Abstract
Immune checkpoint inhibitors (ICIs) have revolutionized cancer therapy; however, their use is frequently associated with immune-related adverse events (irAEs). In this study, anti-PD-L1 therapy exacerbates muscle wasting in tumor-bearing male mice despite its anti-tumor efficacy, accompanied by an accumulation of CD8+ T cells in muscle. Single-cell RNA sequencing identifies these cells as tissue-resident memory-like CD49a+ CD8+ T cells. While CD8+ T cell depletion prevents muscle wasting, it compromises the anti-tumor efficacy of anti-PD-L1. To resolve this paradox, we identify cathepsin L (CTSL) as a dual-target capable of suppressing both tumor progression and CD8+ T cell-mediated muscle wasting, through integrative transcriptomic analysis. Pharmacological inhibition of CTSL not only mitigates anti-PD-L1-induced muscle wasting but also further suppresses tumor growth, potentially via downregulation of BNIP3. Here, we show that CTSL is a dual-action target to uncouple anti-tumor efficacy from muscle-specific irAEs, offering a strategy to improve clinical outcomes of ICIs.
Similar content being viewed by others
Introduction
Immune checkpoint inhibitors (ICIs), such as monoclonal antibodies (mAbs) targeting programmed cell death protein 1 (PD-1) or its ligand PD-L1, have revolutionized cancer treatment by harnessing the immune system to combat various cancer types, including non-small cell lung cancer (NSCLC)1,2. By unleashing cytotoxic T cells, ICIs significantly improve tumor control, but this immune overactivation also triggers immune-related adverse events (irAEs), which can affect organs as diverse as the lung, colon, liver, and skin3,4,5. Notably, skeletal muscle has emerged as a target for irAEs, with anti-PD-1 (αPD-1) or anti-PD-L1 (αPD-L1) therapy linked to myositis, myasthenia gravis, and other myopathies6,7. These muscle-specific irAEs can severely impair quality of life in cancer patients, necessitating immunosuppressive treatments, which may diminish the therapeutic efficacy of ICIs8,9. Thus, there is an urgent need for strategies that prevent muscle-specific irAEs without compromising anti-tumor immunity.
Cancer cachexia is a multifaceted syndrome marked by involuntary weight loss, systemic inflammation, and wasting of fat and skeletal muscle10,11. It arises from complex tumor-host crosstalk involving the central nervous system (CNS), liver, white adipose tissue (WAT), and skeletal muscle, and is driven in part by elevated pro-inflammatory cytokines such as interleukin-6 (IL-6) and tumor necrosis factor-α (TNF-α)12,13,14,15,16. Cachexia directly contributes to roughly 20% of cancer deaths and is associated with poor responses to ICIs10,11,17,18,19, whereas obesity often predicts better outcomes following ICI therapy20,21. Despite the profound impact of cachexia on immunotherapy outcomes, the bidirectional interplay between these processes and their underlying mechanisms remains largely unexplored.
Skeletal muscle wasting, a hallmark of cancer cachexia, involves decreased protein synthesis, enhanced ubiquitin-proteasome activity, dysregulated autophagy, and apoptosis22,23. Beyond these intrinsic pathways, tumor-educated immune cells, including macrophages, neutrophils, and cytotoxic T cells, drive muscle atrophy via cytokine release and direct cytotoxicity24,25,26,27,28,29. Given that ICIs reinvigorate cytotoxic T cells and muscle is a documented irAE target, ICI treatment might exacerbate cancer-associated muscle wasting as a muscle-specific irAEs.
Cathepsin L (CTSL) is a lysosomal cysteine protease ubiquitously expressed across tissues. Under certain conditions, it is secreted into the extracellular matrix (ECM) to remodel tissue architecture30,31,32. In many cancers, CTSL is upregulated and promotes angiogenesis, invasion, and metastasis, features linked to poor clinical outcomes33,34,35. Paradoxically, CTSL can also suppress tumorigenesis by modulating growth signaling pathways36,37, triggering oncogene-induced senescence38,39, and participating in chromatin remodeling40. Beyond cancer, CTSL regulates immune function by enhancing the pore-forming activity of cytotoxic T cells and natural killer (NK) cells41, guiding NKT cell and CD4+ T cell development42,43, and processing major histocompatibility complex class II (MHC-II)44. In skeletal muscle, CTSL exacerbates cancer-associated muscle wasting by enhancing autophagy-dependent proteolysis45,46. Collectively, these findings highlight CTSL’s context-dependent roles in tumor progression, immune regulation, and muscle homeostasis.
Here, using a syngeneic orthotopic lung tumor-bearing (TB) mouse model, we show that αPD-L1 treatment suppresses tumor growth but concurrently aggravates muscle wasting by recruiting a distinct subset of CD49a+ CD8+ tissue-resident memory-like T cells into skeletal muscle. Although CD8+ T cell depletion mitigates muscle wasting, it also abolishes ICI’s anti-tumor effects, revealing a critical therapeutic trade-off. Through unbiased integrative transcriptomic analysis, we identify CTSL as a dual-action target. Its inhibition preserves muscle mass by blocking T cell-mediated atrophy and simultaneously augments ICI efficacy. Our findings provide a framework to prevent muscle-specific irAEs while enhancing the benefits of ICIs.
Results
αPD-L1 exacerbates muscle wasting while suppressing tumorigenesis
To assess whether ICIs contribute to cancer cachexia, we analyzed body weight changes in cancer patients treated with ICIs and compared with patients receiving other cachexia-inducing therapies, such as doxorubicin and pemetrexed47,48,49,50, at Yonsei University Severance Hospital (Supplementary Fig. 1A). Patients treated with ICIs, including αPD-1 and αPD-L1, exhibited body weight loss comparable to that observed in patients receiving other chemotherapies (two one-sided tests [TOST], p-value < 0.0001; Supplementary Fig. 1B). This observation was validated in an independent cohort from Pusan National University Hospital (Supplementary Fig. 1C).
To determine whether the observed weight changes were driven by ICI treatment itself rather than simply reflecting insufficient tumor control, we reanalyzed a preclinical subcutaneous tumor model in which tumor growth was effectively suppressed by αPD-L1 therapy51. Remarkably, mice that responded to αPD-L1 still lost tumor-free body weight, whereas untreated tumor-bearing controls gained net body weight regardless of tumor size (Fig. 1A and Supplementary Fig. 1D, E). These data indicate that αPD-L1 can directly exacerbate cachexia as an irAE, independent of its anti-tumor efficacy.
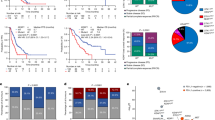
A Tumor volume and tumor-free body weight (BW) changes at 13 weeks in CT26 mice. αPD-L1 responders (green, n = 5; per original study51; Supplementary Fig. 1D, E) versus IgG controls (black, n = 8); BW normalized to initial BW (IBW). B Male C57BL/6 mice were intratracheally (i.t.) injected with KrasIKKαL cells, treated intraperitoneally (i.p.) with the indicated antibodies or IgG three times weekly, and euthanized when any group reached ≥ 10% BW loss versus IBW. Lung size (C), tumor burden (D), BW changes (E), and gastrocnemius (GA) and tibialis anterior (TA) muscle weights (F) in TB versus CON mice (n = 7 mice; BW and muscle weights normalized to IBW). Representative H&E of TA (G) and averaged myofiber CSA per mouse (H; n = 7; ≥100 myotubes per field; three fields per mouse). I Percentage of CD3+ CD8+ cells within CD45+ infiltrates in tumor and GA by flow cytometry (n = 5). J Principal component analysis (PCA) of TA transcriptomes (CON, n = 5; IgG, n = 4; αPD-L1, n = 3; αCD8a, n = 3). Top five enriched Gene Ontology (GO) terms for differentially expressed genes (DEGs) in muscles: IgG vs. CON (K); αPD-L1 vs. IgG (L); αCD8a vs. IgG (M). Enrichment score represents −log10-transformed adjusted p-values; FDR false discovery rate. N WB analysis of indicated proteins in GA. Vinculin muscle-specific loading control. Samples derive from the same experiment; different gels for MuRF-1, LC3B-I/II, and another for vinculin were processed in parallel. O Final grip strength normalized to IBW (n = 7 mice). Co-culturing of C2C12 myotubes (P) with splenic CD8+ T cells from CON or TB mice in KrasIKKαL cell-conditioned medium (CM); myotube diameter (Q) from 12 myotubes per well, biological triplicate wells, each dot means averaged diameter per well. R WB analysis of C2C12 myotube proteins after co-culture. For A, data are mean ± SEM, one-sided t-test; for D–F, H, I, O, and Q, mean ± SEM, two-sided unpaired t-test; for K–M, hypergeometric test with Benjamini–Hochberg correction. Source data are provided as a Source Data file.
To validate this in vivo, we used a syngeneic orthotopic lung adenocarcinoma (LUAD) model in male C57BL/6 mice. Tumors were established by intratracheal instillation of KrasIKKαL cells (Fig. 1B), which develop an immunosuppressive tumor microenvironment (TME) with low CD8+ T cell and high regulatory T cell (Treg) infiltration52,53, ideal for ICI studies. Then, mice received intraperitoneal injections of αPD-L1 with or without anti-CD8α (αCD8a), or isotype-control immunoglobulin G (IgG). Once any treatment group first reached ≥ 10% body weight loss relative to its own baseline, a standard cachexia threshold54,55,56,57, all cohorts were sacrificed on the same day.
Strikingly, αPD-L1-treated TB mice reached this threshold earliest, despite having significantly smaller tumors and lower lung weights than IgG controls (Fig. 1C–E, and Supplementary Fig. 2A, B). At the endpoint, the αPD-L1 group had profound loss of epididymal WAT (eWAT) and gastrocnemius (GA) and tibialis anterior (TA) muscles, along with reduced muscle fiber cross-sectional area (CSA), compared to the IgG group (Fig. 1E–H and Supplementary Fig. 2C, blue vs. red). By contrast, the IgG group, despite larger tumors, maintained near-baseline body weight, while still losing fat and muscle compared to healthy controls (CON) (Fig. 1C–H and Supplementary Fig. 2A–C, red vs. black), indicating a bona fide cachectic phenotype rather than simple growth arrest. Comparable food intake across TB groups (Supplementary Fig. 2D) and lack of αPD-L1-induced weight loss in CON mice (Supplementary Fig. 2E, F) rule out anorexia or overt toxicity. Of note, both IgG and αPD-L1 slightly delayed weight gain in CON mice (Supplementary Fig. 2E, F), likely reflecting a non-specific effect of chronic antibody exposure. Only TB mice exhibited pronounced hypersensitivity to αPD-L1, manifesting accelerated cachexia independently of tumor control.
ICI-potentiated cytotoxic CD8+ T cells can also provoke irAEs4,58. In lung tumors of TB mice, αPD-L1 reversed immunosuppression by boosting CD8+ T cells and lowering Foxp3+ Tregs (Fig. 1I and Supplementary Fig. 2G). By contrast, CD8+ T cell infiltration was already elevated in the skeletal muscle of TB mice relative to CON (Fig. 1I, red vs. black), and further amplified following αPD-L1 treatment (Fig. 1I, blue vs. red), suggesting a role for CD8+ T cells in muscle wasting. To test causality, we depleted CD8+ T cells with anti-CD8α (αCD8a) antibodies. As expected, CD8+ T cell depletion abolished the anti-tumor efficacy of αPD-L1 (Fig. 1C, D and Supplementary Fig. 2A, yellow vs. blue), yet failed to rescue body weight, fat, or muscle loss by αPD-L1 (Fig. 1E–H and Supplementary Fig. 2B, C, yellow vs. blue), reflecting a trade-off: removing CD8+ T cells spares muscle wasting but permits unchecked tumor growth, which itself further drives cachexia. Critically, CD8+ T cell depletion alone, without αPD-L1, preserved body weight and prevented fat and muscle wasting, while tumor size remained unchanged (Fig. 1C–I and Supplementary Fig. 2A–C, purple vs. red), underscoring CD8+ T cells as crucial mediators in cancer-induced muscle atrophy.
To uncover underlying molecular changes, we performed RNA-seq on skeletal muscle tissues. Principal component analysis (PCA) showed that IgG and αPD-L1 groups clustered far from CON, whereas αCD8a monotherapy shifted the muscle transcriptome back toward CON (Fig. 1J). Functional enrichment analysis revealed that genes involved in ECM organization were downregulated, whereas autophagy-related genes were upregulated in the IgG group relative to CON (Fig. 1K and Supplementary Fig. 3A, B). αPD-L1 treatment further suppressed the expression of structural muscle genes compared to IgG (Fig. 1L and Supplementary Fig. 3C). Notably, CD8⁺ T cell depletion reversed these transcriptional alterations (Fig. 1M and Supplementary Fig. 3A–C). Uniquely, αPD-L1-induced gene signatures associated with leukocyte migration and chemotaxis (Fig. 1L and Supplementary Fig. 3D), implicating immune infiltration in muscle wasting.
