Abstract
Artemisinin and its semisynthetic derivatives (ART) are crucial medicines in artemisinin-based combination therapies worldwide. Despite ART’s efficacy, small proportions of young intraerythrocytic ring stage parasites can survive the drug’s short half-life, and dormant forms can cause recrudescence if not cleared by partner drugs. Certain mutations in the Kelch propeller region of P. falciparum protein (PfK13) are linked to the higher ring-stage survival (RS), which above 1% can be a feature of ‘artemisinin partial resistance’. Emerging evidence indicates epigenetic modulators may contribute to RS. Here, we report systematic evaluations of all putative histone acetyltransferases (HATs) of P. falciparum in 30 culture-adapted field isolates and 43 subcloned field isolates. Only PfMYST shows a full association with RS phenotype modulations. Knockdown experiments confirm the linkage of Pfmyst expression to these modulations, with evidence of altered metabolic processes. Through single-cell RNA sequencing, ChIP-seq analysis, and CRISPR/cas9 genetic manipulation, PfMYST-targeted RS-related genes have been identified and functionally validated. Multi-omics analysis indicates significant interplay of PfMYST and PfK13 mechanisms in RS. PfMYST epigenetic modulation extends to other antimalarials, including amodiaquine, pyrimethamine, chloroquine, and pyronaridine. Collectively, our findings provide important information on the epigenetic regulatory mechanism of P. falciparum RS after pulses of ART and other antimalarials.
Similar content being viewed by others
Introduction
Artemisinin and its semisynthetic derivatives (collectively termed ART) are crucial first-line drugs against falciparum malaria. From 2000–2015, an estimated 22% (17%–28%) of the 663 (542–752) million clinical cases averted by malaria control measures were attributed to the use of an ART drug in combination with a partner drug from a different antimalarial class (artemisinin-based combination therapy, or ACT)1. ACT is effective in preventing the recrudescence of parasitemia that can occur as Type I drug failure (WHO RI level resistance) after ART treatment alone (monotherapy)2,3,4. The partner drug works to eliminate the small fraction of blood-stage parasites after they persist in a ‘dormant’ state, while the bulk of the parasitemia is killed off by ART5,6,7. Persisters that are not completely eliminated can give rise to renewed parasitemia and recrudescent infection. This phenomenon was dramatically exemplified in 2004 by an International Artemisinin Study Group meta-analysis demonstrating the inefficacy of chloroquine (CQ) as a partner drug. In trials using two different ACTs in six African countries, 28-day cure rates were achieved in only 7–53% (PCR uncorrected) of patients treated with artesunate-chloroquine in the Ivory Coast, Burkina Faso, and Sao Tome–Principe, whereas cure rates were achieved in 68–85% (PCR uncorrected) of patients treated with artesunate-amodiaquine in Kenya, Senegal, and Gabon8. More recently in Africa, increasing clinical failure rates of sulfadoxine/pyrimethamine-containing ACTs due to antifolate mutations (in dhfr and dhps) of P. falciparum required switches to alternative partner drugs such as lumefantrine (LMF), amodiaquine (AQ) or piperaquine (PPQ)9,10,11,12. In Cambodia, the rapid development of PPQ resistance due to plasmepsin 2/3 genes amplification and mutations in the pfcrt gene mandated the discontinuation of PPQ and a return to mefloquine (MEF) as a partner drug for ACT13,14,15.
ART-treated Plasmodium falciparum malaria parasites also exhibit a phenotype of ring-stage survival (RS)16. This phenotype can be elicited in RS assays (RSA0-3h) that expose 0–3 h early-stage ring-infected erythrocytes to a high-concentration pulse (e.g., 700 nM) of dihydroartemisinin (DHA) for 6 hours, after which the cells are then returned to culture without DHA and the surviving parasites are counted at 72 h. RS levels, calculated from the ratio of parasite counts from DHA-treated ring stages to counts from control untreated ring stages, can range from ca. 0.1% to > 10%, and RS levels ≥ 1% are considered a feature of ‘partial’ artemisinin resistance (ART-R)17. RSA0-3h have come into favor over conventional half-maximal inhibitory or lethal dose measures (IC50, LD50) of parasite susceptibility, which show little variation of ART response among parasite strains (typically at low nM levels of DHA) because these measures cover the full 2-day P. falciparum life cycle and growing intraerythrocytic trophozoites and schizonts of all the strains are killed16. Three-day courses of an ART with an effective partner drug thus remain effective, despite RS variations6,18,19,20. Indeed, in recent studies, the inheritance of high- or low-RS phenotypes in a P. falciparum cross did not show any linkage of RS > 1% to higher IC50 values or to recrudescences from the dormant persister stages after non-human primate infections were treated with 3 days of artesunate alone21. Whether malaria parasites with RS phenotypes may evolve and give rise in the future to P. falciparum strains with frank RII or RIII ART-R in the trophozoite and schizont stages warrants careful monitoring.
Certain polymorphisms or mutations in the Kelch-repeat propeller of the P. falciparum protein K13 (PfK13) are linked to high RS levels21,22,23. Of 260 identified non-synonymous polymorphisms in PfK1324,25 (132 of these polymorphisms are entered into PlasmoDB; https://plasmodb.org/plasmo/app/record/gene/PF3D7_1343700), around 15 mutations with high RS rate have been associated with longer parasitemia clearance half-life (PC½ > 5 h) in Southeast Asia25, although exceptions to these associations are observed26,27. One well-known example, C580Y, is associated with a parasite clearance half-life > 5 hours in Southeast Asia23,24, but prolonged clearance times have yet to be demonstrated where this mutation occurs in a few isolates in regions of New Guinea and South America28,29,30,31. Nevertheless, the linkage of these PfK13 mutations to higher RS levels and the evidence for their spread under the wide use of ACT32,33,34 raises intriguing questions about the PfK13’s role in the parasite and any selective advantages the mutations may offer to the parasite under ART pressure. PfK13 has been found to regulate digestive vacuole (DV) biogenesis and mediate the endocytosis of host erythrocyte cytosol; mutations that reduce the activity of PfK13 may curb the digestion of hemoglobin, consequently reducing heme-dependent ART activation and increased RS levels after ART treatment35,36,37.
Also, effects on hemoglobin digestion or uptake may help to explain changes in the RS phenotype from falcipain mutations38,39,40,41 or mutations in UBP1, AP-2μ, and coronin42,43,44,45,46. Mutants with reduced PfK13 activity have also been observed to decelerate (delay) ring stage development by several hours before returning to a normal pace of development in trophozoites and schizonts37,47. Rings with several hours delayed development are not the same as persister stages, which have been shown to be morphologically and developmentally distinct from rings in recrudescence studies and a parasite cross7,21,48,49,50.
It is noteworthy that the WHO RI phenotype of ART-treated P. falciparum infections has been stable since the advent of ART in the 1970s, without progression to RII or RIII levels of resistance6. Some studies have identified parasites with increased tolerance to ART after sustained exposure to the drug in vitro; yet, these parasites lose their tolerance after the drug exposure is discontinued, indicating the possibility of reversible epigenetic mechanisms23,51. Frequent recrudescence of ART-treated P. falciparum infections occurs with or without K13 mutations21. These behaviors are analogous to epigenetically regulated adaptive resistance observed in bacteria, where phenotypic variability exists in isogenic populations due to intrinsic noise in gene expression, and survival rates at subinhibitory concentrations suggest non-mutation origins52. In addition, the detailed mechanism for artemisinin resistance still remains widely divided due to different research perspectives. For example, recent studies have also illustrated PfK13-independent mechanisms for artemisinin resistance, including other genetic factors, epigenetic alterations and epitranscriptomic-based RNA modification53,54,55. All these results have validated that artemisinin resistance can result from genetic or epigenetic factors which regulate gene expression collectively and produce a specific phenotype in response to ART. These evidences explain many aspects of RS, while why only a subset of parasites enters quiescence state remains less investigated. Due to the complexity of ART resistance, we thus focused on the detailed investigation for epigenetic mechanism based on our previous results in which a subpopulation of parasites can tolerate stress without genetic modifications.
In P. falciparum, epigenetic mechanisms play a pivotal role in regulating chromatin structure, gene expression, and stress responses56. Recent studies highlight the involvement of histone acetyltransferases (HATs) such as GCN5 and MYST in modulating stress responses to ART exposure. Lucky et al. found that inhibiting the P. falciparum GCN5 homolog PfGCN5 could reduce the RS level of parasites subjected to DHA treatment57. In addition, Liang et al. showed that manipulating PfK13 and PfMYST levels affected parasitemia recovery in ring stages exposed to 1000 nM DHA for 24 h58. Decreasing PfK13 and increasing PfMYST led to earlier recovery, while the opposite delayed recovery, consistent with previous observations on PfK13’s role in the rates of ART-treated RS35,37,59. Considering these findings, we have conducted further studies of members of the HAT family, including PfMYST, for their mediation of histone acetylation and effects on the RS phenotype. Here, we examine potential associations of the RS phenotypes and HAT expression levels detected in early rings of culture-adapted field isolates and subcloned field isolates of P. falciparum. Genetic manipulation and chemical inhibition experiments provide rigorous evaluations of the association of RS survival with Pfmyst expression. Resulting histone modifications are identified by ChIP-seq analysis, and single-cell RNA sequencing (scRNA-seq) is used to compare cell clusters from PfMYST knockdown parasites and various PfK13 mutant parasites exposed to DHA. Results from these experiments are used to examine 27 PfMYST-associated candidate genes for potential modulation of RS. Finally, we explore the effects of Pfmyst expression on RS after exposure to other well-known antimalarial drugs.
Results
Histone acetyltransferase expressions vary in P. falciparum parasites, with PfMYST linked to RS survival, regardless of PfK13 mutations
For our investigations of RS phenotypes in this work, we chose two clonal PfK13-mutant lines of P. falciparum for comparison to counterparts with the wild-type (WT) gene (Supplementary Data 1). Clonal line A8 carries the PfK13 C580Y mutation (A8C580Y), a codon change that is widely prevalent in Southeast Asia and is found at low levels in New Guinea, South America, and Africa30,31,60; Clonal line A5 carries the G538V mutation (A5G538V) from the China-Myanmar border61,62. To study the contribution of these PfK13 mutations to the RS phenotypes of A8 and A5, we used site-specific mutagenesis to revert the codons to the WT PfK13 sequence, thereby producing the C580 line A8Rev and the G538 line A5Rev (Fig. 1a). RS assays (RSA0-3h) using 700 nM DHA showed lower survival levels of 0.5 ± 0.2 % for the A8Rev parasites compared to 7.9 ± 0.8 % for the A8 C580Y parasites (P < 0.001); however, survival rates in these assays remained high for the A5Rev parasites, for which we obtained RSA0-3h levels of 4.4 ± 0.8 % compared to 5.5 ± 0.7 %, a non-significant difference (P = 0.072) (Fig. 1b). Our comparison of survival rates using the standard RSA0-3h assay confirmed a statistically significant difference between A8 and A5. For subsequent analyses, we used their respective revertant strains as controls, making the baseline differences acceptable within the study’s framework.
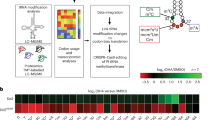
a Two isolates bearing mutations in the PfK13 gene, A5 (G538V) and A8 (C580Y), were reverted to the WT sequences via CRISPR-Cas9 gene editing technique, leading to the creation of A5_R (G538) and A8_R (C580), respectively. b RSA0-3h for the PfK13 mutant parasites: A8 (C580Y) and A5 (G538V); PfK13 WT parasites: A8_R (C580) and A5_R (G538). Data were presented as mean ± SEM from four independent experiments with technical duplicates. P-values were determined using the two-tailed student’s t test. c RSA0-3h for WT 3D7 strain and subclones of A8 (C580Y), A8_R (C580), A5 (G538V) and A5_R (G538) isolates with 50 nM, 200 nM and 700 nM DHA, respectively. Data were presented as the mean value of survival rate (n = 4, 18, 10, 25 and 10 for each group with 3 technical replicates). d Correlation analysis between the transcription levels of 10 genes encoding histone acetyltransferases and ART susceptibility in the subclones of A5 and A8 field isolates. Parasites carrying G538V and C580Y mutations in PfK13 were adapted and subcloned for RT-qPCR assay and 0–3 h ring-stage survival assay. The 0–3 h ring-stage parasites were exposed to a 6-hour treatment with 700 nM DHA. Data were presented as mean value from three independent experiments with technical triplicates. Correlation analysis was conducted using GraphPad Prism 8.0 software. Source data are provided as a Source Data file.
To investigate additional genetic variations, we performed whole-genome sequencing (WGS) on A8, A5, and their revertants. Our sequencing yielded high-quality data, with an average genome coverage depth exceeding 150X and > 95% of the genome covered at ≥ 10X across all samples. To assess genetic relatedness, we calculated PI_HAT values, which estimate the proportion of the genome shared due to identity by descent (IBD). A PI_HAT value greater than 0.9 is commonly used to confirm genetic identity63. In our dataset, all pairwise PI_HAT values between A8, A5, and their respective revertants exceeded 0.9, confirming that these subclones are indeed isogenic. As illustrated in (Supplementary Fig. 1a), we identified the G538V mutation in A5 and the C580Y mutation in A8, as well as successful reversion of these mutations in A5Rev and A8Rev. Within each strain, subclones exhibited identical or synonymous mutations, further supporting their isogenic status. Regarding the differing reversion patterns as well as resulting resistance phenotype after site-specific mutagenesis between A5Rev and A8Rev, we hypothesize that both genetic and epigenetic factors may contribute to the observed RS phenotypes64. Thus, we performed a modified RSA0-3h assay to assess survival rate variations among these subclones.
Eighteen subclones were obtained from A8, and all were verified by PCR and DNA sequencing to have the PfK13 C580Y mutation, consistent with their isogenic background; likewise, 25 subclones were obtained from A5 and verified to all have the G538V mutation (Supplementary Data 2 and Supplementary Data 3). RSA0-3h levels after 6 h 700 nM DHA exposures ranged between 0.9 % and 14.2 % for the A8 subclones, and between 1.2 % to 6.4 % for the A5 subclones (Fig. 1c). The resistant level of obtained subclones of A5Rev and A8Rev has also been quantified to further validate the variation of survival rate (Fig. 1c, Supplementary Fig. 2 and Supplementary Data 4). Interestingly, at 50 and 200 nM DHA exposures, the RSA0-3h levels of these subclones elevated significantly, ranging between 4.2 % to 70.1 % (A8: 50 nM DHA), 8.1 % to 41.4 % (A5: 50 nM DHA), 2.9 % to 22.1 % (A8: 200 nM DHA) and 3.5 % to 15.9 % (A5: 200 nM DHA), suggesting the potential involvement of concentration-dependent factors in epigenetic regulation. Next, to examine the expression of P. falciparum HAT genes and test for potential correlations with these observed RSA0-3h variations, we used RT-qPCR to determine the transcription levels of 10 putative HAT genes in each of the A8 and A5 subclones (Fig. 1d and Supplementary Fig. 1b). In these analyses, significant negatively sloped correlations were detected in the A8 subclones between the RSA0-3h levels and the transcription of two genes: Pfmyst (PF3D7_1118600; P = 0.0015) and a putative acetyltransferase (PF3D7_1020700; P = 0.0223).
