Abstract
Understanding species-level abundance dynamics in complex microbial communities is key to managing microbial ecosystems, yet it remains a major challenge. In wastewater treatment plants (WWTPs), the presence and abundance of process-critical bacteria are essential for removing or recycling pollutants. However, individual species can fluctuate without recurring patterns. Accurately forecasting these dynamics is critical for preventing failures and guiding process optimization. We have developed a graph neural network-based model that uses only historical relative abundance data to predict future dynamics. Each model is trained and tested on individual time-series from 24 full-scale Danish WWTPs (4709 samples collected over 3–8 years, 2–5 times per month). It accurately predicts species dynamics up to 10 time points ahead (2–4 months), sometimes up to 20 (8 months). The approach, implemented as the “mc-prediction” workflow, is also tested on other datasets, including a human gut microbiome, showing its suitability for any longitudinal microbial dataset.
Similar content being viewed by others
Introduction
Complex microbial communities are ubiquitous on the planet, playing an essential role across all natural ecosystems1 and engineered biotechnological systems. The most widespread engineered ecosystem is biological wastewater treatment2, which removes organic and inorganic pollutants to avoid eutrophication and pollution of receiving waters. Simultaneously, resources and energy are recovered to ensure sustainability3. The structure of microbial communities in wastewater treatment plants (WWTPs) influences treatment performance4, highlighting the importance of continuous monitoring, which is now possible on-site within few hours5. The main factors involved in shaping the structure of microbial communities have been studied extensively over the years6,7, with a general consensus that both stochastic factors (such as immigration) and deterministic factors (such as temperature, nutrients, and predation) have a significant influence8,9,10,11, however, their relative contributions can vary. The interplay among these factors adds a significant challenge to understanding WWTP ecosystems. Although the microbial community structure can be characterized in detail by DNA-based methods, it has not yet been possible to develop accurate models to predict temporal microbial dynamics in full-scale WWTPs, or other ecosystem. The ability to predict the community composition in the near future would be very valuable to plant operators, allowing them to circumvent emerging problems in time, optimize plant operation and performance, and gain insights into the microbial community dynamics.
Temporal dynamics of individual community members can vary greatly over time in any aquatic ecosystem12,13,14,15, which limits the usefulness of community snapshots from only one or a few points in time. Seasonal variation and recurring patterns in microbial community dynamics have long been documented in lakes16 and in activated sludge (AS) WWTPs, where species-specific patterns are often observed. For example, different temporal patterns can be observed even among different species within the same genus, such as filamentous Candidatus Microthrix15.This process-critical filamentous bacteria causes poor settling properties of AS flocs resulting in poor performance, which is the most widespread operational problem in WWTPs globally10,17,18. Community structure can, to some extent, be controlled by process design, addition of specific chemicals, or operational changes. However, in most cases, a reliable cause-effect relationship has not been established.
Instead of relying on established cause-effect relationships, prediction may be built on the advances in computational methods such as machine- and deep learning models based on artificial neural networks (ANN). Combined with increasing computational performance, these methods provide unexploited opportunities compared to traditional approaches. Recent successful examples within the field of microbial ecology are predictions of microbial interactions19, the functional potential of a metagenome20, the future community structure21,22, or transient short-term dynamics23,24. Despite these efforts, no study has successfully predicted the dynamics of all individual members of the microbial community across multiple future time points (e.g., weeks or months). Only one study of a marine area from the Western English Channel predicted the bacterial community assemblages as a function of environmental parameters25 by extrapolating the bacterial abundances. However, it was only performed with low taxonomic resolution (order level), thus only giving a very rough insight into the temporal dynamics.
Our goal was to implement a graph neural network-based machine learning approach for accurately predicting the future dynamics of individual microorganisms in complex microbial communities at the highest possible resolution26 (amplicon sequence variant, ASV level). We aimed to develop a model based only on historical relative community composition over time, since consistent, reliable and detailed environmental parameters can be difficult or impossible to obtain for many ecosystems, including WWTPs. Additionally, the limited understanding of other abiotic or biotic interactions, e.g., microorganisms’ growth rates or predation, makes it challenging to include mechanistic components. As a case study, we used the activated sludge ecosystem, using our comprehensive microbial dataset from 24 different full-scale WWTPs across Denmark comprising 4709 samples collected over 3–8 years at 2–5 time points per month. This extensive time series captures both operational fluctuations and seasonal variations15, without incorporating environmental variables.
Our approach differs from previous studies modeling community structure21 or prediction of community structure temporal dynamics27. We chose to use a graph neural network-based approach specifically designed for multivariate time series forecasting that considers relational dependencies between the individual variables, which is well-suited for prediction of complex microbial community dynamics28. Since each WWTP is unique in terms of microbial community structure, influent wastewater, design, and operation, developing a universal predictive model for the entire activated sludge ecosystem is not feasible. Therefore, models were trained and tested independently for each site. However, the graph neural network-based approach is generic, and we showed its applicability to other microbial ecosystems, including human gut microbiota. Finally, we developed a software workflow “mc-prediction” for community prediction in any ecosystem, publicly available at https://github.com/kasperskytte/mc-prediction29, following best practices for scientific computing30.
Results
The microbial community structure of the 4709 samples collected from 24 full-scale WWTPs with nutrient removal was obtained using 16S rRNA amplicon sequencing, and ASVs were classified using the MiDAS 4 ecosystem-specific taxonomic database31 to provide high-resolution classification at species level. The top 200 most abundant ASVs (corresponding to around 125 species, all classified) in each dataset were selected (out of a total of 76,555 unique ASVs across all datasets), amounting to 52–65% of all DNA sequence reads per dataset, representing more than half of the biomass in the plants. The remaining reads were primarily representing very low abundant taxa and numerous singletons. The sampling intervals varied between the datasets, but in most cases, they were between 7–14 days (Figure S1). Although the sampling interval in the datasets should preferably be consistent, in practice this was difficult to obtain in long-term longitudinal experiments that span many years. The number of samples in each dataset varied significantly between 92 and 344, where few WWTPs had periods with no sampling, which could impact the prediction accuracy (Fig. S2). For the model training, we made a chronological 3-way split of each dataset into training, validation, and test datasets, where the latter was used to evaluate the prediction accuracy compared to true historical data (Fig. 1a).
