Abstract
Duchenne muscular dystrophy (DMD) is characterized by progressive muscle wasting and weakness. Serum proteins may offer insight into disease processes and clinical decline. This observational study uses the 7 K SomaScan® assay to discover serum proteins associated with muscle function and disease milestones. In total 702 serum samples from 153 male patients, collected across two centers (2009–2022), are analyzed. Using linear mixed effects modelling, we evaluate age and corticosteroid use as covariates affecting protein levels and assess protein correlations with longitudinal clinical function. Here we show 318 aptamers (294 proteins) significantly associated with motor performance across the two sites, with most associations found with lower limb functional tests (NSAA, 10MRW, and 6MWT). Thirty-six proteins are associated with milestones including RGMA, ART3, ANTXR2, and DLK1. These proteins show promise as prognostic biomarkers, and could potentially be used for patient stratification in clinical trial design and for monitoring interventions.
Similar content being viewed by others
Introduction
Duchenne muscular dystrophy (DMD) is the most common form of muscular dystrophy1, caused by variants in the DMD gene that result in an absence of functional dystrophin2. Without dystrophin, skeletal muscles are repeatedly damaged and replaced by fibrofatty tissue over time3. Symptoms typically begin in early childhood with delayed motor milestones and include progressive muscle weakness. Untreated, most individuals lose the ability to walk by the age of 10 years, followed by loss of the ability to use the upper extremities4,5.
Current treatment for DMD consists of chronic corticosteroids (CS), including prednisone or deflazacort. CS use has been shown to improve strength and delay loss of ambulation, and when combined with non-invasive ventilation, CS can increase life expectancy to 30–40 years of age4,5. There is variability in CS dosing strategies between physicians and centers. Higher CS doses and daily administration slow motor decline to a greater extent than lower doses and intermittent dosing schedules6,7; however, higher cumulative doses increase the risk of adverse effects such as negative metabolic consequences, bone fragility, and delayed puberty7,8.
Despite the use of CS, progress in understanding the biology of the disease, and the large investment in pre-clinical research, there is currently no cure for DMD. Multiple investigational drugs have failed to show significant improvement in participants’ performance in clinical trials, with only a few disease-modifying drugs receiving commercial approval9,10,11,12. While drug trial failures may be related to limited drug potency, it has also become clear that inter-individual variation in disease trajectories and limited sensitivity of motivation-sensitive outcome measures complicate the detection of significant and clinically meaningful treatment effects in clinical trials. Additionally, participants are typically enrolled in clinical trials before the age of 8, during the early stages of disease progression, when minimal decline is expected within the duration of an interventional study. Therefore, being able to objectively predict long-term clinical outcomes based on biological evidence would greatly facilitate investigational drug trial design and conduct.
Numerous serum biomarkers have been proposed for DMD13,14,15. However the focus has mostly been on the identification of cross-sectional differences between unaffected controls and individuals with DMD. Research efforts have focused on the effects of CS use on biomarkers15, muscle damage biomarkers16,17,18, and monitoring response to (micro-)dystrophin restoration therapies19,20. However, serum biomarkers with prognostic or predictive value are currently lacking. Initial attempts to determine longitudinal trajectories in serum biomarkers related to disease progression18,21,22 have been limited by small samples, incomplete clinical and follow-up data, and limited numbers of proteins assessed. Therefore, discovery studies in large and well-characterized cohorts are needed to identify biomarkers that correlate with clinical performance and that predict clinically meaningful disease milestones.
In this retrospective, multicohort study, we aimed to identify proteins in serum associated with clinical function and predictive of meaningful disease milestones in individuals with DMD, while accounting for the effects of age and corticosteroid treatment. Using the 7 K SomaScan®, we analysed 702 longitudinal serum samples obtained from 153 individuals with DMD. We present a collection of serum proteins that are associated with, and predictive of, clinical function in DMD.
Results
Serum samples from 74 males with DMD from the Leiden University Medical Center (LUMC) cohort and 79 males with DMD from the University of Florida (UF) cohort were included in the study. In total, 693 of 702 samples (98.7%) passed quality control standards for inclusion in analyses (see methods). Table 1 gives details of participant and sample characteristics for both cohorts. LUMC participants were significantly younger at the first sample visit compared to UF participants (mean[SD]; 8.4[3.4] vs 10.9[3.2], p < 0.001). An average of 4.3 serum samples per participant were analysed, with longer follow-up duration in the LUMC cohort (5.7[3.6] vs 3.4[2.5] years, p < 0.001). An intermittent CS regimen (10 days on/10 days off) was most common in the LUMC cohort, while daily dosing was most common in the UF cohort. Age at initiation of CS treatment was comparable across sites. Only a few individuals remained CS naïve for the entire study period (4.1% vs 3.2%).
Serum protein associations with age and corticosteroid use
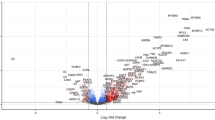
Given the progressive nature of DMD, we first identified proteins associated with age. Overall, 4796 probes (4436 proteins) were associated with age in the LUMC cohort and 2668 probes (2498 proteins) in the UF cohort (FDR < 0.05)(Fig. 1a, b). A total of 2317 probes (2186 proteins) were shared between cohorts, with 2251 probes showing concordant directional change (Fig. 1c, d). Multiple muscle proteins were negatively associated with age, including creatine kinase (CK-MM), myomesin 3 (MYOM3), titin (TTN), and troponin I2 fast skeletal type (TNNI2) (Fig. 1e–h). Other notable negative associations with age include proteins involved in fibrosis, extracellular matrix as well as bone morphogenetic proteins and protein synthesis pathways.
