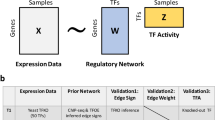
Abstract
Transcription factors (TFs) and transcriptional coregulators are emerging therapeutic targets. Gene regulatory networks (GRNs) can evaluate pharmacological agents and identify drivers of disease, but methods that rely solely on gene expression often neglect post-transcriptional modulation of TFs. We present Epiregulon, a method that constructs GRNs from single-cell ATAC-seq and RNA-seq data for accurate prediction of TF activity. This is achieved by considering the co-occurrence of TF expression and chromatin accessibility at TF binding sites in each cell. ChIP-seq data allows motif-agonistic activity inference of transcriptional coregulators or TF harboring neomorphic mutations. Epiregulon accurately predicted the effects of AR inhibition across different drug modalities including an AR antagonist and an AR degrader, delineated the mechanisms of a SMARCA4 degrader by identifying context-dependent interaction partners, and prioritized drivers of lineage reprogramming and tumorigenesis. By mapping gene regulation across various cellular contexts, Epiregulon can accelerate the discovery of therapeutics targeting transcriptional regulators.
Similar content being viewed by others
Introduction
Transcription factors and transcriptional coregulators shape cell fates and lineage commitment, and their dysregulation drives congenital diseases and tumor growth. TFs bind to specific DNA sequences, whereas coregulators interact with TFs in a context-specific manner and lack defined motifs. These transcriptional modulators represent an important class of therapeutic targets in oncology and beyond. Current therapeutics targeting transcriptional regulators inhibit their activity by either blocking the ligand binding domains1, degrading the protein2,3 or disrupting protein–protein interactions4. Delineating the functions of transcriptional regulators can accelerate our understanding of disease biology and drug discovery.
Gene regulatory networks model the underlying circuitry of gene regulation and have been applied to understand lineage commitment, plasticity and drug response5. Early methods to construct gene regulatory networks relied exclusively on gene expression data and sought to identify an association between the expression of TFs and their target genes6,7. Now, single-cell multiomics technologies provide chromatin accessibility information in addition to gene expression, and recent GRN inference methods leverage both modalities to improve performance. Most of these methods, including CellOracle8, FigR9, Pando10 and GRaNIE11, rely on linear relationships between the mRNA levels of transcription factors and their target genes to model gene regulation. In addition, SCENIC+ utilizes random forest regression to model the regulatory relationships between TFs, regulatory elements and target genes12.
Several challenges still impede the broader utilization of these GRN inference methods in basic biology and drug discovery. First, none of these methods were specifically designed to predict changes in which the TF activity is decoupled from its gene expression. These include drug perturbations that disrupt protein complex formation or localization, post-translational modifications that can impact TF activity and genetic alterations (e.g. neomorphic mutations and CRISPR genome editing) that can silence TF function or add new functions without changing gene expression. Second, the use of motif sequences in GRN construction precludes activity inference of transcriptional coregulators, an important class of regulators without defined motifs. This is problematic for methods that use motifs to filter target genes based on putative TF binding sites.
Here, we present Epiregulon, a method that constructs GRNs from single-cell multiomics and TF occupancy data to infer activity of transcriptional regulators. Epiregulon uses the co-occurrence of TF mRNA expression and chromatin accessibility at TF binding sites to accurately determine the relevance of potential target genes in a given biological context. Unlike most other GRN tools, Epiregulon also leverages ChIP-seq data to infer the activity of TFs and transcriptional coregulators lacking defined motifs. Functional interpretation of the built GRN is performed by computing the Jaccard similarity of target genes and known pathway annotations. We applied Epiregulon to several real and simulated datasets where it successfully recovered the ground truth, detected novel regulators, and matched or outperformed existing GRN methods. Our results indicate that Epiregulon is well-suited to infer single-cell TF activity in the context of drug perturbation and lineage reprogramming. Epiregulon is implemented as a suite of open-source R packages that are available from the Bioconductor project.
Results
Epiregulon constructs GRNs to infer TF activity at the single-cell level
Epiregulon is designed to infer the activity of a transcription factor or a transcriptional coregulator (collectively referred to as TFs for brevity) under a variety of biological scenarios: (1) regulator activity driven by overexpression, (2) regulator activity decoupled from mRNA expression, (3) a context-dependent co-regulator interacting with different TFs and (4) gain of function due to neomorphic mutations or hijacking by other factors (Fig. 1a).
a Epiregulon can infer regulator activity for lineage development, drug perturbations, motif-lacking co-regulators or regulators harboring neomorphic mutations. Correlation and mutual information weight estimation methods are appropriate for the first scenario, whereas co-occurrence is applicable to all cases. b If users provide scRNA-seq and scATAC-seq, Epiregulon can construct GRNs either from ChIP-seq data or motif annotations. Pan-cell-type, tissue-specific and sample-specific ChIP-seq data compiled from ChIP-Atlas and ENCODE are available through the scMultiome package. Epiregulon outputs regulator activity at the single cell level, a pruned and weighted gene regulatory network and differential activity analysis to identify potential drivers of cell states. c For benchmarking, we downloaded the paired scATAC-seq and scRNA-seq PBMC data from 10x Genomics. We identified cell types using SingleR and known marker genes. Shown is the UMAP representation of the various cell types present in the data. d Gene expression of known lineage markers. e Activities of the same lineage markers were calculated using Epiregulon (correlation weight estimation method). f Area under the receiver operating characteristic curve (AUROC) is calculated based on whether TF expression or TF activity can distinguish cells of the matching lineage vs. the remaining cells based on a total of 20 factors. g Gene expression changes after depletion of 7 individual factors (ELK1, GATA3, JUN, NFATC3, NFKB1, STAT3 and MAF) were obtained from the knockTF database and genes with absolute logFC > 0.5 and corrected p-value < 0.05 (two-sided moderated t-test, limma) were considered ground truth target genes. GRNs obtained from the shown packages were evaluated for precision and recall of target genes. h Run time and memory use of the GRN construction from the PBMC data were evaluated with 64GB and 20 cores on the high-performance computing (HPC) cluster. In the case of GRaNIE, the memory allocation needs to be increased to 128 GB and for FigR, the memory allocation was increased to 256 GB. Each package was run 5 times. Source data are provided as a Source Data file. Created in BioRender. Yao, X. (2025) https://BioRender.com/x50fdft.
Epiregulon infers TF activity from a single-cell multiomics dataset containing paired RNA-seq and ATAC-seq counts (Fig. 1b). The ATAC-seq data are first used to identify regulatory elements (REs) from regions of open chromatin. The REs are filtered to those that overlap the binding sites of the TF, typically determined from external ChIP-seq data. Epiregulon provides a pre-compiled list of ChIP-seq binding sites from ENCODE and ChIP-Atlas spanning 1377 factors, 828 cell types/lines and 20 tissues (refer to Supplementary Data 1 and the “Methods” section for statistics and quality control). This list of sites can be further stratified by cell line or tissue, depending on the user’s biological context. We also provide options to identify sites from motif annotations (see the “Methods” section, Fig. 1b) or to use user-supplied binding sites.
Once a list of relevant REs has been obtained, each RE is tentatively assigned to all genes within a distance threshold. A gene is considered a target gene (TG) if the correlation between ATAC-seq and RNA-seq counts across metacells in the paired single-cell data is strong. Each RE-TG edge is then assigned a weight using the “co-occurrence method”, defined as the Wilcoxon test statistic from the comparison of the TG expression between “active” cells (that both express the TF and have open chromatin at the RE) to all other cells. The co-occurrence method is the default weighting scheme as it is less reliant on the degree of TF expression and thus can handle situations where TF activity is decoupled from expression. However, other weights can be used for TFs where activity is driven by expression (e.g., correlation or mutual information between TF and TG expression). Simulations indicate that Epiregulon’s results are robust to the choice of weighting scheme, even with sparse data and false connections (Supplementary Methods, Supplementary Figs. 1, 2).
After repeating the above steps for all TFs of interest, we obtain a weighted tripartite graph spanning the TFs, the REs overlapping their binding sites and the neighboring TGs. This weighted graph is the final GRN that is returned by Epiregulon. For each cell, the predicted activity of a TF is defined as the RE-TG-edge-weighted sum of the expression values of its TGs in the GRN, divided by the number of TGs. Epiregulon can also test for differential activity between conditions via total activity or edge subtraction of the GRN (Supplementary Methods, Supplementary Fig. 3) to identify potential context-dependent interaction partners of each TF. A more detailed description of the entire Epiregulon algorithm is provided in the Methods and Supplementary Fig. 4.
Epiregulon yields a high recall of target genes in PBMC data
We first evaluated the performance of Epiregulon using a human peripheral blood mononuclear cell (PBMCs) dataset obtained from 10x Genomics (Fig. 1c). The estimated activities of known lineage factors aligned to their respective lineages with greater specificity compared to TF expression alone (Fig. 1d–f). These lineage factors include TCF713, GATA314, BCL11B15, RUNX116, RUNX316, FOXP317, and IKZF118 (T cells), SPI119, CEBPA20,21, CEBPB22 and KLF423 (myeloid cells), EBF124, PAX525, POU2AF126 and TCF327 (B cells), EOMES28, TBX2128 and PRDM129 (NK cells) and IRF830 and TCF431 (dendritic cells, DC). Epiregulon also captured the multi-lineage nature of transcription factors. In addition to being a lineage factor of NK cells, TBX21 exhibited heightened activity in CD8+ memory T cells, consistent with the depletion of this cell type in Tbx21−/− mice32. IRF8 activity was elevated in DCs and moderate in monocytes, consistent with its known functions in myeloid development33.
Next, we benchmarked Epiregulon and other GRN methods (Supplementary Table 1) based on their ability to predict target genes. From the knockTF database, we identified 7 factors that were depleted in human blood cells—ELK1, GATA3, JUN, NFATC3, NFKB1, STAT3 and MAF. Genes with altered expression upon depletion of each TF were considered true target genes of that TF. Epiregulon detected more of these altered genes than other GRN methods in the PBMC dataset, at the cost of a modest loss in precision (Fig. 1g, Supplementary Fig. 5). SCENIC+ was the most precise method but failed to return a GRN for 3 of the 7 lineage factors (Fig. 1g). These results indicate that each method achieves a different compromise between recall and precision and Epiregulon is most suited for high recovery of target genes. Epiregulon also used the least computational time and memory (Fig. 1h), which is advantageous for iterative analyses.