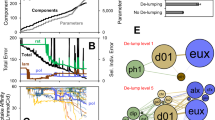
a The datasets are divided chronologically into 3 parts, where the first part is the training dataset, the second is the validation dataset, and the last is the test dataset. The train and validation datasets are used to train and optimize each neural network model until the value calculated by the loss function no longer improves. The final model is then tested on a separate test dataset and a numeric error is calculated between the real values and the predicted values. b Schematic of the overall model architecture. The historical values for each Amplicon Sequence Variant (ASV) (1) are first clustered by a few different methods (2): a graph neural network is used to infer putative interactions between ASVs and cluster them accordingly. Additionally, clustering by known biological functions, Improved Deep Embedded Clustering (IDEC), and ranked abundances were also tested. Then, a temporal convolution network is trained separately on each cluster to extract temporal features (3). Finally, a chosen number of predicted consecutive sample points are obtained through the output layer (4).
To optimize the prediction accuracy of the model, four different pre-clustering methods were tested, followed by a graph neural network model designed and trained on each cluster. The cluster size was set to 5 ASVs for all methods except IDEC, which decided for itself (see next section). The prediction accuracy and the predicted relative abundances for each ASV were then obtained. Briefly summarized, the graph model design consisted of a few different layers. First, the graph convolution layer learns the interaction strengths and extracts interaction features among ASVs (Fig. 1b). Then, the temporal convolution layer extracts the temporal features across time, and lastly, the output layer with fully connected neural networks uses all features to predict the relative abundances of each ASV. Moving windows of 10 historical consecutive samples from each multivariate cluster of 5 ASVs are the inputs to the graph models, and the 10 future consecutive samples after each window are the outputs. This is iterated throughout the train, validation, and test datasets for each of the 24 datasets.
General prediction accuracy and the effect of pre-clustering ASVs
To maximize the prediction accuracy, we chose to test and evaluate the effect of pre-clustering before model training using four different methods. One method was to cluster into biological functions. Where known, the ASVs were pre-clustered into 5 important biological functions in wastewater treatment at genus level according to the MiDAS Field Guide32: polyphosphate accumulating organisms (PAOs), glycogen accumulating organisms (GAOs), filamentous bacteria, ammonia oxidizing bacteria (AOB), and nitrite oxidizing bacteria (NOB). Additionally, we tested pre-clustering the ASVs using the Improved Deep Embedded Clustering (IDEC) algorithm33 pre-clustering by using the time-varying graphical clustering algorithm on the graph network interaction strengths from the graph neural network model itself34 (from here on referred to as “graph”, see Figure S3 for a few examples of proposed interaction strengths), and lastly using clusters by ranked abundances from the top in groups of 5 ASVs.
The prediction accuracy of each model was evaluated using 3 different metrics (Bray-Curtis, mean absolute error, and mean squared error) for each cluster type and for each WWTP test dataset. Only the Bray-Curtis metric is shown in boxplots in Fig. 2, because the other two metrics showed the same overall trends (Fig. S4). The graph models were trained on moving windows of 10 consecutive samples at a time and predicted 10 consecutive time points into the future after each window. Depending on the sampling interval of the datasets (Fig. S1), this corresponds to 2.0–3.5 months into the future.
The values are calculated using the Bray-Curtis dissimilarity measure (0.0 means identical and 1.0 completely different) of each cluster (group of ASVs). The different clustering types are indicated by colors according to the legend. The x-axis has been square-root transformed to better reveal differences at the lower values. Data are presented as median, 25th–75th percentiles, and whiskers extending to 1.5×IQR; outliers shown as points. The datasets are ordered by the number of samples in each dataset, which is indicated in parentheses. Source data is provided as a Source Data Fig. 2.
Overall, the models showed a good to very good prediction accuracy as later illustrated in Figs. 3 and 4. The best prediction accuracies across all datasets were generally obtained when the model was trained on clusters (of 5 ASVs) defined by either the graph network interaction strengths or by ranked abundances. However, there was not a large difference when assessing median accuracies. Notably, it was clear that clustering by biological function generally resulted in a lower prediction accuracy compared to the other clustering methods, except for the Ejby Mølle and Hirtshals datasets (Fig. 2). IDEC clustering enabled some of the highest possible accuracies, but it produced a larger spread in the prediction accuracy between the individual clusters for many datasets. Graph pre-clustering based on network interaction strengths was chosen for the remaining results because it achieved the best overall accuracy.
Left column: The real relative abundances are indicated by the black points alongside the model predictions from both the train, validation, and test datasets, indicated by red, yellow, and green points, respectively. The model training and evaluation was done in windows of 10 samples predicting 10 samples into the future. The models used here have been trained on graph clusters. Right column: observed (real historic) values plotted against the predicted values with two different linear regression lines added: A linear regression between the observed and predicted relative abundance values (blue), and a 1.0-slope line (black) to represent the theoretical perfect prediction. The R2 values of each model are indicated in the top left corner, calculated based on the residuals of the linear regression model fitted to the data as well as the 1.0-slope line, respectively. Source data is provided as a Source Data Fig. 3.
Both the real and predicted abundances are shown in pairs for each time point and the genus and species level classifications of each Amplicon Sequence Variant (ASV) separated by semicolon are indicated on the y-axis. The relative abundances are indicated by a color gradient (blue through yellow to red, low to high). The model was trained on graph clusters. Source data is provided as a Source Data Fig. 4.
The prediction accuracy varied significantly between the individual clusters. At first glance, there was no apparent trend between the number of samples in the datasets and their median prediction accuracy, as it could have been expected due to more training data with more samples (R-squared value for a linear model between the two was only 0.008406). To further investigate how the number of samples affects prediction accuracy, we selected the longest dataset, Aalborg W, and separated it into 3 smaller subsets containing every second, third, and fourth sample and processed each dataset separately. A clear trend was observed with better overall prediction accuracy when the number of samples increased (Fig. S5). The 4 different Damhusåen datasets were from 4 parallel treatment lines, all with the same influent wastewater and operation, and the accuracy between these was very similar, as expected. While it is difficult to describe the prediction accuracy by a single metric value, the primary goal of model training is to minimize a single metric value calculated by a chosen loss function. For this study, we chose the Bray-Curtis dissimilarity measure. The lowest median value obtained across all clusters was 0.147 when pre-clustering by graph network interaction strengths. However, this single value is not sufficient to express the overall prediction accuracy of all the models because there is significant variation within each cluster. Therefore, it is important to further inspect each time series prediction for each ASV individually, as demonstrated later.
