Abstract
SLC15A4, an endolysosomal solute carrier family transporter, plays a critical role in TLR7/8/9-induced immune responses through assembling a complex with the downstream adaptor TASL in a conformation-dependent manner. Despite its close functional association and promising therapeutic potential in infections, tumors, and autoimmune diseases, the development of conformation-specific antibodies for human SLC15A4 (hSLC15A4) remains challenging. Here, using a systematic screening and validation approach, we identify a pair of conformation-selective antibodies, clones 107 and 235, targeting the endolysosomal lumen surface of hSLC15A4 with opposite conformation-regulatory activities. Specifically, clone 107 selectively binds to hSLC15A4 in a TASL binding-incompetent luminal-open state; whereas clone 235 stabilizes hSLC15A4 in a TASL binding-competent cytoplasmic-open state. Our research identifies antibodies that recognize distinct conformations of hSLC15A4, potentially enabling modulation of the TLR7/8/9 pathway and contributing to the development of targeted therapies and research tools selectively targeting hSLC15A4.
Similar content being viewed by others
Introduction
The innate immune system is the first line of defense against invading pathogens. Host cells produce a variety of pattern recognition receptors (PRRs) to recognize pathogen- (PAMPs) or damage- (DAMPs) associated molecular patterns and initiate the innate immune response, leading to the production of proinflammatory cytokines and the induction of cell death1,2. Toll-like receptors (TLRs), located on the plasma or endolysosomal membrane, are an evolutionarily conserved family of PRRs that are crucial for defense against various types of PAMPs and DAMPs3,4,5,6. Among the multiple TLRs, TLR7/8/9 are responsible for recognizing aberrant nucleic acids within the endolysosome, making them as the critical pathways in response to both pathogen infection and cellular stress3,7,8,9. Proper activation of these pathways helps fight pathogen infection and boosts the host’s anti-tumor immunity3,10,11,12; however, excessive activation can lead to severe autoimmune diseases, such as systemic lupus erythematosus (SLE)10,13,14,15,16.
SLE is a chronic autoimmune disease that can be life-threatening in severe cases. The disease is primarily caused by the hyperactivation of the type-I interferon (IFN-I) pathway, particularly in plasmacytoid dendritic cells (pDCs), and activation of autoreactive B cells17. Several studies have shown that SLC15A4 is highly expressed in pDCs and B cells and is closely associated with SLE disease18,19,20,21,22,23,24. SLC15A4 is a member of the SLC15 family of solute carriers, which is an endolysosome-resident amino acid transporter in immune cells that mediates the transport of L-histidine, small peptides, and NOD1/NOD2 ligands19,25,26,27,28,29,30. Systemic studies show that SLC15A4 recruits TASL, another SLE-associated protein, to the endolysosome through its conformational changes and then activates the transcription factor IRF5 via the C-terminal conserved pLxIS motif (where p represents a hydrophilic residue and x represents any residue) of TASL, which in turn regulates TLR7/8/9-dependent production of IFN-I and other proinflammatory cytokines20,22,31,32,33,34,35. In addition, the production of IFN-I, tumor necrosis factor α (TNFα), IL (interleukin)−6 and IL-12 upon TLR7/8/9-stimulation is largely abolished in immune cells from either SLC15A4 loss-of-function or knockout mice18,23,30. Accordingly, SLC15A4 loss-of-function or knockout mice exhibit significant reductions in SLE manifestations23,36.
Given its central role in both anti-pathogen/anti-tumor immune responses and SLE pathogenesis, SLC15A4 represents a promising therapeutic target for the treatment of infections, tumors, and autoimmune diseases. Based on its mechanism in inflammatory responses, targeting specific conformations of hSLC15A4 offers a strategy for disease treatment. Previous studies have explored a variety of small molecule inhibitors targeting SLC15A434,37. However, to date, no macromolecules, such as antibodies, that specifically target SLC15A4 with conformational specificity have been identified.
In this study, we used single-cell BCR sequencing technology to screen conformation-specific murine monoclonal antibodies against human SLC15A4 (hSLC15A4). Our biochemical and structural analyses show that clones 107 and 235 recognize distinct conformations of hSLC15A4. Specifically, clone 107 selectively binds to hSLC15A4 in a TASL binding-incompetent luminal-open state and potentially functions as a blocking antibody that negatively regulates the hSLC15A4-mediated TLR7/8 pathway. In contrast, clone 235 stabilizes hSLC15A4 in a TASL binding-competent cytoplasmic-open state and potentially acts as an activating antibody that positively influences this pathway. In addition, our study reveals the mechanism of TASL recruitment, which is consistent with previous studies32,33,34, and highlights the highly conserved mechanism of the SLC15A4-TASL adapter complex. These findings provide molecular insights into the mechanisms by which clones 107 and 235 target hSLC15A4, which are critical for the development of hSLC15A4 modulators, tracing tools for biotechnological applications, and SLC15A4-targeting macromolecular drugs for potential treatment of infections, tumors, and autoimmune diseases.
Results
Identification of conformation-selective antibodies for hSLC15A4
Considering the close connection of SLC15A4 with various pathological conditions and the pivotal role of conformational changes in modulating the TLR7/8/9 pathway, we opted to take a systematic screening and validation approach to identify conformation-selective monoclonal antibodies targeting hSLC15A4. To ensure a diverse repertoire of antibodies recognizing different conformations and epitopes of hSLC15A4, we prepared two types of samples for mouse immunization: hSLC15A4 alone and a complex of hSLC15A4 with its downstream adapter protein TASL. Given that antibodies targeting the endolysosomal lumen surface (topologically similar to the extracellular surface) rather than the cytoplasmic surface have broader application potential, the immunogenic N- and C-terminal flexible tails of hSLC15A4 located on the cytoplasmic side were removed. In addition, to minimize the generation of anti-TASL antibodies and to increase protein expression levels, we designed a truncated construct of mouse TASL (deletion of aa: 35-203) with the SLC15A4-interacting N-terminus (aa: 1–18) replaced by the sequence of human TASL (Supplementary Fig. 1a, b). Mice were then immunized with either the apo hSLC15A4 or the hSLC15A4-TASL complex carrying these designed modifications (Fig. 1a and Supplementary Fig. 1c, d).
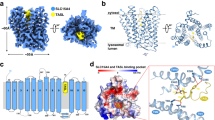
a Scheme of recombinant antibody screening. FACS, fluorescence-activated cell sorting. b Flow cytometry analyses of HEK293T cells surface-stained with Ab107 or Ab235. Cells transfected with indicated hSLC15A4 constructs (blue: Ab107; green: Ab235). EV empty vector, FL full-length, LL/AA hSLC15A4-L14A/L15A, 32-558 hSLC15A4-32-558, ISO isotype control, Ab antibody. c Schematic diagram of endolysosomal lumen loop substitutions of hSLC15A4 (left) and interactions between the substitutions and Ab107 or Fab235 (right). d Coomassie brilliant blue-stained SDS–PAGE gel of hSLC15A4 or hSLC15A4-TASL in complex with Fabs. Fractions marked by black star in Supplementary Fig. 2c–f were used for sample preparation. Data are representative of three independent experiments.