Epiregulon predicts the responses of AR-modulating drugs
A more challenging task is to predict changes when TF activity is decoupled from its gene expression. We generated a single-cell multiomics dataset to evaluate changes in AR activity following drug treatment. Six prostate cancer cell lines (4 AR-dependent and 2 AR-independent) were treated with 3 AR-modulating agents (Fig. 2a, b). The first agent is the clinically approved AR antagonist, enzalutamide, which interferes with AR protein function by blocking its ligand-binding domain1. The second agent is ARV-110, a degrader of AR protein which acts by bringing an E3 ubiquitin ligase in close proximity to an AR protein3 (Fig. 2c). We also synthesized a third agent, SMARCA2_4.1, a degrader of SMARCA2 and SMARCA4. These two mutually exclusive paralog proteins encode the ATPase subunit of the SWI/SNF chromatin remodeler, which is crucial for recruiting AR to the chromatin34 (Fig. 2d, refer to Supplementary Methods for the method of synthesis). These pharmacological agents are not known to directly or consistently suppress AR mRNA levels.
a Mechanisms of action of 3 AR-modulating agents. b Six prostate cancer cell lines were treated and profiled for changes in their gene expression and chromatin accessibility by paired scATAC-seq and scRNAseq. Shown is the UMAP representation (5028 VCaP cells, 5958 LNCaP cells, 3568 22Rv1 cells, 945 MDA-PCa-2b cells, 3639 NCI-H660 cells and 3980 DU145 cells). Cells were merged from 2 technical replicates. Cells were treated for 24 h at 1 µM of enzalutamide or ARV-110 or 0.1 µM of SMARCA2_4.1. c Immunoblotting of AR and HDAC as a loading control after 24 h of treatment. This is a representative result from 2 biological replicates. d Chemical structure of SMARCA2_4.1. e Prostate cancer cell lines were treated for 5 days and cell viability was measured by CellTiter-Glo. Dotted line indicates the concentrations used in the scATAC-seq/scRNA-seq experiment. Data are presented as mean ± standard deviation (s.d.) based on 4 biological replicates. f Shown are the AR gene expression and the AR activity computed by Epiregulon (co-occurrence weight estimation method). Numbers of cells are as follows: VCaP (DMSO 1392 cells, Enza 1266 cells, ARV-110 1377 cells, SMARCA2_4.1 993 cells); LNCaP (DMSO 1499 cells, Enza 1966 cells, ARV-110 758 cells, SMARCA2_4.1 1735 cells); 22Rv1 (DMSO 970 cells, Enza 1002 cells, ARV-110 554 cells, SMARCA2_4.1 1042 cells); MDA-PCa-2b (DMSO 306 cells, Enza 76 cells, ARV-110 188 cells, SMARCA2_4.1 375 cells). Boxplots presented as median values ± 25%. Lower whisker is the smallest observation ≥25% quantile −1.5 × interquantile range (IQR). Upper whisker represents the largest observation ≤75% + 1.5 × IQR. g Each cell was identified by the HTO tag corresponding to treatment. A cell was classified into either the DMSO or AR inhibitor-treated group based on AR activity. Bar plots show the median receiver operating characteristic curve (AUROC) in the two sensitive cell lines, LNCaP and VCaP, treated with enzalutamide or ARV-110 for a total of 4 samples. Source data are provided as a Source Data file. Created in BioRender. Yao, X. (2025) https://BioRender.com/x50fdft.
We measured the response of all 6 cell lines to these pharmaceutical targets using cellTiterGlo at 1 and 5 days of treatment. At 1 day post-treatment, there was minimal cell death (Supplementary Fig. 6a), allowing us to profile their gene expression and chromatin changes. After 5 days of treatment, LNCaP and VCaP exhibited substantial cell death after enzalutamide treatment but MDA-PCa-2b and 22RV1 remained resistant (Fig. 2e). Similarly, LNCaP and VCaP showed the greatest sensitivity towards AR degrader ARV-110 while MDA-PCa-2b and 22RV1 were mildly responsive. Neither of the AR-independent cell lines responded to enzalutamide or ARV-110. All 6 cell lines responded to SMARCA2_4.1 treatment (Fig. 2e).
We used Epiregulon to predict changes in AR activity as a result of drug treatment. We leveraged publicly available AR ChIP-seq data for LNCaP, VCaP and 22Rv1 and generated our own ChIP-seq for MDA-PCa-2b. Consistent with the observed drug efficacy, Epiregulon predicted decreased AR activity following enzalutamide and ARV-110 treatment in the known sensitive cell lines LNCaP and VCaP, and minimal changes in AR activity in the resistant cell lines 22Rv1 and MDA-Pca-2b (Fig. 2f). Epiregulon predicted AR activity correctly in VCaP cells despite discordant trends in AR expression (Fig. 2f), highlighting the utility of the GRN approach.
We benchmarked Epiregulon against other GRN inference methods based on the ability of each method’s AR activity estimates to discriminate between DMSO- and drug-treated cells. Epiregulon achieved the highest area under the receiver operating characteristics curve (AUROC) when averaged across 2 cell lines and 2 drugs, implying that it is accurate at distinguishing control and treated cells in the sensitive cell lines (Fig. 2g, Supplementary Fig. 6b). Pando was comparably accurate in LNCaP but performed poorly in VCaP (Supplementary Fig. 6b); this is in part because enzalutamide treatment increased AR expression in VCaP cells despite AR inhibition (Fig. 2f). Lack of response to enzalutamide and a slight response to ARV-110 were observed for the resistant cell lines MDA-PCa-2b and 22Rv1 and (Supplementary Fig. 6c). These results indicate that Epiregulon should be the method of choice to predict drug efficacy from regulator activity.
Epiregulon estimates activity of AR harboring neomorphic mutations
An obvious alternative method to quantify AR activity is to compute gene set scores for existing AR signatures35,36,37,38,39. This assumes that the GRN of our biological system of interest is similar to that of the system from which the signatures were derived. For example, the AR signature predicted AR suppression consistent with cell fitness data of VCaP cells (Fig. 3a, b), likely because the amplification of wild-type AR is a frequent event in AR-dependent tumors, and the AR target genes derived from AR amplified samples are somewhat conserved in VCaP cells35,39. In contrast, the MDA-PCa-2b cell line harbors two mutations (L702H, T787A) in AR that change its ligand specificity (Supplementary Fig. 7a), and the combination of these 2 mutations is not well represented in patient tumors. We hypothesized that these mutations would also alter AR’s regulatory behavior; indeed, many canonical AR targets such as KLK3 and TMPRSS2 were not suppressed by any of the AR modulating agents (Fig. 3c, d). The change in AR function compromises the use of existing signatures, all of which failed to predict a decrease in AR activity upon treatment (Supplementary Fig. 7b, Supplementary Data 2).
a Shown are the AR activity estimated from the signature score in Bluemn et al.35 and its correlation with AR activity estimated by Epiregulon in VCaP cells, which harbor the amplification of the wildtype AR. Numbers of cells are as follows: DMSO 1392 cells, Enza 1266 cells, ARV-110 1377 cells, SMARCA2_4.1 993 cells. Boxplots presented as median values ± 25%. Lower whisker is the smallest observation greater than or equal to 25% quantile −1.5 × interquantile range (IQR). Upper whisker represents the largest observation ≤75% + 1.5 × IQR. b Normalized expression of genes in the Bluemn AR signature for VCaP. c Same as a, but for MDA-PCa-2b, which harbors two mutations in the AR gene and as a result has enhanced specificity for hydrocortisone over 5α-DHT. Numbers of cells are as follows: DMSO 306 cells, Enza 76 cells, ARV-110 188 cells, SMARCA2_4.1 375 cells. Boxplots presented as median values ±25%. Lower whisker is the smallest observation greater than or equal to 25% quantile −1.5 × interquantile range (IQR). Upper whisker represents the largest observation ≤75% + 1.5× IQR. d Same as b, but for MDA-PCa-2b. e Normalized expression of putative AR targets of MDA-PCa-2b as inferred by Epiregulon. f AR occupancy as measured by ChIP-seq at ATAC-peaks containing the regulatory elements mapped to AR target genes in MDA-PCa-2b cells treated with DMSO or the SMARCA2/4 degrader, SMARCA2_4.1. Center represents the center of the regulatory elements. g The ground truth regulator activity was computed using all the publicly available ChIP-seq obtained in LNCaP cells. This activity was then correlated (Pearson’s) with activity computed either from pan-cell-type ChIP-seq (red) or motif annotations (blue) for each of the regulators. h Same as g, but using ChIP-seq generated in MDA-PCa-2b. i Shown is the Jaccard similarity between the target genes derived from LNCaP ChIP-seq vs. target genes derived from pan-cell-type ChIP-seq (red) or motif annotations (blue). j Same as h, but using ChIP-seq generated in MDA-PCa-2b. Source data are provided as a Source Data file.
We previously observed that Epiregulon predicted a decrease in AR activity after SMARCA2_4.1 treatment in MDA-PCa-2b cells (Fig. 2f). Further investigation of this result revealed that Epiregulon predicted a different set of AR targets for MDA-PCa-2b compared to VCaP, many of which were downregulated by SMARCA2_4.1 treatment (Fig. 3e, Supplementary Fig. 7c, d). ChIP-seq confirmed reduced AR occupancy at the regulatory elements of AR target genes upon treatment (Fig. 3f). This explains the improved performance of Epiregulon over signature scores at predicting drug response in the presence of neomorphic mutations.
Pan-cell-type ChIP-seq outperforms motifs for accurate estimation of TF activity
Ideally, ChIP-seq data is available for the system of interest, as this provides the most accurate information about the TF’s binding sites. In the absence of such data, Epiregulon’s precompiled list of pan-cell-type ChIP-seq binding sites allows users to use information from other cell types, cell lines or tissues for exploratory analysis. Activities predicted by Epiregulon using the pan-cell-type binding sites were highly correlated with the activities obtained using the gold standard cell-line-matched ChIP-seq data (Fig. 3g, h, Supplementary Fig. 7e, f). In contrast, activities predicted by motif annotations did not always correlate well with cell-line-matched ChIP-seq (Fig. 3g, h, Supplementary Fig. 7e). These results suggest that unmatched ChIP-seq data should be generally preferred to motif annotations for estimating TF activity.
To further investigate the performance of Epiregulon with the pan-cell-type list, we examined the degree of overlap between the target genes identified with the pan-cell-type sites and those identified with cell-line-matched ChIP-seq. We observed good overlap for several factors in multiple cell lines (Fig. 3i, j, Supplementary Fig. 7g), indicating that the pan-cell-type list can often be good enough for target gene identification when cell-line-matched ChIP-seq data is not available. However, other factors exhibited a weaker overlap (Fig. 3i, j, Supplementary Fig. 7g), suggesting that cell-line-matched ChIP-seq is still necessary for pinpointing specific target genes.
Epiregulon uncovered context-dependent effects of SMARCA4 degradation
SMARCA4 is a transcriptional coregulator that is responsible for proliferation of prostate cancer cell lines40. SMARCA2_4.1 effectively depleted SMARCA4 protein expression in all 6 cell lines at 24 hours (Fig. 4a), so we would expect to see a decrease in SMARCA2_4.1 activity from each GRN method. SMARCA4 does not have a well-defined motif as it can interact with different TFs depending on the cellular context. This precludes the use of existing methods that rely on motif annotations for GRN construction. In contrast, Epiregulon can use public SMARCA4 ChIP-seq data to determine the most likely target genes without relying on motifs, upon which it correctly predicts decreased SMARCA4 activity in all cell lines (Fig. 4b).
a Immunoblotting of SMARCA4 after 24 h of treatment and HDAC as a loading control. b SMARCA4 activity computed by Epiregulon for all 6 prostate cell lines after 24 h of treatment. Boxplots presented as median values ± 25%. Lower whisker is the smallest observation ≥25% quantile −1.5 × interquantile range (IQR). Upper whisker represents the largest observation ≤75% + 1.5 × IQR. c Altered regulon size indicates the number of altered target genes mapped to each regulator. Altered genes are defined by genes with logFC > 0.5 and FDR < 0.05 after SMARCA2_4.1 treatment. LogFC in activity indicates the changes in regulator activity estimated by Epiregulon. d ChIP-seq of AR, SMARCA4 and FOXA1 in MDA-PCa-2b treated with the SMARCA2/4 degrader, SMARCA2_4.1 for 24 h at 0.1 µM at SMARCA4 binding sites. e Representative regions of SMARCA4, AR and FOXA1 ChIP-seq in MDA-PCa-2b cells treated with SMARCA2_4.1. f Same as c, but for DU145 cells. g ChIP-seq of SMARCA4, TEAD1 and FOSL1 in DU145 cells treated with the SMARCA2/4 degrader, SMARCA2_4.1 for 24 h at 0.1 µM. h Representative regions of SMARCA4, TEAD1 and FOSL1 ChIP-seq in DU145 cells treated with SMARCA2_4.1. Created in BioRender. Yao, X. (2025) https://BioRender.com/x50fdft.
