Abstract
Engineering genetic circuits to process complex biological signals remains a significant challenge due to non-orthogonal signal responses that limit precise control. In this study, we introduce a framework that integrates orthogonal operational amplifiers (OAs) into standardized biological processes to enable efficient signal decomposition and amplification. By engineering σ/anti-σ pairs, varying ribosome binding site (RBS) strengths, and utilizing both open-loop and closed-loop configurations, we design scalable OAs that enhance the precision, adaptability, and signal-to-noise ratio of genetic circuits. Additionally, we present a prototype whole-cell biosensor capable of detecting transcriptional changes in response to growth conditions, enabling growth-state-responsive induction systems. These systems provide dynamic gene expression control without external inducers, offering significant advantages for metabolic engineering applications. We also apply our framework to mitigate crosstalk in multi-signal systems, ensuring independent control over each signal channel within complex biological networks. Our approach enhances synthetic biology systems by robust signal processing and precise dynamic regulation.
Similar content being viewed by others
Introduction
Cells naturally process external signals through intricate genetic circuits, enabling adaptation to diverse environments1. These circuits rely on complex, non-linear interactions among genes and proteins to regulate cellular behavior, maintain homeostasis, and mediate environmental responses2,3. Synthetic biology harnesses these mechanisms for applications like biosensors4,5,6,7, metabolic engineering8,9, and therapeutics10,11. However, traditional synthetic circuits often use binary (ON/OFF) signals, contrasting sharply with natural systems that adeptly manage complex, non-orthogonal signals—interdependent signals prone to interference2,12,13,14,15,16,17. This non-orthogonality causes crosstalk in engineered systems, complicating precise control and limiting their effectiveness in processing rich biological information18. While sophisticated concepts like neural networks theoretically could manage such complexity, their practical biological implementation with limited components remains a significant hurdle19,20,21.
To advance synthetic biology, robust methods are needed to decompose non-orthogonal biological signals into distinct, orthogonal components17. This would enable precise control over gene expression and cellular behavior, crucial for developing reliable synthetic systems22,23. Principles from analog electronics, especially operational amplifiers (OAs), offer a promising approach24,25,26. OAs are fundamental for signal processing tasks like amplification and subtraction, making them ideal for signal decomposition27. Although prior studies have explored OA-like motifs in biological contexts28, comprehensive investigations into their parametric behaviors, standardized connections for modularity, and particularly their application to multi-dimensional signal decomposition in synthetic biology have been lacking.
This gap is especially evident when addressing key goals like autonomous dynamic control or multi-signal processing. For instance, achieving dynamic regulation of metabolic pathways often relies on traditional strategies like external inducers29, quorum sensing (QS)30,31,32,33, metabolic factor integration9,34,35,36, or growth-stage promoters37,38,39,40,41,42. However, these methods frequently present significant limitations, including challenges with cost, scalability, precision, and robust integration. Similarly, resolving crosstalk in multi-signal networks, such as those found in bacterial QS systems, is a persistent problem. Conventional strategies like directed evolution or promoter engineering to minimize this interference often involve performance trade-offs, typically sacrificing sensor sensitivity for improved specificity. These challenges highlight the need for advanced regulatory solutions that can process complex biological signals with high fidelity.
In this work, we present a comprehensive framework for decomposing multidimensional, non-orthogonal biological signals using rationally designed synthetic Operational Amplifier (OA) devices. Our approach employs a signal processing strategy based on vector operations and matrix multiplication, implemented with orthogonal extracytoplasmic function (ECF) σ factors and their cognate repressors43,44,45. We show that fine-tuning ribosome binding sites and implementing both open-loop and closed-loop configurations (the latter via negative feedback) allows for precise control over linear signal processing, stability, and signal-to-noise ratio. This framework enables the orthogonalization of intertwined biological signals. We first apply this for dynamic control, engineering inducer-free, growth-stage-responsive circuits in Escherichia coli that amplify regulatory signals up to 153/688-fold (Fig. 1a, b). Furthermore, we use the system to resolve three-dimensional signal crosstalk in bacterial quorum sensing systems by implementing an orthogonal signal transformation (OST) matrix (Fig. 1c). In conclusion, this work establishes a versatile framework of synthetic operational amplifier circuits that effectively decompose complex biological signals. This provides a powerful toolkit to mitigate crosstalk and engineer more predictable genetic systems, enabling precise control over the whole-cell response to its environment.
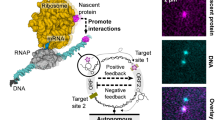
a OA System Overview and Applications. The diagram illustrates the design and application of synthetic operational amplifiers (OAs) for biological signal processing. Each OA module processes two input signals and generates a linear output signal, which is used to control downstream actuators, such as growth regulation or production pathways. The OA circuit performs signal subtraction and amplification to produce precise, orthogonal outputs. The green and red circuits depict two distinct orthogonal outputs used for different biological processes, such as exponential-phase-specific growth and stationary-phase-specific production. b OA Design, Testing, and Optimization. Circuit Design: Each OA circuit consists of an activator (+) and a repressor (-), regulated by specific ribosome binding sites (RBSs) and controlled by IPTG or CA inducers. The outputs are linked to a reporter gene, such as YFP, for signal measurement. Modeling and Optimization: Experimental data are used to fit models, enabling the analysis and optimization of key system parameters, including linear response area, growth pressure, and promoter performance. This iterative process refines OA circuit performance and informs the selection of optimal components for orthogonal signal transformation (OST). Fine-tuning of promoter and RBS combinations further optimizes OA functionality, ensuring compatibility with input signals and enhancing output precision. c Multi-Dimensional Signal Processing via OST Circuits. Input Signal: The input signals, represented as a matrix (X), include various overlapping environmental signals or transcriptional activities. OST Filtering: The OST circuit applies a coefficient matrix to transform input signals into orthogonal components. This involves subtracting and scaling signals using OA modules configured with specific coefficients (A and R values). Output Signal: The resulting orthogonal output matrix (O) features only diagonal elements, ensuring independent, interference-free outputs suitable for precise regulation of multiple biological pathways.
Results
Decomposing multidimensional signals by synthetic orthogonal signal transformation circuits
Gene expression in biological systems is influenced by various environmental signals, such as nutrient availability, temperature, and stress. These signals often result in overlapping expression profiles that complicate their separation. For example, during the exponential and stationary growth phases, multiple factors simultaneously regulate transcriptional activity, leading to non-orthogonal signal overlap46,47,48. To address this challenge, we examined two promoters with overlapping activities during these growth stages (Fig. 2a). These non-orthogonal signals make phase-specific transcriptional control more challenging, highlighting the need for methods to break them down into orthogonal components. This decomposition allows for the precise identification and regulation of distinct biological states, providing a foundation for robust signal processing systems.
a Orthogonal decomposition and linear transformation. The growth dynamics of an exponential-phase-specific promoter (PEXP) and a stationary-phase-specific promoter (PSTA) are shown, with RPU intensity (yellow) and OD600 (gray) plotted over time. The exponential phase (pink) and stationary phase (green) are highlighted. PEXP exhibits high expression during the exponential phase and low expression during the stationary phase, while PSTA shows the opposite trend. Promoters with these characteristics are classified as exponential-phase-specific or stationary-phase-specific promoters, respectively. The data correspond to experimental measurements of PS284 (PEXP) and PS190 (PSTA). Transformation through Exponential-phase OA maps input vectors to the X-axis (signals reflecting exclusive exponential-phase activity), while Stationary-phase OA maps input vectors to the Y-axis (signals reflecting exclusive stationary-phase activity). The curves show the mean value of 6 parallel experimental results. Source data are provided as a Source Data file. b Matrix operation format and transfer function. The OST performs linear transformations using matrix operations. Input vectors are mapped to output vectors according to coefficients \(\alpha\) and \(-\beta\). For exponential-phase-specific signals, the output satisfy \({X}_{o}\gg 0\) and \({Y}_{o}\le 0\); for stationary-phase-specific signals, \({X}_{o}\le 0\) and \({Y}_{o}\gg 0\). This ensures strict condition-dependent outputs, achieving precise signal decomposition. c N-dimensional signal processing with an orthogonal OST. The OST framework extends to \(N\)-dimensional inputs, utilizing, for each input, one activator (A) and \(N-1\) repressors (R). Input signals are transformed by a coefficient matrix to produce an output matrix. When the output matrix is diagonal, the signals are orthogonal, representing independent biological responses specific to each input condition.
To control gene transcription across exponential and stationary growth phases, we designed a genetic circuit capable of distinguishing these states. Promoter activity during the exponential and stationary phases was represented as 2D vectors, with x and y denoting the respective expression levels. This vector representation allowed us to visualize and address the overlap between transcriptional activities in these two phases (Fig. 2a)49. The overlapping signals were then decomposed into orthogonal components through linear transformations involving subtraction and scaling, as illustrated in Fig. 2b.
