Abstract
WNT signaling is an essential pathway regulating tissue morphogenesis and regeneration. However, harnessing the pathway for regenerative medicine has been challenging due to the lack of approaches to identify and deliver specific WNT ligands to the target tissue. Herein, we reported that WNT and R-spondin (RSPO) proteins could be transported on engineered exosomes and activate the pathway synergistically. We showed that WNT3A and RSPO1 co-treatment could effectively regulate hepatic cell fate and uncovered functional crosstalk with the PPARα signaling pathway. Moreover, dual-ligand-carrying exosome (exoWNT3A/RSPO1) hyperactivated the WNT signaling and promoted efficient hepatic organoid growth compared to the small molecule inhibitor CHIR99021. Importantly, the exosome can be efficiently delivered for robust WNT signaling activation in the liver. Remarkably, exoWNT3A/RSPO1 could accelerate liver repair and regeneration under various conditions, including acute and chronic injuries and aging-associated phenotypes. Collectively, our work revealed the broad therapeutic effects of WNT signaling activation in the liver through the dual-ligand-carrying exosomes.
Similar content being viewed by others
Introduction
The liver possesses a remarkable regenerative capacity, fully regenerating even after surgical removal of up to 70% of its mass following partial hepatectomy1,2. The WNT signaling pathway plays a critical role in this process, as evidenced by its robust activation during liver regeneration and the diminished regenerative capacity observed when the canonical WNT/β-catenin pathway is genetically disrupted3,4. These observations have prompted researchers to develop therapeutic modalities to activate the WNT signaling pathway to achieve liver regeneration in end-stage liver disease or during liver transplantation5. On the other hand, WNT pathway-activating modalities for efficient liver repair under acute and chronic injury remain largely unexplored. Moreover, recent lineage-tracing studies have revealed a limited physiological role for WNT-responsive Axin2-labeled hepatocyte populations in the liver during injury repair and homeostasis, further undermining the potential of WNT pathway activation for treating diseased or injured livers6,7. Hence, it would be of great interest to systematically evaluate the capacity of WNT signaling activation as a therapeutic strategy for liver regeneration under multiple conditions.
Native WNT ligands are attractive for targeted pathway activation due to their ligand-receptor specificity. Among the 19 WNT ligands, each exhibits preferential binding to one or more of the 10 Frizzled receptors, enabling cell- and tissue-specific activities8. While small-molecule inhibitors of GSK3β could strongly induce canonical WNT/β-catenin signaling, their pleiotropic effects often lead to significant toxicity. Moreover, such modalities neglect the functional contribution of non-canonical WNT pathways9. However, functional WNT ligand contains a palmitate modification that poses challenges for protein production in scale and, more importantly, limits their working range as morphogens due to the hydrophobic property10.
Besides WNT ligands, the R-spondin (RSPO) proteins are potent co-activators of the WNT signaling pathway. Through binding to LGR receptors, RSPO proteins stabilize the WNT-Frizzled complex, amplifying and prolonging the WNT stimulus11,12,13. The non-equivalent role of WNT ligands and RSPO proteins in regulating the signaling pathway and stem cell activity in adult tissue has been well-demonstrated in the intestinal crypt and is likely conserved across other organs14,15. Considering the above, developing a delivery system capable of effectively packaging WNT signaling-activating ligands, enabling long-range delivery, and preserving their functional integrity would be critical for harnessing these molecules for therapeutic applications.
Exosomes are nano-sized extracellular vesicles (EVs) that can carry a variety of biomolecules and are vital in cell-cell transport16. These vesicles are increasingly recognized as excellent drug/therapeutic carriers due to the observed efficacy in delivering payloads to target cells in mice with low immune clearance and high biocompatibility17,18. Although some WNT ligands have been detected on exosomes in Drosophila and mammalian systems19, their physiological function when released on exosomes remains poorly understood20. Notably, WNT3A has been engineered onto exosomes to treat lung and cartilage injury, although the underlying mechanisms remain to be elucidated21,22. Moreover, the observed WNT signaling activation from these exosome-based approaches has been modest compared to the strong induction achieved by GSK3β inhibitors such as CHIR99021 and LiCl, which likely accounts for their limited therapeutic efficacy.
Herein, we developed a panel of engineered exosomes with active loading of individual 19 WNT and 4 RSPO ligands. These exosomes exhibit comparable activities to recombinant proteins, even in the presence of WNT antagonists. We also observed a synergistic activation of WNT signaling when combining WNT- and RSPO1-carrying exosomes. Using human liver progenitor cell cultures, we identified that WNT3-RSPO1 or WNT3A-RSPO1 combinations could regulate liver progenitor cell fate, partially mediated by the PPARα signaling pathway. Importantly, we demonstrated that dual WNT3A/RSPO1 ligands carried on a single exosome (exoWNT3A/RSPO1) efficiently induced WNT activity in an Axin2-mGFP mouse model, rescued mice from acute and chronic liver injuries, and reversed aging-related liver phenotypes. This platform of WNT signaling-activating exosomes opens up opportunities for evaluating WNT ligand activity across various biological systems and offers a promising strategy for tissue regeneration in injured, diseased, and aged tissues.
Results
Active loading of all 19 WNT ligands on engineered exosomes via WLS
To achieve active loading of hydrophobic WNT ligands on exosomes, a carrier protein needs to be engineered to interact with all 19 WNT ligands and function as an exosome scaffold protein. As the addition of protein tags influences the activity of WNT ligands23, conventional fusion protein approaches with exosome scaffolds such as LAMP2b and CD63 are not considered. Wntless (WLS) is an eight-pass transmembrane protein that interacts with lipidated WNT ligands and is essential for their delivery to the membrane for secretion24,25,26,27. Homozygous mutations in WLS result in deficient WNT secretion and are associated with human pleiotropic genetic disorders28. Hence, WLS could function as a promising carrier protein to load all 19 WNT ligands on exosomes efficiently.
To evaluate this hypothesis, we first generated HEK293 cells co-expressing WLS and WNT3A, one of the most widely studied WNT ligands (Supplementary Fig. 1a). To achieve large-scale exosome production with high purity, we established a workflow comprising serum-free suspension culture and an ultrafiltration-based exosome purification approach (EXODUS)29 (Supplementary Fig. 1b). The purified engineered exosomes exhibited bilayer-membrane structures with diameters <200 nm (Supplementary Fig. 1c, d), a particle-to-protein ratio of more than 1012 particle/mg (Supplementary Fig. 1e), and a vesicle purity exceeding 95% (Supplementary Fig. 1f). Intriguingly, WLS protein could be efficiently loaded on exosomes (exoWLS) indicating that WLS could serve as an exosome scaffold protein and significantly increase the WNT3A level on purified exosomes (exoWNT3AWLS) (Fig. 1a, b). In contrast, large EVs showed minimal WLS loading, with no significant difference in WNT3A loading. Immunogold labeling confirmed the increased WNT3A loading on exoWNT3AWLS (Fig. 1c) and highlighted that WNT3A molecules could be localized on the exosome surface, which was further validated with co-immunoprecipitation assay (Supplementary Fig. 1g, h).
a WLS and WNT3A levels in large extracellular vesicles (EVs) and exosomes from engineered cell lines. b The estimated number of WNT3A molecules in each exosome particle (n = 5) (see methods). c Representative TEM images of immunogold-labeled exosomes using anti-WNT3A antibody. Red arrows show WNT3A on the exosome surface. d TOPFlash reporter assay in HEK293FT treated with the same exosome particles (1 × 109). The relative normalized activity to untreated group is shown (n = 3). e TOPFlash reporter assay in HEK293FT treated with the same protein amount (100 ng/mL). The relative normalized activity to untreated group is shown (n = 3). f Schematic illustrating WNT signaling antagonists and pathway component interactions (Left, Created in BioRender. Chan, Y. (2025) https://BioRender.com/yxnywpj). TOPFlash reporter assay with similar WNT3A protein levels (100 ng/mL) using unmodified HEK293FT (No) or HEK293FT overexpressing DKK1, WIF1, SFRP1, FRZB, or NOTUM. The relative normalized activity to untreated samples is shown (n = 3) (Right). g Protein expression of 15 WNT ligands on engineered exosomes. h R-spondin1 (RSPO1) protein level in cell lysate, conditioned media, and large EVs and exosomes from His-tagged RSPO1 expressing cells. i TEM images of immunogold-labeled exosomes using an anti-RSPO1 antibody. Red arrows show RSPO1 on the exosome surface. j TOPFlash reporter assay in HEK293FT cells treated with the same exosome particle number (1 × 109). The relative activity normalized to untreated samples is shown (n = 3). k Levels of unphosphorylated (Active) β-Catenin (Ser45) and total β-Catenin protein levels in HEK293FT cells treated with exoWNT3AWLS, exoRSPO1 or exoWNT3AWLS+ exoRSPO1 for 24 hrs. l AXIN2 and CCND1 expression in HEK293FT cells treated with exoWNT3AWLS and exoRSPO1 alone or in combination for 24 hrs. The relative fold-change normalized to exoWT control is shown (n = 3). All statistical data are presented as mean ± SD. Two-tailed unpaired Student’s t tests are used to compare the two groups. The p values are calculated and shown as: ns p ≥ 0.05, *p < 0.05, **p < 0.01, ***p < 0.001. n indicates different independent sample/cell experiments.
Next, we evaluated the functional properties of the exoWNT3AWLS using the T cell factor (TCF)-Optimized Promoter-luciferase (TOPFlash) reporter assay30. Corresponding with the increased WNT3A loading, exoWNT3AWLS treatment resulted in higher WNT signaling-inducing activity compared to exoWNT3A (no WLS engineered) or exoWLS at similar particle number (Fig. 1d). Furthermore, at a similar amount of WNT3A protein, exoWNT3AWLS exhibited a comparable level of WNT signaling-inducing activity compared to human recombinant WNT3A protein (hrWNT3A) (Fig. 1e). Besides the conserved WNT-activating capacity, we also hypothesized that the exosome-bound WNT3A, either enveloped or directly interacting with WLS on the surface, could be shielded from antagonists (Fig. 1f). To validate this, WNT antagonists (DKK1, WIF1, SFRP1, FZRB, and NOTUM) expressing cells (Supplementary Fig. 1i) were generated and used for TOPFlash reporter assays with hrWNT3A and exoWNT3AWLS at similar WNT3A protein level. Remarkably, while WNT antagonists inhibited the reporter activity of hrWNT3A, exosome-bound WNT3A maintained comparable levels of reporter activity in the presence of WIF1, SFRPs, FZRB, and NOTUM, but not DKK1 which targets WNT coreceptors LRP5/6 (Fig. 1f). These results highlighted that our WNT-carrying exosomes were capable of maintaining and protecting the functional activity of WNT ligands, even in the presence of WNT antagonists.
