Abstract
Heterodimerization of opioid receptors (ORs), MOR, KOR, and DOR, is implied in their functional regulation and diversification, and thus its understanding is crucial for developing better analgesic treatments. However, our knowledge on OR heterodimerization/heterodimers remains limited. Here, using single-molecule imaging and functional analysis, we find that MOR, the main morphine receptor, repeatedly forms transient (≈250 ms) heterodimers with DOR every 1-10 seconds, but not with KOR, whereas DOR and KOR also form transient heterodimers. We obtain all the heterodimer-monomer equilibrium constants and rate constants with/without agonists. We identify the critical heterodimer binding sites in the extracellular domains, in addition to the less-specific transmembrane domains, and develop soluble peptide blockers for MOR-DOR and DOR-KOR heterodimerization, using amino-acid sequences mimicking the extracellular binding sites. With these peptide blockers, we dissect the monomer/dimer roles in OR internalization and signaling. The soluble MOR-DOR heterodimer blocker reduces the development of long-term morphine tolerance in mice.
Similar content being viewed by others
Introduction
As elucidated in our companion paper1, the transient homodimerization of μ-, κ-, and δ-opioid receptors (MOR, KOR, and DOR, respectively) provides a critical layer of regulation within the opioid signaling paradigm. Theories and methods for quantitative analyses of single-molecule imaging data developed there revealed that all ORs undergo transient homodimerizations for brief periods of 120–180 ms (koff = 6.7–8.5 s−1), and they repeat dissociation and rebinding to another molecule every few seconds or less, directly detectable at expression levels over 0.3 copies/µm2, with homodimer-dissociation equilibrium constants of 6.0–16.6 copies/µm2. The homodimerization is predominantly driven by unique 9–26 amino-acid stretches within the C-terminal cytoplasmic domain of each receptor, with less specific contributions from the transmembrane (TM) domains. Soluble but membrane-permeable peptides mimicking these C-terminal regions reduced homodimerization, enabling us to dissect the functions of homodimers. Compared to their monomeric counterparts, KOR-KOR and DOR-DOR homodimers (KK and DD homodimers)—unlike MOR-MOR homodimers (MM homodimers)—activate downstream signals differently in response to agonist binding, without affecting receptor internalization.
In this paper, we extend these advanced single-molecule imaging studies described in the companion paper to investigate OR heterodimerization. Given its crucial role in morphine response and the significant medical and social implications thereof, MOR has been the focus of extensive OR research2,3,4,5,6,7, including its heterodimerization with DOR and KOR (DM and MK heterodimers, respectively)2,5,8,9. Extensive studies have reported DM heterodimerization10,11,12,13 and downstream signals suggesting DM interactions14,15,16,17, although DM co-expression might be limited to small populations of neurons, such as excitatory interneurons, projection neurons in the spinal cord dorsal horn, and nociceptive neurons in dorsal root ganglia1,16,18,19,20,21,22,23. In contrast, evidence for MK heterodimerization remains sparse, which suggests a dependence on the physiological context9,24,25.
The DK coupling exhibits unique signaling and functional regulation, making the DK heterodimer as another promising therapeutic target for pain treatment3,26,27. Indeed, DK heterodimerization and observations suggesting DK interactions have also been reported3,25,26,28,29, further highlighting its significance.
Despite these studies, the fundamental properties of heterodimers, including heterodimer-monomer dissociation equilibrium constants and rate constants (KD, koff, which is inverse lifetime, and kon), heterodimer interaction sites, and cellular- and tissue-level functions, remain enigmatic. This research addresses these fundamental issues by leveraging the advanced single-molecule image analysis described in the companion paper1. Such understanding would form the basis for developing opioid analgesics with heightened efficacy and minimized tolerance.
The heterodimer-monomer equilibrium constant, KD (hetero-KD), can only be obtained after the homodimer-monomer equilibrium constants KDs (homo-KDs) for the two constituent ORs are evaluated, as detailed in the companion paper1. We also compare the properties and dimerization sites between homo- and hetero-dimers of ORs.
This study lays the groundwork for future explorations of dimerization phenomena across the broader GPCR family30,31. Heterodimerization among class-A GPCRs has been increasingly observed, offering insights into novel signaling pathways and pharmacological profiles8,32,33,34,35,36. Examples include heterodimers between MOR and V1b vasopressin receptor37, MOR and somatostatin receptor 2, CCR5 and CXCR436, dopamine D2 receptor and neurotensin NTS1 receptor38, angiotensin II AT1 receptor and norepinephrine α2C-adrenergic receptor33, cannabinoid receptors CB1 and CB239, dopamine receptors D1 and D240, and others. Class-C GPCRs, such as the γ-aminobutyric acid type B (GABAB) receptor, exhibit prominent dimerization properties41,42,43, marking an emergent frontier in GPCR research with profound implications for drug discovery.
Here, we unequivocally demonstrate that, even under expression conditions as low as 1 copy/µm2 for each OR species, DOR-MOR and DOR-KOR form metastable heterodimers, whereas MOR-KOR heterodimers are not detectable. These findings have been substantiated through examinations conducted with a level of quantification unparalleled in the existing biomedical literature, with the exception of the OR homodimer research described in our companion paper. Furthermore, we discovered that the extracellular N-terminal domain interactions and extracellular loop 3 (EL3) interactions are specifically responsible for DM and DK heterodimerizations, respectively. In contrast, the TM domain interactions contribute less specifically, challenging the common belief that GPCR dimerizations are primarily mediated by TM domain interactions8,13,27,44,45,46. In addition, allosteric conformational changes involving both extracellular domains and TM domains might be involved. Building on these findings, we demonstrated that soluble peptides mimicking the extracellular amino-acid sequences implicated in DM and DK heterodimerizations effectively inhibit heterodimer formation. In murine models, the administration of a soluble peptide blocker targeting the DM heterodimers into the cerebral ventricles diminishes the development of long-term morphine tolerance. These results strongly suggest that DM heterodimerization based on the N-terminal domain sequence occurs in neuronal tissues. These findings underscore potential avenues for improving opioid drug administration strategies aimed at mitigating tolerance and dependence.
Results
Metastable DM and DK, but not MK, heterodimers are detected in the PM
To investigate heterodimer formation among ORs, SNAPf- and Halo-tagged ORs were co-expressed in CHO-K1 cells, which lack endogenous ORs (Supplementary Fig. 1a)47. These tagged ORs are functional (Supplementary Fig. 1a, b) and labeled with SNAP-Surface 549 and Halo-SaraFluor650T dyes (Supplementary Fig. 1c–d)1. The labeled OR molecules were simultaneously observed in two colors at the single molecule level at video rate (30 Hz) at 37 °C, using a home-built total internal reflection fluorescence (TIRF) microscope. The cells expressing each OR species at fluorescence spot densities between 0.5–1 spot/µm2 (total number densities between 1–2 spots/µm2) after fluorescence labeling were selected for microscope observations. Virtually all of the OR fluorescent spots exhibited diffusion in the PM.
When DOR and MOR were expressed in the same cell, they exhibited frequent brief colocalization and co-diffusion, suggesting metastable DM heterodimer formation (Supplementary Movies 1 and 2; Fig. 1a). For quantifying colocalization, we employed a pair cross-correlation function (PCCF; Supplementary Fig. 1e; also see companion paper)1, which provides both a simple measure for heterodimer formation, the colocalization index, and a fundamental constant, the heterodimer-monomer dissociation equilibrium constant KD1. As a control for incidental fluorescent spot colocalizations, the video frames in the green channel were rotated 180° before overlaying1.
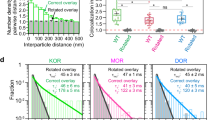
a Typical (among 20 replicates) image sequence of simultaneous two-color single fluorescent molecule observations and trajectories of the molecules shown in the images. These molecules undergo transient hetero-colocalization and co-diffusion (SNAP-Surface 549-labeled SNAPf-DOR, green; SaraFluor650T-labeled MOR-Halo, magenta). b DM and DK heterodimers form, but MK heterodimers do not, as detected by colocalization indexes for correct and rotated overlay1. In the box plots, horizontal bars, crosses, boxes, and whiskers indicate median values, mean values, interquartile ranges (25–75%), and 10–90% ranges, respectively. * and ns represent significant (p < 0.05) and non-significant (p ≥ 0.05) differences, respectively (two-sided Welch’s T test). All of the statistical parameters and analysis results including sample size n and p values are provided in Supplementary Data 2. d Histograms showing the distributions of hetero-colocalization durations for correct and rotated overlays. The control histograms for rotated overlays (gray) were fitted by single exponential functions (black), providing the lifetimes of the incidental overlap events between the magenta and green spots (τinci). The histograms for correct overlays (colors) were fitted by the sum of two exponential functions: The faster decay time (τ1) was close to the lifetime of incidental overlaps (τinci), and the slower decay time provided the heterodimer lifetime (τ2). The heterodimer lifetime after correction for the trackable duration lifetime is shown in each box. The koff values (1/τ2) calculated from τ2 are shown in c. c Summary of KD and koff values for OR heterodimers. e Comparison of the amino acid sequences among the three classical ORs (rat) in the domains where the amino acid homologies are lower. Asterisks mark the identical amino acids. Source data are provided as a Source Data file.
The colocalization index clearly indicated that DOR and MOR form DM heterodimers and DOR and KOR form DK heterodimers in the PM. However, at the expression levels employed in this study, MK heterodimers were not detected (Fig. 1b and Supplementary Fig. 1e), consistent with the previous result25. The heterodimer-monomer dissociation equilibrium constants KDs were obtained from the PCCFs (Fig. 1c).
As we explained in the companion paper1, the dissociation rate constant, koff (=1/τ2; Fig. 1c), was determined from the heterodimer lifetime (τ2) which was obtained by fitting the distribution of colocalization durations with the sum of two exponential functions and corrected with the trackable duration lifetimes (Supplementary Fig. 1f). The τ2 values are 260 ± 12 ms for DM heterodimers and 241 ± 10 ms for DK heterodimers, after corrections for trackable duration lifetimes (Fig. 1d and Supplementary Fig. 1f). The histogram for the MK colocalization durations did not exhibit any τ2 component (the decay component representing the heterodimer loss), consistent with the lack of detectable heterodimers for this pair (only incidental colocalization with a lifetime of τ; Fig. 1d).
