Abstract
Replication stress (RS), if not effectively and timely addressed, could result in DNA damage in mitosis. However, the relationship between RS and other mitotic events, such as nuclear envelope (NE) breakdown and reassembly, remains poorly understood. Here we report that RS can lead to NE defect. Importantly, rather than de novo NE rupture, the defect per se is a result of nuclear envelope reassembly defect (NERD) during mitosis. Interestingly, NERD is associated with mitotic DNA damage, and repair of the damage by DNA polymerase theta (Polθ)-mediated end joining (TMEJ) ameliorates NERD. Genomic mapping of lamina associated domains (LADs) by cleavage under targets and tagmentation (CUT&Tag) identifies a population of replication stress-sensitive LADs (RESSLADs). Strikingly, a substantial portion of RESSLADs reside in the common fragile sites (CFSs). The loss of RESSLADs-NE interaction under RS might be attributed to the sustained phosphorylation of Lamin A/C at the sites of NERD. In addition, prominent NE defect is observed under multiple conditions of synthetic lethality. Altogether, these findings establish a link between genome instability and nuclear vulnerability under replication stress.
Similar content being viewed by others
Introduction
DNA replication in the S phase and chromosome segregation during mitosis are the two main events of the cell cycle. In normal cells, these two processes are highly regulated and temporally separated, making them incompatible. However, under replication stress (RS) caused by oncogene activation, exposure to DNA damaging agents, or nucleotide depletion1,2,3, perturbed replication originating in the S phase can persist into mitosis and lead to detrimental outcomes such as chromosome pulverization, micronucleation, and the formation of chromosome bridges. These events are often accompanied by replication fork collapse, DNA double-strand breaks (DSBs), and catastrophic genome rearrangements including chromothripsis and kataegis4,5,6. The underlying mechanisms, however, remain largely unknown.
One outstanding question is how RS and the resulting DSBs are addressed during mitosis. During interphase, DSBs can be repaired by homologous recombination (HR) and non-homologous end-joining (NHEJ). However, these canonical pathways are largely inactive in mitosis7. Two models have been proposed to solve the “replication problem” during mitosis. First, entry of replication forks into prophase activates mitotic DNA synthesis (MiDAS), which utilizes break-induced replication (BIR) pathway to process the replication forks and trigger conservative replication. Multiple enzymes, including MUS81-EME1, SLX4, POLD3 and RAD52, have been reported to play important roles in MiDAS8,9. Second, DNA polymerase theta (Polθ, encoded by the POLQ gene) mediated end joining (TMEJ) joins the DSB ends from collapsed replication forks during premature mitosis4. Additionally, it repairs chromosome breaks at common fragile sites (CFSs)10 and resolves the replication-blocking agents during mitosis11,12,13,14, albeit in an error-prone manner. A recent study also demonstrates that Polθ/TMEJ but not MiDAS mediates RS-induced structural variant formation throughout CFSs during mitosis14. Although these pathways operate differently, MiDAS and TMEJ are not necessarily mutually exclusive; together they act as a final safeguard to prevent chromosome instability before the birth of daughter cells.
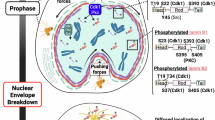
In addition to chromosome segregation, another significant morphological change in mitosis is the breakdown and reassembly of the nuclear envelope (NE). Upon entry into mitosis, filamentous lamina proteins, including Lamin A/C and Lamin B1, are phosphorylated, causing them to depolymerize and release the chromatin they bind to at the lamina associated domains (LADs), which leads to NE breakdown and chromosome condensation. Shortly after chromosome separation during anaphase, lamina proteins are dephosphorylated and repolymerized for NE reassembly, allowing the NE to rebind with chromatin to form LADs15,16. Notably, the assembling NE transiently forms two distinct domains during telophase: the “core” and “non-core” domains. The NE membranes assembling over chromatin in contact with the mitotic spindle form the “core” domain and accumulate core proteins, including Lamin A/C, barrier-to-autointegration factor (BAF) and LEM domain-containing proteins that interact with BAF. Membranes away from the spindle form the “non-core” NE domain and recruit non-core proteins, which include the nuclear pore complexes (NPCs), Lamin B1 and its receptor LBR17,18,19. At the end of mitosis, the ESCRT (endosomal sorting complexes required for transport) protein CHMP7 and LEM2, a transmembrane ESCRT adaptor, undergo a liquid-like condensation to form a macromolecular O-ring seal at the confluence between membranes, chromatin and the spindle, finalizing nuclear envelope reassembly20.
NE defect often causes DNA damage at the rupture sites21,22, and contributes to tumor invasion23 and various human diseases, including laminopathies and aging18. But how NE integrity and dynamics respond to genome instability under replication stress remains unclear. Here, we report an interesting phenomenon in which mild RS induces a nuclear envelope reassembly defect (NERD) in mitosis. The NERD is associated with mitotic DNA damage and can be suppressed by Polθ/TMEJ. Genomic mapping of LADs using CUT&Tag identifies a subset of LADs sensitive to replication stress, which we term RESSLADs (REplication Stress Sensitive LADs). Sustained phosphorylation of Lamin A/C and degradation of Lamin B1 occur at the NERD sites. Therefore, these discoveries bridge the gap between genome instability and NE defect under RS.
Results
Replication stress results in nuclear envelope defect
To investigate the impact of RS on NE integrity, we applied a moderate concentration (0.4 μM) of the DNA polymerase inhibitor aphidicolin (APH)8,24 to human RPE-1 cells stably expressing GFP tagged BAF, a marker of nuclear integrity25. Consistent with previous reports26, mild RS induced by APH caused chromosome missegregation and the formation of micronuclei. Intriguingly, APH treatment also resulted in a four-fold increase (from ~6.7% to ~27.1%) of cells with GFP-BAF foci compared to DMSO treatment (Fig. 1a, b, and Supplementary Fig. 1a), suggesting that mild RS could induce NE defect. The NE defect was positively correlated with the dose of APH, peaking at 0.4 μM (Supplementary Fig. 1b). However, the defect rate decreased when cells were accumulated at the S phase with high doses of APH (Supplementary Fig. 1b, c). Furthermore, the RS-induced NE defect was observed in multiple cell lines including HeLa, U2-OS and HCT-116 (Supplementary Fig. 1d), although the ratios varied.
a RPE-1 TP53−/− cells stably expressing GFP-BAF were treated with 0.4 μM APH (Aphidicolin) followed by Hoechst staining and imaging (top), percentage of cells with GFP-BAF foci or micronuclei were quantified (bottom) (n = 3, N > 200). b Representative images of GFP-BAF and Lamin B1 (left) from 2 independent experiments. Yellow arrows: MN; white arrowheads: GFP-BAF foci. Right: Line-scan intensity (boxed regions); note BAF-Lamin B1 anti-correlation. c Representative images (left) and quantification (right, n = 3, N > 200) of cGAS-mNG foci. Arrowhead: cGAS-mNG foci. d Representative TEM images of nuclear envelope (n = 2, N = 5). Boxed regions: Magnified views; white arrows: NE bilayer; yellow arrows: NE defects. Scale bar, 2 μm. e Representative images of GFP-BAF and Lamin A/C (left) from 2 independent experiments. Arrowheads: GFP-BAF foci. Right: Line-scan intensity (dashed line); note BAF-Lamin A/C colocalization. f Localization of endogenous BAF and Lamin B1. Arrowheads: endogenous BAF foci. The bar graph shows quantification of cells with endogenous BAF foci (n = 3, N > 200). g Representative images of endogenous BAF and Lamin A/C from 3 independent experiments. White arrowheads: endogenous BAF foci. h Western blots of the indicated proteins (left). The bar graph indicates the quantification of protein levels normalized to DMSO group with GAPDH as internal control. Data are presented from 4 independent experiments. i Localization of GFP-BAF under DMSO or PDS (pyridostatin) treatment for 48 h (left). The bar graph shows the quantification of cells with BAF foci (n = 3, N > 200). j Western blots of the indicated proteins in IMR-90 cells with or without doxycycline (Dox) treatment for HRas-G12V induction for 7 days. The bar graph indicates the quantification of protein levels normalized to DMSO group with GAPDH as internal control. k Localizations of GFP-BAF, Lamin B1 and Lamin A/C with or without HRas-G12V induction for 7 days. Arrowheads indicate the BAF foci. The bar graph shows the percentage of cells with GFP-BAF foci (n = 3, N > 200). Data are presented as mean ± SD. The p-values are from two-tailed unpaired Student’s t-test. Scale bars for all images except (d), 10 μm. Source data are provided as a Source Data file.
Next, immunofluorescence assays against the lamina proteins were performed to confirm the NE defect indicated by GFP-BAF foci. The results demonstrated that Lamin B1 was absent from the GFP-BAF foci (Fig. 1b, region 1) or “patches” (severer NE defect, Fig. 1b, region 2) under APH treatment, contrasting with the continuous nuclear periphery in DMSO-treated cells.
In addition to GFP-BAF, other reporters including RFP-NLS, a RFP-NLS GFP-BAF dual reporter, and cGAS-mNG (mNeonGreen) were used to confirm the effect of RS on NE integrity. Cells with cytosolic RFP-NLS increased by APH treatment from 6.6% to ~20% which was unaffected by GFP-BAF (Supplementary Fig. 1e, left and middle panels). Strikingly, the RFP-NLS GFP-BAF dual reporter (cytosolic RFP-NLS with GFP-BAF foci) dramatically reduced the background to 1.5%, and APH treatment increased NE defect to 15.1% (Supplementary Fig. 1e, left and right panels). The high background of RFP-NLS might be due to NE disassembly and re-assembly during mitosis. Meanwhile, APH treatment induced a significant increase of cells with cGAS-mNG foci (4.2% to 19.5%, Fig. 1c) which colocalized with BAF foci (Supplementary Fig. 1f). It is of note that the baseline of NE defect varies with different reporters, suggestive of differential sensitivities, intrinsic limitations, and potential side effects. Nevertheless, all these systems validate that RS triggers significant increase of NE defect.
Moreover, the structure of the NE was examined by transmission electron microscopy (TEM) at a higher resolution. The bilayer structure of NE was intact in control cells, whereas gaps were observed in cells subjected to mild RS (Fig. 1d). Surprisingly, unlike Lamin B1, Lamin A/C demonstrated strong enrichment and co-localization with GFP-BAF foci (Fig. 1e), indicating a distinct role of Lamin A/C in response to RS-induced NE defect. GFP-BAF showed ~10-fold increase of expression compared to the endogenous BAF protein and APH induced ~1.4-fold increase (Supplementary Fig. 1g, h), but it did not affect cell proliferation (Supplementary Fig. 1i). To further exclude the potential side effects of GFP-BAF, immunofluorescence of endogenous BAF and lamina proteins was performed in WT cells without GFP-BAF. The results demonstrated that APH treatment induced the formation of endogenous BAF foci with the depletion of Lamin B1 (Fig. 1f) and enrichment of Lamin A/C (Fig. 1g).
Further, the expression levels of lamina proteins were examined by immunoblotting. APH-treated cells showed higher expression of both GFP-BAF and endogenous BAF (Fig. 1h and Supplementary Fig 1g, h), while the levels of both Lamin B1 and Lamin A/C were reduced (Fig. 1h and Supplementary Fig. 1j), anti-correlated with the duration of APH treatment (Supplementary Fig. 1k). Since the expression of lamina proteins was quite stable during cell cycle (Supplementary Fig. 1l), it is less likely that the reduced lamina protein levels is due to the change of S-phase accumulation under RS.