Next, we explored whether a similar strategy could be employed to load other WNT family ligands. As there is limited direct evidence to support the interaction of WLS with all 19 human WNT ligands for the active loading of these ligands onto exosomes, we generated fluorescent protein-tagged WNT3A (WNT3A-mCherry) and WLS (WLS-ZsGreen) to investigate potential protein interactions. Intriguingly, we observed that the WLS-ZsGreen fusion protein was highly localized in vesicles. Notably, a drastic shift in WNT3A-mCherry localization to these vesicles was observed in the presence of WLS-ZsGreen compared to WNT3A-mCherry alone (Supplementary Fig. 2a). Using this visualization strategy, we generated 18 other WNT-mCherry and WLS-ZsGreen to assay the potential interaction between WLS and WNT family ligands. Remarkably, all WNT ligands showed similar re-localization to the vesicles in the presence of WLS-ZsGreen (Supplementary Fig. 2b), indicating that WLS likely interacts with all WNT ligands. We then co-expressed WLS with each of the other 18 untagged WNT ligands and purified the engineered exosomes (exoWNTWLS). The size and morphology of these exosomes were largely intact and well-preserved (Supplementary Fig. 3a). Importantly, the successful loading of 15 WNT ligands onto exosomes was validated using antibodies recognizing the endogenous WNT proteins (Fig. 1g and Supplementary Fig. 3b).
Overall, we reported the unprecedented efficient active loading of all 19 WNT ligands onto engineered exosomes and further demonstrated that this approach offered a protective advantage against WNT antagonists.
Active RSPO proteins carried on exosomes exhibited synergistic activity in WNT signaling activation
Similar to previous reports10, we observed that the WNT3A ligand alone has limited canonical WNT signaling-inducing capacity (Supplementary Fig. 4a), which could be a major challenge for efficient pathway activation in vivo to achieve therapeutic effectiveness. Hence, we explore the potential to load the co-agonist RSPO onto exosomes. Surprisingly, significant levels of RSPO1 were detected on both large EVs and exosomes derived from His-tagged RSPO1-producing HEK293 cells (Fig. 1h), which was previously unreported. Notably, the RSPO1-loaded exosomes (exoRSPO1) could induce TOPFlash activity comparable to a similar amount of human recombinant RSPO1 (hrRSPO1) proteins (Supplementary Fig. 4b). Immunogold labeling confirmed the loading of RSPO1 on the exosomes and indicated that RSPO1 could be carried on the surface of the exosome (Fig. 1i). We further verified this localization with limited proteinase K and trypsin treatment assays (Supplementary Fig. 4c, d). In addition, we detected the loading of active RSPO1 on exosomes using Chinese hamster ovary (CHO) cells, a commonly used system to produce hrRSPO1 (Supplementary Fig. 4e, f). Moreover, we showed that other RSPO family members31 could also be loaded onto exosomes, albeit at lower levels than RSPO1 (Supplementary Fig. 4g–j).
Importantly, we demonstrated that the exoRSPO1 synergizes with exoWNT3AWLS to enhance WNT signaling activation using the TOPFlash reporter assay (Fig. 1j). This increase in canonical WNT/β-catenin signaling pathway was confirmed by Western blot with antibody against unphosphorylated (Active) β-Catenin (Ser45) and total levels of β-Catenin (Fig. 1k) and the induction of downstream genes AXIN2 and CCND1 (Fig. 1l) in HEK293FT cells. Collectively, we demonstrated that functional RSPO proteins could be released on exosomes, and exoRSPO1 exhibits synergistic WNT signaling-inducing activity when co-treated with exoWNT3AWLS.
exoWNTWLS and exoRSPO1 synergistically regulate hepatic progenitor cell fate in vitro
WNT signaling pathway is widely reported to play a role in liver tissue morphogenesis and is activated during tissue regeneration32,33. Using previously published methods34,35,36, we set up a multi-step induced pluripotent stem cell (iPSC) differentiation protocol where the induction of the liver progenitor cell (LPC), resembling previously described iPSC-derived LPC36, was dependent on canonical WNT signaling activation (Supplementary Fig. 5a, b). As the cells transit from the hepatic endoderm (HE) to the LPC stage, the cellular and nuclear size significantly enlarged and expressed key hepatoblast markers, including ALB, AFP, and HNF4α (Supplementary Fig. 5c, d). Using our panel of WNT-carrying exosomes, exoWNT3WLS and exoWNT3AWLS, co-treated with exoRSPO1, could strongly induce LGR5 and MYC expression in the HE cells (Supplementary Fig. 5e, f). Correspondingly, HE cells treated with exoWNT3WLS+exoRSPO1 or exoWNT3AWLS+exoRSPO1 exhibited morphological changes resembling LPC, in comparison to exoWNTWLS or exoRSPO1 alone (Fig. 2a). Profiling of WNT signaling downstream genes LGR5 and MYC, and hepatoblast marker genes AFP, ALB, and HNF4α further supported the synergistic activity of both exoWNTWLS and exoRSPO1 to induce the LPC state (Fig. 2b, c). Moreover, the LPC cultured with the exosomes could be maintained long-term (Supplementary Fig. 5g). Importantly, we could further validate this result with the hrWNT3A and hrRSPO1 treatment (Supplementary Fig. 5h). Transcriptome profiles of cells under different exosome treatments support significant change in cellular states only in the presence of both WNT and RSPO1 ligands (Fig. 2d and Supplementary Fig. 6a). The results showed strong regulation of WNT signaling-associated genes and enrichment of reported hepatoblast markers in exoWNT3WLS+exoRSPO1- or exoWNT3AWLS+exoRSPO1-treated HE cells (Fig. 2e).
a Bright-field images of iPSC-derived hepatic endoderm (HE) cells treated with exoWT, exoWNT3WLS, exoWNT3AWLS, exoRSPO1, exoWNT3WLS+exoRSPO1 or exoWNT3AWLS+exoRSPO1 for 6 days. b Expression level of WNT signaling downstream genes LGR5 and MYC and hepatoblast marker genes AFP, ALB, and HNF4α in HE cells treated with indicated exosomes for 6 days. The relative fold change was obtained via normalization with the sample treated with exoWT (n = 3). c Immunofluorescence staining for AFP, ALB, and HNF4α in HE cells treated with the indicated exosomes for 6 days. d Principal Component Analysis (PCA) of transcriptome profiles of HE cells treated with indicated exosomes for 6 days (n = 2). e Heatmap showing relative expression levels of genes associated with WNT signaling and hepatoblast across different samples (n = 2). f KEGG pathway analysis for genes upregulated in HE cells treated with exoWNT3AWLS+exoRSPO1 compared to exoWT. g Immunofluorescence staining for ALB in HE cells treated with exoWNT3A+exoRSPO1 and different PPAR inhibitors. Cells were treated with DMSO, PPARα inhibitor GW6471 (10 μM), or PPARβ/δ inhibitor GSK3787 (10 μM). All statistical data are presented as mean ± SD. Two-tailed unpaired Student’s t tests are used to compare the two groups. The p values are calculated and shown as: ns p ≥ 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. n indicates different biological replicates using independent cell experiments.
Interestingly, besides the WNT signaling pathway, the Peroxisome proliferator-activated receptor (PPAR) signaling pathway was highly active in the cells treated with exoWNT3AWLS+exoRSPO1 (Fig. 2f and Supplementary Fig. 6b). While the PPAR signaling pathway is widely reported to regulate metabolic activity in the human liver37, the role of PPAR signaling in regulating human hepatic progenitor cell fate has not been reported. We first confirmed that the LPC expressed PPARA and PPARD (Supplementary Fig. 6c). Next, we showed that PPAR agonist treatment alone could not replace exoWNT3A+exoRSPO1 to induce the LPC state (Supplementary Fig. 6d, e). On the other hand, using isoform-specific inhibitors38,39,40 to block specific PPAR signaling identified PPARα activity to be important for the induction of LPC state in the presence of exoWNT3AWLS+exoRSPO1 treatment (Fig. 2g and Supplementary Fig. 6f). We further confirmed the role of specific PPAR isoforms in LPC induction using shRNAs targeting specific PPAR isoforms (Supplementary Fig. 6g, h). These results supported the utility of our exosome panel to study WNT ligands regulating hepatic cell fate and identified the PPARα signaling pathway to potentially play a role in this process. Notably, the results also further supported our hypothesis that WNT3A/WNT3 ligand alone is likely insufficient to modulate hepatic cell fate, and synergistic activity with RSPO overcomes this hurdle.
