Abstract
The Mre11 complex comprises Mre11, Rad50 and Nbs1 (Xrs2 in S. cerevisiae). The core components, Mre11 and Rad50 are highly conserved, with readily identifiable orthologs in all clades of life, whereas Nbs1/Xrs2 are present only in eukaryotes. In eukaryotes, the complex is integral to the DNA damage response, acting in DNA double strand break (DSB) detection and repair, and the activation of DNA damage signaling. We present here a 3.2 Å cryo-EM structure of the S. cerevisiae Mre11-Rad50 complex with bound dsDNA. The structure provided a foundation for detailed mutational analyses regarding homo and heterotypic protein interfaces, as well as DNA binding properties of Rad50. We define several conserved residues in Rad50 and Mre11 that are critical to complex assembly as well as for DNA binding. In addition, the data reveal that the Rad50 coiled coil domain influences ATP hydrolysis over long distances.
Similar content being viewed by others
Introduction
The Mre11 complex is integral to the DNA damage response (DDR). The complex promotes DNA double-strand break (DSB) repair and in eukaryotes, it activates the Tel1/ATM kinase to initiate DNA damage signaling. The complex is comprised of two molecules each of Mre11 and Rad50, and one molecule of Nbs1 (Xrs2 in budding yeast)1. Orthologs of Mre11 and Rad50 are found in all clades of life, whereas Nbs1/Xrs2 is exclusive to Eukarya. Mre11 and Rad50 (MR) specify enzymatic functions (nuclease and ATPase, respectively), whereas Nbs1/Xrs2 lacks catalytic functions. Recent studies demonstrate that the essential function of Nbs1/Xrs2 is to facilitate proper assembly of the complex, which in turn promotes nuclear localization2,3. Nevertheless, there is some evidence for Nbs1/Xrs2 independent functions of the MR complex in S. cerevisiae4.
The Mre11 complex is a distant member of the Structure Maintenance of Chromosomes (SMC) family of proteins; however, there are two important distinctions between the Mre11 complex and classical SMC proteins. First, whereas SMC proteins interact via a hinge domain that lies within their respective coiled coil regions, Rad50 dimerizes in the analogous region through a zinc-binding domain assembled from two Rad50 protomers termed the Rad50 Zn hook domain. Second, the complex does not contain kleisin subunits, which are found in SMC complexes such as cohesin, condensin and Smc5/6 associated with the Walker A and B ATPase domains5. Instead, the Rad50 Walker A and B domains from each of the two Rad50 protomers associate with each other in part by binding two ATP molecules6.
The Mre11 complex appears to function in two forms. In the ATP-bound closed form, the Rad50 Walker A and B domains from each of the two Rad50 protomers associate with each other, with the adjacent coiled coils adopting a rod-shaped structure. This form appears to primarily influence Tel1/ATM activation and non-homologous end joining7,8,9. Upon ATP hydrolysis, the Mre11 complex assumes an open form in which the two ATP-binding domains rotate outward, such that the Mre11 dimer interface, in which the nuclease active sites reside, becomes accessible. Accordingly, the open form has been implicated in Mre11 complex-mediated DSB end resection and repair by homologous recombination6.
In contrast, recent cryo-EM analysis of the SbcCD complex, the E. coli ortholog of the eukaryotic MR complex, presents a different view. The structures obtained reveal that in the absence of DNA, the Rad50 coiled coils are in an open, ring-shaped form with the Mre11 dimer underneath the Rad50 ATPase domains. This form senses a DNA end, which subsequently induces “zippering” of the coiled coils into a closed rod-shaped form. Zippering of the coiled coils is accompanied by SbcD (Mre11) relocalizing to the side of Rad50, where it binds the DNA end and creates a nuclease channel in which DNA ends or hairpin structures are processed prior to homology-directed DSB repair10,11. This raises the possibility that the structural transitions themselves, as opposed to the open and closed endpoints, may be functionally important.
The mechanisms underlying the transition from the closed to open forms of the Mre11 complex require further investigation. In previous studies, we determined the crystal structure of a 182-aa fragment spanning the human Rad50 Zn hook domain. Those data demonstrate that the apex of the Rad50 coiled coils form a rod-shaped structure that is unlikely to come apart during the transition from the closed to open forms of the complex. In addition to the zinc-binding interface situated at the apex, the adjacent coiled coils form a second interface that is roughly equivalent in size to the zinc-binding interface (ca., 900 Å2). This coiled coil interface is situated between the apical zinc hook interface and a highly flexible five-residue “hinge” loop (CGSQD)12, suggesting that this flexible region permits mobility of this region of the coiled coils. Given that the apical region of Rad50 is rigid and stable, this hinge domain, and possibly additional regions of coiled coil flexibility, are presumably required for the opening of the closed complex.
The available evidence suggests at least two modes of DNA binding by the Mre11 complex that are specific to the closed and open forms of the complex. We used cryo-EM to examine the globular domain and the proximal coiled coil domain of the S. cerevisiae MR complex to gain insights regarding DNA engagement, the interfaces that mediate assembly of the complex, and to probe the mechanism of ATM/Tel1 activation. We report herein a 3.2 Å structure of the closed form of the MR complex with bound dsDNA. Reminiscent of the E. coli. structure of the MR-dsDNA complex10, the coiled coils encircle the DNA and come together above it. Mre11 does not make contact with the dsDNA, suggesting that this DNA-bound structure represents a precursor to the open form of the complex, as predicted previously13. A recently reported structure of the C. thermophilum MRN complex bound to ATPγS revealed a similar rod-shaped structure that did not include DNA11.
Results
The domain organization of the Mre11 and Rad50 protomers is outlined in Fig. 1A. Rad50 contains a tripartite filamentous architecture in which the N- and C-terminal Walker A and B domains are separated by two long arms (ca. 400 aa) of coiled coil segments. The N-terminal arm terminates at a 100 aa domain comprising the zinc hook domain, which constitutes a Zn-dependent dimerization interface. The C-terminal coiled coil arm begins at the zinc hook and extends to the Walker B domain. The Rad50 protein folds at the hook domain such that the N- and C-arms of the coiled coils are in an antiparallel configuration and the Walker A and B domains interface to form part of the Mre11 complex globular domain. Mre11 has an N-terminal Mn2+ dependent phosphoesterase (nuclease) domain, access to which is regulated by a capping domain. There is a helix-turn-helix (HTH) motif C-terminal to the capping domain that appears to bind the Rad50 coiled coils proximal to the globular domain, while the remainder of the C-terminus is largely disordered9. Nbs1 (Xrs2 in S. cerevisiae) is found only in Eukarya. It has N-terminal FHA and BRCT domains and a highly conserved ten amino acid motif (RKNFKTFVKV in S. cerevisiae) that is necessary and sufficient for interaction with the Mre11 dimer. The remaining protein segments appear to be disordered2,14.
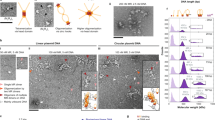
A Domain architecture of S. cerevisiae Rad50 and Mre11. B, C The 3.2 Å cryo-EM structure of MREQ-DNA complex in electron density (B) and ribbon (C) representations. The symmetry related Rad50 subunits are colored in light and dark green, the symmetry related Mre11 subunits are colored in light and dark orange, and the bound dsDNA is in yellow. The bound ATPs are shown in space-filling representation (C). D, E The 3.5 Å cryo-EM structure of apo-MREQ in electron density (D) and ribbon (E) representations. The bound ATPs are shown in space-filling representation in panel E.
We purified the Mre11-Rad50-Xrs2 (MRX) complex from S. cerevisiae to carry out structural studies. We were unable to obtain useable MRX particles with or without DNA and crosslinking. To minimize the spatial dynamics of the complex, we next purified the Mre11-Rad50 (MR) complex, in which the E1235 residue of Rad50 was mutated to Q (REQ) to prevent ATP hydrolysis9, which likely drives the transition from the closed to the open form of the complex13,15.
Cryo-EM structure of the MR-dsDNA complex
MREQ, with Mre11 containing a C-terminal 1xFLAG tag, was purified from yeast extracts by sequential affinity purification to FLAG-Agarose, Heparin-Sepharose and peak fractionation by gel filtration chromatography (Fig. S1A) to obtain highly pure MREQ (Fig. S1B). MREQ was incubated with a 83-mer double stranded DNA (dsDNA) in the presence of ATP, MgCl2 and MnCl2 prior to making the cryo-EM grids. We also made cryo-EM grids without dsDNA; however, we were unable to obtain high-quality homogeneous 2D class averages suitable for further processing.
The structure of the MR-dsDNA complex at an overall resolution of 3.2 Å (cryo-EM workflow listed in Fig. S2, FSC curve in Fig. S3A, angular distribution plot in Fig. S3B, and statistics in Table S1) is shown in density and ribbon representations in Fig. 1B, C, respectively. The central component of the complex spanning the head segment of Rad50 and the nuclease with capping domains of Mre11 can be monitored at the highest resolution (Fig. S3C), as reflected in the tracing of side chains of these segments (Fig. S3D-S3F). The correct density tracing of these segments was facilitated by incorporation of secondary structure elements derived from AlphaFold2 and Schrödinger program-based homology modeling. The head-proximal coiled coil arms of Rad50 and the HTH domain of Mre11 can be monitored at medium resolution (3.5–4.0 Å; Fig. S3C), with these segments suitable for structural rigid body fit and refinement. By contrast, traceable coiled coil arm segments of Rad50 (residues 220–239 and 1089–1102) are observed at lower resolution, while the remainder of the coiled coil arm and Zn hook segments (residues 239–1089) of Rad50, the segment (residues 413–441) connecting the capping and HTH domains of Mre11 and the C-terminus of Mre11 (residues 508–692), are flexible and cannot be traced in the complex (Figs. 1B, 1C and S3C). Notably, we can also monitor a pair of bound ATPs and about two turns of bound dsDNA in the MR-DNA complex (Figs. 1B and 1C).
We were also able to elucidate the structure of apo-MR in the same data set at an overall resolution of 3.5 Å (cryo-EM workflow listed in Figure S2 and statistics in Table S1). The density and ribbon representations are shown in Fig. 1D, E, respectively. Despite the lower resolution, we were able to trace both Mre11 domains, both Rad50 head domains, a pair of bound ATPs, and one of the two head-proximal coiled-coil segments.
Rad50 head domain alignment and Mre11 nuclease pocket
The pair of ATPase domains of Rad50 in the S. cerevisiae MR (data not shown) and MR-dsDNA complex (Figure S3G) in this study both adopt an “engaged” alignment, following comparison with the “engaged” alignment observed in the S. cerevisiae Cohesin (PDB: 6ZZ6) versus the “juxtaposed” alignment observed in the S. cerevisiae Condensin (PDB: 6YVU).