In the A5 subclones, a significant, negatively sloped correlation of the RSA0-3h levels was also detected with Pfmyst transcription (P = 0.0449), but the correlation was not significant for PF3D7_1020700 (P = 0.1169). No significant correlations were found between the transcription level of any of the remaining 8 putative HAT genes and RSA0-3h levels of the A8 or A5 subclones exposed to the 6 h 700 nM DHA pulses. These remaining genes included Pfgcn5 (PF3D7_0823300), which has been reported to influence chromatin structure and gene expression in stress responses and various parasite processes including erythrocyte invasion, virulence, and drug responses57,65,66,67,68. In addition, the a significant negatively sloped correlation between the RSA0-3h levels and the transcription of Pfmyst was also observed in resistant A5rev subclones, while no significant correlation was found in sensitive A8rev subclones (Supplementary Fig. 3).
In light of the possible effect of DHA concentration on HAT gene expression57,68,69,70, we also obtained RSA0-3h levels using a 6 h pulse of 200 or 50 nM of DHA. Results of these experiments (Supplementary Figs. 4, 5) confirmed negatively sloped correlations between these RSA0-3h levels and Pfmyst transcription in the A8 and A5 subclones (P = 0.0024 and P = 0.0027 at 50 nM, P = 0.0007 and P = 0.0051 at 200 nM, respectively), as well as between these RSA0-3h levels and PF3D7_1020700 transcription in the A8 and A5 subclones (P = 0.0042 and P = 0.0312 at 50 nM, P = 0.008 and P = 0.0882 at 200 nM, respectively). For the remaining 8 genes, some significant negatively sloped correlations were also observed from the A5 or A8 subclones, e.g., between transcription levels of two genes (PF3D7_1323300 and PF3D7_1227800) and RSA0-3h results from the A8 subclones after 6 h 50 nM DHA exposures. however, no correlations were consistently present in all groups with various DHA exposures. Intriguingly, among the ten HATs genes, we observed a tendency of positive-sloped correlation only between the Pfgcn5 expression and RSA0-3h levels without statistical significance (all P > 0.05).
Considering the consistently strong, negatively sloped correlations of Pfmyst transcription with RSA0-3h levels at different DHA concentrations in both A8 and A5 subclones, we studied an additional 30 culture-adapted P. falciparum isolates from African countries and the Thai-Myanmar border (Supplementary Data 1). By direct DNA sequencing of PCR products, all of these parasite lines were found to carry wild-type sequences in the PfK13 β-propeller domain. The correlation analysis between Pfmyst transcription levels and ART susceptibility was conducted using survival rates obtained from RSA0-3h assays, where parasites were exposed to 50 nM, 200 nM, and 700 nM DHA. RSA0-3h levels after 6 h pulses of either 50 nM or 200 nM DHA indicated a strong, negatively sloped correlation association with Pfmyst transcription (Fig. 2a and Supplementary Fig. 6aP = 0.0008 and 0.0209, respectively), confirming PfMYST-mediated epigenetic regulation of P. falciparum RS levels, in PfK13 WT as well as PfK13 mutant parasites (Supplementary Fig. 6b).
a Correlation analysis between the transcription levels of Pfmyst and ART susceptibility in field-adapted parasites (n = 28) carrying the WT PfK13 gene. Parasites were exposed to 50 nM DHA and then cultured normally for 66 hours. The correlation analysis was performed using GraphPad Prism 8.0 software, and data were presented as mean value from three independent experiments with technical triplicates. b Schematic illustrating the CRISPR/Cas9-mediated fusion of the glmS sequence and Ty1 tag at the 3’ terminus of PfMYST, resulting in the PfM-KD parasite line (Pfmyst-ty1-glmS). The fusion event was verified by PCR with gDNA extracted from the transfectant. Similar trends were observed in two independent experiments. c Intraerythrocytic growth curves of two subclones (C4, D5) of the Pfmyst-ty1-glmS line and the WT 3D7 control, with and without 5 mM GlcN treatment for 6 h. Data were presented as mean ± SEM from three independent replicates with technical triplicates. d Western Blot analysis of the Pfmyst-ty1-glmS line and WT 3D7 parasites, in which parasites were pretreated with 5 mM GlcN for 6 h and total protein extracts were obtained from ring, trophozoite, and schizont-stage parasites. The analysis utilized a commercial antibody against the Ty1 epitope, alongside Aldolase as the internal control. Similar trends were observed in two independent experiments; RT-qPCR analysis of Pfmyst transcript level at ring (0-6 hpi), trophozoite (20–26 hpi), and schizont-stage (36–42 hpi) with/without the exposure of GlcN. Data were presented as mean ± SEM from four independent experiments with technical triplicates. P-values were determined using the two-tailed student’s t test. e RSA with DHA concentrations ranging from 50 nM to 700 nM was tested on different parasites, including PfM-KD parasites with or without GlcN treatment, WT and Rrp6 knockdown parasites (Pfrrp6-Ty1-glmS). Data were presented as mean ± SEM from four independent experiments with technical duplicates. f The effects of histone acetyltransferase inhibitors on parasite RSA0-3h induced by DHA. WT parasites were pre-incubated with PfMYST specific inhibitor (NU9056) and PfGCN5 inhibitors (curcumin, and anacardic acid) for 12 h before RSA0-3h. Various DHA concentrations (50 nM, 200 nM, 700 nM) were tested. Data were presented as mean ± SEM from three independent experiments with technical triplicates. P-values were determined using One-way ANOVA with Bonferroni correction compared with vehicle control. CCM: curcumin, AA: anacardic acid. Source data are provided as a Source Data file.
Reduction of PfMYST activity in P. falciparum confers increased RS rates and slower ring-stage development after artemisinin treatment
In budding yeast, Saccharomyces cerevisiae, the histone acetyltransferase complex consists of multiple subunits, with Esa1 being the catalytic subunit, representing the sole essential acetyltransferase in yeast71. Esa1 belongs to the MYST family of acetyltransferases. In P. falciparum, the orthologous enzyme of Esa1, PfMYST, has been demonstrated to fulfill the conserved function of H4 acetylation and is indispensable for the parasite during intraerythrocytic development72. To investigate the effects of the PfMYST-mediated RS phenotype in P. falciparum, we utilized CRISPR-Cas9-editing to generate Pfmyst-Ty1-glmS for ribozyme knockdown of Pfmyst expression in P. falciparum line 3D7 (designated as PfMYST-KD or PfM-KD line from hereon) (Fig. 2b). The 3D7 parasites were readily transformed with this construct and subcloned although, as reported previously for the glmS system73, propagation of the Pfmyst-Ty1-glmS-transformed subclones was about 50% inhibited in routine culture without GlcN treatment (subclones C4 and D5, Supplementary Fig. 7a). In comparison, application of GlcN treatment completely suppressed propagation of these subclones (Supplementary Fig. 7a). To minimize the impact of GlcN on parasite growth in RS experiments, we optimized the timing of GlcN addition before the enrichment of the late-stage schizonts as previously described37. In the optimized method where the pretreatment time was shortened to 6 h, the growth inhibition effect was minimal, with only a 50% reduction in growth compared to the WT parasites (Fig. 2c).
To address the issue whether the KD effect, as well as the related phenotype, could maintain after the removal of GlcN, either the resistance or the mRNA expression level was equivaled by RSA0-3h and RT-qPCR. As illustrated in Supplementary Fig. 7b, resistant phenotype could be observed for parasites exposed to at least 6 h GlcN treatment. Meanwhile, according to the results from RT-qPCR, the transcript level was down-regulated at 36 h post removal for parasites subjected to either 6 h or 24 h treatment (Supplementary Fig. 7c). Taken together, these results indicated that 6 h-preatment of GlcN was enough for gene KD and resistance induction. GlcN was added to the culture 6 h prior to purifying the schizonts with Percoll and maintained until 1 h after the addition of DHA in our RSA0-3h assays.
Western-blot assays revealed reductions of PfMYST protein production in ring, trophozoite, and schizont stages in the GlcN-treated relative to untreated PfM-KD line (Fig. 2d). The mRNA expression level of Pfmyst at different developmental stages was further validated using RT-qPCR to investigate the inducible KD effect, and consistent with results from Western blot assay, significantly decreased transcript level was achieved, indicating the reliability for our KD strategy (Fig. 2d). For further confirmation of the ribozyme’s knockdown effect, we compared Pfmyst expression detected from the Ty1 tag in PfM-KD line to Pfmyst expression in a control line transformed with a TH-Pfmyst construct encoding Ty1 and HA at the N-terminus of PfMYST. In experiments with no GlcN added to the cultured parasites, western blots showed that Pfmyst expression level was significantly lower in the PfM-KD line relative to Ha-Ty1-Pfmyst-transformed parasites (Supplementary Fig. 1c). This suggests that the fusion of the glmS sequence at the 3’ terminus of the Pfmyst gene may destabilize transcripts by disrupting its mRNA processing, thereby reducing the PfMYST product at the translational level. In order to investigate whether the resistance phenotype could maintain upon the removal of GlcN, both modified RSA0-3h and consecutive RSA0-3h were performed. Results from these experiments further validated a possible epigenic resistance mechanism (Supplementary Fig. 7d, e).
Figure 2e presents the RSA0-3h results from three parasite lines with or without this exposure to GlcN; the PfM-KD line; the untransformed 3D7 line; and Pfrrp6-Ty1-glmS, a previously described glmS transformant of the Pfrrp6 gene74. These RSA0-3h results after 700 nM DHA treatment showed a significant increase of the survival rate from 1.1 % +/− 0.3 % to 7.9 % +/− 2.0 % in the GlcN-exposed PfM-KD parasites, providing direct evidence for PfMYST’s effect on RS. Significant increases of RS from 3.2 % +/− 1.4 % to 12.4 % +/− 1.0 % and 6.4 % +/− 1.0 % to 20.3 % +/− 2.8 % were likewise observed after 200 nM and 50 nM DHA treatment in these assays. We note that increased survivals were also evident in the PfM-KD parasites not exposed to GlcN (e.g., the 1.1 % RS rate of these parasites after 700 nM DHA treatment, Fig. 2e), consistent with reduced expression of Pfmyst from transcript destabilization by the glmS sequence noted above. Knockdown of Pfrrp6 expression caused little or no increase in the RS rates, confirming that the substantial RS increases after PfMYST knockdown were from reduced Pfmyst expression and not from a growth defect or unexpected effect of the glmS ribozyme system (Fig. 2e). To further validate the impact of the Pfmyst expression on P. falciparum ART susceptibility, we designed experiments to inhibit PfMYST activity using a selective MYST inhibitor (NU9056), along with control experiments that used two known HAT inhibitors specific to GCN5 (curcumin and anacardic acid)75,76,77 (Fig. 2f and Supplementary Fig. 8). NU9056, but not the GCN5 inhibitors, caused a significant increase in RSA0-3h, consistent with the findings of PfMYST-specific knockdown in our experiments.
Previous studies found that the RS phenotype of ART-R lines, conferred by PfK13 C580Y and certain other PfK13 mutations, exhibits a ring-stage development delay of up to 6 hours78,79; this delay, along with higher RS rates of the mutant parasites, increases reliance on subsequent drug doses to clear the drug-sensitive trophozoite and schizont stages. To test if PfMYST knockdown can confer a ring-stage development delay comparable to PfK13 C580Y, we exposed highly synchronized PfM-KD parasites to GlcN as described above and followed the stage timing of development after a pulse treatment of 0–3 h rings with 50 nM DHA. Initially, we also attempted to evaluate the growth arrest effect of parasites exposed to higher DHA concentrations78. However, due to the lethal effects of higher doses, particularly with the 3D7 parasite strain, the fluorescence signal was substantially weakened and overlapped with background noise. This significant interference made accurate measurement difficult. Therefore, we reduced the concentration to 50 nM, as previously demonstrated in studies80,81. This concentration was chosen to balance the induction of a detectable growth arrest effect while ensuring that the fluorescence signal remained distinguishable from the background, allowing for reliable quantification and interpretation of the experimental results.
In the upper panel of Fig. 3a, we first examined the life cycle of the corresponding parasite strains in the absence of DHA exposure. As shown in the representative Giemsa-stained smears, all parasite strains exhibited similar IDC durations of approximately 46–48 h, which helped exclude any potential effects of GlcN on parasite growth. However, when parasites were exposed to 50 nM DHA for 6 h, a temporal growth arrest was observed. A distinction was noted between sensitive strains (3D7 without GlcN and PfM-KD without GlcN) and resistant strains (PfM-KD pretreated with GlcN for 24 h and A8 field isolate carrying the C580Y mutation). Compared to parasites treated with GlcN or DHA but not both of these compounds together, only parasites with resistance treated with DHA showed a ring stage development delay of approximately 6 h (Fig. 3a–c).
a Intra-erythrocytic developmental stages of parasites observed in Giemsa-stained blood smears. Blood smears were prepared every 6 h to monitor the changes in stage development. The WT 3D7, Pfmyst-ty1-glmS without GlcN (PfM-KD (-), Pfmyst-ty1-glmS with GlcN (PfM-KD (+) and A8 (K13_C580Y) lines were monitored without or after 50 nM DHA pulse, respectively. A solid line was applied to mark the prolonged ring stage development. Similar trends were observed in two independent experiments. b Intra-erythrocytic development patterns of WT 3D7 and PfM-KD parasites given different exposures to GlcN followed by DHA treatment. Blood smears were conducted every 4 h, and the percentage of each stage parasite was obtained from two independent experiments. Curves were fitted to visualize the differences in stage developmental patterns, and data were presented as mean ± SEM from three independent experiments with technical duplicates. c Impact of 50 nM DHA treatment on parasite growth cycles. Changes in the SYTO-61 staining profile of viable parasites after drug exposure were monitored by FACS in the subsequent cycle. Growth retardation was observed in the DHA-treated PfMYST-KD parasites that were pre-treated with GlcN. d Assessment of stage-specific susceptibility for 3D7 and PfMYST-KD parasites using ART survival assays. Highly synchronized parasites were exposed to DHA during the intraerythrocytic developmental cycle (IDC) within specific time windows (RSA0-3h, RSA9-12h, and TSA18-21h), and the survival rates of the parasites were calculated. In parallel, control experiments were conducted using the same parasite strain without GlcN treatment. Data were presented as mean ± SEM from four independent experiments with technical duplicates. P-values were determined using One-way ANOVA with Bonferroni correction compared with 0–3 h samples. e The effect of repeated ART selection on RSA0-3h of PfMYST knockdown (PfM-KD (+)) and A8 (K13_C580Y) parasites, respectively. Data were presented as mean ± SEM from three independent experiments with technical triplicates. P-values were determined using One-way ANOVA with Bonferroni correction. No significant difference in RSA0-3h was observed among parasites selected from different rounds of 700 nM DHA treatment. Source data are provided as a Source Data file.