Predicting the dynamics of key bacteria in wastewater treatment
We evaluated the prediction accuracy of 4800 individual time series predictions (200 ASVs * 24 datasets). There were in total 1182 different ASVs in the 24 WWTPs belonging to the top 200 in each plant. Of these, 188 belonged to one of the defined functional groups. When calculated individually for each ASV, the median of the mean absolute percentage error (MAPE) between the real and predicted values on the test datasets ranged between 14.2% and 17.7%. This means that, if the real relative abundance of an ASV was 10.0% and the MAPE value was 15.0%, the predicted relative abundance would be between 8.5–11.5%. To further evaluate the prediction accuracy for each individual ASV, R21:1 values of each model were calculated based on the residuals of the linear regression model fitted to the data and compared to 1.0-slope line (see Fig. 3, right). In general, we observed that models for the most abundant ASVs were the most accurate, except for a few cases. Importantly, the prediction accuracy of ASVs within process-critical taxa was consistently high across all datasets (Fig. S6). ASVs with low abundance or those that were not observed in significant quantities across the entire dataset had the lowest accuracy models.
To illustrate these findings, we investigated in detail some process-critical bacteria, using Mariagerfjord WWTP dataset as an example (Fig. 3). We selected 3 relevant taxa with distinct ecological functions and abundance patterns, Ca. Microthrix parvicella (ASV2), a common problematic filamentous species35, Tetrasphaera (ASV1), the most abundant PAO in WWTPs globally31,36,37, and Nitrospira defluvii (ASV24), the most prevalent NOB worldwide31,38,39,40.
Proper settling of activated sludge flocs ensures an effective solid-liquid separation in the final clarifier, which is essential for the overall performance of WWTPs. Some filamentous bacteria directly impair settling properties or can produce unwanted foam on the surface, which can reduce the overall plant performance and effluent water quality. Key filamentous genera in AS include Ca. Microthrix, Ca. Amarolinea, Gordonia, Ca. Villigracilis, and Ca. Sarcinithrix. Some of these filaments contribute to floc formation, but others cause settling problems. Thus, a reliable prediction of the near future abundance profile of these species is of great importance for plant operators to be prepared and minimize operational problems. For example, Ca. Microthrix is the most abundant and troublesome filamentous genus in many Danish WWTPs during the winter and early spring, and its relative abundance is highly correlated with settling problems35. The prediction accuracy for Ca. Microthrix parvicella (ASV2) from Mariagerfjord performed well on the test dataset (Fig. 3). Noticeably, there was a clear recurring seasonal pattern for ASV2 in the training data, which the model captured well, but the magnitude of the recurring pattern was almost two-fold in the test dataset compared to the training set. Regardless, the model was still able to accurately predict the dynamics using relative abundances as the only input variable.
Another important functional group in full-scale WWTPs with enhanced removal of phosphate is PAOs, in particular the genus Tetrasphaera. In contrast to the temporal dynamics of ASV2 (Ca. Microthrix parvicella), there was no apparent recurring seasonal pattern for ASV1 (Tetrasphaera). However, large relative abundance variations between 1-10% were observed. Nevertheless, the model was still able to accurately predict these fluctuations (Fig. 3). Lastly, the abundance dynamics of one of the NOBs, ASV24 (Nitrospira defluvii) were captured well even though it had lower abundance in the training set compared to the test set. Although the prediction was lower than in the other two examples, a good prediction accuracy was obtained (Fig. 3)
The microbial community composition of Danish WWTPs was relatively stable over the years for the top-most abundant ASVs, as exemplified with Mariagerfjord dataset (Fig. 4). However, there were clear differences in the stability of the different ASVs. By interpreting the numerical differences alone, it is clear that the models seem to capture the overall trends for most ASVs, but a few in the top 30 were consistently inaccurate to varying degrees, such as ASV305, ASV2974, and ASV9. Similar observations were made for the other 23 WWTP.
Predicting the dynamics of entire microbial communities
To evaluate how the approach can be used to predict the dynamics of the entire microbial communities, we combined the predicted values from the individually trained models for the 200 most abundant ASVs for each WWTP. Principal Coordinate Analysis (PCoA) based on the Bray-Curtis dissimilarity measure was used to visualize the relative differences between the real and predicted microbial communities based on the distances between the sample points (Fig. 5). There was a noticeable difference between the real and predicted communities, and in most cases the predicted samples did not overlap exactly with the real samples. The combined effects of inaccurate predictions for some ASVs impacted the calculated dissimilarities between the real and the predicted samples even though most ASVs showed accurate predictions. Therefore, predicting entire communities accurately requires that every individual model performs well, hence one must instead select the ASVs of interest individually.
The distance measure used is the Bray-Curtis dissimilarity measure. The sample points are colored according to which part they belong (training = red, validation = yellow, testing = green) alongside the true samples (black). The plot area has been scaled symmetrically with an aspect ratio of 1. The predicted community profiles are based on models trained on graph clusters and combined into a single multivariate dataset. Source data is provided as a Source Data Fig. 5.
PCoA allows to visualize immediately the relative differences between the real and predicted communities. The two principal components captured 46.6% of the variance, which is a significant portion, but does not show the complete picture. To test whether there was a significant difference between the real and predicted communities, we used the analysis of similarities (ANOSIM) test on the test datasets with the Mariagerfjord as an example. The test was based on the Bray-Curtis dissimilarity measure and the R statistic was 0.0102, which means the difference was quite small numerically. The mean R statistic for all datasets was 0.057.
The effect of prediction window lengths on prediction accuracy
The predictions presented so far have been performed with a prediction window of 10 samples into the future for all datasets. To evaluate the effect of different prediction window lengths on prediction accuracies and how far into the future it is possible to predict without sacrificing too much accuracy, we tested prediction window lengths of 3, 5, 10, 15, and 20 samples into the future. These windows represent chronologically ordered samples rather than exact days, as the temporal resolution depends on the sampling frequency of each dataset. Considering a sampling frequency of 7 days, this corresponds to predictions ranging from 21 days (3 samples) to 140 days (20 samples) into the future. We also tested whether the inclusion of the sampling intervals in days as an extra input variable to the model would improve the prediction accuracy, but the results were very similar or slightly worse (Fig. S7). When including time intervals between samples we achieved a median Bray Curtis dissimilarity of 0.151, compared to 0.147 without. As expected, a longer prediction window resulted in lower overall prediction accuracies (Fig. 6), however, some clusters were still accurate enough to be useful even with 20 samples predicted.
The values are calculated using the Bray-Curtis dissimilarity measure (0.0 means identical and 1.0 completely different) of each cluster (group of 5 ASVs). The y-axis has been square-root transformed to better reveal differences at the lower values. Data are presented as median, 25th–75th percentiles, and whiskers extending to 1.5×IQR; outliers shown as points. The models were trained on graph clusters from the Mariagerfjord dataset. Source data is provided as a Source Data Fig. 6.
