Abstract
Inflammatory bowel disease (IBD) arises from genetic-environmental interactions. Adenosine deaminases acting on RNA 1 (ADAR), an RNA-editing enzyme converting adenosine (A) to inosine (I), is essential for tissue homeostasis. Here we report that intestinal ADAR deficiency contributes to IBD pathogenesis in humans with reduced ADAR expression in patient intestinal crypts. Genetic or pharmacological inhibition of ADAR in mice causes spontaneous ileitis and colitis. Organoid studies show that ADAR loss leads to double-strand RNA (dsRNA) and endogenous retroviruses (ERVs) accumulation, disrupting intestinal homeostasis via melanoma differentiation-associated protein 5 (MDA5)-mediated dsRNA sensing and Janus kinase (JAK)-signal transducer and activator of transcription (STAT) signaling. Editome analyses identify Mda5 as an ADAR target, and edited Mda5 exhibits impaired dsRNA sensing. The human ADAR p.N173S mutation is a loss-of-function variant that fails to rescue IBD in intestinal Adar deficient mice, whereas JAK1/2 inhibitor Ruxolitinib attenuates IBD. We conclude that the ADAR-dsRNA/ERVs-MDA5-JAK/STAT axis is a potential therapeutic target for IBD.
Similar content being viewed by others
Introduction
Inflammatory bowel disease (IBD), including Crohn’s disease (CD) and ulcerative colitis (UC), is characterized by chronic inflammation of the digestive tract. IBD affects as many as 1.6 million North Americans, 2 million Europeans, and 2.6 million Chinese1,2,3. IBD can manifest in individuals of diverse ethnic backgrounds4. Prolonged inflammation results in structural and functional damage to the intestinal tract and represents a key risk factor for colon cancer5.
The etiology of IBD remains to be clearly defined. Epidemiological observations have implicated both genetic and environmental factors in the pathogenesis of IBD6,7. The worldwide increase in the incidence of IBD is undeniably associated with environmental factors, including smoking, diet, medication, urban living, socioeconomic status, stress, microbial exposure, intestinal permeability, appendectomy, and tonsillectomy8. Among genetic factors, family history is a risk factor for IBD9. Twin concordance studies provide robust evidence for the significant contribution of genetic factors to the susceptibility of CD and UC10,11. Genome-wide association studies (GWAS) have identified multiple genes and rare variants associated with IBD, such as those of the NOD2, IL23R, ATG16L1, SLC39A8, FUT2, TYK2, IFIH1 (MDA5), SLAMF8, and PLCG2 genes12,13. Understanding the genetic basis of IBD will help to combat this disease.
RNA editing, a post-transcriptional RNA modification, refers to the enzymatic conversion of RNA nucleotides. Adenosine deaminases acting on RNA 1 (ADAR) is an RNA editing enzyme that catalyzes the conversion of adenosine (A) to inosine (I)14,15. ADAR-mediated A-to-I RNA editing is essential to suppress cellular innate immune interferon responses to double-stranded RNA (dsRNA)16, which underlies the genetic variants of ADAR associated with common inflammatory diseases17. In humans, loss-of-function mutations in ADAR and gain-of-function mutations in its downstream dsRNA sensor MDA5 have been identified in several autoimmune diseases, such as Aicardi–Goutières syndrome (AGS)18,19. However, whether and how intestinal ADAR plays a role in IBD has not been reported.
In this work, we identify a loss-of-function human ADARN173S variant that activates MDA5-mediated dsRNA sensing and innate immune response and triggers bowel inflammation. The intestinal ADAR-dsRNA/ERVs-MDA5-JAK/STAT axis can be explored as a therapeutic target for IBD.
Results
Identification of ADARN173S as a loss function mutation that correlates with IBD incidence in patients
The mouse Adar1 has been reported to be essential for organ development and tissue homeostasis20. The ADAR gene is highly conserved across mammalian species, with 83% amino acid similarity between humans and mice, suggesting that ADAR may play a similar fundamental role in human biology. Indeed, the Genome Aggregation Database (gnomAD) (22) analysis showed that the human ADAR exhibits low and very low observed-to-expected ratios of missense and predicted loss-of-function (pLoF) variants, respectively (Fig. 1a), suggesting that ADAR is an indispensable gene. To determine the relevance of ADAR mutations in human IBD, we analyzed the association of ADAR gene single-nucleotide polymorphisms (SNPs) with IBD traits in human populations derived from a European IBD database (23). We identified two missense variants p.Asn (N) 173 Ser (S) (1:154574600T/C variant) and p.Gln (Q) 139 Pro (P) (1:154574702T/G variant), and one synonymous variant (1:154557848A/G variant) that were over-represented in samples from persons with IBD compared to the non-IBD cohort, and the carrier rates of ADARN173S and ADARQ139P in a Finnish cohort are approximately 1 in 126 and 1 in 9930, respectively (Fig. 1b and Supplementary Table 1). We went on to investigate the role of these variants in bowel inflammation. The structures of human ADAR and its Gln139 and Asn173 sites were predicted by AlphaFold (Fig. 1c).
a Observed-to-expected ratio of variants in ADAR based on the Genome Aggregation Database (gnomAD, https://gnomad.broadinstitute.org). pLoF predicted loss-of-function variants. b Allele frequency of the 1: 154574702T/G variant, (p.Gln139Pro), 1:154557848A/G variant, and 1:154574600T/C variant (p.Asn173Ser) in the Finnish (FIN) IBD (UC and CD) and non-IBD population as obtained through the IBD exomes browser (http://ibd.broadinstitute.org). Shown below are the carrier rates of different variants in a Finnish cohort. OR: odds ratio. c The structure of human ADAR and the Asn173 and Gln139 sites (right) predicted by AlphaFold. d The A-to-I RNA editing activity (left) and ADAR protein expression (right) in HCT116 cells transfected with shControl or shADARp110 with or without reconstitution by ectopic expression of WT human ADAR or ADARN173S (n = 3). e Sequence chromatograms of the GLI1 gene transcript in the indicated cell groups. The A to G editing efficiencies are labeled. Data are presented as the mean ± SEM. The statistical significance was analyzed using Fisher’s exact test (Fig. 1b). The remaining statistical differences were determined using unpaired two-tailed t-test. Source data are provided in the Source data file.
To determine whether ADARN173S affects its A-to-I RNA editing activity, we performed the reconstitution experiment in shADAR knockdown human colon cancer HCT116 cells using the expression vector of either the WT human ADAR or ADARN173S. Our results showed that reconstitution with ADARN173S exhibited lower RNA editing activity compared with the WT ADAR, as measured by the edited/WT RNA ratio (Fig. 1d) and Sanger sequencing (Fig. 1e) of the GLI1 gene, a well-established ADAR editing target21. The successful knockdown of the endogenous ADAR and reconstitutions of the WT and N173S mutant were confirmed by Western blotting. These results demonstrated that ADARN173S is a loss-of-function variant, suggesting that ADAR and its genetic variants are potentially involved in IBD.
The expression of ADAR is specifically decreased in the gut epithelium of IBD patients and mice
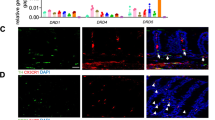
To define the cellular source of ADAR in the gut, we first analyzed a publicly available single-cell RNA sequencing (scRNA-seq) dataset derived from normal human gut (https://www.proteinatlas.org)22 and Crohn’s disease patients (https://singlecell.broadinstitute.org). Among the cell clusters in the normal human gut, the enterocytes, also known as simple columnar epithelial cells, had the highest expression of ADAR in both the normal small intestine (Supplementary Fig. 1a) and colon (Supplementary Fig. 1b). In the colon tissues of Crohn’s disease patients, the expression of ADAR is positively corrected with the expression of CD19 (a B cell marker), CD68 (a macrophage marker), and CD3D (a T cell marker) (Supplementary Fig. 1c). To determine the in-situ expression of ADAR, we analyzed a human IBD tissue array that includes 10 normal colonic tissues and 9 inflamed colitis (4 CD and 5 chronic colitis) by immunohistochemistry (Supplementary Table 2). The expression of ADAR was restricted to the epithelial area in the normal colon tissues (Fig. 2a). In IBD samples, the expression of ADAR decreased in crypts, but increased in the underneath non-epithelial cells (Fig. 2a). Consistent with the human results, the protein expression of Adar was decreased in crypts but increased in non-epithelial cells in mouse colitis induced by dextran sodium sulfate (DSS), as shown by immunofluorescence (Fig. 2b) and immunohistochemistry (Fig. 2c). The increased expression of ADAR in non-epithelial cells was likely secondary to the inflammatory response, because ADAR is an inflammatory responsive gene23.
a Representative IHC staining of ADAR in normal human colon (n = 10) and surgical specimens from patients with IBD (Crohn’s disease n = 4, chronic colitis n = 5). Red arrowheads indicate crypts. Shown right are the quantifications of relative ADAR intensity. Scale bars: 100 μm. Representative immunofluorescence (IF) (b) and IHC (c) staining of Adar in colon tissues of vehicle control mice or colitis mice induced by 6-day treatment with 2% DSS (n = 8 for Vehicle, n = 10 for DSS-6d). Red arrowheads indicate crypts. Shown on the right are the quantifications of relative Adar intensity. Scale bars: 100 μm. The statistical differences were determined using unpaired two-tailed t-test. Source data are provided in the Source data file.