A dual WNT3A and RSPO1 carrying exosome for effective induction of WNT signaling activity and regulation of hepatic cell fate
The synergistic effects of exoWNT3AWLS and exoRSPO1 prompt us to investigate the possibility of developing an exosome carrying both proteins (Fig. 3a). In addition, for in vivo applications, it would be challenging to control the simultaneous delivery of two proteins on separate exosomes. In this aspect, we successfully engineered cells (Supplementary Fig. 7a) to produce exosomes carrying both WNT3A and RSPO1 (exoWNT3A/RSPO1). WNT3A and RSPO1 were highly enriched on exoWNT3A/RSPO1, and protein levels were comparable to exoWNT3AWLS and exoRSPO1 (Fig. 3b and Supplementary Fig. 7b). Similar to the single ligand exosomes, the dual ligands were enriched on the surface of the exosome (Supplementary Fig. 7c, d). Using the STORM system and immunogold labeling, we confirmed the existence of both proteins on a single exosome (Fig. 3c, d). Importantly, the exoWNT3A/RSPO1 exhibited a close to two-fold increase in TOPFlash activity (Fig. 3e) and enhanced levels of both total and active forms of β-Catenin (Supplementary Fig. 7e) compared to exoWNT3AWLS+exoRSPO1, suggesting that having both proteins in the same exosome further facilitates their synergistic activity. In addition, we observed a dosage-dependent increase in TOPFlash reporter activity with the exoWNT3A/RSPO1 treatment (Supplementary Fig. 7f). We further determined with mass spectrometry that the proportion of WNT3A to RSPO1 on the exosomes was approximately 1:2, and this ratio was favorable for the induction of TOPFlash activity (Supplementary Fig. 7g, h).
a Schematic illustration of an exosome engineered to carry WNT and RSPO protein (Created in BioRender. Chan, Y. (2025) https://BioRender.com/yxnywpj). b WNT3A and RSPO1 levels on exoWT, exoWNT3AWLS, exoRSPO1, and exoWNT3A/RSPO1). c Images with the STORM system showing WNT3A and RSPO1 on exoWNT3A/RSPO1. Exosomes were stained with antibodies specific for WNT3A, RSPO1, and CD63. d Immunogold-labeling of WNT3A and RSPO1 on exosomes with anti-WNT3A antibody (10-nm gold particle, blue arrow) and anti-RSPO1 antibody (6-nm gold particle, green arrow). e TOPFlash reporter assay in HEK293FT cells treated with exoWT, exoWNT3AWLS+exoRSPO1, or exoWNT3A/RSPO1 at the same particle number (1 × 109) or CHIR99021 (3 μM). The relative fold change in activity normalized to exoWT control is shown (n = 3). f Bright-field image and immunofluorescence staining of AFP, ALB, and HNF4α in HE cells after exoWNT3A/RSPO1 treatment for 6 days. g Expression level of LGR5, MYC, ALB, AFP, and HNF4α in HE cells after different treatments for 6 days. The relative fold-change normalized to exoWT control is shown (n = 3). h Bright-field images of hepatic organoids cultured with exoWT, exoWNT3A/RSPO1 or CHIR99021 (3 μM) for 10 days. i Expression level of LGR5 and AXIN2 in hepatic organoids after different treatments for 10 days. The relative fold-change normalized to exoWT control is shown (n = 3). j PCA of transcriptome profiles of hepatic organoids under different treatments for 10 days (n = 2). k Venn diagram showing the intersection of differentially expressed genes (DEGs) between exoWNT3A/RSPO1 vs. exoWT and CHIR99021 vs. exoWT. l GSEA of common and specific DEGs in k against the KEGG gene dataset. m Bright-field images of hepatic organoids treated with exoWT, or exoWNT3A/RSPO1 in the presence of DMSO, SB202580 (10 μM), or PAN-RAS-IN-1 (10 μM). All statistical data are presented as mean ± SD. Two-tailed unpaired Student’s t tests are used to compare the two groups. The p values are calculated and shown as: ns p ≥ 0.05, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. n = 3 indicates 3 biological replicates using independent cell experiments.
In corroboration with the functional WNT signaling-inducing capacity (Fig. 3e), comparable to the exoWNT3AWLS+exoRSPO1 and 3 µM CHIR99201, a concentration used in most studies to maintain LPC state, the exoWNT3A/RSPO1 could also efficiently support the LPC fate and induce the PPAR signaling pathway (Fig. 3f, g and Supplementary Fig. 8a, b). To further evaluate the exosome’s utility in regulating hepatic cell fate, we extended our investigation to demonstrate that exoWNT3A/RSPO1 treatment effectively induced WNT signaling and maintained hepatic organoids (Supplementary Fig. 8c, d)41. Interestingly, we found that while WNT pathway genes such as LGR5 and AXIN2 were induced to comparable levels in the organoids treated with our exosomes or CHIR99201, the organoid growth was significantly more robust in the exoWNT3A/RSPO1-treated culture (Fig. 3h, i). Next, we profiled the transcriptome of the organoids cultured in the exoWT, exoWNT3A/RSPO1 or CHIR99201 to reveal molecular differences. We observed significant gene expression differences between the exoWNT3A/RSPO1 or CHIR99201 treated organoids (Fig. 3j, k), and detected a significant enrichment of RAS-MAPK signaling and cell cycle pathway genes in exoWNT3A/RSPO1 treated organoids (Fig. 3l and Supplementary Fig. 8e, f). Treatment with RAS-MAPK signaling inhibitors (SB203580 and PAN-RAS-IN-1) significantly inhibited the growth of the exoWNT3A/RSPO1-treated organoids (Fig. 3m). Overall, we generated a dual ligand exosome that demonstrated highly efficient WNT signaling activation capacity and hepatic cell fate modulation comparable to a widely used small molecule inhibitor.
Efficient delivery and activation of WNT-signaling pathway in mouse liver with exoWNT3A/RSPO1 treatment
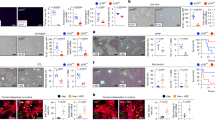
The remarkable WNT-signaling-inducing effect and hepatic cell fate regulation capacity of the exoWNT3A/RSPO1 prompt us to investigate the exosome’s functional capacity in vivo. First, to assess the biodistribution of the exosomes, we injected ~2 × 1010 particles exoWNT3A/RSPO1 intravenously into the mice. The exosomes are almost undetectable in the peripheral blood within an hour and rapidly accumulate in the liver (Supplementary Fig. 9a, b). 24 hrs after administration, the exoWNT3A/RSPO1 were primarily detected in the liver (Fig. 4a, b), and were taken up by both the liver parenchymal (such as ALB+ hepatocyte) and non-parenchymal cells (such as F4/80+ macrophage) (Fig. 4c).
a Fluorescence images showing the biodistribution of DiR-labeled exoWNT3A/RSPO1 after intravenous administration for 24 hrs. b Fluorescent images of tissue sections showing the distribution of PKH26-labeled exoWNT3A/RSPO1 in various organs. c Fluorescent images of tissue sections showing the cellular uptake of PKH26-labeled exoWNT3A/RSPO1 by different liver cell types, including hepatocytes (ALB+), resident Kupffer cells (F4/80+), cholangiocytes (CK19+), endothelial cells (CD31+), and stellate cells (α-SMA+). d Representative images showing Axin2-mGFP signals in liver central vein (CV) (Left) and schematic illustrating the treatment of Axin2-mGFP reporter mice with 2 × 1010 particles exosomes (Right, Created in BioRender. Chan, Y. (2025) https://BioRender.com/yxnywpj). e Fluorescent images of tissue sections showing the colocalization of Axin2-mGFP signals and zonation markers, including GS (zone 3), CYP2E1 (zone 2 and zone 3), and E-cadherin (zone 1). f Quantification of the percentage of area positive for mGFP and GS (n = 5). All statistical data are presented as mean ± SD. Two-tailed unpaired Student’s t tests are used to compare the two groups. The p values are calculated and shown as: ns p ≥ 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. n = 5 indicates 5 biological replicates using individual mice.
Next, to evaluate whether the exoWNT3A/RSPO1 could induce WNT signaling in vivo, exosomes were similarly introduced into the Axin2-mGFP reporter mice (Fig. 4d). Around the central vein (CV) in the liver, reportedly enriched for AXIN2 expressing hepatocytes42, we observed a significant increase in mGFP signal for the exoWNT3A/RSPO1-treated mice (Fig. 4e, f). In comparison to the liver, we observed modest levels of WNT-activity in other organs (Supplementary Fig. 9c). In line with a previous report15, we also observed that exoRSPO1 could induce WNT signaling activity in the liver, albeit at lower intensity compared to exoWNT3A/RSPO1. Notably, Zone 3 Glutamate synthetase-positive (GS+) hepatocytes were significantly increased in exoWNT3A/RSPO1-treated mice, highlighting potential GS+ hepatocyte proliferation (Fig. 4e). As WNT-signaling is reportedly activated in multiple organs, we further evaluated the safety of exoWNT3A/RSPO1 for long-term weekly intravenous administration (Supplementary Fig. 10). No significant histopathological changes were observed in multiple organs for mice treated for 1 month and even up to 3 months. However, mild alterations in the small intestine emerged when treatment was extended to 6 months, suggesting that prolonged exposure beyond a defined therapeutic window may impact extrahepatic organs. Taken together, our data thus far support the functional capacity of exoWNT3A/RSPO1 to induce WNT signaling in vivo and its potential as a therapeutic for liver injury treatments and regeneration.
exoWNT3A/RSPO1 accelerates liver repair and regeneration in mice after acute injury
We first evaluated the therapeutic potential of the exosome to rescue acute liver injury. exoWNT3A/RSPO1 or exoWT ( ~ 2 × 1010 particles per administration) were administered into acetaminophen (APAP)-induced acute liver injury model mice (Fig. 5a). The liver was harvested 24 hrs after the last exosome administration. Histological analysis revealed a significant reduction in necrotic tissue in the exoWNT3A/RSPO1-treated group, compared to the untreated and exoWT-treated group (Fig. 5b and Supplementary Fig. 11a, b). Correspondingly, the AST and ALT levels in the blood are also markedly lower in the exoWNT3A/RSPO1-treated group (Fig. 5c). Immunofluorescence staining for hepatocyte marker HNF4α and zonation markers CYP2E1 and GS supported the successful regeneration of the hepatocytes and restoration of liver zonation in mice treated with exoWNT3A/RSPO1 (Fig. 5d). Transcriptome profiling shows significant global expression changes, and notably, the WNT signaling pathway was induced in the liver tissues treated with exoWNT3A/RSPO1, validating the functional effect of the exosomes (Supplementary Fig. 11c, d). In corroboration with the staining results, we observed the enrichment of the liver function markers and a reduction in genes associated with inflammation and cell death in exoWNT3A/RSPO1-treated group (Fig. 5e). Similar to the observation in LPC, we also noticed an enrichment of the PPAR signaling pathway in mice treated with exoWNT3A/RSPO1 (Supplementary Fig. 11e, f). To determine if the therapeutic effect was dose-dependent, 5 × 109 or 2 × 1010 particles exoWNT3A/RSPO1 were administered intravenously into APAP injury mice. We observed that reduced exosome particle number results in lower Axin2-mGFP signals and a corresponding decrease in the rescue effect (Supplementary Fig. 11g–i).