Consistent with these transcriptomic shifts, IgG-treated TB mice upregulated the muscle-specific E3 ligase MuRF-1 and autophagy marker LC3B-II in muscle, both of which were suppressed by CD8+ T cell depletion (Fig. 1N and Supplementary Fig. 2H). Functionally, grip strength was reduced in TB mice (Fig. 1O, red vs. black), and further impaired by αPD-L1 (Fig. 1O, blue vs. red), but restored by CD8+ T cell depletion (Fig. 1O, purple vs. red). These results support that cytotoxic CD8+ T cells are required for cancer-associated muscle wasting and its exacerbation by PD-L1 blockade.
To test this directly, we co-cultured fully differentiated C2C12 myotubes with splenic CD8+ T cells from either CON or TB mice in the presence of KrasIKKαL cell-conditioned medium (CM) (Fig. 1P). CM alone reduced myotube diameter, and addition of CD8+ T cells exacerbated atrophy regardless of their source, with TB-derived cells more potent (Fig. 1Q and Supplementary Fig. 4A, B). In contrast, CD4+ T cells had no impact, even with CM (Supplementary Fig. 4C, D). Western blot (WB) analysis confirmed that CM upregulated the muscle-specific E3 ligase atrogin-1 and autophagy marker LC3B-II (Fig. 1R and Supplementary Fig. 4E). Co-culture with CD8+ T cells further increased atrogin-1 (Fig. 1R and Supplementary Fig. 4E) and worsened atrophy (Fig. 1Q, blue vs. red). Thus, cytotoxic CD8+ T cells can directly induce muscle wasting, acquiring enhanced atrophy-inducing capacity in the tumor milieu, and likely underlie the muscle-specific irAEs seen with αPD-L1 therapy.
Cytotoxic CD49a+ CD8+ T cells accumulate in skeletal muscle of TB mice
To dissect the immune landscapes of tumor versus muscle in cachexia, we performed single-cell RNA-sequencing (scRNA-seq) on CD45+ cells isolated from both lung tumors and skeletal muscles of TB mice exhibiting cachectic features (Supplementary Fig. 5A–E). This degree of wasting phenotype resembled that observed in αPD-L1-treated TB mice (Fig. 1E, F and Supplementary Fig. 2C) and was accompanied by elevated pro-cachectic cytokines Tnf-α and Il-6 in lung, muscle, eWAT, and liver of TB mice, as well as reduced grip strength (Supplementary Fig. 5E–I). Similar results were obtained in a well-established cancer cachexia model using intratracheal injection of Lewis lung carcinoma (LLC) cells (Supplementary Fig. 5K–Q), validating our model.
Among 14,028 CD45+ cells captured, 11,348 single-cell transcriptomes passed quality control for analysis. We identified eight major immune clusters with distinct tissue biases (Fig. 2A–C and Supplementary Fig. 6A, B). Lung tumors were dominated by neutrophils and macrophages, then classical T and B cells (Fig. 2B). Strikingly, skeletal muscle showed selective enrichment of a unique T cell subset: CD49a+ CD8+ T cells (Fig. 2C). This CD49a+ CD8+ T cell population formed its own transcriptional cluster, separate from classical T cells (Fig. 2A and Supplementary Fig. 6B). Although both subsets expressed T cell receptor (TCR)-related genes, the CD49a+ CD8+ subset showed lower and more heterogeneous Cd3 and Cd8 expression and a muscle-enriched signature, including Nr3c1, Nrp1, Entpd1, Cd200, Ly6c1, Zbtb16, and Itga1 (Fig. 2D and Supplementary Fig. 6B–D). Notably, Itga1, which encodes CD49a, marks tissue-resident memory CD8+ T (TRM) cells, was robustly and broadly expressed within this population; Ly6c1 and Zbtb16 are linked to tissue homing and innate-like T cell identity (Fig. 2D and Supplementary Fig. 6D)59,60,61,62,63,64,65.
A UMAP of scRNA-seq profiles from CD45+ cells isolated from lung tumors or GA muscles of TB mice (pooled from three animals), colored by tissue origin (left) and by immune cell types (right). Relative proportions of each CD45+ immune cluster in lung tumors (B) and GA muscle (C). D UMAP expression overlays for selected T cell-related and cytotoxicity genes in CD45+ cells isolated from tumors or muscle tissue (purple indicates expression; scale bar at right). Top five enriched GO/Kyoto Encyclopedia of Genes and Genomes (KEGG) terms for genes upregulated in classical T cells versus CD49a+ CD8+ T cells (E) and vice versa (F). Enrichment scores represent −log10 of the adjusted p-values from the hypergeometric test with Benjamini–Hochberg correction. G and H Percentage of CD49a+ CD8+ T cells (G, CD45+ CD3+ CD8+ CD49a+) and CD8+ T cells (H, CD45+ CD3+ CD8+) within CD45+ infiltrating tumor and GA muscle tissues by flow cytometry (n = 6). I qRT-PCR of Itga1, Prf1, Gzma, and Gzmb in CD8+ T cells co-cultured with C2C12 myotubes in the presence or absence of CM as described in Fig. 1Q. Vinculin, a muscle-specific housekeeping gene for normalization (n = 4 independent biological replicates per group). For E and F, hypergeometric test with Benjamini–Hochberg correction; for G–I, data are mean ± SEM, two-sided unpaired t-test. Source data are provided as a Source Data file.
Comparative transcriptomic analysis revealed that, unlike classical T cells (enriched in genes related to leukocyte activation and TCR signaling), CD49a+ CD8+ T cells upregulated genes for epithelial migration and Rap1 signaling (Fig. 2E, F and Supplementary Fig. 6E–H). Given that Rap1 enhances integrin-dependent tissue homing66,67, this reinforces their identity as a TRM-like cytotoxic T cell subset in skeletal muscle.
Flow cytometry confirmed that CD49a+ CD8+ T cells were significantly enriched in the muscle of TB mice, but were reduced in lung tumors, where Tregs and myeloid-derived suppressor cells (MDSCs) were increased compared to CON mice (Fig. 2G and Supplementary Figs. 5J, 7A, B). In muscle, most CD8+ T cells co-expressed CD49a, whereas in tumors, only a fraction did (Fig. 2G, H). The LLC cachexia model similarly showed muscle enrichment of CD49a+ CD8+ T cells (Supplementary Fig. 5R), underscoring the cross-model reproducibility of this immune feature.
Finally, in ex vivo co-cultures, CD8+ T cells exposed to C2C12 myotubes plus CM upregulated Itga, Prf1, Gzma, and Gzmb (Fig. 2I), mirroring the in vivo signature (Fig. 2D). Given emerging links between TRM cells and irAEs68,69,70, our data suggest that CD49a+ CD8+ TRM-like cells contribute to cancer-induced muscle wasting.
CTSL as a target for tumorigenesis and T cell-mediated muscle wasting
Cytotoxic CD8+ T cells are critical drivers in cancer-associated muscle wasting. However, their systemic depletion abolishes anti-tumor immunity (Fig. 1C), and selectively targeting the CD49a+ subset remains technically challenging. To uncover druggable nodes that could simultaneously curb tumor progression and muscle atrophy, we performed integrative transcriptomic analysis of tumor and skeletal muscle from TB mice.
We first identified 55 genes that were upregulated in both tumor and muscle during cachexia and downregulated in muscle upon CD8⁺ T cell depletion (Fig. 3A). Protein–protein interaction (PPI) network71 revealed a densely connected “Community 1” enriched for genes associated with proteasome and autophagy, processes central for muscle catabolism (Fig. 3B and Supplementary Fig. 8A, B). These genes were consistently upregulated across multiple murine cachexia models (Fig. 3C and Supplementary Fig. 8C), underscoring their broader relevance.
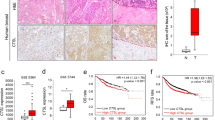
A Venn diagram of genes upregulated in muscles (TA, GA) and lung tumor from TB versus CON, and genes downregulated in TA from αCD8a versus IgG groups. B PPI network of 55 overlapping DEGs from (A), highlighting Communities 1 and 2 (nodes > 5; STRING combined score > 400); KEGG pathways enriched in Community 1 (adjusted p < 0.05). C Heatmap of Community 1 genes in muscle across multiple murine cancer cachexia models (Supplementary Fig. 8C). Color shows log2-fold change (FC) in cachectic (CAC) versus control (CON) groups: red, up; blue, down; *adjusted p < 0.05; **adjusted p < 0.005. D Representative WB in tumor and GA. Gapdh (lung) and vinculin (muscle) as loading controls. Two mice per group are shown (n = 4 per group); quantification for all mice in Supplementary Fig. 9A, B. For GA, samples derive from the same experiment; different gels for Cstl, and another for vinculin, were processed in parallel. Expression changes of Ctsl and neighbors in PPI network in lung (E) and muscle (F, G); color intensity shows log2 FC; red, up; blue, down; bold edge, significant DEGs (orange, up; dark blue, down). H Correlation coefficients among indicated genes in TB versus CON lung (E). I Top five KEGG pathways for upregulated Ctsl neighbors in the PPI network (F). J Top five KEGG pathways for downregulated Ctsl neighbors in PPI network (G). K Kaplan–Meier plot of TCGA LUAD patients stratified by CTSL expression (high, red, n = 254; low, black, n = 288). Correlation of CTSL versus CD274 expression in TCAG LUAD (L, n = 541; log2(TPM + 1)) and GSE283829 NSCLC (M, n = 27; log2(read count+1)). N CTSL mRNA expression in skeletal muscle of NSCLC cachexia patients (CAC) versus healthy controls (CON); E-MTAB-12781; n = 5. Box plots show median (center line), interquartile range (IQR), and whiskers to 1.5× IQR. For B, F, G, I, and J, hypergeometric test with Benjamini–Hochberg correction; for C, Wald test with Benjamini–Hochberg correction; for K, log-rank test; for L and M, two-sided t-test for Pearson correlation; for N, one-sided unpaired t-test. Source data are provided as a Source Data file.
Within this core cluster, Ctsl emerged as a particularly promising target due to its robust and consistent upregulation across datasets (Fig. 3C and Supplementary Fig. 8C). WB confirmed elevated Ctsl expression in both tumor and muscle of TB mice relative to CON (Fig. 3D and Supplementary Fig. 9A, B). In muscle, Ctsl levels rose further with αPD-L1 therapy and fell down with CD8+ T cell depletion, whereas in tumors, only CD8+ T cell depletion reduced Ctsl expression (Fig. 3D). These patterns implicate CTSL in both tumor progression and T cell-mediated muscle wasting, highlighting it as a dual-action therapeutic target.
Pathway analysis of PPI partners co-expressed with Ctsl highlighted its tissue-specific roles (Fig. 3E–J). In tumors, Ctsl was co-upregulated with matrix metalloproteinases Mmp7 and Mmp13 (Fig. 3E, H) that drive ECM remodeling and metastasis72. In contrast, in skeletal muscle, Ctsl was co-expressed with genes involved in autophagy, apoptosis, and lysosome pathways (Fig. 3F, I and Supplementary Fig. 8D). These muscle-associated co-expression signatures were reversed by CD8+ T cell depletion (Fig. 3G, J, and Supplementary Fig. 8D), suggesting that CTSL may contribute to T cell-mediated muscle wasting.
To explore clinical relevance, we analyzed CTSL expression across human cancer datasets. The Cancer Genome Atlas (TCGA) data revealed widespread CTSL expression in multiple tumor types, prominently in NSCLC (both LUAD and lung squamous cell carcinoma [LUSC]) (Supplementary Fig. 10A). High CTSL expression predicted poorer overall survival in LUAD, LUSC, and pan-cancer cohorts (Fig. 3K and Supplementary Fig. 10B). CTSL levels also strongly correlated with CD274 (PD-L1) expression across tumors (Fig. 3L and Supplementary Fig. 10C) and in an independent NSCLC dataset (Fig. 3M). Beyond its role in tumors, our reanalysis of human muscle transcriptome73 showed that CTSL was significantly upregulated in muscle from cachectic NSCLC patients compared to healthy individuals (Fig. 3N), underscoring its involvement in human cancer cachexia. Collectively, these data identify CTSL as a clinically relevant target for suppressing tumor progression and T cell-driven muscle wasting.
CTSLi prevents muscle wasting by αPD-L1, further suppressing tumor
To test whether targeting CTSL could both enhance αPD-L1 efficacy and prevent its muscle-specific toxicity, we treated TB mice with a CTSL inhibitor (CTSLi) alone or combined with αPD-L1 (Fig. 4A). Combination therapy further reduced lung tumor burden compared to αPD-L1 monotherapy (Fig. 4B, C, and Supplementary Fig. 11A), indicating additive anti-tumor activity via distinct but complementary mechanisms. Crucially, CTSLi, whether used alone or with αPD-L1, significantly prevented body weight loss, fat depletion, and muscle wasting seen with αPD-L1 (Fig. 4D–H and Supplementary Fig. 11B, C). Moreover, combination therapy markedly reduced muscle levels of cachexia-associated cytokines Tnf-α and Il-6 (Supplementary Fig. 11D, E), supporting its ability to suppress αPD-L1-induced muscle wasting.
