Abstract
Obesity poses a substantial threat to human health but lacks effective management. Recent advancements in large-scale deep sequencing and cryo-electron microscopy (cryo-EM) have transformed drug discovery paradigms. Leveraging prior genetics investigation, we pinpointed Leucine-rich repeat-containing G protein–coupled receptor 4 (LGR4) as a promising target for combating obesity. Here, we present cryo-EM structures of full-length LGR4 alone and in complex with RSPO2(FU). Notably, we develop an inhibitory nanobody (NB21) that blocks the binding of RSPO1/2 to LGR4, and we also determine the structure of the LGR4-NB21 complex. NB21-mFc (NB21 fused with mouse IgG2) effectively inhibits the canonical Wnt signaling pathway, thereby enhancing mitochondrial respiration and thermogenesis in beige adipocytes. In vivo, NB21-mFc increases energy expenditure by promoting the browning of white fat, conferring resistance to both diet-induced and genetic (ob/ob) obesity. Furthermore, LGR4 deficiency abolishes the effects of NB21-mFc in boosting the browning program and subsequent weight reduction. In summary, our study unveils structural insights into the LGR4-RSPOs and LGR4-NB21 complexes, paving the way for the development of an LGR4–targeting nanobody for weight loss.
Similar content being viewed by others
Introduction
Obesity threatens global health due to its escalating prevalence and its correlation with a multitude of complications, including diabetes, cardiovascular diseases, and cancers1,2,3,4,5,6. It is projected that by the year 2025, approximately one-fifth of the global adult population will be afflicted with obesity7. Various approaches, from lifestyle intervention to bariatric surgery, have been employed in the prevention and management of obesity, but these methods often exhibit low effectiveness or high risk8,9,10. While several anti-obesity medications are available, most are prescribed for short-term use due to severe psychiatric and cardiovascular adverse effects8,11,12,13. Therefore, identifying novel targets and developing safe and effective weight loss therapeutics are urgently needed.
The breakthrough in human genome study with high-throughput deep sequencing has ushered in a new era in drug development, spanning from genetic target discovery to clinical drug intervention14. A typical example is GPR75, an exciting new anti-obesity target first discovered by human molecular genetics from the UKBiobank study15. Notably, AstraZeneca and other companies have initiated the development of GPR75 antagonists as novel therapeutic agents16. Concurrently, advancements in cryo-electron microscopy (cryo-EM) have greatly accelerated our understanding of ligand recognition and receptor activation, especially in various G protein-coupled receptors (GPCRs)17,18, which have been employed in structure-based drug design (SBDD)19. The integration of new target identification with deep sequencing and the development of their structure-aided selective blockers using cryo-EM seems to be a promising strategy that merits investigation.
A gain-of-function (p.A750T) mutation in Leucine-rich repeat-containing G protein–coupled receptor 4 (LGR4, also known as GPR48) was identified as a predisposing factor for severe obesity in young individuals from the Genetics of Obesity in Chinese Youngs (GOCY) study, while, almost at the same time, the deCODE study in Iceland unveiled that over 300 individuals harboring a rare loss-of-function mutation (p.R126X) in LGR4 exhibited a reduction of body weight20,21,22. The ablation of LGR4 enhanced energy expenditure and reduced adiposity by promoting the conversion of preadipocytes from white to brown adipocytes21,23. LGR4, a new member of GPCR superfamily, owes crucial roles in maintaining endocrine and cardiometabolic homeostasis and has recently gained extensive acknowledgment24,25. Genetic and biological evidence suggest that LGR4 could be a potential therapeutic target for obesity; however, it is currently widely accepted that developing small molecule antagonists targeting the ectodomain (ECD) of the LGR4 appears to be challenging due to the presence of two too-smooth curved β sheets on its concave surface26.
LGR4 consists of a large ECD rich in 17 leucine-rich repeats (LRRs), a cysteine-rich hinge region, a 7-trans-membrane domain (7-TMD), and a short intracellular domain25,27,28. At least four different endogenous ligands of LGR4 have been reported, including R-spondin (RSPO) 1/2/3/4, Norrin, Nidogen-2, and RANKL29,30,31,32,33,34. Noteworthy, our previous studies, along with others, have shown that the binding of RSPO1/2 to LGR4 can inhibit beige fat thermogenesis or promote adipogenesis of white adipocyte by amplifying canonical Wnt/β-catenin signaling pathway, consequently contributing to obesity35. RSPOs comprise an N-terminal signal peptide, two furin-like cysteine-rich domains (FU1 and FU2), a thrombospondin type I repeat (TSR) domain, and a C-terminal domain36. Crystal structure analysis has revealed that FU2 of RSPO1 binds to the concave side of horseshoe LGR4(ECD) via electrostatic and hydrophobic interactions28,37. Although many studies have focused on the structure of the LGR4 and the LGR4-RSPOs complex, an intact structure of full-length LGR4, which is crucial for a comprehensive understanding of cellular signal transduction and drug development, remains unresolved due to several technical bottlenecks18,26,28,37.
In this condition, the employment of neutralizing nanobodies to inhibit LGR4, with the assistance of structural confirmation, could potentially serve as a valuable approach for the purpose of weight reduction. Additionally, focusing on antibodies that directly block LGR4 binding sites for RSPOs could help minimize side effects and improve specificity for anti-obesity therapeutic effects.
In this work, we successfully resolve the structure of the full-length LGR4 protein and the LGR4-RSPO2(FU) complex using cryo-EM. Furthermore, we identify a nanobody NB21 as an LGR4 blocker, which disrupts the binding of RSPO1/2 to LGR4. NB21-mFc (NB21 fused with mouse IgG2) promotes thermogenesis to combat obesity by inhibiting the canonical Wnt signaling pathway in vitro and in vivo. Our study highlights the potential for LGR4–targeting nanobody NB21 as a drug for weight loss.
Results
Cryo-EM structures of full-length LGR4 and LGR4-RSPO2(FU) complex
LGR4(ECD) structure exhibited a distinctive horseshoe shape, comprising 17 LRR motifs, thereby imparting dynamic flexibility26,28. To facilitate structural determination, we generated a high-affinity nanobody (NB52) targeting the ECD of LGR4, with a dissociation constant (KD) of 3.88 ± 0.47 nM38. This nanobody was further expanded into a larger “megabody” (MB52) to enhance particle size and optimize orientation, effectively serving as a fiducial marker. The structure of full-length LGR4 (LGR4-apo) was obtained in the presence of MB52 with an approximately 3.03 Å resolution, which produced a clear map that enables us to place its ECD and 7-TMD into a structural model and allow the unambiguous assignment of side chains of most amino acids (unless otherwise stated, the stabilizing MB52 is always added for other LGR4 complex) (Fig. 1a and Supplementary Fig. 1a, b). The overall structure of LGR4 bears resemblance to those of luteinizing hormone receptor (LHCGR/LGR2) and thyrotropin receptor (TSHR/LGR3) (Supplementary Fig. 2a, b)39,40. However, LGR4(ECD) forms an extended and curved arc spanning 180 degrees. This structure exhibits an upright conformation of the ECD relative to the membrane layer, resembling a right-handed solenoid with a slight twist to the right. Meanwhile, the 7-TMD of LGR4 is similar to the inactive conformation seen in class A GPCRs like rhodopsin and β2 adrenergic receptors41,42,43,44.
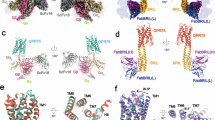
a Structure of the LGR4-MB52 complex. The cryo-EM map contoured at 0.157 (13.7σ) (left) and atomic model (right) are shown. b Structure of the MB52-LGR4-RSPO2(FU) complex. The cryo-EM map contoured at 0.153 (7.87σ) (left) and atomic model (right) are shown. c Interface between LGR4 and RSPO2(FU). d The cryo-EM densities for the interaction interfaces between LGR4 and RSPO2(FU) contoured at 0.153 (7.87σ). Interacting residues are drawn as sticks. LGR4 cyan, RSPO2(FU) purple, MB52 gray. The same color scheme is used throughout the manuscript unless stated otherwise.
To explore whether RSPO2 induces changes in the LGR4 conformation, we determined the cryo-EM structure of the LGR4-RSPO2(FU) complex with an overall resolution of 3.06 Å (Fig. 1b and Supplementary Fig. 1c, d). This revealed that LGR4 forms a 1:1 stoichiometric complex with RSPO2(FU). RSPO2(FU) binds to the concave inner surface near the top of the ECD through its FU2 domain (Fig. 1b), consistent with the previously reported structure of the LGR4(ECD)-RSPO1 complex26,28. Comparing the structures of LGR4 of LGR4-apo and the LGR4-RSPO2(FU) binary complex, we observed a high degree of structural similarity, except for minor interference from the side chains of certain interacting residues (Supplementary Fig. 2c). Aligning the 7-TMD of LGR4-apo with that in the LGR4-RSPO2(FU) complex demonstrated a strong structural overlap (Supplementary Fig. 2c). Although the ECDs of LGR4 in both structures largely coincide, there is a slight alteration in the rotation of the ECD. In the complex, the LGR4(ECD) exhibits a 2.5-degree rotation toward the 7-TMD compared to its position in the unbound LGR4, which may be due to the flexibility of the LGR4(ECD) relative to the 7-TMD (Supplementary Fig. 2c). The orientation of the LGR4(ECD) in this complex also aligns with that seen in the recently reported structures of inactive LHCGR/LGR2 and TSHR/LGR339,40, suggesting that RSPO2(FU) binding alone does not induce a transition to an active conformation in LGR4.
The LGR4-RSPO2(FU) interaction primarily centers on repeats LRR3–LRR7 within LGR4, which bind to residues from two FU domains of RSPO2(FU) (Fig. 1b). It buries approximately 2200.96 Å2 of solvent-accessible surface area. The interface can be dissected into two distinct regions. The first binding site is stabilized by multiple polar interactions. Specifically, salt bridges are formed between R86 of RSPO2(FU) and D137, D161, and D162 of LGR4, and between K58 from the FU1 domain of RSPO2(FU) and D162 of LGR4 (Fig. 1c, d). Additionally, R123 and K127 from the FU2 domain of RSPO2(FU) establish salt bridges with E228 and E252 of LGR4, respectively. Hydrogen bonds are also observed between H76, R121, and D108 of RSPO2(FU) and Q113, N226, and T229 of LGR4 (Fig. 1c, d). The second site is characterized by a hydrophobic patch, where interleaved stacking binding occurs. This interaction involves F105 and F109 from the FU2 domain of RSPO2(FU), and H157, W159, A181, V204, V205, and H207 from LGR4 (Fig. 1c, d). The most critical binding residues at the RSPO1/2 and LGR4 interface are highly conserved. In contrast, the conformation of their FU1 domain varies significantly (Supplementary Fig. 3a–c).