a Schematic illustrating the experimental setup for APAP-induced acute liver injury and exosome administration in mice, livers, and serum were harvested at day 3 (Created in BioRender. Chan, Y. (2025) https://BioRender.com/yxnywpj). b Bright-field (Top) and H&E staining (bottom) of livers from healthy and APAP-injury mice and APAP-injury mice treated with exoWT or exoWNT3A/RSPO1. c Serum ALT and AST levels in mice from each treatment group (n = 5). d Immunofluorescent staining of GS, CYP2E1, and HNF4α in liver sections from each treatment group. Selected regions around the CV are shown. e Heatmap of relative expression of genes associated with hepatocyte cell fate, liver function, inflammation, and cell death in the mice liver from APAP-injury treated with exoWT or exoWNT3A/RSPO1 (n = 3). f Schematic illustrating the experimental setup for APAP-induced acute liver injury and exosome administration in mice. Livers and serum were harvested on days 1, 2, and 3 post-APAP injury (Created in BioRender. Chan, Y. (2025) https://BioRender.com/yxnywpj). g TUNEL staining of liver sections from each treatment group on the indicated day. h Quantification of TUNEL+ cells in i (n = 5). i Immunofluorescent staining of Ki67+ cells of the liver sections from each treatment group on the indicated day. j Quantification of the proportion of Ki67+ cells in g (n = 5). k Immunofluorescent staining of Ki67+ primary human hepatocytes treated with exoWT or exoWNT3A/RSPO1. l Quantification of the proportion of Ki67+ cells in k (n = 5). All statistical data are presented as mean ± SD. Two-tailed unpaired Student’s t tests are used to compare the two groups. The p values are calculated and shown as: ns p ≥ 0.05, ** p < 0.01, ***p < 0.001, ****p < 0.0001. n indicates different biological replicates using individual mice.
To gain insights into how the exosomes promote liver repair after acute injury, we repeated the experiment and harvested tissues at the early time point for investigation (Fig. 5f). We observed many TUNEL+ apoptotic cells at day 1 (D1) after APAP injury, and the numbers remained high at D2 and D3. In contrast, after 24 and 48 hrs of exosome administration, exoWNT3A/RSPO1 treatment significantly decreased the apoptotic cells (Fig. 5g, h and Supplementary Fig. 12a). In parallel, we observed a large increase in the number of Ki67+ cells only in the exoWNT3A/RSPO1-treated mice (Fig. 5i, j and Supplementary Fig. 12b). We also observed a significant induction of hepatocytes markers including Zone 1 and 2 hepatocyte specific markers, suggesting induction of hepatocyte regeneration during early exoWNT3A/RSPO1 treatment (Supplementary Fig. 12c, d). To further evaluate the direct effect of exoWNT3A/RSPO1 on hepatocytes, we cultured primary human hepatocytes (PHH) with exoWNT3A/RSPO1. Consistently, we observed around a 2-fold increase in the proportion of Ki67+ hepatocytes treated with exoWNT3A/RSPO1 at days 1 and 5 (Fig. 5k, l). Transcriptome profiling further supports the induction of the WNT signaling pathway and increased expression of cell cycle genes in the PHH treated with exoWNT3A/RSPO1 (Supplementary Fig. 12e, f). Besides the hepatocytes, we also observed significant changes to the non-parenchymal cells in the APAP-induced injury mice after exoWNT3A/RSPO1 treatment (Fig. 5e and Supplementary Fig. 13a, b). Specifically, the exoWNT3A/RSPO1 treatment significantly reversed the loss of liver-resident macrophages, reduced the level of monocyte infiltration, and decreased the amount of fibrosis, similar to previous acute liver repair studies43,44. Overall, these results indicated that exoWNT3A/RSPO1 could promote hepatocyte proliferation and reduce necroptosis to accelerate liver repair and regeneration after APAP-induced acute injury.
The effective therapeutic response prompts us further to validate the effects with a different acute liver injury model induced by CCl4, another commonly used reagent for inducing hepatoxicity in mice. The CCl4-treated mice with exoWNT3A/RSPO1 treatment had significantly reduced necrotic and inflammatory areas (Supplementary Fig. 14a-c). Correspondingly, levels of serum markers ALT and AST decreased, and a restoration of liver zonation and hepatocyte populations in the exoWNT3A/RSPO1-treated group was observed (Supplementary Fig. 14d, e). These results demonstrated that exoWNT3A/RSPO1 could effectively rescue acute liver injury induced by different pharmacological agents.
exoWNT3A/RSPO1 repairs and restores liver function and physiology after chronic liver injury and in aging mouse liver
To further explore the therapeutic potential of exoWNT3A/RSPO1 in treating long-term chronic liver injury, we subjected mice to CCl4 treatment twice a week for up to 6 weeks to induce the development of advanced liver fibrosis (Supplementary Fig. 15a). In parallel, we treated the mice with exoWNT3A/RSPO1 once a week. We found that exoWNT3A/RSPO1 treatment reduced the granulated surfaces and the amount of fibrous scarring, as evident from the Masson and Sirius red staining (Supplementary Fig. 15b, c). Correspondingly, the serum ALT and AST levels were significantly reduced in the exoWNT3A/RSPO1 group (Supplementary Fig. 15d). These results showed that the exoWNT3A/RSPO1 treatment has protective effects against CCl4-induced chronic injury.
Given the therapeutic effects observed thus far, we explored whether exoWNT3A/RSPO1 treatment could reverse liver injury in an intervention setup (Fig. 6a). Following three weeks of CCl4 treatment, we started the exosome treatment, where liver fibrosis was significantly detected (Supplementary Fig. 15e). The mice were harvested after three additional weeks of CCl4 and exosome co-treatment. Remarkably, we observed that the pathological features were also significantly mitigated in mice treated with exoWNT3A/RSPO1. Specifically, the liver surface was less granulated with much lower levels of fibrotic scar in the exoWNT3A/RSPO1-treated group (Fig. 6b, c). In corroboration, the serum liver injury markers decreased after exoWNT3A/RSPO1 treatment (Fig. 6d). We observed a significant increase in hepatocyte population around the CV, and profiling of zonation markers further supports the regeneration of liver tissue physiology in exoWNT3A/RSPO1-treated group (Fig. 6e). The global expression profile of the livers from mice treated with CCl4+exoWNT3A/RSPO1 in each setup closely clustered with the non-injured control group treated with Oil (Supplementary Fig. 15f, g). Specifically, we observed significant upregulation of liver function-associated genes and downregulation of genes associated with inflammatory response, cell death, and fibrosis in exoWNT3A/RSPO1-treated group (Fig. 6f). Interestingly, we also observed the upregulation of PPAR signaling pathway in these chronically injured mice livers treated with exoWNT3A/RSPO1 (Supplementary Fig. 15h, i), supporting a potential role of this pathway in the WNT signaling-mediated liver regeneration. Of note, our exosome treatment regimen resulted in a decrease in α-SMA population detected in the CCl4-treated mice (Supplementary Fig. 16a), similar to the treatment of APAP-induced liver injury model (Supplementary Fig. 13a, b). Using the Axin2-GFP mice, we observed that the exosome could activate WNT signaling in α-SMA-positive cells, but the overall population of α-SMA cells is comparable (Supplementary Fig. 16b), suggesting that our exoWNT3A/RSPO1 treatment regimen has a limited fibrogenesis effect45,46,47 in the liver injury models. In summary, the evidence thus far supported that exoWNT3A/RSPO1 could rescue long-term chronic liver injury. More importantly, the intervention study suggested that exoWNT3A/RSPO1 treatment holds the potential for repairing and regenerating the chronically injured liver.
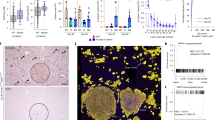
a Schematic illustrating the intervention treatment setup for CCl4-induced chronic liver injury (Created in BioRender. Chan, Y. (2025) https://BioRender.com/yxnywpj). b Image of whole liver (Top), Masson (Middle), and Sirius red (Bottom) staining for fibrotic areas in liver sections from mice injected with Oil only (Oil), CCl4 without exosomes (CCl4), CCl4 with exoWT, and CCl4 with exoWNT3A/RSPO1. c Quantification of Masson (Left) and Sirius red (Right) staining in fibrotic area of mice liver from each treatment group (n = 5). d Serum ALT and AST levels of mice liver from each treatment group (n = 5). e Immunofluorescent staining of GS, CYP2E1, and HNF4α in mice liver from each treatment group. Selected regions around the CV are shown. f Heatmap of relative expression of genes associated with hepatocyte cell fate, liver function, inflammation, and cell death in mice liver from CCl4-treated mice administrated with exoWT or exoWNT3A/RSPO1 for 3 weeks (n = 3). g Schematic illustrating the experimental setup for aging-mice (18-month-old) treatment (Created in BioRender. Chan, Y. (2025) https://BioRender.com/yxnywpj). h H&E (Top) and β-galactosidase (bottom) staining in liver tissue from each mice group. i Immunofluorescent staining of Ki67 (Top) and p21 (Bottom) cells in the liver from each mice group. j PCA of the transcriptome profile of liver tissues from young mice, old mice, and old mice treated with exoWT or exoWNT3A/RSPO1 (n = 3). k Heatmap of relative expression of genes associated with liver functions and aging in liver tissue from each mice group (n = 3). All statistical data are presented as mean ± SD. Two-tailed unpaired Student’s t tests are used to compare the two groups. The p values are calculated and shown as: ***p < 0.001, ****p < 0.0001. n indicates different biological replicates using individual mice.