A Schematic of the animal study. Lung size (B), tumor burden (C), BW changes (D), and GA and TA muscle weights (E) in mice (n = 8; weights normalized to IBW). F Representative H&E and Masson’s Trichrome images of TA muscle. G Muscle CSA quantification (n = 8 mice per group; ≥ 100 myotubes per field, three fields per section from each mouse). H Final grip strength normalized to IBW (n = 8 mice per group). I CTSL concentration in tumor and TA muscle (n = 3 mice per group; normalized to tissue mass). J and K Percentage of indicated immune subsets within CD45+ infiltrating tumor and GA muscle tissues by flow cytometry (n = 8). L Correlation of CTSL and HLA-B expression in human skeletal muscle: NSCLC (E-MTAB-12781; CON, n = 4; CAC, n = 5) and PDAC (GSE133523; CON, n = 3; CAC, n = 3), log2(TPM + 1). M Correlation of Ctsl and H2-D1 expression in lung tumor and muscle (tumor: CON, n = 3; TB, n = 5; muscle: CON, n = 5; IgG, n = 4; αPD-L1, n = 3; αCD8a, n = 3), log2(TPM + 1). N WB of MHC-I expression in GA muscle. Two mice per group are shown (n = 4 per group); quantification of all four mice is presented in Supplementary Fig. 11H. Samples derive from the same experiment; different gels for MHC-I and another for vinculin were processed in parallel. O WB analysis of MHC-I expression in C2C12 myotubes transfected with si-RNA against Ctsl (si-Ctsl) or a non-targeting si-control (si-Cont). Samples derive from the same experiment; different gels for Ctsl, and another for MHC-I, vinculin, were processed in parallel. P C2C12 myotube diameter after transfection with siRNA, followed by co-culturing with splenic CD8+ T cells from TB mice in CM (12 myotubes per well, biological triplicates; total 36 myotubes per group). For C–E, G–K, and P, data are mean ± SEM, two-sided unpaired t-tests; for L and M, two-sided t-test for Pearson correlation. Source data are provided as a Source Data file.
Next, we examined the link between CTSL and cytotoxic T cells. In both tumor and muscle, Ctsl concentration rose in TB mice and was effectively lowered by CTSLi (Fig. 4I). In skeletal muscle, Ctsl levels strongly correlated with infiltration of CD8+ and CD49a+ CD8+ T cells; both were amplified by αPD-L1 (Fig. 4I–K, blue vs. red) and reversed by CTSLi (Fig. 4I–K, yellow vs. blue). In contrast, in lung TME, CTSLi modestly increased CD8+ T cell infiltration under αPD-L1 therapy (Fig. 4J, K, yellow vs. blue), suggesting that CTSL modulates immune cell dynamics differently in muscle versus tumor. These divergent effects align with tissue-specific immune contexture in TB mice: tumors were rich in immunosuppressive MDSCs and Tregs, whereas muscle exhibited a more immunostimulatory profile with elevated CD8+ T cells (Fig. 4J, K and Supplementary Fig. 11F, G). Interestingly, CTSLi shifted CD8+ T cell and Treg frequencies in opposite directions in tumor and muscle (Fig. 4J, K and Supplementary Fig. 11G), underscoring its context-dependent immune modulation.
MHC-I molecules, critical for CD8+ T cell recognition via TCRs, are generally low in healthy muscle but are upregulated in inflammatory conditions, promoting CD8+ T cell-mediated tissue damage74,75,76. We thus examined the relationship between CTSL and MHC-I expression in skeletal muscle. In public human muscle datasets73,77, CTSL expression correlated with HLA-B (encoding MHC-I) levels in both healthy and cachectic individuals (Fig. 4L and Supplementary Fig. 12A, B). In TB mice, Ctsl expression positively correlated with H2-D1 (the murine MHC-I homolog) in muscle, but not in tumors (Fig. 4M). MHC-I expression was elevated at both transcript and protein levels in skeletal muscle of TB mice and was further upregulated by αPD-L1; this upregulation was effectively reversed by CTSLi (Fig. 4N and Supplementary Fig. 11H, 12C). To rule out off-target effects of CTSLi, we performed Ctsl knockdown in C2C12 myotubes: siRNA-mediated Ctsl silencing reduced MHC-I expression at both protein and mRNA levels and protected against CD8+ T cell-induced atrophy (Fig. 4O, P and Supplementary Fig. 11I–K). Collectively, these results indicate that CTSL inhibition can uncouple anti-tumor immunity from muscle-specific irAE: it augments αPD-L1 efficacy while suppressing cytotoxic T cell-driven muscle wasting via an MHC-I-dependent mechanism.
BNIP3 mediates CTSL-induced tumorigenesis and muscle wasting
To uncover how CTSL drives both tumor progression and muscle wasting, we focused on its top interactors within our integrative PPI network (Fig. 5A and Supplementary Fig. 13A, B). Among these candidates, Bnip3 emerged as the most consistently correlated with Ctsl across both tumors and muscles, based on data from our TB mouse model and additional murine cancer cachexia models (Fig. 5B, C). Importantly, Bnip3 was the only direct Ctsl-interacting partner whose expression was tightly co-regulated with Ctsl in both tumor and muscle, and uniquely downregulated in muscle upon CD8+ T cell depletion (Fig. 5D and Supplementary Fig. 13C, D). Moreover, as shown in Fig. 3B, it was identified as a core member of Community 1.
A Scatter plots of genes ranked by Pearson’s correlation with Ctsl expression across lung tumor and skeletal muscle (TB versus CON). B Comparative ranking of these correlation coefficients in lung tumor versus muscle for Ctsl’s direct neighbors in PPI network (as shown in Fig. 3E). C Correlation of Ctsl versus Bnip3 expression in lung tumor and muscle (log2 TPM + 1). D Three-way Venn diagram of Ctsl’s direct neighbors that are upregulated in lung tumor, upregulated in muscle (TB versus CON), and downregulated in muscle (αCD8a versus IgG). Subnetwork of Ctsl-Bnip3 neighbors upregulated in tumors (E, TB versus CON; node color represents log2 FC). Bar plot (F) of KEGG pathways enriched among genes (adjusted p-value < 0.05). Subnetwork of Ctsl-Bnip3 neighbors (G) upregulated in muscle (left in the box: TB versus CON) and downregulated in muscle (right in the box: αCD8a versus IgG group). Node color indicates log2 FC. Bar plots (H) of KEGG pathways enriched among genes (adjusted p-value < 0.05). For C, two-sided t-test for Pearson correlation; for F and H, hypergeometric test with Benjamini–Hochberg correction. Source data are provided as a Source Data file.
BNIP3, a mitophagy receptor, regulates mitochondrial quality control and energy homeostasis, but its effects diverge depending on tissue contexts. In tumors, BNIP3 promotes cancer cell survival, metabolic adaptation, and metastasis78,79,80. By contrast, in skeletal muscle, BNIP3 contributes to atrophy by triggering autophagy and/or mitophagy81,82,83. Accordingly, the Ctsl-Bnip3 network displayed tissue-specific co-expression patterns: in tumors, Bnip3 was co-upregulated with genes involved in mitochondrial function (e.g., Bid, Bok, Pgam5, Ppif, Pmaip1, Vdac1) and energy metabolism (e.g., Bid, Ddit4, Gapdh, Ldha, Ppif, Pgk1, Phb2, Tpi1) (Fig. 5E, F), whereas in muscle, it was co-upregulated with autophagy and mitophagy regulators (e.g., Akt1, Bnip3l, Foxo3, Sqstm1, Map1lc3b, Mtor, Npc1), but these patterns were reversed by CD8+ T cell depletion (Fig. 5G, H). These contrasting co-expression modules suggest that BNIP3 serves as a context-dependent effector of CTSL signaling in tumor and muscle.
CTSL enhances tumor metastatic potential via BNIP3 upregulation
To evaluate the clinical relevance of the CTSL-BNIP3 network in LUAD, we first analyzed TCGA data and observed that the majority of CTSL-BNIP3 network genes were significantly upregulated in tumor versus matched normal tissues (Fig. 6A), mirroring our TB model (Fig. 5E). Immunofluorescence (IF) co-staining of a human LUAD tissue microarray confirmed that BNIP3 expression was markedly higher in CTSL-high tumors than in CTSL-low tumors (Fig. 6B and Supplementary Fig. 14A, B). Strikingly, when LUAD patients were stratified by their combined CTSL and BNIP3 expression levels, the CTSL-BNIP3-high subgroup exhibited significantly worse overall survival compared to the CTSL-BNIP3-low subgroup (HR = 1.9; log-rank p = 0.0013) (Fig. 6C and Supplementary Fig. 15A), exceeding the prognostic power of CTSL alone (Fig. 3K, HR = 1.53; p = 0.0035). Functional enrichment analysis further revealed that metastasis-associated gene signatures were markedly enriched in the CTSL-BNIP3-high patients both in TCGA (Fig. 6D, left) and in an independent validation cohort (Supplementary Fig. 15B–D), as well as in tumors from TB mice (Fig. 6D, right). Together, these data support a model that CTSL and BNIP3 cooperate to drive metastatic programs in LUAD.
A Human PPI network of CTSL-BNIP3 neighbors colored by log2 FC (tumor versus matched normal lung) in TCGA LUAD. B Immunofluorescence of CTSL (green) and BNIP3 (red) in LUAD tissue microarray cores: normal adjacent, n = 22; tumor cores, n = 128. Bar graph shows proportions of BNIP3-high/-low within the CTSL-high and CTSL-low tumors. C Kaplan–Meier overall survival for TCGA LUAD stratified by combined CTSL and BNIP3 expression (CTSL-BNIP3-high, red, n = 183; CTSL-BNIP3-low, black, n = 156). D GSEA plots of ALONSO_METASTASIS_UP in CTSL-BNIP3 high versus low groups in TCGA LUAD (left) and in TB mice (right). E Representative WB of indicated proteins in TB muscle. Two mice per group are shown (n = 4 per group); quantification of all four mice in Supplementary Fig. 16A. Samples derive from the same experiment; different gels for Ctsl, N-Cad, Gapdh, and another for Bnip3 were processed in parallel. F WB of indicated proteins in A549 cells treated with or without CTSLi; GAPDH as loading control; quantification from four independent biological replicates in Supplementary Fig. 16B. Samples derive from the same experiment; different gels for CTSL and another for N-Cad, Bnip3, GAPDH, were processed in parallel. Migration (G) and invasion (H) assays for A549 cells treated with or without CTSLi. Bar graphs represent quantified gap closure (G) and invaded cells (H); biological triplicates per group. WB of indicated proteins in A549 cells transfected with si-CTSL (I) or si-BNIP3 (J); quantification from four independent biological replicates in Supplementary Fig. 16E, F. For I, samples derive from the same experiment; different gels for CTSL and another for N-Cad, Bnip3, GAPDH, were processed in parallel. For J, samples derive from the same experiment; different gels for CTSL, N-cad, GAPDH, and another for BNIP3 were processed in parallel. Quantified migration after si-CTSL (K) or si-BNIP3 (L) transfection; biological triplicates per group. M Working model: combined αPD-L1 and CTSLi therapy suppresses tumorigenesis. For B, one-sided Fisher’s exact test; for C, log-rank test; for D, GSEA; for G, H, K, L, data are mean ± SEM, two-sided unpaired t-test. Source data are provided as a Source Data file.
In TB mice, lung tumors showed coordinated upregulation of Ctsl and Bnip3 alongside N-cadherin (N-cad), a key driver of epithelial-to-mesenchymal transition (EMT) (Fig. 6E and Supplementary Fig. 16A). Whereas αPD-L1 monotherapy had only a modest and nonsignificant effect on N-cad, leaving Ctsl and Bnip3 levels unchanged, addition of CTSLi to αPD-L1 markedly suppressed Ctsl, Bnip3, and N-cad expression (Fig. 6E and Supplementary Fig. 16A). This indicates that the CTSL–BNIP3 axis promotes the EMT process independently of αPD-L1 signaling. Consistently, CTSLi treatment of human LUAD A549 cells reduced BNIP3 and N-cad levels in human LUAD A549 cells (Fig. 6F and Supplementary Fig. 16B) and significantly impaired both migration and invasion of A549 and KrasIKKαL cells (Fig. 6G, H and Supplementary Fig. 16C, D). Genetic knockdown of CTSL in A549 cells recapitulated these effects, downregulating BNIP3 and N-cad expression (Fig. 6I and Supplementary Fig. 16E), whereas BNIP3 silencing decreased N-cad without altering CTSL (Fig. 6J and Supplementary Fig. 16F). Functionally, silencing either CTSL or BNIP3 markedly inhibited A549 cell migration (Fig. 6K, L). Together, these data suggest that CTSL promotes LUAD cell motility and invasiveness through a BNIP3-dependent EMT program. Targeting CTSL could additively enhance anti-tumor efficacy by engaging complementary pathways alongside αPD-L1 treatment (Fig. 6M).
