Abstract
KhpB, also known as EloR, is a recently discovered global RNA-binding protein in various pathogenic bacteria that regulates critical cellular processes. KhpB is unique in containing both an R3H domain and a KH domain, which are universal RNA/DNA-binding domains found across various proteins involved in diverse cellular functions. However, the precise roles of these domains in KhpB’s RNA-binding mechanism remain unclear, particularly as no structural data of the R3H domain bound to RNA/DNA have been reported for any protein. In this study, we present the crystal structures of both the RNA-free and RNA-bound forms of Thermus thermophilus KhpB dimer. These structures reveal that the KH and R3H domains cooperate to form a composite RNA-binding site capable of binding a single RNA molecule. Notably, the coordinated interaction requires RNA molecules that are at least 7 nucleotides long. This interaction induces conformational changes, including the closure of the RNA-binding cleft between the two domains. The structural data further reveal that KhpB primarily interacts with the phosphate backbone of RNA, while most of the base moieties remain solvent-exposed. These findings provide structural insights into the molecular function of KhpB and shed light on the RNA-binding strategies of other R3H domain-containing proteins.
Similar content being viewed by others
Introduction
Post-transcriptional regulation of gene expression plays a critical role in bacterial physiology and virulence. Bacterial post-transcriptional regulation has been extensively studied in Gram-negative bacteria, where it is primarily mediated by three key RNA-binding proteins: CsrA (also known as RsmA), Hfq, and ProQ. CsrA binds to the Shine-Dalgarno sequence in the 5’ UTR of mRNAs1, either repressing or enhancing translation. This regulation is modulated by sRNAs binding to CsrA. Hfq and ProQ, on the other hand, act as RNA chaperones that facilitate intra and inter molecule base-pairing interactions of various kinds of RNAs2,3,4,5. Hfq recognizes A-rich region in the 5’ UTR of mRNAs and the complementary U-rich region in the 3’ ends of sRNAs6 to promote hybridization of these RNA molecules. In contrast, ProQ binds and stabilizes specific secondary structures in both target mRNAs and sRNAs, thereby assisting their proper folding and structural organization1,5.
In addition to those RNA-binding proteins, KhpA and KhpB/EloR have recently been identified as global RNA-binding proteins in some Gram-positive and -negative bacteria. KhpB was initially discovered as a regulator of cell elongation in Streptococcus pneumoniae7. Subsequent studies demonstrated that the deletion of the khpB gene leads to increased expression of the ftsA gene in S. pneumoniae and that this regulatory effect requires a 74-nt sequence in the 5’ UTR of the ftsA mRNA. RNA immunoprecipitation experiments further revealed that KhpB interacts with the 5’ UTR of the ftsA mRNA8. These findings suggest that KhpB binds to mRNA and regulates translation at the post-transcriptional level. Cross-linking experiments provided evidence for a direct intracellular interaction between KhpB and KhpA, highlighting the importance of this interaction in post-transcriptional regulation9. KhpA has been shown to be capable of dimer formation and to exist as both a homodimer and a heterodimer with KhpB9. In some other pathogens such as Clostridioides difficile, Enterococcus faecalis, and Enterococcus faecium, studies employing the Grad-seq technique have revealed that KhpB is a global RNA-binding protein that interacts with a wide variety of RNAs and regulates the expression of diverse genes, including those involved in pathogenicity10,11.
More recently, in a Gram-negative bacterium, Fusobacterium nucleatum, KhpA and KhpB were found to interact with various types of RNAs including 5’ UTR, coding sequence, 3’ UTR, rRNA, tRNA, and sRNA, as revealed by RNA immunoprecipitation experiments12. KhpB binds to a particularly wide range of sRNAs among these RNAs and affects their stability12. Gel shift assays showed that KhpB directly binds to RNA and that KhpA enhances KhpB’s RNA binding capacity, even though KhpA itself lacks RNA-binding ability12. F. nucleatum, one of the human oral bacteria13, has recently been implicated in the progression of breast and colorectal cancers by localizing within the tumors14,15,16. Notably, F. nucleatum lacks the RNA-binding proteins Hfq and ProQ12, making KhpB the main global RNA-binding protein and a central regulator of their morphology and virulence. Thus, KhpB has been a critical target for understanding the global regulation of gene expression in this clinically significant bacterium.
The amino acid sequence of KhpB includes the K-homology (KH) and R3H domains in its middle and C-terminal regions, while the N-terminal region varies among species (Fig. 1A). The KH domain is a highly conserved RNA-binding domain found in various RNA-binding proteins across all kingdoms of life17,18. This domain is characterized by a conserved GXXG amino acid sequence motif located between two helices, which are flanked by β-strands. KhpA also contains a KH domain but lacks additional domains8,12. Similar to the KH domain, the R3H domain is known as a single-stranded DNA/RNA-binding domain that is typically found in combination with other nucleotide-binding domains in proteins with diverse functions19,20,21. The R3H domain is defined by a conserved R3H (RXXXH) motif within an α-helix. Three-dimensional structures of the KH domains from various proteins in complex with RNA have been extensively studied18,22,23,24, revealing their RNA-binding mechanisms. On the other hand, although the structure of the R3H domain of human Sμbp-2 in complex with dGMP has been solved21, its complex with DNA/RNA has not been solved and its binding mode has not been elucidated.