Both the catalytic residues and divalent cations in the Mre11 nuclease active site of the cryo-EM structure of the S. cerevisiae MR-dsDNA complex reported in this study (Figure S3H, in color) superpose well with those of the X-ray structure of the P. furiosus Mre11 (PDB: 1S8E; Figure S3H, in silver).
Protein-protein contacts involving Mre11 capping domain with Rad50 head domain
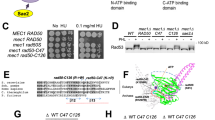
Our structure of the MR-dsDNA complex revealed details of the interaction interface between Rad50 head domain and Mre11 capping domain, with five residues from Rad50 and six from Mre11 contributing to the interface (two alternate views in boxed panels of Fig. 2A). Rad50-E1155 makes hydrogen bond contacts with Mre11-K410 and Mre11-R412, while Rad50-D1167 interacts with Mre11-K322 and Mre11-R390. In addition, the interface is further stabilized by side chain hydrogen bond contacts between Rad50-E1243 and Mre11-R389. Rad50 residues E1155, D1167 and E1243 and Mre11 residues N387, R390, K410 and R412 are widely conserved (Fig. 2B, top and bottom, respectively), supporting their functional significance. Accordingly, mutation of these three Rad50 acidic residues (E1155, D1167 and E1243) to alanine (to create rad50-MRI-3A) imparts temperature-dependent sensitivity to camptothecin (CPT) (Fig. 2C, top panel). Note that in the rad50-MRI-3A mutant, the Rad50-G1163 and -T1164 backbone interactions with Mre11-N387 and -R390 remain intact (boxed panels, Fig. 2A). All three possible double mutants from the rad50-MRI-3A triad exhibit modest CPT sensitivity at 37 °C, whereas only mild sensitivity was observed for single mutants at 37 °C (Fig. 2C, top panel).
A The boxed panels show two views rotated by 90° of protein-protein interactions between the Mre11 capping domain and Rad50 head domain in the MR-dsDNA complex. Conserved Rad50 E1155, D1167 and E1243 residues and Mre11 N387, R389, R390, K410 and R412 residues are labeled in the boxed segments. B Evolutionary conservation of Rad50 (top) and Mre11 (bottom) residues mediating protein-protein interactions. Tm, Thermotoga maritima; Ct, Chaetomium thermophilum; Sc, Saccharomyces cerevisiae; Sp, Schizosaccharomyces pombe; Dm, Drosophila melanogaster; Ce, Caenorhabditis elegans; Mm, Mus musculus; Hs, Homo sapiens. Rad50 and Mre11 tumor alleles and their number of instances (in brackets) are depicted below the alignments. C CPT-survival of rad50 (top) and mre11 (bottom) mutants in the Mre11-Rad50 interaction interface. Plates were either incubated at 30 °C (left panel) or 37 °C (right panel). The following abbreviations were used: rad50-MRI-3A (rad50-E1155A D1167A E1243A); mre11-MRI-5A1 (K322A N387A R389A R390A R412A); mre11-MRI-5A2 (K322A N387A R389A R390A K410A); mre11-MRI-6A (K322A N387A R389A R390A K410A R412A). D Assessment of the Mre11-Rad50 and Mre11-Xrs2 interaction by co-immunoprecipitation. Cells were either grown at 30 °C (left panel) or 37 °C (right panel) and Rad50, Mre11 or Xrs2 were immunoprecipitated from the prepared cell extracts (labeled on the left) and probed with anti-Rad50, anti-Mre11 or anti-Xrs2 antibodies (labeled on the right) by Western blot. Molecular size markers in kDa are indicated on the left. Images shown are representative of three independent experiments with comparable results. Uncropped Western blots are provided as a source data file. E Assessment of the Mre11-Rad50 and Mre11-Xrs2 interaction by the yeast 2-hybrid assay. Rad50, Mre11 and Xrs2 were expressed as a fusion with the Gal4-DNA Binding Domain (pGBD; TRP1) or the Gal4-Activation Domain (pGAD; LEU2) as indicated. Reporter activation assessed by growth in the absence of histidine (Do-TLH) and in the absence of adenine and histidine (Do-TLAH) were assessed in WT, mre11-MRI-6A and mre11-E38K mutants. Empty plasmids were included as a negative control.
mre11-MRI-6A contains alanine mutations of the six Mre11 residues noted above (K322A, N387A, R389A, R390A, K410A, and R412A). As Mre11-K410 and Mre11-R412 both form contacts with Rad50-E1155 (boxed panels, Fig. 2A), mutations of those Mre11 residues are mre11-MRI-5A1 (K322A N387A R389A R390A R412A) and mre11-MRI-5A2 (K322A N387A R389A R390A K410A). mre11-MRI-6A phenocopied rad50-MRI-3A, with markedly enhanced CPT sensitivity at 37 °C (Fig. 2C, bottom panel). That mre11-MRI-5A1 and −5A2 exhibited equivalent sensitivity is consistent with the interpretation that Mre11-K410 and Mre11-R412 both contact Rad50-E1155 (boxed panels, Fig. 2A).
The CPT sensitivity of rad50-MRI-3A, mre11-MRI-5A1 and mre11-MRI-6A mutants (Fig. 2C, lower panel) results from disruption of interaction with Xrs2 in those mutants. Co-immunoprecipitation from cells grown at 30 °C or 37 °C revealed that the abundance of Rad50-MRI-3A was strongly reduced both at 30 °C and 37 °C, but the interaction with Mre11 was intact at both incubation temperatures (Fig. 2D).
Similarly, Mre11-MRI-5A1, −5A2 and Mre11-MRI-6A proteins had only a modest decrement in Rad50 interaction as inferred from co-immunoprecipitations (Fig. 2D). To ask whether disruption of Mre11 dimerization accounted for the lack of Xrs2 interaction, a yeast two hybrid assay was carried out to determine if homodimerization was impaired. The data showed that Mre11-MRI-6A homodimerization was strongly compromised, as was interaction with Xrs2 and Rad50 (Fig. 2E).
Xrs2 contains the nuclear localization signal (NLS) and is essential for the nuclear import of the Mre11 complex4. While we observe that rad50 and mre11 mutants with diminished Xrs2 association exhibit CPT-sensitivity (Fig. 2C), we have not determined whether this sensitivity results from impaired nuclear import of the MRX complex, destabilization of protein-protein interactions within the complex, or a combination of both.
Protein-protein contacts involving Mre11 HTH domain with Rad50 head-proximal coiled-coil domain
Previous studies have noted that the Mre11 HTH domain lies across the Rad50 coiled coils proximal to the globular domain largely through hydrophobic interactions comprising approximately 970 Å213,16. Within the HTH domain, the structure revealed an interaction between the Rad50-D1126 side chain and the backbones of Mre11-S460, -L461, and -L462 (Fig. 3A, top left boxed panel). Changing D1126, which is highly conserved (Fig. 3B), to alanine, asparagine or glutamate phenocopied the CPT sensitivity of the rad50∆ (Fig. 3C) and completely disrupted the Mre11-Rad50 association (Fig. 3D). These data indicate that the interface of the Rad50 coils, particularly the interactions of Rad50-D1126 with Mre11 HTH motif (Fig. 3A, top left boxed panel) are necessary and sufficient to retain the Rad50-Mre11 interaction.
A The boxed panels show protein-protein and ATP-protein contacts in the MR-dsDNA complex. (Top left panel) The Rad50 head proximal coil residue D1126 forms hydrogen bonds with the main chain of Mre11 HTH domain residues S460, L461 and L462. (Bottom left panel) Mre11 residue E38 forms hydrogen bonds with the main chain of Mre11 residues E299 and T300. (Right panel) Ligand-protein contacts between bound ATP and Walker A and B motifs of Rad50 in the complex. Encapsulation of Mg2+-coordinated ATP by conserved Walker A (orange) and Walker B (blue) motifs from one Rad50 D-loop (magenta) and P-loop (pink) from the other Rad50. Hydrogen bonds amongst ATP-protein and within the Rad50 dimer are shown as dashed lines. Critical residues mediating the Rad50-Rad50 interaction within this nucleotide binding domain (NBD) are shown. B Rad50-D1126 in the Rad50 head proximal coils is highly conserved. C CPT-sensitivities of rad50-mutants in D1126 and W1157 residues. D Mre11-Rad50 complex integrity in Rad50 coils (D1126A/N/E) and Rad50 ATPase (W1157A/Y, R13A, N36A) mutants assessed by co-immunoprecipitation. Representative images from three independent experiments are shown. Molecular size markers in kDa are given on the left. Uncropped Western blots are provided as a source data file. E Rad50 ATP-interacting residues R13 and N36 are critical for CPT-survival. F Rad50-W1157 lining the ATP-binding pocket is highly conserved. G Positioning and van der Waals contacts of Rad50-W1157 within the hydrophobic pocket. H Minor CPT-sensitivities at 37 °C of mutants in Rad50 residues (E159, K1209) mediating nucleotide-independent Rad50-Rad50 interaction.
Collectively, these data reveal the critical role of the Mre11 HTH domain, particularly that of Rad50-D1126 in the Rad50 coils (Fig. 3A, top left boxed panel), in promoting Mre11-Rad50 interaction. Single mutations distal to the Mre11 HTH domain, such as those involving the capping domain, imparted at best partial temperature sensitivity (Fig. 2C), indicating that those residues do not strongly influence the interaction of Mre11 and Rad50.
Protein-protein and ATP-protein contacts centered on the ATP-binding pocket of Rad50
In addition to homodimerization at the apical Zinc hook domain, Rad50 homodimerization is also mediated by Walker A and B motifs, with the Walker A from one protomer interacting with Walker B from the other17. This interaction is stabilized by ATP coordination between each Walker A and B pair such that two ATP molecules reside within the interface.
Within the Walker A motif, Rad50-R13 forms a hydrogen bond with the adenosine moiety of ATP and Rad50-K40 coordinates with the β and γ phosphates of ATP, whereas Rad50-N36 interacts with both the ATP γ-phosphate and Rad50-D1241 in the D loop of the opposing Walker B motif (Fig. 3A, right boxed panel). Mutation of the corresponding residues in bacteriophage T4 Rad50 (gp46) severely reduces ATP binding affinity and hydrolysis, as well as Mre11 (gp47) dependent nuclease activity17. Consistent with those data, rad50-R13A and rad50-N36A phenocopy the CPT sensitivity of rad50∆ (Fig. 3E). Notably, this outcome was not due to disruption of the Mre11-Rad50 interaction, as co-immunoprecipitation of these Rad50 mutants and Mre11 was indistinguishable from WT (Fig. 3D).