After three rounds of immunization, ~10,000 hSLC15A4-binding B cells were isolated from the spleen and lymph nodes of immunized mice via flow cytometric analysis. These cells were then subjected to single-cell B cell receptor sequencing (scBCR-seq) using the 10x Chromium Single Cell platform (10x Genomics) (Fig. 1a). Subsequent sequence analysis, antibody cloning and expression, hSLC15A4-binding characterization, and functional validation resulted in the identification of two monoclonal antibodies targeting hSLC15A4 with desired properties, namely clone 107 (from immunization with apo hSLC15A4) and clone 235 (from immunization with hSLC15A4-TASL complex). Cell surface staining and flow cytometric analysis using HEK293T cells expressing wild-type (WT) hSLC15A4 show that both clones can recognize the extracellular surface of hSLC15A4, indicating a dynamic distribution of hSLC15A4 in endolysosomal and plasma membranes and an extracellular (or endolysosomal luminal) targeting property of these antibodies (Fig. 1b). Given that mutation (L14A/L15A) or deletion (Δ1-31) in the N-terminus of hSLC15A4 is known to trigger its dominant plasma membrane localization20, we indeed observed a significant increase in cell surface antibody binding for HEK293T cells expressing these hSLC15A4 mutants (Fig. 1b). These data show that clones 107 and 235 are specific antibodies targeting the endolysosomal lumen surface (topologically similar to the extracellular surface) of hSLC15A4 and may have the potential to interfere with SLC15A4-mediated immune responses within the endolysosome.
Binding region mapping for clone 107 and clone 235
To elucidate the molecular basis of clone 107 and clone 235 binding to hSLC15A4, we performed systematic mutagenesis and flow cytometric assays to delineate the specific regions of hSLC15A4 targeted by clone 107 and by the Fab fragments of clone 235 (referred to as Fab235), respectively. Since both antibodies recognize the endolysosomal lumen surface of hSLC15A4 (Fig. 1b), we generated a series of glycine-serine (GS) substitutions covering each of the lumen-side loop (LL) connecting transmembrane helix (TM)1/TM2 (ΔLL1-2), TM3/TM4 (ΔLL3-4), TM5/TM6 (ΔLL5-6), TM7/TM8 (ΔLL7-8), TM9/TM10 (ΔLL9-10), and TM11/TM12 (ΔLL11-12) (Fig. 1c). Subsequent flow cytometric analyses show that both clone 107 and clone 235 can recognize the LL1-2, LL3-4, and LL5-6 mutants to a similar extent as wild-type hSLC15A4, whereas the LL7-8 and LL9-10 mutants are virtually unable to bind either clone 107 or clone 235 (Fig. 1c and Supplementary Fig. 2a, b). Furthermore, the interactions between the LL11-12 mutant and antibodies are also significantly affected, with ~85% reduction for clone 107 and ~50% reduction for clone 235 (Fig. 1c and Supplementary Fig. 2a, b). These results indicate that clone 107 and clone 235 primarily engage with the C-terminal portion of lumen-side regions of hSLC15A4, specifically the loops connecting TM7/TM8, TM9/TM10, and TM11/TM12.
Different conformational preferences of Fab107 and Fab235 for hSLC15A4
Although both clone 107 and clone 235 target a potentially overlapping surface on hSLC15A4 (Fig. 1c and Supplementary Fig. 2a, b), we evaluated the interactions of these two antibodies with either the apo-form hSLC15A4 or the TASL-bound hSLC15A4 to better understand the recognition mechanisms. Although hSLC15A4 itself can freely adopt different conformations, previous studies have shown that the formation of SLC15A4-TASL complex represents an immune active state and requires SLC15A4 to be specifically in its cytoplasmic-open conformation20,31,32,33,34. In line with flow cytometric results (Fig. 1c and Supplementary Fig. 2a, b), the size-exclusion chromatography (SEC) and SDS-PAGE experiments demonstrate that both antibodies can form stable complexes with the apo-form hSLC15A4 in vitro (Fig. 1d and Supplementary Fig. 2c, e, g, h). However, when incubated with the hSLC15A4-TASL complex, Fab107 and Fab235 exhibit disparate binding capabilities. Fab235 demonstrates a consistent and stable interaction with both the apo-form and TASL-bound hSLC15A4, whereas Fab107 does not exhibit a clear binding with the hSLC15A4-TASL complex (Fig. 1d and Supplementary Fig. 2d, f, g, h). Together, these assays suggest that Fab107 and Fab235 may prefer to recognize different conformational states of hSLC15A4. Fab107 seems to favor a TASL-binding-incompetent luminal-open conformation; while Fab235 tends to recognize a TASL-binding-competent cytoplasmic-open conformation.
Fab107 stabilizes hSLC15A4 in luminal-open state
To elucidate the structural details underlying Fab107’s preference for stabilizing a TASL-binding-incompetent luminal-open conformation of hSLC15A4, we purified the sample of hSLC15A4-Fab107 complex and successfully determined its cryo-EM structure to a resolution of 3.15 Å (Fig. 2a, b, Supplementary Fig. 3 and Supplementary Table 1). The cryo-EM map is of sufficient quality to build the atomic model for most of the transmembrane region of hSLC15A4 (residues 33–559), except for some potentially disordered sections in the loops connecting TM3/TM4 (residues 134-149) and TM6/TM7 (residues 264–303) (Fig. 2a, b and Supplementary Fig. 3). Notably, the cryo-EM density corresponding to Fab107 was also clearly resolved, allowing unambiguous model building for the variable domains of both the heavy (VH) and light (VL) chains (Fig. 2a, b and Supplementary Fig. 3). The transmembrane region of hSLC15A4 adopts a canonical Major Facilitator Superfamily (MFS) fold comprising 12 transmembrane helices (TMs) divided into N-terminal (NTD: TM1-TM6) and C-terminal (CTD: TM7-TM12) domain (Fig. 2b). A bundle bridge motif, inserted between the NTD and CTD, harbors a lateral helix connecting TM6 and TM7 (Fig. 2b), a feature also observed in other SLC15 family members38. Interestingly, although the apo-form hSLC15A4 has the ability to freely adopt different conformations, the presence of Fab107 induces hSLC15A4 to dominantly adopt the luminal-open conformation, with virtually no particles identified in the cytoplasmic-open conformation during the cryo-EM data process (Supplementary Fig. 3c).
a Cryo-EM density map of hSLC15A4-Fab107 complex. b Overall structure of Fab107 in complex with hSLC15A4 in luminal-open conformation. NTD and CTD are colored in palegreen and lightblue respectively, bundle bridge is colored in cyan, and light chain and heavy chain of Fab107 are colored in orange and violet respectively in (a, b). c Hydrogen bonding interactions between the endolysosomal lumen loops of hSLC15A4 and the light chain of Fab107. Key residues are shown as sticks and hydrogen bonds are represented as yellow dashed lines. d Hydrogen bonding interactions between the endolysosomal lumen loops of hSLC15A4 and the heavy chain of Fab107. Key residues are shown as sticks and hydrogen bonds are represented as yellow dashed lines. e Hydrophobic interactions between hSLC15A4 and the heavy chain of Fab107. Key residues are shown as sticks and dots. f Flow cytometric analyses showing the binding ability of wild type (WT) hSLC15A4 and its mutants to Ab107. Data are representative of two independent experiments. g CTD-centered structural superimposition of the cytoplasmic-open hSLC15A4 and the luminal-open hSLC15A4-Fab107. Sterically hindered residues are shown as sphere.