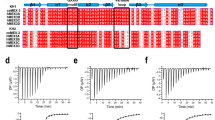
A Amino acid sequence alignment of KhpB proteins from T. thermophilus (accession number BAD70265.1), C. symbiosum (WP_227031341), S. pneumoniae (COD10839.1), and F. nucleatum (AAL94218.1). The amino acid sequences of KhpB proteins were aligned by CLUSTALW42 and visualized using the ESPript3 program43. For simplicity, the N-terminal 41 residues of F. nucleatum KhpB were omitted. Black bars beneath the sequences indicate the GXXG and R3H amino acid sequence motifs of the KH and R3H domains. B Crystal structure of the ttKhpB monomer. Two orthogonal views of the RNA-free ttKhpB monomer are displayed. The positions of the N- and C-termini are labeled as “N” and “C”. C Gel filtration analysis of ttKhpB. The elution profile of ttKhpB is shown as a plot of relative absorbance at 280 nm versus elution volume. The void volume of the column is indicated by “vo”. Source data are provided as a Source Data file. D The crystal structure of ttKhpB dimer is shown in two orthogonal views. The two subunits are distinguished by color intensity: one subunit is shown in darker colors and the other in lighter colors. The KH and R3H domains are shown in red and blue colors, respectively. E Hydrophobic dimeric interface of ttKhpB. The surface of the KhpB monomer is color-coded based on hydrophobicity/hydrophilicity, illustrating the predominantly hydrophobic nature of the dimeric interface.
KhpB is unique in possessing both a KH domain and an R3H domain, making it an intriguing candidate for investigating an RNA-binding mechanism involving these domains. Previously, the crystal structure of RNA-free KhpB homodimer from Clostridium symbiosum was determined and deposited in the Protein Data Bank (PDB ID: 3GKU), although it has not been described in a published study. However, the structure of KhpB in complex with RNA remains to be solved. Thermophilic bacterial proteins are often employed as model systems in structural biology due to their enhanced stability and high success rates in crystallization25. In this study, we determined the crystal structure of KhpB from an extremely-thermophilic Gram-negative bacterium Thermus thermophilus (ttKhpB) in both RNA-free and RNA-bound forms. These structures unveil the mechanism of coordinated RNA binding by the KH and R3H domains and provide valuable insights into the substrate recognition of KhpB.
Results and discussion
Crystal structure of the RNA-free form of the ttKhpB dimera
The full length of ttKhpB (189 amino acid residues) was purified and crystallized in an RNA-free form. The 2.99-Å diffraction data were phased by molecular replacement using the model structure predicted by AlphaFold226. The model structure was refined to the Rwork and Rfree values of 24.9% and 27.0%, respectively (Fig. 1, Supplementary Fig. 1A, and Table 1). The N-terminal 37 residues were invisible, indicating that the region is flexible or unstructured in the crystal. This observation aligns with the AlphaFold2-predicted structure, which also indicates that the N-terminal 37 residues of ttKhpB is likely unstructured (Supplementary Fig. 2). The KH domain of ttKhpB consisted of the α1, α2, and α3 helices and β1, β2, and β3 strands, while the R3H domain comprises the α4 and α5 helices and β4 and β5 strands (Fig. 1A, B). The overall structure of the KH and R3H domains of ttKhpB closely resembled that of C. symbiosum KhpB (PDB ID: 3GKU) with a Cα RMSD value of 4.1 Å (Supplementary Fig. 3A).
The crystal of the RNA-free form of ttKhpB contained one molecule per asymmetric unit. In the gel filtration analysis, the elution profile of ttKhpB (a calculated molecular mass of 21 kDa) exhibited a peak corresponding to an apparent molecular mass of 45 kDa, indicating that ttKhpB predominantly exists as a dimeric form in solution (Fig. 1C). In the crystal, the KhpB molecule interacted with neighboring molecules, with the KH domain contributing a particularly large interacting surface. The interface at the KH domain was composed of dense clusters of hydrophobic amino acid residues located on the α1 and α3 helices (Fig. 1D, E), which was proposed to be a biological dimeric interface by the PISA (Proteins, Interfaces, Structures and Assemblies) program27 (Table S1). AlphaFold328 also predicted a dimeric interface consisting of the α1 and α3 helices (Supplementary Fig. 2B). This dimeric interface is conserved in C. symbiosum KhpB (Supplementary Fig. 3B) and the RNA-bound form of ttKhpB, which is described later. Furthermore, the same interface has been reported as the biologically relevant dimer interface in the structurally related protein KhpA9. Collectively, the present results, together with previous reports, suggest that the α1 and α3 helices of the KH domain form the dimeric interface in ttKhpB.
Coordinated RNA-binding by the KH and R3H domains of ttKhpB
Prior to the crystallization of the ttKhpB-RNA complex, the RNA-binding capacity of ttKhpB was evaluated using a gel shift assay. Cy5-labeled RNAs of various lengths were mixed with ttKhpB and analyzed by native polyacrylamide gel electrophoresis. As shown in Fig. 2, a shift in the RNA signals, indicating ttKhpB binding, was observed for RNAs longer than 7 nucleotides, with the binding intensity saturating at 9 nucleotides. Based on these results, a 10-mer single-stranded RNA (5’-CCCCCCCCCC-3’) was selected for cocrystallization with ttKhpB. The model structure for the 2.65-Å diffraction data was refined to the Rwork and Rfree values of 20.3% and 26.2%, respectively (Table 1 and Supplementary Fig. 1B). Clear electron density for the RNA molecule was observed (Supplementary Fig. 4), enabling the construction of a detailed ttKhpB-RNA complex model for the data (Fig. 3A).
A An electrophoretic mobility shift assay was performed to investigate the RNA-binding property of ttKhpB. 5’-Cy5-labeled single-stranded RNAs of various lengths (5-mer to 10-mer) were tested. The RNA-protein complexes were resolved on a 10–20% gradient polyacrylamide gel, and shifted signals were visualized. B Quantitative analysis of RNA binding. The percentages of shifted RNA signals (%Shifted) from (A) were quantified and plotted against ttKhpB protein concentrations. The data reveal the binding behavior of ttKhpB toward RNAs of various lengths. Average values from three independent experiments are shown with standard deviations. Source data are provided as a Source Data file.