Finally, in addition to APH treatment, the effects of other types of replication stress were investigated. First, treatment of pyridostatin (PDS), a stabilizer of G-quadruplex (G4) induced the formation of GFP-BAF foci in a dose-dependent manner (Fig. 1i). Second, a genetic model with tetracycline inducible TetOn-HRas-G12V was established to trigger replication stress27,28 in both RPE-1 cells and IMR-90 (human lung epithelial) cells. After doxycycline (Dox) induction, the expression of Lamin B1 and Lamin A/C was decreased, while the GFP-BAF level was slightly increased (Fig. 1j and Supplementary Fig. 1m), similar to the APH treatment. Accordingly, the percentage of cells with GFP-BAF foci was significantly increased (Fig. 1k), Lamin B1 depletion and Lamin A/C enrichment were observed at the BAF foci (Fig. 1k). Therefore, NE defect appears to be a general response to RS.
Does DNA damage cause NE defect? To address this question, DNA-damaging agents such as Camptothecin (CPT, 5 μM for 1 h), Zeocin (50 μg/mL for 1 h) and ionizing radiation (IR, 10 Gy) were applied to induce acute DNA damage and test their effects on NE integrity. Although drastic DNA damage, as indicated by γH2AX, was observed after treatments, the cells did not show significant increase in BAF foci (Supplementary Fig. 1n, o), in contrast to a recent report that prolonged (24 h) DNA damage causes NE rupture reported by cGAS during interphase29. It is of note that the level of DNA damage induced by RS was lower than DNA damaging reagents (Supplementary Fig. 1p). Collectively, these data suggested that mild RS, rather than acute DNA damage could induce an increase in NE defect.
RS-induced NE defect is due to NE reassembly defect (NERD) in mitosis
To track the dynamics of RS-induced NE defect, we first quantified the percentage of cells with GFP-BAF foci. Compared to the DMSO-treated control cells, cells treated with APH accumulated GFP-BAF foci in a time-dependent manner (Fig. 2a and Supplementary Fig. 2a–c), suggesting that cell cycle progression may play a role in NE defect. To test this possibility, synchronization assay was performed. RO-3306, a selective CDK1 inhibitor (CDK1-i)30 was applied to arrest cells in the G2 phase. Although CDK1-i treatment alone showed the same level of BAF foci as the control, the increased NE defect induced by APH treatment was abolished by CDK1-i (Fig. 2b). Therefore, mitotic entry appears to be crucial for RS-induced NE defect. In addition, treatment of PHA-767491, a selective DDK (Dbf4-dependent kinase) inhibitor (DDK-i) that inhibits DNA replication initiation and causes G1/S accumulation (Supplementary Fig. 2d)31, also reversed NE defect in a dose-dependent manner (Fig. 2c). Consistently, the effects of these two inhibitors were confirmed by endogenous BAF staining (Supplementary Fig. 2e, f). Together, these results indicated that RS-induced NE defect is coupled with cell cycle progression, particularly mitotic entry.
a Quantification of cells with GFP-BAF foci with indicated treatments. Statistical analysis is performed by ordinary two-way ANOVA with Tukey’s multiple comparisons test (n = 6, N > 200). b, c The effect of CDK inhibitor (CDK-i) (b, n = 6, N > 200) or DDK inhibitor (DDK-i) (c, n = 3, N > 200) on GFP-BAF foci formation under APH treatment. d Dynamics of GFP-BAF and H2B-RFP under indicated treatments. The GFP-BAF and H2B-RFP are stably expressed in RPE-1 cells and imaged at intervals of 5 min. The time of metaphase is referred to as time “0”, due to its round morphology. This is distinct from the time point of NE resealing in telophase. Arrows indicate the formation of GFP-BAF foci. Arrowheads indicate the accumulation of smaller GFP-BAF puncta. e Quantification of mitotic events with or without BAF foci formation. N > 50 mitotic events were quantified for each experiment. f Single cell tracking for BAF foci and cell fate. Cells (53 for DMSO and 83 for APH) were imaged immediately upon treatment for 72 h at 5 min intervals. Cell fate after each mitosis is analysed for single cells. The number of cells undergoing specific event is labelled. g The NE subdomains reassembled in telophase. h Relative abundance of the indicated ‘core’ and ‘non-core’ proteins at NE defect sites. The fluorescent intensity of individual proteins at BAF foci under APH treatment is normalized to the DMSO group after background subtraction. Left to right, n = 24, 16, 20, 10, 12, 26 and 12. i Localization of LEM2. Note the colocalization of GFP-LEM2 with BAF foci in APH-treated cells. Bar graph at the bottom shows the quantification of cells with LEM2 and BAF foci (n = 3, N > 200). Unless otherwise stated, statistical analysis is performed by two-tailed, unpaired Student’s t-test for interested pairs (ns p > 0.05, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001). Scale bar, 10 μm. Source data are provided as a Source Data file.
To further confirm the temporal relationship between cell cycle and NE defect, an RPE-1 cell line stably expressing GFP-BAF and RFP-H2B (Histone 2B, a marker of chromatin mass) was constructed for live-cell time-lapse imaging. As expected, GFP-BAF in cells with DMSO treatment did not form foci either before or after metaphase, which was referred to as “time 0” due to its featured morphology (Fig. 2d top panels and Supplementary Movie 1). However, GFP-BAF in APH-treated cells was diffusive before metaphase and quickly formed foci (indicated by arrows) during telophase. The BAF foci increased in number and size, presumably through the fusion of small foci or enlargement of the defect. It is of note that the paired daughter cells exhibited heterogeneity in the number of BAF foci, as well as the time of BAF accumulation (Fig. 2d bottom panels and Supplementary Movie 2). Since telophase is accompanied by NE reassembly, the RS-induced NE defect per se is a result of mitotic NE reassembly defect (NERD), rather than de novo NE defect in interphase. BAF accumulation occurred about 86 min post-metaphase on average (Supplementary Fig. 2g), and nearly 50% of the mitotic events under RS generated NERD (Fig. 2e). Lastly, we assessed the cell fate after mitosis by single cell tracking. Overall, the APH-treatment cells showed much more heterogeneity than the DMSO-treated population. With APH treatment, the cell divided more slowly and the duration of cell cycle became longer (Supplementary Fig. 2h, i). This may explain why BAF foci under RS accumulated later (Supplementary Fig. 2g) than the time of normal NE resealing at ~60 minutes post-metaphase (Fig. 2d, top panel). Intriguingly, only a small amount of cells could re-sealed the defective NE. The more BAF foci in the parental cells, the worse defects would be observed in the daughter cells. Cells with no less than 3 BAF foci tended to undergo cell cycle arrest or mitotic catastrophe (Fig. 2f).
As NE reassembly is highly organized in telophase (Fig. 2g), the core and non-core domain proteins were examined by immunofluorescence. The results showed that the core proteins, including Lamin A/C, Emerin and LAP2β (Fig. 1e and Supplementary Fig. 2j, k) were enriched and colocalized with GFP-BAF foci, whereas the non-core proteins including LBR, Nup62 and TPR (Supplementary Fig. 2l–n) were depleted at the GFP-BAF foci under APH treatment (Fig. 2h). Moreover, LEM domain-containing proteins such as LEM1, LAP2α, MAN1, Ankle1 and Ankle2 can interact with BAF25. We therefore tagged these proteins with RFP or Halo tag and stably expressed them in RPE-1 GFP-BAF cells. As expected, they all displayed co-localization with GFP-BAF foci under RS (Supplementary Fig. 2o–s). Although RS could induce mitotic errors including micronucleation and formation of ultra-fine bridge (UFB) (Fig. 1a and Supplementary Fig. 2t), as reported32,33,34. High-resolution imaging showed that the sites of NERD in telophase were distal to UFB in APH group (Supplementary Fig. 2u). Quantification of the cells with UFBs/NERD showed that RS induces 1.94 ± 0.66% UFBs and 31.34 ± 0.62% NERD in telophase cells (Supplementary Fig. 2u, right). It is noteworthy that co-localization between UFB and NERD was not observed, suggesting that NERD might not be a direct outcome of mitotic errors such as UFBs. However, the ESCRT-III protein CHMP7 (endogenously tagged with Halo tag35) and LEM2, two NE sealing factors during telophase36, showed enrichment at the BAF foci (Fig. 2i, Supplementary Fig. 2v). These observations are reminiscent of abnormal NE reassembly and sealing problems in telophase, supporting the notion that RS-induced NE defect is resulted from NERD.
Repair of mitotic DNA damage by TMEJ ameliorates NERD
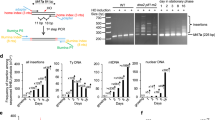
In addition to NERD, RS also triggers DNA damage and repair synthesis through MiDAS or TMEJ pathways in early mitosis (Fig. 3a). This raises a question of whether these two processes are related, and what effect mitotic DNA damage has on NERD. To address this, we used siRNA to target MiDAS-related genes, including MUS81 and POLD3, as well as the TMEJ factor POLQ to knock down their expression, and then investigated their effects on RS-induced NE defect. The designed siRNAs effectively suppressed the expression of these factors (Supplementary Fig. 3a, b). Although the suppression of MUS81 and POLD3 significantly reduced the mitotic DNA repair synthesis visualized by EdU (5-ethynyl-2′-deoxyuridine) staining (Supplementary Fig. 3c), they had almost no effect on NE defect under APH treatment (Fig. 3b). In contrast, POLQ knock-down caused a significant increase in NE defect (~39%) compared to the control siRNA (~22%) with APH treatment (Fig. 3b). It is of note that MUS81 depletion increased the rate of NE defect in the absence of RS, consistent with its role in countering intrinsic RS and stabilizing replication forks37,38,39. The depletion of MUS81 itself causes RS. To further validate these RNAi results, we used CRISPR-Cas9 with dual sgRNAs flanking the start codon to knock out the POLQ gene (Supplementary Fig. 3d). The genotypes of POLQ−/− clones were confirmed by PCR and sequencing, and their mRNA expression was abolished (Supplementary Fig. 3e–g). Compared to wild type (WT), POLQ−/− cells with APH treatment showed a significant increase in NE defect (Fig. 3c). Meanwhile, both endogenous BAF and cGAS-mNG foci exhibit significant increase in POLQ−/− cells following APH treatment (Fig. 3d, Supplementary Fig. 3h). These data suggest that the TMEJ pathway, instead of MUS81-POLD3 mediated MiDAS pathway, plays an important role in RS-induced NERD.
a RS induces mitotic DNA damage and NERD, respectively. It is unknown whether and how mitotic DNA repair pathways such as MiDAS and TMEJ affect NERD. b Representative images of GFP-BAF under DMSO or APH treatment with knockdown of the indicated genes by siRNA. Scale bar, 10 μm. Bar graph on the right shows the quantification of cells with BAF foci. N > 200 for each experiment. c Representative images of GFP-BAF in WT or POLQ−/− cells with DMSO or APH treatment. Scale bar, 10 μm. Bar graph underneath shows the quantification of cells with BAF foci. N > 200 for each experiment. d Representative images of endogenous BAF in WT or POLQ−/− cells with DMSO or APH treatment. Scale bar, 10 μm. Bar graph underneath shows the quantification of cells with BAF foci. N > 200 for each experiment. e Alkaline comet assay for WT and POLQ−/− cells with or without APH treatment. Scale bar, 10 μm. The right panel shows the quantification of tail moment. N > 25 for each experiment. f Experimental workflow for detection of mitotic DNA damage (g) and mitotic DNA repair synthesis (h). g The γH2AX foci on mitotic chromosomes in WT and POLQ−/− cells with or without APH treatment. Scale bar, 10 μm. Quantification of γH2AX foci is shown on the right. N > 50 for each experiment. h The EdU foci on mitotic chromosomes in WT and POLQ−/− cells with or without APH treatment. Scale bar, 10 μm. Quantification of EdU foci is shown on the right. N > 35 for each experiment. Unless otherwise stated, the p-values of two-tailed unpaired Student’s t-test are indicated for interested pairs. Data are presented as mean ± SD, n = 3 independent experiments. Source data are provided as a Source Data file.