To address whether parasites at different stages post-invasion have comparable susceptibilities to DHA, we performed stage-specific survival assay (RSA0-3h, RSA9-12h, TSA18-21h) in which RS assays on highly synchronized PfM-KD parasites that were incubated with GlcN at schizont-early ring stages as described above and then treated with 6 h pulse of 700 nM DHA at 0–3, 9–12, or 18–21 h post-invasion (hpi). In parallel, a control experiment was carried out on the same parasite strain without the addition of GlcN. Results showed RS rates of 7.8% +/− 0.6 % for the 0–3 h stage, 2.4% +/− 0.6 % for the 9–12 h stage, and 0.6 % +/− 0.2 % for the 18–21 h stage, suggesting greatest effect of the PfMYST knockdown in the ring stage immediately after red blood cell invasion (Fig. 3d). Likewise, for parasites without the incubation of GlcN, the susceptibility varied similarly, i.e. early-stage parasites were less sensitive to DHA. Finally, we compared the outcomes of three sequential RSA0-3h assays on GlcN-treated PfM-KD parasites vs. A8 (K13 C580Y, not GlcN exposed) parasites. Results from these assays showed similar survival levels for both parasite populations (P > 0.05) (Fig. 3e), suggesting that RS may have pathways in common with features of the PfMYST knockdown phenotype.
Recrudescence and metabolomic features of DHA-treated PfMYST knockdown parasites
To assess the impact of PfMYST knockdown on parasites in DHA-treated cultures, we conducted in vitro recrudescence assays on parasite lines of this study with or without GlcN exposure of the starting populations. To address whether the PfK13 mutations could have an impact on the recrudescence profile, in vitro recrudescence assay was also performed on A5 and A8 and their respective revertant strains. Figure 4a presents the recrudescence curves from the these lines as well as PfM-KD and 3D7, thus providing a representation of different PfK13 mutations82. All DHA-treated flasks showed a rapid decline of parasitemia, with elimination of the bulk of the actively-replicating parasites within 24 hours. Recrudescence from persister parasites in the flasks then occurred roughly 3 weeks later, but with notable differences. The time until recrudescence was shortened by about 3 days with PfM-KD parasites that had been exposed to GlcN. In comparison, the time to recrudescence of the WT 3D7 parasites was not shortened after exposure to GlcN but was, in fact, lengthened by about 4 days (Fig. 4a). No faster recrudescence was observed with PfK13 mutation in either A8C580Y or 3D7-K13C580Y-transformed line.
a Recrudescence assay of the PfMYST-KD, WT 3D7, PfK13-modified 3D7 (K13_C580Y), A5 (K13_G538V), A5Rev (K13_G538), A8 (K13_C580Y) and A8Rev (K13_C580) lines. Assays of the 3D7 and PfMYST-KD lines were performed with (+) or withour (-) exposure of the starting populations to GlcN. Ring-stage parasites were exposed to three 6 h pulses at 700 nM DHA. Recrudescence curves were obtained based on parasitemia determined from Giemsa-stained smears. Data were presented as mean ± SEM from three independent experiments with technical duplicates. b Schematic workflow for the targeted metabolomics study. Elements were Created in BioRender (https://www.biorender.com/) with CC-BY 4.0 license (https://BioRender.com/e0ufn5x). (1) WT and PfMYST knockdown parasites (with GlcN treatment) were synchronized and exposed to 50 nM DHA or 0.1% DMSO (control) for 3 h/6 h. (2) After thorough washing, parasite pellets were swiftly quenched in dry ice, and the extracted metabolome was stored at − 80 °C. (3) High-resolution MS-based targeted metabolomics was conducted to analyze the alterations in energy metabolism. Two biological replicates were performed for each group. c Global analyses of metabolomic data revealed an alerted metabolic profile in parasites with either PfMYST-mediated or PfK13-associated resistance after DHA exposure. Highly synchronized parasites (0–3 hpi) were pre-treated with GlcN for 24 h and then subjected to a 3 or 6 h DHA pulse at a dosage of 50 nM in two independent biological replicates. Extracted metabolome for each group was then subjected to targeted metabolic profiling in which 19 key compounds involved in tricarboxylic acid cycle (TCA cycle), glycolytic pathway, pentose phosphate pathway and corresponding cofactors were simultaneously detected to assess alterations in energy metabolism. Statistical comparisons between samples subjected to vehicle control were conducted using a two-tailed student’s t test and significantly alerted metabolome were highlighted. The first - or + is corresponding to the exposure of GlcN, and the second − or + is corresponding to the exposure of DHA. Source data are provided as a Source Data file.
After ART treatment and in the hours immediately afterward, Plasmodium parasites exhibit mitochondrial swelling along with dramatic changes in the rough endoplasmic reticulum, nuclear envelope, and parasitophorous vacuole83,84. These changes are features of the great number of parasites that die and rapidly clear from the culture within the first day. Small numbers of the parasites that become persisters also show mitochondrial enlargement, with the movement of the mitochondria into closer association with the nucleus indicative of stress response and retrograde signaling50,85. To gain deeper insights into the parasite condition from DHA exposure, we conducted a targeted metabolomics analysis with a specific focus on central carbon metabolism, a vital component in parasite development, and compared the global metabolic profile of DHA-treated PfMSYT-KD and PfK13 mutated parasites. In brief, highly synchronized ring-stage (0–3 hpi) parasites were exposed to 50 nM DHA for 3 or 6 h, the metabolites extracted from each group were subjected to targeted metabolic profiling, simultaneously detecting 19 key compounds involved in the tricarboxylic acid cycle (TCA cycle), the glycolytic pathway, the pentose phosphate pathway, and their corresponding cofactors using ultra-performance liquid chromatography-tandem mass spectrometry (Fig. 4b).
Considering the DHA-induced growth arrestment was mainly responsible for the enhanced survival, we aim to investigate the similarities and dissimilarities in the metabolic profile of PfM-KD and PfK13 mutant parasites subjected to both 3 and 6 h exposure. Targeted metabolomics has revealed that following DHA exposure, metabolic pathways associated with energy metabolism were disrupted, and the integrated peak area for each sample was listed in Supplementary Data 5. The global metabolomic profile (19 metabolites) from parasites of the RS resistance phenotype exposed to DHA was distinct from that of the control at 3 and 6 h (Fig. 4c, Fig. S9). Both PfM-KD parasite lines and PfK13 mutants exhibited a diminished profile in energy metabolism, consistent with entry into the metabolically quiescent state of dormancy. Sample-to-sample correlation of the metabolite profiles showed that either PfM-KD samples or PfK13 mutant samples clustered separately from WT controls, indicating substantial changes in parasite metabolism throughout the IDC after DHA exposure. Besides, the overall trend toward lower levels of energy-related metabolites was accompanied by the arrested phenotype, and the alteration in metabolite pools was not a delayed response but was a consistent change that became more apparent during the DHA exposure. The most notable alterations included increased levels of glycolytic and pentose phosphate pathway intermediates for PfMYST-regulated resistance, whereas increased levels of TCA cycle intermediates were observed in PfK13 mutants. It has been reported that P. falciparum absorbs a significant amount of glucose during the IDC and relies heavily on glycolysis to provide energy for rapid growth and proliferation, resulting in an increase in the production of pyruvic acid and lactic acid86.
It is worth highlighting that glucose, when phosphorylated irreversibly into glucose-6-phosphate in the cytosol, exhibited significantly higher levels in the DHA-treated PfMYST knockdown parasites, while downstream metabolite fructose 1,6-bisphosphate levels decreased, potentially contributing to reduced glycolysis. Given that glucose-6-phosphate can be catabolized into by either the glycolysis pathway or the pentose phosphate pathway, the observed alert in the pentose phosphate pathway may reflect an altered redox balance, possibly related to the generation of reactive oxygen species induced by DHA. However, several intermediates of glycolysis and the pentose phosphate pathway were suppressed for PfK13 mutant parasites, and DHA exposure also affected certain metabolic pathways in a complex manner that was not confined to decreased intermediates. For example, most of the TCA metabolism intermediates elevated toward the end of the time course, and the elevation was concomitant with suppressed levels of compounds involved in glycolysis. Contrarily, the glycolytic intermediates were elevated in PfM-KD parasites, whereas other intermediates of the same pathway were suppressed for PfK13 mutants. Thus, the quiescent state induced at the young ring stage by DHA in PfM-KD and PfK13 mutant parasites likely seems to involve the same energy-associated metabolic mechanisms, but the PfK13 mutation shifts the intra-population distribution to the TCA cycle instead of glycolysis.
In summary, the suppression of these pathways in DHA-induced quiescence is consistent with the morphological disruption in which prolonged ring-stage as well as growth arrestment was observed. Distinct metabolic profiles of the PfK13 mutant vs. PfMYST mediated RS phenotypes suggest differential involvement of the glycolytic and TCA pathways in these parasites. The molecular mechanisms underlying these shifts of metabolism and their role in dormancy are subjects for future investigation.
Single-cell RNA sequence reveals genes under PfMYST regulation in artemisinin-treated P. falciparum
Given the robust RS phenotype of altered ART susceptibility associated with PfMYST knockdown, we next aimed to elucidate the genomic targets of PfMYST regulation by single-cell RNA sequencing experiments. Single-cell transcriptomes were generated at 0, 6, and 9 hpi under 200 nM DHA treatment for WT and PfM-KD parasites treated with GlcN or DMSO control (Fig. 5a). After filtering out low-quality transcriptomes, a total of 78,262 high-quality transcriptomes remained, with transcripts from an average of 265 genes per cell being detected. The number of expressed genes or mRNAs increases substantially from 0 h to 6 h, corresponding to the progression in intra-erythrogenic development (Supplementary Fig. 10a, b and Supplementary Data 6, 7). Marker gene expression in each sample consistently aligned with the corresponding time of parasite collection and treatment. Early ring-stage marker genes were highly expressed in the 0 and 6 h samples, whereas late ring-stage marker genes were predominantly expressed in the 9 h samples. Trophozoite- and schizont-specific marker genes were infrequently detected. Notably, ART-responsive genes were identified in DHA-treated parasites (Supplementary Fig. 10c). Results from uniform manifold approximation and projection (UMAP) and pseudotime trajectory analysis revealed that the cells self-organized along a developmental trajectory (Fig. 5b), segregating into two distinct directions as determined by DHA treatment (Supplementary Fig. 11a). Pseudotime analysis quantitatively demonstrated that PfM-KD parasites exhibited delayed development compared to WT controls, both in the presence and absence of DHA treatment (Supplementary Fig. 11b). This developmental delay was evidenced by consistently lower pseudotime values in PfM-KD parasites following GlcN-induced knockdown, indicating these parasites remained closer to their initial state at 0 h. This observation provides the molecular basis for the previously observed growth arrest in early ring-stage parasites with PfMYST knockdown (Fig. 3a).
a Experimental design for single-cell RNA sequencing. Parasites with various genetic backgrounds, including WT3D7, PfMYST knockdown, PfK13-mutated 3D7 (Y493H), 3D7 (C580Y), A5 (G538V), A8 (C580Y), and A8Rev (C580Y reversion) strains, were treated with or without DHA (200 nM), and RNA was harvested at distinct time intervals. Elements were Created in BioRender (https://www.biorender.com/) with CC-BY 4.0 license (https://BioRender.com/k7aqvkw). GlcN: glucosamine. b UMAP visualization of PfMYST-associated scRNA-seq samples, colored by sample origin. 0 h samples predominantly cluster in the upper region, DHA-treated 6- and 9 h samples cluster to the right, and untreated 6- and 9 h samples cluster to the left. c UMAP plot of PfMYST knockdown and DHA treatment samples, revealing 12 distinct clusters. Clusters 6 and 12 identified as DHA target clusters. d Stacked bar chart depicting the percentage of each cell cluster in different samples. e Normalized cluster ratios between PfMYST knockdown (PfM-KD (+/+)_6h) and control parasites (PfM-KD (-/+)_6h) samples following the 6-hour DHA treatment. Higher cluster ratios observed in clusters 6 and 12 for PfM-KD ( + / + )_6. f Venn diagram showing differentially expressed genes (DEGs) between the target cluster and other clusters in PfMYST knockdown parasites (PfM-KD (+/+)_6h) and control parasites (PfM-KD (-/+)_6h) after the 6 h DHA treatment. The 46 DEGs unique to PfM-KD (+/+)_6h are identified as “PfMYST target genes” potentially contributing to prolonged ring-stage in PfMYST knockdown parasites. g Venn diagram showing overlap between “PfMYST target genes” and three PfK13-associated target gene sets. h Characterization of 21 PfMYST target genes overlapping with PfK13-associated targets, showing: gene IDs (first panel), gene annotations (second panel), fold change in expression between target cluster and other clusters for each sample (third and fifth panels), and fold change of ChIP signals after PfMYST knockdown for each histone modification in ring stage (fourth panel). i Gene ontology analysis of “PfMYST target genes” that are either PfMYST-specific or shared with PfK13-associated target genes. P-values were calculated using Fisher’s exact test, with Benjamini-Hochberg-adjusted and Bonferroni-adjusted P-values provided in Supplementary Data 11 and Supplementary Data 14. Source data are provided as a Source Data file.
To identify the genes that contribute to the RS phenotype in PfM-KD parasites, we initially categorized the cell populations based on the whole transcriptomic profiles, which aligned generally with the timing of drug treatment (0 h: parasites before the DHA treatment, 6 h, parasites after 6 h DHA treatment, and 9 h: 3 h after removing DHA treatment). We hypothesized that the DHA-treated PfMYST knockdown parasite [PfM-KD (+/+)_6h], which aligns with the RSA experiment timing and exposure to DHA, would contain a larger cluster of surviving cells compared to the DHA-treated but GlcN-untreated PfM-KD parasites parasite [PfM-KD (-/+)_6h]. To find these surviving cells, all cells were segregated into 12 clusters (Fig. 5c, Supplementary Data 8). The proportions of each cluster were calculated and compared between two samples, namely the [PfM-KD (+/+)_6h] and control parasites without GlcN treatment [PfM-KD (-/+)_6h] (Fig. 5d, e). Notably, PfM-KD (+/+)_6h exhibited a higher proportion of cells in clusters 6 and 12 compared to PfM-KD (-/+)_6h (P = 1.737763e-26, Chi-squared test). Moreover, the proportions of clusters 6 and 12 were higher in PfMYST knockdown samples than in natural samples at 6 h, both with or without DHA treatment, as well as samples at 9 h with DHA treatment. (Supplementary Fig. 11c). Therefore, clusters 6, and 12 were defined as “target clusters” from the ring stages, which are suspected to survive the DHA drug pulse (Fig. 5e and Supplementary Fig. 11d). Intriguingly, cells within the “target clusters” appeared to maintain a state similar to 0 h, with the greatest percentage observed from 0-hour cells, and smaller percentages in the 6 h and 9 h samples after PfMYST knockdown, consistent with the earlier observation that PfMYST knockdown delays parasite development (Supplementary Fig. 11e).