a Volcano plots showing the strength and significance of the association with age in the LUMC cohort, and b in the UF cohort. For each probe (point), the -log10 FDR (y-axes) and coefficient (x-axes) of age as determined by an LMEM probe expression from age are shown. The horizontal line represents the threshold for significance, FDR < 0.05. c Venn diagram showing the overlap of protein numbers associated with age in the LUMC (orange) and UF cohorts (blue). d Scatterplot of the coefficients for age shown in (a, b) in the LUMC (x-axis) versus UF (y-axis) cohorts. Dark green dots show proteins significantly associated with age in both cohorts. e–h Trajectory plots of selected probes decreasing with age such as CK-MM, MYOM3, TTN, and TNNI2, colored by site. i–l Trajectory plots of selected probes increasing with age such as CNTN3, CNDP1, LEP, and TIMP4, colored by site. (RFU relative fluorescence units, DMD Duchenne muscular dystrophy, LUMC Leiden University Medical Center, UF University of Florida, CK-MM muscle creatine kinase, MYOM3 myomesin 3, TTN, titin TNNI2, troponin I2 fast skeletal type, CNTN3 contactin 3, CNDP1 carnosine dipeptidase 1, LEP leptin, TIMP4 tissue inhibitor of metalloproteinase 4). Source data are provided as a Source Data file.
Proteins positively associated with age include central nervous system proteins such as contactin 3 (CNTN3) and carnosine dipeptidase 1 (CNDP1), adipose tissue proteins such as leptin (LEP) and tissue inhibitor of metalloproteinase 4 (TIMP4), and various cytokines (Fig. 1i–l). Other noteworthy findings include opposing age associations for two insulin-like growth factor 1 (IGF1) probes, as well as discordant trajectories for several insulin-like growth factor binding proteins, with IGFBP5 and IGFBP6 increasing and IGFBP1 and IGFBP2 decreasing with age. A full list of all significant concordant proteins and their coefficients, and whether they were previously reported15,21,23 or newly discovered, can be found in Supplementary Data 1. Pathway analysis of proteins associated with age shows the sustained inflammation and pro-fibrotic components related to progression in DMD (Supplementary Data 2).
Given that the majority of participants had chronic exposure to CS, we next identified protein expression related to CS exposure. Because of the association of a large number of proteins with age, we treated age as a covariate to isolate the effects of CS treatment. In the LUMC cohort, 846 probes (790 proteins) were significantly associated with CS use compared to 396 probes (364 proteins) in the UF cohort (FDR < 0.05)(Fig. 2a, b). Furthermore, 244 probes (227 proteins) were shared across sites (Fig. 2c, d). Among the significant probes, we observed previously reported proteins like CD23 and matrix metalloproteinase-3 (MMP3)8, as well as newly identified proteins such as immunoglobulin lambda-like polypeptide 1 (IGLL1) and repulsive guidance molecule A (RGMA) (Fig. 2e, h).
Volcano plots showing strength and significance of the associations between proteins and corticosteroid use in a the LUMC cohort, and b the UF cohort. For each probe (point), the −log10 FDR (y-axes) and coefficient (x-axes) of CS use as determined by an LMEM probe expression from CS use and age are shown. A few probes are highlighted by arrows. The horizontal line represents the threshold for significance, FDR < 0.05. c Venn diagram showing the overlap of proteins associated with corticosteroid use in common between LUMC (orange) and UF (blue) cohorts. d Scatterplot of the coefficients for CS use shown in (a, b) in LUMC (x-axis) and UF (y-axis) cohorts. Box plots of probe expression in CS treated versus untreated patients (LUMC 405 samples, UF 79 samples) for two proteins previously reported in association with CS use (e, f) and two proteins identified in this study (g, h). For each box, the center line represents the median and the lower and upper edges represent the 25th (Q1) and 75th (Q3) percentiles respectively. Whiskers extend to the most extreme probe expression values within 1.5 × interquartile range from Q1 and Q3. i Scatterplot showing the relationship between age coefficients (x-axis) and CS coefficients (y-axis) for probes significantly associated with CS treatment. Proteins showing discordant coefficients for age and CS treatments were considered as efficacy biomarkers, while those with concordant coefficients were considered as safety biomarkers (shaded gray). Orange dots represent estimates for the LUMC cohort, while blue dots represent estimates for the UF cohort. (CS corticosteroids, DMD Duchenne muscular dystrophy, LUMC Leiden University Medical Center, UF University of Florida, RFU relative fluorescence units, MMP3 matrix metalloproteinase-3, IGLL1 immunoglobulin lambda like polypeptide 1, RGMA repulsive guidance molecule A). Source data are provided as a Source Data file.
To assess whether these associations were potentially related to CS efficacy or safety effects, we analysed the direction of the association with age and CS. Proteins showing opposite trends (such as declining with age as disease progresses and increasing with CS treatment) were considered potential CS efficacy biomarkers, while those with concordant effects (such as decreasing with both age and CS treatment) were considered safety biomarkers (Fig. 2i). A total of 44 proteins were identified as efficacy biomarkers, including previously identified proteins such as angiopoietin-2 (ANGPT2) and proteins related to lipoprotein transport, as well as newly identified proteins such as ADP-ribosyltransferase 3 (ART3) and RGMA (Fig. 2h) (Supplementary Data 3). A total of 172 proteins were found as potential safety biomarkers, including previously identified MMP3, Afamin, IGFBP5, and proteins involved in SMAD signaling, as well as new proteins including IGF binding proteins and metalloproteinases (Supplementary Data 4). Pathway analysis of proteins associated with corticosteroids treatment was enriched with pathways related to cholesterol and lipids metabolism, as well as immunomodulation and inflammation (Supplementary Data 2).