Notably, the loop region (residues 472–520) within the ECD was not resolved in our cryo-EM structure due to insufficient electron density. AlphaFold3 modeling showed that RSPO2(FU) binding sites are spatially distant from this unmodeled region (https://alphafoldserver.com/), indicating a mechanism distinct from the follicle-stimulating hormone (FSH)–induced activation of follicle-stimulating hormone receptor (FSHR/LGR1), where this region is essential (Supplementary Fig. 4)45,46. Additionally, unlike the G protein-dependent signaling of FSHR-FSH complex, our data indicate that RSPOs modulate LGR4 activity likely in a G protein-independent manner.
Development of a nanobody blocking the LGR4-RSPO1/2 complex
Based on our well-established LGR4–targeting nanobody phage display library38, we next screened out NB21 exhibiting significantly inhibitory properties against LGR4 in TOPFlash experiments (Supplementary Fig. 5a). NB21 was found to exhibit a high binding affinity to LGR4(ECD) with a KD of 1.12 ± 0.04 nM and effectively competed with RSPO1/2 binding (Supplementary Fig. 5b–d). Subsequently, we solved the cryo-EM structure of the LGR4-NB21 complex with a nominal resolution of 3.64 Å (Fig. 2a and Supplementary Fig. 6). The density map exhibited remarkable quality, enabling the construction of both the main and side chains of the protein. NB21 attached itself to the concave inner surface near the upper extremity of the LGR4(ECD) through its complementarity-determining regions (CDRs) (Fig. 2a–c). This interaction resulted in the burial of approximately 2260.04 Å2 of solvent-accessible surface area on LGR4. Notably, NB21’s orientation was nearly perpendicular to the cell surface, with the opposite end of its CDRs facing downwards (Fig. 2a). Comparing the LGR4-NB21 complex with the unbound LGR4 structure, we observed a high degree of structural congruence, except for minor perturbations in the sidechains of the specific residues that were induced by NB21 (Supplementary Fig. 2d).
a Structure of the MB52-LGR4-NB21 complex. The cryo-EM map contoured at 0.136 (12.2σ) (left) and the atomic model (right) are shown. b Interface between LGR4 and NB21. c The cryo-EM densities for the interaction interfaces between LGR4 and NB21 contoured at 0.136 (12.2σ). d Superposition of LGR4(ECD) in LGR4-RSPO2(FU) complex and LGR4-NB21 complex. Interacting residues are drawn as sticks. LGR4 cyan, NB21 pink, MB52 gray, RSPO2(FU) purple. The same color scheme is used throughout the manuscript unless stated otherwise.
Upon superimposing the LGR4-RSPO2(FU) complex with the LGR4-NB21 complex, it became evident that the binding epitope of NB21 overlaps with the binding sites of RSPO1/2 on LGR4 (Fig. 2d and Supplementary Fig. 3b). This overlap effectively hindered the interaction between LGR4 and RSPO1/2, which was critical for RSPO1/2’s role in bridging LGR4 and ZNRF3/RNF43 to enhance canonical Wnt signaling38. Thus, NB21 disrupted this association, theoretically resulting in impaired canonical Wnt signaling transduction.
The structural analysis reveals that NB21 established interactions with LGR4 through its three CDRs, specifically targeting LRR4/5/8/10 on LGR4 (Fig. 2a–c). The binding achieved by NB21 was facilitated by shape complementarity, preserving the conformation of LGR4 without disruption. The primary constituent of this interface was a network of hydrophilic interactions. At site I of this interaction, the side chain of K31 of NB21 forms two salt bridge with the side chains of both D137 and D161 on LGR4, which is also the RSPO1/2's binding residues (Fig. 2b, c and Supplementary Fig. 3b). Additionally, K31 also engaged in a σ-π interaction with the aromatic ring of W159 from LGR4. Furthermore, the phenol hydroxyl group of Y32 of NB21 donated a hydrogen bond to the side chain carboxyl of D162 in LGR4 (Fig. 2b, c). Y27’s aromatic ring and H157 in LGR4 participated in π-π interactions and formed a hydrophobic interaction with W159 in LGR4 (Fig. 2b, c). In site II of this interaction, R100 of NB21 formed a salt bridge with E228 in LGR4, while S102 of NB21 formed two hydrogen bonds with D231 and N233 in LGR4 (Fig. 2b, c). Moreover, these residues exhibited complete identity between human and mouse LGR4 (Supplementary Fig. 3b), indicating the possibility of NB21 being a species cross-reactive nanobody. We also found that NB21 has a high binding affinity to mouse LGR4, with a KD of 0.82 ± 0.01 nM (Supplementary Fig. 3d). To extend the half-life and increase the bioavailability of NB21, we engineered and expressed the NB21-mFc (NB21 fused with mouse IgG2), which was applied in the subsequent in vitro and in vivo functional experiments.
NB21-mFc blunts the effects of RSPO1/2 on beige adipocytes
We previously revealed that the germline gain-of-function mutations in human RSPO1 and LGR4 inhibit the browning capacity of white adipocytes and consequently promote adiposity partially via the canonical Wnt signaling pathway21,22,35,47. To evaluate the involvement of NB21 in the process, stromal vascular fractions (SVFs) of subcutaneous white adipose tissue (sWAT) were first treated with hRSPO1 alone or in addition to NB21-mFc. Increased levels of active (non-phosphorylated) alongside total β-catenin observed with hRSPO1 treatment were weakened with the addition of NB21-mFc (Fig. 3a). Consistently, NB21-mFc inhibited the nuclear β-catenin accumulation triggered by hRSPO1 (Fig. 3b) and mitigated its effects on the transcription of Tcf7l2, a marker of canonical Wnt signaling pathway activation, as well as other targets such as Axin2, Wisp2, Nkd1, CyclinD1, and c-Myc (Fig. 3c–h). RSPO2 also activated the canonical Wnt signaling pathway and inhibited browning progress in beige adipocytes (Supplementary Fig. 7a–d). In line with that NB21 blocks the binding sites shared by RSPO1/2 on LGR4 (Supplementary Fig. 3b), NB21-mFc suppressed the activation of the canonical Wnt signaling pathway and thermogenic gene expression induced by RSPO1 or RSPO2 in beige adipocytes in a dose-dependent manner (Fig. 3i–l and Supplementary Fig. 7e–h). These findings suggest that NB21-mFc can blunt the effects of RSPO1/2 on the canonical Wnt signaling pathway and the thermogenic gene expression in beige adipocytes.
a–h The SVFs derived from sWAT of C57BL/6 J mice treated with hRSPO1 (0.1 μg mL−1) alone or in addition to NB21-mFc (0.1 μM) for 48 h. a Immunoblot analysis (left) and quantification (right) of active (non-phosphorylated) and total β-catenin (n = 3 independent cell cultures per group). HSP90 was used as an internal control. b Representative images (left) of immunofluorescence staining of β-catenin and quantitative analysis (middle and right) of average fluorescence intensity of β-catenin in the nucleus. Analysis was performed on individual cells from randomly selected fields (n = 598 for PBS, n = 458 for hRSPO1, and n = 456 for hRSPO1 + NB21-mFc). Scale bar, 50 μm. c–h qPCR analysis of the downstream genes of canonical Wnt signaling pathway, including Tcf7l2 (c), Axin2 (d), Wisp2 (e), Nkd1 (f), CyclinD1 (g), and c-Myc (h) (n = 3 independent cell cultures per group). i–l qPCR analysis of Tcf7l2 (i) and thermogenic genes, including Ucp1 (j), Cidea (k), and Cox8b (l), in induced beige adipocytes pre-treated with hRSPO1 (0.1 μg mL−1) alone or in addition to different concentrations of NB21-mFc (0.01, 0.1, and 1.0 μM) from day -2–2 (n = 3 independent cell cultures per group). For a, c–l, values are presented as mean ± SEM. Kernel density plot (b, middle) shows the data distribution with the central line indicating the median; violin plot overlaid with box plot (b, right) show the median (center line), the 25th and 75th percentiles (box bounds), and whiskers extending to 1.5× IQR. Statistical analysis was performed using one-way ANOVA followed by Dunnett’s multiple-comparison test (a, c–l), and Kruskal–Wallis test followed by Bonferroni–Dunn test (b). SVFs stromal vascular fractions, sWAT subcutaneous white adipose tissue, qPCR quantitative PCR, IQR interquartile range, ANOVA analysis of variance. Source data are provided as a Source Data file.
NB21-mFc boosts thermogenesis in vivo
Next, we aimed to evaluate whether NB21-mFc could promote thermogenesis in vivo. With intraperitoneal injection of NB21-mFc for 7 days (Fig. 4a), mice displayed mild weight loss and fat mass reduction (Supplementary Fig. 8a, b). Using body weight as a covariant in ANCOVA analysis48,49,50, we observed a significant increase in total energy expenditure (EE) in NB21-mFc–treated mice (Fig. 4b and Supplementary Fig. 8c, d). Additionally, a slight increase in respiratory exchange ratio (RER) was observed in the NB21-mFc–treated group (Fig. 4c). No difference was detected in cumulative food intake, physical activity, or fecal calorie loss between the two groups (Supplementary Fig. 8e–g). Under acute cold exposure, NB21-mFc–treated mice exhibited higher EE and core temperature (Fig. 4d, e). Moreover, after chronic exposure to coldness for 7 days (Fig. 4f), smaller adipocytes were observed in visceral white adipose tissue (vWAT), sWAT, and brown adipose tissue (BAT), along with increased UCP1 protein expression in vWAT and sWAT in NB21-mFc–treated group (Fig. 4g, j, and Supplementary Fig. 8h). Consistently, the mRNA levels of thermogenic genes, including Ucp1, Cidea and Dio2, and mitochondrial respiratory genes, such as Pgc1α and Ndufa8, were significantly elevated in these fat depots in NB21-mFc–treated group (Fig. 4h, k and Supplementary Fig. 8i). Moreover, the protein levels of UCP1 together with mitochondrial respiratory chain, including UQCRC2 (complex Ⅲ), MTCO1 (complex Ⅳ), and NDUFB8 (complex Ⅰ), were increased in NB21-mFc–treated group (Fig. 4i, l). Together, these results reveal that NB21-mFc enhances adipose thermogenesis and energy expenditure in mice.
a–e 8-week-old female C57BL/6 J mice fed HFD were treated by intraperitoneal injection with PBS or NB21-mFc (0.2 mg kg−1) once daily (n = 8 mice per group). a Schematic diagram of treatment. b, c 24 h EE trace (b, left) and normalized EE with body weight as covariant (b, right), 24 h RER (c) in room temperature (22 °C) at day 7. d After the abovementioned experiments, EE of mice in (a) before and after acute cold exposure (5 °C) were recorded. The values under cold were calculated between 3–4 h. e After 21 days of treatment, body core temperature changes after 6 h cold exposure (5 °C). f–l 8-week-old female C57BL/6 J mice fed HFD were treated by intraperitoneal injection with PBS or NB21-mFc (0.2 mg kg−1) once daily under chronic cold stimulation (5 °C) for 7 days (n = 6 mice per group). f Schematic diagram of treatment. g, j Representative images of H&E staining, immunofluorescence staining of UCP1/Perilipin, and quantification of adipocyte area in vWAT (g) and sWAT (j). Scale bar, 100 μm. h, k qPCR analysis of thermogenic and mitochondrial respiratory complex genes in vWAT (h) and sWAT (k). i, l Immunoblot analysis and quantification of UCP1 and mitochondrial respiratory complexes in vWAT (i) and sWAT (l). For b–e, h, i, k, l, values are presented as mean ± SEM. Kernel density plot (g, j) shows the data distribution; violin plots overlaid with box plots (g, j) show the median (center line), the 25th and 75th percentiles (box bounds), and whiskers extending to 1.5× IQR. Statistical analysis was performed using two-sided ANCOVA (b), two-way ANOVA (c–e), two-sided Wilcoxon rank-sum test (g, j), and unpaired two-tailed Student’s t test (h, i, k, l). HFD high-fat diet, EE energy expenditure, RER respiratory exchange ratio, vWAT visceral white adipose tissue, sWAT subcutaneous white adipose tissue, qPCR quantitative PCR, H&E hematoxylin-eosin, IQR interquartile range, ANCOVA analysis of covariance, ANOVA analysis of variance. Source data are provided as a Source Data file.
