Abstract
Sorangium cellulosum is a cellulolytic myxobacterium that produces a vast array of complex natural products with diverse chemical scaffolds and biological activities. However, biosynthetic investigations of these metabolites have been hindered by the scarcity of genetic manipulation tools available for their producing microorganisms. Here, we develop an efficient electroporation method for transforming foreign DNA into various Sorangium strains, enabling effective genetic engineering via homologous recombination. This facilitates delineation of the biosynthetic pathway to ambruticin, unveiling several previously undisclosed steps. Notably, AmbK is identified as the elusive epoxide hydrolase responsible for the formation of the tetrahydropyran ring during post-polyketide synthase (PKS) modification, while the terminal PKS module AmbH is shown to catalyse dual rounds of chain elongation during polyketide assembly. Our findings provide significant insights into the intricate molecular machinery governing myxobacterial natural product biosynthesis and greatly enhance our ability to further engineer Sorangium strains to unlock their biosynthetic potentials.
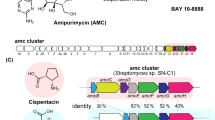
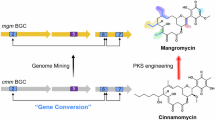
Similar content being viewed by others
Introduction
Natural products derived from microbial sources have long been a cornerstone of pharmaceutical drug discovery due to their structural diversity and therapeutic potential1,2,3. Among the myriad of microbial natural product producers, such as fungi and actinomycetes, the Gram-negative myxobacteria have gained increasing recognition for their rich secondary metabolism in recent decades4,5,6. Sorangium cellulosum, a cellulolytic member of the myxobacteria, accounts for nearly half of all known metabolites produced by this bacterial group7. These include the anticancer epothilones8, the antifungal soraphens9, the antibacterial sorangicins10 and the antiviral ratjadones11 (Fig. 1a). Strains of Sorangium are distinguished by their unusually large genomes, reaching up to 15 Mbp, which encode a wide array of secondary metabolite biosynthetic gene clusters. This genomic richness underscores their substantial potential for natural product discovery, offering a diverse chemical repertoire for biotechnological and pharmaceutical applications12,13. Prior studies have suggested that Sorangium harbours biosynthetic machinery unique to myxobacteria, featuring rare and unusual enzymology not typically observed in other microbial organisms14.
a Examples of natural products from S. cellulosum with diverse chemical scaffolds and biological activities. b Colonies of S. cellulosum So ce10 showing a distinctive red colour on agar plate and the structures of ambruticins. c Genetic organisation of the amb gene cluster. d Previously proposed biosynthetic pathway of ambruticin showing key steps involving tetrahydropyran ring formation, ACP = acyl carrier protein, see Supplementary Fig. 1 for the full pathway.
Genetic manipulation of Sorangium strains is crucial for exploiting their biosynthetic potential and gaining an in-depth understanding of these intricate pathways. However, such efforts have been extremely challenging due to the lack of robust genetic modification tools15,16. These organisms are slow-growing and possess a complex social lifecycle, including fruiting body formation, which complicates their cultivation and genetic manipulation. The extensive GC-rich genome of S. cellulosum, containing many repetitive sequences and intrinsic antibiotic resistance genes, poses further difficulties for genomic editing12,17. Previously, plasmids that could insert into the middle of the target gene were mainly introduced into Sorangium cells through conjugation with a helper E. coli strain, and gene disruptions were achieved via a single-crossover recombination event18,19. The conjugation approach is time-consuming and has been found to be strain-specific; protocols developed for one particular strain often cannot be applied to another20. Electroporation has been reported scarcely and has not seen widespread application21.
The ambruticins were first reported in the late 1970s and are among the earliest characterised metabolites produced by Sorangium strains22,23. They have since attracted sustained interest due to their unusual chemical architectures and potent antifungal activities against a variety of fungal pathogens, including Coccidioides immitis, Histoplasma capsulatum, and Blastomyces dermatitidis24,25,26,27,28,29,30. The ambruticins share several key structural features including a dihydropyran (DHP) and tetrahydropyran (THP) with an unsaturated hydrocarbon linker and an unusual trisubstituted cyclopropyl ring, but differ mainly in the functionality at C-5 (Fig. 1b). In 2006, Reeves and co-workers reported analysis of the ambruticin biosynthetic gene cluster (BGC) in S. cellulosum So ce10 and proposed several unusual features of the modular biosynthetic pathway and post-PKS tailoring based on extensive bioinformatic analysis, selected gene disruption experiments and isotopic labelling studies (Fig. 1c)31. Interestingly, one of the PKS modules in AmbE was hypothesised to catalyse three iterative chain elongations and downstream processes, including a Favorskii rearrangement and a carbon atom excision event would afford the cyclopropyl intermediate for further modifications (Supplementary Fig. 1)31. However, these steps remain speculative and await further experimental validation.
Since 2014, Hahn and colleagues have undertaken a series of biosynthetic investigations on the ambruticins and the closely related jerangolids32,33,34,35,36,37,38. Through in vitro assays using synthetic substrate surrogates, their studies on key enzymes have elucidated several steps involved in ambruticin biosynthesis. The DHP is formed via an intramolecular oxa-Michael addition catalysed by AmbDH3 of linear polyketide (Fig. 1d, structure i) to give THP (Fig. 1d, structure ii)32. AmbDH3 has broad substrate specificity and has been used in the total synthesis of the diarylheptanoid centrolobine39. More recent studies indicate that oxidation of the THP to the DHP occurs prior to generation of the C3 to C7 dihydroxylated tetrahydropyran in ambruticin biosynthesis, but the precise timing remains unknown40.
Using a combination of homology modelling, phylogenetic analysis, and in vitro experiments, Hahn and co-workers reannotated AmbG as a fatty acyl:adenylate ligase (FAAL)–acyl carrier protein (ACP) didomain enzyme with an unusually broad substrate tolerance, providing an important advancement in the understanding of the biosynthesis of the ambruticin middle fragment37. Nevertheless, much of the fundamental biosynthetic machinery of ambruticin, including formation of the central cyclopropane ring and generation of the C3-C7 THP, remains elusive. This underscores the need for further exploration utilising advanced genetic engineering techniques to elucidate the remaining complexities involved in its biosynthesis.
Using the ambruticin-producing strain S. cellulosum So ce10 as a model system, we here present an efficient approach to genetically manipulate Sorangium strains and delineate the intricate molecular machinery of ambruticin biosynthesis. Through targeted genetic engineering of the ambruticin BGC and characterisation of key metabolites from generated mutant strains, combined with feeding studies and whole-cell biotransformation, we have established the roles of the otherwise cryptic AmbK in THP formation and AmbH in PKS module iteration in ambruticin biosynthesis. Our work marks a significant step towards a deeper understanding of the complex chemistry and biology of ambruticin biosynthesis, laying the groundwork for further investigations aimed at elucidating the mechanistic details of unusual enzymology in polyketide assembly and modifications.
