Abstract
Targeting cancer cell plasticity through chromatin organization is an emerging research area, yet the molecular mechanisms that govern chromatin loop formation remain unclear. Here, we develop a CRISPR screen based on our engineered live-cell CTCF-cohesin contact reporters to identify regulators of chromatin loops. Our findings reveal that tousled-like kinase 2 (TLK2) functions as a key regulator of chromatin loop formation during the cancer stemness transition. Mechanistically, TLK2 phosphorylates DYNLL1, enhancing its interaction with CTCF to promote CTCF-cohesin hub formation at the KLF4 locus. Suppressing TLK2 impairs cancer stemness plasticity, sensitizes cancer cells to cytotoxic stress in vitro, and reduces lung metastases and enhances immunotherapy response in breast cancer mouse models. Clinically, elevated TLK2 expression correlates with poor prognosis in breast cancer patients. Collectively, these findings identify TLK2 as a potential therapeutic target for mitigating cancer stemness plasticity, highlighting chromatin loop-targeting therapy as a promising strategy to eradicate cancer stem cells.
Similar content being viewed by others
Introduction
Cancer stem cells (CSCs) pose a critical challenge in cancer therapy due to their central role in therapy resistance and tumor relapse, rendering this sub-population an attractive target for intervention1. However, efforts to target CSCs in clinical trials have yielded suboptimal outcomes2. Emerging evidence has revealed intricate inter-transitions between CSCs and non-CSCs, complicating therapeutic strategies that focus on CSCs3. Therefore, interventions targeting CSCs should also consider preventing cellular plasticity transitions. Yet, the underlying mechanisms governing cancer cell plasticity and the optimal strategies for its therapeutic targeting remain to be fully elucidated.
Conrad Waddington’s epigenetic landscape theory suggests that chromatin organization serves as a barrier restricting cellular identity4. Advances in genome organization affirm its role in establishing and maintaining distinct cell identities during development5. Additionally, aberrant chromatin organization has been linked to cancer progression and serves as potential targets6,7,8. Thus, a strategy to target cancer cell plasticity by intervening chromatin organization appears promising.
Our understanding of chromatin organization signals has so far primarily focused on intranuclear chromatin-associated factors such as CTCF and cohesin. CTCF establishes contact points for long-range chromatin loops9,10. Cohesin, loaded at NIPBL binding sites, extrudes chromatin fibers and halts extrusion until engaging CTCF anchors to shape chromatin into loops11,12. These loops are eventually released by the action of WAPL13,14. Cohesin loop extrusion is believed to play a central role in establishing chromatin architecture. Conversely, our understanding of regulators that modulate chromatin loop formation is still limited.
Here we uncover extensive changes in chromatin states including CTCF-cohesin contact sites, H3K27ac modification and chromatin accessibility during non-CSC to CSC transition. We thus develop a CRISPR screen using CTCF-cohesin contact reporters to explore key mechanisms regulating chromatin looping process. We identify tousled-like kinase 2 (TLK2) as a targetable regulator of chromatin loop formation in cancer stemness transition, supporting chromatin loop-targeting therapy to eliminate the plastic CSCs.
Results
CTCF-cohesin hubs changes during non-CSC to CSC transition
To investigate whether chromatin state is reprogrammed during the non-CSC to CSC transition, we enriched CSCs by mammosphere formation assays and compared the genome-wide binding patterns of chromatin architectural proteins CTCF and cohesin (RAD21) between CSCs and non-CSCs. We defined the co-binding sites of CTCF and cohesin as CTCF-cohesin hubs (Fig. 1a, b). Notably, during the non-CSC to CSC transition, 41.91% and 47.57% of CTCF-cohesin hubs in MDA-MB-231 and MCF7 cells, respectively, displayed either a loss or a gain (Supplementary Data 1, 2, Fig. 1c, and Supplementary Fig. 1a, b). Similarly, 46.68% and 54.58% of the identified H3K27ac peaks displayed either a loss or a gain (Fig. 1d and Supplementary Fig. 1c), and 41.53% and 39.60% of the identified open chromatin (as assessed by ATAC-seq) regions showed an increase or a decrease in accessibility during CSC formation in MDA-MB-231 and MCF7 cells, respectively (Supplementary Data 3, Fig. 1e and Supplementary Fig. 1d, left panels).
a Representative examples of consensus, loss, and gain of CTCF-cohesin hubs in CSCs and non-CSCs. CTCF and RAD21 ChIP-seq were performed in adherent (non-CSCs-enriched) or mammosphere (CSCs-enriched) MDA-MB-231 cells. Triangles represent CTCF binding motifs. b, ChIP-qPCR and ReChIP-qPCR validation of CTCF and RAD21 binding at indicated sites. (panel 1, p = 0.0072; 0.0258; <0.0001; 0.0126; 0.0299; 0.0067; <0.0001; 0.0043; <0.0001; <0.0001; 0.0342; 0.0335; panel 2, p = 0.0320; 0.0040; 0.0206; 0.0202; 0.0174; 0.0061; 0.0381; 0.0049; panel 3, p = 0.0060; 0.0221; <0.0001; <0.0001; 0.0063; 0.0252; <0.0001; 0.0036; 0.0018; 0.0085; 0.0034; panel 4, p = < 0.0001; 0.0040; 0.061; 0.0068) c Heatmap of consensus, loss, and gain of CTCF-cohesin hubs. d Heatmap of H3K27ac changes. e Left panel: Heatmap of accessible chromatin regions. Right panel: De novo motif enrichment of accessible chromatin regions using HOMER. f GO analysis of the top 1,000 dynamically accessible chromatin regions using GREAT. Statistical analysis was performed using the GREAT platform with default settings. The binomial test used is two-sided, and adjusted p values were calculated by GREAT to account for multiple comparisons. g Correlation analysis between the number of differentially expressed genes (DEGs) and the number of dynamic CTCF-cohesin hubs per megabase across individual chromosomes in non-CSC to CSC transition. Linear regression and nonparametric Spearman correlation coefficient were used to assess the relationship between differentially expressed genes (DEGs) and dynamic CTCF-cohesin hubs per chromosome. The Spearman correlation coefficient (r), exact p value, and regression line were reported in the plot. h ALDH assay for CSC frequency after siRAD21 or CTCF-cohesin proximity induction. Means and standard deviation from biological triplicates (n = 3) are shown. (p = 0.0001; <0.0001) i, Western blot analysis of stemness markers in MDA-MB-231 cells treated with siRAD21 or chemical induction of CTCF-cohesin proximity. Error bars represent mean ± SD (b, h). The merged images are shown from two biological replicates (a, c, d). Statistical significance was assessed using one-way ANOVA with Bonferroni correction (b, h); *P < 0.05; **P < 0.01; ***P < 0.001. Source data are provided as a Source Data file.
Analyzing differentially accessible chromatin regions (DARs) unveiled that both gained and lost DARs are enriched for CTCF or CTCFL binding motifs (Fig. 1e and Supplementary Fig. 1d, right panels). Additionally, the CSC gained DARs are enriched for FOXM1 binding motif, which is essential for the self-renewal of CSCs15 and is a major predictor of adverse outcomes in a variety of different cancer types16. Further analysis of differential chromatin accessibility across cell cycle phases indicated that the FOXM1 binding motif increases accessibility in all phases (Supplementary Fig. 2). The GREAT (Genomic Regions Enrichment of Annotations Tool) gene ontology (GO) analysis illustrated that the CSC-gained DARs were relevant to stemness-related processes, including mesenchymal cell proliferation, cell fate determination, telomerase activity, and Wnt signaling pathway, whereas the CSC-lost DARs were relevant to differentiated mammary epithelium processes, including hormone receptor binding, keratinocyte development, and growth factor binding (Fig. 1f). Furthermore, the number of dynamic CTCF-cohesin hub per Mb (mega base pairs) and the number of differentially expressed genes (DEGs) in individual chromosome showed strong positive linear correlations (Supplementary Data 4–6, Fig. 1g, and Supplementary Fig. 3a–c).
We next measured aldehyde dehydrogenase (ALDH) activity and the expression of cancer stemness markers in cells where CTCF-cohesin hubs were disturbed using siRNA targeting RAD21 or the chemically induced proximity (CIP) assay to immobilize CTCF-cohesin hubs (Supplementary Fig. 3d–f). The interventions reduced the proportion of ALDH-positive cells and decreased the expression of cancer stemness markers FOXM1, c-Myc, KLF4, and TERT (Fig. 1h, i). Collectively, these results indicate that CTCF-cohesin hubs exhibit dynamic changes during non-CSC to CSC transition.
CRISPR screen identifies TLK2 as CTCF-cohesin hub regulator
To identify the regulators governing CTCF-cohesin hub formation, we devised an approach to report the CTCF-cohesin hubs in living cells using bimolecular fluorescence complementation (BiFC) assay (Fig. 2a and Supplementary Fig. 4a). In this system, yellow fluorescence is activated when VC155 and VN173 are brought together in close proximity by the association of CTCF and cohesin at chromatin contact sites. This fluorescence intensity serves as a readout of the overall abundance of CTCF-cohesin hubs. The flow cytometry results showed that knocking down NIPBL significantly reduced the fluorescence intensity of CTCF-cohesin hubs, whereas knocking down WAPL significantly increased CTCF-cohesin complex fluorescence intensity (Fig. 2b and Supplementary Fig. 4b). This observation establishes the utility of CTCF-cohesin BiFC reporters as a valid way to score CTCF-cohesin interactions.