A Crystal structure of the RNA-bound form of the ttKhpB dimer. ttKhpB and RNA are shown in the cartoon and stick models, respectively. The KH and R3H domains of each subunit are color-coded as in Fig. 1D. The residues in the observed 7 nucleotides of the RNA are labeled from C1 to C7 in the direction from the 5’ to 3’ end. B Surface representation of the RNA-bound form. The same structure as in (A) is shown with ttKhpB in a sphere model, highlighting the RNA-binding groove. C Electrostatic surface was calculated using the Adaptive Poisson-Boltzmann Solver program and visualized with PyMOL. A positively charged surface patch (in blue) is observed at the RNA-binding site.
The crystal structure contained two ttKhpB molecules in each asymmetric unit, similar to the RNA-free structure, forming dimers with the α1 and α3 helices of the KH domain as the dimeric interface (Fig. 3A). The ttKhpB dimer had two RNA-binding sites: we observed electron density for 7 nucleotides of the 10-mer RNA bound at one site, while only a single nucleotide was visible at the other site (Supplementary Fig. 5A). Apparently, crystal packing interfered with one of the two RNA-binding sites, likely displacing most of the bound RNA and leaving only one nucleotide of the RNA visible (Supplementary Fig. 5B). Therefore, our discussion of the RNA-binding mode of ttKhpB will focus hereafter on the binding site where 7-mer RNA binding was observed.
The RNA was bound within a groove formed by one R3H domain and two KH domains in the ttKhpB dimer (Fig. 3B). The 5’-terminal side of the RNA interacted with the R3H domain of one subunit, while the 3’-terminal side of the RNA was bound by the KH domains of the same subunit and the other subunit. Thus, the RNA was captured by coordinated binding of the KH and R3H domains. In this binding mode, RNA needed to be at least 7 nucleotides long to bridge the two binding sites. This structural feature is consistent with the gel shift assay results, which demonstrated that ttKhpB required an RNA length of at least 7 nucleotides for binding. In this paper, the residues in the observed 7-mer RNA will be referred to as C1 through C7, in order from 5’ to 3’. Furthermore, electrostatic surface representation revealed a positively charged surface patch that coincides with the RNA-binding site (Fig. 3C).
The solvent-accessible surface area buried upon RNA binding was calculated using the PISA program. The total buried surface area of the RNA was 658.5 Ų. The buried surface area per nucleotide is summarized in Supplementary Table 2. Notably, nucleotides C1 to C4 exhibited relatively large buried surface areas, suggesting that these residues are more tightly engaged in the interaction with KhpB.
Specific interactions with RNA in the KH and R3H domains
At the RNA-binding site of the R3H domain, Arg153 and His157, key residues of the R3H motif, were directly involved in the RNA binding (Fig. 4A, left panel). The positively charged side chain of Arg153 formed an electrostatic interaction with the negatively charged backbone phosphate group of the RNA. Additionally, the imidazole ring of the His157 side chain stacked with the pyrimidine ring of C1. The RNA backbone phosphate group of C1 also interacted with the side chain of Arg177, contributing to the RNA recognition. These interactions are consistent with those observed between the R3H domain of human Sμbp-2 and dGMP21. To confirm the involvement of the R3H motif in RNA binding, the R153A/H157A double mutant form of ttKhpB was generated, and its RNA-binding ability was evaluated using electrophoretic mobility shift assay. As shown in Fig. 4B, C, the R153A/H157A double mutation impaired the RNA-binding ability of ttKhpB. Circular dichroism (CD) spectroscopy showed that the mutant protein had a spectrum nearly identical to that of the wild-type protein (Supplementary Fig. 6), suggesting that the overall structure was preserved despite the mutations. These results indicate that the integrity of the R3H motif is important for RNA binding by ttKhpB. However, the R3H domain interacted with only a single nucleotide of the longer RNA, indicating that other domains are required for stable RNA binding. This may explain why the R3H domain does not exist alone but instead coexists with other nucleic acid-binding domains.
A Detailed view of the RNA-binding sites. The RNA-binding sites in the R3H domain (left) and the KH domain (right) are shown. Residues involved in the interactions with RNA are depicted as stick models, with dotted lines indicating specific interactions. Arg 153 and His157 are components of the R3H motif. The prime symbol indicates that the residue is from the opposite subunit. B Effects of mutations on the RNA-binding ability of ttKhpB. Electrophoretic mobility shift assay was performed using a 10-mer RNA to assess the effect of the G83D and R153A/H157A mutations in ttKhpB. C The %Shifted values in the panel B were quantified and plotted against protein concentrations. For reference, the binding curve of the wild type (reproduced from Fig. 2B) is included. Average values from three independent experiments are shown with standard deviations.
At the RNA-binding site formed by the two KH domains, the phosphate groups of the C2 to C6 backbone were captured by the side chains of multiple basic amino acid residues, including His94, Arg97, and Arg102 from one KH domain and Arg80 from the other KH domain (Fig. 4A, right panel). Thus, the formation of the composite RNA-binding site relies on the dimerization of the two ttKhpB molecules.
Interestingly, the GXXG motif-containing loop appeared to interact only loosely with the RNA through van der Waals interactions without forming any specific interactions (Supplementary Fig. 7). In another KH domain protein, it has been reported that replacing the first Gly in the GXXG motif by an Asp decreased the RNA-binding ability29. To test this in ttKhpB, the G83D mutation was introduced into ttKhpB to replace the first Gly in the GXXG motif by an Asp. CD spectroscopy showed that the spectrum of the G83D mutant protein was essentially identical to that of the wild-type protein (Supplementary Fig. 6). This mutation, however, did not affect the RNA-binding ability of ttKhpB (Fig. 4B, C and Table 2). These findings suggest that the GXXG motif of ttKhpB does not participate in specific interaction with RNA.
