Abstract
Voltage-gated potassium ion (K+) channels perform critical roles in many physiological processes, while gain- or loss-of-function mutations lead to life-threatening pathologies. Here, we establish the high-resolution structure of a pivotal intermediate state of the Kv7.1 (KCNQ1) channel using cryogenic electron microscopy. The 3.53 Å resolution structure reveals straightened upper S1 and S2 voltage sensor helices, distancing them from the pore filter helix compared to fully activated channels. The outward translation of the S4 voltage sensor is essentially complete in this intermediate state, and the S4-S6 helices and the S4-S5 linker do not change position significantly between intermediate and activated states. The PIP2 ligand can bind in both states. Movement of S1 and S2 helices towards the filter helix from intermediate to activated states may explain smaller components of KCNQ1 voltage sensor fluorescence, differential Rb+/K+ selectivity, and pharmacological responses to activators and inhibitors. Single channel recordings and the location of long QT mutations suggest the potential physiological and disease importance of the intermediate state.
Similar content being viewed by others
Introduction
Voltage-gated potassium ion (Kv) channels are highly specialized proteins that regulate excitability throughout the human body1. The human Kv7 channel subfamily, and hKv7.1 (hKCNQ1) channels in particular, are good examples of the widespread functionality of Kv channels in general, with critical roles in the repolarization of the heart2,3, fluid transport in the inner ear4, and transmembrane flux regulation in epithelial tissues2,5,6. Their importance is underscored by the existence of clinical syndromes such as deafness7, long and short QT syndromes8,9, and diabetic nephropathy10, arising from polymorphisms or mutations in hKCNQ1 channels and their accessory subunits.
KCNQ1 alpha subunits assemble as tetramers, with each subunit comprising six transmembrane helices (TMHs, S1 to S6). While the S1-S4 helices of each subunit comprise the voltage sensor domain (VSD), the S5-S6 helices of all four subunits come together to form the pore domain (PD, Fig. 1a)11,12, and as KCNQ1 is a domain-swapped channel, the VSD of one subunit interacts with the pore helices of an adjacent subunit. In addition to calmodulin (CaM), a structural obligate of the channel that binds its C-terminus to promote proper channel folding and gating13,14, PIP2 binding is essential for electro-mechanical coupling between the VSD and the pore, to allow ion conduction15,16,17,18. Additional accessory KCNE1-5 β-subunits can assemble with the channel complex to modulate the current kinetics in ways unique to each channel complex that suit their specific physiological roles throughout the body2,3,6,19,20,21.
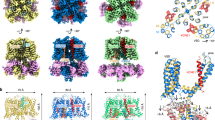
a KCNQ1 transmembrane helices S1-S6, S4-S5 linker (S4-S5L), filter helix (FH), selectivity filter (SF), and intracellular helices HA-HD that bind calmodulin (CaM). S1-S4 comprise the VSD, while S5, S6, and FH form the PD. Gating residues in Xenopus (+10 for hKCNQ1): E1 (xE150), E2 (xE160), D3 (xD192), R1 (xR218), R2 (xR221), Q3 (xQ224), R4 (xR227), H5 (xH230), and R6 (xR233). b Cartoon of S2 and S4 orientations in resting, intermediate, and activated states, showing xE150 interactions with positive charges R1, R2, and R4. F0 refers to xF157, part of the CTC. c tsA201 cells expressing human (h) and Xenopus (x) variants of WT and E1R/R2E KCNQ1, transfected with either hWT-GFP, xWT-GFP, xE1R/R2E-GFP, hE1R/R2E + GFP, hE1R/R4E + GFP or KCNE1-hWT + GFP. Currents from -80 to +60 mV for 4 s, then 2 s at −40 mV. d Tail current G-V relationships. WT (hWT, Boltzmann V1/2 = −23.3 ± 1.2 mV, k = 9.0 ± 1.1 mV, n = 7; and xWT, V1/2 = −30.8 ± 2.3 mV, k = 7.9 ± 0.99 mV, n = 5). E1R/R2E, linear regression slopes were not different from zero (hE1R/R2E, p = 0.99, n = 5; and xE1R/R2E, p = 0.94, n = 5). e, f Rb+/K+ permeability ratio, tail currents at −60 mV after 5 s + 60 mV pulse. e Currents in 140 mM K+ (black) and Rb+ (color). Dotted line: 0 pA. f Summary, log base 2 y-scale: hWT, 2.51 ± 0.22, n = 6; xWT, 2.43 ± 0.32, n = 6; hE1R/R2E, 1.95 ± 0.10, n = 8; xE1R/R2E, 1.7 ± 0.20, n = 6; KCNE1-hWT, 0.60 ± 0.06, n = 6. Using ANOVA, multiple pairwise comparison and a Bonferroni test, KCNE1-hWT p < 0.0001 vs. hWT and p = 0.0001 vs. hE1R/R2E). g Currents before (black) and with 300 nM XE991 (color). h Peak reduction fit to Hill equation. IC50s for: hWT (67 nM, n = 4), xWT (119 nM, n = 4), hE1R/R2E (235 nM, n = 4–5), xE1R/R2E (166 nM, n = 4), and hE1R/R4E (~ 135 μM, n = 3-4). No differences between WT and E1R/R2E (one-way ANOVA). All error bars in d, f, h, are mean ± SEM.
The mechanistic pathways underlying the voltage sensitivity of tetrameric Kv channels and the coupling of voltage activation to gate opening and ion conduction have been of interest since the first quantitative descriptions of membrane excitability22,23,24,25,26, but the quest to understand structurally and electrically discrete intermediate steps in the Kv channel activation pathway is hampered by the lack of easily-studied voltage sensor states intermediate between resting and fully-activated27. Unusually, the presence of a stable, functionally distinct, and physiologically relevant intermediate state is a hallmark of KCNQ117,28,29. Hence, VSD activation occurs in two formally defined steps: first from the resting to the intermediate state, and then to the activated state (Fig. 1b)28,29,30. These steps involve the translocation of a series of positive gating charges in S4 across the electric field. For human KCNQ1 (hKCNQ1), most of the gating charge is concentrated in R228 (R1; which corresponds to R218 in Xenopus KCNQ1, (xKCNQ1)), R231 (R2; R221 in xKCNQ1) and R237 (R4; R227 in xKCNQ1)31. The positive charges are stabilized by negative charges in S2 (E1: E160 and E2: E170 in hKCNQ1) and S3 (D3: D202 in hKCNQ1). A highly conserved F0 (F167 in hKCNQ1) forms a hydrophobic seal (Fig. 1, Supplementary Fig. 1). Together, E2, D3, and F0 form the charge transfer center (CTC) that facilitates the nearly switch-like movement of positive charges across the electrical field11,32. Mutations reversing the charges in specific gating residue pairs have been engineered to stabilize hKCNQ1 in specific VSD states. The charge reversal mutations E160R/R231E (hE1R/R2E) are thought to lock the channel in the intermediate state, whereas the E160R/R237E (hE1R/R4E) mutations hold it in the activated state28,29,30. These mutants show different pore properties, including higher Rb+/K+ permeability and sensitivity to the pharmacological compound XE991 for hE1R/R2E compared to hE1R/R4E (Fig. 1, Supplementary Table 1)29, and have been accepted as bona fide representations of the intermediate and fully activated states, respectively17,28. Note that these are VSD mutants so that the channel pore gate may be closed in intermediate and activated voltage sensor states, or open in the presence of PIP2.
In recent years, cryo-EM structures of the activated states of KCNQ111,33, and KCNQ1 + KCNE314 channels have been obtained, and recently, low-resolution intermediate and resting state structures have been reported18. Here, we characterize the Xenopus KCNQ1 E150R/R221E intermediate state structure (xE1R/R2E, equivalent to human E1R/R2E) at high resolution and show that the electrophysiological properties of xE1R/R2E mirror those of hKCNQ1 channels in the intermediate state. The xE1R/R2E structures show marked differences in the orientation of the S1 and S2 TMHs within the voltage sensor domain in comparison with the fully activated xKCNQ1 structure, while the overall S4 position is similar. We address the relationship between our intermediate state structure of KCNQ1, represented by the E1R/R2E mutant, to prior KCNQ1 structures. We suggest how fluorescence changes and gating currents that are observed upon activation of hKCNQ1 channels may be explained in terms of the intermediate activated state structure34,35, and how the E1R/R2E structure helps us understand the unusual permeability and pharmacological properties of the intermediate and activated states.
Results
Functional validation of xE1R/R2E as a bona fide model of the intermediate state
The charge reversal mutations E150R and R221E (xE1R/R2E) in the S2 and S4 TMHs were generated in the xKCNQ1-GFP linked construct previously used for structural studies (referred to below as xWT, Xenopus wild type, residues 67-61033, see Methods). Note that in these experiments, the human (h) channel construct hWT was GFP-tagged, while the KCNE1-hWT (KCNE1 linked to KCNQ1), hE1R/R2E, and hE1R/R4E constructs were instead co-expressed with GFP to aid identification of transfected cells (see Methods). Whole cell patch clamp recordings of cells expressing the xE1R/R2E construct show a constitutively open channel with similar kinetics and voltage dependence to the homologous mutation in the full-length hKCNQ1 channel (hE1R/R2E, Fig. 1c). This contrasts with the hWT and xWT channels which show voltage-dependent activation with half-activation voltages (V1/2s) of -23.3 and -30.8 mV, respectively (Fig. 1d). Further validation of the xKCNQ1 and xE1R/R2E constructs was carried out by comparing their Rb+ currents during repolarizing pulses (Fig. 1e-f) and XE991 sensitivity (Fig. 1g-h) to the full-length human isoforms. Almost identical relative Rb+/K+ current ratios were measured at the tail currents for hE1R/R2E (1.95 ± 0.10) and xE1R/R2E (1.74 ± 0.20). These were, in turn, similar to xWT (2.43 ± 0.32) and hWT (2.51 ± 0.22), but quite different to KCNE1-hWT (0.60 ± 0.06, Fig. 1f)29,36. Likewise, xE1R/R2E showed an XE991 sensitivity similar to its human counterpart (IC50 = 235 nM and 166 nM for hE1R/R2E and xE1R/R2E, respectively), and close to that of WT (IC50 = 67 nM for hWT and 119 nM for xWT), but much higher sensitivity than hE1R/R4E (IC50 above 30 µM, the highest concentration tested, Fig. 1h, Supplementary Table 1). Overall, the functional similarity between Xenopus and human E1R/R2E demonstrates that xE1R/R2E replicates the electrophysiological properties of the intermediate state of hKCNQ1, validating its use as a structural model.
Structures of xE1R/R2E and xWT KCNQ1
The channel structures were determined along with CaM, in the presence of 0.03% (w/v) glyco-diosgenin (GDN) and 4 mM EGTA. This protocol resulted in higher yield, purity, number of particles and higher resolution structures in comparison to the n-dodecyl-β-D-maltoside (DDM)/cholesteryl hemisuccinate (CHS) that we used before33. Local resolution estimation maps reveal consistent high-resolution density for PDs (Fig. 2) and less consistent resolution for the VSDs and the intracellular parts, in common with previous KCNQ1 maps11,33. The xWT structure was elucidated at 3.45 Å with local resolution between 2.9-3.3 Å for the PD and 3.3-6.0 Å for the VSD (Fig. 2a, Supplementary Fig. 2, Supplementary Table 2 and detailed sample maps in Supplementary Fig. 3). This structure was found to be very similar to the xWT structure we described previously in DDM/CHS33 with a RMSD of 1.44 Å between the two protomers for all the Cα atoms and 0.68 Å for the Cα atoms in the PD (residues 249-349). The xE1R/R2E structure was determined at 3.5 Å resolution with local resolution between 2.8-3.3 Å for the PD and 3.5-6.0 Å for the VSD (Fig. 2a, Supplementary Fig. 4, Supplementary Table 2 and detailed sample maps in Supplementary Fig. 5). In both maps, the lower parts of the VSD are better resolved than the upper sections (Supplementary Figs. 2b, 4d), as in other previously reported KCNQ1 structures11,14,33,37. However, the local resolution for the upper part of the VSD is significantly lower for xE1R/R2E, suggesting that the charge reversal mutation also unexpectedly increases the conformational flexibility of this region. The xWT and xE1R/R2E structures adopt a domain-swapped configuration where the VSD of the one subunit interacts with the PD of the adjacent subunit (PD’, Fig. 2b), rather than its own. CaM, wrapped around the intracellular helices HA and HB (Fig. 1a), is attached to the VSD of each subunit (Fig. 2b).
a Cryo-EM density maps for WT and xE1R/R2E colored according to the local resolution (legend below, scale in Å). Density maps are viewed from side and above (lower panel). CaM, anchored below the TMHs is indicated. b Model structures show two of the four domain-swapped subunits colored in different shades of blue for the xWT and orange for the xE1R/R2E. CaM, which co-purifies with KCNQ1, is colored yellow and the other two subunits are shown in white. The channels have a configuration where the VSD of one subunit (VSD, light color) interacts with the PD of the adjacent subunit (PD’, dark color).