The decomposition was mathematically implemented by applying a coefficient matrix to the input signals, corresponding to the operations performed by an operational amplifier (OA) circuit (Fig. 3a). Ideally, this transformation produces a diagonal matrix, where only diagonal elements retain expression levels, while all off-diagonal elements are zero. This diagonalization distinctly represents each growth state, ensuring accurate signal separation. In real biological systems, however, negative values from the operation are either replaced with zero or adjusted according to biological constraints. These modifications ensure that the circuit remains functional under practical conditions.
a Illustration of orthogonal decomposition of two transcriptional input vectors under distinct physiological conditions using engineered operational amplifier (OA) filters. To isolate the transcriptional signal specific to condition 1, the Orthogonal-X filter applies a transformation using weights \([{\alpha }_{1},-{\beta }_{1}]\), yielding an output vector aligned along the X-axis and suppressing the Y-axis component (\(Y\le 0\)). Conversely, the Orthogonal-Y filter with weights \([{\alpha }_{2},-{\beta }_{2}]\) extracts the signal corresponding to condition 2 by ensuring the X-axis output is minimized (X ≤ 0), producing a vector along the Y-axis. b Theoretical abstraction of the signal decomposition process and its generalization to N-dimensional transcriptional inputs. The OA computation is reformulated as a matrix operation that transforms a non-orthogonal input matrix into a diagonalized output. This formulation supports both forward calculation (output from known inputs and weights) and inverse computation (deriving the input or weight matrix from a desired output), enabling rational circuit design for signal decomposition in high-dimensional transcriptional spaces.
Operational amplifiers are critical to implementing these linear transformations27. These devices systematically execute operations such as subtraction and scaling, and their design ensures a linear relationship between activation and repression without cooperative interactions. By fine-tuning parameters such as RBS strength, we optimized the OA circuits to reliably perform operations of the form \(\alpha \, \cdot \, {I}_{1} - \beta \, \cdot \, {I}_{2}\). These optimizations allowed us to achieve robust decomposition of overlapping signals.
Expanding this framework, we demonstrated its scalability to multidimensional signal inputs. As shown in Fig. 2c, the system processed \(N\)-dimensional inputs, each comprising \(N\) distinct conditional values, and decomposed them into orthogonal components. This required \(N\) orthogonal regulatory pairs that functioned independently and avoided interference with the host cell’s native regulatory systems. Theoretically, \(N\) is limited by available, biologically compatible, and truly orthogonal regulatory pairs. Practically, increasing \(N\) is constrained by cumulative host cell effects like metabolic burden. Importantly, to ensure compatibility within the biological framework, both input and output signals must adopt the same biochemical format—in this case, normalized transcription signals.
We further validated the versatility of the system by constructing orthogonal signal transformation (OST) circuits to handle complex, multidimensional inputs. Using matrix operation principles outlined in Fig. 3b, we demonstrated bidirectional functionality: the coefficient matrix could be used to design OA circuits for specific target outputs, and known OA circuit parameters could reconstruct input signal matrices. These capabilities underscore the adaptability of OST circuits as tools for signal decomposition and synthesis, enabling the design of robust genetic systems capable of processing complex biological signals.
Synthetic OA circuits construction and performance evaluation
To implement orthogonal signal decomposition, we constructed synthetic operational amplifier (OA) circuits designed to separate overlapping transcriptional inputs into linear components. The OA circuit performs the operation \(\alpha \, \cdot \, {X}_{1}-\beta \, \cdot \, {X}_{2}\), where \({X}_{1}\) and \({X}_{2}\) are input transcription signals that regulate the production of the activator (A) and repressor (R), respectively. This design ensures a linear activation-repression relationship, avoiding cooperative interactions, as shown in Fig. 4a. Both inputs and outputs are normalized transcription signals, providing compatibility with the host’s biochemical environment.
a–c Open-loop amplifier circuit. a The open-loop circuit design. Left: The genetic circuit construction. Right: The corresponding mathematical model. Input signals \(({X}_{1},{X}_{2})\) regulate the production of activator (A) and repressor (R), respectively, each driven by specific RBSs. The effective activator concentration (\({X}_{E}\)) is computed within the OST device (gray box) and subsequently activates the output promoter (PA) to produce the final output (\(O\)). b Experimental validation of the open-loop circuit. The data points represent the measured values of \({X}_{1}\) and \({X}_{2}\), which are used to calculate effective activator concentration \({X}_{E}=3.06{X}_{1}-1.51{X}_{2}\), with the corresponding output \(O\) plotted as a function of \({X}_{E}\). The input-output relationship is fitted to the Hill function (solid black line). The first-order Taylor expansion of the Hill function (blue dashed line) approximates the linear region. The effective bandwidth (<3 dB) is indicated by gray dashed lines, confirming that the circuit operates linearly within \({X}_{E}\) \(\in ({\mathrm{0,1.2}})\). c SNR heatmap for the open-loop circuit. Input signals: \({{input}}_{1}={P}_{S284}\), \({{input}}_{2}={P}_{S190}\). The non-functional area \({O}_{{target}}\ge 0 \, {{\mbox{ and }}} \, {O}_{{off}-{target}}\le 0\) is shown in white. Orange-shaded regions indicate suboptimal bandwidth (>3 dB). The red circle marks experimentally derived circuit parameters (3.06, 1.51). d–f Closed-loop amplifier circuit. d Closed-loop circuit design. An additional repressor regulated by PECF22M1 modifies the effective activator concentration to \({X}_{E}^{{\prime} }={X}_{E}-K^{\prime} \cdot \beta {\cdot X}_{A}\). Left: schematic circuit design. Right: mathematical representation. e Experimental validation of the closed-loop circuit. Black data points represent measurements, fitted to the Hill function (solid black line) and first-order Taylor approximation (blue dashed line). The effective bandwidth expanded to \({X}_{E}\) \(\in ({\mathrm{0,36.6}})\). f SNR heatmap for the closed-loop circuit. The functional area is significantly expanded compared to the open-loop configuration, demonstrating enhanced fidelity and robustness. The red circle indicates experimentally derived circuit parameters (3.06, 1.51). For data presented in (b, e), experiments were independently repeated on three separate days. Source data are provided as a Source Data file.
The circuit was constructed using ECF σ factors and their cognate anti-σ factors43, as well as T7 RNA polymerase (T7 RNAP) and its inhibitor, T7 lysozyme50,51. These pairs were selected for their well-characterized linear interactions and orthogonality. Input \({X}_{1}\) regulates the production of activator \(A\) via RBSs with a translation rate \({r}_{1}\) and degradation rate \({\gamma }_{1}\), resulting in the following activator concentration:
Similarly, \({X}_{2}\) regulates \(R\) production, yielding:
The effective activator concentration (\({X}_{E}\)) is computed as:
Notably, this formula is universally applied in this study to compute \({X}_{E}\) for all circuit configurations and analyses. The circuit’s output follows an equation with a Hill coefficient of 1:
Here, \({K}_{2}\) is the activator binding constant, and \({O}_{\min }\approx 0\) was neglected during the following calculation. This relationship indicates that while \({X}_{E}\) is a linear combination of inputs \({X}_{1}\) and \({X}_{2}\), the output remains linear only when \({X}_{E}\ll {K}_{2}\). Outside this range, the output deviates from linearity. The linear range of the output is positively correlated with the binding coefficient \({K}_{2}\), which determines the extent of the linear regime. Additionally, the OA circuit’s gain \(({O}_{\max })\) is determined by the activator’s binding strength to the output promoter. The operational range of the circuit is characterized by the standard −3dB bandwidth, which corresponds to the frequency range where the output signal is reduced to half of its maximum value, typically representing the effective bandwidth within which the system operates with minimal error52 (Fig. 4e).
To validate the design, we constructed an open-loop circuit OAO(E22.14.21.PECF22) and tested under various IPTG and CA gradient conditions, which correspond to induction strengths for input+ and input- respectively (Supplementary Fig. 1). The experimental data were fitted to the theoretical model, with a correlation coefficient (\({R}^{2}=0.887\)) indicating that the model effectively captures the linear relationship between the two inputs (Fig. 4b, Supplementary Fig. 2). However, the open-loop configuration exhibited limitations, particularly a restricted linear range and increased sensitivity to noise, which could affect its performance under more complex conditions.