Rad50-W1157 is conserved from S. cerevisiae to humans (Fig. 3F). It lies just adjacent to the ATP binding pocket in the Walker B motif (Fig. 3A, right boxed panel). The W1157 residue extends into a hydrophobic pocket within the same protomer where it makes multiple van der Waals contacts (Fig. 3G). It appears critical that this residue is aromatic (Fig. 3F). It is substituted with phenylalanine in T. maritima (Fig. 3F), and whereas rad50-W1157A is as CPT sensitive as rad50∆, rad50-W1157Y is only minimally sensitive to CPT (Fig. 3C). The CPT sensitivities observed are associated with destabilization of Rad50 itself, as well as its interaction with Mre11. Co-immunoprecipitation experiments reveal that the abundance of Rad50-W1157A and -W1157Y is reduced, but some residual interaction with Mre11 is observed only for Rad50-W1157Y (Fig. 3D). Similarly, the interaction between Rad50-E159 and -K1209 (Fig. 3A, right boxed panel) plays a less significant role in Mre11 complex function, as rad50-E159A and rad50-K1209A exhibit mild CPT sensitivity only at 37 °C (Fig. 3H).
Mre11 and Rad50 are each stabilized by intramolecular interactions within individual domains. Mre11-E38 forms side chain and backbone contacts with Mre11-T300 and E-299 within the same protomer (Fig. 3A, lower left boxed panel). In a previous study, the mre11-E38K mutation was modeled in yeast after the corresponding human residue was found to be recurrently mutated in ovarian and endometrial cancers. mre11-E38K exhibited strongly reduced levels of Mre11-E38K protein and sensitivity to CPT18. As above, a two hybrid assay showed that Mre11-E38K homodimerization is compromised, as is interaction with Rad50 and Xrs2 (Fig. 2E). This indicates that disruption of the contacts with Mre11-T300 and -E299 alters the overall structure of Mre11 and thereby has global effects on the structure of the Mre11 complex.
Protein-dsDNA contacts
The DNA binding contacts of the eukaryotic Rad50-dsDNA complex from C. thermophilum have been previously described19. In our structure of the S. cerevisiae MR-dsDNA complex, we noted that 18-bp of dsDNA [modeled as an (dA)n-(dT)n duplex] is encapsulated by elements of the pair of Rad50 domains in the MR-dsDNA complex (Figs. 1B, C and 4A). The 18-bp DNA footprint of MR is consistent with that seen for the C. thermophilum Rad50-dsDNA19 and E. coli MR (SbcCD)-dsDNA10 complexes. The dsDNA is positioned above the pair of Rad50 head domains and sandwiched between the pair of Rad50 head-proximal coiled coil arms, anchored in place through a network of hydrogen bonding interactions. We can trace intermolecular hydrogen bonds between Rad50 side chains (-T111, -S169 and -R1201) and main chain (-N58, -F109, -T111 and -S169) with backbone phosphates of bound dsDNA (Fig. 4A, boxed panel). Notably, the side chain of Rad50-K60 is inserted deep into the minor groove of bound dsDNA and is likely positioned to form hydrogen bonds with bases of dsDNA. The bound dsDNA is positioned within a basic channel lined primarily by lysine side chains originating from the pair of encapsulating Rad50 domains in the complex (Fig. 4B).
A The boxed panel shows details of intermolecular protein-DNA contacts (dashed black lines) between the side and main chain residues of Rad50 and phosphates of dsDNA in the complex. The basic side chains lining the positively-charged DNA binding pocket are shown in stick representation, except for the untraceable side chains of Lys1181 and Lys1183, which are shown in ball representation. B The dsDNA is encapsulated within a basic channel lined primarily by basic lysine and arginine residues. Two views rotated by 90° of the surface electrostatic potential of MR in the region surrounding the bound dsDNA in the complex, with positively charged basic surfaces labeled and shown in blue and green from each Rad50 protomer. C CPT-survival of rad50 mutants targeting potential dsDNA binding residues assessed at 30 °C and 37 °C. D CPT-survival of rad50 mutants targeting potential dsDNA binding residues (panel C) in rad50-R1201A background assessed at 30 °C and 37 °C. E Quantitation of DNA binding of WT and Rad50-mutant proteins. Indicated amounts (0–600 nM) of purified Rad50 proteins (see Fig. S4A) were incubated with 5 nM 83-mer dsDNA (same substrate as used in cryo-EM) in the presence of 2 mM cold ATP, 0.1 mM γ32P-ATP and 5 mM MgCl2. Bound and free substrates were separated by the electrophoretic mobility assay (EMSA). Examples of EMSA gels are given in Fig. S4B. Data are presented as mean values ± SD. Error bars denote standard deviation from four (Rad50-WT and Rad50-R1201A) or three independent experiments (Rad50-K60A, -R131A, -K173A K174A, -K1181A K1183A, -3KA, -K60A R1201A, -R131A R1201A and -K1181A K1183A R1201A), respectively. Source data are provided with this paper. F Mre11 complex integrity of Rad50-dsDNA binding residue mutants assessed by Rad50-and Mre11-co-immunoprecipitation (labeled Rad50-IP; Mre11-IP) and Rad50 and Mre11 Western blotting (anti-Rad50; anti-Mre11). The integrity of rad50-single and double mutants were assessed both in rad50-R1201A (blot on left side) and WT (R1201)-background (blot on right side). Representative images from one of three independent experiments with similar results are shown. Numbers on the left indicate molecular size markers (in kDa). Uncropped Western blots are provided as a source data file. G CPT-survival of rad50-K192A, K195A and K196A single, as well as double mutants and triple mutant (rad50-3KA) assessed at 30 °C and 37 °C.
It is well established that defects in dsDNA binding by Rad50 confer sensitivity to clastogenic insult. Therefore, we assessed CPT sensitivity of alanine mutants of dsDNA-interacting residues described above. As with alterations of protein-protein interfaces, phenotypic assessment was carried out at 30 °C and 37 °C, because partial destabilization of Rad50 dsDNA binding may confer a temperature-sensitive phenotype.
Basic residues within the Rad50 channel that point towards the dsDNA phosphate backbone and are situated within 2 to 4 Å were changed to alanine (Fig. 4A, boxed panel). With the exception of rad50-R1201, single mutations of these residues had virtually no impact on CPT sensitivity (Fig. 4C). We purified Rad50 WT and mutant proteins from yeast cells (Fig. S4A) and assessed Rad50 ATP-dependent dsDNA binding by EMSA using a radiolabeled 83-mer dsDNA substrate (Fig. S4B). Rad50-K60A, -R131A, -K173A K174A, -K1181A K1183A and -R1201A exhibited essentially WT dsDNA binding activity (Fig. 4E and S4B), consistent with the lack of CPT sensitivity (Fig. 4C). Some of these residues are conserved from archaea to human and were previously shown to interact with dsDNA13,15,19. They include Rad50-K60, -K103, -K104, -R131 and -R1201 and were previously assessed phenotypically following charge reversal substitution with glutamate and shown to impart strong defects in DNA repair15,19. The mutants shown in Fig. 4C were next combined with rad50-R1201A. All of the mutant combinations were nearly as CPT sensitive as rad50Δ, with the exception of rad50-K103A K104A R1201A, rad50-K186A R1201A and rad50-S169A R1201A (Fig. 4D). As expected, this was correlated with reduced dsDNA binding (Fig. 4E and S4B).
Decrements in DNA binding were not due to loss of Mre11 complex integrity (Fig. 4F). Rad50 protein levels were comparable in any single and R1201A double mutants. Interactions between Rad50 and Mre11 were slightly reduced in rad50-K103A K104A R1201A, rad50-T111A R1201A and rad50-K192A R1201A and were partially impacted in rad50-K195A R1201A (Fig. 4F). Nevertheless, these data support the view that dsDNA binding defects underlie the phenotypes observed.
A transient Rad50 DNA binding site?
As described below, Molecular Dynamics (MD) simulations suggested that residues of Rad50 somewhat distal to the dsDNA binding domain (Rad50-K192, -K195 and -K196) in the 3.2 Å structure of the MR-dsDNA complex (Fig. 4A, boxed panel) may transiently engage DNA prior to the binding mode observed in the structure.
The Rad50-K196 side chain points towards the dsDNA at a distance of 4.3 Å from the backbone phosphate, while Rad50-K192 and -K195 positioned on the same helix, are rotated away from the DNA phosphate backbone by a distance of 5.3 and 10.1 Å, respectively (Fig. 4A, boxed panel). Rotation of the coiled coils, suggested by MD simulations (see below), may bring Rad50-K192 and -K196 into contact with the dsDNA, at least transiently, while K195 has a structural role in positioning K192 and K195 for DNA-interaction.
To address the significance of these residues, alanine substitutions were carried out and assessed phenotypically. The single alanine substitutions had very little impact on CPT sensitivity. In contrast, the triple mutant rad50-K192A K195A K196A (hereafter referred as rad50-3KA) was almost as sensitive as rad50∆ (Fig. 4G). This strong CPT-sensitive phenotype was not due to a compromised Mre11 interaction, nor disruption of Rad50-3KA dsDNA binding, as WT-levels of Rad50-3KA were immunoprecipitated with Mre11 (Fig. 4F), and Rad50-3KA dsDNA binding was unaffected in vitro (Fig. 4E and S4B).
Intragenic suppressors of rad50-3KA
We performed an intragenic suppressor screen to identify mutations that could bypass the rad50-3KA phenotype, perhaps by promoting the handoff of dsDNA from Rad50 to Mre11, an event that would not be detectable in vitro. A centromeric plasmid containing the rad50-3KA ORF was chemically mutagenized and transformed into rad50Δ/RAD50 diploid cells. Following sporulation, random rad50Δ spores containing the mutagenized plasmid were recovered. Among 960 colonies analyzed for growth on selective plates containing 30 µM CPT, 18 clones showed increased resistance to CPT relative to rad50-3KA (Fig. 5A).