CTSL induces mitochondrial dysfunction and muscle wasting via BNIP3
To investigate the involvement of the CTSL-BNIP3 axis in muscle physiology, we analyzed skeletal muscle transcriptomic data from 803 healthy individuals in the GTEx RNA-seq database. Despite being derived from non-cachectic tissue, this large-scale cohort revealed a strong positive correlation between CTSL and BNIP3 expression (Supplementary Fig. 17A). Notably, individuals with high co-expression of CTSL and BNIP3 exhibited muscle gene expression profiles that closely resembled those of TB mice (Figs. 5G, 7A). Functional enrichment analysis revealed significant alterations in biological processes related to muscle metabolism and proteostasis in the CTSL-BNIP3-high group (Fig. 7B). Particularly, the CTSL-BNIP3-high group showed coordinated upregulation of proteasomal degradation-related genes and downregulation of mitochondrial oxidative phosphorylation genes (Fig. 7C and Supplementary Fig. 17B), implying that the CTSL-BNIP3 axis may impair mitochondrial function and contribute to muscle wasting.
A Human PPI network of CTSL-BNIP3 neighbors colored by log2 FC in skeletal muscle from CTSL-BNIP3-high versus -low individuals (GTEx). B GSEA of DEGs in GTEx muscle: top five positively and negatively enriched pathways (NES, normalized enrichment scores). C Representative GSEA plots for WP_PROTEASOME_DEGRADATION (top) and WP_ELECTRON_TRANSPORT_CHAIN_OXPHOS_SYSTEM_IN_MITOCHONDRIA (bottom) in CTSL-BNIP3-high versus -low muscles. D Representative WB of indicated proteins in TB muscle. Two mice per group are shown (n = 4 per group); quantification of all four mice in Supplementary Fig. 18A. Samples derive from the same experiment; different gels for Ctsl, another for Bnip3, another for vinculin, were processed in parallel. E WB and quantification of Bnip3 in C2C12 myotubes 24 h after si-Ctsl or si-Cont transfection; biological triplicates for each group. Vinculin as a loading control. F C2C12 myotube diameter after 24 h transfection with si-Bnip3 or si-Cont and 72 h co-culture with CD8+ T cells in CM; 12 myotubes per well, biological triplicates for each group. Relative mtDNA copy number in GA muscle from TB mice (G, n = 8); in C2C12 myotubes co-cultured with CD8+ T cells in CM plus or minus CTSLi (H, biological triplicates for each group); and transfected with si-Ctsl, si-Bnip3, or si-Cont then co-cultured with CD8+ T cells in CM (I, biological triplicates for each group). C2C12 myotube diameter (J) and mtDNA copy number (K) after co-culture with CD8+ T cells in CM plus or minus mouse recombinant Gzma (10 ng/ml); biological triplicates for each group. L Working model illustrating how the CTSL-BNIP3 network promotes muscle wasting in cooperation with cytotoxic T cells. For B and C, GSEA for functional enrichment analysis; for E–K, data are mean ± SEM, two-sided unpaired t-test. Source data are provided as a Source Data file.
Consistent with our findings from human datasets, TB muscle exhibited coordinated upregulation of Ctsl and Bnip3 (Fig. 7D and Supplementary Fig. 18A). Importantly, while αPD-L1 further elevated Bnip3 expression, CTSLi administration reversed this increase (Fig. 7D). Moreover, knockdown of Ctsl reduced Bnip3 expression in C2C12 myotubes (Fig. 7E), and silencing Bnip3 in C2C12 myotubes significantly attenuated CD8+ T cells-induced atrophy (Fig. 7F and Supplementary Fig. 18B, C), implying a role for the CTSL-BNIP3 axis in T cell-mediated muscle wasting, possibly through the promotion of mitochondrial dysfunction.
Mitophagy regulates mitochondrial DNA (mtDNA) copy number, critical for muscle function84,85,86. We observed a marked reduction in mtDNA copy number in TB muscle, which was exacerbated by αPD-L1 and restored by CTSLi (Fig. 7G). Notably, mtDNA copy number (Fig. 7G) inversely correlated with both Ctsl expression and CD8+ T cell infiltration (Fig. 4I–K), linking cytotoxic T cell activity to mitochondrial loss. In C2C12 co-cultures, CD8+ T cells reduced both myotube diameter and mtDNA copy number, and these two parameters were rescued by CTSLi (Fig. 7H and Supplementary Fig. 18D, E). Moreover, siRNA knockdown of either Ctsl or Bnip3 similarly restored mtDNA copy number in the presence of CD8+ T cells (Fig. 7I), supporting that the CTSL-BNIP3 axis underlies T cell-mediated mitochondrial dysfunction.
To explore how the CTSL-BNIP3 axis and CD8+ T cells converge on mitochondrial dysfunction, we examined the contribution of GZMA, a cytotoxic mediator known to induce mitochondrial damage87. Recombinant Gzma treatment of C2C12 myotubes reduced both diameter and mtDNA copy number even without CD8+ T cells, and these effects were abolished by CTSLi (Fig. 7J, K and Supplementary Fig. 18F). Given our single-cell data showing expression of Gzma, Gzmb, and Prf1 in CD49a+ CD8+ T cells (Fig. 2D), we propose that GZMA cooperates with the CTSL-BNIP3 axis to deplete mtDNA and drive muscle wasting.
Taken together, during cachexia, CTSL upregulation in muscle induces aberrant MHC-I expression, recruiting TRM-like CD49a+ CD8+ T cells. These T cells release GZMA, which, together with BNIP3, leads to mtDNA depletion, mitochondrial dysfunction, and ultimately muscle atrophy (Fig. 7L). Thus, CTSL promotes muscle wasting through complementary mechanisms, not only by proteasomal degradation and autophagy45,46, but also by T cell-mediated mitochondrial injury.
Discussion
Our study reveals a paradox of αPD-L1 immunotherapy: while it potently suppresses tumor growth, it exacerbates systemic cachexia and muscle wasting. We show that TB mice accumulate a unique subset of TRM-like CD49a+ CD8+ T cells in skeletal muscle. These cells bear a potent cytotoxic signature and directly induce muscle atrophy. αPD-L1 further amplifies their infiltration, indicating that cytotoxic T cells not only mediate anti-tumor response but also provoke muscle-specific irAEs. Although CD8+ T cell depletion rescues muscle mass, it abolishes tumor control, highlighting a critical therapeutic trade-off.
By integrating transcriptomic profiles from muscle and tumor, we identify CTSL as a dual-action therapeutic target. Its pharmacologic inhibition not only preserves muscle mass but also enhances the anti-tumor efficacy of αPD-L1, which was further validated with genetic CTSL knockdown in vitro. Although TNF inhibitors have been shown to mitigate irAEs without compromising ICI efficacy88, CTSLi outperformed TNFα blockade in both tumor suppression and muscle preservation (Supplementary Fig. 19A–D). CTSLi co-treatment normalized muscle Tnf-α levels and further lowered Il-6 below baseline (Supplementary Fig. 11D, E), implying that IL-6 drives intrinsic cancer-induced muscle wasting, whereas TNF-α mediates ICI-related toxicity. Therefore, CTSL inhibition offers a broader therapeutic index, uncoupling anti-tumor immunity from muscle-specific irAEs.
Cancer cachexia is orchestrated by complex tumor-host crosstalk among CNS, skeletal muscle, liver, and adipose tissues12,89,90. Aberrant tumor-muscle communication disrupts metabolic homeostasis and accelerates muscle wasting, ultimately impairing clinical outcomes91,92, often mediated by immune dysregulation through pro-inflammatory cytokines and direct immune cell damage93,94. While previous studies implicate tumor-educated macrophages and muscle-infiltrating neutrophils in cancer-associated muscle atrophy via IL-1α, IL-6, lipocalin-2, or ferroptosis25,26, our findings extend this paradigm to CD49a+ CD8+ T cells. We show that CTSL upregulates MHC-I expression on muscle fibers, normally low in healthy muscle but inducible under inflammation74,75,95,96, thereby recruiting cytotoxic CD49a+ CD8+ T cells. CTSLi suppresses muscle MHC-I and CD8+ T cell infiltration, breaking this pathogenic loop.
Interestingly, we observed earlier and more severe adipose tissue loss than muscle loss during cachexia progression, similar to prior works57,97,98. Notably, this was accompanied by CD8+ T cell infiltration into epididymal fat (Supplementary Fig. 20A, B), suggesting a shared, systemic immune-driven wasting mechanism across metabolic organs. The heightened adipose vulnerability may reflect organ-specific cytokine sensitivity or immune microenvironments.
In TB tumors, CTSL drives metastatic potential through tumor-intrinsic mechanisms, rather than by modulating the immune microenvironment. Although CTSL can play tumor-suppressive roles in certain contexts36,37,40, its overexpression more commonly correlates with poor clinical outcomes across diverse malignancies35,99,100,101,102. CTSL expression is regulated by transcription factors such as signal transducer and activator of transcription 3 (STAT3)103,104,105. In turn, CTSL can activate oncogenic pathways, including STAT3100,106 and NF-κB32,107, establishing potential feed-forward loops. Through these interconnected circuits, CTSL contributes to tumor growth, chemoresistance, and metastasis99,100,101,102,108,109.
We further show that BNIP3 serves as a context-dependent effector of CTSL signaling, driving distinct outcomes in tumor versus muscle. In tumors, the CTSL–BNIP3 axis enhances metastatic potential via EMT signaling, whereas in skeletal muscle, it promotes mtDNA depletion and exacerbates T cell-mediated wasting. Importantly, GZMA released by CD8+ T cells cooperates with BNIP3 to impair mitochondrial integrity, linking cytotoxic activity directly to muscle wasting87. These tissue-specific effects likely arise from contrasting immune landscapes (Figs. 6M, 7L). The immunosuppressive TME limits cytotoxic T cell activity, while chronically inflamed muscle, normally immune-privileged, becomes permissive to CD8+ T cell infiltration110,111,112. Thus, in cancer, sustained inflammation may facilitate aberrant CD8+ T cell-mediated muscle damage via GZMA release113,114,115,116, with BNIP3 serving as a critical mediator of the resulting mitochondrial dysfunction and muscle atrophy.
Several questions remain. First, it is not yet clear whether muscle-infiltrating CD8+ T cells are antigen-specific or activated in an antigen-independent, cytokine-driven manner (e.g., IL-15)114,117,118. Elevated CTSL in muscle could generate novel peptide fragments, potentially creating neoantigens that drive T cell responses119. Alternatively, bystander activation via cytokines such as IL-15 may instead explain the expansion of CD49a+ CD8+ T cells, which express the bystander-marker Entpd1120. Second, the upstream signals that induce CTSL and how CTSL regulates BNIP3 remain to be defined. STAT3, activated by IL-6, is a compelling candidate regulator of CTSL103,104,121; BNIP3 is a well-known hypoxia-inducible factor 1α target79,122, implicating hypoxia signaling in the CTSL–BNIP3 axis. Deciphering these transcriptional and post-translational regulatory mechanisms will be a priority for future work.
In conclusion, our work uncovers cachexia as a potential muscle-specific irAE driven by CD49a+ CD8+ T cells under ICI therapy. We identify CTSL as a dual-action therapeutic target. CTSL inhibition preserves muscle mass, mitigates irAEs, and enhances the anti-tumor efficacy of ICI, offering a clinically translatable strategy to uncouple ICI efficacy from debilitating muscle toxicity.
methods
Ethics statement
This study was approved by the Institutional Review Boards of the Yonsei University Dental Hospital (IRB# 2-2024-0048) and Pusan National University Hospital (PNUH) Ethics Committee (IRB# 2306-016-128). Written informed consent was obtained from all participants for the use of their data in research and was in compliance with the Declaration of Helsinki. All mice procedures in this study were approved by Yonsei University Institutional Animal Care and Use Committee (IACUC; protocol# 2023-0074) and conducted in accordance with international guidelines.
Human patient data
Electronic health records (EHRs) were collected from two tertiary hospitals in South Korea: Yonsei University Severance Hospital and PNUH. Patients diagnosed with various cancers who received αPD-1 (pembrolizumab, nivolumab), αPD-L1 (atezolizumab, durvalumab), or other anti-cancer drugs (gemcitabine, docetaxel, doxorubicin, pemetrexed, bevacizumab, trastuzumab) were included. The index date was defined as the first day of drug administration. Pre-treatment body weight was measured within 30 days before the index date, while post-treatment body weight was defined as the lowest weight recorded within 18 months following the index date. Differences between pre- and post-treatment weights were analyzed using a paired t-test, with statistical significance set at p < 0.05. No gender-specific recruitment criteria were applied.
Mice
Six-week-old male C57BL/6 (C57BL/6NCrlOri) were purchased from Orient Bio (Seongnam, Korea) and acclimated for one week before experiments. Mice were housed in specific-pathogen-free conditions under a 12-h light/12-h dark cycle, and climate-controlled conditions, with ad libitum access to standard chow and water.