In the crystal structure, the 2’-OH groups of RNA are not recognized by KhpB. Consistent with this, we found that it binds to a single-stranded DNA with comparable affinity to a single-stranded RNA (Supplementary Fig. 8 and Table 2). A similar phenomenon has been reported for other RNA-binding proteins, such as heterogenous nuclear ribonucleoprotein, protein kinase R, and bacterial cold shock protein B, that bind to both RNA and DNA with comparative affinities30,31,32. In cells, free single-stranded DNAs are thought to be rarely present33, whereas single-stranded RNAs are overwhelmingly abundant. Therefore, ttKhpB is presumed to preferentially bind to single-stranded RNAs in the cellular context.
As described earlier, ttKhpB primarily interacts with the backbone phosphate groups of RNA, with limited interactions involving the base moieties. Among the interactions, only Thr106 forms a contact with the amino group of cytosine base at C6. To assess the importance of the exocyclic amino group of cytosine at C6, we tested an RNA that lacks cytosine (Sequence_3: 5’-GGUGGUUGUG-3’). The dissociation constant (Kd) value for this RNA was comparable to that of the cytosine-rich RNA (Sequence_1: 5’-CCCCCCCCCC-3’) (Supplementary Fig. 9B and Table S3), indicating that the amino group of C6 does not play a critical role in the KhpB–RNA binding.
We further performed the same binding analysis using 8 additional RNA molecules of an identical length but different sequences. These included RNAs composed exclusively of pyrimidine bases, purine bases, or various combinations thereof. All of them exhibited Kd values comparable to that for Seq_1 (Supplementary Fig. 9 and Table S3). These results support the previous reports that KhpB binds a wide variety of RNA sequences in vivo10,11. However, we cannot exclude the possibility that KhpB preferentially recognizes specific RNA sequences with higher affinity. To identify such sequence preferences, comprehensive approaches such as SELEX would be required in future studies.
RNA-dependent conformational changes of ttKhpB
A comparison of the RNA-free and RNA-bound forms of ttKhpB revealed conformational changes upon RNA binding. Specifically, the orientation of the α4 helix in the R3H domain shifted, closing the groove between the R3H and KH domains (Fig. 5A). As the KH and R3H domains approached each other, a hydrogen bond was formed between the main chain of Glu175 in the R3H domain and the side chain of Glu85 in the GXXG motif of the opposite subunit (Supplementary Fig. 7). Changes were also observed in the dimeric interface of the KH domain upon RNA binding (Fig. 5B, C). In the RNA-bound form, the contact area of the dimeric interface, composed of the α1 and α3 helices, increased. The interface areas, estimated using PISA, were 657 Ų in the RNA-free form and 702 Ų in the RNA-bound form. In the RNA-free structure, the imidazole group of His94 and hydroxy group of Thr88 were 8.5 Å apart, whereas, in the RNA-bound structure, the distance between them was 2.7 Å, suggesting the formation of a hydrogen bond (Fig. 5C, right panel).
A Comparison of the RNA-free (orange) and RNA-bound (blue) forms of ttKhpB monomer. ttKhpB and RNA are shown in the cartoon and stick models, respectively. The movement of the R3H domain upon RNA binding is indicated by an arrow. B The dimeric interface (α1 and α3 helices) of the RNA-free form of ttKhpB is shown in the cartoon model. The helices from the two subunits are color-coded orange and yellow-orange. Amino-acid side chains contributing to hydrophobic interactions between the monomers are indicated in the stick model. C The dimeric interface of the RNA-bound form of ttKhpB is shown in two orthogonal views. Side chains of Thr88 and His94 are also shown in the stick model. The hydrogen bonds between these two residues are indicated by dotted lines. The red underlined residues represent interaction sites that were not observed in the RNA-free form. Source data are provided as a Source Data file. D The overall dimeric structures of the RNA-free (upper) and RNA-bound (lower) forms of ttKhpB were superimposed based on the structural similarity of the KH domain of subunit A and displayed separately. Red triangles indicate the positions of the C-termini, showing the conformational shift induced by RNA binding. E The Cα-Cα distance between the RNA-free and RNA-bound forms of ttKhpB in (D) was calculated for each amino acid residue. Source data are provided as a Source Data file.
These changes in intra- and intermolecular interactions resulted in alterations to the quaternary structure (Fig. 5D). Superimposing the RNA-free and RNA-bound structures of ttKhpB based on the structural similarity of the KH domain in subunit A (the subunit involved in 7-mer RNA binding) revealed a positional shift in the other subunit (subunit B) (Fig. 5E). The distance between the C-termini of the free and RNA-bound forms was 36 Å. To further quantify the conformational differences between the RNA-free and RNA-bound forms, we conducted a difference-distance matrix analysis34 (Supplementary Fig. 10). This analysis revealed that the major difference arises from the relative positioning between the KH and R3H domains, while maintaining their domain structures.
Interestingly, the two subunits in the RNA-bound form of ttKhpB exhibited identical structures, despite only a single nucleotide of the RNA being identified in one subunit (Supplementary Fig. 11). As noted earlier, a single RNA-binding site in ttKhpB is formed by the KH and R3H domains from one subunit and the KH domain from the other subunit. Given this structural property of the composite RNA-binding site, RNA binding on one side of the dimer likely induces conformational changes in both subunits. This observation raises the possibility of positive cooperativity between the two binding sites of ttKhpB. Among other KH proteins, some are known to exhibit positive cooperativity based on multimerization at the KH domain interface35. To examine the cooperativity between the two RNA-binding sites of ttKhpB, we analyzed the RNA concentration dependence of complex formation using electrophoretic mobility shift assay (Supplementary Fig. 12). The binding data were fitted to the Hill equation, which yielded a Hill coefficient of 1.7. This result suggests that the two RNA-binding sites exhibit positive cooperativity, where binding of RNA to one site enhances the affinity at the second site.
