHMT2 was highly expressed in stage II, III, IV and V compared to that in stage I WT samples in database GSE31403 (Supplementary Fig. 7d).
Given the functional significance of EHMT2, we performed in vitro experiments to investigate its functional role and underlying mechanism. First, high expression of EHMT2 in WT was verified by Western blot analysis of 18 paired WT and NAT samples (Fig. 7d, Supplementary Fig. 7e). Second, EHMT2 knockdown using EHMT2-specific small interfering RNA (siRNA) in WT cells resulted in the downregulation of di-methylation levels of H3K9 (Fig. 7e, Supplementary Fig. 7f). Functional assays demonstrated that EHMT2 silencing induced G1 arrest and inhibited cell proliferation (Supplementary Fig. 7g, h, i). Furthermore, qPCR analyses showed that EHMT2 silencing led to a decrease in pre-rRNA levels in WT cells, indicating that EHMT2 was involved in rRNA regulation (Fig. 7f). Moreover, we conducted RNA-seq on WT cells with or without EHMT2 knockdown to assess the impact of EHMT2 on gene expression in WT. The results revealed significant downregulation of genes involved in Wnt signaling, cell cycle regulation, and nephrogenesis, while upregulated genes were associated with autophagy, apoptosis, and programmed cell death (Fig. 7g). Specifically, key regulators of the cell cycle, Wnt/β-catenin signaling, and nephrogenesis were markedly downregulated following EHMT2 inhibition (Fig. 7h), suggesting that EHMT2 may regulate cell proliferation and cell cycle progression through these pathways.
To further elucidate the potential therapeutic targeting of EHMT2 in WT, we treated the WT cell line with different doses of the EHMT2 small molecule inhibitor BIX-01294 for 24 h. As shown in Fig. 7i, H3K9 di-methylation, the primary methylation target of EHMT2, was significantly reduced, while the total histone H3 protein level remained unchanged. These findings indicated the crucial role of EHMT2 in H3K9 dimethylation in WT and highlighted EHMT2 as a promising biomarker and potential drug target for WT.
Discussion
In this study, to obtain a comprehensive molecular characterization of WT, we generated genomic, transcriptomic, proteomic, and phosphoproteomic data of WT and NAT samples, presenting a thorough exploration of molecular attributes, as well as identifying potential diagnostic and prognostic biomarkers and therapeutic targets of WT.
To explore the impact of genomic alterations on mRNA and protein expression, we delved into not only genomic alteration but also the interplay between gene alterations and their corresponding effects on both mRNA and protein levels. In addition to known CNAs correlated with prognosis, such as deletions of 1p, 16q51, 11q, and 11p1552 and gain of 1q, our results revealed that 6p and 12q were frequently co-amplified with 1q and had a strong cis-effect. In addition, amplifications of 1q, 6p and 12q resulted in the activation of cell cycle progression, and genes located in these regions were anti-correlated with OS and EFS in WT. The identification of frequent amplifications in chromosomes 6p and 12, in conjunction with 1q gain, not only broadens our understanding of the genomic landscape of WT but also raises the possibility of utilizing these alterations as practical risk classification indicators. Currently, evidence suggests that patients with 1p/16q LOH might benefit from intensified therapy to regimen DD4A (vincristine, dactinomycin and doxorubicin) or regimen M (vincristine, dactinomycin, and doxorubicin alternating with cyclophosphamide and etoposide), although the underlying mechanisms of 1p/16q LOH remain elusive11,51. However, this intensified treatment approach comes with a significant level of toxicity. Our results revealed significant reprogramming of energetic metabolism at the mRNA and protein levels in samples with 1p LOH and/or 16q LOH. This metabolic reprogramming has been reported to have a close association with cancer progression and chemoresistance, primarily through the Warburg effect, which disrupts normal metabolism and diminishes the metabolic fitness of tumor-infiltrating immune cells53,54,55. Several drugs that inhibit glycolysis are currently undergoing preclinical and clinical studies exploiting the glycolytic activity of tumor cells, such as antiglycolytic drugs targeting glucose transporters or glycolytic enzymes (HK2, GAPDH, LDH-A, or PDK)56,57. Understanding how these pathways contribute to cancer progression and resistance to chemotherapy can pave the way for targeted therapies that disrupt these mechanisms, potentially improving treatment outcomes for WT patients. Last, the association of mutations in the Wnt/β-catenin pathway with a stromal-like signature and mesenchymal phenotype in WT cells underscores the importance of elucidating the molecular underpinnings of specific subgroups of this tumor. This knowledge not only sheds light on potential tumorigenic mechanisms but also has direct clinical relevance, as it opens up the possibility of targeted therapies, such as clinical trials involving small molecule inhibitors such as Tegavivnt58,59, which hold promise for the treatment of WT cases characterized by Wnt/β-catenin pathway mutations. These findings collectively underscore the vital role of understanding the genomic and molecular intricacies of WT in advancing our ability to diagnose, classify, and treat this pediatric malignancy effectively.
The classification of WT is now mainly based on histopathology, while tumor heterogeneity on genomic or gene expression levels within histopathology groups cannot be ignored60. Although most of our WT samples were classified as mixed type by HE and IHC staining, unsupervised clustering based on protein and mRNA expression data classified them into 3 molecular subgroups, each with distinct molecular features, prognostic relevance, and potential targeting strategies. As reported, WT may originate from early stages of embryonic kidney, and our study further suggested different cellular origins and tumorigenesis for the main pathological types in subgroups based on the correspondence between the molecular features of different embryonic kidney developmental stages and the molecular subgroups36,61. Among these three subgroups, S1 was correlated with chromosomal instability, a high proportion of blastemal cells, and a worse prognosis. Furthermore, our results showed that S1 was characterized by the highest frequency of CNAs, hyperactivation of DDR and epigenetic regulation pathways, and the lowest immune infiltration, which was consistent with previous reports that the high expression of DDR-related genes was associated with the lack of immune infiltration in tumor tissues62,63. Combined with mesenchymal features in S2 and immunological features in S3, these findings imply that distinct targeted therapeutic approaches can be tailored according to molecular subgroups, laying the theoretical groundwork for the development of precision therapies in WT. These multifaceted insights collectively underscore the significance of our study in advancing the understanding and treatment of this pediatric malignancy.