Orthotopic tumor transplantation
The lung tumor-bearing (TB) mouse model was established using KrasIKKαL cells52,53. Cells were maintained in RPMI1640 medium (Gibco, Waltham, MA, USA) supplemented with 10% fetal bovine serum (FBS, Gibco), 1% antibiotic–antimycotic mixture (Gibco), and 100 mg/L Normocin (InvivoGen, San Diego, CA, USA). Seven-week-old male C57BL/6 mice were randomized into treatment groups, ensuring comparable mean body weights. Each mouse received an intratracheally (i.t.) instillation of 5 × 106 KrasIKKαL cells in 50 μL phosphate-buffered saline (PBS). Body weight and food intake were monitored weekly. For the Lewis lung carcinoma (LLC) model, an identical protocol was followed using 5 × 106 LLC cells. Mice were sacrificed when any group reached ≥ 10% loss in initial body weight (IBW), in accordance with our IACUC-approved humane endpoint and the standard cachexia threshold in murine studies54,55,56,57. At necropsy, lungs, epididymal white adipose tissues (eWATs), and skeletal muscles (GA and TA) were harvested for downstream analyses, including weight measurement, histology, Western blotting (WB), flow cytometry, RNA-seq, or scRNA-seq.
Treatment with αPD-L1 and αCD8a in TB mice
Beginning three days after i.t. tumor cell instillation, mice received intraperitoneal (i.p.) injection of 10 mg/kg anti-mouse PD-L1 antibody (αPD-L1, clone 10 F.9G2™, BE0101, BioXCell, Lebanon, NC, USA) with or without 10 mg/kg anti-mouse CD8α antibody (αCD8a, clone 2.43, BE0061, BioXCell) three times a week. Control received 10 mg/kg isotype IgG (rat IgG2b, clone LTF-2, BE0090, BioXCell).
Combination therapy with αPD-L1 and CTSLi in TB mice
Starting 3 days after tumor cell injection, mice received i.p. injections of 10 mg/kg αPD-L1 with or without 0.5 mg/kg of Z-Phe-Tyr-CHO (CTSLi, 23249, Cayman, Ann Arbor, MI, USA) three times a week. Control received isotype IgG.
Combination therapy with αPD-L1 and TNF blockade in TB mice
Following the tumor injection, mice received i.p. injections of 10 mg/kg αPD-L1 with or without 6.2 mg/kg of anti-mouse TNF-α antibody (αTNF-α, clone XT3.11, BE0058, BioXCell) three times a week.
Grip strength test
Forelimb grip strength was assessed using a digital grip strength meter (JD-A-22, JEUNGDO BIO&PLANT, Seoul, Korea). Each mouse was gently placed on a mesh bar, allowing it to grasp the bar with its forelimbs. Once the grip was secured, its tail was gently pulled backward in a horizontal plane until it released the bar. The peak force (in newtons, N) was recorded by the meter. Ten consecutive measurements were taken per mouse with brief rest intervals, and the mean of the ten trials was calculated as the absolute grip strength. Values in newtons (N) were converted to grams (1 N = 101.97 g), then normalized to each animal’s IBW (g).
Histology, lung tumor burden, and muscle cross-sectional area measurement
Tissue samples were harvested and fixed in 10% neutral buffered formalin (NBF) for 24 h, and dehydrated (muscle/lung in 70% ethanol; WAT in 20% sucrose). Paraffin-embedded (muscle, lung) or frozen (WAT) sections were prepared by the Avison Biomedical Research Center (ABMRC) at Yonsei University. Then, sections were stained with hematoxylin and eosin (H&E) or Masson’s Trichrome for morphology and fibrosis assessment, and immunohistochemistry (IHC) was performed overnight at 4 °C using primary antibodies (1:100), followed by appropriate secondary detection. Antibodies are listed in Table 1. For IHC quantification, positively stained cells were counted in three randomly chosen fields per section for each mouse under an inverted microscope (Leica DMi1, Leica Biosystems, Wetzlar, Germany). To assess lung tumor burden, lung tumor areas in H&E-stained sections were quantified using ImageJ software (National Institutes of Health, Bethesda, MD, USA) and displayed as a percentage of the total lung area occupied by tumors52,53. To measure muscle cross-sectional area (CSA), H&E-stained muscle sections were measured using ImageJ software. The mean CSA of fibers (at least 100) per field (three fields per section from each mouse) was analyzed, and the average CSA was calculated for each mouse.
Immunofluorescent staining in a human LUAD tissue microarray
Human LUAD tissue microarrays (LC1504) were purchased from US Biomax, Inc. (Rockville, MD, USA). Briefly, tissue sections were incubated overnight at 4 °C with a primary antibody against CTSL (1:100) and BNIP3 (1:100). Then, these sections were washed and stained with the fluorescence-conjugated secondary antibodies for 1 h at room temperature. Antibodies are listed in Table 1. Finally, the sections were mounted with the diamidino-2-phenylindole (DAPI) mounting medium (P36962, Invitrogen).
Fluorescence images were captured on a ZEISS LSM700 confocal microscope (Carl Zeiss, Oberkochen, Germany). For quantitative analysis, the mean fluorescence intensity (MFI) of each marker was measured in all 150 tissue cores (22 adjacent normal lung tissues; 128 LUAD malignant and tumor-adjacent lung tissues, representing 50 cases) using ZEISS ZEN 3.11 software (Carl Zeiss). The overall average MFI across all cores was 5.50549 for CTSL and 15.19499 for BNIP3. Cores were classified based on MFI: CTSL MFI ≥ 5.50549 as CTSL-high; CTSL MFI < 5.50549 as CTSL-low; BNIP3 MFI ≥ 15.19499 as BNIP3-high; BNIP3 MFI < 15.19499 as BNIP3-low. We then calculated the proportions of BNIP3-high and BNIP3-low cores within the CTSL-high and CTSL-low groups and plotted them in a bar graph. Fisher’s exact test was used to assess the statistical association between CTSL and BNIP3 expression levels.
Isolation of tissue-infiltrating CD45+ cells
Freshly isolated tissues were finely minced and enzymatically digested in RPMI1640 medium containing 0.3 mg/ml Liberase TL (5401119001, Sigma-Aldrich, St. Louis, MO, USA) and 0.4 μg/ml deoxyribonuclease I (DN25, Sigma-Aldrich) for 2 h at 37 °C with gentle agitation. Following enzymatic digestion, cells were mechanically disaggregated using 18 and 21 G syringes (Korea Vaccine Co., Ltd, Seoul, Korea) and passed through a 100-μm cell strainer (93100, SPL Life Sciences, Pocheon, Korea) to obtain a single-cell suspension. Red blood cells (RBCs) were depleted with ammonium-chloride-potassium (ACK) lysis buffer (A10492-01, Gibco), and CD45+ immune cells were positively selected using magnetic beads (130-052-301, Miltenyi Biotec, Bergisch Gladbach, Germany) in accordance with the manufacturer’s instructions. The isolated CD45+ cells were subsequently subjected to flow cytometry analysis or scRNA-seq.
Flow cytometry
CD45+ cells were counted using LUNA-II™. The Ultimate Brightfield Cell Counter (Logos Biosystems, Anyang, Korea) and stained with fluorescence-conjugated antibodies using a total 1 × 106 (muscle) or 5 × 107 (lung) of CD45+ cells. The following antibodies (listed in Table 2) were used: for lymphocytes, CD45 (1:100), CD3ε (1:100), CD8α (1:200), CD49a (1:100), CD4 (1:100), Foxp3 (1:100); for myeloid-derived suppressor cells (MDSCs), CD45 (1:100), CD11b (1:200), Ly6G (1:100); all from BioLegend (San Diego, CA, USA). Following surface staining, Foxp3 was detected by intracellular staining using the Foxp3/Transcription Factor Staining Buffer Set (00-5521-00, eBioscience, San Diego, CA, USA) per the manufacturer’s instructions. Data were acquired on a BD FACS Symphony A5 (BD Biosciences, Franklin Lakes, NJ, USA) and analyzed with FlowJo™ Software (BD Biosciences).
WB analysis
Cells and tissues were lysed using RIPA buffer (9806, Cell Signaling Technology, Danvers, MA, USA) containing 1 mM phenylmethylsulfonyl fluoride (PMSF, PS2064, Gabiochem, Norcross, GA, USA) and protease inhibitor cocktail (11836170001, Roche, Basel, Switzerland) for 1 h on ice. Lysates were centrifuged at 15,000×g for 15 min, and protein concentrations were determined. Protein lysates (30–50 μg) were mixed with protein gel-loading buffer (1610737, Bio-Rad Laboratories, Hercules, CA, USA), heated at 95 °C for 5 min, and separated on 10–15% sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS–PAGE). The resolved proteins were transferred to polyvinylidene difluoride (PVDF) membranes (Millipore, Burlington, MA, USA), blocked with 5% skim milk, and probed with primary antibodies. Then, secondary antibodies were applied, and proteins were detected using a chemiluminescence reagent (Biomax, Seoul, Korea) and visualized using an image analyzer (Amersham, GE Healthcare, Little Chalfont, UK). Antibodies are listed in Table 1. Band intensities were quantified using ImageJ software.
Enzyme-linked immunosorbent assay for CTSL
Mouse Cathepsin L ELISA Kit (NBP2-89172, Novus Biologicals, Centennial, CO, USA) was utilized to determine the concentrations of CTSL in tissue lysates, according to the manufacturer’s protocols. Briefly, the tissue lysates were added to the antibody-coated wells and incubated for 2 h at 37 °C. Then, the wells were washed and then incubated with biotinylated detection antibody for 1 h at 37 °C. After rinsing, HRP-conjugated streptavidin was added for 30 min at 37 °C. Following a final wash step, substrate solution was added to each well and incubated for 15 min. Absorbance was measured at 450 nm using a microplate reader. CTSL concentration was interpolated from a standard curve and normalized to the mass of the starting tissue.
Quantitative real-time polymerase chain reaction (qRT-PCR)
Total RNA was extracted from tissues and cells using the TRIzol reagent (15596026; Invitrogen). The cDNA was synthesized using a cDNA synthesis kit (25081, iNtRON Biotechnology, Seongnam, Korea). qRT-PCR was performed to quantify gene expression using Power SYBR™ Green PCR Master Mix (4367659, Applied Biosystems, Waltham, MA, USA) on a QuantStudio3 Real-Time PCR System (Applied Biosystems) according to the manufacturer’s protocols. Primers are listed in Table 3. Gene expression levels were normalized to the housekeeping genes Gapdh or Vinculin. Data analysis was conducted using the ΔΔCt method, and the results were expressed as fold-changes in mRNA expression relative to control samples.
DNA extraction and mtDNA copy number measurement
Genomic DNA was extracted from cells or tissues using AccuPrep Genomic DNA Extraction Kit (Bioneer), according to the manufacturer’s instructions. To quantify mtDNA copy number, qRT-PCR was performed using primers specific for mtDNA (Cytochrome c oxidase subunit 2, Cox2). As an internal standard, the ribosomal protein S18 (Rsp18) gene was amplified using primers listed in Table 3. The amount of mtDNA was adjusted to the amount of nuclear DNA.
Ex vivo co-culture of C2C12 myotubes with splenic T cells
C2C12 myoblasts (4 × 104 cells per well, CRL-1772, ATCC, Manassas, VA, USA) were plated in six-well plates and cultured in DMEM media (Gibco) containing 10% FBS and 1% antibiotic-antimycotic mixture at 37 °C with 5% CO2. To induce myotube differentiation, cells were incubated in the differentiation medium (DM, DMEM medium containing 2% horse serum) for 5 days to promote the formation of multinucleated myotubes. Following differentiation, the C2C12 myotubes were cultured with or without KrasIKKαL cell-derived conditioned medium (CM, diluted 1:4 into DM) to stimulate muscle wasting conditions. CD8+ T cells were isolated from the spleen and lymph nodes of CON or TB mice. After tissue disaggregation and RBC depletion, CD8+ T cells were positively selected using the EasySep Mouse CD8α Positive Selection Kit (18953, STEMCELL Technologies, Vancouver, BC, Canada) according to the manufacturer’s protocol. For co-culture with CD4+ T cells, CD4+ T cells were positively selected using the EasySep Mouse CD4+ T cell Isolation Kit (19852, STEMCELL Technologies). Isolated CD8+ or CD4+ T cells (4 × 105 cells per well) were co-cultured with C2C12 myotubes in either DM or CM for 96 h. Myotubes were imaged using an ECLIPSE Ts2 with an LED microscope (Nikon Corporation, Tokyo, Japan), and the myotube diameter was measured using ImageJ software. Randomly selected 12 myotubes per well were analyzed from biological triplicates for each group, and the average diameter was used for statistical analysis.
C2C12 siRNA knockdown assay
Following differentiation, C2C12 myotubes were transfected with 20 nM of siRNAs against Ctsl (SR411229, OriGene Technologies, Rockville, MD, USA) or Bnip3 (SR404627, OriGene Technologies) using Lipofectamine RNAiMax (13778150, Invitrogen) for 24 h. Then, C2C12 myotubes were co-cultured with splenic CD8+ T cells in the presence of either DM or CM for an additional 72 h.
