Abstract
Intratumoral low oxygen tension promotes cancer cell invasion and metastasis. Hypoxia-Inducible Factor 1-alpha (HIF1α) is the principal transcription factor orchestrating cellular responses to hypoxic stress, mediating the regulation of genes implicated in adapting to perturbations in oxygen homeostasis. Here, we describe our findings that functionally demonstrate a nucleolar localization domain in HIF1ɑ that enables HIF1ɑ to translocate to the nucleolus. Nucleolar HIF1ɑ binds the ribosomal DNA promoter and upregulates RNA Polymerase I activity leading to dysregulated ribosomal RNA transcription and consequently enhanced ribosome biogenesis. Ribosome biogenesis is important in supporting cellular metabolic processes and invasion and metastasis. Our findings are recapitulated in breast tumors wherein upregulated HIF1ɑ and rRNA biogenesis are associated with poor prognosis. Finally, our studies demonstrate that inhibition of RNA Polymerase I impedes aggressive traits of hypoxia-driven cancer progression, highlighting the potential of this approach as a therapeutic strategy in breast cancer. Cumulatively our work unravels an unprecedented role of HIF1ɑ in regulating rRNA biogenesis in breast cancer.
Similar content being viewed by others
Introduction
Metastasis represents a pivotal phase in cancer progression, where cancer cells disseminate from the primary site to establish secondary tumors in distant organs, posing a critical challenge in cancer treatment. Breast cancer, serving as a model system, epitomizes this challenge, being one of the primary contributors to mortality associated with cancer in women globally1,2. Annually, metastasis accounts for the majority of the over 40,000 fatalities attributed to breast cancer in the US alone3,4,5. This complex process entails cancer cells detaching from the primary tumor, migrating, invading through the extracellular matrix, and colonizing distant sites6, a progression intricately influenced by the tumor microenvironment7. The tumor milieu includes stressors such as acidic waste, immune infiltrates, nutrient deprivation, and hypoxia.
Intratumoral hypoxia is an important driver of invasion and metastasis in breast cancer. Hypoxia results from an insufficient supply of oxygen and low oxygen tension due to rapid tumor growth. Hypoxia, as a major cellular stressor, modulates and activates numerous signaling pathways and promotes the cellular metabolism shift from oxidative phosphorylation and tricarboxylic acid cycle to glycolysis by upregulating a plethora of glucose transporters and glycolytic enzymes8,9,10,11. This shift modulates tumor cells’ metabolic heterogeneity and facilitates tumor progression, invasion, recurrence, epithelial–to-mesenchymal transition (EMT), and therapy resistance12,13,14. The molecular program of EMT enables transition to invasive cancer and is typified by the loss of cell-cell adhesion and polarity and the development of a mesenchymal phenotype15.
The progression of cancer involves a cascade of molecular changes, driving the acquisition of new traits within tumor cells. Central to this process are extensive gene expression programs that facilitate the synthesis of proteins essential for the altered cellular state. This intricate biosynthetic process heavily relies on the molecular machinery of ribosomes. Consequently, the process of ribosome biogenesis presents an important event for facilitating EMT and promoting cancer cell proliferation and progression16,17,18.
Ribosome biogenesis constitutes a complex, multistage process initiated within the nucleolus, culminating in the assembly of functional ribosomes. Ribosomes are complex assemblies of ribosomal RNA (rRNA), ribosomal proteins, and ribosome-associated proteins, which function as translation machines. Ribosomal DNA (rDNA) is present as multiple repeats on five acrocentric chromosomes and is transcribed by RNA Polymerase I (RNA Pol I) to 47S pre-rRNA. The three rRNA species—28S, 5.8S, and 18S—are produced by splicing and processing of the 47S pre-rRNA19, which together with 5S rRNA, and several ribosomal proteins make up the functional ribosome. Several signaling pathways are implicated in regulating ribosome biogenesis20. Upregulated ribosome biogenesis fulfills the need for increased protein synthesis in cancer. This is associated with increased RNA Pol I activity and rDNA transcription and is evident as prominent nucleoli21,22,23. High nucleolar scoring in breast cancer, defined as increased nucleolar number and/or size, is associated with higher tumor grade and poor survival23. Hyperactive ribosome biogenesis has been reported in multiple cancer types, and as such, represents a promising target for novel anti-cancer drugs24,25. Given the essential role of ribosomes to enable cell survival and plasticity, it is important to note that the process of ribosome biogenesis is acutely sensitive to the tumor milieu. Hypoxia alters the processing of ribosomal proteins and their incorporation into the ribosome26 and prompts a subset of ribosomes with differentially methylated rRNA that facilitate the translation of specific transcripts27. However, the impact of hypoxia-driven signaling events on ribosome biosynthesis and ensuing mesenchymal plasticity and metastasis remain unclear.
Here, we report that breast cancer patients with an elevated hypoxia signature exhibit unfavorable progression-free survival. In these patients, there was an enrichment of ribosome and rRNA biogenesis-related signatures. As the principal mediator of hypoxia, upregulated expression of HIF1α positively correlated with ribosome biogenesis-related signatures. Pursuing these leads, we used multiple model systems to demonstrate that hypoxia upregulates RNA Pol I transcription and rRNA biogenesis. We report a previously unknown role for HIF1α in localizing to the nucleolus to directly regulate rRNA biogenesis and alter the translatome. Targeting ribosome biogenesis effectively suppresses tumor cell invasion and metastasis, offering a promising therapeutic strategy to limit the metastatic potential of hypoxic tumors, particularly in aggressive triple-negative breast cancer (TNBC) and basal-like subtypes.
Results
Hypoxia and ribosome biogenesis are upregulated and correlated in breast cancer
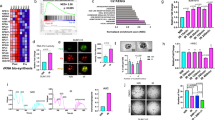
Hypoxia is a feature of most solid tumors and has a powerful impact on tumor progression. rRNA and ribosome biogenesis are necessary to support proliferation and tumor growth20. Using breast cancer as a model system, we tested the relationship between ribosome biogenesis and hypoxia. We applied Gene Set Enrichment Analysis (GSEA) on breast cancer RNA-seq data from The Cancer Genome Atlas (TCGA). Breast tumors demonstrated a significant enrichment of hypoxia and ribosome biogenesis-related signatures in comparison to normal tissues (Figs. 1A and S1A, B). Next, we stratified breast cancer samples from multiple databases (TCGA BRCA, METABRIC cohort, and GSE202203 cohort) into top and bottom hypoxia score quartiles using the Buffa mRNA-based hypoxia signature28. Consistent with previous studies10,28,29,30, patients with high Buffa hypoxia scores had worse progression-free and disease-free survival compared to patients with low Buffa hypoxia scores (Figs. 1B and S1C–E). GSEA revealed a significant enrichment of several ribosome and rRNA biogenesis-related signatures in tumors with the highest quartile of hypoxia scores compared to those in the lowest quartile across the TCGA BRCA, METABRIC, and GSE202203 cohorts (Figs. 1C, D and S1F–J). GSEA also highlighted key tumor promoting processes such as upregulation of cell cycle and checkpoints, MTORC1 signaling, DNA replication, and glycolysis signatures in tumors within the highest quartile of hypoxia scores compared to those in the lowest quartile across various databases (Fig. S2A–I). Leveraging the group-level insights from GSEA, we then applied single-sample Gene Set Enrichment Analysis (ssGSEA) to examine the variability of ribosome biogenesis pathways across individual samples, emphasizing intra-group heterogeneity and enabling a more personalized analysis of biological processes. ssGSEA revealed an upregulation of ribosome and rRNA biogenesis gene sets in tumor samples with the highest quartile Buffa hypoxia scores compared to those with the lowest quartile (Fig. S3A–C). Linear regression analysis demonstrated a strong correlation between Buffa hypoxia scores and ssGSEA scores of rRNA biogenesis signatures in TCGA BRCA, METABRIC, and GSE202203 cohorts (Figs. S4A, B and S5A).
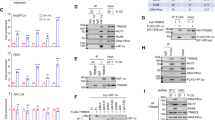
A Heatmaps and GSEA enrichment plots showing enrichment of Buffa hypoxia (normalized enrichment score (NES) = 2.071, P < 0.0001) and GOBP ribosome biogenesis gene sets (NES = 1.477, P < 0.01) in TCGA BRCA tumors (n = 1101) versus normal tissues (n = 139). NES and P values were calculated using GSEA (1000 permutations, two-sided). B Kaplan–Meier curves for disease-specific and progression-free survival in TCGA BRCA patients stratified by Buffa hypoxia score (top vs bottom quartile, n = 257 each). Hazard ratios (HR) with 95% confidence intervals and two-sided log-rank P values are shown. C Bubble plot of ribosome biogenesis-related gene sets enriched in TCGA BRCA high vs low hypoxia groups (n = 257/group). NES, gene set size, and GSEA nominal P values are indicated. D GSEA enrichment plots for GOBP Ribosome Biogenesis (NES = 1.854, P = 0.0253) and GOCC Preribosome (NES = 1.808, P = 0.0247) in TCGA BRCA stratified by Buffa hypoxia score. E Linear regression of RNA45S5 expression level versus Buffa hypoxia score in GSE28844 (n = 61), GSE42568 (n = 121), GSE5460 (n = 123), and GSE36771 (n = 107). Pearson’s r and two-sided P values are reported. F Correlation between RNA18SN5, RNA28SN5 expression and Buffa hypoxia scores in METABRIC (n = 1980). Pearson’s r and two-sided P values are shown. G Experimental design of SUM1315 cells exposed to hypoxia for 24 h prior to RNA-seq. H GSEA plots showing enrichment for Hallmark Hypoxia (NES = 2.896, FDR < 10−4) and KEGG Ribosome (NES = 2.013, FDR < 0.001) gene sets in RNA-seq data from SUM1315 cells exposed to hypoxia for 24 h, as outlined in (G).
In-depth analysis of multiple breast cancer microarray databases (GSE28844, GSE42568, GSE5460, GSE36771, GSE162228, GSE102484, and GSE21653) revealed that the levels of the RNA45S5 (225767_at) probe were moderately but significantly positively correlated with Buffa hypoxia scores (Figs. 1E and S5B). Furthermore, several hypoxia-upregulated genes also positively correlated with the expression of RNA45S5 probe in these three datasets (Fig. S5C–E), solidifying that hypoxia is associated with rRNA transcription and ribosome biogenesis. Additionally, the expression levels of RNA18SN5 and RNA28SN5 were positively correlated with Buffa hypoxia scores in METABRIC samples (Figs. 1F and S5F). Spatial gene expression mapping was then performed on human breast cancer tissue to investigate the spatial relationship between hypoxia and ribosome biogenesis signatures in tumors. Using the Visium HD Spatial Gene Expression platform, our analysis revealed a significant overlap between upregulated hypoxia-related genes and ribosome biogenesis signatures in both ductal carcinoma in situ (DCIS) and invasive ductal carcinoma (Fig. S6A–D). These findings highlight the relevance of ribosome biogenesis in hypoxic regions of breast cancer tissues.
Led by this collective evidence, we proceeded to validate the functional impact of hypoxia on ribosome and rRNA biogenesis. We undertook RNA-sequencing analysis of human TNBC SUM1315 cells cultured in hypoxic versus normoxic conditions (Fig. 1G) and identified significant enrichment of ribosome biogenesis signatures in hypoxia (Figs. 1H and S7A). We also performed ssGSEA analysis of publicly available data from cell lines representing breast cancer, hepatoma, glioma, and prostate cancer that were cultured in normoxic or hypoxic conditions. The data demonstrated an upregulation of multiple ribosome and rRNA biogenesis gene sets in hypoxia compared to normoxia (Fig. S7B–F). Our findings demonstrate that hypoxia and ribosome biogenesis are intimately linked and upregulated in breast cancer, underscoring a pronounced correlation that influences tumor progression and survival outcomes.
Hypoxia upregulates RNA Pol I activity
To expand and determine the mechanistic underpinnings of the interplay between hypoxia and rRNA biogenesis, we queried rRNA biosynthesis in multiple breast cancer cell lines representing several subtypes including SKBR3 (Luminal HER2+), T47D (Luminal A; ER+, PR+, and HER2−), SUM1315 (triple-negative basal-like), EMT6 (mouse; triple-negative basal-like) (Fig. 2A). We quantified RNA Pol I transcription using two independent assays: (i) the activity of RNA Pol I was quantified by the transient 5′ external transcribed spacer (5′ETS) and (ii) nascent RNA synthesis was assessed following a very short pulse with 5-ethynyl uridine (EU). Hypoxia clearly upregulated RNA Polymerase I (Pol I) activity as seen by the increase in the 5′ETS in hypoxia-treated cells when compared to cells in normoxia (Figs. 2B and S8A–C). Similar outcomes were noted in cells treated with CoCl2 or deferoxamine (DFO), which stabilize HIF1α and induce pseudohypoxia under normoxic conditions (Fig. S8D). To further examine how different oxygen levels affect Pol I activity, we exposed SKBR3, T47D, and SUM1315 cells to physioxia (for pre-conditioning the cells to the physiological O2 tension of breast tissue) or hypoxia. When cells were pre-incubated under physioxia and then exposed to hypoxia, there was a marked increase in RNA Pol I activity in T47D and SUM1315 cells (Fig. S8E), suggesting that hypoxia enhances RNA Pol I activity following physioxic preconditioning.
A Schematic of short-term hypoxia exposure and assays performed on breast cancer cell lines SKBR3, T47D, and SUM1315. Cells were incubated under hypoxia or normoxia for 8 h, followed by Pol I activity assay, EU incorporation, polysome profiling, and nucleolar staining. B Pol I activity was quantified by qRT-PCR targeting the 5′ ETS region following 8-h hypoxia exposure. β-Actin (ACTB) was used as the internal reference gene for relative normalization. Data are presented as mean ± SEM of three biological replicates. Statistical significance was assessed using a two-sided Student’s t-test. C–E EU incorporation assay showing increased nucleolar rRNA synthesis (EU, green; Fibrillarin, red; merged) after hypoxia in SKBR3 (C), T47D (D), and SUM1315 (E). For each condition, ≥ 8 random fields were analyzed in three independent biological experiments. Bar graphs show mean ± SD; each dot corresponds to the mean EU intensity from one experiment. Two-sided Student’s t-tests were used for statistical comparison. Scale bar = 10 µm. F–H Nucleolar counts per nucleus were quantified in SKBR3 (F), T47D (G), and SUM1315 (H) cells following 8-h exposure to normoxia or hypoxia using Nucleolar-ID staining. Bar plots display the mean ± SD across three independent experiments; each dot represents the mean nucleolar count per condition from one independent experiment. Statistical significance was assessed using a two-sided Student’s t-test. Scale bar = 10 μm. I Representative polysome profiles from sucrose gradients (10–50%) in SKBR3, T47D, and SUM1315 cells after 8-h normoxia or hypoxia. 40, 60, and 80S ribosomal subunits are indicated. Representative images from n = 2 independent experiments.