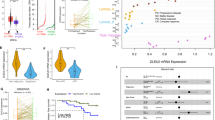
a Schematic of the CTCF-cohesin hub reporter system based on bimolecular fluorescence complementation (BiFC). Yellow fluorescent protein (YFP) Venus is cleaved into N-terminal (VN173/YFPn) and C-terminal (VC155/YFPc) fused with cohesin subunits (RAD21 or SMC3) and CTCF, respectively. YFPn and YFPc form intact fluorescent protein Venus upon CTCF-cohesin interaction. b Validation of CTCF-cohesin hub reporters using siRNA-mediated knockdown of NIPBL and WAPL. Left panel: Flowchart of validation assay. Middle panel: Representative flow cytometry plot of reporter validation in MDA-MB-231 cells. Right panel: Median fluorescence intensity (MFI) of Venus in siNC, siNIPBL and siWAPL cells. Means and standard deviation from biological triplicates (n = 3) are shown. (p = 0.0055; 0.0004; 0.0457; 0.0083) c Flowchart illustrating the CRISPR screening approach used to identify epigenetic regulators of CTCF-cohesin hubs. d Heatmap of genes potentially regulating CTCF-cohesin hubs. The gene mutation frequency in BRCA patients was labeled. e Flow cytometry validation of candidate genes using CTCF-RAD21 BiFC assays. f Normalized counts of 10 sgRNAs targeting TLK2 in BiFC-high and BiFC-low populations from the CRISPR screen. Each point represents a distinct sgRNA targeting TLK2. Read counts were normalized to total sequencing depth per group. sgRNA-level variation was used to assess enrichment differences between groups. Statistical significance was assessed using the Wald test, and false discovery rates were calculated by MAGeCK. g MA plot showing differential CTCF and RAD21 binding in TLK2 knockdown versus control cells, analyzed using DiffBind. Magenta points represent significantly differentially bound sites (FDR < 0.05), while the blue shading indicates the overall density of all binding events across the range of log concentration. The results were acquired with two biological replicates. h Heatmap showing consensus and loss of CTCF-cohesin hubs in TLK2 knockdown and control MDA-MB-231 cells. The merged peaks are shown from two biological replicates. Error bars represent mean ± SD (b). Statistical significance was determined using one-way ANOVA with Bonferroni correction (b); ns not significant; *P < 0.05; **P < 0.01; ***P < 0.001. Source data are provided as a Source Data file.
Subsequently, we screened a self-constructed guide RNA (gRNA) library of human epigenetic factors (see “Methods” section) with the screening workflow as illustrated in Fig. 2c. We applied the MAGeCK-MLE package to analyze the read counts of each sgRNA, and calculated beta score to evaluate the regulatory genes (Supplementary Data 7 and Supplementary Fig. 4c–f). The top 20 ranked genes that potentially regulated CTCF-cohesin hub formation in a direct or indirect way were displayed in the heatmap (Fig. 2d). The sgTLK2, sgPHF17, sgNONO, sgNIPBL, sgPRMT5 and sgELAVL1 displayed significant differences when compared between the CTCF-cohesin and the control BiFC groups. We then annotated these genes with mutation frequencies and functional GO using expression and co-expression data across TCGA breast cancer types. Gene expression co-expression analysis is a well-established approach used to infer putative gene functions based on the functional annotation of genes that are co-expressed with each other, as demonstrated in previous studies17. Specifically, DKC1, TLK2, NIPBL, BRD2, NONO, and ELAVL1 showed a high correlation with chromatin-related GO terms, such as sister chromatid cohesion, as identified through this co-expression analysis (Supplementary Data 8, 9).
We validated 15 candidate genes using BiFC-Flow cytometry assay. The knockout of USP16, ERBB4, TP53, DKC1, TLK2, UBE2A and NONO significantly reduced CTCF-cohesin hub abundance, whereas deletion of ELAVL1, YEATS2, CPEB1 and RAG2 significantly increased CTCF-cohesin hub abundance (Fig. 2e and Supplementary Fig. 4g). We further examined the effect of these genes on mammosphere formation. As shown in Supplementary Fig. 4h, the knockout of USP16, DKC1, TLK2, ELAVL1 and CPEB1 significantly inhibited mammosphere formation in MDA-MB-231 cells.
Among the array of identified hits, TLK2 stands out with a compelling negative selection beta-score (βCTCF&RAD21 = −0.147, βCTCF&SMC3 = −0.349, βYFPn&YFPc = 0.046), and a high frequency mutation score (Data retrieved from cBioPortal18: 9% in METABRIC cohorts19; 10% in Breast Invasive Carcinoma cohorts20; 25% in The Metastatic Breast Cancer Project cohorts21, primarily due to gene amplification, Fig. 2d, f). ChIP-seq analyses affirmed that the chromatin occupancy of CTCF-cohesin hubs was reduced in response to TLK2 knockdown (Fig. 2g, h and Supplementary Fig. 4i).
TLK2 promotes CTCF-cohesin hubs via DYNLL1 phosphorylation
Since the kinase activity of TLK2 is necessary for CTCF-cohesin hub formation and cancer stemness (Fig. 3a and Supplementary Fig. 5a, b), we initially examined ASF1A and ASF1B, which are well-known substrates of TLK222,23,24,25,26,27,28,29,30, in regulating CTCF-cohesin hub formation in mammosphere cells. TLK2 knockdown reduced about 50% of the CTCF-cohesin BiFC signal, while double knockdown of both ASF1A and ASF1B resulted in a 15% reduction in the CTCF-cohesin BiFC signal (Supplementary Fig. 5c, d), suggesting the existence of another TLK2 substrate involved in regulating CTCF-cohesin hub formation.
a CTCF-cohesin BiFC assays in TLK2 knockdown MDA-MB-231 cells expressing either wild-type (WT) or kinase-dead (KD) shRNA-resistant TLK2. Means and standard deviation from biological triplicates (n = 3) are shown. (p = < 0.0001; <0.0001; 0.9599; <0.0001; <0.0001; <0.0001; 0.6822; <0.0001) b APEX2-DIA proteomic workflow identifying TLK2 interactors; empty vector as control. Volcano plot from triplicates (n = 3), two-sided unpaired t-test. c Venn diagram showing interacting partners of TLK2 and CTCF. Statistical details same as above. d APEX2-IP assay validating TLK2-DYNLL1 interactions. e, f Co-IP using nuclear fractions and PLA confirming DYNLL1-TLK2-CTCF interactions. For each condition, 50 cells from three random fields were analyzed to generate the plot. Data are presented as mean ± SD; center indicates the mean. Scale bar: 10 μm. g, h CTCF-cohesin BiFC assays in DYNLL1 knockdown MDA-MB-231 cells with or without TLK2 overexpression. Data are presented as mean ± SD; center indicates the mean. (g p = < 0.0001; <0.0001; <0.0001; <0.0001; h p = 0.0045; 0.0007; 0.0006; 0.0014; 0.0017; 0.0009) i Western blot of DYNLL1 phosphorylation in TLK2 knockdown cells. j In vitro kinase assay showing TLK2-mediated phosphorylation of DYNLL1. Arrow shows phosphorylated DYNLL1. k LC-MS/MS confirming phosphorylation of DYNLL1 at Ser14. l CTCF-cohesin BiFC assays in TLK2 knockdown MDA-MB-231 cells rescued with DYNLL1 S14D or S14A mutants. (p = < 0.0001; <0.0001; 0.0531; 0.0013; 0.0012; 0.1322) m Western blot showing enhanced binding of the DYNLL1 S14D mutant to CTCF compared with WT and S14A forms. n Western blot demonstrating reduced CTCF binding to DYNLL1 upon TLK2 knockdown. o Domain mapping assays showing DYNLL1 binding domains and motifs of CTCF. wt wild-type motif, mut mutant motif. p Schematic depicting the TLK2-DYNLL1 axis in regulating CTCF-cohesin hub formation. Error bars represent mean ± SD (a, g, h, l). Statistical significance was determined using one-way ANOVA with Bonferroni correction (a, g, h, l); ns not significant; *P < 0.05; **P < 0.01; ***P < 0.001. Source data are provided as a Source Data file.
We next conducted proximity labeling immunoprecipitation assay mediated by engineered ascorbic acid peroxidase 2 (APEX2) fused with TLK2, and combined with mass spectrometry (MS) to explore its substrates (Fig. 3b). A set of well-known TLK2 interactors, such as ASF1A, UBN1, UBN2, CEBPB and DYNLL1, were identified in our assay (Supplementary Data 10). Upon cross-referencing the list with known CTCF-interacting proteins, DYNLL1, a previously characterized TLK2 interactor26,28,31, was identified as a prominent candidate for further investigation (Fig. 3c). DYNLL1 has been demonstrated to function like a molecular glue to stabilize dimeric structures with many binding partners32,33. The interaction between TLK2 and DYNLL1 was also validated by APEX2-IP, nuclear co-IP, and proximity ligation assay (PLA) assays (Fig. 3d–f and Supplementary Fig. 5e, f). The knockdown of DYNLL1 significantly impaired CTCF-cohesin hub abundance (Fig. 3g, h and Supplementary Fig. 5g).
Next, we ask whether activation of TLK2 would lead to DYNLL1 phosphorylation during chromatin hub formation. TLK2 knockdown significantly decreases serine phosphorylation of DYNLL1 (Fig. 3i). The in vitro autoradiography shows that TLK2 phosphorylates DYNLL1 and ASF1A (Fig. 3j and Supplementary Fig. 5h). The highly conserved DYNLL1 contains 4 serine residues and 4 threonine residues (Supplementary Fig. 5i). The group-based phosphorylation site predicting and scoring (GPS34, version 3.0) algorithm predicted that the Ser14 residue is the potential TLK2 target (Supplementary Fig. 5j). The LC-MS/MS further confirms the presence of a modified peptide containing phosphorylated Ser14 (Fig. 3k). The S14D (phosphomimetic mutation, Ser to Asp) mutant of DYNLL1 rescues CTCF-cohesin hub formation, while the S14A (non-phosphorylatable mutation, Ser to Ala) mutant cannot (Fig. 3l and Supplementary Fig. 5k). Furthermore, the S14D mutant of DYNLL1 binds CTCF more effectively than the wild-type (WT) and S14A forms (Fig. 3m). The knockdown of TLK2 was found to reduce the binding of CTCF to DYNLL1 (Fig. 3n).
Moreover, we utilized the ChatGPT o3-mini model and reported DYNLL1 binding motifs35,36 to predict potential binding motifs within CTCF (see “Methods” section). The predicted motifs are listed in Supplementary Data 11. The high-scoring motifs, ITLQVVNME (N-terminal) and RFTQSG (sixth zinc finger domain), were mutated as ITAAAVNME and RFAASG to validate their binding specificity. As shown in Fig. 3o, Co-IP assays revealed a reduced binding of DYNLL1 to the mutant CTCF proteins compared to wild-type CTCF. These results confirm that both the ITLQVVNME and RFTQSG motifs are critical for DYNLL1 binding.
In conclusion, our results suggest that TLK2 phosphorylates DYNLL1, potentially at Ser14, to promote its interaction with CTCF and facilitate CTCF-cohesin hub formation (Fig. 3p).