Results
Development of the genetic engineering method for S. cellulosum
Strains of S. cellulosum typically exhibit bright pigmentation attributed to the presence of carotenoids (Fig. 1b). These pigments are usually biosynthesised through a well-conserved biosynthetic pathway (Fig. 2a)41. Inactivation or knockout of the crtB gene, which encodes phytoene synthase, a key enzyme in carotenoid biosynthesis, has been observed to cause an orange-to-white phenotype change in many microorganisms42,43,44. Our comparative analysis of all available whole-genome sequences of S. cellulosum revealed a high level of conservation in their carotenoid biosynthetic gene clusters (Supplementary Fig. 2 and Supplementary Table 1), indicating that crtB could potentially serve as a reporter gene for Sorangium.
a Biosynthesis of carotenoids in S. cellulosum So ce10. b Construction of the pXZ-crtBKO plasmid and possible outcomes from homologous recombination. c Verification of crtB disruption by phenotype (i) and genotype (ii). The colour of the numbers above the gel lanes indicates the colony appearance: orange numbers indicate colonies with an orange phenotype, and black numbers indicate colonies with a white phenotype. A total of 42 white colonies from three independent biological replicates (N = 3) were validated by genotype, uncropped gels are provided in the Source data file. d Optimisation of the electroporation conditions. Error bars represent mean ± SD. N = 3 biological replicates for each temperature condition; N = 4, 3, 4, 5 replicates for OD600 of 0.5, 1, 1.5, 2, respectively; N = 5, 4, 4, 5 replicates for Voltage of 12, 15, 18, 21 kV/cm, respectively; N = 5, 10, 5, 9, 5 replicates for pulse number of 1, 2, 3, 4, 5, respectively. e, f Impact of homology arm length on overall transformation efficiency and ratio of double-crossover events. Error bars represent mean ± SD, N = 4 biological replicates. g Disruption of the crtB gene in S. cellulosum So ce307 and So ce836. Statistical analysis was performed using two-tailed Student’s t test or one-way ANOVA followed by Tukey’s multiple comparisons test (***p < 0.001; ****p < 0.0001). Source data are provided as a Source data file.
To construct the plasmid for targeted inactivation of crtB in S. cellulosum So ce10, an antibiotic selection marker must be inserted between the upstream and downstream homology arms of the target site. Due to intrinsic antibiotic resistance genes in their genome, strains of Sorangium are resistant to many commonly used antibiotics12. Our preliminary screening indicated that S. cellulosum So ce10 is sensitive to hygromycin B; therefore, the hygromycin-resistance gene hyg was initially chosen as the selection marker. For the plasmid backbone, the pEX18 suicide vector, which has been widely used for Gram-negative bacteria45,46, was selected. Consequently, the hyg marker flanked by upstream and downstream homology arms of ~1500 bp was ligated with the backbone vector to create the knockout plasmid targeting crtB, designated as pXZ-crtBKO (Fig. 2b).
We next aimed to deliver the plasmid pXZ-crtBKO into S. cellulosum So ce10 by electroporation. Electrocompetent cells prepared in 10% glycerol were mixed with plasmid DNA, electroporated, and immediately transferred to recovery medium. The recovered cells were then plated on hygromycin-containing selection plates. After 5-7 days of incubation, both orange and white colonies were observed, indicating successful transformation (Fig. 2c, i). By homologous recombination, a single-crossover event would integrate the plasmid into the genome without disrupting crtB, thus retaining the orange phenotype (Fig. 2bi, ii). In contrast, a double-crossover event would result in the replacement of crtB by the hygromycin selection marker, leading to a white phenotype (Fig. 2biii). PCR verification confirmed that the red and white colonies are indeed single-crossover and double-crossover transformants, respectively (Fig. 2cii).
To enhance our workflow efficiency, we extensively screened and optimised electroporation conditions. This included adjusting optical density and the temperature of competent cells, as well as refining electroporation voltage and the number of pulses (Supplementary Table 2). A transformation efficiency of up to 1.2 × 10³ colony-forming units (CFU)/μg DNA was achieved when electrocompetent cells were prepared at room temperature47 using cells at OD600 0.5, with electroporation carried out at a voltage of 15 kV/cm for 3 times (Fig. 2d). In addition, the impact of homology arm length was explored. We determined that a minimum arm length of 1000 bp is necessary for efficient homologous recombination (Fig. 2e). However, an arm length of ~1500 bp is preferred to maximise the occurrence of double-crossover events (Fig. 2f). Similar results were observed when using other plasmid backbones such as pKC1139 and pYH7, which are commonly employed for gene knockout studies in Streptomyces strains48,49. For sequential genetic manipulation, alternative antibiotic selection markers were investigated. Both the tetracycline-resistance gene and the chloramphenicol-resistance gene were found to be effective markers for So ce10, albeit with lower efficiency compared to the hygromycin marker.
The versatility of our electroporation method was tested on other Sorangium strains, including the jerangolid producer So ce307 and the epothilone producer So ce836. In both cases, successful disruption of the crtB gene was achieved with comparable efficiency to that of So ce10 (Fig. 2g and Supplementary Fig. 3). Notably, previous attempts to genetically manipulate So ce307 using conjugation methods were unsuccessful31 while no prior genetic manipulation studies have been reported for So ce83650,51,52. To directly benchmark our electroporation protocol against the traditional conjugation-based method, we investigated crtB inactivation in these strains using both approaches in parallel. Despite repeated efforts, the conjugation method failed to yield any transformants in So ce307 or So ce836. In the case of So ce10, only a few single-crossover mutants were obtained, and no double-crossover mutants could be recovered (Supplementary Fig. 4). Additionally, the conjugation process was complicated by substantial background growth of the donor E. coli S17.1 strain. In contrast, our electroporation protocol consistently achieved efficient transformation and a reproducible frequency of double-crossover events across strains, highlighting both its technical advancement and broader applicability (Supplementary Table 3).