As a functional read-out of rRNA synthesis, we quantified nascent RNA synthesis following a short pulse with EU, followed by evaluating the intensity of EU incorporation. Most of the nascent RNA synthesis is observed in the nucleolus; there was a significant increase (52–90%) in EU incorporation in hypoxic cells compared to normoxic cells (Figs. 2C–E and S9A–C). Independent public prostate-cancer dataset (GSE196634) similarly showed elevated 18, 28, and 45S rRNA transcript levels in prostate epithelial cells (Fig. S9D), collectively supporting that hypoxia upregulates an rRNA biogenesis program in tumor cells. An increase in Pol I activity presents as transcriptionally active Nucleolar Organizer Regions that are evident as nucleoli31. Aligned with the functional readout, hypoxic cells showed a significantly greater number of nucleoli per nucleus compared to normoxic cells (Figs. 2F–H and S9E).
Next, we investigated if hypoxia impacts rRNA processing. For this, we employed qPCR analysis using primers that monitor the 45S rRNA precursor and key processing junctions of mature rRNAs32. Our results demonstrate that hypoxia notably upregulates rRNA transcription, as evidenced by the increased levels of the 45S rRNA precursor across all three breast cancer cell model systems (Fig. S9F–H). In addition to enhanced transcription, hypoxia stimulates rRNA processing, particularly at the early junctions of 18S and 28S rRNAs. While 5.8S rRNA processing remained steady in SUM1315 cells, it was notably upregulated in T47D and SKBR3 cells under hypoxic conditions, providing compelling evidence that hypoxia promotes rRNA biogenesis by increasing RNA Pol I activity and enhancing selective processing of rRNA.
We conjectured that the rRNA being synthesized would be incorporated into ribosomes. To address this, we enriched the ribosomal subunits from the cells exposed to hypoxia. As evidence of increased rRNA synthesis, we registered increased 40S and 60S ribosomal subunits from hypoxic cells as compared to normoxic cells, providing further evidence of elevated rRNA biogenesis in hypoxia (Figs. 2I and S10A–C). To assess the effect of hypoxia on protein synthesis, we conducted a puromycin incorporation assay, labeling cells after returning them to normoxia following 8 h of hypoxia (Fig. S10D). There was a modest reduction in protein synthesis in SKBR3 and T47D cells (Fig. S10E), suggesting that overall protein synthesis is marginally impacted post-hypoxia. Overall, these data converge on indicating that hypoxia upregulates rRNA biogenesis and cumulatively highlight the pivotal role of hypoxia in enhancing RNA Pol I transcription and rRNA biogenesis, emphasizing the intricate relationship between cellular oxygen levels and rRNA biogenesis.
HIF1α regulates RNA Pol I activity in hypoxia
Hypoxic conditions stabilize HIF1α protein; however, the stabilized steady state levels tend to vary in a time dependent manner33,34. Thus, to determine an optimal level of exposure we conducted a time course investigation of HIF1α levels in hypoxia. HIF1α levels gradually increase after exposure to hypoxia with peak levels at 4–8 h following induction of hypoxia (Fig. 3A–C). The mouse EMT6 cell line showed a similar pattern with peak HIF1α levels between 4 and 8 h of hypoxia (Fig. 3D). In agreement with this, SUM1315, T47D, and EMT6 cells exhibited HIF1α stabilization after 8 h of treatment with CoCl2 or DFO (Fig. S11A–C). Knockdown of HIF1α (Fig. 3E, F and H) resulted in a notable decrease in RNA Pol I activity in SUM1315 and SKBR3 under hypoxic conditions (Fig. 3G and I).
A–C Immunoblot analysis of HIF1α protein levels in SKBR3 (A), T47D (B), and SUM1315 (C) cells exposed to hypoxia (1% O2) for 0, 4, 8, and 24 h. D Immunoblot analysis of HIF1α stabilization in EMT6 cells exposed to hypoxia for 0, 4, 8, 16, and 24 h. E Schematic representation of HIF1α knockdown experimental design under short-term hypoxia. F, H Immunoblot analysis of HIF1α knockdown efficiency following siRNA transfection in SUM1315 (F) and SKBR3 (H) cells after 8-h hypoxia exposure. G, I Pol I activity and pre-rRNA synthesis measured by 5′ ETS qRT-PCR following HIF1α knockdown in SUM1315 (G) and SKBR3 (I) cells under hypoxia. Relative expression levels were normalized to β-Actin. Data are presented as mean ± SEM of three biological replicates. Statistical significance was assessed using a two-sided Student’s t-test. J Immunoblot analysis of HIF1α protein levels in SUM1315 and SKBR3 cells transfected with HIF1α overexpression (OE) or empty vector (CTRL EV). K Pol I activity quantified by 5′ ETS qRT-PCR in HIF1α OE and CTRL EV cells. Data are presented as mean ± SEM of three biological replicates. Two-sided Student’s t-test was used. For all immunoblots, densitometric quantifications were first normalized to β-Actin loading control and then further normalized to corresponding control conditions (either normoxia, empty vector, or non-target siRNA control), as appropriate. Immunoblotting was repeated in at least two independent biological experiments with comparable results; a representative blot is shown.
Next, we established stable transfectants engineered to constitutively express HIF1α that was resistant to degradation (Fig. 3J); These transfected cells demonstrated a marked elevation in RNA Pol I transcription compared to control transfections in normoxic conditions (Fig. 3K), thereby establishing a role for HIF1α in regulating RNA Pol I activity. To query the clinical relevance of HIF1α in exerting a regulatory role on RNA Pol I transcription, we queried TCGA breast cancer cohort using the ssGSEA_ROC tool. In this cohort, RNA Pol I associated gene signatures were found to be significantly enriched in patients exhibiting high HIF1α expression levels relative to those with low expression (Fig. S11D, E). We also applied correlation-based gene set enrichment analysis (corGSEA) on RNA-seq data from multiple mammary cell lines35 and registered a similar positive correlation between high levels of HIF1α and ribosome biogenesis-related signatures (Fig. S11F). These findings underscore the critical regulatory function of HIF1α in rRNA transcription and ribosome biogenesis.
HIF1α localizes to the nucleolus
After uncovering HIF1α’s role in regulating rRNA biogenesis, we investigated the sub-cellular localization of HIF1α in hypoxia using high-resolution confocal microscopy. Intriguingly, HIF1α demonstrated nucleolar localization. Figure 4A depicts the nucleolar localization of HIF1α concurrent with POLR1A (RPA194) and UBTF, which are bona fide nucleolar markers36. The fluorescence intensity signals of HIF1α, POLR1A, and UBTF exhibit an overlap in the nucleolar compartment, suggesting their co-existence in the nucleolus (Fig. 4B). The degree of co-localization between HIF1α and POLR1A, as well as HIF1α and UBTF, was quantitatively assessed using Mander’s overlap coefficient, revealing mean values of 0.67–0.78 for HIF1α-POLR1A and 0.78–0.89 for HIF1α-UBTF across SKBR3, T47D, and SUM1315 cell lines (Fig. 4C, D). This robust co-localization coefficient of HIF1α with POLR1A and UBTF implies that HIF1α may be directly involved in the rDNA transcription machinery. To ascertain the nucleolar localization of HIF1α, we analyzed nucleolar extracts from hypoxia-treated breast cancer cell lines (SKBR3, T47D, and SUM1315) for HIF1α. As seen in Figs. 4E, F and S12A, HIF1α was detected in the nucleolar extract of hypoxia-treated cells. We also registered a clear nucleolar presence of HIF1α when we immunoprecipitated HIF1α from the nucleolar fractions of hypoxic T47D cells compared to normoxic cells (Fig. S12B). These data collectively demonstrate that HIF1α localizes to the nucleolar compartment.
A Confocal images of SKBR3 (top), T47D (middle), and SUM1315 (bottom) cells exposed to 8-h hypoxia (1% O2), stained for HIF1α (green), POLR1A (RPA194; red), UBTF (blue), and DAPI (gray), showing nucleolar localization of HIF1α. Representative images from n = 2 independent experiments. Scale bar = 2.5 μm. B Fluorescence intensity line profiles illustrate the overlap between HIF1α (green), POLR1A (red), and UBTF (blue) within nucleoli. Representative images are shown in insets. Scale bar = 1 μm. C, D Quantification of HIF1α co-localization with POLR1A (C) and UBTF (D), assessed by Manders’ Overlap Coefficient (MOC) in SKBR3, T47D, and SUM1315 cells. Box-and-whisker plots display the minimum-to-maximum range with the median indicated by the center line. Data shown are from one representative of two independent biological experiments, with n = 9–10 nucleoli analyzed per group by confocal microscopy. Scale bar = 1 μm. E, F Western blot analysis of HIF1α distribution across whole-cell (WCF), cytoplasmic (CPF), nuclear (NUF), and nucleolar (NOF) fractions in T47D (E) and SUM1315 (F) cells following 8-h normoxia or hypoxia. Fibrillarin and α-Tubulin serve as nucleolar and cytoplasmic markers, respectively, confirming efficient subcellular fractionation. Immunoblotting was repeated in two independent biological experiments with comparable results; a representative blot is shown.
The HIF1α protein has a nucleolar localization domain and binds to the rDNA promoter and induces chromatin remodeling
We next investigated if nucleolar localization of stabilized HIF1α protein is due to its intrinsic characteristics. The Nucleolar Localization Sequence Detector (NoD)37 was employed to identify potential nucleolar localization sequences (NoLSs) in the HIF1α protein sequence. The NoD showed that HIF1α contains a putative NoLS in its N-domain (between residues 5 and 34). This NoLS is conserved across multiple species (Figs. 5A and S12C). To assess the functionality of this predicted HIF1α NoLS (Fig. S12D, E), we engineered two constructs that expressed the wild-type (WT) or mutant (Mut) form of HIF1α NoLS at the N-terminal of the EGFP protein (Fig. 5A). Live-cell imaging revealed that the WT HIF1α NoLS effectively mediated the accumulation of EGFP in the nucleolus in normoxic conditions in SUM1315 and T47D cells (Fig. 5B). On the contrary, the mutant NoLS led to the exclusion of EGFP from the nucleolus, indicating the governing role of this NoLS in nucleolar localization of HIF1α (Fig. 5C).
A Schematic representation of HIF1α protein domains. The predicted nucleolar localization signal (NoLS; residues 5–34) and the engineered mutant (Mut-NoLS) are indicated. Additional functional domains include MP-NLS (monopartite nuclear localization signal), BP-NLS (bipartite nuclear localization signal), bHLH (basic helix–loop–helix), PAS-A, PAS-B (Per/ARNT/Sim domains), ODD (oxygen-dependent degradation domain), and CAD (C-terminal activation domain). B, C The predicted NoLS directs nucleolar accumulation of EGFP. SUM1315 and T47D cells were transfected with wild-type or mutant HIF1α NoLS fused to pEGFP-N1. Representative live-cell images show EGFP (green) and DAPI (blue). Data shown are from one representative of two independent experiments. Scale bar = 5 μm. D Schematic diagram of the human 45S rDNA promoter region, indicating locations of predicted hypoxia response elements (HREs) and external transcribed spacer (ETS). E, F Chromatin immunoprecipitation (ChIP)-qPCR analysis showing HIF1α binding to two regions (site A and site B) of the 45S rDNA promoter in SUM1315 (E) and SKBR3 (F) cells exposed to 8-h hypoxia. Isotype IgG served as negative control. Data are presented as mean ± SEM from n = 3 technical replicates. The experiment was independently performed two times, each yielding consistent results; data from one representative independent experiment is shown. G Flowchart outlining proteomic screening strategy for HIF1α-associated nucleolar proteins interacting with RNA Pol I subunit RPA194, including nucleolar fractionation, immunoprecipitation, and LC-MS analysis in SUM1315 and T47D cells. H Pol I activity in SUM1315 cells after NPM1 knockdown under 8-h hypoxia. 5′ ETS qRT-PCR was performed; data are mean ± SEM from n = 3 technical replicates. The experiment was independently performed two times, each yielding similar results; data from one representative independent experiment is shown. I Proposed model illustrating NPM1 interaction with HIF1α and RPA194, promoting Pol I activity under hypoxic conditions.
To assess the functional relevance of the NoLS on the ability of HIF1α to regulate rRNA biogenesis, we transfected SUM1315 cells with wild-type (WT-NoLS) or mutant (Mut-NoLS) HIF1α constructs (Fig. S13A) and evaluated the colocalization of HIF1α with fibrillarin (FBL), a nucleolar marker. Immunofluorescence analysis revealed that WT-NoLS HIF1α strongly co-localized with FBL in the nucleoli, while Mut-NoLS HIF1α exhibited impaired co-localization (Fig. S13B, C). Next, we queried the ability of the WT-NoLS or Mut-NoLS HIF1α constructs to regulate RNA Pol I activity. qRT-PCR analysis demonstrated that WT-NoLS HIF1α robustly increased rRNA synthesis, whereas Mut-NoLS HIF1α only moderately upregulated RNA Pol I activity, confirming the importance of NoLS in regulating nucleolar function (Fig. S13D).
To further characterize the functional consequences of NoLS disruption, we examined the expression of canonical HIF1α target genes involved in glycolysis and metabolic adaptation (LDHA, CA9, SLC2A1, PDK1, and HK2). qRT-PCR analysis revealed no notable difference between WT-NoLS and Mut-NoLS HIF1α in the induction of these genes, indicating that disruption of the NoLS domain does not impact the nuclear transcriptional activator function of HIF1α (Fig. S13E). This suggests that while NoLS is essential for HIF1α nucleolar localization and RNA Pol I regulation, it is dispensable for its broader transcriptional activity within the nucleus.
We next examined the number of nucleoli per nucleus in WT-NoLS and Mut-NoLS HIF1α-expressing cells. WT-NoLS HIF1α expression led to a pronounced increase in nucleolar counts per nucleus (Fig. S13F–H). In contrast, Mut-NoLS HIF1α-expressing cells displayed a nucleolar count similar to non-transfected controls, indicating that NoLS is required for HIF1α-mediated regulation of nucleolar number. This underscores the critical role of NoLS in HIF1α nucleolar localization and its capacity to modulate rRNA biogenesis.