Intestinal epithelial- or stem cell-specific ablation of ADAR triggers spontaneous bowel inflammation and compromises intestinal organoid growth
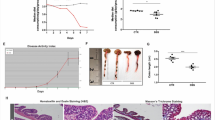
We then used Adar knockout mice to study the role of Adar in IBD. We have previously reported that the whole body Adar knockout mice were embryonically lethal20. In order to study the intestine-specific function of Adar in IBD, we generated tamoxifen (TMX)-inducible and gut epithelial cell-specific Adar knockout (AdariΔgut) mice by crossing Adarfl/fl mice with Villin-CreERT2 mice as outlined in Supplementary Fig. 2a. The efficiency of Adar knockout in the gut of AdariΔgut mice was confirmed by real-time PCR (Supplementary Fig. 2b), immunohistochemistry (Supplementary Fig. 2c), and decreased A-to-I RNA editing activity (Supplementary Fig. 2d) at 3 days post-induction (dpi). AdariΔgut mice exhibited marked lethality within 6 days of TMX induction (Fig. 3a). Gut ablation of Adar triggered spontaneous and severe bowel inflammation, as evidenced by shortened intestine and colon lengths (Fig. 3b), decreased body weight (Supplementary Fig. 2e), and increased colitis activity (Supplementary Fig. 2f). Mouse inflammatory antibody array showed that most of the inflammation factors were increased in the serum of AdariΔgut mice at 3 dpi, among which IFNγ had the highest induction (Fig. 3c and Supplementary Table 3). At the histologic level, AdariΔgut mice showed increased epithelial damage (Fig. 3d) and TUNEL staining (Fig. 3e) and decreased staining of the epithelial markers E-cadherin (Fig. 3f) and mucin 2 (Muc2) (Fig. 3g). Consistently, Western blotting showed that the protein expression of E-cadherin and Muc2 was decreased, whereas the protein expression of IFNγ, DNA damage marker γH2X, p-Stat1, and cleaved caspase3 was elevated in AdariΔgut mice (Fig. 3h).
a Survival curves of Adarfl/fl and AdariΔgut mice after tamoxifen induction and representative appearance of intestinal edema (insert) (n = 8). b Representative images of the gastrointestinal tract at 3 days post tamoxifen induction (3 dpi) with the quantifications of colon and small intestine length shown on the right (n = 6 for Adarfl/fl, n = 7 for AdariΔgut). c Dot blots of 40 inflammatory factors in the serum by mouse inflammation antibody array with the heatmap quantifications of dot intensity shown below (n = 3 mice combined serum in each group, and shown are duplicate results of each factor, array map information is provided in Supplementary Table 3). d H&E staining of intestinal sections at 3 dpi. Scale bars: 100 μm. Graphs below are histopathological scores and lengths of ileal crypts and villi (n = 5 for Adarfl/fl, n = 3 for AdariΔgut). e Representative images of TUNEL staining (green) on the ileal sections at 3 dpi. Scale bars: 100 μm. f Immunofluorescence of E-cadherin (red) staining at 3 dpi. Scale bars: 100 μm. g Immunofluorescence of Lysozyme (green) and Muc2 (red) staining at 3 dpi. Scale bars: 100 μm. h Western blot analysis of ileal samples at 3 dpi (n = 3). i Top hallmark categories of upregulated genes based on RNA-seq results derived from ileal crypts of Adarfl/fl and AdariΔgut mice at 3 dpi. j 3D PCoA score plot of different samples (n = 3). k Volcano plot of differentially expressed genes (Log2 (Fold change) > 1, P < 0.05), blue and red indicate downregulated and upregulated genes, respectively. The number of genes changed are shown in the pie chart (upper). l GSEA of the pathways related to type I IFN and IFNγ production. m Heatmap of the expression of inflammation-related genes from RNA-seq (n = 3). n Relative mRNA expression of inflammatory genes in the ileum measured by qRT-PCR (n = 3 for Adarfl/fl, n = 4 for AdariΔgut). o Representative images and Caspase 3/7 activity (green) and propidium iodide (PI, red) staining of organoids isolated from the ileum of Villin-CreERT2 Adarfl/fl mice and treated with or without 4-OHT (200 nM) at 6 day. Shown on the right are the quantifications of relative PI and Caspase 3/7 signals. n = 5, Scale bars: 100 μm.Data are presented as the mean ± SEM. The statistical significance was analyzed using Log-rank (Mantel-Cox) test (Fig. 3a). The remaining statistical differences were determined using unpaired two-tailed t-test. Source data are provided in the Source data file.
To determine the mechanism by which intestinal ablation of Adar triggered bowel inflammation, Hallmark pathway analysis was performed on our RNA-seq data derived from intestinal crypts of Adarfl/fl and AdariΔgut mice at 3 dpi. Transcriptome profiling showed the upregulation of inflammatory response and TNF, IL6, IFNα/β/γ related pathways (Fig. 3i and Supplementary Fig. 2g), as well as a good separation in 3D principal coordinate analysis (PCoA) plots (Fig. 3j) in AdariΔgut crypts. The volcano plot showed 1198 genes were downregulated and 1550 genes were upregulated in AdariΔgut crypts (Fig. 3k). GSEA analysis also showed robust upregulation of type I IFN response, IFNγ (type II IFN), IL-6, and TNF production in AdariΔgut crypts (Fig. 3l and Supplementary Fig. 2g). The activation of innate gut immunity in AdariΔgut mice was also manifested by increased mRNA expression of inflammatory and interferon-stimulated genes (ISG) by RNA-seq heatmap (Fig. 3m) and RT-qPCR (Fig. 3n and Supplementary Fig. 2h). IFNγ has been reported to causatively contribute to the pathogenesis of IBD24. Consistently, we found that treatment of AdariΔgut mice with IFNγ neutralizing antibody effectively attenuated mucosal epithelial damage induced by Adar ablation in vivo (Supplementary Fig. 2i), suggesting that the induction of IFNγ was mechanistically required for the inflammatory intestinal injury caused by Adar ablation.
Stem cell-derived intestinal organoid is a widely used model for IBD research25. We thus used intestinal organoids to further characterize the effects of Adar inhibition on intestinal growth and differentiation. Organoids isolated from AdariΔgut mice and treated with 4-OHT, or from WT mice and infected with adenovirus expressing shAdar failed to form a crypt-villus structure and exhibited cell death as shown by propidium iodide (PI) staining (Fig. 3o and Supplementary Fig. 2j). In an independent pharmacological model, organoids isolated from WT mice and treated with the ADAR inhibitor 8-azaadenosine (8-Aza)26 also showed inhibition of normally balanced growth and even cell death at a high dose (Supplementary Fig. 2k). GSEA of RNA-seq data showed a robust upregulation of apoptosis in AdariΔgut crypts (Fig. 3i and Supplementary Fig. 3a). Activation of the apoptosis pathway was further confirmed by increased expression of cleaved caspase 3 in the ileum of AdariΔgut mice (Fig. 3h) and increased caspase 3/7 activity in the AdariΔgut organoids (Fig. 3o). We then used cell programmed death inhibitors screen assay to define the mode of cell death in Adar deficient organoids. Our results showed that only the Rip1 inhibitor necrostatin-1 rescued Adar deficiency-induced organoid death, suggesting that the cell death was Rip1 dependent (Supplementary Fig. 3b). In contrast, treatment with the Caspase 8 inhibitor IETD-FMK, Caspase 3 inhibitor Z-DEVD-FMK, necroptosis: RIP3 inhibitor GSK-872, pan-caspase inhibitor Z-VAD-FMK, autophagy inhibitor Bafilomycin A1, or necrosis inhibitor IM-54 failed to rescue the cell death (Supplementary Fig. 3c).