AmbK is an epoxide hydrolase catalysing THP ring formation
Our next goal was to investigate key steps in the ambruticin biosynthetic pathway in S. cellulosum So ce10 using the genetic engineering strategy we had established, beginning with formation of the C3-C7 THP ring. In their biosynthetic studies, Reeves and co-workers disrupted ambJ, which encodes a flavin-dependent epoxidase and isolated ambruticin J (1) as the putative PKS product lacking the 6,7-epoxide31. Thus, it is hypothesised that AmbJ catalyses selective epoxidation of the 6,7-double bond to give 6,7-epoxy-ambruticin J (2), followed by epoxide-opening at C-7 through nucleophilic attack by the 3-hydroxyl group to form the six-membered THP ring and thereby generating ambruticin F (3) (Fig. 1d). Similar epoxide formation/epoxide-opening cascades have been identified in the biosynthesis of lasalocid53,54, mupirocin46 and xiamenmycin55, and in each case an additional epoxide hydrolase (Lsd19, MupZ and XimE respectively) is required to favour formation of the typically disfavoured THP ring56. In the absence of these epoxide hydrolases, spontaneous cyclisation products with tetrahydrofuran (THF) rings are isolated from the epoxide intermediates (Supplementary Fig. 5).
To date, no epoxide hydrolase has been identified within the amb BGC. Hence, it has been speculated that either the epoxidase AmbJ may be multifunctional, or alternatively, the presence of the 8,9-alkene moiety could increase the susceptibility of C-7 of the oxirane to nucleophilic attack57,58. Indeed, our recent model studies using truncated surrogates of ambruticin J indicated that both the 5-OH and the 8,9-alkene may influence the intramolecular ring opening, leading to the formation of a THP product59. However, details of the biosynthetic mechanism of tetrahydropyran ring formation in the ambruticins remain unknown.
With this in mind, we carefully examined the genomic neighbourhoods of the ambJ gene in the BGC to search for a potential epoxide hydrolase. Interestingly, a previously unidentified short region of ca. 500 bp in length immediately downstream of ambJ was found (Fig. 3a). Sequence analysis of this region identified two potential overlapping open reading frames (ORFs) in the same orientation, which could be translated into peptide sequences of 136 (ORF1) and 128 (ORF2) amino acids, respectively (Supplementary Fig. 6). BLAST searches of these two sequences suggested that ORF1 might be non-functional as no significant similarities were found, whereas ORF2 appears to encode for a YciI family protein, which we designated as AmbK. Protein structure prediction of AmbK by AlphaFold360 indicated a similar α + β overall fold to that of Lsd1954, MupZ46, and XimE55, despite low sequence homology (Fig. 3a).
a Identification of ambK from the amb BGC. b HPLC traces of the So ce10 ΔambJ and ΔambK mutants. c Structural elucidation of ambruticin K (4). d Confirmation of AmbK as the epoxide hydrolase responsible for THP ring formation.
We hypothesised that AmbK might be a candidate for the cryptic epoxide hydrolase implicated in THP formation and so investigated the function of this putative enzyme by first disruption of the corresponding gene in the ambruticin-producing strain S. cellulosum So ce10 (Supplementary Fig. 7). Using our developed method, ambJ and ambK knock-out mutant strains were successfully generated. As expected, cultures of the resultant ΔambJ mutant produced ambruticin J (1, m/z 457 [M-H]-) as the major metabolite (Fig. 3bii)31. In contrast, cultures of the ΔambK mutant strain contained none of the usual metabolites (e.g. ambruticin S and the VS series of compounds) found in the wild-type strain (Fig. 1b) but instead yielded a compound with a mass of 492 Da (4, m/z 491 [M-H]-) as the major metabolite (Fig. 3biii). 1H and 13C NMR data analysis of compound 4, compared with those of ambruticin J (1) and F (3), showed that a DHP ring and 3,5-dihydroxy acid were present but there were no signals which could be attributed to the cyclopropyl group (Supplementary Fig. 8). Further detailed analysis of its 2D NMR data, including 1H-1H COSY, HSQC and HMBC, suggested that compound 4, named ambruticin K, bears hydroxyl groups at C-6 and C-12, and features a conjugated 7E,9E-diene (Supplementary Table 4), as opposed to the 6E,8E-diene present in ambruticin J (Fig. 3c). Acetylation of ambruticin K (4) gave tetraacetate 5 and the 1H-NMR spectrum showed with characteristic downfield shifts of the signals assigned to 3-H, 5-H, 6-H and 12-H in accord with the proposed structure (Fig. 3c and Supplementary Fig. 9). The absolute configuration at C-12 was determined to be R by Mosher ester analysis of the acetonide derivative of 4 (Supplementary Methods 1 and 2, Supplementary Fig. 10 and Supplementary Table 5)61. We propose that ambruticin K is formed via addition of water to C-12 of the proposed intermediate 6,7-epoxy-ambruticin J (2), leading to opening of the cyclopropane ring, and concomitant formation 7,9-diene and opening of the 6,7-epoxide to give the 7,9-dien-6-ol 4 (Fig. 3d). Whilst 2 was not seen in crude extracts, it is well established that in many biosynthetic pathways (e.g. lasalocid53,54 and mupirocin46) and synthetic studies59 involving epoxide forming/epoxide opening cascades that epoxide intermediates are unstable and not detectable being readily transformed to a more stable product.
Isolation of ambruticin K (4) from cultures of the ΔambK mutant of S. cellulosum indicated that AmbK is the elusive epoxide hydrolase required for THP ring formation in ambruticin biosynthesis. To verify the function of AmbK, in vivo biotransformations were undertaken using E. coli cells overexpressing AmbJ and AmbK. Soluble expression of both proteins was achieved after condition optimisations (Supplementary Fig. 11a). First, ambruticin J (1) was incubated with AmbJ-expressing E. coli cells, which gave the expected ambruticin K (4) (Supplementary Fig. 11b, ii). Pleasingly, incubation of ambruticin J with E. coli cells co-expressed with both AmbJ and AmbK gave ambruticin F (3) with the characteristic THP ring as confirmed by NMR (Supplementary Fig. 11biii). Furthermore, ambruticin K (4) with the co-expressed AmbJ and AmbK simply returned the starting material 4 (Supplementary Fig. 11biv) in accord with the proposal that ambruticin K is not an intermediate in ambruticin biosynthesis but arises through addition of water to the proposed intermediate 6,7-epoxy-ambruticin J (2). Overall, these studies confirm that AmbJ catalyses selective epoxidation of ambruticin J (1) to produce 6,7-epoxy-ambruticin J (2), followed by the epoxide hydrolase AmbK catalysing regioselective 6-membered ring formation to generate the THP ring in ambruticin biosynthesis (Fig. 3d).