Through comprehensive analysis of frequently mutated genes and CNAs with cis effects, differently expressed mRNAs, proteins and kinases, we identified potential therapeutic targets with FDA-approved drugs in WT, especially in S1. These include eribulin for CDK2 in breast cancer64, dasatinib for BRAF in lung cancer65, and trabectedin for PARP1 in sarcomas66. In addition to these well-known drugs targeting common activating pathways, such as the cell cycle and DDR, in malignancies, we also found that epigenetic regulation was highly activated in WT, especially in S1. Further investigation revealed that EHMT2, a histone methyltransferase, was not only amplified at the genomic level and highly expressed at the mRNA, protein, and phosphorylation levels, especially in S1, but also correlated with a worse prognosis, highlighting EHMT2 as a candidate drug target for the study. EHMT2 is highly expressed in a variety of malignant tumors, associated with poor prognosis,s and activates the Wnt/β-catenin signaling pathway through transcriptional repression of APC protein in hepatocellular carcinoma49,67,68. Further experiments in our study confirmed the high expression of EHMT2 in WT and its regulation of H3 histone methylation, Wnt/β-catenin pathway activation, transcription of rDNA, and cell proliferation in the WT cell line, indicating the potential role of EHMT2 inhibition in the treatment of WT. Further validation of the therapeutic potential of EHMT2 as well as other candidates is still needed.
In conclusion, our integrated analysis based on multi-omics data illustrated the regulatory mechanisms of known key genomic events in WT, established molecular subgroups and provided potential biomarkers and drug targeting modalities. However, our conclusions or hypothesis still need further validation in a large cohort and experiments. We believe that the multi-omics dataset of WT and the integrated results demonstrated in this article will become a rich resource for further research on WT and yield additional insights.
Methods
Our research complies with all relevant ethical regulations and was approved by Ethics Committees of Xinhua Hospital Affiliated to Shanghai Jiao Tong University School of Medicine (XHEC-D-2022-119) and Beijing Children’s Hospital (2023-E-187-Y).
Experiment subject
Tumor samples and clinical information
The samples of WT and NAT were surgical specimens of patients with WT who were diagnosed and treated in Xinhua Hospital and Beijing Children’s Hospital. Informed consent of guardians of participants was obtained in all cases for clinical information collection, sample collection and analysis. Sex or gender was not considered in the study design. The sex of participants was determined based on biological sex, and gender information was not involved. The obtained Wilms tumor and kidney tissue samples were placed in liquid nitrogen for quick freezing and then stored in a −80 °C refrigerator for further use. The clinical information of WT patients was obtained by retrospective analysis of clinical electronic medical records, and the prognosis information was obtained through telephone follow-up.
Cell lines
SK-NEP-1 and 293 T cells were purchased from the National Collection of Authenticated Cell Cultures. SK-NEP-1 cells were cultured in McCoy’s 5 A medium (Cat No. 16600082, Thermo Fisher Scientific, CA, USA) with 15% fetal bovine serum (FBS, Cat No. 10270106, Gibco, NY, USA), 100 units/ml penicillin, and 100 μg/ml streptomycin (Cat No. 15140122, Thermo Fisher Scientific). 293 T cells were maintained in DMEM (Cat No. 10-013-CVR, Corning, Corning, NY, USA) with 10% FBS, penicillin, and streptomycin. Cells were all cultured at 37 °C in a 5% CO2 incubator.
Quantitative proteomics analysis
Protein extraction and tryptic digestion
A total of 159 samples were subjected to mass spectrometry experiments, including 88 WT samples and 71 NAT samples, each sample has two technical replicates. The WT and NAT samples were cut to small pieces and washed with PBS. Samples were manually pulverized on ice in the room at 4 °C. The total protein of WT and NAT samples was extracted using 8 M urea lysis solution (8 M urea and 50 mM ammonium bicarbonate) with protease inhibitor (Cat No. 05892970001, Roche, Basel, Switzerland) using tissue homogenizer. After being quantified using the BCA assay, 150 µg of protein was reduced with 5 mM dithiothreitol at 55 °C for 30 min. Cys residues were then alkylated with 15 mM iodoacetamide at room temperature in the dark for 30 min. −20 °C pre-cooled acetone and 4 °C pre-cooled TCA solution (volume ratio: tissue protein lysate: acetone: TCA = 1:8:1) was used to purify protein samples at 4 °C overnight with rotation. The protein precipitation was washed with 0.1% HCl-acetone (once) and acetone (twice), and then air dry. The protein samples (150 μg) were dissolved in 156 μl 50 mM TEAB and 24 μl trypsin solution (100 ng/ul), and then incubated at 37 °C overnight. The digestion was continued with adding 12 μl more of trypsin solution (100 ng/μl) at 37 °C for 4 h.
TMT-label
48 μl (30 μg) of digested peptides from each sample were labeled with 6-plex Tandem Mass Tag (TMT) reagents according to the manufacturer’s instructions (Cat No.90066, Thermo Fisher Scientific). In brief, peptides (30 μg) from each of the samples were mixed with different 10 μl of TMT reagent (0.8 g) that was dissolved freshly in 41 μl of anhydrous acetonitrile. After 1 h incubation at room temperature (800 rpm), 4.8 μl of 5% hydroxylamine was added and incubated for 15 min at room temperature (800 rpm) to quench the reaction. Peptides labeled by different TMT reagents were then mixed into one sample.
Peptide pre-fractionation by high-pH HPLC
Samples were dried using Speed-Vac, resolved in 1 ml of 0.1% trifluoroacetic acid (TFA), and then desalted using a Sep-Pak cartridge according to the manufacturer’s instructions (Waters, Milford, MA, USA). Pre-fractionation by reverse-phase chromatography at high pH into 80 fractions per sample was conducted using Agilent 1260 Infinity II liquid chromatograph (Agilent, CA, USA). In detail, 180 μg of desalted, 6-plex TMT-labeled peptides was reconstituted in 80 μl 2% acetonitrile (pH 10, with ammonium formate), loaded on a 4.6 × 250 mm, Peptide BEH C18 column, 130 A, 5μm (Waters, Framingham, Massachusetts, USA), and then separated on an Agilent 1260 Infinity II liquid chromatograph (Agilent, CA, USA). Peptides were separated using Solvent A (2% acetonitrile, pH 10) and a nonlinear increasing concentration of solvent B (98% acetonitrile). The 85 min separation liquid chromatograph gradient followed this profile: (min: %B) 0:5; 2:12; 10:33; 67:95; 82:95; 85:5. The flow rate was set at 1 mL/min. For each 180 μg separation, 80 fractions were collected and then combined into 20 fractions for further analysis.
Liquid chromatography
An UltiMate 3000 HPLC (Thermo Fisher Scientific) was used to perform online separation. 1/3 of each peptide fraction containing 0.5 μg peptide was dissolved in a 4 μl injection volume with 0.1% formic acid and then injected onto an in-house packed 20 cm x 75 um diameter C18 silica picofrit capillary column (inspire C18 100 A 3 μm, DIKMA, Beijing, China, No. 85111; TSP standard FS Tubing, Polymicro, No. TSP075375). Solvent A was composed of 0.1% formic acid, and solvent B was composed of 80% acetonitrile and 0.1% formic acid. A 75 min LC-MS/MS method was used with the following gradient profile: (min: %B) 0:2; 5:7; 52.5:30; 62:48; 62.5:99; 69.5:99; 70:5; 75:5. The flow rate was set at 300 nL/min.