The S1 and S2 helices sit apart from the filter helix in the intermediate state
An overview of the xE1R/R2E subunit colored according to the RMSD values calculated between the two aligned models (xE1R/R2E and xWT) is shown in the center panel of Fig. 3, while the surrounding panels show detailed comparisons of the different regions of the protomers. The VSD is comprised of S0-S2 helices, the S2-S3 linker which interacts with CaM, and helices S3 and S4 (Fig. 3a). The upper halves of the S1 and S2 helices adopt a curved conformation in the xWT structure (blue) but appear much straighter in the xE1R/R2E structure and thus distant from the filter helix (Fig. 3a, orange; Movie 1). In comparison to the xWT structure, the lower and upper parts of S1 in the xE1R/R2E structure are shifted laterally by ~1.4 Å and ~6.0 Å, respectively, and the helix is tilted by ~12.0°. Accordingly, the lower and upper parts of S2 are shifted ~0.7 Å and ~6.0 Å, respectively, and the helix is tilted by ~11.5°. In the xE1R/R2E structure, the S0, S2-S3L and S3 helices of the VSD occupy the same 3D space as they do in the xWT structure relative to the pore domain (Fig. 3a) and thus are already in a fully activated state. The S4 and the S4-S5 linker (S4-S5L) appear to largely preserve the activated state conformation despite a small rigid shift downwards (~ 1 Å) and a ~ 2o tilt (measured at residue xG235) of the S4-S5L in xE1R/R2E.
The protomers were superposed based on the pore domains (residues 249-349), which resulted in an RMSD of 1.92 Å for all the Cα atoms. The shaded center panel shows the xE1R/R2E monomer colored according to the RMSD values calculated between the two aligned models. Blue indicates small movements and high similarity to the xWT structure, while orange represents larger movements highlighting the profound changes in the VSD in comparison to the other domains of the protein. a, left The VSD rotated by 30 degrees with regards to the center panel showing only the S0-S2 helices, xWT in blue and xE1R/R2E in orange. The Cα-Cα distances between the two structures are measured at the black lines and the angles at the dashed magenta curved line. a, middle The S2-S3L, and S3-S4 helices of the VSD and the S4-S5L. a, right An enlarged view of the S4/S4-S5L loop. The distance between the Cα atoms of xG235 is shown in black and the angle in magenta curved dashed line. b-e show the superposed structures focusing on the different domains. b The pore domain rotated by 90 degrees for clarity. c Distances between Cα atoms of residues Gly335, Ser339 and Leu343, from subunits across from one another, reveal a closed inner gate for both xE1R/R2E and xWT. d The intracellular helices HA and HB that interact with CaM (not shown in the panel for clarity), and HC. e CaM is attached to the VSD through interactions with the S2-S3L loop of the VSD in all structures. In the inset is an enlarged representation of the interactions between the residues of CaM, shown as ball and stick, and the VSD, shown as sticks.
The RMSD between the protomers of the xE1R/R2E and xWT structures was only 0.37 Å for the Cα atoms in the PD (residues 249-349), suggesting that no substantial movements occur in the pore domain helices S5 and FH, the P-loop, the selectivity filter, helix S6 (Fig. 3b). Both structures have a closed inner gate formed by a bundle crossing of the S6 helices at the intracellular end of the pore (key residues; xG335, xS339, xL343, Fig. 3c) with only minor differences between xWT and xE1R/R2E, which extends to the C-terminus, helices HA-HC (Fig. 3d). Unsurprisingly therefore, CaM remains attached to the S2-S3 linker (S2-S3L) of the VSDs (Fig. 3e) as it does in all closed channel structures obtained in the absence of PIP233.
The architecture of the charge transfer center (CTC) in the intermediate state
A closer examination of the density maps for the xWT (blue) and xE1R/R2E (orange) S4 helix is shown in Fig. 4, and a density-modified (DM) map generated from the xE1R/R2E cryo-EM map is also shown. The DM map was generated using EMReady38, which applies deep learning to enhance structural details and improve map interpretability. Comparison of the maps for the S4 residues between the xWT and xE1R/R2E clearly shows a rearrangement in the lower part of the S4 with displacement and rotation of different parts of the S4 helix backbone (Fig. 4a, b, region between the horizontal dashed lines). In xE1R/R2E, the backbone of R4 is pushed down by ~ 2.7 Å and the side chain occupies the position of the xWT side chain of H5, which in turn is displaced by ~180o and comes close to the S5’ from the adjacent subunit (Fig. 4b, Movie 2). These changes are represented in the model structures where the backbones are shown as tubes for clarity (Fig. 4c, d, and superposed in Supplementary Fig. 6a). R4 and H5 change their orientations compared with xWT and are placed below the CTC (F0) along with R6. R1 and R2E remain above the CTC in the xE1R/R2E structure, while in xWT R4 is placed above the CTC to interact with E1 (Fig. 4c, Supplementary Fig. 6a). The R4 above the CTC in xWT shows that the channel is in the fully activated state, as in all the other KCNQ1 structures determined in detergent or nanodiscs.
a Cryo-EM maps for xWT (blue), xE1R/R2E (orange) and density-modified map (DM map) for xE1R/R2E (orange), clipped on the S4 residues, highlight the differences in the R4-H5 area. The DM map for the xE1R/R2E dataset was used to facilitate model building. b Sample maps for the S4 residues in xWT (blue), and xE1R/R2E (orange). Dashed lines show the approximate position of the Cα atoms of R4, H5 and R6 in the xWT structure. c, d S2, S4 and S5’ helices of xWT in blue (c) and xE1R/R2E in orange (d) highlighting the gating residues of the VSD. To assist with orientation, the tetrameric structures in the insert are colored gray, and the displayed helices in color. The adjacent S5 (S5’) is also illustrated in a darker hue. The CTC residues (xE160 (E2), xF157 (F0), and xD192 (D3)) are shown, as are the relative positions of the S4 residues xR218 (R1), xR221E (R2E), xQ224 (Q3), xR227 (R4), xH230 (H5), and xR233 (R6). Cα atoms are highlighted as yellow spheres, backbones are shown as tubes, and helices S1 and S3 were removed for clarity. D3 in the S3 is shown in sticks.
A further exploration of the residue interrelationships in the VSDs of our xWT, xE1R/R2E, and other recently published KCNQ1 structures may be found in the Supplementary text and Supplementary Figs. 6–8.
The role of R221 and E150 in the intermediate state
In the xWT structure, the side chains of xT134 and xV131 in the S1 helix interact with the main and side chain, respectively, of xY289’ in the filter helix from the adjacent subunit (FH’, Fig. 5a). The distance between the side chain of xT134 and the main chain of Y289’ increases from 3.5 Å (hydrogen bond distance, Cα-Cα distance: 6.5 Å) in xWT, to 11 Å (Cα- Cα distance: ~13 Å) in xE1R/R2E, setting the S1 away from the FH’ at a distance that is too large to allow any side chain interactions (Fig. 5b). The S1-FH’ contact in the xWT structure may be enabled by interactions between the xR221 in the S4 and residues in the S1. Analysis of potential hydrogen bond formation and polar interactions between the S1 and the S4 helices ( < 4 Å) reveals that in the xWT structure, xR2 could form a hydrogen bond with xS133 (2.9 Å), polar-polar interactions with the side chain of xS130 (2.9 Å), and Van der Waals interactions with xT134 (Fig. 5a). These interactions provide rigidity to the activated state of the VSD by allowing tighter packing and offsetting the charged and polar groups. With this approach of S1 to S4, the S2 of the xWT structure also adopts its curved conformation and S1 is brought closer to interact with the selectivity filter helix (FH’) through xT134 (Fig. 5a, Supplementary Fig. 7a). Substitution of the R2 with glutamic acid (R2E) in the xE1R/R2E structure arrests the channel in the intermediate state and prevents those interactions from occurring. Consequently, S1 remains distant from the S4 helix and the FH’ (Fig. 5b, Supplementary Fig. 7b). The positions within the main chains of xE1R and xR2E (Cα-Cα distance; ~11 Å) and residual density for their side chains in the upper part of the VSD can allow a salt bridge interaction between them, but the lower resolution of this region indicates that multiple conformations are likely, and therefore that this interaction is dynamic (see Methods, Movie 3, Fig. 5b).
a-b Top view representations of one VSD and its interactions with the filter helix from the adjacent subunit (FH’). In the inset, the tetrameric structures are shown to facilitate orientation. The FH’ interacting with the VSD is shown in a darker shade of blue for xWT (a), and orange for xE1R/R2E (b). Residues interacting with 221 and 150 are shown as gray and yellow sticks, respectively. Residues that may interact with R227 in the xE1R/R2E structure (b) are shown as orange sticks. The salt-bridge interaction in (a) between E150 and R227 is shown as a black line, polar interactions as blue dashed lines and the potential salt-bridge interaction in (b) as a black dashed line.
In the xWT structure E1 forms a salt bridge with R4 (R227, Fig. 5a) and the E1R substitution would likely lead to repulsive forces between the two positive charges. As a result, the R4 is placed in the CTC in the xE1R/R2E structure (Fig. 4d, Supplementary Fig. 6a), while S2 cannot approach the S4 and instead appears to tilt at xF0 to form a straight helix (Supplementary Figs. 5a, 6b). Our data show that the E1 and R2 charge reversal mutations limit the approach of the S1 and S2 towards the S4 and the pore filter helix, which prevents the formation of crucial interactions with residues in the selectivity filter helix that form normally in the activated state. This lack of interaction likely underlies the increased flexibility of the upper part of the VSD.
Molecular Dynamics Simulations validate the xE1R/R2E VSD conformation
To further assess the stability of our xE1R/R2E model we ran three independent 500 ns-long molecular dynamics (MD) simulations of the protein in a solvated membrane, monitoring both overall stability and more specifically the z position of side chains of the charged residues surrounding the CTC (Fig. 6a). By the end of the 500 ns, the backbone RMSD of the whole protein had plateaued at ~3.6 Å, with the RMSD of the VSDs remaining stable within 2-3 Å of their starting structure (Supplementary Fig. 9a). Residues surrounding the CTC maintained their conformations throughout the trajectories, with the z coordinates of each side chain fluctuating minimally (Fig. 6a, Supplementary Fig. 9c). Traces from the single replicates show occasional vertical flips of H5 and R4, such that the sidechain of H5 does briefly move above that of R4 in the z-plane, while still remaining out of and below the CTC (Supplementary Fig. 10a). The flipping was relatively infrequent and short lived across the four VSDs in the three separate simulations (~ 24% of total aggregated time), and still consistent with our placement of R4 and H5 sidechains in the model, which supports the validity of their conformations.
a Time evolution of the z-coordinates of selected VSD side chains in xE1R/R2E, with F0 (F157) set as reference. In all plots, dark lines represent averages over all four VSDs from three simulation replicates, while shaded areas indicate the standard deviation. b Same for xWT-R. c Time evolution of the R150-E221 salt bridge (red) and the R150-R227 (black) distance in xE1R/R2E. d Time evolution of the E150-R221 salt bridge (red) and the E150-R227 (black) distance in xWT-R. e, left Superposition of S1 and S2 in xWT (blue) and in xE1R/R2E (orange, last frame of one replicate). e, right Time evolution of distances between xE1R/R2E T145Cα and V131Cα and their respective positions in xWT; the dashed lines represent distances between residues from the cryo-EM structures. f, left Superposition of S1 and S2 in xWT (blue) and in xWT-R (green, last frame of one replicate). f, right Time evolution of distances between xWT-R T145Cα and V131Cα and their respective positions in xWT; the dashed lines represent distances between residues from the cryo-EM structures. g Superposition of xE1R/R2E (orange), xWT-R (green) and xWT structures (blue). The inset shows a single VSD (VSD1) from each of the three structures seen from above and the side from the final frame of the simulations for xE1R/R2E and xWT-R.