A CPT-survival of 18 intragenic rad50-3KA suppressors isolated in a genetic screen. Plasmids of the original screen suppressor clones were recovered, sequenced and re-transformed in a rad50Δ strain. CPT-survival after retransformation is shown. Plates were either incubated at 30 °C or 37 °C. B Localization of the rad50-3KA suppressor mutations in Rad50 primary sequence (top) and on the Rad50 structure (bottom). Note that the P168S suppressor previously isolated in a screen suppressing rad50-46 hook mutant phenotypes20 was also independently isolated for rad50-3KA here. Residues A1206 and A1249 are not depicted. C Rad50-ATPase activity of purified WT and mutant MR complexes. Indicated concentrations (0–2 µM) of MR-proteins were incubated in the presence of both ss and dsDNA in a buffer containing cold and hot ATP and MgCl2. The hydrolyzed radiolabeled phosphate was separated from γ32-ATP by thin-layer chromatography. Data are presented as mean values ± SD. Error bars denote standard deviation from four experiments. Source data are provided with this paper. D Meiosis in W303+ background of WT, rad50-3KA without and with suppressors (P168S, E356K and S1173L). Sporulation efficiencies (top panel; percentage of tetrads of sporulated cells) and spore viability (bottom panel; determined by tetrad dissection) are shown. Representative pictures of DAPI-stained sporulated cultures are shown. The spore viability of rad50-3KA without suppressors could not be determined (ND) due to the lack of tetrads. Scale bars (5 µm) are denoted. E Meiotic DSB formation and repair by meiotic recombination at the HIS4-LEU2 hotspot by Southern blot. The migration level of the crossover recombinant fragments (Recs, recombinants; 5.6 and 4.6 kb, red arrows) above and below the parental band24,45 and the unprocessed 3.3 and 3.0 kb meiotic double-strand break fragments (DSB, green arrows) are denoted. Cells were cultivated in sporulation media (SPM) for 0, 4, 8, 10 and 24 (WT, rad50-3KA, rad50-3KA P168S, rad50-C1G and rad50-46) or for 6, 8, 12 and 24 hours (dmc1∆). One representative Southern blot from three independent experiments is shown. Molecular size markers were detected by the probe and are given on the left (in kDa). F Cell survival of RAD50 WT, rad50-46, rad50-3KA without and with P168S or E356K suppressor mutants in a Mec1- and/or Sae2-deficient background. All strains were in the mec1Δ sml1Δ background.
Ten of the eighteen rad50-3KA suppressors were located either within the Walker A and B motifs or in the proximal coiled coils (Fig. 5B, top panel); Walker A (P57A, M92I, D136N, P140S, P168S), Walker B (R1201K, A1206T, A1249T), the N terminal coils (V209I) and the C terminal coils (T1119I). Two are located on β hairpin proximal to head domains (S1173L, E1197K), while the remaining six lie within the coils distal to the Walker A and B motifs (E350K, E356K, T357I, E438K, E886K, and A953V). With the exception of P168S and R1201K, most suppressors in Walker A/B motifs were surface exposed and localized >10 to 20 Å below the bound dsDNA (Fig. 5B, bottom panel), indicating that suppression of rad50-3KA can be effected at mid- and long-range distances.
Mutations in the coiled coil regions as well in the Walker A and B motifs were previously shown to suppress the phenotype of a mutation in the Rad50 hook domain, rad50-46. Notably, P168S, which was recovered in the screen for rad50-3KA suppressors was also previously recovered in an independent screen for rad50-46 intragenic suppressors20.
Defects in Rad50 ATPase activity
ATP hydrolysis mediates the transition from the closed to the open form of the Mre11 complex13, which is a prerequisite for the engagement of dsDNA in the active site of Mre116. The rad50-3KA pronounced CPT sensitivity (Fig. 4G) and normal dsDNA binding (Fig. 4E), could therefore be attributable to a defect in ATP hydrolysis. To test that interpretation, we purified WT and mutant MR complexes (Fig. S4C) and measured ATP hydrolysis as a function of MR concentration (0-2 µM) after 90 minutes of incubation with ATP-γ32P (Fig. 5C and S4D). We observed at least a 50% reduction in ATPase activity of Rad50-46 compared to WT, whereas Rad50-3KA, Rad50-K60A R1201A and Rad50-R131A R1201A were as defective as the Rad50-EQ mutant (Fig. 5C).
P168S reverted the CPT sensitivity of both rad50-46 and rad50-3KA to that of WT (Fig. S5A). Although the ATPase activity specified by Rad50-46 is 2.5 higher than that of Rad50-3KA (Fig. 5C), P168S suppressed each by the same magnitude – roughly two fold. This result indicates that Rad50-46 and Rad50-3KA are both defective in the same DNA repair mechanism dependent on Mre11 nuclease activity and Sae2 (Fig. S5B and S5C).
In meiosis, the Mre11 nuclease activity is required to remove Spo11 from meiotic DSB ends21,22. rad50-46 accumulates unprocessed meiotic DSBs due to failure to remove Spo11. P168S rescued rad50-46 meiotic phenotypes essentially to WT levels20. Meiotic progression in rad50-3KA was assessed (Fig. 5D, top panel). Essentially no tetrads could be detected in rad50-3KA sporulated cultures, and DAPI signals appeared heavily fragmented as previously observed in spo11Δ/Δ cells and resulting in meiotic chromosome missegregations23. This meiotic defect was rescued almost to WT levels by all three rad50-3KA suppressors tested (P168S, E356K and S1173L; Fig. 5D, top panel), with both sporulation efficiencies (14-17%) and spore viabilities (90-93%), comparable to WT (24% and 95% respectively; Fig. 5D, bottom panel).
We asked whether rad50-3KA meiotic defect is due to a failure to remove Spo11 from meiotic DSBs. A rad50-3KA strain without and with P168S was constructed and used to visualize unprocessed meiotic DSBs and meiotic DSB repair products (recombinants)24. P168S rescued the rad50-3KA meiotic DSB processing defect as previously observed for rad50-4620, evident by the bands now appearing to represent the meiotic recombinants products (Fig. 5E). A rad50 hook mutant defective in zinc coordination (rad50-C1G), and a Rad50 ATPase catalytic mutant (rad50-E1235Q), failed to efficiently form meiotic DSBs and did not exhibit appreciable unprocessed DSBs or recombinants (Fig. 5E and data not shown), and did not produce viable spores (data not shown).
The Rad50-3KA ATPase defect is more severe than that of Rad50-46 (Fig. 5C), making it likely that additional functions may be compromised. The rad50-46 mutant is defective in Tel1 activation20, as inferred from the inability of rad50-46 sae2Δ to suppress Mec1 deficiency25. That phenotype is suppressed by P168S (Fig. 5F). Defective Tel1 activation results in telomere shortening, and comparable telomere lengths were observed in rad50-46 and rad50-3KA. Whereas P168S restored normal telomere length to rad50-46, it failed to do so in rad50-3KA (Fig. S5D). P168S suppresses the ATP hydrolysis defects in rad50-46 and rad50-3KA by approximately the same magnitude (two fold), but the latter double mutant remains two fold less active than the former. These data indicate that a threshold of ATPase activity may be required for Tel1 activation (Fig. 5F).
Molecular modeling and MD simulations of MR bound to dsDNA
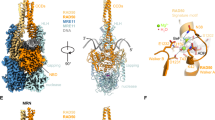
Only a small segment of the head-proximal coiled coils of Rad50 were resolved in the cryo-EM structure of the S. cerevisiae MR-dsDNA complex (Fig. 1B, C). Thus, we constructed a model of full-length S. cerevisiae MR-dsDNA complex (Fig. 6A), using the computational protocol described in the Methods section (Figs. S6A-S6L and S7A-S7C).
A, B The topology of the full-length (panel A) and coiled-coil truncated (B) DNA bound MR complex. C, D Details of amino acid K192, K196, K1181, D1128, and P168 side chain conformations before (C) and after (D) 30 ns of MD simulations. Salt bridges are shown as dashed lines. The side chain of P168 is shown in space-filling representation. Lysine residues involved in hydrogen bond formation are shown by red arrows.
To facilitate MD simulations, we constructed a model of the MR-dsDNA complex containing truncated Rad50 coiled-coils ending with the Zn hook (Fig. 6B), using the protocol described in the Methods section (Fig. S7D and S7E).
The MR-dsDNA complex containing truncated coiled-coils was subjected to MD simulations (Fig. 6C, D). We observe a coaxial rotational movement of the Rad50 coiled coils along their axes in the WT MR-dsDNA complex during the 30 ns simulation (Movie S1). The dsDNA is initially positioned with K196 of individual Rad50 monomers hydrogen bonded to the phosphates of opposing strands of the DNA duplex (Fig. 6C). At the end of 30 ns, K192 and K196 (on an α-helix) and K1181 (on a β-hairpin) from one Rad50 monomer (in light green) form an asymmetric interaction with consecutive phosphate backbone of one strand of the DNA duplex (Fig. 6D). The mild CPT sensitivity of K195A (Fig. 4G) is likely due to the loss of the salt bridge between the side chains of N-terminal K195 and C-terminal D1128 of Rad50 (Fig. 6C, D) that provides rigid support for the interactions of K192 and K196 with consecutive phosphates on one DNA strand (Fig. 6D). It should also be noted that Rad50 β-hairpins (Thr1168 to Lys1193; shown in light and dark magenta colors in Fig. 6C, D) move toward the same minor groove of dsDNA from two opposite Rad50 monomers, albeit closer to a different DNA strand, at the end of the 30 ns MD simulation, as a result of which K1181 (on the light magenta colored β-hairpin) is positioned to form a hydrogen bond with the backbone phosphate (Fig. 6D).
ATP-hydrolysis is restored in Rad50-3KA P168S
MD simulations were carried out using coiled coil truncated WT, Rad50-3KA and Rad50-3KA P168S MR-dsDNA complexes (Fig. 7). Hydrolysis of the ATP β-γ phosphodiester bond occurs via nucleophilic attack of an activated hydroxyl ion of a water molecule, resulting in hydrolysis of the β-γ bond and release of the γ-phosphate26,27. Conserved Rad50-D1234, -E1235, -H1272 and Q158 line the ATP-binding pocket and mediate hydrolysis (Fig. 7A).
(A) Sequence alignment of the catalytic residues of three ABC ATPases. The conserved D, E and H residues in ABC-transporters of Hemolysin B, C. thermophilum and S. cerevisiae are highlighted in blue background. (B-D) The conformations of catalytically-competent amino acid side chains at the beginning (left panel) and end (right panel) of 30 ns MD simulation of the dsDNA-bound truncated MR complex: WT (panel B), Rad50-3KA (panel C), and Rad50-3KA P168S (panel D). Dashed lines indicate the distances between negatively charged oxygen (as computed by the Schrödinger program) on the catalytically competent E1235 and the positively charged Mg2+ and ammonium group of K40. The negatively charged carboxylate oxygens of D1234 and E1235 residues are shown by blue and red arrows, respectively. The partially negatively charged amide oxygen of Q158 is shown by a green arrow.