Mass spectrometry
Samples were analyzed with a Q Exactive™ Plus mass spectrometer (Thermo Fisher Scientific). Data-dependent acquisition was performed using Thermo Scientific Xcalibur v4.2.47 software at a spray voltage of 2 kV. Full MS spectra were measured with a resolution of 70,000, an AGC target of 3e6 and a mass range from 350 to 1800 m/z. dd-MS2 spectra were measured with a resolution of 17,500, an AGC target of 1e6, an isolation window of 1.6 m/z, a maximum injection time of 45 msec.
Protein identification and quantification
Raw MS/MS spectra were searched against the UniProt Knowledgebase for Homo sapiens (download date: 2019-01-06) using Maxquant (Max-Planck-Institute of Biochemistry, Version 1.6.17.0). And intensity-based TMT-6plex labeled quantification was used in protein quantification. The settings of protein quantification and identification were as follows. Trypsin/P was set as the proteolytic enzyme with two missed cleavages permitted. Carbamidomethyl (C) was set as fixed modification and Acetyl (protein N-term) and Oxidation (M) were set as variable modifications. The mass tolerance for precursor ions was set as 20 ppm in First search and 4.5 ppm in Main search, and that for fragment ions was set as 0.5 Da. The minimum peptide length was seven amino acids. Match between runs was enabled.
Phosphoproteomics analysis
Phosphoproteomics was conducted according to published nature protocol, and the brief procedures were as follows69:
Protein extraction and digestion
A total of 46 samples were subjected to mass spectrometry experiments, including 23 WT samples and 23 NAT samples, each sample has 1 technical replicate. The WT and NAT samples were cut to small pieces and washed with PBS. The total proteins of WT and NAT samples were washed with TBS, extracted using SDC lysis buffer (4% SDC and 100 mM Tris-HCl) using tissue homogenizer and then heat-treated for 5 min at 95 °C immediately. After being quantified using the BCA assay, 500 µg of protein were reduced with reduction/alkylation buffer at 45 °C for 5 min. The protein samples were digested with trypsin solution (an enzyme-to-substrate ratio of 1:100) at 37 °C overnight.
Phosphorated peptides enrichment
Add 400 µl of ISO and 100 µl of EP enrichment buffer to each sample in order and mix thoroughly between steps. Then add TiO2 beads (a bead-to-protein ratio of 12:1) resuspended in EP loading buffer and incubated at 40 °C with shaking (2000 r.p.m.) for 5 min. The TiO2 beads were washed with EP wash buffer for five times and then transferred into a C8 StageTip using 150 μl EP transfer buffer, following by centrifugation (1500 g for 8 min at RT). Elute the phosphopeptides with 30 µl of EP elution buffer by centrifugation (1500 g for ~4 min at RT) and dry the elution under vacuum at 45 °C until ≤15 µl of sample remains. The phosphopeptides in remaining EP elution buffer were desalted using SDB-RPS StageTip according to the aforementioned protocol.
Liquid chromatography
An UltiMate 3000 HPLC (Thermo Fisher Scientific) was used to perform online separation. 1/2 of each phosphopeptides was dissolved in a 4 μl injection volume with 0.1% formic acid and then injected onto an in-house packed 20 cm x 75 um diameter C18 silica picofrit capillary column (inspire C18 100 A 3 μm, DIKMA, Beijing, China, No. 85111; TSP standard FS Tubing, Polymicro, No. TSP075375). Solvent A was composed of 0.1% formic acid, and solvent B was composed of 80% acetonitrile and 0.1% formic acid. A 120-minute LC-MS/MS method was used with the following gradient profile: (min: %B) 0:1; 5:6.5; 100:27.5; 105:43.5; 105:99; 110:99; 110.5:5; 120:5. The flow rate was set at 300 nL/min.
Mass spectrometry
Samples were analyzed with a Q Exactive™ Plus mass spectrometer (Thermo Fisher Scientific). Data-dependent acquisition was performed using Thermo Scientific Xcalibur v4.2.47 software at a spray voltage of 2 kV. Full MS spectra were measured with a resolution of 70,000, an AGC target of 3e6 and a mass range from 350 to 1800 m/z. dd-MS2 spectra were measured with a resolution of 35,000, an AGC target of 1e6, an isolation window of 1.6 m/z, a maximum injection time of 110 msec.
Protein identification and quantification
Raw MS/MS spectra were searched against the UniProt Knowledgebase for Homo sapiens (download date: 2019-01-06) using Maxquant (Max-Planck-Institute of Biochemistry, Version 1.6.17.0). And intensity-based label-free quantification was used in protein quantification. The settings of protein quantification and identification were as follows. Trypsin/P was set as the proteolytic enzyme with two missed cleavages permitted. Carbamidomethyl (C) was set as fixed modification and Acetyl (protein N-term), Oxidation (M), Phospho (STY) were set as variable modifications. The mass tolerance for precursor ions was set as 20 ppm in First search and 4.5 ppm in Main search, and that for fragment ions was set as 0.5 Da. The minimum peptide length was seven amino acids.
Whole exome sequencing
The genomic DNA was extracted using DNeasy Blood & Tissue Kit (QIAGEN, Hilden, Germany) according to the manufacturer’s instructions. DNA degradation and contamination were monitored on 1% agarose gels. DNA concentration was measured by Qubit® DNA Assay Kit in Qubit® 2.0 Flurometer (Invitrogen, USA). A total amount of 0.6 μg genomic DNA per sample was fragmented to an average size of 180 ~ 280 bp and subjected to DNA library creation using established Illumina paired end protocols. The Agilent SureSelect Human All ExonV6 Kit (Agilent Technologies, Santa Clara, CA, USA) was used for exome capture according to the manufacturer’s instructions. The Illumina Novaseq platform (Illumina Inc., San Diego, CA, USA) was utilized for genomic DNA sequencing in Personal Biotechnology Co., Ltd (Shanghai, China) to generate 150 bp paired end reads.
Somatic mutation and germline variants detection
The somatic short variants were discovered by VarScan v2.3.970. Briefly, the paired-end reads of WES were aligned to the human reference genome (hg19) with BWA-mem (0.7.17-r1188)71. The bam files were further processed by reordering reads, removing PCR duplicates, and converting the alignments to mpileup format with samtools (v1.4.1)72. Consequently, the single-nucleotide variants (SNVs) and small insertions and deletions (INDELs) were called by VarScan ‘somatic’ mode. The somatic mutations and germline variants with high confidence were identified by VarScan ‘processSomatic’ mode, and annotated by ANNOVAR73 using the databases including refGene, 1000g2012apr_asn, dbnsfp30a, clinvar_20170130, snp138, ljb26_all, exac03nontcga, gnomad_exome, gnomad_genome, and mcap. The mutation data was converted to maf format. The TMB was calculated by dividing the total number of non-silent mutations by the sequenced regions (60 Mb). These analyses were performed by R maftools74 package.