Consistent with the flow cytometry (Fig. 1b), the cryo-EM structure again confirms that Fab107 engages with the endolysosomal lumen surface and dominantly binds to the CTD of hSLC15A4 (Fig. 2a, b). The interface between Fab107 and hSLC15A4 covers a relatively large area (~1000 Å2) (Fig. 2b), which can be simply divided into the light chain- and heavy chain-involved interactions (Fig. 2c–e). The light chain-involved interactions are mainly mediated by the complementary determining region 1 (CDRL1) and CDRL3 of the light chain and the endolysosomal lumen loops LL7-8 and LL9-10 of hSLC15A4 (Fig. 2c). Specifically, multiple hydrogen bonds are formed through both main- and side-chain interactions, involving residues Q27L/N28L/N32L of CDRL1, N92L of CDRL3, T360 of LL7-8, and T438/N441 of LL9-10 (hereafter, light and heavy chain residues are designated by the superscript identifiers L and H, respectively) (Fig. 2c). Compared to the light chain, the heavy chain forms more extensive interactions with hSLC15A4, involving all three CDR regions and all endolysosomal lumen loops of the CTD (Fig. 2d, e). A hydrogen bond network is formed by T30H/R31H/W33H of CDRH1, E57H of CDRH2, D106H/D108H of CDRH3, as well as R350 of LL7-8, K431/E432/T434/N436/H445 of LL9-10, and N526 of LL11-12 (Fig. 2d). In addition, hydrophobic contacts further enhance the interaction, which are formed by Y102H/Y103H/Y105H/W107H of CDRH3 and F524 of LL11-12 (Fig. 2e). In line with the structural observation, alanine or serine substitutions of key interacting residues within the three endolysosomal lumen loops (R350A/P361A, N436A/T438A/N441A/H445A, F524S, and N526A) either abolish or reduce the binding ability with Fab107 (Fig. 2f and Supplementary Fig. 5a), highlighting the critical role of these interactions.
The presence of Fab107 robustly induces hSLC15A4 to adopt the luminal-open conformation, suggesting that the cytoplasmic-open conformation of hSLC15A4 is likely not suitable for Fab107 binding. Based on our Fab235-bound hSLC15A4-TASL structure (detailed in the following section; Fig. 3a, b, Supplementary Fig. 4 and Supplementary Table 1) and the previously reported cytoplasmic-open structure33, we were able to perform structural comparisons between the cytoplasmic-open state and the Fab107-bound luminal-open state. The CTD-centered structural superimposition shows that the bound of Fab107 will form steric hindrance with hSLC15A4 in the cytoplasmic-open conformation, specifically involving the clashes between LL7-8 and the light chain, and LL9-10 and TM9 with the heavy chain (Fig. 2g). This analysis provides a structural rationale for the reduced binding capabilities observed between Fab107 and the hSLC15A4-TASL complex (Fig. 1d and Supplementary Fig. 2e, f, h). Taken together, these data suggest that Fab107 binding may stabilize hSLC15A4 in its luminal-open conformation, thereby preventing TASL recruitment and potentially impeding downstream immune signaling.
a Cryo-EM density map of hSLC15A4-TASL-Fab235 complex. b Overall structure of Fab235 in complex with TASL-bound hSLC15A4 in cytoplasmic-open conformation. NTD, CTD, TASL are colored in palegreen, lightblue and yellow respectively, light chain and heavy chain of Fab235 are colored in orange and violet respectively in (a, b). c Hydrophobic interactions between the endolysosomal lumen loops of hSLC15A4 and Fab235. d Top view of the endolysosomal lumen loops (colored in cyan) of hSLC15A4 for hydrogen bonding interactions. e–g Hydrogen bonding interactions between the endolysosomal lumen loops (colored in cyan) of hSLC15A4 and Fab235. h Flow cytometric analyses showing the binding ability of wild type (WT) hSLC15A4 and its mutants to Fab235. Data are representative of two independent experiments. The interface between NTD (i) and CTD (j) of hSLC15A4 and Fab235 in CTD-centered structural superimposition of luminal-open hSLC15A4 and cytoplasmic-open hSLC15A4-Fab235. Wide variations of interface between two different conformations with Fab235 are circled by black dashed line. NTD of cytoplasmic-open hSLC15A4 is colored in palegreen, CTD of cytoplasmic-open hSLC15A4 is colored in lightblue and luminal-open hSLC15A4 is colored in yellow respectively.
Fab235 stabilizes hSLC15A4 in cytoplasmic-open state
The biochemical analyses show that Fab235 likely favoring a TASL-binding-competent cytoplasmic-open conformation of hSLC15A4 (Fig. 1d and Supplementary Fig. 2c, d, g). To better understand the molecular basis of Fab235-mediated conformational stabilization of hSLC15A4, we purified the sample of the hSLC15A4-TASL-Fab235 complex and successfully determined its cryo-EM structure to a resolution of 2.82 Å (Fig. 3a, b, Supplementary Fig. 4 and Supplementary Table 1). The high-resolution cryo-EM map is of sufficient quality to build the atomic model for the critical interaction regions of hSLC15A4, TASL and Fab235 (Fig. 3a, b and Supplementary Fig. 4). Similar to Fab107 and consistent with the flow cytometry (Fig. 1b), the cryo-EM structure confirms that Fab235 engages with the endolysosomal lumen surface of hSLC15A4 (Fig. 3a, b). As expected, hSLC15A4 adopts the cytoplasmic-open conformation, which is preferred for interactions of both TASL and Fab235 (Fig. 3a, b).
The interface between Fab235 and hSLC15A4 is mediated by both the heavy and light chains of Fab235 with four endolysosomal lumen loops of hSLC15A4 (LL1-2 of the NTD; LL7-8, LL9-10, and LL11-12 of the CTD) (Fig. 3c–g). The interactions involve hydrophobic contacts and hydrogen bonds, with the hydrophobic contacts occupying most of the interface and the hydrogen bonds surrounding the hydrophobic center further strengthening the interactions (Fig. 3c–g). Specifically, residues A65 of LL1-2, I435/V442/V443 of LL9-10, and F524 of LL11-12 form extensive hydrophobic contacts with Y33L of CDRL1, Y92L/H93L of CDRL3, F32H of CDRH1, and I102H/I103H/A105H of CDRH3 (Fig. 3c). In addition, Fab235-mediated hydrogen bonding interactions involve the LL7-8, LL9-10, and LL11-12 regions of hSLC15A4, details of which are as follows: 1) E353 and T359 of LL7-8 form hydrogen bonds with S30L of CDRL1 and R104H of CDRH3, respectively (Fig. 3d, e); 2) T434 and D448 of LL9-10 form hydrogen bonds with Y59H of the heavy chain framework region 3 (FWRH3) and R94L of CDRL3, respectively (Fig. 3d, f); 3) D523/N526/N528 of LL11-12 form a hydrogen bond network with S52H/S53H/S56H/N57H of CDRH2 (Fig. 3d, g).
To evaluate the importance of hSLC15A4-Fab235 interactions, we performed cellular experiments to assess the effect of hSLC15A4 mutants on Fab235 binding. Despite the relatively extensive interface, mutations at each of these lumen-side loops markedly reduce Fab235 binding, albeit to different extents (Fig. 3h and Supplementary Fig. 5b). Notably, alanine substitutions at LL9-10 (T434A/D448A) and LL11-12 (D523A/N526A/N528A) completely abolish Fab235 binding, whereas alanine substitutions at LL7-8 (E353A/T359A) reduce binding by approximately 50% (Fig. 3h and Supplementary Fig. 5b). Furthermore, among the hydrophobic interactions, serine substitutions of A65 on LL1-2 and F524 on LL11-12 cause a reduction of ~25–35%, while the multiple point mutation in the hydrophobic pocket (V442S/V443S/I435S on LL9-10) leads to a complete loss of Fab235 binding (Fig. 3h and Supplementary Fig. 5b). These combined structural and cellular findings highlight the critical role of hSLC15A4-Fab235 interactions.