Even though the same starting SMARCA4 ChIP-seq data were used for all cell lines, Epiregulon constructed a different GRN for each cell line to capture the context-dependent effects of SMARCA2_4.1 treatment. In MDA-PCa-2b (an AR-dependent cell line), AR and FOXA1 were amongst the factors with the largest altered regulon size and differential activity (Fig. 4c). ChIP-seq experiments indicated that SMARCA4 loss led to concomitant eviction of AR and FOXA1 at SMARCA4 binding sites (Fig. 4d, e). In the AR-independent cell line DU145, FOSL1 and TEAD1 were amongst the top perturbed factors while AR was not (Fig. 4f). ChIP-seq further validated the loss of FOLS1 and TEAD1 at SMARCA4 binding sites following SMARCA4 degradation (Fig. 4g, h). This is consistent with their role in shaping the epigenomic landscape of stem-cell-like prostate cancer39. These results demonstrate how Epiregulon’s GRN can be inspected to identify context-specific cofactors of the targeted factor.
Epiregulon identifies drivers of lineage reprogramming
To evaluate Epiregulon’s ability to identify drivers of cell states, we performed Reprogram-Seq41 to model the lineage transition of prostate adenocarcinomas by overexpressing defined factors. Lentivirus encoding transcription factors were introduced into LNCaP cells using two independent constructs: (1) pLenti9-reprogram-seq-V2-Cbh-UTR2-3 (UTR) driven by the chicken beta-actin promoter containing a puromycin-resistant cassette and (2) pLenti9-reprogram-seq-V2 (V2) driven by EF-1alpha promoter (Fig. 5a). We overexpressed 4 factors, NKX2-1, GATA6, FOXA1 and FOXA2, along with a negative control mNeonGreen. NKX2-1 is a known driver of AR-independence and neuroendocrine transition in prostate cancer42, whereas GATA6 has unknown function in the prostate. FOXA2 promotes neuroendocrine transition in a genetically engineered mouse model43. None of these factors is expressed in parental LNCaP. FOXA1 is already expressed in parental LNCaP, but we still introduced it since it is required for neuroendocrine transition42. We verified the expression of these factors using flow cytometry and immunoblotting. Except for NKX2-1 V2, other constructs achieved sufficient TF expression in LNCaP, with 30–70% of cells demonstrating overexpression above an uninfected control measured by flow cytometry (Supplementary Fig. 8a–c).
a Lentiviral constructs used to introduce TFs into LNCaP cells in the reprogram-seq assay. b UMAP representation of 3903 LNCaP cells transduced with virus encoding GATA6, NKX2-1, FOXA1, FOXA2 and mNeonGreen. The cells were infected in individual wells, hashtagged with HTO and then pooled into a single run. c Distribution of HTO tags in each of the clusters. d Motif enrichment in cluster-specific peaks was performed by ArchR using the CisBP motif database. e Chromatin accessibility at neuroendocrine (NE)—and stem cell-like (SCL)-specific regions defined by Tang et al.39 computed by ChromVAR. f Shown are the distribution of HTO tag assignment for GATA6 and NKX2-1, the level of TF gene expression, chromatin accessibility at GATA6- and NKX2-1 binding sites estimated by ChromVAR and TF activity computed by Epiregulon. g Gene expression (left) and Epiregulon-inferred activity (right) of NKX2-1 and GATA6. h Spearman correlation of regulon weights vs. log-fold changes of putative target genes of GATA6 (left) or NKX2-1 (right) with respect to mNeonGreen-infected cells. Shown is the confidence interval of 95%. P-values are calculated from 2-sided t-test. i Each cell was identified by the HTO tag corresponding to the well receiving virus encoding GATA6 or mNeonGreen. A cell was classified into either expressing GATA6 or not based on its TF activity. AUROC - area under the receiver operating characteristic curve. j Same as i but for NKX2-1. k Number of TFs detected in the GRN computed by the different packages. Source data are provided as a Source Data file.
Expression of these TFs altered the cell state of LNCaP cells, leading to the formation of distinct clusters (Fig. 5b). Cluster 1 was composed exclusively of GATA6-expressing cells while Cluster 3 contained only NKX2-1-expressing cells (Fig. 5c). Furthermore, peaks upregulated in Cluster 1 and 3 compared to mNeonGreen controls were highly enriched for GATA6 and NKX2-1 motifs respectively (Fig. 5d). NXK2-1 and GATA6 overexpression resulted in profoundly different cluster distributions from mNeonGreen (Supplementary Fig. 8d). Most strikingly, overexpression of NKX2-1 and GATA6 increased chromatin accessibility at neuroendocrine and stem-cell-like specific regions, respectively, and decreased accessibility at AR-dependent regions (Fig. 5e, Supplementary 8e) without overt changes in cell fitness (Supplementary Fig. 8f). We focused the rest of our analysis on GATA6 and NKX2-1 since their overexpression resulted in distinct reprogramming effects.
We tested Epiregulon’s ability to quantify the activity of GATA6 and NKX2-1 in this dataset. This is an interesting use case as the long distance between the polyA tail and the transcription factor cassette (1054 bp in UTR and 2110 bp in v2) hinders the efficient capture of exogenous TF mRNA by the 3’ protocol used in scRNA-seq. As a result, the observed expression is a poor representation of TF activity (Fig. 5f, g). In contrast, Epiregulon uses the expression of inferred target genes, correctly predicting increased activity for GATA6 in cluster 1 and NKX2-1 in cluster 3 (Fig. 5f, g). A subset of GATA6 targets were exclusively expressed in cluster 1 and NKX2-1 targets were exclusively expressed in cluster 3 (Supplementary Fig. 8g). In a differential expression analysis comparing cells in cluster 1 or 3 versus mNeonGreen control, GATA6 and NKX2-1 targets were highly ranked in cluster 1 and cluster 3, respectively (Supplementary Fig. 8h). Weights of target genes were also significantly correlated with their log-fold changes (Fig. 5h).
We performed a systematic benchmarking exercise to evaluate the performance of Epiregulon. Most GRN methods could distinguish GATA6-expressing cells from mNeonGreen-expressing cells with similar accuracy (Fig. 5i). However, Epiregulon was the only method that was able to predict NKX2-1 activity (Fig. 5j). This improved sensitivity is attributed partially to the use of ChIP-seq data instead of motif annotations; the GRN derived from ChIP-seq data yielded 118 NKX2-1 targets, whereas no targets were obtained by Epiregulon with motif annotations. Epiregulon’s GRN detected a large number of TFs mapped to their putative target genes (Fig. 5k), highlighting the benefit of using ChIP-seq to empirically determine binding sites.
We used deep neural networks (DNNs) to determine the importance of sequence information within Epiregulon’s GRN. DNNs are capable of learning complex patterns in the input data associated with accessible regions to make predictions of ATAC-seq coverage44,45,46. We trained two DNN models with ATAC-seq signals from cluster 1 and cluster 3 cells. respectively. We then occluded sequences within the REs in the regulons inferred by Epiregulon and compared the changes in predicted accessibility (Fig. 6a). Occlusion of GATA6 REs resulted in greater alterations of chromatin accessibility in cluster 1 than cluster 3, while the converse was true for NKX2-1 (Fig. 6b). This suggests that the REs inferred by Epiregulon contain important sequence information for determining accessibility.
a We train a DNN model on the ATAC-seq signals from cluster 1 and cluster 3 cells, respectively. We compare the predicted accessibility of either the wildtype sequence or the occluded sequence in the regulatory elements from the regulons inferred by Epiregulon. b Normalized changes in predicted accessibility if we occlude the sequences found in the regulatory elements of GATA6 and NKX2-1 regulons in either the DNN model trained on cluster 1 or cluster 3 cells. The number of regulatory elements for GATA6 is 402, and the number of regulatory elements for NKX2-1 is 134. Boxplots presented as median values ± 25%. Lower whisker is the smallest observation ≥25% quantile −1.5 × interquantile range (IQR). Upper whisker represents the largest observation ≤75% + 1.5 × IQR. c Each cell was identified by the HTO tag corresponding to the well receiving virus encoding GATA6 or mNeonGreen, and this information served as the true cell labels. For each TF, a cell was classified into either expressing GATA6 or not. GATA6 ChIP-seq was obtained by merging ChIP-seq from ChIP-atlas and ENCODE. ChIP+ motif refers to ChIP-seq peaks that contain GATA6 motifs. We trained a DNN model on cells expressing GATA6 (cluster 1) using Basenji2 and predicted an importance score for each motif based on the difference between the original sequence and the motif occluded sequence. We filtered for those motifs with importance scores higher than the quartiles of scores.
We also considered whether the motif annotation approach could be improved by deep learning models. For GATA6, we computed motif importance scores with two independent sequence deep learning models, Basenji and ChromBPNet (see the “Methods” section). We could not perform this analysis in NKX2-1 due to the lack of REs passing significance based on motif annotations. We applied thresholds on these scores to identify the motifs that are most likely to correspond to binding sites. However, the use of thresholded motifs did not improve Epiregulon’s estimation of TF activity; in fact, overly stringent thresholding reduced the number of target genes and degraded performance (Fig. 6c and Supplementary Fig. 8i). This motivates the continued use of the pan-cell-type ChIP-seq data, which still provides the accurate predictions of TF occupancy for activity inference.
Epiregulon predicts known and novel drivers of the cancer state from clinical samples
We further applied Epiregulon to clinical specimens to evaluate its ability to discover regulators in heterogenous and complex samples. We obtained scATAC-seq and scRNA-seq data from primary tumors and normal adjacent tissues47 and used Epiregulon to construct a GRN on both normal epithelial and tumor cells for 3 different cancer indications (renal cell carcinoma, glioblastoma and pancreatic adenocarcinoma). Epiregulon detected many well-known factors, including ZHX2, PAX8, N3RC1 in renal cell carcinoma, FOSL2 and SOX2 in glioblastoma and KLF5 in pancreatic adenocarcinoma (Fig. 7a-c and refer to Supplementary Table 2 for a full list of references). We also found that KLF9 was suppressed in pancreatic adenocarcinoma, consistent with its role in tumor suppression48. Interestingly, the changes in TF activity are more pronounced than changes in gene expression (Fig. 7a–c). This implies that Epiregulon can identify drivers of the tumorigenic state even in the absence of strong changes in gene expression.
ScATAC-seq and scRNA-seq data of primary tumors and normal adjacent tissues were obtained from Terekhanova et al.47 and paired using Seurat’s label transfer function. Only tumor cells and matching normal cell types were used for GRN construction by Epiregulon (co-occurrence weight method). The top regulators were identified using Epiregulon’s findDifferentialActivity function. Shown are the expression and activity of the top regulators in a renal cell carcinoma, b glioblastoma and c pancreatic adenocarcinoma. Red stars mark regulators, which are known to promote tumor growth, and blue stars mark regulators, which are known to inhibit tumor growth. Created in BioRender. Yao, X. (2025) https://BioRender.com/nn27yx0.