Somatic CNA analysis
Somatic copy number alteration (SCNA) analysis used VCF files that included both somatic and germline variants. In preparation for CNV calling, we retained only the variants where the read depth of normal samples was greater than or equal to 20. The CNV calling was conducted using R saasCNV package75. Specifically, the variant allele frequency (VAF) and read depth (RD) were used to identify segments with potential CNV by joint segmentation. The CNVs were called with a p-value cutoff of 0.05. To retrieve gene-level CN values and identify the significant CNA regions in WT, we performed GISTIC analysis (version 2)76 on Gene Pattern platform77 (https://www.genepattern.org). The SCNAs with false discovery rate (FDR) less than 0.25 were considered as significantly amplified or deleted regions.
The CNV scores were calculated based on the log2 ratios of all segments. Specifically, the absolute log2 ratios of all segments (indicating the CN aberration of these segments) within a chromosome were weighted by the segment length and summed up to derive the instability score for the chromosome. The genome-wide chromosome instability index was derived by summing up the instability score of all 22 autosomes.
RNA extraction, library construction, and sequencing
Total RNA was isolated using the Trizol Reagent (Invitrogen Life Technologies), after which the concentration, quality, and integrity were determined using a NanoDrop spectrophotometer (Thermo Scientific). Three micrograms of RNA were used as input material for the RNA sample preparations. Sequencing libraries were generated using the TruSeq RNA Sample Preparation Kit (Illumina, San Diego, CA, USA). Briefly, mRNA was purified from total RNA using poly-T oligo-attached magnetic beads. Fragmentation was carried out using divalent cations under elevated temperature in an Illumina proprietary fragmentation buffer. First-strand cDNA was synthesized using random oligonucleotides and SuperScript II. Second strand cDNA synthesis was subsequently performed using DNA Polymerase I and RNase H. Remaining overhangs were converted into blunt ends via exonuclease/polymerase activities and the enzymes were removed. After adenylation of the 3′ ends of the DNA fragments, Illumina PE adapter oligonucleotides were ligated to prepare for hybridization. To select cDNA fragments of the preferred 200 bp in length, the library fragments were purified using the AMPure XP system (Beckman Colter, Beverly, CA, USA). DNA fragments with ligated adapter molecules on both ends were selectively enriched using Illumina PCR Primer Cocktail in a 15-cycle PCR reaction. Products were purified (AMPure XP system) and quantified using the Agilent high-sensitivity DNA assay on a Bioanalyzer 2100 system (Agilent). The sequencing library was then sequenced on a Hiseq platform (Illumina) by Shanghai Personal Biotechnology Cp. Ltd.
Gene expression quantification and normalization
We preprocessed the raw reads of RNA-seq using fastp with default options, removing low-quality reads. Subsequently, we aligned them to the hg19 reference genome using HISAT v2.2.178 and annotated the genes using GENCODE v19 gene annotation79. Next, we converted the SAM files to BAM files and sorted the reads using SAMtools v1.4.1. Consequently, we quantified gene expression using StringTie v2.2.080. The gene expression was normalized to Fragments Per Kilobase Million (FPKM) by R ballgown v2.32.0 package81.
Normalization of proteomic and phospho-proteomic data
The proteomic data were normalized by internal reference scaling (IRS) method following the previous study82. Briefly, a global scaling value is first computed for each sample to represent the average expression value across all samples. The raw data is then sample-loading normalized within each sample using the sample-loading normalization factors to ensure uniform total expression values. Subsequently, the Trimmed Mean of M values (TMM) method is employed to normalize using the SL-normalized data. Furthermore, we computed the average protein intensities across the 6 samples as the reference intensity for each TMT experiment using TMM-normalized data. We computed the average reference intensity across all TMT experiments and adjusted the reference value in all TMT experiments. The IRS normalization factors were calculated by dividing the reference intensities by the average reference intensity for the TMT experiments. For each TMT experiment, the expression data was IRS-normalized by multiplying the TMM-normalized data with the IRS normalization factor. Consequently, the ComBat method in R SVA package83 was employed to remove the batch effect between the TMT experiments.
Tumor versus normal differential transcriptomic, proteomic, and phospho-proteomic analyses
The transcriptomics, proteomics, and phosphor-proteomics data were used to perform differential expression analysis between WT and NAT samples. The standardized expression profiles of transcriptome, proteome, and phosphor-proteome were logarithmically transformed with base 2. Subsequently, the R limma package84 was employed to compare the WT and NAT tissues, resulting in the identification of differentially expressed genes (adjusted p < 0.05 and fold change > 2), proteins (adjusted p < 0.05 and fold change > 1.2), and phosphorylation sites (adjusted p < 0.05 and fold change > 2). The p-values were adjusted by the Benjamini & Hochberg method.
Gene-wise correlation between transcriptomic and proteomic data
We calculated gene-wise correlations between transcriptomic and proteomic data in WT samples and NAT samples, respectively. Specifically, we included the genes detected in all WT or NAT samples at both the transcriptome and proteome levels in this analysis. Spearman’s correlation between transcriptomic and proteomic data was calculated for each gene across all the samples.
Multiomics-based subgroup identification in WT
The WT subgroups based on the mutli-omics data were identified by R CancerSubtypes package85. Briefly, we selected 5000 genes and 4000 proteins using a ‘topk’ approach based on the log-transformed gene expression data and protein expression data. Subsequently, we executed a spectral clustering-based similarity network fusion (SNF) analysis on the selected gene expression and protein expression data. We used specific parameters, including clusterNum = 3, K = 55, alpha = 0.5, t = 20, maxK = 5, pItem = 0.9, reps = 500, finalLinkage = ‘average’ for the SNF analysis. We selected the maximal cluster number that has an average consensus score over 0.8 as the optimal cluster number, which was calculated by R ConsensusPlus package.
Impact of CNA on gene and protein abundance
To investigate the impact of copy number variation on mRNA and protein expression, we employed the R package multiOmicsViz (https://www.bioconductor.org/packages/release/bioc/html/multiOmicsViz.html) to carry out a correlation analysis between CNA and mRNA/protein expression. Specifically, we first identified significant CNV genes based on a threshold (greater than 0.4 or less than −0.4) in at least 10% samples (n = 36). Subsequently, multiOmicsViz function in the R package was used to calculate the Spearman correlation and visualize the significant CNA-mRNA/protein pairs. The p-values were adjusted for multiple-testing correction by applying the Benjamini-Hochberg procedure. The significant CNA-mRNA/protein pairs were identified if the adjusted p-values < 0.25.