The interface between Fab235 and hSLC15A4 buries ~1086 Å2 of surface area, distributed over both the NTD (~68 Å2) and CTD (~1018 Å2) domains of hSLC15A4 (Fig. 3b). Structural superimposition focusing on the CTD reveals that, when the conformation of hSLC15A4 changes from the cytoplasmic-open to luminal-open, the interface between Fab235 and the NTD is absent, and the interface between Fab235 and the CTD is reduced to almost half (~764 Å2) of the original (Fig. 3i, j). This observation suggests that Fab235 preferentially binds to hSLC15A4 in the cytoplasmic-open state, and the robust interactions between Fab235 and hSLC15A4 ensure the stabilization of this specific conformational state. Taken together, these results suggest that Fab235 binding may stabilize hSLC15A4 in the cytoplasmic-open conformation, thereby facilitating TASL recruitment and potentially promoting downstream immune signaling.
Conformational differences in Fab107- and Fab235-bound states
To gain insight into the conformational differences of hSLC15A4 upon binding to different antibodies, we compared the structures of hSLC15A4 in its Fab107- and Fab235-bound states. Structural superimposition of individual NTDs or CTDs reveals minimal changes in the transmembrane helices of each domain, except for slight shifts in TM7 and TM12 of the CTD (Fig. 4a, b). The endolysosomal lumen loops of the CTD, however, display dynamic alterations, possibly due to the binding to different antibodies (Fig. 4b). Notably, while both Fab107 and Fab235 primarily bind to the CTD of hSLC15A4, their binding modes and positions are significantly different (Figs. 2a, b, 3a, b and 4c), highlighted by an ~90-degree rotation of the overall structures between Fab107 and Fab235 (Fig. 4c).
Rocking motions of NTD (a) and CTD (b) of hSLC15A4 (luminal-open state: gray, cytoplasmic-open state: palegreen and lightblue) after structural alignment of both transporter units. Red lines show the shift of the TMs. c Structural superimposition of hSLC15A4-Fab107 complex (colored in lightpink) and hSLC15A4-TASL-Fab235 complex (colored in palecyan) in lateral view (left) and lysosomal lumen view (right). Lateral view (d), endolysosome lumen view (e), and cytoplasmic view (f) of hSLC15A4 in luminal-open state. g NTD-centered structural superimposition of cytoplasmic-open hSLC15A4 (colored in palegreen and lightblue) and luminal-open hSLC15A4 (colored in gray). Black arrows indicate the oscillation of the CTD. Lateral view (h), endolysosome lumen view (i), and cytoplasmic view (j) of hSLC15A4 in cytoplasmic-open state.
In the Fab107-bound luminal-open conformation, the luminal ends of TM1 and TM2 in the NTD are apart from TM7 and TM8 in the CTD, creating a cavity for substrate entrance and exposing a surface suitable for Fab107 binding (Fig. 4d, e). Furthermore, the cytoplasmic ends of TM4 and TM5 in the NTD pack tightly against TM10 and TM11 in the CTD, sealing hSLC15A4 from the cytoplasmic side and narrowing the TASL binding pocket (Fig. 4d, f). In contrast, in the Fab235-bound cytoplasmic-open conformation, the CTD undergoes a rigid rotation of approximately 30 degrees counterclockwise relative to the NTD (Fig. 4g), consistent with the ‘rocker switch’-like alternating-access mechanism of MFS transporters39,40. Moreover, the 90-degree rotation of Fab235 enables it to target both LL1-2 of the NTD and the three endolysosomal lumen loops of the CTD, thereby stabilizing hSLC15A4 in the cytoplasmic-open conformation (Figs. 3c–g and 4c). In this state, the NTD and CTD are in close proximity at their endolysosomal lumen ends (Fig. 4h, i). Specifically, TM1 and TM2 in the NTD are in close proximity to TM7 and TM8 in the CTD, effectively sealing the central cavity from the lumen side and creating a substantial surface comprising four endolysosomal lumen loops and facilitating the binding of Fab235 (Fig. 4h, i). On the cytoplasmic side, TM4 and TM5, along with TM10 and TM11, diverge to create the entrance of a polar cavity within the transporter, exposing the TASL-binding pocket and facilitating TASL recruitment (Fig. 4h, j). Together, the varied conformations of hSLC15A4 facilitate its accommodation of different antibodies, and the distinct binding modes of Fab107 and Fab235 enable them to effectively target and stabilize unique conformations of hSLC15A4.
Interactions between hSLC15A4 and TASL
The cryo-EM structure of the hSLC15A4-TASL-Fab235 complex not only reveals the molecular basis of Fab235-mediated conformational stabilization, but also provides an opportunity to analyze the interaction between SLC15A4 and TASL in the presence of a bound conformation-specific antibody. Residues 1–19 of TASL can be readily traced based on the well-defined density, whereas no density was observed for residues from 20 to the C-terminus, potentially due to the dynamic nature and lack of interaction with hSLC15A4 (Fig. 3a, b). Similar to the recently reported structure33, TASL binds within the cytoplasmic cavity of hSLC15A4, with its N-terminus inserting into the cavity and forming numerous contacts with hSLC15A4 (Fig. 5a, b). The intermolecular interactions between TASL and hSLC15A4 are mediated by both hydrogen bonds and hydrophobic interactions (Fig. 5b), including: 1) E4 of TASL interacts with R48 on TM1 and K172 on TM4 of hSLC15A4 (Fig. 5b); 2) S3, Y6, and E11 of TASL individually interact with E465 on TM10, D385 on TM8, and Q485 on TM11 of hSLC15A4, respectively (Fig. 5b); 3) L7 of TASL forms hydrophobic interactions with L473 on TM10 and F492 on TM11 of hSLC15A4, further strengthening the interaction (Fig. 5b). To validate the interface between hSLC15A4 and TASL, co-immunoprecipitation assays were conducted using HEK293T cells expressing different mutants of hSLC15A4 or TASL. Mutations of key residues of hSLC15A4 (R48A, K172A, D385A, E465A, Q485A, L473S, and F492S) significantly reduce the binding affinity to TASL (Supplementary Fig. 5c). In addition, mutations of TASL residues E4A, Y6A, E11A, and L7S, but not S3A, also affect binding to hSLC15A4 (Supplementary Fig. 5d). Although a longer model of TASL was traced and built in the current structure, the interaction interface between hSLC15A4 and TASL can be well aligned with the previously reported structure33, indicating a stable hSLC15A4-TASL interaction and confirming an active conformation of the Fab235-bound complex (Supplementary Fig. 5e). The main difference between the current and previously reported structures lies in the orientation of the C-terminus of the bound TASL fragment, which points in different directions (Supplementary Fig. 5e). Consequently, E11 of this region forms a hydrogen bond with hSLC15A4 (Fig. 5b), which was not reported in the previous study33.
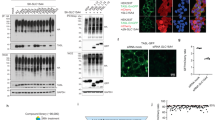
a Cut-open side view (left) and cytosolic bottom view (right) of electrostatic potential surface of hSLC15A4-TASL complex in cytoplasmic-open structure. TASL is shown as cartoon and the binding pocket is highlighted using dashed yellow oval. b Interactions between TASL (colored in yellow) and hSLC15A4 (colored in palegreen and lightblue) in cytoplasmic-open conformation. c Immunoblots showing whole-cell extracts (WCE) and immunoprecipitates (IP, Flag tag) from Flag-tagged hSLC15A4 transiently transfected HEK293T cells co-expressing Myc-tagged TASL from different species. hTASL Homo sapiens TASL, mTASL Mus musculus TASL, xTASL Xenopus tropicalis TASL, nmTASL Northern mallard TASL, gTASL Gallus TASL. Data are representative of three independent experiments. d TNFα and IL6 production of indicated THP1 cells stimulated with R848. gRNA1 and gRNA2, two different gRNAs targeting CXorf21. Mean ± s.d. (n = 4 biological replicates). e IL6 production of TASL-knockout THP1 cells stably reconstituted with TASL from various species unstimulated or R848 stimulated. Whole-cell extracts from indicated THP1 cells were analyzed by immunoblotting. Mean ± s.d. (n = 6 biological replicates). hTASL Homo sapiens TASL, mTASL Mus musculus TASL, xTASL Xenopus tropicalis TASL, nmTASL Northern mallard TASL, gTASL Gallus TASL.