The TLK2-DYNLL1 axis facilitates non-CSC to CSC transition
Consistent with previous reports that TLK2 regulates cell proliferation27,37,38, we also found that knockdown of TLK2 inhibited cell growth and cell cycle progression to varying degrees in multiple cancer cell lines (Supplementary Fig. 6a–c). For cancer stemness, the knockdown of TLK2 significantly reduced the proportion of ALDH-positive cells and impairs mammosphere formation in MDA-MB-231 cells (Fig. 4a, b). Long-term inhibition of TLK2 (up to 21 days) demonstrated that both CTCF-cohesin hubs and the proportion of ALDH-positive cells persistently remained at low levels (Supplementary Fig. 6d, e). Similarly, the knockdown of DYNLL1 displayed a similar function in inhibiting ALDH-positive cells and mammosphere formation (Fig. 4a, b). The S14D mutant of DYNLL1 rescued these functions, whereas the S14A mutant cannot (Fig. 4c, d). Limiting dilution assays (LDA) in NOD/SCID mice further confirmed that TLK2 knockdown significantly reduced the CSC frequency (Fig. 4e). Together, these results indicate that TLK2 plays a critical role in regulating cell proliferation, cancer stemness, and CTCF-cohesin hubs.
a Flow cytometry analysis of ALDH-positive cell rates in MDA-MB-231 cells with or without TLK2 or DYNLL1 knockdown. Means and standard deviation from biological triplicates (n = 3) are shown. (p = <0.0001; <0.0001; <0.0001; <0.0001) b Mammosphere formation assay of MDA-MB-231 cells with or without TLK2 or DYNLL1 knockdown. Scale bar, 50 μm. Means and standard deviation from biological triplicates (n = 3) are shown. (Diameter: p = <0.0001; <0.0001; <0.0001; <0.0001; Number: p = <0.0001; <0.0001; <0.0001; <0.0001) c Flow cytometry analysis of ALDH-positive cell rates in TLK2 knockdown MDA-MB-231 cells with DYNLL1 S14D or S14A mutant rescue. Means and standard deviation from biological triplicates (n = 3) are shown. (p = <0.0001; 0.1406; <0.0001) d Mammosphere formation assay in TLK2 knockdown MDA-MB-231 cells with S14D or S14A DYNLL1 mutant rescue. Scale bar, 50 μm. Means and standard deviation from biological triplicates (n = 3) are shown. (Diameter: p = <0.0001; 0.0505; <0.0001; Number: p = <0.0001; 0.0702; <0.0001) e Limiting dilution assay (LDA) in NOD/SCID mice comparing tumor formation in TLK2 knockdown versus control cells. Resected tumor xenografts were subjected to mammosphere formation assays to evaluate mammosphere-forming capacity. Means and standard deviation from biological triplicates (n = 3) are shown. (p = <0.0001) f ALDH-low (ALDHlo) MDA-MB-231 cells were sorted, re-seeded, and monitored for ALDH-positive rates over 6 days. Means and standard deviation from biological triplicates (n = 3) are shown. (p = 0.0003; 0.0011; <0.0001; 0.0004; 0.0010) g Co-culture of mNeonGreen-labeled control MDA-MB-231 cells with mRuby3-labeled TLK2 knockdown cells under 2D normal or 3D mammosphere conditions, analyzed by flow cytometry for proportional changes. Error bars represent mean ± SD (a, b, c, d, f). Statistical significance was determined by one-way ANOVA with Bonferroni correction (a, b, c, d); two-tailed unpaired Student’s t test (e, f); ns not significant; *P < 0.05; **P < 0.01; ***P < 0.001. Source data are provided as a Source Data file.
We next explored whether TLK2 was essential for non-CSC to CSC plastic transition. MDA-MB-231 cells with low ALDH activities (ALDHlo) were isolated using flow cytometry and re-seeded in plates. TLK2 inhibition significantly slowed the recovering efficiency and frequency of ALDH positive cells (Fig. 4f), indicating its vital role in acquiring cancer stem cell characteristics and promoting their proliferation.
To prove the conjecture that the reciprocal transition between non-CSCs and CSCs can determine the cell fate in response to stress, we labeled wild-type MDA-MB-231 cells with mNeonGreen and TLK2 knockdown cells with mRuby3, then mixed them and co-cultured them under stress conditions, and analyzed the proportional changes. The results showed that TLK2 knockdown MDA-MB-231 cells were marginally decreased in normal culture condition, whereas they were significantly decreased in mammosphere culture condition (Fig. 4g). These results indicate that TLK2 knockdown enhances the vulnerability of MDA-MB-231 cells to stress, particularly in mammosphere culture conditions, suggesting that TLK2 plays a crucial role in maintaining cancer stem cell characteristics and their ability to withstand adverse environments.
The TLK2-DYNLL1 axis regulates CTCF-cohesin hubs near KLF4
To identify the cancer stemness genes regulated by the TLK2-DYNLL1 axis, we merged the lists of downregulated genes from TLK2 knockdown, upregulated genes during mammosphere formation, CTCF-cohesin-bound genes, and stemness genes. The Yamanaka factor Krüppel-like factor 4 (KLF4) known for enhancing breast cancer stemness39 was identified as a prominent candidate for further investigation (Fig. 5a). CTCF ChIA-PET data from H1-hESC (4DN Data Portal40) highlight that the KLF4 promoter interacts with an enhancer located at the end of the LINC01509 gene (Fig. 5b).
a Venn diagram illustrating candidate stemness genes regulated by the TLK2-DYNLL1 axis. b CTCF ChIA-PET data from H1-hESC (4DN Data Portal) indicating interaction between the KLF4 promoter and an enhancer located within the LINC01509 gene. Red circle presents the potential interaction. c Analysis of RNA expression, chromatin accessibility, and binding of CTCF, RAD21, DYNLL1, and H3K27ac at the KLF4 locus in TLK2 knockdown versus control cells. The merged peaks are shown from two biological replicates. d, e qPCR and Western blot showing KLF4 expression in TLK2 and DYNLL1 knockdown cells. Means and standard deviation from biological triplicates (n = 3) are shown. (p = <0.0001; <0.0001; <0.0001; <0.0001) f Chromosome conformation capture (3C-qPCR) assays measuring interaction frequency between the KLF4 promoter and its enhancer. Means and standard deviation from biological triplicates (n = 3) are shown. (p = 0.0003; 0.0019; 0.0030; 0.0324) Error bars represent mean ± SD (d, f). Statistical significance assessed by one-way ANOVA with Bonferroni correction (d, f); ns not significant; *P < 0.05; **P < 0.01; ***P < 0.001. Source data are provided as a Source Data file.
Our omics data showed that the knockdown of TLK2 reduced the binding of CTCF, RAD21, and DYNLL1, as well as reduced accessibility and H3K27ac modification at both the KLF4 promoter and enhancer (Fig. 5c and Supplementary Fig. 7). We further validated that the knockdown of TLK2 and DYNLL1 reduces KLF4 expression (Fig. 5d, e). Two available but non-specific TLK kinase inhibitors, inhibitors THD41,42 and Go698328,37 also reduced CTCF-cohesin hubs and KLF4 expression (Supplementary Fig. 7b–d). Moreover, we examined the interaction frequency between the KLF4 promoter and its enhancer using chromosome conformation capture assays (3C-qPCR). The interaction frequency was significantly reduced in cells with knockdown of TLK2 and DYNLL1 (Fig. 5f). These data suggest that the TLK2-DYNLL1 axis may contribute to enhancing KLF4 regulatory interactions and expression near the CTCF-cohesin hub.
Targeting TLK2 enhances cancer therapeutic responsiveness
We assessed the clinical potential of targeting TLK2 for tumor therapy. Notably, TLK2 inhibition led to a marked decrease in cell viability under docetaxel (DTX) chemotherapy and T cell-dependent peripheral blood mononuclear cell (PBMC) cytotoxicity (Fig. 6a–c). We then investigated whether TLK2 inhibition could enhance the efficacy of PD-L1 blockade immunotherapy in a breast cancer mouse model (Fig. 6d). Consistent with recent report43, the combination of TLK2 inhibition and anti-PD-L1 antibodies resulted in a significant enhancement of primary tumor inhibition (Fig. 6d and Supplementary Fig. 8). The combination also significantly reduced the lung metastasis (Fig. 6e). THD and Go6983 also effectively reduced the ratio of ALDH-positive cells and mammosphere formation (Supplementary Fig. 8b, c). Collectively, the data suggest that inhibiting TLK2 enhances the responsiveness of cancer cells to therapeutic interventions.
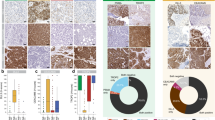
a Co-culture of mNeonGreen-labeled control and mRuby3-labeled TLK2 knockdown MDA-MB-231 cells treated with docetaxel (DTX), followed by flow cytometry analysis of proportional changes. Representative results from three biological replicates performed are shown. b Representative immunofluorescence of peripheral blood mononuclear cell (PBMC) killing assays in TLK2 knockdown and control cells, showing apoptosis through caspase-3 staining (Ac-DEVD-AFC). Scale bar, 50 μm. c Flow cytometry analysis of PBMC-mediated killing effect (left) and statistical summary of repeated killing assays (right). Means and standard deviation from biological triplicates (n = 3) are calculated. (p = 0.0004) d, e Evaluation of TLK2’s effect on immunotherapy in a mammary fat pad spontaneous lung-metastasis mouse model. BALB/c mice bearing TLK2-stable knockdown or control 4T1 cells were treated with anti-PD-L1 antibodies or isotype control. Primary tumors were resected on day 18, and mCherry/GFP ratio in the lungs was quantified on day 30. Representative images are shown. The center line of the box plots represents the median; box bounds represent the interquartile range (25th–75th percentile); whiskers extend to the minimum and maximum values. (d p = 0.0001; 0.0431; 0.0082; e p = < 0.0001) f Box plots showing the fraction of patients with TLK2 genetic alterations across TCGA PanCancer Atlas Studies (cBioPortal). g Correlation between TLK2 mRNA expression and specific copy-number alterations (cBioPortal data). Data are presented as mean ± SD; center indicates the mean. h, Kaplan-Meier plots evaluating disease-free survival (DFS) and overall survival (OS) based on TLK2 expression determined by IHC. Error bars represent mean ± SD (c, d, e, g). Statistical significance was assessed using one-way ANOVA with Bonferroni correction (c, d); two-tailed unpaired Student’s t test (e); ns not significant; *P < 0.05; **P < 0.01; ***P < 0.001. Source data are provided as a Source Data file.
To evaluate the clinical relevance of TLK2, we examined the available datasets from The Cancer Genome Atlas (TCGA) for copy number alterations (CNAs), relative expression, and prognosis of TLK2. By analyzing data from 10,945 patient samples of TCGA PanCancer Atlas Studies, consistent with previous findings37,38, we also observed that TLK2 frequently exhibited copy number gain and upregulated expression in breast cancer (Fig. 6f, g). Additionally, the high expression of TLK2 was associated with reduced relapse-free survival (RFS) in a variety of cancer types from the Kaplan-Meier plotter database (Supplementary Fig. 9).