AmbH is an iterative module for polyketide backbone assembly
Our next goal was to gain a deeper understanding of assembly of the polyketide backbone. The chain assembly has been proposed to initially follow the canonical type I PKS biosynthetic logic by the loading module and elongation modules 1–6 in AmbA-AmbE (Supplementary Fig. 1)31. Module 7 in AmbE then performs three iterative chain elongations, followed by further downstream processes involving AmbF, AmbG, and AmbI to generate the cyclopropane ring-containing backbone. It was postulated that this backbone then undergoes another round of elongation by the terminal PKS module AmbH, and is subsequently hydrolysed by its thioesterase (TE) to release ambruticin J31. For many years, these steps lacked detailed, direct experimental evidence to support the proposed assembly process in ambruticin biosynthesis. However, important insights were revealed from studies by Hahn and co-workers using extensive bioinformatic analysis and in vitro enzymatic assays37. They reannotated AmbG as a fatty acyl:adenylate ligase (FAAL)–acyl carrier protein (ACP) didomain and showed that it adenylates a wide range of free carboxylic acids with broad structural diversity (Fig. 4a).
a Previous in vitro studies of AmbG by Hahn and co-workers37. b HPLC trace of the So ce10 ΔambG mutant. c Structural elucidation of ambruticin G (6). d Feeding study of ambruticin G (6) by the ΔambA-I/ΔambJ mutant. e Feeding study of ambruticin G (6) by the ΔambA-I mutant. f The iterative module AmbH converts ambruticin G to ambruticin J, which is further metabolised by AmbJ and AmbK along with downstream enzymes to the final ambruticin VS compounds.
Inspired by Hahn’s work, we hypothesised that inactivating AmbG may facilitate capture of cryptic intermediates during polyketide chain assembly. With this in mind, the FAAL domain of AmbG was disrupted and the ΔambG mutant strain obtained (Supplementary Fig. 7c). A compound with a mass of 370 Da was isolated from cultures of the ΔambG mutant strain of S. cellulosum (Fig. 4b). Structural elucidation by HRMS and extensive NMR analysis confirmed that this product included an unsaturated carboxylic acid with intact cyclopropane and dihydropyran rings. Signals were apparent for 4 double bonds in the chain, including a 2E,4E-conjugated diene (Supplementary Table 6). The compound, named ambruticin G (6) (Fig. 4c), is shorter than the mature chain of ambruticin J, effectively lacking two acetate units in accord with its potential role as a biosynthetic precursor to ambruticin J (Supplementary Fig. 12). Since AmbH is presumed to be the sole PKS module acting after AmbG (Supplementary Fig. 1), we hypothesised that AmbH could extend ambruticin G by two rounds of chain elongation to afford ambruticin J. This hypothesis is supported by the KS-AT-KR-ACP-TE domain architecture of the AmbH module, where recursive chain extension and ketoreductase-catalysed reduction would be expected to yield the C-3 and C-5 hydroxyl groups found in ambruticin J62.
To test our hypothesis, the entire continuous region of the amb BGC upstream of ambG (ambA-ambI) was knocked out from the ΔambJ mutant to give the ΔambA-I/ΔambJ multi-domain mutant (Supplementary Fig. 7d). As expected, cultures of this strain produced no ambruticin-related metabolites as the main PKS modules were absent. However, AmbG and AmbH remained intact and functional in this strain. Subsequent feeding studies of ambruticin G (6) into this ΔambA-I/ΔambJ mutant were conducted, and full conversion to ambruticin J (1) was observed after 3 hours of incubation (Fig. 4d). In a parallel experiment, the entire region of ambA-ambI was deleted from the wild-type strain. Feeding ambruticin G into the resulting ΔambA-I mutant showed conversion to ambruticin S and the VS series compounds, as is found in the wild-type strain (Fig. 4e). These results clearly reveal that ambruticin G is a biosynthetic intermediate and highlight AmbH as an iterative module catalysing dual chain elongation of ambruticin G to afford ambruticin J (Fig. 4f).
An updated pathway for ambruticin biosynthesis
Reeves and co-workers proposed that starting from AmbF in ambruticin biosynthesis, unusual steps occur, including a Favorskii rearrangement and a carbon atom excision event, which would deliver a cyclopropyl polyketide intermediate to the terminal PKS module AmbH for a final round of chain elongation, followed by chain release to afford ambruticin J for further post-PKS modifications (Fig. 5a)31. Collectively, the results described herein have revealed the critical roles of the otherwise cryptic AmbH and AmbK, providing significant insights into both the assembly of the polyketide backbone and the post-PKS tailoring processes involved in ambruticin biosynthesis in S. cellulosum So ce10. These findings have led us to propose a revised biosynthetic pathway for ambruticin (Fig. 5b).
a Previously proposed biosynthetic pathway of ambruticin31. b Revised biosynthetic pathway of ambruticin based on this study.
With identification of ambruticin G (6) as a biosynthetic intermediate, it is now evident that the terminal module AmbH is iterative and catalyses two rounds of chain elongation (Fig. 5b). Subsequent THP ring formation from ambruticin J (1) requires AmbJ, which functions as the epoxidase, and in this work, an additional epoxide hydrolase, AmbK, has been identified. In the absence of AmbK, hydrolysis of epoxide 2 occurs giving ambruticin K (4), which is not an intermediate in ambruticin biosynthesis. Ambruticin F (3) contains all 3 rings (THP, DHP, and trisubstituted cyclopropane) characteristic of the ambruticins, and further tailoring modifications at C-5 occur to yield the final metabolites (Fig. 5b).
A further important feature to note is that the structure of ambruticin G (6) incorporates the DHP ring characteristic of the ambruticins. This result is in accord with our recent studies, which indicate that desaturation of the THP to a DHP catalysed by the Rieske oxygenase AmbP occurs prior to formation of the C3-C7 THP ring40. The results described herein indicate that the timing of the desaturation step can be as early as the PKS assembly stage.
The mechanism of formation of the cyclopropane ring moiety in ambruticin biosynthesis remains unknown. However, since the truncated metabolite ambruticin G (6) contains an intact cyclopropane ring, it is now apparent that the cyclopropane ring must be formed prior to AmbG channelling the free carboxylic acid intermediate back onto the assembly-line for additional PKS processing. The hydrolase domain (H) and pyridoxal cofactor binding motif domain (Px) in the AmbF module, along with the flavin-dependent monooxygenase AmbI, are most likely to play pivotal roles in this process (Fig. 5b). Studies are currently underway in our laboratories to investigate these fascinating biotransformations.
Discussion
The studies reported herein have led to the development of an efficient electroporation method for transforming foreign DNA into various Sorangium cellulosum strains, enabling effective genetic engineering via double-crossover homologous recombination within a single step in these genetically intractable myxobacteria. By using different antibiotic selection markers, we also achieved sequential deletions in these organisms. Nevertheless, a potential limitation of our study is the current unavailability of a broader panel of Sorangium strains to comprehensively assess the generalisability of our electroporation approach. While our method demonstrates high efficiency in Sorangium strains that grow in suspension, its applicability to strains exhibiting clumped or aggregated growth remains to be fully evaluated. Our preliminary experiments with the recently isolated icumazole-producing strain Sorangium cellulosum MBD-1, which displays dense clumping during growth, suggest that transformation is still possible, albeit with reduced efficiency. These observations underscore the influence of morphological characteristics in transformation outcomes and highlight the need for further strain-specific optimisation to extend the applicability of this method to a wider range of Sorangium isolates.