Model structure of the KhpA/KhpB heterodimer
KhpA is a small protein composed solely of a KH domain. Previous studies reported that KhpAs from F. nucleatum and S. pneumoniae can form both homodimers and heterodimers with KhpBs. This raises the possibility that KhpB may exist in the cell as a heterodimer with KhpA, as well as in its homodimeric form. T. thermophilus has KhpA (ttKhpA) in addition to ttKhpB. To investigate the potential formation of the ttKhpA/ttKhpB heterodimer in vitro, we tried to express recombinant ttKhpA in E. coli. However, this attempt was unsuccessful, preventing structural analysis of the heterodimer.
Thus, the structure of the ttKhpA/ttKhpB heterodimer is discussed based on a prediction made by AlphaFold328. ttKhpA was predicted to have a structure highly similar to the KH domain of ttKhpB (Supplementary Fig. 13A). AlphaFold3 also predicted a structural model of the ttKhpA/ttKhpB heterodimer, in which the two proteins interacted through the interface formed by the α1 and α3 helices of their KH domains (Supplementary Fig. 13B). This suggested that the dimerization mode of the ttKhpA/ttKhpB heterodimer is nearly identical to that of the ttKhpB homodimer.
Based on this prediction, it is expected that the RNA-binding mode of the ttKhpA/ttKhpB heterodimer would be similar to that of the ttKhpB homodimer. However, the ttKhpA/ttKhpB heterodimer contains only one R3H domain, and, therefore, the heterodimer has only one RNA-binding site. This structural difference might lead to variations in RNA-binding specificity or affinity between the ttKhpB homodimer and the ttKhpA/ttKhpB heterodimer. In particular, it should be noted that positive cooperativity, which might occur in RNA binding of the ttKhpB homodimer due to its symmetrical composite RNA-binding sites, is unlikely to operate in RNA binding of the ttKhpA/ttKhpB heterodimer.
Comparison of KhpB with other bacterial global RNA-binding proteins
Hfq and ProQ function as global RNA chaperones: Hfq facilitates the annealing of sRNAs to their mRNA targets by stabilizing transient RNA–RNA interactions, whereas ProQ promotes the folding and structural stabilization of individual RNA molecules. Structural analyses have revealed that Hfq primarily interacts with the RNA phosphate backbone, binding RNA in a manner that leaves several base moieties solvent-exposed36,37. Similarly, ProQ interacts predominantly with the RNA backbone and makes only limited sequence-specific contacts with base moieties38. These binding modes allow the bound RNA to remain partially accessible and capable of forming base-pairing interactions both within the same RNA molecule and between different RNA molecules.
Our structural analyses, in this study, indicate that ttKhpB adopts a similar RNA recognition strategy. ttKhpB primarily interacts with the phosphate backbone of RNA, without accommodating the nucleobases in specific binding pockets. In our RNA-bound structure, a large part of the base moieties is exposed to solvent, resembling the binding modes observed for Hfq and ProQ. This spatial configuration may allow the bound RNA to engage in duplex formation. It is therefore possible that ttKhpB also functions as an RNA chaperone, facilitating either the annealing of two distinct RNA molecules or the formation of secondary structures within a single RNA molecule, in a manner analogous to the proposed mechanisms of Hfq and ProQ. Further biochemical and functional studies will be required to test this hypothesis.
In this study, we elucidated the structural basis for RNA recognition by the bacterial RNA-binding protein KhpB from Thermus thermophilus. The crystal structures of RNA-free and RNA-bound forms revealed that the tandem KH and R3H domains cooperatively bind a single-stranded RNA molecule with each domain contributing to interactions along the phosphate backbone. The positively charged surface spanning both domains forms a continuous RNA-binding interface, enabling recognition of 7 nucleotides. Notably, RNA binding induced conformational changes in both protomers of the dimer, supporting a positively cooperative manner of binding. These results would provide a basis for further studies on the role of KhpB in global RNA regulation.
Methods
Construction of expression plasmids
The DNA fragment coding ttKhpB was amplified by PCR using the T. thermophilus HB8 genomic DNA as the template with the following primers: 5’-TTCATATGAACGAGCGCAAGAAGAG-3’ (forward) and 5’-TTAGATCTTTACACCACGTGCCGCTCC-3’ (reverse). The forward and reverse primers contained NdeI and BglII recognition sites, respectively (underlines). The amplified DNA fragment was digested by the restriction enzymes and ligated into the NdeI-BamHI sites of the pET-11a vector (Novagen). The expression plasmids for the G83D and R153A/R157A mutant forms of ttKhpB were constructed by PrimeSTAR mutagenesis procedure (Takara) using the expression plasmid for the wild-type ttKhpB as the template. The primer sets used for introduction of G83D, R153A, and R157A mutations were 5’-TTCATCGACAAGGAGGGGCGGACCCTG-3’ and 5’-CTCCTTGTCGATGAAGCGGCCCAGGTC-3’, 5’-GGGGAGGCGCGGATCGTGCACATGCTCCTCAAG-3’ and 5’-GATCCGCGCCTCCCCGGGGGGCATGGG-3’, and 5’-ATCGTGGCCATGCTCCTCAAGAACCACCCCCGG-3’ and 5’-GAGCATGGCCACGATCCGCCGCTCCCCGGGGGG-3’, respectively. The underlines indicate the codons carrying the mutations. All primers were purchased from Eurofins co. DNA sequencing analyses revealed that the constructions were error-free.