The predictions for some ASVs remained very accurate even with 20 predicted sample points, such as ASV2 (Ca. M. parvicella) from Mariagerfjord (Fig. 7). On the contrary, the prediction accuracy was low for some ASVs, for example ASV24 (N. defluvii) (Fig. S8). The fact that prediction accuracies vary more, and generally decrease, with more predicted samples again highlights the importance of always inspecting each prediction individually and, in practice, hand picking relevant ASVs for a particular WWTP. Generally, the dynamics of many of the most abundant and thus, presumably, the most important ASVs were captured well, while the low-abundant ASVs were less accurately predicted.
Left column: The real relative abundances are indicated by the black points alongside the model predictions from both the training (red), validation (blue), and test (green) datasets. The model training and evaluation was done in windows of 10 samples predicting varying number samples into the future. The models used here have been trained on graph network clusters from the Mariagerfjord dataset. Right column: observed (real historic) values plotted against the predicted values with two different linear regression lines added: A linear regression between the observed and predicted relative abundance values (blue), and a 1.0-slope line (black) to represent the theoretical perfect prediction. The R21:1 values of each model are indicated in the top left corner, calculated based on the residuals of the linear regression model fitted to the data as well as the 1.0-slope line, respectively. Source data is provided as a Source Data Fig. 7.
Predicting community dynamics in other environments
To evaluate the prediction accuracy of our workflow in other ecosystems, we tested three different datasets from other studies: a one-year dataset from the human gut41, a 17-year time series of fungal environmental DNA from a coastal marine ecosystem42, and a 5-year dataset from another municipal WWTP in Hong Kong11.
The human gut dataset consisted of self-collected stool samples from a self-reported healthy male (30–35 years old) living in Boulder, Colorado, US43. The study aimed to define temporal variations of normal gut microbiota to improve clinical methods for detecting changes due to lifestyle or illness41. The dataset included 332 samples collected at an average interval of 1.33 days, with a few at longer intervals of 3–6 days and two separated by a 17-day gap for unspecified reasons. The data could only be retrieved at genus-level taxonomic resolution based on the original Greengenes database assignments. No other metadata related to diet, lifestyle, or medications was reported. Overall, their results showed that the community structure was stable over the sampling period, but dynamic variations occurred for specific taxa. As such, our workflow was able to provide good prediction accuracies for the human gut dataset, generally comparable to our 24 WWTP datasets (Bray Curtis dissimilarity 0.21-0.22, Figure S9), as well as for individual gut relevant genera44 as shown with three examples in Fig. 8, with good predictions for at least 10 points, corresponding to 2-3 weeks.
Left column: The real relative abundances are indicated by the black points alongside the model predictions from both the training (red), validation (blue), and test (green) datasets. The model training and evaluation was done in windows of 10 samples predicting varying number samples into the future. The models used here have been trained on graph network clusters from a dataset from the human gut39. Right column: observed (real historic) values plotted against the predicted values with two different linear regression lines added: A linear regression between the observed and predicted relative abundance values (blue), and a 1.0-slope line (black) to represent the theoretical perfect prediction. The R21:1 values of each model are indicated in the top left corner, calculated based on the residuals of the linear regression model fitted to the data as well as the 1.0-slope line, respectively. Source data is provided as a Source Data Fig. 8.
In contrast, the workflow proved incapable of predicting the fungal community dynamics in the coastal marine dataset, even though it was a very long and consistently sampled dataset (540 samples, average frequency 12.3 days). The main reason was that the dataset was extremely sparse and zero-inflated compared to the other two datasets and had extreme fluctuations in the relative abundances ranging from near 0% to as much as 50% within a very short time for a single microorganism (See Fig. S10). Lastly, the prediction accuracies obtained for the Hong Kong WWTP dataset were comparable to the ones obtained for our 24 WWTP datasets with a median Bray-Curtis dissimilarity between 0.21–0.22 (Fig. S9). The prediction accuracy of the community members was acceptable despite the lower number of samples (58 samples; average frequency 32 days (14 – 91 days range)), especially for the most abundant species (a few examples are shown in Fig. S11).
Discussion
We present a novel workflow “mc-prediction” for predicting the temporal dynamics of complex microbial communities at species (ASV level) using only historical relative abundance time series of all community members. Built on a graph neural network-based approach specifically designed for multivariate time-series forecasting, we could predict the dynamics of all abundant species in WWTPs ecosystem with high accuracy for 10-15 future time points (corresponding to 2-3 months ahead). Some species could be predicted 4–6 months ahead when trained on 3–8 years high-resolution datasets, without requiring additional metadata. Due to the generalizability of the graph neural network-based approach, we show that the workflow can be applied to any microbial ecosystem if good quality, high-resolution time series of microbial abundance data are available. For example, mc-prediction successfully predicted individual genus dynamics in the human gut with good accuracy up to 10 future time points (~2–3 weeks). This is a significant advancement for the surveillance and management of microbial communities in both engineered and natural ecosystems, where the prediction of individual species dynamics can improve the operational performance in engineered ecosystems or support health monitoring in clinical contexts.
Predicting the future abundance dynamics of the individual microbes in complex communities in nature or engineered systems has remained a main challenge, even with other computational or statistical methods than machine- or deep learning. Leveraging the 24 newly presented WWTP datasets, we developed a predictive workflow relying solely on historical microbial abundance data, further applying it to three additional published datasets (human gut41, fungal environmental DNA from a coastal marine ecosystem42, and another municipal WWTP in Hong Kong11). Instead of aiming to develop a universal model for an entire ecosystem type (e.g., all AS-WWTPs), we trained individual models for each site (e.g., each WWTP), as microbial relative abundance patterns often vary even between individual species within the same genus15,35, a pattern also observed in this study. Additionally, environmental factors (such as temperature, nutrient concentrations) or operational changes can contribute to site-specific differences. However, obtaining high-resolution, high-quality metadata is rarely possible from full-scale WWTPs, making its incorporation impractical compared to other ecological models. This limitation can be extended to other ecosystems, such as the human gut, where comprehensive metadata on health parameters, diet, and lifestyle is nearly impossible to obtain. Therefore, solely relying on historical abundance data to achieve accurate site-specific predictions is an accessible strategy to extend the workflow to broader applications. Moreover, the workflow, “mc-prediction”, (available at https://github.com/KasperSkytte/MC-prediction29) is portable, reproducible, and applicable to any other community time series dataset of good quality. It is also highly efficient and can run on a normal laptop. The processing time for each dataset was between 1-12 hours, depending on whether all or only a single clustering method was used, when using only 4 CPUs and max 8GB of memory usage.