This method has been applied to genetic engineering in S. cellulosum So ce10, leading to a greater understanding of the biosynthesis of the antifungal polyketides, the ambruticins. Through a combination of targeted gene knockouts, whole-cell biotransformation, and feeding experiments, we identified a truncated metabolite, ambruticin G (6), and revealed the role of the iterative PKS module AmbH in polyketide backbone assembly of ambruticin biosynthesis. Furthermore, a previously unknown epoxide hydrolase, AmbK, has been identified and shown to be crucial for THP ring formation during post-PKS modification. Building on these findings, we have proposed an updated pathway for ambruticin biosynthesis (Fig. 5b), which lays the foundation for further investigations aimed at elucidating mechanistic details of its unique enzymology, particularly the cyclopropyl ring formation step.
Despite myxobacteria being renowned for producing a wealth of structurally diverse and biologically active natural products, we are still only beginning to uncover the full extent of their capabilities6. In future, using the tools to genetically manipulate Sorangium strains reported herein will significantly contribute to unlocking the vast and largely untapped biosynthetic potential of myxobacteria. Such efforts hold immense promise for the creation of novel analogues with altered chemical structures, potentially leading to compounds with enhanced or entirely new biological activities, such as improved antimicrobial, anticancer, or antiviral properties. As we refine these tools and gain deeper insights into the complex enzymology of these biosynthetic pathways, we may also uncover entirely new classes of natural products, further expanding the chemical and biological diversity accessible from myxobacteria.
Methods
General experimental
Strains of Sorangium cellulosum were purchased from the German Collection of Microorganisms and Cell Cultures (DSMZ) and maintained on HS agar (0.4% glucose, 0.15% Casein peptone, 0.1% KNO3, 0.1% MgSO4·7H2O, 0.008% NaFe-EDTA, 0.2% HEPES, 0.00625% K2HPO4, 0.0075% CaCl2·2H2O, 1.5% agar, pH 7.2). Oligonucleotides for polymerase chain reactions (PCR) were purchased from Tsingke (China). PCR amplification was performed using Phanta Flash Super-Fidelity DNA Polymerase (Vazyme, China). Plasmids were purified using a HighPure Rapid Mini Plasmid Kit (Biomed, China), and PCR products were purified with the HiPure Gel Pure DNA Mini Kit (Magen, China). Restriction enzymes were purchased from New England Biolabs (Ipswich, MA, USA). DNA ligation was performed using ClonExpress Ultra One Step Cloning Kit (Vazyme, China). Molecular biology experiments were conducted following the manufacturer’s instructions. Strains, plasmids, and oligonucleotides used in this study are listed in Supplementary Tables 7 and 8 and Supplementary Data 1.
LC-MS data were obtained on a SHIMADZU LC-MS system comprising a SIL-20AXR autosampler, LC-20ADXR liquid chromatograph, SPD-M40 photodiode array detector, and SHIMADZU LCMS-2010 EV liquid chromatography-mass spectrometry. HPLC-grade H2O and MeCN were added with 0.1% formic acid as the solvent system. Analytical LC-MS data were obtained using a Phenomenex Kinetex column (C18, 250 × 4.6 mm, 5 µm) at a flow rate of 1 mL min-1. Column chromatographic separations were carried out using Sephadex LH-20 (Cytiva) as packing materials. Preparative HPLC purification was carried out using a SilGreen CH0525010C 18AQ column (C18, 250 × 10.0 mm, 5 µm) at a flow rate of 4.7 mL min−1. High-resolution electrospray ionisation mass spectrometry (HR-ESI-MS) was performed on a Waters ACQUITYI-Class PLUS/Xevo G2-XS Tof instrument. 1H, 13C NMR and 2D NMR data were collected on a Bruker Avance Ⅲ 500 MHz or Bruker Ascend 800 MHz NMR spectrometer.
Genetic engineering of S. cellulosum
The upstream and downstream fragments of the target gene or region were amplified from genomic DNA via PCR. Knockout plasmids were constructed by inserting the hygromycin, tetracycline, or chloramphenicol selection marker, flanked by the upstream and downstream homology arms, into the KpnI/HindIII-linearised pXZ vector. These plasmids were introduced into Sorangium cells via electroporation, and double-crossover transformants were selected on HS agar supplemented with appropriate concentrations of hygromycin, tetracycline, or chloramphenicol. Sequences of the antibiotic resistance genes and the working concentrations of antibiotics are shown in Supplementary Data 2.
For electrocompetent cells preparation, a 1% (v/v) inoculum from mid-exponential phase preculture of S. cellulosum strain was inoculated into 100 mL HS broth, cultivated in shake flasks at 30 °C, 200 rpm. Upon reaching an OD600 of 0.5 (approximately 30 hours after inoculation), cells were harvested by centrifugation at 4500 × g. The pellet was then washed three times with 10 mL 10% (v/v) glycerol at 15,000 × g. The cell pellet was then resuspended in 10% glycerol to a final volume of 900 μL and aliquoted into 90 µL/ EP tubes for storage at −80 °C.
For electroporation, the electrocompetent cells were thawed on ice, and 500 ng–5 µg of plasmid was mixed with 90 µL of competent cells. The mixture was then transferred into a precooled 1 mm cuvette, and subjected to electroporation using a Bio-Rad MicroPulser, at a voltage of 15 kV/cm for 3 times. Immediately after pulsing, 500 µL of precooled liquid HS medium was added, followed by transferring into 3 mL of liquid HS medium. Cells were allowed to recover at 30 °C and 200 rpm for 2 days. Subsequently, cells were collected by centrifugation and spread on HS agar plates supplemented with appropriate antibiotics. Transformants typically appear after incubation at 30 °C for 5–7 days. Knockout mutants were screened by PCR and further confirmed by sequencing.
For conjugative transformation of plasmid pXZ-crtBKO into Sorangium cellulosum, the plasmid was first introduced into chemically competent E. coli S17.1 cells via heat shock. The resulting E. coli donor strain carrying pXZ-crtBKO was cultured overnight in LB medium containing kanamycin (50 µg/mL) at 37 °C. A 1% v/v inoculum was then transferred to 50 mL of fresh kanamycin-containing LB medium and grown to early exponential phase (OD600 ≈ 0.4–0.6), while the recipient Sorangium cultures were grown in 50 mL HS medium at 30 °C until reaching an OD600 of 0.5. Both cell types were harvested by centrifugation and washed with LB and S42 medium31, respectively. The Sorangium cells were resuspended and subjected to a 10-min heat shock at 50 °C in pre-warmed S42 medium. Equal volumes (typically 1 mL each) of donor and recipient were mixed, centrifuged, and resuspended in 200 μL of glucose-free S42 medium. The cell mixture was spotted onto an antibiotic-free S42 agar plate and incubated at 30 °C for 48 h. The resulting mating puddle was resuspended in 0.5 mL S42 liquid medium and plated onto S42 agar supplemented with appropriate concentrations of hygromycin and carbenicillin. Plates were incubated at 30 °C for 7–10 days until Sorangium transformant colonies emerge.