Given its function as a transcription factor, HIF1α interacts with the promoter elements of its target genes. Therefore, we explored the possibility of HIF1α binding to the rDNA promoter. Interestingly, there are two predicted hypoxia response element (HRE) sequences in the promoter region of human 45S rDNA at positions −231 and −350 relative to the transcription start site upstream of the 5′ ETS. We found that HIF1α binds to the proximal promoter region of rDNA, at positions −350 (Site A) and −231 (Site B) (Fig. 5D). Using chromatin immunoprecipitation (ChIP), we validated that HIF1α occupies both sites in hypoxia-treated breast cancer cells (Fig. 5E, F). To confirm that the HIF1α-bound rDNA promoter regions supported transcription, we evaluated the promoter region for evidence of active/inactive chromatin marks. ChIP-qPCR analyses revealed an enrichment of distinct active chromatin marks H3K4me3 and H3K27ac at the rRNA locus, specifically within the HIF1α-bound proximal promoter regions of rDNA, highlighting an open chromatin state conducive to transcription (Fig. S14A–C). In contrast, enrichment of the heterochromatin-associated markers H3K36me3 and H3K9me remained unchanged under hypoxic conditions at the rDNA promotor (Fig. S14D, E), suggesting a selective epigenetic response to hypoxic conditions.
We next sought to identify the molecular partners of HIF1α within the nucleolar compartment. For this, we used a multipronged approach—we performed ChIP for RNA Pol I catalytic subunit RPA194 coupled with mass spectrometry (ChIP-MS) and independently, immunoprecipitated (IP) HIF1α from the nucleolar fraction (IP-MS). As depicted in Fig. 5G, ChIP-MS identified proteins that were enriched by more than twofold in hypoxic conditions; these were sorted based on the magnitude of change in hypoxia, and then curated to focus on confirmed nucleolar proteins as reported in the Human Protein Atlas (HPA) database38. A cohort of seven proteins was identified by (nucleolar) HIF1α IP-MS, including Nucleophosmin 1 (NPM1), Non-POU domain-containing octamer-binding protein (NONO), and Thymopoietin (TMPO) as prime candidates for HIF1α interaction (Fig. 5G). Subsequent gene knockdown approaches (Fig. S15A–C) revealed that NPM1 knockdown effectively mitigated hypoxia-driven upregulation of RNA Pol I transcription in SUM1315 cells, suggesting a unique role for NPM1 in mediating HIF1α regulation of RNA Pol I transcription (Figs. 5H and S15D–F).
We further investigated the interaction between HIF1α and NPM1 and its impact on rDNA transcription under hypoxic conditions. In SUM1315 cells engineered to express HIF1α, co-immunoprecipitation provided confirmation of the association between HIF1α and NPM1 (Fig. S15G, H). Further, ChIP followed by quantitative PCR (ChIP-qPCR) demonstrated that knockdown of NPM1 markedly reduced the binding of HIF1α to HREs at the rDNA promoter (Fig. S15I), underscoring an important role for NPM1 in facilitating HIF1α-mediated transcriptional regulation of rDNA.
Overall, our findings delineate HIF1α nucleolar localization as an intrinsic event facilitated by specific localization sequences, enabling direct rDNA promoter binding and transcriptional activation. This mechanism, coupled with the identification of the NPM1 nucleolar protein as a key interactor (Fig. 5I), underscores a sophisticated molecular network through which HIF1α orchestrates the activation of rRNA biogenesis, presenting a nuanced understanding of cellular adaptations to hypoxia.
Hypoxia and rRNA biogenesis drive cancer progression and metastasis
To elucidate the intricate dynamics of cancer progression and invasion, we assessed the role of rRNA biogenesis in modulating these processes under hypoxic conditions. We analyzed data from the METABRIC cohort and found that expression levels of HIF1α, RNA28SN5, and RNA18SN5 were significantly greater in grade 3 neoplasms, as well as in ER-negative and HER2-negative subtypes, with the most pronounced upregulation observed in HER2-enriched and basal subtypes (Fig. S16A–C). We next sought to determine the prognostic relevance of the correlation between HIF1α and rRNA levels. Application of Kaplan–Meier plotter (KMplot) revealed that high expression of HIF1α and rRNA (RNA45S5 and RNA5-8SN1) was associated with worse overall survival and relapse-free survival in breast cancer patients (Fig. S17A, B)39. In addition, Kaplan–Meier estimator analysis (GSE42568, GSE162228, GSE29271, and GSE21653) revealed similar prognostic outcomes of HIF1α and rRNA levels in breast cancer patients using the R2 genomics analysis platform (Fig. S17C, D). TNMplot database analysis also revealed that HIF1α transcript levels were significantly higher in metastatic samples compared to primary tumors (Fig. S18A)40. Complementing this, ssGSEA analysis of RNA-seq data from the AURORA US Metastasis Project (GSE209998), which includes 51 primary tumors and 102 metastases from 55 metastatic breast cancer patients, revealed significant positive correlations between the Buffa hypoxia signature and ribosome biogenesis-related signatures (Fig. S18B–E). This trend of increased hypoxia and ribosome biogenesis in metastatic tumors suggests that enhanced ribosome biogenesis under hypoxic conditions may contribute to metastasis.
Further stratification of breast cancer subtypes in TCGA BRCA, METABRIC, and GSE202203 cohorts revealed that the basal subtype exhibits the highest hypoxia and ribosome biogenesis signatures. Analysis of Buffa, Hu, and West hypoxia scores demonstrated that basal tumors consistently have the highest levels of hypoxia-associated signatures compared to luminal A, luminal B, and HER2+ subtypes (Fig. S19A–C). In parallel, gene signatures related to ribosome biogenesis and rRNA processing, including GOBP ribosome biogenesis, Bryant rRNA biogenesis, and Tafforeau rRNA processing, were significantly enriched in basal breast cancer across all three cohorts (Fig. S19D–F). Collectively, these findings establish that basal/TNBC subtypes of breast cancer are characterized by the highest levels of hypoxia and ribosome biogenesis, reinforcing the hypothesis that hypoxia-driven ribosome biogenesis plays a crucial role in aggressive breast cancer progression and metastasis.
To elucidate the influence of hypoxia on the characteristics of actively translated transcripts and subsequent protein networks in breast cancer, we performed polysome profiling analysis in T47D, SUM1315, and SKBR3 cell lines. Using a sucrose gradient fractionation system, we enriched polysomes from these cells and carried out RNA-seq analysis of the polysome fractions to identify the transcripts being translated by the ribosomes. We identified 41 actively translated transcripts that were consistently enriched under hypoxic conditions compared to normoxic conditions (Fig. S20A). Functional enrichment analysis of these hypoxia-associated transcripts using the STRING database revealed a significant enrichment in biological processes such as cellular response to hypoxia, mitochondrial outer membrane permeabilization, and monosaccharide metabolic processes (Fig. S20B, C), highlighting potential targets for therapeutic intervention in hypoxia-related events in breast cancer. Furthermore, BRCA Clinical Proteomic Tumor Analysis Consortium (CPTAC) samples showed a significant positive correlation between hypoxia scores, HIF1α expression, and protein levels of polysome-enriched transcripts in T47D, SUM1315, and SKBR3 cells, particularly for proteins PGK1, P4HA1, KDM3A, PDK1, GYS1, and SLC2A1, which are key metabolic regulators that promote tumor progression (Fig. S21A–F). As such these studies highlight the translational impacts of hypoxia-driven enhanced ribosome biogenesis on proteins that influence cellular metabolic processes and chromatin status.
BMH-21 inhibits RNA Pol I activity and reduces cancer invasiveness under hypoxia
Our data indicate that in hypoxic conditions, stabilized HIF1α displays its presence in the nucleolus, where it interacts with NPM1, binds to the rDNA promoter, and activates RNA Pol I. Given the lack of direct pharmacological interventions to alleviate or target tumor hypoxia, we evaluated BMH-21, a compound that selectively inhibits rRNA synthesis by intercalating into rDNA and blocking RNA Pol I transcription41,42,43,44. To determine the effect of a low dose of BMH-21 (50 nM; much below the reported dose for 50% growth inhibition for various cell lines) on hypoxic cells, we exposed SUM1315 cells to 8 h of either hypoxia or normoxia and treated them with BMH-21, DMSO (control), or Actinomycin D (positive control). BMH-21 did not inflict nucleolar stress in either condition, as evidenced by sustained colocalization of HIF1α and NPM1 (Fig. S22A, B). Similarly, cells engineered to overexpress stable HIF1α (HIF1α OE) did not exhibit nucleolar stress when treated with BMH-21 (Fig. S22C, D). In contrast and expectedly, Actinomycin D noticeably disrupted nucleolar structure under both normoxic and hypoxic conditions.
Independently, we queried the effect of BMH-21 on cells engineered to constitutively express HIF1α (OE). BMH-21 consistently inhibited cell viability and RNA Pol I activity in HIF1α-OE cells compared to controls harboring an empty vector (EV) (Fig. S22E, F). Notably, in the EV group, RNA Pol I activity showed recovery after 12 h, emphasizing this differential effect without compromising nucleolar integrity.
We then assessed the impact of BMH-21 on hypoxia-driven tumor cell migration and invasion. BMH-21 treatment significantly decreased transmembrane migration of SUM1315, EMT6, and T47D cells in hypoxic conditions (Figs. 6A, B and S23A). Similarly, the invasive potential of these cells through a Matrigel invasion chamber was markedly reduced in the presence of BMH-21 (Figs. 6C, D and S23B). These findings suggest that BMH-21 effectively impedes the hypoxia-triggered invasive behavior commonly associated with aggressive cancer progression.
A, B Representative images and quantification of transwell migration (4 h) in SUM1315 and EMT6 cells after 8-h hypoxia with or without BMH-21. Bar plots display mean ± SD of three biological replicates. Scale bar = 100 μm. C, D Representative images and quantification of Matrigel invasion (16 h) in SUM1315 and EMT6 cells under the same conditions. Bar plots display mean ± SD of three biological replicates. Scale bar = 100 μm. E, F Transwell migration (8 h) and Matrigel invasion (16 h) in SUM1315 HIF1α-overexpressing (OE) or empty vector (EV) cells treated with BMH-21 or vehicle. Bar plots display mean ± SD of three biological replicates. Scale bar = 100 μm. G, H Pulmonary metastasis assay (PuMA) of SUM1315 HIF1α OE and EV cells following BMH-21 or vehicle. Corrected total cell fluorescence (CTCF) quantified using ImageJ. Box-and-whisker plots show minimum-to-maximum range, median (center); each dot represents one lung section (n = 7 per group). The assay was repeated twice with consistent results; data from one representative experiment are shown. Scale bar = 50 px. I, J 3D spheroid assays for SUM1315 HIF1α OE and EV cells ± BMH-21. Circularity quantified by ImageJ. Bar plots display mean ± SD of three biological replicates. Scale bar = 50 μm. K Schematic of experimental design and treatment timeline. L, M Tumor growth (volume and diameter) in mice injected with SUM1315 HIF1α OE cells treated with BMH-21 or vehicle. Data are presented as mean ± SEM; n = 15 mice per group. N Tumor weight at endpoint. Box-and-whisker plots: min-to-max range, median (center); n = 15 mice per group. O Lung metastasis counts at endpoint. Box-and-whisker plots: min-to-max range, median (center); n = 15 mice per group. P Proposed model illustrating RNA Pol I inhibition counteracts HIF1α-driven invasion. Statistical significance was assessed using one-way ANOVA followed by Tukey’s multiple comparisons test. For (L, M), comparisons were restricted to HIF1α OE groups. Significance thresholds: P < 0.05 (*), < 0.005 (**), < 0.0005 (***); ns not significant.
To explore the role of nucleolar localization in HIF1α-mediated invasion, we assessed the invasive behavior of SUM1315 cells expressing either WT-NoLS or Mut-NoLS HIF1α, with corresponding non-transfected controls (Fig. S23C). Both WT-NoLS and Mut-NoLS enhanced invasion relative to controls, but WT-NoLS showed a notably stronger effect, underscoring the importance of HIF1α’ nucleolar localization in potentiating HIF1α-driven invasive behavior. Furthermore, BMH-21 markedly curtailed migration and invasion of SUM1315 cells engineered for constitutive expression of WT-NoLS HIF1α (Fig. 6E, F). Aligned with these findings, the pulmonary metastasis assay (PuMA) revealed a marked reduction in metastatic outgrowth in HIF1α-OE cells treated with BMH-21 compared to EV controls (Figs. 6G, H and S23D). Next, we employed a 3D culture system to assess the suppressive impact of BMH-21 on the invasive behavior of HIF1α-driven cancer cell spheroids. Compared to vehicle control-treated clusters, the cell clusters in BMH-21-treated HIF1α OE cells demonstrated significantly greater intact circularity, suggestive of reduced invasive potential (Figs. 6I, J and S23E, F).
To solidify these observations, we established stable transfectants of EMT6 cells that constitutively expressed (OE) HIF1α and confirmed that HIF1α-OE resulted in upregulated RNA Pol I activity (Fig. S24A, B). Using these cells, we performed an ex vivo PuMA assay, where lung sections from mice injected with HIF1α-OE or control cells were cultured with or without BMH-21 (Fig. S24C). Bioluminescence imaging (BLI) at days 6 and 8 showed a pronounced increase in metastatic outgrowth of HIF1α-OE cells, while BMH-21 treatment effectively reduced metastatic burden, particularly in the HIF1Α-OE group, consistent with our findings in the SUM1315 PuMA assay (Fig. S24D, E). These results further support the role of HIF1α in promoting metastatic potential and the efficacy of RNA Pol I inhibition in suppressing this process.
BMH-21 suppresses tumor growth and metastasis in HIF1α triple-negative breast cancer models
Basal-like/TNBC breast cancer exhibits the highest levels of hypoxia and ribosome biogenesis, making these subtypes highly dependent on rRNA synthesis for tumor progression. These findings provide a strong rationale for targeting RNA Pol I as a therapeutic strategy in TNBC, where tumor growth and metastatic potential may be particularly reliant on enhanced ribosomal activity. Therefore, the studies were advanced to pre-clinical assessment, where we orthotopically injected SUM1315 HIF1α-OE cells into the mammary fat pad of NSG mice. The animals were administered BMH-21 at a low dose of 10 mg/kg, which is substantially lower than previously reported dosages in murine models (Fig. 6K)41,45. Remarkably, BMH-21 treatment resulted in a significant suppression of tumor growth and a reduced incidence of tumor recurrence post-surgical resection in the BMH-21-treated cohort (Figs. 6L–N and S25A–E). We also assessed lung metastases in these mice (Figs. 6O and S26). BMH-21 treatment notably decreased the number of metastatic nodules, particularly in the context of HIF1α overexpression, demonstrating its efficacy in curtailing metastatic progression through Pol I inhibition.