To develop this workflow, initially we developed a neural network model that was based on the Long Short-Term Memory (LSTM) method45, which performed well when trained on a single ASV at a time. However, as complex interactions are known to exist between species in microbial communities9,46, instead we decided to implement a graph neural network approach, which generally outperformed LSTM in our experiments. The interaction strengths in the graph neural network may also represent hypothetical biological interactions between the individual ASVs in the group, which should be verified experimentally. Model design and implementation required many decisions governed by trial and error until no more significant improvements in prediction performance were obtained across the datasets. For example, decisions included setting the values of all the different hyperparameters, such as batch size, number of training epochs, iterations, the group size for each clustering method, and the sizes of the individual layers in the neural network. With hundreds of different combinations to test, we settled on a configuration that performed consistently well across all datasets, although further optimization may be needed if applied to other datasets.
In a first attempt, we aimed to predict the future abundances of all microbial community members using a single model for each WWTP, but the obtained prediction accuracy was very poor. Subsequently, we tested various clustering methods to improve prediction accuracy. Prediction accuracy was similar between models trained on clusters based on graph network interaction strengths and models trained on ranked abundance groups, suggesting that most abundant ASVs had the most influence on predicting other abundant ASVs. The lower prediction accuracy obtained when clustering by biological function (at genus level) highlights that species carrying out the same overall function in the wastewater treatment process may show very different physiological traits and dynamics and should be analyzed separately15,35.
Generally, the prediction accuracy declined as more ASVs were grouped together, with an optimal group size around 5 ASVs. Regardless of clustering type, the individual models showed varying prediction accuracies on both the training- and test datasets, where most were good or at least acceptable, but few were very poor. This limits the ability to predict the future dynamics of the entire microbial community with the current workflow, highlighting the need for manual inspections of predictions on both the training- and test datasets for each model (e.g., as illustrated in Fig. 3), and avoid predictions from models that perform poorly. Manual inspection is also important because the relative error between the observed and predicted abundances may seem numerically significant for a machine learning algorithm but is negligible in an ecological context, especially for low-abundance species with minimal impact on wastewater treatment performance. It is recommended that the most relevant species are selected beforehand, as relying on a single metric to evaluate overall prediction performance of all models is not ideal. The reasons why a few species were not easily predicted remain speculative, but across datasets we observed that species with very low abundance, where instrument and technical variations have a larger impact, and species exhibiting drastic changes, clearly exemplified by the fungal dynamics from coastal marine ecosystems40, were particularly difficult to predict.
The total number of samples in our WWTP datasets and the human gut dataset41 were proven sufficient for predicting 10 time points into the future with acceptable accuracy. However, more frequent sampling or longer time series can improve the prediction accuracy, extend the prediction window, and potentially capture short-term fluctuations better. The number of time points predicted into the future was set to 10 because the overall prediction accuracy was acceptable for most ASVs, at least the most abundant ones. In the case of WWTPs, the 10 time points provide adequate time for WWTP operators to benefit from the predictions of 2–3 months (depending on sampling interval), while considering that one week is the minimum time that it takes to sample, transport, prepare, and sequence the DNA, unless performed on-site at the WWTP. The number of predicted time points could be extended for certain ASVs where the trained model performs well or decreased to improve the accuracy at the expense of a shorter prediction window. The small variations observed in the predicted values when predicting different numbers of time points into the future are likely related to the inherent randomness and heuristics of neural networks, but generally, accuracy decreased with a larger prediction window. Surprisingly, we did not find a significant improvement in the overall prediction accuracy when including time intervals between the samples as an additional variable in the model training (Fig. S7). This may be because, overall, our datasets are consistently sampled without long gaps, so incorporating additional variables will only increase the input complexity and thus also overfitting the model. However, we have made this feature optional in the workflow, since it may be beneficial to other datasets.
In particular at wastewater treatment plants, obtaining accurate predictions of key bacteria can be of great value for management and operation. For example, severe performance problems due to bulking and poor settling of the sludge are common globally, which are primarily caused by the excessive growth of filamentous bacteria such as Ca. Microthrix35. A common solution is to add expensive chemicals such as polyaluminium chlorides (e.g., PAX-14)47 to improve solid-liquid separation, but it can take up to 3 weeks before it is effective48. Here we have demonstrated that it is now possible to accurately predict the future abundance dynamics of Ca. Microthrix as far as 20 sample points ahead (4-8 months), allowing WWTP operators to act in time and both avoid problems and reduce operating costs. Chemical dosage, other control interventions, or operational changes only result in gradual changes in community structure49. Despite these changes, our models trained on periods without such interventions they still learned from the new abundance patterns and captured the resulting effect on community dynamics (Figure S12).
Similar practical applications could be developed for other ecosystems, for example the human gut, where predicting the dynamics of key microorganisms may improve personalized health management related to imbalances in gut microbiota, such as inflammatory bowel disease and irritable bowel syndrome, obesity or type 2 diabetes41,44.
Based on our findings testing the prediction capabilities on several ecosystems, we have the following recommendations to obtain accurate predictions using the “mc-prediction” workflow: a) If the dataset is obtained through modern high-throughput DNA sequencing methods like in this study, a consistent and highly standardized method must be used to reduce technical variation and bias throughout all the steps from sampling to wetlab work, DNA sequencing, and bioinformatic processing; b) Genus-level or preferably species-level resolution is needed not to aggregate taxa that may have different dynamics; c) The dataset must contain at least 100 samples, preferably with, but not required, a consistent sampling interval between sampling timepoints; d) The dataset must be continuous, and the abundance values can show significant changes between time points, but it must not be largely zero-inflated where species are completely absent for long periods or show sudden extremely high abundances for just a few sampling points; e) If the dataset is obtained from a natural environment, it should span at least one year, preferably several years, to capture any potential seasonal variations.
Methods
Sampling
A survey of 29 Danish municipal activated sludge WWTPs was conducted between 2015 and 2023 as part of the ongoing Microbial Database of Activated Sludge (MiDAS) project and samples were collected and handled as previously described49. Protocols are available at https://midasfieldguide.org. More than 5306 samples in total have been collected from the 29 WWTPs, however only 4,709 samples from 24 different WWTPs were suited for this study, mainly because of a too short timespan for 5 datasets. All the WWTPs had biological nitrogen (N) removal and either enhanced biological phosphorus (P) removal (EBPR) or chemical P removal. Samples were taken from the aeration tanks on average every 10 days (see Figure S2 for exact sampling dates, and Figure S1 for the sampling interval distribution).