To overcome these limitations, we introduced a negative feedback loop to create a closed-loop configuration, inspired by the open- and closed-loop designs of electronic OAs (Fig. 4d)27,53. A similar design strategy for constructing inverse functions was demonstrated, which constructed inverse functions with negative feedback in signal processing54. In our system, the additional repressor (R), regulated by an internal feedback promoter (PECF22M1, a mutant of PECF22) modulated the effective activator concentration45:
where \({X}_{A}\) represents the concentration of the additional repressor introduced by the feedback loop, and \(K^{\prime}\) accounts for promoter-specific binding coefficients (see Supplementary Note 2&3, Supplementary Table 6). Note that the two input promoters responding to the σ factor—PA and PA′—are different, as they correspond to distinct regulatory inputs. However, the RBSs which drive anti-σ expression, are the same. This design ensures that a single β parameter can be used for both feedback branches. If different RBSs are used, another β′ would need to be introduced, increasing modeling complexity. This feedback mechanism significantly stabilized the output and extended the linear operating range, resulting in improved fidelity and robustness across a wider range of input conditions compared to the open-loop system.
We further analyzed the signal-to-noise ratio (SNR) for both configurations. SNR, defined as:
illustrates the circuit’s ability to distinguish desired signals from noise and reliably process inputs. Here, \({O}_{{target}}\) denotes the output component aligned with the intended signal direction, while \({O}_{{off}-{target}}\) captures leakage along the orthogonal direction. For example, in a two-dimensional OA circuit, if the goal is to amplify signal \({X}_{1}\) while suppressing \({X}_{2}\), the output corresponding to \({X}_{1}\) is considered \({O}_{{target}}\), and that from \({X}_{2}\) is \({O}_{{off}-{target}}\). The open-loop configuration (Fig. 4c) demonstrated a constrained functional area, with areas of suboptimal performance under certain input conditions. By contrast, the closed-loop circuit (Fig. 4f) significantly expanded the functional area, achieving higher fidelity and robustness under dynamic input conditions.
These results demonstrate that the synthetic OA circuit can achieve orthogonal signal processing with high precision. While the open-loop configuration provides accurate decomposition within a defined range, the closed-loop design extends the operational range and enhances noise tolerance. Both the open-loop and closed-loop designs demonstrated strong performance. To compare their application-level performance, we constructed a closed-loop version of STA-ON2 by introducing a negative feedback loop (Fig. 5e, Supplementary Figs. 12, 13). As shown in Supplementary Fig. 10, although the closed-loop activation starts slightly later, the final expression output is comparable to the open-loop design. In scenarios where input signals fall outside the linear range of open-loop OAs, the closed-loop configuration offers a broader linear regime and improved reliability. However, for inputs within the linear range, open-loop OAs remain preferable due to their simplicity and minimal resource demand. Together, these designs offer versatile tools for processing dynamic and complex biological signals. To further demonstrate generalizability, we constructed and validated additional OA circuits with varying parameters and architectures. Experimental results are provided in Supplementary Note 5, with design parameters summarized in Supplementary Table 1-2. Detailed theoretical analyses and experimental data are provided in Supplementary Notes 1–3.
a Analysis of native and synthetic promoters. Left: Volcano plot of native E. coli RNA-seq data. Promoters with SE ratio >1.5 (orange) or <0.67 (blue) were classified as stationary- or exponential-phase-specific, respectively. Selected promoters are labeled with their locus tag IDs. Differential expression was calculated using PyDESeq2 with a two-sided Wald test. Right: Sequence logos of stationary-phase (top) and exponential-phase (bottom) promoters highlight distinguishing motifs. b Promoter scoring for OA circuits. For each OA configuration, all possible promoter pairs were systematically evaluated as inputs to identify the pair with the highest OA score, representing the optimal performance achievable by that OA. Example: the best-performing combination for OAO(E11.2.2.PECF11) was NSP4 (a native stationary-phase promoter, A) and NEP7 (a native exponential-phase promoter, R). c Synthetic vs. native promoter performance. OA scores were simulated for all combinations of native and synthetic promoters using a fitted model. Each data point represents one promoter pair (n = 36 per group). The embedded box plots show the median (center line), the 25th and 75th percentiles (box bounds), and the minimum and maximum values excluding outliers (whiskers). d Fine-tuning of OA performance via RBS modification. The orange star represents the initial configuration of PS459 and PS394 with RBSs BCD2 and BCD2, which fell into a non-functional region (target output <0). By modifying the RBSs to BCD7 (activator) and BCD18 (repressor), shifting the OA configuration (\(\alpha /\beta\) ratio) into the functional area (green star). e Functional circuits for EXP-ONs and STA-ONs. Input: Promoter activity under exponential (\(X\)) and stationary (\(Y\)) phases. Coefficient: OA parameters for signal processing. Prediction vs. Measurement: Modeled vs. experimental outputs (in RPU). STA-ON1 and STA-ON2 were constructed in both open- and closed-loop forms using configurations. All EXP-ON reporters are tagged by ssrA to capture the real time expression84. f Applications in continuous culture systems. Fold changes were calculated using a minimum expression value of 0.01. The expression levels were measured in three independent biological replicates (n = 3). Source data are provided as a Source Data file.
Synthetic growth-responsive device
As discussed earlier, genetic devices can be designed to generate clean expression patterns under either exponential or stationary growth conditions. The final output of such systems depends on two critical components: the input matrix (promoters or sensors) and the operational matrix (OA circuits). While the influence of OA circuit parameters on performance has been previously analyzed, the selection of input promoters is equally critical, as it directly impacts the system’s specificity and fidelity. Clear phase-specific promoter activities, with minimal overlap between exponential and stationary phase expression, are essential for optimal circuit function, as demonstrated by the importance of promoter signal separation on OA performance (Supplementary Fig. 4).
In E. coli, transcription across growth phases, regulated by σ factors like σ70 and σ38, is inherently non-orthogonal46,47,48,55,56,57. To identify growth-responsive promoters in E. coli, we used RNA sequencing data to quantify promoter activity during exponential and stationary growth phases (Fig. 5a). Promoters were classified as stationary- or exponential-phase-specific based on a defined threshold (\(n\), default 1.5). Stationary-phase promoters were identified when the ratio of stationary-phase to exponential-phase transcription strength (SE ratio) exceeded \(n\), while exponential-phase promoters were defined when the SE ratio was less than \(1/n\). Transcription strength was calculated from RNA-seq data as the normalized read depth around the transcription start site (TSS) (Methods).
We further analyzed the sequence motifs of these phase-specific promoters from native RNAseq and identified distinct features consistent with prior studies46,48,58,59 (Supplementary Fig. 5–8): stationary-phase promoters favored AT-rich discriminator regions, cytosine enrichment at –13 nt and –36 nt, and a “TATACT” –10 motif; exponential-phase promoters displayed GC-rich discriminators and the canonical “TATAAT”. To validate these features, we selected a subset of promoters from Fig. 5a (highlighted with green circles) that showed large SE ratio differences and matched the identified motif patterns. Their experimentally measured phase-specific expression strengths are shown in Supplementary Fig. 9 and Supplementary Table 5, confirming their predicted behavior.
Based on these results, we engineered a synthetic promoter library incorporating these phase-specific elements. Experimental tests (Supplementary Fig. 10) confirmed that synthetic promoters achieved significantly higher SE ratios and broader angular separation between input vectors than native promoters. To further quantify the performance ceiling of each OA configuration, we systematically screened all possible native promoter pairs as inputs to each OA, computing the best achievable OA score per combination (Fig. 5b). We repeated this analysis for synthetic promoters, allowing direct comparison between the native and synthetic promoter performance across OAs. This comparison, shown in Fig. 5c, demonstrated that synthetic inputs consistently outperformed native ones.
Functional OAs were then constructed for applications requiring phase-specific outputs, such as exponential-phase-ON (EXP-ON) and stationary-phase-ON (STA-ON) circuits (Fig. 5e, Supplementary Fig. 11, Supplementary Table 9). Measured outputs closely matched theoretical predictions. The framework used for input-output matching is detailed in Supplementary Note 4 (OD predictions in Supplementary Fig. 3). Some OA configurations were initially non-functional due to suboptimal α/β ratios (e.g., Fig. 5d, orange star). To resolve this, we tuned RBS strengths via bicistronic designs60, a robust strategy for improving translation predictability and minimizing RBS–CDS context dependence. Axes in Fig. 5d are labeled “RBS +” and “RBS −” to represent activator/repressor translation strength changes. This figure illustrates how tuning RBSs (e.g., BCD2 → BCD7, BCD2 → BCD18, RBS expression data in Supplementary Table 8) shifted the circuit into the functional zone (green star) without altering the underlying promoter or coding sequence.
Finally, we evaluated the system’s robustness in continuous culture experiments. Using identical promoters as inputs but processed through distinct OAs (e.g., OAO(E38.7.18.PECF38) and OAO(E22.7.18.PECF22)), we achieved divergent expression outputs tailored to specific applications. Promoter pairs such as PS284 and PS190 generated dynamic expression patterns, achieving fold-change amplification from 2–4 fold in native systems to 153/688-fold with synthetic circuits (Fig. 5f). This improvement highlights the system’s ability to amplify signals and achieve distinct phase-specific outputs, consistent with the signal decomposition framework in Fig. 2 and Fig. 3a, where overlapping signals are separated into orthogonal components for precise processing.