NB21-mFc’s inhibition on browning depends on LGR4
RSPO1 binds to LGR4 yielding a robust inhibition on browning21,35, while the effective concentration of RSPO2 was approximately 20 folds higher than that of RSPO1 (Supplementary Fig. 7a–d), in consistency with previous findings51. Further analysis of a published single-cell dataset from mouse white adipose tissue (WAT) indicated that the expression level of Rspo1 was much higher than that of Rspo2 throughout the whole WAT (Supplementary Fig. 9a–c)52. This trend was also observed in the adipose stem and progenitor cells (ASPCs) subpopulation (Supplementary Fig. 9c), which play a crucial role in maintaining adipose tissue homeostasis and facilitating adipogenesis53. Similarly, qPCR analysis demonstrated that Rspo1 exhibits the highest expression in vWAT SVFs, in which ASPCs represent the predominant cell population54,55, significantly surpassing the expression levels of Rspo2 across all three types of adipose tissue (Supplementary Fig. 9d). Next, we verified that the neutralizing effects of NB21-mFc on the browning suppression of LGR4-RSPO1 complex were abolished in beige adipocytes deficient of LGR4 (Lgr4m/m), which is supported by the changes in the gene expression of thermogenesis and mitochondrial respiration (Fig. 5a–c and Supplementary Fig. 10a–f), as well as oxygen consumption capacity of induced beige adipocytes (Fig. 5d and Supplementary Fig. 10 g). Despite that NB21 was specifically designed to target the ECD of LGR4, sequence alignment reveals conserved NB21 binding regions among LGR4, LGR5 and LGR6, raising the possibility of cross-reactivity (Supplementary Fig. 11a). Although Lgr4 is also expressed at significantly higher levels compared to Lgr5 and Lgr6 in adipose tissue (Supplementary Fig. 9e, f)35, to further exclude the possibility of LGR5 or LGR6 mediating the effects of NB21 in beige adipocytes, we conducted lentivirus-mediated knockdown experiments targeting Lgr5 and Lgr6 in SVFs (Supplementary Fig. 11b, f). Remarkably, regardless of the effective silence of either Lgr5 or Lgr6, NB21-mFc consistently mitigated the inhibitory effects of RSPO1 on beige adipocyte thermogenesis (Supplementary Fig. 11c–e, g–i). These findings demonstrate that NB21-mFc enhances thermogenic capacity in adipose tissue primarily through LGR4, rather than LGR5 or LGR6.
a–d Induced beige adipocytes derived from WT and littermate Lgr4m/m mice treated with hRSPO1 (0.1 μg mL−1) alone or in addition to NB21-mFc (0.1 μM) from day −2–2. a–c qPCR analysis of Ucp1 (a), Cidea (b), and Cox8b (c) (n = 4 independent cell cultures per group). d OCR measurement (n = 8 independent cell cultures per group). e–j 8-week-old female WT and Lgr4m/m mice fed HFD were treated by intraperitoneal injection with PBS or NB21-mFc (0.2 mg kg−1) every other day for 6 weeks (n = 6 mice per group for WT, and n = 4 mice per group for Lgr4m/m). e Schematic diagram of treatment. f, g Body weight gain (f) and tissue weight of vWAT and sWAT (g) of two genotypes after 6-week treatment. h Representative images of H&E staining, immunofluorescence staining of UCP1/Perilipin, and quantification of adipocyte area in vWAT. Scale bar, 100 μm. i qPCR analysis of thermogenic and mitochondrial respiratory complex genes in vWAT. j Immunoblot analysis (left) and quantification (right) of UCP1 and mitochondrial respiratory complexes in vWAT. For a–d, f, g, i, j, values are presented as mean ± SEM. Kernel density plot (h) shows the data distribution; violin plots overlaid with box plots (h) show the median (center line), the 25th and 75th percentiles (box bounds), and whiskers extending to 1.5× IQR. Statistical analysis was performed using one-way ANOVA followed by Dunnett’s multiple-comparison test (a–d), unpaired two-tailed Student’s t test (f, g, i, j), and two-sided Wilcoxon rank-sum test (h). ns not significant, WT wild-type, OCR oxygen consumption rate, HFD high-fat diet, H&E hematoxylin-eosin, vWAT visceral white adipose tissue, sWAT subcutaneous white adipose tissue, qPCR quantitative PCR, IQR interquartile range, ANOVA analysis of variance. Source data are provided as a Source Data file.
Then, we aimed to test whether NB21-mFc could combat adiposity in vivo. We first evaluated the potential weight-loss effects of NB21-mFc in diet-induced obesity (Fig. 5e–j and Supplementary Figs. 12, 13). Under high-fat diet (HFD), female 8-week-old mice were intraperitoneally injected with either NB21-mFc (0.2 mg kg−1) or PBS once every other day for 6 weeks (Fig. 5e). Intriguingly, NB21-mFc significantly reduced body weight in female wild-type (WT) mice (Fig. 5f), mainly attributable to the reduction of vWAT and sWAT (Fig. 5g) featured with smaller adipocytes in parallel with more intense staining of the browning marker (UCP1) (Fig. 5h and Supplementary Fig. 12c). Consistent with in vitro findings, NB21-mFc enhanced the expression of genes involved in thermogenesis and mitochondrial respiration in vWAT and sWAT (Fig. 5i, j and Supplementary Fig. 12d). Consistent but relatively mild changes were observed in BAT (Supplementary Fig. 12e, f). Along with decreased diet-induced adiposity, NB21-mFc also reduced liver lipid accumulation (Supplementary Fig. 12g, h). Importantly, NB21-mFc lost the above browning-augment and anti-obesity effects in Lgr4m/m littermates (Fig. 5f–j and Supplementary Fig. 12a, c–h). Meanwhile, we observed that NB21-mFc yielded similar metabolic benefits in male WT mice but not in Lgr4m/m littermates (Supplementary Fig. 13). Overall, these findings suggest that NB21-mFc attenuates diet-induced adiposity by promoting thermogenic capacity mainly in an LGR4–dependent way.
NB21-mFc reduces adiposity in ob/ob mice
Lgr4 ablation could reduce adiposity even in severely obese models induced by Leptin monogenic mutation21. We next investigated the anti-obesity effect of NB21 in Lep-deficient (ob/ob) mice. 8-week-old ob/ob mice were treated with NB21-mFc at relatively low dosages (0.03 mg kg−1 or 0.1 mg kg−1) once every other day (Fig. 6a). After 7 days of treatment, NB21-mFc–treated ob/ob mice exhibited a significant increase in total EE and a decrease in RER (Fig. 6b, c and Supplementary Fig. 14a, b). No difference was detected in cumulative food intake, physical activity, or fecal calorie loss between the two groups (Supplementary Fig. 14c–e). After 7 weeks of treatment, NB21-mFc–treated ob/ob mice exhibited a dose-dependent decrease in body weight, along with decreased fat mass and increased lean mass (Fig. 6d–f). Specifically, the weights of three adipose tissues including BAT, sWAT and vWAT were all significantly decreased after NB21-mFc treatment (Fig. 6g, j and Supplementary Fig. 14f). Smaller adipocytes as well as enhanced UCP1 protein staining were observed in BAT, sWAT and vWAT after NB21-mFc treatment (Fig. 6h, k and Supplementary Fig. 14g). Consistently, the mRNA levels of thermogenic genes and mitochondrial respiration-related genes were elevated in BAT and sWAT of NB21-mFc–treated (0.1 mg kg−1) ob/ob mice (Fig. 6i, l), and an increase trend was observed in vWAT (Supplementary Fig. 14h). Meanwhile, severe ectopic lipid accumulation in the liver was attenuated by NB21-mFc (Supplementary Fig. 14i, j). Additionally, NB21-mFc–treated ob/ob mice showed a reduction in inflammation markers, especially Ccl2, Tnfα, and Il6, in the three adipose tissues (Supplementary Fig. 14k–m). These findings suggest that a relatively long-term use of NB21-mFc does not induce obvious immune responses, but rather alleviates obesity-related chronic inflammation, further supporting its therapeutic potential. Together, these data suggest that the NB21-mFc treatment reduces adiposity even in severely obese models caused by genetic defects.
8-week-old male ob/ob mice fed NCD were treated by intraperitoneal injection with PBS or NB21-mFc (0.03 mg kg−1 or 0.1 mg kg−1) every other day for 7 weeks (n = 6 mice per group). a Schematic diagram of treatment. b, c 24 h EE trace (b, left) and normalized EE with body weight as covariant (b, right), 24 h RER (c) in room temperature (22 °C) at day 7. d–f Body weight (d), fat mass (e), and lean mass (f) after 7-week treatment. g, j Tissue weight of BAT (g) and sWAT (j). h Representative images of H&E staining and immunofluorescence staining of UCP1/Perilipin in BAT. Scale bar, 100 μm. i, l qPCR analysis of thermogenic and mitochondrial respiratory complex genes in BAT (i) and sWAT (l). k Representative images of H&E staining, immunofluorescence staining of UCP1/Perilipin, and quantification of adipocyte area in sWAT. Scale bar, 100 μm. For b–g, i, j, l, values are presented as mean ± SEM. Kernel density plot (k) shows the data distribution; violin plots overlaid with box plots (k) show the median (center line), the 25th and 75th percentiles (box bounds), and whiskers extending to 1.5× IQR. Statistical analysis was performed using two-sided ANCOVA (b), two-way ANOVA (c), one-way ANOVA followed by Tukey’s multiple-comparison test (d–g, j), unpaired two-tailed Student’s t test (i, l), and Kruskal-Wallis test followed by Bonferroni–Dunn test (k). NCD, normal chow diet, EE energy expenditure, RER respiratory exchange ratio, BAT brown adipose tissue, sWAT subcutaneous white adipose tissue, qPCR quantitative PCR, H&E hematoxylin-eosin, IQR interquartile range, ANCOVA analysis of covariance, ANOVA analysis of variance. Source data are provided as a Source Data file.