We also tested the stability of the xE1R/R2E conformation when the mutations in the VSD were reversed back to the xWT sequence in the pdb file, xR221 and xE150 (referred to as xWT-R, for Wild-Type-Reversion). While this construct showed slightly more pronounced backbone fluctuations during the simulations, the VSD RMSDs remained stable within 2-3 Å (Supplementary Fig. 9b). The residues surrounding the CTC were once again stable above (R1, R2, Q3) and below (R4, H5, R6) the CTC (Fig. 6b, Supplementary Fig. 9d), with occasional flips of H5 and R4 (Supplementary Fig. 10b). The salt bridge between xR221E and xE150R was also quite stable throughout the simulations of xE1R/R2E (Fig. 6c, red trace) and more so when the charges were reversed back in xWT-R, with the two residues approaching more closely and the distance fluctuating less overall (Fig. 6d, red trace). xE150R (E1R) and xR227 (R4) did not flip to approach one another throughout the simulations (Fig. 6c, black trace) even when the charges were reversed in xWT-R (Fig. 6d, black trace), and there were no substantial shifts in the center of mass positions of the S4 helices (Supplementary Fig. 11a, b). Moreover, no major changes were detected along the pore axis in any of the three simulations for each construct (Supplementary Fig. 11c).
To gauge the flexibility of the upper S1-S2 helices in E1R/R2E and xWT-R, the distance between the Cα atoms of xT145 and xV131 and their reference positions in xWT were measured over time (Fig. 6e, f). Interestingly, reversing the charge in xWT-R allowed for greater flexibility and sometimes further straightening of the S2 helix than was observed with the E1R/R2E structure. This is highlighted in the superpositions of xWT (in blue), xE1R/R2E (in orange) and xWT-R (in green) (Fig. 6g–inset). The upper S2 helix in xWT shows the greatest curvature, while xE1R/R2E occupies an intermediate position and the xWT-R S2 helix has completely straightened by the last frame of the simulation in this VSD and moved further away from the FH’. Flexibility for the upper S1-S2 was also indicated by our 3D variability analysis, but the conformations were strikingly different than the xWT (Supplementary text, Supplementary Fig. 8). The MD simulations support the stability of our xE1R/R2E model and suggest that the outer S1-S2 helices could undergo an even greater degree of motion in moving from resting or intermediate states to the fully activated states in the WT channel, further supporting the physiological relevance of our model.
PIP2 opens the xE1R/R2E gate
Electrophysiology experiments have shown that hE1R/R2E and xE1R/R2E are constitutively open channels in the absence of KCNE129 (Fig. 1c), but in our xE1R/R2E structure the gate is closed (Fig. 3c). This suggests that the xE1R/R2E channel is still a ligand-regulated channel and that the binding of PIP2 is mandatory for its opening. High-resolution structures of hKCNQ1-PIP2 have only been obtained in the presence of KCNE314 or ML27737, where PIP2 binding triggers a rearrangement of the S6-loop-HA segment into a continuous helix. This is followed by a translocation of helix HB and a rotation of CaM by ~180o, which abolishes its interactions with the VSD and leads to dilation of the pore14,37.
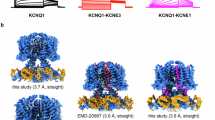
To determine the open structure of xE1R/R2E, we added 1 mM PIP2 (08:0 PI(4,5)P2, Avanti Research) to the purified sample 20 min prior to cryo-plunging. Data analysis (Supplementary Fig. 12) revealed two low-resolution xKCNQ1 classes (~ 8 Å) which both adopt a wider gate configuration compared to E1R/R2E without PIP2 (Fig. 7). Structural heterogeneity leading to low resolution structures is consistent with previous attempts to capture KCNQ1 alone in the open state using PIP214,37 but the clear densities of the PD allowed us to fit the hKCNQ1-KCNE3-PIP2 open structure (PDB-ID: 6V01)14 into the Class 1 map to show an open channel gate (Fig. 7a). The distance between the Cα atoms of two hL353 residues is 19.3 Å (Fig. 7a, purple dashed line) in comparison to the xE1R/R2E (xL343) where it is 11.7 Å (Fig. 3c)14. Notably, the S6 helices are straightened and CaM is detached from the S2-S3 linker conformation, as expected from an open gate structure.
Analysis of a dataset collected in the presence of 1 mM PIP2 shows two low resolution xE1R/R2E classes (~ 8 Å). The inset panels are bottom views of the lower part of the S6 (res h349-359 and x339-349), which moves away from the pore in the PIP2-bound structure14. a The hKCNQ1-KCNE3-PIP2 open channel structure (PDB-ID: 6V0114, KCNE3 was removed for clarity), colored purple, fits in the Class 1 map. CaM is detached from the VSD and the distance between the Cα atoms of hL353 in S6 helix is 19.3 Å (purple dashed line in the insert). b The xE1R/R2E structure, colored orange, fits in map 2 with CaM remaining attached to the VSD, but with the lower part of the S6 suggesting a wider gate conformation. Building of the S6 in the class 2 map, colored in black, shows that the distance between the Cα atoms of xL343 increases from 11.7 Å (orange dashed line in the insert) to 15.7 Å (black line in the insert).
The Class 2 map shows that CaM remains attached to the loop connecting helices S2 and S3 in the VSD and helices HA and HB preserve their apo state conformation (Fig. 7b, orange) as has been previously reported37. Docking of the apo xE1R/R2E structure into the Class 2 map shows how the lower part of S6 in the model (Fig. 7b, orange) would need to splay wider to fit in the density for this class. Building of the S6 helix in the class 2 map (Fig. 7b, black), suggests that the map can likely fit an almost open-gate S6 with the distance between the Cα atoms of two xL343 resides being 15.7 Å (Fig. 7b, black line) in comparison to the xE1R/R2E and the open hKCNQ1-KCNE3-PIP2 (PDB-ID: 6V01)14 structures, where the distance is 11.7 Å (Fig. 7b, orange dashed line) and 19.3 Å (Fig. 7a, purple dashed line), respectively. The results demonstrate that xE1R/R2E can adopt an open conformation similar to the one described for WT hKCNQ1.
Fluorescence emission during VSD activation support an initial large and secondary small VSD displacement
Our structural data (Figs. 2–4) place the S4 helix of xE1R/R2E in a similar position within the membrane to the fully activated xWT structure. Dynamic measurements of VSD movement in KCNQ1 using voltage clamp fluorometry of Alexa Fluor 488-labeled residues at the top of the S429,39 would therefore lead us to expect a large initial component of fluorescence (denoted F1) as the S4 moves outward and channels transition from resting to intermediate states, and then a much smaller or absent second fluorescence component (F2) when channels transition to the fully activated state at more positive voltages, as the top of the S1 and S2 domains move closer toward the pore filter helix. Fluorescence records from Alexa-488-labeled G219C-hKCNQ1 obtained at voltages between -180 to +80 mV were better fit with a bi-exponential than a mono-exponential activation time course at voltages positive to -40 mV (Fig. 8a), and the fluorescence-voltage (F-V) relations from individual oocytes were best fit assuming a double Boltzmann distribution. This allowed separation of the F-V from each oocyte into its F1 and F2 components (Fig. 8b, c). As expected from a channel that conducts in the intermediate state39, the F1 component of hKCNQ1 fluorescence tracks closely with the G-V (mean F1 V1/2 = -52 mV; G-V V1/2 = -45 mV) and is slightly steeper. It comprises ~80% of the total fluorescence report, much the same as in KCNE1 + hKCNQ1 (Fig. 8c, e). In KCNE1 + hKCNQ1 though, the F1-V relationship is displaced to more negative voltages and does not overlay the G-V V1/2 (Fig. 8d), as KCNE1 is thought to largely prevent conduction in the intermediate state29. The F2 components of fluorescence from hKCNQ1 and KCNE1 + hKCNQ1 are much smaller, around 20% of the total, and displaced to positive voltages (mean hKCNQ1 F2 V1/2 = 11 mV; KCNE1 + hKCNQ1 V1/2 = 3 mV, Figs. 8c, 8e). These findings are entirely consistent with the structural studies described above and suggest that the F1 components of fluorescence reflect the displacement of the S4 outward across the electric field, while the much smaller F2 component reflects the reorientation of the S1 and S2 towards the filter helix and the top of S4 itself (Figs. 3a, 5, 8f).
a Fluorescence from Alexa-Fluor 488-labeled hKCNQ1 (C214A/G219C/C331A) during a series of 2 s voltage steps, from -180 to +80 mV, following a 4 s -140 mV pre-pulse. Traces from a representative experiment fit by a biexponential (left, green) or a monoexponential equation (middle, purple). A comparison between both fits at +80 mV is shown in the right panel. The normalized fluorescence at the end of the 2 s pulse was fit to a double Boltzmann equation in independent experiments (n = 6) to obtain the F1 (mean V1/2 = -51.9 mV, k = 15.5 mV) and F2 (mean V1/2 = 11.4 mV, k = 16.8 mV) components of fluorescence. b Overlay of the overall fluorescence-voltage relation (F-V) and its F1 and F2 fluorescence components, together with the conductance-voltage (G-V) curve (Boltzmann fit: V1/2 = -44.6 mV, k = 13.7 mV, n = 4). c Relative amplitudes of the F1 (red) and F2 (blue) components (n = 6). d, e Analysis of F-V relationships of KCNE1 + hKCNQ1 from previously reported experiments88 (n = 24) individually fit with a double Boltzmann equation (F1: mean V1/2 = -96.9 mV, k = 26.2 mV; F2: mean V1/2 = 3.0 mV, k = 18.6 mV). d Overlay of the fluorescence components with the G-V (Boltzmann fit: V1/2 = 4.0 mV, k = 16.6 mV, n = 16). e Relative amplitudes of the F1 (red) and F2 (blue) components. f Updated cartoon representations of the intermediate and activated states based on the high-resolution xWT and xE1R/R2E states reported in this work. Helix S3 has been removed for clarity. Arrow shows movement of S1 and S2 towards the filter helix in the activated state. All error bars in Figure represent mean ± SEM.
Intermediate state regulation by the KCNE1 β-subunit
It is difficult to separate overlapping currents through intermediate and fully activated open states in WT hKCNQ1 channels and appreciate their separate contributions to the overall current waveform (Fig. 1c). In the Y46A-KCNE1 mutant, however, intermediate and activated state conductances are separated. A fast-activating current component is apparent at voltages up to +40 mV and a slower activating current is seen positive to +60 mV (Fig. 9a), which gives two clear phases to the G-V relationship (Fig. 9b, red circles). The tail currents also show different morphologies after pulses to lower and higher voltages (Fig. 9a, right), which suggests occupancy of different open states during the prior depolarizations.
a Whole cell currents during 4 s pulses from -90 mV to between +120 and −80 mV, and 2 s tail pulse to −40 mV for Y46A-KCNE1-hKCNQ1. Red traces denote current during pulse to +60 mV and subsequent tail at -40 mV. b G-V relations for Y46A-KCNE1-hKCNQ1 (red; n = 8), KCNE1-hKCNQ1 (black; n = 17), and hKCNQ1 (blue; n = 12) constructs. For all data presented in this figure, hKCNQ1 is GFP linked and all other constructs are co-expressed with GFP c Eight representative single channel sweeps for Y46A-KCNE1-hKCNQ1 at +60 mV, 4 s pulses. d All-points histograms from Y46A-KCNE1-hKCNQ1 traces showing major peaks at 0.046 and 0.104 pA. e All-points histograms from 2:4 stoichiometry KCNE1:hKCNQ1 complexes showing major peaks at 0.040 and 0.122 pA. Fits were made in Clampfit 10.5 (Molecular Devices) and plotted in Graphpad Prism 10.4. f Rb+/K+ permeability ratio from relative tail current amplitudes at −60 mV for indicated constructs plotted on a log base 2 y-scale (n = 5-8). Crosshatch bar indicates Y46A-KCNE1-hKCNQ1 pulsed to 0 mV. g ML277 (1 μM) increase in tail current at -40 mV plotted on a log 2 y-scale (n = 3–12), for hKCNQ1, KCNE1-hKCNQ1, hE1R/R2E and hE1R/R4E (data from Eldstrom et al., 202142). All error bars in Figure represent mean ± SEM. Cells were pulsed to +60 mV for 4 s followed by a 0.9 s repolarization to -40 mV. Statistical comparisons for bar graph data in (f) and (g) are shown in Supplementary Fig. 13.