Discussion
We present Epiregulon, a computational method to construct GRNs from single-cell multiomics data in a motif-agnostic manner. Epiregulon performs robustly across a multitude of datasets, identifying target genes and accurately quantifying TF activity even in the presence of neomorphic mutations. It is an accurate tool for predicting response to drug perturbations and is the only tool that can infer the activity of a chromatin remodeler amongst the tools we benchmarked. We show that Epiregulon predictions based on ChIP-seq data outperform those from motif annotations, even after prioritization of the latter by DNN models. Our analyses demonstrate that Epiregulon can be reliably applied to evaluate TF-targeting pharmaceutical agents or to study epigenomic drivers of tumorigenesis and cell state changes.
Recent advances in chemical biology offer exciting new targeting strategies for the previously undruggable TFs and transcriptional coregulators. It will be important to develop a unified method to evaluate and compare various drug modalities, which may include direct modulators of the effector domain, degraders of protein and indirect modulators49. However, we have shown that TF expression is not a reliable measure of TF activity as negative feedback loops can still induce gene upregulation despite compromised TF function. Canonical gene signatures or motif-dependent gene regulatory network (GRN) methods also fail to account for alterations in the TF cistrome caused by gain-of-function mutations or TF hijacking50,51,52. By constructing disease- or/and lineage-specific GRNs, Epiregulon helps identify the most likely targets for a particular model system and determines whether drugs are on-target and/or sufficiently potent. Furthermore, as demonstrated with the SMARCA4 degrader, Epiregulon can uncover therapeutically impactful co-targets. This GRN method can be applied to a broad range of motif-independent transcriptional regulators, including chromatin readers, transcriptional kinases and histone-modifying enzymes, when identification of interaction partners is often challenging.
Epiregulon has some important limitations that may affect its performance. Acute perturbations can alter gene expression without substantial changes in chromatin accessibility53, reducing the effectiveness of Epiregulon and other GRN methods. Epiregulon does not explicitly model cooperativity between different TFs, which simplifies GRN construction but may reduce the accuracy of the activity estimates. Epiregulon relies on the availability of high-quality ChIP-seq data for accurate estimates of TF activity. The current quality filters on peak numbers and number of reads may be insufficient to remove low-quality ChIP-seq datasets and introduce erroneous estimates of TF occupancy. Future versions of Epiregulon may incorporate enrichment filters, including Fraction of Reads in Peaks and strand correlation metrics. Furthermore, while the pan-cell-type list is often satisfactory for TF prioritization, validation for the functional importance needs to be achieved by performing ChIP-seq on TFs of interest in the relevant biological systems. Finally, our benchmarking was limited to the few TFs for which we have ground truth data, so it is difficult to generalize conclusions about Epiregulon’s performance to all TFs. Nevertheless, we envision that Epiregulon will become a useful tool for drug discovery, cancer biology and beyond.
Methods
The research conducted here complies with all relevant ethical regulations of Genentech Inc.
Epiregulon
The Epiregulon workflow consists of several components: GRN construction, pruning of networks (optional), estimation of weights, calculation of activity, functional annotation of regulons, differential network analysis and identification of interaction partners. Epiregulon is designed to work seamlessly with ArchR but can also accept SingleCellExperiment objects. It is available as three related R packages. Epiregulon is the core package that performs GRN construction and activity inference. Epiregulon.extra contains differential analysis and plotting functions. Epiregulon.archr contains functions interfacing with ArchR. ChIP-seq data is available through scMultiome. All packages except Epiregulon.archr are available through Bioconductor.
Data preprocessing
Epiregulon assumes that prior preprocessing of the data has been performed by users’ methods of choice, requiring paired gene expression matrix (scRNA) and peak matrix (scATAC) along with dimensionality reduction matrix as the input. If gene expression and peak matrix are not paired, integration of scATAC-seq and scRNA-seq data must be performed prior to Epiregulon using methods such as ArchR’s addGeneIntegrationMatrix function.
For the datasets included in this manuscript, reads were mapped by the Cell Ranger ARC 2.0 and were further processed with the use of ArchR54. Briefly, cells were filtered based on TSS enrichment (>3) and number of ATAC-seq reads mapped to the nuclear genome (>1000). Moreover, doublets were removed using ArchR’s filterDoublets function. ATAC-seq data were represented as a 500 bp tile matrix on which iterative latent semantic indexing (LSI) was performed. Similarly, normalized gene expression data were subject to iterative LSI. The dimensionality reductions from both modalities were then combined into a single matrix that was used for cell clustering by graph clustering approach implemented in Seurat55,56,57. The ATAC-seq peaks were called separately for each cluster and, after normalization, were merged to produce one peak set for all cells (peak × cell matrix). The cells were assigned to their sample barcodes using demuxEM58. Enrichment of chromatin accessibility at each peak or motif was computed using the chromVAR59 function provided through ArchR. Briefly, chromVAR computes the bias-corrected deviation of per-cell accessibility for a given motif or TF binding sites from the average of all cells.
Network construction
Epiregulon provides two similar methods to establish peak to gene links using the calculateP2G function. If there is an existing ArchR project, Epiregulon retrieves the peak to gene links that have been previously assigned by ArchR. Briefly, ArchR creates 500 cell aggregates, resampling cells if needed. ArchR computes the correlation between chromatin accessibility and target genes within a size window (default ± 250 kb) and retains peak-gene pairs that exceed a correlation threshold (default Epiregulon cutoff is 0.5). In the absence of an ArchR project, Epiregulon defines cell aggregates using k-means clustering based on the reduced dimensionality matrix and performs correlation in the same manner as ArchR does. If cluster labels are provided, cluster-specific correlations are reported in addition to overall correlations.
TF occupancy data
Each regulatory element is then interrogated for TF occupancy based on a compilation of public TF ChIP-seq binding sites. ChIP-seq data were downloaded from ChIP-Atlas (chip-atlas.org) and ENCODE (encodeproject.org).
We created sample- and tissue-specific ChIP-seq peak sets for each factor to allow for tissue or sample matched analysis.
For ChIP-Atlas data, we only retained ChIP-seq samples that met the following criteria:
-
Total number of unique reads ≥ 20M
-
Number of peaks (FDR < 1 × 10−5) ≥ 1000
For ENCODE data, we remove any samples with Audit.NOT_COMPLIANT or Audit.ERROR. flags. We retain only samples with IDR thresholded peaks ≥ 1000.
We also created a pan-cell-type ChIP-seq peakset using the merged ChIP-seq peaks provided by ChIP-Atlas and the ENCODE ChIP-seq data, yielding 1376 unique factors (human) and 626 unique factors (mouse).
The mode of action in Supplementary Data 1E was annotated based on gene ontology terms with “GO:0045944_positive_regulation_of_transcription_by_RNA_polymerase_II” as activator and “GO:0000122_negative_regulation_of_transcription_by_RNA_polymerase_II” as repressor. Peak sets were merged across each sample or each tissue for every TF. Data is provided in the scMultiome package as GRanges list objects and can be accessed using the getTFMotifInfo function from the Epiregulon package or directly from the scMultiome package using the tfBinding function.
If desired, ChIP-seq peaks can be further annotated for the presence of motifs using the addMotifScore function. If motif annotation has been performed previously using ArchR’s addMotifAnnotations function, motif annotations can be easily retrieved and appended to the peak matrix. In the ArchR independent workflow, Epiregulon can annotate peak matrix using motifmatchr’s motif matching function59 and cisbp as the reference motif database60. Alternatively, users can start entirely from motif annotations and leverage deep learning motifs to select motifs based on motif importance scores (see section on DNN models).
Network pruning (optional)
Epiregulon prunes the network by performing tests of independence on the observed number of cells jointly expressing transcription factor (TF), regulatory element (RE) and target gene (TG) vs. the expected number of cells if TF/RE and TG are independently expressed using the pruneRegulon function.
We define n as the total number of cells, k as the number of cells jointly expressing TF, TG and RE above a set threshold, g as the number of cells jointly expressing TF and RE above a threshold and h as the number of cells expressing TG above a threshold. p, the expected probability of cells jointly expressing TF, TF and RE above a threshold is defined in Eq. (1):
Two tests of independence are implemented, the binomial test and the χ2-test. In the binomial test, the expected probability is p, the number of trials is the total number of cells n, and the observed number of successes is k, the number of cells jointly expressing all three elements.
In the χ2-test, the expected probability for having all 3 elements active is also \(p\). The observed cell count for the active category is \(k\), and the cell count for the inactive category is \(n-k\). P-values are calculated from a chi-squared distribution with degree of freedom equal to 1. Cluster-specific p-values are calculated if users supply cluster labels. Finally, multiple hypothesis testing was performed using the Holm method.
Estimation of weights
While network pruning provides statistics on the joint occurrence of TF-RE-TG, we would like to further estimate the strength of regulation using the addWeights function. Biologically, this can be interpreted as the magnitude of gene expression changes induced by changes in transcription factor activity. Epiregulon provides 3 different methods to estimate weights. Two measures (correlation and co-occurrence (Wilcox)) give both the magnitude and directionality of changes, whereas weights computed by mutational information (MI) are always non-negative. Within 2 of the methods (correlation and MI), there is an option of modeling the TG expression based only on TF expression, or on the product of TF expression and RE chromatin accessibility. Consideration of both TF expression and RE chromatin accessibility is highly recommended, especially for scenarios in which TF activity and TF gene expression are decoupled (as in the case of drug perturbations or CRISPR genome editing). The correlation and mutual information statistics are computed on pseudobulks by user-provided cluster labels and yield a single weight across all clusters per each TF–RE–TG triplet. In contrast, the Wilcoxon method groups cells based on the joint expression of TF, RE, and TG in each single cell or in cell aggregates. Cell aggregation uses a default value of 10 cells and can help overcome sparsity and speed up computation. If cluster labels are provided, we can obtain cluster-specific weights using the Wilcoxon method.
Co-occurrence (Wilcox)
Cells were divided into two groups, with the first group jointly expressing TF gene expression and chromatin accessibility at RE, and the remaining cells in the second group. The default cutoff is 1 for normalized gene expression and 0 for normalized chromatin accessibility. There is also an option to use the median of each feature as an adaptive cutoff.
Correlation
If we consider only TF expression (tf_re.merge set to FALSE), the weight is the correlation coefficient between the TF gene expression and TG gene expression. If we consider both TF expression and RE chromatin accessibility (tf_re.merge set to TRUE), the weight is the correlation coefficient between the product of TF gene expression and RE chromatin accessibility vs the TG gene expression.
Mutual information between the TF and target gene expression
If we consider only TF expression (tf.re_merge is set to FALSE), the weight is the mutual information between the TF gene expression and the TG gene expression. If we consider both TF expression and RE chromatin accessibility (tf.re_merge is set to TRUE), the weight is the mutual information between the product of TF gene expression and RE chromatin accessibility vs. the TG gene expression.
Calculation of TF activity
The activities for a specific TF in each cell are computed by averaging the weighted expressions of target genes linked to the TF in Eq. (2)
where \({K}_{t,c}\) is the activity of a TF \(t\) for a cell \(c\), \(\left|{R}_{t}\right|\) is the total number of target genes for a TF \(t\), \({Y}_{g}\) is the normalized count of target gene \(g\) where \(g\) is regulated by TF \(t\) and \({\beta }_{t,g}\) is the regulatory weight of TF \(t\) on target gene \(g\). \({R}_{t}\) is the regulon of TF \(t\). If cluster labels are provided, cluster-specific weights are used.
Gene set enrichment of regulons
Gene set enrichment of a regulon is performed by testing whether the target genes of a TF are over-represented in known gene signatures such as those provided by MSigDB using a hypergeometric test. Target genes can be refined by filtering the regulons on user-defined weights.