Lastly, to further probe the capacity of exoWNT3A/RSPO1 to restore liver physiological states, 18-month-old mice were treated with the exosomes twice a week for 2 weeks (Fig. 6g). In the exoWT-control group, we observed a significant increase in the number of polyploid hepatocytes and the number of hepatocytes expressing senescence-associated β-Galactosidase compared to liver from younger mice (8-week-old). In contrast, these phenotypes were markedly reduced in the liver treated with exoWNT3A/RSPO1 (Fig. 6h and Supplementary Fig. 17a, b). We also noted a marked increase in Ki67+ hepatocytes in exoWNT3A/RSPO1-treated aged liver (Fig. 6i and Supplementary Fig. 17c). A reduction in the senescence cell population with p21 staining and reduction in overall p16 and p21 gene expression were also detected in exoWNT3A/RSPO1-treated mice (Fig. 6i and Supplementary Fig. 17c). Besides, we further observed an increase in marker expression for key parenchymal and non-parenchymal cell types in the liver treated with exoWNT3A/RSPO1, suggesting a restoration of the liver physiology (Supplementary Fig. 17c). Remarkably, PCA analysis and hierarchical clustering of transcriptome profiles of the treatment groups showed that the global expression profile of the aged liver treated with exoWNT3A/RSPO1 more closely resembled the young liver tissue (Fig. 6j and Supplementary Fig. 17d). Significant upregulation of liver function-associated genes and a decrease in aging and senescence-associated markers were observed in exoWNT3A/RSPO1-treated and young liver tissue compared to exoWT-treated and old liver tissue (Fig. 6k). These results supported the functional regeneration of the aged mouse liver after treatment with the exoWNT3A/RSPO1.
Discussion
The WNT signaling pathway has been extensively studied for its role in organ morphogenesis and tissue regeneration3,10,32. However, there has been a lack of tools for evaluating the therapeutic function of WNT-activating ligands for tissue repair and regeneration. Herein, we reported a novel strategy for in vivo delivery of WNT signaling-activating proteins, including WNT ligands and RSPO proteins, via engineered exosomes. We demonstrated that these exosomes and their combinations effectively regulate cell fate and robustly activate WNT signaling in vitro and in vivo. Remarkably, we uncovered the broad therapeutic potential of dual protein-carrying exosomes in treating mice with acute and chronic liver injuries and aging-related liver phenotypes. These findings underscore the transformative potential of this exosome-based platform in regenerating diseased and aged tissues via WNT signaling activation, offering a promising avenue for advancing regenerative medicine in adult tissues.
Spatial transcriptomic profiling studies have further revealed the robust activation of the WNT signaling pathway in the regenerating liver with acute injury or partial hepatectomy43. Macrophage-secreted WNT2 has been shown to stimulate hepatocyte proliferation through activating WNT/β-catenin signaling48. Additionally, macrophage-mediated engulfment of hepatocyte debris induces WNT3A expression and promotes the differentiation of liver progenitor cells into hepatocytes49. Furthermore, endogenous WNT9B and WNT7A are induced, likely playing distinct roles in the regeneration process4,43,50. These studies underscore the critical involvement of the WNT signaling pathway in liver tissue repair and regeneration, suggesting that modulating this pathway could serve as a promising therapeutic strategy for liver-associated diseases and injuries. However, current treatment modalities, such as WNT or RSPO protein-mimicking biologics or small molecule inhibitors that prevent β-catenin degradation, may fail to fully replicate the complex and nuanced functions of native WNT proteins. The limitation in WNT native protein use lies primarily in the hydrophobic nature of the WNT ligands, which complicates protein purification and, together with the presence of antagonists in the liver and circulatory system, hampers their long-range transport in vivo and restricts therapeutic utility10. Notably, our engineered exosomes demonstrated a protective role in preserving the activity of WNT ligands, even in the presence of WNT antagonists. This capability enhances the delivery of WNT ligands and promotes their functional efficacy. Of note, this study bridged the gap between profiling studies and therapeutic developments, where our platform could be used to screen and identify WNT signaling-inducing ligands for the regeneration of the liver and other organs.
Similar efforts to load specific WNT ligands, such as WNT3A and WNT5A, on exosomes have been reported22,51. Of note, Gao et al.21 reported that functional WNT3A-carrying exosomes required the co-expression of an engineered glypican. Surprisingly, we found that WLS alone was sufficient for the efficient loading of functional WNT ligands and further demonstrated the capacity of the molecule to load the entire WNT family of ligands on exosomes. In addition, our study found WNT ligands or RSPO protein alone has a limited capacity to induce hepatic cell fate and pathway induction in the tissue, while the combination of RSPO1, with either WNT3 or WNT3A, had the potential to induce and maintain the differentiation from HE cells to LPC (Fig. 2). Our study highlighted the importance of RSPO proteins for the WNT ligands to activate the pathway effectively on a single exosome. Consistent with this, we showed that the WNT signaling-inducing activity of exoWNT3A/RSPO1 was even higher than that of individual exosomes used in combination. Furthermore, we observed in the LPC induction experiments that the single ligand exosomes and dual ligand exosomes have differential effects in regulating downstream WNT signaling genes (Fig. 2e), highlighting the potential of our system for future mechanistic studies on WNT signaling.
The striking observation that exoWNT3A/RSPO1 could reverse various forms of liver damage suggested that these exosomes were likely achieving this exceptional feat through the induction of hepatocyte proliferation and, to a lesser extent, through the modulation of non-parenchymal cells that likely play different roles in different injury conditions52. Indeed, we observed a high proportion of Ki67+ cells after exoWNT3A/RSPO1 treatment, suggesting the activation of hepatocyte proliferation. We further demonstrated the direct effects of exoWNT3A/RSPO1 on PHH. In the mouse liver, a population of Axin2+, WNT-responsive hepatocytes around the CV has been well-described42, and we similarly observed strong induction of the WNT activity in these cells with exoWNT3A/RSPO1. Moreover, the importance of WNT signaling in inducing hepatocyte proliferation was also well-illustrated in studies that generated stable proliferating hepatocyte cultures from mouse and human livers33,53. Specifically, the efficiency of deriving proliferating mouse hepatocyte organoids is significantly increased through the sorting and enrichment of Axin2+ hepatocytes33. We found that these Axin2+ pericentral regions were diminished after liver injury and were restored after the administration of the exoWNT3A/RSPO1 (Fig. 5d). In parallel, we observed the recovery of the liver zonation from the re-emergence of the GS+ and CYP2E1+ hepatocytes. Notably, in comparison to specific zonal activity observed with exoWNT3A/RSPO1, Sun et al. demonstrated that the intra-peritoneal injection of recombinant RSPO1-Fc could further induce WNT signaling and metabolic reprogramming in Zone 1 hepatocytes15, suggesting that WNT-signaling activation could modulate hepatocytes across multiple zones. Overall, our study provided strong evidence that the activation of hepatocyte proliferation with WNT-inducing therapeutics could repair and regenerate the damaged and deteriorated liver induced by various conditions.
Previous studies have highlighted that PPARα activation promotes progenitor cell-mediated liver regeneration in zebrafish and induces hepatomegaly around the CV and hepatocyte proliferation around the portal vein in mice54,55. In this study, we observed strong activation of the PPAR signaling pathway in hepatic progenitor cells and various injured liver tissues treated with exosomes. Inhibiting PPARα signaling in the hepatic endoderm prevented the generation of hepatoblast, whose activation failed to induce the phenotype observed after WNT3A/RSPO1 treatment. These observations suggest that PPAR signaling likely works downstream with WNT signaling for hepatic cell fate determination. These results underline likely signaling crosstalk between the WNT and PPAR pathways to modulate liver repair and regeneration. In addition to its widely reported role in regulating metabolic functions in the liver, our study further highlighted the potential role of the PPAR signaling pathway in human and mouse liver tissue regeneration.
Our studies demonstrated that a systematic engineering approach enables the eventual development of an exosome with WNT ligand or RSPO protein loading and provides an approach to screen tissue-specific WNT agonists. This proof-of-concept study highlighted the immense potential for using engineered exosomes to overcome challenges in delivering biomolecules to target cells for therapeutic applications. Of note, we also established a workflow that enabled the efficient production of high-quality exosomes for future translational applications. In summary, this paradigm study highlights the immense potential of engineered WNT-activating exosomes in WNT-related research and potential clinical applications for tissue and organ regeneration.
Methods
Animal studies
C57BL/6 J mice (wild-type, male, 6 ~ 8 weeks for injury model or 18 months for aging experiment) were purchased from Gempharmatech (Guangdong, China). Axin2-mGFP mice were generated and supplied by Yi Arial Zeng laboratory. Mice were raised in a 12 h light/dark cycle at 25 °C and 60% humidity with free access to food and water. All experimental procedures were approved by the Ethics Committee of Guangzhou National Laboratory (AUCP-2022-12-A02) and performed by the Guide for the Care and Use of Laboratory Animals.
Engineered exosome preparation, isolation, and characterization
HEK293 (Procell system, China) and HEK293T (Procell system, China) cells were cultured in DMEM supplemented with 10% FBS and 1×penicillin-streptomycin. Cells were incubated at 37 °C with 5% CO2 in a humidified atmosphere. For virus packaging, HEK293T cells were seeded into a 10 cm dish and transfected with the indicated plasmids with lipo8000 for 48 hrs before the virus was harvested. HEK293 cells were infected with lentivirus with polybrene for 48 hrs, and the stable clonal cells were obtained through flow cytometry and puromycin selection. Engineered cells were cultured and expanded in serum-free media (Celer-S201) using a shaker flask. When the cell density reached 1×107 cells/mL, detected by the EVE™ PLUS automatic cell counter, the supernatant was collected and subjected to a series of centrifugation at 200 g for 5 mins to remove floating cells, 2000 g for 10 mins to remove cell debris, and 10,000 g for 20 mins to remove large EVs. After passing through a 0.22 μm filter, exosomes were isolated from the supernatant using EXODUS (HuiXin Bio., China) according to the manufacturer’s protocol. The purity, morphology, diameter, and concentration of exosomes were characterized by TEM (Talos L120C, Thermo Scientific) and Nano-FCM (nanofcm, China).