We carried out 30 ns MD simulations on truncated versions of WT (Fig. 7B), Rad50-3KA (Fig. 7C) and Rad50-3KA P168S (Fig. 7D) MR-dsDNA complexes. Side chain alignments of Rad50-E1235 (red arrow), -D1234 (blue arrow) and -Q158 (green arrow) with triphosphate chain of bound ATP are shown before and after 30 ns. Similar side chain alignments of D1234, E1235 and Q158 lining the ATP pocket were observed for WT (Fig. 7B, right panel) and Rad50-3KA P168S (Fig. 7D, right panel) at the end of the 30 ns. The same side chains align differently in the Rad50-3KA MR-dsDNA complex at the end of the simulation (Fig. 7C, right panel). The relevant distances between the side chains and Mg2+ before and after the 30 ns MD simulation and the sum of the differences in their conformational changes are summarized in Table S2. Notably, the partially negatively charged amide oxygen on the side chain of Q158 retains its position in WT (Fig. 7B, right panel) and Rad50-3KA-P168S (Fig. 7D, right panel) but undergoes a large shift away from the Mg2+ at the end of the 30 ns simulation in Rad50-3KA (Fig. 7C, right panel). In contrast, the negatively charged atoms in the Rad50-3KA P168S have become closer to the catalytic center by 11.1 Å (Table S2). Hence, the ATP hydrolysis mechanism is effectively restored to that of wild type Rad50 in Rad50-3KA P168S.
Rotation of Rad50 coiled coils
The 30 ns MD simulations of WT (Movie S1) and Rad50-3KA P168S (Movie S3) also revealed a clockwise coaxial rotation of the coiled coils (as viewed from the Rad50 globular domain toward the Zn hook). Clockwise rotation was also observed for Rad50-3KA, but the trajectory deviated from a coaxial rotation (Movie S2) revealing the impact of Rad50-3KA on the coiled coils. Notably, the same direction of rotation was also observed in the WT MR-dsDNA complex with full-length Rad50 coiled coils; thus, it was not the result of the truncation. Morphing the transition from DNA-free C. thermophilum MR complex with rod-like coiled coil11 to dsDNA-bound complex presented here is also accompanied by clockwise rotation of the coiled coils (data not shown).
Discussion
Here we report on a cryo-EM structure of the S. cerevisiae MR-dsDNA complex at 3.2 Å resolution, partially augmented by molecular modeling and molecular dynamics simulations. Our detailed structure- function analysis of protein mutants revealed new features of the Rad50-dsDNA interaction interface, as well as novel inter- and intra- molecular protein-protein contacts that are critical for the assembly and dynamics of the MR-dsDNA complex. Data obtained over the last ten years indicate formation of two major states of the MR-dsDNA complex, namely an ATP bound “closed” complex with dsDNA bound by Rad50 dimers important for DNA end stabilization/sensing and activation of the Tel1 checkpoint kinase, and an “open” complex which forms upon ATP hydrolysis, and in which the Mre11 nuclease active site is accessible to the dsDNA substrate28.
The structure of the stable S. cerevisiae MR-dsDNA complex presented here appears to represent a “closed” complex, having a rod-shaped structure that is similar to that recently reported for the C. thermophilum MRN complex in presence of ATPγS, but without DNA11. The structure presented here also strongly resembles a recent structure of the human complex bound to dsDNA29. Of the 83-mer dsDNA used for complex formation in our study, 18 base pairs of duplex DNA are bound in a central basic DNA binding groove formed by the ATP-bound Rad50 dimer, with the Rad50 coils encapsulating the dsDNA from the top (Figs. 1B, 1C, 4A and 4B). The dsDNA length and Rad50 DNA interacting residues align well with the previously reported crystal-structure of the dsDNA bound T. maritima Rad5019, with the exception that the coils remained open in that structure. It is thus conceivable that the closed structure presented herein and that of the T. maritima represent different stages in the engagement of dsDNA by Rad50.
The distal coiled coils of Rad50 were not visible in our cryo-EM structure of the MR-dsDNA complex (Figs. 1B and 1C), but molecular modeling (Fig. 6A) suggests that the full-length coiled coils are in a closed configuration, essentially spanning the segment from the globular to the hook domain but punctuated with short flexible regions which are analogous to the “hinge” loop (CGSQD) identified in the human Rad50 hook structure12. This is consistent with atomic force microscopy (AFM) studies of the human MR complex which showed that upon dsDNA binding, the coiled coils adopt a rod-shaped parallel conformation30. Hence, it appears that at least partially opened coils are required for initial DNA binding by Rad50, likely loaded proximal to the head domains of Rad50, in agreement with our previous Rad50 hook structure, as well as AFM data regarding the human complex12,31.
The major point emerging from this study is that the coiled coils influence Rad50 ATP hydrolysis. It is clear from previous work that ATP hydrolysis by Rad50 governs the transition from the “closed” to “open” forms of the complex13,28. Hence, there is a reciprocal relationship between ATP hydrolysis and the disposition of the coiled coils. The Rad50 ATPase in Rad50-3KA is essentially equivalent to the Rad50-E1235Q hydrolysis defective mutant (Fig. 5C), and blocks cleavage of Spo11 from meiotic DSB ends (Fig. 5E). Spo11 cleavage, and therefore the ATPase activity of Rad50-3KA appears to be restored by E365K (in the coiled coils) and S1173L (in the conserved β-hairpin loop11), both of which suppress the spore inviability of Rad50-3KA (Figs. 5C and 5D). Additional suppressors or Rad50-3KA were identified in the coiled coils underscoring the mid- and long- range influence of the coiled coil domain (Figure S8). In this regard, it is perhaps notable that recent proteomic data reveal multiple Tel1 phosphorylation sites exclusively within the Rad50 coiled coils32, suggesting a mechanism for transient modulation of coiled coil behavior.
A previously described Rad50 hook domain mutation, rad50-46 exhibits CPT sensitivity that was suppressed by two mutations in the coiled coils, N607Y and N873I20. Given the role of Rad50 ATPase activity in potentiating Mre11 nuclease function, we presume that the ATPase defect in rad50-46 (Fig. 5C) underlies both the CPT sensitivity and defects in meiotic DSB processing observed in that mutant20. In addition to suppressors in the coiled coils, both rad50-3KA and rad50-46 are suppressed by P168S in the Rad50 head domain. It is remarkable that rad50-46 and rad50-3KA, which are at least 300 Å apart, are both suppressed by P168S which lies within the Walker A domain and very far from the hook domain.
Extensive mutational analyses and MD simulations reveal previously unappreciated dynamics of the coiled coils as they engage DNA. The simulations suggest that the Rad50 coiled coils undergo a clockwise coaxial rotation (Movie S1) which coincides with transient contacts between Rad50 K192 K195 K196 (Rad50-3K residues) and dsDNA. This rotation appears to impel the DNA deeper into the cleft formed below the coiled coils, ultimately leading to the disposition captured in the cryo-EM structure of the complex. This “zippering” motion of the coiled coils around DNA is reminiscent of that recently described for the SbcCD complex10,11. The Rad50-3KA containing Mre11 complex does not exhibit defects in DNA binding in vitro, consistent with the view that the interaction of dsDNA with Rad50 K192 K195 K196 may be transient and does not represent a stable DNA binding surface. In the C. thermophilum MRN complex structure lacking dsDNA, residues corresponding to Rad50 K192 K195 K196 face into the channel between the coiled coils and interact with a β-hairpin loop that is not well conserved in sequence11,19, but present in all Rad50 structures so far. That most of the β-hairpin loop is no longer visible in the structure of the MR-dsDNA complex presented here, suggesting that it folds away upon DNA binding or becomes disordered (Movie S1).
The residues at the Rad50-dsDNA interface are highly conserved; many residues assessed here were previously reported residues in E. coli and T. maritima9,10,19, and are in close proximity to the 18 base pair DNA duplex in our structure. Previous studies made charge reversal mutants in various yeast Rad50 DNA binding residues, resulting in CPT sensitivity and in reduced DNA binding in vitro15,19. Alanine substitutions of these residues were individually inconsequential, as mutants showed WT CPT-survival and dsDNA binding (Fig. 4C, E). In agreement with these previous reports, when two or more residues that included R1201A were mutated (Fig. 4D), these mutants were strongly sensitized to CPT almost to the level of rad50∆ and showed reduced dsDNA binding in vitro (Fig. 4E) underscoring the fact that multiple contacts between Rad50 and dsDNA contribute to DNA binding. DNA binding stimulates Rad50-ATPase. Accordingly, we found that both Rad50-K60A R1201A and Rad50-R131A R1201A showed ATPase activities reduced to the levels of Rad50-E1235Q ATPase deficient mutant (Fig. 5C).
As noted above, at the end of 30 ns MD simulations, three lysine residues of Rad50 were identified that either directly interacted with the phosphodiester backbone of the dsDNA (K192 and K196) or stabilized the coils through interaction between K195 and D1128 (Fig. 6D); however, in our structure, these are too far situated (>4.5 Å) to reveal a DNA interaction. Both K192 and K195 are highly conserved in evolution and are found to be repeatedly mutated (24 tumors in MSK IMPACT analyses33), while K196 is not conserved in mammals. Within the E. coli MR complex, two lysine residues (K194 and K890) interact with DNA and charge reversal mutants showed also reduced DNA binding10. The crystal structure of T. maritima Rad50-dsDNA revealed three lysine residues within the coils (K175, K178 and K182) that interacted with the DNA and charge reversal mutations also resulted in reduced DNA binding15. Except K182, which aligns with K192, these residues do not align with the S. cerevisiae K192, K195 and K196, but are located in the same part of the coils (α7 helix). In this context, we propose that the S. cerevisae Rad50 K192, K195 and K196 residues, amongst others, allow the coils to properly position the duplex DNA in the basic DNA binding groove which is required to activate the Rad50 ATPase. The failure to activate the Rad50 ATPase would in turn block opening of the complex to provide DNA access to the Mre11 catalytic center10,34.
Collectively, the data reported here strongly illustrate the influence of the Rad50 coiled coil domains on the enzymatic activities specified within the globular domain of the Mre11 complex. Moreover, the extensive structure-based functional analyses presented provide a highly granular view of the interfaces and residues that mediate complex assembly as well as function.