Knowing the normal intestinal crypts had an abundant expression of Adar, we went on to generate intestinal stem cell (ISC)-specific Adar knockout (AdariΔISC) mice by crossing the Adarfl/fl mice with the Lgr5-CreERT2 mice27 (Supplementary Fig. 4a), and the knockout efficiency was confirmed by immunohistochemistry (Supplementary Fig. 4b). Similar but less severe spontaneous bowel inflammatory pathology was observed in AdariΔISC mice, as evidenced by animal death (Supplementary Fig. 4c), decreased body weight (Supplementary Fig. 4d), increased colitis activity (Supplementary Fig. 4e), and increased histologic epithelial damage in the ileum and colon (Supplementary Fig. 4f).
Adar deletion leads to the accumulation of endogenous retroviruses
ADAR regulates the sensing of self and non-self RNA, and loss-of-function of ADAR can cause AGS, a severe encephalopathy with IFN signature18,28. We used GO analysis of the RNA-seq data to comprehensively evaluate the intracellular environment in the intestinal crypts and found that multiple viral infections and antiviral response pathways were robustly upregulated in AdariΔgut crypts (Fig. 4a). Consistent with these findings, RNA-seq showed increases in the expression of multiple endogenous retroviruses (ERVs) in AdariΔgut crypts (Fig. 4b), which were further confirmed by qPCR (Fig. 4c). In the clinical samples, we also observed the upregulation of ERVs in patients with UC and CD through an analysis of a public database of repeat sequencing29,30 (Fig. 4d and Supplementary Table 4).
a GSEA of the pathways related to anti-virus from RNA-seq results derived from ileal crypts of Adarfl/fl and AdariΔgut mice (3 dpi). b RNA-seq results to show the level of ERVs in the ileal crypts of Adarfl/fl and AdariΔISC mice at 3dpi (n = 3). c Relative expression of ERV species in the ileum as determined by qRT-PCR (n = 3). d Biopsies from patients with IBD and unaffected controls were analyzed for virus mimicry repeats (analysis of a public database). Data are presented as the mean ± SEM. The statistical differences were determined using unpaired two-tailed t-test. Source data are provided in the Source data file.
MDA5-mediated dsRNA sensing is required for intestinal Adar deficiency-induced bowel inflammation and organoid death
MDA5 is a well-established dsRNA sensor, and Adar knockout neuronal progenitor cells exhibited MDA5-dependent spontaneous interferon production28,31. As expected, ablation of Adar accumulated dsRNA in organoids (Fig. 5a) and ileum (Fig. 5b) as shown by the immunofluorescence with the dsRNA specific J2 antibody. We then generated Mda5-deficient AdariΔgut (AdariΔgutMda5−/−) mice to determine whether Mda5 was required for Adar deficiency-triggered bowel inflammation. Mda5 knockout was verified by immunohistochemistry (Supplementary Fig. 5a) and qPCR (Supplementary Fig. 5b). Indeed, AdariΔgutMda5-/-mice were rescued from Adar deficiency induced animal death (Fig. 5c), body weight loss (Supplementary Fig. 5c), colon and intestine length shortening (Fig. 5d), mucosal epithelial damage (Fig. 5e), TUNEL staining increase (Fig. 5f), loss of E-cadherin and Muc2 expression (Fig. 5g–i), and intestinal apoptosis (Fig. 5i). Knockout of Mda5 also prevented organoid death and abnormal differentiation, and ERVs accumulation induced by Adar knockout (Fig. 5j and Supplementary Fig. 5d), Adar shRNA knockdown, or pharmacological inhibition of Adar (Supplementary Fig. 5e). Gene Expression Profiling Interactive Analysis web server analyses of the normal human colon tissues (GTEx) showed the expression of ADAR has a strong negative correlation with expression of MDA5 (Supplementary Fig. 5f). We also used the human colon cancer cell line SW480 to evaluate the human relevance the ADAR-MDA5 signaling. We showed that shRNA knockdown of ADAR inhibited the growth of SW480 as measured by Cell Counting Kit-8 (CCK8), but this effect was abolished when MDA5 was simultaneously knocked down by shRNA (Supplementary Fig. 5g). The knockdown efficiencies were verified by qRT-PCR (Supplementary Fig. 5h).
a Representative images of Adar (green) and J2 (red) staining in organoids isolated from the ileum of Villin-CreERT2 Adarfl/fl mice and treated with or without 4-OHT (200 nM) for 3 days.Scale bars: 100 μm. b Representative images of J2 staining (green) on ileal sections of Adarfl/fl and AdariΔgut mice (3 dpi). Scale bars: 100 μm. c Survival curves of Adarfl/fl, AdariΔgut, and AdariΔgutMda5−/−mice after tamoxifen induction (n = 5 for Adarfl/fl, n = 8 for AdariΔgut, n = 6 for AdariΔgutMda5−/−). d, h Adarfl/fl, AdariΔgut and AdariΔgutMda5−/− mice after tamoxifen induction at 3-dpi. d Small intestine and colon length (n = 5 for Adarfl/fl, n = 7 for AdariΔgut, n = 6 for AdariΔgutMda5−/−). e H&E staining of intestinal sections. Scale bars: 100 μm. Shown on the right are quantifications of histopathological scores, and lengths of ileal crypts and villi (n = 5). f Representative images of TUNEL staining (green) on the ileal sections. Scale bars: 100 μm. g Immunofluorescence of E-cadherin (red) staining. Scale bars: 100 μm. h Immunofluorescence of Lysozyme (green) and Muc2 (red) staining. Scale bars: 100 μm. i Western blot analysis of ileal samples (n = 3). j Representative images of the growth process of organoids isolated from the ileum of AdariΔgut and AdariΔgutMda5−/− mice treated with or without 4-OHT. Scale bars: 100 μm. Data are presented as the mean ± SEM. The statistical significance was analyzed using Log-rank (Mantel-Cox) test (Fig. 5c). The remaining statistical differences were determined using one-way ANOVA with multiple comparisons. Source data are provided in the Source data file.
Mda5 is a target of Adar-mediated RNA editing, which decreases its dsRNA sensing activity
Although ADAR-mediated A-to-I RNA editing is known to be essential for mRNA regulation, only a small number of ADAR RNA editing target genes have been experimentally defined. In order to mechanistically understand the role of ADAR-mediated RNA editing in bowel inflammation, we performed RNA editome analysis on our RNA-seq transcriptome data derived from crypts isolated from Adarfl/fl and AdariΔgut mice at 3 dpi. A total of 3460 A-to-I RNA editing sites were identified, which were defined by their decreased editing levels in AdariΔgut crypts compared to Adarfl/fl crypts. To explore the functional relevance of ADAR-mediated RNA editing, GSEA on these editing site-containing genes was performed. We found that the editing target RNA species were largely clustered in apoptosis and inflammation-related pathways, as shown by the lists of Hallmark (Fig. 6a) and KEGG gene sets (Fig. 6b). The large number of A-to-I editing sites allowed us to define common sequences important for editome function. The nucleotides 5′ and 3′ to the editing site (−1 and +1 positions) show a strong preference for G enrichment (Fig. 6c), consistent with a previous report32. The consensus motif identified by Homer in the ±50 nt neighborhood centered around each predicted A-to-I editing sites was shown in Fig. 6d. Figure 6e is the schematic illustration of the unbiased edited RNA species screen, using RNA editome analysis, GSEA, and RNA transcriptome profile to identify genes that participate in Adar deficiency facilitated bowel inflammation.
a, b The top 30 HALLMARK (A) and top 20 KEGG (B) enrichment analysis of A-to-I editing site containing genes from RNA-seq results derived from ileal crypts of Adarfl/fl and AdariΔgut mice (3 dpi). c Two sample logo (http://www.twosamplelogo.org) visualization of the differences between two sets of aligned samples of amino acids of A-to-I editing sites. Editing sites (the A nucleotide at position 0) are aligned together. d Consensus motif identified by Homer in the ±50 nt neighborhood centered around each of the predicted A-to-I editing sites (n = 3). e Strategy for combined RNA A-to-I editome and RNA transcriptome analysis. f RNA-seq at the Mda5 locus indicating positions of A-to-I editing site (62440874) from Adarfl/fl and AdariΔgut mice. The editing efficiency is labeled. g Domain structures of Mda5 and the alignment of sequences. h IFN-β reporter activity stimulated by flag-tagged WT Mda5 and mutant Mda5I480V in the absence or presence of poly (I:C) dsRNA (0, 250, 500 ng/mL) in HEK 293 T cells (n = 4). i Representative images of the organoids isolated from the ileum of AdariΔgutMda5−/− mice infected with lenti-Ctrl, lenti-Mda5, and lenti-Mda5I480V and then treated with 4-OHT at 6 days. Shown on the right are the quantifications of relative PI signals (n = 5). Scale bars: 100 μm. Data are presented as the mean ± SEM. The statistical significance was analyzed using unpaired two-tailed t-test (Fig. 6h) and one-way ANOVA with multiple comparisons (Fig. 6i). Source data are provided in the Source data file.