Kinase activity analysis
The kinase activity analysis is divided into the identification of tumor/subgroup-related kinases and the estimation of kinase activity. First, sites annotated as activating kinase activity in Signor25 and those in PhosphoSitePlus26 were used for this analysis. To identify the cancer- and subgroup-associated kinases, we excluded the sites with a significant protein expression change between WT and NAT samples or between the WT subgroups. The retained sites were ordered by the statistics for differential expression analysis and were subjected to phosphosite set enrichment analysis based on the kinase-phosphosite relationships by R clusterProfiler (GSEA function)86. In addition, kinase activity was estimated following the method of single-sample gene set enrichment analysis. The enrichment scores based on the phosphosite abundance and kinase-phosphosite relationships were calculated by R GSVA package87 and used as the relative activities of kinases.
Subgroup prediction in TARGET cohort
To demonstrate the performance of our multi-omics-based subgroups on WT samples with longer follow-up time, we collected publicly available gene expression profiles and clinical information from the TARGET dataset (n = 125). First, we used the signature genes of our subgroups identified by both RNA-seq and proteomics data as the template of the nearest template prediction (NTP) algorithm33. Next, based on these signature genes, the NTP algorithm was applied to predict the subgroups of WT patients from the TARGET cohort.
Subgroup-specific RNA and protein identification
The subgroup-specific RNA and proteins were identified by comparing any one of the three subgroups against the other two subgroups using the R limma (linear modeling for microarray data) method84 to conduct differential analysis. RNAs or proteins that were upregulated in this comparison were considered subgroup-specific features (adjusted p < 0.05, and fold change > 1.5 for RNAs and >1.2 for proteins).
Quantification of immune and stromal cell infiltration
We integrated and performed batch correction on pan-renal cancer data from multiple sources including Xinhua-WT, TARGET-WT, TCGA-KICH, TCGA-KIRC, and TCGA-KIRP cohorts. Next, we identified a common set of genes shared across RNA expression datasets and aggregated gene expression data. We applied batch correction using the ComBat method83 to address batch effects. The batch-corrected gene expression data were saved in a GCT file. Finally, we estimated immune and stromal scores for the samples of pan-renal cancer using R ESTIMATE package88. In addition, we estimated the immune cell proportion using bulk RNA-seq data by CIBERSORTx method45.
Functional enrichment analysis of signature RNAs/proteins/phospho-proteins
To gain further insight into biological implications, we performed functional enrichment analysis of the signature RNAs/proteins/phospho-proteins, which was carried out in R clusterProfiler package86 using the Fisher’s exact test with an adjusted p-value cutoff of 0.05. The signature RNAs/proteins/phospho-proteins were unified to official gene symbols, and the enriched pathways were obtained from MSigDB C289 using R msigdbr package.
Statistics and reproducibility
Quantification methods and statistical analysis methods for proteomic, genomic, transcriptomic and integrated analyses were mainly described above. The statistical tests used for each analysis and whether they were one-sided or two-sided were indicated in the respective figure legends. No statistical method was used to predetermine sample size.
Functional experiments
siRNA interference
Scramble siRNA and different specific siRNAs targeting EHMT2, were chemically synthesized (RiboBio). SK-NEP-1 cells and Wit49 cells were seeded in six-well plates at 40% confluency and transfected with siRNAs using RFectSP siRNA transfection reagent (Cat No. 11025, Changzhou Bio-generating Biotechnologies Corp., Changzhou, China) or Lipofectamine 2000 transfection reagent (Cat No. 11668019, Thermo Fisher Scientific, CA, USA) according to the manufacturer’s instructions, respectively.
The target sequences were as follows:
siEHMT2-1: GAGTGATGATGTCCACTCA
siEHMT2-2: CTCCAGGAATTTAACAAGA
Western Blot
Protein extraction and Western Blot were conducted as previously described90. In brief, protein samples were loaded onto a polyacrylamide gel for electrophoresis and then electrotransferred onto polyvinylidene difluoridemembranes (Bio-Rad). Blots were blocked with 5% bovine serum albumin at room temperature for 2 h and incubated with the selectedprimary antibodies at 4 °C overnight. After washing the membranes forthree times, the membranes were incubated with second antibody for1 h at room temperature. Bands were visualized by electrogeneratedchemiluminescence (Pierce Biotechnology) with the Bio-Rad ChemiDoc XRS imaging system. Primary antibodies specific for GAPDH (Cat No. 5174, 1:1000), RB (Cat No. 9309, 1:1000), p-RB S780 (Cat No. 8180, 1:1000), p-RB S795 (Cat No. 9301, 1:1000), p-RB S807/811 (Cat No. 8516, 1:1000), CDK2 (Cat No. 2546, 1:1000), Histone H3 (Cat No. 4499, 1:1000), Di-Methyl-Histone H3 (Lys9) (Cat No. 4658, 1:1000), and EHMT2 (Cat No. 3306, 1:1000) were purchased from Cell Signaling Technology (Beverly, USA). The results of Western Blot were quantified using ImageJ (v 1.8.0).
RNA isolation and qPCR
TRIzol (Cat No. T9108, Takara, Beijing, China) was used to isolate total RNAs, and a PrimeScript™ RT reagent Kit (Cat No. RR037A, Takara, Beijing, China) was used to reverse-transcribe RNA products into cDNAs. Quantitative real-time PCR was performed using SYBR Green Master Mix (Cat No. 11198ES03, Yeasen, Shanghai, China) and specific primers produced by GENEWIZ (Suzhou, China) as follows.
EHMT2-Forward: 5’-GAGAACATCTGCCTGCACTG-3’
EHMT2-Reverse: 5’-GTTGACAGCATGGAGGTCAC-3’
GAPDH-Forward: 5’-CATGAGAAGTATGACAACAGCCT-3’
GAPDH-Reverse: 5’-AGTCCTTCCACGATACCAAAGT-3’
Pre_rRNA_1-Forward: 5’-CCTGCTGTTCTCTCGCGCGTCCGAG-3’
Pre_rRNA_1-Reverse: 5’-AACGCCTGACACGCACGGCACGGAG-3’
Pre_rRNA_2-Forward: 5’-GAACGGTGGTGTGTCGTTC-3’
Pre_rRNA_2-Reverse: 5’-GCGTCTCGTCTCGTCTCACT-3’
5-Ethynyl-2’-deoxyuridine (EdU) incorporation assay
The EdU incorporation assay was performed using a Cell-Light EdU Apollo488 In Vitro Kit (Cat No. C10310-3, RiboBio, Guangzhou, China). Cells transfected with siRNAs were cultured for 72 h or treated with inhibitors for indicated times, and then treated with complete culture medium containing 50 μM EdU for 2 h before harvest. After incubation, cells were centrifuged at 1000 rpm for 3 min and then washed with 1× PBS. After that, cells were fixed using 4% paraformaldehyde for 20 min at room temperature, and the fixation was stopped by incubation with 2 mg/ml glycine for 5 min. Cells were then washed with PBS and permeabilized with 0.5% Triton X-100 for 10 min. After washing cells with PBS, Apollo staining solution (Fluor 488) was used for DNA staining. Cells were then washed 1–3 times with 0.5% Triton X-100 and resuspended with 1× PBS before being subjected to flow cytometry using CytoFLEX flow cytometer (Beckman Colter life science, CA, USA) according to the manufacturer’s instructions within 24 h.