hSLC15A4 interacts with diverse TASL homologs
Sequence alignment reveals a high degree of conservation for the TASL-binding surface of different SLC15A4 proteins (Supplementary Fig. 6). Most TASL homologs also preserve the essential SLC15A4-contacting N-terminal residues and the C-terminal pLxIS motif for downstream signaling (Supplementary Fig. 7). This suggests that hSLC15A4 may interact with various TASL homologs and subsequently trigger TLR7/8-induced inflammatory responses. To confirm this hypothesis, cellular assays were conducted to evaluate the interaction of hSLC15A4 with TASL proteins from different species. The results show that, in addition to human TASL (Homo sapiens), hSLC15A4 also interacts with the TASL proteins from Mus musculus, Xenopus tropicalis, Northern mallard, and Gallus (Fig. 5c). Furthermore, TLR7/8-induced inflammatory responses remain active (albeit to a different extent) in TASL knock-out (CXorf21−/−) THP-1 cells stably expressing various TASL homologs, suggesting that these TASL homologs could be recruited by hSLC15A4 to elicit TLR7/8-induced inflammatory responses (Fig. 5d, e). These findings highlight the highly conserved interaction between SLC15A4 and TASL, indicating the widespread conservation of TASL as an adapter in TLR-mediated inflammatory responses across species.
Discussion
SLC15A4 has emerged as a central player in endolysosomal TLR activation in pDCs and B cells18,19,20,21,22,23,24, suggesting its therapeutic potential in autoimmune diseases, antitumor responses, and antipathogen immunity. In this study, we identified two endolysosomal lumen surface antibodies, clones 107 and 235, which bind to a partially overlapped epitope on hSLC15A4, but target hSLC15A4 in its distinct conformational states. Our biochemical and structural analyses elucidate how these antibodies selectively target different conformations of hSLC15A4, indicating their potential to ether inhibit or activate the hSLC15A4-mediated endolysosomal TLR pathway by affecting TASL recruitment.
Fab107 forms robust and extensive interactions with the endolysosomal lumen surface of hSLC15A4, which stabilize hSLC15A4 in its luminal-open conformation and prevent it from transitioning to the cytoplasmic-open conformation due to potential steric hindrance. Importantly, the luminal-open conformation of hSLC15A4 is incompatible with TASL binding, indicating that Fab107 has the potential to block the hSLC15A4-mediated immune responses. In contrast, the binding of Fab235 helps to keep hSLC15A4 stable in its cytoplasmic-open conformation, as demonstrated by a larger buried interface (1086 Å2) with both the NTD and CTD of hSLC15A4. On the other hand, the luminal-open conformation of hSLC15A4 likely significantly weakens its interaction with Fab235 through a loss of NTD-mediated contacts and a reduced interaction area (764 Å2). Given that the cytoplasmic-open state is necessary for SLC15A4 to recruit the downstream adapter TASL, Fab235 may potentiate hSLC15A4-mediated immune signaling by stabilizing hSLC15A4 in the TASL-binding-competent conformation. This property positions Fab235 as a potential candidate for antitumor and antipathogen therapies that specifically target SLC15A4 by coupling it with an endolysosomal TLR agonist. In addition to being developed as potential therapeutics for diseases such as tumors, infections, and autoimmunity, Fab107 and Fab235 also function as conformation-specific antibodies that recognize hSLC15A4 either in the luminal- or cytoplasmic-open conformation, offering prospects for their utility as tracking tools to monitor the inhibitory or stimulatory state of hSLC15A4.
Notably, the two conformation-selective antibodies in this study were identified through direct screening without modification or optimization. While these initial candidates exhibit promising specificity, their therapeutic potential could be further enhanced through systematic optimization. Key areas for improvement include: 1) increasing binding affinity through structure-guided mutagenesis; 2) enhancing pH stability to ensure broader physiological compatibility; and 3) improving intracellular delivery efficiency—for example, by generating bispecific antibodies that fuse Fab107 or Fab235 with rapidly internalizing antibodies, or by conjugating them with cell-penetrating peptides (CPPs). The current proof-of-concept study provides valuable insights into the rational design of hSLC15A4-targeting modulators.
Furthermore, our research elucidates the molecular recruitment of TASL by hSLC15A4, aligning with previous studies32,33,34. Additionally, our findings highlight the broad-spectrum interaction between hSLC15A4 and TASL homologs. This is possible because of the relatively conserved N-terminus of TASL, which interacts with SLC15A4, and the cytoplasmic-open channel of SLC15A4, which accommodates TASL. Importantly, the complex formation between hSLC15A4 and various TASL homologs can successfully trigger downstream immune signaling, demonstrating the highly conserved nature of the SLC15A4-TASL adapter complex across species in terms of both interaction and function. This feature holds promise for the development of drugs based on the SLC15A4-TASL adapter complex for various diseases34.
In summary, our work identifies two conformation-specific antibodies targeting different conformations of hSLC15A4 and elucidates the detailed mechanisms of these two antibodies targeting hSLC15A4. This work may also serve as a valuable reference for the development of effective therapies and research tools that selectively target different functional states of hSLC15A4.
Methods
Ethical statement
Mouse experiments were approved by the Institutional Animal Care and Use Committees (SYXK2021070) at the Institute of Biophysics, Chinese Academy of Sciences.
Cloning, expression and purification of hSLC15A4 and TASL
The gene encoding human SLC15A4 (hSLC15A4) and TASL were synthesized and codon-optimized for expression in Spodoptera frugiperda (Sf9) insect cells. Full length hSLC15A4 or truncated hSLC15A4 (residues 32–558) was cloned into a pFastBac vector (Invitrogen) with a C-terminal 8x His-tag. The designed truncated construct of TASL was cloned into a pFastBac vector with a C-terminal 2x Strep-tag. The constructs were expressed individually in Sf9 insect cells following the instructions for the Bac-to-Bac baculovirus expression system (Invitrogen).
For apo hSLC15A4, Sf9 insect cells at a density of 2–2.5 ×106 cells/mL were infected with P4 baculovirus and grown at 27 °C. After 72 h, cells were harvested and resuspended in lysis buffer (50 mM Tris-HCl pH 7.5, 150 mM NaCl, 20 mM imidazole, and 10% glycerin) supplemented with 1 mM PMSF and homogenized by sonication on ice. hSLC15A4 was extracted with 1% (w/v) n-dodecyl-b-d-maltoside (DDM, Anatrace) and 0.1% (w/v) cholesteryl hemisuccinate (CHS, Anatrace) at 4 °C for 2 h. The insoluble fraction was removed by centrifugation at 30,000 × g at 4 °C for 30 min, and the supernatant was incubated with Ni-NTA resin with gentle agitation for 1 h. The resin was then washed using lysis buffer supplemented with 0.06% glyco-diosgenin (GDN, Anatrace). hSLC15A4 was eluted with buffer including 50 mM Tris-HCl pH 7.5, 500 mM NaCl, 500 mM imidazole, 10% glycerin, and 0.12% GDN. The protein was concentrated and further purified by size exclusion chromatography (SEC) on a Superdex 200 Increase10/300 GL column (GE Heathcare) in SEC buffer (20 mM HEPES pH 7.5, 150 mM NaCl, and 0.01% GDN). The peak fractions were pooled, concentrated to 5–10 mg/mL, and stored at −80 °C for later use.