To identify the amino-acid sequences responsible for specific heterodimerization, we initially examined the OR domains with lower amino-acid identities/homologies among the three ORs. The candidate sites we initially selected are summarized in Fig. 1e. The amino-acid sequence homologies in the cytoplasmic C-terminal domains are lower, but they are not included in this figure as they are involved in KK, MM, and DD homodimerizations1 and we unequivocally showed that they are not involved in DK and DM heterodimerizations (Figs. 2a, 3a).
a The N- and C-terminal domains of DOR and KOR are not involved in DK heterodimerization. DK colocalization indexes of N/C-terminal deletion mutants of DOR and full-length KOR (top) and N/C-terminal deletion mutants of KOR and full-length DOR (bottom). b Summary of the names and exact amino-acid ranges of the OR domains and TM1-based peptides employed in this work. c DOR’s EL3, TM1, and TM4, but not EL2 and N-term, are involved in DK heterodimerization. d Both EL3 domains of DOR and KOR are involved in DK heterodimerization. Soluble peptides (1 µM) with the aa sequences of the EL3 domains (EL3-domain peptides) of DOR and KOR, but not that of MOR, block DK heterodimerization. e DOR’s TM1 and TM4 bind to KOR. Colocalization indexes show that the affinities to WT-KOR are greater in the order of WT-DOR > DOR’s TM1 > DOR’s TM4. Because the TM4 orientation in the PM is from the PM cytoplasmic surface toward outer surface, to maintain this orientation, the N-terminus of TM4 was linked to the C-terminus of TMLDLR, which is oriented in the opposite way and does not interact with ORs. In the box plots (a, c–e), horizontal bars, boxes, and whiskers indicate mean values, interquartile ranges (25–75%), and 10-90% ranges, respectively. * and ns represent significant (p < 0.05) and non-significant (p ≥ 0.05) differences, respectively (Tukey’s multiple comparison test). The data set used for multiple comparison is indicated by the group of lines in each figure, and different sets are indicated by different colors. All of the statistical parameters and analysis results including sample size n and p values are provided in Supplementary Data 2. Source data are provided as a Source Data file.
a Involvement of the extracellular N-terminal domains of DOR and MOR in DM heterodimerization, particularly that of DOR’s aa22-42 and MOR’s aa32-61. The figures show that mutants with various deletions in the N-terminal domains of DOR and MOR exhibit lower colocalization indexes with full-length (WT) MOR and DOR, respectively, compared to the colocalization index between full-length DOR and MOR. At the bottoms of both panels are the results for the C-terminal deletion mutants, indicating the lack of cytoplasmic C-terminal domain involvement in DM heterodimerization. b Histograms showing the colocalization duration distributions for the pair of WT-MOR and DOR(∆22-42) (left, blue) and the pair of WT-DOR and MOR(∆32-61) (right, orange). Histograms for the control and rotated overlays of WT DOR-MOR images are the same as those shown in Fig. 1d. c The extracellular N-terminal domains of DOR and MOR are involved in DM heterodimerization (left and right panels, respectively), as are DOR’s TM1 and TM4 domains, but not the TM6 domain (left panel). (Left panel) In the box plots (a, c), horizontal bars, boxes, and whiskers indicate mean values, interquartile ranges (25–75%), and 10-90% ranges, respectively. * and ns represent significant (p < 0.05) and non-significant (p ≥ 0.05) differences, respectively (Tukey’s multiple comparison test). The data set used for multiple comparison is indicated by the group of lines in each figure, and different sets are indicated by different colors. All of the statistical parameters and analysis results including sample size n and p values are provided in Supplementary Data 2. Source data are provided as a Source Data file.
DOR’s EL3, TM1, and TM4, but not EL2, N- or C-terminal domain, contribute to DK heterodimerization
To identify the domains essential for DK heterodimer formation, we first examined the heterodimerization of wild-type (WT) KOR and N- or C-terminal deletion mutants (∆N or ∆C mutants) of DOR, as well as WT DOR and N- or C-terminal deletion mutants (∆N or ∆C mutants) of KOR (Fig. 2a). The colocalization index analysis indicated that neither the N- nor C-terminal domains of DOR and KOR were involved in the DK heterodimerization.
Given the scarce occurrence of MK heterodimers (colocalization index = 1.15 ± 0.03), we tested whether MOR mutants with some domains substituted with DOR counterparts could heterodimerize with WT-KOR (Fig. 1e; see Fig. 2b for domain names and amino-acid numbers; results shown in Fig. 2c left). Substitution with DOR’s TM1 or TM4 domain in MOR (MOR[TM1DOR] or MOR[TM4DOR]) facilitated heterodimer formation with WT-KOR, qualitatively consistent with a previous result27. Similarly, introducing DOR’s EL3 into MOR (MOR[EL3DOR]) also promoted heterodimerization with WT-KOR. In contrast, the substitution of MOR’s EL2 or N-terminal domain (MOR[EL2DOR] or MOR[NtermDOR]) failed to induce heterodimerization with WT-KOR. These results indicate the involvement of DOR’s EL3 domain, alongside its TM1 and TM4 domains, in DK heterodimer formation.
The lifetime distributions for the MOR(EL3DOR)-KOR and MOR(TM1DOR)-KOR heterodimers revealed the appearance of the τ2 component, which was absent in the distribution for the WT-MK pairs (Supplementary Fig. 2a). Although the τ2 values are smaller than that for WT-DK heterodimers (164 ± 25 ms or 131 ± 15 ms vs. 241 ± 9.7 ms), the presence of this longer-lived component implicates DOR’s EL3 and TM1 domains in DK heterodimerization.
Experiments replacing various DOR domains with the MOR counterparts and examining heterodimerization with WT-KOR (based on the lack of MK heterodimerization) showed reduced heterodimerization for TM1, TM4, and EL3 domain replacements, but not for EL2 or N-terminal domain substitutions (Fig. 2c right). This further indicates that DOR’s EL3, TM1, and TM4 domains are integral to DK heterodimer formation.
Soluble EL3-domain peptides of DOR and KOR both block DK dimerization
To address the possibility that a variety of domain deletions (Fig. 2a) and substitutions (Fig. 2c) in WT-DOR could alter its conformation and thereby indirectly affect heterodimerization, we synthesized soluble peptides derived from EL3 sequences of DOR and KOR (Fig. 1e). Addition of either peptide (1 µM, final concentration) to cells co-expressing DOR and KOR significantly reduced DK heterodimerization, unlike the peptide with the MOR EL3 sequence (Fig. 2d). This highlights the critical roles of the DOR and KOR EL3 domains in DK heterodimerization. Given that DOR’s EL2 domain and the N-terminal domains of both DOR and KOR are not involved in DK heterodimerization (Fig. 2a, c), these results indicate that the interaction between the DOR and KOR EL3 domains is crucial for DK heterodimerization, although that with KOR’s EL2 domain has not been ruled out. In addition, DOR’s EL3 domain or MOR’s EL3 domain failed to block DM heterodimers (Supplementary Fig. 2b), indicating the specificities of DOR’s and KOR’s EL3 domains for DK heterodimerization.
Furthermore, we found that DOR’s TM1 and TM4 peptides expressed in the PM became colocalized with KOR, supporting their involvement in DK heterodimerization (Fig. 2b, e). These results are consistent with those shown in Fig. 2c and a previous observation28, and indicate that the interactions between the DOR and KOR EL3s, and those of DOR’s TM1 and TM4 with KOR’s TM domains are involved in DK heterodimerization.
DM dimerization is predominantly mediated by N-terminal domain interactions
To identify the amino-acid sequences responsible for DM heterodimerization, first, we systematically deleted partial sequences from the N- and C-terminal domains of DOR and MOR (Fig. 3a). We found that DOR’s N-terminal amino acids 22-42 (with the critical amino acids 22-27) and MOR’s N-terminal amino acids 32-61 play key roles in DM heterodimerization, likely by binding to each other (Fig. 3a). The key DOR sequence was further confirmed by replacing DOR’s amino acids 21-27 with an arbitrarily selected amino-acid sequence, rather than deleting the original sequence (Supplementary Fig. 3). Therefore, these sequences are likely to be the binding interfaces for DM heterodimerization.
The histograms of the colocalization durations for DOR∆22-42 with WT-MOR and MOR∆32-61 with WT-DOR both revealed only small fractions of the τ2 components, with significantly reduced heterodimer lifetimes of 88 ± 12 ms and 91 ± 8 ms, respectively (Fig. 3b).
We further explored the specific regions mediating DM heterodimerization by taking advantage of that MK heterodimerization hardly occurs at the expression levels we employed. Thus, we examined WT-MOR interactions with DOR mutants with various DOR domains substituted with their KOR counterparts (Fig. 3c left). The colocalization index upon replacement of the N-terminal domain substantially decreased, consistent with the previous demonstration that the N-terminal domains of DOR and MOR are instrumental in DM heterodimer formation (Fig. 3a, b).
Conversely, the replacement of MOR’s N-terminal domain with that from KOR (NtermKOR) greatly reduced the colocalization with DOR (Fig. 3c right). This result further supports (1) that MOR’s N-terminal domain is important for the DM interaction and (2) that KOR’s N-terminal domain is not responsible for the DK interaction. Importantly, this result (Fig. 3c right), combined with the DOR(NterrmKOR) data (Fig. 3c left), further demonstrates that the predominant interactions for DM heterodimerization occur between the N-terminal domains of both DOR and MOR, consistent with the N-terminal deletion results (Fig. 3a, b). For DM heterodimerization, the N-terminal domain of DOR or MOR might interact with the EL2 and EL3 domains of the partner molecules, but this would be far less important compared with the interaction between the two N-terminal domains, because the lack of either DOR residues 22-42 or MOR residues 32-61 is sufficient to significantly lower the colocalization index (Fig. 3a) and eliminate the second colocalization lifetime component (Fig. 3b).