Cell cycle analysis
Cell cycle analysis was conducted using a Cell Cycle and Apoptosis Analysis Kit (Cat No. 40301ES50, Yeasen, Shanghai, China). After siRNA transfection for 72 h or treatment with inhibitors for indicated times, cells were collected and washed with 1× PBS, and fixed with 70% cold ethanol at 4 °C overnight. Cells were then cultured with staining solution (0.5 ml of staining buffer with 10 μl of propidium iodide and 10 μl of Rnase A) for 30 min at 37 °C in dark before being sieved through a 400-mesh screen and subjected to flow cytometry using CytoFLEX flow cytometer (Beckman Colter life science, CA, USA) according to the manufacturer’s instructions. The data were then processed using FlowJo LLC (https://www.flowjo.com/solutions/flowjo/).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The mass spectrometry proteomics data generated in this study have been deposited in the ProteomeXchange Consortium via the iProX partner repository91,92 under accession code PXD063650. The raw data generated in this study regarding genome and transcriptome of WT and NAT samples have been deposited in the Genomic Sequence Archive (GSA) for Human database93,94 under accession code HRA005718 [https://ngdc.cncb.ac.cn/gsa-human/browse/HRA005718]. The raw sequencing data are available under controlled access due to data privacy laws related to patient consent for data sharing and the data should be used for research purposes only. Access can be obtained by approval via their respective DAC (Data Access Committees) in the GSA-human database. According to the guidelines of GSA-human, all non-profit researchers are allowed access to the data and the Principle Investigator of any research group is allowed to apply for Controlled access of the data. The user can register and login to the GSA database website (https://ngdc.cncb.ac.cn/gsa-human/) and follow the guidance of “Request Data” to request the data step by step (https://ngdc.cncb.ac.cn/gsa-human/document). The approximate response time for accession requests is about 2 weeks. The access authority can be obtained for Research Use Only. The user can also contact the corresponding author directly. Once access has been granted, the data will be available to download for 3 months. The transcriptomic data of the WT cell line in this study have been deposited in the GEO (Gene Expression Omnibus) under accession code GSE298590. The public datasets of TARGET cohort were downloaded from GDC data portal (https://portal.gdc.cancer.gov/). The gene expression data of GSE31403 dataset was downloaded from Gene Expression Omnibus. Source data are provided with this paper.
Code availability
The code for data analysis and processed data of mutation, mRNA, protein, and phosphoprotein, as well as RNA-seq data of WT cell line can be accessed at github (https://github.com/zhangli-tools/Wilms-tumor) and Zenodo (https://doi.org/10.5281/zenodo.15542663, https://zenodo.org/records/15542663)95. The repository contains all scripts necessary to reproduce the analysis presented in this paper. There are no restrictions on use beyond those described in the MIT License. The code depends on publicly available R packages such as Seurat, dplyr, ggplot2, etc., each distributed under their respective open-source licenses.
References
Breslow, N., Olshan, A., Beckwith, J. B. & Green, D. M. Epidemiology of Wilms tumor. Med. Pediatr. Oncol. 21, 172–181 (1993).
Nakata, K. et al. Incidence of childhood renal tumours: an international population-based study. Int J. Cancer 147, 3313–3327 (2020).
Ni, X. et al. Socioeconomic inequalities in cancer incidence and access to health services among children and adolescents in China: a cross-sectional study. Lancet 400, 1020–1032 (2022).
Wilms Tumor and Other Childhood Kidney Tumors Treatment (PDQ(R)): health Professional Version. In: PDQ Cancer Information Summaries) (PDQ, 2002).
Ooms, A. H. A. G. et al. Significance of TP53 mutation in wilms tumors with diffuse anaplasia: a report from the Children’s Oncology Group. Clin. Cancer Res. 22, 5582–5591 (2016).
Deng, C., Dai, R., Li, X. & Liu, F. Genetic variation frequencies in Wilms’ tumor: a meta-analysis and systematic review. Cancer Sci. 107, 690–699 (2016).
Gadd, S. et al. A Children’s Oncology Group and TARGET initiative exploring the genetic landscape of Wilms tumor. Nat. Genet 49, 1487–1494 (2017).
Huff, V. Wilms’ tumours: about tumour suppressor genes, an oncogene and a chameleon gene. Nat. Rev. Cancer 11, 111–121 (2011).
Treger, T. D., Chowdhury, T., Pritchard-Jones, K. & Behjati, S. The genetic changes of Wilms tumour. Nat. Rev. Nephrol. 15, 240–251 (2019).
Grobner, S. N. et al. The landscape of genomic alterations across childhood cancers. Nature 555, 321–327 (2018).
Dix, D. B. et al. Augmentation of therapy for combined loss of heterozygosity 1p and 16q in favorable histology Wilms Tumor: a Children’s Oncology Group AREN0532 and AREN0533 study report. J. Clin. Oncol. 37, 2769–2777 (2019).
Gratias, E. J. et al. Association of chromosome 1q gain with inferior survival in favorable-histology Wilms Tumor: a report from the Children’s Oncology Group. J. Clin. Oncol. J. Am. Soc. Clin. Oncol. 34, 3189–3194 (2016).
Lorenzo, A. & Iglesias Lopes, R. Faculty Opinions recommendation of Gain of 1q as a prognostic biomarker in Wilms tumors (wts) treated with preoperative chemotherapy in the international society of paediatric oncology (SIOP) WT 2001 trial: A SIOP renal tumours biology consortium study. In: Faculty Opinions – Post-Publication Peer Review of the Biomedical Literature). (Faculty Opinions Ltd., 2017).
Rodriguez, H. & Pennington, S. R. Revolutionizing precision oncology through collaborative proteogenomics and data sharing. Cell 173, 535–539 (2018).
Dagogo-Jack, I. & Shaw, A. T. Tumour heterogeneity and resistance to cancer therapies. Nat. Rev. Clin. Oncol. 15, 81–94 (2018).
Bekker-Jensen, D. B. et al. An optimized shotgun strategy for the rapid generation of comprehensive human proteomes. Cell Syst. 4, 587–599 (2017).
Hein, M. Y. et al. A human interactome in three quantitative dimensions organized by stoichiometries and abundances. Cell 163, 712–723 (2015).