Furthermore, we assessed TLK2 protein expression in 262 primary human breast cancer patient samples from Sun Yat-Sen University Cancer Center, utilizing immunohistochemistry (IHC). Clinical parameters and features of the patients and their samples were summarized in Supplementary Data 12. Aligning with prior studies that identify TLK2 as a predictor of poor prognosis27,37,38, our analysis showed that high TLK2 expression significantly predicted worse clinical disease-free survival (DFS) and overall survival (OS) in the patient cohort (Fig. 6h).
Discussion
A compelling hypothesis is that highly plastic cancer cells exhibit more flexible chromatin, allowing the transition in the gene expression or cellular identity in response to external stimuli and thereby enhancing tumor survival against cytotoxic therapies. Consequently, targeting chromatin dynamic may effectively hinder cancer cell plasticity and overcome therapeutic resistance. In this context, we reveal TLK2 functions as a regulator of chromatin loop formation in cancer stemness transition, offering a potential therapeutic approach, chromatin loop-targeting therapy, to target cancer stemness plasticity (Fig. 7).
Upper left, Schematic summarizing the identification of TLK2 as a regulator of CTCF-cohesin hub formation through CTCF-cohesin hub reporter and CRISPR screening. Upper right, Schematic illustrating the role of the TLK2-DYNLL1 axis in regulating CTCF-cohesin hub formation in proximity to the KLF4 stemness gene. Lower, Schematic depicting the suppression of CTCF-cohesin hub formation by targeting TLK2, inhibiting cancer stemness plasticity.
Recent advancements in high-throughput chromatin profiling technologies, such as Hi-C44, ChIA-PET45, and related methodologies, have dramatically broadened our knowledge on chromatin organization. However, experimental limitations have impeded the interrogation of the chromatin organization regulators. One of the major challenges in chromatin research is the development of structure-specific reporters that are compatible with high-throughput screening assays. Previously reported strategies such as Perturb-ATAC (coupling CRISPR screen with single-cell ATAC-seq)46 and HiDRO (an automated imaging pipeline Oligopaints by quantitative measurement of chromatin interactions in single cells)47, have significantly advanced the field but are primarily suited for analyzing chromatin accessibility or specific chromatin interaction across dozens to hundreds of genes. Here, we utilize the CTCF-cohesin BiFC assay to quantify specific chromatin loops in live cells, providing a targeted and complementary approach to existing screening methodologies. While our BiFC system excels in monitoring dynamic CTCF-cohesin hubs, it does not interrogate other chromatin loop types (e.g., mediator-cohesin loops) or higher-order structures like topologically associating domains. The development of chromatin structure reporters that reflect different levels of chromatin organization, especially those compatible with CRISPR-based high-throughput screening, will greatly advance our understanding of chromatin structure regulation.
Interestingly, our integrated chromatin profiling and mechanistic studies reveal that TLK2 regulates distinct modes of CTCF-cohesin hub dynamics during CSC transition. Loss-type hubs are primarily characterized by cohesin (RAD21) disengagement, whereas gain-type hubs are defined by increased CTCF binding. This suggests a dual-mechanism model of chromatin loop remodeling, in which loop loss and gain may be governed by distinct molecular mechanisms. Notably, we found that TLK2 promotes gain-type hub formation by phosphorylating DYNLL1, which enhances its interaction with CTCF and facilitates its binding at new genomic loci. This defines a TLK2-DYNLL1 axis that enhances the formation CTCF-cohesin hubs, contributing to chromatin architecture reprogramming during stemness acquisition. Together, these findings explain the CTCF-dominant pattern at gain-type hubs observed during the non-CSC to CSC transition and position TLK2 as a key signaling integrator linking kinase activity to nuclear loop organization.
TLK2 is particularly notable in cancer biology due to its frequent overexpression in human cancers and reduced DFS27,28,37,38,48. TLK2 has emerged as a crucial chromatin-assembly kinase, with its activity primarily mediated through its substrates histone chaperones ASF1A and ASF1B22,23,24,25,26,27,28,29,30. TLK2-ASF1A/B axis is essential for proper histone deposition, key processes in cell proliferation. Consistently, we also observed that the inhibition of TLK2 disrupts the cell cycle and suppresses cell proliferation. Thus, targeting TLK2 in cancer may produce a “one target, two hits” effect through distinct pathways: the TLK2-ASF1 axis in histone deposition, leading to suppression of cancer cell proliferation, and another TLK2-DYNLL1 pathway in regulating CTCF-cohesin hub formation, restraining cancer stemness transition. Additionally, other roles of TLK2 have been revealed, including regulation of DNA damage response27,29,38,49,50, innate immune signaling48, and immune infiltration43. Further exploration is needed to understand the comprehensive benefits of targeting TLK2 in anti-cancer therapy.
Notably, current available TLK kinase inhibitors, such as Go698328,37, and phenothiazines or phenothiazine analogs41,42,51,52, including THD, lack sufficient effectiveness or specificity for TLK2, as demonstrated by existing experimental evidence. Nonetheless, our observations indicate that both compounds can impair CTCF-cohesin hub formation and suppress CSC-like phenotypes. However, given their limited selectivity and potential off-target effects, these agents were not used to draw definitive mechanistic conclusions in this study. The development of selective and validated TLK2 inhibitors remains a priority for future mechanistic and translational research.
While our study provides evidence of the role of TLK2 in modulating cancer cell plasticity and stemness, we acknowledge that our reliance on breast cancer cell lines in 2D and 3D culture conditions has limitations. The use of 3D culture conditions, such as mammosphere assays, is a well-established method for enriching CSCs; however, it may not fully recapitulate the complexity of the tumor microenvironment in vivo. Additionally, while we have employed several stemness markers and functional assays, such as ALDH activity and LDA, to assess cancer stemness, these markers and assays have their own limitations. For example, ALDH activity is not specific to CSCs and can be influenced by metabolic changes in cancer cells. Similarly, LDAs provide a quantitative measure of stemness but may not capture the heterogeneity of CSC populations. Furthermore, our study focuses on the regulation of chromatin dynamics by TLK2, which is an important aspect of cancer cell plasticity. However, cancer cell plasticity is a multifaceted phenomenon influenced by various factors, including epigenetic modifications, signaling pathways, and interactions with the tumor microenvironment. Therefore, our findings represent one piece of the puzzle, and further studies using complementary models, such as patient-derived xenografts or genetically engineered mouse models, are needed to fully understand the role of TLK2 in cancer cell plasticity and stemness.
Collectively, our study holds implications for comprehending how TLK2 potentially impacts cancer cell plasticity through regulating CTCF-cohesin hub formation. This underscores the prospect of targeting chromatin looping as a promising avenue for eradicating plastic CSCs.
Methods
Ethics approval
All experimental protocols involving human and animal materials were reviewed and approved by the Institutional Research Ethics Committee of Sun Yat-sen University Cancer Center (Approval No. G2022-070-01).
Reagents
Chemicals, antibodies, cell lines and oligonucleotides are listed in Supplementary Data 13.
Cell lines
Human breast cancer cell lines (MDA-MB-231, MCF-7, T47D, HCC1806), human embryonic kidney HEK293T cells, and mouse mammary carcinoma 4T1 cells were acquired from the American Type Culture Collection (ATCC). Human osteosarcoma U2OS cell was given by Yuanzhong Wu from SYSUCC. Each cell line was cultured in standard medium at 37 °C in a humidified incubator with 5% CO2. Drosophila Schneider 2 cells were given from Meizhen Zheng’lab. Regular mycoplasma testing was performed on the cell lines.
siRNA transfection assay
The siRNA sequences are provided in Supplementary Data 13. Cells at 60% confluence were collected and adjusted to a density of 5 × 105 cells mL−1 in Pen/Strep-free growth medium. One microliter of siRNA in 50 µL of Opti-MEM (Life Technologies) was incubated with Lipofectamine 2000 (2 µL per µL of siRNA) in another 50 µL of Opti-MEM for 20 min at room temperature. Subsequently, one milliliter of cell suspension was combined with the siRNA/Lipofectamine 2000 mixture and plated in 6-well plates. After a 6-h incubation, the medium was changed to complete growth medium.
Construction of plasmids
For all knockdown or knockout assays, the sgRNA and shRNA sequences are presented in Supplementary Data 13. The annealed sgRNAs were cloned into Bbs1-digested pSpCas9 BB-2A-Puro (PX459) V2.0 (Addgene #62988) or lentiCRISPR v2 (Addgene #52961) vectors, while the annealed shRNAs were cloned into Age1- and EcoR1-digested pLKO-Tet-On (Addgene #21915) vectors.
For protein expression plasmids used in the CIP assay, FRET assay, APEX2 assay, in vitro kinase assay, and TLK2 overexpression assay, the cDNA of FKBP, Frb, mNeonGreen, mRuby3, and APEX2 were synthesized by the company based on sequences from the fluorescent protein database FPbase or Addgene, as detailed in Supplementary Data 13. The cDNAs of TLK2, CTCF, DYNLL1 and ASF1A were amplified from the cDNA of MDA-MB-231 according to the sequence in GenBank. The “NM” accession numbers are shown in Supplementary Data 13. These genes were cloned into pLVX vectors for eukaryotic expression or pET28 vectors for prokaryotic expression as needed.
Lentiviral packaging assay
Approximately 4 × 106 HEK293T cells were seeded in 10 cm dishes and provided with 10 mL of standard DMEM medium 1 day prior to transfection. The lentiviral constructs, packaging plasmid (psPAX2), and envelope plasmid (pMD2.G) were co-transfected using FuGene HD (Promega) at a ratio of 4:3:1. Media was refreshed after 6 h. Viruses were collected 48 h post-transfection, snap-frozen, and stored at −80 °C.
Quantitative RT-PCR assay
RNA was extracted using the HiPure Total RNA Plus Mini Kit (Magen). cDNA was synthesized from RNA using the TransScript All-in-One First-Strand cDNA Synthesis SuperMix for qPCR (TransGen). Transcripts were amplified using ChamQ Universal SYBR qPCR Master Mix (Vazyme), and quantitative PCR was conducted on a LightCycler 480 (Roche) instrument. Relative expression levels were calculated through the 2−ΔΔCt method. Primers were designed using the Primer-BLAST software. The list of primers is available in Supplementary Data 13.
RNA sequencing assay and data analysis
RNA was extracted with the HiPure Total RNA Plus Mini Kit (Magen) and then sent to the Novogene company to build libraries and sequence with Illumina HiSeq2000 (150 bp, paired-end). RNA integrity numbers were determined using Agilent 2100, and all samples used for RNA-seq library preparation had RIN numbers greater than 9.5. RNA-seq data was analyzed using the RNACocktail framework53. DEGs were subjected to volcano plots using the “ggplot2” visualization package in R. The complete DEG list is provided in Supplementary Data 5 and 10. The coverage track (bigWig) files of RNA-seq data were generated from BAM files and normalized by bins per million mapped reads (BPM, equivalent to TPM).