Applications of functional amplifiers for dynamic metabolic control
Dynamic control systems are crucial in metabolic engineering for optimizing chassis cells by decoupling their growth and production functions61,62,63. However, conventional approaches to achieve such dynamic regulation often face significant limitations, including reliance on costly external inducers, issues with scalability and dynamic range, challenges in integrating with cellular metabolic states, or a lack of robustness and precision, all of which can restrict their broader application.
Our OA circuits, based on OST, address these challenges by enabling precise, growth-phase-specific regulation without relying on external inducers. This makes OA circuits a powerful tool for dynamic control, as demonstrated in two key applications: heterologous protein expression in E. coli BL21 and dynamic regulation of the shikimate acid (SA) pathway in E. coli MG1655. These two applications were chosen to illustrate the versatility of OA circuits in enabling precise dynamic ON/OFF control: EXP-ON for activating gene expression during the exponential phase and STA-ON for switching it on during the stationary phase.
For protein expression, E. coli BL21 is a widely used strain due to its capacity for expressing toxic or high-level proteins29,64. However, unregulated protein expression imposes metabolic stress, inhibiting growth and often resulting in expression failure without external inducers64. Conventional systems resolve this by using inducers like IPTG to delay expression until cells reach sufficient density, but this increases costs and complicates scalability65. To address these challenges, we planned to use a STA-ON circuit for dynamic regulation. As illustrated in Fig. 6a, the OA-based controller plays a key role by suppressing expression during the exponential phase—when cells are most sensitive to stress—and enabling high-level expression in the stationary phase. This strategy minimizes toxicity while maximizing protein output. Notably, the same promoters exhibit different expression levels in BL21 compared to MG1655 with medium from M9-glucose to LB, and the parameters of the OA circuit also require recalibration66. To address this, we performed induction tests for OAO(E22.18.21.PECF22) and OAO(E22.7.21.PECF22) in BL21 to determine the corresponding OA coefficients and constructed a new STA-ON5 application based on these parameters. Experimental comparisons (Fig. 6b, Supplementary Fig. 14) confirmed significant differences in performance between the same construct in MG1655 and BL21. Both open-loop and closed-loop designs maintained functionality across hosts. Notably, a few circuits displayed reversed phase preference, as detailed in Supplementary Fig. 14. STA-ON2 in BL21 suppressed protein expression during the exponential phase to minimize stress and activated expression in the stationary phase for efficient production. As shown in Fig. 6d, both open-loop (OL) and closed-loop (CL) configurations of STA-ON2 successfully expressed LacI and T7RNAP protein under growth-phase-specific control. SDS-PAGE analysis confirmed that STA-ON2 OL and CL circuits achieved expression levels comparable to aTc-inducible systems when aTc was added at OD600 ~ 0.5, thereby avoiding early growth stress in BL21. In contrast, no expression was observed in wild-type BL21 or in the inducible system when aTc was added at the start of cultivation (t = 0), which caused significant growth stress. The growth profiles in Fig. 6c further demonstrate this effect: when aTc was added at the start of cultivation (t = 0), the induced system showed severely inhibited growth, consistent with toxic burden from premature expression. In contrast, both the STA-ON2 OL and CL circuit and late aTc induction (at OD ~ 0.5) maintained growth curves comparable to the BL21 wild-type control, confirming the effectiveness of OA-based dynamic regulation. These results highlight STA-ON5’s ability to reduce growth stress while enabling cost-effective and scalable protein expression without inducers. To further demonstrate the generalizability of the STA-ON design, we employed ECF22-, ECF11-, and ECF38-based circuits to express four distinct output proteins, including LacI, T7RNAP, lysozyme, and sfGFP (Supplementary Fig. 16). Among them, sfGFP exhibited strong expression even under early induction (aTc-ON(OD ~ 0)), likely due to its minimal cytotoxicity, as supported by the corresponding growth profiles in Supplementary Fig. 15. While lysozyme expression of STA-ON2 appeared relatively weaker in SDS-PAGE, this is consistent with its lower quantified expression levels (Fig. 5e, Fig. 6b), as shown in Supplementary Table 10.
a Protein expression in BL21. Sensor: stationary-phase promoter as input1, exponential-phase promoter as input2. Controller: OAO(E22.7.21.PECF22). Actuator: protein expression module. The STA-ON circuit alternates between exponential-phase OFF and stationary-phase ON states, suppressing protein expression during the exponential phase to avoid growth inhibition and activating production during the stationary phase. b Cross-strain performance comparison of STA-ON circuits. STA-ON circuits were evaluated in two E. coli MG1655 and BL21 under M9-glucose and LB media. c Growth curve analysis of STA-ON device in BL21. The STA-ON device demonstrates growth comparable to BL21wt and the induced system. Samples with circuits induced by aTc (t = 0) exhibited significant growth inhibition, likely due to metabolic stress. Data are presented as mean values ± standard deviation (SD) from three independent biological replicates. d SDS-PAGE analysis of His-tagged protein expression. The STA-ON2 circuit was used to express His-tagged LacI (38 kDa) and T7RNAP (98 kDa) in both open-loop (OL) and closed-loop (CL) configurations with expression comparable to the induced systems, confirming effective stationary-phase-specific activation without external inducers. Experiment was independently repeated three times with similar results. Full uncropped scans are available in the Source Data file. e Dynamic regulation of SA production using the EXP-ON circuit. Sensor: exponential-phase promoter as input1, stationary-phase promoter as input2. Controller: OAO(E22.14.21.PECF22). Actuator: protein expression module. Implemented in an E. coli \(\triangle\)aroK \(\triangle\)aroL strain. During the exponential phase, the circuit expresses AroK & AroL to support growth. In the stationary phase, it switches OFF, allowing SA accumulation. f Growth and SA production dynamics. OD600 (green) increases during the growth phase and plateaus in the production phase, while SA production (black) rises sharply, reflecting efficient metabolic flux redirection. Error bars represent the standard deviation of three biological replicates. g Temporal dynamics of EXP-ON1 circuit activity and growth. The yellow line represents EXP-ON1 circuit activity, with high expression during the growth phase and a rapid OFF switch in the production phase, redirecting metabolic flux toward SA accumulation. Source data are provided as a Source Data file.
The second application focuses on the dynamic regulation of the shikimate pathway in E. coli MG1655 for shikimate acid (SA) production30. In wild-type MG1655, SA is rapidly metabolized into aromatic amino acids via the aroK & aroL genes, preventing its accumulation. While inducible systems can redirect flux toward SA accumulation, they are limited by high costs, resource consumption, and poor scalability30. We used the EXP-ON1 circuit, to dynamically regulate aroK & aroL, activating these genes during the exponential phase to support growth and switching them off in the stationary phase to halt the pathway, enabling SA accumulation. In an aroK & aroL knockout strain, EXP-ON1 effectively controlled SA production dynamics. During exponential growth, the circuit supported cell proliferation, while in the stationary phase, it redirected metabolic flux to achieve high SA accumulation (Fig. 6e). System performance was validated by OD600 and SA quantification measurements (Fig. 6f, Supplementary Fig. 17), showing steady cell growth during the exponential phase and sharp increases in SA levels during the production phase. Temporal regulation of AroK&AroL expression (Fig. 6g) demonstrated a precise ON/OFF switch, achieving SA yields comparable to those reported for other systems, without requiring costly inducers.
These applications highlight the flexibility and scalability of OA circuits for dynamic metabolic control. By addressing the limitations of traditional strategies, OA circuits enable cost-effective, inducer-free regulation of complex metabolic processes and high-level protein expression, offering significant advantages for biotechnological applications.