Interactions of some ORs’ TM domains enhance both homo- and hetero-dimerizations of ORs with less specificities
DM heterodimerization might be aided by TM domain interactions, as suggested previously13,46,48. N-terminal domain deletions and specific modifications decreased the colocalization index to 1.15, but not to ≈1.0 (Fig. 3a, c), and reduced the heterodimer lifetimes to ≈90 ms compared to the 260-ms lifetime of the WT DM heterodimers, but failed to totally eliminate the longer τ2 component (Fig. 3b), suggesting the presence of other weaker binding sites.
Substitution of DOR’s TM1 or TM4 with its KOR counterpart, but not TM6 (Fig. 1e), yielded moderate reductions in colocalization with WT-MOR, but not as much as the N-terminal domain substitution (with NtermKOR) (Fig. 3c left). This result suggests that DOR’s TM1 and TM4 play ancillary roles in DM heterodimer stability. These results, but not those of TM6, are consistent with previous computational modeling predictions48,49. The DOR mutant bearing the three replacements of KOR’s N-terminal domain, TM1, and TM4 exhibited a lower colocalization index with WT-MOR, almost to the level of MK interaction (1.15).
Importantly, DK heterodimerization also involves the same TM domains of DOR; i.e., DOR’s TM1 and TM4 (Fig. 2c, left and right, and e). Furthermore, MOR’s TM1 was proposed to participate in MM homodimerization1. These results suggest that some OR TM domains support various OR homo- and hetero-dimerizations; namely, interactions between some of the ORs’ TM domains occur but with lower specificities, thus generally enhancing OR dimerizations. This finding is aligned with previous controversial reports: The conclusion we obtained from the reported results in the literature is that the same TM domains contribute to both homo- and hetero-dimerizations of ORs and other GPCRs to various extents44,46,48,50,51.
Soluble N-terminal-domain peptides block DM heterodimerization
We then utilized the soluble peptides corresponding to the N-terminal sequences of both DOR and MOR implicated in DM heterodimerization, as we did for confirming the DK heterodimer binding sites. These soluble peptides from DOR and MOR are called Dpep(m-n)DM and Mpep(m-n)MD, respectively, where m and n indicate the amino acid numbers in the original OR sequences (see “Location in DOR” and “Location in MOR” in Fig. 4a, respectively). Among the various DpepDMs and MpepMDs, Dpep(20-42)DM and Mpep(32-61)MD most potently lowered the colocalization indexes to 1.2–1.3 (Fig. 4a), mirroring the effects observed with the deletion and replacement mutants (Fig. 3a, c).
a Effect of the addition of 1 µM soluble peptides with the same amino-acid sequences as various parts of the N-terminal domains of DOR and MOR (DpepDM and MpepDM, respectively) on the DM colocalization index. In the box plots, horizontal bars, boxes, and whiskers indicate mean values, interquartile ranges (25–75%), and 10-90% ranges, respectively. b With an increase of the Dpep(20-42)DM concentration, the DM colocalization index decreased. The gray keys at 0 µM Dpep(20-42)DM represent the data points obtained by using rotated overlays. The colocalization index reduces to about the middle value between no addition and the addition of 1 or 10 µM of the Dpep(20-42)DM (saturating conditions) at a Dpep(20-42)DM concentration of 0.1 µM (these are normal 3D concentrations because the peptide is very soluble). Therefore, we conclude that KD for Dpep(20-42)DM binding to MOR is on the order of 0.1 µM. In the box plots, horizontal bars, crosses, boxes, and whiskers indicate median values, mean values, interquartile ranges (25–75%), and 10–90% ranges, respectively. c Dpep(20-42)DM reduces the τ2 of DM colocalization (blue). Histograms for the control (no addition) and rotated overlays of WT DOR-MOR images are the same as those shown in Fig. 1d. In the box plots (a, b), * and ns represent significant (p < 0.05) and non-significant (p ≥ 0.05) differences, respectively (Tukey’s multiple comparison test). The data set used for multiple comparison is indicated by the group of lines in each figure. All of the statistical parameters and analysis results including sample size n and p values are provided in Supplementary Data 2. Source data are provided as a Source Data file.
The colocalization index for DM heterodimers decreased with an increase in the Dpep(20-42)DM concentration to 1 µM (where the peptide effect is almost saturated), indicating that the affinity of the DM heterodimers (three-dimensional KD because the peptide is very soluble) would be on the order of 0.1 µM in three-dimensional space (Fig. 4b and its caption). This 3D concentration could not readily be converted to a two-dimensional molecular density, as discussed in the companion paper1. Briefly, in two-dimensional space, the dimerization efficiency is far greater, by a factor of 106, than that in three-dimensional space52. Therefore, the efficacy of the peptides in blocking OR dimerization, even at the 0.1 µM-order concentrations required for blocking DM heterodimerization, is deemed plausible given the nanomolar affinity range of OR ligands (1-10 nM).
Dpep(20-42)DM’s introduction substantially shortened the DM heterodimer lifetime from 260 ms to 124 ms (Fig. 4c), supporting that the amino acid sequence 20-42 in DOR is critical for DM heterodimer formation. Dpep(20-42)DM exhibited excellent stability in the cell culture medium (virtually no degradation) and in 50% mouse plasma ( ≤ 10% degradation) at 37 °C for 80 min (Supplementary Fig. 4).
The membrane-integral peptide TM1MOR suppresses both DM heterodimerization and MM homodimerization
Next, we examined whether MOR’s TM1 (TM1MOR) is involved in DM heterodimerization, as previously described13,46,48. In addition to TM1MOR (amino acids 71-99), a second construct containing amino acids 53-70 before the TM1 domain (Nterm-TM1MOR) was used (see Fig. 2b for construct names and exact amino-acid ranges), following a previous report13. When these peptides were co-expressed with DOR, both TM1MOR and Nterm-TM1MOR colocalized with DOR, like the full-length MOR (all molecules expressed at densities of 0.5–1.0 fluorescent spots/µm2; Fig. 5a, b).
a Experimental design for examining the interactions of TM1MOR, Nterm-TM1MOR, and TM4DOR (Fig. 2b) with WT-ORs (top). For the correct orientation of TM4DOR in the PM, it was linked to the transmembrane domain of LDL receptor (TMLDLR). Representative (among 20 replicates) single-molecule images of DOR-mGFP (green), MOR-Halo (magenta), and SNAPf-TM1MOR (cyan) co-expressed at similar levels of 0.5-1.0 fluorescent spots/µm2 (bottom). b TM1MOR and Nterm-TM1MOR form dimers with WT-DOR as efficiently as WT-MOR does. TM1DOR and TM4DOR form dimers with WT-MOR to lesser extents. Expression levels of all molecules were adjusted to 0.5-1.0 fluorescent spots/µm2 individually. c Reduction of the DM colocalization index with an increase in the number densities of the TM peptides (TM1MOR, Nterm-TM1MOR, and TM4DOR) relative to that of MOR, expressed in the PM. DOR and MOR were both expressed at levels of 0.5-1.0 fluorescent spots/µm2. The effects of TM1MOR and Nterm-TM1MOR are stronger than that of TM4DOR. The rectangular box without the peptides (x = 0) indicates the result of the box plot without datapoints (to avoid excessive complexity in the plot), where the cross indicates the mean value and the box indicates the interquartile range (25–75%). d TM1MOR and Nterm-TM1MOR reduce MM and DD homodimers (in addition to DM heterodimers), but 1 µM Dpep(20-42)DM does not. TM1MOR and Nterm-TM1MOR expression levels were ≈10x of MOR and DOR, following the results shown in panel c. TMLDLR is a negative control, representing non-interacting TM domains. In the box plots (b, d), horizontal bars, boxes, and whiskers indicate mean values, interquartile ranges (25–75%), and 10-90% ranges, respectively. * and ns represent significant (p < 0.05) and non-significant (p ≥ 0.05) differences, respectively (Tukey’s multiple comparison test). All of the statistical parameters and analysis results including sample size n and p values are provided in Supplementary Data 2. Source data are provided as a Source Data file.
To identify which DOR’s TMs interact with MOR, we examined the colocalization of WT-MOR with TM1DOR and TM4DOR, which were previously suggested to interact with TM1MOR46,48. TM1DOR and TM4DOR moderately colocalized with MOR when expressed at similar levels (both at densities of 0.5–1.0 fluorescent spots/µm2; Fig. 5b).
In the peptide-disrupting heterodimer experiment, we found that all TM1MOR, Nterm-TM1MOR, and TM4DOR suppressed DM heterodimerization in an expression level-dependent manner, and TM1MOR exhibited a greater efficacy than TM4DOR (Fig. 5c). These results indicated TM1MOR and TM4DOR’s significant contribution to the DM interaction but also suggesting the presence of other significant binding sites, including TM1DOR and other TM domains as well as the N-terminal domain.
Notably, the addition of either Dpep(20-42)DM (also Mpep(32-61)MD) or TM1MOR alone reduced the colocalization index to near-baseline levels (1.2–1.3) (Figs. 4a, b and 5c). These results suggest that DM heterodimerization might not be induced by simple bindings at two sites, but instead mediated by cooperative allosteric conformational changes of the extracellular N-terminal domains and TM domains in both OR molecules. In this case, our description of the dynamic dimer-monomer equilibrium should be considered as a simplified representation of the actual molecular events.
Despite the TM contributions to heterodimerization, the DM homodimerization mediated by TM1MOR interaction with TM1DOR and TM4DOR appears to be weaker and less specific compared to the N-terminal interactions between DOR and MOR: both TM1DOR and TM4DOR are involved in both DM (Fig. 3c left) and DK (Fig. 2c, e) heterodimerizations, and TM1MOR is involved in both DM heterodimerization (Fig. 3c right; Fig. 5d as it blocks DD homodimerization) and MM homodimerization1. In striking contrast, Dpep(20-42)DM had no influence on either MM or DD homodimerization (Fig. 5d). A control TM-peptide, TMLDLR, with the sequence of the LDL receptor’s TM domain, exhibited no effect on homodimer blocking (Fig. 5d).
In conclusion, the N-terminal domains of DOR and MOR are specifically involved in DM heterodimerization, whereas the TM domains contribute but interact less specifically: TM1MOR in MOR enhances both DM heterodimerization and MM homodimerization, and TM1DOR and TM4DOR in DOR participate in both DM and DK heterodimerization. Meanwhile, DM heterodimerization might involve allosteric interactions between the N-terminal and TM domains.