General fermentation procedure
Wild-type or mutant strains of S. cellulosum So ce10 were inoculated on HS agar plates and incubated at 30 °C for 3 days. Colonies from these plates were scraped and used to inoculate 100 mL of liquid HS medium in a 500 mL flask. This seed culture was incubated at 30 °C, 200 rpm for 2–3 days. For production fermentation, 20% of the seed culture was used to inoculate SF1-P medium (0.3% soy peptone, 0.6% fructose, 0.1% MgSO4·7H2O, 0.1% CaCl2·2H2O, 0.008% ferric citrate, and 0.05 M HEPES, pH 7.6; ferric citrate and HEPES were filter sterilised and added after autoclaving). The culture was supplemented with 1 g L-1 of fructose every 24 h, starting on the second day, and incubated at 30 °C, 200 rpm for 7 days. The supernatant was extracted three times with ethyl acetate (EtOAc). The combined EtOAc extracts were evaporated in vacuo to yield a crude extract, which was then analysed by LC-MS or subjected to further purification.
Isolation and purification of ambruticin J, ambruticin K and ambruticin G
A 2.0 L scale fermentation of the ΔambJ mutant of S. cellulosum So ce10 was performed following the general fermentation procedure. The crude extract was initially purified using Sephadex LH-20 column chromatography with MeOH as the eluent to give a crude fraction of ambruticin J. This fraction was further purified by HPLC eluting with a gradient elution of 40% to 95% acetonitrile in water over 20 min, with ambruticin J eluting at ~85% acetonitrile, yielding 1.6 mg of the compound. Similarly, ambruticin K (eluting at ~70% acetonitrile) was isolated from the ΔambK mutant of S. cellulosum So ce10, following the same purification protocol, with a yield of 1.4 mg L−1. Ambruticin G (eluting at approximately 90% acetonitrile) was obtained from the ΔambG mutant strain with a yield of 1 mg L−1. For HR-ESI-MS and NMR data of the compounds, see in Supplementary Tables 4 and 6 and Supplementary Figs. 13–38.
Chemical derivatisation of ambruticin K
For acetylation of ambruticin K, 4 was dissolved in 25 µL of pyridine and mixed with 25 µL of acetic anhydride. The reaction was stirred at room temperature overnight and then quenched with water. The reaction mixture was extracted with EtOAc, and the organic layer was dried under vacuum to obtain a crude extract. This extract was then purified using HPLC with a gradient elution of 5% to 95% MeCN in water over 20 min to yield compound 5.
For preparation of the acetonides derivatives of ambruticin K and their Mosher esters, see Supplementary Methods 1 and 2.
Feeding studies of ambruticin G
A 50 mL culture of the So ce10 ΔambA-I or So ce10 ΔambA-I/ΔambJ strain in SF1-P medium was incubated at 30 °C for 48 h. Subsequently, 0.1 mg of ambruticin G dissolved in 10 μL of MeOH was added to each culture. For the So ce10 ΔambA-I strain, incubation continued at 30 °C for an additional 30 min, while for the So ce10 ΔambA-I/ΔambJ strain, 0.1 mg of ambruticin G was added every 30 min, with a total incubation period of 6 h. Three volumes of EtOAc were used for extraction, and the solvent was removed under vacuum. The dried samples were redissolved in 200 μL MeOH and analysed by LC-MS.
Expression of AmbJ and AmbK in E. coli
The ambJ and ambK genes were amplified by PCR from the genomic DNA of S. cellulosum So ce10 using primers listed in Supplementary Data 1. The resulting PCR products were introduced into pET151 and pET28a vectors, respectively. For AmbJ expression, the pET151-ambJ construct and pGro7 (expressing GroES-GroEL) were co-transformed into E. coli BL21(DE3) cells. The cells were grown overnight in 5 mL of lysogeny broth (LB) medium supplemented with 50 μg mL-1 carbenicillin and 25 μg mL-1 chloramphenicol at 37 °C with shaking at 200 rpm. 1 mL of this pre-culture was used to inoculate 100 mL of LB medium containing 50 μg mL-1 carbenicillin, 25 μg mL-1 chloramphenicol, and 0.5 g L-1 L-arabinose. The culture was incubated at 37 °C with shaking at 200 rpm until the OD600 reached 0.5-0.6. Protein expression was induced by adding 0.1 mM isopropyl β-D-1-thiogalactopyranoside (IPTG), and the cells were incubated overnight at 16 °C with shaking at 120 rpm. For AmbK expression, the pET28a-ambK construct was transformed into E. coli BL21(DE3). Protein expression followed similar procedure with LB medium supplemented with 50 μg mL-1 kanamycin. For co-expression of AmbJ and AmbK, the same procedure used for AmbJ expression was followed, but with the LB medium supplemented with 50 μg mL-1 carbenicillin, 50 μg mL-1 kanamycin, and 25 μg mL-1 chloramphenicol.
Whole-cell biotransformation
An overnight culture (50 mL) of E. coli BL21 (DE3) cells overexpressing AmbJ or both AmbJ and AmbK was grown in LB medium. The cells were centrifuged at 4 °C and 2400 × g for 5 min, and the pellets were resuspended in 2 mL of 100 mM potassium phosphate buffer (pH 7.2) containing 20 mM glucose. Subsequently, 0.5 mg of ambruticin J or ambruticin K dissolved in 50 μL of MeOH was added. The mixtures were incubated at 30 °C and 200 rpm for 12 h. Reactions were quenched by adding three volumes of ethyl acetate, followed by vortexed and centrifugation. The ethyl acetate layer was dried under vacuum, dissolved in 400 μL methanol, and subjected to LC-MS analysis.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Whole genome sequences of Sorangium cellulosum So ce10 and Sorangium cellulosum So ce307 have been deposited at NCBI’s GenBank under accession number CP162580 and CP162579, respectively. Other data supporting the findings of this study are available in the main text, supplementary information and from corresponding author(s) upon request. Source data are provided with this paper.
References
Newman, D. J. & Cragg, G. M. Natural products as sources of new drugs over the nearly four decades from 01/1981 to 09/2019. J. Nat. Prod. 83, 770–803 (2020).