Three-dimensional signaling processing with external input matrix
Building upon the established challenge of crosstalk in multi-signal systems like quorum sensing (QS), we specifically investigated the LuxR (3-OC6HSL), LasR (3-OC12HSL), and CinR (3-OHC14HSL) QS pathways67,68. These systems are frequently used in synthetic biology but are well-documented for exhibiting severe crosstalk and overlapping transcriptional responses17. Ideally, their cognate promoters—Plux, Plas, and Pcin—should operate orthogonally, responding exclusively to their respective N-acyl-homoserine lactone (AHL) signals69,70,71,72. However, as illustrated in experimental characterizations (e.g., Fig. 7a), the natural encoding mechanisms within these systems lead to significant signal overlap, resulting in complex, non-orthogonal outputs72,73.
a Resolving crosstalk through signal decomposition and decoding in the QS system. The input signal matrix shows promoter responses to 3OC6-HSL, 3OC12-HSL, and 3OC14-HSL, with crosstalk arising from natural encoding by the QS system (QS matrix). Using the ideal orthogonal output matrix and Fig. 3b matrix operations, a decoding matrix was derived to resolve crosstalk. To address unattainable \(\alpha /\beta\) ratios, the decoding matrix was decomposed into a promoter scaling matrix and an OA coefficient matrix via the Hadamard product. b Design of the three-dimensional OST circuit. Based on Fig. 7a calculations, ECF22 is linked to Plux with S1-weighted Plas driving anti-ECF22, ECF11 to Plas with \({S}_{3}\)-weighted Pcin driving anti-ECF11, and ECF38 to Pcin with \({S}_{2}\)-weighted Plas driving anti-ECF38. Effective outputs of ECF22, ECF11, and ECF38 drive YFP, CFP, and mCherry, respectively. c Input, predicted, and experimental output signal matrices. Heatmaps compare input signals with crosstalk, predicted orthogonal outputs, and experimental results. Red boxes in the input heatmap highlight crosstalk regions—weak signals where strong responses are expected or unintended activation. Predicted and experimental results align closely, confirming that the decoding matrix resolved crosstalk and ensured intended responses. d Comparison of predicted and measured results. The scatter plot shows predicted values on the x-axis and experimental results on the y-axis, demonstrating a strong correlation (R² = 0.855). This confirms the OA circuit’s ability to resolve crosstalk and restore orthogonal signal patterns. The expression levels were measured in three independent biological replicates (n = 3). Source data are provided as a Source Data file.
To address this specific instance of three-dimensional signal interference without altering the native QS components, we applied our three-dimensional Orthogonal Signal Transformation (3D-OST) framework. Our approach focused on computationally decoding the mixed transcriptional responses from these interacting pathways to reconstruct clear, orthogonal output signals.
Based on Fig. 3b, the decoder matrix can be calculated by relating the desired output matrix (orthogonal signals) to the input matrix (AHL signal combinations) through the inverse of the encoding matrix, which represents natural signal transmission in the QS system. However, direct implementation of this decoder using our OA library was not feasible, as none of the existing α/β combinations satisfied the full decoder parameters.
To resolve this, we decomposed the decoder matrix into two components: an OA coefficient matrix and a promoter scaling matrix, which were then combined via the Hadamard (element-wise) product. This approach allowed us to retain the functional logic of OA computation while adjusting signal amplitude at the input level. Promoter scaling was achieved by modifying the -10 and -35 core motifs of each promoter without altering their regulatory logic or signal specificity. This strategy follows the motif substitution method6, which enables strength tuning via defined motif combinations (e.g., a–g, A–H). All variants used in this study—such as PlasO-cC, PlasO-bE, and PcinO-bC—share the same binding architecture and differ only in motif region (Supplementary Table 11).
Using these scaled promoters and feasible OA coefficients, we implemented the full decoder matrix and constructed the 3D OST system (Fig. 7b). The results demonstrate that the OST-based decoder effectively mitigates crosstalk, restoring orthogonality in QS systems and achieving strong alignment between predicted and experimental outputs (Fig. 7c, d). These findings validate the ability of the OST framework to decode and separate cross-interacting signals with high precision. By extending its scalability from 2D to 3D systems, the OST framework provides a robust and versatile solution for resolving crosstalk in complex signaling networks, enabling the engineering of orthogonal multi-signal genetic circuits.
Discussion
Logic circuits have achieved remarkable success in processing electronic signals, evolving into sophisticated systems capable of managing diverse tasks. In contrast, genetic circuits in biological systems face inherent limitations due to the small number of molecules involved and slower reaction kinetics, which constrain scalability and operational efficiency15. Analog circuits are essential for addressing various signaling challenges and are foundational in numerous applications26,74. Among these, operational amplifiers (OAs) stand out as critical devices, providing key capabilities for linear signal processing and serving as fundamental components in advanced operations such as integration and square root calculations27,53. While preliminary designs of OAs have been developed for integral control systems28, comprehensive research on their parameter behaviors and the establishment of standard connections for signal processing remains scarce. Our research addresses these gaps by presenting the first design and demonstration of OAs in a standard connectable format, ensuring compatibility between input and output signals as RPUs. This standardized format facilitates the integration of OAs into automated design platforms, potentially paving the way for the development of more complex analog and analog-digital hybrid circuits for synthetic biology23,75.
We developed a standardized framework for engineering OAs, integrating σ/anti-σ pairs, optimizing ribosome binding site (RBS) strengths, and utilizing both open-loop and closed-loop configurations. Open-loop OAs effectively handled discrete signal decomposition and amplification, while closed-loop configurations extended linearity, enabling continuous signal processing. These designs ensure compatibility between input and output signals, facilitating seamless integration into synthetic circuits. Our approach was validated through theoretical analyses and experimental results, demonstrating improved signal-to-noise ratios (SNRs) and operational efficiency. By aligning OA parameters with matrix operations, we further automated the design and screening process, enabling efficient circuit optimization for diverse applications.
Building on our signal processing framework and standardized OAs, we developed a prototype universal whole-cell biosensor capable of detecting transcriptional changes associated with different cell growth stages. This biosensor demonstrated high specificity and significant signal amplification by processing transcriptional changes through orthogonal filters with growth-phase-responsive signals. The dynamic range was amplified from 2/10-fold to 153/688-fold across various growth stages, matching or surpassing the performance of traditional specific biosensors6,69. When integrated into growth-state-responsive induction systems, it enabled dynamic gene expression control without the need for external inducers, offering a scalable and cost-effective solution for metabolic engineering. Additionally, this biosensor design is compatible with other whole-cell sensor blueprints that detect diverse signals, facilitating the detection of inputs that lack dedicated sensing proteins76.
A persistent challenge in synthetic biology is crosstalk in multi-signal systems, where overlapping pathways compromise signal fidelity and hinder precise control23. To address this, we developed a three-dimensional framework for signal decomposition and orthogonalization, enabling accurate and independent regulation of each signal channel in complex networks. This approach significantly reduces interference between signals, ensuring reliable processing of overlapping biological inputs. By leveraging native sensors, our system mitigates crosstalk while maintaining compatibility with existing biological components, contributing to the development of more robust and efficient multi-signal networks. Recent efforts have explored neural network-based strategies for processing complex signals19,20,21; however, our framework offers distinct advantages. First, the internal steps can be quantitatively controlled using well-characterized transcriptional or translational parts. Second, it has the potential to integrate with multi-layer neural networks, facilitating linearized weighting operations and expanding its capacity for advanced signal processing. Our synthetic promoter design strategy, while effective in E. coli, is currently based on transcriptional features such as SE ratio, -10, extended -10 and -35 motifs, and discriminator regions. Although these elements are well-supported in E. coli, their transferability to other organisms remains limited. To address this and expand the applicability of our OA framework, future work will explore more generalized promoter design strategies. In particular, leveraging large-scale transcriptomic datasets and integrating language models or transformer-based architectures may enable species-specific promoter optimization in a data-driven and automated manner. This would improve scalability, reduce manual intervention, and facilitate application across a wider range of microbial hosts.
Despite its advancements, the OA framework relies on specific design constraints to ensure functionality. A precise 1:1 ratio between activators and inhibitors and adherence to a Hill coefficient of 1 are critical for maintaining balanced operation and signal linearity. Additionally, while our σ/anti-σ system offers multiple orthogonal pairs, transcriptional toxicity and variability in translation strength across different chassis cells present challenges that require further optimization. Addressing these issues is necessary to expand the applicability of OAs across various species and environments. Applying this \(N\)-dimensional OA framework to higher \(N\) values increases genetic part numbers, impacting construct complexity and cellular burden. The genetic encoding presents a key design trade-off: systems with up to N2 operons offer fine-grained coefficient control but are genetically intensive. Alternatively, strategies like tandem-promoter designs could reduce operon requirements to around 2\(N\), enhancing genetic economy, though potential issues like transcriptional roadblocking would then need careful management in future developments of complex biological signal processors13,77.
Looking forward, the potential for applying single-molecule activation and inhibition systems holds promise for advancing computational speed and reducing toxicity. These systems could expand the OA framework to protein-based computation20,73, RNA-based computation10,78, or Cas/anti-Cas systems79, enhancing versatility and scalability. Furthermore, integrating this framework into eukaryotic systems could provide a platform for dynamic regulation and analog-digital hybrid circuits in higher organisms, making it a valuable tool for synthetic biology and biotechnological innovations.
In conclusion, our study provides a robust and scalable solution for processing complex biological signals using orthogonal operational amplifiers. By integrating standard linear transformations, matrix operations, and quantitative component design, this framework enables precise signal decomposition and reliable integration with downstream genetic functions. Importantly, this approach operates without the need for external inducers, relying on the differential changes in cellular responses to external signal for dynamic regulation, which enhances its cost-effectiveness and scalability. While this approach may not be the simplest for specific signal operations, its versatility and adaptability establish a foundation for advanced analog-digital hybrid circuits, fostering innovation in next-generation synthetic biology applications.