Cell fractionation and protein extraction
WCL (whole cell lysates) was extracted using RIPA buffer (50 mM Tris pH 7.4, 1 mM EDTA, 150 mM NaCl, 1% NP-40, 0.5% sodium deoxycholate, 0.1% SDS) for 30 min, followed by protein concentration test and established Western blotting protocols.
For co-IP assay, WCL or nuclear lysates were prepared by the following methods. To acquire WCL, cells were lysed in NETN buffer (20 mM Tris-HCl pH 8.0, 100 mM NaCl, 1 mM EDTA, 0.5% NP-40) at 4°C for 1 h, then centrifuged with 12,000 rpm for 10 min to get the supernatant. In case of nuclear lysates, cells were firstly incubated with cold hypotonic buffer (10 mM HEPES-NaOH, pH 7.9, 10 mM KCI, 0.5 mM β-mercaptoethanol, 1.5 mM MgCl2) on ice for 20 min. Then, 10% NP-40 was added to the cell suspension to a final concentration of 0.2%. After incubation on ice for 2 min, the samples were centrifuged at 4 °C at 2400 × g for 15 min, and the supernatant (cytoplasmic lysates) was carefully discarded. The cell pellets (nuclei) were washed with cold hypotonic buffer and then lysed with nuclei lysis buffer (10 mM Tris-HCI, pH 7.6, 2 mM MgCl, 150 mM NaCl, 0.5% NP-40, 1 mM DTT, 1 mM PMSF, 0.5 mM EDTA) for 1 h, followed by a centrifugation step at 12,000 rpm for 10 min, the supernatant was collected as nuclear lysates for co-IP assay.
All of the lysis buffer were supplemented with Phosphatase Inhibitor Cocktail and protease inhibitor cocktail (TargetMol). All procedures were conducted at 4 °C.
Co-immunoprecipitation assay
Protein extracts were first incubated with primary antibody at 4 °C overnight. This was followed by an additional incubation with 20 µL of protein A/G-agarose beads for 2 h. The beads with bound proteins were washed three times using ice-cold PBS supplemented with a protease inhibitor cocktail (PI) and a 500-fold dilution of Phosphatase Inhibitor Cocktail 3 (Sigma). The beads were then denatured by heating with loading buffer at 100 °C. The products were subsequently analyzed by Western blotting.
Cell cycle analysis
Cell cycle distribution was assayed by PI (Propidium Iodide) staining. Briefly, cells were collected, washed with PBS and fixed with 70% ethanol at 4 °C overnight. Then, the cells were centrifuged and stained with PI staining buffer (50 μg mL−1 PI, 100 μg mL−1 RNase A, 0.2% Triton X-100 in PBS) at 37 °C for 30 min in dark. The cells were then washed and resuspended in PBS for flow cytometry detection. Data were analyzed with CytExpert and FlowJo software.
Cell proliferation assays
Cell proliferation was measured by using CCK-8 cell viability assay. Briefly, approximately 1 × 103 cells were distracted into each well of a 96-well plate, and cultured in an incubator at 37 °C with 5% CO2. At days 1–3, supernatants were removed, and the cells were incubated with 10 μL CCK-8 (Beyotime) reagent at 37 °C for 1 h. The 96-well plates were shaken for 10 s and the OD450 values were detected.
Colony formation assay
Five hundred cells were seeded into 6-well plates and cultured at 37 °C in a humidified incubator. After 7 days, the colonies were fixed and stained with 0.1% crystal violet.
Mammosphere formation assay
Mammosphere formation was conducted in ultralow attachment plates (Corning) using a medium supplemented with 2% B27, 20 ng mL−1 bFGF, and 20 ng mL−1 EGF. Cells were seeded at a density of approximately 2 cells μL−1 and cultured at 37 °C with 5% CO2. After 5 days, spheres with a diameter greater than 40 μm were counted at ×40 magnification (Olympus).
ALDH+ cell staining and sorting assays
The ALDEFLUOR kit (StemCell Technologies) was utilized to analyze and isolate cells into subpopulations with high or low ALDH enzymatic activity. Briefly, cells were trypsinized to dissociate them into single cells and then resuspended in ALDEFLUOR assay buffer containing the ALDH substrate (BAAA, 1 µM per 1 × 106 cells) and incubated for 45 min at 37 °C. As a control, cells in a buffer containing the substrate were incubated with 15 mM diethylaminobenzaldehyde, which inhibits ALDH activity. After washing with ALDEFLUOR assay buffer, cells were passed through a BD Falcon cell strainer cap tube for sorting. The percentage of ALDH-positive cells was determined using a BD FACSAria II flow cytometer.
BiFC assay
Cells were seeded on a 35-mm glass-bottom culture dish 16 h before transfection. Transfection was performed with Venus-N and Venus-C using Lipofectamine 2000 in Opti-MEM, following the manufacturer’s instructions. After a 6-h incubation, the medium was replaced with fresh medium, and cells were further incubated for 30 h. For nuclear staining, cells were washed three times with 1 × PBS, followed by incubation in 2 ml of Hoechst 33258 (Sigma-Aldrich) in 1 × PBS solution for 30 min, and then washed three times with 1 × PBS.
FRET fluorescence imaging assay
FRET experiments were carried out using a ZEISS LSM880 microscope with a ×63 oil immersion objective. Cells were transfected with FRB-mNeonGreen-CTCF and FKBP-mRuby3-RAD21/SMC3 using Lipofectamine 2000 (Life) following the manufacturer’s protocol. The FRET efficiency was calculated using the sensitized emission method. Fluorescence imaging of each sample was performed at room temperature after Rapamycin was added for 24 h. The FRET efficiency was acquired as the enhanced mRuby3/mCherry emission due to energy transfer according to recommendations from ZEISS.
ChIP-qPCR and Re-ChIP-qPCR
For Chromatin immunoprecipitation followed by qPCR (ChIP-qPCR) assay, cells were fixed, hypotonically lysed, and sonicated in a Covaris E220 sonicator in 0.1% SDS/TE. An aliquot was diluted in RIPA and incubated with 5 μL of each antibody at 4 °C overnight. The antibodies were retrieved with ChIP-grade ProteinA/G Magnetic Beads (Thermo Fisher), and bound chromatin was washed and eluted. After reverse cross-linking, the immunoprecipitated and input DNA was used to perform the qPCR analysis. For Re-ChIP-qPCR, the initially eluted material, which was immunoprecipitated by the indicated antibody, underwent additional immunoprecipitation using another antibody and IgG antibodies. The percentage of bound DNA was quantified relative to the original DNA input. Real-time PCR was conducted using the primers specified in Supplementary Data 13. All data were normalized based on the input.
ChIP-seq and CUT&Tag assays and data analysis
For Chromatin immunoprecipitation sequencing (ChIP-seq) assay, cells were fixed, hypotonically lysed, and sonicated in a Covaris E220 sonicator in 0.1% SDS/TE. An aliquot was diluted in RIPA and incubated with 5 μL of each antibody at 4 °C overnight. The antibodies were retrieved with ChIP-grade ProteinA/G Magnetic Beads (Thermo Fisher), and bound chromatin was washed and eluted. After reverse cross-linking, the amount of immunoprecipitated and input DNA was determined with Qubit (Thermo Fisher). Libraries were prepared with the NEBNext ChIP-Seq Library Prep Kit for Illumina. The libraries were sent to the Novogene company for sequencing with Illumina HiSeq2000 or NovaSeq platform (150 bp, paired-end).
Cleavage under targets and tagmentation (CUT&Tag) assay was performed with the following procedure54 with NovoNGS CUT&Tag 2.0 High-Sensitivity Kit (Novoprotein) following manufacturer’s protocol. Briefly, 5 × 104 MDA-MB-231 cells with 5 × 103 Drosophila Schneider 2 cells as a spike-in control were incubated with 10 μL of Concanavalin A beads. Subsequently, 1 μg of primary antibodies with 1 μL of Drosophila melanogaster Histone variant H2Av antibody (Active Motif cat#61686) were added and incubated overnight at 4 °C with gentle rotation. After washing, the secondary antibody was added and incubated for 1 h at room temperature, using either goat anti-mouse (Abcam) or anti-rabbit antibody (Abcam) at a 1:100 dilution in Dig-Wash buffer. The pA-Tn5 adapter complex was added and incubated with the samples for 1 h at room temperature. The fragmented DNA was reverse-crosslinked in Dig-300 buffer. DNA purification was performed using AMPure beads (Beckman Coulter), followed by amplification with 5 PCR cycles using NEBNext MasterMix. qPCR was conducted to determine the additional number of PCR cycles required. Libraries were purified using the MinElute PCR Purification Kit (QIAGEN) and assessed for nucleosome patterning on the QSep1 (Bioptic). Finally, the libraries were sent to Novogene for sequencing on the Illumina HiSeq2000 or NovaSeq platform (150 bp, paired-end).
Raw 150-nucleotide sequencing reads were filtered using trim_galore and were subjected to quality assessment using FastQC. Reads were then aligned to the reference human genome (GRCh37/hg19) or the Drosophila reference genome dm6 using HISAT2 with X, no-spliced-alignment, and no-temp-splicesite parameters according to the manual55. PCR duplicates were removed using Picard MarkDuplicates. Alignment files were then down-sampled to ensure each file contains the same amount of unique Drosophila reads. Enriched peak regions were determined using MACS256 with both ChIP and control (input) samples, or CUT&Tag samples. Peak annotation was performed using the Homer software57. Overlapped unique peaks and genes were identified using BEDtools. The density heat map of CTCF-cohesin hubs was generated using Deeptools58. Differentially bound sites were called using DiffBind59. The bigWig files of ChIP-seq were created from BAM files and normalized by 1× depth (reads per genome coverage, RPGC). The bigWig files of CUT&Tag data were created from the down-sampled BAM files.