Human WNT3A and RSPO1 protein enzyme-linked immunosorbent assay
WNT3A and RSPO1 protein loading efficiency on exosomes was measured by enzyme-linked immunosorbent assay (ELISA). Briefly, 96-well microplates were pre-coated with capture antibody and incubated overnight at 4 °C. Before use, the coating antibody was aspirated, and the plate was rinsed with washing buffer. After blocking with 1% BSA, exosome samples or standards were incubated for 1.5 hrs at room temperature. After removing the blocking buffer, the protein-specific antibody was added and incubated for 1.5 hrs at room temperature. The antibody was removed, and samples were rinsed with the washing buffer three times. Subsequently, streptavidin-HRP was added and incubated for 20 mins at room temperature. After aspiration and washing, the TMB substrate was added to each well and incubated for 5 mins before adding the stop solution. The luminescence was measured using a microplate reader set to 450 nm.
WNT3A molecule estimation in each exosome
The number of WNT molecules was calculated by averaging the protein amount detected in ELISA with the molecular weight of the protein. Subsequently, the average number of particles for each exosome is determined by averaging with the number of particles measured using nano-flow cytometry (Nano-FCM).
TOPFlash activity assay
HEK293FT (Procell system, China) cells were cultured in DMEM supplemented with 10% FBS and 1×penicillin-streptomycin and seeded in a 96-well plate. Cells were transfected with TOPFlash plasmid with lipo8000 for 24 hrs and incubated with exosomes (1×109 particles/mL) or 50 ng/mL protein for WNT3A or RSPO1. After treatment for 24 hrs, cells were collected and analyzed using a Luciferase Report Assay System according to the manufacturer’s instructions. The absorbance was determined at 560 nm.
Hepatic endoderm cell culture and exosomes treatment
The human induced pluripotent stem cell line (iPSC) was kindly gifted from Dr. Igor Samokhvalov and cultured in mTeSR1 at 37 °C in 5% CO₂ as previously reported56. The iPSC was differentiated to posterior foregut (PFG) as previously reported34. Briefly, the iPSCs were digested into single cells using Gentle Cell Dissociation Reagent for 5 mins at 37 °C, and approximately 400,000 iPSCs were seeded in a 24-well plate pre-coated with 30×diluted Matrigel in mTeSR1 with Y-27632. After 24 hrs post-seeding, the media was changed to the definitive endoderm differentiation media (STEMdiff™) following the manufacturer’s instructions. After 4 days of differentiation, the cells were cultured for another 6 days in the PFG differentiation media containing Advanced DMEM/F12, 1% L-Glutamine, 20 ng/mL BMP4, 10 ng/mL bFGF, and 2% B27 supplement. For the generation of HE, PFG cells were digested as single cells using TrypLE Express, and seeded in culture plates pre-coated with Collagen I. Cells were seeded for 24 hrs in PFG media supplied with Y-27632, and media was changed to HE media containing Advanced DMEM/F12 containing 0.5% N2 supplement, 1% B27 supplement, 1% L-Glutamine, 1% HEPES, 1% penicillin/streptomycin, 10 mM nicotinamide, 1.25 mM N-Acetylcysteine, 0.1 nM Cholera endotoxin, 2 nM T3, 1 μM Dexamethasone, 0.5 μM Dasatinib, 50 ng/mL KGF, 50 ng/mL FGF10, 20 ng/mL TGFα, 50 ng/mL HGF, 250 ng/mL R-spondin1, supplied with 2 μM A-8301. Cells were treated for 8 days, and the media were replaced every 2-3 days. Stable HE cells were passaged every 6-7 days. After approximately five passages, cells were harvested for the detection of the HE markers and further experiments. To obtain LPC, cultured HE cells were digested into single cells using TrypLE Express, and 100,000 cells were seeded in 24-well collagen I pre-coated plate. Cells were seeded for 24 hrs in PFG media supplied with Y-27632, and subsequently incubated with HE media (without RSPO1) and with 3 μM CHIR99201 added for 8 days. For testing of exosomes and protein, the CHIR99201 was replaced with different exosomes (1×109 particles/mL for each kind of exosome) or 100 ng/mL protein for WNT3A or RSPO1. For stable LPC cultured with exosomes, the media were replenished every 2-3 days, and cells were passaged upon confluency.
Primary human hepatocyte (PHH) culture and exosome treatment
PHH cells were purchased from LV-BioTech (Lot# 386545, Shenzhen, China). Frozen cells were thawed using 10 mL recovery media, centrifuged at 50 × g for 5 mins at room temperature, and resuspended in coating media. Cells were seeded into 24-well plates pre-coated with Collagen I. After 24 hrs, the medium was replaced with maintenance media, which was changed every two days. For exosome treatment, 1 × 10⁹ particles/mL exoWT or exoWNT3A/RSPO1 particles were added to the maintenance media and cultured for 1 week, with media changed every two days.
Adult liver organoid culture and exosomes test
Adult liver organoids (Huayi Regeneration, China) were cultured as previously described41. Cells were maintained at 37 °C in 5% CO₂, with media replenished every 3-4 days, and passaged every 7-10 days upon confluency. For exosome treatments, liver organoids were dissociated into single cells and resuspended in 10 µL Matrigel (~40,000 cells/droplet). 1×109 particles/mL exoWT or exoWNT3A/RSPO1 were used to replace respective proteins in the media. Organoids were cultured for 10 days, with media and exosomes replenished every 3-4 days. Bright-field images were captured on day 10 post-seeding for comparison.
Exosome cellular uptake and body distribution tracing
Exosomes were stained with PKH26 according to the manufacturer’s instructions, and the residual dye was removed through EXODUS purification. Labeled exosomes (1×109 particles/mL) were incubated with HE cells or liver organoids for 24 hrs. After fixing with 4% PFA and staining with DAPI, exosome cellular uptake was observed using confocal microscopy. To determine the biodistributions of exosomes after intravenous administration, exosomes were stained with 10 μM DiR for 20 mins at room temperature, and the residual dye was removed through EXODUS. 200 μL labeled exosomes (1×1011 particles/mL) were injected into mice through the tail vein. The biodistribution images were captured using IVIS Lumina system (PerkinElmer), and the tissue section images were obtained using the confocal system.
shRNA-mediated knockdown of PPARA and PPARD and LPC induction
shRNAs targeting PPARA and PPARD were synthesized by Igebio (Guangzhou, China) and subcloned into the pLKO.1-puro vector between AgeI and EcoRI sites. Lentiviral particles were packaged in HEK293T cells using lipo8000. HE cells were infected with equal amounts of virus in the presence of 10 μg/mL polybrene. After 24 hrs, the medium was replaced with HE medium, in which RSPO1 was replaced by 1 × 10⁹ particles/mL of exoWNT3A/RSPO1, and cultured for 6 days with medium changed every two days. Knockdown efficiency was assessed by real-time PCR. The shRNA target sequences were as follows: PPARA-shRNA1: GAACAGAAACAAATGCCAGTA; PPARA-shRNA2: GTAGCGTATGGAAATGGGTTT; PPARA-shRNA3: GCTTTACGGAATACCAGTATT; PPARD-shRNA1: CCGCAAACCCTTCAGTGATAT; PPARD-shRNA2: GATCAAGAAGACCGAAACCGA; PPARD-shRNA3: GTGTGGAAGCAGTTGGTGAAT.
Immunocytochemistry
HE cells and adult liver organoids were fixed with 4% PFA and blocked with 1% BSA. Primary antibodies were incubated overnight at 4 °C, respectively. After washing with PBS, corresponding 488-conjugated or 594-conjugated secondary antibodies were incubated for 2 hrs at room temperature. Samples were washed twice before staining with DAPI. The fluorescence images were captured using confocal microscopy.
Acetaminophen (APAP) treatment and exosome administration
Mice were fasted overnight for a period of 16 hrs before APAP injections. APAP was freshly dissolved in sterile PBS (at 56 °C) at a concentration of 20 mg/mL. After complete dissolution, the stock was stored at 37 °C to avoid precipitation. A single intraperitoneal injection of APAP at a dosage of 600 mg/kg body weight was administered to male mice. Mice were maintained on a normal chow diet and water after APAP injections. ~2×1010 particles of exoWNT3A/RSPO1 or exoWT were intravenously administered into mice after APAP treatment for 24 hrs and 48 hrs. Mice were euthanized 24 hrs after the last exosome administration.
CCl4 treatment and exosome administration
Mice were treated with 25% v/v CCl4 (reconstituted in olive oil) or olive oil only (2.5 μL/g body weight) at a single dose to induce acute liver injury or twice per week for up to 6 weeks to induce chronic injury. For acute liver injury repair, ~2×1010 particles of exoWNT3A/RSPO1 or exoWT were intravenously administered into mice after CCl4 treatment for 24 hrs and 48 hrs. For the chronic liver injury repair, ~2×1010 particles of exoWNT3A/RSPO1 or exoWT were intravenously administered into mice once per week for 3 weeks or 6 weeks. Mice were euthanized 24 hrs after the last exosome administration.
Aging mice and exosome administration
18-month-old C57BL/6 J mice were treated with ~2 × 1010 particles of exoWNT3A/RSPO1 or exoWT through the tail vein twice a week for 2 weeks. Mice were euthanized 24 hrs after the last exosome administration.
Serum ALT/AST assay
Mouse blood was collected immediately after euthanizing and before liver separation and kept at room temperature for 4 hrs. Serum was separated by centrifugation at 2500 g for 10 mins. Serum ALT and AST levels were detected using the Aspartate aminotransferase Assay Kit or Alanine aminotransferase Assay Kit, according to the manufacturer’s protocol. Briefly, 10 μL serum diluted in PBS was mixed with the substrate at 37 °C for 30 mins, and kit reagents were added at 37 °C for 20 mins. The OD value was measured at 505 nm using a microplate reader.