To our surprise, we found that Mda5 had an A-to-I editing site (genome location 62440874, exon 7: c.1438A to G, leading to the change of amino acid 480 from Ile (I) to Val (V)) with a high editing frequency (19.5%, or 8/41 reads) in the Adarfl/fl crypts, but decreased to 0% in the AdariΔgut crypts (Fig. 6f). The structure of the mouse Mda5 and its Ile480 site was predicted by AlphaFold (Supplementary Fig. 6a). The domain structures of Mda5 and the Ile480 sites and the surrounding sequences in different species are outlined in Fig. 6g. To define the functional relevance of the edited form of Mda5, we overexpressed Flag-tagged WT mouse Mda5, or its I480V mutant in HEK293T cells, and found the basal and Poly(I:C) stimulated dsRNA sensing activity of Mda5I480V was decreased in cells co-transfected with the Mda5-responsive IFNβ promoter-luciferase reporter gene, IFNβ-Luc (Fig. 6h). Mechanistically, the protein stability of Mda5 was decreased in cycloheximide-treated MC38 murine colon cancer cells infected with lentivirus expressing Flag-tagged Mda5I480V compared to its WT counterpart (Supplementary Fig. 6b). Moreover, lentiviral reconstitution of Mda5 led to the death of organoids isolated from AdariΔgut Mda5−/− mice, but this effect was blunted in AdariΔgut Mda5−/−organoids reconstituted with Mda5I480V (Fig. 6i and Supplementary Fig. 6c), suggesting that the edited form Mda5I480V had decreased dsRNA sensing activity.
Ruxolitinib attenuates Adar deficiency-induced organoid death and bowel inflammation
Canonical type I IFN signaling activates the Janus kinase (JAK)-signal transducer and activator of transcription (STAT) pathway, leading to the transactivation of IFN-stimulated genes33. The JAK inhibitors are used in the clinic to treat IBD34. GSEA of RNA-seq data showed robust upregulation of IFN and JAK-STAT signaling in AdariΔgut crypts (Fig. 3i and Supplementary Fig. 7a). Using the organoid screen system, we found the JAK1/2 dual inhibitor Ruxolitinib showed the best efficacy in rescuing Adar deficiency-induced organoid death, whereas the JAK1 inhibitor Itacitinib, JAK2 inhibitor Fedratinib, and JAK3/2 inhibitor Tofacitinib were not effective (Fig. 7a and Supplementary Fig. 7b). Consistently, Ruxolitinib, but not other JAK inhibitors, showed the best effect in preventing Adar ablation induced animal death (Fig. 7b), body weight loss (Supplementary Fig. 7c), ERVs accumulation (Supplementary Fig. 5d) and mucosal epithelial damage (Fig. 7c) in vivo. Treatment with Ruxolitinib also reversed intestinal TUNEL staining (Fig. 7d), loss of E-cadherin and Muc2 expression (Fig. 7e–g), and increased expression of cleaved caspase 3, p-Stat1, p-Stat3, and p-Stat6 in AdariΔgut mice (Fig. 7g and Supplementary Fig. 7d). These results suggested that JAK1 and JAK2 inhibitors may have synergistically prevented Adar ablation-induced organoid death and bowel inflammation. In the human colon cancer SW480 cells, the growth inhibitory effect of ADAR knockdown was also abolished when the ADAR knockdown cells were treated with Ruxolitinib (Supplementary Fig. 7e), consistent what was observed in the intestinal Adar knockout mice.
a Representative images and Caspase 3/7 activity (green) and PI (red) staining of organoids isolated from the ileum of Villin-CreERT2 Adarfl/fl mice and treated without or with 4-OHT (200 nM) and co-treated with 1 μM Ruxolitinib (Rux), Tofacitinib (Tof), Fedratinib (Fed), or Itacitinib (Ita) for 5 days. Shown on the right are the quantifications of relative PI and Caspase 3/7 signals. (n = 5). Scale bars: 100 μm. b Survival curves of Adarfl/fl and TMX induced AdariΔgut mice co-treated with Rux (60 mg/kg/day), Tof (40 mg/kg/day), Fed (60 mg/kg/day), or Ita (30 mg/kg/day) by daily i.p. injections for 5 days (n = 5 for Adarfl/fl, n = 8 for AdariΔgut, n = 5 for AdariΔgut+Rux, n = 4 for AdariΔgut+Tof, n = 7 for AdariΔgut+Fed, n = 4 for AdariΔgut+Ita). c H&E staining of intestine and colon sections. Scale bars: 100 μm. Shown on the right are quantifications of histopathological scores and on the lower left are quantifications of lengths of ileal crypts and villi (n = 5 for Adarfl/fl, n = 5 for AdariΔgut, n = 5 for AdariΔgut+Rux, n = 4 for AdariΔgut+Tof, n = 5 for AdariΔgut+Fed, n = 4 for AdariΔgut+Ita). PC proximal colon, DC distal colon. d Representative images of TUNEL staining (green) on the ileal sections. Scale bars: 100 μm. e Immunofluorescence of E-cadherin (red). Scale bars: 100 μm. f Immunofluorescence of Lysozyme (green) and Muc2 (red). Scale bars: 100 μm. G Western blot analysis of ileal samples. The relative values of protein expression are labeled (n = 3). Data are presented as the mean ± SEM. The statistical significance was analyzed using Log-rank (Mantel-Cox) test (Fig. 7b). The remaining statistical differences were determined using one-way ANOVA with multiple comparisons. Source data are provided in the Source data file.
Characterization of ADARN173S as a loss function mutation that may have triggered bowel inflammation in patients and mice
To confirm ADARN173S as a loss function mutation that may have triggered bowel inflammation in vivo, we reconstituted the WT human ADAR or ADARN173S in the gut of AdariΔgut mice for 5 weeks by infecting mice with adeno-associated virus 8 (AAV8) under the control of gut-specific GC-C promoter35, during which the mice were treated with TMX to induce the knockout of endogenous Adar, as outlined in Fig. 8a. The success of ADAR reconstitution was confirmed by immunohistochemistry (Fig. 8b). Compared with mice infected with AAV8-hADAR, mice infected with AAV8-hADARN173S showed exacerbated bowel inflammation induced by endogenous intestinal Adar knockout, as evidenced by higher incidence of animal death (Fig. 8c), accelerated body weight loss (Fig. 8d), aggravated mucosal epithelial damage (Fig. 8e), and accumulation of dsRNA in the ileum and colon tissues (Fig. 8f). These results suggested that ADARN173S is a loss-of-function mutation that likely contributes to the IBD incidence in human populations.
a The scheme of overexpressing WT human ADAR or ADARN173S in the gut of AdariΔgut mice by infecting the mice with adeno-associated viruses 8 (AAV8) for 5 weeks under the control of the gut-specific GC-C promoter and then treated with TMX to induce the knockout of the endogenous Adar. b Representative immunohistochemical staining of Adar in intestine and colon sections. Scale bars: 100 μm. c Survival curves after tamoxifen treatment (n = 11 for AAV-hADAR, n = 12 for AAV-hADARN173S). d Relative body weight (n = 9 for AAV-hADAR, n = 10 for AAV-hADARN173S). e H&E staining of intestine and colon sections. Scale bars: 100 μm. Shown below are the quantifications of histopathological scores, and lengths of ileal crypts and villi (n = 5 for AAV-hADAR, n = 4 for AAV-hADARN173S). f Representative images of J2 staining (green) on the ileum and colon sections. Scale bars: 100 μm. g Summary of ADAR loss-of-function triggered spontaneous IBD through the intestinal ADAR-ERVs/dsRNA-MDA5-IFNγ-JAK/STAT axis. Pharmacological inhibition of JAK1/2 by Ruxolitinib attenuated IBD in Adar-deficient mice. Summary model of ADAR in IBD created in BioRender. Xu, M. (2025) https://BioRender.com/ad9fbmq. Data are presented as the mean ± SEM. The statistical significance was analyzed using Gehan-Breslow-Wilcoxon test (Fig. 8c). The remaining statistical differences were determined using unpaired two-tailed t-test. Source data are provided in the Source data file.