Overexpression and purification of the proteins
The E. coli Rosetta2(DE3) pLysS cells were transformed with the pET-11a/ttkhpB plasmid. One liter of LB medium (Difco) containing 100 μg/ml ampicillin and 30 μg/ml chloramphenicol was inoculated with 1 ml of overnight preculture. After 3 h of cultivation at 37 °C, isopropyl-β-D-thiogalactopyranoside (FUJIFILM Wako) was added to a final concentration of 0.2 mM. The cells were further cultivated at 37 °C for 4 h and harvested by centrifugation. Cells were resuspended with 20 ml of 50 mM Tris-HCl (pH 8.0) (eLANT) containing 100 mM NaCl (nacalai) and stored at –80 °C until use. The cells were thawed in a room temperature water bath to be lysed and treated at 70 °C for 20 min. After the heat treatment, the lysate was immediately chilled on ice for 10 min, and centrifuged at 48,000 × g for 20 min at 4 °C. The supernatant was filtered and loaded onto a TOYOPEARL SuperQ-650 (TOSOH) column (bed volume 20 ml) equilibrated with 50 mM Tris-HCl (pH 8.0). The column was washed with 50 ml of 200 mM NaCl in 50 mM Tris-HCl (pH 8.0), and then the protein was eluted with 100 ml of 400 mM NaCl in 50 mM Tris-HCl (pH 8.0). Ammonium sulfate (nacalai) was added to the protein solution to a final concentration of 1.0 M. The protein solution was loaded onto a TOYOPEARL Phenyl-650M (TOSOH) column (bed volume 20 ml) equilibrated with 50 mM Tris-HCl (pH 8.0) containing 1.0 M ammonium sulfate. The column was washed with 100 ml of 0.4 M ammonium sulfate in 50 mM Tris-HCl (pH 8.0), and then the protein was eluted with 60 ml of 0.2 M ammonium sulfate in 50 mM Tris-HCl (pH 8.0). The protein solution was dialyzed against 50 mM Tris-HCl (pH 8.0). The purity of the recombinant ttKhpB proteins was evaluated by SDS-PAGE analysis with Coomassie Brilliant Blue (nacalai) staining (Supplementary Fig. 14).
Crystallization and structure determination of ttKhpB
To obtain the crystal of the RNA-free ttKhpB, 1 µl of 12.0 mg/ml ttKhpB solution was mixed with an equal volume of the reservoir solution containing 2.0 M sodium formate. To obtain the crystal of the RNA-bound ttKhpB, 1 µl of 12.0 mg/ml ttKhpB solution containing 580 µM single-stranded RNA (5’-CCCCCCCCCC-3’) (BEX co.) was mixed with an equal volume of the reservoir solution containing 100 mM HEPES (pH 7.5) (Hampton Research), 200 mM ammonium sulfate (Hampton Research), and 25% (w/v) polyethylene glycol 3350 (Hampton Research). The drops were equilibrated against 30 μl of the corresponding reservoir at 20 °C using the sitting drop vapor diffusion method. The crystals of the RNA-free and RNA-bound forms were soaked in the reservoir solutions containing 30% glycerol or PEG3350, respectively, before being frozen in liquid nitrogen. The X-ray diffraction data of the crystals were collected at BL38B1 in SPring-8 (Hyogo, Japan) at a wavelength of 1.000 Å at −173 °C. The diffraction data were processed by the HKL2000 program package version 71539. The structures were solved by molecular replacement using PHASER program in the PHENIX suite (version 19.2)40. The model structure of ttKhpB was predicted by AlphaFold2 and used as the search model for molecular replacement. The model was refined using COOT (version 0.7.1)41 and PHENIX programs. The statistics for data collection and refinement are shown in Table 1. Ramachandran plot statistics for the RNA-free and RNA-bound structures of ttKhpB are as follows: For the RNA-free structure, 97.2% of residues are in the most favored regions, 2.8% in additionally allowed regions, and 0% in generously allowed or disallowed regions. For the RNA-bound structure, 97.7% of residues are in the most favored regions, 2.3% in additionally allowed regions, and 0% in generously allowed or disallowed regions. The atomic coordinates and structure factors of the RNA-free and RNA-bound forms have been deposited in the Protein Data Bank with the IDs 9LRG and 9LRI, respectively.
Gel filtration analysis
Gel filtration analysis was performed at room temperature using a Superdex 200 HR 10/30 column (GE Healthcare Biosciences) on an AKTA system (GE Healthcare Biosciences). A 100 µl solution of 10 µM ttKhpB in 50 mM Tris-HCl (pH 8.0) (eLANT) containing 100 mM NaCl (nacalai) was applied to the column. Elution was carried out at a flow rate of 0.1 ml/min with 50 mM Tris-HCl (pH 8.0) containing 100 mM NaCl. The elution profile was monitored by measuring absorbance at 280 nm. The column was calibrated using molecular weight standards: apoferritin (440 kDa), aldolase (160 kDa), conalbumin (75 kDa), ovalbumin (44 kDa), carbonic anhydrase (29 kDa), and ribonuclease A (12.4 kDa).