Serum protein associations with longitudinal motor performance
To identify proteins associated with clinical severity, we next evaluated associations with performance on common functional assessments used to monitor motor function in DMD, including the North Star Ambulatory Assessment (NSAA), timed ten-meter walk/run test (10MRW), six minute walk test (6MWT), and Performance of Upper Limb 2.0 (PUL). Individual trajectories of performance on these tests are shown in Fig. 3a–d, and age and CS use were modeled as covariates in analyses. In the LUMC cohort 2,005 probes (1882 proteins) showed a significant association with at least one motor function assessment compared to 483 probes (454 proteins) in the UF cohort (FDR < 0.05). Of these, 318 probes (294 proteins) were shared across the two sites, with most associations found with lower limb functional tests (NSAA, 10MRW, and 6MWT). This is consistent with the larger number of data points and the larger degree of functional decline in the cohorts on these assessments compared to the PUL. A scatterplot of coefficients from each cohort shows concordant directionality (Fig. 3e–h).
a–d Longitudinal trajectory plots of NSAA scores, 10MRW velocity, 6MWT distance, and PUL 2.0 scores from both cohorts. Error bands represent 95% confidence intervals. e–h Scatterplots of coefficients, indicating the direction of protein association with motor function test performance (subpanels) in the LUMC (x-axes) versus UF (y-axes) cohorts. Bolded data points represent significant associations with the number of significant associations listed at the top-left in each subpanel. i UpSetR plot of significant probes for test of motor function. Bar plots at the bottom-left show the number of probes associated with each individual motor function test, colored by site. j Trajectory plots showing the relationship between ART3 levels (y-axes) and NSAA score (left panels) or 6MWT distance (right panels) by age for the LUMC cohort (top panels) and the UF cohort (bottom panels). The color code (depicted on the top-right in each panel) represents NSAA scores and 6MWT distances. (CS corticosteroids, DMD Duchenne muscular dystrophy, LUMC Leiden University Medical Center, NSAA North Star Ambulatory Assessment, PUL Performance of the Upper Limb 2.0, UF University of Florida, RFU relative fluorescence units). Source data are provided as a Source Data file.
To assess the generalizability of models, we applied the models trained on each cohort to predict clinical tests results of the second cohort. Validation results showed highest reconstruction accuracies for the NSAA, 10MRW, and 6MWT with Q2 values comparable across both cohorts for the 10MRW. Models trained on the LUMC cohort performed better for the 6MWT, while models trained on the UF cohort were more accurate for the NSAA. In the LUMC cohort, 122 probes (116 proteins) significantly and accurately predicted all three scales used to monitor lower limb function (NSAA, 10MRW, 6MWT) compared to 109 probes (105 proteins) in the UF cohort (Fig. 3i). Notably, ART3, RGMA and dihydrolipoamide dehydrogenase (DLD) showed highly significant associations and accurate predictions across multiple motor function assessments (Fig. 3j). A list of all proteins with significant associations across functional assessments and cohorts, and whether they are previously reported15,21,23 or newly discovered, is provided in Supplementary Data 5. Pathway analysis showed that proteins associated with motor function were related to energy metabolism and skeletal muscle contraction (Supplementary Data 2).
Prediction of clinical milestones
Events such as loss of the ability to ambulate (LoA), loss of the ability to reach overhead (OHR), and loss of the ability to bring the hand to the mouth (HTM) are clinically relevant disease milestones in DMD. Survival curves for these milestone events are shown in Fig. 4a for each cohort. We sought to determine whether the proteins associated with the NSAA, 10MRW, and 6MWT, could predict LoA and whether proteins associated with PUL could predict OHR and HTM. We identified 41 probes (36 proteins) significantly associated with one or more milestones (Fig. 4b). Three probes, targeting RGMA, ectonucleotide pyrophosphatase/phosphodiesterase family member 5 (ENPP5), and regulator of G protein signaling 21 (RGS21), were associated with two different milestones (FDR < 0.05). RGMA was associated with milestones in both cohorts, while ENPP5 (LUMC) and RGS21 (UF) were cohort specific. Associations for delta-Like Non-Canonical Notch Ligand 1 (DLK1) and ART3 were confirmed by two independent probes: DLK1 was associated with loss of OHR in the UF cohort and ART3 with LOA in the LUMC cohort (Fig. 4c). The direction of loge hazard ratios (lnHR) remained consistent across probes and outcomes. Notably, proteins like RGMA and anthrax toxin receptor 2 (ANTXR2) had large negative lnHRs, while receptor-type tyrosine-protein phosphatase delta (PTPRD) and neural cell adhesion molecule (NCAM) had large positive lnHRs. Kaplan–Meier plots for RGMA across all three milestones are presented for both cohorts (Fig. 4d). Supplementary Fig. 1 shows trajectory plots for RGMA, DLK1, ART3, and ANTXR3, while Supplementary Data 6 lists all proteins significantly predicting lower and upper limb milestones.
LoA: LUMC n = 74, UF n = 78; OHR LUMC n = 74, UF n = 76; HTM LUMC n = 74, UF n = 77. a Survival curves showing the probability (y-axis) of never achieving LoA, loss of OHR, and loss of HTM (solid/dashed/dotted lines) by age in each cohort. b UpSetR plot of probes significantly associated with each clinical milestone by the Cox model. Bar plots at the bottom-left show the number of probes associated with each individual milestones, color coded by site. c Forest plot showing lnHR (x-axes) of protein probes (y-axes) significantly associated (FDR < 0.05) with at least one clinical milestone. Points represent the estimated lnHR, error bars represent the 95% confidence intervals. Each panel shows the proteins associated with an individual milestone. d Survival curves for the 3 clinical milestones (columns) stratified by RGMA expression quartiles at first visit (colors), by years since first visit (x-axes). The first row represents curves for LUMC, the second row represents curves for UF. (HTM hand-to-mouth, LoA loss of ambulation, OHR overhead reach, UF University of Florida, LUMC Leiden University Medical Center). Source data are provided as a Source Data file.