The N173S mutation occurs in the Zα domain of ADAR. We recently reported that the human ADARP193A is also a Zα domain variant, but with a largely intact RNA editing activity36. We analyzed mice with the knock-in of the mouse AdarP195A, equivalent of the human ADARP193A, and found that the AdarP195A knock-in mice did not exhibit spontaneous inflammatory epithelial damage in the ileum and colon tissues (Supplementary Fig. 8a, b). These results strongly suggested that the pro-IBD effect of ADARN173S may have been accounted for by its loss of editing activity. Figure 8g summarizes our proposed model by which ADAR deficiency triggered spontaneous IBD through the intestinal ADAR-ERVs/dsRNA-MDA5-IFNγ-JAK/STAT axis.
Discussion
The current study investigated the critical role of ADAR in bowel inflammation. Although we previously reported that the whole-body Adar knockout mice exhibited compromised intestinal tissue homeostasis, the mechanism and human IBD relevance were not explored37. Knowing the role of ADAR in the tissue homeostasis of multiple organs38, the use of systemic Adar knockout mice was another major limitation in our previous report37, because the secondary effect cannot be ruled out. In the current study and through the use of two independent intestinal specific and tamoxifen inducible Adar knockout mouse models, we demonstrated that the intestinal Adar is essential for maintaining intestinal stem cell homeostasis, promoting differentiation, and ensuring survival. Gut-specific ablation or pharmacological inhibition of Adar led to impaired intestinal stem cell differentiation, reactivation of ERVs, and triggering of an innate immune response characterized by the upregulation of interferon signature genes, likely as a result of dsRNA accumulation and its sensing by MDA539.
We observed an increased level of γH2AX in the ileal tissues of AdariΔgut mice. γH2AX is a well-established marker of DNA damage and plays a critical role in checkpoint-mediated cell cycle arrest and DNA repair following double-strand breaks. The dsDNA-STING pathway has been reported to mediate the type I-IFN response40,41, but further investigations are necessary to determine whether the dsDNA-STING pathway is mechanistically involved in the inflammatory intestinal injury in response to Adar ablation. We are aware of that ADAR expression was reported to be upregulated in the entire colon of colitis-associated colorectal neoplasm (CAN) cases, but it was downregulated in non-CAN cases42. In the current study, we observed that ADAR expression in normal colon tissues was mainly restricted to the epithelium. In IBD samples, ADAR expression decreased in crypts but increased in the underlying non-epithelial cells. The discrepancies between these two studies may be explained by the distinct disease stages and tissue context. More specifically, while the study by Kazutaka and colleagues primarily investigated CAN cases42, our study was focused exclusively on non-CAN IBD.
Our observation that Ruxolitinib, a JAKs 1 and 2 dual inhibitor, was effective in attenuating Adar deficiency-induced bowel inflammation and mortality was clinically important. Cytokines play a critical role in maintaining intestinal homeostasis34. Several cytokines, including IFNγ, IL-12, IL-23, and IL-9, have been shown to act as mediators of pathological responses in UC and/or CD43. While cytokine-targeting therapies have revolutionized the treatment of IBD, they have predominantly focused on individual cytokines. There remains an unmet medical need for patients who fail to respond or develop resistance to cytokine-targeting treatment. Knowing that many of these cytokines converge on the JAK-STAT signaling pathway, JAKs, including JAK1, JAK2, JAK3, and tyrosine kinase 2 (TYK2), have been explored as therapeutic targets, as their blockade has the potential to disrupt the signaling of over 50 cytokines44. Clinical development is underway for several small-molecule inhibitors of JAKs that possess the ability to modulate multiple pro-inflammatory cytokine-dependent pathways. Tofacitinib, a JAK3/2 inhibitor, has received approval for the treatment of UC45, while other agents have shown promising efficacy in early phase clinical trials46,47. Our organoid screen and in vivo drug treatment demonstrated that Ruxolitinib exhibited the best efficacy in attenuating Adar deficiency-induced bowel inflammation and mortality in mice. Ruxolitinib was the first JAK inhibitor FDA approved for myelofibrosis, polycythemia vera, graft-versus-host disease, atopic dermatitis, and vitiligo48,49,50,51,52. The lack of a rescue effect of Tofacitinib in our mouse models and mouse intestinal organoids was a surprise, considering that this drug has been approved for the treatment of UC45. Nevertheless, our findings suggested that simultaneous inhibition of JAK1 and JAK2 is an effective approach to mitigate ADAR deficiency-induced bowel inflammation. The future uses of human IBD patient samples or organoids derived from IBD patients are necessary to determine whether a reduced ADAR expression may influence patient’s responses to JAK inhibitor treatment.
Our finding that MDA5 was a target of ADAR-mediated RNA editing is intriguing. MDA5 is a RIG-I-like receptor and functions as a dsRNA helicase. MDA5 can recognize long dsRNA, the genomic RNA of dsRNA viruses, and replicative intermediates of both positive-sense and negative-sense RNA viruses53. ADAR plays a specific and crucial role as a negative regulator of the MDA5-MAVS RNA sensing pathway31. Mutations in ADAR are responsible for the autoimmune disease AGS and MDA5-dependent encephalopathy with IFN pathway activation in the brain54,55. ADAR has been reported to interact with Z-RNA to promote the editing of endogenous dsRNA and prevent MDA5-dependent immune activation56. MDA5 is a critical gene implicated in immune-related diseases, and gain-of-function mutations in MDA5 lead to a spectrum of autoimmune and undefined neurological and immunological phenotypes characterized by upregulated type I interferon signaling19. Here, we found the expression of MDA5 was upregulated in AdariΔgut mice and MDA5 was required for intestinal Adar deficiency-induced bowel inflammation and death of intestinal organoids and mice. We found the edited form of Mda5, Mda5I480V is a loss-of-function variant, suggesting that Mda5 editing is a feedforward mechanism to prevent dsRNA-induced innate immune response. It should be noted that although our animal data revealed a regulatory link between ADAR and MDA5, which was supported by our cell culture results using the human colon cancer SW480 cells, further studies using human IBD patient samples or organoids are necessary to confirm whether MDA5 is a direct target of ADAR-mediated RNA editing in humans.
Our most exciting and clinically relevant finding is the characterization of the loss-of-function ADARN173S variant, a SNP associated with IBD incidence in a Finnish cohort that contains 11389 individuals. There were 89 heterozygotes and 1 homozygous ADARN173S mutation in this cohort, so the carrier rate is approximately 1 in 126. The N173S variant in the Zα domain of ADAR has been identified as crucial for its interaction with both Z-DNA and Z-RNA57,58, consistent with the loss-of-function phenotype of ADARN173S. At the functional level, intestinal reconstitution of ADARN173S in AdariΔgut mice was less efficient than the WT ADAR in rescuing the IBD phenotypes. The loss-of-function nature of ADARN173S was consistent with our observations that the expression of ADAR was decreased in the crypts of IBD patients and mice, and inhibition of ADAR was sufficient to cause spontaneous bowel inflammation. In contrast, the Zα domain variant ADARP193A with an intact RNA editing activity36 was not associated with IBD in patients, nor did it cause IBD phenotype when knocked into the mice. We acknowledge that although our findings in human participants suggest an association between these variants and IBD, the current results have their limitations, such as limited patient numbers in the control and patient cohorts, and the lack of access to tissue samples from the variant carriers for functional validations. Future, more comprehensive in vivo functional characterization of these variants in animal models and case-control studies in human participants is necessary to fully establish the pathogenic and mechanistic roles these ADAR variants in IBD.