Discussion
Recent achievements in genetic study in a large-scale population or extreme cases with deep sequencing have yielded numerous novel potential anti-obesity targets, notably including druggable GPCRs14. Despite widespread application of SBDD and optimization techniques over the past decade, successful clinical implementations for targeting GPCRs have been limited56,57. The high challenge of routine GPCRs crystallization constitutes a key factor18,57. Here, we have successfully resolved the cryo-EM structure of full-length LGR4 and further described the molecular details of LGR4 complexed with one of its ligands, RSPO2. In this context, we developed a neutralizing nanobody (NB21) that precisely targets the LGR4-RSPO1/2 binding domain, thereby specifically blocking RSPO1/2 binding to LGR4. Importantly, both in vitro and in vivo experiments confirmed the efficacy of NB21-mFc in promoting thermogenesis and weight loss.
Historically, drugs targeting GPCRs have typically been small-molecule drugs or peptides with low molecular weight58. However, there is an increasing interest in exploring the potential of therapeutic antibodies for modulating the function of GPCRs, primarily due to their remarkable selectivity and specificity58,59. Some advancements have been achieved in the field of drug development pertaining to LGRs, such as BNC101 targeting LGR5 for the treatment of gastrointestinal disease58,60, and the antibody targeting LGR1 (FSHR) for the treatment of obesity61,62.
Novel findings in human genetics facilitate the identification and clinical translation of innovative therapeutic approaches, with GPR75 serving as a representative instance14. In 2013, the deCODE study initially disclosed the presence of a rare loss-of-function mutation of LGR4 (p.R126X) within the Icelandic population20. It was observed that heterozygous carriers of this mutation showed a reduction in body weight. Simultaneously, we identified a low-frequency activating mutation LGR4 (p.A750T) associated with adiposity in GOCY study21,22. More recently, we successfully identified an activating mutation p.R219W in RSPO1, one of the LGR4 ligands, and several activating mutations (p.T59A, p.R124H, p.R274H, and p.G708E) in CTNNB1/β-catenin, the core component in the canonical Wnt signaling pathway, in young individuals with severe obesity35,47,63. Taken together, aberrant activation of the RSPO1–LGR4–CTNNB1/β-catenin pathway exhibits a strong association with adiposity, partly due to the suppression of beige adipose thermogenesis21,25,35,47,63. With valid human genetic evidence, this novel pathway holds promise for investigating weight-loss medications.
The transition from identifying molecular targets to drug approval is typically lengthy and costly, averaging 25 years14. In the field of weight-loss drugs, GLP-1R agonists were approved for marketing approximately 18 years after the discovery of GLP-164,65,66,67. Given the increasing availability of genetically validated drug targets, it is imperative to employ innovative methodologies such as structural biology information to mitigate the expenses and duration associated with drug development14,18. Within a concise timeframe, we successfully discerned a potential target LGR4 via deep sequencing in severely obese cases, subsequently leading to the screening and identification of a potential neutralizing nanobody, NB21, employing cryo-EM–assisted validation. We anticipate that this strategy will expedite the drug development pipeline for weight loss and the important implications for subsequent research on the transformation of candidate targets for obesity.
The LGR4-RSPOs-ZNRF3/RNF43 module plays a crucial regulatory function in preserving homeostasis of the canonical Wnt and other signaling pathways, thereby governing the state and development of stem cells, along with physiological processes such as energy homeostasis25,68,69. Consistent with our findings that RSPO1 promotes obesity, Dong et al. reported that RSPO2 overexpression contributes to obesity and insulin resistance, which is dependent on LGR470. Theoretically, it is plausible that the development of RSPO1/2-blocking drugs, such as neutralizing antibodies, may potentially lead to enhanced energy expenditure and weight reduction, but we needed at least two effective antibodies targeting RSPO1 and RSPO2, respectively, to block LGR4 activation. In addition, the absence of RSPO2 results in dysplasia of the fingertips and fetal demise, a process that depends on ZNRF3/RNF43 rather than LGR470, suggesting that NB21’s specific targeting LGR4-RSPO1/2 complex could potentially circumvent these adverse effects. Thus, the prioritization of developing neutralizing antibodies with high selectivity assumes greater significance in the context of weight-loss drug design.
Nonetheless, certain unresolved issues warrant attention. Firstly, the increased mass of the mFc fusion of NB21 could potentially cause steric hindrance. To mitigate this, a GGGGS linker was introduced between NB21 and mFc. Importantly, our functional results confirm that this modification does not affect the ability of NB21-mFc to promote browning and weight loss. Secondly, in the short-term HFD-fed model receiving NB21-mFc treatment, while we observed reduced fat mass and increased energy expenditure, the absence of RER decrease might be attributable to the insufficient fat stores used for fatty acid oxidation in these mice after only 7 days of HFD feeding. Importantly, in ob/ob mice with a high amount of fat storage, NB21-mFc significantly decreased RER along with reduced fat mass, indicating an increased fat oxidation. These data support the potent role of NB21-mFc in combating adiposity with a higher fatty acid oxidation capacity, especially in already obese conditions. Thirdly, the Norrin binding site on LGR4 overlaps with that of RSPO1/2, though not identically71. Norrin, recognized as another endogenous ligand of LGR4, is known for its role in retinal vascular development72. While NB21-mFc cannot penetrate the blood-retina barrier, its impact on the retina is speculated to be minimal. Fourthly, our previous findings indicated that LGR4 engages in competition with RANK for RANKL binding, which is crucial for maintaining bone homeostasis33. As the structure of the LGR4-RANKL complex remains elusive, computational modeling using AlphaFold3 suggests potential spatial overlap with the LGR4-RSPOs interaction site. However, computational predictions need validation by experimental evidence. The ability of NB21-mFc to induce browning and reduce adiposity in the ob/ob model at low concentrations suggests that optimizing its dosage and tissue distribution may aid in mitigating the potential adverse effects. Fifthly, the sequence homology among LGR4/5/6 suggests that NB21 may exhibit some cross-reactivity, potentially leading to off-target effects. However, due to the higher and more specific expression of LGR4 in adipose tissue and Lgr5/6 silence showing no effects on NB21, we expect the impact of cross-reactivity with LGR5/6 to be minimal in this context. Future studies will focus on optimizing NB21 targeting for LGR4, for instance, localized or targeted drug delivery methods could improve NB21’s selective targeting and enhance its therapeutic potential in adipose tissues. Finally, the gain-of-function A750T mutation in LGR4, located in the transmembrane helix 6 (TM6) region critical for GPCR activation, warrants further investigation. This mutation may stabilize an active-like conformation of LGR4, enhancing its role in canonical Wnt/β-catenin signaling or facilitating alternative downstream mechanisms, potentially contributing to obesity susceptibility. In addition, our findings also indicate that RSPO2 may not act as a conventional ligand for LGR4 and highlight the complexity of LGR4 activation mechanism. In a very recent study, we resolved the structure of the LGR4-RSPO2(FU)-ZNRF3 complex at a 2:2:2 stoichiometry38, in which RSPO2 appears to function as a linker facilitating interactions between LGR4 and ZNRF3 to promote canonical Wnt signaling. Further studies are needed to clarify the impact of this mutation on LGR4–mediated pathways and its implications for obesity-targeted therapies.
In conclusion, this study resolves the structures of full-length LGR4-apo and in complex with RSPO2(FU), which provides essential structural guidance for future drug design. Additionally, we successfully developed an LGR4–specific nanobody, named NB21, and determined its complex structure with LGR4. The structure reveals that the NB21 epitope overlaps with the RSPO1/2 binding sites on LGR4, offering a selective mechanism to inhibit this interaction. NB21-mFc represents a promising approach for targeting LGR4 to reduce adiposity by increasing energy expenditure. We propose that this proof-of-concept strategy of drug development, involving the identification of targets with valid human genetic evidence and the development of antibodies with the cryo-EM–assisted validation, will likely emerge as a prevailing trend in the field of anti-obesity drug investigation and development21,22,23.
Methods
Ethics statement
All animal procedures were approved by the Institutional Animal Care and Use Committee (IACUC) of Shanghai Model Organisms Center and conducted in accordance with the Guide for the Care and Use of Laboratory Animals.
Construct design
The DNA sequence of the human LGR4 (residues 25–833) was inserted into a modified pEG BacMam vector (160451, Addgene, USA) with N-terminal influenza haemagglutinin (HA) signal peptide and a C-terminal Flag tag. The DNA sequences of the FU domain of RSPO2 (residues 37–143) or RSPO1 (residues 35–143) were cloned into the pEG BacMam vector, incorporating an N-terminal Gaussia signal peptide, a C-terminal HRV-3C protease site, and concluding with a human IgG Fc domain. The DNA sequence of the entire human LGR4 was cloned into the pcDNA3.1(+) vector for use in cell-based assays. Furthermore, the DNA sequence of the mouse LGR4 (residues 1–951) with a C-terminal 3*Flag tag was cloned into the pcDNA3.1-IRES-GFP vector.
Construction of camelid nanobody phage-display library
The process of camel immunization and nanobody library construction were carried out following established protocols38,73. Two camels were immunized subcutaneously with approximately 1 mg of human LGR4(ECD) protein mixed with an equal volume of FAMA adjuvant (#3030, Gerbu, Germany). The immunizations were administered weekly over seven weeks. Three days after the final immunization, peripheral blood lymphocytes (PBLs) were extracted from whole blood using Ficoll-Paque™ PLUS density gradient media (17144003, Cytiva, USA), adhering to the manufacturer’s instructions. Total RNA was isolated from the PBLs and converted to cDNA using the SuperScript III First-Strand Synthesis System (18080400, Invitrogen, USA). To amplify VHH-encoding sequences, a two-step enriched nested PCR was performed with VHH-specific primers. The amplified VHH sequences were inserted into the pMECS vector (pMECS, NTCC, China) using PstI and BsteII restriction sites. The resulting constructs were transformed into electrocompetent E. coli TG1 cells (60502, Lucigen, USA).
The transformed E. coli TG1 cells were superinfected with M13KO7 helper phages to produce a VHH-displaying phage library. This library underwent two rounds of bio-panning to selectively enrich phages presenting LGR4–specific VHHs. For the bio-panning procedure, LGR4(ECD) was employed as the target antigen and immobilized on streptavidin-coated plates via enzymatic biotinylation using the BirA biotin-protein ligase system. This system ensures site-specific biotinylation at a designated biotin acceptor sequence. The antigen concentration was optimized for binding (typically 10–20 µg mL−1) and performed in solution. Following two rounds of bio-panning, positive clones were identified through ELISA screening using LGR4(ECD)-coated plates to assess the binding specificity of the selected nanobodies. Absorbance readings from the positive clones were five times higher than the control, confirming strong specificity for LGR4(ECD). The VHH sequences of the positive clones were subsequently determined via DNA sequencing.