The extended coiled coil structure is a conserved feature of Rad50 across all clades of life. This begs a question not addressed here: why are the coiled coils so long? We showed previously that shortening of the coiled coils compromised Mre11 complex function, even when Rad50 with shortened coils was co-expressed with Rad50 having a corresponding increase in coiled coil length35. This indicates that the coiled coil length per se is not paramount. Recent work suggests that bridging of sister chromatids during DSB repair is mediated via coiled coil interactions in trans11. That mode of bridging with S. cerevisiae Rad50 could span up to 120 nm. That distance is far greater than would be required to accommodate two 30 nm fibers of heterochromatic stretches of chromatids, but it is conceivable that loops or other structures of sister chromatids in euchromatic regions could necessitate a longer coiled coil structure to effect tethering during DSB repair. As the granularity of the globular domain structure increases, and the influence of the coiled coils becomes clearer, the need to understand the dynamics and structure(s) relevant to the Rad50 coiled coils becomes more acute.
Methods
Purification of S. cerevisiae Mre11-Rad50 complex
S. cerevisiae MREQ was overexpressed in yeast upon addition of 2% galactose for 18 h in Do-Trp-Leu lactic acid media containing 1% sucrose. Induced yeast cells were pelleted and lysed using a freezer Mill in MR-buffer (50 mM HEPES pH 7.6, 300 mM NaCl, 10% glycerol, 0.025% CHAPSO, 2 mM DTT, 1 mM PMSF and Roche cOmplete™ Protease Inhibitor Cocktail). The clarified yeast extract was purified by FLAG-affinity followed by Heparin. The 800 mM NaCl Heparin-eluate was buffer exchanged to 200 mM NaCl on a Micro Bio-SpinTM 6 column (Bio-rad).
The MREQ sample was concentrated and applied to a Superpose 6 Increase column equilibrated with 200 mM NaCl, 50 mM HEPES, 2 mM DTT, pH 7.5, containing 2% glycerol and 0.025% CHAPSO (3-([3-Cholamidopropyl]dimethylammonio)-2-hydroxy-1-propanesulfonate) to obtain peak fractions containing the complex as shown in Figure S1B.
Rad50 expression and purification from yeast cells and DNA binding assays
Rad50-1FLAG proteins were expressed and purified as previously described18. Electrophoretic Mobility Shift Assay (EMSA) was performed as described before18.
Mre11-Rad50 ATPase assay
MR-complexes were expressed in yeast and purified as described for MREQ by FLAG-affinity only without the Heparin binding and elution step. Protein concentrations were measured using the Bio-Rad DC Protein Assay using the FLAG-elution buffer as blank reference. The ATPase reaction mixture (5 µl) contained the MR-proteins of the indicated concentrations (0–2 µM) in ATPase buffer (25 mM HEPES pH 7.6, 100 mM NaCl, 1 mM DTT, 10 mM MgCl2, 0.1 µM γ-32P-ATP, 2 mM ATP, 50 µM ssDNA (a 50 nt long oligonucleotide) and 400 ng dsDNA (1 kb). After 90 min at 30 °C, 0.8 µl was loaded on a thin layer chromatography (TLC) plate (TLC PEI Cellulose F, EMD Millipore). The air-dried TLC plate was developed in a mobile phase of 0.5 M lithium chloride and 0.5 M formic acid and exposed to a phosphor-imager screen. The percentage of ATP hydrolysis was calculated as ratio of released Pi/total radioactivity per lane following quantification with ImageGauge software (GE).
Yeast strains and manipulations
Mutants were introduced by site-directed mutagenesis in the plasmids Ycp50-RAD50 or pRS316-ADE2-MRE11. Yeast strains used in this study are given in Table S3. The rad50 and mre11 mutant ORFs were confirmed by sequencing. The URA3 plasmids were transformed in JPY1836 (rad50∆KanMX ura3-) or JPY7128 (mre11∆LEU2 ura3-) and plated on Do-Ura plates. Single clones were picked and grown overnight in Do-Ura media. For damage- sensitivity assays, five-fold serial cell dilutions (250,000 - 80 cells per spot) were spotted on plates without or with camptothecin (CPT) and incubated for 2 days at 30 °C or 37 °C. To assess Mre11 and Rad50 protein levels and complex integrity, 50 ml of YPD media was inoculated with yeast Do-Ura cultures and grown at 30 °C or 37 °C to exponential phase. Yeast cell lysis (Buffer: 25 mM Tris-Cl pH 7.6, 150 mM NaCl, 10% Glycerol, 0.4% Igepal, 1 mM EDTA, 1 mM DTT, 1 mM PMSF, Complete Protease Inhibitor Cocktail), immunoprecipitations and western blotting were done as described before35. Western blotting was performed using rabbit anti-Rad50 (#64911), rabbit anti-Mre11 (#59567) and rabbit anti-Xrs2 (UWM45) antisera and protA/G-HRP and the target proteins were detected using ECL prime (Cytiva).
Screen for intragenic suppressors of rad50-3KA CPT-survival
10 µg of Ycp50-rad50-3KA plasmid was chemically randomly mutagenized by incubation for 90 min at 70 °C in pH-adjusted (pH 7.0) freshly prepared 0.5 ml 1 M hydroxylamine solution. The plasmid DNA was re-purified using Qiagen Miniprep columns according to QIAprep Miniprep Handbook. The chemically modified plasmid was transformed in Bacteria, and the Ycp50-rad50-3KA* random mutant library purified by Maxiprep (Qiagen). The library was transformed in a heterozygote diploid Magic strain JPY8425 (rad50∆KanMX/RAD50 lyp1∆::pSTE3-LEU2/LYP1 can1::pSTE2-Sp-HIS5/CAN), allowing following sporulation selection of rad50∆ Ycp50-rad50-3KA* random spores on Do-Ura-Leu-Arg-Lys +G418 (250 µg/ml) +Canavanine (200 µg/ml) +Thialysine (200 µg/ml). 960 random spores were assessed for survival in presence of 30 µM CPT. From 22 surviving clones the plasmids were recovered and rad50-3KA* ORF sequenced to determine the intragenic suppressor mutations. 18 plasmids retained the rad50-3KA allele along with single-intragenic suppressor mutations and following re-transformation in JPY1836 (rad50∆) were confirmed to suppress rad50-3KA CPT-survival.
Genetic assessment of rad50-3KA without and with intragenic suppressors
To assess meiotic phenotypes, a rad50∆ /∆ homozygote diploid (JPY8566) was transformed with Ycp50-RAD50, Ycp50-rad50-3KA, Ycp50-rad50-3KA-P168S, Ycp50-rad50-3KA-E356K and Ycp50-rad50-3KA-S1173L and transformants on Do-Ura+G418 plates were grown oN in Do-Ura+G418 media, cells were pelleted and washed with water, then incubated for 16 hours in Do-Ura pre-sporulation media, followed by 42 hours in sporulation media. Sporulated cultures were fixed with 80% ethanol, washed with water, DAPI-stained and the percentage of tetrads were determined by counting tetrads with visible 3 or 4 DAPI-stained spores.
To assess telomere lengths, a RAD50/rad50∆ heterozygote diploid strain (JPY8425) containing the Ycp50-rad50 plasmids were sporulated and random spores selected as described above. Two-PCR genotyped and verified single clones of each genotype were grown for about 20 generations in Do-Ura+G418 media. Genomic DNA was isolated, PstI-digested and run on a 1.5% agarose gel. Telomere lengths were determined by Southern-blot with a telomere specific probe as described previously35.
To assess MMS-survival in Mec1-deficient cells, random spores were obtained from a heterozygote diploid strain (JPY7361; rad50∆/RAD50 mec1∆/MEC1 sml1∆/SML1) containing the indicated Ycp50-rad50 plasmids. To assess MMS-survival in Mec1- and Sae2-deficient background, a rad50∆ mec1∆ sml1∆ sae2∆ spore (JPY2137) was transformed with the Ycp50-rad50 plasmids.
To assess CPT-survival in mre11-H125N background, a rad50∆ mre11-H125N spore (JPY8574) was transformed with the Ycp50-rad50 plasmids.
Cryo-EM analyses of ATP-bound MR-DNA complex
MREQ (0.5 mg/ml) was incubated with 0.25 mM ATP, 0.5 mM MgCl2, 0.25 mM MnCl2 and 0.5 μM dsDNA (5’TTGATAAGAGGTCATTTTTGCGGATGGCTTAGAGCTTAATTGCTG AATCTGGTGCTGTAGCTCAACATGTTTTAAATATGCAA-3’ and its complementary strand) in 200 mM NaCl, 50 mM HEPES, 2 mM DTT, pH 7.5, containing 2% glycerol and 0.025% CHAPSO for 1 hr at 4 °C. This sample was applied onto glow-discharged UltrAuFoil 300 mesh R1.2/1.3 grid. Grids were blotted for 2.5 s at 4 °C, under 100% humidity, and flash frozen in liquid ethane using a FEI Vitrobot Mark IV. All images were collected on a FEI Titan KRIOS electron microscope operated at an acceleration voltage of 300 kV with a Gatan K3 camera with a 1.083 Å pixel size. Movies were recorded in counting mode at an electron dose rate of 30 e − /pixel/s with a total exposure time of 2 s and frames rate of 40 ms/frame, for an accumulated electron dose of 51.22 e − /Å2. Motion correction was performed with MotionCor236, and contrast transfer function parameters were estimated by Ctffind437. All other steps of image processing were performed by RELION 3.038 and Cryosparc v3.3.039. After particles auto picking from 5820 images and initial 2D classification, a total of 4,181,422 particles were selected for 3D classification and the initial models were reconstructed from 2,000,000 particles. After four rounds of 3D classification, a total of 324,671 particles from one class were selected to reconstruct two initial models and 3D classification. Finally, 232,521 particles were selected for homogeneous refinement, NU refinement and local refinement to yield a 3.5 Å electron microscopy map of apo-MR in Cryosparc v3.3.039. After multiple rounds of 3D classification, a total 242,558 particles were selected for homogeneous refinement, NU refinement and local refinement to yield a 3.2 Å electron microscopy map of MR-DNA complex in Cryosparc v3.3.039. All reported map resolutions are from gold-standard refinement procedures with the Fourier shell correlation cutoff being 0.143 criterion by applying a soft mask. Model-building of apo-MR and MR-DNA complex structures were performed manually based on the cryo-EM density map and computed model40 by using COOT441. The model was then refined against the cryo-EM density map using phenix.real_space_refine42. All figures were prepared by PyMOL (https://pymol.org) or UCSF Chimera43. Details of data collection, image processing and model building are shown in Table S1 and Figure S2.