In summary, our study has revealed the critical role of ADAR-mediated RNA A-to-I editing in maintaining intestinal stem cell homeostasis and preventing bowel inflammation. We propose that the expression and/or genetic variation can be diagnostic markers for IBD. The intestinal ADAR-ERVs/dsRNA-MDA5-JAK/STAT pathway represents a fundamental mechanism in ADAR deficiency triggered spontaneous IBD, which offers opportunities for clinical interventions.
Methods
Human patient datasets and human cohorts, and scRNA-seq analyses
Human normal colon and IBD tissue arrays were purchased from US Biomax (Derwood, MD) with appropriate ethics and consent and used for H&E and immunohistochemical staining. Details of the patient information are shown in Supplementary Table 2. For human intestine and colon scRNA-seq analyses, the human small intestine and colon dataset was downloaded from the Human Protein Atlas browser (https://www.proteinatlas.org) and human Crohn Disease (https://singlecell.broadinstitute.org) to identify cell clusters and ADAR gene expression in different cell lineages.
Experimental animals
The use of mice in this study adhered to the guidelines set by the University of Pittsburgh Institutional Animal Care and Use Committee (Protocol numbers: 19044749 and 22030802). Mice were housed at 25 °C in a 12-h light-dark cycle in the animal facility at the University of Pittsburgh. Food and water were available ad libitum. Villin-CreERT2 mice (Strain #: 020282), Lgr5-CreERT2 mice (Strain #: 008875), and Mda5−/− mice (Strain #: 015812) were obtained from Jackson Laboratory (Ronkonkoma, NY). Adarfl/fl mice in the C57BL/6 genetic background were custom-made and described previously59. Adarfl/fl mice were crossed with Villin-CreERT2 mice to create tamoxifen (TMX)-inducible gut-specific Adar knockout mice (AdariΔgut) and then crossed with Mda5−/− mice to generate gut epithelial-specific Adar deficiency and Mda5-knockout (AdariΔgut Mda5−/−) mice. Adarfl/fl mice were crossed with Lgr5-CreERT2 mice to create TMX-inducible intestinal stem cell-specific Adar knockout mice (AdariΔISC). Adeno-associated viruses 8 (AAV8) expressing WT-hADAR or hADARN173S under the control of the gut-specific GC-C promoter35 were custom-made by Vector Biolabs (Malvern, PA). When necessary, AdariΔgut mice were given a tail-vein injection of AAV-hADAR or AAV-hADARN173S at the dose of 1 × 1011 viral particles/mouse 5 weeks before TMX-induced knockout of the endogenous Adar. To induce Adar knockout, 8-week-old AdariΔgut, AdariΔISC, and AdariΔgutMda5−/− mice received five consecutive intraperitoneal (i.p.) injections of 50 mg/kg/day TMX in corn oil. TMX-treated Adarfl/fl littermates were used as the controls. Mice were administered with Ruxolitinib (Rux, 60 mg/kg/day), Tofacitinib (Tof, 40 mg/kg/day), Fedratinib (Fed, 60 mg/kg/day), or Itacitinib (Ita, 30 mg/kg/day) by daily i.p. injections 30 min before TMX injection for 5 days. All chemicals were purchased from MedChemExpress (Monmouth Junction, NJ). When necessary, Adarfl/fl and AdariΔgut mice received intraperitoneal injections of IFNγ neutralizing antibody or control antibody (150 µg/mouse) every other day, starting on the same day as TMX treatment. Sample sizes for mouse experiments were determined empirically, and mice were randomly assigned to either the control or experimental groups. Blinding was not required for the mouse experiments conducted in this study. The mouse model of bowel inflammation used in this study did not exhibit a sex preference.
Intestinal crypt isolation, organoid culture, drug treatment, and lentiviral transduction
Intestinal crypts were isolated and cultured following a previous protocol with minor modifications60. Briefly, the ileums were washed in ice-cold PBS, sliced into 1–2 mm segments, and subjected to 15–20 washes in PBS to remove impurities. Subsequently, Gentle Cell Dissociation Reagent (GCDR, cat #7174, Stem Cell Technologies, MA) was applied following the manufacturer’s instructions. For intestinal organoid culture, the crypts were counted, embedded in Matrigel, and maintained in IntestiCult organoid growth medium (cat #6005, Stem Cell Technologies, MA). When applicable, 4-hydroxytamoxifen (4-OHT, 200 nM), Rux (1 μM), Tof (1 μM), Fed (1 μM), Ita (1 μM), 8-Azaadenosine (8Aza, 1, 2 μM), Necrostatin-1 (RIP1i, 30 μM), IETD-FMK (Cas8i, 10 μM), Z-DEVD-FMK (Cas3i, 10 μM), GSK-872 (RIP3i, 50 μM), Z-VAD-FMK (Z-VAD, 40 μM), Bafilomycin A1 (BafA1, 100 μM), or IM-54 (10 μM) were added to the medium for a duration of 3–5 days, with medium changes every 2 days. For organoid lentiviral transduction, the organoids were harvested and transferred into 15 mL tubes. After centrifugation, the supernatant was discarded, and lentivirus containing the vector (lenti-Ctrl), WT-Mda5 (lenti-Mda5), or Mda5I480V (lenti-Mda5I480V) (1 × 106 PFU/100 crypts) was introduced. The organoid-virus mixture was incubated in a 37 °C incubator for 1 h to facilitate transduction. Subsequently, the organoids were plated and cultured following the previously mentioned procedure. To analyze the dead cells, organoids were incubated with 4 μM of the CellEvent Caspase-3/7 Detection Reagents (cat #C10423, Thermo Fisher Scientific, Waltham, MA) for 30 min following the manufacturer’s instructions. During the final 5 min of incubation, PI (Sigma Aldrich, St. Louis, MO) was added directly to the wells at the concentration of 50 μg/mL. Images of the organoids were captured under the bright field.
Transcriptome RNA sequencing and RNA-seq for ERVs analysis
Ileal crypts were isolated, and total RNA was extracted using the RNeasy Mini Kit before being used for library preparation. RNA-seq was performed at the Health Sciences Sequencing Core at the Children’s Hospital of Pittsburgh, and the results were analyzed as previously described61. For ERVs analysis, aligned reads were analyzed using RepEnrich2 with the Repeat Library 20140131 Annotation. EdgeR (version 3.26.839) was utilized to identify differentially expressed repetitive elements62.
RNA-seq for A-to-I editome analysis
The RNA A-to-I editome analysis was custom-conducted by Yanke Biotechnology Co., Ltd (Guangzhou, China). In brief, RNA editing sites were identified using REDItools. Trimmed and filtered RNA-seq reads were aligned to the Mus_musculus reference genome (GRCm39) using BWA. Two sample logos were used to visualize differences between two sets of aligned samples of amino acids at A-to-I editing sites. Consensus motifs were identified by Homer in the ±50 nt neighborhood centered around each predicted A-to-I editing site.
Cytokine measurement
Serum samples were collected for cytokine measurements. The semi-quantitative detection of 40 types of mouse inflammatory cytokines was performed using c-series mouse inflammation antibody arrays from RayBiotech (Peachtree Corners, GA). Inflammatory antibody array map information is provided in Supplementary Table 3. The dot blots were quantified using the NIH Image J software.
Quantitative real-time PCR and RNA editing activity assay
Total RNA was extracted using the Trizol reagent from Invitrogen (Carlsbad, CA). Two micrograms of total RNA were reverse transcribed into cDNA using a kit from Thermo Fisher Scientific (Waltham, MA). Real-time PCR was performed using SYBR Green reagent on the QuantStudio 6 Flex Real-Time PCR System from Applied Biosystems, following previously described protocols60. Data analysis was conducted using the comparative ΔΔCT method, with Cyclophilin or Gapdh as housekeeping gene controls. The relative RNA editing activity (edit/WT RNA ratio of Azini or Gli1 gene) was calculated using the formula 2-(CtEdit – CtWT) as previously reported21. The real-time PCR primer sequences are listed in Supplementary Table 5.