To uncover the potential genes involved in the RS phenotype, we further performed DEGs comparing the “target clusters” with other clusters for both knockdown [PfM-KD (+/+)_6h] and control parasite [PfM-KD (−/+)_6h]. DEGs listed in Supplementary Data 9 represent the intersection of DEGs identified by two independent software: Seurat and Monocle. In Seurat, differential expression was determined using the Wilcoxon Rank Sum test. P-value adjustment was performed using the Bonferroni correction method, based on the total number of genes tested in the dataset. In Monocle, DEGs were identified using a generalized linear model assuming a negative binomial distribution (“negbinomial” family), and the resulting q-values (adjusted P-values) reflect false discovery rate (FDR) correction. The results showed that the majority of the DEGs (23 out of 37) in PfM-KD (−/+)_6h were contained in the DEGs (total of 69) of PfM-KD (+/+)_6h, indicating that these common DEGs were likely influenced by DHA exposure in both samples (Fig. 5f and Supplementary Data 9). We, therefore, speculate that the 46 DEGs unique to PfM-KD (+/+)_6h are correlated with PfMYST knockdown and underlie the RS phenotype; we refer to these as “PfMYST target genes” (Supplementary Fig. 12a and Supplementary Data 10). In the 6-hour samples subjected solely to DHA treatment or PfMYST knockdown, we observed similar expression differences for “PfMYST target genes” between the “target clusters” and other clusters, although most of these differences did not attain statistical significance (Fig. 5h).
Previous studies have linked PfMYST to the acetylation of H4K5, H4K8, H4K12, and H4K16, yet the specific genomic loci for these modifications remained elusive72. To address this issue, through CRISPR-Cas9 tagging, we generated a PfMYST-HA-GFP transgenic line for ChIP-seq assays. The genome-wide distribution profiles of PfMYST itself and PfMYST-mediated H4 acetylation in P. falciparum are described in Supplementary Text and Supplementary Figs. 12–15. Our data revealed significantly reduced H4K5ac and H4K8ac levels upon PfMYST knockdown in the ring stage. This decrease was prominent in gene promoter regions, impacting pathways critical for ring-stage functions, such as endocytosis, DNA transcription, ribosomal biogenesis, and localization in the cell (Supplementary Fig. 15a and Supplementary Fig. 16d). As expected, upon PfMYST knockdown, the acetylation modification level decreased in most “PfMYST target genes” associated with RS, with a more pronounced decrease in the ring stage compared to the schizont stage (Fig. 5h and Supplementary Fig. 12b, c). This suggests that PfMYST knockdown likely exerts a greater impact on “PfMYST target genes” in the rings than schizonts.
scRNA-seq uncovers the interplay between RS effects resulting from PfMYST knockdown and PfK13 mutation
Previous research has identified the parasite’s Kelch13 gene (PfK13) as a genetic determinant of the RS phenotype23,37,87,88. Several PfK13 mutations, such as Y493H, G538V, and C580Y, have been shown to confer significant RS levels up to 10%. To explore the interplay between RS effects resulting from PfMYST knockdown and PfK13 mutation, we generated PfK13Y493H and PfK13C580Y mutant parasites from the laboratory 3D7 strain using CRISPR/Cas9 site-directed gene editing. We then collected single-cell transcriptomes from 3D7, PfK13Y493H, PfK13C580Y line, and the A5 line (PfK13G538V) from a field isolate, employing methods similar to those described earlier. Since significant gene expression differences exist between laboratory strains and field strains, we conducted separate analyses for these two groups. Following stringent quality control, we obtained a total of 36,148 cells from laboratory strains, with an average detection of 249 genes per cell, and 22,747 cells from the A5 strain with an average detection of 320 genes per cell (Supplementary Fig. 17).
In the case of laboratory strain, cells were primarily grouped based on time of collection and clustered into 8 groups, with 0 h and 6 h samples distinguished (Supplementary Fig. 18a, b). Notably, the percentage of cells within each cluster and their distribution did not show a significant difference in the WT 3D7 strain with GlcN and DHA treatment, compared to the WT (-/+)_6h and WT (+/+)_6h samples (Supplementary Fig. 18c, d). Furthermore, fewer than 200 DEGs (Supplementary Data 9) were identified between WT (−/+)_6h and WT (+/+)_6h samples, a much smaller number compared to the impact of GlcN and DHA treatment on WT 3D7 parasites. Collectively, these results indicate that GlcN treatment has minimal effects on the WT 3D7 clone.
A similar approach was employed to characterize surviving DHA-treated cells within PfK13 mutant samples. Cluster 6 showed a higher proportion in PfK13 mutant samples than in 3D7 samples (P = 3.301106e-214 for C580Y mutant, and 3.239777e-65 for Y493H mutant) and was consequently designated as the “target cluster” (Supplementary Fig. 18e, f). It has been previously reported that parasites with inactivated PfK13 or a mutated PfK13 reduce hemoglobin endocytosis, thereby limiting ART activation and promoting higher RS levels37. We found that PfK13-associated endocytosis genes displayed lower expression levels in the “target cluster” compared to other clusters in PfK13 mutant samples, providing further evidence that cluster 6 represents mainly the surviving cells (Supplementary Fig. 18g). A similar trend was observed in PfM-KD (+/+)_6h (Supplementary Fig. 18h). Several DEGs were shared among PfK13 mutant samples, which show consistent expression trends between the “target cluster” and other clusters. These genes with consistent trends were defined as “PfK13Y493H/C580Y target genes” for RS > 1% (Supplementary Fig. 18i, j).
To identify the potential “RS target genes” in K13G538V parasites, we performed the same analysis using the obtained single-cell transcriptomic data. These samples were divided into three clusters, roughly corresponding to the parasite developmental stage and drug treatment conditions (Supplementary Fig. 19a, b). Consistent with the previous observations, the presence or absence of DHA treatment determines the developmental trajectory of the parasite (Supplementary Fig. 19c). Interestingly, ~ 90% of cells of no DHA treatment parasites in PfK13G538V(-)_6h fell into cluster 3, which may represent a natural development state (Supplementary Fig. 19d, e). Surprisingly, around 13% of PfK13G538V(+)_6h cells were also located in cluster 3, suggesting that these cells continued to develop normally despite the ART pressure, although their pseudotime was smaller than that of PfK13G538V(-)_6h cells within cluster 3 (Supplementary Fig. 19f). Subsequently, we divided the cells of PfK13G538V(+)_6h into the “target cluster” and other clusters based on their presence in cluster 3 (Supplementary Fig. 19g). Similar to PfK13Y493H/C580Y results, the expression of PfK13-associated endocytosis genes was relatively lower in the “target cluster”, (Supplementary Fig. 19h). The DEGs between the “target cluster” and other clusters in PfK13G538V(+)_6h were defined as “PfK13G538V target genes”. Both sets of PfK13-associated target genes suggested that ATP metabolism, endocytosis, and response to stress were closely associated with RS (Supplementary Fig. 18j and Supplementary Fig. 19i).
To extend our findings, we supplemented our dataset with single-cell transcriptomes from the A8 strain (PfK13C580Y) and A8Rev (PfK13C580 reversion mutant). Samples segregated into two distinct clusters based on drug treatment (Supplementary Fig. 20a, b). Untreated samples predominantly mapped to cluster 2, representing normal developmental status. Notably, 9% of cells from the drug-resistant A8(+)_6h samples also mapped to cluster 2, while the reversion mutant A8Rev (+)_6h, which has lost the RS phenotype with reversion to PfK13 C580, showed minimal representation in this cluster (Supplementary Fig. 19c, d). Importantly, A8(+)_6h cells that mapped to cluster 2 exhibited downregulation of Kelch13-associated endocytosis genes (Supplementary Fig. 20e). These observations suggested that A8(+)_6h cells in cluster 2 likely represent the resistant subpopulation, which we subsequently defined as the “target cluster” (Supplementary Fig. 15b). DEGs between this “target cluster” and other clusters in A8(+)_6h were classified as “A8 target genes.”
To distinguish between epigenetically-mediated (PfMYST) and genetically-mediated (PfK13 mutant) drug resistance mechanisms, we compared the identified target gene sets from different samples (Fig. 5g). 25 PfMYST-specific target genes were enriched in pathways related to cell cycle regulation and host-parasite interactions, representing distinctive features of PfMYST-mediated resistance (Fig. 5i and Supplementary Data 11). Notably, ~ 46% of “PfMYST target genes” overlapped with PfK13-associated target genes, with 9 genes shared across all target gene sets. These common putative resistance genes exhibited similar expression patterns within the “target cluster” across different samples (Fig. 5h). Gene Ontology analysis revealed that beyond classical resistance-associated pathways such as apoptosis regulation, protein stability, and stress response, both epigenetic and genetic factors relied on histone acetylation regulation to mediate the RS phenotype, further emphasizing the critical role of epigenetic mechanisms in antimalarial resistance.
Functional validation of potential resistance genes in DHA response and the effects of PfMYST on susceptibility to other antimalarials
To functionally validate the potential resistance genes identified by our single-cell transcriptome analysis, we conducted genetic manipulation experiments using CRISPR/cas9 technology (Fig. 5g). Due to the gene editing efficiency as well as functional complexity for above-identified potential resistance genes, we successfully overexpressed two genes (PF3D7_1010300 and PF3D7_1102700) in the 3D7 strain and knocked down or knocked out 8 genes coding for the knob-associated histidine-rich protein (KAHRP, PF3D7_0202000), proliferation-associated protein 2g4 (PF3D7_1428300), ATP-dependent RNA helicase DDX5 (PF3D7_1419100), Plasmodium exported protein PHISTb (PF3D7_1477500), putative large subunit GTPase 1 (PF3D7_1431300), Plasmodium exported protein PHISTa (PF3D7_1372000), Plasmodium exported protein hyp9 (PF3D7_0220600), and conserved Plasmodium protein (PF3D7_0915600) (Fig. 6a). The transgenic parasite lines were validated by diagnostic PCR with primers listed in Supplementary Data 2 and the level of corresponding proteins were measured by either western blot or qPCR (Supplementary Fig. 21). RSA0-3h was performed to assess the impact of these genes on RS levels. As shown in Fig. 6c, d, no discernible effect on RS was found with the overexpression of two genes, while significantly increased RS occurred with the knockdown of six genes, including KAHRP, 2G4, DDX5, PHISTa/b and GTPase 1. Importantly, in vitro recrudescence was shortened by 2–4 days for two transgenic lines, including with knocked-down expression of Plasmodium exported protein PHISTb (PF3D7_1477500) and proliferation-associated protein 2g4 (PF3D7_1428300), but not others (Fig. 6b). The KAHRP, was consistent with previous findings from ART selection experiment results.
a Ten potential resistant genes in response to DHA treatment are functional validated by episomal over-expression (OE), knockout (KO) or knockdown (KD). The overall results of RSA0-3h assays of these transgenic lines are summarized in the last column. b Recrudescence Assay of PHISTb_KO, 2G4_KO, PHISTa_KO, DDX5_KO with the parent 3D7 line of transfection as control. Ring-stage parasites were exposed to three 6 h pulses at 700 nM DHA. Recrudescence curves were obtained based on parasitemia determined from Giemsa-stained smears. Data were presented as mean ± SEM from three independent experiments with technical duplicates. c,d RSA0-3h for over-expression (OE), knockout (KO) or knockdown (KD) of “PfMYST-targeted genes”. Two OE, four KO and four KD lines were tested for their RSA0-3h under 700 nM DHA treatment with WT 3D7 strain as a negative control. Data were presented as mean ± SEM from four independent experiments with technical duplicates. Statistical comparisons between vehicle control and GlcN-treated samples were conducted using a two-tailed student’s t test. Statistical comparisons between WT and mutant parasites were conducted using One-way ANOVA with Bonferroni correction, and the exact P-value was provided in Source Data. (*P < 0.05; ** P < 0.01; *** P < 0.001). e The impact of PfMYST knockdown on RSA0-3h for seven antimalarial drugs, including AQ, PYR, CQ, PYA, LMF, MEF, and PPQ. RSA0-3h was modified according to the IC50 of each drug, with drug concentrations set at 10 x IC50. Survival rates were calculated using the standard method. Data were presented as mean ± SEM from four independent experiments with technical duplicates. Statistical comparisons between vehicle control and GlcN-treated samples were conducted using a two-tailed student’s ttest. AQ: Amodiaquine, PYR: Pyrimethamine, CQ: Chloroquine, PYA: Pyronaridine, LMF: Lumefantrine, MEF: Mefloquine, PPQ: Piperaquine. f Relationship between the alerted resistance level and pharmacokinetic parameters, including drug half-life and Tmax. Drugs with increased/decreased resistance were displayed in red/green circles, respectively. Source data are provided as a Source Data file.
Given the variable expression of Pfmyst within the natural parasite population and its broad target gene family across different stages, we hypothesized that its effects, i.e., extending ring stage and shortening trophozoite-schizont stages, on RS levels might be applicable to other antimalarial drugs, particularly those with less susceptibility in rings and relatively shorter half-life in vivo. To investigate this, we tested the RSA0-3h of 7 additional antimalaria drugs, most of them act on the trophozoite stage, using the PfM-KD parasite. As shown in Fig. 6e, f, PfMYST knockdown significantly enhanced the survival for four drugs, including Amodiaquine (AQ), Pyrimethamine (PYR; 3 daily doses), Chloroquine (CQ), and Pyronaridine (PYA), but not for Lumefantrine (LMF), Mefloquine (MEF), and Piperaquine (PPQ). Similar to DHA, no IC50 changes for these drugs could be detected between PfMYST knockdown and WT parasites (Supplementary Fig. 8 and Supplementary Fig. 22). It is worth noting that RSA0-3h levels for MEF in the PfM-KD parasites without GlcN treatment increased, suggesting a potential regulatory role for the expression level of PfMYST in RSA0-3h outcomes. Optimization of the RSA protocol based on drug IC50 or half-life is warranted to further evaluate parasite responses to these drugs. Nevertheless, these findings raise concern about epigenetically-driven drug resistance and dormant forms of pathogens as reported in bacteria (Fig. 7)89.
Various levels of Pfmyst expression exist in the natural P. falciparum parasite population, affecting the expression of genes through epigenetic regulation. After exposure to DHA, some parasites in the targeted cluster with low Pfmyst expression display altered cell cycle timing and metabolic quiescence, promoting ring-stage survival and recrudescence after a period of dormancy. Elements were Created in BioRender (https://www.biorender.com/) with CC-BY 4.0 license (https://BioRender.com/o6aevoo.).
Discussion
Drug treatment failures due to ART resistance pose a major challenge for malaria control. Since both the PfK13-dependent and independent mechanism has been reported, ART resistance could be a complex and cumulative trait which is influenced by multiple factors including genetic factors, epitranscriptomic modifications, tRNA modifications etc53,54,55. Thus, these findings have implicated that there may not be a single universal mechanism of ART resistance which contribute to a combination of the above-mentioned factors that collectively regulate gene expression and modulate distinct phenotypic responses to the drug. In this study, we have revealed that epigenetic factor (PfMYST) has been implicated in the regulation for parasite growth. Based on the distinct RS phonotype resulted from the regulation of PfMYST of P. falciparum, we have focused on uncovering the epigenetic regulatory mechanism of P. falciparum RS after pulses of ART and other antimalarial drugs due to growth arrestment.