Differential TF activity by total activity
This differential analysis compares the differences in the activity of each transcription factor between conditions. This analysis is well suited for identifying factors that have contrasting levels of activities, for instance, lineage factors that are turned on or off during certain developmental stages or cell types. TF activities are compared between groups using any standard statistical methods. Here we use scran’s findMarkers function to find differential activity between user provided groups of cells (https://rdrr.io/bioc/scran/man/findMarkers.html).
Differential TF activity by network topology
A second approach to investigate differential TF activity is to compare target genes or network topology. This is useful when a transcription factor differs in the target genes it regulates, while maintaining a similar level of total activity. This can happen when a transcription factor redistributes to a different set of genomic regions, due to mutations in the transcription factors or changes in the interaction partners. Differences in network topology are calculated by taking the degree centrality of the edge-subtracted graphs between two conditions, with cluster-specific regulon weights representing edge weights in each condition.
Consider networks G(1) and G(2) with identical node sets N and respective adjacency matrices A(1) and A(2). Then then the edge-subtracted network G′ induced by G(1) and G(2) has the adjacency matrix whose entries are calculated as \({e}_{{ij}}^{{\prime} }={|}{e}_{{ij}}^{(1)}-{e}_{{ij}}^{(2)}{|}\), where e(j)ij represents the weight of edge connecting ith transcription factor with jth gene in network G(j).
We simplify the tripartite TF–RE–TG graph to a bipartite TF–TG graph by taking the maximum of TF–RE–TG weights of the same TF–TG pairs. Degree centrality is the sum of the weights associated with the ith transcription factor: \({\sum }_{i}{e}_{{ij}}^{{\prime} }\). Degree centrality is further normalized to account for differences in the number of target genes of each transcription factor. The default normalization method is dividing degree centrality by the square root of the number of target genes. This strikes a balance between penalizing TFs with an abundance of target genes and prioritizing TFs with differential target genes. Transcription factors are ranked by normalized degree centrality.
Benchmarking
Benchmarking using PBMC data
PBMC dataset was downloaded from 10x Genomics (https://www.10xgenomics.com/datasets/pbmc-from-a-healthy-donor-granulocytes-removed-through-cell-sorting-10-k-1-standard-2-0-0). The Cell Ranger output was processed using ArchR as described in the “Data preprocessing” section. As a result, 9,702 out of the 11,582 cells were kept for downstream analysis. We used the peak matrix retrieved from the ArchR project as input to GRN inference tools. Gene expression data was retrieved from ArchR or directly from Cell Ranger output, depending on the benchmarked tool.
Clustering was performed using LSI dimensionality reduction, which combined information from both chromatin accessibility and gene expression data. We used marker genes to determine naive CD4+ T cells (IL7R, CCR7), CD14+ monocytes (CD14, LYZ), and CD4+ memory cells (S100A4, IL7R) and SingleR with BlueprintEncodeData from the celldex package as a reference for other cell types. Cell clusters were annotated into one of the following types: naive CD4+ T, memory CD4+ T, naive CD8+ T, memory CD8+ T, monocytes, CD14+ monocytes, FCGR3A+ monocytes, B, NK, DC. 24 cells were left unannotated and were excluded from further analyses.
We used PBMC data from the KnockTF database61 as the ground truth to test the accuracy of target gene assignment to transcription factors. We collected data presenting the results of seven knockdown experiments, each one targeting a different transcription factor. Target genes were determined using quality filters (absolute value of logFC > 0.5, corrected p-value < 0.05). For each package we benchmarked, we calculated precision and recall based on the predicted and ground truth target genes. Precision is defined as the number of predicted target genes that are altered by the knockdown of TF/total number of predicted target genes. Recall is defined as the number of predicted target genes that are altered by the knockdown of TF/total number of altered genes.
All packages were tested for run time and memory use. Data preprocessing was excluded from these measurements (including topic analysis in SCENIC+). We assigned 64 GB and 20 cores on HPC for each run. In the case of GRaNIE the memory allocation had to be increased to 128 GB and for FigR the memory allocation was increased to 256 GB. Each package was run 5 times and median run time and memory use were recorded.
Benchmarking using Reprogram-Seq data
We introduced 4 transcription factors into LNCaP cells - FOXA1, FOXA2, NKX2-1, and GATA6 and obtained paired gene expression and chromatin accessibility information. Because FOXA1 was already highly expressed in LNCaP cells at the basal level, its introduction did not have any profound impact on lineage plasticity, and therefore, FOXA1 was excluded from subsequent analyses. We focused on NKX2-1 and GATA6 because their expression resulted in distinct cell clusters. Each cell is identified by the HTO tag corresponding to the well receiving virus encoding a particular TF, and this information serves as the ground truth cell labels. For each TF, a cell is classified into either expressing this TF or not. Area under the receiver operating characteristic curve (AUROC) was computed for all the packages being benchmarked, as well as for gene expression of the TF being evaluated. Note that because each TF has different transduction efficacy and not all cells are able to take up the virus or express the TF, the theoretical maximum of AUROC will not reach 1.
Single-cell data was processed using Cell Ranger as described for PBMC data. Differential peak analysis was performed using ArchR’s getMarkerFeatures. Briefly, an equal number of cells with a similar distribution of TSS enrichment and fragment numbers were sampled from treatment conditions (cluster 1 or 3) and background conditions (mNeonGreen). Differential peaks were identified using two-sided t-tests with FDR correction on the chromatin accessibility counts at the peak regions. Cutoffs used were FDR ≤ 0.01 and absolute Log2FC ≥ 1. Differential Peaks were then annotated with motifs from the CisBP database.
Benchmarking using AR antagonist data
For the drug treatment dataset, we treated 6 prostate cancer cell lines with 3 different therapeutic agents and obtained paired gene expression and chromatin accessibility information. Only the 2 AR-dependent cell lines, LNCaP and VCaP, were included in the benchmark since they are known to respond to AR-targeting agents. Similarly, only enzalutamide1 and ARV-1103 were used in the benchmark studies since they are known to specifically inhibit AR activity. Cells treated with Enzalutamide and ARV-110 are supposed to show reduced AR activity compared to cells treated with the DMSO control. Each cell line was analyzed separately by retaining only peaks found in each cell line. Each cell is identified by the HTO tag corresponding to the well receiving DMSO or one of the two AR inhibitors, and this information serves as the ground truth. For each of the two drugs (enzalutamide or ARV-110), a cell is classified into either treated with DMSO control or an AR inhibitor. AUROC is computed for all the packages being benchmarked, based on the AR activity values, as well as for AR gene expression. In the Epiregulon workflow, we used the Wilcoxon method to estimate weights with cell-line AR ChIP-seq.
We used 4 GRN inference tools (FigR, SCENIC+, GRaNIE and Pando) to benchmark the performance of Epiregulon. To ensure consistency, we applied the same gene expression and chromatin accessibility matrices across all tools. We first used ArchR to determine the peaks, i.e., DNA regions with frequent Tn5 transposase insertion events from the fragment files output by CellRanger. Then the peak x cell matrix was produced by counting the number of insertions per peak per cell. From the same ArchR project, we also retrieved normalized gene expression to be used by Epiregulon, GRaNIE, and FigR. For the remaining tools, we used a subset of cells in the gene expression data that matched the cells in the ArchR project to make sure that all tools work on the same cells. Below, we describe the workflows used in each tool. We followed the tutorial examples provided on official websites with only minor changes to the default settings.
FigR
We performed benchmarking against FigR9. In the first step of GRN construction, the correlation between peak accessibility and target gene expression was determined using runGenePeakcorr. We used a non-default search range around TSS (250 kb) to be consistent across all the benchmarked tools. Correlation coefficients were also computed with background peaks, and the significance of gene–peak association was determined with one-tailed z-test. Only gene–peak associations that show positive correlation and are statistically significant were retained (p ≤ 0.05). DORaCs (High-density domains of regulatory chromatin) correspond to genes with ≥7 associated peaks and were calculated by summing the ATAC-seq reads in peaks matched to each gene. Each TF is associated with a single motif by selecting the motifs most highly correlated with other motifs of the same TF. Each DORC is evaluated for enrichment of TFs by comparing the match frequency in its peakset vs the match frequency in a set of background peaks; the p-values of the enrichment were obtained by z-test (\({P}_{{{{\rm{E}}}}{{{\rm{n}}}}{{{\rm{r}}}}{{{\rm{i}}}}{{{\rm{c}}}}{{{\rm{h}}}}{{{\rm{m}}}}{{{\rm{e}}}}{{{\rm{n}}}}{{{\rm{t}}}}}\)). Smoothing of gene expression matrix and chromatin accessibility data summarized across target genes was performed after determination of k nearest-neighbor cells based on LSI space retrieved from the ArchR project and constructed using ATAC-seq data. The Spearman correlations between the smoothed DORC accessibility and smoothed TF gene expression were computed (\({{{\rm{C}}}}{{{\rm{o}}}}{{{\rm{r}}}}{{{\rm{r}}}}{{{\rm{e}}}}{{{\rm{l}}}}{{{\rm{a}}}}{{{\rm{t}}}}{{{\rm{i}}}}{{{\rm{o}}}}{{{\rm{n}}}}\)) and their significance was obtained using z-test (\({P}_{{{{\rm{C}}}}{{{\rm{o}}}}{{{\rm{r}}}}{{{\rm{r}}}}{{{\rm{e}}}}{{{\rm{l}}}}{{{\rm{a}}}}{{{\rm{t}}}}{{{\rm{i}}}}{{{\rm{o}}}}{{{\rm{n}}}}}\)). From the output of the main function (getFigRGRN) we retrieved scores indicating the strength of association between transcription factors and target genes, which were computed as Eq. (3).
The regulation scores were used as weights when calculating activity with Epiregulon.
SCENIC+
We performed benchmarking against SCENIC+12. Briefly, we used pycisTopic, which uses Latent Dirichlet Allocation to group regulatory elements into topics. The model evaluation indicated 20, 20, 10, 10 as the number of topics for VCaP, LNCaP, MDA and 22Rv1 cells from the AR dataset, respectively, and 20 for the Reprogram-Seq dataset. The input peak matrix was retrieved from the AchR project. We then used pycistarget to identify TF-region links. Pycistarget identifies motif matches in the peak regions using HOMER, scores each region for motif importance and identifies differentially enriched motifs above background regions. Only regions showing NES > 3.0 and motifs with adjusted p-value < 0.05 and log2FC > 0.5 were retained. TF-gene importance scores were calculated using gradient-boosting machine regression by predicting TF expression from target gene expression. Region-gene importance scores were calculated using gradient-boosting machine regression by predicting target gene expression from region accessibility. Genes were ranked by TF-gene importance scores, and only genes in the leading edge of the gene set enrichment were used for the eRegulon. Gene set enrichment was also performed for region-gene pairs. Peaks were ranked by imputed chromatin accessibility, and genes were ranked by gene expression counts per cell. Enrichment score was defined as the AUC at 5% of the ranking and was calculated using AUCell. The enrichment score was used as the activity score in the benchmark assessment.
GRaNIE
We performed benchmarking against GRaNIE11. Briefly, GRaNIE overlapped TF binding sites obtained from HOCOMOCO-based TF motifs with ATAC-seq peaks. GRaNIE identified TF-peak connections using Pearson correlation between TF expression and the peak accessibility across samples. The cutoffs for the correlation were chosen based on an empirical FDR calculated from the ratio of TF-peaks in the background peaks over the total number of TF-peaks in both the background and foreground. Peak-gene connections were identified using the correlation between the gene expression and chromatin accessibility. All the GRN edges have a weight of 1. The default threshold is FDR < 0.2 for TF-peak links and FDR < 0.1 for peak-gene links. Activities were computed using Epiregulon’s calculateActivity.