Our comprehensive analysis, integrating linear regression and Pearson correlation analysis, uncovered intricate relationships between tumor progression, ribosomal RNA expression (quantified using RNAscope), hypoxia (measured via CA9 staining), and cell proliferation (assessed via PCNA immunoreactivity) within primary tumors. A strong positive correlation was identified between the abundance of lung metastases and RNAscope signal intensity, demonstrating a clear link between elevated RNA Pol I activity and increased metastatic potential, particularly in HIF1α-OE tumors (Fig. S27A–C and Supplementary Data 1). The impact of BMH-21 treatment was evident, as it significantly reduced both RNAscope signal and metastasis in these models, supporting the hypothesis that Pol I inhibition can mitigate tumor aggressiveness. Reflective of cellular proliferation, PCNA immunoreactivity showed a strong positive correlation with tumor size and lung metastasis (Fig. S27D–F). BMH-21 treatment reduced PCNA levels and metastatic burden, particularly in HIF1α-OE cells. CA9 staining, a marker of hypoxia, demonstrated a positive association with lung metastasis in HIF1α-OE tumors (Fig. S27G–I). Additionally, aligned with decreased metastasis, we see that BMH-21-treated HIF1α-OE tumors exhibited prominently networked collagen fibers compared to controls (Fig. S27J), suggesting that BMH-21 likely influences the architecture of the extracellular matrix to impede metastasis.
To further evaluate the therapeutic potential of BMH-21 in TNBC and to complement the findings with SUM1315-xenografts, we employed an orthotopic EMT6 murine model. BALB/c mice were injected with EMT6 cells expressing either HIF1α or EV into the mammary fat pad. BMH-21 (10 mg/kg) was administered intraperitoneally five times per week for five weeks and tumor growth was monitored using caliper measurements (Fig. S28A, B). While BMH-21 treatment did not noticeably alter tumor growth (Fig. S28B), upregulated HIF1α expression led to a significant increase in lung metastases, as indicated by elevated bioluminescent flux (Fig. S28C–F). These findings were corroborated by post-mortem inspection of lungs, which showed a marked increase in visible metastatic nodules in the HIF1α-expressing group compared to controls. Importantly, BMH-21 treatment significantly reduced metastatic burden, particularly in HIF1α-OE tumors, demonstrating its efficacy in suppressing metastasis of TNBC (Fig. S29A, B). In agreement with the SUM1315 tumors, the EMT6 HIF1α-OE tumors demonstrated notably greater PCNA staining that correlated positively with the abundance of lung metastases, consistent with an aggressive proliferative phenotype that was attenuated by BMH-21 (Fig. S29C–F). Collectively, these results highlight BMH-21’s efficacy in reducing RNA Pol I-driven transcription, proliferation, and metastasis, particularly in aggressive hypoxia-associated TNBC tumors (Fig. 6P).
Discussion
Metastasis and hypoxia are intrinsically linked phenomena in the progression of solid tumors, driving the spread of cancer cells from the primary site to distant organs46. Hypoxia, characterized by reduced oxygen availability, plays a pivotal role in tumor biology, influencing processes such as EMT, invasion, stemness, resistance to radiation therapy, and ultimately, metastasis10,47. In the context of breast cancer, which serves as an illustrative model, healthy breast tissue is characterized by a relatively high oxygen concentration of approximately 8.5%. However, this level significantly decreases within breast tumors, often dropping to 1.5% or lower48,49. This stark contrast highlights the adaptability of cancer cells to hypoxic conditions, underscoring the importance of understanding the mechanisms by which hypoxia facilitates metastasis50,51. Hypoxia was previously reported to impact RNA Pol I activity and alter the epitranscriptomic modifications of rRNA compared to normoxia, underscoring the complexity of the cellular response to low oxygen levels27. Our study highlights the dynamic interplay between hypoxia and ribosome biogenesis, revealing how hypoxic environments in breast cancer can significantly influence cellular processes12,29. Our findings demonstrate that hypoxia-induced changes in ribosome biogenesis are a crucial aspect of tumor biology. This extends our understanding of hypoxia beyond its established roles in metabolic reprogramming52,53 and underscores the significance of modifying protein synthesis machinery in response to hypoxic stress in breast cancer cells.
We demonstrated the integral association between hypoxia and ribosome biogenesis in breast cancer. Analysis of TCGA RNA-seq data revealed a significant enrichment of hypoxia and ribosome biogenesis signatures in tumors, as evidenced by poor survival outcomes in patients with elevated Buffa hypoxia scores28. This correlation underscores the critical role of hypoxia in modulating rDNA transcription, further supported by our analyses across multiple datasets and ssGSEA, which highlighted the upregulation of ribosome and rRNA biogenesis in tumors with high hypoxia scores. Our in vitro experimental models confirmed hypoxia’s capacity to upregulate RNA Pol I activity, emphasizing HIF1α regulatory role in this process, where its stabilization under hypoxic conditions directly augments RNA Pol I activity. This complements our findings that suggest potential therapeutic targets in the nexus of hypoxia, HIF1α, and ribosome biogenesis to combat tumor growth and progression.
Perhaps the most striking finding from our study is the nucleolar localization of HIF1α during hypoxia. Traditionally recognized for its role in the cytoplasm and nucleus, where it regulates transcription of genes involved in survival under low oxygen, HIF1α presence in the nucleolus suggests a direct involvement in the control of rRNA synthesis. This finding opens unexplored avenues for research into how HIF1α might influence other nucleolar processes, such as ribosomal assembly and the cellular stress response, potentially reshaping our understanding of how cancer cells adapt to hypoxic environments. Our approach of leveraging ChIP-MS and IP-MS has unearthed a network of protein interactors associated with HIF1α in the nucleolus. Among these, the interaction with NPM1 is particularly noteworthy, given NPM1's known roles in ribosome maturation and cellular stress responses54,55. These interactions hint at a broader regulatory landscape where HIF1α might influence gene expression and cellular adaptation strategies through direct engagement with key protein players.
Our study also investigated the nuanced mechanisms through which hypoxia modulates ribosome biogenesis in breast cancer, highlighting a pivotal role for HIF1α in orchestrating ribosome biogenesis. Through comprehensive analyses, including sucrose gradient fractionation and RNA sequencing of polysome-enriched fractions from T47D, SUM1315, and SKBR3 breast cancer cells, we identified 41 actively translated transcripts consistently enriched under hypoxic conditions. Functional enrichment analysis underscored significant associations with cellular responses to hypoxia and metabolic alterations, suggesting these pathways as potential therapeutic targets.
Investigating the therapeutic potential of RNA Pol I inhibition using BMH-21, a selective RNA Pol I inhibitor, yielded a notable reduction in cell viability and RNA Pol I activity in tumor cells engineered to overexpress HIF1α. Building on the evidence from 3D cultures and ex vivo PuMA models the orthotopic mouse model further substantiated the efficacy of BMH-21 at a low dose of 10 mg/kg. Even at this reduced dosage, BMH-21 effectively attenuated HIF1α-dependent cancer progression, resulting in significant reductions in tumor volume and metastatic spread. Notably, the data affirmed that BMH-21’s selective targeting capability is not due to the induction of nucleolar stress in hypoxia, further distinguishing it as a precise inhibitor of RNA Pol I activity, also evidenced in decreased RNAscope signal intensity in HIF1α-OE tumors. These results collectively position RNA Pol I inhibition as a promising therapeutic approach in targeting aggressive, hypoxia-driven cancer evolution and progression.
Hypoxia upregulates rRNA biogenesis and RNA Pol I activity, suggestively enhancing cellular adaptation to hypoxic stress by modulating the ribosome biogenesis27. Inhibiting RNA Pol I activity disrupts this adaptive response, impairing cellular survival and aggressiveness under hypoxic conditions. BMH-21 selectively inhibits RNA Pol I transcription by targeting the RPA194 subunit, without significantly affecting RNA Pol II or III activity, distinguishing it from other compounds and making it a highly specific tool for studying rRNA biogenesis41,42,43,44. Our study provides insights into HIF1α’s role in localizing to the nucleolus and directly upregulating RNA Pol I transcription during hypoxia. This opens unexplored therapeutic avenues, suggesting that targeting RNA Pol I may counteract the hypoxia-driven progression of aggressive tumors, particularly TNBC. Further research is needed to investigate the precise dynamics of ribosome biogenesis under hypoxic conditions and its implications for cancer progression and resistance to therapy. In summary, our work establishes a direct link between hypoxia, HIF1α, and ribosome biogenesis in breast cancer, offering insights into the cellular adaptations that support tumor survival and growth in hypoxic environments, and identifying potential targets for therapeutic intervention in hypoxia-driven aggressive cancer phenotypes.
Methods
These studies comply with the relevant ethical regulations. All animal experiments were performed in accordance with protocols approved by the Institutional Animal Care and Use Committee (IACUC) at the University of Alabama at Birmingham. All experimental materials, including detailed information on cell lines, antibodies, reagents, animal strains, and software tools, are comprehensively reported in the “Methods” section and summarized in Supplementary Data 5 to ensure transparency and reproducibility.
Patient data analysis
TCGA Breast Cancer processed RNA-seq data, comprising 1247 normal tissues and primary breast cancer tumors, was downloaded from Xena browser in September 202356. This data, derived using the Illumina HiSeq 2000 RNA Sequencing platform, represents transcription levels presented as log2(x + 1) transformed RSEM normalized count. Additionally, we accessed data from the METABRIC study, which included mRNA expression profiles using the Illumina HT-12 v3 microarray and clinical phenotype data for 1980 patients, sourced from the cBioPortal website in October 202357. RNA sequencing data for 3207 breast cancer patients were obtained from the NCBI Gene Expression Omnibus (GEO) database, identified by accession number GSE202203. Expression microarray datasets encompassing multiple breast cancer cohorts were retrieved using the R2: Genomics Analysis and Visualization Platform (available at http://r2.amc.nl). The datasets include GSE28844, which features 61 breast cancer patient samples; GSE42568, which contains 104 breast cancer samples and 17 normal mammary tissue samples; GSE5460, which comprises 123 breast cancer samples; GSE36771, which includes 107 breast cancer samples; and GSE209998, which includes specimens from 55 metastatic breast cancer (MBC) patients, representing a total of 153 unique tumors, consisting of 51 primary tumors and 102 matching metastases. cBioPortal for Cancer Genomics was used to retrieve the CPTAC of TCGA breast cancer patients57. Spatial gene expression data were explored using Loupe Browser v7.0, with log2 as the scale value, and feature averaging used for combining data. The dataset files were downloaded from 10× Genomics Spatial Data Sets (www.10xgenomics.com/datasets). These analyses enabled the identification of differentially expressed genes and provided detailed insights into the molecular dynamics within breast cancer tissues (gene sets used are listed in Supplementary Data 2). Processed RNA sequencing and microarray expression data for various cell lines under hypoxic and normoxic conditions were obtained from NCBI GEO. This included T47D breast cancer cells (GSE149132) cultured under 0.5% O2 (hypoxic) and 20% O2 (normoxic) for 48 h, HepG2 hepatoma cells (GSE18494) maintained at 19% O2 with treatments of 0.5% O2 for 4 and 8 h, MCF-7 breast cancer cells (GSE47533) subjected to 1% O2 for 16, 32, and 48 h versus a 21% O2 control, IDH1 wild-type glioma stem cells (GSE118683) under matched conditions of 20% O2 and 1.5% O2 for 72 h, and RWPE-1 prostate epithelial cell (GSE196634) exposed to 1% O2 for 6, and 12 h versus normoxic conditions.
Hypoxia scores calculation
To assess hypoxia in breast cancer patients, we employed Buffa and Winter mRNA signatures that were previously established for hypoxia (Supplementary Data 3)28,30. For the computation of hypoxia score, the expression level of each gene in a patient’s profile was compared to the median expression of that gene across the cohort. A gene score of +1 was assigned if a patient’s gene expression exceeded the median, while a score of −1 was given if it fell below. The overall RNA Hypoxia Score for a patient was then calculated as the sum of scores across all genes within a signature58. Patients were stratified into low or high hypoxia groups based on quartiles derived from their RNA Hypoxia Scores.
Bioinformatic tools and analysis
Gene sets related to ribosome biogenesis and RNA Polymerase I (Pol I) activity were sourced from the Molecular Signatures Database, complemented with published gene sets recognized for their relevance in various stages of ribosome and rRNA biogenesis, namely Metge rRNA Biosynthesis59, Tafforeau rRNA Processing Factors60, and Bryant rRNA Biogenesis61 gene sets. Moreover, we manually curated genes associated with Pol I core subunits, its regulation, and ribosome biogenesis, designating these as Pol I Core Machinery, Positive Pol I Regulation, and Samant Ribosome Biogenesis Signature (all gene sets listed in Supplementary Data 4). Using the stratification based on the hypoxia scores, we conducted Kaplan–Meier survival analyses and GSEA to study the impact of hypoxia on patient survival outcomes and the differential pathway enrichment between patients with high and low hypoxia scores, respectively. To visualize the differential enrichment of gene sets and signatures between tumors with the highest and lowest quartile hypoxia scores, bubble plots were created using the ggplot2 package in R Studio. We also carried out ssGSEA on RNA sequencing data from these patients, focusing on gene sets related to ribosome and rRNA biogenesis (reported in Supplementary Data 4). Utilizing the GenePattern platform62, we calculated ssGSEA scores, further applying Z-score normalization to guarantee the data’s consistency and comparability. We then correlated these hypoxia scores with ssGSEA scores for ribosome and rRNA biogenesis signatures through linear regression analysis to explore their association. For visual representation, heatmaps were generated by Morpheus software (https://software.broadinstitute.org/morpheus).
We also employed a multifaceted analytical approach to explore the relationship between HIF1α transcript levels and gene signature enrichment across various mammary cell lines and breast cancer patient data. Utilizing RNA-seq data from the ARCHS4 database35, we conducted correlation analyses using the Correlation AnalyzeR package63. This analysis aimed to investigate how HIF1α transcript levels correlate with the enrichment of diverse gene signatures, particularly those related to ribosome biogenesis. To further delve into the association between HIF1α expression and ribosome biogenesis in breast cancer, we turned to the TCGA BRCA patient dataset. Here, we employed the ssGSEA_ROC tool on the GenePattern platform62, categorizing patients based on high and low HIF1α expression levels. This tool facilitated the generation of receiver operating characteristic (ROC) curves and the computation of the area under the curve from ssGSEA results, focusing on a binary phenotype classification of ribosome biogenesis-related signatures. This robust analysis provided not only statistical assessments but also visual representations of the top-scoring gene sets for each phenotype, enhancing our understanding of the link between HIF1α expression and ribosomal gene enrichment.