To unravel the unique mechanisms behind the RS phenotype, we investigated the expression patterns of histone acetyltransferases (HATs) in Plasmodium parasites. HATs serve as pivotal regulators of chromatin modification and gene expression, influencing critical processes like cell cycle progression, antigenic variation, and immune evasion90. Our study revealed variable gene expression profiles among well-synchronized field isolates and cloned parasite lines, with one standout, PfMYST, exhibiting a compelling link to ring survival assays. Notably, it has been reported that PfMYST expression was positively associated with DHA resistance due to enhanced DNA repairment which was opposite to the conclusion in our study58. Recent evidence indicates that gene expression fluctuates dynamically over the period of dormancy from induction to recrudescence7,91,92,93. Therefore, continuous modulation of Pfmyst expression as performed in previous studies would likely impact the expression of various parasite genes differently across the range of time points through dormancy and recrudescence, potentially influencing the overall outcome of parasite survival58,94. In contrast, in our system (CRISPR/Cas9-glmS), GlcN was applied for only 6 h prior to artemisinin exposure, with no further modulation of Pfmyst expression during the recovery period. This approach more closely reflects the natural scenario, where parasites are transiently exposed to artemisinin without ongoing modulation of gene expression during their recovery. Meanwhile, studies have also validated that there was no direct correlation between the resistance and RS phenotype21,49. Thus, the inconsistency between our results and previous studies could be explained due to the above-mentioned reasons. To further validate the PfMYST-mediated RS phenotype, we also employed an independent knockdown system (termed knocksideways) targeting PfMYST and PfGCN5. A negative correlation between Pfmyst expression and RSA0-3h levels was again observed (Supplementary Fig. 23), consistent with the findings from our CRISPR/Cas9-glmS approach, thus further supporting our hypothesis that the discrepancy can be attributed to the different KD strategy in which continuous knockdown differently affected the state of the parasite.
Herein, in our study, all mechanistic analyses were conducted in early ring-stage parasites, thus, we further hypothesized that Pfmyst transcript levels fluctuate dynamically in response to ART exposure. To investigate this, we measured Pfmyst transcript levels at multiple time points (0 h, 6 h, 9 h, 12 h, and 18 h post-DHA exposure) using RT-qPCR in 3D7, A8/A8Rev, and A5/A5Rev parasite strains. Interestingly, we observed significant temporal fluctuations in Pfmyst transcript post-ART exposure, suggesting a dynamic regulatory response rather than a static expression pattern (Supplementary Fig. 24). This is further evidence that the discrepancy with previous studies arises primarily from differences in the timing of gene expression measurements and the specific recovery phases being examined.
PfMYST, in particular, demonstrated a precise targeting of histone modifications, including H4K5ac and H4K8ac, pivotal for pathways governing the parasite’s life cycle, metabolic activities, ring survival, and recrudescence. Notably, the knockdown of PfMYST exerted a significant influence across multiple pathways, including cell cycle arrest, ribosomal biogenesis, and endocytosis, vital for parasite functions, resulting in the delay of parasite development and retention in the ring stage. Intriguingly, these effects seemed to interplay with various facets of ART’s mechanisms of action. ART primarily relies on heme activation sourced mainly from hemoglobin digestion within the food vacuole95. Once activated, ART exhibits a broad reactivity with cellular targets, disrupting the homeostasis of cellular proteins87,96,97.
PfMYST knockdown affects the heme catabolic process and may reduce heme production, resulting in lower ART activation and decreased cellular stress. Furthermore, both PfMYST knockdown and PfK13 mutant parasites displayed altered energy metabolism profiles, particularly in the TCA cycle78,81. The primary goal of growth-arrested parasites is to maintain the basic supply of energy and sustain active regulatory mechanisms for sensing and responding to environmental stress. During the arrested state with reduced rates of both anabolic and catabolic metabolism, parasites were forced to rely on either an alternative source of energy supply or metabolic pathways re-routing. In addition, the metabolic profile of glycolysis and the pentose phosphate pathway were also differentially altered in PfMYST knockdown parasites, in which was opposite to that of PfK13 mutants. Glucose-6-phosphate levels increased, while fructose 1,6-bisphosphate levels decreased, potentially contributing to reduced glycolysis. This may help maintain redox balance against DHA-induced reactive oxygen species. We linked this phenomenon to the growth arrestment, during which the parasite is able to decrease its metabolic rate dramatically to conserve energy and resources, ultimately leading to increased survival in stress conditions. We propose that P. falciparum adjusts its metabolic growth accordingly, allowing it to survive and persist. This response appears to be specific to either PfMYST or PfK13-regulated resistance. Thus, the quiescent state induced at the young ring stage by DHA in PfMYST and PfK13 parasites likely seems to involve the same energy-associated metabolic mechanisms, but the PfK13 mutation shifts the intra-population distribution to the TCA cycle instead of glycolysis. We conclude that the biological mechanisms for either PfMYST or PfK13 regulated resistance may differ, at least in the metabolism of glycolysis and the TCA cycle. Nevertheless, it would be of great interest to further investigate the impact of PfMYST-specific regulation on the glycolytic activity of intraerythrocytic parasites by measuring the extracellular acidification rate to validate the significantly attenuated glycolysis process and glycolysis capacity in a time-dependent manner. Although we have demonstrated that measuring the steady-state metabolome pools of parasites received DHA exposure is capable to reveal the perturbed pathways, while how the flux through each pathway causes the metabolic alterations in steady-state metabolite pools remains to be further investigated. Thus, a stable-isotope labeling strategy could be a powerful alternative to overcome this limitation and provide a detailed pattern for how the flux through these corresponding metabolic pathways causes the elevation and suppression of intermediates in metabolic pathways. In summary, DHA-exposed ring-stage PfMYST knockdown parasites and PfK13 mutant parasites experienced significant reductions in energy metabolism, and adaptation to these stress-associated conditions was accompanied by a dramatic rewiring of metabolism, from a highly glycolytic metabolism to increased dependence on TCA metabolism.
Single-cell transcriptomic analyses have provided invaluable insights into the response of PfMYST knockdown parasites to ART (DHA) treatment. This approach has not only illuminated specific cellular clusters that react to DHA but has also shed light on the intricate dynamics of drug responses at the cellular level. These findings align with previous observations in clinical patients, reaffirming the clinical relevance of the study. Comparable expression profile changes have been observed in clinical patients, consistent with previous findings47,98,99. Remarkably, the overlap between approximately 40% of “PfMYST target genes” and those associated with PfK13 highlights a shared molecular basis in the RS phenotype. In addition, the knob-associated histidine-rich protein, a top-hit gene, was also detected in previous studies of ART-selected parasites51. Its presence in those parasites suggests a possible role in the RS phenotype and survival strategies or stress responses of the parasites.
It is noteworthy that epigenetically regulated mechanisms of the RS phenotype affect parasite responses to other antimalarial drugs, including several partner drugs of ART. In the present study, PfMYST knockdown parasites exhibited altered RS rates in response to five out of seven malaria drugs tested, with the exception of PPQ and LMF; however, we note that the specific protocol employed may influence these outcomes. For instance, the PPQ survival assay (PSA) was designed to assess parasite survival rates following exposure to a pharmacologically relevant dose of PPQ (200 nM) for 48 hours100,101. Notably, in vitro assays of PPQ-resistant parasites have displayed a bimodal dose-response curve, featuring a second peak at high drug concentrations (0.1–10 μM), suggesting that the area under the curve for this second peak could serve as an alternative indicator of PPQ resistance102. Refining the drug treatment timing and duration within the protocol may provide further insights into the precise effects of PfMYST on the susceptibility of malaria parasites to various antimalarial drugs.
Extensive efforts have been dedicated to unraveling the intricate mechanisms underlying ring-stage survival after ART treatment. These mechanisms may involve diverse processes such as mitochondrial dysfunction, oxidative stress, DNA damage, proteasome disruption, proteostasis disturbance, and hindrance of hemoglobin digestion by the parasite38,50,103,104,105. In addition to certain mutations of PfK13, mutations in genes like coronin, pcubp1, pcap243,45, AP-2μ (adapter protein-2 μ subunit,106, UBP-1 (ubiquitin-binding protein-1,107, and falcipain 2a40,41 have been shown to influence RS by altering hemoglobin uptake and digestion processes. Transcriptional changes in genes related to redox regulation, heat shock response, proteasome activity, DNA processes, metabolism, and endocytosis also contribute to the complexity of ART response99,108, reflecting the interplay between genetic factors, transcriptional responses, and metabolic activities. However, RS phenotypes of these parasites often diminish rapidly upon removal of ART pressure, suggesting potential epigenetic rather than genetic origins23,51,55, akin to the epigenetically regulated adaptive resistance observed in bacteria52. Here, our findings confirm a distinction between the RS phenotype affected by PfK13 mutations and the dormancy phenotype of persisters, which has been genetically dissociated from those mutations21,49. With the RS phenotype, a small fraction of the ring-stage population can survive a single ART pulse, whereas, with the dormancy phenotype, persisters can survive daily ART treatment courses of 3 days or more to regenerate a population of parasites with the same features as before6,7. These observations highlight the importance of understanding the underlying mechanisms.
In summary, the complexity of RS susceptibility in malaria involves a combination of genetic factors, transcriptional responses, metabolic alterations, and epigenetic regulation. Our research uncovered a PfMYST-mediated epigenetic mechanism that regulates RS phenotype in P. falciparum. Naturally, varying levels of Pfmyst expression exist in P. falciparum parasite populations, influencing the expression of numerous downstream genes involved in vital pathways, such as the heme catabolic process and oxidative phosphorylation regulation. Upon exposure to DHA, certain parasites with lower Pfmyst expression undergo modified parasite cell cycle and metabolism, influencing ring-stage survival and recrudescence in P. falciparum. Continued exploration into the genetic determinants, cellular pathways, and epigenetic modifications linked to RS, as well as dormancy, will contribute to the development of potential therapeutic strategies and the preservation of ART’s efficacy as a frontline antimalarial medication. The roles of transcriptional and epigenetic responses governing these phenotypes, as well as the signaling processes involved, remain to be further explored.
Methods
Ethics statements
This study was conducted in strict accordance with the guidelines for the application of biosamples from Jiangsu Institute of Parasitic Diseases (JIPD). The experimental protocol was reviewed and approved by the Research Ethics Committee of JIPD (IRB00004221).
Parasite culture, plasmid construction and transfection
P. falciparum strain 3D7-G7 and field strain A8/A5 were cultured in human red blood cells in culture medium (10.44 g/L RPMI-1640, 25 mM HEPES, 10% v/v Albumax I, 0.1 mM hypoxanthine, 20 μg/ml gentamicin) under conditions of 5% O2, 5% CO2, 90% N2 at 37 oC. Synchronization of parasites in the ring stage was achieved using a 5% sorbitol solution, while purification of schizont-stage parasites was carried out through a 40/70% Percoll discontinuous gradient.
In this study, we designed various plasmids to facilitate gene editing and overexpression experiments in 3D7-G7 strain for pfmyst gene edition and tagging, we constructed the pfmyst-ha-gfp, pfmyst-ty1-ribo, and ht-pfmyst plasmids based on the pL6cs-sgRNA. To functionally validate the PfMYST-regulated genes identified by our single-cell transcriptome analysis, we constructed the KO or KD plasmids based on the pL6cs-sgRNA. The pL6cs-sgRNA plasmid contains the sgRNA expression cassette, homologous arms, and a positive selection marker (hDHFR) resistant to WR99210. To create the corresponding pL6cs plasmids for specific gene editing, we amplified the homologous arms and target insertions using the primers provided in Supplementary Data 2, followed by ligation using the AscI and AflII restriction sites. In addition, the 20-bp guide sequences were cloned into the respective pL6cs-sgRNA plasmids using the AvrII and XhoI restriction sites. The expression of the SpCas9 endonuclease was achieved using the pUF1-Cas9 plasmid, which includes two nuclear location sequences (NLS), a 3 × FLAG tag, and a blasticidin S deaminase (BSD) resistance marker. For overexpression experiments, plasmids were generated based on the pLN plasmid backbone.
For transfection, 100 µg of each plasmid were electroporated into the fresh human-type O erythrocytes, which were resuspended in cytomix (120 mM KCl, 10 mM KH2PO4, 25 mM HEPES, 2 mM EGTA, 0.15 mM CaCl2, 5 mM MgCl2, pH 7.6), under standard electroporation conditions. Subsequently, the electroporated erythrocytes were mixed with enriched late schizonts and immediately transferred to the standard culture environment. For gene editing by the CRISPR/Cas system, pL6cs-sgRNA and pUF1-Cas9 plasmids were co-transfected. To support parasite growth and selection, we refreshed the media daily during the initial week and every second day thereafter. Positive selection drugs were introduced 72 h post-transfection, with the following working concentrations: 2.5 µg/ml for BSD (Gibco, R21001), 2.5 nM for WR99210 (Sigma, SML2976), and 1.5 µM for DSM1 (Millipore,5.33304). To confirm successful transfection and integration, we conducted PCR analysis and DNA sequencing. Parasites were harvested and lysed with 0.15% saponin for genome extraction with the TIANamp genomic DNA kit (DP304, Tiangen, China) as per the provided instructions. PCR products were purified using the TIANgel Midi Purification Kit (DP209, Tiangen, China) and subjected to Sanger sequencing. Detailed information about the primers used for these verifications can be found in Supplementary Data 1.
RT-qPCR analysis
To assess the baseline transcription levels of Pfmyst and other histone acetyltransferases in the obtained subclones, ring-stage parasite samples (~ 10 h) were collected without DHA exposure. The infected red blood cells (iRBCs) were pelleted, washed with 5 mL PBS, and then resuspended in 2 mL TRIzol reagent. The samples were incubated at 37 °C for 5 min. RNA was extracted using the Pure Total RNA Isolation Kit (Vazyme Biotech Co., Ltd), and 500 ng of total RNA was reverse transcribed into cDNA using HiScript Q RT Super Mix (Vazyme Biotech Co., Ltd) with an oligo(dT) primer. The reaction mixture was diluted to a final volume of 200 μL for qPCR analysis, following the protocol provided with the SYBR Green I kit (Vazyme Biotech Co., Ltd). Primers for each gene are listed in Supplementary Data 2, and seryl-tRNA synthetase was used as the internal reference gene. Gene expression levels were quantified using the 2−ΔCT method for normalization.
Whole-genome sequencing data analysis
Raw sequencing reads were quality-controlled using TrimGalore (v0.6.4) to remove adapter sequences and low-quality reads with default parameters. The high-quality reads were then aligned to the reference Plasmodium falciparum 3D7 genome (PlasmoDB v68) using the Burrows-Wheeler Aligner (BWA-mem v0.7.17) with default settings. The resulting alignments in SAM format were converted to position-sorted BAM files using SAMtools (v1.9).
Quality assessment revealed excellent sequencing depth across all samples, with average genome coverage exceeding 180X and more than 96% of genomic positions covered at ≥ 10X, confirming the high quality of our sequencing data. Variant calling was performed using bcftools mpileup followed by bcftools call to generate raw BCF files. These variants were subsequently filtered using bcftools view with stringent quality thresholds (read depth between 100–300 and QUAL score > 25).
Functional annotation of the identified variants was performed using ANNOVAR to determine their potential impact on protein-coding genes. To assess genetic relatedness between samples, we calculated pairwise identity-by-descent estimates (PI_HAT values) using PLINK (v1.9.0). These values, representing the probability of identical allele sharing across the genome, were visualized using R with the pheatmap package.