Clark, D. J. et al. Integrated proteogenomic characterization of clear cell renal cell carcinoma. Cell 180, 207 (2020).
Ortiz, M. V. et al. Prohibitin is a prognostic marker and therapeutic target to block chemotherapy resistance in Wilms’ tumor. JCI Insight 4, e127098 (2019).
Zhang, Q., Wang, J., Dong, R., Yang, S. & Zheng, S. Identification of novel serum biomarkers in child nephroblastoma using proteomics technology. Mol. Biol. Rep. 38, 631–638 (2010).
Zhang, J. et al. Screening and identification of non-inflammatory specific protein markers in Wilms’ tumor tissues. Arch. Biochem. Biophys. 676, 108112 (2019).
Ellis, M. J. et al. Connecting genomic alterations to cancer biology with proteomics: the NCI clinical proteomic tumor analysis consortium. Cancer Discov. 3, 1108–1112 (2013).
Wang, L. B. et al. Proteogenomic and metabolomic characterization of human glioblastoma. Cancer Cell 39, 509–528.e520 (2021).
Li, Y. et al. Histopathologic and proteogenomic heterogeneity reveals features of clear cell renal cell carcinoma aggressiveness. Cancer Cell 41, 139–163.e117 (2023).
Lo Surdo, P. et al. SIGNOR 3.0, the SIGnaling network open resource 3.0: 2022 update. Nucleic Acids Res. 51, D631–D637 (2023).
Hornbeck, P. V. et al. PhosphoSitePlus: a comprehensive resource for investigating the structure and function of experimentally determined post-translational modifications in man and mouse. Nucleic Acids Res. 40, D261–D270 (2012).
Ooms, A. et al. Renal tumors of childhood-a histopathologic pattern-based diagnostic approach. Cancers 12, 729 (2020).
Bamford, S. et al. The COSMIC (Catalogue of Somatic Mutations in Cancer) database and website. Br. J. Cancer 91, 355–358 (2004).
Li, S. S. et al. Targeting the Wnt/beta-catenin signaling pathway as a potential therapeutic strategy in renal tubulointerstitial fibrosis. Front. Pharm. 12, 719880 (2021).
Feng, Y. et al. Wnt/beta-catenin-promoted macrophage alternative activation contributes to kidney fibrosis. J. Am. Soc. Nephrol. 29, 182–193 (2018).
Edeling, M., Ragi, G., Huang, S., Pavenstadt, H. & Susztak, K. Developmental signalling pathways in renal fibrosis: the roles of Notch, Wnt and Hedgehog. Nat. Rev. Nephrol. 12, 426–439 (2016).
Munoz-Felix, J. M. & Martinez-Salgado, C. Dissecting the involvement of Ras GTPases in kidney fibrosis. Genes 12, 800 (2021).
Hoshida, Y. Nearest template prediction: a single-sample-based flexible class prediction with confidence assessment. PLoS ONE 5, e15543 (2010).
Metsuyanim, S. et al. Accumulation of malignant renal stem cells is associated with epigenetic changes in normal renal progenitor genes. Stem Cells 26, 1808–1817 (2008).
Dekel, B. et al. Multiple imprinted and stemness genes provide a link between normal and tumor progenitor cells of the developing human kidney. Cancer Res. 66, 6040–6049 (2006).
Li, H., Hohenstein, P.& Kuure, S. Embryonic kidney development, stem cells and the origin of Wilms Tumor. Genes 12, 318 (2021).
Young, M. D. et al. Single-cell transcriptomes from human kidneys reveal the cellular identity of renal tumors. Science 361, 594–599 (2018).
Young, M. D. et al. Single cell derived mRNA signals across human kidney tumors. Nat. Commun. 12, 3896 (2021).
Horster, M. F., Braun, G. S. & Huber, S. M. Embryonic renal epithelia: induction, nephrogenesis, and cell differentiation. Physiol. Rev. 79, 1157–1191 (1999).
Chaffer, C. L., Thompson, E. W. & Williams, E. D. Mesenchymal to epithelial transition in development and disease. Cells Tissues Organs 185, 7–19 (2007).
Tham, M. S. & Smyth, I. M. Cellular and molecular determinants of normal and abnormal kidney development. Wiley Interdiscip. Rev. Dev. Biol. 8, e338 (2019).
Miao, Z. et al. Single cell regulatory landscape of the mouse kidney highlights cellular differentiation programs and disease targets. Nat. Commun. 12, 2277 (2021).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Liberzon, A. et al. The Molecular signatures database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Newman, A. M. et al. Determining cell type abundance and expression from bulk tissues with digital cytometry. Nat. Biotechnol. 37, 773–782 (2019).
Brohl, A. S. et al. Immuno-transcriptomic profiling of extracranial pediatric solid malignancies. Cell Rep. 37, 110047 (2021).
Boldrini, R. et al. Tumor-infiltrating T cells and PD-L1 expression in childhood malignant extracranial germ-cell tumors. Oncoimmunology 8, e1542245 (2019).
Vasaikar, S. et al. Proteogenomic analysis of human colon cancer reveals new therapeutic opportunities. Cell 177, 1035–1049 (2019).
Guo, Y. et al. EHMT2 promotes the pathogenesis of hepatocellular carcinoma by epigenetically silencing APC expression. Cell Biosci. 11, 152 (2021).
Kato, S. et al. Gain-of-function genetic alterations of G9a drive oncogenesis. Cancer Discov. 10, 980–997 (2020).
Grundy, P. E. et al. Loss of heterozygosity for chromosomes 1p and 16q is an adverse prognostic factor in favorable-histology Wilms tumor: a report from the National Wilms Tumor Study Group. J. Clin. Oncol. 23, 7312–7321 (2005).
Fernandez, C. V. et al. Clinical outcome and biological predictors of relapse after nephrectomy only for very low-risk wilms tumor: a report From Children’s Oncology Group AREN0532. Ann. Surg. 265, 835–840 (2017).
Li, Z. & Zhang, H. Reprogramming of glucose, fatty acid and amino acid metabolism for cancer progression. Cell Mol. Life Sci. 73, 377–392 (2016).
Tan, Y. et al. Metabolic reprogramming from glycolysis to fatty acid uptake and beta-oxidation in platinum-resistant cancer cells. Nat. Commun. 13, 4554 (2022).
Xia, L. et al. The cancer metabolic reprogramming and immune response. Mol. Cancer 20, 28 (2021).
Abdel-Wahab, A. F., Mahmoud, W. & Al-Harizy, R. M. Targeting glucose metabolism to suppress cancer progression: prospective of anti-glycolytic cancer therapy. Pharm. Res. 150, 104511 (2019).
Jagust, P., de Luxan-Delgado, B., Parejo-Alonso, B. & Sancho, P. Metabolism-based therapeutic strategies targeting cancer stem cells. Front. Pharm. 10, 203 (2019).