From the many proteins associated with functional tests and disease milestones, we shortlisted those with the most consistent and strongest associations. The selection was based on significance across multiple clinical scales and milestones within and across cohorts, and based on associations detected by more than one aptamer (Table 2). Proteins ANTXR2, ART3, euchromatic histone lysine methyltransferase 2 (EHMT2) and RGMA were linked to the risk of reaching disease milestones, with risks increasing between 136 and 981% for every one-unit decrease in the ln-transformed, standardized expression. In contrast, CS treatment was associated with increased biomarker levels, corresponding to a reduction in risk up to 90%. The greatest risk reduction was observed for the loss of HTM milestone. For the LoA milestone, risk reductions ranged from 22 to 39% in all cases. Additionally, we estimated how monitoring these proteins in blood can help evaluate the yearly risk increase for reaching these milestones, with the increase in risk ranging from 43 to 215% annually, depending on the protein.
Discussion
Large-scale serum biomarker discovery is now possible in DMD with the availability of high-throughput proteomics platforms such as SomaScan®. In this retrospective study, we had a unique opportunity to combine robust and comprehensive clinical data from two large, independent international cohorts of individuals with DMD along with serum levels of 6628 proteins. The availability of extensive longitudinal serum samples and clinical data allowed us not only to analyse protein signatures associated with age and corticosteroid usage, but also to identify novel proteins that could predict clinical function and motor milestones. Proteins like RGMA were found to predict both lower and upper limb clinical milestones including LoA and loss of HTM. Furthermore, several proteins were associated with an increased risk of a specific clinical milestone such as ART3 for LoA. The ability to compare these findings across two independent cohorts demonstrated the generalizability of our results.
Previous serum biomarker studies in dystrophinopathies have enabled the identification of proteins able to discriminate between individuals with and without DMD as well as described proteins associated with age. For instance, it is well-established that serum creatine kinase (CK) protein levels and activity decrease with age, reflecting muscle damage and loss of muscle mass. A few studies have reported SomaScan® data in DMD, identifying additional age-related biomarkers such as LEP21, MYOM38, and complement C4-A23. Complementing these findings, we show how the decline over time of muscle proteins is combined with an increase in adipogenic markers including LEP, growth hormone receptor (GHR), and adiponectin (ADIPOQ), along with proteins involved in complement activation and inflammation. Such observations show how monitoring proteins in serum reflect the active substitution of muscle mass with adipose tissue. Future studies should aim to assess whether protein levels in serum are directly associated with magnetic resonance-derived muscle fat fraction.
Analysis of CS treatment showed MMP3 and IGLL1 as the proteins most consistently associated with CS treatment both in daily and intermittent dosed patients. Previous research has also identified elevated MMP3 in individuals with DMD treated with CS24, while IGLL1 is novel in relation to DMD. However, recent research has shown that weekend CS use in adults with limb girdle muscular dystrophy (LGMD) reduces IGLL1 levels in conjunction with increased MMP325. Given the role of IGLL1 in B cells, its reduction alongside CS treatment may reflect the immunosuppressive effects exerted by the drug. CS treatment had both normalizing and exacerbating effects on disease progression proteins. For instance, it exacerbated age related biomarkers, including previously reported MMP3, Afamin, and IGFBP58, as well as certain apolipoproteins such as APOA2, APOL1, and APOA5. In contrast, a normalization was observed for other apolipoproteins such as APOE4, APOC3, and APOE, suggesting that steroid treatment could on one hand normalize dyslipidemia (with effects on APOE4, APOC3, and APOE) and on another hand affect lipid metabolism and potentially cardiovascular health (with exacerbation of APOA2, APOL1, and APOA5 levels). The compensatory effect of APOE is also supported by the more severe phenotype observed in mdx ApoE double knockout mice26,27. Moreover, we found that ANGPT28, previously associated with disease progression, was elevated in those treated with CS compared to those untreated. Likewise, RGMA, DLK1, ANTXR2, and ART3, which decreased with disease progression in both UF and LUMC cohorts, were increased by treatment with CS.
A major strength of this study was the ability to identify proteins associated with clinical function. RGMA and ART3 were directly related to patients’ performance, as measured by outcomes assessing both upper (PUL) and lower limb (NSAA, 6MWT, 10MRV) function. RGMA, ANTXR2, EHMT2, DLK1, and ART3 had large negative lnHR when considering clinical milestones. To further illustrate this finding, we stratified the population by RGMA levels and found that lower levels of RGMA corresponded to earlier disease milestones (LoA and OHR).
RGMA is part of the repulsive guidance molecule family of glycoprotein-1 (GP1) anchor proteins, primarily expressed in the central nervous system and muscle tissue, according to the Human Protein Atlas gene expression data. Initially recognized for its role in neurogenesis, guiding axonal growth, and serving as a key target for neuronal survival, RGMA has since been implicated in myogenesis28,29. It is proposed to play a central role in regulating cellular hypertrophy and hyperplasia30. Furthermore, RGMA has been identified in association with several conditions, including spinal and bulbar muscular atrophy (SBMA)31, Parkinson’s disease, Alzheimer’s disease, multiple sclerosis, and cerebrovascular accidents, as well as in association with upper limb function measured by elbow flexion in DMD21,28. Importantly, Somascan cannot discriminate between RGMA and RGMB, so future analyses should aim to refine such associations using orthogonal validation analyses.
ANTXR2, also known as capillary morphogenesis protein 2 (CMG2) plays an important role in cellular interaction by binding collagen IV and laminin, suggesting involvement in extracellular matrix adhesion. It is expressed in various tissues, including muscles. Loss-of-function variants in ANTXR2 cause hyaline fibromatosis syndrome, and ANTXR2 knockout mice show collagen VI accumulation in the uterus32 suggesting a potential involvement in muscle homeostasis.