ChIP-seq and data analysis
ChIP-seq library preparation
The ChIP-seq assay was conducted following established protocols with some minor adjustments. Parasites were synchronized and harvested at different stages. To initiate cross-linking, 1%formaldehyde (Sigma, F8775) was immediately mixed with parasites and gently rotated for 10 min at 37 °C. The cross-link reaction was quenched with 0.125 M glycine for 5 min on ice. RBCs were lysised with 0.15% saponin for 10- 15 min on ice and then subjected to three washes. Next, the parasites were incubated with 2 ml of Lysis Buffer (10 mM HEPES pH 7.9, 10 mM KCl, 0.1 mM EDTA pH8.0, 0.1 mM EGTA pH8.0, 1 mM DTT, 0.25% NP40, 1 × protease inhibitors cocktail) for 30 min on ice, followed by dounce homogenization for 100 strokes. Following centrifugation, the precipitated nuclei were resuspended in 200 μl of SDS Lysis Buffer (containing 1% SDS, 10 mM EDTA, 50 mM Tris-HCl pH8.0). The chromatin was sonicated using the Covaris M220 Sonicator with the following settings: 5% duty factor, 200 cycles per burst, and 75 W of peak incident power for 20 min to yield chromatin fragments at 200–500 bp in size. The chromatin fragments were diluted tenfold in a dilution buffer (0.01% SDS, 1.1% Triton X-100, 1.2 mM EDTA, 16.7 mM Tris-HCl, pH 8.0, 150 mM NaCl, 1× protease inhibitors cocktail) and co-incubated with Protein A/G magnetic beads (Pierce) for two hours. The supernatant was then co-incubated with Protein A/G magnetic beads and the corresponding antibody at 4 °C overnight. The antibodies used included 1 μg rabbit anti-GFP (Abcam, ab290), 1 μg rabbit IgG (Abcam, ab171870), 3 μg anti-H3K9ac (Millipore, 07-352), anti-H3K9me3(Abcam, ab8898), and anti-H4K8ac (Millipore, 07-328). The beads were subjected to several washes: once at 4 °C for 5 min with Low Salt Immune Complex Wash Buffer (0.1% SDS, 1% Triton X-100, 2 mM EDTA, 20 mM Tris-HCl pH 8.0, 150 mM NaCl), High Salt Immune Complex Wash Buffer (0.1% SDS, 1% Triton X-100, 2 mM EDTA, 20 mM Tris-HCl pH 8.0, 500 mM NaCl) and LiCl Immune Complex Wash Buffer (0.25 M LiCl, 1% NP-40, 1% Deoxycholate, 1 mM EDTA, 10 mM Tris-HCl pH 8.0), followed by two washes at room temperature for 5 min each with TE Buffer (10 mM Tris-HCl pH 8.0, 1 mM EDTA pH 8.0). The target protein-DNA complexes were eluted with 200 μL fresh Elution Buffer (1% SDS, 0.1 M NaHCO3). These complexes were then incubated at 45 °C overnight, followed by incubation at 37 °C for 30 min with RNase A and at 45 °C for two hours with Proteinase K. ChIP-DNA was extracted using the MinElute PCR purification kit (Qiagen, 28006). In the ChIP library preparation process, the ChIP-DNA was first subjected to end repair using End-it DNA End-repair Kit (Epicenter No.ER81050), followed by the addition of 3’ A base using Klenow Fragment (3´ → 5´ exo-) kit (NEB No.M0212L). Subsequently, adapter ligation was performed with Quick Ligation Kit (NEB No.M2200L), and the libraries were size-selected and amplified using KAPA Biosystems KB2500. The amplification was carried out with a PCR program: including 1 min at 98 °C, 12 cycles of 10 s at 98 °C, and 1 min at 65 °C, followed by a final extension of 5 min at 65 °C. These amplified libraries were sequenced using an Illumina HiSeq Xten system, and each ChIP-seq assay included two biological replicates.
ChIP-seq data analysis
Initially, all the ChIP-seq reads underwent processing using TrimGalore (v0.6.4) to trim adapter sequences and eliminate low-quality reads. Subsequently, the adapter-trimmed reads were mapped to the P. falciparum 3D7 genome (PlasmoDB v68) using Bowtie2 (v2.3.5.1) with the following parameters: -t -N 0 -q --no-mixed --no-discordant. The mapping results in SAM format were then converted to position-sorted BAM format using Samtools (v1.9). To further refine the dataset, unmapped reads and those with mapping quality scores below 20 were removed using sambamba (v0.7.0). In addition, multiple mapped reads and PCR duplicates were eliminated utilizing a JavaScript tool (MarkDuplicates.jar) from Picard (v2.20.7). Subsequently, the BAM files were converted into bigwig files using bamCoverage from the deeptools suite (v3.3.0) with parameters: --normalizeUsing RPKM. To facilitate comparative analysis, the bigwig files derived from the ChIP sample were normalized to the input sample employing bigwigCompare from the deeptools suite (v3.3.0) with parameters: –operation ratio. Finally, the Integrative Genomics Viewer (IGV) was used to visualize the histone modification signal in specific genomic regions within a track view. Additional steps that contribute to the comprehensive understanding of histone modification and PfMYST binding patterns are detailed as follows:
-
(1)
Validation of ChIP-seq data reproducibility: The gene body regions (from ATG to stop codon) were extracted from the GTF file, sourced from PlasmoDB (release-68). These gene body regions were extended by 1000 bp in both upstream and downstream directions. The normalized ChIP signals within these extended gene body regions were generated using the deeptools suite multiBigwigSummary (v3.3.0). Pearson correlation and hierarchical clustering were employed to assess the reproducibility and correlation of the ChIP data. Following reproducibility validation, replicates were pooled for downstream analysis.
-
(2)
Global histone modification (HM) signals on each chromosome: Each chromosome was split into bins using makewindows from bedtools (v2.28.0) with parameter -w 3000. The normalized signal of HM ChIP-seq within each bin was calculated by multiBigwigSummary in BED mode, with the default parameters from the deeptools suite (v3.3.0). The normalized signal was plotted using geom_bar from ggplot2 in R.
-
(3)
Distribution of ChIP signals around genes’ 5’UTR: Genes’ 5’UTR or promoter was defined as regions spanning from 1500 bp upstream to 500 bp downstream of the ATG start codon. The normalized ChIP signals within 5’UTR regions were generated by the deeptools suite multiBigwigSummary (v3.3.0). A heatmap of ChIP signals in genes’ 5’UTR was generated using computeMatrix and plotHeatmap from the deeptools suite (v3.3.0). The input data files consisted of bigwig files of the ChIP signals normalized by the input sample.
-
(4)
Peak calling: The enrichment regions corresponding to histone modification and PfMYST binding were identified using the peak calling algorithm integrated into MACS (v2.1.2). Specific parameters were applied for peak calling, including --gsize 2.3e + 7 --format BAMPE -q 0.05 --keep-dup all. An additional parameter (--broad) was used for histone modification peak calling. The common peaks that exhibited overlap between biological replicates were retained for subsequent annotation. Peaks were annotated using a Python script (annotatePeaks.pl) from the HOMER suite (v4.11.1). To be considered as peaks-annotated genes, peaks had to meet two criteria: they were the nearest gene to the peak, and the distance between their TTS and the peak was < 3000 bp.
Single-cell RNA sequencing and data analysis
Parasites were synchronized using 5% sorbitol for two rounds, then pre-incubated with GlcN for 24 h. Young rings, with a ~ 3 h window, were harvested using the RSA0-3h protocol and subsequently exposed to 200 nM DHA, alongside vehicle-treated 0.05% DMSO controls, for the designated time points (0 h, 6 h, 9 h) as depicted in Fig. 5a. After a 6 h exposure, parasites were washed three times with complete medium to remove any residual DHA, then returned to normal culturing conditions prior to scRNA sequencing.
For cell sorting, synchronized parasites were stained with 2 μM Hoechst 33342 for 10 min at 37 °C, washed, and re-suspended in complete medium (without AlbuMAX) at 0.1% hematocrit. ~ 10,000 Hoechst 33342-positive cells were then sorted using a FACSAria II sorter (BD Biosciences) as previously described109,110,111. The sorted cells underwent two washes with 0.05% BSA/PBS, were re-suspended in 100 μL 0.05% BSA/PBS, and counted using a cell counting chamber. To confirm the sorted cells viability and counts, parasites were stained with 0.4% trypan blue and measurements were taken with an Automated Cell Counter (Thermo Fisher Scientific) before the scRNA library preparation and sequencing. Subsequently, libraries were prepared using the Chromium Single Cell 3ʹReagent Kits v3 (10x genomics) following the instructions provided in the user guide. Single-cell RNA seq data were obtained using the Illumina HiSeq Xten system.
ScRNA-seq data analysis
In the analysis of single-cell RNA-seq (ScRNA-seq) data, the following steps were undertaken:
-
(1)
Processing of raw single-cell RNA-seq data: Raw sequencing reads were assessed using quality metrics and analyzed using the Cell Ranger software (v7.1.0). The “count” command was utilized to create a filtered count matrix with the default parameters.
-
(2)
Quality control of single-cell RNA-seq data: To ensure the analysis was conducted on high-quality cells, a series of quality control measures were applied. Cells with low quality, characterized by unique detected genes less than 50, unique detected genes exceeding 400, or mitochondrial content surpassing 0.1%, were filtered out. For the remaining cells, additional outliers were removed. Cells that exhibited a median absolute deviation from the median for the total number of UMIs or unique detected genes twice the median were excluded. To address potential duplicates, the R package DoubletFinder (version 2.0.2) was employed. The mean-variance-normalized bimodality coefficient (BCMVN) was calculated for each sample to determine the neighborhood size (pK). The number of artificial doublets (pN) was set based on the cell number of each sample. The detection of doublets was excluded using the “doubletFinder_v3” function. Detailed information regarding the quality control for each sample is shown in Supplementary Data 8.
-
(3)
Single-cell RNA-seq data clustering and annotation:
Clustering analyses were performed separately for different sample groups (PfMYST samples, 3D7 and PfK13Y493H/C580Y mutation samples, and geographical strain samples). After filtering, the data were processed using the standard workflow implemented in the Monocle3 (v0.2.2) package. Briefly, datasets were analyzed with the following steps:
-
a)
Combining filtered count matrices: The filtered count matrices were combined across datasets using the “combine_cds” function with the default parameters.
-
b)
normalization, dimensionality reduction, and clustering: The “preprocess_cds” function was applied to normalize the data by log transformation and size factor adjustment, addressing variation in sequencing depth. A lower-dimensional space was calculated for subsequent dimensionality reduction. Dimensionality reduction was performed using the uniform manifold approximation and projection (UMAP) method via the ‘reduce_dimension’ function. Unsupervised clustering of cells was achieved using the Leiden community detection method through the “cluster_cells” function. All visualizations based on the UMAP were generated by the “plot_cell” function.
-
c)
Principal Trajectory Graph: Cells were fitted into a principal trajectory graph using the learn_graph function. The root of the trajectory, predominantly occupied by 0-hour cells, was programmatically selected as the starting point for erythrocytic development.
-
d)
Differential Expression Analysis: Differential expression analysis was performed using both Monocle3 and Seurat (v5.2.1). Monocle3 employed a regression model via the “fit_models” function, with subsequent adjustment of P-values using the Benjamini and Hochberg method by the “coefficient_table” function. Genes expressed in more than 500 cells with an adjusted P-value < 0.05 were retained. Seurat’s FindMarkers function, based on the Wilcoxon rank sum test, was used for differential expression analysis, with P-values adjusted using Bonferroni correction. Genes with an adjusted P-value < 0.05 were retained. Common differential expression genes obtained by both packages were used for further analysis.
-
e)
cell markers collection and cell cluster annotation: different parasite developmental stage markers and DHA-respond markers were collected from previously published literature for help in cell cluster annotations.
This comprehensive workflow allowed for the exploration and clustering of single-cell RNA-seq data, enabling the identification of differentially expressed genes within distinct sample groups.
Gene Ontology (GO) term enrichment analysis
GO term enrichment analysis was conducted for specific genes utilizing the Gene Ontology Enrichment tool available at PlasmoDB (https://plasmodb.org/). GO terms were considered statistically significant if they exhibited a P-value of ≤ 0.05. The results were visualized using the ggplot2 R package, aiding in the interpretation and presentation of enriched GO terms associated with the analyzed genes.
Gene set enrichment analysis
Gene set enrichment analysis was conducted using the “wilcoxauc” function from the R package presto (v1.0.0) to determine the average log2 fold change for each gene when comparing the target cluster with other clusters. Subsequently, all genes were ranked based on their average fold change, creating an ordered gene list. This ordered gene list was then employed as input for the GSEA software (v4.2.3), utilizing the GSEAPreranked tool with the classic enrichment statistic method. This comprehensive analysis allowed for the identification of significant gene set enrichments, providing valuable insights into the dataset.
Immunofluorescence Assay (IFA) and Live cell imaging
For the immunofluorescence assay, parasites were lysed with 0.15% saponin at 37 °C for 5 min and fixed with 4% polyformaldehyde for 20 minutes on ice. The fixed parasites were carefully deposited on microscope slides, allowed to air dry, and sealed with 1% bovine serum albumin (BSA) in PBS for 1 h at room temperature. Subsequently, the primary antibodies (mouse anti-Ty1, Sigma, SAB4800032; rabbit anti-HA, Abcam ab9110) were co-incubated with the parasites for 1 h at room temperature. Following three thorough washes, corresponding secondary antibodies (Alexa 488 goat anti-mouse antibody, Thermo A11029; Alexa 568 goat anti-rabbit antibody, Thermo A11036) were applied and co-incubated with the parasites for an additional 1 hour at room temperature. After three washes, the parasites were sealed with a mounting medium containing DAPI. Image capture was performed using a Nikon A1R confocal microscope. For live cell imaging, iRBCs were carefully placed on a slide and covered with a coverslip, after which images were immediately acquired using the Nikon A1R confocal microscope.
0–3 h Ring stage survival assay (RSA0-3h)
RSA0-3h was performed following established procedures. Highly synchronized parasites, with a parasitemia of ~ 5%, were obtained through two rounds of 5% sorbitol treatment at 40-hour intervals. These parasites were pretreated with GlcN for 24 h before the assay. When the proportion of mature schizonts (characterized by clearly observable segmented nuclei) exceeded 0.5%, a 40/70% Percoll enrichment was performed. The collected parasites were cultured for an exact 3 h period and then subjected to a 5% sorbitol treatment to obtain the 0–3 h ring-stage parasites. These highly synchronized parasites were exposed to dihydroartemisinin (DHA) at a concentration of 700 nM, or to 0.1% dimethyl sulfoxide (DMSO) as the solvent control, for an exact 6-hour period. Subsequently, the drug was removed, and the parasites were cultured for an additional 66 hours under standard conditions. The survival rates of the parasites were assessed by microscopic examination of blood films. To assess the response of genetically modified parasites to DHA, various dosages, including 50 nM, 200 nM, and 700 nM, were used in the RSA0-3h. Furthermore, a modified ring-stage survival assay was conducted to evaluate the effects of multiple antimalarial drugs. Parasites were pretreated with 5 mM GlcN for 6h and then subjected to two to three rounds of antimalarial exposure. Seven antimalarials, including amodiaquine, pyrimethamine, chloroquine, lumefantrine, piperaquine, pyronaridine, and mefloquine, were applied at a concentration of approximately 10 times their IC50, which was been previously determined. The parasites were exposed to these antimalarials for 6 h in a 24-well culture plate, followed by thorough washing to remove excess drugs. The parasite cultures were then maintained under standard conditions, after which microscopic examination was performed to determine the parasite survival rate.