The single channel records at +60 mV from Y46A-KCNE1-hKCNQ1 reveal the kinetic basis for these two types of behavior. In traces 1, 3, 5, 6 (Fig. 9c), the channel shows opening bursts after a long first latency, as we have previously described for WT KCNE1-KCNQ1 currents40,41. Traces 2, 4, 7, and 8 show much shorter latencies and the openings remain very small, similar to what we have observed with hE1R/R2E complexes28,42. The Gaussian fits of the all-points histogram for the records shown detect two separate peaks at 0.046 pA and 0.104 pA (Fig. 9d), with means of 0.032 ± 0.004 pA and 0.081 ± 0.013 pA for 3 different patches. Very similar values for the two conductance levels are seen in unsaturated 2:4 KCNE1:hKCNQ1 complexes where the peaks of the Gaussian distribution are seen at 0.04 and 0.12 pA (Fig. 9e).
The presence of two distinct conducting states in Y46A-KCNE1 mutants is supported by measurements of the Rb+/K+ permeability ratio and response to ML277. hKCNQ1 alone has a high Rb+ permeability ratio to K+ (2.5x, Fig. 9f) and high sensitivity to the activator ML277 (~8x, Fig. 9g, Supplementary Fig. 13), as do all hE1R/R2E (locked in the intermediate state) and Y46A-KCNE1 constructs. Linkage of KCNE1 to KCNQ1 or the hE1R/R4E (E160R/R237E) mutant, which locks the VSD in the fully activated state, reduces the Rb+/K+ permeability ratio to 1.0 or less, and abolishes the response to ML277.
KCNE3-hKCNQ1 channels occupy intermediate open states comparable with that of E1R/R2E co-expressed with KCNE3
Physiologically, the KCNE3-KCNQ1 complex may preferentially occupy the intermediate state and conduct current, based on the arrangement of the arginines in the CTC in the cryo-EM structure of hKCNQ1-CaM-KCNE314, an NMR structure of the VSD alone, and from the overlap of the F1 component of fluorescence with the GV curve43. Superposition of the hKCNQ1-CaM-KCNE3 complex (Fig. 10a), which is the only available closed structure of KCNQ1 with an accessory regulatory subunit (PDB-ID: 6V0014), onto the xWT and xE1R/R2E structures shows that the interactions between the upper S1 in the xE1R/R2E structure and KCNE3 would be lost in the xE1R/R2E-KCNE3 model because of the tilt of the S1-S2 helices (Fig. 10b–d).
a Cryo-EM structure of the hKCNQ1-CaM-KCNE3 complex (PDB-ID: 6V0014). hKCNQ1 is in shades of gray, and KCNE3 in yellow. b Side (above) and top (below) views of the binding cleft of KCNE3 in the KCNE3-hKCNQ1 complex structure. KCNE3 interacts with residues in the VSD, the S5’ and FH’ from the adjacent subunit and the S6” from the diagonal subunit. c-d xWT (blue) and xE1R/R2E (orange), were superposed onto 6V00 with KCNE3 shown in yellow. e hKCNQ1 residues that interact with KCNE3-Y60 in the KCNE3-hKCNQ1 complex structure preserve their conformation in xWT and xE1R/R2E. f-g For all data presented in this figure, hKCNQ1 and co-expressed KCNE3 are GFP-linked; all other constructs are co-expressed with GFP. Whole cell and single channel currents from KCNE3-hKCNQ1 and KCNE3 + hE1R/R2E constructs during 4 s pulses from -90 mV to between +80 and −80 mV, 0.8 s tail pulse to −40 mV. The protocol for single channel sweeps is shown above data in (g). h All-points histograms with Gaussian fits for data in (g). i Tail current density from tsA201 cells expressing hKCNQ1 (n = 8-9), KCNE3-hKCNQ1 (n = 3) and KCNE3 + hE1R/R2E (n = 4-6) constructs. Protocol as in (f). j Tail current response to 1 µM ML277 (n = 3-12) plotted on a log 2 y-scale. k Rb+/K+ permeability ratio from relative tail current amplitudes at −60 mV (n = 3-8) plotted on a log 2 y-scale. All error bars in the Figure represent mean ± SEM. Statistical comparisons for bar graph data in (j) and (k) are shown in Supplementary Fig. 13.
Whole cell currents, G-V relationships, and single channel openings (Fig. 10f–i) from WT KCNE3-hKCNQ1 and KCNE3 + hE1R/R2E are very similar and support the idea that KCNE3-KCNQ1 complexes conduct significantly in the intermediate state. Expression of KCNE3 linked to hKCNQ1 or KCNE3-GFP co-expressed with hE1R/R2E in tsA201 cells both resulted in small constitutive currents with a slightly decaying time dependence and overlaying G-V relationships (Fig. 10f, i). The single channel openings show little latency, appearing immediately on depolarization and sometimes again later in the traces, with some long closings during depolarization (Fig. 10g). A fit of the all-points histograms (Fig. 10h) for the records shown indicates a non-zero peak in WT KCNE3-hKCNQ1 of 0.08 pA, with a mean of 0.092 ± 0.01 pA for three separate cells. The fit to the all-points histogram for the records of KCNE3-GFP+hE1R/R2E identifies a non-zero peak of 0.075 pA, with a mean of 0.085 ± 0.005 pA for three cells.
KCNE3-containing complexes, like hKCNQ1 alone, are very sensitive to 1 μM ML277 whether paired with WT hKCNQ1 or hE1R/R2E (Fig. 10j, Supplementary Fig. 13). This is further evidence that KCNE3-hKCNQ1 shows a dominant intermediate state residence and that the published structure14 may represent an activated state where F68 in KCNE3 clashes with any movement of F279 in KCNQ1 to prevent accommodation of the drug33. KCNE3 has previously been shown to reduce Rb+ permeability in the WT hKCNQ1 channel44, but unlike KCNE1, KCNE3 maintains an ability to suppress the high Rb+ permeability of even the hE1R/R2E intermediate state (Fig. 10k, Supplementary Fig. 13).
Discussion
In this work we characterized an intermediate gating structure of the potassium channel KCNQ1 (Kv7.1). xKCNQ1 E150R/R221E (xE1R/R2E) stabilizes the channel in a conformation that experimentally reproduces intermediate voltage sensor activation in hKCNQ1 and its electrophysiological properties, including its Rb+/K+ tail current ratio and its XE991 and ML277 sensitivities (Fig. 1 and Supplementary Table 1). The use of the xE1R/R2E mutations allowed us to capture this intermediate state with improved resolution through reduction of sample variability (Fig. 2). By comparing it with the fully activated WT xKCNQ1 structure, we obtain insight into the structural differences between these states. Conservatively, in our xE1R/R2E structural model, the extracellular halves of S1 and S2 show a displacement of 6 Å compared to the WT activated structure (Fig. 3). Consequently, the number of intra-subunit S1/S2-S4 and inter-subunit S1-FH’ interactions is reduced in the mutant, which leads to increased flexibility in the upper half of the VSD (Supplementary Fig. 7). Replicated MD simulations indicate that the xE1R/R2E structure is stable over the sub-μs timescale, and more specifically the z positions of side chains of the charged residues surrounding the CTC do not change significantly relative to the CTC itself (Fig. 6). Reversion of the xE1R/R2E mutations back to WT results in slightly increased atomic fluctuations in the simulations, but the VSD RMSD remains stable. Although the ion conduction pathway itself is relatively unchanged between the WT activated and the xE1R/R2E intermediate-closed structures, changes in dynamics, affected by the reduced interactions between the VSD and the PD, could underlie the differences in Rb+/K+ apparent permeability and drug sensitivity of the intermediate state once the channel opens. By defining the structure of the intermediate activated state of KCNQ1, this work provides insights into KCNQ1 gating and modulation.
A recent study of hKCNQ1 in electrically polarized lipid vesicles revealed a high resolution activated state similar to previously described KCNQ1 structures and to the xWT structure described here18. In addition, they determined a low resolution intermediate state (PDB-ID: 8SIM) which was in the closed state due to the lack of PIP2, and in which the VSD helices occupy the same 3D space as in our xWT and previously determined structures (Supplementary Fig. 6)11,14,18,33. Comparison of the xE1R/R2E structure (Fig. 4) with their low-resolution (6.2 Å) intermediate structure (PDB-ID: 8SIM18) highlights the pronounced straightened conformation of S1 and S2 in our xE1R/R2E structure (Supplementary Fig. 6). Even though 8SIM is low resolution, comparison between xE1R/R2E and 8SIM shows somewhat similar conformations for R1, R2, Q3, and R4 relative to residues F0, D3 and E2 in the CTC (Supplementary Fig. 6). More specifically, in both structures the R4 occupies the CTC, but in 8SIM, H5 occupies the position in which we have placed the R6 side chain of xE1R/R2E, and the lower part of the S4 is further placed ~2.5 Å downwards.
Apart from the 8SIM structure, there is another WT cryo-EM structure of full-length hKCNQ1 determined by Ma et al.37 (PDB-ID: 7XNI) where the VSD adopts a conformation different from previous xWT structures, including the xWT KCNQ1 in our study (Supplementary Fig. 6). Even though the upper VSD in 7XNI was resolved at low resolution, it appears that the S1 and S2 domains also tilt37 towards the orientation we see in the xE1R/R2E structure, but then adopts a classical activated-state VSD configuration in the presence of ML277, similar to the one described here for xWT. Overall superposition of 7XNI onto xWT shows that the S1 has not approached the S4 and the FH’ in the 7XNI structure and that the S2 tilts at E1, whereas in the xE1R/R2E structure it tilts at the F0 residue (Supplementary Fig. 6a). In the xE1R/R2E structure this tilt can be attributed to a loss of interaction of xE221 with the S1 helix, repulsive forces between R4 and xR150, and pairing between R150 and E221 in the xE1R/R2E structure (Fig. 5). This shows that even though the side chain of E1 appears flexible in the 7XNI structure, it can likely still form a salt bridge with R4 and therefore the tilt of the S2 cannot happen below R4 (Supplementary Fig. 6). The authors attributed the conformation of the VSD to the different detergent used in their work, to the use of the full-length construct instead of a truncated one, or to the fact that at 0 mV some particles in the S4 may have been in the activated or the intermediate state37. This supports the idea that the xE1R/R2E charge reversal mutation is stabilized in an intermediate state that the full-length hKCNQ1 can also access, at least in detergent conditions, and it validates that the observed changes in the xE1R/R2E structure can occur in the absence of an introduced mutation, and therefore that our structure likely represents a physiological intermediate state of KCNQ1.
The xE1R/R2E intermediate state structure provides insight into the function of the KCNQ1 channel as it transitions from resting to activated states and how its interactions with the accessory subunits KCNE1 and KCNE3 modify its functional properties. The intermediate gating state has interesting kinetic, permeation, and pharmacological properties that set it apart from fully activated channel states (Fig. 1) and make it a specific target for therapies aimed at regulating the function of KCNQ1 channel complexes in a variety of different situations.