Immunofluorescence and histology image analysis
After euthanizing, the liver was separated from the mice, and the resected tissue was fixed in 4% PFA directly. For cryopreservation, liver tissues were washed thrice with PBS, immersed in 15% sucrose solution for 15 mins, and transferred to and kept in 30% sucrose solution until they settled to the bottom. The tissues were then embedded in OCT solution and stored at −80 °C. Liver tissues were sectioned at 5 μm thickness using a cryostat (Leica) and allowed to defrost and dry at room temperature for 30 mins. The sections were dipped in PBS for 10 mins and permeabilized using 0.3% Triton-X100 for 10 mins. The sections were rinsed thrice in PBS and blocked with 3% normal sheep serum for 30 mins. Then, the sections were incubated with primary antibodies at 4 °C overnight. After three washes with PBS, samples were incubated with the corresponding fluorescent-labeled secondary antibodies for 2 hrs at room temperature, protected from light. The sections were finally washed with PBS and incubated with DAPI for 5 mins. The fluorescence images were captured using confocal microscopy. Image J was used for image analysis as well as quantification.
Single-EV Protein Localization by dSTORM
EVs were captured on tetraspanin antibody-coated coverslips (Smart EV kit, Abbelight) to isolate single vesicles. Samples were labeled with Alexa Fluor 647-conjugated anti-CD63, CD9, and CD81 antibodies (Abbelight), along with antibodies against WNT3A and RSPO1, which were then labeled with Alexa Fluor 532 and 488. Sequential excitation with 640, 532, and 488 nm lasers was performed, and 10,000 frames per ROI were acquired at 20 ms exposure. Auto-focus lock minimized z-drift during imaging. Data were processed with Abbelight NEO software for super-resolution reconstruction and drift correction. Imaging was carried out on an Abbelight SAFe 180 STORM system mounted on an Evident IX83 inverted microscope with a 100× 1.49 NA TIRF objective.
H&E staining
Mouse liver sections were deparaffinized in xylene and rehydrated through an ethanol gradient. After hydration in distilled water, sections were immersed in hematoxylins for 1 min, rinsed in running tap water, and differentiated in Scott’s tap water for 3 mins. After being rinsed in tap water, the sections were stained with Eosin-Y for 3 mins and dehydrated in absolute alcohol.
Masson Trichrome staining
Mice liver sections were deparaffinized in xylene and rehydrated through an ethanol gradient. After hydration with distilled water, sections were fixed in Bouin’s solution at room temperature overnight and washed in running tap water to remove the yellow color. The sections were stained in working Gills Hematoxylin Solution for 5 mins, washed in running tap water for 5 mins, and then rinsed in deionized water. The sections were further stained in Biebrich Scarlet-Acid Fucshin for 5 mins and rinsed in deionized water. Subsequently, the sections were placed in the working Phosphotungstic/Phosphomolybdic Acid Solution for 5 mins and in Anilline Blue Solution for 20 mins. Lastly, the sections were rinsed in acetic acid for 2 mins and dehydrated in absolute alcohol.
Sirius red staining
Mice liver sections were deparaffinized in xylene and rehydrated through an ethanol gradient. After hydration with distilled water, sections were immersed in Picro-Sirius red solution for 1 hr at room temperature and dehydrated in absolute alcohol.
TUNEL staining
TUNEL staining was performed using a one-step TUNEL cell apoptosis assay kit (green fluorescence, Beyotime) according to the manufacturer’s instructions. Briefly, frozen liver tissue sections were cut at a thickness of 10 μm and fixed with 4% paraformaldehyde for 30 mins at room temperature. The fixed sections were rinsed twice with PBS and permeabilized using 3% Triton X-100 in PBS for 20 mins, followed by two PBS washes. The sections were then incubated with TUNEL assay solution for 1 h at 37 °C. After staining, the sections were mounted with mounting medium containing DAPI, and images were captured using a Zeiss LSM 800 confocal microscope with 405 nm and 488 nm lasers.
Gene expression analysis
Total RNA was extracted from HEK293 cells, HE cells, adult organoids or part of mice liver tissues using TRIzol reagent and quantified by Nanodrop (Thermo). Genomic DNA was removed, and cDNAs were synthesized using the HiScript II 1st Strand cDNA Synthesis Kit ( + gDNA wiper). Samples were then used for real-time PCR with SYBR Green Mix, and gene expression was normalized to ACTB. Primers were obtained from Sangon Biotech., and the sequences are listed in Supplementary information Table S2.
Protein expression analysis
Total protein was extracted from HEK293 cells, large EVs, and exosomes using a RIPA lysis buffer supplemented with protease inhibitors. Protein concentration was quantified using BCA assay kit and denatured with 5×loading buffer at 100 °C for 10 mins. About 30 μg of each protein sample was loaded into 4-20% XPAGE™ gel. After SDS-PAGE, proteins were transferred onto PVDF membrane and blocked with 5% non-fat milk. Membranes were incubated with primary antibody overnight at 4 °C. After washing thrice with TBST solution, membranes were incubated with HRP-conjugated secondary antibody for 1 h at room temperature. The signal was detected using a High Sensitivity ECL Substrate Kit.
Protein mass spectrum analysis of exosomes
Extracellular vesicle proteins were reduced with 10 mM dithiothreitol (30 min, room temperature) and alkylated with 20 mM iodoacetamide (30 min, RT, dark). Proteins were digested sequentially with Lys-C (enzyme/protein, 1:100, w/w, 4 h) and trypsin (enzyme/protein, 1:50, w/w, 12 h, 37 °C). Peptides were quenched with 10% trifluoroacetic acid to a final concentration of 1% and desalted using C18 StageTips.
Peptide mixtures were analyzed on a Vanquish Neo UHPLC coupled online to an Orbitrap Astral mass spectrometer (Thermo Fisher Scientific) using a 150 mm Aurora Elite C18 column (75 μm ID, IonOptics). Separation was performed with mobile phase A (0.1% FA in water) and mobile phase B (0.1% FA in 80% ACN) under a 24-min gradient: 1–4% B (500 nL/min, 0.1 min), 4–10% B (500 nL/min, 1.8 min), hold at 10% B (flow from 500 nL/min to 300 nL/min, 0.1 min), 10–25% B (300 nL/min, 11 min), 25–45% B (300 nL/min, 7 min), 45–99% B (300 nL/min, 2 min), followed by 99% B (500 nL/min, 2 min).
The Orbitrap Astral was operated in data-independent acquisition mode with a FAIMS Pro Duo interface (compensation voltage −45 V). MS1 spectra were acquired in the Orbitrap (AGC target 500%, maximum injection time 5 ms, resolution 240,000, scan range 380–980 m/z). MS2 spectra were acquired after HCD fragmentation (normalized collision energy 25%) using the Astral analyzer (2-Th isolation window, AGC target 500%, maximum injection time 3 ms) across 380–980 m/z.
Raw data were processed in Spectronaut v19 (Biognosys) using “Direct-DIA” mode against the UniProt human database (20,397 entries, downloaded 16 October 2022). Trypsin was specified as the protease with up to two missed cleavages. Carbamidomethylation (C) was set as a static modification, while oxidation (M) and protein N-terminal acetylation were dynamic. Default parameters were applied otherwise.
Bulk RNA sequencing
mRNA sequencing was performed by Novogene Co., Ltd. (Beijing, China). For short, RNA integrity was assessed using the RNA Nano 6000 Assay Kit of the Bioanalyzer 2100 system (Agilent Technologies). mRNA was purified from total RNA using poly-T oligo-attached magnetic beads. After fragmentation, the first strand cDNA was synthesized using random hexamer primers, followed by the second strand cDNA synthesis. The library was checked with Qubit and real-time PCR for quantification and bioanalyzer for size distribution detection. Sequencing was performed on an Illumina Novaseq platform and 150 bp paired-end reads were generated. Clean data/reads were generated from raw data/reads using fastp software, and mapped to reference genome using Hisat2 (v2.0.5). For HE cells, adult hepatic organoids and primary human hepatocytes, reads were aligned against the GRCH38 human reference genome, while for mouse livers, reads were aligned against the mm10 mouse reference genome. Reads numbers mapped to each gene was calculated using featureCounts (v1.5.0-p3).
Sequencing data analysis was performed using the DESeq2 package (v1.44.0) in R environment (v4.4.0). Sample variability was performed by Principal Component Analysis (PCA) using top 20,000 genes. Differential gene expression analysis was performed using DESeq2 package (v1.44.0) and visualized by ComplexHeatmap package (v4.4.0). Gene set enrichment analysis (GSEA) and pathway enrichment were performed using clusterProfiler package (v4.12.2) against the KEGG gene sets or gene ontology (GO) gene sets57. All data analysis and visualization were performed using R studio (v2024.04.0).
Statistical analysis
GraphPad Prism 9 was utilised for statistical analysis. All statistical data are presented as mean ± SD. Two-tailed unpaired Student’s t tests were used to compare the two groups. The p-values are calculated and shown as: ns p ≥ 0.05, * p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data supporting this paper are presented within the paper and the Supplementary Materials. The mass spectrometry data for exosome composition produced by this research were deposited in the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD067589. The RNA-seq data generated in this study were deposited to the NCBI GEO database under accession codes GSE305923. Other previously published RNA-seq data used in this study are available in the NCBI GSE database under accession codes: GSE233413 and GSE112330.The original datasets are also available from the corresponding author upon request. Source data are provided with this paper.
References
Michalopoulos, G. K. & Bhushan, B. Liver regeneration: biological and pathological mechanisms and implications. Nat. Rev. Gastroenterol. Hepatol. 18, 40–55 (2021).
Huang, R., Zhang, X., Gracia-Sancho, J. & Xie, W. F. Liver regeneration: Cellular origin and molecular mechanisms. Liver Int 42, 1486–1495 (2022).
Pu, W. et al. Bipotent transitional liver progenitor cells contribute to liver regeneration. Nat. Genet 55, 651–664 (2023).
Xu, J. et al. A spatiotemporal atlas of mouse liver homeostasis and regeneration. Nat. Genet 56, 953–969 (2024).
Zhang, C., Sun, C., Zhao, Y., Ye, B. & Yu, G. Signaling pathways of liver regeneration: Biological mechanisms and implications. iScience 27, 108683 (2024).
Sun, T. et al. AXIN2(+) Pericentral Hepatocytes Have Limited Contributions to Liver Homeostasis and Regeneration. Cell Stem Cell 26, 97–107 e106 (2020).
May, S. et al. Absent expansion of AXIN2+ hepatocytes and altered physiology in Axin2CreERT2 mice challenges the role of pericentral hepatocytes in homeostatic liver regeneration. J. Hepatol. 78, 1028–1036 (2023).