16S rRNA amplicon library preparation and sequencing
DNA extraction, sample preparation, and amplicon sequencing was done largely as described in McIIroy et al.50. In summary, DNA was extracted using the FastDNA Spin Kit for soil (MP Biomedicals) with four times repeated bead beating on a FastPrep-96 (MP Biomedicals). DNA concentrations were measured before and after PCR using a Qubit 2.0 Fluorometer (Invitrogen). The DNA libraries were barcoded, and PCR amplification was performed using the V1-V3 primer set with the following DNA sequences: 27F 5′-AGAGTTTGATCCTGGCTCAG-3′51; 534R 5′-ATTACCGCGGCTGCTGG-3′52. The amplicon libraries were purified using the Agencourt AMPure XP bead protocol (Beckmann Coulter), validated on a TapeStation 2200 system using D1000 ScreenTape (Agilent Technologies) and then pooled in equimolar concentrations. The pooled amplicon libraries were then sequenced using paired-end (2 × 301 bp) sequencing on a MiSeq (Illumina), though only the forward reads were used for analysis because merged reads did not provide any significant improvement to taxonomic classification in the V1-V3 region but instead increased the number of chimeric and spurious Amplicon Sequence Variants (ASVs/zOTUs).
Bioinformatics and amplicon data pre-processing
A complete shell script workflow was developed to process the basecalled and demultiplexed fastq files to produce an abundance table alongside taxonomic classification, available at https://github.com/kasperskytte/asv_pipeline. All the following steps were performed using usearch v.11.0.66753. Only the forward reads were used covering mostly the V1-V2 regions, and not V3. In summary, PhiX genome spike-in sequences were filtered using the filter_phix command, fastq files were quality filtered using the fastq_filter command with the fastq_maxee option set to 1.0 and the sequences were truncated to 250 bp by setting the fastq_trunclen option to 250. The sequences were then de-replicated using the fastx_uniques command, and chimeras and bad reads were filtered and ASVs identified using the UNOISE algorithm54. ASVs with less than 60% identity to any sequence in the taxonomic database were removed using the usearch_global command. An abundance table counting the individual ASVs per sample was then generated by mapping to the raw, unfiltered reads using the otutab command. Taxonomic classification was then performed using both strands with the SINTAX algorithm55 with sintax_cutoff set to 0.8. The taxonomic reference database used was the MiDAS ecosystem-specific database version 4.8.131, which is based on full-length 16S rRNA gene sequences obtained directly from the activated sludge and anaerobic digesters from treatment plants across the world and processed using the AutoTax pipeline56. This results in high coverage and species-level classification for between 66.5-76.5% of ASVs across the datasets.
Samples with less than 5000 total reads were filtered from the abundance table (though 75% of the samples had more than 25,000 reads), and the read counts were normalized by sample totals (i.e., percent relative abundance of total reads per sample). ASVs that were not present in more than 0.1% relative abundance in any sample or had near-zero abundance (0.01%) in 60% or more of the samples were removed. To reduce the size of the data only the top 200 most abundant ASVs in each dataset (each WWTP) were used. The total abundance of the most abundant ASVs made up between 52–62% of the total reads per sample (median across all samples per dataset). The remaining ASVs are believed to be less relevant to the activity in wastewater treatment processes57,58. Furthermore, the duration between sample points showed some variations across datasets. The exact dates or duration between sampling points were not used for the model training as any potential extrapolation would not be reliable, so they have just been sorted chronologically. Based on the sampling interval, the dates are used to project time points onto the predicted values.
Initial experimentation was done to find the best way to preprocess the data to optimize the prediction accuracy of the models. Standardization (subtracting the mean and dividing by standard deviation) as well as division by maximum, mean, standard deviation, and applying different smoothing factors were tested. The best preprocessing showed to be division by the mean relative abundance of each ASV across all samples and applying a smoothing factor of 4. Smoothing was done using the lfilter function from the scipy python module (v1.8.0).
Graph neural network model and pre-clustering
A complete containerized pipeline with all required software and scripts preinstalled as well as GPU support for Tensorflow was developed and is available as a GitHub package at https://github.com/KasperSkytte/MC-prediction/pkgs/container/mc-prediction. The exact release version used for this study including analysis scripts used to create figures and calculate statistics is v1.0.1. The Tensorflow 2.7.0 and the Keras API 2.7.0 modules were used extensively to implement the prediction pipeline in Python 3.8.10 as well as several other utility python modules. Refer to the GitHub repository for details and exact versions of every single package used. The abundance table, taxonomy, and sample metadata files are initially loaded using the ampvis2 R package 2.8.659 to allow a standardized format and for compatibility with the output from the most common amplicon data processing tools such as QIIME60, mothur61, DADA262, and USEARCH53. The dataset was split chronologically into 3 separate training, validation, and test datasets in a three-way split in the respective proportions 80%, 5%, and 15%. To optimize the prediction accuracy of the model, four different pre-clustering methods were tested individually, and a graph neural network model was designed and then trained on each cluster. The cluster size was set to 5 ASVs for all methods except IDEC, which decided for itself. The 4 clustering methods were 1) clustering by known metabolic function at genus level according to the MiDAS field guide (retrieved 2022 Jul 25 from www.midasfieldguide.org), 2) clustering by using the Improved Deep Embedded Clustering (IDEC)33 encoder-decoder algorithm, 3) ranked abundance clusters ranking from the most abundant to the least abundant, and lastly 4) by using the time-varying graphical clustering algorithm on the graph network interaction strengths34. The 5 metabolic functional groups used were polyphosphate accumulating organisms (PAO), glycogen accumulating organisms (GAO), filamentous microorganisms, ammonia oxidizing bacteria (AOB), and nitrite oxidizing bacteria (NOB). The hyperparameters used for the IDEC model were adjusted where a batch size of 32 was used with max 200 pre-training epochs and a tolerance of 0.001. The individual IDEC clusters were evaluated using the cluster accuracy metric (ACC).
For each of the 4 pre-clustering methods mentioned above, the graph neural network machine learning model was used as described previously28 and then was trained on the individual clusters through up to 30 iterations to find the best fit, where the training was stopped early if the model did not improve within 5 iterations. To evaluate the model performance, three different loss functions were used to calculate the error per cluster per time point between the true and predicted values; first of all, the Bray Curtis dissimilarity metric as it provides an ecologically meaningful comparison, and then also the mean absolute error (MAE) and mean squared error (MSE).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw DNA sequencing data from the 5307 environmental samples from activated sludge used for this study has been deposited to the NCBI SRA under accession ID PRJNA1021975. Furthermore, the community data produced from the raw data as well as all files used for the analyses and figures in this study is available in the github repository (https://github.com/kasperskytte/asmc-prediction) and data deposition at figshare (https://doi.org/10.6084/m9.figshare.25288159.v1. The source data underlying the figures are provided with this paper. Source data are provided with this paper.