During activation, translation of the gating charges across the electric field is accomplished by the transition of the channel complex from resting to the intermediate state, and is tracked by gating current recordings34,35 and the overlaying large F1 component35 of Alexa-488 fluorescence (Fig. 8). We find no structural evidence, though, for a secondary external displacement of the S4 domain of the channel, as the position of the S4 in the xE1R/R2E structure is almost the same as in the WT structure reported in this paper (Figs. 3a, 4, Supplementary Fig. 6a) and previously published by ourselves and others14,33. There is a secondary F2 component of fluorescence from hKCNQ1 (Figs. 8b, 8c), and we propose that this minor component, which is also observed in KCNE1 + hKCNQ1 (Fig. 8e) and in KCNE3 + hKCNQ145, results from the reorientation of the top of S1 and S2 towards the pore as the channel moves from the intermediate state to the fully activated state (Fig. 8f). This results in a change of the overall quenching environment at the top of S4 where Alexa-488 is located. This conclusion is in agreement with the inability of some prior studies to detect a second component of gating charge attributable to the F2 component of fluorescence39. The F2 fluorescence component has a V1/2 close to +10 mV and provides a voltage range spanning the isoelectric line over which the secondary reorientation of the S1 and S2 domains towards the pore occurs. This explains how, in the absence of any applied electric field, the 7XNI structure37 might have captured a population of channels in transition from the intermediate state to the fully activated state.
The intermediate state is a functionally relevant state in hKCNQ1 channels and KCNE1 + hKCNQ1 channels in the lower half of the activation range of voltages between -40 and 0 mV, before the fully activated state is reached at more positive voltages. The Y46A-KCNE1-hKCNQ1 mutant, which hinders the ability of KCNE1 to fully integrate its amino terminus into the cleft between KCNQ1 subunits, illustrates this clearly (Fig. 9). The interaction of Y46 at the extracellular surface of KCNE1 with W323 in hKCNQ1 is critical to stabilize the outer interface of KCNE1 between two subunits of the complex (Y46 in KCNE1 is equivalent to Y60 in KCNE3, Fig. 10e), which allows KCNE1 to exert its suppression of the intermediate state and increase channel complex conductance in the activated state46. Mutations of Y46 to Cys increase the Rb+/K+47 and Cs+/K+ conductance ratios, particularly after labeling with MTSET48, and destabilize the extracellular binding pocket for activators49. As well, Ala and Leu mutants at Y46 shift activation of KCNE1 + hKCNQ1 complexes to more negative potentials and have faster activation kinetics50, as if they preferentially allow intermediate state conductance. By pushing the stabilization of the activated state to much more positive voltages in Y46A-KCNE1-hKCNQ1, the intermediate state conductance is observed at the whole cell and single channel level clearly separate from the activated state conductance in terms of its voltage-dependence, higher Rb+/K+ permeability, higher sensitivity to ML277, lower amplitude single channel openings, and shorter latencies to opening (Fig. 9). The Y46A-KCNE1-hKCNQ1 complex can conduct through the intermediate state at potentials around 0 mV and through both intermediate and fully activated states at more positive voltages. The same separation of the intermediate and activated state conductances can be observed in the single channel histogram of the 2:4 stoichiometry KCNE1:hKCNQ1 channel complex (Fig. 9e) and suggests that intermediate state conductance is a physiologically relevant contributor to the overall KCNE1 + KCNQ1 current waveform until the N-terminus of KCNE1 is tightly bound to hKCNQ1. The results emphasize the importance of tyrosine at position 46 in KCNE1 in stabilizing the hKCNQ1 pore and enhancing the fully activated open state conductance.
Our work also supports the recent suggestion that the active conducting form of KCNE3-KCNQ1 is the intermediate state43,45. The comparison of the electrophysiological and pharmacological properties of WT KCNE3-hKCNQ1 with KCNE3 + hE1R/R2E emphasizes their similarities with respect to expressed current density, current waveforms, voltage-dependence, ML277 sensitivity, and single channel currents (Fig. 10f–j). Interestingly, none of the KCNE3-hKCNQ1 constructs showed an increased Rb+/K+ permeability ratio (Fig. 10k). This difference between the effects of KCNE1 and KCNE3 on the Rb+/K+ permeability ratios of hKCNQ1 may be due to their structures. The KCNE1 TMH has a greater degree of curvature than KCNE351 and may make fewer contacts with the hKCNQ1 pore. The interactions that KCNE3 makes with hKCNQ1 allow a placement of the β-subunit that enables it to maintain its modulation of that pore property.
Since xE1R/R2E is a constitutively open channel, it begs the question whether this conformation of the VSD can illuminate mechanisms for loss-of-function or gain-of-function hKCNQ1 mutants. Many missense LQT1 mutations have been found in the VSD and the position of several of these residues is different between the xWT and xE1R/R2E structures (Supplementary Fig. 14). Interestingly, some of the residues that appear important for coupling of the S1 to the filter helix, which no longer interact in the xE1R/R2E structure, have been reported as pathological mutants (human E160K, R231C, S140G, T144A, V141M; Supplementary Fig. 14), with hS140G and hV141M being associated with atrial fibrillation. It has been proposed that both of these mutants have some intermediate state conductance in the presence of KCNE1, which is particularly evident during deactivation52. Further complexity is introduced by a 2:4 stoichiometry where the mutant hKCNQ1 subunit may or may not be associated with a β-subunit53, which allows for variable expression of the mutant phenotype.
Importantly, previous work has shown that substitution of the arginine alone at position h231 (R2) can lead to constitutively open channels with F46, M46, E54, A55, W56, Q54, and C57 (a gain-of-function mutant also due to supertrafficking to the plasma membrane46, Supplementary Fig. 14). In addition, substitution of arginine with the positively charged lysine makes the VSD sensitive to voltage changes54, highlighting the importance of a positive charge at position R2 for normal voltage dependence54. Although the structures may be different, it is likely that the substitution of the arginine with a non-positively charged residue leads to loss of interactions with residues in the S1, and as a result S1 and S2 sit apart from the S4 and the pore, as described in xE1R/R2E (Fig. 5b). In support of this observation, hR231C which was thought to lock the S4 in the activated state, displays a high Rb+/K+ permeability ratio in the absence of KCNE1 but a low ratio in the presence of KCNE158,59. However, when expressed with KCNE1, hR231C currents57 show constitutive and voltage-dependent components similar to Y46A-KCNE1-hKCNQ1, whereas hR231E currents are constitutive only54. Given the possibility for some clinically relevant mutations to enhance either the intermediate or activated states, our structural determinations are crucial tools to understand the pathologies of and improve therapeutic strategies for human KCNQ1-related disorders with state-specific channel modulators33,60.
Methods
All research involving biological hazards was conducted in accordance with the University of British Columbia Policies and Procedures, Biosafety Practices and Public Health Agency of Canada guidelines, under Biohazard approval certificate B21-0006, of the UBC Biosafety Committee.
Electrophysiology
tsA201 cells (Mycoplasma free, Catalog #96121229, Sigma-Aldrich, St Louis, MO) were cultured and plated as previously described25,27 and transfected with LipofectamineTM 2000 (Thermo Fisher Scientific, Waltham, MA, USA). For data in Fig. 1, 2 µg of either hE1R/R2E or hE1R/R4E, or 1 μg linked KCNE1-hWT were co-transfected with 500 ng GFP. For hWT (hKCNQ1), xWT and xE1R/R2E, 2 µg were transfected alone, as the constructs used are already GFP tagged. For data in Figs. 9 and 10, constructs that are an α-subunit linked to a β-subunit have a dash between subunits (eg. KCNE1-hE1R/R2E), while subunits that are co-expressed have a + sign between subunits (eg. KCNE3 + hE1R/R2E). Cells were transfected with 1.5 µg of the following constructs: KCNE1-hKCNQ1, KCNE3-hKCNQ1, hE1R/R2E, hE1R/R4E, KCNE1-hE1R/R2E, KCNE1-hE1R/R4E, Y46A-hKCNE1-hKCNQ1, and Y46A-hKCNE1-hE1R/R2E, along with 0.5 µg of GFP. hE1R/R2E (1 μg) was co-expressed with KCNE3-GFP (3 µg). For the hKCNQ1-GFP linked construct, 1.5 μg was used.
Whole-cell patch clamp experiments were performed 24 h after transfection at room temperature (22 °C). GFP fluorescence was used to identify successfully transfected cells. The pipette solution contained (in mM): 130 KCl, 5 EGTA, 1 MgCl2, 4 Na2-ATP, 0.1 GTP, 10 HEPES (pH 7.2 with KOH). The bath solution contained (in mM): 135 NaCl, 5 KCl, 1 MgCl2, 2.8 Na-Acetate, 10 HEPES, 1 mM CaCl2 (pH 7.4 with NaOH). For Rb+/K+ permeability experiments, the bath solutions used contained either high K+ (in mM: 140 KCl, 1 MgCl2, 2.8 Na-Acetate, 10 HEPES, 0.1 CaCl2, pH 7.4 with KOH) or high Rb+ (in mM: 140 RbCl, 1 MgCl2, 2.8 Na-Acetate, 10 HEPES, 0.1 CaCl2, pH 7.4 with RbOH). Data were acquired using an Axopatch 200B amplifier, Digidata 1440 A digitizer, and pCLAMP 10.3 software (Molecular Devices, LLC, San Jose, CA), sampled at 10 kHz and filtered at 5 kHz. Electrode pipettes for whole-cell recordings were pulled from thin-walled borosilicate glass (World Precision Instruments, Sarasota, FL) using a linear multistage electrode puller (Sutter Instrument, Novato, CA). Pipettes were fire-polished prior to use and had resistances ranging from 1-3 MΩ. Series resistances were <5 MΩ and compensation of ~75% was applied, leaving a calculated voltage error of ~1.25 mV/nA current.
Data analysis was performed using Prism 8 or Prism 10 for Mac (GraphPad Software, Inc., San Diego, CA). Conductance-voltage (G-V) curves were obtained from the peak tail current, normalized (G/Gmax), and plotted as a function of voltage. For hWT and xWT, the resulting relationships were fitted to a Boltzmann equation of the form:
G/Gmax = Gmin + (Gmax – Gmin)/(1 + exp((V1/2 – V)/k), where V1/2 represents the voltage of half-maximal activation and k is the slope value. For hE1R/R2E and xE1R/R2E, the G-V plots were fitted to a linear regression and divergence of the slope from 0 was tested with an F-test. Data are mean ± SEM of n = 5-7 independent cells per condition.
Rb+/K+ current ratios were measured at the negative peak of the tail current. In Fig. 1, differences were assessed using an ANOVA and post hoc Bonferroni tests for selected comparisons: hWT vs. xWT, hE1R/R2E vs. xE1R/R2E, hWT vs. hE1R/R2E, xWT vs. xE1R/R2E, and hWT vs. KCNE1-hWT. Only hWT vs. KCNE1-hWT and hE1R/R2E were statistically significant (p-value < 0.0001 and = 0.0001, respectively).
For the pharmacology experiments, XE991 dihydrochloride aliquots were dissolved in water and stored at -20 °C. One aliquot was used per experimental day to prevent impact from thaw-freeze cycles and perfused to the bath at concentrations from 30 nM to 30 µM. The current level after application of the drug was normalized to the control peak and results fitted using the Hill equation with slope value = 1. Normalized response (Y) = Ymin + (Ymax – Ymin)/(1 + [XE991]/IC50). No differences between the IC50s of WT and hE1R/R2E constructs were observed with ANOVA. hE1R/R4E was not considered in the statistical analysis due to being poorly estimated. ML277 aliquots were dissolved in DMSO according to the supplier’s instructions, stored at -20 °C and diluted into the bath to reach a final concentration of 1 µM. The tail current after application of the drug was normalized to the control current and results are expressed as fold current increase.
Single channel recordings
Recordings were made from mouse ltk- fibroblasts (ATCC Cat# CCL-1.3; Cedarlane Labs, Burlington, Ontario, Canada), plated on glass coverslips, and transfected 24 h later using Lipofectamine 2000 (ThermoFisher Scientific). Cells were transfected with 1.5 μg of channel DNA and 0.75 μg of GFP:pcDNA3 as a marker for transfected cells or 1.5 μg of channel DNA and 4.5 μg of KCNE3-GFP. Recordings were acquired as previously described42,61.