The selected VHHs were inserted into a pMECS vector, which included a PelB signal peptide, a hemagglutinin (HA) tag, and a His6 tag, and then expressed in E. coli TOP10F’ cells (WXR15-100S, Huayueyang Biotech, China). After cell lysis and protein purification, over twenty nanobodies were obtained. These candidates were initially screened for their binding affinity to LGR4(ECD) using Biolayer Interferometry (BLI). Nanobodies exhibiting strong binding affinity were further assessed for their functional activity in the TOPFlash reporter assay to identify those that inhibited canonical Wnt signaling. Finally, BLI competitive assays were performed to confirm the ability of the selected nanobodies to block LGR4-RSPOs interactions.
Biolayer interferometry (BLI) binding assay
BLI was used to determine the binding affinity (KD) of nanobodies, including NB21, to human or mouse LGR4(ECD). BLI experiments were performed using an Octet RED96e system (ForteBio, Sartorius, Germany) at 25 °C.
Enzymatic biotinylated LGR4(ECD) was immobilized onto Streptavidin (SA) Dip and Read™ Biosensors (18-5019, Sartorius, Germany). The biosensors were pre-equilibrated in kinetics buffer (PBS with 0.02% Tween−20) for 10 min before use. The LGR4(ECD) was prepared at a concentration of 5 µg mL−1 in the same buffer and immobilized for 120 seconds until the response reached approximately 1 nm relative units (RUs).
The baseline was established in kinetic buffer (PBS with 0.02% Tween 20), followed by incubations of the sensors with nanobody at two-fold serial dilutions (100 nM to 3.125 nM) for 300 seconds. Dissociation was measured in the kinetic buffer for 600 seconds. Data were acquired at a rate of 10 Hz using the Octet RED96e instrument. To account for nonspecific binding and signal drift, appropriate negative controls were included in each experiment. SA biosensors immobilized with biotinylated LGR4(ECD) were dipped into the kinetic buffer without the analyte following the baseline step. The response from this control was subtracted from the sample sensorgrams during data analysis to eliminate background signals. In addition, uncoated SA biosensors dipped into nanobody solution were used to confirm minimal nonspecific binding to the sensor surface.
All binding curves were analyzed using ForteBio Data Analysis Software v3.43 (Fortebio, Sartorius, Germany). All binding curves were fitted using a 1:1 Langmuir binding model, with global fitting applied across all concentrations using a shared Rmax value. The equilibrium dissociation constant (KD) was calculated as the ratio of Koff to Kon. No additional normalization or data transformation was applied.
Luciferase reporter assay
The TOPFlash assay was performed according to standard protocols. In brief, HEK293T cells were transiently transfected with plasmids encoding LGR4, along with super 8×TOPFlash firefly luciferase (D2501, Beyotime, China) and pRL-SV40-renilla luciferase reporter plasmids (D2762, Beyotime, China) at a mass ratio of 1:1:0.1. 12 h post-transfection, cells were serum-starved overnight and then treated with RSPO1(FU)-mFc, either alone or pre-incubated with nanobodies, at 37 °C for 2 h. The TOPFlash signal was quantified using the Dual-Glo luciferase assay kit (11402ES80, YEASEN, China), following the manufacturer’s protocol.
BLI competitive assay
BLI was also employed to assess whether NB21 competes with RSPOs for binding to the same epitope. Biotinylated LGR4(ECD) (5 µg mL−1) was immobilized on SA Dip and Read™ Biosensors for 120 seconds until the response reached approximately 1 nm RUs. The baseline was measured in kinetic buffer (PBS buffer with 0.02% Tween 20), and the sensors were then exposed to 500 nM NB21 until the response curve plateaued. Following this, the sensor was incubated with a mixture of NB21 and RSPO1/2(FU)-mFc until the curve stabilized at the plateau phase.
For comparison, another LGR4(ECD)-coated sensor’s baseline was measured in kinetic buffer, and the sensor was exposed to 500 nM RSPO1/2(FU)-mFc until the curve plateaued. Binding curves were analyzed using ForteBio Data Analysis Software v3.43, and the Rmax values were compared to evaluate NB21’s effect on RSPO1/2(FU)-mFc binding. If NB21 competes with RSPO1/2(FU)-mFc, the Rmax for the mixture would be lower than that for the RSPOs alone, suggesting overlapping binding sites and reduced binding when both are present.
Expression and purification of RSPO1(FU)-mFc, RSPO2(FU)-mFc, and RSPO2 (FU) proteins
The plasmids encoding RSPO1(FU)-mFc or RSPO2(FU)-mFc protein were transiently transfected into HEK293 F cells at a density of 2 × 106 cells/mL using Polyethyleneimine (PEI; 24765−2, Polysciences, USA). 24 h post-transfection, sodium butyrate was added to achieve a final concentration of 10 mM. Cells were then incubated at 30 °C for 72 h, after which the supernatant was collected by centrifugation at 5000 × g for 30 min. RSPO1(FU)-mFc or RSPO2(FU)-mFc protein were then purified from the supernatant using protein A affinity chromatography (SA012500, Smart-Lifesciences, China). Elution was carried out with acidic buffer (0.1 M glycine-HCl, pH 3.0), followed by immediate neutralization with 1 M Tris-HCl (pH 8.0) to ensure protein stability. For the purification of RSPO2(FU) domain, the eluted RSPO2(FU)-mFc was digested with HRV-3C protease (P2303, Beyotime, China) and further purified by HiLoad 16/600 Superdex 75 column chromatography (28-9893-33, Cytiva, USA) to remove the mFc domain.
Expression and purification of MB52 and NB21 proteins
NB52 or NB21 (sequences provided in Supplementary Table 1) was cloned into a pMECS vector featuring a PelB signal peptide, an HA tag, and a His6 tag at the C-terminus. The nanobodies were expressed in the periplasm of E. coli strain TOP10F’ cells. After induction, the culture supernatant was collected by centrifugation at 18,000 × g for 30 min to remove cells and debris. The clarified supernatant was loaded onto a HisTrap HP column (29-0510−21, Cytiva, USA) pre-equilibrated with buffer A (20 mM Tris-HCl, pH 8.0, 150 mM NaCl, 10 mM imidazole). The column was washed with 10 column volumes of buffer A containing 20 mM imidazole (V900153, Vetec, Brazil) to remove nonspecifically bound proteins. Nanobodies were eluted with buffer A containing 250 mM imidazole. The eluted proteins were concentrated and further purified by size-exclusion chromatography using a Superdex 75 Increase 10/300 GL column (29-0915-96, Cytiva, USA) equilibrated with gel filtration buffer (20 mM HEPES, pH 7.5, 150 mM NaCl). The purity and monodispersity of the nanobodies were assessed by sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) and analytical size-exclusion chromatography.
For MB52 generation, the VHH gene of NB52 was fused to the circularly permuted extracellular adhesin domain of Helicobacter pylori (HopQ, 58 kDa) to generate a megabody (MB52). MB52 expression was performed in the periplasm of E. coli strain TOP10F’ cells, and purification followed the same protocol used for NB21.
Expression and Purification of LGR4 alone and in complex with RSPO2(FU) or NB21
Human LGR4 was expressed in HEK293 GnTI- cells cultured in FreeStyleTM 293 medium (12338018, Gibco, USA). 24 h after baculovirus infection, the cells were supplemented with 10 mM sodium butyrate to enhance protein expression and incubated at 30 °C for 72 h before harvest. Cell lysis was performed using a homogenizer in a buffer containing 20 mM HEPES (pH 7.4), 10 mM NaCl, and 5% glycerol, supplemented with an EDTA-free protease inhibitor cocktail (P1005, Beyotime, China). To form the complex, 1 μM RSPO2(FU) or NB21, together with1 μM MB52, was added to the lysate. After lysing, the membrane was solubilized in the same buffer with the addition of 1% Lauryl maltose neopentyl glycol (LMNG; NG310, Anatrace, USA) and 0.1% cholesteryl hemisuccinate (CHS; GDN101, Anatrace, USA) at 4 °C for 2 h.
The supernatant was collected by centrifugation at 18,304 × g for 1 h and applied to an anti-DYKDDDDK Affinity chromatography column (SA042100, Smart-Lifesciences, China). The column was washed with the same buffer, gradually decreasing the detergent concentration to 0.02% LMNG and 0.002% CHS, and finally eluted with 20 mM HEPES (pH 7.4), 10 mM NaCl, 5% glycerol, 0.02% LMNG, 0.002% CHS, and 0.2 mg/ml Flag-peptide (53577, GL Biochem, China). The eluted complex protein was concentrated and further purified by size-exclusion chromatography in 20 mM HEPES (pH 7.4), 50 mM NaCl, 0.002% LMNG, and 0.0002% CHS using a Superose™ 6 Increase 10/300 GL column. For the expression and purification of LGR4 alone, the procedure was the same as for the complexes, with the exception that RSPO2(FU)or NB21 was not included.
Cryo-EM data collection
To prepare cryo-EM grids, 3 μL of purified complexes at 2–3 mg mL−1 concentration was applied to freshly glow-discharged 300-mesh holey gold grids (R1.2/1.3 UltrAufoil, Quantifoil, Germany, or ANTcryo™ Au300-1.2/1.3, Nanodim, China) under 100% humidity at 4 °C. After a 5-second wait, the grids were blotted for 3 seconds with a force setting of 2 and rapidly plunged into liquid ethane using a Vitrobot Mark IV (Thermo Fisher Scientific, FEI), which was maintained at cryogenic temperatures using liquid nitrogen.
Data acquisition was performed on a Titan Krios electron microscope equipped with a K3 direct electron detector. Micrographs were recorded at a calibrated pixel size of 0.5355 Å, with a dose rate of 23.3 electrons per pixel per second. Defocus values ranged from -1.5 μm to -2.5 μm, and each exposure lasted 3 seconds, capturing 36 frames per micrograph, with a total dose of 70 electrons per Ų.
The datasets comprised 8,184 movies for LGR4 alone, 5941 movies for the LGR4-RSPO2(FU) complex, and 3885 movies for the LGR4-NB21 complex, all collected under these optimized conditions.
Cryo-EM data processing, model building and refinement
The dataset was imported into cryoSPARC v3.3.2 (Structura Biotechnology, Canada) for processing. Image stacks were aligned using the patch motion correction module, and the contrast transfer function (CTF) parameters for each micrograph were determined by patch CTF estimation. Micrographs with a CTF fit resolution better than 4 Å were selected for further analysis. Particles were auto-picked, extracted, and subjected to two rounds of 2D classification. Well-defined particles were selected for 3D ab initio reconstruction, followed by multiple rounds of 3D heterogeneous refinement to remove poorly resolved classes. The refined particles then underwent additional 3D ab initio reconstruction, homogeneous refinement, and non-uniform refinement to generate a high-quality model. Finally, global and local CTF refinement were applied to generate the final map.