Electrophoretic mobility shift assay
The 5’-Cy5-labeled single-stranded RNAs of 5-, 6-, 7-, 8-, 9-, and 10-mer lengths (5’-CCCCC-3’, 5’-CCCCCC-3’, 5’-CCCCCCC-3’, 5’-CCCCCCCC-3’, 5’-CCCCCCCCC-3’, and 5’-CCCCCCCCCC-3’ (BEX Co.) were synthesized. The RNAs employed for assessing the RNA sequence specificity of KhpB were synthesized by Fasmac Co. The sequences of these RNAs are listed in Table S3. The 10-mer ssDNA 5’-CATGCCTGAA-3’ was also synthesized by BEX Co. Each RNA or DNA (250 nM) was incubated with varying concentrations of the wild-type or mutant forms of ttKhpB in a reaction buffer containing 50 mH HEPES (pH 7.5) (eLANT), 100 mM NaCl (nacalai), 2.5 mM MgCl2 (nacalai), 1 mM DTT (FUJIFILM Wako), and 0.2 mg/ml bovine serum albumin (nacalai) for 30 min at room temperature. The protein concentrations used are indicated at the top of the figure panels. In the experiments to examine the RNA concentration dependence of ttKhpB’s RNA-binding ability, the ttKhpB concentration was set to 3μM, and the RNA concentration ranged from 0 to 2.2 μM. The reaction mixtures were loaded onto a 10–20% gradient polyacrylamide gel (ATTO) and electrophoresed in EzRun TG buffer (ATTO). RNAs were visualized by using a LuminoGraph I imaging system (ATTO). Signal intensities were quantified using CS analyzer version 3.0 software (ATTO). The %Shifted values were calculated as the percentage of shifted signals relative to the total signals in each lane.
For the binding assays performed by varying protein concentrations, the percentage of RNA shifted was plotted against the monomer concentration of the protein, and the data were fitted to the equation, %Shifted = 100 × [E]0/(Kd + [E]0), where [E]0 is the total molar concentration of the protein, to determine the Kd value. To evaluate the cooperativity of KhpB binding, RNA-binding assays were also performed by varying the RNA concentration. The percentage of RNA shifted was plotted against the total RNA concentration ([S]0), and the data were fitted to the equation, %Shifted = 100 × [S]0n/(Kd + [S]0n), where n is the Hill coefficient.
CD spectrometry
Measurements were performed with a Jasco spectropolarimeter, model J-720W (Jasco), in a solution comprised of 50 mM potassium phosphate (pH 7.0) (nacalai) and 10 μM protein using a 0.1-cm cell at 25 °C. The residue molar ellipticity [θ] was defined as 100θobs/(lc), where θobs is the observed ellipticity, l is the length of the light path in centimeters, and c is the residue molar concentration of the protein.
Difference-distance matrix analysis. To quantitatively assess the conformational differences between the RNA-free and RNA-bound forms of the ttKhpB dimer, we performed a difference-distance matrix analysis34. The Cα–Cα distance matrices were calculated for each subunit, and the absolute differences between equivalent residue pairs were computed. Regions with larger inter-residue distance differences indicate conformational changes or domain movements. Visualization was performed using the matplotlib and seaborn libraries in Python.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data generated or analyzed during this study are included in this published article (and its supplementary information files). The atomic coordinates and structure factor amplitudes of the RNA-free and RNA-bound forms of ttKhpB are publicly available at the Protein Data Bank under the accession numbers 9LRG and 9LRI, respectively. Source data are provided with this paper.
References
Holmqvist, E. et al. Global RNA recognition patterns of post-transcriptional regulators Hfq and CsrA revealed by UV crosslinking in vivo. EMBO J. 35, 991–1011 (2016).
Ding, Y., Davis, B. M. & Waldor, M. K. Hfq is essential for Vibrio cholerae virulence and downregulates sigma expression. Mol. Microbiol. 53, 345–354 (2004).
Melamed, S., Adams, P. P., Zhang, A., Zhang, H. & Storz, G. RNA-RNA interactomes of ProQ and Hfq Reveal Overlapping And Competing Roles. Mol. Cell 77, 411–425 e7 (2020).
Sittka, A. et al. Deep sequencing analysis of small noncoding RNA and mRNA targets of the global post-transcriptional regulator, Hfq. PLoS Genet 4, e1000163 (2008).
Westermann, A.J. et al. The major RNA-binding protein ProQ impacts virulence gene expression in Salmonella enterica Serovar Typhimurium. mBio 10, 1110–1128 (2019).
Dimastrogiovanni, D. et al. Recognition of the small regulatory RNA RydC by the bacterial Hfq protein. Elife 3, e05375 (2014).
Stamsas, G. A. et al. Identification of EloR (Spr1851) as a regulator of cell elongation in Streptococcus pneumoniae. Mol. Microbiol. 105, 954–967 (2017).
Zheng, J. J., Perez, A. J., Tsui, H. T., Massidda, O. & Winkler, M. E. Absence of the KhpA and KhpB (JAG/EloR) RNA-binding proteins suppresses the requirement for PBP2b by overproduction of FtsA in Streptococcus pneumoniae D39. Mol. Microbiol. 106, 793–814 (2017).
Winther, A. R., Kjos, M., Stamsas, G. A., Havarstein, L. S. & Straume, D. Prevention of EloR/KhpA heterodimerization by introduction of site-specific amino acid substitutions renders the essential elongasome protein PBP2b redundant in Streptococcus pneumoniae. Sci. Rep. 9, 3681 (2019).
Lamm-Schmidt, V. et al. Grad-seq identifies KhpB as a global RNA-binding protein in Clostridioides difficile that regulates toxin production. Microlife 2, uqab004 (2021).
Michaux, C., Gerovac, M., Hansen, E. E., Barquist, L. & Vogel, J. Grad-seq analysis of Enterococcus faecalis and Enterococcus faecium provides a global view of RNA and protein complexes in these two opportunistic pathogens. Microlife 4, uqac027 (2023).
Zhu, Y., Ponath, F., Cosi, V. & Vogel, J. A global survey of small RNA interactors identifies KhpA and KhpB as major RNA-binding proteins in Fusobacterium nucleatum. Nucleic Acids Res. 52, 3950–3970 (2024).
Brennan, C. A. & Garrett, W. S. Fusobacterium nucleatum - symbiont, opportunist and oncobacterium. Nat. Rev. Microbiol. 17, 156–166 (2019).