Immunofluorescence, immunohistochemistry, and Western blot analysis
Paraffin sections were subjected to deparaffinization and rehydration. Immunofluorescence (IF) was performed using antibodies against Adar (GeneTex), E-cadherin, dsRNA(J2), Lysozyme, and Muc2. For immunohistochemistry, ADAR and Mda5 antibodies were used and incubated overnight at 4 °C. Fluorescent secondary antibodies or the VECTASTAIN ABC Kit were utilized, and the DAB Peroxidase Substrate Kit from Vector Laboratories was employed for visualization of the staining. The percentage of positive area per mouse was determined by quantifying the positive stain areas using Image J software. Tissue protein samples for Western blot analysis were extracted using RIPA lysis buffer supplemented with a protease inhibitor cocktail. The membranes were incubated with primary antibodies, including β-actin, Adar (Santa Cruz, CA), Caspase3, Cleaved Caspase 3, E-cadherin, γH2X, IFNγ, Lysozyme, Muc2, p-Stat1, Stat1, p-Stat3, Stat3, p-Stat6, and Stat6 followed by incubation with appropriate secondary antibodies. Detailed antibody information is provided in Supplementary Table 6.
Macroscopic and histological analysis
Colon and intestine tissues were rinsed with phosphate-buffered saline (PBS) and subsequently fixed overnight in 10% formalin. After paraffin embedding, tissue sections of 4–5 μm thickness were prepared using a microtome. Subsequently, the sections were deparaffinized, rehydrated, and subjected to histological analysis by H&E staining. Assessment of IBD disease severity and histological scoring was performed as previously described60. For TUNEL staining, the sections were stained with the In Situ Cell Death Detection Kit from Roche. Images were acquired using an all-in-one fluorescence microscope (BZ-X810) from KEYENCE (Itasca, IL).
Plasmid construction, cell transfection, and luciferase reporter gene assays
The pGenlenti-flag-Mda5 (#160110) and pmGFP-ADAR (#117927) expression vectors were obtained from GenScript (Piscataway, NJ) and Addgene (Watertown, MA), respectively. The pLKO-shCtrl, pLKO-shADAR, and pLKO-shMDA5 vectors were obtained from Sigma-Aldrich. Mutations of Mda5I480V and ADARN173S were performed using the QuikChange Lightning Site-Directed Mutagenesis Kit from Agilent Technologies (Santa Clara, CA). All cloned sequences were verified by DNA sequencing at GENEWIZ (South Plainfield, NJ). Lentiviral particle packaging and transduction were performed as previously reported63. The MDA5-responsive luciferase reporter plasmid pGL3-IFNβ-luc (#102597) was acquired from Addgene. Transfection and luciferase reporter gene assay were performed using 293T cells, as previously described64. The relative cell growth was measured by the Cell Counting Kit-8 (CCK-8) (HY-K0301) from MedChemExpress.
Statistics
Statistical analysis was performed using Prism GraphPad 9.0 software (San Diego, CA). Experimental data were presented as either means ± SEM or violin plots. Statistical significance between two groups was assessed using a two-tailed t-test. Multiple comparisons were performed using one-way ANOVA with Tukey’s test. Survival curves were plotted using the Kaplan-Meier method, and statistical differences were determined using either the Log-Rank test or the Gehan–Breslow–Wilcoxon test. A P value of less than 0.05 was considered statistically significant. At least three independent experiments were repeated with similar results.
See Supplementary Materials and Methods. Information on patients, inflammatory antibody array, primer sequences, and antibodies is provided in Supplementary Tables 2, 3, 5, and 6, respectively.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The RNA-seq datasets generated in this study have been deposited in GEO database under accession code GSE290331. The remaining data generated in this study are available in the main text or the supplementary materials or from the corresponding author on request. Source data are provided with this paper.
References
Jairath, V. & Feagan, B. G. Global burden of inflammatory bowel disease. Lancet Gastroenterol. Hepatol. 5, 2–3 (2020).
Collaborators, G. B. D. I. B. D. The global, regional, and national burden of inflammatory bowel disease in 195 countries and territories, 1990-2017: a systematic analysis for the Global Burden of Disease Study 2017. Lancet Gastroenterol. Hepatol. 5, 17–30 (2020).
Qiu, Y. et al. Disease burden of inflammatory bowel disease in China from 1990 to 2017: findings from the global burden of diseases 2017. EClinicalMedicine 27, 100544 (2020).
Baumgart, D. C. & Carding, S. R. Inflammatory bowel disease: cause and immunobiology. Lancet 369, 1627–1640 (2007).
Na, Y. R., Stakenborg, M., Seok, S. H. & Matteoli, G. Macrophages in intestinal inflammation and resolution: a potential therapeutic target in IBD. Nat. Rev. Gastroenterol. Hepatol. 16, 531–543 (2019).
Bisgaard, T. H., Allin, K. H., Keefer, L., Ananthakrishnan, A. N. & Jess, T. Depression and anxiety in inflammatory bowel disease: epidemiology, mechanisms and treatment. Nat. Rev. Gastroenterol. Hepatol. 19, 717–726 (2022).
Patankar, J. V. et al. E-type prostanoid receptor 4 drives resolution of intestinal inflammation by blocking epithelial necroptosis. Nat. Cell Biol. 23, 796–807 (2021).
Ananthakrishnan, A. N. et al. Lifestyle, behaviour, and environmental modification for the management of patients with inflammatory bowel diseases: an International Organization for Study of Inflammatory Bowel Diseases consensus. Lancet Gastroenterol. Hepatol. 7, 666–678 (2022).
Liu, Z. et al. Genetic architecture of the inflammatory bowel diseases across East Asian and European ancestries. Nat. Genet. 55, 796–806 (2023).
Bengtson, M. B., Aamodt, G., Vatn, M. H. & Harris, J. R. Concordance for IBD among twins compared to ordinary siblings-a Norwegian population-based study. J. Crohns Colitis 4, 312–318 (2010).
Noble, A. J., Nowak, J. K., Adams, A. T., Uhlig, H. H. & Satsangi, J. Defining interactions between the genome, epigenome, and the environment in inflammatory bowel disease: progress and prospects. Gastroenterology 165, 44–60 (2023).
Jostins, L. et al. Host-microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491, 119–124 (2012).
Sazonovs, A. et al. Large-scale sequencing identifies multiple genes and rare variants associated with Crohn’s disease susceptibility. Nat. Genet. 54, 1275–1283 (2022).
Nishikura, K. Functions and regulation of RNA editing by ADAR deaminases. Annu. Rev. Biochem. 79, 321–349 (2010).
Hao, X. et al. ADAR1 downregulation by autophagy drives senescence independently of RNA editing by enhancing p16(INK4a) levels. Nat. Cell Biol. 24, 1202–1210 (2022).
Mannion, N. M. et al. The RNA-editing enzyme ADAR1 controls innate immune responses to RNA. Cell Rep. 9, 1482–1494 (2014).
Li, Q. et al. RNA editing underlies genetic risk of common inflammatory diseases. Nature 608, 569–577 (2022).
Rice, G. I. et al. Mutations in ADAR1 cause Aicardi-Goutieres syndrome associated with a type I interferon signature. Nat. Genet. 44, 1243–1248 (2012).
Rice, G. I. et al. Gain-of-function mutations in IFIH1 cause a spectrum of human disease phenotypes associated with upregulated type I interferon signaling. Nat. Genet. 46, 503–509 (2014).
Wang, Q., Khillan, J., Gadue, P. & Nishikura, K. Requirement of the RNA editing deaminase ADAR1 gene for embryonic erythropoiesis. Science 290, 1765–1768 (2000).
Crews, L. A. et al. An RNA editing fingerprint of cancer stem cell reprogramming. J. Transl. Med. 13, 52 (2015).
Karlsson, M. et al. A single-cell type transcriptomics map of human tissues. Sci. Adv. 7, eabh2169 (2021).
de Reuver, R. et al. ADAR1 prevents autoinflammation by suppressing spontaneous ZBP1 activation. Nature 607, 784–789 (2022).
Ito, R. et al. Interferon-gamma is causatively involved in experimental inflammatory bowel disease in mice. Clin. Exp. Immunol. 146, 330–338 (2006).
Angus, H. C. K., Butt, A. G., Schultz, M. & Kemp, R. A. Intestinal organoids as a tool for inflammatory bowel disease research. Front. Med. 6, 334 (2019).
Veliz, E. A., Easterwood, L. M. & Beal, P. A. Substrate analogues for an RNA-editing adenosine deaminase: mechanistic investigation and inhibitor design. J. Am. Chem. Soc. 125, 10867–10876 (2003).
Ishikawa, K. et al. Identification of quiescent LGR5(+) stem cells in the human colon. Gastroenterology 163, 1391–1406.e1324 (2022).