Code availability
The code used to train the models, perform the analyses, and generate results in this study is publicly available and has been deposited in the mc-prediction repository at https://github.com/kasperskytte/mc-prediction, under the GNU General Public License v3.0 license. The specific version of the code associated with this publication is archived in Zenodo and is accessible via https://doi.org/10.5281/zenodo.1684027029.
References
Bar-On, Y. M., Phillips, R. & Milo, R. The biomass distribution on Earth. Proc. Natl. Acad. Sci. 115, 6506–6511 (2018).
Jones, E. R., van Vliet, M. T. H., Qadir, M. & Bierkens, M. F. P. Country-level and gridded estimates of wastewater production, collection, treatment and reuse. Earth Syst. Sci. Data 13, 237–254 (2021).
Grady, C. P. L., Daigger, G. T., Love, N. G. & Filipe, C. D. M. Biological Wastewater Treatment. (CRC Press, Boca Raton, https://doi.org/10.1201/b13775. 2011).
Yang, Y., Wang, L., Xiang, F., Zhao, L. & Qiao, Z. Activated sludge microbial community and treatment performance of wastewater treatment plants in industrial and municipal Zones. Int. J. Environ. Res. Public. Health 17, 436 (2020).
Andersen, M. H. et al. Fast DNA-analyses for surveillance of microbial communities in full-scale deammonification tanks: Potential for control and troubleshooting. Water Res. 236, 119919 (2023).
Sun, H. et al. Effects of influent immigration and environmental factors on bacterial assembly of activated sludge microbial communities. Environ. Res. 205, 112426 (2022).
Xia, Y., Wen, X., Zhang, B. & Yang, Y. Diversity and assembly patterns of activated sludge microbial communities: A review. Biotechnol. Adv. 36, 1038–1047 (2018).
Bairey, E., Kelsic, E. D. & Kishony, R. High-order species interactions shape ecosystem diversity. Nat. Commun. 7, 12285 (2016).
Faust, K. & Raes, J. Microbial interactions: from networks to models. Nat. Rev. Microbiol. 10, 538–550 (2012).
Gao, P. et al. Correlating microbial community compositions with environmental factors in activated sludge from four full-scale municipal wastewater treatment plants in Shanghai, China. Appl. Microbiol. Biotechnol. 100, 4663–4673 (2016).
Ju, F. & Zhang, T. Bacterial assembly and temporal dynamics in activated sludge of a full-scale municipal wastewater treatment plant. ISME J. 9, 683–695 (2015).
Faust, K., Lahti, L., Gonze, D., de Vos, W. M. & Raes, J. Metagenomics meets time series analysis: unraveling microbial community dynamics. Curr. Opin. Microbiol. 25, 56–66 (2015).
Fuhrman, J. A., Cram, J. A. & Needham, D. M. Marine microbial community dynamics and their ecological interpretation. Nat. Rev. Microbiol. 13, 133–146 (2015).
Gonze, D., Coyte, K. Z., Lahti, L. & Faust, K. Microbial communities as dynamical systems. Curr. Opin. Microbiol. 44, 41–49 (2018).
Peces, M. et al. Microbial communities across activated sludge plants show recurring species-level seasonal patterns. ISME Commun. 2, 1–11 (2022).
Hullar, M. A. J., Kaplan, L. A. & Stahl, D. A. Recurring seasonal dynamics of microbial communities in stream habitats. Appl. Environ. Microbiol. 72, 713–722 (2006).
Nielsen, P. H., Kragelund, C., Seviour, R. J. & Nielsen, J. L. Identity and ecophysiology of filamentous bacteria in activated sludge. FEMS Microbiol. Rev. 33, 969–998 (2009).
Eikelboom, D. H. Filamentous organisms observed in activated sludge. Water Res. 9, 365–388 (1975).
Li, C., Lim, K. M. K., Chng, K. R. & Nagarajan, N. Predicting microbial interactions through computational approaches. Methods San. Diego Calif. 102, 12–19 (2016).
Zuñiga, C., Zaramela, L. & Zengler, K. Elucidation of complexity and prediction of interactions in microbial communities. Microb. Biotechnol. 10, 1500–1522 (2017).
Ansari, A. F., Reddy, Y. B. S., Raut, J. & Dixit, N. M. An efficient and scalable top-down method for predicting structures of microbial communities. Nat. Comput. Sci. 1, 619–628 (2021).
Liu, X., Nie, Y. & Wu, X.-L. Predicting microbial community compositions in wastewater treatment plants using artificial neural networks. Microbiome 11, 93 (2023).
Baranwal, M. et al. Recurrent neural networks enable design of multifunctional synthetic human gut microbiome dynamics. eLife 11, e73870 (2022).
Thompson, J. C., Zavala, V. M. & Venturelli, O. S. Integrating a tailored recurrent neural network with Bayesian experimental design to optimize microbial community functions. PLOS Comput. Biol. 19, e1011436 (2023).
Larsen, P. E., Field, D. & Gilbert, J. A. Predicting bacterial community assemblages using an artificial neural network approach. Nat. Methods 9, 621–625 (2012).
Callahan, B. J., McMurdie, P. J. & Holmes, S. P. Exact sequence variants should replace operational taxonomic units in marker-gene data analysis. ISME J. 11, 2639–2643 (2017).
Delogu, F. et al. Forecasting the dynamics of a complex microbial community using integrated meta-omics. Nat. Ecol. Evol. 8, 32–44 (2024).
Zhao, K. et al. Multiple Time Series Forecasting with Dynamic Graph Modeling. Proc. VLDB Endow. 17, 753–765 (2023).
Andersen, K. S. & Zhao, K. KasperSkytte/MC-prediction: Version used for the paper. Zenodo https://doi.org/10.5281/ZENODO.16840270 (2025).
Wilson, G. et al. Best Practices for Scientific Computing. PLOS Biol. 12, e1001745 (2014).
Dueholm, M. K. D. et al. MiDAS 4: A global catalogue of full-length 16S rRNA gene sequences and taxonomy for studies of bacterial communities in wastewater treatment plants. Nat. Commun. 13, 1908 (2022).
Nierychlo, M. et al. MiDAS 3: An ecosystem-specific reference database, taxonomy and knowledge platform for activated sludge and anaerobic digesters reveals species-level microbiome composition of activated sludge. Water Res. 182, 115955 (2020).