Model building of MR complex containing full-length coiled coils
We have used AlphaFold2 (AF2) to build the Rad50 monomer, Rad50 dimer and Mre11 dimer and then used the experimentally derived cryo-EM structure of the MR-dsDNA complex as the core for the building of a more complete MR complex in which the missing elements were incorporated. The program computed a complete Rad50 monomer (Fig. S6A), but for the Rad50 dimer, it generated several fragments encompassing head domains with short coiled coils, ending in the range E252 to L288 in the N-terminal and in the range from L1029 to Q1061 at their C-terminal ends. Three groups of predicted dimeric models ranked from 0 to 24, were characterized by one or two superhelical twists of an individual coiled coil, with the ends of coiled coil fragments of the two monomers being apart to a varying degree from each other (Fig. S6B, S6C) and bent inwardly (Fig. S6B) or outwardly (Fig. S6D) in each of the three models. We used the dimeric arrangement of the head-domain with the bound ATP molecules and extended this arrangement with the coiled coils built by AF2. For these extensions, we ran separate AF2 predictions. For the dimeric segment from A181 to M1139 (Fig. S6E) the AF2 predictions resulted in divergent structures encompassing the sequence range from F400 through T951. Therefore, for the diverging part, we ran AF2 prediction covering the region I250 to Q526 on the N-terminus and N824 to T1069 on the C-terminus. The loop from Q526 to N824 was excluded in this computation, and therefore, we ran a tetrameric prediction for this region. The results were well-converged. A pair of chains held with each other by numerous hydrophobic interactions has been treated as representing a fragment of an individual monomer of the Rad50 dimeric molecule (Fig. S6F). At this point, we have taken steps to straighten up the coiled coils stitched together and to symmetrize the Rad50 dimer. This was achieved by iterative computing an average between two monomers superpositioned by their Cα atoms and using this average as two individual monomers in the next iteration (Fig. S6G).
Next, we assembled the Mre11 + Rad50 as the complex, using the framework of the cryo-EM structure, without the HTH segment. We undertook symmetrization steps for this tetramer of two heterodimers (Fig. S6H) achieving r.m.s.d 0.003 Å by Cα atoms of two superpositioned symmetric halves of the complex (Rad50 monomer + Mre11 monomer, 974 superimposed atoms in each half).
After augmenting the Rad50 dimer built so far and ending at Q526 to N824 with a computed as a tetrameric fragment from L464 to T890 converted to a dimer based on hydrophobic interactions, we introduced the apical Zn hook segment. This modeled Zn hook segment we obtained after converting the human Zn hook crystal structure (PDB: 5GOX) into its S. cerevisiae sequence homolog. A β-hairpin which mediates the interaction between the head domains and coiled coils of Rad50 (from T1168 to K1193) that was disordered in the cryo-EM structure and was introduced in the position observed in the C. thermophilum Rad50 structure using an AF2-generated module where it extends along the interface between the N-terminal and C-terminal fragments of the coiled coil at the point of their contacts with head domain19. We ran another symmetrization round for the complete Rad50 dimer situated on Mre11 dimer.
The Rad50-binding domain of Mre11, Mre11 HTH - Rad50 interface, was introduced using segment computed by AF2 with high confidence. This interface was not well-resolved in the cryo-EM structure. We positioned the D430 to E439 helical segment of the Mre11 connecting this HTH domain with the capping domain by docking computation [HDOCK]44 using this helix as a ligand and the interface of the assembled Rad50-Mre11 as a receptor. To connect this helix with the rest of Mre11, we have used the loop-building functionality of the Schrödinger software (Fig. S6I).
The insertion of DNA into the full-length Rad50 dimer required adjustment of coiled coils from their position in the truncated cryo-EM segment (Fig. S6J), dictated by the introduction of the β-hairpin and smooth extension of the coiled coils into their full-length form (Fig. S6K, boxed in Fig. S6H). We repaired the elements of the perturbed coiled coils by alignment procedures with corresponding elements originating from a full-length monomeric Rad50 AF2 model. The resulting full-length MR complex with dsDNA inserted is shown in Fig. S6L.
The all-Cα-atoms symmetry between chains A + D and B + C (two halves composed of Mre11 ([A and B] and full-length Rad50 [D and C] each) was in the final hybrid model characterized by an rmsd of 0.55 Å (for 1748 atoms), as compared to an rmsd 0.39 for the experimentally obtained cryo-EM structure of our MR-dsDNA complex with the coiled coils truncated just above the DNA (for 744 atoms).
We compared the position of ATP with coordinated Mg and catalytic water with those in the structure of C. thermophilum Rad50 (PDB: 5DA9), in which an oxygen of ATP is replaced the γS atom. The positions of the ATPs in our cryo-EM structure of the MR-dsDNA complex and 5DA9.pdb superposed well, except for the unresolved water molecules in the cryo-EM structure (Fig. S4E). In addition, we checked the conformation of the WT E1238 side chain in 5DA9.pdb that is engaged with γS atom to define the reverse substitution conformation Q1235E required to adequately study the effects of 3KA and 3KA P168S mutants on ATP hydrolysis by MD simulation.
The full-length of the extended Rad50 dimer, as measured between Cα of Y1312 and N696 of the same monomer, is 68.6 nM (Fig. S7A). Each folded-back monomeric antiparallel coiled coil is held by intramolecular hydrophobic interactions between the stretches of hydrophobic heptad repeats, whereas the intermolecular interactions between monomeric coiled coils are predominantly governed by the electrostatic, hydrogen-bonded and π-π interactions of constituent amino acid side chains at their interfaces. Partial residue labeling of the symmetric coiled coils is listed in Fig. S7A.
Going toward the turn at the Zn hook, there are eight alpha-helical stretches punctuated by intervening loops in each N-terminal arm (N180 to S291, K299 to F376, D386 to L464, T466 to E490, I492 to N557, D558 to F565, D575 to L637, E645-to D684, Fig. S7A). Going from the Zn hook toward the head domain, there are seven α-helical stretches punctuated by intervening loops in the C-terminal arm (N696 to T711, A713 to S790, I792 to S821, T832 to K975, G976 to R1047, D1049 to T1101, D1105 to T1161, Fig. S7A). The intervening loop sizes vary from as short as a single residue to as long as 12 residues. At the six superhelical left-handed crossings, a stretch of a continuous α-helix in the antiparallel coiled coil coincides with the intervening loop connecting α-helical stretches of the coiled coil partner running in the opposite direction (Fig. S7A, numbered 1 through 6). Except for a short helical segment from D558 to F565 running perpendicular to the helical axis, marked by an ‘x’ at crossing 5 in Figure S7A, all other helices form a braid-like interaction with their partners.
The two folded-back monomeric coiled coils run predominately side-by-side, but near the Zn hook region, their axes cross (Fig. S7B, marked l), resulting in a right-handed superhelical half-turn. Before this turn at the Zn hook, the projections of the axes cross twice, so that when viewing along the DNA axis, a right-handed crossing is observed at I430/I920 (Figure S7B, marked i), that is followed by a left-handed crossing at Y480/L877 (Fig. S7B, marked k). Notably, the segment A221/L1099 to A371/I1959 is a non-intertwined stretch. The overall half-helical right-handed turn close to the Zn hook is evident by the dark green chain switching the position at the right side of the molecule near to head domains to the position at the left side near the Zn hook domain (Fig. S7B).
Notably, among most of the contacts between identical helices of two monomeric coiled coils running in parallel, there are a few hubs (shown as red boxes in Fig. S7C) at which all four constitutive helices engage, resulting in an enriched pattern of the amino acid side chain interactions with enhanced interactions at residues G349-T406/G976-K948, K270-Q319/R1047-Q1000, and an extended stretch spanning L464-A592/D772-S887 near to Zn hook.
Model building of MR complex containing truncated coiled coils
In the full-length coiled coils thus constructed, we have noticed a structurally repetitive pattern centered at I430/I920 and A608/T764. To reduce the size of the molecule during MD simulations focused on ATP hydrolysis, we have produced a truncated version of Rad50 by superposition of the fragment A532 to I551 + N577 to E635 + H739 to D808 onto the fragment R346 to A371 + A388 to H449 + R903 to D971 by Cα atoms. In this alignment, the loops coordinating Zn turned by ~120° from their Zn-coordinating position. To reinstate Zn-coordination, the fragment of the full model with correct Zn hook geometry encompassing Cα atoms of residues Y646 to D684 + Y531 to N557 + I792 to F799 was superimposed onto the Cα atoms of the residues Y646 to D684 + R347 to H374 + D954 to F962 of the truncated version obtained in the previous step. After these operations, the peptide bonds could smoothly be generated between residues Y358 and N542 at the N-terminal and L795 and I959 at the C-terminal coiled coils, respectively (Figures S7D and S7E). This operation resulted in the removal of 2 × 184 + 2 × 164 = 696 residues of coiled coils without compromising their geometries (Fig. 6B). The computed rmsd of Cα atoms of the MR complex with shortened coiled coils onto full-length complex (2976 atoms aligned and 2168 atoms used in the final alignment) was 0.49 Å, indicative of a good representation of the shortened coiled coil version of the characteristics of the full-length model.
Model building of MR-dsDNA complex containing full-length coiled coils
We used a GC-rich DNA duplex from PDB: 6S85, as we considered it better suited for MD computations. The coiled coil contacts in the region around bound DNA in the full-length molecule differed from the shorter coiled coil fragment observed in our cryo-EM structure of the MR-dsDNA complex. We remodeled the region Q180 to N290/V1050 to Y1161 in the AF2 model obtained without dsDNA to introduce the tight contacts with dsDNA observed experimentally. In our cryo-EM structure of the MR-dsDNA complex, the geometry of coiled coil fragments S219 to E239/Q1089 to D1102 needed to be reconfigured to retain the DNA contacts while propagating into the full-length secondary structures of coiled coils distal to DNA-bound regions.
Model building of dsDNA bound Rad50-3KA and Rad50-3KA P168S complexes containing truncated coiled coils
We obtained the mutant series Rad50-3KA, as well as Rad50-3KA P168S, by mutating the corresponding residues of truncated dsDNA bound MR complexes with the WT sequence and running Protein Preparation routine on Maestro GUI of Schrödinger software until RMSD of all heavy atoms 0.3 Å. The Zn ion in the zinc hook was held in place via pseudo bonds with sulfur atoms of C687 and C690 formed during Protein Preparation. One Mg ion in the head domain was held in place via pseudo bonds formed with O2β and O2γ of ATP molecule, oxygens of two coordinated water molecules, and atoms Oε1 of Q158 and Oγ1 of T41 formed during Protein Preparation. The Mg ion coordinating the second ATP molecule was kept unbound. The amino acid side chain dynamics around ATP and Mg unbound with pseudo bonds were analyzed. The complexes prepared with no pseudo bonds for Mg ions in both ATP-bound sites and with both ATP + Mg molecules fixed by pseudo bonds in their corresponding binding sites have retained the rotation of the coiled coils. The amplitude of rotation decreased in the case of both ATP molecules fixed in their binding sites by pseudo bonds.