ART3, known as ADP-ribosyltransferase 3, is mainly expressed in skeletal muscle tissue, according to GTEx and Human Protein Atlas databases. Gene expression data from both human individuals and Chinese Meishan pigs showed that ART3 is primarily expressed in muscles rich in fast twitch fibers33,34, which are more susceptible to damage in DMD. ART3 has been shown to decrease in serum across multiple dystrophies and myopathies25 and in SBMA31. In SBMA, decreased RGMA, myostatin, and ART3 expression correlated with higher thigh MRI muscle fat fraction, akin to patterns observed in DMD31. Additionally, research in Wannanhua pigs suggests ART3 is involved in fat deposition in muscle35, further strengthening the biological rationale behind the association with DMD identified in this study. This is the first study to show the relationship between ART3’s and CS status and clinical function in DMD.
Finally, EHMT2, known as euchromatic histone lysine methyltransferase 2, and DLK1 known as delta homolog 1 as well as Pref-1 (preadipocyte factor 1) showed interesting associations. Both EHMT2 and DLK1 declined with age and were associated with disease milestones. EHMT2 was normalized by CS treatment. EHMT2 was previously associated with renal fibrosis36, atrial fibrosis37, cardiomyocytes hypertrophy38 and high fat diet induced obesity and hepatic insulin resistance39, all of which align with the ongoing pathogenic processes in DMD. The reduction of EHMT2 with age may reflect the diminishing magnitude of pathological processes as muscle mass is lost progressively. DLK1, a transmembrane protein involved in cell growth during development, is expressed at low levels in adults, primarily in endocrine tissues. It regulates the differentiation of multiple cell types, including adipocytes, and plays an important role in skeletal muscle biology during fetal development and postnatal growth40. Although the role of DLK1 in adult skeletal muscle regeneration is less clear, upregulated expression has previously been observed in DMD and Becker muscular dystrophy41. In contrast, reduced DLK1 expression in fibroadipogenic progenitors corresponded to increased adipogenic committment42. This reduction in DLK1 levels may be associated with increased adipogenic commitment in DMD.
One limitation of this study was the variability in cohort characteristics, due to differences in participant pools and standards of care. The LUMC samples consisted of participants seen clinically, who were generally younger and primarily treated with a 10 days on, 10 days off CS regimen. No data were available on whether a patient was in the “on” or “off” phase of treatment at the time of sample collection. In contrast, the UF cohort samples were from participants in a natural history research study, who were generally older and primarily on a daily CS regimen, with several individuals co-enrolled in investigational drug trials receiving either placebo or the investigated drug (11.6% of samples in the LUMC cohort and 29.2% of samples in the UF cohort). Daily treatment with steroid most likely led to the observed slower motor decline in the UF cohort as well as the later occurrence of disease milestones compared to the LUMC cohort (where intermittent treatment was mostly used). The smaller proportion of individuals reaching milestones and the slower motor decline in the UF cohort may explain why fewer probes were associated with clinical outcomes and milestones in that cohort. As these different CS dosing strategies coincided with cohort effects, we were unable to directly compare the impact of intermittent CS compared to daily CS. This comparison could potentially be explored in prospective research, such as the FOR-DMD clinical trial. Additionally, we did not correct for steroid type, as a substantial number of patients had switched between CS types during the studied time period. We also did not correct for CS dose, as we found that site-specific effects, likely driven by the variations in treatment between the two cohorts, had a more substantial impact on the data than dose itself. Lastly, due to the retrospective nature of this study, we had a number of missing data points. Despite these limitations, the relatively large patient population and number of samples enabled the identification and validation of multiple serum proteins significantly associated with function across both cohorts.
In conclusion, we identified proteins associated with clinically meaningful outcomes in individuals with DMD across two independent cohorts. RGMA, DLK1, ANTXR2, EHMT2 and ART3 emerged as potential prognostic biomarkers based on the strength of their associations with clinical milestones, significance of the findings across scales, and their biological plausibility in connection to disease processes. A serum biomarker panel that accurately detects these proteins could enable the connection of short-term changes to disease stabilization and a decreased risk of decline in the mid- to long-term.
These biomarkers could have significant potential for both clinical management and clinical trials. In clinical practice, they could act as prognostic tools, helping to predict individual disease trajectories, enabling earlier interventions, and improving patient monitoring. Additionally, they could support more personalized treatment strategies by identifying individuals at risk of reaching disease milestones, allowing for timely adjustments in care. In the context of clinical trials, these proteins could refine participant selection criteria, better stratify participants based on their likelihood of disease progression, and provide an alternative readout to monitor treatment effects. These findings open the door for serum biomarkers to play a critical role in both clinical care and research in DMD.
Methods
Study cohort, design, and outcomes
This was a retrospective, multicenter, cohort study including serum samples and clinical data collected from individuals with DMD participating in research protocols at the Leiden University Medical Center (LUMC) and at the University of Florida (UF) between 2009 and 2022. We included 407 serum samples from 74 individuals aged 4–24 years at LUMC and 295 serum samples from 79 individuals aged 5–22 years at UF. Samples were included based on availability. At LUMC, blood samples were collected during annual outpatient clinic visits as part of routine clinical care, though the exact timing of blood collection was not standardized. At UF, blood sample collection was an optional addition for research participants enrolled in the ImagingDMD natural history study (NCT01484678), which included muscle magnetic resonance imaging and functional data collection. Blood samples were typically obtained at the conclusion of these visits. Written informed consent was obtained from all participants or their caregivers as described in protocol B22.013 at LUMC and IRB201500981 and IRB201700056 at UF, which were approved by the respective regulatory boards at both sites.