Pando
We performed benchmarking against Pando10. Briefly, ATAC-seq peaks were intersected with PhastCons conserved elements and cCREs derived from ENCODE. TFs present in the 4000 most variable genes were included in the downstream analysis. Motifs for the TFs were obtained from JASPAR2020 and CISBP. Gene expression and chromatin accessibility counts were smoothed by averaging cells within a neighborhood. Pando then models target gene expression as the weighted sum of the product of TF expression and the chromatin accessibility of the region where the TF binds according to Eq. (4).
where \({Y}_{g}\) is the expression of the target gene \(g\), \({X}_{t}\) is the expression of the transcription factor \(t\), \({A}_{r}\) is the chromatin accessibility at region \(r\), \(\beta\) is the fitted coefficient and \(c\) is the intersection.
Fitted coefficients were tested for significance using analysis of variance (ANOVA). Only edges with FDR < 0.05 were retained in the final GRN. Activities were calculated using Epiregulon’s calculateActivity function with the fitted coefficients \({\beta }_{t,g,r}\) as the weights.
GRN construction from patient tumors
Unpaired scATAC-seq fragment files and Seurat objects containing author-processed scRNA-seq counts of primary tumors and normal adjacent tissues were downloaded from NCI Human Tumor Atlas Network as indicated in Terekhanova et al.47 scATAC-seq data were preprocessed by ArchR as described in the data preprocessing section. Pairing of scATAC-seq and scRNA-seq was performed using Seurat’s label transfer function implemented within ArchR with patient ID as the restraint. Only tumor cells and matching normal cell types were retained and were down-sampled to 50,000 cells for each indication. GRN was constructed using Epiregulon’s co-occurrence (wilcox) weight estimation method, and the top regulators were identified using Epiregulon’s findDifferentialActivity function. Only TFs with an altered regulon size >35 were retained. Altered regulon size refers to the number of genes showing an absolute normal–tumor log fold change > 0.5 and FDR < 0.05.
DNN training
Steps for DNN-based TF-RE mapping involved dataset processing, model training, motif scoring and thresholded GRN construction.
Dataset processing
The reference genome hg38 and the pseudo-bulk ATAC (taken from ArchR) of individual clusters (cluster 1 or cluster 3) were collected. The whole genome, excluding unmappable regions62, was split into 3072 bp regions. Using the coordinates of each region, the corresponding DNA sequence and bigwig coverage were obtained. The DNA sequence was converted to one-hot encoded form. The DNA and coverage information were saved in tf records files for faster I/O during training. Training, validation and test splits were done by chromosomes, taking chr8 as the test set and chr9 as the validation set.
Model training
We used Basenji2 architecture with 32 base-pair resolution (binned coverage at 32 bp). The model was trained by randomly sampling a 2048 bp segment from the input DNA and taking the corresponding coverage (one cluster per model). We used Poisson NLL as loss, used reverse complement augmentation and trained for a maximum of 50 epochs with early stopping.
We additionally trained models with base-pair resolution using the ChromBPNet architecture. We first trained a custom Tn5 sequence bias model for this dataset on non-peak regions. We ensured that these models learned Tn5 bias by applying DeepSHAP to the model outputs. Using this bias model to regress out Tn5 sequence bias, we then trained ChromBPNet TF models. These models were trained on 2114 bp sequences centered on ATAC-seq peaks.
Motif scoring
For each of the motif regions (taken from the ArchR motif positions file), we obtained the DNA sequence (denoted as wild type) by centering at the motif and extending to 2048 bp. We occluded the motif by replacing motif nucleotides with ‘N’ (or 0.25 in one-hot encoding) (denoted as mutant). We computed the score−mean fraction change in predicted coverage by subtracting mutant from wild type and normalizing by predicted wild type coverage.
Entire RE occlusion
Similarly, we assessed the importance of entire REs for model comparison. We used ChIP-based regulons to identify genomic coordinates containing GATA6 (or NKX2-1) binding sites, at 500 bp resolution. We then obtained wild-type (WT) predictions for sequences centered at these REs and mutant sequence predictions by occluding the entire 500 bp RE. We quantified the importance of each RE by subtracting the sum of the mutant predictions from the WT predictions and normalizing by the WT value.
Thresholded GRN construction
We computed several quartiles of scores as thresholds. We filtered for those motifs with importance scores higher than the threshold corresponding to bigger differences in prediction. Using the RE remaining we constructed the GRN as before.
Reprogram-Seq
Construction of lentivirus plasmids for TF over-expression
All lentivirus plasmids were generated by GenScript. Briefly, the ORF of transcriptional factors was codon optimized, synthesized and cloned after the hEF1a promoter. The puromycin resistance gene is driven by a separate Cbh promoter to enable antibiotics selection. Maxi-prep of each plasmid was performed to maximize the transfection efficiency.
Cell culture and virus packaging
LNCaP Clone FGC cells were obtained from ATCC and cultured with RPMI-1640 media with 10% FBS and 2 mM L-glutamine. The cells were split every 4–5 days to maintain the appropriate density. 293T cells were cultured in DMEM with 10% FBS, 100 μM NEAA, 2 mM Glutamine. They were split every 2–3 days. One day before transfection, the cell culture dish was treated with 5 ml 1% gelatin in PBS, incubated for 10 min, then aspirated. 3.5 × 106 293T cells were seeded into each 10 cm dish to reach ~80% confluence. On the day of transfection, 20 µl Lipofectamine 2000 was added to 480 µl OptiMEM. In a new tube, the plasmid was mixed in 500 µl Opti-MEM at the following ratio: carrier plasmid, 5 µg; delta8.9, 16 µg; VSVG, 1 µg. Both mixes were combined and incubated for 20 min before adding to the dish. The dish was incubated at 37 °C for 6 h and then 6 ml of complete media was added. Virus was then harvested 48 h after transfection. Briefly, all supernatants were harvested and filtered through a 0.45 µm filter bottle. The virus was concentrated by using the Lenti-X concentrator (TAKARA Bio.) following the manufacturer’s instructions and resuspended in 1 ml 1% BSA in PBS per dish. The concentrated virus was divided into 200 µl aliquots and stored in -80 °C until infection.
Infection of LNCaP cells by lentivirus
Two days before infection, 4 × 105 LNCaP cells were seeded into each well of a 6-well plate. On the day of infection, aspirate the media and change to 0.5 ml RPMI + 10% FBS + 1X Glutamax with 8 µg/ml Polybrene. A total of 200 µl concentrated lentivirus was added to each well to achieve high MOI, and then the plate was centrifuged at 800 × g for 45 min at room temperature. After that, the plate was put into the incubator for another 3 h before 2 ml of the full media was added to each well. Two days after infection, the media was refreshed with 1 µg/ml puromycin. The cells were then grown for another 7 days before harvesting for the single-cell analysis, during which the cells were split accordingly when the confluence reached 100%.
Validation of protein expression
Sufficient protein expression of exogenous TFs was first validated by flow cytometry. LNCaP cells were fixed at 8% paraformaldehyde at room temperature for 30 min. Cells were permeabilized by 0.2% Triton/PBS at room temperature for 20 min and blocked with 2% BSA/PBS at room temperature for 1 h. Cells expressing the specific TFs along with uninfected cells were incubated with the following antibodies: NKX2-1-APC (Miltenyi Biotec, 130-118-309), GATA6-PE (Cell Signaling Technology, 26452) and FOXA2-AlexaFluor 488 (Abcam, catalog number ab208376). Cells were analyzed on a cell sorter (Sony SH800S).
Protein expression was further validated by immunoblotting against GATA6 and NKX2-1. For NKX2-1, LNCaP-FGC, NKX2-1 overexpressing LNCaP-FGC cells and NCI-H660 cells were lysed using RIPA buffer supplemented with HALT protease and phosphatase inhibitor (Thermo Scientific, Cat #: 78440), and lysates were quantified using Pierce BCA Protein assay kit (Thermo Scientific, Cat #: 23225) according to the instructions for the microplate procedure. Gel electrophoresis was performed with 20 μg of protein lysates, on 4–12% NuPAGE Novex Bis–Tris midi gels (Thermo Fisher, Cat #: WG1402) at constant voltage of 100 V in NuPage MOPS SDS running buffer (Invitrogen, catalog number NP0001) for 1.75 h, followed by transfer to PVDF membranes using Trans-blot Turbo (Biorad) for 13 min at 1.3 A and 25 V. Membranes were incubated in 3% milk in TBST (13.7 mM NaCl, 2 mM Tris pH 7.5, Tween20) for 1 h before o/n incubation shaking at 4 °C with anti-NKX2-1 antibody (Cell Signaling Technologies, Cat #: 12373), diluted 1:1000 in 1%BSA in TBST. Following 4×5 min washes in TBST, the membranes were incubated for 1 h incubation, with 1:5000 diluted goat anti-rabbit HRP conjugated secondary antibody (Thermo Fisher Scientific, Cat #: 31460). Following 4 × 5 min washes in TBST, Supersignal West Pico Chemiluminescent Substrate (Thermo Fisher, Cat #: 34080) was used as the detection reagent prior to film exposure. Membranes were stripped using Pierce™ Restore™ PLUS Western Blot Stripping Buffer, Thermo Scientific PI46430r, prior to performing washing, incubation and detection steps as above for the loading control. For this, a beta-actin antibody (Cell Signaling Technology, Cat #: 3700S) was used at 1:5000 dilution, in combination with an HRP-conjugated goat anti-mouse antibody (Pierce, Cat #: 1858413,).
For GATA6, 1 × 106 cells were lysed with 100 µl cold RIPA Buffer and normalized by the protein quantification kit. A total of 20 μg of protein was used to perform the immunoblotting with Jess Automated Western Blot System (proteinsimple, Cat #: 004-650) following the standard manufacturer’s protocol. GATA6 (Cell Signaling Technology, Cat #: 5851) and beta-actin antibodies were used at 1:50 dilutions.
Incucyte growth assays
A total of 3000 LNCaP cells in 100 µl of RPMI + 10% FBS were seeded into a 96-well plate per condition (5 replicates each). Plates were read in an IncuCyte S3. Phase object confluence (percentage area) for cell growth was measured every 4 h.
Drug treatment with AR inhibitors and SMARCA2/4 degrader
The following cell lines were obtained from commercial sources as indicated: LNCaP Clone FGC (ATCC, Cat #: CRL-1740), VCAP (ECACC, Cat #: 6020201), DU145 (ATCC, Cat #: HTB-81), 22Rv1 (ATCC, Cat #: CRL-2505), NCI-H660 (ATCC, Cat #: CRL-5813) and MDA-PCa-2b (ATCC, Cat #: CRL-2422). Cell line authentication was routinely conducted by SNP-based genotyping using Fluidigm multiplexed assays at the Genentech cell line core facility. All cell lines used in this study tested negative for mycoplasma contamination. LNCaP, VCaP, DU145 and 22Rv1 were cultured with RPMI-1640 media with 10% FBS and 2 mM L-glutamine. NCI-H660 cells were cultured with DMEM/F12 with 0.005 mg/ml insulin, 0.01 mg/ml Transferrin, 30 nM Sodium selenite,10 nM Hydrocortisone,10 nM beta-estradiol, 4 mM L-glutamine. MDA-PCa-2b cells were cultured with HCP1 medium (Enzo, Cat #: AES-0403), 20% FBS, 1x Glutamine and pen/strep. The cells were split every 4–5 days to maintain the appropriate density. For the drug response assay, cells were seeded in six-well format. The cells were split every 4–5 days to maintain the appropriate density.