Atanasov, A. G. et al. Natural products in drug discovery: advances and opportunities. Nat. Rev. Drug Discov. 20, 200–216 (2021).
Hemmerling, F. & Piel, J. Strategies to access biosynthetic novelty in bacterial genomes for drug discovery. Nat. Rev. Drug Discov. 21, 359–378 (2022).
Weissman, K. J. & Müller, R. Myxobacterial secondary metabolites: bioactivities and modes-of-action. Nat. Prod. Rep. 27, 1276 (2010).
Saggu, S. K., Nath, A. & Kumar, S. Myxobacteria: biology and bioactive secondary metabolites. Res. Microbiol. 174, 104079 (2023).
Wang, C. et al. Recent advances in discovery and biosynthesis of natural products from myxobacteria: an overview from 2017 to 2023. Nat. Prod. Rep. 41, 905–934 (2024).
Gerth, K., Perlova, O. & Müller, R. Sorangium cellulosum. Myxobacteria: Multicellularity and Differentiation (ed. Whitworth, D. E.) 329–348 (ASM Press, 2014).
Höfle, G. et al. Epothilone A and B-novel 16-membered macrolides with cytotoxic activity: isolation, crystal structure, and conformation in solution. Angew. Chem. Int. Ed. 35, 1567–1569 (1996).
Naini, A., Sasse, F. & Brönstrup, M. The intriguing chemistry and biology of soraphens. Nat. Prod. Rep. 36, 1394–1411 (2019).
Lilic, M. et al. The antibiotic sorangicin A inhibits promoter DNA unwinding in a Mycobacterium tuberculosis rifampicin-resistant RNA polymerase. Proc. Natl. Acad. Sci. USA 117, 30423–30432 (2020).
Fleta-Soriano, E. et al. The myxobacterial metabolite ratjadone A inhibits HIV infection by blocking the Rev/CRM1-mediated nuclear export pathway. Microb. Cell. Fact. 13, 17 (2014).
Schneiker, S. et al. Complete genome sequence of the myxobacterium Sorangium cellulosum. Nat. Biotechnol. 25, 1281–1289 (2007).
Han, K. et al. Extraordinary expansion of a Sorangium cellulosum genome from an alkaline milieu. Sci. Rep. 3, 2101 (2013).
Wenzel, S. C. & Müller, R. Myxobacterial natural product assembly lines: fascinating examples of curious biochemistry. Nat. Prod. Rep. 24, 1211 (2007).
Julien, B. & Rodriguez, E. Genetic engineering of myxobacterial natural product biosynthetic genes, Manual of Industrial Microbiology and Biotechnology (eds. Baltz, R. H. et al.) 426–437 (ASM Press, 2014).
Yue, X., Sheng, D., Zhuo, L. & Li, Y. Genetic manipulation and tools in myxobacteria for the exploitation of secondary metabolism. Eng. Microbiol. 3, 100075 (2023).
Whitworth, D. E., Sydney, N. & Radford, E. J. Myxobacterial genomics and post-genomics: a review of genome biology, genome sequences and related ‘omics studies. Microorganisms 9, 2143 (2021).
Jaoua, S., Neff, S. & Schupp, T. Transfer of mobilizable plasmids to Sorangium cellosum and evidence for their integration into the chromosome. Plasmid 28, 157–165 (1992).
Pradella, S. et al. Characterisation, genome size and genetic manipulation of the myxobacterium Sorangium cellulosum So ce56. Arch. Microbiol. 178, 484–492 (2002).
Kopp, M. et al. Critical variations of conjugational DNA transfer into secondary metabolite multiproducing Sorangium cellulosum strains So ce12 and So ce56: development of a mariner-based transposon mutagenesis system. J. Biotechnol. 107, 29–40 (2004).
Kopp, M., Irschik, H., Pradella, S. & Müller, R. Production of the tubulin destabilizer disorazol in Sorangium cellulosum: biosynthetic machinery and regulatory genes. ChemBioChem 6, 1277–1286 (2005).
Ringel, S. M. et al. Ambruticin (W7783), a new antifungal antibiotic. J. Antibiot. 30, 371–375 (1977).
Connor, D. T., Greenough, R. C. & Von Strandtmann, M. W-7783, a unique antifungal antibiotic. J. Org. Chem. 42, 3664–3669 (1977).
Knauth, P. & Reichenbach, H. On the mechanism of action of the myxobacterial fungicide ambruticin. J. Antibiot. 53, 1182–1190 (2000).
Shubitz, L. F. et al. Efficacy of ambruticin analogs in a murine model of coccidioidomycosis. Antimicrob. Agents Chemother. 50, 3467–3469 (2006).
Vetcher, L. et al. The antifungal polyketide ambruticin targets the HOG pathway. Antimicrob. Agents Chemother. 51, 3734–3736 (2007).
Masschelein, J., Jenner, M. & Challis, G. L. Antibiotics from Gram-negative bacteria: a comprehensive overview and selected biosynthetic highlights. Nat. Prod. Rep. 34, 712–783 (2017).
Marcos-Torres, F. J., Volz, C. & Müller, R. An ambruticin-sensing complex modulates Myxococcus xanthus development and mediates myxobacterial interspecies communication. Nat. Commun. 11, 5563 (2020).
Hahn, F. & Guth, F. M. The ambruticins and jerangolids – chemistry, biology and chemoenzymatic synthesis of potent antifungal drug candidates. Nat. Prod. Rep. 37, 1300–1315 (2020).
Ma, S., Mandalapu, D., Wang, S. & Zhang, Q. Biosynthesis of cyclopropane in natural products. Nat. Prod. Rep. 39, 926–945 (2022).
Julien, B., Tian, Z., Reid, R. & Reeves, C. D. Analysis of the ambruticin and jerangolid gene clusters of Sorangium cellulosum reveals unusual mechanisms of polyketide biosynthesis. Chem. Biol. 13, 1277–1286 (2006).
Berkhan, G. & Hahn, F. A dehydratase domain in ambruticin biosynthesis displays additional activity as a pyran-forming cyclase. Angew. Chem. Int. Ed. 53, 14240–14244 (2014).
Friedrich, S. et al. Characterisation of the broadly-specific O-methyl-transferase JerF from the late stages of jerangolid biosynthesis. Molecules 21, 1443 (2016).
Berkhan, G., Merten, C., Holec, C. & Hahn, F. The interplay between a multifunctional dehydratase domain and a C-methyltransferase effects olefin shift in ambruticin biosynthesis. Angew. Chem. Int. Ed. 55, 13589–13592 (2016).
Sung, K. H. et al. Insights into the dual activity of a bifunctional dehydratase-cyclase domain. Angew. Chem. Int. Ed. 57, 343–347 (2018).