Voltage clamp fluorometry
Mature female Xenopus laevis frogs (Xenopus 1, Dexter, MI, USA) were anaesthetized in a solution containing: 2 g/L tricaine methanesulfonate, 2 g/L HEPES (pH 7.4 with NaOH). Under anesthesia, the animal was euthanized in accordance with the University of British Columbia animal care protocols. The ovarian lobes were extracted, divided into smaller sections and digested for 2-4 h in a solution containing: 3 mg/mL collagenase type 4 (Worthington Biochemical Corporation, Lakewood, NJ, USA), 82.5 mM NaCl, 2.5 mM KCl, 1 mM MgCl2, 5 mM HEPES (pH 7.6 with NaOH). The oocytes were washed and stored in a media containing: 500 ml Leibovitz’s L-15 medium (Thermo Fisher Scientific), 15 mM HEPES, 1 mM L-glutamine, 250 mg/L gentamycin (pH 7.6 with NaOH). Stage IV and V oocytes were selected and stored at 18 °C. Oocytes were injected with cRNA synthesized using the Ambion mMessage mMachine T7 transcription kit (Applied Biosystems, Foster City, California, USA). 10 ng of 214 A/G219C/C331A-hKCNQ1 pcDNA3.1+ (G219C-hKCNQ1; a gift from Dr. Jianmin Cui) cRNA was injected, alone or with 5 ng KCNE1 pBSTA. The pGEMHE vector was a gift from Dr. Yoshihiro Kubo. Experiments were performed 3-4 days post injection at room temperature.
For two electrode voltage clamp fluorometry (VCF) experiments, the bath solution contained (in mM): 96 NaCl, 3 KCl, 1 MgCl2, 2 CaCl2, 0.1 LaCl3, 5 HEPES (pH 7.4 with NaOH). The pipette solution contained 3 M KCl. Oocytes were labeled with 10 μM Alexa-Fluor 488 C5-maleimide in a depolarizing high potassium solution containing (in mM): 98 KCl, 1.8 CaCl2, 5 HEPES (pH 7.6 with KOH) for 30 min on ice. The oocytes were washed with the bath solution and left on ice prior to recording. Fluorescence and ionic recordings were obtained simultaneously as previously described62,63 with an Omega XF100-2 filter set (Omega Optical Inc, Brattleboro, VT, USA). Fluorescence recordings from the same oocyte were averaged to reduce signal noise errors. To correct for photobleaching, a fluorescence signal was recorded in the absence of a voltage step and was subtracted from the signal. The obtained traces were downsampled from 0.15 ms to 1 ms to facilitate curve fitting by averaging every 6-7 datapoints. This reduced noise while preserving the kinetics of interest, which were orders of magnitude slower than the filter cutoff.
Protein production
A construct encoding truncated Xenopus laevis KCNQ1 (NP_001116347.1, residues 67-610), suitable for cryo-EM studies11,33, as first described by Sun and MacKinnon11, was purchased from Genewiz (South Plainfield, NJ, USA). The C-terminus of the protein was followed by a preScission protease site and a green fluorescent protein (GFP)-6xHis tag to monitor gene expression levels and facilitate protein purification. Both CaM and xKCNQ1 constructs were cloned into the pcDNA3 expression vector and used for protein production in mammalian cells.
tsA201 adherent cells were cultured in 5-layer flasks in culture media [minimum essential media (MEM) supplemented with 10% (v/v) fetal bovine serum (FBS) and 1x antibiotic-antimycotic] at 37 °C, 5% CO2 until confluence. Cells were harvested with trypsin, which was deactivated using culture media and removed by centrifugation. Cells were resuspended in culture media, plated in 150 mm culture dishes (8.7 × 106 cells/plate) and incubated for 48 h at 37 °C, 5% CO2 in culture media supplemented with 5% (v/v) FBS. Before transfection, the culture media was replaced (5% FBS) and DNA samples were co-transfected into the tsA201 cells at a ratio of xKCNQ1:CaM 5:1 using polyethylenimine (1.5 μg PEI per 1 μg DNA). After 48 h, the transfected cells were washed with cold PBS, scrapped, collected with centrifugation (250 ×g) and stored at −80 °C.
Protein purification
Purification of xKCNQ1-CaM was performed using an anti-GFP nanobody-based affinity method. The 6×His-thioredoxin (Trx) nanobody fusion protein was produced in E.coli Rosetta gami 2 (DE3) cells as previously described33. Cells were suspended in lysis buffer (50 mM HEPES pH 7.4, 300 mM NaCl, 10% w/v sucrose, lysozyme, and DNase I), lysed using a Fisher Scientific Sonic Dismembrator Model 500 (1 s on/off, 50% amplitude, 2 min, on ice, twice) and the suspension was spun down using a JA25.5 fixed-angle rotor (Beckman Coulter) at 39,000 × g for 45 min. The supernatant was filtered through a 0.45 μm filter and loaded on a HisTrap FF column (Cytiva) using an AKTA purifier (Cytiva). The column was pre-equilibrated in buffer A (50 mM HEPES pH 7.4 and 300 mM NaCl) and after loading the sample it was washed with buffer A supplemented with 5% buffer B (50 mM HEPES pH 7.4, 300 mM NaCl, and 500 mM imidazole) and the bound proteins were eluted using 100% buffer B. Fractions containing 6×His-TRX-nanobody were pooled and dialyzed over-night at 4 °C in buffer A in presence of Tobacco Etch Virus protease (TEV; ~1 mg TEV per 20 mg total protein concentration). The sample was filtered through a 0.45 μm filter, loaded on a HisTrap FF column (Cytiva) and the flow-through containing the nanobody was collected and dialyzed overnight at 4 °C against buffer C (20 mM Tris-HCl pH 8.0, 20 mM NaCl). The sample was filtered through a 0.45 μm filter, loaded on an anion exchange column (HiTrapQ, Cytiva) and the flow-through containing pure nanobody was dialyzed against buffer D (50 mM NaPi buffer pH 7.5 and 300 mM NaCl). Nanobody at 1 mg/mL concentration was coupled with NHS-activated sepharose FF (Cytiva), as described before33. Briefly, the NHS-activated Sepharose FF (Cytiva) was washed with 1 mM ice-cold HCl and then equilibrated in buffer D. The nanobody was added at 1 mg/mL concentration at a ratio of 1 ml resin per 5 mg nanobody followed by over-night stirring at 4o C. The resin was washed with buffer D and then excessively with 0.1 M Tris-HCl pH 8.0 and stored in 0.1 M Tris-HCl pH 7.4.
For xKCNQ1-CaM WT and xE1R/R2E purifications, ~25 g cell pellets were suspended in 100 mL buffer E (40 mM HEPES pH 7.2, 150 mM KCl, 4 mM EGTA, 1× protease inhibitor cocktail set III (Millipore), 1 mM PMSF and benzonase) and homogenized using a Dounce homogenizer. Cells were lysed using a Fisher Scientific Sonic Dismembrator Model 500 (1 s on/off, 50% amplitude, 20 s) on ice and cell debris was removed with centrifugation (4,400×g, 20 min, 4 °C). Subsequently, membranes were collected with centrifugation (100,000×g, 60 min, 4 °C, Ti45 Beckman Coulter rotor), suspended in buffer E, homogenized using a Dounce homogenizer, and solubilized in 1.5% (w/v) GDN [ratio; (5 mL membrane suspension-GDN)/(g of membranes)] while stirring at 4 °C. After 2 h, insoluble material was removed with centrifugation (100,000 × g, 60 min, 4 °C, 70.1Ti Beckman Coulter rotor), the supernatant was filtered through a 0.45 μm membrane and incubated with 1.5 mL of the nanobody-resin under rotation at 4 °C for 4 h. The slurry was poured in a gravity column and washed with 5 CV buffer F (20 mM HEPES pH 7.2, 150 mM KCl, 4 mM EGTA, 2 mM DTT, 1× protease inhibitor cocktail set III(Millipore)) supplemented with 0.8% w/v GDN, 5CV buffer F supplemented with 0.08% w/v GDN and 10CV buffer F supplemented with 0.03% w/v GDN. Finally, PreScission protease was added, and the slurry was incubated overnight at 4 °C under gentle rotation. Cleaved xKCNQ1 was concentrated (Amicon Ultra-0.5 100 kDa MWCO) and applied on a Superose 6 Increase 10/300 GL (Cytiva) equilibrated in buffer F supplemented with 0.03% w/v GDN at 4 °C. Fractions containing xKCNQ1 were pooled, concentrated to ~3 mg/mL (Amicon Ultra-0.5 100 kDa MWCO) and plunged fresh for cryo-EM studies
Cryo-EM sample preparation, data collection, and analysis
For the PIP2 dataset, 1 mM PIP2 (08:0 PI(4,5)P2, AvantiResearch) was added in the sample 20 min prior to cryo-plunging. Quantifoil R2/2 copper grids with holey carbon (Electron Microscopy Sciences) were glow-discharged (PELCO easiGlow) and using a Vitrobot Mark IV (Thermofisher Scientific), 2.5 μL of the sample was applied to the grids at 4 °C and 100% humidity, blotted using a blot force of 4 for 2 s, and plunged in liquid ethane. Grids were screened using a 200 kV Glacios microscope (ThermoFisher Scientific) equipped with a Falcon III detector and data were collected on a 300 kV Titan Krios (ThermoFisher Scientific) equipped with a Falcon 4i Direct Electron Detector and Selectris energy filter at the High resolution Macromolecular Cryo-Electron Microscopy (HRMEM) facility at UBC. Data were collected with a pixel size of 0.96 Å, magnification of x130,000, total dose of 50 electrons per Å2 and a defocus range between -0.5–2 μm. Data processing was performed using Cryosparc (V4.6.0 for xE1R/R2E datasets and V4.2.1 for the xWT dataset, Supplementary Figs. 2, 4, 12). Gain-corrected images were patch motion corrected and the CTF estimation was done using the patch CTF estimation tool.
For the xWT dataset, 5500 movies were collected, and blob picker was used for initial particle picking and the xKCNQ1-CaM like 2D classes were used for template-based particle picking. Particles were extracted using a box size of 336 pixels and cleaned up using 2D classification and 3D classification. For 3D classification, xKCNQ1-CaM particles were used for ab-initio reconstruction and heterogeneous refinement. After 3 rounds of 3D classification, 47,918 particles were used for non-uniform refinement with C4 symmetry imposed to obtain a high-resolution map.
For xE1R/R2E, two datasets were collected, consisting of 32340 (Dataset #1) and 11044 (Dataset #2) movies. Both datasets were upsampled by x2 (pixel size 0.38 Å). Each dataset was processed separately, and after ab-initio and heterogeneous refinement, particles were combined and further cleaned up to obtain a high-resolution map. Images with CTF fit worse than 6 Å were omitted leading to 29620 and 10982 micrographs for datasets #1 and #2, respectively. Blob picker was used to generate classes for template-based picking and particles were extracted using a box size of 720 pixels which was Fourier-cropped to 120 pixels to minimize computational time. For dataset #1 and #2, 7495136 and 2600208 particles underwent 2 rounds of 2D classification culminating in 1550069 and 965336 particles, respectively. Highest quality 2D classes were used to generate ab-initio maps and subsequently, all particles were classified with heterogeneous refinement. xE1R/R2E-CaM particles from the 2 datasets were re-extracted to a box size of 720 pixels which was Fourier-cropped to 360 pixels (pixel size 0.96 Å) and combined. The 833,545 particles were subsequently classified using heterogeneous refinement and a 3.82 Å map was generated from 388,672 particles after non-uniform refinement with C4 symmetry imposed (Initial map, Supplementary Fig. 4b) which was used later for 3D variability analysis. Three more rounds of ab-initio and heterogeneous refinement followed by non-uniform refinement with C4 symmetry imposed led to 95597 particles and a 3.53 Å (Final map, Supplementary Fig. 4c). An additional classification step beyond this point was tested but not pursued, as it resulted in only KCNQ1 classes with decreased overall and VSD resolution. The final map was used for model building as explained below.