For hSLC15A4-TASL, Sf9 insect cells at a density of 2–2.5 × 106 cells/mL were infected with P4 hSLC15A4 and TASL baculovirus simultaneously. After a two-step affinity chromatography (Ni-NTA resin and Strep-Tactin resin), the protein was further purified by SEC on a Superdex 200 Increase10/300 GL column in SEC buffer. The peak fractions were pooled, concentrated to 5–10 mg/mL, and stored at −80 °C for later use.
Antibody generation
All the mice used in this study are BALB/c background female mice and maintained in a specific-pathogen-free facility (temperature 20–22 °C, humidity 50–60%, 12 h light–dark cycle). All mice were used at age of 6–20 weeks.
BALB/c mice were immunized monthly with hSLC15A4 (residues 32-558) or the hSLC15A4-TASL complex plus Mn2+ adjuvant (MnStarter Biotechnology). Sera from immunized mice were tested by flow cytometry for reactivity towards hSLC15A4-expressing HEK293T cells. Approximately 10,000 antigen-specific B cells (hSLC15A4 or hSLC15A4-TASL complex-bound CD3−B220+CD19+CD38−GL7+ B cells) from the spleens and lymph nodes of immunized mice were isolated by the BD FACSAria III cell sorter and then barcoded through the 10x Chromium Single Cell platform (10x Genomics). The Chromium Single Cell V(D)J Reagent kit was used to prepare single-cell RNA libraries according to the manufacturer’s instructions. After quality control, the libraries were sequenced on Illumina Novaseq X Plus platform in 150 bp pair-ended manner (Berry Genomics Corporation, Beijing, China). Recombinant monoclonal antibodies against hSLC15A4 were synthesized using our published methods41. Briefly, BCR sequences were assembled and assessed using the V(D)J tool of Cell Ranger suit (v.6.0.1) against the GRCm38 reference genome (parameters: --denovo). Assembled contigs labeled as low-confidence, non-productive, or with UMIs <2 were discarded. Only cells with at least one light chain (IGL) and one heavy chain (IGH) were retained. In each filtered B cell, light and heavy chains were pairwise mated and ranked according to their mean clone levels. The selected VJ sequences of IGL were synthesized and cloned into vector with κ light chain, and the selected VDJ sequences of IGH were synthesized and cloned into vectors with constant regions of mouse IgG2a heavy chain or CH1 region of mouse IgG2a fused with a Myc-tag and 9x His-tag. The antibodies or Fab fragments were produced in HEK293F cells and purified with a protein A agarose or Ni-NTA resin prepacked column. The binding of purified antibodies to hSLC15A4 was confirmed by FACS. The sequences of Fab fragments are provided in Supplementary Table 2.
hSLC15A4-Fab and hSLC15A4-TASL-Fab complexes assembly
To prepare the hSLC15A4-Fab107 and hSLC15A4-Fab235 complexes, the purified hSLC15A4 (~6 mg/mL) was incubated with about 1.2-fold excess of Fab107 and Fab235, respectively, in SEC buffer (20 mM HEPES pH 7.5, 150 mM NaCl, and 0.01% GDN) on ice. After 30 min of incubation, the samples were loaded onto a Superdex G200 Increase10/300 GL column equilibrated with SEC buffer. Peak fractions were analyzed by SDS-PAGE.
To prepare the hSLC15A4-TASL-Fab107 and hSLC15A4-TASL-Fab235 complexes, the purified hSLC15A4-TASL (~6 mg/mL) was incubated with about 1.2-fold excess of Fab107 and Fab235, respectively, in SEC buffer (20 mM HEPES pH 7.5, 150 mM NaCl, and 0.01% GDN) on ice. After 30 min of incubation, the samples were loaded onto a Superdex G200 Increase10/300 GL column equilibrated with SEC buffer. Peak fractions were analyzed by SDS-PAGE.
Reconstitution of nanodiscs
MSP1D1 was expressed and purified according to a published protocol42. 1-Palmitoyl-2-oleoyl-sn-glycero-3-phospho-(1′-rac-glycerol) (POPG, Avanti) was solubilized using 2% DDM and 0.02% CHS in SEC buffer to a final concentration of 10 mM. hSLC15A4, Fab, MSP1D1, and POPG were mixed at a molar ratio of 1:1.1:2.2:120 and incubated at 4 °C for 1 h with gentle agitation. Detergents were removed by Bio-beads (Bio-Rad) at 4 °C overnight with constant rotation. After clarification by centrifugation, the mixture was loaded onto a Superdex 200 Increase 10/300 GL size-exclusion column equilibrated with 20 mM HEPES pH 7.5, 150 mM NaCl. The peak fractions were pooled and concentrated to about 15 mg/mL for cryo-EM sample preparation.
Cryo-EM sample preparation and data collection
3 μL aliquots of purified nanodisc-reconstituted hSLC15A4-Fab107 or hSLC15A4-TASL-Fab235 complexes were applied to glow-discharged holey carbon grids (Quantifoil Au R1.2/1.3, 300 mesh) for 3 s. Grids were blotted for 3 s at 4 °C and 100% humidity, and then plunge-frozen into liquid ethane using a Vitrobot Mark IV (Thermo Fisher Scientific). The grids were transferred to a Talos Arctica 200 kV FEG (Thermo Fisher Scientific) equipped with a Gatan K2 Summit detector and a GIF Quantum energy filter (slit width 20 eV). The movie datasets were collected in super-resolution mode, yielding a pixel size of 0.5 Å with a set defocus range of −0.8 μm to −1.2 μm. Each movie was dose-fractioned into 32 frames with a dose rate of 9.2 e−Å−2s−1, generating a total dose of 60 e−Å−2. Three datasets for hSLC15A4-Fab107 (7499 micrographs in total) and two datasets for hSLC15A4-TASL-Fab235 (3633 micrographs in total) were automatically collected using the SerialEM software package43.
Cryo-EM data processing
For the hSLC15A4-Fab107 datasets, all movie stacks were motion-corrected using MotionCor2 and binned twofold44. The dose-weighted micrographs unsuitable for further data processing were manually removed, and the remaining micrographs were processed in cryoSPARC345. For dataset 1, the parameters of the contrast transfer function (CTF) were estimated using Patch CTF. Thousands of particles were initially picked by blob picker from several micrographs, generating 2D averages for further template picking. A total of 1,202,084 particles were picked using template picker and extracted in a 256-pixel box. After two rounds of 2D classification, 407,127 particles with clear features were selected. Then, an Ab-Initio reconstruction of 50,000 particles in selected 2D classes was performed, producing 4 maps as 3D references. Two rounds of Hetero refinement generated 233,802 particles, which were further refined in Non-uniform (NU) refinement, yielding an overall resolution of 3.67 Å. To further improve the resolution, the best particles were transferred to Relion-3.1, where auto-refine and Bayesian polishing were performed46. The polished particles were imported into cryoSPARC3 and subjected to NU refinement. After CTF refinement, the dataset gave rise to a reconstruction at 3.25 Å. For dataset 2 and 3, a total of 173,320 and 146,232 particles were generated, respectively, using a similar procedure. All the particles were combined and applied for another rounds of refinement, including NU refinement, CTF refinement, and local refinement, resulting in a final reconstruction at 3.15 Å. The hSLC15A4-TASL-Fab235 datasets were processed by a similar procedure and yielded a reconstruction at 2.82 Å.
Model building and refinement
Predicted models of hSLC15A4 and Fab were generated by AlphaFold as initial models47. The models were then docked into the density maps and merged using Chimera48. The resulting model was then manually rebuilt in COOT49 and further refined by real-space refinement in PHENIX50. The model stereochemistry was evaluated using the comprehensive validation (cryo-EM) utility in PHENIX. Structural figures were generated using PyMOL51 and ChimeraX52.