Under the conditions where only DOR and MOR are expressed, the addition of Dpep(20-42)DM or Mpep(32-61)MD will decrease DM heterodimers. This will increase the numbers of monomeric DOR and MOR molecules available for DD and MM homodimerization, leading to increased numbers of DD and MM homodimers.
M-agonists’ effect on DM heterodimerization
We examined the effects of MOR agonists (M-agonists), morphine and [D-Ala2, N-MePhe4, Gly-ol]-enkephalin (DAMGO), on DM heterodimerization (Fig. 6a–d). At an M-agonist concentration of 0.5 µM in the cell culture medium, MOR would be rapidly bound by morphine and DAMGO (denoted as M*mor and M*DAMGO, respectively)53, and therefore we examined the M-agonist effect on DM heterodimerization within 5 min after the agonist addition, prior to any detectable internalization. As detailed in the companion paper, DAMGO enhanced MM homodimerization and slightly elongated the MM dimer lifetime1.
a Experimental PCCFs for DM heterodimerization in the presence and absence of 0.5 µM M-agonists and 1 µM Dpep(20-42)DM (mean ± SEM; n = 20 replicates) and their fitting curves using KD for DM heterodimerization and σ (precisions of single-molecule localizations for the two probes and spatial precision for overlaying two-color images) as fitting parameters. MD-cells expressing both DOR and MOR at 0.5-1.0 fluorescent spots/µm2 were employed. b Colocalization indexes obtained from the PCCFs shown in a. Together with the results shown in a, the results indicate that morphine hardly affects DM heterodimerization whereas DMAGO enhances it. The Dpep(20-42)DM addition significantly reduces DM heterodimers under all conditions examined here. In the box plots, horizontal bars, boxes, and whiskers indicate mean values, interquartile ranges (25–75%), and 10-90% ranges, respectively. * and ns represent significant (p < 0.05) and non-significant (p ≥ 0.05) differences, respectively (Tukey’s multiple comparison test). All of the statistical parameters and analysis results including sample size n and p values are provided in Supplementary Data 2. c Morphine shortens and DAMGO prolongs DM heterodimer lifetimes, whereas the further addition of Dpep(20-42)DM reduces the lifetimes, making them shorter than those for DM heterodimers without any treatment. d Schematic summary of KD and koff values for DM heterodimers in the presence and absence of DAMGO and DpepDM(20-42)DM. Source data are provided as a Source Data file.
M*mor exhibited almost the same colocalization index with DOR as non-ligated MOR, although the DM*mor heterodimer lifetime was shorter (Fig. 6a–c). Dpep(20–42)DM suppressed DM heterodimerization to a similar level for both M*mor and non-ligated MOR (Fig. 6a, b). In contrast, M*DAMGO exhibited a greater colocalization index with DOR (DM*DAMGO heterodimers) with a prolonged heterodimer lifetime, compared to MOR in the basal state (Fig. 6a–c). The presence of Dpep(20-42)DM in the medium markedly reduced both the colocalization index and lifetime of the DM*DAMGO heterodimers. We were unable to evaluate the effects of the DOR agonist SNC-80 (0.5 µM) on DM heterodimerization, due to rapid internalization of both MOR and DOR following ligand application, as detailed in subsequent sections.
Summary of K D, k off, and k on values and expected percentages of OR protomers existing as homo- and hetero-dimers at various expression levels
The equilibrium constants KDs were deduced by fitting the experimental PCCF histograms for DM and DK heterodimers with theoretical models (Fig. 6a), using hetero-KD and σ as fitting parameters1. The derived KD, koff, and kon values for KK, MM, and DD homodimers as well as DM and DK heterodimers, with or without M-agonists and Dpep(20-42)DM, are summarized in Fig. 6d and Supplementary Tables 1 and 2 (for the influence of expression levels on the evaluated KD values with actual curve fitting results, see Supplementary Fig. 5; all statistical data are summarized in Supplementary Data 2).
Based on the KD values obtained here, the percentages of OR protomers existing as monomers, DD and MM homodimers, and DM and DK heterodimers in the presence and absence of morphine and DAMGO, at various number densities of ORs expressed in the PM, are calculated and shown in Fig. 7 and Supplementary Tables 2-3 for the cases where DOR and MOR expression levels are equalized. In particular, Supplementary Table 3 presents the fractions of MOR or DOR protomers existing as monomers, homodimers, and heterodimers. For the cases where expression levels are freely varied, see Fig. 7. Without knowing the KD values for both homo- and hetero-dimerizations, such evaluations would have been impossible.
In a-c, from left to right, no addition, + morphine, and + DAMGO. a Overall percentages of MOR and DOR protomers existing as DM heterodimers. b Percentages of MOR protomers existing as DM heterodimers. c Percentages of DOR protomers existing as DM heterodimers. d Overall percentages of DOR and KOR protomers existing as DK heterodimers (left), percentages of DOR protomers existing as DK heterodimers (middle), and percentages of KOR protomers existing as DK heterodimers (right).
Evaluation of the numbers of homo- and hetero-dimers in the presence of all three ORs and the numbers of MMD and DDM hetero-trimers at various expression levels of MOR and DOR
In some neurons, three ORs might be simultaneously expressed and/or their expression levels might be quite high. In these situations, five types of dimers (KK-, MM-, DD-, DM-, and DK-dimers) and three monomers (K, M, and D) would coexist and oligomers greater than dimers would appear. The KK, MM, and DD homodimers are likely to co-exist because their respective major dimerization sites are distinct, although they are all located in the cytoplasmic C-terminal domains. The major binding sites for DM heterodimers are in their N-terminal domains and those for DK heterodimers are in their EL3 domains, which are extracellular, and thus steric hindrance to exclude each other might occur to certain extents. Meanwhile, interference between the heterodimerization sites in the extracellular domain and the homodimerization sites in the cytoplasmic domain would be limited. All ORs might interact with each other via TM domain interactions, but the interactions are likely to be weaker and less specific, although they might act cooperatively to induce greater oligomers. G-proteins might bind to ORs both before and after stimulation, but this effect is included in the measurements of KD, koff, and kon in the case of the CHO-K1 cells used in this study.
Here, although considerations are required for complications by the secondary binding sites as explained in the previous paragraph, by assuming that the five binding sites are distinct and independent (KK, MM, DD, DM, and DK), we obtained a snapshot view of how the three ORs expressed in the same PM distribute among monomers and various dimers in the PM by calculating the fractions of various particle species and creating their distributions by Monte Carlo simulations (Supplementary Note 1), using the kinetic constants evaluated here (Fig. 8a). The results before and after the DAMGO addition are shown in Fig. 8b with the fractional percentages of monomers and five types of dimers, giving a useful schematic view of their (random) distributions in the two-dimensional PM space. As described, DAMGO binding enhances MM dimers when MOR alone is expressed, and it enhances DM dimers when only DOR and MOR are co-expressed, but in the presence of three OR species, the fractional increase of MM is minimal and that of DM is only about a factor of 1.5–2 under these low expression conditions (1–10 copies/µm2). Due to the five-way interactions (excluding the MK interaction), the co-existence of three ORs acts like a buffer system, moderating the effects of DAMGO and other agonists on various coexisting species in a quite stable manner.
Snapshot of OR distributions as monomers, homodimers, and heterodimers at selected low physiological concentrations (number densities) in the PM expressing KOR, MOR, and DOR at a 1:1:1 ratio by simulation (a, b), and estimation of the number of heterotrimers (2 DOR and 1 MOR or 1 DOR and 2 MOR molecules) in cells co-expressing various concentrations of DOR and MOR at a 1:1 ratio calculated by using koff and kon evaluated in these studies (c, d). a Schematic figure showing the kinetic parameters obtained in this work at the steady state (left) and after DAMGO stimulation (right). b Simulated distributions of ORs expressed at 1 (left) and 10 (right) copies/µm2. Note that the area sizes are different (100 and 10 µm2 for 1 and 10 copies/µm2, respectively) to match the total copy number to 100 in each box. For the detailed mathematical model, see Supplementary Note 1. c Schematic diagram of trimer formation for MOR and DOR under conditions with the absence of interference between the homodimer and heterodimer formation sites, using the kinetic parameters of homo- and hetero-interactions of MOR and DOR obtained in these studies. For the details of conditions and calculations, see Supplementary Note 1. d Estimated fractions of monomers (M + D), dimers (MM + DD + DM), and trimers (DMM + DDM) in terms of the total numbers of protomers.
Furthermore, we calculated how the number of oligomers greater than dimers increases with an increase in expression levels. We examined the case where only MOR and DOR are expressed at the same levels and homo- and hetero-interactions occur independently without the contributions from other secondary interaction sites, like the case for the results shown in Fig. 8a, b. The fractions of MOR and DOR protomers existing as MMD and DDM trimers, as well as dimers (DD, MM, and DM) and monomers, were calculated based on the kinetic constants (kon and koff values) evaluated here (Fig. 8c, d; Supplementary Table 2). The results show that at the expression levels employed in this study (2–3 copies/µm2 by combining data from the homo- and hetero-dimer papers), trimers will scarcely exist, which is consistent with our experimental observations. At a quite high local number densities of 10 copies/µm2, approximately one-third of OR (DOR + MOR) molecules will exist as dimers and approximately 7% of OR (DOR + MOR) molecules will exist as trimers.
Virtually all OR molecules alternate between existing as homo- and hetero-dimers and monomers during short periods on order of 1–10 s
Cautious interpretations are required for the small percentages of OR protomers, 3.8–20.8%, existing as homo- and hetero-dimers at any moment at low expression conditions (1 copy/µm2 for each OR; see columns of expression levels of 1 copy/µm2 and 1 + 1 copies/µm2 in Supplementary Tables 1 and 2, respectively; including the presence of morphine and DAMGO, but not peptide dimer blockers). Based on these values, one might consider the functional influence of dimers to be small, but this would be incorrect. Note that the dimer lifetimes are in the range of 118-369 ms, indicating that these dimers fall apart quickly, but then the monomers will again form dimers with different (and sometimes the same) partner molecules1. This process is continually repeated. Therefore, although at any time the percentages of protomers existing as dimers are limited, virtually all OR molecules rapidly interconvert among homodimer-, monomer-, and heterodimer-states and monomers during short periods on order of 1–10 s, even at low expression levels of ≈1 copy/µm2 for both molecules (Supplementary Tables 1 and 2).