For the xE1R/R2E-PIP2 dataset, 9414 movies were collected and images with a CTF fit worse than 7 Å were omitted. 2,598,205 particles were extracted using a box size of 336 pixels and used for 2D classification. 475,867 xKCNQ1 particles were used for ab-initio and heterogeneous refinement (3D classification). After two rounds of 3D classification and NU-refinement with C4 symmetry imposed, 2 low resolution classes of xKCNQ1 were obtained showing a typical open configuration (Class 1, 33200 particles, 7.75 Å, Supplementary Fig. 12) and an open gate configuration with CaM attached to the VSD (Class 2, 34976 particles, 8.17 Å, Supplementary Fig. 12).
xE1R/R2E 3D variability analysis
To explore the heterogeneity in the xE1R/R2E dataset, we performed 3D Variability Analysis (3DVA) using CryoSPARC (v4.6.0). The initial map at 3.82 Å resolution (Supplementary Fig. 4b), along with 388672 particles, served as the starting point for the analysis. To further improve particle alignment, sequential local refinements were performed using masks encompassing the tetrameric xKCNQ1-CaM and the tetrameric transmembrane domain. Subsequently, C4 symmetry particle expansion (1554688 particles) and local refinement of the transmembrane region of a single xKCNQ1 protomer was conducted which yielded a map at 3.42 Å resolution and was used as input for 3DVA. During all alignment steps, the pose/gaussian prior was applied, with a standard deviation of 5° and 3° for the prior over rotations and 2.5 Å and 1.5 Å for the prior over shifts, for the tetrameric xKCNQ1-CaM, the tetrameric xKCNQ1 TM region, and the TM region of a single xKCNQ1 protomer, respectively. Additionally, rotations and shifts were re-centered at each iteration. The 3DVA was run with three modes of variability. Clustering of the particles followed by local refinement showed potential small movements of the upper part of the S1 helix. Combinations of particles with similar S1 conformations did not improve the resolution to build structural models but further highlighted the upper S1 movement.
Model building and refinement
For the xWT structure determined in GDN, our previous published xKCNQ1 WT structure determined in DDM:CHS (PDB-ID: 7TCP)33 was used as a reference for model building. For the xE1R/R2E structure, the xWT in GDN was used as a starting model. The reference structures were docked in the cryo-EM maps and built manually using Coot64. For xE1R/R2E, model-angelo65 was used to produce an unbiased model which placed the R4 in the CTC, as we expected, the R6 occupying its unambiguous density and H5 clashing on R6. Since there is no density for the side chain of H5, this suggests that it is pushed out from the CTC. Model building was facilitated by a density-modified map generated from the final xE1R/R2E cryo-EM map using EMReady38 (DM map, Fig. 4, Supplementary Fig. 5), and the model was further optimized with ISOLDE66. ISOLDE was run at 300 K and helped position the side chains by performing interactive molecular dynamics simulation, optimizing conformations based on the local environment, reducing clashes and improving overall stability and fit in the cryo-EM map. The structures were built from residues 95-555, with disorder regions 67-94, 206-215, and 385-496. CaM was built from residues 10-146. Models went through rounds of manual building using Coot and real-space refinement with secondary restraints using PHENIX67. Model quality was assessed using Molprobity68 and figures and morph were prepared using ChimeraX69 and Chimera70.
Molecular dynamics simulations
The cryo-EM structure of the xE1R/R2E mutant, with closed pore and VSDs in the intermediate conformation was investigated using MD simulations. The segment of each subunit defining the TM pore and VSD domains (residues N95 to Q349) were retained for MD simulations, while the calmodulin binding domains were removed. The xWT-R structure was obtained from xE1R/R2E by reversing the mutations using Chimera70. SWISS-MODEL71 was used to model missing loops. The CHARMM-GUI Membrane Builder server72 was employed to prepare all necessary files for the simulations. Each of the protein structures was inserted in a 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine (POPC) membrane and solvated with explicit water molecules. The total system charge was neutralized with a 150 mM KCl solution, obtaining around 192,000 atoms in total. Simulations were performed with the NAMD software73 and the CHARMM36m74 force field for proteins and lipids, with a modified TIP3P75 water model and NBFIX parameters76,77. Periodic boundary conditions (PBCs) were applied using a tetragonal unit cell, and long-range electrostatic interactions were calculated via the Particle Mesh Ewald (PME) algorithm78. A 12 Å cutoff was used for short-range electrostatic and van der Waals interactions by applying a force-based switching function starting to take effect at 10 Å79. An integration time step of 2 fs was employed, with the SHAKE algorithm80 to constrain covalent bonds involving hydrogen atoms, except in water molecules, where SETTLE was used81. Before production, all systems were relaxed with a ∼30 ns-long equilibration beyond the CHARMM-GUI equilibration protocol, allowing proper hydration of solvent-exposed regions of the pore cavity82. Then, three independent replicates of each system were simulated for 500 ns at constant pressure (1 atm) and temperature (310 K), NPT ensemble, using the Nosé-Hoover Langevin piston method (oscillation period 50 fs and damping time scale 25 fs)83 and a Langevin thermostat (damping coefficient 1 ps−1). The simulated systems’ setups are summarized in Supplementary Table 3. The Molecular Dynamics Simulations checklist is provided as Supplementary Table 4.
Calculations of the channel pore radius
The pore radius of each structure was calculated using the HOLE84,85 program. For replicated trajectories, average and standard deviation values were obtained from aggregating the three 500 ns-long runs and taking a snapshot every 50 ns. The structures were aligned to the starting conformation using the backbone atoms as a reference. For all HOLE calculations, AMBER van der Waals radii were adopted with a cutoff of 6 Å.
Molecular graphics and analysis
Molecular graphics and analysis were performed with UCSF ChimeraX86 and Visual Molecular Dynamics (VMD)87 and custom Tcl scripts.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The cryo-EM maps have been deposited in the Electron Microscopy Data Bank (EMDB) under accession codes EMD-48725 (xWT-CaM), and EMD-48726 (xE1R/R2R-CaM). The atomic coordinates have been deposited in the Protein Data Bank (PDB) under accession codes 9MY3 (xWT-CaM) and 9MY4 (xE1R/R2R-CaM). Previously published PDB files mentioned in this manuscript PDB-ID: 7TCP (xWT-CaM in DDM:CHS); PDB-ID: 6V01 (KCNQ1-KCNE3-CaM complex with PIP2); PDB-ID: 6V00 (KCNQ1-KCNE3-CaM); PDB-ID: 7XNI (Full length human KCNQ1-CaM); PDB-ID: 8SIM (human KCNQ1-CaM in the intermediate state in electrically polarized vesicles). All MD details can be found in the MD checklist, included as Supplementary Table 4. Initial and final MD conformations and all simulation input files may be found in a data repository (https://doi.org/10.5281/zenodo.16615177), referenced in the MD checklist. A README file describes the repository contents. All other data supporting the findings of this work are available within the paper, or provided in the Supplementary information files and the Source data file for Figs. 1, 8, 9 and 10. Source data are provided with this paper.
References
Shieh, C. C., Coghlan, M., Sullivan, J. P. & Gopalakrishnan, M. Potassium channels: molecular defects, diseases, and therapeutic opportunities. Pharm. Rev. 52, 557–594 (2000).
Sanguinetti, M. C. et al. Coassembly of K v LQT1 and minK (IsK) proteins to form cardiac I Ks potassium channel. Nature 384, 80–83 (1996).
Barhanin, J. et al. K v LQT1 and IsK (minK) proteins associate to form the I Ks cardiac potassium current. Nature 384, 78–80 (1996).
Hibino, H., Nin, F., Tsuzuki, C. & Kurachi, Y. How is the highly positive endocochlear potential formed? The specific architecture of the stria vascularis and the roles of the ion-transport apparatus. Pflug. Arch. 459, 521–533 (2010).
Liin, S. I., Barro-Soria, R. & Larsson, H. P. The KCNQ1 channel - remarkable flexibility in gating allows for functional versatility. J. Physiol. 593, 2605–2615 (2015).
Schroeder, B. C. et al. A constitutively open potassium channel formed by KCNQ1 and KCNE3. Nature 403, 196–199 (2000).
Jervell, A. & Lange-Nielsen, F. Congenital deaf-mutism, functional heart disease with prolongation of the Q-T interval and sudden death. Am. Heart J. 54, 59–68 (1957).
Kapplinger, J. D. et al. Spectrum and prevalence of mutations from the first 2,500 consecutive unrelated patients referred for the FAMILION long QT syndrome genetic test. Heart Rhythm 6, 1297–1303 (2009).
Bellocq, C. et al. Mutation in the KCNQ1 gene leading to the short QT-interval syndrome. Circulation 109, 2394–2397 (2004).
Ohshige, T. et al. A single nucleotide polymorphism in KCNQ1 is associated with susceptibility to diabetic nephropathy in japanese subjects with type 2 diabetes. Diabetes Care 33, 842–846 (2010).
Sun, J. & MacKinnon, R. Cryo-EM Structure of a KCNQ1/CaM Complex Reveals Insights into Congenital Long QT Syndrome. Cell 169, 1042–1050 e1049 (2017).
Tristani-Firouzi, M. & Sanguinetti, M. C. Structural determinants and biophysical properties of HERG and KCNQ1 channel gating. J. Mol. Cell Cardiol. 35, 27–35 (2003).
Shamgar, L. et al. Calmodulin is essential for cardiac I-KS channel gating and assembly - Impaired function in long-QT mutations. Circulation Res. 98, 1055–1063 (2006).
Sun, J. & MacKinnon, R. Structural Basis of Human KCNQ1 Modulation and Gating. Cell 180, 340–347 e349 (2020).
Zhang, H. et al. PIP(2) activates KCNQ channels, and its hydrolysis underlies receptor-mediated inhibition of M currents. Neuron 37, 963–975 (2003).
Zaydman, M. A. et al. Kv7.1 ion channels require a lipid to couple voltage sensing to pore opening. Proc. Natl Acad. Sci. U. S. A 110, 13180–13185 (2013).
Hou, P. et al. Two-stage electro-mechanical coupling of a KV channel in voltage-dependent activation. Nat. Commun. 11, 676 (2020).
Mandala, V. S. & MacKinnon, R. The membrane electric field regulates the PIP(2)-binding site to gate the KCNQ1 channel. Proc. Natl Acad. Sci. USA 120, e2301985120 (2023).
Tinel, N., Diochot, S., Borsotto, M., Lazdunski, M. & Barhanin, J. KCNE2 confers background current characteristics to the cardiac KCNQ1 potassium channel. EMBO J. 19, 6326–6330 (2000).
Angelo, K. et al. KCNE5 induces time- and voltage-dependent modulation of the KCNQ1 current. Biophysical J. 83, 1997–2006 (2002).
Grunnet, M. et al. KCNE4 is an inhibitory subunit to the KCNQ1 channel. J. Physiol. 542, 119–130 (2002).
Hodgkin, A. L. & Huxley, A. F. A quantitative description of membrane current and its application to conduction and excitation in nerve. J. Physiol. 117, 500–544 (1952).
Bezanilla, F. The voltage sensor in voltage-dependent ion channels. Physiological Rev. 80, 555–592 (2000).
Armstrong, C. M. Voltage-gated K channels. Sci.STKE. 2003, re10 (2003).
Armstrong, C. M. & Hollingworth, S. Na(+) and K(+) channels: history and structure. Biophys. J. 120, 756–763 (2021).
Kurata, H. T. & Fedida, D. A structural interpretation of voltage-gated potassium channel inactivation. Prog. Biophys. Mol. Biol. 92, 185–208 (2006).
Lacroix, J. J. et al. Intermediate state trapping of a voltage sensor. J. Gen. Physiol. 140, 635–652 (2012).
Hou, P. et al. Inactivation of KCNQ1 potassium channels reveals dynamic coupling between voltage sensing and pore opening. Nat. Commun. 8, 1730 (2017).
Zaydman, M. A. et al. Domain-domain interactions determine the gating, permeation, pharmacology, and subunit modulation of the IKs ion channel. Elife 3, e03606 (2014).
Wu, D. et al. State-dependent electrostatic interactions of S4 arginines with E1 in S2 during Kv7.1 activation. J. Gen. Physiol. 135, 595–606 (2010).
Silva, J. R. et al. A multiscale model linking ion-channel molecular dynamics and electrostatics to the cardiac action potential. Proc. Natl Acad. Sci. USA 106, 11102–11106 (2009).
Tao, X., Lee, A., Limapichat, W., Dougherty, D. A. & MacKinnon, R. A gating charge transfer center in voltage sensors. Science 328, 67–73 (2010).
Willegems, K. et al. Structural and electrophysiological basis for the modulation of KCNQ1 channel currents by ML277. Nat. Commun. 13, 1–17 (2022).