Cell lines
SUM1315 cells (obtained from Asterand Bioscience) were cultured in DMEM-F12 medium (Thermo Fisher, Cat # 11330032) supplemented with 5% heat-inactivated fetal bovine serum (FBS) (Thermo Fisher, Cat # A5256701), 10 μg/mL insulin (Millipore Sigma, Cat # I5500-500MG), and 20 ng/mL epidermal growth factor (EGF) (Millipore Sigma, Cat # E9644-.2MG). T47D cells (ATCC # HTB-133) were cultured in RPMI medium (Thermo Fisher, Cat # 22400105) supplemented with 10% FBS and 10 μg/mL insulin. SkBr3 cells (ATCC # HTB-30) were cultured in McCoy’s 5 A medium (Thermo Fisher, Cat # 16600082) supplemented with 10% FBS. The EMT6 murine mammary carcinoma cell line (ATCC # CRL-2755) was cultured in DMEM (Thermo Fisher, Cat # 11995065) supplemented with 5% FBS, 1% Glutamax (Thermo Fisher, Cat # 35050061), and 1% non-essential amino acids (Thermo Fisher, Cat # 11140050). All culture media were devoid of antibiotics or antimycotics and cells were maintained at 37 °C in a humidified environment containing 5% CO2. For hypoxic culture conditions, 37 °C cells were incubated in a modular incubator chamber (Billups Rothenberg Inc.) containing 1% O2, 5% CO2, and 94% N2. Following this setup, cells were exposed to hypoxic conditions for durations ranging from 4 to 24 h. To create physioxia, an oxygen sensor (Forensics Detectors, model FD-103-O2) was used to precisely mix atmospheric gas with hypoxia gas to achieve a final concentration of 8–8.5% O2 in a hypoxia chamber (Billups Rothenberg Inc.). The chamber was then incubated at 37 °C for the specified duration to maintain stable physioxic conditions for the experiment.
Stable SUM1315 cell lines overexpressing HIF1α were established through the transfection of HA-HIF1α P402A/P564A-pcDNA3 (Addgene #18955) or an empty control vector (pcDNA3.1+) (Invitrogen, Cat # V79020) using Lipofectamine 2000 (Thermo Fisher) following the supplier’s instructions. Selection was achieved using G418 Sulfate (Thermo Fisher, Cat # 10131027) at a concentration of 500 µg/mL. The cell culture was performed in media devoid of antibiotics and antimycotics, except when specified, in a controlled atmosphere at 37 °C and 5% CO2 to ensure optimal growth conditions.
In our study, SUM1315 and SKBR3 cells were transfected with HIF1α siRNA (ON-TARGETplus Human HIF1α siRNA—SMARTpool, Horizon Discovery) or a non-targeting control siRNA (ON-TARGETplus Non-targeting Pool, Horizon Discovery), employing Lipofectamine™ 2000 Transfection Reagent for the process. Following a 16-h incubation period, cells were subjected to an 8-h hypoxia treatment. Subsequently, RNA and protein lysates were harvested for analysis.
In our experiments, SUM1315 cells were transfected with siRNAs targeting NONO (ON-TARGETplus Human NONO siRNA—Horizon Discovery), TMPO (ON-TARGETplus Human TMPO siRNA, Horizon Discovery), and NPM1 (ON-TARGETplus Human NPM1 siRNA—Horizon Discovery), alongside a non-targeting control siRNA (ON-TARGETplus Non-targeting Pool, Horizon Discovery) as a baseline comparison, utilizing Lipofectamine™ 2000 Transfection Reagent. After transfection, cells were allowed to incubate for 16 h prior to exposure to either 8 h of hypoxic or normoxic conditions. Subsequent to this exposure, RNA and protein lysates were extracted for further analysis.
Lentiviral vector production and transduction
Lentiviral vectors encoding WT-NoLS and Mut-NoLS HIF1α were procured from GenScript (Catalog No. SC1394-VP-1, Lot Nos. U714WHBBGO-3/P2JH001 and U714WHBBGO-6/P2JH001). The viral titers for each preparation were determined via quantitative fluorescence assays. The titers were 8.03 × 108 IFU/mL for the Mut-NoLS HIF1α (P402A/P564A) variant and 5.63 × 108 IFU/mL for the WT-NoLS HIF1α (P402A/P564A) variant. Both preparations were tested for mycoplasma contamination using commercial kits.
SUM1315 cells were transduced with lentiviral constructs encoding either WT-NoLS or Mut-NoLS HIF1α, while EMT6 cells were transduced with the WT-NoLS HIF1α lentiviral vector. Cells were seeded in 6-well plates and transduced with 5–10 µL of viral supernatant per well. To enhance transduction efficiency, polybrene (Sigma-Aldrich, Cat#TR-1003-G) was added to the transduction medium at a final concentration of 1:1000. The cells were incubated with the viral supernatant for 24 h under standard conditions (37 °C, 5% CO2). The transduction was carried out under standard culture conditions (37 °C, 5% CO2) for 24 h. Following incubation, the viral medium was replaced with fresh growth medium, and cells were allowed to recover and expand for subsequent assays. To generate stable cell lines, transduced cells were subjected to selection using puromycin at the appropriate concentration for each cell type. All lentiviral transductions were performed under strict sterile conditions to maintain consistency, ensure reproducibility, and minimize contamination across experiments.
EU incorporation assay
As previously described64,65, the rate of rRNA synthesis was estimated as a readout of 5-ethynyl uridine (EU) incorporation into nascent rRNA transcripts. SkBr3, T47D, and SUM1315 cells were plated 150–200 K cells in 35 mm plates with coverslips. Cells were treated with hypoxia for 8 h. Click-iT™ RNA Alexa Fluor™ 488 HCS Assay kit (Invitrogen# C10327) was used. After the end of treatment, EU was added to the media directly to the final concentration of 2 mM and incubated for 1 h at 37 °C. After EU incubation, media was removed, and cells were fixed with 3.7% Formaldehyde in PBS for 15 min. The fixative was then removed, and plates were washed two times with PBS. 500 µl of Click-iT reaction cocktail was added and kept in room temperature for 30 min. After the removal of Click-iT reaction cocktail, 500 µl of Click-iT reaction rinse buffer was added and then washed with PBS two times. 5% BSA/0.3% Triton in PBS solution was used to as a blocking solution for 1 h. Primary antibody for Fibrillarin (FBL) (1:500; Abcam# ab166630) was added to the plates and incubated overnight at 4 °C. After washing, goat anti-Rabbit Secondary Antibody, Alexa Fluor™ 594 (1:400; Invitrogen # A-11012) was added and incubated in the dark for 1 h and then washed with PBS for 3 times. Coverslips were mounted with VECTASHIELD® PLUS Antifade Mounting Medium with DAPI (Vector Laboratories # H-2000). Images were acquired by Nikon Eclipse Ti-U and analyzed according to the manufacturer’s protocol to calculate the binary intersection mean intensity using NIS-Elements AR (Nikon) software.
Western blotting
Cell lysates were obtained by treating cells with 2× Laemmli buffer containing β-mercaptoethanol, which were then heated at 95 °C for five minutes. Following this, proteins were separated via SDS-PAGE and subsequently transferred onto polyvinylidene difluoride membranes. These membranes were then subjected to overnight incubation at 4 °C with primary antibodies HIF1α (human 1:500, BD Bioscience # 610959; mouse 1:1000, Calbiochem # 400080). Immunoblotting with β-actin (1:50000; Sigma-Aldrich, Cat# A3854, Sigma-Aldrich) and Fibrillarin (1:1000; abcam Cat # ab166630) were used to confirm equal loading for the lysates of whole-cell and the nucleolar fraction, respectively. Anti-rabbit or anti-mouse HRP-conjugated secondary antibody (1:5000; GE Healthcare) were used for detection, and blots were developed with Pierce™ ECL Western Blotting Substrate (Thermo Scientific Cat # 32209 or ECL™ Prime Western Blotting System (Cytiva Cat # RPN2232) and imaged using an Amersham Imager 600. Image Studio Lite (LI-COR Biosciences) software was used for densitometric analysis.
Puromycin incorporation assay for protein synthesis
To investigate the effect of hypoxia on protein synthesis, we conducted a puromycin incorporation assay using SKBR3 and T47D cell lines. Cells were treated with either normoxia or hypoxia for 8 h, followed by two washes with PBS. Puromycin dihydrochloride (Sigma-Aldrich, Cat# P8833) was then added to the culture medium at a final concentration of 1 µM, and cells were incubated for 30 min at 37 °C to label newly synthesized proteins. Following incubation, cells were lysed in RIPA buffer, and whole-cell lysates were subjected to Western blot analysis. Nascent proteins labeled with puromycin were detected using Anti-Puromycin Antibody clone 12D10 (1:5000; Sigma-Aldrich, cat # MABE343), with α-tubulin used as a loading control using anti-α-Tubulin (DM1A) (1:10000; Cell Signaling, Cat # 12351). Densitometric analysis was performed to quantify the incorporation of puromycin, and the values were normalized to control conditions.
Nucleolar staining
SkBr3, T47D, and SUM1315 Cells were plated 1–2 × 105 cells on FluoroDishes (World Precision Instruments, Cat# FD35-100) and allowed to incubate overnight. Following this, the cells were subjected to hypoxia for 8 h. After that, nucleoli were labeled by NUCLEAR-ID® Green detection kit (Enzo Life Sciences # ENZ-51009-500) and the nuclei were stained using NucBlue™ Live ReadyProbes™ Reagent (Invitrogen # R37605) per manufacturer’s protocol. Live cells imaging was captured using a Nikon Eclipse Ti-U microscope, maintaining consistent exposure times for all captured images. By analyzing 120 cells across 10 random fields, each nucleus in the field was assessed for nucleoli, and the overall count of nucleoli per nucleus was determined using NIS-Elements AR (Nikon) software.
Nucleolar isolation
Nucleoli were enriched following established protocols66,67. After plating cells, a 36-h incubation period was followed by exposure to either normoxic or hypoxic conditions for 8 h. Subsequently, cells underwent washing steps and were collected in 1× PBS followed by centrifugation. Osmotic shock was induced, and cell lysis was achieved with the addition of NP-40, followed by homogenization using a tight Dounce homogenizer. Nuclei were then purified through a 250 mM sucrose cushion, subsequently sonicated to release the nucleoli. These nucleoli were further purified through a 340 mM sucrose cushion. The lysate from the nucleolar fraction was then utilized for immunoblotting analysis.
Confocal microscopy
SkBr3, T47D, and SUM1315 cells were plated 1–2 × 105 cells in 35 mm plates with coverslips. Cells were treated with hypoxia for 8 h. Following this, the culture media was aspirated, and the cells underwent fixation using 3.7% formaldehyde in PBS for a duration of 10 min. Subsequently, the fixative was carefully aspirated, and the plates were rinsed twice with PBS. 0.3% Triton in PBS was added for 15 min for permeabilization, then 5% BSA solution was added for 1 h for blocking. After washing, cells were incubated with primary mouse anti- HIF1α (1:50; BD Bioscience # 610959) and rabbit anti-POLR1A (1:250; Invitrogen # PA5-56766) overnight at 4 °C. Following the washing step, coverslips were exposed to a secondary antibody solution comprising anti-Rabbit, Alexa Fluor™ 594 (dilution 1:250; Invitrogen # A-11012) and anti-Mouse, Alexa Fluor™ 488 (dilution 1:250; Invitrogen # A-11001) for a duration of 1 h. Following PBS washing, coverslips underwent an incubation period with UBF (F-9) Alexa Fluor® 647 (dilution 1:50; Santa Cruz Biotechnology # SC-13125) for a duration of 1 h. Subsequently, coverslips were mounted using VECTASHIELD® PLUS Antifade Mounting Medium with DAPI (Vector Laboratories, Cat # H-2000). Cells were imaged with Nikon A1R Confocal Microscope using the Plan Apo 60× Oil λS DIC N2 objective to capture high-resolution images of the nucleoli. For Z stacks, images were taken at step intervals of 0.2 µm thickness, and overlayed in the NIS-Elements software to generate 3D images. The intensity profile and co-localization analysis, quantified as Manders’ Overlap Coefficient, were assessed utilizing NIS-Elements AR (Nikon) software.
Immunofluorescence
SUM1315 cells, including EV and HIF1α-overexpressing stable cells, were plated on poly-L-lysine-coated coverslips at a density of 75,000–80,000 cells per well. After 24 h of incubation to ensure cell attachment, cells were treated with either DMSO (control), 50 nM BMH-21 (Selleckchem, Cat # S7718), or Actinomycin D (5 µg/ml) (Enzo Life Sciences, Cat # 51006-ACD) for 8 h. Treatments were conducted under normoxic (21% O2) or hypoxic (1% O2) conditions. Following treatment, cells were fixed with 3.7% formaldehyde for 10 min at room temperature, washed twice with PBS, and permeabilized using 0.3% Triton ×-100 in PBS for 15 min.
Cells were blocked with 5% BSA in 0.3% Triton ×-100/PBS for 1 h at room temperature, followed by overnight incubation at 4 °C with primary antibodies in 1% BSA/PBS containing 0.3% Triton ×-100. The primary antibodies used were Mouse Monoclonal Anti-Human HIF-1α (1:50 dilution; BD, Cat # 610959), Rabbit Polyclonal Anti-Nucleophosmin 1 (1:100 dilution; Abcam, Cat # ab15440), and Rabbit Recombinant Anti-Fibrillarin (1:500 dilution; Abcam, Cat # ab166630). After three washes with PBS (10 min each), cells were incubated with the appropriate secondary antibodies, Alexa Fluor 488 (1:250 dilution; Goat anti-Mouse IgG, Invitrogen, Cat # A-11001) and Alexa Fluor 594 (1:250 dilution; Goat anti-Rabbit IgG, Invitrogen, Cat # A-11012), for 1 h at room temperature. Following secondary antibody incubation, cells were washed three times with PBS (10 min each). Nuclei were counterstained with VECTASHIELD PLUS Antifade Mounting Medium containing DAPI (Vector Laboratories, Cat # H-2000). Coverslips were mounted onto glass slides, sealed, and imaged using a Nikon Eclipse Ti microscope. Colocalization coefficients and visualization were analyzed using NIS-Elements Advanced Research software.