Next, we explored whether the increased NE defect in POLQ−/− was due to mitigated DNA damage repair. Indeed, higher levels of DNA damage were detected in POLQ−/− clones under RS, whereas no difference between POLQ−/− and WT was found under DMSO treatment in the comet assays (Fig. 3e and Supplementary Fig. 3i). To further examine RS-induced DNA damage in mitosis, cells were synchronized at G2/M boundary by RO-3306 and subsequently released into mitosis with Edu labelling for 0.5 h, after which the mitotic cells were harvested for either γH2AX or EdU staining (Fig. 3f). Compared to WT cells, POLQ−/− cells with APH treatment showed more γH2AX foci (Fig. 3g) but fewer EdU foci from the mitotic repair synthesis (Fig. 3h). Furthermore, EdU foci were mostly co-localized with γH2AX foci on mitotic chromosomes under APH treatment (Supplementary, Fig. 3j). To investigate the role of DNA damage response in NERD, cells were treated with KU60019, an ATM kinase inhibitor (ATM-i). The result showed that ATM-i, particularly at high concentrations, slightly increased NE defects in both DMSO- and APH-treated cells (Supplementary, Fig. 3k). Similar effect was observed in POLQ−/− cells (Supplementary, Fig. 3l). Therefore, ATM-mediated DNA damage response might not account for RS-induced NERD. Indeed, ATR has been reported to play an important role in mitotic DNA damage response40. Meanwhile, it is noticeable that the POLQ-deficient cells appeared to be more fragmented with more BAF foci (Fig. 3b–d). Indeed, POLQ depletion resulted in ~2.5-fold increase of cell death with APH treatment, detected by FACS (Fluorescence-Activated Cell Sorting) after Annexin V and PI (propidium iodide) staining (Supplementary Fig. 3m). Together, these results identify TMEJ as a dominant repair pathway for RS-triggered mitotic DNA damage, which mitigates the adverse impact of mild RS on NE integrity.
RS could cause loss of LADs at CFSs
The meshwork-like nuclear lamina not only safeguards the NE against mechanical stress but also provides a scaffold for chromatin binding, interacting with hundreds of large genomic regions known as lamina-associated domains (LADs)41. Does RS affect the LADs formation? To address this question, Cleavage Under Targets and Tagmentation (CUT&Tag) was performed against Lamin A/C and Lamin B1 in RPE-1 cells with or without APH treatment. The results revealed that the total chromatin binding sites of both Lamin A/C and Lamin B1 were reproducibly and significantly reduced under RS compared to DMSO treatment (Fig. 4a–c). Further examination of LADs with reduced signals under RS identified 485 LADs from Lamin A/C and 486 LADs from Lamin B1. Intriguingly, a majority of them (355 LADs) were shared between Lamin A/C and Lamin B1 (Fig. 4d and Supplementary Data 1), which we termed RESSLADs (REplication Stress-Sensitive LADs). RESSLADs were distributed across all chromosomes but appeared to have a higher density at chromosome ends closer to the telomeres (Supplementary Fig. 4a). These data indicated that the chromatin binding to NE was reduced under RS.
a, b Traces of overall signal intensity in CUT&Tag experiments for Lamin A/C (a) or Lamin B1 (b) with DMSO or APH treatments in RPE-1 cells. Each treatment has 3 repeats (rep 1 to 3). The colour intensities are scaled to the maximum read count at peaks in each dataset. c Quantification of total binding sites of Lamin A/C (left) and Lamin B1 (right), respectively. d Venn diagram showing the overlap of LADs between Lamin A/C and Lamin B1. The overlapped LADs are defined as RESSLADs. e Distribution RESSLADs in Common Fragile Sites (CFS), Rare Fragile Sites (RFS) and others. f Representative profile of peaks for Lamin A/C and Lamin B1 at FRA1A locus. The RESSLADs (#200, #1, #23 and #33) are indicated in purple. Lamin A/C peaks are indicated in pink and Lamin B1 peaks are in blue. The intensity maps at the bottom represent the average of three repeats. g The binding sites of Lamin A/C and Lamin B1 at FRA1A locus. h Venn diagram showing the overlap of LADs (RESSLADs) between Lamin A/C and Lamin B1 in HCT-116 cells. i Distribution of RESSLADs in Common Fragile Sites (CFS), Rare Fragile Sites (RFS) and others in HCT-116 cells. j Venn diagram showing the overlap of RESSLADs between RPE-1 cells and HCT-116 cells. k Relative mRNA expression of genes located in RESSLAD#1 (ACOT7 and GPR153) and RESSLAD#2 (SLURP1, LYNX1 and LY6D) in RPE-1 cells with/without APH treatment. The values are normalized to DMSO group with GAPDH as internal control. l CUT & Tag qPCR for γH2AX. The workflow was on the top. The bar graph shows the binding of RESSLADs to γH2AX with the indicated treatments. m ChIP-qPCR analysis of the binding of RESSLAD#1 to Lamin A/C in RPE-1 WT and POLQ−/− cells with or without APH treatment. Data are presented as mean ± SD, n = 6 independent experiments. Unless otherwise stated, the p-values of two-tailed unpaired Student’s t-test are indicated for interested pairs. Data are presented as mean ± SD, n = 3 independent experiments. Source data are provided as a Source Data file.
Since RS, such as APH treatment, induces the “expression” and fragility of the common fragile sites (CFSs) in the genome, we investigated whether RESSLADs were located in CFSs. Indeed, 99 RESSLADs (~28%) in RPE-1 cells were located in CFSs and 18 RESSLADs were colocalized with rare fragile sites (RFSs) (Fig. 4e). However, some CFSs including FHIT (Supplementary Fig. 4b) and WWOX (Supplementary Fig. 4c) did not show alterations in LADs between DMSO and APH, possibly due to the genetic background of RPE-1 in which the fragility of CFSs might be different from other cell lines42. The mapping of RESSLADs to CFSs showed that the RESSLAD#1 with the most reduced signal resided in FRA1A which spans a 2.2 Mb region on chromosome 1. Indeed, another 3 RESSLADs (#29, #33 and #200) were also mapped to FRA1A (Fig. 4f). Accordingly, FRA1A exhibited an approximately 25% decrease in binding peaks of Lamin A/C and Lamin B1, respectively (Fig. 4g). Consistent with the results in non-transformed RPE-1 cell line, CUT&Tag experiment using HCT-116 cell line also showed reduced binding of LADs to Lamin A/C and Lamin B1 under RS (Supplementary Fig. 4d and Supplementary Data 1). A total of 382 RESSLADs were identified, with ~24% in fragile sites (Fig. 4h, i). However, only 32 RESSLADs (<10%) were shared between RPE-1 and HCT-116 (Fig. 4j), indicative of a cell line-dependent sensitivity and responses of RESSLADs to RS. The difference between RPE-1 and HCT-116 might reflect their different genetic backgrounds. Nevertheless, these data support the notion that RS causes loss of LADs/CFSs from NE, which might be a property of NERD.
Finally, we investigated the potential consequences of such reduced RESSLADs-NE interaction. To do this, we examined the expression of genes in RESSLADs. Under APH treatment, the expression of genes ACOT7 and GPR153 in RESSLAD#1 (FRA1A) and SLURP1, LYNX1 and LY6D in RESSLAD#2 (FRA8D) were significantly down-regulated (Fig. 4k). This down-regulation is possibly due to the RS-induced DNA damage at these loci. Consistent with this idea, CUT&Tag qPCR for γH2AX in mitotic cells showed that APH treatment increased DNA damage in RESSLAD#1 and RESSLAD#2 (Fig. 4l). Furthermore, chromatin immunoprecipitation (ChIP)-PCR for Lamin A/C demonstrated that binding of RESSLADs to NE was exacerbated in POLQ−/− cells compared to the WT cells under RS (Fig. 4m), suggesting an important role of TMEJ in maintaining genome stability and chromatin-NE interaction.
Lamin A/C phosphorylation is sustained during NERD
In mitosis, lamina proteins undergo phosphorylation during NE breakdown and dephosphorylation to facilitate LADs formation during NE reassembly15,16. We asked whether the phosphorylation of Lamin A/C was involved in the reduced interaction between RESSLAD and NE under RS. Although more than 20 phosphorylation sites on Lamin A/C have been reported43, we focused on the phosphorylation of Serine 22 (pS22), which is predominant during the NE remodelling in mitosis44. Indeed, APH treatment caused ~5-fold increase in pS22-Lamin A/C while reduced the total Lamin A/C protein level (Fig. 5a). Intriguingly, pS22-Lamin A/C signal was highly enriched at NERD sites (Fig. 5b), with >72% of the BAF foci containing pS22-Lamin A/C (Supplementary Fig. 5a). To further understand these dynamics, we examined the changes in pS22-Lamin A/C during mitosis under RS using a synchronization assay. Cells were treated with DMSO or APH for 32 h, followed by RO-3306 treatment for another 16 h before release into mitosis. Before release when cells were in G2 phase, no pS22-Lamin A/C signal was detectable. Upon mitotic entry, the DMSO-treated group exhibited a rapid and strong increase in the mitotic marker pS10-H3 (phosphorylation of Serine 10 on Histone H3) and pS22-Lamin A/C, with both signals disappearing between 60 min and 120 min (Fig. 5c). In contrast, the APH-treated group maintained high levels of pS22-Lamin A/C after 120 min, despite the disappearance of the pS10-H3 signal (Fig. 5c), correlating with the time of NERD. The reduced pS10-H3 in APH-group might suggest a delayed mitotic entry of the treated cells. To overcome this problem, mitotic shake-off was performed to harvest same amount of mitotic cells. At the end, pS22-Lamin A/C levels in APH group were always higher than the control, at individual time points (Supplementary Fig. 5b).
a Immunoblotting of the indicated proteins with or without APH treatment. Note the pS22-Lamin A/C level under APH treatment. Bar graphs show the quantification of total Lamin A/C (middle) or pS22-Lamin A/C level (right). Values under APH treatment are normalized to DMSO treatment with GAPDH as internal control. Data are presented as mean ± SD, n = 4 independent experiments. Statistical analysis is performed by two-tailed unpaired Student’s t-test. b Representative Immunofluorescence images of GFP-BAF and pS22-Lamin A/C with DMSO or APH treatment. Arrowheads indicate the BAF foci. Scale bar, 10 μm. Right, quantification of the percentage of cells with pS22-Lamin A/C foci. Data are presented as mean ± SD, n = 3 independent experiments, N > 200 for each experiment. Statistical analysis is performed by two-tailed unpaired Student’s t-test. c Immunoblotting of the indicated proteins with or without APH treatment at different time points after mitotic entry. Cells are harvested at 0, 15, 30, 60, 120 and 180 min after release from G2 phase. d Localizations of GFP-BAF and Lamin A WT or S22 mutants tagged with N-terminal mCherry. HCT-116 cells stably expressing GFP-BAF are transfected with Lamin A WT, S22A or S22D vectors and treated with DMSO or APH for 48 h. Arrows indicate the BAF foci. Note the severe NE defect in S22D with APH treatment. Scale bar, 5 μm. e Quantification of cells with BAF foci in (d). Data are shown as mean ± SD, n = 3 independent experiments, N > 400 in each experiment. Statistical analysis is performed by two-tailed unpaired Student’s t-test. Source data are provided as a Source Data file.