This indicates that ORs would perform their functions while undergoing rapid monomer-dimer interconversions. If the functionalities of monomers and dimers are different, this raises the possibility to fine-tune the OR functions by modulating monomer-dimer interconversions by natural agonists and synthetic drugs.
Lack of morphine-induced internalization of MOR and DOR even in MD cells
Transient DM heterodimerization might affect agonist-induced intracellular signaling and internalizations of MOR and DOR, thereby affecting both the analgesic efficacy and the tolerance development. To assess how transient DOR–MOR (DM) heterodimerization affects receptor internalization (Fig. 9a), we analyzed agonist-induced responses using three CHO-K1-derived cell lines: (1) M-cells (SNAPf-MOR), (2) D-cells (SNAPf-DOR), and (3) MD-cells co-expressing both MOR and DOR (tagged with SNAPf and Halo7; see Methods). The total expression levels of the receptors are 1.5 ± 0.5 fluorescent spots/µm². Receptors were labeled for TIRF-based internalization assays, with Dpep(20–42)DM (1 µM) used to disrupt DM heterodimers.
a Schematic figure showing that Dpep(20-42)DM reduces DM heterodimers, helping to dissect the DM heterodimer and MOR monomer functions, including signaling and internalization, upon M-agonist addition. b Time courses of MOR and DOR internalization before and after the addition of 0.5 µM morphine or DAMGO in the presence and absence of 1 µM DpepDM (mean ± SEM; 10 cells for each data point). MOR internalization was examined in MD-cells and M-cells, whereas DOR internalization was observed only in MD-cells c Schematic illustration showing immensely enhanced efficiencies of binding to internalization machineries such as GRKs and β-arrestins and of DM*DAMGO heterodimer internalization. d (Top) Experimental design for observing Ca2+ mobilization after M-agonist addition. Ca2+ mobilization was monitored by Fluo-4 fluorescence intensity in M-, D-, and MD-cells. (Bottom) Typical (among 20 cells) Fluo-4 images and single-molecule images of DOR and MOR. e Ca2+ mobilization parametrized by using [FMax-Fb]/Fb for Fluo-4 signals before and after the addition of morphine and DAMGO, in the presence and absence of Dpep(20-42)DM. Horizontal bars, crosses, boxes, and whiskers indicate median values, mean values, interquartile ranges (25–75%), and 10-90% ranges, respectively. * and ns represent significant (p < 0.05) and non-significant (p ≥ 0.05) differences, respectively (Tukey’s multiple comparison test). f Schematic illustration showing the osmotic pump implantation for intracerebroventricular injection of Dpep(20-42)DM and subcutaneous morphine injection in a mouse [Created in BioRender. Aladag, A. (2025) https://BioRender.com/0tevzb7]. g Tail flick test results on day 11, showing that morphine-induced analgesia was maintained by the continuous administration of Dpep(20-42)DM (10 µg/day). The analgesia effect lasted for at least 120 min. h Dpep(20-42) reduces morphine tolerance in mice. For g and h, * represents significant (p < 0.05) difference (two-way ANOVA test; the data show mean ± SEM; n = 17 DpepDM group, n = 8 control peptide group). All of the statistical parameters and analysis results including sample size n and p values are provided in Supplementary Data 2. Source data are provided as a Source Data file.
The time courses of OR internalizations were monitored for 55 min by measuring the OR molecules remaining in the PM. The OR amounts in the PM were measured by TIRF microscopy, by evaluating the signal intensities of the OR molecules in the TIRF illumination range (in PM plus cytoplasm within ≈100 nm from the PM) before and after the addition of a membrane-impermeable fluorescence quencher, Mn(III) meso-tetra(4-sulfonatophenyl)porphine (Mn3+-TSP) (i.e., the intensity of the fluorescence signal from the cytoplasm in the TIRF illumination range), and then subtracting the latter from the former see Methods)1. Each time course obtained under various conditions could be fitted with a single exponential function plus a constant. The constant provided the OR fraction whose internalization is undetectable within 55 min, and the exponential decay constant provided the residency lifetime in the PM for the OR’s internalized fraction observed during 55 min (Fig. 9b and Supplementary Figs. 6 and 7; for summaries and statistical test results see Supplementary Table 4 and Supplementary Data 2).
MOR internalization before and after morphine addition was hardly detectable in either M- or MD-cells and was unaffected by Dpep(20–42)DM (Fig. 9b, Left column, Top and middle; Supplementary Fig. 6, Top-fourth rows), indicating that morphine-bound MOR (M*mor), whether as monomer, homodimer, or heterodimer, is internalized inefficiently.
However, these observations require cautious interpretations due to the low fractions of M and M*mor existing as DM and DM*mor heterodimers, respectively, under the employed expression levels ( ≈ 5% as shown in Supplementary Table 2; see the column of 1 + 1 copies/µm2; i.e., the case approximating experimental observation conditions of 0.5–1.0 fluorescent spots/µm2 for both MOR and DOR), although, as described in the previous subsection, virtually all OR molecules would experience dimer periods within 1–10 s. In certain PM domains in some neurons, the MOR and/or DOR concentrations may far exceed those used in this study (e.g., MOR in primary cilia)54, and the morphine effect on the MM and/or DM dimer internalizations could be more pronounced.
Highly-enhanced M*DAMGO internalization during transient binding to DOR
In contrast to morphine, DAMGO strongly induced DAMGO-bound MOR (M*DAMGO) internalization in both M- and MD-cells (Fig. 9b, Supplementary Fig. 6, and Supplementary Table 4). Internalization was greater in MD-cells (48%) than M-cells (30%), despite only ~9% of M*DAMGO existing as DM*DAMGO heterodimers. Blocking DM dimerization with Dpep(20–42)DM reduced internalization to 26%. Although the co-internalization of MOR and DOR following DAMGO exposure has been documented previously13,55,56,57,58, the critical insight obtained here is that DM*DAMGO heterodimers, despite their very low fraction (9.0%) and short lifetime (369 ± 11 ms; Fig. 6c, d) even at a low expression level of 1 copy/µm2 for each molecular species, contribute to a significant proportion of M*DAMGO internalization.
These observations suggest an unanticipated regulatory mechanism for M*DAMGO internalization. Namely, at the moment of DM*DAMGO heterodimer formation, its ability to bind to cellular internalization machineries, such as GRKs and β-arrestin 259, might suddenly rise dramatically, and then the binding affinity would rapidly decline due to DM*DAMGO heterodimer dissociation, which would occur within a fraction of a second (1/koff = 0.37 s; Supplementary Table 2). Hence, the cycle of DM*DAMGO heterodimerization and dissociation (which occurs every few seconds or so) would function analogously to a car driver adjusting the speed by frequently and briefly pushing on and releasing the gas pedal, to regulate the M*DAMGO internalization rate (Fig. 9c). Although the enhancement of internalization might occur only during the brief periods when DM*DAMGO heterodimers are formed (Fig. 9c), the total frequency of heterodimerization across the PM ensures that the overall internalization of M*DAMGO as DM*DAMGO heterodimers is significant. This correlates with the area under the curve consisting of many short pulses, depicted in Fig. 9c60,61,62. Thus, the overall internalization rate of M*DAMGO could be modulated by the frequencies and durations of transient DM*DAMGO heterodimerization events.
M*DAMGO monomer/homodimer internalization would occur much less efficiently at the level of individual molecules, but since larger fractions of M*DAMGO exist as monomers at any moment, the overall contributions of M*DAMGO monomers/homodimers to M*DAMGO internalization are quite significant (Fig. 9c). If the expression levels of MOR and DOR are elevated in certain cell types, then M*DAMGO internalization would predominantly occur through DM*DAMGO heterodimers.
We also examined the effects of the DOR-specific agonist SNC-80 on receptor internalization63. The treatment led to extensive internalization of SNC-80-bound DOR (D*SNC) in both D- and MD-cells, with similar extents and rates (Supplementary Fig. 7, middle and right columns; Supplementary Table 4), indicating that DOR*SNC is internalized without the influence of heterodimerization with MOR. Meanwhile, SNC-80 induced significant MOR internalization in MD-cells, which was reduced by the further addition of Dpep(20-42)DM (Supplementary Fig. 7, left column; Supplementary Table 4), indicating that MOR is internalized when it forms heterodimers with D*SNC in MD-cells.
Transient DM*Ago heterodimers produce strong short pulse-like signals
We examined the cytoplasmic signals triggered by M-agonists, morphine and DAMGO, by monitoring the intracellular Ca2+ mobilization in cells stably expressing Gqi5 and MOR, which were further transiently transfected to express DOR or KOR (called M-, MD-, and MK-cells in this subsection; Fig. 9d; cf Supplementary Fig. 1a and Methods)1. All ORs were expressed at the same low expression levels employed for single-molecule detection of dimers (0.5–1.0 fluorescent spots/μm2 for each OR).
In M-cells, both morphine and DAMGO induced comparable Ca²⁺ signals, unaffected by Dpep(20–42)DM. No response was evident in D-cells or K-cells. In contrast, MD-cells exhibited enhanced Ca²⁺ responses to both agonists (Ca2+ signal increase was greater with DAMGO than morphine) compared to M-cells under the same conditions. This heterodimer-dependent enhancement was mitigated by Dpep(20-42)DM, but not by a peptide with a scrambled sequence derived from Dpep(20-42)DM, indicating that DM heterodimerization amplifies M*Ago signaling (M*Ago denotes agonist-bound MOR, collectively indicating M*mor and M*DAMGO). To further validate this observation of Ca2+ mobilization via Gqi5, we examined cAMP reduction via Gi proteins using a cAMPinG1-based cAMP assay64,65,66,67,68,69. Unlike the Ca2+ mobilization assay, both MOR and DOR were expressed at levels of 5-10 fluorescent spots/µm2 (5-10 times higher than for Ca2+ assays and single-molecule observations). Consistent with the Ca2+ mobilization data, the DAMGO-induced cAMP concentration decrease via Gi was greater when DOR was co-expressed, and this enhancement was suppressed by Dpep(20–42)DM (Supplementary Fig. 8). These results are consistent with a previous report indicating that morphine and DAMGO activate MOR with greater efficacy when DOR is co-expressed over MOR alone, but without explicitly estimating the percentages of MOR existing as DM heterodimers53.