C2C12 myotube treatment with recombinant granzyme A
Following differentiation, C2C12 myotubes were treated with 10 ng/ml of mouse recombinant granzyme A (Gzma, HY-P76377, MedChemExpress, Monmouth Junction, NJ, USA) with or without CTSLi (1 μg/ml) in the presence of either DM or CM for 96 h.
Cell invasion assay
Cell invasion was assessed using a 6.5-mm transwell chamber with an 8.0-μm pore polycarbonate membrane (Corning Coster, Lowell, MA, USA). The lower and upper surfaces of the membrane were coated with 10 μl of gelatin (1 mg/ml in distilled water) and 40 μl of Matrigel (1 mg/ml in PBS), respectively. Cells (A549, 1 × 105 cells per well; KrasIKKαL, 5 × 104 cells per well) were seeded into the insert in media containing 5% FBS with or without 10 μg/ml of CTSLi. Cells were allowed to invade for 24 h. After incubation, invasive cells on the bottom of the membrane were stained with Mayer’s hematoxylin (Cancer Diagnostics Inc., Durham, NC, USA). Non-invaded cells on the upper membrane were removed using cotton swabs. The number of invaded cells was counted under a light microscope, and the percentage of invasion was calculated by comparing the number of invaded cells to the control.
Cell migration assay
Cell migration was evaluated using a Culture-Insert (ibidi, Gräfelfing, Germany) to create a 500-μm cell-free gap. Cells (7 × 103 cells per well) were seeded into the insert, which was removed after cell attachment to leave a 500-μm gap. Cells were then treated with CTSLi (10 μg/ml) for 6 h (KrasIKKαL cells) or 24 h (A549 cells). For gene silencing, A549 cells were seeded and then transfected with 20 nM siRNAs against CTSL (SR319840, OriGene Technologies) or BNIP3 (SR319494, OriGene Technologies) using Lipofectamine RNAiMax for 48 h following the manufacturer’s protocol. The gap closure, representing cell migration, was measured as the change in distance between the initial and final positions of the cell using ImageJ software.
Preprocessing of bulk RNA-seq
RNA-seq was performed on the following samples: TA muscles from CON mice (n = 5), TB mice treated with IgG (n = 4), αPD-L1 (n = 3), or αCD8a (n = 3); GA muscles from CON mice (n = 3), TB mice (n = 3); lung tissues from CON mice (n = 3); and lung tumors from TB mice (n = 5). Approximately 10 µg of total RNA was isolated from each sample using TRIzol reagent. Then, mRNA libraries were prepared using the MGIEasy RNA Directional Library Prep Kit (MGI Tech Co., Ltd., Shenzhen, China) and sequenced on the MGISEQ-2000 system (MGI Tech Co., Ltd.) to generate 100 bp paired-end reads. The quality of the raw sequencing data (FASTQ files) was assessed using FastQC (v0.11.9). Adapter sequences, low-quality bases, and reads with excessive ambiguous bases were removed using TrimGalore (v0.6.6). Trimmed reads were aligned to the Mus musculus reference genome (GRCm39) using the STAR aligner (v2.7.9a)123. Transcript abundance was quantified for each gene using RSEM (v1.3.3), and reported as read counts or transcripts per million (TPM)124.
Differential gene expression analysis and functional annotation
Differential gene expression between sample groups (e.g., TB vs. CON) was performed using DESeq2 (v4.3.1). The following criteria were applied to identify differentially expressed genes (DEGs): αPD-L1 vs. IgG: p-value < 0.005; αCD8a vs. IgG or IgG vs. CON: false discovery rate (FDR)-adjusted p-value < 0.01, absolute log2 fold-change (FC) > 1; TB vs. CON: FDR-adjusted p-value < 0.01, absolute log2 FC > 1.5. Functional enrichment analysis was conducted using Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) databases. A hypergeometric test was implemented through the clusterProfiler R package (v4.10.1) to assess the overrepresentation of specific biological processes, molecular functions, and pathways among DEGs. Additionally, gene set enrichment analysis (GSEA) was performed using fgsea (v1.28.0) to identify pathways significantly overrepresented among the top- or bottom-ranked genes based on the differential gene expression analysis. Parameters of GSEA were set as follows: minSize = 10, maxSize = 500, and nperm = 10,000. Single-sample gene set enrichment analysis (ssGSEA) was conducted to evaluate the overall expression of metastasis signature genes (MsigDB, C2: ALONSO_METASTASIS_UP) for individual lung tumor samples (TCGA and TB mice) via the R package gsva (v1.50.5).
Sample preparation for scRNA-seq
Single-cell suspensions from lung tumors and skeletal muscles of TB mice were processed using CD45+ cell isolation protocols as mentioned. A total of 4 × 106 CD45+ cells were collected from skeletal muscle and 2 × 108 CD45+ cells were obtained from lung tumors. The scRNA-seq was performed by GENINUS Inc. (Seoul, Korea) on the BGI MGISEQ-2000 platform. Barcoded libraries for scRNA-seq were generated using Chromium Single Cell 3’ v2 Reagent Kits (10 × Genomics, Pleasanton, CA, USA) and sequenced on the Illumina NovaSeq 6000 platform (San Diego, CA, USA).
scRNA-seq data analysis
Raw sequencing reads were aligned to the Mus musculus reference genome (GRCm39) using the CellRanger (v7.0.1). The gene-barcode count matrix was analyzed using the Seurat R package (v5.0.3). Quality control filters were applied to exclude cells expressing fewer than 300 or more than 4000 genes to avoid low-quality cells and potential doublets. Cells with high mitochondrial gene expression (>30%) were also discarded, as these likely indicated damaged cells. To mitigate batch effects, datasets from lung and GA muscle tissues were integrated using Seurat’s standard integration pipeline. After quality control, 11,348 out of the original 14,028 cell barcodes were retained for downstream analysis. Data normalization was performed using the NormalizeData function, which log-normalizes gene expression values, scales them by a factor of 10,000, and applies a log-transformation. Principal component analysis (PCA) and uniform manifold approximation and projection (UMAP) were used for dimensionality reduction and to visualize distinct cell populations. The optimal number of principal components (PCs) was determined using ElbowPlot, with 24 PCs selected for further analysis. Clustering was performed using the FindNeighbors and FindClusters functions to group cells based on similar gene expression profiles. Marker genes for each cluster were identified using the FindAllMarkers function, which selects genes with a minimum log2 FC of 0.25 (FDR-adjusted p-value < 0.05) and detection in at least 25% of cells in the cluster. Cell type annotations were assigned based on well-established marker genes from literature and resources like GPT-4o, GPTCelltype (v1.0.1), and Azimuth (https://azimuth.hubmapconsortium.org/)125.
Gene network analysis
To identify functional gene modules within a given gene set, we utilized protein-protein interaction (PPI) data from the STRING database (v12.0) for Mus musculus71. STRING integrates various sources of evidence, including co-expression, co-occurrence, literature mining, pathway databases, and experimental data. Accordingly, links in the resulting network represent predicted or known functional associations rather than direct physical interactions.
To identify densely connected subnetworks, we applied the Walktrap community detection algorithm (cluster_walktrap) implemented in the igraph R package (v2.0.3). Functional enrichment analysis of selected gene modules (e.g., Community 1 in Fig. 3B) was performed using the enrichKEGG function from the clusterProfiler R package (v4.10.1).
For network visualization, the igraph object was converted to a Cytoscape-compatible format using the createNetworkFromIgraph function in the RCy3 R package (v2.26.0), and rendered in Cytoscape (v3.10.2). Interaction pairs with medium confidence (combined score >500) were retained for PPI network construction, unless otherwise specified. To map human gene expression data (TCGA or GTEx) onto the mouse-derived network, mouse gene symbols were converted to their human orthologs using the homologene R package (v1.4.68.19.3.27).
Public transcriptome data analysis
To explore gene expression patterns in different cachectic models, publicly available mouse transcriptome data were retrieved from the GEO database (http://www.ncbi.nlm.nih.gov/geo/) using the keyword ‘cachexia’, yielding four relevant datasets: GSE107470, GSE51931, GSE144567, and GSE114820 (listed in Supplementary Fig. 8C). Given the technological variability across datasets (microarray and RNA-seq), each underwent independent pre-processing appropriate for its respective platform. For RNA-seq data, raw FASTQ files were retrieved from the European Nucleotide Archive (ENA, https://www.ebi.ac.uk/ena/) and processed according to the RNA-seq preprocessing protocol outlined earlier. Differential gene expression between the two groups was assessed using the Wald test from the R package DESeq2. For microarray data, normalized gene expression profiles for each sample were obtained using the R package GEOquery (v2.72.0). Differential gene expression between two groups was evaluated using the moderated t-test implemented in the eBayes function of the R package limma (v3.60.4). To assess the clinical relevance of CTSL expression in human cancer cachexia, human muscle transcriptome data were retrieved from publicly available repositories, the GEO database (GSE133523) and EMBL-EBI ArrayExpress (E-MTAB-12781). Raw FASTQ files were preprocessed by trimming low-quality reads and adapter sequences, mapping the cleaned reads to the human reference genome (GRCh38), and quantifying gene expression in TPM using the RSEM (v1.3.3) for individual samples.
TCGA transcriptome data analysis
RNA-seq data from tumor tissues covering 33 cancer types were retrieved from the TCGA legacy gene expression dataset using the GDCquery function within the R package TCGAbiolinks (v2.32.0). The dataset includes read counts and TPM values for 19,969 protein-coding genes across individual patient samples. For survival analysis, patients were stratified into two groups based on CTSL expression levels (high and low). To estimate the overall survival difference between groups, Kaplan–Meier (K–M) survival curves were generated using the survfit function from the R package survival (v3.6.4), and statistical significance was evaluated using the log-rank test. Cox proportional hazard regression models were constructed using the coxph function in R, and hazard ratios (HRs) were derived to estimate the corresponding relative risk.
To investigate transcriptional alterations associated with the CTSL-BNIP3 axis specifically in LUAD cohorts, tumor samples were stratified based on expression levels of CTSL and BNIP3. To define CTSL-high and CTSL-low groups, we used the median CTSL expression value across the individual TCGA cohorts as the cutoff: individuals with expression values above the median were classified as CTSL-high, while those with expression values below the median were classified as CTSL-low. The same approach was applied to define BNIP3-high and BNIP3-low groups. The intersection of the CTSL-high and BNIP3-high subsets defined the CTSL-BNIP3-high group, whereas the overlap between CTSL-low and BNIP3-low samples defined the CTSL-BNIP3-low group. Differential gene expression analysis between the CTSL-BNIP3-high and CTSL-BNIP3-low groups was performed using the DESeq2 package in R. Subsequent pathway enrichment analysis was conducted using GSEA with gene sets curated from the WikiPathways database.
Genotype-tissue expression (GTEx) data analysis
Human skeletal muscle RNA-seq data were obtained from the genotype-tissue expression (GTEx) project (v8 release) via the GTEx portal (https://gtexportal.org/). A total of 803 skeletal muscle samples from healthy individuals were downloaded along with corresponding gene expression quantification files (TPM and read count formats). Analogous to the TCGA LUAD analysis, samples were stratified based on CTSL and BNIP3 expression levels. The top 50% and bottom 50% of samples for each gene were categorized as high and low, respectively. The intersection of CTSL-high and BNIP3-high samples was defined as the CTSL-BNIP3-high group, while the intersection of CTSL-low and BNIP3-low samples comprised the CTSL-BNIP3-low group. Differential gene expression analysis between the two groups was performed using DESeq2, followed by GSEA using WikiPathways gene sets to identify enriched biological pathways.
Statistics and reproducibility
All experiments were performed with at least three independent biological replicates, and consistent results were obtained across replicates, unless stated otherwise. All statistical parameters, including the exact number of samples (n) for each group, measures of dispersion and precision (reported as mean ± standard error of the mean, [SEM]), and statistical significance, are presented in the figures and figure legends. Statistical significance was determined using a two-sided unpaired t-test for two-group comparisons, one-way analysis of variance (ANOVA) for multiple group comparisons, and the log-rank test for survival analysis, unless stated otherwise. Equivalence of body weight change between treatment groups was assessed using the two one-sided test (TOST) with an equivalence margin defined as 50% of the pooled standard deviation. A p-value of <0.05 was considered statistically significant. The level of significance is indicated in the figures. All statistical analyses were performed using GraphPad Prism (v8.0.2, GraphPad Software, La Jolla, CA, USA) and R software (v4.3.2).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The bulk RNA-seq and scRNA-seq datasets generated in this study have been deposited in the GEO under the following accession codes: GSE278013, GSE278396, and GSE278765. The public datasets reutilized in this study can be accessed via GEO (GSE107470 [https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE107470], GSE51931, GSE144567, GSE114820, and GSE133523) and ArrayExpress (E-MTAB-12781 [https://www.ebi.ac.uk/biostudies/ArrayExpress/studies/E-MTAB-12781?query=e-mtab-12781]). Source data are provided with this paper.