To further demonstrate the role of S22-Lamin A/C phosphorylation in NERD under RS, mutations of S22 were introduced to Lamin A with N-terminal RFP tag. The mutants were examined in HCT-116 cells by transient transfection. Compared to the Lamin A-WT (wild type), phospho-dead Lamin A-S22A mutant could still localize to BAF foci under RS (Fig. 5d), but it significantly reduced the percentage of cells with BAF foci (Fig. 5e). In contrast, the phospho-mimetic Lamin A-S22D mutant readily caused increased BAF foci formation even in DMSO treatment, and the effect was exacerbated under APH treatment (Fig. 5d, e). ATR-dependent phosphorylation of Lamin A/C at S282 has been reported to play a role in DNA damage-induced NE defect in interphase29. However, the phospho-dead Lamin A-S282A mutant showed almost no difference with Lamin A-WT under both DMSO and APH treatments (Supplementary Fig. 5c, d). Altogether, these data confirmed a critical role of pS22-Lamin A/C in NERD and chromatin organization under RS.
Prominent NE defect occurs under conditions of synthetic lethality
Lastly, we investigated the effect of NERD on clinical implications. RS can activate ATR (Ataxia-Telangiectasia Mutated and Rad3-related protein kinase) to maintain replication fork stability and checkpoint function45. Exploiting the reliance on ATR and DNA damage response under RS has become a strategy for inducing synthetic lethality in anti-cancer drug discovery. In the field of DNA damage response, synthetic lethality predominantly emphasizes the accumulation of chromosomal instability and DNA damage46, but the integrity of NE has remained largely unexplored. Although treatment of VE-821, an ATR inhibitor (ATR-i) had a comparable effect to DMSO treatment alone, it significantly exacerbated NE defect induced by APH treatment (Fig. 6a, b). Accordingly, the synergistic effect of ATR-i and APH led to decreased cell viability (Fig. 6c). The synthetic lethality of ATR-i in APH-treated cells may be due in part to NE defect. These results also suggested that ATR plays a more important role than ATM in suppressing mitotic DNA damage and NERD.
a Representative images of GFP-BAF with the indicated treatments for 48 h. b Quantification of cells with BAF foci for (a). Data are presented as mean ± SD, n = 3 independent experiments, N > 200 for each experiment. Statistical analysis is performed by two-tailed unpaired Student’s t-test. c Viability assay of RPE-1 cells with the indicated concentrations of ATR-i (VE-821) ± APH treatment for 48 h. Values under different conditions are normalized to the value in DMSO treatment without ATR-i. Data are presented as mean ± SD, n = 9 independent experiments. Statistical analysis is performed by two-way ANOVA. d Representative images of GFP-BAF in WT or BRCA2−/− cells after treatment with DMSO or 5 μM PARP-i (Olaparib) for 48 h. Two BRCA2−/− clones with different genotypes are included. e Quantification of WT or BRCA2−/− cells with BAF foci with indicated treatments. Data are presented as mean ± SD, n = 3 independent experiments, N > 250 for each experiment. Two-way ANOVA is performed for different groups. Two-tailed unpaired Student’s t-test is performed for interested pairs (ns p > 0.05, *p < 0.05, **p < 0.01, ***p < 0.001). f Viability assay for WT or BRCA2−/− cells with treatment of the indicated concentrations of PARP-i. Data are presented as mean ± SD, n = 3 independent experiments. Statistical analysis is performed by two-way ANOVA. g Representative images of GFP-BAF in WT or BRCA2−/− cells after treatment of DMSO or 5 μM Polθ-i (ART558) for 48 h. h Quantification of WT or BRCA2−/− cells with BAF foci after treatment with the indicated concentrations of Polθ-i for 48 h. Data are presented as mean ± SD, n = 3 independent experiments, N > 250 for each experiment. Two-way ANOVA is performed for different groups. Two-tailed unpaired Student’s t-test is performed for interested pairs (ns p > 0.05, *p < 0.05, **p < 0.01, ***p < 0.001). i Viability assay for WT or BRCA2−/− cells with treatment of the indicated concentrations of Polθ-i. Data are presented as mean ± SD, n = 3 independent experiments. Statistical analysis is performed by two-way ANOVA. For (a, d, g), scale bar = 10 μm. Source data are provided as a Source Data file.
Because poly(ADP-ribose) polymerase (PARP) inhibitors (PARP-i) induce death of cells deficient in homologous recombination (HR)47, we examined the NE integrity in HR-deficient BRCA2−/− cells treated with PARP-i. To do this, dual sgRNAs were used to knock out the BRCA2 gene using CRISPR-Cas9 (Supplementary Fig. 6a). Two independent BRCA2−/− clones were confirmed by genotyping (Supplementary Fig. 6b, c), and their mRNA expression was abolished (Supplementary Fig. 6d). Compared with the WT cells, both BRCA2−/− clones showed decreased proliferation (Supplementary Fig. 6e, f) and higher ratios of micronucleation (Supplementary Fig. 6g), indicating a higher level of RS. Consistently, the BRCA2−/− cells exhibited a higher rate of NE defect than WT cells under DMSO treatment (Fig. 6d). When treated with the PARP-i Olaparib, WT cells showed no significant response, whereas BRCA2−/− cells exhibited exacerbated NE defects (Fig. 6d, e) and reduced viability (Fig. 6f). These data suggested that deathly synthetic interactions could cause severe NE defects.
Interestingly, DNA polymerase Polθ and the TMEJ pathway also show synthetic lethality with BRCA248. We assessed NE integrity in BRCA2−/− cells treated with the Polθ inhibitor (Polθ-i) ART558. Similar to the results observed under PARP-i treatment, ART558-treated BRCA2−/− cells, but not WT cells, exhibited increased NE defects (Fig. 6g, h) and reduced cell viability (Fig. 6i). Notably, even a very low concentration (0.1 μM) of Polθ-i caused a pronounced reduction (~60%) in the viability of BRCA2−/− cells (Fig. 6i). Collectively, these results suggested that NERD and subsequent NE dysfunction may be a general cellular response under conditions of synthetic lethality.
Finally, we examined the consequence particularly cellular metabolisms in cells with RS-induced NE defect. The results showed that the APH-treated group showed elongated mitochondria. In the same population, cells with BAF foci had significantly longer mitochondria than those without BAF foci (Supplementary Fig. 6h). In addition, the autophagic flow was also altered by RS, as the number of GFP-LC3 foci was increased by more than 2 folds (Supplementary Fig. 6i). Therefore, RS-induced NE defect are accompanied by metabolic changes such as mitochondrial dysfunction and autophagic alteration, highlighting the complex and multifaceted impacts of RS on cell functions.
Discussion
Studies of RS have predominantly focused on DNA damage response and genome instability, with limited exploration on its effects on cellular structures such as the nuclear envelope. Here, we establish a link between genome instability and NE defect under RS using multiple reporters of NE integrity. The baselines of NE defect varied in different reporters, reflecting their intrinsic noise and limitations. Notably, the endogenous BAF staining and RFP-NLS GFP-BAF dual reporter both show ~1% NE defect, which is consistent with the reported large datasets with endogenous BAF staining49. Nevertheless, all reporter used in this study could detect significant increase in NE defect upon RS. It is of note that RS-induced NERD in mitosis appears to be quite different from NE rupture triggered by prolonged DNA damage in interphase29. In addition, RS could induce mitotic DNA damage and cause sustained phosphorylation of Lamin A/C, leading to NERD. DNA Polθ/TMEJ suppresses NERD by repairing mitotic DNA damage. Conversely, Polθ deficiency exacerbates NERD (Fig. 7).
In WT cells, Lamin A/C phosphorylation by CDK1 in prophase triggers NE breakdown, and its dephosphorylation in telophase allows the reassembly of NE with normal LADs (top). Under replication stress, TMEJ actively repairs mitotic DNA damage, rescues NE reassembly and ameliorates NERD (middle). However, with POLQ-deficiency, mitotic damage could not be efficiently repaired, leading to sustained Lamin A/C phosphorylation and exacerbated NERD.
As a result of RS, the mitotic entry of replication machinery causes the disassembly of the replisome and exposure of under-replicated DNA to nucleases, resulting in DSBs4,8,50. The repair of DSBs during mitosis is challenging, as the canonical DSB repair pathways including HR and NHEJ are largely inactive during mitosis51. Previous studies have suggested that MiDAS effectively repairs RS-induced mitotic DNA damage8,52,53. Although depletions of MUS81 and POLD3 reduce, rather than abolish mitotic DNA synthesis (Supplementary Fig. 3c), they do not influence NERD under RS (Fig. 3b). Strikingly, we found that Polθ/TMEJ plays a critical role in reducing both RS-induced mitotic DNA damage and NERD. Consistent with our findings, a recent study demonstrates that TMEJ but not MiDAS mediates the repair of CFSs in mitosis14. One possible explanation for the distinct roles of MiDAS mediated by MUS81-POLD3 and TMEJ could be temporal resolution: MiDAS may extend replication into prophase, whereas TMEJ repairs DNA damage in late mitosis until NE reassembly. This suggests that MiDAS pathway and TMEJ may target different types of DNA substrates or genomic regions. Additionally, we cannot exclude the possibility that Polθ plays a direct role in maintaining NE integrity or regulating NE reassembly during mitosis, which remains to be investigated. In addition to Polθ, RS-induced NERD may involve other factors in microhomology-mediated end joining pathway that regulate NE reassembly54.
During NE breakdown in prophase, inner nuclear membrane proteins, including Lamin A/C, undergo progressive phosphorylation. Phosphorylation of S22 and S392 of Lamin A/C by CDK1/Cyclin B leads to Lamin A/C depolymerization and release of chromatin binding55,56. Interestingly, sustained pS22-Lamin A/C at NERD sites, identified in this study, implies functional defect of Lamin A/C in organizing NE structure and LADs-NE interaction during NERD. Future studies will address how mitotic DSBs regulate Lamin A/C phosphorylation, and whether this failure in dephosphorylation is due to persistent kinase activation or phosphatase suppression during NERD (Fig. 7). It is noteworthy that Lamin A/C can also be phosphorylated, in ATR dependent manner, during prolonged DNA damage or in micronuclei57. However, these events are thought to take place in interphase, in contrast to the RS-induced NERD in mitosis. These observations together suggest that lamina proteins can be modified diversely in response to different types of stress. In addition, RS caused elevated expressions of BAF, indicating a general response of BAF to RS. One possible explanation would be the enrichment and/or phase separation of BAF at NERD sites, as a NE repair factor25,58,59, that protects it from degradation.