We found very small fractions of M*Ago molecules existing as DM*Ago heterodimers (4.9% for DM*mor and 9.1% for DM*DAMGO at low experimental expression levels of ≈1 + 1 copies/µm2 in Supplementary Table 2) for very brief durations (369 ± 11 ms for DM*DAMGO and 187 ± 20 ms for DM*mor; Fig. 6c). The M*DAMGO monomer and homodimer generated similar Ca²⁺ signals, suggesting that each M*AGO molecule in a DM*Ago heterodimer exhibits a markedly higher signaling rate compared to monomeric or homodimeric M*Ago molecules. This enhancement may arise from conformational changes in M*Ago upon DOR binding and/or those in DOR upon M*Ago binding, potentially allowing non-ligated DOR to participate in signaling. The strong signals from a small population of short-lived DMAgo heterodimers (369 ms) suggest a fundamentally important signaling mechanism, similar to the enhanced internalization with DMDAMGO heterodimers (previous subsection).
Namely, at the moment of DM*Ago heterodimer formation, the probability that the M*Ago molecule and possibly the M*Ago-bound DOR molecule bind to and activate Gqi5 or Gαi, consequently triggering downstream signaling cascades, might rise steeply, but this activation is short-lived because the DM*Ago heterodimer dissociates quickly within fractions of a second (1/koff = 0.19 s and 0.37 s for DM*mor and DM*DAMGO, respectively; Supplementary Table 3). Hence, a DM*Ago heterodimer will create brief, pulse-like signals, and thus the signal in the entire cell is produced as the sum of the brief, stronger pulse-like signals from individual DM*Ago heterodimers plus the weaker signals from M*Ago monomers (Fig. 9c)60,61,62. As we proposed previously60,61,62, the regulation of the signal time courses of an entire cell is more readily achieved when signaling entities (individual signaling molecules and molecular complexes) emit short pulse-like signals as opposed to prolonged ones: only simple summation (aggregation) of brief signals suffices to generate the overall cellular signal response, while if individual signals are prolonged, complex integration rather than simple summation must be performed to produce the required cellular signal time courses60. In short, the cellular signal intensities might be regulated by adjusting the frequencies of the short pulse-like formation of DM*Ago heterodimers. In addition, in cells or PM domains with elevated but still physiological OR molecular densities, the signals emanating from DM*Ago heterodimers could be quite substantial, governing the cellular signal.
Dpep(20-42) reduces morphine tolerance in mice
As described previously, Dpep(20-42)DM moderately reduces M*Ago-induced signaling, suggesting potential impacts not only at the cellular level but also in nervous tissues and systemic functions. To explore this, we tested whether the soluble DM heterodimer blocker peptide Dpep(20-42)DM could modulate morphine-mediated analgesia in murine models.
The peptide, dissolved in artificial cerebrospinal fluid (aCSF), was administered directly into the cerebral ventricles at a dose of 10 µg/day continuously for two days prior to the commencement of daily subcutaneous morphine injections (10 mg/kg/injection; one injection/day) (Fig. 9f). As a control, we used the peptide derived from the scrambled sequence of Dpep(20-42)DM. We utilized the tail-flick test to evaluate the analgesic efficacy of morphine and track the progression of antinociceptive tolerance.
Our results revealed that the analgesic potency of morphine was enhanced by Dpep(20-42)DM, which was especially evident between 60 and 120 minutes after morphine application over the course of the study (days 1, 5, 8, and 11; Supplementary Fig. 9). Notably, on day 11, the analgesic effect in the Dpep(20-42)DM group was approximately 3.5-fold greater than that in the control group (Fig. 9g).
The administration of Dpep(20-42)DM reduced morphine tolerance development, with effects becoming evident from day 8 after treatment initiation. Dpep(20-42)DM-treated mice retained >87% of their initial analgesic efficacy through days 8–11, whereas the control group retained only ≈27% (Fig. 9h). Mice receiving half the peptide concentration exhibited intermediate tolerance levels (Supplementary Fig. 9). These results suggest that inhibiting DM* heterodimerization by Dpep(20-42)DM diminishes morphine tolerance, implicating MOR–DOR co-expressing neurons, consistent with previous studies10,13,14,17. Suppression of DM* heterodimers by Dpep(20-42)DM also diminished the signal levels in cells in vitro (Fig. 9e). Accordingly, maintaining proper signal levels in cells might be important for sustained morphine analgesia, which will be discussed further.
Discussion
The significance of OR heterodimerization in pain therapy and neuropsychiatric disorders is garnering increasing attention34,35,70. Although there is controversy in the OR research field as to whether different ORs co-exist in the same neuron, the coexistence of MOR and DOR, which has been most extensively investigated, is becoming clear in several domains or neuronal circuits in the brain and peripheral nervous system under various physiological, pathological, and pharmacological conditions16,20,21,22,23. The heterodimerization of three classical ORs introduces a new dimension to receptor interactions with G proteins, GRKs, and arrestins, providing additional opportunities for modulating OR functions13,58,71,72,73. This modulation could be clinically utilized for managing chronic pain and treating neuropsychiatric disorders2,34,35,70.
However, our understanding of OR heterodimers and their clinical implications is limited due to the paucity of fundamental knowledge on heterodimer formation dynamics and mechanisms and the lack of specific blockers. Using our advanced single-molecule imaging and analysis methods1, we obtained the fundamental parameters KD, koff, and kon, describing OR homo- and hetero-dimerizations. Crucially, these parameters are independent of OR expression levels, and thus we firmly established the formation of transient DM and DK heterodimers, alongside KK, MM, and DD homodimers, in the PM of cells expressing these molecules. These parameters were determined in the presence and absence of M-agonists and Dpep(20-42)DM (Supplementary Tables 1 and 2), and the percentages of protomers existing as DM and DK heterodimers at various expression levels were calculated from the KD values (Supplementary Tables 2–3 and Fig. 7), which will be useful for future studies to reveal the biological and pharmacological consequences of OR homo- and hetero-dimers. We hardly detected transient OR trimers and tetramers, indicating that at low physiological expression levels, the percentage of OR protomers existing as oligomers greater than dimers would be limited (Fig. 8).
We did not detect MK dimers at expression levels of 1.5 ± 0.5 total fluorescent spots (for two OR molecules) employed here. However, since MK heterodimers have been proposed to exist in female spinal cords24, it is possible that the MOR and KOR expression levels may be much higher and MK heterodimers may form there.
This study first documented the transient nature of OR heterodimers, revealing lifetimes of DM and DK heterodimers of approximately 250 ms, roughly twice as long as those of homodimers (120 ms for DD and MM and 180 ms for KK). Previous single-molecule imaging studies likely missed the OR dimers due to their scarce presence at the low expression levels typically employed in these studies ( ≈ 1 copy/µm2) and their brief lifetimes74,75,76. Indeed, experiments performed at higher protein densities using bioluminescence resonance energy transfer (BRET), fluorescence resonance energy transfer (FRET), and biochemical assays detected the dimers, although these methods could not elucidate the dynamic nature of OR dimerization44,75,77,78,79. Interestingly, pull-down assays, which require minute-order procedures, detected homo- and hetero-dimers despite their 100-ms-order lifetimes10,80, probably due to the rapid reassociation kinetics.
In a separate study using a Gs-coupled GPCR, β2-adrenergic receptor (β2AR), our group found that it also undergoes intermittent, frequent, transient dimerizations in similar time scales, and interestingly, Gs proteins are recruited to both β2AR homodimers and monomers with binding lifetimes shorter than 50 ms81. This time scale is shorter than the receptor dimer lifetimes, suggesting that these short-lived dimers formed frequently can affect Gs-downstream signals and might be important for regulating GPCR downstream signals.
The interaction domains critical for heterodimerization have been identified as both the N-terminal domains for DM and both the EL3 domains for DK heterodimers (Figs. 2–4). While TM1MOR is involved in DM heterodimerization by binding to TM1DOR, TM4DOR, and other TMs of DOR (Fig. 5b–d), it is also involved in MM homodimerization by binding to another MOR (perhaps mainly the TM1 domain). Therefore, although TM1MOR contributes to dimerization, its contribution is less specific and would primarily be to augment overall affinity. The nature of lower-specificity TM interactions is also found in the literature. The involvement of TM domains in DM heterodimerization13,48 and DK heterodimerization was previously proposed28, whereas the same TM domains such as TM4DOR and TM1MOR were also proposed to be involved in DD82 and MM50,83 homodimerizations. The TM interactions with lower specificities were previously proposed as the rolling dimer interface model, in which multiple TM interaction sites co-exist and interconvert44. In line with these observations, while MOR homodimerization was previously found to depend on the interaction between TM5 and TM684, in the presence of galanin Gal1 receptor, another GPCR, the homodimerization site shifts to the TM4/TM5 interface and MOR-Gal1 receptor heterotetramer is formed via TM5/TM6 interface84.
Since the addition of either Dpep(20-42)DM (also Mpep(32-61)MD) or TM1MOR alone reduced the colocalization index for DM heterodimers to near-baseline levels (1.2–1.3) (Figs. 4a, b, and 5c), we suggest that DM heterodimerization might also involve cooperative allosteric conformational changes between the N-terminal domains and TM domains in DOR and MOR, which might affect their functions85. In addition, although the binding of nucleotide-free Gα to GPCR was proposed to affect GPCR conformations86, we could not include investigations to clarify the effect of G-proteins on GPCR dimerization in the present research. Examination of this effect will be a very important next step for further advancing our understanding of OR dimerization and its functions. The possible involvement of these processes suggests the complexity of OR dimerization, which we have simplified using first-order kinetics to aid our understanding at this stage of research.