Code availability
The R scripts and corresponding datasets used to generate the figures in this study are available at https://github.com/202397101/Ctsl and have been archived on Zenodo under the https://doi.org/10.5281/zenodo.17102995126.
References
Kraehenbuehl, L., Weng, C. H., Eghbali, S., Wolchok, J. D. & Merghoub, T. Enhancing immunotherapy in cancer by targeting emerging immunomodulatory pathways. Nat. Rev. Clin. Oncol. 19, 37–50 (2022).
Lu, Y., Zhang, X., Ning, J. & Zhang, M. Immune checkpoint inhibitors as first-line therapy for non-small cell lung cancer: a systematic evaluation and meta-analysis. Hum. Vaccin. Immunother. 19, 2169531 (2023).
Maher, V. E. et al. Analysis of the association between adverse events and outcome in patients receiving a programmed death protein 1 or programmed death ligand 1 antibody. J. Clin. Oncol. 37, 2730–2737 (2019).
Martins, F. et al. Adverse effects of immune-checkpoint inhibitors: epidemiology, management and surveillance. Nat. Rev. Clin. Oncol. 16, 563–580 (2019).
Okiyama, N. & Tanaka, R. Immune-related adverse events in various organs caused by immune checkpoint inhibitors. Allergol. Int 71, 169–178 (2022).
Matas-Garcia, A. et al. Emerging PD-1 and PD-1L inhibitors-associated myopathy with a characteristic histopathological pattern. Autoimmun. Rev. 19, 102455 (2020).
Makarious, D., Horwood, K. & Coward, J. I. G. Myasthenia gravis: an emerging toxicity of immune checkpoint inhibitors. Eur. J. Cancer 82, 128–136 (2017).
Schneider, B. J. et al. Management of immune-related adverse events in patients treated with immune checkpoint inhibitor therapy: ASCO Guideline update. J. Clin. Oncol. 39, 4073–4126 (2021).
Tomsitz, D., Ruf, T., S. Zierold, S., French, L. E. & Heinzerling, L. Steroid-refractory immune-related adverse events induced by checkpoint inhibitors. Cancers (Basel) 15, https://doi.org/10.3390/cancers15092538 (2023).
Meza-Valderrama, D. et al. Sarcopenia, malnutrition, and cachexia: adapting definitions and terminology of nutritional disorders in older people with cancer. Nutrients 13, https://doi.org/10.3390/nu13030761i (2021).
Baracos, V. E., Martin, L., Korc, M., Guttridge, D. C. & Fearon, K. C. H. Cancer-associated cachexia. Nat. Rev. Dis. Prim. 4, 17105 (2018).
Altea-Manzano, P., Decker-Farrell, A., Janowitz, T. & Erez, A. Metabolic interplays between the tumour and the host shape the tumour macroenvironment. Nat. Rev. Cancer 25, 274–292 (2025).
Ferrara, M., Samaden, M., Ruggieri, E. & Venereau, E. Cancer cachexia as a multiorgan failure: reconstruction of the crime scene. Front Cell Dev. Biol. 10, 960341 (2022).
Argiles, J. M., Busquets, S., Stemmler, B. & Lopez-Soriano, F. J. Cancer cachexia: understanding the molecular basis. Nat. Rev. Cancer 14, 754–762 (2014).
Patel, H. J. & Patel, B. M. TNF-alpha and cancer cachexia: molecular insights and clinical implications. Life Sci. 170, 56–63 (2017).
Strassmann, G., Fong, M., Kenney, J. S. & Jacob, C. O. Evidence for the involvement of interleukin 6 in experimental cancer cachexia. J. Clin. Investig. 89, 1681–1684 (1992).
Jo, H. et al. Prognostic significance of cachexia in advanced non-small cell lung cancer patients treated with pembrolizumab. Cancer Immunol. Immunother. 71, 387–398 (2022).
Miyawaki, T. et al. Desensitizing effect of cancer cachexia on immune checkpoint inhibitors in patients with advanced NSCLC. JTO Clin. Res. Rep. 1, 100020 (2020).
Yu, Y., Yan, L., Huang, T., Wu, Z. & Liu, J. Cancer cachexia reduces the efficacy of immune checkpoint inhibitors in cancer patients. Aging (Albany, NY) 16, 5354–5369 (2024).
Kichenadasse, G. et al. Association between body mass index and overall survival with immune checkpoint inhibitor therapy for advanced non-small cell lung cancer. JAMA Oncol. 6, 512–518 (2020).
A. Cortellini et al. Baseline BMI and BMI variation during first line pembrolizumab in NSCLC patients with a PD-L1 expression >/= 50%: a multicenter study with external validation. J. Immunother. Cancer 8, (2020). https://doi.org/10.1136/jitc-2020-001403
Park, S. Y., Hwang, B. O. & Song, N. Y. The role of myokines in cancer: crosstalk between skeletal muscle and tumor. BMB Rep. 56, 365–373 (2023).
Sartori, R., Romanello, V. & Sandri, M. Mechanisms of muscle atrophy and hypertrophy: implications in health and disease. Nat. Commun. 12, 330 (2021).
Liu, M. et al. The crosstalk between macrophages and cancer cells potentiates pancreatic cancer cachexia. Cancer Cell https://doi.org/10.1016/j.ccell.2024.03.009 (2024).
Shukla, S. K. et al. Macrophages potentiate STAT3 signaling in skeletal muscles and regulate pancreatic cancer cachexia. Cancer Lett. 484, 29–39 (2020).
Wang, D. et al. LCN2 secreted by tissue-infiltrating neutrophils induces the ferroptosis and wasting of adipose and muscle tissues in lung cancer cachexia. J. Hematol. Oncol. 16, 30 (2023).
Hayashi, Y. et al. IL36G-producing neutrophil-like monocytes promote cachexia in cancer. Nat. Commun. 15, 7662 (2024).
Ng, S. K. et al. The protective effect of cannabinoids against colorectal cancer cachexia through modulation of inflammation and immune responses. Biomed. Pharmacother. 161, 114467 (2023).
Baazim, H. et al. CD8(+) T cells induce cachexia during chronic viral infection. Nat. Immunol. 20, 701–710 (2019).
Bonnans, C., Chou, J. & Werb, Z. Remodelling the extracellular matrix in development and disease. Nat. Rev. Mol. Cell Biol. 15, 786–801 (2014).
Vizovisek, M., Fonovic, M. & Turk, B. Cysteine cathepsins in extracellular matrix remodeling: extracellular matrix degradation and beyond. Matrix Biol. 75-76, 141–159 (2019).
Pranjol, M. Z. I. et al. Cathepsin L-induced galectin-1 may act as a proangiogenic factor in the metastasis of high-grade serous carcinoma. J. Transl. Med 17, 216 (2019).
Olson, O. C. & Joyce, J. A. Cysteine cathepsin proteases: regulators of cancer progression and therapeutic response. Nat. Rev. Cancer 15, 712–729 (2015).
Sudhan, D. R. & Siemann, D. W. Cathepsin L targeting in cancer treatment. Pharm. Ther. 155, 105–116 (2015).
Lu, J. et al. Cathepsin L in lung adenocarcinoma: prognostic significance and immunotherapy response through a multi omics perspective. Cancer Inf. 23, 11769351241307492 (2024).
Reinheckel, T. et al. The lysosomal cysteine protease cathepsin L regulates keratinocyte proliferation by control of growth factor recycling. J. Cell Sci. 118, 3387–3395 (2005).
Dennemarker, J. et al. Deficiency for the cysteine protease cathepsin L promotes tumor progression in mouse epidermis. Oncogene 29, 1611–1621 (2010).
Duarte, L. F. et al. Histone H3.3 and its proteolytically processed form drive a cellular senescence programme. Nat. Commun. 5, 5210 (2014).
S. Chang et al. Hypoxia increases methylated histones to prevent histone clipping and heterochromatin redistribution during Raf-induced senescence. Nucleic Acids Res 53, (2025). https://doi.org/10.1093/nar/gkae1210
Li, Z. et al. KDM4C inhibition blocks tumor growth in basal breast cancer by promoting cathepsin L-mediated histone H3 cleavage. Nat. Genet. https://doi.org/10.1038/s41588-025-02197-z (2025).
Konjar, S. et al. Human and mouse perforin are processed in part through cleavage by the lysosomal cysteine proteinase cathepsin L. Immunology 131, 257–267 (2010).
Honey, K. et al. Thymocyte expression of cathepsin L is essential for NKT cell development. Nat. Immunol. 3, 1069–1074 (2002).
Honey, K., Nakagawa, T., Peters, C. & Rudensky, A. Cathepsin L regulates CD4+ T cell selection independently of its effect on invariant chain: a role in the generation of positively selecting peptide ligands. J. Exp. Med. 195, 1349–1358 (2002).
Nakagawa, T. et al. Cathepsin L: critical role in Ii degradation and CD4 T cell selection in the thymus. Science 280, 450–453 (1998).
Martin, A., Gallot, Y. S. & Freyssenet, D. Molecular mechanisms of cancer cachexia-related loss of skeletal muscle mass: data analysis from preclinical and clinical studies. J. Cachexia Sarcopenia Muscle 14, 1150–1167 (2023).
Tardif, N., Klaude, M., Lundell, L., Thorell, A. & Rooyackers, O. Autophagic-lysosomal pathway is the main proteolytic system modified in the skeletal muscle of esophageal cancer patients. Am. J. Clin. Nutr. 98, 1485–1492 (2013).
Novinger, L. J., Weinzierl, N. M. & Bonetto, A. Diversity in chemotherapy-induced cachexia. Am. J. Physiol. Cell Physiol. 328, C139–C147 (2025).
Tohgo, A., Kumazawa, E., Akahane, K., Asakawa, A. & Inui, A. Anticancer drugs that induce cancer-associated cachectic syndromes. Expert Rev. Anticancer Ther. 2, 121–129 (2002).
Hiensch, A. E. et al. Doxorubicin-induced skeletal muscle atrophy: elucidating the underlying molecular pathways. Acta Physiol. (Oxf.) 229, e13400 (2020).
Pensec, C. et al. Impact of pemetrexed chemotherapy on the gut microbiota and intestinal inflammation of patient-lung-derived tumor xenograft (PDX) mouse models. Sci. Rep. 10, 9094 (2020).
Nutsch, K. et al. TIGIT and PD-L1 co-blockade promotes clonal expansion of multipotent, non-exhausted antitumor T cells by facilitating co-stimulation. Nat. Cancer 5, 1834–1851 (2024).
Song, N. Y. et al. IKKalpha-deficient lung adenocarcinomas generate an immunosuppressive microenvironment by overproducing Treg-inducing cytokines. Proc. Natl Acad. Sci. USA 119, https://doi.org/10.1073/pnas.2120956119 (2022).
Song, N. Y. et al. IKKalpha inactivation promotes Kras-initiated lung adenocarcinoma development through disrupting major redox regulatory pathways. Proc. Natl Acad. Sci. USA 115, E812–E821 (2018).
Arora, G. K. et al. Cachexia-associated adipose loss induced by tumor-secreted leukemia inhibitory factor is counterbalanced by decreased leptin. JCI Insight 3, https://doi.org/10.1172/jci.insight.121221 (2018).
Goncalves, M. D. et al. Fenofibrate prevents skeletal muscle loss in mice with lung cancer. Proc. Natl Acad. Sci. USA 115, E743–E752 (2018).
Kim-Muller, J. Y. et al. GDF15 neutralization restores muscle function and physical performance in a mouse model of cancer cachexia. Cell Rep. 42, 111947 (2023).
Liu, X. et al. Activation of GPR81 by lactate drives tumour-induced cachexia. Nat. Metab. 6, 708–723 (2024).
Shiravand, Y. et al. Immune checkpoint inhibitors in cancer therapy. Curr. Oncol. 29, 3044–3060 (2022).
Buggert, M., Price, D. A., Mackay, L. K. & Betts, M. R. Human circulating and tissue-resident memory CD8(+) T cells. Nat. Immunol. 24, 1076–1086 (2023).
Cheuk, S. et al. CD49a expression defines tissue-resident CD8(+) T cells poised for cytotoxic function in human skin. Immunity 46, 287–300 (2017).
Reilly, E. C. et al. CD49a identifies polyfunctional memory CD8 T cell subsets that persist in the lungs after influenza infection. Front Immunol. 12, 728669 (2021).
Smolders, J. et al. Tissue-resident memory T cells populate the human brain. Nat. Commun. 9, 4593 (2018).
Kumar, B. V. et al. Human tissue-resident memory T cells are defined by core transcriptional and functional signatures in lymphoid and mucosal sites. Cell Rep. 20, 2921–2934 (2017).
Hanninen, A., Maksimow, M., Alam, C., Morgan, D. J. & Jalkanen, S. Ly6C supports preferential homing of central memory CD8+ T cells into lymph nodes. Eur. J. Immunol. 41, 634–644 (2011).
Park, J. Y., DiPalma, D. T., Kwon, J., Fink, J. & Park, J. H. Quantitative difference in PLZF protein expression determines iNKT lineage fate and controls innate CD8 T cell generation. Cell Rep. 27, 2548–2557 e2544 (2019).