For the LGR4-MB52 complex, 984,615 particles underwent 2D classification, and classes with good features were selected to generate an ab initio model with five classes. One of the classes, with a resolution of 3.84 Å, underwent non-uniform refinement. Further global CTF refinement and local CTF refinement were applied to generate an LGR4-MB52 map with a final resolution of 3.03 Å.
For the MB52-LGR4-RSPO2(FU) complex, approximately 3,663,000 particles were picked. After 2D classification, particles with good features were selected for 3D classification with five classes. The particles of two classes were combined and reserved for non-uniform refinement and CTF refinement, resulting in the MB52-LGR4-RSPO2(FU) complex map with a final resolution of 3.06 Å.
For the MB52-LGR4-NB21 complex, approximately 1,836,475 particles were picked. After 2D classification, particles with good features were selected for 3D classification with five classes. The particles of four classes were combined and reserved for non-uniform refinement and CTF refinement, yielding the map of the MB52-LGR4-NB21 complex with a final resolution of 3.64 Å.
In the cryo-EM reconstruction of the LGR4-MB52 complex, the predicted structure of LGR4 from AlphaFold2 (DeepMind, USA) and the initial template of MB52, generated using SWISS-MODEL (https://swissmodel.expasy.org/docs/references), were fitted into the composite cryo-EM map of the LGR4-MB52 complex as a starting model using UCSF Chimera v1.15 (https://www.cgl.ucsf.edu/chimera/). Model refinement and manual docking of cholesterols into the cryo-EM density were performed using Coot v0.9.4 (http://www2.mrc-lmb.cam.ac.uk/personal/pemsley/coot/). Statistics of data processing and structural refinement are listed in Supplementary Table 2.
Expression and purification of NB21-mFc
NB21 was cloned into the pEG BacMam vector, which contains an N-terminal IL2 signal peptide, a C-terminal GGGGS linker, and a mouse IgG2 (mFc) domain. The plasmid was transiently transfected into HEK293 F cells using PEI at a cell density of 2 × 106 cells mL−1. After 24 h, sodium butyrate was added to a final concentration of 10 mM, and the cells were incubated at 30 °C for an additional 72 h. The cell supernatant was then collected by centrifugation at 5000 × g for 30 min. NB21-mFc was purified from the cell supernatant using protein A affinity chromatography, and further purified using a HiLoad 16/600 Superdex 75 column.
Single-cell sequencing and analysis
Single-cell RNA-seq matrix of mouse white adipose tissues were obtained from the Gene Expression Omnibus (GEO) database (GSE176171)52. The data underwent processing utilizing the Seurat v5.0.1 package within the R v4.3.2 programming environment74. The expression matrix was imported, resulting in the creation of a Seurat object. Subsequently, cells were subjected to quality control metrics, normalization, and scaling. Principal component analysis (PCA) and t-distributed stochastic neighbor embedding (t-SNE) were employed for dimensionality reduction and visualization. Cluster identities were determined by matching enriched genes to a previous study52. Gene distribution plots were generated using the FeaturePlot commands within the Seurat package.
Stromal vascular fractions (SVFs) isolation and beige adipocyte differentiation
Mouse SVFs were isolated from the sWAT of 6–8-week-old mice. Briefly, the adipose tissues were finely chopped and subjected to enzymatic digestion using collagenase type II (C9407, Sigma‑Aldrich, USA) at a concentration of 2 mg mL−1 in PBS (MA0015, Meilunbio, China) solution containing 10 mM HEPES (15630-080, Gibco, USA) and 1 % bovine serum albumin (0332, GBCBIO, China). This process was carried out for a duration of 30 min at a temperature of 37 °C. Next, the reaction was halted by adding complete medium (MA0212, Meilunbio, China). The suspension was subjected to centrifugation, washing, and filtration using a 40-μm cell strainer (352340, Falcon,USA). Subsequently, the filtered material was plated onto 6-cm dishes. The SVFs were cultured in DMEM/F12 (21331-020, Gibco, USA) supplemented with 10 % fetal bovine serum (FBS; 10099141C, Gibco, USA), 1 mM L-glutamine (25030-081, Gibco, USA), 1 % penicillin/streptomycin (15140-122, Gibco, USA), and 10 ng mL−1 recombinant mouse fibroblast growth factor basic protein (FGF; 3139-FB, R&D, USA). Then, SVFs were cultured on 48-well plates and permitted to attain confluence. Beige adipocyte differentiation was induced using a cocktail medium containing 5 μg mL−1 insulin (Novo nordisk, USA), 0.5 mM 3-Isobutyl-1-methylxanthine (I7018, Sigma‑Aldrich, USA), 1 μM dexamethasone (MA1536, Meilunbio, China), 1 nM 3,3′,5-Triiodo-L-thyronine (T3; T2877, Sigma‑Aldrich, USA), and 1 μM rosiglitazone (R2408, Sigma‑Aldrich, USA) for day 0–2 followed by the addition of growth medium supplemented with insulin, T3 and rosiglitazone for day 3–6, depending on experimental requests. This protocol was adapted from our previous study with appropriate modifications as described above21.
In order to assess the impact of NB21 on the process of browning, NB21 was administered on two occasions (on days −2 and 0, respectively) both prior to and during the induction procedure. In the antagonizing experiments, recombinant human RSPO1 (4645-RS, R&D, USA) or RSPO2 (3266-RS, R&D, USA) proteins were co-administered with NB21-mFc.
Lentivirus production and infection
Lentiviral constructs targeting mouse Lgr5 (shLgr5: CCCATGACTGAAAGCTGTCAT, NCBI RefSeq: NM_010195.2) and Lgr6 (shLgr6: AGTCTGATGGAACCAAGTTTG, NCBI RefSeq: NM_001033409.4) were generated using the pSLenti-U6-shRNA-CMV-EGFP-F2A-Puro-WPRE lentiviral vector. SVFs were infected with lentivirus when cell confluency reached 30%–50%, at a multiplicity of infection (MOI) of 50–100, following the manufacturer’s protocol. To enhance infection efficiency, 5 μg mL−1 Polybrene (107689-10 G, Sigma‑Aldrich, USA) was added. The culture medium was replaced with standard growth or induction medium within 8–12 h after infection.
Immunofluorescence staining of cells
SVFs were cultured in 8-well glass chamber slides (PEZGS0816, Millipore, USA) and treated with hRSPO1 (0.1 μg mL−1) and(or) NB21-mFc (0.1 μM) for 48 h, followed by fixation with 4% paraformaldehyde (P1110, Solarbio, China). SVFs are incubated with a blocking solution (S3022, DAKO, USA) to reduce the non-specific binding of antibodies, followed by incubation overnight at 4 °C with active β-catenin (non-phosphorylated Ser33/37/Thr41) primary antibody (1:500; 8814s, Cell Signaling Technology, USA). On the next day (48 h later), the SVFs were incubated with an Alexa Fluor 555-conjugated goat anti-rabbit IgG secondary antibody (1:500; A-21428, ThermoFisher, USA) for 120 min at room temperature. The slides were then sealed using 4’,6-diamidino−2-phenylindole (DAPI) Fluoromount-G mounting media (0100−20, SouthernBiotech, USA) and imaged using a confocal microscope (LSM880, Zeiss, USA).
RNA isolation and qPCR analysis
RNAs were isolated from cells or adipose tissues by using a total RNA isolation kit (LS1040, Promega, China) according to the manufacturer’s protocols. Next, isolated RNAs were reverse transcribed into cDNA templates using a HiScript_III 1st Strand cDNA Synthesis (R312-02, Vazyme, China), following the instructions in the manuals. cDNA was measured using quantitative real-time PCR (qPCR) using ChamQ Universal SYBR qPCR Master Mix (Q711-02, Vazyme, China). qPCR was conducted on an ABI system (Life Technology, USA). The primers used in this study are provided in Supplementary Table 3. The data were subjected to analysis using the comparative threshold cycle (ΔΔCt) method, with 36B4 serving as the reference gene.
Protein preparation and western blotting
Total protein extraction from SVFs and adipose tissues was conducted using cold radioimmunoprecipitation assay (RIPA) lysis buffer (R20095, Biocolors, China) supplemented with a Protease Inhibitor Cocktail (78444, Thermo Scientific, USA). The concentration of extracted protein was determined by employing the BCA Protein Assay Kit (23227, Thermo Scientific, USA). Protein simples underwent separation via sodium dodecyl SDS-PAGE and subsequent transfer onto a 0.45 μm polyvinylidene difluoride (PVDF) membrane (IPVH00010, Millipore, USA) at 200 mA for 120 min while maintaining cold temperatures. PVDF membranes were blocked with 5% BSA solution (332, GBCBIO, China) for 90 min at room temperature, subsequently undergoing an overnight incubation with the primary antibody at 4 °C. Horseradish peroxidase (HRP)-conjugated secondary antibody incubation was performed at room temperature for 90 min. Results were visualized using an eBlot luminescent image analyzer (GenScript, China) with enhanced chemiluminescence (ECL) luminous fluid (NEL104001EA, Perkin Elmer, USA). HSP90 (1:1000; 4877 s, Cell Signaling Technology, USA) was used as the internal control. The following primary antibodies were used: anti-active β-catenin (non-phosphorylated Ser45) (1:1000; 19807, Cell Signaling Technology, USA), anti-active β-catenin (non-phosphorylated Ser33/37/Thr41) (1:1000; 8814s, Cell Signaling Technology, USA), anti-total β-catenin (1:1000; 9562 s, Cell Signaling Technology, USA), anti-UCP1 antibody (1:1000; ab10983, Abcam, UK), and anti-UQCRC2 & MTCO1 & NDUFB8 (1:1000; ab110413, Abcam, UK). The following secondary antibodies were used: HRP-linked anti-rabbit IgG (1:1,000; 7074 s, Cell Signaling Technology, USA) and HRP-linked anti-mouse IgG (1:1,000; 7076 s, Cell Signaling Technology, USA).
Oxygen consumption rate (OCR) measurements
SVFs were plated in Seahorse XF96 V3 PS Cell Culture Microplates (103792-100, Agilent, USA) coated with poly-L-lysine (P4707, Sigma‑Aldrich, USA). With the exogenous hRSPO1 or NB21-mFc treatment from day −2–2, the cells were induced into beige adipocytes for 3 days, followed by OCR measurement at 37 °C was assessed using the Seahorse Bioscience XFe96 Flux analyzer (Agilent, USA) with Seahorse XF Cell Mito Stress Test Kit (103015-100, Agilent, USA) according to the manufacturer’s instructions. During the assay, 1 μM oligomycin (to detect coupled respiration), 2 μM Carbonyl cyanide-4-phenylhydrazone (FCCP; to assess uncoupled respiration), and 0.5 μM rotenone/antimycin (to measure non-mitochondrial respiration) were injected into the well sequentially.