Soluble peptides with sequences mirroring the extracellular heterodimer formation domains (EL3DOR and EL3KOR peptides for DK heterodimers and DpepDM and MpepMD for DM heterodimers) can specifically reduce DM and DK heterodimers (Figs. 2d, 4, and 5d). In particular, we extensively utilized the Dpep(20-42)DM peptide to dissect the functions of M*Ago monomers/homodimers and DM*Ago heterodimers after the application of M-agonists, morphine and DAMGO (where M*Ago collectively denotes agonist-bound MOR). DM*Ago dimerization was enhanced by DAMGO but not morphine, and reduced by Dpep(20-42)DM (Fig. 6). Morphine failed to induce the internalization of M*mor or DOR even in MD-cells, where DM*mor heterodimerization is induced. M*DAMGO monomers and M*DAMGOM*DAMGO homodimers are internalized in similar manners1, and DM*DAMGO heterodimerization further enhanced M*DAMGO internalization (Fig. 9b, c). DM*Ago heterodimerization also enhanced the downstream Ca2+ signal (Fig. 9e). Monomers of a prototypical GPCR, the β2-adrenergic receptor, were previously found to efficiently activate a stimulatory heterotrimeric G protein, Gs87, but our results indicate that certain GPCRs, including MOR, might acquire enhanced activities upon heterodimer formation.
In our study, conducted under the low receptor expression levels ( ≈ 1 + 1 copies/µm2), only small fractions ( ≤ 9.1%) of M*Ago will exist as DM*Ago heterodimers at any moment (Supplementary Table 3). Combining this with the short 369-ms lifetime of DM*Ago heterodimers, we advanced an argument that each DM*Ago heterodimer will create a brief, intense pulse-like signal, contrasted with the weaker signals from M*Ago monomers, and the overall cellular signal is produced as the sum of these signals (Fig. 9c, e)60,61,62. Note that all M*Ago molecules form DM*Ago heterodimers every 1–10 s even under low expression levels (Supplementary Table 2). Enhanced internalization of M*DAMGO might also follow a similar enhancement pattern. These results were observed under expression levels far lower than those typically used in bulk optical and biochemical assays (by factors of 10-1000)77,78,79, suggesting a unique signaling mechanism in native physiological conditions.
In specific neural tissue domains or certain cell types where OR expression levels are higher54, DM*Ago heterodimers could play a more dominant role in cellular responses, dictated by the frequency of their transient active phases. Building upon this hypothesis, and recognizing the possibility that small quantities of DM*Ago heterodimers might induce critical signals in key neurons, we explored the in vivo effects of Dpep(20-42)DM administered into murine cerebral ventricles to assess its impact on morphine tolerance, unlike a previous study using systemically applied Nterm-TM1MOR-TAT13, which would non-specifically block multiple dimer forms (Fig. 5). We found that cerebral ventricle-administered Dpep(20-42)DM significantly curtails the development of long-term tolerance (Fig. 9f–h). However, directly comparing the effectiveness of Dpep(20-42)DM to Nterm-TM1MOR-TAT is challenging due to differing administration methods and the inherent specificity of Dpep(20-42)DM, which likely contributes to its reduced side effects.
Our in vivo findings align with previous findings that a genetic or pharmacological blockade of DOR can enhance MOR-agonist induced analgesia, highlighting the role of DM heterodimers in regulating MOR-mediated pain pathways10,14,17,72,88,89,90 The newly identified specific extramembrane domains for DM heterodimer-blocking present targets for modulating OR function, offering strategic avenues for the development of opioid therapies with reduced tolerance and dependence.
These in vivo results should be compared with our findings in cultured cells, where Dpep(20-42)DM, facilitating DM*mor dissociation into monomers, reduces the Ca2+ signal from Gqi5 (Fig. 9e). These apparently contrasting findings made in vivo and in vitro suggest that finely tuning cellular signals upon morphine application is crucial for preventing long-term tolerance in mice. Agents that modulate the levels of DM*mor heterodimers in the PM, like Dpep(20-42)DM, could offer more precise control over drug effects than simply varying morphine dosage.
In a recent study, small molecules that can specifically interfere with the TM5/TM6 interface of the cannabinoid CB1 receptor-serotonin 5-HT2A receptor heteromer have been developed91 based on the TM5/TM6 interface found by the TM peptide blockers92. In light of the therapeutic potential of short peptides and small molecules for clinical use, development of chemical compounds that mimic Dpep(20-42)DM and the TM peptides engaging dimer-specific interfaces and are capable of crossing the blood–brain barrier would represent a promising strategy for advancing opioid therapies with reduced tolerance and dependence.
Methods
cDNA construction
All of the newly generated cDNA constructs and other constructs obtained from outside sources, including gifts and constructs from commercial sources, were sequenced to examine their exact DNA sequences. The cDNA encoding rat MOR tagged with GFP was a gift from Dr. R. Schülz of the University of Münich, Germany93. The cDNA encoding rat KOR and DOR was a gift from Dr. Hiroshi Takeshima of the Kyoto University94. The mCherry was a gift from Prof. R. Y. Tsien of the University of California San Diego95. The cDNA encoding CD47 was a gift from Eric C. Brown of Genentech96. The cDNA encoding CD28 was a gift from Simon J. Davis of University of Oxford97. The cDNAs encoding SNAPf and mGFP (A206K) were obtained from New England Biolabs and Clontech, respectively. To generate plasmids for expressing SNAPf-ORs and ORs-Halo7 in CHO-K1 cells, the tag protein SNAPf or Halo7 was attached to the N-/C-terminus of the ORs, an additional signal sequence of interleukin 6 was attached to the N-/C-terminus of the tag protein, and a 21 amino-acid linker (SGGGSGG × 3) was inserted between the ORs and the tag protein. The deletion and point mutants of ORs were generated using a Q5® Site-Directed Mutagenesis Kit (NEB, Cat. #E0554). The detailed cDNA information is provided in the DNA sequence file (Supplementary Data 1).
Mice
Male C57BL/6 N mice (30-40 g; n = 25) were purchased from CLEA Japan, Inc. and maintained on a 12-h light/dark cycle with rodent chow and water available ad libitum. They were housed in groups of four until testing. Animal studies were performed according to protocols approved by the Nagasaki University Animal Care and Use Committee (Approved number: 2002181596).
Cell culture, transfection, and microscope observations
CHO-K1 cells (Dainippon Pharma) were confirmed free of mycoplasma contamination by MycoAlert (Lonza)98. CHO-K1 cells were routinely cultured in Ham’s Nutrient Mixture F12 (Sigma-Aldrich) supplemented with 10% (v/v) fetal bovine serum (FBS, Life Technologies), 100 units/mL penicillin (Sigma-Aldrich), and 0.1 mg/mL streptomycin at 37 °C under a 5% CO2 atmosphere. For cDNA transfections for single-molecule tracking studies of ORs (exhibiting 0.5 ~ 1 fluorescent spots/µm2), approximately 5 × 106 cells were mixed with 200 ng OR cDNA plasmids in 100 µL transfection buffer, and electroporation was performed according to the manufacturer’s instructions (4D-Nucleofector, Lonza; SF Cell Line solution and program for CHO-K1 and T24 cells). For cDNA transfections for Ca2+ mobilization and cAMP assays with Gqi5 and cAMPinG1, approximately 2 × 105 cells were mixed with 1 µg cDNA plasmids encoding Gqi5 and cAMPinG1 in 100 µL transfection buffer. The transfected cells were seeded in glass-base dishes (35-mm in diameter with a 12-mm diameter glass window, 0.15-mm-thick glass; Iwaki, Tokyo; 2 × 105 cells/dish) and cultured for 24–48 h before fluorescence microscopy observations. All microscope observations were performed at 37°C by placing the entire microscope, except for the far ends of the excitation arms and the detection arms, in a home-built microscope environment chamber made with thermo- and electric-field-insulating plastic sheets and equipped with four heating circulators (SKH0-112-OT, Kokensya, Tokyo, Japan). The Ham’s F12 medium used for microscope observations was free of sodium bicarbonate and phenol red, and buffered with 2 mM N-[tris(hydroxymethyl)methyl]-2-aminoethanesulfonic acid (TES, Sigma-Aldrich) at pH 7.4 (called Ham’s F12 observation medium).
Fluorescence labeling of ORs
The SNAPf-tagged wild-type and mutant ORs expressed in the PM (SNAPf tag located at the extracellular N-terminus) were covalently conjugated by simultaneously incubating the cells with two fluorescent SNAP ligands, SNAP-Surface 549 (New England Biolabs) and SNAP-CF660R (Shinsei Kagaku), both at 300 nM, in the growth medium at 37 °C in the CO2 incubator for 30 min. The cells were washed three times with fresh medium (5-min incubation each time), and then the Ham’s F12 observation medium was added. The Halo7-tag protein fused to the intracellular C-terminus of the OR (Supplementary Fig. 1a) was covalently conjugated with SaraFluor650T-Halo-ligand (Goryo Kayaku), by incubating the cells with 100 nM Halo ligand. The SNAPf-OR and OR-Halo7 proteins were simultaneously labeled in the growth medium, at 37 °C in the CO2 incubator for 30 min. The cells were washed three times with fresh medium (5-min incubation each time), and then the Ham’s F12 observation medium was added. Determination of the labeling efficiencies of the SNAPf-tag protein attached to the extracellular N-termini and Halo7-tag protein attached to the cytoplasmic C-termini of various proteins under monomeric conditions. CD47, TMLDLR, and MOR’s 1-70 aa sequence linked to the N-terminus of TMLDLR expressed at a fluorescent spot number density of ≈1 spot/µm2 and MOR and MOR’s ∆358-382 mutant (monomeric mutant developed in this study) expressed at a spot number density of <0.1 spot/µm2. For determining the Halo7-tag labeling efficiency, these proteins are linked to the monomeric StagyGold (mSG)-tag at their N-termini99. Results showed that the labeling efficiencies are very similar among the five proteins employed here, providing efficiencies of 0.6631 ± 0.0107 for SNAP-Surface 549 at a SNAP-dye concentration of 300 nM and 0.8585 ± 0.0046 for SaraFluor650T (mean ± SEM of five proteins) at a Halo-dye concentration of 100 nM. These results suggest that the labeling efficiencies of Halo-SaraFluor650T in terms of binding to the Halo7-tag protein would generally not be affected much by the domains of the proteins of interest conjugated to the SNAP-tag protein, like the case where SNAP-Surface 549 and SNAP-CF660R binding to the SNAPf-tag protein would generally not be affected much by the domains of the proteins of interest conjugated to the SNAP-tag protein. We suspect that the somewhat long 12-amino acid linker we employed for conjugating the Halo7-tag protein to the protein of interest (SGSG ×3) might help to provide similar labeling efficiencies for different proteins. By controlling the expression levels and using these labeling conditions, the spot numbers of SNAPf-OR and OR-Halo7 proteins were approximately equalized (0.75 ± 0.25 spots/µm2 for each color; i.e., a total spot number density of 1.5 ± 0.5 spots/µm2).