Methods
Strain, media and reagents
E. coli DH10B or DH10B Marionette strain (containing the Marionette sensor array71) was used for cloning. Plasmid-based promoter test systems were characterized in E. coli MG1655 and E. coli BL21. The OA induction circuit was evaluated in both the E. coli MG1655 Marionette strain (containing the Marionette sensor array71) and the E. coli BL21 Marionette strain. The BL21 Marionette strain was additionally utilized for protein expression. The shikimate fermentation strain MGC2 (E. coli strain MG1655 ΔaroK ΔaroL) was derived from the wild-type MG1655 using the λ Red recombination system80,81 (Supplementary Fig. 21). MG790 (MG1655 attB:: cinR luxR lasR) (Supplementary Fig. 20) was utilized as a testing strain for evaluating baseline promoters and functional circuits in crosstalk genetic networks.
Cloning was performed using a hierarchical Golden Gate Assembly82 (GGA) strategy with type IIS enzymes (BsaI, BpiI) (Supplementary Fig. 18). Recognition sites were removed by synonymous codon substitution. Genetic parts (promoters, ORFs, terminators) were assembled into plasmids with distinct selection markers and transformed into cloning strain.
Luria-Bertani (LB) medium contained 1% tryptone, 0.5% yeast extract, and 1% NaCl. M9(CA)-glu medium was prepared using 5× M9 Salt Solution (3.39% Na2HPO4, 1.5% KH2PO4, 0.25% NaCl, 0.5% NH4Cl), 10 mg/mL thiamine, 2 mM MgSO4, 0.2% casein hydrolysate (CA), 0.1 mM CaCl2, 0.4% glucose used as the sole carbon source, and deionized H2O to a final volume of 1 L.
Appropriate antibiotics were used at the following concentrations for cell selection and maintenance: 100 μg/mL ampicillin, 50 μg/mL kanamycin, 25 μg/mL chloramphenicol, 50 μg/mL spectinomycin.
Induction was achieved using the following chemicals: cuminic acid (Ca, sigma, 268402); isopropyl-β-D-thiogalactopyranoside (IPTG, Macklin, I811719); anhydrotetracycline (aTc, GLPBIO, GC16227); N-(β-Ketocaproyl)-L-homoserine lactone(3-O-C6-HSL, sigma-aldrich, K3007); N-(3-Oxododecanoyl)-L-homoserine lactone(3-O-C12-HSL, sigma-aldrich, O9139); N-(3-Hydroxytetradecanoyl)-DL-homoserine lactone(C14-HSL, sigma-aldrich, 51481).
All plasmids and strains are listed in Supplementary Data 1 & Supplementary Table 3.
Parts and circuits measurement
The expression of σ and anti-σ factors was regulated by two distinct inducible promoters: Ptac (induced by IPTG) and PcymRc (induced by Cumate). Induction was performed using a gradient of inducer concentrations (Supplementary Fig. 1). Input RPU (relative promoter units) was calculated based on the yellow fluorescent protein (YFP) expression measured under identical experimental conditions.
OA induction system measurements were conducted according to the following protocol: frozen stocks were streaked onto plates and grown overnight in Luria-Bertani (LB) medium. Overnight culture was diluted 200-fold in a 96-well plate with M9(CA) (without inducers) and incubated at 37 °C and 800 rpm for 5 h (PST-60HL-4). This culture was then diluted 20-fold into fresh M9(CA) medium supplemented with inducers and grown under the same conditions (Supplementary Fig. 19). After 3 h of incubation, OD600( ≈ 0.2-0.3) and fluorescence were measured.
For promoter and circuits characterization, the overnight culture (LB) was diluted 40-fold in a 96-well plate (Corning; 3799) with M9(CA) medium and incubated at 37 °C in a Digital Shaker (PST-60HL-4) at 800 rpm for 2 h. The culture was then diluted 40-fold a second time and grown under the same conditions. OD600 and fluorescence measurements were recorded during the exponential phase and the stationary phase of cell growth, corresponding to OD values of ~0.3 and ~1.6, typically occurring at around 3 h and 9 h, respectively49.
Continuous measurements were processed by a Synergy H1 plate reader (BioTek). The overnight cultures (LB) were diluted 200-fold into pre-warmed M9(CA) media in 96-well plates. After 2 h, the cultures were diluted 100-fold into pre-warmed M9(CA) media in 96-well plates. This plate was incubated at 37 °C in plate reader with high orbital shaking. OD600 and fluorescence measurements were recorded every 10 minutes for 24 h.
Calculation of Relative Promoter Units (RPU)
To characterize a promoter, the median fluorescence values (YFP) were measured along with the fluorescence of the RPU reference strain (E. coli Reference) (\({{YFP}}_{RE}\)) and that from white cells (E. coli MG1655) (\({{YFP}}_{0}\)), all under the same growth conditions as described above. The promoter activity in RPU was calculated using the following formula:
If CFP or mCherry were used as reporters, their relative promoter units (RPUs) were calculated using their respective reference plasmids. The excitation and emission wavelengths for the fluorescent proteins were as follows: YFP (500 nm/530 nm), CFP (433 nm/470 nm), and mCherry (580 nm/610 nm).
Shikimate fermentation
Shikimate production trials were conducted in M9 minimal medium (without CA) containing 10 g/L glucose as the sole carbon source. Strains were initially grown overnight in LB medium and subsequently transferred to fresh M9 minimal medium (1% glucose) at 1% inoculum ratio. Fermentation was carried out in 50 mL working volumes with an initial OD of 0.05. Flasks were incubated under controlled conditions: 30 °C, 220 rpm shaking, and 80% relative humidity for 96 h(ZQZY-CF8). Samples were collected periodically to analyze extracellular metabolites.
Supernatants were analyzed by high-performance liquid chromatography (HPLC) using an Agilent 1260 series instrument equipped with an Aminex HPX-87H column (300 mm by 7.8 mm; Bio-Rad Laboratories, Hercules, CA). An isocratic 25-minute method with a 5 mM sulfuric acid mobile phase at a flow rate of 0.6 mL/min was employed. The column temperature was maintained at 55 °C, and the refractive index detector (RID) was set to 35 °C.
His-tagged protein expression, purification, quantification, and SDS-PAGE analysis
To evaluate circuit-dependent protein expression, His-tagged target proteins were expressed in Escherichia coli BL21(DE3) under both induced and non-induced conditions. Monoclonal colonies were inoculated into LB medium and cultured overnight at 37 °C with shaking at 220 rpm (Bluepard THZ-103B). The following day, cultures were diluted 1% (v/v) into fresh LB medium for pre-culture expansion.
For non-induced systems, cells were grown to OD₆₀₀ ≈ 0.3, then further inoculated 1% (v/v) into 50 mL LB medium in 250 mL baffled flasks and cultivated for 24 h at 37 °C, 220 rpm(ZQZY-CF8). For induced systems, cells were grown to OD₆₀₀ ≈ 0.5 before addition of anhydrotetracycline (aTc, final concentration 200 ng/mL), followed by continued incubation under identical conditions for 24 h. Cultures then were centrifuged at 3928×g for 5 min to collect cell pellets, which were washed twice with sterile PBS buffer. Cell disruption was performed by high-pressure homogenization, and lysates were clarified by centrifugation at 15712×g for 10 min at 4 °C to collect soluble protein fractions.
His-tagged proteins were purified from the supernatants using Ni-NTA affinity chromatography (Ni-NTA 6FF resin, Sangon Biotech, Cat. No. C600033) via gravity-flow columns, following the manufacturer’s protocol. The procedure involved column equilibration, sample binding, washing with buffer containing imidazole, and elution of bound proteins with high-imidazole elution buffer.
Protein concentration was quantified directly from the eluted fractions using a microvolume nucleic acid/protein spectrophotometer (e.g., NanoDrop One, Thermo Fisher). Absorbance at 280 nm was measured, and protein concentrations were calculated using instrument software with appropriate extinction coefficients.
Purified proteins were analyzed by SDS-PAGE. A 40 μL aliquot of each sample was mixed with 10× SDS-PAGE loading buffer and denatured at 100 °C for 10 min. Samples were loaded onto 15% YEASEN Precast Protein Plus Gels (Cat. No. 36248ES10), with 10 μL of protein marker (Thermo, 10–170 kDa, Cat. No. 26616). Electrophoresis was run at 100 V for 1 h followed by 120 V for 2–3 h. Gels were stained overnight with Coomassie Brilliant Blue R-250 (50% methanol, 10% acetic acid, 0.025% dye), then destained twice using 5% acetic acid and 5% ethanol for 3–4 h per round. Gels were imaged using a Bio-Rad ChemiDoc Imaging System, and protein band intensities were quantified using ImageLab software.