ATAC-seq and data analysis
Assay for transposase-accessible chromatin sequencing (ATAC-seq) assay was performed with Hyperactive ATAC-Seq Library Prep Kit for Illumina (Vazyme Biotech) following manufacturer’s protocol. Briefly, bulk cells were harvested, or G1, S, and G2/M phase cells were sorted using Hoechst 33342 staining. Dead cells were removed using the MACS® Dead Cell Removal Kit (Miltenyi Biotec). Then, 20,000 cells with 2 × 103 Drosophila Schneider 2 cells as a spike-in control were treated with 50 μL of lysis buffer (10 mM Tris-HCl, pH 7.4, 3 mM MgCl2, 10 mM NaCl, 0.1% NP-40) for 2 min to isolate nuclei. The nuclei were incubated in 50 μL of transposition buffer (Vazyme) to tag the accessible chromatin for 45 min at 37 °C with 1000 rpm shaking. Transposed DNA was purified using the MinElute PCR Purification Kit (QIAGEN) and amplified with 5 cycles using NEBNext MasterMix. qPCR was performed to determine the additional number of PCR cycles needed. Libraries were purified using the MinElute PCR Purification Kit (QIAGEN) and assessed for nucleosome patterning on the QSep1 (Bioptic). The libraries were sent to Novogene for sequencing with the Illumina HiSeq2000 or NovaSeq platform (150 bp, paired-end).
ATAC-seq data were analyzed following the guidelines of Harvard FAS Informatics (https://informatics.fas.harvard.edu/atac-seq-guidelines.html). Optimally, reads were aligned to the reference human genome (GRCh37/hg19) or the Drosophila reference genome dm6 using HISAT2 with X, no-spliced-alignment, and no-temp-splicesite parameters, following the manual55 to replace bowtie2. PCR duplicates were removed using Picard MarkDuplicates. Alignment files were then down-sampled to ensure each file contains the same amount of unique Drosophila reads. Enriched peak regions were determined using MACS2. Peak annotation was performed using HOMER. The differentially accessible regions were called using DiffBind. The list of peaks is provided in Supplementary Data 4. The bigWig files of ATAC-seq data were created from the down-sampled BAM files.
3C-qPCR
Quantitative Chromosome Conformation Capture Analysis (3C-qPCR) was performed with the following procedure60. Briefly, 5 × 106 cells were cross-linked with 1% formaldehyde for 10 min at room temperature, followed by quenching with 0.125 M glycine for 5 min. Cells were hypotonically lysed in 500 µL of lysis buffer (10 mM Tris-HCl, pH 8.0, 10 mM NaCl, 0.2% Igepal CA630, and a protease inhibitor cocktail) for 15 min at 4 °C. The nuclei were collected by centrifugation at 1000 × g for 5 min at 4 °C and then washed twice in lysis buffer. Nuclei were then resuspended in 50 µL of 0.5% SDS and incubated at 62 °C for 10 min to permeabilize them. The SDS was neutralized by adding 25 µL of 10% Triton X-100 and 145 µL of distilled water, followed by incubation at 37 °C for 15 min. For chromatin digestion, 25 µL of 10× NEB CutSmart buffer and 20 µL of MseI restriction enzyme were added, and the mixture was incubated overnight at 37 °C with rotation. Following digestion, the chromatin was added to a solution consisting of 663 µL of distilled water, 120 µL of NEB T4 DNA ligase buffer, 100 µL of 10% Triton X-100, 12 µL of 10 mg mL−1 BSA, and 0.5 µL of T4 DNA ligase. Ligation was carried out overnight at 16 °C with gentle rotation. To reverse cross-links, 50 µL of 20 mg mL−1 proteinase K and 120 µL of 10% SDS were added, followed by incubation at 55 °C for 30 min. An additional 130 µL of 5 M NaCl was added, and the mixture was incubated at 65 °C for 4 h to reverse the cross-link. DNA was then purified by PCR Purification Kit (Magen). qPCR amplification was performed using the primer sequences in Supplementary Data 13.
CRISPR pool construction and screening
A total of 1087 epigenetic genes, which directly or indirectly regulate DNA, RNA, or histone modification, were collected from the EpiFactors database61 and GO database62. For each gene, 10 sgRNAs with the highest predicted on-target knockout (KO) efficiency and lowest off-target efficiency were designed. The designs were based on the Human Activity-Optimized CRISPR Knockout Library (David Sabatini, Eric Lander’s labs), Human CRISPR Knockout Pooled Library (GeCKO v2, Feng Zhang’s lab), and Toronto KnockOut (TKO) CRISPR Library (Jason Moffat’s lab). In cases where we could not retrieve 10 sgRNAs from the three reference libraries, we designed them using the Graphical User Interface for DNA Editing Screens (GUIDES) software. Oligo sgRNA sequences with flanking adaptors were synthesized using the Syno® 3.0-oligo pool synthesis platform (Synbio Technologies). Subsequently, the oligo pool was amplified by PCR using primers with homologous arms to the lenti-CRISPR V2 vector. The PCR product was then inserted into the lenti-CRISPR V2 vector using Gibson assembly, following the manufacturer’s instructions. The plasmid library pool was sequenced, confirming sgRNA representation and pool complexity with ~6-fold changes in abundance between the 10th and 90th percentiles. The final epimone-wide library comprised 10,916 sgRNAs targeting 1104 human epigenetic factors and 200 non-targeting control sgRNAs.
The lentivirus library was packaged in HEK293T cells and filtered through a 0.22 μm filter. MDA-MB-231 cells were infected at an MOI of 0.3 and selected with puromycin for 7 days. Subsequently, the cells were divided into three groups. The first group was used to extract genomic DNA using the TaKaRa MiniBEST Universal Genomic DNA Extraction Kit as the Input group; the second group was transfected with the CTCF-YFPn and Cohesin-YFPc BiFC plasmids; the third group was transfected with empty YFPn and YFPc BiFC control vectors. Thirty-six hours later, the top 10 percent and bottom 10 percent of G1 phase cells were sorted and collected based on yellow fluorescence intensity. The sgRNA cassettes were then amplified from the genomic DNA and subsequently subjected to next-generation sequencing (NGS) using the Illumina HiSeq2000 or NovaSeq platform (150 bp, paired-end). The data were analyzed using the MAGeCK-MLE package63 and visualized with the “pheatmap” visualization package in R. The gene essentiality estimation beta score is provided in Supplementary Data 7.
APEX2 proximity ligation assay and mass spectrometry
Cells were incubated with 500 µM biotin-phenol for 30 min in the incubator. Equivoluminal 2 mM media-diluted H2O2 was added to the cells to initiate the labeling reaction for 30 s and was removed immediately. The reaction was further quenched by washing three times with ice-cold quenching buffer (TBS supplemented with 1 mM CaCl2, 10 mM sodium ascorbate, 1 mM sodium azide, and 1 mM Trolox). Cells were scraped in quenching buffer with protease inhibitors, lysed in RIPA buffer (50 mM Tris, 150 mM NaCl, 1% Triton X-100, 0.5% deoxycholate, 0.1% SDS, pH 7.4) supplemented with 10 mM sodium ascorbate, 1 mM sodium azide, 1 mM Trolox, 1 mM DTT, and protease inhibitors (TargetMol). Samples were briefly sonicated, spun down at 10,000 × g for 10 min. The supernatant was incubated with Streptavidin Magnetic Beads (NEB) at 4 °C overnight. The following day, the beads were washed four times with RIPA buffer, two times with 1 M KCl solution, two times with 50 mM NH4HCO3 solution, two times with 2 M Urea solution, three times with RIPA buffer, and finally two times with PBS buffer.
The beads were sent to to Chi Biomedicine for on-bead digestion and LC-MS/MS analysis. Peptides were desalted, lyophilized, and re-dissolved in solvent A (0.1% formic acid in water). For each sample, 5 μL of peptides was loaded onto a C18 reverse-phase column (Acclaim PepMap C18, 75 μm × 25 cm, 1.9 μm resin, Dr. Maisch GmbH) and separated over a 60-min nonlinear gradient from 4 to 95% buffer B (80% acetonitrile, 0.1% formic acid). The column was maintained at 55 °C, with a flow rate of 300 nL min−1 and an electrospray voltage of 2.0 kV. Mass spectrometry was performed using an Orbitrap Exploris 480 (Thermo Fisher Scientific, MA, USA) equipped with a FAIMS interface and coupled to an EASY-nanoLC 1200 system. The instrument operated in data-independent acquisition (DIA) mode with a hybrid data strategy. MS1 scan settings: resolution 120,000; m/z range 350–1200; AGC target 300%; maximum injection time 50 ms. MS2 scan settings: resolution 30,000; AGC target 200%; injection time 50 ms; normalized collision energy (NCE) 30. FAIMS compensation voltages were set to −45 V and −65 V. DIA acquisition used 80 variable isolation windows.
Raw DIA data were analyzed using Spectronaut v19 (Biognosys AG, Switzerland) with BGS Factory Settings. Database searching was performed against the UniProt Homo sapiens database (version 2024, 20,608 entries), using Trypsin as the digestion enzyme. Fixed modification: carbamidomethylation (C). Variable modifications: oxidation (M), acetylation (protein N-term), and biotin-phenol labeling (Y). False discovery rate (FDR) thresholds were set at 1% for precursor, peptide, and protein levels. The PTM site probability cutoff was set at 0.75. Decoy sequences were generated using a mutated sequence scrambling strategy. Data normalization was performed using the local normalization method. Quantification was based on MaxLFQ using peptides passing 1% FDR. Differential protein expression analysis was conducted using a fold change threshold of >2 and a P < 0.01 (Student’s t test). Proteins with >50% missing values in any group were excluded; those with <50% missing values were imputed using a minimal value strategy. APEX2 alone were used as the control for non-specific interactions. A total of 3 biological replicates per condition were analyzed. The interaction proteins are listed in Supplementary Data 10.
T cell-dependent nonspecific cytotoxicity assays
Cherry-H2B-labeled MDA-MB-231 cells with target shRNAs were pre-seeded in 96-well plates. Human PBMCs were isolated from healthy donors and activated with 100 ng mL−1 CD3 antibody, 100 ng mL−1 CD28 antibody, and 10 ng mL−1 IL2 for 5 days. They were then co-cultured with MDA-MB-231 cells in the presence or absence of the fluorescence caspase-3/7 substrate Ac-DEVD-AFC for 2 days. Finally, the co-cultured cancer cells were imaged using confocal microscopy or collected for flow cytometric analysis.
Tissue samples and immunohistochemical (IHC) staining assay
All clinical specimens were formalin-fixed, paraffin-embedded (FFPE) tumor tissues obtained from breast cancer patients treated at Sun Yat-Sen University Cancer Center, Guangzhou, China. The study was approved by the Institutional Research Ethics Committee of Sun Yat-Sen University Cancer Center (Approval No. G2022-070-01). Given the retrospective and anonymized nature of the study, the requirement for informed consent was waived.