Clinical data were obtained at the same clinic or research visit as serum sample collection. Data included age at sample collection, CS use at the time of sample collection, and performance on tests of function. CS information was categorized by use (treated or untreated), type (deflazacort, prednisone, or other), and regimen (daily or intermittent – defined as 10 days on/10 days off or weekend dosing). Motor function tests included the North Star Ambulatory Assessment (NSAA), 10-m run/walk (10MRW) velocity, 6 min walk test (6MWT), and Performance of the Upper Limb 2.0 (PUL)43,44,45,46. Three disease milestones were recorded: age at loss of ambulation (LoA), age at loss of overhead reach (OHR), and age at loss of hand to mouth (HTM). LoA was defined as patient-reported inability to walk 5 m unaided at home in the LUMC cohort and inability to walk 10 m unaided within 45 s in the UF cohort. Age at loss of OHR and HTM were primarily derived from PUL scores and occasionally from patient-reported data. Cohort characteristics were described using mean and standard deviation (SD). Kaplan–Meier analysis was used to describe age at LoA, OHR, and HTM.
Sample collection and proteomic analysis
Serum samples were collected according to standard phlebotomy procedures, left to clot for ~30 min, and centrifuged (2350 g for 10 min at LUMC and 1000 g for 15 min at UF). Samples were then ultimately frozen at −80 °C for long term store. 150 µL aliquots were simultaneously shipped from both centers to SomaLogic (Boulder, Colorado, USA) and analysed using the SomaScan® proteomic platform as a single batch to avoid batch effects. SomaScan is a high-throughput proteomics platform that uses SOMAmers (Slow Off-rate Modified Aptamers), which are biotinylated DNA molecules designed to bind specifically to target proteins47. The SOMAmers were detected on a microarray, with fluorescence intensity (reported as relative fluorescence units- RFUs) correlating to the quantity of target protein present. This allows for the simultaneous analysis of thousands of proteins with high sensitivity and precision. The 7 K SomaScan® platform used in this study included 7596 aptamers that detect 6628 proteins. SomaLogic’s extensive quality control metrics have been previously published48 and include pre-assay quality control, hybridization normalization, interpolate median signal normalization, plate scaling, calibration, and adaptive normalization. Finally, pooled matrix-matched samples are run alongside clinical samples to quantify the quality of each assay. A total of 9 samples (2 LUMC/7 UF) did not pass SomaLogic’s quality control standards and were excluded from further analysis, results, and data tables.
Identification of protein probes associated with motor function tests
As the collected data were longitudinal, linear mixed effects models (LMEMs) were constructed using the pymer4 package, version 0.7.8, a Python interface for the lme4 R package, version 1.1-3149,50. For each site and probe, as a baseline LMEM, the relationship between probe expression values and age were modeled, grouping samples by patient. Probe expression values were loge-transformed and standardized, and ages were standardized. The strength and direction of the associations were represented by the fixed effects coefficients, with their significance indicated by the p value. Coefficients represent the change in transformed protein levels per unit change in standardized age. Next, to determine the effects of CS use, an LMEM was conducted with CS statuses (yes/no) as an additional covariate. Coefficients for CS use represent the change in transformed protein levels with CS use.
For each site, motor function test, and protein, an LMEM was then used to predict scores from loge-transformed, standardized probe expression values, again grouping samples by patient. Standardized ages and CS statuses were included. Coefficients represent the change in clinical scale per unit change in transformed protein levels. To determine the generalizability of this model, the resulting LMEM was used to predict scores of that motor function tests for the other site. That is, for LUMC models, UF-predicted values were generated using UF data, and for UF models, LUMC-predicted values were generated using LUMC data. Predicted values were compared against original values via reconstruction accuracy:
Significant probes whose validation Q2 values were within the top 5% of Q2 values (i.e., at or above the 95th percentile) were retained to predict clinical milestones. To account for multiple hypothesis testing, Benjamini-Hochberg false discovery rates (FDRs) were computed in all analyses51. FDR < 0.05 was considered to be significant.
Identification of protein probes associated with clinical milestones
For each site and clinical milestone, candidate probes for downstream analyses were selected as follows: for LoA, probes associated with NSAA, 10MRV, and/or 6MWT were considered. For loss of HTM and OHR, probes associated with PUL 2.0 were considered. Then, for each candidate probe, a Cox proportional hazards model was constructed, using the lifelines Python package, version 0.2752, to predict that milestone using loge-transformed, standardized probe expression values and CS statuses as covariates. Age was used as the time component and samples were grouped by patient.
P values were computed using log-rank tests. All p values underwent FDR adjustments to account for multiple hypothesis testing. FDR < 0.05 was considered to be significant.
Changes in risk of achieving milestones
To compute changes in risk of achieving milestones (Table 2), for a single probe, site, and milestone, the loge hazard ratio (lnHR), ln H, was extracted from the Cox proportional model above. A coefficient for CS status, βCS, and a coefficient for age, βage, were obtained from the associated LMEM predicting probe expression from CS status and age. Changes in risk were computed as follows:
Pathway analysis
Pathway analysis was performed using the enrichR package in R53.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Raw data cannot be deposited in a public or restricted-access repository, even after anonymization, due to privacy restrictions imposed by local data privacy officers. Access can be obtained by qualified investigators by applying to the local data privacy officers through the corresponding authors Dr. P. Spitali (p.spitali@lumc.nl) and Prof. G. Walter (glennw@ufl.edu). Access to the data will be granted after the application has been reviewed and approved by the institutional review board and privacy officers of the Leiden University Medical Center and University of Florida. A response can be expected within a few weeks. Source data are provided with this paper.
Code availability
The code supporting this study is available on Zenodo at https://zenodo.org/records/14968701.