Kostic, A. D. et al. Genomic analysis identifies association of Fusobacterium with colorectal carcinoma. Genome Res. 22, 292–8 (2012).
Parhi, L. et al. Breast cancer colonization by Fusobacterium nucleatum accelerates tumor growth and metastatic progression. Nat. Commun. 11, 3259 (2020).
Han, Y. W. Fusobacterium nucleatum: a commensal-turned pathogen. Curr. Opin. Microbiol. 23, 141–7 (2015).
Olejniczak, M., Jiang, X., Basczok, M. M. & Storz, G. KH domain proteins: another family of bacterial RNA matchmakers?. Mol. Microbiol. 117, 10–19 (2022).
Nicastro, G., Taylor, I. A. & Ramos, A. KH-RNA interactions: back in the groove. Curr. Opin. Struct. Biol. 30, 63–70 (2015).
Grishin, N. V. The R3H motif: a domain that binds single-stranded nucleic acids. Trends Biochem Sci. 23, 329–330 (1998).
Fukita, Y. et al. The human S mu bp-2, a DNA-binding protein specific to the single-stranded guanine-rich sequence related to the immunoglobulin mu chain switch region. J. Biol. Chem. 268, 17463–17470 (1993).
Jaudzems, K. et al. Structural basis for 5’-end-specific recognition of single-stranded DNA by the R3H domain from human Smubp-2. J. Mol. Biol. 424, 42–53 (2012).
Yeoh, Z. C. et al. A minimal complex of KHNYN and zinc-finger antiviral protein binds and degrades single-stranded RNA. Proc. Natl. Acad. Sci. USA 121, e2415048121 (2024).
Korn, S. M., Ulshofer, C. J., Schneider, T. & Schlundt, A. Structures and target RNA preferences of the RNA-binding protein family of IGF2BPs: an overview. Structure 29, 787–803 (2021).
Feracci, M. et al. Structural basis of RNA recognition and dimerization by the STAR proteins T-STAR and Sam68. Nat. Commun. 7, 10355 (2016).
Yokoyama, S. et al. Structural genomics projects in Japan. Nat. Struct. Biol. 7, 943–945 (2000).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Paxman, J. J. & Heras, B. Bioinformatics tools and resources for analyzing protein structures. Methods Mol. Biol. 1549, 209–220 (2017).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Barnes, M. et al. Molecular insights into the coding region determinant-binding protein-RNA interaction through site-directed mutagenesis in the heterogeneous nuclear ribonucleoprotein-K-homology domains. J. Biol. Chem. 290, 625–639 (2015).
Mayo, C. B. & Cole, J. L. Interaction of PKR with single-stranded RNA. Sci. Rep. 7, 3335 (2017).
Tolnay, M., Vereshchagina, L. A. & Tsokos, G. C. Heterogeneous nuclear ribonucleoprotein D0B is a sequence-specific DNA-binding protein. Biochem. J. 338, 417–25 (1999).
Sachs, R., Max, K. E., Heinemann, U. & Balbach, J. RNA single strands bind to a conserved surface of the major cold shock protein in crystals and solution. RNA 18, 65–76 (2012).
Kruger, N. J. & Stingl, K. Two steps away from novelty-principles of bacterial DNA uptake. Mol. Microbiol. 80, 860–867 (2011).
Chen, J. E., Huang, C. C. & Ferrin, T. E. RRDistMaps: a UCSF Chimera tool for viewing and comparing protein distance maps. Bioinformatics 31, 1484–1486 (2015).
Paziewska, A., Wyrwicz, L. S., Bujnicki, J. M., Bomsztyk, K. & Ostrowski, J. Cooperative binding of the hnRNP K three KH domains to mRNA targets. FEBS Lett. 577, 134–40 (2004).
Link, T. M., Valentin-Hansen, P. & Brennan, R. G. Structure of Escherichia coli Hfq bound to polyriboadenylate RNA. Proc. Natl. Acad. Sci. USA 106, 19292–19297 (2009).
Kovach, A. R., Hoff, K. E., Canty, J. T., Orans, J. & Brennan, R. G. Recognition of U-rich RNA by Hfq from the Gram-positive pathogen Listeria monocytogenes. RNA 20, 1548–1559 (2014).
Kim, H. J. et al. Structural basis for recognition of transcriptional terminator structures by ProQ/FinO domain RNA chaperones. Nat. Commun. 13, 7076 (2022).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D. Struct. Biol. 75, 861–877 (2019).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. Biol. Crystallogr. 66, 486–501 (2010).
Thompson, J. D., Higgins, D. G. & Gibson, T. J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994).
Gouet, P., Robert, X. & Courcelle, E. ESPript/ENDscript: extracting and rendering sequence and 3D information from atomic structures of proteins. Nucleic Acids Res. 31, 3320–3323 (2003).
Acknowledgements
The experiments at SPring-8 BL38B1 were carried out with the approval of JASRI (JASRI proposal nos. 2016A2522, 2016B2522, 2017A2544, and 2017B2544). The authors thank beamline scientists for help during data collection. This work was supported by JSPS KAKENHI grant number 24K08718 (K.F).
Author information
Authors and Affiliations
Contributions
K.F. and T.Y. designed the research; K.F., T.M., and S.B. performed the research; S.B. and T.K. developed the methodology; K.F., T.M., S.B., T.K., and T.Y. analyzed data and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Ning Jia, Daniel Straume and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Fukui, K., Murakawa, T., Baba, S. et al. KH–R3H domain cooperation in RNA recognition by the global RNA-binding protein KhpB. Nat Commun 16, 8028 (2025). https://doi.org/10.1038/s41467-025-62302-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-62302-y