30 ns MD simulations on MR-dsDNA complexes containing truncated coiled coils
MD computations were run with DESMOND software on a High-Performance Computing Cluster with protocol as previously described18, except for the duration of the productive run, 30 ns. During system preparation for MD computations, Na and Cl atoms within 10 Å of ATP were excluded to avoid the direct effect of charged ions on ATP dynamics.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Source data are provided with this paper. The atomic coordinates of the S. cerevisiae apo-MREQ and dsDNA bound MREQ complex have been deposited in the Research Collaboratory for Structural Bioinformatics Protein Data Bank with the codes 9BI5 and 9BI4, respectively. Cryo-EM density maps of the S. cerevisiae apo-MREQ and dsDNA bound MREQ complex have been deposited in the Electron Microscopy Data Bank with accession codes EMD-44559 and EMD-44558, respectively. Any other data are available from the corresponding author upon request. Source data are provided with this paper.
References
Stracker, T. H. & Petrini, J. H. The MRE11 complex: starting from the ends. Nat. Rev. Mol. Cell Biol. 12, 90–103 (2011).
Kim, J. H., Penson, A., Taylor, B. S. & Petrini, J. H. J. Nbn-Mre11 interaction is required for tumor suppression and genomic integrity. Proc. Natl Acad. Sci. USA 116, 15178–15183 (2019).
Kim, J. H. et al. The Mre11-Nbs1 interface is essential for viability and tumor suppression. Cell Rep. 18, 496–507 (2017).
Oh, J., Al-Zain, A., Cannavo, E., Cejka, P. & Symington, L. S. Xrs2 dependent and independent functions of the Mre11-Rad50 complex. Mol. Cell 64, 405–415 (2016).
Nasmyth, K. & Haering, C. H. The structure and function of SMC and kleisin complexes. Annu Rev. Biochem 74, 595–648 (2005).
Syed, A. & Tainer, J. A. The MRE11-RAD50-NBS1 complex conducts the orchestration of damage signaling and outcomes to stress in DNA replication and repair. Annu Rev. Biochem 87, 263–294 (2018).
Deshpande, R. A. et al. ATP-driven Rad50 conformations regulate DNA tethering, end resection, and ATM checkpoint signaling. EMBO J. 33, 482–500 (2014).
Liu, Y. et al. ATP-dependent DNA binding, unwinding, and resection by the Mre11/Rad50 complex. EMBO J. 35, 743–758 (2016).
Lammens, K. et al. The Mre11:Rad50 structure shows an ATP-dependent molecular clamp in DNA double-strand break repair. Cell 145, 54–66 (2011).
Kashammer, L., et al. Mechanism of DNA end sensing and processing by the Mre11-Rad50 complex. Mol. Cell 76, 382–394 (2019).
Rotheneder, M. et al. Cryo-EM structure of the Mre11-Rad50-Nbs1 complex reveals the molecular mechanism of scaffolding functions. Mol. Cell 83, 167–185.e169 (2023).
Park, Y. B. et al. Eukaryotic Rad50 functions as a rod-shaped dimer. Nat. Struct. Mol. Biol. 24, 248–257 (2017).
Lim, H. S., Kim, J. S., Park, Y. B., Gwon, G. H. & Cho, Y. Crystal structure of the Mre11-Rad50-ATPgammaS complex: understanding the interplay between Mre11 and Rad50. Genes Dev. 25, 1091–1104 (2011).
Schiller, C. B. et al. Structure of Mre11-Nbs1 complex yields insights into ataxia-telangiectasia-like disease mutations and DNA damage signaling. Nat. Struct. Mol. Biol. 19, 693–700 (2012).
Rojowska, A. et al. Structure of the Rad50 DNA double-strand break repair protein in complex with DNA. EMBO J. 33, 2847–2859 (2014).
Hopfner, K. P. et al. Structural biochemistry and interaction architecture of the DNA double- strand break repair Mre11 nuclease and Rad50-ATPase. Cell 105, 473–485 (2001).
De la Rosa, M. B. & Nelson, S. W. An interaction between the Walker A and D-loop motifs is critical to ATP hydrolysis and cooperativity in bacteriophage T4 Rad50. J. Biol. Chem. 286, 26258–26266 (2011).
Hohl, M. et al. Modeling cancer genomic data in yeast reveals selection against ATM function during tumorigenesis. PLoS Genet. 16, e1008422 (2020).
Seifert, F. U., Lammens, K., Stoehr, G., Kessler, B. & Hopfner, K. P. Structural mechanism of ATP-dependent DNA binding and DNA end bridging by eukaryotic Rad50. EMBO J. 35, 759–772 (2016).
Hohl, M. et al. Interdependence of the rad50 hook and globular domain functions. Mol. Cell 57, 479–491 (2015).
Keeney, S., Giroux, C. N. & Kleckner, N. Meiosis-specific DNA double-strand breaks are catalyzed by Spo11, a member of a widely conserved protein family. Cell 88, 375–384 (1997).
Moreau, S., Ferguson, J. R. & Symington, L. S. The nuclease activity of Mre11 is required for meiosis but not for mating type switching, end joining, or telomere maintenance. Mol. Cell Biol. 19, 556–566 (1999).
Klapholz, S., Waddell, C. S. & Esposito, R. E. The role of the SPO11 gene in meiotic recombination in yeast. Genetics 110, 187–216 (1985).
Hunter, N. & Kleckner, N. The single-end invasion: an asymmetric intermediate at the double-strand break to double-holliday junction transition of meiotic recombination. Cell 106, 59–70 (2001).
Usui, T., Ogawa, H. & Petrini, J. H. A DNA damage response pathway controlled by Tel1 and the Mre11 complex. Mol. Cell 7, 1255–1266 (2001).
Zaitseva, J., Jenewein, S., Jumpertz, T., Holland, I. B. & Schmitt, L. H662 is the linchpin of ATP hydrolysis in the nucleotide-binding domain of the ABC transporter HlyB. EMBO J. 24, 1901–1910 (2005).
Kozlova, M. I., Shalaeva, D. N., Dibrova, D. V. & Mulkidjanian, A. Y. Common patterns of hydrolysis initiation in P-loop fold nucleoside triphosphatases. Biomolecules 12, 1345 (2022).
Hopfner, K. P. Mre11-Rad50: the DNA end game. Biochem Soc. Trans. 51, 527–538 (2023).
Fan, Y. et al. Structural basis for DNA double-strand break sensing by human MRE11-RAD50-NBS1 and its TRF2 complex. bioRxiv, 2025.2003.2014.643254.
Moreno-Herrero, F. et al. Mesoscale conformational changes in the DNA-repair complex Rad50/Mre11/Nbs1 upon binding DNA. Nature 437, 440–443 (2005).
Tatebe, H. et al. Rad50 zinc hook functions as a constitutive dimerization module interchangeable with SMC hinge. Nat. Commun. 11, 370 (2020).
Comstock, W., Sanford, E., Navarro, M. & Smolka, M. B. Profiling Tel1 signaling reveals a non-canonical motif targeting DNA repair and telomere control machineries. J. Biol. Chem. 301, 108194 (2025).
Cheng, D. T. et al. Memorial sloan kettering-integrated mutation profiling of actionable cancer targets (MSK-IMPACT): a hybridization capture-based next-generation sequencing clinical assay for solid tumor molecular oncology. J. Mol. Diagn. 17, 251–264 (2015).
Gut, F. et al. Structural mechanism of endonucleolytic processing of blocked DNA ends and hairpins by Mre11-Rad50. Mol. Cell 82, 3513-+ (2022).
Hohl, M. et al. The Rad50 coiled-coil domain is indispensable for Mre11 complex functions. Nat. Struct. Mol. Biol. 18, 1124–1131 (2011).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Varadi, M. et al. AlphaFold protein structure database in 2024: providing structure coverage for over 214 million protein sequences. Nucleic Acids Res. 52, D368–D375 (2024).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr D Biol. Crystallogr 66, 213–221 (2010).
Pettersen, E. F. et al. UCSF Chimera–a visualization system for exploratory research and analysis. J. Comput Chem. 25, 1605–1612 (2004).
Yan, Y., Tao, H., He, J. & Huang, S. Y. The HDOCK server for integrated protein-protein docking. Nat. Protoc. 15, 1829–1852 (2020).
Schwacha, A. & Kleckner, N. Identification of double holliday junctions as intermediates in meiotic recombination. Cell 83, 783–791 (1995).
Acknowledgements
We are grateful to Robert Reid and Rodney Rothstein (Columbia University), Patrick Sung (Yale University), Xiaolan Zhao, Scott Keeney and Dirk Remus (MSKCC) for yeast strains, plasmids, reagents, and technical support. We thank current and past members of the Petrini and Patel laboratories and Tom Kelly for helpful discussion throughout the course of this work. We thank Tom Kelly and Scott Keeney for critical reading of the manuscript. This work was supported by R35 GM136278-04 award (J.P.), Leukemia Lymphoma SCOR 7021020 and Maloris Foundation Awards (D.J.P.), MSKCC Basic Research Innovation Award (J.P. and D.J.P.) and Cycle for Survival (J.P.). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
M.H. expressed yeast proteins and M.H. and Y.Y. purified the proteins. Y.Y. made cryo-EM grids and collected and analyzed data. V.K. refined the cryo-EM structure, performed MD simulations and computational molecular modeling. M.H. did yeast genetic and biochemical experiments. D.J.P. and J.P. designed overall research. D.J.P. and J.P. wrote the manuscript. All authors were involved in the interpretation of data, final manuscript preparation, and approval.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Limitations of the study
In vivo, the S. cerevisiae Mre11 complex consists of Mre11, Rad50 and Xrs2. We were unable to obtain particles of MRX for our studies. This restricted our structure guided genetic analyses to Mre11 and Rad50. Also, we only got particles for MREQ and not for MRWT, thus preventing us from getting structural insights of the complex after ATP-hydrolysis. In addition, the flexibility of the extended Rad50 coiled coil domain precluded us from obtaining structural information regarding suppressor mutations that reside within the coils, thus limiting our ability to determine the mechanism(s) of suppression. The molecular defect of rad50-3KA was mostly informed from Molecular Dynamics simulations and future studies are required to test the suggested model.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hohl, M., Yu, Y., Kuryavyi, V. et al. Structure guided functional analysis of the S. cerevisiae Mre11 complex. Nat Commun 16, 7469 (2025). https://doi.org/10.1038/s41467-025-62583-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-62583-3
This article is cited by
-
Structural basis for DNA break sensing by human MRE11-RAD50-NBS1 and its regulation by telomeric factor TRF2
Nature Communications (2025)