Walz, A. L. et al. Tumor biology, biomarkers, and liquid biopsy in pediatric renal tumors. Pediatr. Blood Cancer 70, e30130 (2023).
Spreafico, F. et al. Wilms tumour. Nat. Rev. Dis. Prim. 7, 75 (2021).
Cresswell, G. D. et al. Intra-tumor genetic heterogeneity in wilms tumor: clonal evolution and clinical implications. EBioMedicine 9, 120–129 (2016).
Wang, P. et al. Dissecting the global dynamic molecular profiles of human fetal kidney development by single-cell RNA sequencing. Cell Rep. 24, 3554–3567.e3553 (2018).
Higgs, E. F., Bao, R., Hatogai, K. & Gajewski, T. F. Wilms tumor reveals DNA repair gene hyperexpression is linked to lack of tumor immune infiltration. J. Immunother. Cancer 10, e004797 (2022).
Kciuk, M. et al. PD-1/PD-L1 and DNA damage response in cancer. Cells 12, 530 (2023).
Kaklamani, V. G. et al. Phase II neoadjuvant clinical trial of carboplatin and eribulin in women with triple negative early-stage breast cancer (NCT01372579). Breast Cancer Res. Treat. 151, 629–638 (2015).
Sen, B. et al. Kinase-impaired BRAF mutations in lung cancer confer sensitivity to dasatinib. Sci. Transl. Med. 4, 136ra170 (2012).
Grignani, G. et al. Trabectedin and olaparib in patients with advanced and non-resectable bone and soft-tissue sarcomas (TOMAS): an open-label, phase 1b study from the Italian Sarcoma Group. Lancet Oncol. 19, 1360–1371 (2018).
Nachiyappan, A., Gupta, N. & Taneja, R. EHMT1/EHMT2 in EMT, cancer stemness and drug resistance: emerging evidence and mechanisms. FEBS J. 289, 1329–1351 (2022).
Saha, N. & Muntean, A. G. Insight into the multi-faceted role of the SUV family of H3K9 methyltransferases in carcinogenesis and cancer progression. Biochim Biophys. Acta Rev. Cancer 1875, 188498 (2021).
Humphrey, S. J., Karayel, O., James, D. E. & Mann, M. High-throughput and high-sensitivity phosphoproteomics with the EasyPhos platform. Nat. Protoc. 13, 1897–1916 (2018).
Koboldt, D. C. et al. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 22, 568–576 (2012).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Mayakonda, A., Lin, D. C., Assenov, Y., Plass, C. & Koeffler, H. P. Maftools: efficient and comprehensive analysis of somatic variants in cancer. Genome Res. 28, 1747–1756 (2018).
Zhang, Z. & Hao, K. SAAS-CNV: a joint segmentation approach on aggregated and allele specific signals for the identification of somatic copy number alterations with next-generation sequencing data. PLoS Comput. Biol. 11, e1004618 (2015).
Mermel, C. H. et al. GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol. 12, R41 (2011).
Reich, M. et al. GenePattern 2.0. Nat. Genet 38, 500–501 (2006).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Frankish, A. et al. Gencode 2021. Nucleic Acids Res. 49, D916–D923 (2021).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Frazee, A. C. et al. Ballgown bridges the gap between transcriptome assembly and expression analysis. Nat. Biotechnol. 33, 243–246 (2015).
Plubell, D. L. et al. Extended multiplexing of tandem mass tags (TMT) labeling reveals age and high fat diet specific proteome changes in mouse epididymal adipose tissue. Mol. Cell Proteom. 16, 873–890 (2017).
Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E. & Storey, J. D. The SVA package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28, 882–883 (2012).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Xu, T. et al. CancerSubtypes: an R/Bioconductor package for molecular cancer subtype identification, validation and visualization. Bioinformatics 33, 3131–3133 (2017).
Wu, T. et al. clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innovation 2, 100141 (2021).
Hanzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinforma. 14, 7 (2013).
Yoshihara, K. et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat. Commun. 4, 2612 (2013).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Cheng, C. et al. P300 Interacted With N-Myc and regulated its protein stability via altering its post-translational modifications in neuroblastoma. Mol. Cell Proteom. 22, 100504 (2023).
Chen, T. et al. iProX in 2021: connecting proteomics data sharing with big data. Nucleic Acids Res. 50, D1522–D1527 (2022).
Ma, J. et al. iProX: an integrated proteome resource. Nucleic Acids Res. 47, D1211–D1217 (2019).
Chen, T. et al. The genome sequence archive family: toward explosive data growth and diverse data types. Genomics Proteom. Bioinforma. 19, 578–583 (2021).
Members, C.-N. & Partners Database resources of the national genomics data center, china national center for bioinformation in 2024. Nucleic Acids Res. 52, D18–D32 (2024).
Cheng, C. et al. Integrative proteogenomic characterization of Wilms tumor. Zenodo https://doi.org/10.5281/zenodo.15542663 (2025).
Acknowledgements
This work was carried out under the National Natural Science Foundation of China (No. 82472715) and the National Key Research and Development Plan Project of China (No. 2022YFC2705000) to Z.W., the Medical-engineering Crossing Research Program of the Shanghai Jiao Tong University (No. YG2023ZD11) to Z.W. and J.L., Shanghai Science and Technology Committee (No. 20YF1430300) to K.C., the Medical-engineering crossing research program of the Shanghai Jiao Tong University (No. YG2021QN46) to Q.C., Beijing Municipal Public Welfare Development and Reform Pilot Project for Medical Research Institutes (No. JYY2023-4) to X.N.
Author information
Authors and Affiliations
Contributions
Conceptualization, C.C., L.Z., X.C., Y.W., Y.J., and Z.W.; Methodology, C.C., L.Z., K.C., J.S., Q.C. and D.R.; Validation, C.C., L.Z., X.C., T.H., L.P., and Y.W.; Formal analysis, C.C., L.Z., X.C., Y.G., W.Z., H.W., T.S. and X.N.; Investigation, C.C., L.Z., X.C., Y.G. and D.R.; Resources, K.C., T.H., J.S., F.L., L.P., Y.W., and Q.C.; Data curation, C.C. and L.Z.; Writing, C.C., L.Z., X.C., K.C., Y.J., and Z.W.; Visualization, C.C., L.Z., K.C., J.L., Y.J., and Z.W.; Supervision, H.W., T.S., X.N., Y.W., J.L., Y.J., and Z.W.; Funding acquisition, Q.C., K.C., Y.W., X.N., J.L., and Z.W.
Corresponding authors
Ethics declarations
Competing interests
All authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Gabriel Malouf, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Cheng, C., Zhang, L., Chang, X. et al. Integrative proteogenomic characterization of Wilms tumor. Nat Commun 16, 7715 (2025). https://doi.org/10.1038/s41467-025-62234-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-62234-7