The identification of RESSLADs and their overlap with CFSs highlights the potential changes in chromatin structure and gene transcription under RS. The distribution of RESSLADs to non-CFS regions suggested that RS induces a broader effect on the whole genome, not limited to CFSs. As CFSs vary across different cell lines42,60, it is possible that fragile sites are waiting to be discovered in RESSLADs. In addition, our finding shows that the phosphorylation at S22 of Lamin A/C is significantly elevated in response to RS and highly localized to NERD sites. Given that three-dimensional chromatin architecture plays a critical role in gene regulation and is mainly regulated by the nuclear lamina, this phosphorylation reduces the chromatin-binding capacity of Lamin A/C, thereby altering the cell’s epigenetic landscape16,61,62. In addition to Lamin A/C phosphorylation, CFS “expression” often leads to large deletions in genomic loci, which may also reduce Lamin binding to CFSs regions. The alterations of LADs are associated with pathological transcription patterns in multiple diseases including progeria63 and cardiovascular diseases64,65. Due to the prevalence of RS in cancers, the relationship between RESSLAD and CFS may have implications in tumorigenesis under stressed conditions. However, the precise molecular pathways leading to these modifications (Fig. 7), as well as the downstream changes in chromatin structure and gene expression remain to be elucidated. Moreover, the enrichment of phosphorylated Lamin A/C at NERD sites may also indicate its engagement in subsequent NE repair. Indeed, BAF and Lamin A/C are rapidly recruited to sites of NE defect in response to mechanical force and irradiation59,66,67. In vitro experiments illustrate that the phosphorylated Lamin A/C shows higher intra-nuclear mobility, which may facilitate a quicker response to NE defect68.
In summary, these findings not only enhance our understanding of NE dynamics in response to RS, but also open avenues for therapeutic strategies targeting NE vulnerability in cancer and other genome instability-related diseases.
Methods
Cell culture
All cells were cultured at 37 °C in a humidified atmosphere containing 5% CO2. RPE-1 cells were maintained in Dulbecco’s Modified Eagle Medium (DMEM)/Nutrient Mixture F-12 medium supplemented with 10% fetal bovine serum (FBS) and 1% penicillin-streptomycin (Gibco). Aneuploid osteosarcoma (U2-OS), cervical cancer cells (HeLa) and HEK293FT cells were maintained in DMEM medium supplemented with 10% FBS and 1% penicillin-streptomycin (Gibco). HCT-116 cells were cultured in RPMI-1640 medium supplemented with 10% FBS and 1% penicillin-streptomycin. Human lung epithelia IMR-90 cells were cultured in Prigrow X Series Medium (Applied Biological Materials Inc., Cat. #T9167) supplemented with 10% FBS and 1% penicillin-streptomycin. Mycoplasma was routinely monitored and only the mycoplasma-free cells were used for experiments. Low doses of aphidicolin (APH) (Sigma-Aldrich, Cat. #A0781) were added into media for 48 h to induce replication stress, unless otherwise stated. DNA damages were induced by Camptothecin (5 μM for 1 h, Abcam, Cat. #ab120115), Zeocin (50 μg/mL for 1 h, Thermo Fisher Scientific, Cat. #25005), or 10 Gy ionizing radiation. ATM inhibitor (ATM-i) KU60019 (TargetMol, Cat. #T2474) was used to suppress DNA damage response mediated by ATM.
For synchronization assays, RPE-1 cells were treated with 9 μM CDK inhibitor (CDK-i) RO-3306 (Beyotime, Cat. #SC6673-10mM) for 8 or 16 h for G2 arrest. HeLa cells were treated with 2 mM thymidine for 14 h followed by 330 nM nocodazole for 10 h to arrest cells in M phase. The cell cycle was confirmed by immunoblotting of specific markers. DDK inhibitor (DDK-i) PHA-767491 (MCE, Cat. #HY-13461) was used to inhibit DNA replication initiation.
DNA construction and transfection
To construct RFP fusion plasmids, LEM1, MAN1, Emerin, Ankle1, Ankle2, LEM2, and LAPβ were amplified from cDNA of RPE-1, HEK293 or HeLa cells, then cloned into pDONR223 using gateway BP Clonase (Thermo Fisher Scientific, Cat. #11791019) to generate pENTR223 entry vectors. The genes in pENTR223 were recombined into pLenti-DEST-CMV-N-RFP destination vector using gateway LR Clonase II Enzyme (Thermo Fisher Scientific, Cat. #11791019). To construct the mCherry fusion plasmid, Lamin A was amplified from cDNA and introduced into the pCMV-mCherry vector via NheI and XhoI restriction sites. Overlap extension PCR was performed to introduce S22A, S22D and S282A mutations to Lamin A.
To construct dual sgRNA expression vectors for CRISPR-Cas9 system, PCR fragments with sgRNA sequences flanking the dual H1 & U6 promoters were cloned into pLenti-CRISPRv2-2×Scaffold vector (modified from pLenti-CRISPRv2) through type II DNA endonuclease Esp3I sites. More details can be shared upon request. All vectors were verified by Sanger sequencing. All oligonucleotides and plasmids used in this study refer to Supplementary Data 2 and 3, respectively.
DNA constructs were transfected into cells using Lipo8000 (Beyotime, Cat. #C0533) or Lipofectamine 2000 (Thermo Fisher Scientific, Cat. #11668019) according to the manufacturer’s instructions.
Virus packaging and construction of stable cell lines
HEK293FT cells were seeded in a 6-well plate at 70-80% confluence and incubated overnight to allow for cell attachment. Packaging plasmids (pMD2.G and psPAX2 for lentivirus, gag/pol and pVSV-G for retrovirus) together with a target viral vector were transfected into the HEK293FT using Lipofectamine 2000 according to the manufacturer’s instructions. After an overnight incubation, the medium was replaced with 3 mL of fresh medium. The culture media were collected twice, at 48 h and 72 h post transfection, respectively, and filtered through a 0.45 μm syringe filter.
Stable cell lines were generated by transduction with lentivirus or retrovirus carrying the genes of interest. RPE-1 TP53−/− cells stably expressing GFP-BAF or RFP-BAF or cGAS-mNG (mNeonGreen) were selected with G418 (500 μg/mL). Cells stably expressing H2B-RFP, RFP-NLS, RFP-LEM1, RFP-MAN1, RFP-Emerin, RFP-Ankle1, Ankle2-RFP, LAP2α-Halo or GFP-LEM2 were selected with puromycin (10 μg/mL). cGAS-mNeonGreen and RFP-BAF RPE-1 cells were generated by viral transduction and fluorescence-activated cell sorting (FACS) with FACS Aria III (BD). HeLa, U2-OS and HCT-116 cells stably expressing GFP-BAF were generated by viral transduction and fluorescence-activated cell sorting (FACS) with FACS Aria III (BD). All cell lines are listed in Supplementary Data 4.
Live cell imaging and time-lapse movie
For live cell imaging, cells were seeded onto glass-bottomed 35-mm imaging dishes (NEST, Cat. #801001) or 24-well plates (Cellvis, Cat. #P24-1.5 P) overnight. CKX53 inverted microscope (OLYMPUS, Japan) and NHQLiveTM (Bayjayray, China) were used for wide-field imaging, and Nikon Confocal Laser Scanning Microscope (A1) (Nikon, Japan) was used for confocal imaging. Halo-CHMP7 knock-in cells were incubated with JF646 HaloTag Ligand (Promega, Cat. #GA1120) (1:5000, 500 μM stock in DMSO) for 30 min and washed twice with PBS before imaging. Cells were incubated with Mito-Tracker Deep Red FM (Beyotime, Cat. #C1032-50μg) for 15 min to stain mitochondria and washed twice with PBS before imaging. Nuclei were stained by Hoechst 33342 (Invitrogen, Cat. #H1399). Laser excitation conditions were as follows: Hoechst 33342 at 405 nm, GFP at 488 nm, RFP at 561 nm, JF646 HaloTag Ligand at 647 nm. Images were captured with a 60x NA 1.4 oil objective lens. Acquisition parameters, shutters, filter positions and focus were controlled by Nikon software (NIS-Elements AR 5.30.00).
For live cell time-lapse imaging in Fig. 2d, cells were seeded in 24-well μ-plate (ibidi, Cat. #82406) overnight before changing into media with DMSO or APH. After APH treatment, cells were immediately imaged at 5 min intervals using Nikon (Ti-2E) wide-field inverted microscope (Nikon, Japan) equipped with Perfect Focus, a 20x/0.75 NA Plan Apochromat Lambda objective (Nikon), a Zyla 4.2 sCMOS camera (ANDOR) and an environmental enclosure (37 °C, 95% humidity, 5% CO2). NIS-Elements software (Nikon) was used to visualize and analyze the time-lapse videos. ImageJ was used to create annotated images.
Immunofluorescence
Cells were grown on coverslips, treated with indicated reagents and fixed using 4% paraformaldehyde for 20 min at room temperature. Coverslips were washed twice with PBS buffer, permeabilized with 0.5% Triton X-100 for 5 min, followed by blocking with 3% BSA in PBS buffer for 1 h. Primary antibody incubation was performed in PBS buffer with 3% BSA overnight at 4 °C. After washing 3 times with PBS buffer containing 0.05% Triton X-100, secondary antibody staining was performed for 45–60 min in dark at room temperature. The coverslips were washed 3 times with 0.05% Triton X-100 in PBS, incubated with Hoechst 33342 in PBS buffer for 10 min at room temperature, and then followed by washing 3 times with PBS. SlowFade Diamond Antifade (Thermo Fisher Scientific, Cat. #S36963) was used for mounting. Primary antibodies used are as follows: rabbit anti-Lamin A/C (CST, Cat. #2032, 1:200); mouse anti-Lamin B1 (Sangon Biotech, Cat. #D190800-0100, 1:200), rabbit anti-H2AX p-S139 (CST, Cat. #2577 L, 1:400), and rabbit anti-pS22-Lamin A/C (CST, Cat. #13448, 1:200); Rabbit anti-BAF (ProteinTech, Cat. #30525-1-AP, 1:50); Rabbit anti-RPA (Abcam, Cat. #ab76420, 1:100); Mouse anti-beta-tubulin (Abbkine, Cat. #A01030, 1:100). Secondary antibodies used are as follows: goat anti-mouse Alexa647 (Invitrogen, Cat. #A21236, 1:1500), goat anti-rabbit Alexa647 (Thermo Fisher Scientific, Cat #A21245, 1:1500), goat anti-rabbit Alexa568 (Thermo Fisher Scientific, Cat. #A11036, 1:1500), and goat anti-mouse Alexa568 (Thermo Fisher Scientific, Cat. #A11031, 1:1500).