Real-time PCR
The RNeasy Mini Kit was used to extract RNA from cells (Qiagen, Germantown, MA). The rate of RNA Pol I transcription was determined using real-time PCR to assess the levels of the transient 5′ external transcribed spacer (5′ETS) rRNA27,41. 1 μg of total RNA was used with High-Capacity cDNA kit (Thermo Fisher) to generate the cDNA. PCR reactions were performed with 1:100 diluted cDNA, 2× Maxima SYBR Green Master Mix (Thermo Fisher, Waltham, MA) along with the following primer sets: Human 5′ETS 851-961 Forward primer-GAACGGTGGTGTGTCGTT, and Reverse primer-GCGTCTCGTCTCGTCTCACT; Human Actin Forward primer-CATGTACGTTGCTATCCAGGC, and Reverse primer-CTCCTTAATGTCACGCACGAT; mouse 45S rRNA ITS1 Forward primer-CCGGCTTGCCCGATTT, and Reverse primer-GGCCAGCAGGAACGA; mouse Actin Forward primer-GGCTGTATTCCCCTCCATCG, and Reverse primer-CCAGTTGGTAACAATGCCATGT. The PCR reactions were conducted using the Applied Biosystems Step One Plus Real-time PCR machine. Relative fold changes in 5′ETS transcripts were analyzed using the ΔΔCT method.
To evaluate the transcriptional regulation of HIF1α target genes, quantitative real-time PCR (qRT-PCR) was performed using TaqMan™ probe-based assays. Total RNA was extracted from untransfected, WT-NoLS HIF1α, and Mut-NoLS HIF1α-transfected SUM1315 cells using RNeasy Mini Kit (Qiagen, Germantown, MA). RNA concentration and purity were assessed using a Nanodrop spectrophotometer.
cDNA was synthesized using a high-capacity cDNA reverse transcription kit (Thermo Fisher Scientific), following the manufacturer’s protocol. qRT-PCR reactions were set up in a 10-μL reaction volume containing 5 μL of TaqMan™ Fast Advanced Master Mix (Thermo Fisher, Catalog #4444557), 0.5 μL of gene-specific TaqMan™ probe, 2 μL of cDNA (diluted 1:10), and 2.5 μL of nuclease-free water. Reactions were performed in technical triplicates using the QuantStudio™ 3 Flex Real-Time PCR System (Applied Biosystems). The following TaqMan™ Gene Expression Assays were used: LDHA (Hs00855332-g), CA9 (Hs00154208-m), SLC2A1 (Hs00892681-m), PDK1 (Hs01561847-m), and HK2 (Hs00606086-m). ACTB (Hs99999903-m) was used as an endogenous control for normalization. The thermal cycling conditions included an initial enzyme activation step at 95 °C for 20 s, followed by 40 cycles of denaturation at 95 °C for 3 s and annealing/extension at 60 °C for 30 s. Gene expression levels were analyzed using the ΔΔCt method, normalizing each gene to ACTB as the housekeeping control.
Data were plotted as fold change relative to non-transfected controls, with statistical analysis performed using one-way ANOVA with post hoc comparisons to determine differences between groups. Results are presented as mean ± SEM from at least three independent experiments.
Analysis of ribosomal RNA processing
RNA was extracted from SKBR3, T47D, and SUM1315 cell lines following 8 h of exposure to either normoxia (21% O2) or hypoxia (1% O2) using the RNeasy Mini Kit. A total of 1 μg of RNA was reverse transcribed into cDNA using the High-Capacity cDNA kit (Thermo Fisher). qPCR reactions were performed with 1:100 diluted cDNA using the 2× Maxima SYBR Green Master Mix (Thermo Fisher), with beta-actin mRNA serving as the internal control.
The primer sets (as described before ref. 32) for detecting rRNA precursors included:
45S-f (5′-GAACGGTGGTGTGTCGTT-3′) and 45S-r (5′-GCGTCTCGTCTCGTCTCACT-3′); 18S-5′-junction-f (5′-GCCGCGCTCTACCTTACCTACCT-3′) and 18S-5′-junction-r (5′-CAGACATGCATGGCTTAATCTTTG-3′); 18S-3′-junction-f (5′-AGTCGTAACAAGGTTTCCGTAGGT-3′) and 18S-3′-junction-r (5′-CCTCCGGGCTCCGTTAAT-3′); 5.8S-5′-junction-f (5′-TACGACTCTTAGCGGTGGATCA-3′) and 5.8S-5′-junction-r (5′-TCACATTAATTCTCGCAGCTAGCT-3′); 5.8S-3′-junction-f (5′-GAATTGCAGGACACATTGATCATC-3′) and 5.8S-3′-junction-r (5′-GGCAAGCGACGCTCAGA-3′); 28S-5′-junction-f (5′-CCGAGACGCGACCTCAGAT-3′) and 28S-5′-junction-r (5′-TCCGCTGACTAATATGCTTAAATTCA-3′).
To detect mature rRNAs, the following primers were used: 18S-f (5′-GATGGTAGTCGCCGTGCC-3′) and 18S-r (5′-GCCTGCTGCCTTCCTTGG-3′); 5.8S-f (5′-ACTCGGCTCGTGCGTC-3′) and 5.8S-r (5′-GCGACGCTCAGACAGG-3′); 28S-f (5′-GTGACGCGCATGAATGGA-3′) and 28S-r (5′-TGTGGTTTCGCTGGATAGTAGGT-3′). The internal control primers for beta-actin were Human Actin Forward: CATGTACGTTGCTATCCAGGC and Human Actin Reverse: CTCCTTAATGTCACGCACGAT. Data were plotted as normalized fold changes relative to the normoxic control to assess the transcriptional and processing changes in ribosomal RNA under hypoxic conditions.
Cell proliferation assay
SUM1315 cells, either overexpressing HIF1α or carrying the EV, were plated in a 96-well format, with an initial seeding density of 5000 cells per well and cultured in media supplemented with 50 nM BMH-21 (Selleckchem, cat no. S7718) for various intervals (4, 8, 12, 24, 48 h). Subsequently, the media was replaced with fresh media containing CyQUANT Cell Proliferation Assay reagent (Thermo Fisher), followed by an incubation period of 1 h. Fluorescence measurements were performed as per the reagent manufacturer’s instructions to evaluate cell viability. The data was normalized, analyzed, and presented using GraphPad Prism version 10.
Invasion assay
BD Biocoat Matrigel invasion chambers (Corning, 8.0 µm pore size, Cat. No. 354480) were allowed to warm at room temperature. Warm, serum-free growth medium (500 µl) was added into the inserts and the inserts were allowed to rehydrate for 2 hr at 37 °C. Media was carefully removed and then 20,000 cells (in 500 µl serum-free medium) were added to each insert. Growth medium (750 µl, serum-free) containing 10 µg/ml fibronectin was added to the lower wells. Cells were incubated at 37 °C for 8–24 h (as specified in respective figure legends), either in hypoxia or normoxia with BMH-21 or CTRL, and thereafter fixed in 4% paraformaldehyde for 10 min and stained with 0.5% crystal violet (Difco Laboratories) for 15 min. Filters were rinsed in water and air dried. Experiment was done in triplicate and 4 images per insert were captured using Nikon Eclipse E200LED microscope using 10× objective lens. Counting the number of invaded cells per field was performed using ImageJ68.
Migration assay
Cell culture insert filters (8 µM pore-sized, BD Falcon # 353097) were coated with 600 µl of serum-free media containing 6 ng/ml of gelatin overnight then rehydrated with 150 µl of warm serum-free media at room temperature for 90 min. Media was carefully removed, and the inserts were placed in wells containing 750 µl of serum-supplemented media. Cells (50,000) in 500 µl serum-free media were seeded in each insert and incubated at 37 °C for 4–8 h (as specified in respective figure legends) either in hypoxia or normoxia with BMH-21 or CTRL. Cells were fixed in 4% paraformaldehyde for 10 min and stained with 0.5% crystal violet for 15 min. Filters were rinsed in water and air dried. Experiment was done in triplicate and 4 images per insert were captured using Nikon Eclipse E200LED microscope using 10× objective lens. Counting the number of migrated cells per field was performed using ImageJ68.
Three-dimensional (3D) cell cultures
3D cell cultures were established using the Cultrex 3D Culture Matrix Reduced Growth Factor Basement Membrane (Fisher Scientific, Catalog # 3445-010-01), prepared by thawing overnight at 4 °C to ensure it remained liquid until application. We coated eight-well glass chamber slides (Millipore, Catalog# PEZGS0816) with 150 µl of Cultrex, ensuring uniform application without air bubbles, and allowed the matrix to solidify for 30 min in a cell culture incubator. Cell preparation entailed standard detachment protocols, resuspending cells to a density of 25,000 cells/ml and combining them 1:1 with 2× Assay media containing 4% Cultrex. We added 400 µl of this mixture to the coated wells, resulting in a media concentration of 1× with 2% Cultrex and 5000 cells per well. These cultures were incubated for 10 days, receiving media changes every other day.
For staining, we washed the cultures with PBS, fixed them with a 2% paraformaldehyde and 0.1% glutaraldehyde mix, and then permeabilized with 0.1% Triton ×-100 in PBS. Following blocking in a solution containing 5% BSA with 0.1% Triton ×-100, cultures underwent subsequent washes before being mounted with Vectashield containing DAPI (Vector Laboratories, Cat # H-2000). For the visualization of acinar structures, we used a Nikon Eclipse Ti-U microscope equipped with a 20× objective. The circularity of cell clusters, serving as an indicator of their invasive potential, was quantitatively assessed using ImageJ software.
Chromatin Immunoprecipitation quantitative real-time PCR (ChIP -qPCR)
In our experimental setup, we seeded 2.5 million cells onto 10-cm culture dishes, adhering to the protocol specified by the SimpleChIP Plus Enzymatic kit from Cell Signaling Technologies. This procedure commenced with the fixation of cells using 1% formaldehyde at room temperature, which facilitated the isolation of cell nuclei and the subsequent fragmentation of chromatin via enzymatic digestion with micrococcal nuclease and sonication. For immunoprecipitation, we utilized 10 μg of cross-linked chromatin and 2–3 μg of targeted antibody, incubating the mixture overnight at 4 °C. The histone modification marks (H3K4me3, H3K27ac, H3K36me3, and H3K9me) were probed using specific antibodies from Abcam: Anti-Histone H3 (acetyl K27) (ab4729), Anti-Histone H3 (trimethyl K4) (ab8580), Anti-Histone H3 (trimethyl K36) (ab9050), and Anti-Histone H3 (monomethyl K9) (ab9045). Additionally, the binding affinity of HIF1α to the rDNA promoter was examined using an anti-HIF1α antibody from BD Bioscience (#610959).
Following chromatin elution and reversal of cross-linking, DNA purification was carried out from both ChIP and input samples to facilitate subsequent RT-qPCR analysis, utilizing 2× Maxima SYBR Green Master Mix (Thermo Fisher). The analysis was facilitated by primers targeting the rDNA promoter’s proximal regions. Specifically, primers for the upstream −350 region were Forward primer- CTGCCGCGATCCTTTCTGG and Reverse primer-TACGACCGGGACACACGAG; for the promoter −231 region, Forward primer-CCGAAAATGCTTCCGGCTCC and Reverse primer-TCCCCAAAAACGCCACCATC. We quantified the enrichment of histone modifications or HIF1α occupancy at the rDNA promotor by assessing the threshold cycle (C[T]) values between input and immunoprecipitated DNA. Percent input was calculated using the formula 2% × 2^(C[T] 2% Input Sample—C[T] IP Sample), and percent enrichment was determined relative to controls. Each qPCR assay was conducted in triplicate on an Applied Biosystems StepOnePlus system (Thermo Fisher), ensuring rigorous validation of our findings.
Chromatin immunoprecipitation mass spectrometry (ChIP-MS)
SUM1315 cells were subjected to either hypoxia or normoxia conditions for 8 h. Subsequently, cells were fixed with 1% formaldehyde for 10 min to cross-link DNA and proteins, then washed and lysed for chromatin extraction. Chromatin was digested with micrococcal nuclease and sonicated. Protein G beads, pre-washed in PBS with BSA, were incubated with 3 µg of anti-RPA194 antibody or a control isotype, rotating overnight at 4 °C. The antibody-bead complex was then incubated with 10 µg of chromatin overnight at 4 °C. After washing, the chromatin-bound proteins were eluted and prepared for LC-MS analysis. This involved heating in LDS sample buffer, reduction, and denaturation, followed by separation on a Bis-Tris gel and Coomassie staining. Gel lanes were digested with trypsin and analyzed via LC-MS. Data analysis was conducted using Scaffold 5, applying stringent criteria (99% protein threshold, at least two peptides per protein, and 80% peptide threshold) for high confidence identification, and less stringent criteria (80% protein threshold, at least one peptide) for broader coverage. Isotype control-associated proteins were excluded, and a fold change threshold of >2 in protein abundance at the rDNA promoter with RPA194 was used to indicate significant findings.
Immunoprecipitation (IP)
Cells were lysed using ice-cold 1(C[T] 2% Input Sample—C[T] IP Sample), andX RIPA lysis buffer (Millipore, Cat #20-188) supplemented with HALT protease and phosphatase inhibitors (Thermo Fisher, Cat # 78442). Protein lysates were quantified, and 1 mg of total protein was used for immunoprecipitation. The lysates were incubated with 3 µg of anti-NPM1 antibody (Abcam, Cat # ab15440) at 4 °C for 3 h to allow for the formation of immune complexes. Following this, Protein A/G agarose beads (Santa Cruz, Cat # sc-2003) were added to the lysates and incubated overnight at 4 °C to capture the immune complexes.
The agarose beads were washed once with 1× RIPA buffer for 5 min and then washed three times with 1× PBS for 5 min each to remove non-specific binding proteins. After the final wash, bound proteins were eluted from the beads by adding 2× Laemmli buffer, followed by heat denaturation. The eluted proteins were subjected to Western blot analysis. Immunoprecipitation of HIF1α (3 µg; BD Bioscience, Cat # 610959) was followed by probing with an anti-NPM1 antibody to confirm the co-immunoprecipitation of NPM1 with HIF1α. Conversely, immunoprecipitation of NPM1 followed by Western blotting with an anti-HIF1α antibody confirmed the reciprocal interaction.