Chung, H. et al. Human ADAR1 prevents endogenous RNA from triggering translational shutdown. Cell 172, 811–824.e814 (2018).
Wang, W. et al. Metagenomic analysis of microbiome in colon tissue from subjects with inflammatory bowel diseases reveals interplay of viruses and bacteria. Inflamm. Bowel Dis. 21, 1419–1427 (2015).
Wang, R. et al. Gut stem cell necroptosis by genome instability triggers bowel inflammation. Nature 580, 386–390 (2020).
Pestal, K. et al. Isoforms of RNA-editing enzyme ADAR1 independently control nucleic acid sensor MDA5-driven autoimmunity and multi-organ development. Immunity 43, 933–944 (2015).
Bahn, J. H. et al. Accurate identification of A-to-I RNA editing in human by transcriptome sequencing. Genome Res. 22, 142–150 (2012).
Ivashkiv, L. B. & Donlin, L. T. Regulation of type I interferon responses. Nat. Rev. Immunol. 14, 36–49 (2014).
Salas, A. et al. JAK-STAT pathway targeting for the treatment of inflammatory bowel disease. Nat. Rev. Gastroenterol. Hepatol. 17, 323–337 (2020).
Park, J., Schulz, S. & Waldman, S. A. Intestine-specific activity of the human guanylyl cyclase C promoter is regulated by Cdx2. Gastroenterology 119, 89–96 (2000).
Guo, X. et al. ADAR1 Zalpha domain P195A mutation activates the MDA5-dependent RNA-sensing signaling pathway in brain without decreasing overall RNA editing. Cell Rep. 42, 112733 (2023).
Qiu, W. et al. ADAR1 is essential for intestinal homeostasis and stem cell maintenance. Cell Death Dis. 4, e599 (2013).
Song, B., Shiromoto, Y., Minakuchi, M. & Nishikura, K. The role of RNA editing enzyme ADAR1 in human disease. Wiley Interdiscip. Rev. RNA 13, e1665 (2022).
Fischer, S. E. J. & Ruvkun, G. Caenorhabditis elegans ADAR editing and the ERI-6/7/MOV10 RNAi pathway silence endogenous viral elements and LTR retrotransposons. Proc. Natl. Acad. Sci. USA 117, 5987–5996 (2020).
Saitoh, T. et al. Atg9a controls dsDNA-driven dynamic translocation of STING and the innate immune response. Proc. Natl. Acad. Sci. USA 106, 20842–20846 (2009).
AlDaif, B. A., Mercer, A. A. & Fleming, S. B. The parapoxvirus Orf virus inhibits dsDNA-mediated type I IFN expression via STING-dependent and STING-independent signalling pathways. J. Gen. Virol. 104, 001912 (2023).
Takahashi, K. et al. RNA editing is a valuable biomarker for predicting carcinogenesis in ulcerative colitis. J. Crohns Colitis 17, 754–766 (2023).
Villarino, A. V., Kanno, Y. & O’Shea, J. J. Mechanisms and consequences of Jak-STAT signaling in the immune system. Nat. Immunol. 18, 374–384 (2017).
Cordes, F., Foell, D., Ding, J. N., Varga, G. & Bettenworth, D. Differential regulation of JAK/STAT-signaling in patients with ulcerative colitis and Crohn’s disease. World J. Gastroenterol. 26, 4055–4075 (2020).
Sandborn, W. J. et al. Tofacitinib as induction and maintenance therapy for ulcerative colitis. N. Engl. J. Med. 376, 1723–1736 (2017).
Sandborn, W. J. et al. Efficacy and safety of upadacitinib in a randomized trial of patients with Crohn’s disease. Gastroenterology 158, 2123–2138.e2128 (2020).
Dudek, P., Fabisiak, A., Zatorski, H., Malecka-Wojciesko, E. & Talar-Wojnarowska, R. Efficacy, safety and future perspectives of JAK inhibitors in the IBD treatment. J. Clin. Med. 10, 5660 (2021).
Verstovsek, S. et al. A double-blind, placebo-controlled trial of ruxolitinib for myelofibrosis. N. Engl. J. Med. 366, 799–807 (2012).
Kim, B. S. et al. Treatment of atopic dermatitis with ruxolitinib cream (JAK1/JAK2 inhibitor) or triamcinolone cream. J. Allergy Clin. Immunol. 145, 572–582 (2020).
Uppal, S. K., Kearns, D. G., Chat, V. S. & Wu, J. J. Ruxolitinib cream for the treatment of vitiligo. Lancet 396, 1735–1736 (2020).
McLornan, D. P., Pope, J. E., Gotlib, J. & Harrison, C. N. Current and future status of JAK inhibitors. Lancet 398, 803–816 (2021).
Vannucchi, A. M. et al. Ruxolitinib versus standard therapy for the treatment of polycythemia vera. N. Engl. J. Med. 372, 426–435 (2015).
Wu, B. et al. Structural basis for dsRNA recognition, filament formation, and antiviral signal activation by MDA5. Cell 152, 276–289 (2013).
Guo, X. et al. An AGS-associated mutation in ADAR1 catalytic domain results in early-onset and MDA5-dependent encephalopathy with IFN pathway activation in the brain. J. Neuroinflamm. 19, 285 (2022).
Nakahama, T. et al. Mutations in the adenosine deaminase ADAR1 that prevent endogenous Z-RNA binding induce Aicardi-Goutieres-syndrome-like encephalopathy. Immunity 54, 1976–1988 e1977 (2021).
de Reuver, R. et al. ADAR1 interaction with Z-RNA promotes editing of endogenous double-stranded RNA and prevents MDA5-dependent immune activation. Cell Rep. 36, 109500 (2021).
Schade, M., Turner, C. J., Lowenhaupt, K., Rich, A. & Herbert, A. Structure-function analysis of the Z-DNA-binding domain Zalpha of dsRNA adenosine deaminase type I reveals similarity to the (alpha + beta) family of helix-turn-helix proteins. EMBO J. 18, 470–479 (1999).
Herbert, A. Mendelian disease caused by variants affecting recognition of Z-DNA and Z-RNA by the Zalpha domain of the double-stranded RNA editing enzyme ADAR. Eur. J. Hum. Genet. 28, 114–117 (2020).
Wang, Q. et al. Stress-induced apoptosis associated with null mutation of ADAR1 RNA editing deaminase gene. J. Biol. Chem. 279, 4952–4961 (2004).
Xu, P. et al. Intestinal sulfation is essential to protect against colitis and colonic carcinogenesis. Gastroenterology 161, 271–286 e211 (2021).
Xi, Y. et al. The anti-fibrotic drug pirfenidone inhibits liver fibrosis by targeting the small oxidoreductase glutaredoxin-1. Sci. Adv. 7, eabg9241 (2021).
Juznic, L. et al. SETDB1 is required for intestinal epithelial differentiation and the prevention of intestinal inflammation. Gut 70, 485–498 (2021).
Cai, X. et al. Inhibition of the SLC35B2-TPST2 axis of tyrosine sulfation attenuates the growth and metastasis of pancreatic ductal adenocarcinom. Cell Mol. Gastroenterol. Hepatol. 16, 473–495 (2023).
Xu, P. et al. Inhibition of p53 sulfoconjugation prevents oxidative hepatotoxicity and acute liver failure. Gastroenterology 162, 1226–1241 (2022).
Acknowledgements
The authors would like to acknowledge NIH shared instrumentation grant S10OD030254. This work was supported by NIH grants DK135538 and ES030429 (to WX) and a Pilot & Feasibility grant (to PX) from the Pittsburgh Liver Research Center funded by NIH grant P30DK120531. This work is dedicated to the memory of Dr. Qingde Wang.
Author information
Authors and Affiliations
Contributions
W.X. conceived and mentored this research. W.X. and P.X. designed the study. P.X., Y.X., J.W.K., M.Z., C.G., Y.W., and X.W. performed the experiments and analyzed the data. S.R., M.X., Q.W., and D.Y. gave technical support and conceptual advice. P.X. and W.X. wrote the manuscript. All authors approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Konrad Aden and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xu, P., Xi, Y., Kim, JW. et al. A loss-of-function human ADAR variant activates innate immune response and promotes bowel inflammation. Nat Commun 16, 8560 (2025). https://doi.org/10.1038/s41467-025-63554-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-63554-4