Ultra-fine bridge was detected by using anti-RPA. Cells were synchronized in late G2 (using 9 μM RO-3306) and then released into fresh medium (pre-warmed to 37 °C) for 30 min. Metaphase cells were then harvested by mitotic shake-off and centrifugation at 600 g for 5 min. Pelleted cells were re-seeded onto coverslips coated with Poly-D-lysine (Beyotime, Cat. #ST508) and incubated at 37 °C in an atmosphere of 5% CO2. Cells were collected after 5 min for metaphase, 20 min for anaphase and 45 min for telophase, respectively. The samples were fixed and subjected to immunofluorescence with indicated antibodies. All antibodies used in this study are listed in Supplementary Data 5.
EdU labelling
For mitotic EdU labelling, cells were pulse labelled using 20 μM EdU for 0.5 h after cells were released into mitosis. For S-phase labelling, cells were labelled with 10 μM EdU for 2 h before analysis. Simultaneous fixation and permeabilization were performed using 0.5% Triton X-100, followed by EdU detection using Click chemistry according to manufacturer’s instructions (BeyoClick™ EdU Cell Proliferation Kit, Beyotime, Cat. #C0085S). SlowFade Diamond Antifade (Thermo Fisher Scientific, Cat. #S36963) was used for mounting. Images were captured using Nikon A1.
Electronic Microscopy
RPE-1 cells with or without 0.4 μM APH treatment were fixed with 2.5% glutaraldehyde in 0.1 M phosphate buffer, pH 7.4 for 2 h at room temperature. After washing 3 times with 0.1 M phosphate buffer, cells were scraped and collected with 0.1 M phosphate buffer followed by centrifugation at 1000 g. The pellet was resuspended in 0.1 M phosphate buffer, and centrifuged at 1000 g for 10 min. This step was repeated three times. The samples were post-fixed with pre-cold 1% OsO4 in 0.1 M Phosphate buffer for 2–3 h at 4 °C, followed by rinsing with 0.1 M phosphate buffer for 3 times, 20 min each. The samples were dehydrated in graded ethanol (50%, 70%, 85%, 90%, 95%, 100%) for 15 min under each condition. The penetrations were performed in an order of acetone-epoxy (2:1); acetone-epoxy (1:1); epoxy. Each round of penetration was performed at 37 °C for 12 h. The samples were embedded in epoxy resin using a standard protocol69. Sections parallel to the cell monolayer were cut with a Leica EM UC7 with the thickness of 60–100 nm and examined under a Tecnai G2 20 TWIN (FEI) with an accelerating voltage of 200 kV. Nuclei were identified based on their morphology and were traced by hand.
Western blot
Cells were washed once with PBS buffer and lysed in RIPA buffer supplemented with PMSF. The mixture was incubated on the ice for 30 min, followed by centrifugation 15,000 g for 30 min at 4 °C. The supernatant was mixed with 4xSDS loading buffer and heated at 95 °C for 10 min. The samples were separated by SDS-PAGE gel and proteins were transferred to nitrocellulose membrane. The membrane was blocked in TBST buffer with 5% non-fat milk or 3% BSA at room temperature for 1 h before incubation with a primary antibody diluted in TBST buffer containing 3% BSA at 4 °C overnight. After washing with TBST buffer 3 times, the membrane was incubated with indicated secondary antibodies. Finally, Images were captured using LiCOR Odyssey Imaging Systems (LiCOR, USA) or ChemiDoc MP Imaging system (Bio-Rad) and quantified using ImageJ. Primary antibodies were used as follows: rabbit anti-Lamin A/C (CST, Cat. #2032, 1:1000); mouse anti-Lamin B1 (Sangon Biotech, Cat. #D190800-0100, 1:1000); rabbit anti-H2AX p-S139 (CST, Cat. #2577 L, 1:1000); rabbit anti-pS22-Lamin A/C (CST, Cat. #13448, 1:1000); rabbit anti-GAPDH (Abcam, Cat. #ab9485, 1: 5000); mouse anti-GAPDH (Abbkine, Cat. #A01020, 1: 5000); Rabbit anti-pS10-H3 (Abcam, Cat. #ab30127); Rabbit anti-BAF (Sangon Biotech, Cat. #D290148, 1:1000); Rabbit anti-POLD3 (ProteinTech, Cat. #21935-1-AP, 1:1000); Mouse anti-MUS81 (ProteinTech, Cat. #67351-1-Ig, 1:10000); Rabbit anti-Geminin (Abclonal, Cat. #A25819, 1:1000); Rabbit anti-CDT1 (Abcam, Cat. #ab202067, 1:1000); Mouse anti-alpha-tubulin (ProteinTech, Cat. #66031-1-Ig, 1:10000). Secondary antibodies were used as follows: donkey anti-rabbit IRDye 680RD (Li-COR, Cat. #926-68073, 1:10000); donkey anti-mouse IRDye 800CW (Li-COR, Cat. #926-32212, 1:10000); HRP-Goat anti-Rabbit antibody (Sangon Biotech, Cat. #D110058, 1:10000); HRP-Goat anti-mouse antibody (Sangon Biotech, Cat. #D110087, 1:10000). For quantification, GAPDH or alpha tubulin were used as internal control. For antibodies used in this study refer to Supplementary Data 5.
RNA interference
For RNAi knockdown experiments, cells were transfected with indicated siRNAs using Lipofectamine RNAiMAX Transfection Reagent (Thermo Fisher Scientific, Cat. #13778150) according to the manufacturer’s instructions. Samples were collected after 48–72 h post transfection. MUS81 and POLD3 were depleted using an equimolar of 3 pooled siRNAs. POLQ was depleted using an equimolar of 2 pooled siRNAs. For siRNAs used in this study refer to Supplementary Data 2.
Quantitative RT-PCR
Total RNA was extracted using a FastPure Cell/Tissue total RNA isolation kit (Vazyme, Cat. #RC112-01) according to the manufacturer’s instructions. cDNA was synthesized using the first Strand cDNA Synthesis SuperMix for qPCR Kit (Yeasen, Cat. #11141ES60) following the manufacturer’s instruction. GAPDH was used as internal control. Real-time PCR was performed using a qPCR SYBR Green Master Mix (Yeasen, Cat. #11201ES08) on a CFX96 real-time system (Bio-Rad). Relative mRNA expression was calculated as 2−∆∆Ct. At least three independent replicates were performed for each experiment. Detailed primer information is listed in Supplementary Data 2.
Comet assays
The comet assay was performed as previously described in ref. 70. In brief, the lysis solution was prepared and chilled at 4 °C for before use. Agarose was melted in a water bath of boiling water for 5 min and then cooled in a 37 °C water bath for at least 20 min. Cells (1× 105 mL−1) were combined with molten agarose at a ratio of 1:10 (v/v), and 50 μL was placed onto the comet slide. The slides were placed in a 4 °C refrigerator for 10 min before immersion in lysis solution for 1 h, followed by balancing in electrophoresis buffer (pH 13.0 for alkaline comet assay, pH 7.5 for neutral comet assay) for 30 min. The slides were then subjected to electrophoresis at 25 V for 40 min. Thereafter, the samples were dried at 37 °C for 10 min and stained with 50 μg/mL propidium Iodide (PI) (Sigma-Aldrich, Cat. #P4864-10ML) for 10 min. Images were captured under a 20x objective lens of Nikon A1 microscope. The tail moment was analyzed using the ImageJ opencomet plugin.
Mitotic DNA synthesis (MiDAS) and mitotic DNA damage
The MiDAS assay was performed as previously described in ref. 8. Briefly, asynchronously growing cells were synchronized in late G2 phase of the cell cycle by incubation with 9 μM RO-3306 (Beyotime, Cat. #SC6673-10mM) for 8 h. As stated, 0.4 μM aphidicolin was added for 48 h. Cells arrested in G2 were subsequently washed with pre-warmed PBS for 5 min and allowed to progress in pre-warmed medium with 20 μM EdU (5-ethynyl-2′-deoxyuridine; Thermo Fisher Scientific, Cat. #C10638) at 37 °C under a humidified atmosphere containing 5% CO2 for a further 30 min into prometaphase before mitotic shake-off. To collect mitotic cells, pelleted prometaphase cells were re-seeded onto coverslips coated with Poly-D-lysine for 5 min before fixation. EdU was stained using Click-IT Plus EdU Imaging Kit (Thermo Fisher Scientific, Cat. #C10638) according to the manufacturer’s instruction. Mitotic DNA damage was indicated by further incubation of the coverslips with rabbit anti-H2AX p-S139 (CST, Cat. #2577 L, 1:400) for immunofluorescence.
Cell death analysis
Flow cytometric analysis was conducted to assess the cell fate of POLQ−/− cells under treatment of DMSO or APH for 48 h. Cells were harvested using EDTA-free trypsin and stained utilizing the Annexin V-FITC Apoptosis Detection Kit (Beyotime, Cat. #C1062M) according to the manufacturer’s protocol. Briefly, 195 μL Annexin V-FITC binding solution was used to resuspend the cells gently. Then, 5 μL Annexin V-FITC staining solution and 10 μL propidium iodide (PI) staining solution were added, respectively. After gentle mixing and incubation at room temperature in dark for 20 min, cells were incubated in an ice bath. Flow cytometry analysis was performed using a CytoFLEX LX flow cytometer (Beckman Coulter). More than 10,000 cells were analyzed for each experimental condition. Data were analysed using FlowJo software, version 10.6.2.
CRISPR-Cas9-mediated gene editing
To knock out genes, RPE-1 TP53−/− cell lines were transfected with pSpCas9-T2A-GFP-2×sgRNA, modified from PX458 (Addgene, Cat. #48138) using Lipo8000 (Beyotime, Cat. #C0533). Upon transfection, cells were grown in an antibiotic-free medium for 48 h, followed by single-cell sorting by FACSAria III (BD). The cells were sorted into 96-well plate using GFP signal and cultured for 1 ~ 2 weeks. The cells in a 96-well plate were divided into two 96-well plates, one for PCR screening and one for clonal expansion. For PCR and genotyping, DNA was extracted by adding 30 μL 50 mM NaOH solution per well, followed by heating at 95 °C for 10 min. Next, 2.5 μL 1 M Tris-HCl pH 7.5, was added into each well. Two microliters of the extracted DNA were used to perform PCR with DreamTaq Green PCR premix (Thermo Fisher Scientific, Cat. #K1081). To further validate the knockout clones, genomic DNA of the expanded clones was extracted using a Genomic DNA purification Kit (Thermo Fisher Scientific, Cat. #K0512) for PCR. PCR products were run on an agarose gel and purified using PCR gel purification kit (Thermo Fisher Scientific, Cat. #K0701) for Sanger sequencing (Genewiz). Genotyping was performed using Synthego ICE Analysis Tool at https://www.synthego.com/products/bioinformatics/crispr-analysis.
To generate POLQ knockout cells, dual sgRNAs (5′-GGAAGGCTTTTAGGTCAGTA-3′ and 5′-GGCAACGGGGGCAGCTCCGC-3′) were designed to delete a 522-bp region from exon 1 of POLQ71. To generate BRCA2 knock-out cells, dual sgRNA71 (5′-GGTAAAACTCAGAAGCGC-3′ and 5′-GCAACACTGTGACGTACT-3′) were designed to delete a 276-bp region containing the start codon in exon 2. The primers for CRISPR or genotyping refer to Supplementary Data 2.