Flow cytometry analysis
Fluorescence was measured using a FACSCelesta flow cytometer equipped with a high-throughput sampler (HTS, BD Biosciences). For each sample, more than 10,000 single-cell events were collected. YFP fluorescence was detected using the FITC-A channel. Gating was performed based on forward scatter width (FSC-A) and side scatter width (SSC-A) to exclude debris and cell aggregates (Supplementary Fig. 22). Data were analyzed using FlowJo (v10, BD Biosciences), and median fluorescence intensity was extracted for each gated population. Fluorescence values were reported in arbitrary units (a.u.), calculated as the difference between the measured YFP signal and background autofluorescence (YFPmeasure – YFP0).
RNA-seq analysis of E. coli native promoters
Publicly available RNA-seq data from E. coli K-12 cultures (SRA accession: PRJNA640168) were used to analyze native promoter activity under different growth phases. Specifically, the LB 0.4 Term-seq dataset was used to represent the exponential phase, and the LB 2.0 Term-seq dataset was used for the stationary phase. Raw reads were aligned to the E. coli genome, and per-base coverage (depth) was normalized across datasets. Transcription start sites (TSSs) were identified as genomic positions with a sharp increase in normalized depth. Promoter strength was calculated as the average difference in normalized coverage within ±10 bp around the TSS. Promoters were classified as exponential- or stationary-phase-specific based on the SE ratio (stationary/exponential), using thresholds of >1.5 or <0.67.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The RNA-seq data used in this study were previously published and are available in the NCBI Sequence Read Archive (SRA) under accession code PRJNA640168. The source code and processed data used in this study are available on GitHub at https://github.com/ChenYe-Lab/Synthetic-Signal-Transformation. The raw measurement data generated in this study are provided in the Supplementary Information. A list of plasmids used in this study is provided in Supplementary Data 1, and a list of key DNA sequences is provided in Supplementary Data 2. Source data are provided with this paper.
Code availability
Computer code is available from GitHub under https://github.com/ChenYe-Lab/Synthetic-Signal-Transformation and a snapshot of the code is provided on Zenodo83.
References
Gao, Y., Wang, L. & Wang, B. Customizing cellular signal processing by synthetic multi-level regulatory circuits. Nat. Commun. 14, 8415 (2023).
Babu, M. et al. Genetic interaction maps in Escherichia coli reveal functional crosstalk among cell envelope biogenesis pathways. PLoS Genet. 7, e1002377 (2011).
Rajagopala, S. V. et al. The binary protein-protein interaction landscape of Escherichia coli. Nat. Biotechnol. 32, 285–290 (2014).
Wan, X. et al. Cascaded amplifying circuits enable ultrasensitive cellular sensors for toxic metals. Nat. Chem. Biol. 15, 540–548 (2019).
Lee, H. et al. Plug-and-play protein biosensors using aptamer-regulated in vitro transcription. Nat. Commun. 15, 7973 (2024).
Chen, Y. et al. Tuning the dynamic range of bacterial promoters regulated by ligand-inducible transcription factors. Nat. Commun. 9, 64 (2018).
Xu, B. et al. On-chip engineered living materials as field-deployable biosensing laboratories for multiplexed detection. Adv. Funct. Mater. 35, 2416830 (2024).
Guo, S., Du, J., Li, D., Xiong, J. & Chen, Y. Versatile xylose and arabinose genetic switches development for yeasts. Metab. Eng. 87, 21–36 (2025).
Zhang, F., Carothers, J. M. & Keasling, J. D. Design of a dynamic sensor-regulator system for production of chemicals and fuels derived from fatty acids. Nat. Biotechnol. 30, 354–359 (2012).
Zhong, G. et al. A reversible RNA on-switch that controls gene expression of AAV-delivered therapeutics in vivo. Nat. Biotechnol. 38, 169–175 (2020).
Chowdhury, S. et al. Programmable bacteria induce durable tumor regression and systemic antitumor immunity. Nat. Med. 25, 1057–1063 (2019).
Gardner, T. S., Cantor, C. R. & Collins, J. J. Construction of a genetic toggle switch in Escherichia coli. Nature 403, 339–342 (2000).
Tamsir, A., Tabor, J. J. & Voigt, C. A. Robust multicellular computing using genetically encoded NOR gates and chemical ‘wires’. Nature 469, 212–215 (2011).
Weinberg, B. H. et al. Large-scale design of robust genetic circuits with multiple inputs and outputs for mammalian cells. Nat. Biotechnol. 35, 453–462 (2017).
Chen, Y. et al. Genetic circuit design automation for yeast. Nat. Microbiol. 5, 1349–1360 (2020).
Bonnet, J., Yin, P., Ortiz, M. E., Subsoontorn, P. & Endy, D. Amplifying genetic logic gates. Science 340, 599–603 (2013).
Müller, I. E. et al. Gene networks that compensate for crosstalk with crosstalk. Nat. Commun. 10, 4028 (2019).
Purcell, O. & Lu, T. K. Synthetic analog and digital circuits for cellular computation and memory. Curr. Opin. Biotechnol. 29, 146–155 (2014).
Bashor, C. J. et al. Complex signal processing in synthetic gene circuits using cooperative regulatory assemblies. Science 364, 593–597 (2019).
Chen, Z. et al. A synthetic protein-level neural network in mammalian cells. Science 386, 1243–1250 (2024).
Li, X. et al. Synthetic neural-like computing in microbial consortia for pattern recognition. Nat. Commun. 12, 3139 (2021).
Chen, Y., Kim, J. K., Hirning, A. J., Josić, K. & Bennett, M. R. Emergent genetic oscillations in a synthetic microbial consortium. Science 349, 986–989 (2015).
Brophy, J. A. & Voigt, C. A. Principles of genetic circuit design. Nat. methods 11, 508–520 (2014).
Holt, B. A. & Kwong, G. A. Protease circuits for processing biological information. Nat. Commun. 11, 5021 (2020).
Rizik, L., Danial, L., Habib, M., Weiss, R. & Daniel, R. Synthetic neuromorphic computing in living cells. Nat. Commun. 13, 5602 (2022).
Daniel, R., Rubens, J. R., Sarpeshkar, R. & Lu, T. K. Synthetic analog computation in living cells. Nature 497, 619–623 (2013).
Zeng, J. et al. A synthetic microbial operational amplifier. ACS Synth. Biol. 7, 2007–2013 (2018).
Aoki, S. K. et al. A universal biomolecular integral feedback controller for robust perfect adaptation. Nature 570, 533–537 (2019).
Studier, F. W. Protein production by auto-induction in high-density shaking cultures. Protein Expr. Purif. 41, 207–234 (2005).
Gupta, A., Reizman, I. M. B., Reisch, C. R. & Prather, K. L. Dynamic regulation of metabolic flux in engineered bacteria using a pathway-independent quorum-sensing circuit. Nat. Biotechnol. 35, 273–279 (2017).
Doong, S. J., Gupta, A. & Prather, K. L. Layered dynamic regulation for improving metabolic pathway productivity in Escherichia coli. Proc. Natl Acad. Sci. 115, 2964–2969 (2018).
Dinh, C. V., Chen, X. & Prather, K. L. Development of a quorum-sensing based circuit for control of coculture population composition in a naringenin production system. ACS Synth. Biol. 9, 590–597 (2020).
You, L., Cox Iii, R. S., Weiss, R. & Arnold, F. H. Programmed population control by cell–cell communication and regulated killing. Nature 428, 868–871 (2004).
Xu, X. et al. Pyruvate-responsive genetic circuits for dynamic control of central metabolism. Nat. Chem. Biol. 16, 1261–1268 (2020).
Nielsen, J. & Keasling, J. D. Engineering cellular metabolism. Cell 164, 1185–1197 (2016).
Yu, T. et al. Metabolic reconfiguration enables synthetic reductive metabolism in yeast. Nat. Metab. 4, 1551–1559 (2022).
Yang, S., Du, G., Chen, J. & Kang, Z. Characterization and application of endogenous phase-dependent promoters in Bacillus subtilis. Appl. Microbiol. Biotechnol. 101, 4151–4161 (2017).
Wu, Y. et al. CRISPR–dCas12a-mediated genetic circuit cascades for multiplexed pathway optimization. Nat. Chem. Biol. 19, 367–377 (2023).
Groseclose, T. M., Hersey, A. N., Huang, B. D., Realff, M. J. & Wilson, C. J. Biological signal processing filters via engineering allosteric transcription factors. Proc. Natl Acad. Sci. 118, e2111450118 (2021).
Shabestary, K. et al. Design of microbial catalysts for two-stage processes. Nat. Rev. Bioeng. 2, 1039–1055 (2024).