A total of 1.5 × 105 DU145 cells were seeded into each well of a six-well plate and 5 × 103 DU145 cells were seeded into each well of a 96-well plate; 1.5–2 × 106 LNCAP or VCAP cells were seeded into T75 flasks; 5 × 105 NCI-H660 cells were seeded into each well of a six-well plate. After 48 h, the media was removed and cells were treated with DMSO, 1 μM Enzalutamide, 0.1 μM SMARCA2_4.1 or 1 μM ARV-110 in complete media. After 24 h from treatment, the cells were harvested for the single-cell RNA-ATAC Co-assay to evaluate cell fitness, cells were seeded into a 96-well plate per condition. Viability was assessed by CellTiterGlo at 24 h and 5 days after treatment.
Immunoblotting
Cell lines were treated with drugs for 24 h, and cell pellets were lysed in cold RIPA lysis buffer (50 mM Tris pH 8, 150 mM NaCl, 0.1% Triton X-100, 0.5% sodium deoxycholate and 0.1% SDS) containing 0.5 M NaCl with protease inhibitors (Roche) on ice for 5 min, homogenized for 3 min at speed 10 (NextAdavance, Bullet BlenderR 24) and centrifuged at 15,000 × g for 5 min. Protein concentration was measured by Pierce BCA protein assay (Life Technologies). A total of 30 μg protein was resolved in 4–20% Tris–Glycine gel, and transferred to nitrocellulose membranes by iBlot. Membranes were incubated with primary antibodies overnight at 4 °C: SMARCA4 (Abcam, Cat #: ab11064), AR (Cell Signaling Technologies, Cat #: 5153s), HDAC1(Cell Signaling Technologies, Cat #: 34589S), then with IRDyeR secondary antibodies (LIC-926-32211) at room temperature for 1 h. Blots were imaged with Odyssey Imager for detection (LI-COR).
Single-cell RNA-seq and single-cell RNA-ATAC co-assay
The single-cell RNA-seq was performed using the Chromium Single Cell 3’ kit (V3.1) from 10X Genomics with cell hashing. Briefly, the cells were trypsinized into a single-cell suspension from the six-well plates and washed once with PBS with 1% BSA. The cells from different wells were stained with human TotalSeq-A cell hashing antibodies (BioLegend) containing distinct barcodes following the manufacturer’s protocol. The cells were then washed twice with PBS with 1% BSA before combining, and the live cells were sorted using the SONY SH800S FACS machine. For loading of the 10X chip G, we overloaded the channel with the aim of recovering 20K cells. The library construction was following the standard 10X protocol with the following exceptions: (1) the 1 µl of 10 µM HTO additive primer was spiked in during the cDNA PCR step; (2) during the SPRI beads cleanup step, the supernatant from the 0.6X cleanup was saved to recover the HTO fragment. Along with the transcriptome library, the HTO library was amplified from the supernatant using the HTO-specific primer. The detailed protocol can be found on the cell hashing website (https://cite-seq.com/cell-hashing/).
The single-cell RNA-ATAC co-assay was performed using the 10X Genomics Single Cell Multiome ATAC + Gene Expression kit. In a previous test, we found that the standard cell permeabilization protocol used for nuclei preparation does not completely remove the plasma membrane of the LNCaP cells and robust B2M expression can still be detected with FACS (data not shown). Therefore, we speculated that the same cell hashing strategy would work for the 10X Multiome assay. Briefly, we followed the same hashing protocol as used in the scRNA-seq experiment. After the antibody staining and pooling, the cells were permeabilized by following the nuclei isolation protocol from 10X genomics (10X Nuclei Isolation Protocol). The cells were then counted and loaded on the Chip J with a recovery aim of 12K cells per lane. For the library construction, we followed the standard protocol with the following modifications: (1) 1 µl of 10 µM HTO additive primer was spiked in during the pre-AMP and cDNA PCR steps; (2) during the 0.6X cDNA cleanup, the supernatant was saved to amplify the HTO library, as described in the previous section.
All the libraries were sequenced on Illumina NextSeq 2000 or NovaSeq 4000. We were aiming for 20K, 40K and 2K raw reads per cell for the transcriptome, ATAC and HTO libraries, respectively.
ChIP-seq
For the ChIP-Seq assay, chromatin was prepared from two biological replicates of MDA-PCa-2b cells treated with DMSO or SMARCA2_4.1 (100 nM) for 24 h. ChIP-Seq assay was then performed by Active Motif Inc. using antibodies against AR (Active Motif, Cat #: 39781), FOXA1 (Abcam, Cat #: ab5089) and SMARCA4 (Abcam, Cat #: ab110641). DU145 cells were treated under the same conditions, and ChIP-seq was performed at Genentech. ChIP-seq was performed as previously described with the following modifications63. DU145 cells (10 ×106) were crosslinked for 10 min of 1% formaldehyde. Formaldehyde was quenched by the addition of glycine. Nuclei were isolated with ChIP lysis buffer (1% Triton x-100, 0.1% SDS, 150 mM NaCl, 1 mM EDTA, and 20 mM Tris, pH 8.0). Nuclei were sheared with Covaris sonicator (E220) using the following setup: Fill level—10, Duty Cycle—15, PIP—350, Cycles/Burst—200, Time—8 min). Sheared chromatin was immunoprecipitated overnight with the following antibodies (5 µg antibody each): SMARCA4 (AbCAM, Cat #: ab110641), FOSL1 (Invitrogen, Cat #: PA5-66880) and TEAD1 (BD Biosciences, Cat #: 610923). Antibody chromatin complexes were pulled down with Protein A or Protein A/G magnetic beads and washed twice in IP wash buffer I. (1% Triton, 0.1% SDS, 150 mM NaCl, 1 mM EDTA, 20 mM Tris, pH 8.0, and 0.1% NaDOC), twice in IP wash buffer II (1% Triton, 0.1% SDS, 500 mM NaCl, 1 mM EDTA, 20 mM Tris, pH 8.0, and 0.1% NaDOC), twice in IP wash buffer III. (0.25 M LiCl, 0.5% NP-40, 1 mM EDTA, 20 mM Tris, pH 8.0, 0.5% NaDOC) and once in TE buffer (10 mM EDTA and 200 mM Tris, pH 8.0). DNA was eluted from the beads by vigorous shaking for 20-min in elution buffer (100 mM NaHCO3, 1% SDS). DNA was de-crosslinked overnight at 65 °C and purified with the MinElute PCR purification kit (Qiagen). Sequencing-ready libraries were generated using NEBNext ® Ultra™ II DNA Library Prep Kit for Illumina ® (New England Biolabs E7645). Libraries were quantified using the Qubit dsDNA HS Assay Kit (Thermo Fisher Scientific) and profiled using the D5000 ScreenTape on TapeStation 4200 (Agilent Technologies). Libraries were pooled and sequenced on an Illumina sequencer to generate 25 million paired-end 50-base pair reads per library.
ChIP-seq sequence data was processed using an ENCODE-DC/chip-seq-pipeline2-based workflow (https://github.com/ENCODE-DCC/chip-seq-pipeline2). Briefly, fastq files were aligned on the hg38 human genome reference using Bowtie2 (v2.2.6) followed by alignment sorting (samtools v1.7) of resulting bam files with filtering out of unmapped reads and keeping reads with mapping quality higher than 30. Duplicates were removed with Picard’s MarkDuplicates (v1.126) function, followed by indexation of the resulting bam files with samtools. For each bam file, genome coverage was computed with bedtools (v2.26.0), followed by the generation of bigwig (wigToBigWig v377) files. Peaks were called with SPP for each treatment sample using a pooled input alignment (.bam file) as a control. The peaks called are filtered using exclusion lists that contain genomic regions, resulting in anomalous, unstructured, or experiment-independent high signal62. For noise inherent in peak-calling of TF ChIP-seq experiments, the Irreproducible Discovery Rate framework is then used to adaptively threshold and retain peaks that are reproducible and rank-concordant across replicates. To assess data quality, we measured read mapping statistics, enrichment QC metrics, library complexity and reproducibility ratio (Refer to Supplementary Data 3). Only peaks passing q < 1 × 10−5 are retained for subsequent analysis. Replicate peaks were merged using DiffBind (v3.15.0)’s dba function with minOverlap = 2. Refer to Supplementary Data 3 for final peaks. Bigwigs corresponding to the change fold of ChIP-seq with respect to input controls were visualized using Deeptools (v3.5.0), with -b 500 -a 500 --skipZeros -p max/2 for computeMatrix and zMax and yMax set to 10 for SMARCA4, 15 for AR and 20 for FOXA1 with plotHeatmap.
Statistics and reproducibility
For single-cell experiments, we target at least 100 cells of good-quality cells per condition. Low-quality cells were removed based on TSS enrichment (>3) and number of ATAC-seq reads mapped to the nuclear genome (>1000). Moreover, doublets were removed using ArchR’s filterDoublets function. We performed technical replicates for our scMultiome data and generated duplicate ChIP-seq data. Immunoblotting experiments were performed twice. Drug treatments were done in biological replicates of 4 or more. Cells were seeded evenly into wells, and wells were randomly assigned to treatment conditions. Blinding cannot be performed because the identity of the treatment conditions is needed to ascertain the ground truth.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Raw data have been deposited into GEO SuperSeries GSE252883 with the following subseries: GSE251977 (AR drug dataset), GSE251978 (reprogram-seq) and GSE280803 (ChIP-seq). Processed single-cell multiomic data can be accessed via the scMultiome package, which is available through Bioconductor (>1.5.7). AR drug data can be retrieved as scMultiome::AR_drug(), reprogram-seq can be retrieved as scMultiome::reprogramSeq() and PBMC data can be retrieved as scMultiome:: PBMC_10x() Source data are provided with this paper.
Code availability
Epiregulon: https://github.com/xiaosaiyao/epiregulon. Epiregulon.extra: https://github.com/xiaosaiyao/epiregulon.extra. Epiregulon.archr: https://github.com/xiaosaiyao/epiregulon.archr. Epiregulon.book: https://xiaosaiyao.github.io/epiregulon.book/. scMultiome: https://github.com/xiaosaiyao/scMultiome. Epiregulon manuscript: https://github.com/xiaosaiyao/epiregulon.manuscript. Deep neural network model: https://github.com/xiaosaiyao/epiregulon.sequence.modeling. Differential network simulation: https://github.com/xiaosaiyao/epiregulon.diffnetwork.simulation.
References
Tran, C. et al. Development of a second-generation antiandrogen for treatment of advanced prostate cancer. Science 324, 787–790 (2009).
Sakamoto, K. M. et al. Protacs: chimeric molecules that target proteins to the Skp1-Cullin-F box complex for ubiquitination and degradation. Proc. Natl Acad. Sci. USA 98, 8554–8559 (2001).
Snyder, L. B. et al. Discovery of ARV-110, a first in class androgen receptor degrading PROTAC for the treatment of men with metastatic castration resistant prostate cancer. Cancer Res 81, 43 (2021).
Hagenbeek, T. J. et al. An allosteric pan-TEAD inhibitor blocks oncogenic YAP/TAZ signaling and overcomes KRAS G12C inhibitor resistance. Nat. Cancer 4, 812–828 (2023).
Badia, I. M. P. et al. Gene regulatory network inference in the era of single-cell multi-omics. Nat. Rev. Genet. https://doi.org/10.1038/s41576-023-00618-5 (2023).
Alvarez, M. J. et al. Functional characterization of somatic mutations in cancer using network-based inference of protein activity. Nat. Genet. 48, 838–847 (2016).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Kamimoto, K. et al. Dissecting cell identity via network inference and in silico gene perturbation. Nature 614, 742–751 (2023).