Targeted metabolomics analysis and data processing
A targeted metabolomics analysis was conducted to examine central carbon metabolism, critical for parasite development, during DHA-induced growth arrest. The highly synchronized parasites were pre-treated with GlcN for 24 h before exposure to 50 nM DHA for 6 h, alongside vehicle-treated 0.05% control groups. Uninfected red blood cells and cultures treated with 10 μM DHA were used as blank and lethal controls, respectively. Metabolites from saponin-lysed parasites were extracted using a two-step process. Briefly, parasites were pelleted, frozen with liquid nitrogen, and then resuspended in pre-chilled 80% methanol and 0.1% formic acid, vortexed, and centrifuged to separate metabolites from cellular debris. Samples were collected from two independent experiments. Quality control (QC) samples were prepared by pooling samples from each group to correct for technical variations during the sample processing phase. The extracted metabolites were analyzed using ultra-performance liquid chromatography-tandem mass spectrometry (UPLC-MS/MS). Specifically, 19 key compounds involved in the tricarboxylic acid (TCA) cycle, glycolytic pathway, pentose phosphate pathway, and corresponding cofactors were profiled. The separation was achieved using a BEH C18 Column (2.1 × 100 mm) with a 10 min linear gradient at a flow rate of 0.3 mL/min. The QTRAP® 6500+ mass spectrometer (SCIEX) was operated in both positive and negative polarity modes to capture a broad spectrum of metabolites.
Raw data files were processed using TraceFinder software (version 4.1). Technical reproducibility was validated by QC samples. The extraction mass (within 10 ppm), retention time, and signal/blank ratio (minimum of 10,000 ions) were used for peak qualification. Quantitative analysis was performed using the external standard method, with integrated peak areas for each metabolite. Peak detection and alignment across all samples were conducted using an in-house compound library generated from reference standards analyzed under identical conditions. To evaluate the metabolic alterations induced by DHA treatment, the log2 fold change of corresponding parasite strains with and without DHA exposure was calculated. Statistical comparisons between groups were performed using a two-tailed student’s t test. For the epigenetic-specific profile, we compared the samples with or without PfMYST knockdown (KD) following 3-hour or 6-hour DHA exposure. Peak areas across all replicates are listed in the Supplementary Data 5.”
Histone acetyltransferases (HATs) inhibition assay
The in vitro histone acetyltransferase inhibition assay was tested using three different HATs inhibitors: NU9056, a PfMYST-specific inhibitor, as well as curcumin (CCM) and anacardic acid (AA), both targeting PfGCN5. The 50% inhibitory concentration (IC50) value for these inhibitors was determined across specified concentration ranges (0.15625 μM to 25 μM for NU9056, and 3.9 μM to 500 μM for curcumin and anacardic acid) using the standard methods. GraphPad Prism 5.0 was used for IC50 calculation. To assess their impact on the DHA susceptibility, trophozoite-stage parasites were exposed to 10 × IC50 concentration of NU9056, CCM, and AA for 12 hours. Late schizonts were subsequently enriched, followed by a 3-hour culture period at 37 °C. Newly invaded rings were purified by 5% sorbitol, and RSA0-3h with different DHA concentrations was performed as described earlier.
Recrudescence assay
The recrudescence assay was conducted according to the previous report with slight modifications. Briefly, parasites underwent two rounds of synchronization at 40 h intervals using 5% sorbitol and purified using 40/70% Percoll. Parasites were cultured with fresh erythrocytes for 6 h, allowing merozoite invasion. To achieve highly synchronized early ring-stage parasites (~ 6 h), cultures were subjected to an additional synchronization step with 5% sorbitol, ensuring the removal of late-stage parasites. These early ring-stage parasites were then exposed to 700 nM DHA and cultured for 6 h. After this initial exposure, the excess drug was removed through two washes with RPMI 1640 incomplete media. The same DHA exposure regimen was repeated at 24 and 48 h after the first exposure. Parasitemia was quantified at various time points (0, 6, 12, 18, 24, 30, 36, 42, and 48 h) to construct parasite clearance curves. Starting on day 3, Giemsa-stained smears were prepared every 24 h to monitor recrudescence. Cultures were maintained under standard conditions until parasitemia reached approximately 2%.
Statistics & reproducibility
No sample-size calculations were performed, and sample size was determined to be adequate based on the magnitude and consistence of measure differences between groups. No data were excluded from analysis. Experiments were performed in two/three biological replicates with technical replicates to ensure the replication as well as the validity of the conclusion. Samples were allocated randomly into both control and experimental groups. Our study did not require blinding. It did not involve animal or human cohorts or subjective counting.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw and processed high-throughput sequencing data generated in this study have been deposited in the Gene Expression Omnibus (GEO) database under accession number GSE246116 and can be accessed using the following link. https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE246116. The raw and deconvoluted peak area of mass spectrometry-based metabolomics data generated in this study are provided in the Source Data file. All data needed to evaluate the conclusions are present in the paper or the Supplementary Information.
References
Bhatt, S. et al. The effect of malaria control on Plasmodium falciparum in Africa between 2000 and 2015. Nature 526, 207–211 (2015).
Li, G. Q., Arnold, K., Guo, X. B., Jian, H. X. & Fu, L. C. Randomised comparative study of mefloquine, qinghaosu, and pyrimethamine-sulfadoxine in patients with falciparum malaria. Lancet 2, 1360–1361 (1984).
Giao, P. T. et al. Artemisinin for treatment of uncomplicated falciparum malaria: is there a place for monotherapy? Am. J. Tropical Med. Hyg. 65, 690–695 (2001).
World Health Organization. Chemotherapy of malaria: report of a WHO scientific group. In World Health Organization Technical Report Series No. 375. (1967).
Teuscher, F. et al. Artemisinin-induced dormancy in Plasmodium falciparum: duration, recovery rates, and implications in treatment failure. J. Infect. Dis. 202, 1362–1368 (2010).
Wellems, T. E., Sá, J. M., Su, X. Z., Connelly, S. V. & Ellis, A. C. ‘Artemisinin resistance’: something new or old? Something of a misnomer? Trends Parasitol. 36, 735–744 (2020).
Peatey, C. et al. Dormant Plasmodium falciparum parasites in human infections following artesunate therapy. J. Infect. Dis. 223, 1631–1638 (2021).
Adjuik, M. et al. Artesunate combinations for treatment of malaria: meta-analysis. Lancet 363, 9–17 (2004).
Warsame, M. et al. Treatment of uncomplicated malaria with artesunate plus sulfadoxine-pyrimethamine is failing in Somalia: evidence from therapeutic efficacy studies and Pfdhfr and Pfdhps mutant alleles. Trop. Med. Int. Health 20, 510–517 (2015).
Adeel, A. A. et al. High efficacy of artemether-lumefantrine and declining efficacy of artesunate + sulfadoxine-pyrimethamine against Plasmodium falciparum in Sudan (2010-2015): evidence from in vivo and molecular marker studies. Malar. J. 15, 285 (2016).
Warsame, M. et al. High therapeutic efficacy of artemether-lumefantrine and dihydroartemisinin-piperaquine for the treatment of uncomplicated falciparum malaria in Somalia. Malar. J. 18, 231 (2019).
Marwa, K. et al. Therapeutic efficacy of artemether-lumefantrine, artesunate-amodiaquine and dihydroartemisinin-piperaquine in the treatment of uncomplicated Plasmodium falciparum malaria in Sub-Saharan Africa: A systematic review and meta-analysis. PLoS ONE 17, e0264339 (2022).
Saunders, D. L. et al. Dihydroartemisinin-piperaquine failure in Cambodia. N. Engl. J. Med. 371, 484–485 (2014).
Chaorattanakawee, S. et al. Ex vivo drug susceptibility testing and molecular profiling of clinical Plasmodium falciparum isolates from Cambodia from 2008 to 2013 suggest emerging piperaquine resistance. Antimicrob. Agents Chemother. 59, 4631–4643 (2015).
Ross, L. S. et al. Emerging Southeast Asian PfCRT mutations confer Plasmodium falciparum resistance to the first-line antimalarial piperaquine. Nat. Commun. 9, 3314 (2018).
Witkowski, B. et al. Novel phenotypic assays for the detection of artemisinin-resistant Plasmodium falciparum malaria in Cambodia: in-vitro and ex-vivo drug-response studies. Lancet Infect. Dis. 13, 1043–1049 (2013).
World Health Organization. Malaria: artemisinin partial resistance. (2022).
Assefa, A., Fola, A. A. & Tasew, G. Emergence of Plasmodium falciparum strains with artemisinin partial resistance in East Africa and the Horn of Africa: is there a need to panic? Malar. J. 23, 34 (2024).
Meshnick, S. Perspective: artemisinin-resistant malaria and the wolf. Am. J. Trop. Med. Hyg. 87, 783–784 (2012).
Hanscheid, T. & Hardisty, D. W. How “resistant” is artemisinin resistant malaria?-The risks of ambiguity using the term “resistant” malaria. Travel Med. Infect. Dis. 24, 23–24 (2018).
Sá, J. M. et al. Artemisinin resistance phenotypes and K13 inheritance in a Plasmodium falciparum cross and Aotus model. Proc. Natl. Acad. Sci. USA 115, 12513–12518 (2018).
Straimer, J. et al. Drug resistance. K13-propeller mutations confer artemisinin resistance in Plasmodium falciparum clinical isolates. Science 347, 428–431 (2015).
Ariey, F. et al. A molecular marker of artemisinin-resistant Plasmodium falciparum malaria. Nature 505, 50–55 (2014).
World Health Organization Global Malaria Programme. Report on antimalarial drug efficacy, resistance and response: 10 years of surveillance (2010–2019). (2020).
WWARN K13 Genotype-Phenotype Study Group Association of mutations in the Plasmodium falciparum Kelch13 gene (Pf3D7_1343700) with parasite clearance rates after artemisinin-based treatments-a WWARN individual patient data meta-analysis. BMC Med. 17, 1–20 (2019).
Amaratunga, C., Witkowski, B., Khim, N., Menard, D. & Fairhurst, R. M. Artemisinin resistance in Plasmodium falciparum. Lancet Infect. Dis. 14, 449–450 (2014).
Ashley, E. A. et al. Spread of artemisinin resistance in Plasmodium falciparum malaria. N. Engl. J. Med. 371, 411–423 (2014).
Chenet, S. M. et al. Independent emergence of the Plasmodium falciparum Kelch Propeller Domain mutant allele C580Y in Guyana. J. Infect. Dis. 213, 1472–1475 (2016).
Lautu-Gumal, D. et al. Surveillance of molecular markers of Plasmodium falciparum artemisinin resistance (kelch13 mutations) in Papua New Guinea between 2016 and 2018. Int. J. Parasitology: Drugs Drug Resist. 16, 188–193 (2021).
Mathieu, L. C. et al. Local emergence in Amazonia of Plasmodium falciparum K13 C580Y mutants associated with in vitro artemisinin resistance. Elife 9, e51015 (2020).
Miotto, O. et al. Emergence of artemisinin-resistant Plasmodium falciparum with Kelch13 C580Y mutations on the island of New Guinea. PLoS Pathog. 16, e1009133 (2020).
Hemming-Schroeder, E. & Lo, E. Evidence of a hard selective sweep for artemisinin resistant Plasmodium falciparum. Lancet Infect. Dis. 17, 462–463 (2017).
Meier-Scherling, C. P. G. et al. Selection of artemisinin partial resistance Kelch13 mutations in Uganda in 2016-22 was at a rate comparable to that seen previously in South-East Asia. Preprint at https://doi.org/10.1101/2024.02.03.24302209 (2024).
Wasakul, V. et al. Malaria outbreak in Laos driven by a selective sweep for Plasmodium falciparum kelch13 R539T mutants: a genetic epidemiology analysis. Lancet Infect. Dis. 23, 568–577 (2022).
Yang, T. et al. Decreased K13 abundance reduces hemoglobin catabolism and proteotoxic stress, underpinning artemisinin resistance. Cell Rep. 29, 2917–2928 (2019).
Xie, S. C., Ralph, S. A. & Tilley, L. K13, the Cytostome, and Artemisinin Resistance. Trends Parasitol. 36, 533–544 (2020).
Birnbaum, J. et al. A Kelch13-defined endocytosis pathway mediates artemisinin resistance in malaria parasites. Science 367, 51–59 (2020).
Klonis, N. et al. Artemisinin activity against Plasmodium falciparum requires hemoglobin uptake and digestion. Proc. Natl. Acad. Sci. USA 108, 11405–11410 (2011).
Xie, S. C. et al. Haemoglobin degradation underpins the sensitivity of early ring stage Plasmodium falciparum to artemisinins. J. Cell Sci. 129, 406–416 (2016).
Tumwebaze, P. K. et al. Decreased susceptibility of Plasmodium falciparum to both dihydroartemisinin and lumefantrine in northern Uganda. Nat. Commun. 13, 6353 (2022).
Siddiqui, F. A. et al. Plasmodium falciparum falcipain-2a polymorphisms in Southeast Asia and their association with artemisinin resistance. J. Infect. Dis. 218, 434–442 (2018).
Henrici R. C., van Schalkwyk D. A., Sutherland C. J. Modification of pfap2mu and pfubp1 markedly reduces ring-stage susceptibility of Plasmodium falciparum to artemisinin in vitro. Antimicrob. Agents and Chemother. 64, https://doi.org/10.1128/aac.01542-19 (2019).
Demas, A. R. et al. Mutations in Plasmodium falciparum actin-binding protein coronin confer reduced artemisinin susceptibility. Proc. Natl. Acad. Sci. USA 115, 12799–12804 (2018).
Behrens, H. M., Schmidt, S. & Spielmann, T. The newly discovered role of endocytosis in artemisinin resistance. Med. Res. Rev. 41, 2998–3022 (2021).
Sutherland, C. J., Henrici, R. C. & Artavanis-Tsakonas, K. Artemisinin susceptibility in the malaria parasite Plasmodium falciparum: propellers, adaptor proteins and the need for cellular healing. FEMS Microbiol. Rev. 45, fuaa056 (2021).
Ullah, I. et al. Artemisinin resistance mutations in Pfcoronin impede hemoglobin uptake. Preprint at https://doi.org/10.1101/2023.12.22.572193 (2023).
Mok, S. et al. Drug resistance. population transcriptomics of human malaria parasites reveals the mechanism of artemisinin resistance. Science 347, 431–435 (2015).
Tucker, M. S., Mutka, T., Sparks, K., Patel, J. & Kyle, D. E. Phenotypic and genotypic analysis of in vitro-selected artemisinin-resistant progeny of Plasmodium falciparum. Antimicrob. Agents Chemother. 56, 302–314 (2012).
Breglio K. F., Rahman R. S., Sá J. M., Roberts D. J., Wellems T. E. Kelch mutations in Plasmodium falciparum protein K13 do not modulate dormancy after artemisinin exposure and sorbitol selection in vitro. Antimicrob. Agents and Chemother. 62, https://doi.org/10.1128/aac.02256-17 (2018).