Production of CXorf21-knockout cell lines
The lentiCRISPR v2 plasmid, a generous gift from F. Zhang (Addgene plasmid 52961), was used in this study. CXorf21 sgRNA no1, F: GTAGAAATGGAATCCTCCAT, R: ATGGAGGATTCCATTTCTAC; CXorf21 sgRNA no2, F: CTGAATTAATGGCCATCACC, R: GGTGATGGCCATTAATTCAG were inserted into the lentiviral backbone according to the Lentiviral CRISPR Toolbox protocol53,54. THP1 cells (ATCC) were transduced with the lentiviral vector containing the gRNA targeting CXorf21. Transduced cells underwent puromycin selection for 6 days and were then single-cell-sorted into 96-well plates with RPMI 1640 medium. After 8–10 days of culture, genomic DNA was extracted from the clones using a 50 mM NaOH solution, heated at 100 °C for 15 min, and neutralized with 1 M Tris-HCl (pH 8.0). The genomic region flanking the gRNA-target site was PCR-amplified from genomic DNA using the primers as follows (5′ to 3′ orientation): CXorf21, F: TCACAGTGCTGCAGACAAAC, R: GAATCTGGGTAGGAGAACTGTCATG. The PCR products were then sequenced after cloning into trans1-t1 phage-resistant chemically competent cells using the pEASY-T1 Cloning Kit (TransGen Biotech, CT101-01) according to the manufacturer’s instructions. Two THP1 independent clones with out-of-frame mutations at the CXorf21 gRNA target site were chosen for further experimentation.
Generation of cell lines overexpressing TASL
The DNA sequences encoding TASL from different species were inserted into a lentiviral plasmid containing a GFP reporter gene downstream of an IRES sequence. For lentiviral gene transduction, HEK293T cells were transfected with the lentiviral vectors along with packaging plasmids σNRF and vesicular stomatitis virus G (an envelope plasmid) using standard calcium phosphate techniques. After 48 h, culture supernatants were collected, filtered through 0.45-μm polyethersulfone filters, and supplemented with 8 μg/mL polybrene (Sigma-Aldrich). Cells were infected by spinfection (250 × g, 180 min, room temperature). Following 72 h of culture, lentiviral-infected cells with similar levels of GFP expression were isolated using a BD FACSAria III cell sorter (BD Biosciences).
Cell lysis and western blotting
Cells were lysed in CelLytic M buffer (Sigma) with the addition of 10% protease inhibitor cocktail (Roche). Following centrifugation at 13,300 × g for 20 min at 4 °C, the proteins were separated on 10% SDS‒PAGE gels and transferred onto polyvinylidene difluoride membranes. To minimize nonspecific binding, the membranes were blocked with 3% non-fat dry milk in TBS containing 0.1% Tween 20 before being probed with specific antibodies. Detection of binding was achieved using HRP-conjugated secondary antibodies and the MiniChemi® Mini Chemiluminescent Imaging and Analysis System.
Co-immunoprecipitation
Immunoprecipitation experiments were performed by co-transfecting HEK293T cells with expression vectors encoding SLC15A4-Flag and TASL-Myc, along with their respective mutants. Post-transfection, cells were lysed in 300 μL E1A buffer (50 mM HEPES pH 7.4, 250 mM NaCl, 5 mM EDTA, and 1% NP-40) supplemented with a protease inhibitor cocktail (Roche) for 30 min on ice. A portion of the whole-cell lysate (40 microliters) was collected as input, while the rest of the lysate was pre-cleared for 1 h at 4 °C using protein A-Sepharose (Smart-lifesciences) before proceeding to immunoprecipitation. Immunoprecipitation was carried out using either anti-Myc nanobody agarose beads (AlpaLifeBio) or anti-Flag agarose beads (Sigma) overnight at 4 °C. Subsequently, the beads were washed five times with E1A buffer and eluted with SDS sample buffer, followed by analysis via western blot.
Flow cytometry
HEK293T cells were transfected with expression plasmids encoding hSLC15A4 or its mutants. Following transfection, intracellular staining was performed by fixing and permeabilizing the cells with 2% paraformaldehyde (PFA) in PBS and a permeabilization buffer from Thermo Fisher Scientific. The cells were then stained with monoclonal antibodies, including anti-SLC15A4 clone 107 mIgG2a (2 μg/mL) and anti-SLC15A4 Fab235 Myc (2 μg/mL), followed by secondary staining with PE-conjugated anti-mouse IgG or anti-Myc antibodies. For surface staining, the cells were first directly stained with the same monoclonal antibodies (anti-SLC15A4 clone 107 mIgG2a and anti-SLC15A4 clone 235 mIgG2a) for primary staining, then stained with PE-conjugated anti-mouse IgG for the second staining. The flow cytometry gating strategy is shown in Supplementary Fig. 1e–g. The cells were then analyzed using an Attune NxT Flow cytometer (Thermo Fisher Scientific), and the data were processed with FlowJo software.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The cryo-EM maps have been deposited into the Electron Microscopy Data Bank under accession numbers EMD-60808 (hSLC15A4+Fab107) and EMD-60809 (hSLC15A4+TASL+Fab235). The coordinates have been deposited into the Protein Data Bank under accession numbers 9IRB (hSLC15A4+Fab107) and 9IRC (hSLC15A4+TASL+Fab235). Source data are provided with this paper.
References
Wu, J. & Chen, Z. J. Innate immune sensing and signaling of cytosolic nucleic acids. Annu. Rev. Immunol. 32, 461–488 (2014).
Iwasaki, A. & Medzhitov, R. Control of adaptive immunity by the innate immune system. Nat. Immunol. 16, 343–353 (2015).
Fitzgerald, K. A. & Kagan, J. C. Toll-like Receptors and the Control of Immunity. Cell 180, 1044–1066 (2020).
Takeuchi, O. & Akira, S. Pattern recognition receptors and inflammation. Cell 140, 805–820 (2010).
Lind, N. A., Rael, V. E., Pestal, K., Liu, B. & Barton, G. M. Regulation of the nucleic acid-sensing Toll-like receptors. Nat. Rev. Immunol. 22, 224–235 (2022).
Kawai, T. & Akira, S. The role of pattern-recognition receptors in innate immunity: update on Toll-like receptors. Nat. Immunol. 11, 373–384 (2010).
Ewald, S. E. et al. The ectodomain of Toll-like receptor 9 is cleaved to generate a functional receptor. Nature 456, 658–662 (2008).
Diebold, S. S., Kaisho, T., Hemmi, H., Akira, S. & Reis e Sousa, C. Innate antiviral responses by means of TLR7-mediated recognition of single-stranded RNA. Science 303, 1529–1531 (2004).
Heil, F. et al. Species-specific recognition of single-stranded RNA via toll-like receptor 7 and 8. Science 303, 1526–1529 (2004).
Krieg, A. M. & Vollmer, J. Toll-like receptors 7, 8, and 9: linking innate immunity to autoimmunity. Immunol. Rev. 220, 251–269 (2007).
Kawai, T. & Akira, S. Toll-like receptors and their crosstalk with other innate receptors in infection and immunity. Immunity 34, 637–650 (2011).
Rakoff-Nahoum, S. & Medzhitov, R. Toll-like receptors and cancer. Nat. Rev. Cancer 9, 57–63 (2009).
Marshak-Rothstein, A. & Rifkin, I. R. Immunologically active autoantigens: the role of toll-like receptors in the development of chronic inflammatory disease. Annu. Rev. Immunol. 25, 419–441 (2007).