Cell proliferation and viability assay
To detect cell proliferation, 1000 cells are seeded per well in 96-well plate and imaged using 10x (NA = 0.7) objective lens in the IncuCyte system (Sartorius, Germany). Phase contrast images were collected every 3 h in one week to calculate the confluence automatically according to the manufacturer’s instructions72. For cell viability assay, cells were plated in 96-well plate (2000 cells/well). After 48 h, CCK-8 solution (Beyotime, Cat. #C0039) was added into each well and incubated for 2 h. Finally, the absorbance was measured at 450 nm. To test the effect of drugs, cells were seeded in a 96-well plate at a density of 1000 cells per well overnight. Then cells were treated with DMSO or varying concentrations of Olaparib (CST, Cat. #93852 s), ART558 (WuXi AppTec, Cat. #SZWLAB-20210707) or ATR inhibitor VE-821 (Sigma-Aldrich, Cat. #SML1415) for 48 h. Cell viability was determined using CellTiter-Glo® 2.0 Assay kit (Promega, Cat. #G9241) according to the manufacturer’s instruction. Absorbance at a wavelength of 520 nm for each sample was recorded using a Synergy H1 Microplate Reader (BioTek, USA). The surviving fraction of drug-treated cells was normalized to that of DMSO-treated control. At least three independent experiments were performed.
CUT&Tag assay and sequencing
CUT&Tag (cleavage under targets and tagmentation) was performed using Hieff NGS® G-Type In-Situ DNA Binding Profiling Library Prep Kit (Yeasen, Cat. #12598ES48) against Lamin A/C and Lamin B1 following the manufacture’ s instruction. Briefly, for each sample, 1 × 105 RPE-1 or HCT-116 cells were harvested after treatment with DMSO or 0.4 μM APH for 48 h. Cells were washed three times in PBS, resuspended gently in the wash buffer, and incubated with the concanavalin A-coated magnetic beads pre-washed with the binding buffer. Concanavalin A beads-coupled cells were collected and incubated with 2 μg of anti-Lamin A/C (Santa Cruz, Cat. #sc-7292), anti-Lamin B1 (Abcam, Cat. #ab16048) or IgG control at 4 °C overnight. After removing unbound primary antibodies using a magnetic stand, the secondary antibodies, goat anti-rabbit IgG (H + L) (Yeasen, Cat. #34850ES60) or goat anti-mouse IgG (H + L) (Yeasen, Cat. #34851ES60) were incubated with the concanavalin A beads-coupled cells at room temperature for 1 h. Cells were washed 3 times with the wash buffer to remove the unbound antibodies, and incubated with pA/G-Transposome mix for 1 h at room temperature. To terminate tagmentation, termination solution and proteinase K were added to the samples and incubated at 55 °C for 30 min. DNA was purified using DNA-selection beads and eluted in 21 μL ultrapure water.
For library amplification, 8 μL of DNA was mixed with a uniquely barcoded i5 and a uniquely barcoded i7 primer, and amplified using 2x Ultima Amplification (Yeasen, Cat. #12598ES48). The library was then purified using DNA-selection beads and washed twice with 80% ethanol and eluted in 21 μL ultrapure water. Libraries were sequenced by Berry Genomic (Beijing, China) on an Illumina NovaSeq platform for 10 G per sample and 150-bp paired-end reads were generated.
CUT&Tag data analysis
Raw sequencing reads were first preprocessed using Ktrim software73 to remove sequencing adaptors and low-quality cycles. The preprocessed reads were then aligned to NCBI GRCh38 reference human genome using Bowtie2 software74. PCR duplicates (i.e., reads with identical start and end positions) were identified and removed using in-house programs. Reads from mitochondrial DNA were also removed and the remaining (“clean reads”) were kept for downstream analyses. For DSMO and APH treatments, data from input experiments were used as controls to call peaks for Lamin A/C and Lamin B1 using MACS2 software75, respectively. Peaks with adjusted p-values lower than 0.01 were considered statistically significant. CUT&Tag signals around the peaks were visualized using depTools software76.
Identification of RESSLADs
We first stitched the peaks in the Lamin A/C or Lamin B1 that were less than 20 kb in distance and only kept those longer than 100 kb after stitching. For each kept peak after stitching, we extracted the read numbers in the CUT&Tag experiments, which were further normalized using the clean read count of the corresponding experiments. The normalized read counts were compared between DMSO and APH experiments using t-tests for Lamin A/C and Lamin B1 (each with 3 replicates), respectively. The p-values were corrected using the Benjamini & Hochberg method, and peaks with adjusted p-values lower than 0.05 were called RESSLAD regions.
Chromatin immunoprecipitation (ChIP)-qPCR
Chromatin immunoprecipitation was performed using a ChIP Assay Kit (Beyotime, Cat. #P2078) following the manufacture’ s instruction. Briefly, RPE-1 p53−/− cells and RPE-1 p53−/− POLQ−/− cells were treated with or without APH (0.4 μM) for 48 h. Cells were cross-linked with 1% formaldehyde for 10 min at 37 °C. Reaction was terminated by Glycine, and cells were washed twice with cold PBS and then harvested in a SDS lysis buffer. Then, the Covaris S220 Focused Ultrasonicator (Covaris) was used to sonicate the sample until DNA fragmentation reached sizes of 200 bp to 1 kb. Protein A + G Agarose and Salmon Sperm DNA were used to preclear the whole cell lysate for 30 min at 4 °C. Chromatin immunoprecipitation (ChIP) was carried out overnight at 4 °C with 1 μg of anti-Lamin A/C (Santa Cruz, Cat. #sc-7292) or IgG control (Yeasen, Cat. #34851ES60). The beads were subjected to a washing process on four occasions according to the manufacturer’s protocol and eluted with a solution of 0.1 mol/L NaHCO3 and 1% SDS, followed by reverse cross-linking and phenol chloroform extraction. DNA fragments were precipitated by ethanol. A primer pair of RESSLAD #1 was used to amplify the target fragments with a primer pair of ACTIN gene as internal control. The values are normalized to WT with DMSO treatment after subtracting background of non-specific IgG. The sequence of primers used to identify fragments is listed in Supplementary Data 2.
For detection of mitotic DNA damage signal in RESSLADs by anti-γH2AX antibody, Hieff NGS® G-Type In-Situ DNA Binding Profiling Library Prep Kit (Yeasen, Cat. #12598ES48) was used following the manufacture’s instruction. Briefly, 1 × 105 mitotic RPE-1 cells were harvested by shake-off as described above for each sample after indicated treatments. A beads-coupled cells were collected and incubated with 2 μg of anti-γH2AX (Sigma, Cat. # JBW301) or IgG control (Yeasen, Cat. #34851ES60). The library amplification was referred to above. Primer pairs located in RESSLAD#1 (GPR153) and RESSLAD#2 (SLURP2) were used to amplify the target fragments with ACTIN gene as internal control. The values are normalized to DMSO group after subtracting background of non-specific IgG. The primers used are listed in Supplementary Data 2.
Statistical analysis
All statistical analyses were performed using GraphPad Prism 6. For bar graphs, results were shown as mean ± SD and the two-tailed unpaired Student’s t-test was used to determine the p-values, unless otherwise stated. For time-course curves or multi-drug treatments, two-way ANOVA analysis was performed to evaluate the statistical significance between groups. SuperPlots was used to display the entire dataset distribution and report statistics as previously described in ref. 77. Each experiment was independently performed at least three times, with similar results across replicates.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The CUT&Tag data generated in this study have been deposited in the GEO database under access sion code GSE271990. Source data are provided with this paper.
Code availability
The code used for the analysis of RESSLADs is available at the GitHub (https://github.com/hellosunking/RESS-LADs)78.
References
Macheret, M. & Halazonetis, T. D. DNA replication stress as a hallmark of cancer. Annu Rev. Pathol. 10, 425–448 (2015).
Techer, H., Koundrioukoff, S., Nicolas, A. & Debatisse, M. The impact of replication stress on replication dynamics and DNA damage in vertebrate cells. Nat. Rev. Genet 18, 535–550 (2017).
Cortez, D. Preventing replication fork collapse to maintain genome integrity. DNA Repair (Amst.) 32, 149–157 (2015).
Deng, L. et al. Mitotic CDK Promotes Replisome Disassembly, Fork Breakage, and Complex DNA Rearrangements. Mol. Cell 73, 915–929.e916 (2019).
Maciejowski, J., Li, Y., Bosco, N., Campbell Peter, J. & de Lange, T. Chromothripsis and Kataegis Induced by Telomere Crisis. Cell 163, 1641–1654 (2015).
Zhang, C.-Z. et al. Chromothripsis from DNA damage in micronuclei. Nature 522, 179–184 (2015).
Hustedt, N. & Durocher, D. The control of DNA repair by the cell cycle. Nat. Cell Biol. 19, 1–9 (2016).
Minocherhomji, S. et al. Replication stress activates DNA repair synthesis in mitosis. Nature 528, 286–290 (2015).
Bhowmick, R., Minocherhomji, S. & Hickson, I. D. RAD52 Facilitates Mitotic DNA Synthesis Following Replication Stress. Mol. Cell 64, 1117–1126 (2016).
Rogers, C. B. et al. Fanconi anemia-associated chromosomal radial formation is dependent on POLθ-mediated alternative end joining. Cell Rep. 42, 112428 (2023).
Heijink, A. M. et al. Sister chromatid exchanges induced by perturbed replication can form independently of BRCA1, BRCA2 and RAD51. Nat. Commun. 13, 6722 (2022).
Brambati, A. et al. RHINO directs MMEJ to repair DNA breaks in mitosis. Science 381, 653–660 (2023).
Mateos-Gomez, P. A. et al. Mammalian polymerase θ promotes alternative NHEJ and suppresses recombination. Nature 518, 254–257 (2015).
Wilson, T. E., Ahmed, S., Winningham, A. & Glover, T. W. Replication stress induces POLQ-mediated structural variant formation throughout common fragile sites after entry into mitosis. Nat. Commun. 15, 9582 (2024).
Huguet, F., Gokan, E., Foster, H. A., Amin, H. A. & Vagnarelli, P. Repo-Man/protein phosphatase 1 SUMOylation mediates binding to lamin A and serine 22 dephosphorylation. Open Biol. 12, 220017 (2022).
van Steensel, B. & Belmont, A. S. Lamina-Associated Domains: Links with Chromosome Architecture, Heterochromatin, and Gene Repression. Cell 169, 780–791 (2017).
Liu, S. et al. Nuclear envelope assembly defects link mitotic errors to chromothripsis. Nature 561, 551–555 (2018).
Ungricht, R. & Kutay, U. Mechanisms and functions of nuclear envelope remodelling. Nat. Rev. Mol. Cell Biol. 18, 229–245 (2017).
Haraguchi, T. et al. Live cell imaging and electron microscopy reveal dynamic processes of BAF-directed nuclear envelope assembly. J. Cell Sci. 121, 2540–2554 (2008).
von Appen, A. et al. LEM2 phase separation promotes ESCRT-mediated nuclear envelope reformation. Nature 582, 115–118 (2020).
Cho, S. et al. Mechanosensing by the Lamina Protects against Nuclear Rupture, DNA Damage, and Cell-Cycle Arrest. Dev. Cell 49, 920–935.e925 (2019).
Earle, A. J. et al. Mutant lamins cause nuclear envelope rupture and DNA damage in skeletal muscle cells. Nat. Mater. 19, 464–473 (2020).