Paraffin-embedded tissue blocks were sectioned for IHC. Antigen retrieval was performed by heating the samples in Tris-EDTA buffer pH 8.0 in a microwave oven for 10 min, followed by endogenous peroxidase blocking with hydrogen peroxide. The samples were blocked with 10% goat serum for 1 h at room temperature, then incubated with the primary antibody at 4 °C overnight. Antigen detection steps were performed using the Origin detection reagent (PV-6000/ZLI-9017, Origine China).
For TLK2 scoring, the immunoreactive score (IRS) was used. IRS was calculated as the product of the positive cell proportion score (0–4) and the staining intensity score (0–3), resulting in a range of 0–12. Slides were graded by two independent researchers. An IRS score of 0–6 was defined as low expression, while 7–12 was defined as high expression. Slides were graded by two independent researchers.
In situ proximity ligation assay (PLA)
The Duolink® In Situ Red kit (DUO92101, Sigma) was used. Briefly, the tissue slices with antigen retrieval were prepared following the IHC protocol described above and blocked with Duolink® blocking solution for 1 h at room temperature. Primary antibodies were diluted (1:100 each) in Duolink® antibody dilution buffer and incubated overnight. For negative controls, isotype-matched IgG antibodies were used in place of the primary antibodies, following the negative control strategy described by Lin et al.64. After washing, PLA probes were added and incubated for 1 h at 37 °C. Subsequently, the slices were washed, ligated, and finally mounted with Prolong Gold mounting medium containing DAPI. Visualization was performed using a confocal laser-scanning microscope (Zeiss LSM 880).
In vitro kinase assay
Homemade ASF1A fusion proteins were purified from BL21 E. coli that were transfected with pGEX6p1-GST-ASF1A-HIS using HIS or GST beads. Commercial DYNLL1 was purchased from Novoprotein Scientific Inc (Cat# C218). Active TLK2 was purchased from Sino Biological Inc (Cat# 11551-HNCB, aa 397-772). The kinase reactions were performed in kinase buffer (10×) (Cat# 9802, CST) with 200 μM γ−32P ATP (PerkinElmer), 1 µg of soluble DYNLL1 or ASF1A, and 100 ng of TLK2. The reaction was incubated at 30 °C for 30 min and then resolved by SDS-PAGE followed by autoradiography.
Animal experiments
The animal experiments were conducted with approval from the ethical committee of Sun Yat-Sen University Cancer Center (Approval No. G2022-070-01). Mice were housed in a specific pathogen-free (SPF) facility under standard conditions with a 12-h light/dark cycle, ambient temperature maintained at 20–26 °C, and relative humidity of 40–70%. The maximum allowable tumor volume was 2,000 mm³, and this limit was not exceeded during the study.
Tlk2 knockdown 4T1 cells were bilateral injected into the right, shCtrl 4T1 cells were injected into the left mammary fat pads of female BALB/C mice (4–6 weeks old, Hunan SJA Laboratory Animal Company, China) using 1 × 105 cells (5 mice in each group). The anti-mouse PD-L1 antibody (BP0101; BioXcell) and the anti-rat-IgG2b isotype control (BP0090; BioXcell) were intraperitoneally injected on the indicated day at a dose of 100 μg per mouse. Tumor dimensions (length and width) were measured using calipers, and volumes were calculated using the formula V = (length × width2)/2. Primary tumors were resected on day 18, and mCherry/GFP ratio in the lungs was quantified on day 30. mCherry and GFP were detected using antibodies against mCherry (GTX128508, GeneTex) and GFP (GTX628528, GeneTex) followed by secondary antibodies conjugated with Alexa Fluor® 488 or 546 (1:500, Life Technologies). Imaging was performed using a confocal laser-scanning microscope (Zeiss LSM 880). The mCherry or GFP-positive area was quantified using imageJ. The fold change in mCherry/GFP ratio was calculated.
For Limiting Dilution Assay (LDA) in mice, female NOD-SCID mice (4–6 weeks old, Hunan SJA Laboratory Animal Company, China) were pretreated with doxycycline (DOX, 2 mg mL−1 with 2% sucrose) or vehicle for 2 days prior to tumor injection. DOX-inducible TLK2 knockdown MDA-MB-231 cells or control cells were subcutaneously injected into the mice using 1 × 103 cells (8 mice in each group), 1 × 104 cells (8 mice in each group), 1 × 105 cells (8 mice in each group), and 1 × 106 cells (7 mice in each group) respectively. DOX and vehicle control in drinking water were changed daily, and tumor formation was monitored. After 28 days, mice were sacrificed and dissected to confirm tumor growth. The cancer stem cell frequency was calculated using the ELDA online software (http://bioinf.wehi.edu.au/software/elda/index.html). Resected tumor xenografts were immediately dissociated into single cells and subjected to mammosphere formation assays to evaluate mammosphere-forming capacity.
Gene function prediction
The Spearman’s correlations of co-expressed genes were retrieved from the study of Breast Invasive Carcinoma (TCGA, PanCancer Atlas) in cBioPortal. The top 100 most correlated genes, which are located in different chromosome bands (cytoband), were submitted to Database for Annotation, Visualization and Integrated Discovery65 to perform GO analysis. Chromatin related term were selected to present. The gene list and top enriched GO term are listed in Supplementary Data 8 and 9.
Prediction of DYNLL1 binding motifs
To identify potential DYNLL1 (LC8) binding motifs within the CTCF protein, we employed a computational approach leveraging established DYNLL1 binding motifs, particularly the TQT-like sequence. Due to the absence of a dedicated tool for predicting DYNLL1 binding sites, we utilized the ChatGPT o3-mini model to analyze pertinent studies and predict potential motifs. Key literature sources, including PLoS One (2011, Apr 18;6(4):e18818)35 and Life Science Alliance (2019, Jul 2;2(4):e201900366)36, were consulted, with particular attention to supplementary data listing known and potential DYNLL1 binding motifs. Regarding the application of ChatGPT for protein interaction predictions, recent studies have explored the use of large language models in bioinformatics, particularly in predicting protein-protein interactions. Using this approach, we predicted DYNLL1 binding motifs in the CTCF sequence by identifying TQT-like sequences and considering structural features consistent with DYNLL1’s binding preferences. The predicted motifs are listed in Supplementary Data 11.
Statistics and reproducibility
All assays were independently repeated at least three times, and results are presented as means ± S.D. unless stated otherwise. Sample sizes for in vitro and in vivo experiments were determined based on prior experience, standard practices in the field, and to ensure sufficient statistical power to detect biologically meaningful differences. For in vitro experiments, at least three independent biological replicates were performed unless otherwise stated. For in vivo mouse studies, group sizes of 4–6 mice per condition were used, which is commonly accepted in cancer biology to detect significant treatment effects. No statistical method was used to pre-determine sample size, but sample sizes were consistent with similar published studies and sufficient to yield reproducible results with statistical significance. For high-throughput sequencing-based analyses, two biological replicates per condition were used, and reproducibility was confirmed by correlation analysis and principal component analysis. To reduce bias, the sample groups were processed blindly during the experiment and the result evaluation stage. For in vitro experiments, cells were randomly assigned to experimental conditions to minimize technical bias. For in vivo studies, animals were randomly assigned to control and treatment groups, and all animals were age- and sex-matched to control for biological variability. A two-tailed unpaired Student’s t test was used for analysis between two groups, while one-way or two-way Analysis of Variance (ANOVA) with a post hoc test was used for analyses among three or more groups, as indicated. Nonparametric Spearman’s rank correlation was employed for linear correlation analysis. Statistical comparisons were made using the Chi-square test.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw sequence data generated in this study have been deposited in the Genome Sequence Archive66 in National Genomics Data Center67, China National Center for Bioinformation/Beijing Institute of Genomics, Chinese Academy of Sciences, under accession numbers CRA002434 and HRA001605. The mass spectrometry data generated in this study have been deposited in the ProteomeXchange Consortium via the iProX partner repository68,69 under accession code PXD062009 [https://www.iprox.cn/page/project.html?id=IPX0011405000]. The remaining data are available within the Figshare (https://doi.org/10.6084/m9.figshare.24278641), Supplementary Information or Source Data file. Source data are provided with this paper.
References
Gupta, P. B., Pastushenko, I., Skibinski, A., Blanpain, C. & Kuperwasser, C. Phenotypic plasticity: driver of cancer initiation, progression, and therapy resistance. Cell Stem Cell 24, 65–78 (2019).
Saygin, C., Matei, D., Majeti, R., Reizes, O. & Lathia, J. D. Targeting cancer stemness in the clinic: from hype to hope. Cell Stem Cell 24, 25–40 (2019).
Batlle, E. & Clevers, H. Cancer stem cells revisited. Nat. Med 23, 1124–1134 (2017).
Flavahan, W. A., Gaskell E., Bernstein B. E. Epigenetic plasticity and the hallmarks of cancer. Science 357, eaal2380 (2017).
Yadav, T., Quivy, J. P. & Almouzni, G. Chromatin plasticity: a versatile landscape that underlies cell fate and identity. Science 361, 1332 (2018).
Debruyne, D. N. et al. BORIS promotes chromatin regulatory interactions in treatment-resistant cancer cells. Nature 572, 676 (2019).
Chaffer, C. L. et al. Poised chromatin at the ZEB1 promoter enables breast cancer cell plasticity and enhances tumorigenicity. Cell 154, 61–74 (2013).
Johnstone, S. E. et al. Large-scale topological changes restrain malignant progression in colorectal cancer. Cell 182, 1474–1489 e1423 (2020).
Kim, T. H. et al. Analysis of the vertebrate insulator protein CTCF-binding sites in the human genome. Cell 128, 1231–1245 (2007).
Tang, Z. et al. CTCF-Mediated human 3D genome architecture reveals chromatin topology for transcription. Cell 163, 1611–1627 (2015).
Davidson, I. F. et al. DNA loop extrusion by human cohesin. Science 366, 1338–1345 (2019).
Bauer, B. W. et al. Cohesin mediates DNA loop extrusion by a “swing and clamp” mechanism. Cell 184, 5448–5464 e5422 (2021).
Busslinger, G. A. et al. Cohesin is positioned in mammalian genomes by transcription, CTCF and Wapl. Nature 544, 503 (2017).
Haarhuis, J. H. I. et al. The cohesin release factor WAPL restricts chromatin loop extension. Cell 169, 693–707 (2017).
Yang, N. et al. FOXM1 recruits nuclear Aurora kinase A to participate in a positive feedback loop essential for the self-renewal of breast cancer stem cells. Oncogene 36, 3428–3440 (2017).