Animals
Both male and female mice were used in this study. Sex was considered in the experimental design and data interpretation where applicable. Details regarding the sex of animals and sample sizes are provided in the figure legends, and sex-disaggregated raw data are available in the Source Data file. For metabolic cage and acute cold exposure experiments, female 8-week-old C57BL/6 J mice were provided with a high-fat diet (HFD; 60 kcal% fat; D12492i, Research Diets, USA) and subjected to intraperitoneal injections of either PBS or NB21-mFc (0.2 mg kg−1) once every day. For chronic cold stimulation, female 8-week-old C57BL/6 J mice under HFD were subjected to intraperitoneal injections of either PBS or NB21-mFc (0.2 mg kg−1) daily for 7 days in cages at 5 °C. Lgr4m/m mice were maintained through heterozygous breeding21. Both male and female WT and Lgr4m/m mice, aged 8 weeks, were subjected to an HFD and received intraperitoneal injections of either PBS or NB21-mFc (0.2 mg kg−1) every other day for a duration of 4–6 weeks. Lep-deficient (ob/ob) mice (Strain NO. T001461) were acquired from GemPharmatech (Nanjing, China). Male ob/ob mice, aged 8 weeks, were provided with a normal chow diet (NCD; 10 kcal% fat; P1101F-25, SLACOM, China) and subjected to intraperitoneal injections of either PBS or NB21-mFc (0.03 mg kg−1 or 0.1 mg kg−1) every other day for a duration of 7 weeks.
The mice were accommodated in conventional cages located in a specific-pathogen-free (SPF) facility, adhering to a 12-hour light-dark cycle. They were provided with ad libitum access to food and water at a controlled temperature of 22 ± 2 °C, unless stated otherwise. Mice of identical sex and litter were cohabitated in groups of up to five individuals per cage. All procedures were conducted in accordance with protocols approved by the IACUC of Shanghai Model Organisms Center.
Body composition analysis and indirect calorimetry
Body composition, including fat and lean mass, was measured using an Echo MRI-100H composition analyzer (Echo Medical Systems, USA). For energy expenditure (EE) measurement, mice were housed individually in a Comprehensive Laboratory Animal Monitoring System (Promethion GAFR, Sable Systems International, USA) at either 22 °C or 5 °C to evaluate their food intake, O2 consumption, CO2 production and physical activity. The EE and respiratory exchange ratio (RER) were calculated based on the O2 and CO2 data. EE, O2 consumption, and CO2 production were evaluated through analysis of covariance (ANCOVA) with body weight as a covariate48,49,50. After 24 h of adaptation to room temperature (22 °C), basal EE data were collected, after which the cage temperature was decreased to 5 °C for acute cold challenge over the subsequent 4 h period.
Fecal energy content measurement
Mice were singly housed, and fecal samples were collected from the cage floor over 2 consecutive days. The collected feces were dried at 56 °C for 24 h to remove moisture, then subjected to calorimetric analysis using a 6400 Automatic Isoperibol Calorimeter (Parr Instrument Company, USA) to determine energy content.
Rectal temperature measurement
Before the test, the mice were starved 6 h and then moved to individual cages at 5 °C without food. The rectal temperature was measured using a thermal probe pretreated with Vaseline and a model BAT-12 thermometer (Physitemp Instruments, USA).
Hematoxylin and eosin (H&E) staining
The adipose and liver tissues were isolated and subsequently fixed in a 4% paraformaldehyde solution. Following fixation, the tissues were embedded in paraffin and sliced into sections measuring 5 μm in thickness for H&E staining. Tissue sections were scanned by the Tissue FAXS system (TissueGnostics, Austria). The adipocyte area in sWAT and vWAT was quantified using ImageJ (National Institutes of Health, USA).
Immunofluorescence staining of tissue
The paraffin sections of mouse adipose samples were consecutively subjected to immunostaining with anti-UCP1 antibody (1:500; ab10983, Abcam, UK) and anti-Perilipin antibody (1:300; 9349 s, Cell Signaling Technology, USA), employing the Tyramide Signal Amplification (TSA) technique. CF-488-Tyramide and CF-594-Tyramide were used as tyramine conversion reagents. Antigen retrieval was performed using a 1 mM pH 9.0 Tris-EDTA solution (648310; E9884, Sigma‑Aldrich, USA). The samples were sealed using an antifade mounting medium containing 4,6-diamidino−2-phenylindole (P0131, Beyotime, China). A TissueFAXS Plus automated acquisition system (TissueGnostics, Austria) was utilized for the whole-slide scan.
Statistical analysis
Appropriate statistical tests were applied to each figure. Data from bar and line plots are presented as mean ± SEM. Kernel density plots display data distributions with the central line indicating the median, while violin plots overlaid with box plots show the median (center line), the 25th and 75th percentiles (box bounds), and whiskers extending to 1.5× the interquartile range (IQR). The effective concentration 50% (EC50) and binding affinity values were calculated using non-linear regression. In the case of two-group comparison, the unpaired two-tailed Student’s t test was employed to ascertain statistical significance. When comparing more than two groups, the one-way analysis of variance (ANOVA) was initially conducted, followed by the application of Dunnett’s or Tukey’s multiple-comparison test. For non-normally distributed data, the two-sided Wilcoxon rank-sum test was applied for comparisons between two groups, while the Kruskal-Wallis test followed by the Bonferroni-Dunn post hoc test was employed for comparisons among three or more groups. For the measurements of indirect calorimetry, ANCOVA with body weight as a covariate and two-way ANOVA were performed. Rectal temperatures were analyzed by two-way ANOVA. For reproducibility, in vivo experiments (mouse models) were repeated at least twice, and in vitro (SVF/cell lines) and molecular experiments were performed in at least three independent replicates. Statistical tests were performed in Microsoft Excel v16.77.1 (Microsoft, USA) and R v4.3.2 (R Foundation for Statistical Computing, Austria).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Cryo-EM density maps obtained in this study have been deposited in the Electron Microscopy Data Bank (EMDB) under the accession codes (LGR4-MB52) EMDB-38641 [https://www.ebi.ac.uk/emdb/EMD-38641], (MB52-LGR4-RSPO2(FU)) EMDB-38677 [https://www.ebi.ac.uk/emdb/EMD-38677], (MB52-LGR4-NB21) EMDB-54527 [https://www.ebi.ac.uk/emdb/EMD-54527]. Atomic coordinates obtained in this study have been deposited in the Protein Data Bank (PDB) under IDs (LGR4-MB52) 8XT9, (MB52-LGR4-RSPO2(FU)) 8XUM, and (MB52-LGR4-NB21) 9S37. All data generated in this study are included in the main text or Supplementary Information. Source data are provided with this paper.
References
Bjerregaard, L. G. & Baker, J. L. Change in overweight from childhood to early adulthood and risk of type 2 diabetes. N. Engl. J. Med. 378, 2537–2538 (2018).
Calle, E. E. Obesity and cancer. Bmj 335, 1107–1108 (2007).
Calle, E. E., Rodriguez, C., Walker-Thurmond, K. & Thun, M. J. Overweight, obesity, and mortality from cancer in a prospectively studied cohort of U.S. adults. N. Engl. J. Med. 348, 1625–1638 (2003).
Gregg, E. W. & Shaw, J. E. Global health effects of overweight and obesity. N. Engl. J. Med. 377, 80–81 (2017).
Ng, M. et al. Global, regional, and national prevalence of overweight and obesity in children and adults during 1980−2013: a systematic analysis for the Global Burden of Disease Study 2013. Lancet 384, 766–781 (2014).
Twig, G. et al. Body-mass index in 2.3 million adolescents and cardiovascular death in adulthood. N. Engl. J. Med. 374, 2430–2440 (2016).
Mohammed, M. S., Sendra, S., Lloret, J. & Bosch, I. Systems and WBANs for controlling obesity. J. Health. Eng. 2018, 1564748 (2018).
Narayanaswami, V. & Dwoskin, L. P. Obesity: current and potential pharmacotherapeutics and targets. Pharm. Ther. 170, 116–147 (2017).
Polonsky, K. S. & Klein, S. Gastric banding to treat obesity: band-aid or breakthrough?. Nat. Clin. Pr. Endocrinol. Metab. 4, 421 (2008).
Wen, X. et al. Signaling pathways in obesity: mechanisms and therapeutic interventions. Signal Transduct. Target Ther. 7, 298 (2022).
Cruickshank, C. C. & Dyer, K. R. A review of the clinical pharmacology of methamphetamine. Addiction 104, 1085–1099 (2009).
Dietrich, M. O. & Horvath, T. L. Limitations in anti-obesity drug development: the critical role of hunger-promoting neurons. Nat. Rev. Drug Discov. 11, 675–691 (2012).
Kelley, D. E. et al. Clinical efficacy of orlistat therapy in overweight and obese patients with insulin-treated type 2 diabetes: A 1-year randomized controlled trial. Diabetes Care 25, 1033–1041 (2002).
Trajanoska, K. et al. From target discovery to clinical drug development with human genetics. Nature 620, 737–745 (2023).
Akbari, P. et al. Sequencing of 640,000 exomes identifies GPR75 variants associated with protection from obesity. Science 373, eabf8683 (2021).
Murtaza, B., Asghar, F. & Patoli, D. GPR75: An exciting new target in metabolic syndrome and related disorders. Biochimie 195, 19–26 (2022).
Congreve, M., de Graaf, C., Swain, N. A. & Tate, C. G. Impact of GPCR structures on drug discovery. Cell 181, 81–91 (2020).
Yang, D. et al. G protein-coupled receptors: structure- and function-based drug discovery. Signal Transduct. Target Ther. 6, 7 (2021).
Shimada, I., Ueda, T., Kofuku, Y., Eddy, M. T. & Wüthrich, K. GPCR drug discovery: integrating solution NMR data with crystal and cryo-EM structures. Nat. Rev. Drug Discov. 18, 59–82 (2019).
Styrkarsdottir, U. et al. Nonsense mutation in the LGR4 gene is associated with several human diseases and other traits. Nature 497, 517–520 (2013).
Wang, J. et al. Ablation of LGR4 promotes energy expenditure by driving white-to-brown fat switch. Nat. Cell Biol. 15, 1455–1463 (2013).
Zou, Y. et al. Association of a gain-of-function variant in LGR4 with central obesity. Obesity (Silver Spring) 25, 252–260 (2017).
Sun, P. et al. Loss of Lgr4 inhibits differentiation, migration and apoptosis, and promotes proliferation in bone mesenchymal stem cells. J. Cell Physiol. 234, 10855–10867 (2019).
Weng, J. et al. Deletion of G protein-coupled receptor 48 leads to ocular anterior segment dysgenesis (ASD) through down-regulation of Pitx2. Proc. Natl Acad. Sci. USA 105, 6081–6086 (2008).
Zhang, N., Yuan, M. & Wang, J. LGR4: a new receptor member in endocrine and metabolic diseases. Endocr. Rev. 44, 647–667 (2023).
Wang, D. et al. Structural basis for R-spondin recognition by LGR4/5/6 receptors. Genes Dev. 27, 1339–1344 (2013).
de Lau, W. et al. Lgr5 homologues associate with Wnt receptors and mediate R-spondin signalling. Nature 476, 293–297 (2011).