Ishihara, S. et al. Dual functions of Rap1 are crucial for T-cell homeostasis and prevention of spontaneous colitis. Nat. Commun. 6, 8982 (2015).
Sebzda, E., Bracke, M., Tugal, T., Hogg, N. & Cantrell, D. A. Rap1A positively regulates T cells via integrin activation rather than inhibiting lymphocyte signaling. Nat. Immunol. 3, 251–258 (2002).
Paiola, M. et al. Osteoarthritis increases the risk of inflammatory arthritis due to immune checkpoint inhibitors associated with tissue-resident memory T cells. J. Immunother. Cancer 13, https://doi.org/10.1136/jitc-2024-010758 (2025).
Kalinoski, H. et al. Injury-induced myosin-specific tissue-resident memory T cells drive immune checkpoint inhibitor myocarditis. Proc. Natl Acad. Sci. USA 121, e2323052121 (2024).
Thomas, M. F. et al. Single-cell transcriptomic analyses reveal distinct immune cell contributions to epithelial barrier dysfunction in checkpoint inhibitor colitis. Nat. Med. 30, 1349–1362 (2024).
Szklarczyk, D. et al. The STRING database in 2023: protein–protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res. 51, D638–D646 (2023).
Gonzalez-Avila, G. et al. Matrix metalloproteinases participation in the metastatic process and their diagnostic and therapeutic applications in cancer. Crit. Rev. Oncol. Hematol. 137, 57–83 (2019).
van de Worp, W. et al. A novel orthotopic mouse model replicates human lung cancer cachexia. J. Cachexia Sarcopenia Muscle 14, 1410–1423 (2023).
Freret, M. et al. Overexpression of MHC class I in muscle of lymphocyte-deficient mice causes a severe myopathy with induction of the unfolded protein response. Am. J. Pathol. 183, 893–904 (2013).
Pavlath, G. K. Regulation of class I MHC expression in skeletal muscle: deleterious effect of aberrant expression on myogenesis. J. Neuroimmunol. 125, 42–50 (2002).
Bhattarai, S. et al. The immunoproteasomes are key to regulate myokines and MHC class I expression in idiopathic inflammatory myopathies. J. Autoimmun. 75, 118–129 (2016).
Talbert, E. E. et al. Modeling human cancer-induced cachexia. Cell Rep. 28, 1612–1622 e1614 (2019).
Maes, H. et al. BNIP3 supports melanoma cell migration and vasculogenic mimicry by orchestrating the actin cytoskeleton. Cell Death Dis. 5, e1127 (2014).
Sun, J. et al. Hypoxia-induced BNIP3 facilitates the progression and metastasis of uveal melanoma by driving metabolic reprogramming. Autophagy 21, 191–209 (2025).
Wang, S. et al. BNIP3-mediated mitophagy boosts the competitive growth of Lenvatinib-resistant cells via energy metabolism reprogramming in HCC. Cell Death Dis. 15, 484 (2024).
Fornelli, C. et al. BNIP3 Downregulation ameliorates muscle atrophy in cancer cachexia. Cancers (Basel) 16, https://doi.org/10.3390/cancers16244133 (2024).
Mammucari, C. et al. FoxO3 controls autophagy in skeletal muscle in vivo. Cell Metab. 6, 458–471 (2007).
Zhang, Q. et al. PGAM5 interacts with and maintains BNIP3 to license cancer-associated muscle wasting. Autophagy 20, 2205–2220 (2024).
Medeiros, T. C., Thomas, R. L., Ghillebert, R. & Graef, M. Autophagy balances mtDNA synthesis and degradation by DNA polymerase POLG during starvation. J. Cell Biol. 217, 1601–1611 (2018).
Castellani, C. A., Longchamps, R. J., Sun, J., Guallar, E. & Arking, D. E. Thinking outside the nucleus: mitochondrial DNA copy number in health and disease. Mitochondrion 53, 214–223 (2020).
Herbst, A. et al. Skeletal muscle mitochondrial DNA copy number and mitochondrial DNA deletion mutation frequency as predictors of physical performance in older men and women. Geroscience 43, 1253–1264 (2021).
Martinvalet, D., Zhu, P. & Lieberman, J. Granzyme A induces caspase-independent mitochondrial damage, a required first step for apoptosis. Immunity 22, 355–370 (2005).
Perez-Ruiz, E. et al. Prophylactic TNF blockade uncouples efficacy and toxicity in dual CTLA-4 and PD-1 immunotherapy. Nature 569, 428–432 (2019).
Skipworth, R. J., Stewart, G. D., Dejong, C. H., Preston, T. & Fearon, K. C. Pathophysiology of cancer cachexia: much more than host–tumour interaction?. Clin. Nutr. 26, 667–676 (2007).
Wu, Y., Zou, Q., Jiang, P. & Gao, Q. Tumor–host cometabolism collaborates to shape cancer immunity. Cancer Discov. 14, 653–657 (2024).
Penna, F. et al. The skeletal muscle as an active player against cancer cachexia. Front. Physiol. 10, 41 (2019).
Siff, T., Parajuli, P., Razzaque, M. S. & Atfi, A. Cancer-mediated muscle cachexia: etiology and clinical management. Trends Endocrinol. Metab. 32, 382–402 (2021).
Cote, A. L., Munger, C. J. & Ringel, A. E. Emerging insights into the impact of systemic metabolic changes on tumor-immune interactions. Cell Rep. 44, 115234 (2025).
McAllister, S. S. & Weinberg, R. A. Tumor-host interactions: a far-reaching relationship. J. Clin. Oncol. 28, 4022–4028 (2010).
Salaroli, R. et al. Validity of internal expression of the major histocompatibility complex class I in the diagnosis of inflammatory myopathies. J. Clin. Pathol. 65, 14–19 (2012).
Nagaraju, K. et al. Conditional up-regulation of MHC class I in skeletal muscle leads to self-sustaining autoimmune myositis and myositis-specific autoantibodies. Proc. Natl Acad. Sci. USA 97, 9209–9214 (2000).
Luan, Y. et al. Development of ovarian tumour causes significant loss of muscle and adipose tissue: a novel mouse model for cancer cachexia study. J. Cachexia Sarcopenia Muscle 13, 1289–1301 (2022).
Cui, Q. et al. MIF-ACKR3 causes irreversible fat loss by impairing adipogenesis in cancer cachexia. Cell Metab. 37, 954–970 e958 (2025).
Cui, F. et al. Overexpression of Cathepsin L is associated with gefitinib resistance in non-small cell lung cancer. Clin. Transl. Oncol. 18, 722–727 (2016).
Wang, X. et al. CTSL promotes autophagy in laryngeal cancer through the IL6-JAK-STAT3 signalling pathway. J. Cell. Mol. Med. 29, e70364 (2025).
Zhang, L. et al. CTSL, a prognostic marker of breast cancer, that promotes proliferation, migration, and invasion in cells in triple-negative breast cancer. Front. Oncol. 13, 1158087 (2023).
Xiao, L. X. et al. Macrophage-derived cathepsin L promotes epithelial–mesenchymal transition and M2 polarization in gastric cancer. World J. Gastroenterol. 30, 5032–5054 (2024).
Kreuzaler, P. A. et al. Stat3 controls lysosomal-mediated cell death in vivo. Nat. Cell Biol. 13, 303–309 (2011).
Sargeant, T. J. et al. Stat3 controls cell death during mammary gland involution by regulating uptake of milk fat globules and lysosomal membrane permeabilization. Nat. Cell Biol. 16, 1057–1068 (2014).
Jean, D., Rousselet, N. & Frade, R. Expression of cathepsin L in human tumor cells is under the control of distinct regulatory mechanisms. Oncogene 25, 1474–1484 (2006).
Grozdanic, M. et al. Cathepsin L-mediated EGFR cleavage affects intracellular signalling pathways in cancer. Biol. Chem. 405, 283–296 (2024).
Yang, N., Wang, P., Wang, W. J., Song, Y. Z. & Liang, Z. Q. Inhibition of cathepsin L sensitizes human glioma cells to ionizing radiation in vitro through NF-kappaB signaling pathway. Acta Pharm. Sin. 36, 400–410 (2015).
Han, M. L. et al. Cathepsin L upregulation-induced EMT phenotype is associated with the acquisition of cisplatin or paclitaxel resistance in A549 cells. Acta Pharm. Sin. 37, 1606–1622 (2016).
Zheng, X. et al. Cathepsin L inhibition suppresses drug resistance in vitro and in vivo: a putative mechanism. Am. J. Physiol. Cell Physiol. 296, C65–C74 (2009).
Chazaud, B. Inflammation and skeletal muscle regeneration: leave it to the macrophages!. Trends Immunol. 41, 481–492 (2020).
Mourkioti, F. & Rosenthal, N. IGF-1, inflammation and stem cells: interactions during muscle regeneration. Trends Immunol. 26, 535–542 (2005).
Burzyn, D. et al. A special population of regulatory T cells potentiates muscle repair. Cell 155, 1282–1295 (2013).
Casciola-Rosen, L. et al. Enhanced autoantigen expression in regenerating muscle cells in idiopathic inflammatory myopathy. J. Exp. Med. 201, 591–601 (2005).
Huang, P. L. et al. Skeletal muscle interleukin 15 promotes CD8(+) T-cell function and autoimmune myositis. Skelet. Muscle 5, 33 (2015).
Nakamura, K., Arahata, K., Ishiura, S., Osame, M. & Sugita, H. Degradative activity of granzyme A on skeletal muscle proteins in vitro: a possible molecular mechanism for muscle fiber damage in polymyositis. Neuromuscul. Disord. 3, 303–310 (1993).
Goebels, N. et al. Differential expression of perforin in muscle-infiltrating T cells in polymyositis and dermatomyositis. J. Clin. Investig. 97, 2905–2910 (1996).
Kim, J. et al. Innate-like cytotoxic function of bystander-activated CD8(+) T cells is associated with liver injury in acute hepatitis A. Immunity 48, 161–173 e165 (2018).
Younes, S. A. et al. IL-15 promotes activation and expansion of CD8+ T cells in HIV-1 infection. J. Clin. Investig. 126, 2745–2756 (2016).
Onishi, K. et al. Cathepsin L is crucial for a Th1-type immune response during Leishmania major infection. Microbes Infect. 6, 468–474 (2004).
Simoni, Y. et al. Bystander CD8(+) T cells are abundant and phenotypically distinct in human tumour infiltrates. Nature 557, 575–579 (2018).
Johnson, D. E., O’Keefe, R. A. & Grandis, J. R. Targeting the IL-6/JAK/STAT3 signalling axis in cancer. Nat. Rev. Clin. Oncol. 15, 234–248 (2018).
Liu, J. et al. HIF-1 contributes to autophagy activation via BNIP3 to facilitate renal fibrosis in hypoxia in vitro and UUO in vivo. Am. J. Physiol. Cell Physiol. 326, C935–C947 (2024).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 12, 323 (2011).
Hou, W. & Ji, Z. Assessing GPT-4 for cell type annotation in single-cell RNA-seq analysis. Nat. Methods 21, 1462–1465 (2024).
Son, K. 202397101/Ctsl: First release of Ctsl project: initial code base and documentation. GitHub. Zenodo https://doi.org/10.5281/zenodo.17102995 (2025).
Acknowledgements
This work has been supported by National Research Foundation of Korea (NRF) Grants funded by the Korean Government (grant numbers NRF-2020R1C1C1003338 to N.Y.S., NRF-2022M3A9F3016364 to N.Y.S., and RS-2024-00406281 to N.Y.S.; NRF-2021R1C1C1003988 to H.L.) and by the Yonsei University College of Dentistry Fund (6-2024-0004 to N.Y.S.).
Author information
Authors and Affiliations
Contributions
S.Y.P. carried out the experiments, analyzed data, and wrote the paper. K.S. performed the bioinformatic analysis, analyzed data, and wrote the paper. J.K., K.K., S.J., and B.K. carried out the experiments, analyzed data, and wrote the paper. M.L. performed the bioinformatic analysis and wrote the paper. W.K. contributed to the interpretation of the bioinformatic data and reviewed and edited the paper. W.J.J. analyzed PNUH clinical data and wrote the paper. B.K.C. was responsible for the curation and acquisition of PNUH clinical data and wrote the paper. N.J. conceptualized and designed the clinical data analysis and wrote the paper. W.Y.C. and Y.H. contributed to the interpretation of data and reviewed and edited the paper. H.L. conceptualized and designed the bioinformatic analysis, analyzed data, wrote the paper, and acquired the funding. N.Y.S. conceptualized, designed the experiments, analyzed data, wrote the paper, and acquired the funding.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Fabio Penna, who co-reviewed with Giacomo Rubini, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Park, SY., Son, K., Kim, J. et al. Cathepsin L as a dual-target to mitigate muscle wasting while enhancing anti-tumor efficacy of anti-PD-L1. Nat Commun 16, 10706 (2025). https://doi.org/10.1038/s41467-025-64500-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-64500-0