Gentles, A. J. et al. The prognostic landscape of genes and infiltrating immune cells across human cancers. Nat. Med 21, 938–945 (2015).
van Dam, S., Vosa, U., van der Graaf, A., Franke, L. & de Magalhaes, J. P. Gene co-expression analysis for functional classification and gene-disease predictions. Brief. Bioinform 19, 575–592 (2018).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Pereira, B. et al. The somatic mutation profiles of 2,433 breast cancers refines their genomic and transcriptomic landscapes. Nat. Commun. 7, 11479 (2016).
Cancer Genome Atlas N Comprehensive molecular portraits of human breast tumours. Nature 490, 61–70 (2012).
Li, Q. et al. INK4 tumor suppressor proteins mediate resistance to CDK4/6 kinase inhibitors. Cancer Discov. 12, 356–371 (2022).
Sillje, H. H. W. & Nigg, E. A. Identification of human Asf1 chromatin assembly factors as substrates of Tousled-like kinases. Curr. Biol. 11, 1068–1073 (2001).
Klimovskaia, I. M. et al. Tousled-like kinases phosphorylate Asf1 to promote histone supply during DNA replication. Nat. Commun. 5, 3394 (2014).
Carrera, P. et al. Tousled-like kinase functions with the chromatin assembly pathway regulating nuclear divisions. Gene Dev. 17, 2578–2590 (2003).
Li, Z., Gourguechon, S. & Wang, C. C. Tousled-like kinase in a microbial eukaryote regulates spindle assembly and S-phase progression by interacting with Aurora kinase and chromatin assembly factors. J. Cell Sci. 120, 3883–3894 (2007).
Segura-Bayona, S. et al. Differential requirements for Tousled-like kinases 1 and 2 in mammalian development. Cell Death Differ. 24, 1872–1885 (2017).
Lee, S. B. et al. Tousled-like kinases stabilize replication forks and show synthetic lethality with checkpoint and PARP inhibitors. Sci. Adv. 4, eaat4985 (2018).
Mortuza, G. B. et al. Molecular basis of tousled-like kinase 2 activation. Nat. Commun. 9, 2535 (2018).
Segura-Bayona, S. & Stracker, T. H. The tousled-like kinases regulate genome and epigenome stability: implications in development and disease. Cell. Mol. Life Sci. 76, 3827–3841 (2019).
De Benedetti, A. Tousled kinase TLK1B counteracts the effect of Asf1 in inhibition of histone H3-H4 tetramer formation. BMC Res. Notes 2, 128 (2009).
Pavinato, L. et al. Functional analysis of TLK2 variants and their proximal interactomes implicates impaired kinase activity and chromatin maintenance defects in their pathogenesis. J. Med. Genet. 59, 170–179 (2022).
Becker, J. R. et al. The ASCIZ-DYNLL1 axis promotes 53BP1-dependent non-homologous end joining and PARP inhibitor sensitivity. Nat. Commun. 9, 5406 (2018).
He, Y. J. et al. DYNLL1 binds to MRE11 to limit DNA end resection in BRCA1-deficient cells. Nature 563, 522–526 (2018).
Xue, Y. et al. GPS: a comprehensive www server for phosphorylation sites prediction. Nucleic Acids Res. 33, W184–W187 (2005).
Rapali, P. et al. Directed evolution reveals the binding motif preference of the LC8/DYNLL hub protein and predicts large numbers of novel binders in the human proteome. PLoS ONE. 6, e18818 (2011).
Jespersen, N. et al. Systematic identification of recognition motifs for the hub protein LC8. Life Sci Alliance 2, e201900366 (2019).
Kim, J. A. et al. Comprehensive functional analysis of the tousled-like kinase 2 frequently amplified in aggressive luminal breast cancers. Nat. Commun. 7, 12991 (2016).
Kim, J. A. et al. Amplification of TLK2 induces genomic instability via impairing the G2-M checkpoint. Mol. Cancer Res. 14, 920–927 (2016).
Yu, F. et al. Kruppel-like factor 4 (KLF4) is required for maintenance of breast cancer stem cells and for cell migration and invasion. Oncogene 30, 2161–2172 (2011).
Reiff, S. B. et al. The 4D Nucleome Data Portal as a resource for searching and visualizing curated nucleomics data. Nat. Commun. 13, 2365 (2022).
Lee, S. B. et al. Design, synthesis and biological evaluation of bisindole derivatives as anticancer agents against Tousled-like kinases. Eur. J. Med Chem. 227, 113904 (2022).
Bhoir, S. & De Benedetti, A. Targeting prostate cancer, the ‘tousled way’. Int. J. Mol. Sci. 24, 11100 (2023).
Villamor-Payá, M. et al. Abstract A017: targeting Tlk2 impairs breast cancer growth and engages immune responses. Cancer Res. 84, A017–A017 (2024).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Fullwood, M. J. et al. An oestrogen-receptor-alpha-bound human chromatin interactome. Nature 462, 58–64 (2009).
Rubin, A. J. et al. Coupled single-cell CRISPR screening and epigenomic profiling reveals causal gene regulatory networks. Cell 176, 361 (2019).
Park, D. S. et al. High-throughput Oligopaint screen identifies druggable 3D genome regulators. Nature 620, 209–217 (2023).
Segura-Bayona, S. et al. Tousled-like kinases suppress innate immune signaling triggered by alternative lengthening of telomeres. Cell Rep. 32, 107983 (2020).
Bruinsma, W., van den Berg, J., Aprelia, M. & Medema, R. H. Tousled-like kinase 2 regulates recovery from a DNA damage-induced G2 arrest. EMBO Rep. 17, 659–670 (2016).
Lin, H. Y. et al. The TLK-ASF1 histone chaperone pathway plays a critical role in IL-1beta-mediated AML progression. Blood 143, 2749–2762 (2024).
Ronald, S. et al. Phenothiazine inhibitors of TLKs affect double-strand break repair and DNA damage response recovery and potentiate tumor killing with radiomimetic therapy. Genes Cancer 4, 39–53 (2013).
Singh, V. et al. Generation of phenothiazine with potent anti-TLK1 activity for prostate cancer therapy. iScience 23, 101474 (2020).
Sahraeian, S. M. E. et al. Gaining comprehensive biological insight into the transcriptome by performing a broad-spectrum RNA-seq analysis. Nat. Commun. 8, 59 (2017).
Wang, H. et al. Single-cell analyses reveal mechanisms of cancer stem cell maintenance and epithelial-mesenchymal transition in recurrent bladder cancer. Clin. Cancer Res. 27, 6265–6278 (2021).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907 (2019).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol 9, R137 (2008).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Ramirez, F., Dundar, F., Diehl, S., Gruning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Ross-Innes, C. S. et al. Differential oestrogen receptor binding is associated with clinical outcome in breast cancer. Nature 481, 389–393 (2012).
Hagege, H. et al. Quantitative analysis of chromosome conformation capture assays (3C-qPCR). Nat. Protoc. 2, 1722–1733 (2007).
Medvedeva, Y. A. et al. EpiFactors: a comprehensive database of human epigenetic factors and complexes. Database 2015, bav067 (2015).
Carbon, S. et al. The gene ontology resource: 20 years and still GOing strong. Nucleic Acids Res 47, D330–D338 (2019).
Li, W. et al. MAGeCK enables robust identification of essential genes from genome-scale CRISPR/Cas9 knockout screens. Genome Biol. 15, 554 (2014).
Lin, M. Z., Martin, J. L. and Baxter, R. C. Proximity ligation assay (PLA) to detect protein-protein interactions in breast cancer cells. Bio-protocol 5, e1479 (2015).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Chen, T. et al. The genome sequence archive family: toward explosive data growth and diverse data types. Genom. Proteom. Bioinform. 19, 578–583 (2021).
Members C-N, Partners Database resources of the national genomics data center, China national center for bioinformation in 2022. Nucleic Acids Res. 50, D27–D38 (2022).
Ma, J. et al. iProX: an integrated proteome resource. Nucleic Acids Res. 47, D1211–D1217 (2019).
Chen, T. et al. iProX in 2021: connecting proteomics data sharing with big data. Nucleic Acids Res. 50, D1522–D1527 (2022).
Acknowledgements
We extend our gratitude to Meizhen Zheng’s lab at Southern University of Science and Technology for the generous gift of Drosophila Schneider 2 cells and Yu Zhang’s lab at Peking University for the generous gift of dCas9 plasmids. We thank the bioinformatics platform at Sun Yat-sen University Cancer Center, Dr. Sheng Lin (Shenzhen Health Development Research Center), and Mr. Jianming Zeng (University of Macau) for their valuable support and guidance in using bioinformatics tools. We also appreciate Junjun Ding, Jun Chen, Zhonghui Tang and Song Gao at Sun Yat-Sen University, Cheng Li and Yuting Liu at Peking University, Guangshai Jia at Guangzhou Medical University, Xi Chen at Southern University of Science and Technology and Prof. Eric W-F Lam for their contributions to language editing and insightful comments. This research work was supported by the Science Fund for Creative Research Groups of the National Natural Science Foundation of China (82321003 to Q.L.), the National Natural Science Foundation of China (82473123 and 82173367 to Z.F.W., 82341020 to Q.L., 82403653 to F.L., 82302929 to N.N.C. and 82303009 to B.D.), the National Key R&D Program of China (2022YFA1104002 to Q.L.), the Natural Science Foundation of Guangdong (2022A1515010915 to Z.F.W.), Shenzhen Bay Laboratory Research Funds (SZBL2021080601001 to Q.L.), and the Cancer Innovative Research Program of Sun Yat-sen University Cancer Center (CIRP-SYSUCC-0019 to Q.L.).
Author information
Authors and Affiliations
Contributions
Z.F.W., F.L., and Q.L. conceived and designed the study. Z.F.W., F.L., N.N.C., J.J.W., and X.H.L. performed the majority of experiments and data analysis. M.X.F., M.Y., J.Z., B.D., L.L.W., X.W., M.L.L., D.S.Z., Z.Z.Z., B.W., Z.S.Y., and B.H. provided experimental materials, methodology, data analysis support, and conceptual input. Z.F.W. performed the bioinformatics analyses. Z.F.W., F.L., and Q.L. wrote the manuscript. Z.F.W., Q.L., and B.H. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Eneda Toska, Arrigo De Benedetti, and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, Z., Liu, F., Chen, N. et al. Chromatin looping-based CRISPR screen identifies TLK2 as chromatin loop formation regulator in cancer stemness plasticity. Nat Commun 16, 9288 (2025). https://doi.org/10.1038/s41467-025-64066-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-64066-x
This article is cited by
-
KMT2D loss drives adeno-to-squamous transition and sensitizes TKI-resistant lung cancer to AURKA inhibition
Cell Death & Differentiation (2026)