Plasmid construction
Wild-type and mutant NoLS were cloned at the N-terminal of pEGFP-N1. Primers were designed to contain Kozak consensus sequence, and EcoRI and BamhI recognition sites, with the following primer sets: wild-type NoLS Forward primer-AATTCGCCACCATGGGCGGCGCGAACGACAAGAAAAAGATAA GTTCTGAACGTCGAAAAGAAAAGTCTCGAGATGCAGCCAGATCTCGGCGAAGTAAAGAATCTCGG, and Reverse primer-GATCCCGAGATTCTTTACTTCGCCGAGATC TGGCTGCATCTCGAGACTTTTCTTTTCGACGTTCAGAACTTATCTTTTTCTTGTCGTTCGCGCCGCCCATGGTGGCG; mutant NoLS For-AATTCGCCACCATGGGCGG CGCGAACGACGCGGCCGCGATAAGTTCTGAACGTCGAGCGGAAGCCTCTCGAGATGCAGCCAGATCTCGGCGAAGTGCGGAATCTCGG, and Reverse primer-GATCCC GAGATTCCGCACTTCGCCGAGATCTGGCTGCATCTCGAGAGGCTTCCGCTCGACGTTCAGAACTTATCGCGGCCGCGTCGTTCGCGCCGCCCATGGTGGCG. Transient transfection of SUM1315 and T47D cells was performed with prepared plasmids utilizing Lipofectamine 2000 (Thermo Fisher) in accordance with the manufacturer’s guidelines. Live cell imaging was subsequently achieved through the use of a Nikon Eclipse Ti-U microscope, which is fitted with a 40× objective.
Polysome profiling
Cells were exposed to either hypoxic or normoxic conditions before being treated with cycloheximide (100 µg/ml) (Millipore Sigma, Cat# 239764-1GM) five minutes before collection, followed by three washes in cold PBS containing 100 µg/ml cycloheximide. They were then lysed using a polysome lysis buffer comprising 1% Triton ×-100, 20 mM Tris (pH 7.4), 300 mM NaCl, 10 mM MgCl2, 0.5 mg/ml heparin (Fisher Scientific, Cat # H19), 100 µg/ml cycloheximide, 1× Halt protease inhibitor cocktail (Thermo Fisher, Cat# 78442), and 0.1 U/µl SUPERase•In RNase inhibitor (Invitrogen, Cat # AM2694), and incubated at 4 °C with agitation for ten minutes. Following centrifugation at 13,000 RPM for ten minutes, protein concentrations were determined using the Precision Red assay. An equal amount of protein was then applied to sucrose density gradients ranging from 10% to 50% and centrifuged using a Beckman SW41Ti rotor at 35,000 rpm for three hours and twenty minutes at 4 °C. The gradients were dissected by infusing 60% sucrose into each column base, with the absorbance at 254 nm of the separated fractions being monitored and analyzed as per the protocol provided by the Brandel BR-188 density gradient fractionation system. The quantification of 40, 60, and 80S peak areas was performed using Peak Chart software (Brandel) and the results were normalized and presented as ratios in GraphPad Prism 10.
RNA-seq analysis
RNA was extracted from cell pellet or polysome fractions utilizing the RNAeasy Mini kit (Qiagen) in accordance with the manufacturer’s instructions. Subsequent library construction entailed rRNA depletion and the generation of paired-end sequences, each 150 nucleotides in length, employing the Illumina HiSeq platform (GENEWIZ, US). Initial processing of the fastq-format sequencing outputs involved adaptor sequence removal using the Trimmomatic software, alignment via the HISAT2 algorithm, and PCR duplicate elimination with SAMtools, leading to the generation of BAM or SAM files. These files were then subjected to comprehensive analysis using the Partek Genomics Suite (St Louis, MO, USA), incorporating Reads per Kilobase of transcript per Million mapped reads (RPKM) normalization prior to conducting any statistical evaluations to identify differentially expressed genes based on fold change and significance levels.
STRING database (https://string-db.org/) was employed to perform a functional enrichment analysis of the actively translated transcripts in hypoxia, highlighting biological processes involved69. The enrichment strength, as denoted by the logarithmic ratio of observed to expected values, and the significance, adjusted by the False Discovery Rate employing Benjamini–Hochberg correction, were meticulously calculated for each Gene Ontology term. A network graph derived from STRING database interactions was generated to assess if there was high connectivity among the genes preferably translated in hypoxia compared to normoxia.
Pulmonary metastasis assays (PuMA)
PuMA was implemented following an established protocol70. We injected 2 × 105 GFP-tagged SUM1315 cells, either with EV or overexpressing HIF1α, into the tail veins of athymic nude mice. To ensure ethical treatment, mice were placed in a chamber and administering 100% CO2 at a controlled rate to induce rapid unconsciousness and minimize distress according to the euthanasia guidelines outlined by the American Veterinary Medical Association. Following euthanasia, lungs were harvested under sterile conditions, and the trachea was cannulated to instill 0.6% agarose prepared in PuMA assay medium. This assay media was composed of M-199 medium supplemented with essential growth factors and antibiotics. Following excision, the lungs sections were placed on Surgifoam saturated with culture medium, followed by replacement with medium containing 50 nM BMH-21. Subsequently, the media was renewed every two days, with meticulous repositioning of the lung sections.
At day 7, lung sections were imaged for analysis utilizing a Nikon SMZ800 stereo zoom microscope, and subsequent image analysis was performed using ImageJ software. Corrected total cell fluorescence was computed to assess metastatic colonization, with background fluorescence subtracted to account for nonspecific staining59.
The PuMA of EMT6 was performed to assess metastatic colonization following tail vein injection of 1 × 105 EMT6 cells expressing either EV or HIF1α into BALB/c mice. After 15 min post-injection, mice were euthanized, and under sterile surgical conditions, a thoracotomy was performed to expose the lungs. The trachea was cannulated, and 0.6% low-melting agarose in assay media was instilled to preserve the lung architecture. Lungs were then excised, sectioned, and placed onto Surgifoam scaffolds saturated with M-199 medium supplemented with growth factors and antibiotics.
On day 1 post-excision, the culture medium was replaced with media containing 100 nM BMH-21 or DMSO for designated treatment groups. Media changes were performed every other day, ensuring proper maintenance of the lung tissue sections. For metastatic burden assessment, BLI was performed on days 6 and 8 using the IVIS Lumina III (PerkinElmer). Imaging, data acquisition, and analysis were conducted using Living Image software (PerkinElmer). Bioluminescent signals were quantified as total photon flux (p/s) within a defined region of interest (ROI) for each lung section. This research was conducted with the approval of the IACUC at the University of Alabama at Birmingham, adhering to established ethical guidelines for animal experimentation.
Orthotopic murine mammary tumor studies
SUM1315 breast cancer cells, either expressing EV or overexpressing HIF1α, were prepared by suspending in Cultrex Basement Membrane Extract, Type 3, Pathclear (R&D Systems, Catalog #: 3632-010-02). 2.5 × 106 cells were then injected orthotopically into the inguinal mammary fat pad of 7-week-old female NSG mice (NOD.Cg-Prkdcscid Il2rgtm1Wjl/SzJ) obtained from The Jackson Laboratory. Tumor progression was closely monitored through caliper measurements, from which the mean tumor diameter and volume were calculated. Beginning on day 8 post-inoculation, the mice were subjected to intraperitoneal injections of either BMH-21 (10 mg/kg) or vehicle control, administered five times per week over a duration of 5 weeks. After the third week of treatment, survival surgeries were meticulously performed to excise the primary tumors. Following a period of 21 days post-surgery, euthanasia was performed according to the euthanasia guidelines outlined by the American Veterinary Medical Association, as mentioned before.
To investigate the impact of HIF1α overexpression in EMT6 cell line and BMH-21 treatment on tumor progression and metastatic dissemination, 2.5 × 105 EMT6 cells expressing either EV or HIF1α were suspended in HBSS (Thermo Fisher, Catalog #: 14025076) and orthotopically injected into the inguinal mammary fat pad of 7-week-old female BALB/c mice. The mice were randomly assigned into four experimental groups (n = 7 per group). Tumor growth was monitored three times per week using caliper measurements, from which tumor diameter and volume were calculated. BMH-21 (10 mg/kg) was administered intraperitoneally five times per week for five weeks, starting on day 8 post-inoculation. BLI was conducted weekly to assess tumor progression and metastatic dissemination.
After five weeks of treatment, survival surgery was performed to excise primary tumors. Following a 21-day post-surgery period, mice were euthanized in accordance with American Veterinary Medical Association (AVMA) euthanasia guidelines, and lung tissues were collected for ex vivo imaging.
To evaluate metastatic burden, both in vivo and ex vivo BLI were performed using the IVIS Lumina III (PerkinElmer). Imaging, data acquisition, and analysis were conducted using Living Image software (PerkinElmer). Total photon flux (p/s) within a defined ROI was quantified to measure luminescence intensity, providing insights into metastatic tumor burden.
Picrosirius red staining
The Picrosirius red stain kit was used as per the manufacturer’s instructions (Polysciences CAT# 24901, Warrington, PA). The procedure starts with the removal of paraffin and subsequent rehydration to distilled water. Hematoxylin staining was performed to enhance nuclear contrast, followed by thorough rinsing in distilled water. The sections were then sequentially treated with Solution A for 2 min, washed, immersed in Solution B for 60 min for the primary staining, and transitioned to Solution C for 2 min to finalize the staining process. A brief immersion in 70% ethanol for 45 s preceded the final steps of dehydration, clarification, and mounting of the slides. Stained sections were examined under a Leica Aperio ScanScope CS pathology slide scanner using a 40× objective lens to observe the stained fibers in bright field highlighting the detailed architecture and composition of the tissue.
Immunohistochemistry
Mouse tumor and tissue specimens were preserved in formalin and subsequently embedded in paraffin. Thin sections (5 μm) prepared from these paraffin-embedded samples, underwent deparaffinization and hydration through xylene and a series of ethanol dilutions. Antigen retrieval was facilitated by heating in sodium citrate buffer at boiling temperature for 5 min. Tissue sections were then treated with Dako’s Dual Endogenous Enzyme Block for 15 min to block nonspecific binding sites. This was followed by a washing step in a Tris-buffered saline containing 0.01% Triton ×-100. The sections were incubated overnight at 4 °C with PCNA primary antibody (PC10; Cell Signaling #2586, Danvers, MA) at a dilution of 1:2000. Post-primary antibody incubation, sections were exposed to Dako’s EnVision + System-HRP Labelled Polymer Anti-mouse for 40 min at ambient temperature. DAB + Substrate Chromogen System (Dako) was used for signal development for 3 min, and counterstaining was performed with diluted Harris Hematoxylin. Following dehydration, the sections were mounted using Cytoseal XYL (Epredia 8312, ThermoScientific, Cat # 22-050-262) under coverslips.
For CA9 immunohistochemistry (IHC) staining, paraffin-embedded tissue sections were first baked at 60 °C for 1 h to ensure proper adhesion of the sections. After baking, the slides were deparaffinized and rehydrated by immersing them in a series of baths: xylene (3 times), followed by graded ethanol (absolute, 95%, 70%) for 2 min each. Antigen retrieval was performed using citrate buffer, where slides were boiled for 5–7 min, followed by an additional 10-min incubation at 95 °C. The slides were then cooled in citrate buffer at room temperature for 30 min. Sections were thoroughly washed with deionized water and PBS to remove excess buffer and ensure tissue integrity.
Blocking was carried out by incubating each section with 1% BSA in PBS for 1 h at room temperature. Afterward, the primary antibody (Carbonic Anhydrase IX/CA9 Antibody, Novus Biologicals, Catalog # NB100-417, diluted 1:200) was applied to the sections and incubated overnight at 4 °C. The following day, slides were washed in Tris buffer, and HRP-conjugated polymer was applied for 40 min. After further Tris buffer washes, DAB + Substrate Chromogen System (Dako) was used to develop the signal, followed by counterstaining with hematoxylin. Finally, slides were dehydrated using ethanol and xylene and mounted with Cytoseal XYL. The IHC stained sections were visualized using a Leica Aperio ScanScope CS pathology slide scanner, equipped with 40× objectives, integrated within Aperio ImageScope 12.3.3 software, allowing for detailed examination. To quantify the staining intensity for each marker, the Immunoreactive Score method was employed, providing a standardized measure across samples71.
RNAscope for rRNA biogenesis
In our study, we utilized the RNAscope® 2.5 HD Detection Kit (ACD Bio, catalogue # 322370) to estimate the rRNA expression within FFPE tumors sections. The protocol began with slide preparation, where sections were baked at 60 °C for an hour, followed by deparaffinization in xylene and rehydration through a series of ethanol washes. For pretreatment, sections were incubated with RNAscope® Hydrogen Peroxide to quench endogenous peroxidases, subjected to heat-induced target retrieval, and treated with Protease Plus to enhance tissue permeability. The probe hybridization step involved applying a RNAscope® Probe (Hs-45SrRNA-900-154, ACD Bio, catalogue # 410401) to each section and incubating at 40 °C for two hours to allow targeted RNA binding. Signal amplification was achieved through sequential application of preamplifier and amplifier molecules. Detection of the RNA target was conducted using DAB Chromogen, producing a distinct brown precipitate, with sections subsequently counterstained with 50% Gill’s hematoxylin to delineate cellular nuclei. Finally, the slides were dehydrated, cleared, mounted, and examined with Leica Aperio ScanScope CS pathology slide scanner to capture and analyze the intensity of rRNA expression patterns in tumors sections.
To quantify the intensity of rRNA expression, we employed a color thresholding method using ImageJ software. This approach involved converting each image into an HSB (Hue, Saturation, Brightness) stack, followed by the application of specific threshold values for each component. The threshold ranges were set as 0–255 for both Hue and Saturation, and a narrower range of 0–120 for Brightness, to accentuate features of interest. These thresholds were then used to generate individual masks for each channel. By performing an AND operation on these masks, we ensured that only the overlapping areas across all three channels were retained for further analysis. This methodological approach allowed for precise isolation and analysis of specific features within the images, based on the set parameter values.