Kartha, V. K. et al. Functional inference of gene regulation using single-cell multi-omics. Cell Genom. 2, https://doi.org/10.1016/j.xgen.2022.100166 (2022).
Fleck, J. S. et al. Inferring and perturbing cell fate regulomes in human brain organoids. Nature 621, 365–372 (2023).
Kamal, A. et al. GRaNIE and GRaNPA: inference and evaluation of enhancer-mediated gene regulatory networks. Mol. Syst. Biol. 19, e11627 (2023).
Bravo Gonzalez-Blas, C. et al. SCENIC+: single-cell multiomic inference of enhancers and gene regulatory networks. Nat. Methods 20, 1355–1367 (2023).
Weber, B. N. et al. A critical role for TCF-1 in T-lineage specification and differentiation. Nature 476, 63–68 (2011).
Ting, C. N., Olson, M. C., Barton, K. P. & Leiden, J. M. Transcription factor GATA-3 is required for development of the T-cell lineage. Nature 384, 474–478 (1996).
Li, L., Leid, M. & Rothenberg, E. V. An early T cell lineage commitment checkpoint dependent on the transcription factor Bcl11b. Science 329, 89–93 (2010).
Woolf, E. et al. Runx3 and Runx1 are required for CD8 T cell development during thymopoiesis. Proc. Natl Acad. Sci. USA 100, 7731–7736 (2003).
Ono, M. et al. Foxp3 controls regulatory T-cell function by interacting with AML1/Runx1. Nature 446, 685–689 (2007).
Georgopoulos, K., Moore, D. D. & Derfler, B. Ikaros, an early lymphoid-specific transcription factor and a putative mediator for T cell commitment. Science 258, 808–812 (1992).
Anderson, K. L. et al. Myeloid development is selectively disrupted in PU.1 null mice. Blood 91, 3702–3710 (1998).
Wang, D., D’Costa, J., Civin, C. I. & Friedman, A. D. C/EBPalpha directs monocytic commitment of primary myeloid progenitors. Blood 108, 1223–1229 (2006).
Kim, S. et al. Transcription factor C/EBPalpha is required for the development of Ly6C(hi) monocytes but not Ly6C(lo) monocytes. Proc. Natl Acad. Sci. USA 121, e2315659121 (2024).
Tamura, A. et al. C/EBPbeta is required for survival of Ly6C(-) monocytes. Blood 130, 1809–1818 (2017).
Feinberg, M. W. et al. The Kruppel-like factor KLF4 is a critical regulator of monocyte differentiation. EMBO J. 26, 4138–4148 (2007).
Nechanitzky, R. et al. Transcription factor EBF1 is essential for the maintenance of B cell identity and prevention of alternative fates in committed cells. Nat. Immunol. 14, 867–875 (2013).
Mikkola, I., Heavey, B., Horcher, M. & Busslinger, M. Reversion of B cell commitment upon loss of Pax5 expression. Science 297, 110–113 (2002).
Kim, U. et al. The B-cell-specific transcription coactivator OCA-B/OBF-1/Bob-1 is essential for normal production of immunoglobulin isotypes. Nature 383, 542–547 (1996).
Lin, Y. C. et al. A global network of transcription factors, involving E2A, EBF1 and Foxo1, that orchestrates B cell fate. Nat. Immunol. 11, 635–643 (2010).
Gordon, S. M. et al. The transcription factors T-bet and Eomes control key checkpoints of natural killer cell maturation. Immunity 36, 55–67 (2012).
Kallies, A. et al. A role for Blimp1 in the transcriptional network controlling natural killer cell maturation. Blood 117, 1869–1879 (2011).
Sichien, D. et al. IRF8 transcription factor controls survival and function of terminally differentiated conventional and plasmacytoid dendritic cells, respectively. Immunity 45, 626–640 (2016).
Grajkowska, L. T. et al. Isoform-specific expression and feedback regulation of E protein TCF4 control dendritic cell lineage specification. Immunity 46, 65–77 (2017).
Intlekofer, A. M. et al. Effector and memory CD8+ T cell fate coupled by T-bet and eomesodermin. Nat. Immunol. 6, 1236–1244 (2005).
Cytlak, U. et al. Differential IRF8 transcription factor requirement defines two pathways of dendritic cell development in humans. Immunity 53, 353–370.e358 (2020).
Xiao, L. et al. Targeting SWI/SNF ATPases in enhancer-addicted prostate cancer. Nature 601, 434–439 (2022).
Bluemn, E. G. et al. Androgen receptor pathway-independent prostate cancer is sustained through FGF signaling. Cancer Cell 32, 474–489.e476 (2017).
Dong, B. et al. Single-cell analysis supports a luminal-neuroendocrine transdifferentiation in human prostate cancer. Commun. Biol. 3, 778 (2020).
Hieronymus, H. et al. Gene expression signature-based chemical genomic prediction identifies a novel class of HSP90 pathway modulators. Cancer Cell 10, 321–330 (2006).
Nyquist, M. D. et al. Combined TP53 and RB1 loss promotes prostate cancer resistance to a spectrum of therapeutics and confers vulnerability to replication stress. Cell Rep. 31, 107669 (2020).
Tang, F. et al. Chromatin profiles classify castration-resistant prostate cancers suggesting therapeutic targets. Science 376, eabe1505 (2022).
Cyrta, J. et al. Role of specialized composition of SWI/SNF complexes in prostate cancer lineage plasticity. Nat. Commun. 11, 5549 (2020).
Duan, J. et al. Rational reprogramming of cellular states by combinatorial perturbation. Cell Rep. 27, 3486–3499.e3486 (2019).
Baca, S. C. et al. Reprogramming of the FOXA1 cistrome in treatment-emergent neuroendocrine prostate cancer. Nat. Commun. 12, 1979 (2021).
Han, M. et al. FOXA2 drives lineage plasticity and KIT pathway activation in neuroendocrine prostate cancer. Cancer Cell 40, 1306–1323.e1308 (2022).
Kelley, D. R. Cross-species regulatory sequence activity prediction. PLoS Comput. Biol. 16, e1008050 (2020).
Avsec, Z. et al. Base-resolution models of transcription-factor binding reveal soft motif syntax. Nat. Genet. 53, 354–366 (2021).
Avsec, Z. et al. Effective gene expression prediction from sequence by integrating long-range interactions. Nat. Methods 18, 1196–1203 (2021).
Terekhanova, N. V. et al. Epigenetic regulation during cancer transitions across 11 tumour types. Nature 623, 432–441 (2023).
Ji, P. et al. Kruppel-like factor 9 suppressed tumorigenicity of the pancreatic ductal adenocarcinoma by negatively regulating frizzled-5. Biochem. Biophys. Res. Commun. 499, 815–821 (2018).
Henley, M. J. & Koehler, A. N. Advances in targeting ‘undruggable’ transcription factors with small molecules. Nat. Rev. Drug Discov. 20, 669–688 (2021).
Arruabarrena-Aristorena, A. et al. FOXA1 mutations reveal distinct chromatin profiles and influence therapeutic response in breast cancer. Cancer Cell 38, 534–550 e539 (2020).
Liang, J. et al. Giredestrant reverses progesterone hypersensitivity driven by estrogen receptor mutations in breast cancer. Sci. Transl. Med. 14, eabo5959 (2022).
Pomerantz, M. M. et al. The androgen receptor cistrome is extensively reprogrammed in human prostate tumorigenesis. Nat. Genet. 47, 1346–1351 (2015).
Kiani, K., Sanford, E. M., Goyal, Y. & Raj, A. Changes in chromatin accessibility are not concordant with transcriptional changes for single-factor perturbations. Mol. Syst. Biol. 18, e10979 (2022).
Granja, J. M. et al. ArchR is a scalable software package for integrative single-cell chromatin accessibility analysis. Nat. Genet. 53, 403–411 (2021).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902.e1821 (2019).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e3529 (2021).
Gaublomme, J. T. et al. Nuclei multiplexing with barcoded antibodies for single-nucleus genomics. Nat. Commun. 10, 2907 (2019).
Schep, A. N., Wu, B., Buenrostro, J. D. & Greenleaf, W. J. chromVAR: inferring transcription-factor-associated accessibility from single-cell epigenomic data. Nat. Methods 14, 975–978 (2017).
Weirauch, M. T. et al. Determination and inference of eukaryotic transcription factor sequence specificity. Cell 158, 1431–1443 (2014).
Feng, C. et al. KnockTF: a comprehensive human gene expression profile database with knockdown/knockout of transcription factors. Nucleic Acids Res. 48, D93–D100 (2020).
Amemiya, H. M., Kundaje, A. & Boyle, A. P. The ENCODE Blacklist: identification of problematic regions of the genome. Sci. Rep. 9, 9354 (2019).
Daniel, B., Balint, B. L., Nagy, Z. S. & Nagy, L. Mapping the genomic binding sites of the activated retinoid X receptor in murine bone marrow-derived macrophages using chromatin immunoprecipitation sequencing. Methods Mol. Biol. 1204, 15–24 (1204).
Acknowledgements
We would like to thank Xinpeng Wang, Fang Zhang, Mingtao He, Yunxing Cheng and Michael Berlin (Arvinas) for the synthesis of SMARCA2_4.1 and Peter Dragovich for guiding the synthesis. We would like to thank Jayaram Kancherla and Amber Schedlbauer for their help and advice on single-cell data. Parts of Figs. 1a, b, 2a, 4d, g, 7a–c and Supplementary Fig. 4 were created with BioRender.
Author information
Authors and Affiliations
Contributions
X. Yao conceived Epiregulon. T.W., A.L., S.Y.C., A. Chlebowski, and X. Yao wrote the code in the Epiregulon and scMultiome packages, with contribution from T.K. and Y.G.. A.W. contributed to Epiregulon deployment. Y.W., M. He., A. Chibly, Q.Y. and Z.D. generated data and/or provided guidance on Epiregulon benchmarking. R.Y. and S.X. conceived the AR-related experiments, and D.W., M.S., X. Ye, L.W., and W.Z. performed the experiments. L.F.C., B.D., A.H., and Y.L. contributed to ChIP-seq data. C.W.S. and S.X. conceived the reprogram-seq experiments and D.W., J.T., and K.S. performed the experiments. M.R.C. conceived the use of DNN model and S.M. and S.T. performed the analysis. C.M. provided guidance on AR biology and supervised the AR-related experiments. M. Hafner provided guidance on GRN. X. Yao, A.L. and T.W. wrote the manuscript, with contribution from S.T. and T.K.
Corresponding authors
Ethics declarations
Competing interests
T.W., A.L., D.W., M.S., X. Ye, S.T., K.S., L.W., J.T., S.Y.C., T.K., A. Chlebowski, A.W., W.Z., Y.W., Y.G., L.F.C., B.D., A.H., M. He, A.Chibly, Y.L., C.M., M.Hafner., C.W.S., R.Y., S.X. and X.Yao are or were employees of Genentech Inc. or Roche. A.L., D.W., M.S., X. Ye, K.S., L.W., J.T., A.W., W.Z., Y.W., L.F.C., B.D., A.H., M. He, A.Chibly, Y.L., C.M., M.Hafner, C.W.S., R.Y., S.X. and X.Yao were given Roche stocks.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Włodarczyk, T., Lun, A., Wu, D. et al. Epiregulon: Single-cell transcription factor activity inference to predict drug response and drivers of cell states. Nat Commun 16, 7118 (2025). https://doi.org/10.1038/s41467-025-62252-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-62252-5