References
Mercuri, E. & Muntoni, F. Muscular dystrophies. Lancet 381, 845–860 (2013).
Duan, D. et al. Duchenne muscular dystrophy. Nat. Rev. Dis. Prim. 7, 13 (2021).
Peverelli, L. et al. Histologic muscular history in steroid-treated and untreated patients with Duchenne dystrophy. Neurology 85, 1886–1893 (2015).
Bello, L. et al. Prednisone/prednisolone and deflazacort regimens in the CINRG Duchenne natural history study. Neurology 85, 1048–1055 (2015).
McDonald, C. M. et al. Long-term effects of glucocorticoids on function, quality of life, and survival in patients with Duchenne muscular dystrophy: a prospective cohort study. Lancet 391, 451–461 (2018).
Birnkrant, D. J. et al. Diagnosis and management of Duchenne muscular dystrophy, part 2: respiratory, cardiac, bone health, and orthopaedic management. Lancet Neurol. 17, 347–361 (2018).
Ricotti, V. et al. Long-term benefits and adverse effects of intermittent versus daily glucocorticoids in boys with Duchenne muscular dystrophy. J. Neurol. Neurosurg. Psychiatry 84, 698–705 (2013).
Conklin, L. S. et al. Serum biomarkers of glucocorticoid response and safety in anti-neutrophil cytoplasmic antibody-associated vasculitis and juvenile dermatomyositis. Steroids 140, 159–166 (2018).
Hoy, S. M. Delandistrogene Moxeparvovec: first approval. Drugs 83, 1323–1329 (2023).
Keam, S. J. & Keam, S. J. Vamorolone: first approval. Drugs 84, 111–117 (2023).
Lamb, Y. N. Givinostat: first approval. Drugs 84, 849–856 (2024).
Aartsma-Rus, A. The future of exon skipping for Duchenne muscular dystrophy. Hum. Gene Ther. 34, 372–378 (2023).
Szigyarto, C. A. & Spitali, P. Biomarkers of Duchenne muscular dystrophy: current findings. Degener. Neurol. Neuromuscul. Dis. 8, 1–13 (2018).
Molinaro, M., Torrente, Y., Villa, C. & Farini, A. Advancing biomarker discovery and therapeutic targets in duchenne muscular dystrophy: a comprehensive review. Int. J. Mol. Sci. 25, 631 (2024).
Hathout, Y. et al. Disease-specific and glucocorticoid-responsive serum biomarkers for Duchenne muscular dystrophy. Sci. Rep. 9, 12167 (2019).
Wagner, K. R. et al. Safety and disease monitoring biomarkers in Duchenne muscular dystrophy: results from a phase II Trial. Biomark. Med. 15, 1389–1396 (2021).
Zygmunt, A. M. et al. A longitudinal study of creatine kinase and creatinine levels in Duchenne muscular dystrophy. Muscle Nerve 67, 138–145 (2023).
Awano, H. et al. Longitudinal data of serum creatine kinase levels and motor, pulmonary, and cardiac functions in 337 patients with Duchenne muscular dystrophy. Muscle Nerve 69, 604–612 (2024).
Boehler, J. F. et al. Clinical potential of microdystrophin as a surrogate endpoint. Neuromuscul. Disord. 33, 40–49 (2023).
Chamberlain, J. S. et al. Microdystrophin expression as a surrogate endpoint for Duchenne muscular dystrophy clinical trials. Hum. Gene Ther. 34, 404–415 (2023).
Spitali, P. et al. Tracking disease progression non-invasively in Duchenne and Becker muscular dystrophies. J. Cachexia. Sarcopenia Muscle 9, 715–726 (2018).
Strandberg, K. et al. Blood-derived biomarkers correlate with clinical progression in Duchenne muscular dystrophy. J. Neuromuscul. Dis. 7, 231–246 (2020).
Hathout, Y. et al. Large-scale serum protein biomarker discovery in Duchenne muscular dystrophy. Proc. Natl. Acad. Sci. USA 112, 7153–7158 (2015).
Hathout, Y. et al. Serum pharmacodynamic biomarkers for chronic corticosteroid treatment of children. Sci. Rep. 6, 31727 (2016).
Willis, A. B. et al. Serum protein and imaging biomarkers after intermittent steroid treatment in muscular dystrophy. Sci. Rep. 14, 28745 (2024).
Milad, N. et al. Increased plasma lipid levels exacerbate muscle pathology in the mdx mouse model of Duchenne muscular dystrophy. Skelet. Muscle 7, 19 (2017).
White, Z. et al. Cholesterol absorption blocker ezetimibe prevents muscle wasting in severe dysferlin-deficient and mdx mice. J. Cachexia. Sarcopenia Muscle 13, 544–560 (2022).
Do Carmo et al. RGMa can induce skeletal muscle cell hyperplasia via association with neogenin signalling pathway. In Vitro Cell. Dev. Biol. Animal 57, 415–427 (2021).
Copola, A. G. L. et al. Transcriptomic characterization of the molecular mechanisms induced by RGMa during skeletal muscle nuclei accretion and hypertrophy. BMC Genom. 23, 188 (2022).
Martins, A. F. et al. Repulsive guidance molecules a, b and c are skeletal muscle proteins, and repulsive guidance molecule a promotes cellular hypertrophy and is necessary for myotube fusion. Cells Tissues Organs 200, 326–338 (2015).
Tebbenkamp, A. T. N. et al. Protein biomarker signature in patients with spinal and bulbar muscular atrophy. JCI Insight 9, e176383 (2024).
Bürgi, J. et al. CMG2/ANTXR2 regulates extracellular collagen VI which accumulates in hyaline fibromatosis syndrome. Nat. Commun. 8, 15861 (2017).