Guo, X., Gao, L., Liu, X. & Yin, J. Improved Deep Embedded Clustering with Local Structure Preservation. In Proceedings of the Twenty-Sixth International Joint Conference on Artificial Intelligence 1753–1759 (Melbourne, Australia, 2017). https://doi.org/10.24963/ijcai.2017/243.
Hallac, D., Park, Y., Boyd, S. & Leskovec, J. Network Inference via the Time-Varying Graphical Lasso. In Proceedings of the 23rd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining 205–213 (ACM, Halifax NS Canada, 2017). https://doi.org/10.1145/3097983.3098037.
Nierychlo, M. et al. Low global diversity of Candidatus Microthrix, a troublesome filamentous organism in full-scale WWTPs. Front. Microbiol. 12, 1593 (2021).
Nielsen, P. H., McIlroy, S. J., Albertsen, M. & Nierychlo, M. Re-evaluating the microbiology of the enhanced biological phosphorus removal process. Curr. Opin. Biotechnol. 57, 111–118 (2019).
Liu, R., Hao, X., Chen, Q. & Li, J. Research advances of Tetrasphaera in enhanced biological phosphorus removal: A review. Water Res 166, 115003 (2019).
Juretschko, S. et al. Combined molecular and conventional analyses of nitrifying bacterium diversity in activated sludge: Nitrosococcus mobilis and Nitrospira-like bacteria as dominant populations. Appl. Environ. Microbiol. 64, 3042–3051 (1998).
Ushiki, N. et al. Genomic Analysis of Two Phylogenetically Distinct Nitrospira Species Reveals Their Genomic Plasticity and Functional Diversity. Front. Microbiol. 8, (2018).
Daims, H., Nielsen, J. L., Nielsen, P. H., Schleifer, K. H. & Wagner, M. In situ characterization of Nitrospira-like nitrite-oxidizing bacteria active in wastewater treatment plants. Appl. Environ. Microbiol. 67, 5273–5284 (2001).
Caporaso, J. G. et al. Moving pictures of the human microbiome. Genome Biol. 12, R50 (2011).
Chrismas, N., Allen, R., Allen, M. J., Bird, K. & Cunliffe, M. A 17-year time-series of fungal environmental DNA from a coastal marine ecosystem reveals long-term seasonal-scale and inter-annual diversity patterns. Proc. R. Soc. B Biol. Sci. 290, 20222129 (2023).
Costello, E. K. et al. Bacterial Community Variation in Human Body Habitats Across Space and Time. Science 326, 1694–1697 (2009).
Arumugam, M. et al. Enterotypes of the human gut microbiome. Nature 473, 174–180 (2011).
Hochreiter, S. & Schmidhuber, J. Long Short-Term Memory. Neural Comput. 9, 1735–1780 (1997).
Lin, H. et al. The role of the core microorganisms in the microbial interactions in activated sludge. Environ. Res. 235, 116660 (2023).
Rossetti, S., Tomei, M. C., Nielsen, P. H. & Tandoi, V. ‘Microthrix parvicella’, a filamentous bacterium causing bulking and foaming in activated sludge systems: a review of current knowledge. FEMS Microbiol. Rev. 29, 49–64 (2005).
Roels, T., Dauwe, F., Van Damme, S., De Wilde, K. & Roelandt, F. The influence of PAX-14 on activated sludge systems and in particular on Microthrix parvicella. Water Sci. Technol. J. Int. Assoc. Water Pollut. Res. 46, 487–490 (2002).
Wágner, D. S. et al. Seasonal microbial community dynamics complicates the evaluation of filamentous bulking mitigation strategies in full-scale WRRFs. Water Res 216, 118340 (2022).
McIlroy, S. J. et al. MiDAS: the field guide to the microbes of activated sludge. Database 2015, bav062 (2015).
Weisburg, W. G., Barns, S. M., Pelletier, D. A. & Lane, D. J. 16S ribosomal DNA amplification for phylogenetic study. J. Bacteriol. 173, 697–703 (1991).
Muyzer, G., de Waal, E. C. & Uitterlinden, A. G. Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl. Environ. Microbiol. 59, 695–700 (1993).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Edgar, R. C. UNOISE2: improved error-correction for Illumina 16S and ITS amplicon sequencing. bioRxiv 081257 https://doi.org/10.1101/081257 (2016).
Edgar, R. C. SINTAX: a simple non-Bayesian taxonomy classifier for 16S and ITS sequences. 074161 Preprint at https://doi.org/10.1101/074161 (2016).
Dueholm, M. S. et al. Generation of Comprehensive Ecosystem-Specific Reference Databases with Species-Level Resolution by High-Throughput Full-Length 16S rRNA Gene Sequencing and Automated Taxonomy Assignment (AutoTax). mBio 11, e01557–20 (2020).
Saunders, A. M., Albertsen, M., Vollertsen, J. & Nielsen, P. H. The activated sludge ecosystem contains a core community of abundant organisms. ISME J. 10, 11–20 (2016).
Yang, K. et al. Ecological and functional differences of abundant and rare sub-communities in wastewater treatment plants across China. Environ. Res. 243, 117749 (2024).
Andersen, K. S., Kirkegaard, R. H., Karst, S. M. & Albertsen, M. ampvis2: an R package to analyse and visualise 16S rRNA amplicon data. bioRxiv 299537 https://doi.org/10.1101/299537 (2018).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
Schloss, P. D. et al. Introducing mothur: Open-Source, Platform-Independent, Community-Supported Software for Describing and Comparing Microbial Communities. Appl. Environ. Microbiol. 75, 7537–7541 (2009).
Callahan, B. J. et al. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Acknowledgements
The project has been funded by the Villum Foundation (Dark Matter grant 16,578 to P.H.N. and 40,567 to C.G. and P.H.N.). We like to thank Dr. Feng Ju for providing the Hong Kong WWTP dataset, and Dr. Nathan Chrismas for providing the fungal environmental DNA from a coastal marine ecosystem dataset.
Author information
Authors and Affiliations
Contributions
P.H.N., C.G. and K.S.A. designed the study. K.S.A., M.P., and P.H.N. wrote the manuscript and all authors reviewed and approved the final manuscript. K.S.A. performed the bioinformatic processing, produced and analyzed the results. M.N., M.P. and P.H.N. were responsible for obtaining microbial time series. K.S.A., K.Z., A.L.A., C.B.S., T.J.H., C.G. developed the methods at different stages.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Marco Tulio Angulo, Xiao-Lei Wu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Andersen, K.S., Zhao, K., Agerskov, A.d.L. et al. Predicting microbial community structure and temporal dynamics by using graph neural network models. Nat Commun 16, 9124 (2025). https://doi.org/10.1038/s41467-025-64175-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-64175-7