Statistical analysis
For data visualization and statistical analysis, GraphPad Prism 10 was used. All data were statistically analyzed using the one-way ANOVA with Tukey’s multiple comparison test or the student’s t-test (two-tailed). For all analyses, a p-value of 0.05 or below was considered statistically significant. All error bars represent standard deviations (unless otherwise stated as standard error of the mean). Variance was checked and was comparable between different treatment groups. Linear regression analysis and Kaplan–Meier survival curves were conducted utilizing GraphPad Prism 10. The log-rank test was also applied to assess statistical significance in survival data. All in vitro assays were performed in at least two separate biological replicates, with each replicate comprising a minimum of two technical measurements. Exact numbers of biological and technical repeats are detailed in the corresponding figure legends.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The RNA sequencing data generated in this study have been deposited in the NCBI Gene Expression Omnibus (GEO) under the accession codes GSE270358 (bulk RNA-seq) and GSE270359 (polysome-associated RNA-seq). The mass spectrometry proteomics data have been deposited in the PRIDE repository with the accession numbers PXD053134 (for nucleolar HIF1A binding partners) and PXD053040 (for RPA194-interacting proteins at rDNA loci in hypoxic breast cancer cells). Publicly available datasets analyzed in this study include TCGA breast cancer RNA-seq data downloaded from the UCSC Xena Browser, METABRIC microarray data from cBioPortal, and additional GEO datasets including GSE202203, GSE149132, GSE18494, GSE47533, GSE118683, GSE196634, GSE28844, GSE42568, GSE5460, GSE36771, and GSE209998. Spatial transcriptomic data were retrieved from 10× Genomics Spatial Data Sets, including: (1) Visium_HD_Human_Breast_Cancer_Fresh_Frozen, (2) Human Breast Cancer: Visium Fresh Frozen, Whole Transcriptome, (3) Human Breast Cancer: DCIS, Invasive Carcinoma (FFPE), and (4) Human Breast Cancer (Block A Section 1). All datasets were analyzed using Space Ranger pipelines and are covered under the CC BY 4.0 license from 10× Genomics. Source data are provided with this paper.
Code availability
The R scripts used to calculate hypoxia scores and to reproduce all transcriptomic analyses have been deposited in Zenodo (https://doi.org/10.5281/zenodo.15914115).
References
Momenimovahed, Z. & Salehiniya, H. Epidemiological characteristics of and risk factors for breast cancer in the world. Breast Cancer (Dove Med. Press) 11, 151–164 (2019).
Afifi, A. M. et al. Causes of death after breast cancer diagnosis: a US population-based analysis. Cancer 126, 1559–1567 (2020).
Cancer Stat Facts: Female Breast Cancer. SEER Cancer Stat Facts: Female Breast Cancer. (National Cancer Institute, Bethesda, MD, 2020) https://seer.cancer.gov/statfacts/html/breast.html.
Peart, O. Metastatic breast cancer. Radio. Technol. 88, 519M-539M (2017).
Hagemeister, F. B. Jr. et al. Causes of death in breast cancer: a clinicopathologic study. Cancer 46, 162–167 (1980).
Redig, A. J. & McAllister, S. S. Breast cancer as a systemic disease: a view of metastasis. J. Intern Med. 274, 113–126 (2013).
Fares, J. et al. Molecular principles of metastasis: a hallmark of cancer revisited. Signal Transduct. Target Ther. 5, 28 (2020).
Jaakkola, P. et al. Targeting of HIF-alpha to the von Hippel-Lindau ubiquitylation complex by O2-regulated prolyl hydroxylation. Science 292, 468–472 (2001).
Kallio, P. J. et al. Signal transduction in hypoxic cells: inducible nuclear translocation and recruitment of the CBP/p300 coactivator by the hypoxia-inducible factor-1alpha. EMBO J. 17, 6573–6586 (1998).
Tutzauer, J. et al. Breast cancer hypoxia in relation to prognosis and benefit from radiotherapy after breast-conserving surgery in a large, randomised trial with long-term follow-up. Br. J. Cancer 126, 1145–1156 (2022).
Huang, L. E. et al. Activation of hypoxia-inducible transcription factor depends primarily upon redox-sensitive stabilization of its alpha subunit. J. Biol. Chem. 271, 32253–32259 (1996).
Gilkes, D. M. & Semenza, G. L. Role of hypoxia-inducible factors in breast cancer metastasis. Future Oncol. 9, 1623–1636 (2013).
Gilkes, D. M. Implications of hypoxia in breast cancer metastasis to bone. Int. J. Mol. Sci. 17, 1669 (2016).
Vito, A., El-Sayes, N. & Mossman, K. Hypoxia-driven immune escape in the tumor microenvironment. Cells 9, 992 (2020).
Wu, Y., Sarkissyan, M. & Vadgama, J. V. Epithelial-mesenchymal transition and breast cancer. J. Clin. Med. 5, 13 (2016).
Dermit, M. et al. Subcellular mRNA localization regulates ribosome biogenesis in migrating cells. Dev. Cell 55, 298–313.e10 (2020).
Prakash, V. et al. Ribosome biogenesis during cell cycle arrest fuels EMT in development and disease. Nat. Commun. 10, 2110 (2019).
Elhamamsy, A. R. et al. Ribosome biogenesis: a central player in cancer metastasis and therapeutic resistance. Cancer Res. 82, 2344–2353 (2022).
Marchand, V. et al. High-throughput mapping of 2′-O-Me residues in RNA using next-generation sequencing (Illumina RiboMethSeq Protocol). In RNA Methylation: Methods and Protocols (ed. Lusser, A.) 171–187 (Springer New York: New York, NY, 2017).
Pelletier, J., Thomas, G. & Volarevic, S. Ribosome biogenesis in cancer: new players and therapeutic avenues. Nat. Rev. Cancer 18, 51–63 (2018).
Weeks, S. E. et al. Inhibiting beta-catenin disables nucleolar functions in triple-negative breast cancer. Cell Death Dis. 12, 242 (2021).
Harold, C. M. et al. Ribosomal RNA transcription regulation in breast cancer. Genes (Basel) 12, 502 (2021).
Elsharawy, K. A. et al. Prognostic significance of nucleolar assessment in invasive breast cancer. Histopathology 76, 671–684 (2020).
Bursac, S. et al. Dysregulated ribosome biogenesis reveals therapeutic liabilities in cancer. Trends Cancer 7, 57–76 (2021).
Catez, F. et al. Ribosome biogenesis: an emerging druggable pathway for cancer therapeutics. Biochem. Pharm. 159, 74–81 (2019).
Brumwell, A. et al. Hypoxia influences polysome distribution of human ribosomal protein S12 and alternative splicing of ribosomal protein mRNAs. RNA 26, 361–371 (2020).
Metge, B. J. et al. Hypoxia re-programs 2’-O-Me modifications on ribosomal RNA. iScience 24, 102010 (2021).
Buffa, F. M. et al. Large meta-analysis of multiple cancers reveals a common, compact and highly prognostic hypoxia metagene. Br. J. Cancer 102, 428–435 (2010).
Bos, R. et al. Levels of hypoxia-inducible factor-1alpha independently predict prognosis in patients with lymph node negative breast carcinoma. Cancer 97, 1573–1581 (2003).
Winter, S. C. et al. Relation of a hypoxia metagene derived from head and neck cancer to prognosis of multiple cancers. Cancer Res. 67, 3441–3449 (2007).
Panov, K. I. et al. The ribosomal gene loci-the power behind the throne. Genes (Basel), 12, 763 (2021).
Kwon, I. et al. Poly-dipeptides encoded by the C9orf72 repeats bind nucleoli, impede RNA biogenesis, and kill cells. Science 345, 1139–1145 (2014).
Ivan, M. & Kaelin, W. G. Jr The EGLN-HIF O(2)-sensing system: multiple inputs and feedbacks. Mol. Cell 66, 772–779 (2017).
Chilov, D. et al. Induction and nuclear translocation of hypoxia-inducible factor-1 (HIF-1): heterodimerization with ARNT is not necessary for nuclear accumulation of HIF-1alpha. J. Cell Sci. 112, 1203–1212 (1999).
Lachmann, A. et al. Massive mining of publicly available RNA-seq data from human and mouse. Nat. Commun. 9, 1366 (2018).
Potapova, T. A. et al. Distinct states of nucleolar stress induced by anticancer drugs. Elife 12, RP88799 (2023).
Scott, M. S., Troshin, P. V. & Barton, G. J. NoD: a nucleolar localization sequence detector for eukaryotic and viral proteins. BMC Bioinforma. 12, 317 (2011).
Uhlen, M. et al. Towards a knowledge-based Human Protein Atlas. Nat. Biotechnol. 28, 1248–1250 (2010).
Lanczky, A. & Gyorffy, B. Web-based survival analysis tool tailored for medical research (KMplot): development and implementation. J. Med. Internet Res. 23, e27633 (2021).
Bartha, A. & Gyorffy, B. TNMplot.com: a web tool for the comparison of gene expression in normal, tumor and metastatic tissues. Int. J. Mol. Sci. 22, 2622 (2021).
Peltonen, K. et al. A targeting modality for destruction of RNA polymerase I that possesses anticancer activity. Cancer Cell 25, 77–90 (2014).
Colis, L. et al. Design, synthesis, and structure-activity relationships of pyridoquinazolinecarboxamides as RNA polymerase I inhibitors. J. Med. Chem. 57, 4950–4961 (2014).
Jacobs, R. Q. et al. RNA Polymerase I is uniquely vulnerable to the small-molecule inhibitor BMH-21. Cancers (Basel) 14, 5544 (2022).
Peltonen, K. et al. Small molecule BMH-compounds that inhibit RNA polymerase I and cause nucleolar stress. Mol. Cancer Ther. 13, 2537–2546 (2014).
Morral, C. et al. Zonation of ribosomal DNA transcription defines a stem cell hierarchy in colorectal cancer. Cell Stem Cell 26, 845–861.e12 (2020).
Rankin, E. B. & Giaccia, A. J. Hypoxic control of metastasis. Science 352, 175–180 (2016).
Chen, Z. et al. Hypoxic microenvironment in cancer: molecular mechanisms and therapeutic interventions. Signal Transduct. Target Ther. 8, 70 (2023).
Sebestyen, A. et al. Hypoxia signaling in cancer: from basics to clinical practice. Pathol. Oncol. Res. 27, 1609802 (2021).
Vaupel, P. et al. Oxygenation of human tumors: evaluation of tissue oxygen distribution in breast cancers by computerized O2 tension measurements. Cancer Res. 51, 3316–3322 (1991).
Harris, A. L. Hypoxia-a key regulatory factor in tumour growth. Nat. Rev. Cancer 2, 38–47 (2002).
de Heer, E. C., Jalving, M. & Harris, A. L. HIFs, angiogenesis, and metabolism: elusive enemies in breast cancer. J. Clin. Invest. 130, 5074–5087 (2020).
Taylor, C. T. & Scholz, C. C. The effect of HIF on metabolism and immunity. Nat. Rev. Nephrol. 18, 573–587 (2022).
Semenza, G. L. HIF-1: upstream and downstream of cancer metabolism. Curr. Opin. Genet. Dev. 20, 51–56 (2010).
Mitrea, D. M. et al. Self-interaction of NPM1 modulates multiple mechanisms of liquid-liquid phase separation. Nat. Commun. 9, 842 (2018).
Lindstrom, M. S. NPM1/B23: a multifunctional chaperone in ribosome biogenesis and chromatin remodeling. Biochem. Res. Int. 2011, 195209 (2011).
Goldman, M. J. et al. Visualizing and interpreting cancer genomics data via the Xena platform. Nat. Biotechnol. 38, 675–678 (2020).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Lalonde, E. et al. Tumour genomic and microenvironmental heterogeneity for integrated prediction of 5-year biochemical recurrence of prostate cancer: a retrospective cohort study. Lancet Oncol. 15, 1521–1532 (2014).
Metge, B. J. et al. Ribosome biosynthesis and Hedgehog activity are cooperative actionable signaling mechanisms in breast cancer following radiotherapy. NPJ Precis Oncol. 7, 61 (2023).
Tafforeau, L. et al. The complexity of human ribosome biogenesis revealed by systematic nucleolar screening of Pre-rRNA processing factors. Mol. Cell 51, 539–551 (2013).
Bryant, C. J. et al. A high-throughput assay for directly monitoring nucleolar rRNA biogenesis. Open Biol. 12, 210305 (2022).
Reich, M. et al. GenePattern 2.0. Nat. Genet. 38, 500–501 (2006).
Miller, H. E. & Bishop, A. J. R. Correlation AnalyzeR: functional predictions from gene co-expression correlations. BMC Bioinforma. 22, 206 (2021).
Jao, C. Y. & Salic, A. Exploring RNA transcription and turnover in vivo by using click chemistry. Proc. Natl. Acad. Sci. USA 105, 15779–15784 (2008).
Calkins, A. S., Iglehart, J. D. & Lazaro, J. B. DNA damage-induced inhibition of rRNA synthesis by DNA-PK and PARP-1. Nucleic Acids Res. 41, 7378–7386 (2013).
Weeks, S. E. & Samant, R. S. Isolation of nucleoli for characterization of nucleolar contents to uncover clues to metastatic progression. Methods Mol. Biol. 2294, 269–274 (2021).
Kammerud, S. C. et al. Novel role of the dietary flavonoid fisetin in suppressing rRNA biogenesis. Lab. Invest. 101, 1439–1448 (2021).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Szklarczyk, D. et al. The STRING database in 2023: protein-protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res. 51, D638–D646 (2023).
Mendoza, A. et al. Modeling metastasis biology and therapy in real time in the mouse lung. J. Clin. Invest. 120, 2979–2988 (2010).
Fedchenko, N. & Reifenrath, J. Different approaches for interpretation and reporting of immunohistochemistry analysis results in the bone tissue - a review. Diagn. Pathol. 9, 221 (2014).
Acknowledgements
The authors would like to acknowledge the following funding sources for supporting this work: The Breast Cancer Research Foundation of Alabama (BCRFA) awarded to R.S.S. and L.A.S., and BXAl3374 (VA) awarded to R.S.S. The research was also supported by the National Cancer Institute Cancer Center Support Grant P30 CA013148 and utilized the UAB High Resolution Imaging Facility and O’Neal CCC Mass Spectrometry/Proteomics Shared Resource. We would like to thank Dr. Hebaallah Alsheikh and Bhavya Papineni for scientific discussion and Dr. Peter Anderson for assistance with scanning slides.
Author information
Authors and Affiliations
Contributions
All authors have read and approved the manuscript. The project was conceptualized by L.A.S. and R.S.S. A.E., B.J.M., C.A.S., M.H.E., D.C.H., S.C.K., R.S.S. and L.A.S. contributed to experimental design. A.E., B.J.M., D.C.H., C.A.S., D.C.H., S.C.K., M.H.E. and D.C. aided in data acquisition. A.E., B.J.M., C.A.S., S.C.K., M.H.E. and D.C. performed analysis and interpretation of data. A.E., B.J.M., C.A.S., M.H.E., D.C., R.S.S. and L.A.S. were involved in writing and revision of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Vanesa Lafarga, Annika Jogi, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Elhamamsy, A., Metge, B.J., Swain, C.A. et al. Hypoxic stress incites HIF1α-driven ribosome biogenesis that can be exploited by targeting RNA Polymerase I. Nat Commun 16, 8018 (2025). https://doi.org/10.1038/s41467-025-63315-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-63315-3