Anilionyte, O., Liang, H., Ma, X., Yang, L. & Zhou, K. Short, auto-inducible promoters for well-controlled protein expression in Escherichia coli. Appl. Microbiol. Biotechnol. 102, 7007–7015 (2018).
Pham, N. N. et al. Auto-inducible synthetic pathway in E. coli enhanced sustainable indigo production from glucose. Metab. Eng. 85, 14–25 (2024).
Rhodius, V. A. et al. Design of orthogonal genetic switches based on a crosstalk map of σs, anti-σs, and promoters. Mol. Syst. Biol. 9, 702 (2013).
Helmann, J. D. The extracytoplasmic function (ECF) sigma factors. Adv. Microb. Physiol. 46, 47–110 (2002).
Zong, Y. et al. Insulated transcriptional elements enable precise design of genetic circuits. Nat. Commun. 8, 52 (2017).
Typas, A., Becker, G. & Hengge, R. The molecular basis of selective promoter activation by the σS subunit of RNA polymerase. Mol. Microbiol. 63, 1296–1306 (2007).
Wong, G. T. et al. Genome-wide transcriptional response to varying RpoS levels in Escherichia coli K-12. J. Bacteriol. 199, https://doi.org/10.1128/jb.00755-00716 (2017).
Schellhorn, H. E. Function, evolution, and composition of the RpoS regulon in Escherichia coli. Front. Microbiol. 11, 560099 (2020).
Zhang, H. M. et al. Measurements of gene expression at steady state improve the predictability of part assembly. ACS Synth. Biol. 5, 269–273 (2016).
Zhang, X. & Studier, F. W. Mechanism of inhibition of bacteriophage T7 RNA polymerase by T7 lysozyme. J. Mol. Biol. 269, 10–27 (1997).
Jeruzalmi, D. & Steitz, T. A. Structure of T7 RNA polymerase complexed to the transcriptional inhibitor T7 lysozyme. EMBO J. 17, 4101–4113 (1998).
Graeme, J. G. Optimizing op Amp Performance (McGraw-Hill Professional Publishing, 1997).
Teo, J. J. & Sarpeshkar, R. The merging of biological and electronic circuits. Iscience 23 (2020).
Nevozhay, D., Adams, R. M., Murphy, K. F., Josić, K. I. & Balázsi, G. Negative autoregulation linearizes the dose–response and suppresses the heterogeneity of gene expression. Proc. Natl Acad. Sci. 106, 5123–5128 (2009).
Mejía-Almonte, C. et al. Redefining fundamental concepts of transcription initiation in bacteria. Nat. Rev. Genet. 21, 699–714 (2020).
Ishihama, A. Functional modulation of Escherichia coli RNA polymerase. Annu. Rev. Microbiol. 54, 499–518 (2000).
Dong, T., Kirchhof, M. G. & Schellhorn, H. E. RpoS regulation of gene expression during exponential growth of Escherichia coli K12. Mol. Genet. Genomics 279, 267–277 (2008).
Weber, H., Polen, T., Heuveling, J., Wendisch, V. F. & Hengge, R. Genome-wide analysis of the general stress response network in Escherichia coli: σS-dependent genes, promoters, and sigma factor selectivity. J. Bacteriol. 187, 1591–1603 (2005).
Forquet, R., Pineau, M., Nasser, W., Reverchon, S. & Meyer, S. Role of the discriminator sequence in the supercoiling sensitivity of bacterial promoters. Msystems 6, https://doi.org/10.1128/msystems.00978-00921 (2021).
Mutalik, V. K. et al. Precise and reliable gene expression via standard transcription and translation initiation elements. Nat. methods 10, 354–360 (2013).
Ni, C., Dinh, C. V. & Prather, K. L. Dynamic control of metabolism. Annu. Rev. Chem. Biomol. Eng. 12, 519–541 (2021).
Hartline, C. J., Schmitz, A. C., Han, Y. & Zhang, F. Dynamic control in metabolic engineering: Theories, tools, and applications. Metab. Eng. 63, 126–140 (2021).
Ream, M. & Prather, K. L. Engineered autonomous dynamic regulation of metabolic flux. Nat. Rev. Bioeng. 2, 233–243 (2024).
Rosano, G. L. & Ceccarelli, E. A. Recombinant protein expression in Escherichia coli: advances and challenges. Front. Microbiol. 5, 172 (2014).
Dvorak, P. et al. Exacerbation of substrate toxicity by IPTG in Escherichia coli BL21 (DE3) carrying a synthetic metabolic pathway. Microb. Cell Factories 14, 1–15 (2015).
Cardinale, S., Joachimiak, M. P. & Arkin, A. P. Effects of genetic variation on the E. coli host-circuit interface. Cell Rep. 4, 231–237 (2013).
Grant, P. K. et al. Orthogonal intercellular signaling for programmed spatial behavior. Mol. Syst. Biol. 12, 849 (2016).
Kylilis, N., Tuza, Z. A., Stan, G.-B. & Polizzi, K. M. Tools for engineering coordinated system behaviour in synthetic microbial consortia. Nat. Commun. 9, 2677 (2018).
Wang, B., Barahona, M. & Buck, M. A modular cell-based biosensor using engineered genetic logic circuits to detect and integrate multiple environmental signals. Biosens. Bioelectron. 40, 368–376 (2013).
Collins, C. H., Leadbetter, J. R. & Arnold, F. H. Dual selection enhances the signaling specificity of a variant of the quorum-sensing transcriptional activator LuxR. Nat. Biotechnol. 24, 708–712 (2006).
Meyer, A. J., Segall-Shapiro, T. H., Glassey, E., Zhang, J. & Voigt, C. A. Escherichia coli “Marionette” strains with 12 highly optimized small-molecule sensors. Nat. Chem. Biol. 15, 196–204 (2019).
Greenberg, E. P. Bacterial communication and group behavior. J. Clin. Investig. 112, 1288–1290 (2003).
Klumpe, H. E., Garcia-Ojalvo, J., Elowitz, M. B. & Antebi, Y. E. The computational capabilities of many-to-many protein interaction networks. Cell Syst. 14, 430–446 (2023).
Pandi, A. et al. Metabolic perceptrons for neural computing in biological systems. Nat. Commun. 10, 3880 (2019).
Nielsen, A. A. et al. Genetic circuit design automation. Science 352, aac7341 (2016).
Hernández-Sancho, J. M. et al. A versatile microbial platform as a tunable whole-cell chemical sensor. Nat. Commun. 15, 8316 (2024).
Shin, J., Zhang, S., Der, B. S., Nielsen, A. A. & Voigt, C. A. Programming Escherichia coli to function as a digital display. Mol. Syst. Biol. 16, e9401 (2020).
Green, A. A. et al. Complex cellular logic computation using ribocomputing devices. Nature 548, 117–121 (2017).
Bondy-Denomy, J. et al. Multiple mechanisms for CRISPR–Cas inhibition by anti-CRISPR proteins. Nature 526, 136–139 (2015).
Sharan, S. K., Thomason, L. C., Kuznetsov, S. G. & Court, D. L. Recombineering: a homologous recombination-based method of genetic engineering. Nat. Protoc. 4, 206–223 (2009).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl Acad. Sci. 97, 6640–6645 (2000).
Engler, C., Kandzia, R. & Marillonnet, S. A one pot, one step, precision cloning method with high throughput capability. PloS one 3, e3647 (2008).
Cao, W., Liu, L., Sun, Q., Shan, Y. & Chen, Y. ChenYe-Lab/synthetic-signal-transformation. Zenodo (2025).
Andersen, J. B. et al. New unstable variants of green fluorescent protein for studies of transient gene expression in bacteria. Appl. Environ. Microbiol. 64, 2240–2246 (1998).
Acknowledgements
The authors would like to thank Professor Hirotada Mori and Dr. Xi Yang for generously providing the σ38 knockout strain, which was instrumental for validating our σ38-specific promoters. This work was supported by National Key Research and Development Program of China [2024YFA0916700] to L.L.; the National Natural Science Foundation of China [32171419] to Y.C. and [32330084] to Y.S.; and the Strategic Priority Research Program of the Chinese Academy of Sciences [XDA0510200] to Y.C.
Author information
Authors and Affiliations
Contributions
Y.C. conceived the study. L.L. and Y.C. designed the experiments, W.C., L.L. and Q.S. performed the experiments, L.L. and Y.C. conducted the mathematical model and analyzed data. L.L. and Y.C. wrote the manuscript, Y.C. and Y.S. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
Y.C., L.L., and W.C. are listed as inventors on a patent application (CN202510595826.5, PCT/CN2025/093793) related to the synthetic OA circuits described in this study. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Cao, W., Liu, L., Sun, Q. et al. A framework for complex signal processing via synthetic biological operational amplifiers. Nat Commun 16, 7037 (2025). https://doi.org/10.1038/s41467-025-62464-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-62464-9