Nader, G. P. F. et al. Compromised nuclear envelope integrity drives TREX1-dependent DNA damage and tumor cell invasion. Cell 184, 5230–5246.e5222 (2021).
Sheaff, R., Ilsley, D. & Kuchta, R. Mechanism of DNA polymerase alpha inhibition by aphidicolin. Biochemistry 30, 8590–8597 (1991).
Halfmann, C. T. et al. Repair of nuclear ruptures requires barrier-to-autointegration factor. J. Cell Biol. 218, 2136–2149 (2019).
Xu, B. et al. Replication stress induces micronuclei comprising of aggregated DNA double-strand breaks. PLoS One 6, e18618 (2011).
Di Micco, R. et al. Oncogene-induced senescence is a DNA damage response triggered by DNA hyper-replication. Nature 444, 638–642 (2006).
Segeren, H. A., van Liere, E. A., Riemers, F. M., de Bruin, A. & Westendorp, B. Oncogenic RAS sensitizes cells to drug-induced replication stress via transcriptional silencing of P53. Oncogene 41, 2719–2733 (2022).
Kovacs, M. T. et al. DNA damage induces nuclear envelope rupture through ATR-mediated phosphorylation of lamin A/C. Mol. Cell 83, 3659–3668.e3610 (2023).
Vassilev, L. T. et al. Selective small-molecule inhibitor reveals critical mitotic functions of human CDK1. Proc. Natl Acad. Sci. USA 103, 10660–10665 (2006).
Montagnoli, A. et al. A Cdc7 kinase inhibitor restricts initiation of DNA replication and has antitumor activity. Nat. Chem. Biol. 4, 357–365 (2008).
Wilhelm, T. et al. Mild replication stress causes chromosome mis-segregation via premature centriole disengagement. Nat. Commun. 10, 3585 (2019).
Burrell, R. A. et al. Replication stress links structural and numerical cancer chromosomal instability. Nature 494, 492–496 (2013).
Somyajit, K. et al. Redox-sensitive alteration of replisome architecture safeguards genome integrity. Science 358, 797–802 (2017).
Chu, Q. et al. Oligomeric CHMP7 mediates three-way ER junctions and ER-mitochondria interactions. Cell Death Differ. 30, 94–110 (2022).
Gu, M. et al. LEM2 recruits CHMP7 for ESCRT-mediated nuclear envelope closure in fission yeast and human cells. Proc. Natl Acad. Sci. USA 114, E2166–e2175 (2017).
Lai, X. et al. MUS81 nuclease activity is essential for replication stress tolerance and chromosome segregation in BRCA2-deficient cells. Nat. Commun. 8, 15983 (2017).
Pepe, A. & West Stephen, C. MUS81-EME2 Promotes Replication Fork Restart. Cell Rep. 7, 1048–1055 (2014).
Tan, Q. & Xu, X. MUS81 UFMylation at K400 promotes cell survival in response to camptothecin-induced replication stress. Genome Instab. Dis. 5, 154–163 (2024).
Xing, M. et al. Replication Stress Induces ATR/CHK1-Dependent Nonrandom Segregation of Damaged Chromosomes. Mol. Cell 78, 714–724.e715 (2020).
Berman, B. P. et al. Regions of focal DNA hypermethylation and long-range hypomethylation in colorectal cancer coincide with nuclear lamina-associated domains. Nat. Genet 44, 40–46 (2011).
Boteva, L. et al. Common Fragile Sites Are Characterized by Faulty Condensin Loading after Replication Stress. Cell Rep. 32, 108177 (2020).
Liu, S. et al. Phosphorylation of Lamin A/C regulates the structural integrity of the nuclear envelope. J. Biol. Chem. 301, 108033 (2025).
Heald, R. & McKeon, F. Mutations of phosphorylation sites in lamin A that prevent nuclear lamina disassembly in mitosis. Cell 61, 579–589 (1990).
Toledo, L. I. et al. ATR prohibits replication catastrophe by preventing global exhaustion of RPA. Cell 155, 1088–1103 (2013).
Pang, Y. et al. Synthetic lethality in personalized cancer therapy. Genome Instab. Dis. 4, 121–135 (2022).
Zatreanu, D. et al. Polθ inhibitors elicit BRCA-gene synthetic lethality and target PARP inhibitor resistance. Nat. Commun. 12, 3636 (2021).
Ceccaldi, R. et al. Homologous-recombination-deficient tumours are dependent on Polθ-mediated repair. Nature 518, 258–262 (2015).
Coy, S. et al. 2D and 3D multiplexed subcellular profiling of nuclear instability in human cancer. Preprint at bioRxiv. https://doi.org/10.1101/2023.11.07.566063 (2023).
Ying, S. et al. MUS81 promotes common fragile site expression. Nat. Cell Biol. 15, 1001–1007 (2013).
Terasawa, M., Shinohara, A. & Shinohara, M. Double-strand break repair-adox: Restoration of suppressed double-strand break repair during mitosis induces genomic instability. Cancer Sci. 105, 1519–1525 (2014).
Wu, W. et al. Mitotic DNA synthesis in response to replication stress requires the sequential action of DNA polymerases zeta and delta in human cells. Nat. Commun. 14, 706 (2023).
Groelly, F. J. et al. Mitotic DNA synthesis is caused by transcription-replication conflicts in BRCA2-deficient cells. Mol. Cell 82, 3382–3397.e3387 (2022).
Chandramouly, G. et al. Polλ promotes microhomology-mediated end-joining. Nat. Struct. Mol. Biol. 30, 107–114 (2023).
Gerace, L. & Blobel, G. The nuclear envelope lamina is reversibly depolymerized during mitosis. Cell 19, 277–287 (1980).
Georgatos, S. D., Pyrpasopoulou, A. & Theodoropoulos, P. A. Nuclear envelope breakdown in mammalian cells involves stepwise lamina disassembly and microtubule-drive deformation of the nuclear membrane. J. Cell Sci. 110, 2129–2140 (1997).
Joo, Y. K. et al. ATR promotes clearance of damaged DNA and damaged cells by rupturing micronuclei. Mol. Cell 83, 3642–3658.e3644 (2023).
Young, A. M., Gunn, A. L. & Hatch, E. M. BAF facilitates interphase nuclear membrane repair through recruitment of nuclear transmembrane proteins. Mol. Biol. Cell 31, 1551–1560 (2020).
Kono, Y. et al. Nucleoplasmic lamin C rapidly accumulates at sites of nuclear envelope rupture with BAF and cGAS. J. Cell Biol. 221, e202201024 (2022).
Macheret, M. et al. High-resolution mapping of mitotic DNA synthesis regions and common fragile sites in the human genome through direct sequencing. Cell Res. 30, 997–1008 (2020).
Amiad-Pavlov, D. et al. Live imaging of chromatin distribution reveals novel principles of nuclear architecture and chromatin compartmentalization. Sci Adv 7 (2021).
Zheng, X. et al. Lamins Organize the Global Three-Dimensional Genome from the Nuclear Periphery. Mol. Cell 71, 802–815. e807 (2018).
Ikegami, K., Secchia, S., Almakki, O., Lieb, J. D. & Moskowitz, I. P. Phosphorylated Lamin A/C in the Nuclear Interior Binds Active Enhancers Associated with Abnormal Transcription in Progeria. Dev. Cell 52, 699–713.e611 (2020).
Cheedipudi, S. M. et al. Genomic Reorganization of Lamin-Associated Domains in Cardiac Myocytes Is Associated With Differential Gene Expression and DNA Methylation in Human Dilated Cardiomyopathy. Circ. Res 124, 1198–1213 (2019).
Wang, Y. et al. Lamin A/C-dependent chromatin architecture safeguards naive pluripotency to prevent aberrant cardiovascular cell fate and function. Nat. Commun. 13, 6663 (2022).
Denais, C. M. et al. Nuclear envelope rupture and repair during cancer cell migration. Science 352, 353–358 (2016).
Le Berre, M., Aubertin, J. & Piel, M. Fine control of nuclear confinement identifies a threshold deformation leading to lamina rupture and induction of specific genes. Integr. Biol. (Camb.) 4, 1406–1414 (2012).
Kochin, V. et al. Interphase phosphorylation of lamin A. J. Cell Sci. 127, 2683–2696 (2014).
Kumar, S., Ciraolo, G., Hinge, A. & Filippi, M. D. An efficient and reproducible process for transmission electron microscopy (TEM) of rare cell populations. J. Immunol. Methods 404, 87–90 (2014).
Olive, P. L. & Banáth, J. P. The comet assay: a method to measure DNA damage in individual cells. Nat. Protoc. 1, 23–29 (2006).
Zhou, J. et al. A first-in-class Polymerase Theta Inhibitor selectively targets Homologous-Recombination-Deficient Tumors. Nat. Cancer 2, 598–610 (2021).
Wolfe, A., O’Clair, B., Groppi, V. E. & McEwen, D. P. Pharmacologic Characterization of a Kinetic In Vitro Human Co-Culture Angiogenesis Model Using Clinically Relevant Compounds. SLAS Discov. 18, 1234–1245 (2013).
Sun, K. Ktrim: an extra-fast and accurate adapter- and quality-trimmer for sequencing data. Bioinformatics 36, 3561–3562 (2020).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Ramírez, F., Dündar, F., Diehl, S., Grüning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res 42, W187–W191 (2014).
Lord S. J., Velle K. B., Mullins R. D., Fritz-Laylin L. K. SuperPlots: Communicating reproducibility and variability in cell biology. Journal of Cell Biology 219 (2020).
Sun, K. Code of identification of RESSLADs. Zenodo. https://doi.org/10.5281/zenodo.16881072 (2024).
Acknowledgements
We thank members of the Deng and Ji laboratories for helpful discussions, David Pellman and Johannes Walter for valuable supports at the early stage of this study. We thank Yisui Xia, Lingluo Chu and Shangming Tang for critical reading of the manuscript. We thank Jian Liu, Shaohong Rao, and Chunzhuo Liu and the Bio-imaging Core Facility at SZBL, particularly Zhenglong Sun, Mei Yu and Shixian Huang for technical helps. This work is supported by the National Natural Science Foundation of China (32270779), Shenzhen Medical Research Funds (B2301007), National Key R&D Program of China (2022YFA1302800), Guangdong Special Support Plan (2023TQ07A226), Proof of Concept Fund of Shenzhen Bay Laboratory (C1022426003) and Shenzhen Bay Laboratory Start-up Fund (S201100001) to L.D., National Natural Science of China (22207131) to Y.H., Spark Trial Program from Bayjayray Inc. to Y.L. and National Natural Science of China (92354304; 32371343; 32122025) to W.J.
Author information
Authors and Affiliations
Contributions
L.D. and G.Y. conceived the project and designed the experiments. G.Y. Y.H. Y.Z. D.L. W.C. Q.L.G. Q.G. S.H. and C.S. performed the experiments. F.L. H.Z. Q.P. and K.S. provided helps on CUT&Tag analysis. W.J. helped analyze the imaging data and provided valuable suggestions. G.Y. Y.H. Y.Z. Y.L. and L.D. interpreted the results and prepared the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ye, G., He, Y., Zhang, Y. et al. Mitotic DNA repair by TMEJ suppresses replication stress-induced nuclear envelope reassembly defect. Nat Commun 16, 8836 (2025). https://doi.org/10.1038/s41467-025-63942-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-63942-w