Connelly, S. V. et al. Restructured mitochondrial-nuclear interaction in Plasmodium falciparum dormancy and persister survival after artemisinin exposure. mBio 12, https://doi.org/10.1128/mbio.00753-21 (2021).
Witkowski, B. et al. Increased tolerance to artemisinin in Plasmodium falciparum is mediated by a quiescence mechanism. Antimicrob. Agents Chemother. 54, 1872–1877 (2010).
Sandoval-Motta, S. & Aldana, M. Adaptive resistance to antibiotics in bacteria: a systems biology perspective. Wiley Interdiscip. Rev. Syst. Biol. Med. 8, 253–267 (2016).
Kucharski, M., Nayak, S., Gendrot, M., Dondorp, A. M. & Bozdech, Z. Peeling the onion: how complex is the artemisinin resistance genetic trait of malaria parasites? Trends Parasitol. 40, 970–986 (2024).
Hughes, K. R. & Waters, A. P. Plasmodium falciparum artemisinin resistance: something gained in translation. Trends Parasitol. 40, 541–543 (2024).
Small-Saunders, J. L. et al. tRNA modification reprogramming contributes to artemisinin resistance in Plasmodium falciparum. Nat. Microbiol. 9, 1483–1498 (2024).
Duraisingh, M. T. & Skillman, K. M. Epigenetic Variation and Regulation in Malaria Parasites. Annu. Rev. Microbiol. 72, 355–375 (2018).
Lucky, A. B. et al. Plasmodium falciparum GCN5 plays a key role in regulating artemisinin resistance-related stress responses. Antimicrob Agents Chemother. 67, e00577-23 (2023).
Liang, X. et al. A leak-free inducible CRISPRi/a system for gene functional studies in Plasmodium falciparum. Microbiol. Spectr. 10, e02782-21 (2022).
Siddiqui, G., Srivastava, A., Russell, A. S. & Creek, D. J. Multi-omics based identification of specific biochemical changes associated with PfKelch13-mutant artemisinin-resistant Plasmodium falciparum. J. Infect. Dis. 215, 1435–1444 (2017).
Ndwiga, L. et al. A review of the frequencies of Plasmodium falciparum Kelch 13 artemisinin resistance mutations in Africa. Int. J. Parasitol Drugs Drug Resist 16, 155–161 (2021).
Nima, M. K. et al. Assessment of Plasmodium falciparum artemisinin resistance independent of kelch13 polymorphisms and with escalating malaria in Bangladesh. mBio 13, e03444-21 (2022).
Boullé, M. et al. Artemisinin-resistant Plasmodium falciparum K13 mutant alleles, Thailand–Myanmar border. Emerg. Infect. Dis. 22, 1503–1505 (2016).
Staples, J. et al. Profiling and leveraging relatedness in a precision medicine cohort of 92,455 exomes. Am. J. Hum. Genet. 102, 874–889 (2018).
Schmidt, S. et al. The Kelch13 compartment contains highly divergent vesicle trafficking proteins in malaria parasites. Plos Pathog. 19, e1011814 (2023).
Hoeijmakers, W. A. M. et al. Epigenetic reader complexes of the human malaria parasite, Plasmodium falciparum. Nucleic Acids Res. 47, 11574–11588 (2019).
Rawat, M., Malhotra, R., Shintre, S., Pani, S. & Karmodiya, K. Role of PfGCN5 in nutrient sensing and transcriptional regulation in Plasmodium falciparum. J. Biosci. 45, 1–13 (2020).
Miao, J. et al. A unique GCN5 histone acetyltransferase complex controls erythrocyte invasion and virulence in the malaria parasite Plasmodium falciparum. PLoS Pathog. 17, e1009351 (2021).
Rawat, M. et al. Histone acetyltransferase PfGCN5 regulates stress responsive and artemisinin resistance related genes in Plasmodium falciparum. Sci. Rep. 11, 852 (2021).
Ishola, A. A. & Adewole, K. E. In Silico Screening Reveals Histone Deacetylase 7 and ERK1/2 as Potential Targets for Artemisinin Dimer and Artemisinin Dimer Hemisuccinate. Curr. Drug Discov. Technol. 17, 725–734 (2020).
Kumari, K., Keshari, S., Sengupta, D., Sabat, S. C. & Mishra, S. K. Transcriptome analysis of genes associated with breast cancer cell motility in response to Artemisinin treatment. BMC Cancer 17, 858–870 (2017).
Yan, Y., Harper, S., Speicher, D. W. & Marmorstein, R. The catalytic mechanism of the ESA1 histone acetyltransferase involves a self-acetylated intermediate. Nat. Struct. Biol. 9, 862–869 (2002).
Miao, J. et al. The MYST family histone acetyltransferase regulates gene expression and cell cycle in malaria parasite Plasmodium falciparum. Mol. Microbiol. 78, 883–902 (2010).
Prommana, P. et al. Inducible knockdown of Plasmodium gene expression using the glmS ribozyme. PloS ONE 8, e73783 (2013).
Fan, Y. et al. Rrp6 regulates heterochromatic gene silencing via ncRNA RUF6 decay in malaria parasites. mBio 11, https://doi.org/10.1128/mbio.01110-20 (2020).
Sen, U., Nayak, A., Khurana, J., Sharma, D. & Gupta, A. Inhibition of PfMYST histone acetyltransferase activity blocks Plasmodium falciparum growth and survival. Antimicrob. Agents and Chemother. 65, https://doi.org/10.1128/aac.00953-20 (2020).
Cui, L., Miao, J. & Cui, L. Cytotoxic effect of curcumin on malaria parasite Plasmodium falciparum: inhibition of histone acetylation and generation of reactive oxygen species. Antimicrob. Agents Chemother. 51, 488–494 (2007).
Cui, L. et al. Histone acetyltransferase inhibitor anacardic acid causes changes in global gene expression during in vitro Plasmodium falciparum development. Eukaryot. Cell 7, 1200–1210 (2008).
Yu, X. et al. Ring-stage growth arrest: Metabolic basis of artemisinin tolerance in Plasmodium falciparum. Iscience 26, 105275–105299 (2023).
Khoury, D. S., Cao, P., Zaloumis, S. G. & Davenport, M. P. Artemisinin resistance and the unique selection pressure of a short-acting antimalarial. Trends Parasitol. 36, 884–887 (2020).
Allman, E. L., Painter, H. J., Samra, J., Carrasquilla, M. & Llinás, M. Metabolomic profiling of the malaria box reveals antimalarial target pathways. Antimicrob. Agents Chemother. 60, 6635–6649 (2016).
Mok, S. et al. Artemisinin-resistant K13 mutations rewire Plasmodium falciparum’s intra-erythrocytic metabolic program to enhance survival. Nat. Commun. 12, 530 (2021).
Zhao, Y. et al. Optimization of CRISPR/Cas system for improving genome editing efficiency in Plasmodium falciparum. Front. Microbiol. 11, 625862 (2021).
Jiang, J. B., Jacobs, G., Liang, D.-S. & Aikawa, M. Qinghaosu-induced changes in the morphology of Plasmodium inui. Am. J. Trop. Med. Hyg. 34, 424–428 (1985).
Maeno, Y. et al. Morphologic effects of artemisinin in Plasmodium falciparum. Am. J. Trop. Med. Hyg. 49, 485–491 (1993).
Campanella, M. & Kannan, B. Mitochondrial sites of contact with the nucleus. J. Cell Biol. 223, e202305010 (2024).
Mehta, M., Sonawat, H. M. & Sharma, S. Malaria parasite-infected erythrocytes inhibit glucose utilization in uninfected red cells. FEBS Lett. 579, 6151–6158 (2005).
Mbengue, A. et al. A molecular mechanism of artemisinin resistance in Plasmodium falciparum malaria. Nature 520, 683–687 (2015).
Straimer, J. et al. K13-propeller mutations confer artemisinin resistance in Plasmodium falciparum clinical isolates. Science 347, 428–431 (2015).
Bezrukov, F., Prados, J., Renzoni, A. & Panasenko, O. O. MazF toxin causes alterations in Staphylococcus aureus transcriptome, translatome and proteome that underlie bacterial dormancy. Nucleic Acids Res. 49, 2085–2101 (2021).
Kanyal, A. et al. Genome-wide survey and phylogenetic analysis of histone acetyltransferases and histone deacetylases of Plasmodium falciparum. FEBS J. 285, 1767–1782 (2018).
Cheng, Q., Kyle, D. E. & Gatton, M. L. Artemisinin resistance in Plasmodium falciparum: a process linked to dormancy? Int. J. parasitol. Drugs Drug Resist. 2, 249–255 (2012).
Tripathi, J. et al. The artemisinin-induced dormant stages of Plasmodium falciparum exhibit hallmarks of cellular quiescence/senescence and drug resilience. Nat. Commun. 15, 7485 (2024).
Kiboi, D. et al. Isolation and characterization of Plasmodium falciparum blood-stage persisters by improved selection protocols using dihydroartemisinin alone. Antimicrob. Agents Chemother. 69, e00053-24 (2025).
Kalamuddin, M. et al. MYST regulates DNA repair and forms a NuA4-like complex in the malaria parasite Plasmodium falciparum. Msphere 9, e00140-24 (2024).
Siddiqui, F. A., Liang, X. & Cui, L. Plasmodium falciparum resistance to ACTs: Emergence, mechanisms, and outlook. Int. J. Parasitol. Drugs Drug Resist. 16, 102–118 (2021).
Ismail, H. M. et al. Artemisinin activity-based probes identify multiple molecular targets within the asexual stage of the malaria parasites Plasmodium falciparum 3D7. Proc. Natl. Acad. Sci. USA 113, 2080–2085 (2016).
Bridgford, J. L. et al. Artemisinin kills malaria parasites by damaging proteins and inhibiting the proteasome. Nat. Commun. 9, 3801 (2018).
Zhu, L. et al. The origins of malaria artemisinin resistance defined by a genetic and transcriptomic background. Nat. Commun. 9, 5158 (2018).
Zhu, L. et al. Artemisinin resistance in the malaria parasite, Plasmodium falciparum, originates from its initial transcriptional response. Commun. Biol. 5, 274 (2022).
Duru, V. et al. Plasmodium falciparum dihydroartemisinin-piperaquine failures in Cambodia are associated with mutant K13 parasites presenting high survival rates in novel piperaquine in vitro assays: retrospective and prospective investigations. BMC Med. 13, 305 (2015).
Witkowski, B. et al. A surrogate marker of piperaquine-resistant Plasmodium falciparum malaria: a phenotype-genotype association study. Lancet Infect. Dis. 17, 174–183 (2017).
Bopp, S. et al. Plasmepsin II-III copy number accounts for bimodal piperaquine resistance among Cambodian Plasmodium falciparum. Nat. Commun. 9, 1769 (2018).
Lim, P. et al. Ex vivo susceptibility of Plasmodium falciparum to antimalarial drugs in western, northern, and eastern Cambodia, 2011-2012: association with molecular markers. Antimicrob. Agents Chemother. 57, 5277–5283 (2013).
Tandoh, K. Z., Morang’a, C. M., Wilson, M., Quashie, N. B. & Duah-Quashie, N. O. Malaria artemisinin resistance: an extracellular vesicles export hypothesis. Trends Parasitol. 38, 614–617 (2022).
Zhang, M. et al. The apicoplast link to fever-survival and artemisinin-resistance in the malaria parasite. Nat. Commun. 12, 4563 (2021).
Henriques, G. et al. The Mu subunit of Plasmodium falciparum clathrin-associated adaptor protein 2 modulates in vitro parasite response to artemisinin and quinine. Antimicrob. Agents Chemother. 59, 2540–2547 (2015).
Simwela N. V. et al. Experimentally Engineered Mutations in a Ubiquitin Hydrolase, UBP-1, Modulate In Vivo Susceptibility to Artemisinin and Chloroquine in Plasmodium berghei. Antimicrob. Agents Chemother. 64, https://doi.org/10.1128/aac.02484-19 (2020).
Egwu, C. O. et al. Resistance to artemisinin in falciparum malaria parasites: A redox-mediated phenomenon. Free Radic. Biol. Med. 179, 317–327 (2022).
Poran, A. et al. Single-cell RNA sequencing reveals a signature of sexual commitment in malaria parasites. Nature 551, 95–99 (2017).
Walzer, K. A., Kubicki, D. M., Tang, X. & Chi, J.-T. A. Single-cell analysis reveals distinct gene expression and heterogeneity in male and female Plasmodium falciparum gametocytes. MSphere 3, https://doi.org/10.1128/msphere.00130-18 (2018).
Howick, V. M. et al. The Malaria Cell Atlas: Single parasite transcriptomes across the complete Plasmodium life cycle. Science 365, eaaw2619 (2019).
Acknowledgements
Q.Z. received support from the National Natural Science Foundation of China (NSFC) [82230077, W2411080]; Shanghai Blue Cross Brain Hospital Co., Ltd. and Shanghai Tongji University Education Development Foundation. J.C. received support from the National Natural Science Foundation of China (NSFC) [82320108014, 81971967]; National Key R&D Program of China Grant [2023YFA1801004]. C.J. is supported by the major project in the basic research field of Shanghai Science and Technology Innovation Action Plan [22JC1402300]. X.Y. is supported by the National Natural Science Foundation of China (NSFC) [82102424]. X.C. is supported by the Jiangxi Provincial Natural Science Foundation [20242BAB20250]; Open Project of Key Laboratory of Pathogen-Host Interaction, Ministry of Education [KFKT20250004]; Jiangxi Province Early Vocational Young Scientific and Technological Personnel Training Project [20244BCE52258]; Doctoral Research Initiation Fund Project of Nanchang Medical College [NYB24005]. J.M., J.M.S., and T.E.W. received support from the Division of Intramural Research, National Institute of Allergy and Infectious Diseases, US National Institutes of Health. L.P. and D.M. received funding from the Fondation pour la Recherche Médicale (FRM) under the ‘Équipes FRM 2024’ program (EQU202403018026), with D.M. also supported by the Institut Universitaire de France (IUF) Senior Chair program (2024–2029) and the French Government’s Investissements d’Avenir program through the Laboratoire d’Excellence (LabEx) “French Parasitology Alliance For Health Care” (ANR-11-LABX-0024-PARAFRAP).
Author information
Authors and Affiliations
Contributions
Q.Z., J.C., C.J., T.E.W. and D.M. conceived and designed the experiments. C.W., X.C., Y.Z., X.H., W.S. and R.T. generated all gene knock-out, knockdown and knock-in transgenic parasite lines and performed ChIP-seq, RNA-seq and scRNA-seq assays. X.Y., S.L. and J.T. performed ART-R phenotypic analysis and validation, antimalarial assays and metabolism analysis. J.H., C.J., J.M.S., L.P. and J.M. performed the data analysis. Q.Z., T.E.W. and J.M. wrote the manuscript with contributions from other authors. All authors discussed and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Elena Gómez-Díaz, Sachel Mok, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yu, X., He, J., Wang, C. et al. Epigenetically conferred ring-stage survival in Plasmodium falciparum against artemisinin treatment. Nat Commun 16, 8037 (2025). https://doi.org/10.1038/s41467-025-62479-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-62479-2