Agostino, M., Pohl, S. & Dharmarajan, A. Structure-based prediction of Wnt binding affinities for Frizzled-type cysteine-rich domains. J. Biol. Chem. 292, 11218–11229 (2017).
Sato, N., Meijer, L., Skaltsounis, L., Greengard, P. & Brivanlou, A. H. Maintenance of pluripotency in human and mouse embryonic stem cells through activation of Wnt signaling by a pharmacological GSK-3-specific inhibitor. Nat. Med. 10, 55–63 (2004).
Willert, K. et al. Wnt proteins are lipid-modified and can act as stem cell growth factors. Nature 423, 448–452 (2003).
Planas-Paz, L. et al. The RSPO-LGR4/5-ZNRF3/RNF43 module controls liver zonation and size. Nat. Cell Biol. 18, 467–479 (2016).
Hao, H. X. et al. ZNRF3 promotes Wnt receptor turnover in an R-spondin-sensitive manner. Nature 485, 195–200 (2012).
Koo, B. K. et al. Tumour suppressor RNF43 is a stem-cell E3 ligase that induces endocytosis of Wnt receptors. Nature 488, 665–669 (2012).
Yan, K. S. et al. Non-equivalence of Wnt and R-spondin ligands during Lgr5(+) intestinal stem-cell self-renewal. Nature 545, 238–242 (2017).
Sun, T. et al. ZNRF3 and RNF43 cooperate to safeguard metabolic liver zonation and hepatocyte proliferation. Cell Stem Cell 28, 1822–1837.e1810 (2021).
Jeppesen, D. K. et al. Reassessment of Exosome Composition. Cell 177, 428–445 e418 (2019).
Kalluri R., LeBleu V. S. The biology, function, and biomedical applications of exosomes. Science 367, (2020).
Vo, N. et al. A novel multi-stage enrichment workflow and comprehensive characterization for HEK293F-derived extracellular vesicles. J. Extracell. Vesicles 13, e12454 (2024).
Gross, J. C., Chaudhary, V., Bartscherer, K. & Boutros, M. Active Wnt proteins are secreted on exosomes. Nat. Cell Biol. 14, 1036–1045 (2012).
McGough, I. J. et al. Glypicans shield the Wnt lipid moiety to enable signalling at a distance. Nature 585, 85–90 (2020).
Gao, L. et al. Wnt3a-Loaded Extracellular Vesicles Promote Alveolar Epithelial Regeneration after Lung Injury. Adv Sci (Weinh), e2206606 (2023).
Thomas, B. L. et al. WNT3A-loaded exosomes enable cartilage repair. J. Extracell. Vesicles 10, e12088 (2021).
Willert, K. & Nusse, R. Wnt proteins. Cold Spring Harb. Perspect. Biol. 4, a007864 (2012).
Zhong, Q. et al. Cryo-EM structure of human Wntless in complex with Wnt3a. Nat. Commun. 12, 4541 (2021).
Qi, X. et al. Molecular basis of Wnt biogenesis, secretion, and Wnt7-specific signaling. Cell 186, 5028–5040 e5014 (2023).
Nygaard, R. et al. Structural Basis of WLS/Evi-Mediated Wnt Transport and Secretion. Cell 184, 194–206 e114 (2021).
Korkut, C. et al. Trans-synaptic transmission of vesicular Wnt signals through Evi/Wntless. Cell 139, 393–404 (2009).
Chai, G. et al. A Human Pleiotropic Multiorgan Condition Caused by Deficient Wnt Secretion. N. Engl. J. Med 385, 1292–1301 (2021).
Chen, Y. et al. Exosome detection via the ultrafast-isolation system: EXODUS. Nat. Methods 18, 212–218 (2021).
de Lau, W. et al. Lgr5 homologues associate with Wnt receptors and mediate R-spondin signalling. Nature 476, 293–297 (2011).
de Lau, W. B., Snel, B. & Clevers, H. C. The R-spondin protein family. Genome Biol. 13, 242 (2012).
Huch, M. et al. In vitro expansion of single Lgr5+ liver stem cells induced by Wnt-driven regeneration. Nature 494, 247–250 (2013).
Hu, H. et al. Long-Term Expansion of Functional Mouse and Human Hepatocytes as 3D Organoids. Cell 175, 1591–1606 e1519 (2018).
Ramli, M. N. B. et al. Human Pluripotent Stem Cell-Derived Organoids as Models of Liver Disease. Gastroenterology 159, 1471–1486 e1412 (2020).
Wang, S. et al. Human ESC-derived expandable hepatic organoids enable therapeutic liver repopulation and pathophysiological modeling of alcoholic liver injury. Cell Res 29, 1009–1026 (2019).
Lv, Y. et al. The efficient generation of functional human hepatocytes from chemically induced pluripotent stem cells. Cell Prolif. 57, e13540 (2024).
Qiu, Y. Y., Zhang, J., Zeng, F. Y. & Zhu, Y. Z. Roles of the peroxisome proliferator-activated receptors (PPARs) in the pathogenesis of nonalcoholic fatty liver disease (NAFLD). Pharm. Res 192, 106786 (2023).
Xu, H. E. et al. Structural basis for antagonist-mediated recruitment of nuclear co-repressors by PPARalpha. Nature 415, 813–817 (2002).
Palkar, P. S. et al. Cellular and pharmacological selectivity of the peroxisome proliferator-activated receptor-beta/delta antagonist GSK3787. Mol. Pharm. 78, 419–430 (2010).
Shearer, B. G. et al. Identification and characterization of 4-chloro-N-(2-{[5-trifluoromethyl)-2-pyridyl]sulfonyl}ethyl)benzamide (GSK3787), a selective and irreversible peroxisome proliferator-activated receptor delta (PPARdelta) antagonist. J. Med Chem. 53, 1857–1861 (2010).
Huch, M. et al. Long-term culture of genome-stable bipotent stem cells from adult human liver. Cell 160, 299–312 (2015).
Wang, J. et al. Endothelial Wnts control mammary epithelial patterning via fibroblast signaling. Cell Rep. 34, 108897 (2021).
Ben-Moshe, S. et al. The spatiotemporal program of zonal liver regeneration following acute injury. Cell Stem Cell 29, 973–989.e910 (2022).
Kim, S. et al. Dual-mode action of scalable, high-quality engineered stem cell-derived SIRPα-extracellular vesicles for treating acute liver failure. Nat. Commun. 16, 1903 (2025).
Perugorria, M. J. et al. Wnt-beta-catenin signalling in liver development, health and disease. Nat. Rev. Gastroenterol. Hepatol. 16, 121–136 (2019).
Russell, J. O. & Monga, S. P. Wnt/beta-Catenin Signaling in Liver Development, Homeostasis, and Pathobiology. Annu Rev. Pathol. 13, 351–378 (2018).
Liu, J. et al. Wnt/β-catenin signalling: function, biological mechanisms, and therapeutic opportunities. Signal Transduct. Target. Ther. 7, 3 (2022).
Li, J. et al. PGD 2 /DP1 axis promotes liver regeneration by secreting Wnt2 in KCs in mice. Hepatology, (2024).
Boulter, L. et al. Macrophage-derived Wnt opposes Notch signaling to specify hepatic progenitor cell fate in chronic liver disease. Nat. Med 18, 572–579 (2012).
Zhao, L. et al. Tissue Repair in the Mouse Liver Following Acute Carbon Tetrachloride Depends on Injury-Induced Wnt/β-Catenin Signaling. Hepatology 69, 2623–2635 (2019).
Dzialo, E. et al. WNT3a and WNT5a Transported by Exosomes Activate WNT Signaling Pathways in Human Cardiac Fibroblasts. Int. J. Mol. Sci. 20, (2019).
Rizvi, F. et al. Murine liver repair via transient activation of regenerative pathways in hepatocytes using lipid nanoparticle-complexed nucleoside-modified mRNA. Nat. Commun. 12, 613 (2021).
Zhang, K. et al. In Vitro Expansion of Primary Human Hepatocytes with Efficient Liver Repopulation Capacity. Cell Stem Cell 23, 806–819 e804 (2018).
Kim, M., So, J. & Shin, D. PPARalpha activation promotes liver progenitor cell-mediated liver regeneration by suppressing YAP signaling in zebrafish. Sci. Rep. 13, 18312 (2023).
Fan, S. et al. YAP-TEAD mediates PPAR alpha-induced hepatomegaly and liver regeneration in mice. Hepatology 75, 74–88 (2022).
Philonenko, E. S. et al. Recapitulative haematopoietic development of human pluripotent stem cells in the absence of exogenous haematopoietic cytokines. J. Cell Mol. Med 25, 8701–8714 (2021).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Acknowledgements
This work was funded by Guangzhou National Laboratory Fund (MP-GZNL2025C02009-01 (Y.C)), Guangdong Provincial Pearl River Talents program (2023QN10Y489 (Y.C.)), and National Natural Science Foundation of China (82370053 (L.Y.) and 32200637 (X.W.)). We also thank Qing Zhang from Proteomics and Metabolomics Core Facility, Guangzhou National Laboratory, for her assistance with the sample preparation, LC–MS/MS experiment and data analysis.
Author information
Authors and Affiliations
Contributions
L.Y. and S.W. performed and analyzed experiments, prepared the figures, and wrote the manuscript. Z.P., Y.L., X.W., and L.Y. designed and analyzed experiments. C.Y., J.D., M.L., and J.Z. performed the experiments. W.H. and Y.C. contributed to the interpretation of the results. Y.C. designed the experiment and wrote the manuscript. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Bruce Wang who co-reviewed with Cindy Ament; Xiangrui Liu and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yang, L., Wang, S., Qiao, Z. et al. Dual-ligand engineered exosome regulates WNT signaling activation to promote liver repair and regeneration. Nat Commun 16, 9019 (2025). https://doi.org/10.1038/s41467-025-64069-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-64069-8
This article is cited by
-
Bioengineering of extracellular vesicles with scaffold proteins for drug delivery
Journal of Nanobiotechnology (2026)