Lindner, F., Friedrich, S. & Hahn, F. Total synthesis of complex biosynthetic late-stage intermediates and bioconversion by a tailoring enzyme from jerangolid biosynthesis. J. Org. Chem. 83, 14091–14101 (2018).
Hemmerling, F., Lebe, K. E., Wunderlich, J. & Hahn, F. An unusual fatty acyl:adenylate ligase (FAAL)-acyl carrier protein (ACP) didomain in ambruticin biosynthesis. ChemBioChem 19, 1006–1011 (2018).
Guth, F. M. et al. Rieske oxygenase-catalyzed oxidative late-stage functionalization during complex antifungal polyketide biosynthesis. ACS Chem. Biol. 18, 2450–2456 (2023).
Hollmann, T. et al. Biocatalysts from biosynthetic pathways: enabling stereoselective, enzymatic cycloether formation on a gram scale. ACS Catal. 10, 4973–4982 (2020).
Bowen, J. I. et al. Combining total synthesis and genetic engineering to probe dihydropyran formation in ambruticin biosynthesis. Chem. Sci. 15, 5319–5326 (2024).
Liang, M., Zhu, J. & Jiang, J. Carotenoids biosynthesis and cleavage related genes from bacteria to plants. Crit. Rev. Food Sci. Nutr. 58, 2314–2333 (2018).
Du, K. et al. Reprogramming the endogenous type I CRISPR-Cas system for simultaneous gene regulation and editing in Haloarcula hispanica. mLife 1, 40–50 (2022).
Wang, J., Wei, J., Li, H. & Li, Y. High-efficiency genome editing of an extreme thermophile Thermus thermophilus using endogenous type I and type III CRISPR-Cas systems. mLife 1, 412–427 (2022).
Wu, Y. et al. Dissecting the arginine and lysine biosynthetic pathways and their relationship in haloarchaeon Natrinema gari J7-2 via endogenous CRISPR-Cas system-based genome editing. Microbiol. Spectr. 11, e00288–23 (2023).
Hmelo, L. R. et al. Precision-engineering the Pseudomonas aeruginosa genome with two-step allelic exchange. Nat. Protoc. 10, 1820–1841 (2015).
Wang, L. et al. A Rieske oxygenase/epoxide hydrolase-catalysed reaction cascade creates oxygen heterocycles in mupirocin biosynthesis. Nat. Catal. 1, 968–976 (2018).
Tu, Q. et al. Room temperature electrocompetent bacterial cells improve DNA transformation and recombineering efficiency. Sci. Rep. 6, 24648 (2016).
Bierman, M. et al. Plasmid cloning vectors for the conjugal transfer of DNA from Escherichia coli to Streptomyces spp. Gene 116, 43–49 (1992).
Sun, Y. et al. Organization of the biosynthetic gene cluster in Streptomyces sp. DSM 4137 for the novel neuroprotectant polyketide meridamycin. Microbiol 152, 3507–3515 (2006).
Gemperlein, K. et al. Polyunsaturated fatty acid biosynthesis in myxobacteria: different PUFA synthases and their product diversity. Chem. Sci. 5, 1733 (2014).
Boldt, J. et al. Bursts in biosynthetic gene cluster transcription are accompanied by surges of natural compound production in the myxobacterium Sorangium sp. Microbiol. Biotechnol. 16, 1054–1068 (2023).
Xie, F. et al. Insights into the biosynthesis of icumazole, unveiling a distinctive family of crotonyl-CoA carboxylase/reductase. Cell Rep. Phys. Sci. 4, 101394 (2023).
Shichijo, Y. et al. Epoxide hydrolase Lsd19 for polyether formation in the biosynthesis of lasalocid A: direct experimental evidence on polyene-polyepoxide hypothesis in polyether biosynthesis. J. Am. Chem. Soc. 130, 12230–12231 (2008).
Hotta, K. et al. Enzymatic catalysis of anti-Baldwin ring closure in polyether biosynthesis. Nature 483, 355–358 (2012).
He, B. et al. Enzymatic pyran formation involved in xiamenmycin biosynthesis. ACS Catal. 9, 5391–5399 (2019).
Bowen, J. I., Wang, L., Crump, M. P. & Willis, C. L. Synthetic and biosynthetic methods for selective cyclisations of 4, 5-epoxy alcohols to tetrahydropyrans. Org. Biomol. Chem. 20, 1150–1175 (2022).
Hemmerling, F. & Hahn, F. Biosynthesis of oxygen and nitrogen-containing heterocycles in polyketides. Beilstein J. Org. Chem. 12, 1512–1550 (2016).
Trentadue, K., Chang, C. F., Nalin, A. & Taylor, R. E. Enantioselective total synthesis of the putative biosynthetic intermediate ambruticin. J. Chem. Eur. J. 27, 11126–11131 (2021).
Bowen, J. I., Wang, L., Crump, M. P. & Willis, C. L. Ambruticins: tetrahydropyran ring formation and total synthesis. Org. Biomol. Chem. 19, 6210–6215 (2021).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold3. Nature 630, 493–500 (2024).
Hoye, T. R., Jeffrey, C. S. & Shao, F. Mosher ester analysis for the determination of absolute configuration of stereogenic (chiral) carbinol carbons. Nat. Protoc. 2, 2451–2458 (2007).
Hertweck, C. The biosynthetic logic of polyketide diversity. Angew. Chem. Int. Ed. 48, 4688–4716 (2009).
Acknowledgements
This work was supported by the National Key Research and Development Programme of China (2024YFC3407000 to L.W.), the National Natural Science Foundation of China (32270083 to L.W.), the Strategic Priority Research Programme of the Chinese Academy of Sciences (XDB0810000 to L.W.), and the Beijing Municipal Science & Technology Project, China (Z241100007724009 to L.W.). We are grateful to the EPSRC Bristol Chemical Synthesis Centre for Doctoral Training (EP/L015366/1) for a studentship to J.I.B.
Author information
Authors and Affiliations
Contributions
L.W. and C.L.W. conceptualised the project. X.Z., S.L., B.M., K.G., D.J., Y.H., H.C. and J.L. performed experiments. X.Z., S.L., J.I.B., M.P.C., C.L.W. and L.W. analysed data. X.Z., S.L., M.P.C., C.L.W. and L.W. wrote the manuscript with contributions from all the authors. All authors discussed and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Rolf Müller, Kenji Watanabe and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhong, X., Liu, S., Ma, B. et al. Genetic engineering of Sorangium cellulosum reveals hidden enzymology in myxobacterial natural product biosynthesis. Nat Commun 16, 7996 (2025). https://doi.org/10.1038/s41467-025-63441-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-63441-y