Xu, K., Xu, Y., Rajashankar, K. R., Robev, D. & Nikolov, D. B. Crystal structures of Lgr4 and its complex with R-spondin1. Structure 21, 1683–1689 (2013).
Carmon, K. S., Gong, X., Lin, Q., Thomas, A. & Liu, Q. R-spondins function as ligands of the orphan receptors LGR4 and LGR5 to regulate Wnt/beta-catenin signaling. Proc. Natl Acad. Sci. USA 108, 11452–11457 (2011).
Chen, Y. et al. Nidogen-2 is a novel endogenous ligand of LGR4 to inhibit vascular calcification. Circ. Res 131, 1037–1054 (2022).
Deng, C. et al. Multi-functional norrin is a ligand for the LGR4 receptor. J. Cell Sci. 126, 2060–2068 (2013).
Glinka, A. et al. LGR4 and LGR5 are R-spondin receptors mediating Wnt/β-catenin and Wnt/PCP signalling. EMBO Rep. 12, 1055–1061 (2011).
Luo, J. et al. LGR4 is a receptor for RANKL and negatively regulates osteoclast differentiation and bone resorption. Nat. Med. 22, 539–546 (2016).
Ruffner, H. et al. R-Spondin potentiates Wnt/β-catenin signaling through orphan receptors LGR4 and LGR5. PLoS ONE 7, e40976 (2012).
Sun, Y. et al. Human RSPO1 mutation represses beige adipocyte thermogenesis and contributes to diet-induced adiposity. Adv. Sci. (Weinh.) 10, e2207152 (2023).
de Lau, W. B., Snel, B. & Clevers, H. C. The R-spondin protein family. Genome Biol. 13, 242 (2012).
Xu, J. G. et al. Crystal structure of LGR4-Rspo1 complex: insights into the divergent mechanisms of ligand recognition by leucine-rich repeat G-protein-coupled receptors (LGRs). J. Biol. Chem. 290, 2455–2465 (2015).
Wang, L. et al. Structural insights into the LGR4-RSPO2-ZNRF3 complexes regulating WNT/β-catenin signaling. Nat. Commun. 16, 362 (2025).
Duan, J. et al. Structures of full-length glycoprotein hormone receptor signalling complexes. Nature 598, 688–692 (2021).
Duan, J. et al. Hormone- and antibody-mediated activation of the thyrotropin receptor. Nature 609, 854–859 (2022).
Ballesteros, J. A. et al. Activation of the beta 2-adrenergic receptor involves disruption of an ionic lock between the cytoplasmic ends of transmembrane segments 3 and 6. J. Biol. Chem. 276, 29171–29177 (2001).
Cherezov, V. et al. High-resolution crystal structure of an engineered human beta2-adrenergic G protein-coupled receptor. Science 318, 1258–1265 (2007).
Li, J., Edwards, P. C., Burghammer, M., Villa, C. & Schertler, G. F. Structure of bovine rhodopsin in a trigonal crystal form. J. Mol. Biol. 343, 1409–1438 (2004).
Rasmussen, S. G. et al. Crystal structure of the human beta2 adrenergic G-protein-coupled receptor. Nature 450, 383–387 (2007).
Duan, J. et al. Mechanism of hormone and allosteric agonist mediated activation of follicle stimulating hormone receptor. Nat. Commun. 14, 519 (2023).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Chen, M. et al. CTNNB1/β-catenin dysfunction contributes to adiposity by regulating the cross-talk of mature adipocytes and preadipocytes. Sci. Adv. 6, eaax9605 (2020).
Mina, A. I. et al. CalR: a web-based analysis tool for indirect calorimetry experiments. Cell Metab. 28, 656–666.e651 (2018).
Tschöp, M. H. et al. A guide to analysis of mouse energy metabolism. Nat. Methods 9, 57–63 (2011).
Fernández-Verdejo, R., Ravussin, E., Speakman, J. R. & Galgani, J. E. Progress and challenges in analyzing rodent energy expenditure. Nat. Methods 16, 797–799 (2019).
Toh, Y. et al. LGR4 and LGR5 form distinct homodimers that only LGR4 complexes with RNF43/ZNRF3 to provide high affinity binding of R-spondin ligands. Sci. Rep. 13, 10796 (2023).
Emont, M. P. et al. A single-cell atlas of human and mouse white adipose tissue. Nature 603, 926–933 (2022).
Cristancho, A. G. & Lazar, M. A. Forming functional fat: a growing understanding of adipocyte differentiation. Nat. Rev. Mol. Cell Biol. 12, 722–734 (2011).
Gimble, J. M., Bunnell, B. A., Chiu, E. S. & Guilak, F. Concise review: Adipose-derived stromal vascular fraction cells and stem cells: let’s not get lost in translation. Stem Cells 29, 749–754 (2011).
Madonna, R., Geng, Y. J. & De Caterina, R. Adipose tissue-derived stem cells: characterization and potential for cardiovascular repair. Arterioscler Thromb. Vasc. Biol. 29, 1723–1729 (2009).
Noble, M. E., Endicott, J. A. & Johnson, L. N. Protein kinase inhibitors: insights into drug design from structure. Science 303, 1800–1805 (2004).
Wacker, D., Stevens, R. C. & Roth, B. L. How ligands illuminate GPCR molecular pharmacology. Cell 170, 414–427 (2017).
Hutchings, C. J., Koglin, M., Olson, W. C. & Marshall, F. H. Opportunities for therapeutic antibodies directed at G-protein-coupled receptors. Nat. Rev. Drug Discov. 16, 787–810 (2017).
Hutchings, C. J., Koglin, M. & Marshall, F. H. Therapeutic antibodies directed at G protein-coupled receptors. MAbs 2, 594–606 (2010).
Hauser, A. S., Attwood, M. M., Rask-Andersen, M., Schiöth, H. B. & Gloriam, D. E. Trends in GPCR drug discovery: new agents, targets and indications. Nat. Rev. Drug Discov. 16, 829–842 (2017).
Gera, S. et al. First-in-class humanized FSH blocking antibody targets bone and fat. Proc. Natl Acad. Sci. USA 117, 28971–28979 (2020).
Liu, P. et al. Blocking FSH induces thermogenic adipose tissue and reduces body fat. Nature 546, 107–112 (2017).
Chen, N. et al. Ctnnb1/β-catenin inactivation in UCP1-positive adipocytes augments the browning of white adipose tissue. iScience 26, 106552 (2023).
Kreymann, B., Williams, G., Ghatei, M. A. & Bloom, S. R. Glucagon-like peptide-1 7-36: a physiological incretin in man. Lancet 2, 1300–1304 (1987).
Madsbad, S. Exenatide and liraglutide: different approaches to develop GLP-1 receptor agonists (incretin mimetics)-preclinical and clinical results. Best. Pr. Res Clin. Endocrinol. Metab. 23, 463–477 (2009).
Martens, M. D. et al. Semaglutide reduces cardiomyocyte size and cardiac mass in lean and obese mice. JACC: Basic Transl. Sci. 9, 1429–1431 (2024).
Prado, C. M., Phillips, S. M., Gonzalez, M. C. & Heymsfield, S. B. Muscle matters: the effects of medically induced weight loss on skeletal muscle. Lancet Diabetes Endocrinol. 12, 785–787 (2024).
Annunziato, S., Sun, T. & Tchorz, J. S. The RSPO-LGR4/5-ZNRF3/RNF43 module in liver homeostasis, regeneration, and disease. Hepatology 76, 888–899 (2022).
de Lau, W., Peng, W. C., Gros, P. & Clevers, H. The R-spondin/Lgr5/Rnf43 module: regulator of Wnt signal strength. Genes Dev. 28, 305–316 (2014).
Dong, H. et al. Identification of a regulatory pathway inhibiting adipogenesis via RSPO2. Nat. Metab. 4, 90–105 (2022).
Chang, T.-H., Harlos, K., Hsieh, F.-L. & Yvonne Jones, E. Structural basis of Norrin recognition by Lgr4/5/6. bioRxiv https://doi.org/10.1101/2022.09.29.510199 (2022).
Wang, Y. et al. Norrin/Frizzled4 signaling in retinal vascular development and blood brain barrier plasticity. Cell 151, 1332–1344 (2012).
Pardon, E. et al. A general protocol for the generation of Nanobodies for structural biology. Nat. Protoc. 9, 674–693 (2014).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2024).
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (U24A20675 and 92457302 to J.W., 81930021 to G.N., 82370845 to J.H., 82130099 to J.L., 8247031318 to W.G., and 82273983 to H.J.), the National Key Research and Development Program of China (2022YFC2505201 to J.W.), innovative research teams of high-level local universities in Shanghai (J.W. and G.N.), the Natural Science Major Program of Shanghai Municipal Education Commission (2023ZKZD22 to J.W.), Science and Technology Commission of Shanghai Municipality (18JC1415400 to Y.G., 21JC1404400 to G.N.), Shanghai Outstanding Academic Leaders Plan (LJ2023078 to J.W.), and CAS Interdisciplinary Team of “Light of West China” Program (xbzg-zdsys-202405 to H.J.). We would like to express our sincere gratitude to Prof. Chi-Chung Hui for his valuable insights in formulating the manuscript, and to Mrs. Dongqin Gu for providing technical assistance. The cryo-EM data were acquired at the Cryo-Electron Microscopy Research Center, Shanghai Institute of Materia Medica, CAS. The process of camel immunization, generation of nanobody libraries, as well as the subsequent screening and identification of nanobodies, were conducted at Shanghai Kailuo Biotechnology Co., Ltd.
Author information
Authors and Affiliations
Contributions
J.W. and Y.G. conceptualized and designed the study. J.W., Y.G., J.L., and G.N. supervised the study. L.W. contributed to cryo-EM data acquisition and processing. L.W. and Y.G. performed the analysis of cryo-EM data. Z.Z. and H.J. carried out all cell and animal functional experiments and analyzed the experimental data. L.W., H.Q., S.G., and T.G. performed protein purification. Y.L. designed expression conducts and identified the nanobodies. Q.C. and Z.L. carried out the screening of nanobodies. N.Z. assisted in checking experimental data. Z.Z., J.W., and Y.G. wrote the manuscript. Z.Z. and L.W. and carried out the figure preparation. Z.Z., L.W., H.J., H.Q., W.G., J.H., G.N., and R.L. contributed to text revision and discussion. All authors read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
J.W. and G.N. are listed as inventors on a patent application entitled “LGR4-targeting nanobody NB21 and use thereof in anti-obesity treatment” (PCT/CN2025/076673; CN120058937), filed by Ruijin Hospital, Shanghai Jiao Tong University School of Medicine. The patent covers the NB21 antibody and its therapeutic use in obesity. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Alexander Banks, Eric Geertsma, Arun Shukla, Fikadu Tafesse and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, Z., Wang, L., Qiao, H. et al. Cryo-EM structure of the full-length LGR4-RSPOs complex and a targeting nanobody for anti-obesity therapy. Nat Commun 16, 8406 (2025). https://doi.org/10.1038/s41467-025-63410-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-63410-5