Ruscic, K. J. et al. IKs channels open slowly because KCNE1 accessory subunits slow the movement of S4 voltage sensors in KCNQ1 pore-forming subunits. Proc. Natl Acad. Sci. USA 110, E559–E566 (2013).
Barro-Soria, R. et al. KCNE1 divides the voltage sensor movement in KCNQ1/KCNE1 channels into two steps. Nat. Commun. 5, 3750 (2014).
Pusch, M., Bertorello, L. & Conti, F. Gating and flickery block differentially affected by rubidium in homomeric KCNQ1 and heteromeric KCNQ1/KCNE1 potassium channels. Biophysical J. 78, 211–226 (2000).
Ma, D. et al. Structural mechanisms for the activation of human cardiac KCNQ1 channel by electro-mechanical coupling enhancers. Proc. Natl Acad. Sci. USA 119, e2207067119 (2022).
He, J., Li, T. & Huang, S. Y. Improvement of cryo-EM maps by simultaneous local and non-local deep learning. Nat. Commun. 14, 3217 (2023).
Osteen, J. D. et al. KCNE1 alters the voltage sensor movements necessary to open the KCNQ1 channel gate. Proc. Natl Acad. Sci. USA 107, 22710–22715 (2010).
Werry, D., Eldstrom, J., Wang, Z. & Fedida, D. Single-channel basis for the slow activation of the repolarizing cardiac potassium current, I(Ks). Proc. Natl Acad. Sci. U. S. A 110, E996–E1005 (2013).
Eldstrom, J., Wang, Z., Werry, D., Wong, N. & Fedida, D. Microscopic mechanisms for long QT syndrome type 1 revealed by single-channel analysis of I(Ks) with S3 domain mutations in KCNQ1. Heart Rhythm 12, 386–394 (2015).
Eldstrom, J., McAfee, D. A., Dou, Y., Wang, Y. & Fedida, D. ML277 regulates KCNQ1 single-channel amplitudes and kinetics, modified by voltage sensor state. J. Gen. Physiol. 153 https://doi.org/10.1085/jgp.202112969 (2021).
Taylor, K. C. et al. Structure and physiological function of the human KCNQ1 channel voltage sensor intermediate state. Elife 9 https://doi.org/10.7554/eLife.53901 (2020).
Seebohm, G., Sanguinetti, M. C. & Pusch, M. Tight coupling of rubidium conductance and inactivation in human KCNQ1 potassium channels. J. Physiol. 552, 369–378 (2003).
Kasuya, G. & Nakajo, K. Optimized tight binding between the S1 segment and KCNE3 is required for the constitutively open nature of the KCNQ1-KCNE3 channel complex. Elife 11 https://doi.org/10.7554/eLife.81683 (2022).
Huang, H. et al. Disease-linked supertrafficking of a potassium channel. J. Biol. Chem. 296, 100423 (2021).
Wang, Y. et al. Probing the structural basis for differential KCNQ1 modulation by KCNE1 and KCNE2. J. Gen. Physiol. 140, 653–669 (2012).
Xu, Y. et al. Building KCNQ1/KCNE1 channel models and probing their interactions by molecular-dynamics simulations. Biophys. J. 105, 2461–2473 (2013).
Chan, M. et al. A generic binding pocket for small molecule I(Ks) activators at the extracellular inter-subunit interface of KCNQ1 and KCNE1 channel complexes. eLife 12, 1–31 (2023).
Kuenze, G. et al. Allosteric mechanism for KCNE1 modulation of KCNQ1 potassium channel activation. eLife 9 https://doi.org/10.7554/eLife.57680 (2020).
Kang, C. et al. Structure of KCNE1 and implications for how it modulates the KCNQ1 potassium channel. Biochemistry 47, 7999–8006 (2008).
Peng, G., Barro-Soria, R., Sampson, K. J., Larsson, H. P. & Kass, R. S. Gating mechanisms underlying deactivation slowing by two KCNQ1 atrial fibrillation mutations. Sci. Rep. 7, 45911 (2017).
Chan, P. J. et al. Characterization of KCNQ1 atrial fibrillation mutations reveals distinct dependence on KCNE1. J. Gen. Physiol. 139, 135–144 (2012).
Wu, D., Pan, H., Delaloye, K. & Cui, J. KCNE1 remodels the voltage sensor of Kv7.1 to modulate channel function. Biophys. J. 99, 3599–3608 (2010).
Panaghie, G. & Abbott, G. W. The role of S4 charges in voltage-dependent and voltage-independent KCNQ1 potassium channel complexes. J. Gen. Physiol. 129, 121–133 (2007).
Meisel, E. et al. KCNQ1 channels do not undergo concerted but sequential gating transitions in both the absence and the presence of KCNE1 protein. J. Biol. Chem. 287, 34212–34224 (2012).
Bartos, D. C. et al. R231C mutation in KCNQ1 causes long QT syndrome type 1 and familial atrial fibrillation. Heart Rhythm 8, 48–55 (2011).
Yahnell, J., Wu, X., Perez, M. E. & Larsson, H. P. The effects of long QT syndrome associated mutation R231C on the cardiac IKs channel. Biophysical J. 121, 240A (2021).
Judah, Y., Wu, X., Perez, M. E. & Larsson, P. The efects of long QT syndrome associated mutation R231C on the cardiac IKs channel. Biophys. J. 121, 240a (2022).
Hou, P., Shi, J., White, K. M., Gao, Y. & Cui, J. ML277 specifically enhances the fully activated open state of KCNQ1 by modulating VSD-pore coupling. Elife 8 https://doi.org/10.7554/eLife.48576 (2019).
Mkrtchyan, L. et al. Ion permeation through a narrow cavity constriction in KCNQ1 channels: Mechanism and implications for pathogenic variants. Proc. Natl Acad. Sci. USA 121, e2411182121 (2024).
Es-Salah-Lamoureux, Z., Fougere, R., Xiong, P. Y., Robertson, G. A. & Fedida, D. Fluorescence-tracking of activation gating in human ERG channels reveals rapid S4 movement and slow pore opening. PLoS One 5, e10876 (2010).
Sastre, D. & Fedida, D. Voltage Clamp Fluorometry: Illuminating the Dynamics of Ion Channels. Methods Mol. Biol. 2796, 119–138 (2024).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr D. Biol. Crystallogr 66, 486–501 (2010).
Jamali, K. et al. Automated model building and protein identification in cryo-EM maps. Nature 628, 450–457 (2024).
Croll, T. I. ISOLDE: a physically realistic environment for model building into low-resolution electron-density maps. Acta Crystallogr D. Struct. Biol. 74, 519–530 (2018).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr D. Struct. Biol. 75, 861–877 (2019).
Williams, C. J. et al. MolProbity: More and better reference data for improved all-atom structure validation. Protein Sci. 27, 293–315 (2018).
Goddard, T. D. et al. UCSF ChimeraX: Meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Pettersen, E. F. et al. UCSF Chimera-a visualization system for exploratory research and analysis. J. Comput Chem. 25, 1605–1612 (2004).
Waterhouse, A. et al. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res 46, W296–W303 (2018).
Jo, S., Kim, T. & Im, W. Automated builder and database of protein/membrane complexes for molecular dynamics simulations. PLoS One 2, e880 (2007).
Phillips, J. C. et al. Scalable molecular dynamics with NAMD. J. Comput Chem. 26, 1781–1802 (2005).
Huang, J. et al. CHARMM36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods 14, 71–73 (2017).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Noskov, S. Y. & Roux, B. Control of ion selectivity in LeuT: two Na+ binding sites with two different mechanisms. J. Mol. Biol. 377, 804–818 (2008).
Venable, R. M., Luo, Y., Gawrisch, K., Roux, B. & Pastor, R. W. Simulations of anionic lipid membranes: development of interaction-specific ion parameters and validation using NMR data. J. Phys. Chem. B 117, 10183–10192 (2013).
Darden, T., York, D. & Pedersen, L. Particle mesh Ewald: An N ⋅log(N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089–10092 (1993).
Steinbach, P. J. & Brooks, B. R. New spherical-cutoff methods for long-range forces in macromolecular simulation. J. Computational Chem. 15, 667–683 (1994).
Ryckaert, J.-P., Ciccotti, G. & Berendsen, H. J. C. Numerical integration of the cartesian equations of motion of a system with constraints: molecular dynamics of n- alkanes. J. Computational Phys. 23, 327–341 (1977).
Miyamoto, S. & Kollman, P. A. Settle: An analytical version of the SHAKE and RATTLE algorithm for rigid water models. J. Computational Chem. 13, 952–962 (1992).
Nappi, M. et al. Constitutive opening of the Kv7.2 pore activation gate causes KCNQ2-developmental encephalopathy. Proc. Natl Acad. Sci. USA 121, e2412388121 (2024).
Feller, S. E., Zhang, Y., Pastor, R. W. & Brooks, B. R. Constant pressure molecular dynamics simulation: the Langevin piston method. J. Chem. Phys. 103, 4613–4621 (1995).
Smart, O. S., Neduvelil, J. G., Wang, X., Wallace, B. A. & Sansom, M. S. HOLE: a program for the analysis of the pore dimensions of ion channel structural models. J. Mol. Graph 14, 354–360, 376 (1996).
Smart, O. S., Breed, J., Smith, G. R. & Sansom, M. S. P. A novel method for structure-based prediction of ion channel conductance properties. Biophysical J. 72, 1109–1126 (1997).
Meng, E. C. et al. UCSF ChimeraX: Tools for structure building and analysis. Protein Sci. 32, e4792 (2023).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph 14, 33-38–27-38 (1996).
Fedida, D., Sastre, D., Dou, Y., Westhoff, M. & Eldstrom, J. Evaluating sequential and allosteric activation models in IKs channels with mutated voltage sensors. J. Gen. Physiol. 156, 1–32 (2024).
Acknowledgements
We thank Dr. Jianmin Cui for the C214A/G219C/C331A hKCNQ1 construct, Dr. Yoshihiro Kubo for the pGEMHE vector. Grids were prepared and data collected at the High Resolution Macromolecular Electron Microscopy (HRMEM) facility at the University of British Columbia (https://cryoem.med.ubc.ca). We thank Claire Atkinson, Amy Wo, Barathy Deivanayaga, Liam Worrall and Natalie Strynadka. HRMEM is funded by the Canadian Foundation for Innovation and the British Columbia Knowledge Development Fund. This work was supported by: the Natural Sciences and Engineering Research Council of Canada [Grant# RGPIN-2022-03021], the Canadian Institutes of Health Research [Grant# PJT-156181 and PJT-518041], and the Heart and Stroke Foundation of Canada [Grant# G-21-0031566, and G-24-0036478] to D.F.; and by grants from the Canadian Institutes of Health Research [Grant# PJT-153305] to F.V.P. L.M. and A.R. acknowledge ISCRA for awarding this project access to the LEONARDO supercomputer, owned by the EuroHPC Joint Undertaking, hosted by CINECA (Italy). The authors declare no competing financial interests. E.K. acknowledges support from Michael Smith Health Research BC Research Trainee award RT-2022-2735. M.C. holds a graduate research scholarship from the Canadian Institutes of Health Research. The authors acknowledge that this work was carried out on the UBC Point Gray (Vancouver) campus, which sits on the traditional, ancestral, unceded territory of the xʷməθkʷəy̓əm (Musqueam) First Nation.
Author information
Authors and Affiliations
Contributions
J.E. and D.F. conceptualized the study. E.K., S.R., and F.A. produced protein in mammalian cells for structural studies. E.K. and S.R. purified the proteins, and E.K. prepared grids and analyzed Cryo-EM data. S.M. assisted with Cryo-EM data analysis. D.S., J.E., Y.D., and M.C. collected electrophysiology data. A.R. and L.M. performed MD simulations. E.K., D.M., J.E., A.R., L.M., F.V.P., and D.F. reviewed, analyzed, and interpreted data. E.K., D.S., J.E., A.R., L.M., F.V.P., and D.F. wrote, edited, and reviewed the manuscript. All authors approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kyriakis, E., Sastre, D., Eldstrom, J. et al. A physiologically-relevant intermediate state structure of a voltage-gated potassium channel. Nat Commun 16, 8814 (2025). https://doi.org/10.1038/s41467-025-64060-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-64060-3