Abbassi-Daloii, T. et al. A transcriptome atlas of leg muscles from healthy human volunteers reveals molecular and cellular signatures associated with muscle location. eLife 12, e80500 (2023).
Li, Y., Xu, Z., Li, H., Xiong, Y. & Zuo, B. Differential transcriptional analysis between red and white skeletal muscle of Chinese Meishan pigs. Int. J. Biol. Sci. 6, 350–360 (2010).
Li, X. et al. Integrated 4D analysis of intramuscular fat deposition: quantitative proteomic and transcriptomic studies in Wannanhua Pig Longissimus Dorsi Muscle. Animals14, 167 (2024).
Irifuku, T. et al. Inhibition of H3K9 histone methyltransferase G9a attenuates renal fibrosis and retains klotho expression. Kidney Int. 89, 147–157 (2016).
Xiao, Z. et al. MicroRNA-205-5p plays a suppressive role in the high-fat diet-induced atrial fibrosis through regulation of the EHMT2/IGFBP3 axis. Genes Nutr. 17, 11 (2022).
Papait, R. et al. Histone Methyltransferase G9a is required for cardiomyocyte homeostasis and hypertrophy. Circulation 136, 1233–1246 (2017).
Xue, W. et al. Histone methyltransferase G9a modulates hepatic insulin signaling via regulating HMGA1. Biochim. et. Biophys. Acta (BBA) - Mol. Basis Dis. 1864, 338–346 (2018).
Zhang, L. et al. Expression and functional analyses of Dlk1 in muscle stem cells and mesenchymal progenitors during muscle regeneration. Int. J. Mol. Sci. 20, 3269 (2019).
Andersen, D. C. et al. Characterization of DLK1+ cells emerging during skeletal muscle remodeling in response to myositis, myopathies, and acute injury. Stem Cells 27, 898–908 (2009).
Garcia, S. M. et al. Distinct human stem cell subpopulations drive adipogenesis and fibrosis in musculoskeletal injury. bioRxiv: https://doi.org/10.1101/2023.07.28.551038 (2023).
Scott, E. et al. Development of a functional assessment scale for ambulatory boys with Duchenne muscular dystrophy. Physiother. Res. Int. 17, 101–109 (2012).
Arora, H. et al. Longitudinal timed function tests in Duchenne muscular dystrophy: imagingDMD cohort natural history. Muscle Nerve 58, 631–638 (2018).
McDonald, C. M. et al. The 6-minute walk test and other clinical endpoints in duchenne muscular dystrophy: reliability, concurrent validity, and minimal clinically important differences from a multicenter study. Muscle Nerve 48, 357–368 (2013).
Pane, M. et al. Upper limb function in Duchenne muscular dystrophy: 24 month longitudinal data. PLoS ONE 13, e0199223 (2018).
Gold, L. et al. Aptamer-based multiplexed proteomic technology for biomarker discovery. PLOS ONE 5, e15004 (2010).
Candia, J. et al. Assessment of variability in the plasma 7k SomaScan proteomics assay. Sci. Rep. 12, 17147 (2022).
Jolly, E. Pymer4: Connecting R and Python for linear mixed modeling. J. Open Source Softw. 3, 862 (2018).
Bate, D., Mächler, M., Bolker, B. & Walker, S. Fitting linear mixed-effects models using lme4. J. Stat. Softw. 67,1–48 (2015).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B: Stat. Methodol. 57, 289–300 (1995).
Davidson-Pilon, C. Lifelines: survival analysis in Python. J. Open Sour. Softw. 4, 1317 (2019).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Acknowledgements
This study was funded by Parent Project Muscular Dystrophy through the Protein Mapping Project. Research reported in this publication was also partially supported by Spieren voor Spieren (grant number Svs15) and the National Institute of Neurological Disorders And Stroke of the National Institutes of Health under Award Number # R61NS119639 (Co-I: Spitali), R01 AR056973 (PI: K Vandenborne), and R01 AR065943 (PI: GA Walter). Research reported in this publication was also supported by the CTSI Biorepository at the University of Florida Clinical and Translational Science Institute, which is supported in part by the NIH National Center for Advancing Translational Sciences under award number UL1 TR001427. The authors thank the staff and participants for their contributions. Several of the authors are members of the European Reference Network for Rare Neuromuscular Diseases (ERN EURO-NMD).
Author information
Authors and Affiliations
Contributions
N.A.I. contributed to data acquisition, curation, and analysis and drafting of the manuscript; A.M.B contributed to data acquisition, curation, and analysis and drafting of the manuscript; S.W.M.E. contributed to data cleaning, statistical approach development, and conduct of statistical analyses; S.H.V contributed to data cleaning and statistical analysis; K.C.H.H. contributed to the statistical approach and analyses; H.E.K. contributed to data curation and revision of the manuscript for intellectual content; K.V. contributed to study conceptualization, data acquisition, and revision of the manuscript for intellectual content; E.H.N. contributed to study conceptualization, data acquisition, and revision of the manuscript for intellectual content; G.A.W contributed to study conceptualization, data interpretation, and drafting of the manuscript; P.S. contributed to study conceptualization, statistical approach, data analysis and interpretation, and drafting of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
S.W.M.E., S.V.H. and K.CH.H. are employees of BioSymetrics, which has a commercial interest in the results. The remaining authors have no competing interests.
Peer review
Peer review information
Nature Communications thanks Thurman Wheeler, who co-reviewed with Preeti Kumari; Eric (P) Hoffman and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ikelaar, N.A., Barnard, A.M., Eng, S. et al. Large-scale serum protein biomarkers discovery associated with function and clinical milestones in Duchenne muscular dystrophy. Nat Commun 16, 9073 (2025). https://doi.org/10.1038/s41467-025-64146-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-64146-y