Blasius, A. L. & Beutler, B. Intracellular toll-like receptors. Immunity 32, 305–315 (2010).
Brown, G. J. et al. TLR7 gain-of-function genetic variation causes human lupus. Nature 605, 349–356 (2022).
Fillatreau, S., Manfroi, B. & Dorner, T. Toll-like receptor signalling in B cells during systemic lupus erythematosus. Nat. Rev. Rheumatol. 17, 98–108 (2021).
Kirou, K. A. & Gkrouzman, E. Anti-interferon alpha treatment in SLE. Clin. Immunol. 148, 303–312 (2013).
Blasius, A. L. et al. Slc15a4, AP-3, and Hermansky-Pudlak syndrome proteins are required for Toll-like receptor signaling in plasmacytoid dendritic cells. Proc. Natl. Acad. Sci. USA 107, 19973–19978 (2010).
Kobayashi, T. et al. The histidine transporter SLC15A4 coordinates mTOR-dependent inflammatory responses and pathogenic antibody production. Immunity 41, 375–388 (2014).
Heinz, L. X. et al. TASL is the SLC15A4-associated adaptor for IRF5 activation by TLR7-9. Nature 581, 316–322 (2020).
Lopez-Haber, C. et al. The phagosomal solute transporter SLC15A4 promotes inflammasome activity via mTORC1 signaling and autophagy restraint in dendritic cells. EMBO J. 41, e111161 (2022).
Bentham, J. et al. Genetic association analyses implicate aberrant regulation of innate and adaptive immunity genes in the pathogenesis of systemic lupus erythematosus. Nat. Genet. 47, 1457–1464 (2015).
Baccala, R. et al. Essential requirement for IRF8 and SLC15A4 implicates plasmacytoid dendritic cells in the pathogenesis of lupus. Proc. Natl. Acad. Sci. USA 110, 2940–2945 (2013).
Kobayashi, T. et al. Human SLC15A4 is crucial for TLR-mediated type I interferon production and mitochondrial integrity. Int. Immunol. 33, 399–406 (2021).
Wang, Y. et al. Expression and regulation of the proton-coupled oligopeptide transporter PhT2 by LPS in macrophages and mouse spleen. Mol. Pharm. 11, 1880–1888 (2014).
Sakata, K. et al. Cloning of a lymphatic peptide/histidine transporter. Biochem. J. 356, 53–60 (2001).
Yamashita, T. et al. Cloning and functional expression of a brain peptide/histidine transporter. J. Biol. Chem. 272, 10205–10211 (1997).
Song, F., Hu, Y., Wang, Y., Smith, D. E. & Jiang, H. Functional characterization of human peptide/histidine transporter 1 in stably transfected MDCK cells. Mol. Pharm. 15, 385–393 (2018).
Nakamura, N. et al. Endosomes are specialized platforms for bacterial sensing and NOD2 signalling. Nature 509, 240–244 (2014).
Sasawatari, S. et al. The solute carrier family 15A4 regulates TLR9 and NOD1 functions in the innate immune system and promotes colitis in mice. Gastroenterology 140, 1513–1525 (2011).
Zhang, H. et al. SLC15A4 controls endolysosomal TLR7-9 responses by recruiting the innate immune adaptor TASL. Cell Rep. 42, 112916 (2023).
Custodio, T. F. et al. Molecular basis of TASL recruitment by the peptide/histidine transporter 1, PHT1. Nat. Commun. 14, 5696 (2023).
Chen, X. et al. Structural basis for recruitment of TASL by SLC15A4 in human endolysosomal TLR signaling. Nat. Commun. 14, 6627 (2023).
Boeszoermenyi, A. et al. A conformation-locking inhibitor of SLC15A4 with TASL proteostatic anti-inflammatory activity. Nat. Commun. 14, 6626 (2023).
Odhams, C. A. et al. Interferon inducible X-linked gene CXorf21 may contribute to sexual dimorphism in Systemic Lupus Erythematosus. Nat. Commun. 10, 2164 (2019).
Katewa, A. et al. The peptide symporter SLC15a4 is essential for the development of systemic lupus erythematosus in murine models. PLoS One 16, e0244439 (2021).
Chiu, T. Y. et al. Chemoproteomic development of SLC15A4 inhibitors with anti-inflammatory activity. Nat. Chem. Biol. 20, 1000–1011 (2024).
Killer, M., Wald, J., Pieprzyk, J., Marlovits, T. C. & Low, C. Structural snapshots of human PepT1 and PepT2 reveal mechanistic insights into substrate and drug transport across epithelial membranes. Sci. Adv. 7, eabk3259 (2021).
Drew, D. & Boudker, O. Shared molecular mechanisms of membrane transporters. Annu. Rev. Biochem. 85, 543–572 (2016).
Drew, D., North, R. A., Nagarathinam, K. & Tanabe, M. Structures and general transport mechanisms by the major facilitator superfamily (MFS). Chem. Rev. 121, 5289–5335 (2021).
Zhang, Q. et al. Recognition of cyclic dinucleotides and folates by human SLC19A1. Nature 612, 170–176 (2022).
Ritchie, T. et al. Reconstitution of membrane proteins in phospholipid bilayer nanodiscs. Methods Enzymol. 464, 211–231 (2009).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. elife 7, e42166 (2018).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. Sect. D Biol. Crystallogr. 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. Sect. D Biol. Crystallogr. 66, 213–221 (2010).
Schrödinger, L. The PyMOL Molecular Graphics System, Version 1.8 (Schrödinger, L, 2015).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
Acknowledgements
Cryo-EM data collection was carried out at the Center for Biological Imaging, Core Facilities for Protein Science at the Institute of Biophysics, Chinese Academy of Sciences. The computation work was performed using High-performance computing resources, the Center for Biological Imaging (CBI), Institute of Biophysics, Chinese Academy of Science. The plasmid expressing membrane scaffold protein 1D1 (MSP1D1) was a gift from Dr. Zhenfeng Liu’s group. We thank Junying Jia and Shu Meng (Institute of Biophysics, Chinese Academy of Sciences) for their technical assistance in flow cytometry analysis and sorting. This work was supported by grants from National Key R&D Program of China (2024YFA1307400 to P.G.), National Natural Science Foundation of China (32325028 & 32130057 to P.G., 32171219 to A.G., 32200989 to Y.Z., 82371853 to X.Z., 32401005 to Q.Z.), CAS Project for Young Scientists in Basic Research (YSBR-074 to P.G.), Basic Research Program Based on Major Scientific Infrastructures-CAS (JZHKYPT-2021-05 to P.G.), Beijing Natural Science Foundation (Z220018 to P.G.), and China Postdoctoral Science Foundation (BX20230468, 2023M740257 to Q.Z.).
Author information
Authors and Affiliations
Contributions
Y.Z., Q.Z., P.S. and K.L. purified proteins, prepared cryo-EM samples, collected and processed cryo-EM data, and reconstructed density maps. Y.Z., A.G. and P.G. built and refined models. X.Z. and Y.Z. performed antibody screening and validation. X.Z. performed cellular assays. X.N., J.M. and L.Z. assisted with antibody screening. Y.G. assisted with cryo-EM data process and reconstruction. Y.W. and S.L. assisted with cell culture and protein expression. P.G., L.Z. and A.G. initiated the project and directed the research. Y.Z., A.G. and P.G. wrote the manuscript with help of all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Renhong Yan and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhu, Y., Zhang, X., Zhang, Q. et al. Development of conformation-selective antibodies targeting human SLC15A4. Nat Commun 16, 7324 (2025). https://doi.org/10.1038/s41467-025-62759-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-62759-x