Single molecule fluorescence imaging
Fluorescently labeled ORs expressed in the bottom PM (the PM facing the coverslip) at fluorescent-spot number densities of 0.75 ± 0.25 spots/µm2 for each color (a total spot number density of 1.5 ± 0.5 spots/µm2) were observed at the level of single molecules at 37 °C, using a home-built objective lens-type TIRF microscope constructed on an inverted microscope (Olympus IX-83) with a 100x, 1.49 numerical aperture (NA) objective lens, optimized for the present research based on the instrument used previously100,101. ORs tagged with fluorescent probes were excited with TIR illumination using the following power densities: SNAP-Surface 549 at 561 nm (Coherent OBIS 561-100 LS) at 0.35 µW/µm2; and SaraFluor650T at 642 nm (Omicron LuxXPlus 640-140) at 0.50 µW/µm2. The dual-color images were separated by a dichroic mirror ZT640rdc-UF3 (Chroma) and projected into two detection arms with band-pass filters of 575-625 nm (ET600/50 m; Chroma) for the SNAP-Surface 549 dye and 662.5–737.5 nm (ET700/75 m; Chroma) for the SaraFluor650T dye. The fluorescence signal in each channel was first detected and amplified by a two-stage microchannel-plate image intensifier (C9016-02MLP24; Hamamatsu Photonics), and the intensified image was projected onto the scientific CMOS camera (C11440-22CU; Hamamatsu Photonics), operated at 30 Hz, which was synchronized with the same intensifier-camera set(s) placed on another detection arm. The image sequences in each channel were superimposed after correction for spatial distortions, as described previously. The positions (x and y coordinates) of all of the observed single fluorescent molecules were determined by an in-house computer program102.
Evaluating colocalization durations
The colocalization of two fluorescent molecules was defined as the event where two fluorescent spots, representing these molecules, become localized within 200 nm of each other, as described previously100,103. Each time we found a green-magenta pair located within 200 nm (colocalization), we measured the duration in which their distances remained within 200 nm (colocalized duration) (for the results obtained by using two dye molecular species with different excitation/emission wavelengths, we call them green and magenta probes/movies for convenience in this report). After obtaining the colocalization durations for all of the colocalization events, we generated a histogram (distribution) of colocalization durations. The distribution of incidental colocalization durations was obtained by superimposing the magenta image sequences with the 180-degree rotated (doubly flipped) green image sequences1. For the precise analysis of these histograms, we first produced cumulative histograms and fitted them with a single exponential function (C0 – C1exp[-t/τ1]) or the sum of two exponential functions (C0 – C1exp[-t/τ1] - C2exp[-t/τ2]). The choice of the number of exponentials (including the cases of the sum of three exponential functions) was made based on Akaike’s and Bayesian information criteria (they always agreed). Based on these functions, the functions describing the original histograms were derived and overlaid on the histograms.
First, we found that the distribution of incidental colocalization durations could be fitted by a single exponential function, with a time constant representing the incidental colocalization lifetime τinci. Second, the distribution of colocalization durations was obtained for correctly superimposed magenta and green image sequences. This distribution could be fitted by the sum of two exponential decay functions (τ1 and τ2; τ1 < τ2,). These results are consistent with the theory developed here that predicts the distribution of colocalization durations based on the diffusion equation1. Since τ1 was almost the same as τinci, τ2 provided the homodimer lifetime (this value has to be corrected for the trackable duration lifetimes of the two fluorescent probes, to obtain koff). The trackable duration lifetime (τtrack) for SNAP-Surface 549 bound to SNAPf-MOR was 16.3 ± 1.2 s (n = 500; the same as those described in the companion paper) and that for SaraFluor650T-Halo-ligand bound to MOR-Halo was 12.5 ± 0.8 s (n = 500) (expressed in CHO-K1 cells) (Supplementary Fig. 1f). The photobleaching lifetimes for these fluorescent dye molecules adsorbed on the cover glass of the glass-base dish were 36.6 ± 1.0 s (n = 567) for SNAP-Surface 549 and 20.1 ± 0.10 s (n = 598) for SaraFluor650T.
Evaluating the colocalization index
For the quantitative evaluation of the extent of colocalization (representing both the frequency and lifetime of colocalization events) in simultaneous two-color single-molecule imaging movies, we defined a parameter called the colocalization index104. This analysis method is essentially based on a pair cross-correlation analysis1.
Monte Carlo simulations
The colocalization index depends on the number density of fluorescent spots in the PM. In the present experimental observations, we selected the cells exhibiting fluorescent spot number densities in the PM of 0.75 ± 0.25 spots/µm2 for each color (SNAP-Surface 549 and SaraFluor650; total spot number densities of 1.5 ± 0.5 spots/µm2; for conciseness, we describe the number densities as ≈1.5 spot/µm2 throughout this report). These densities were found to be reasonable by the Monte-Carlo simulation results. We developed the theory to evaluate the dimer-monomer dissociation equilibrium constant KD from the PCCF and the total number of spots in the image, refer to Supplementary Note 1 of this paper and Supplementary Note 2 of companion paper1.
Cell treatments with agonists and peptides
M-agonists, DAMGO (Sigma-Aldrich) and morphine (Takeda), and D-agonist SNC-80 (Sigma-Aldrich) were applied to the cells as described in the companion paper. The DpepDM and MpepMD peptides, as well as the peptides with scrambled sequences derived from DpepDM and MpepMD: (1) PSSSSGSTPAGFADPAARFNASS, (2) CVPRCQGSNGQHDGGTSGGNPDDNLLSLTL, custom-synthesized by Cosmo-Bio, are soluble peptides, and were dissolved in the observation medium and added to the cells in the observation medium at a selected final peptide concentration at 37 °C. Since their target sites are located on the cell surface, the complex procedures employed for incorporating the FAM-Xpep-TATs inside the cells, as described in the companion paper, were unnecessary1.
Peptide stability test
Peptide solutions (1 mM in 10% DMSO, 90% H2O) were mixed with mouse serum (Sigma-Aldrich, USA) or the CHO-K1 cell culture medium at 1:1 (v/v), and incubated at 37 °C with gentle shaking. At different timepoints, 120-µL aliquots were taken, and the protease activity was stopped with 20 µL of 10% (v/v) trichloroacetic acid. After 30 min at 4 °C, samples were centrifuged at 13,000 g for 10 min to remove serum proteins, and the supernatants were analyzed by HPLC-UV (Shimadzu HPLC Nexera LC30-AD, equipped with an SPD-M20A (PDA), CTO-20AC (column oven), SIL-30AC (Auto Sampler), and AscenticsExpress ODS 150 x ID 2.1 mm, 2.7 μm column. Flow: 0.3 mL/min; Inj: 5 μl; Column Oven: 40 °C; Detection: λ 215 nm (Cell Temp. 40 °C).
OR internalizations
We examined the effects of DM heterodimerization on downstream signals after M-agonist stimulation and the internalization of MOR and DOR before and after M- and D-agonist stimulation (0.5 µM), using three cell lines produced here: (1) M-cells, CHO-K1 cells stably expressing SNAPf-MOR at 0.5–1.0 fluorescent spots/µm2 after labeling with SNAP-Surface 549; (2) D-cells, CHO-K1 cells stably expressing SNAPf-DOR at 0.5–1.0 fluorescent spots/µm2 after labeling with SNAP-Surface 549; (3) MD-cells based on M-cells (or D-cells), transiently transfected with the cDNA encoding DOR-Halo7 (or MOR-Halo7), which was labeled with SaraFluor 650 T (cells exhibiting 0.5-1.0 fluorescent spots/µm2 were selected for observations). The two types of MD-cells (based on M-cells or D-cells) are biologically the same (thus, in the following, we simply call them MD-cells), apart from the experimental details (i.e., for the assay of MOR and DOR internalizations, SNAPf-MOR and SNAPf-DOR must be used, respectively, because our assay depends on the quenching of fluorescence signal from these molecules located on the cell surface using the membrane impermeable quencher; see below). Dpep(20-42)DM (1 µM) was extensively used to dissect the functions of DM heterodimers from those of D/M monomers and DD/MM homodimers.
Ca2+ mobilization assay
Ca2+ mobilization was parameterized by using [FMax-Fb]/Fb, where FMax is the Fluo-4 peak signal intensity within 75 s after the addition of the stimulants and Fb is the baseline signal intensity1. When we observed Ca2+ responses using the cells expressing SNAPf-MOR, we employed the CHO-K1 cells stably expressing both Gqi5 and SNAPf-MOR (at number densities of 0.75 ± 0.25 spots/µm2 when labeled with SNAP-Surface 549). To observe the effect of DOR-Halo and KOR-Halo co-expression, the cells stably expressing Gqi5 and SNAPf-MOR were transiently transfected with the cDNAs encoding these proteins. The cells exhibiting spot densities of DOR-Halo or KOR-Halo at 0.75 ± 0.25 spots/µm2 (after labeling with Halo-SaraFluor650T) were selected for observations. Therefore, the Ca2+ mobilization assay was performed at about the same low expression levels of ORs as those employed for single fluorescent-molecule imaging experiments. For the comparison of the functions of the SNAPf-ORs with those of the non-tagged ORs, we hoped to compare the cells expressing the SNAPf-ORs and ORs-Halo7 at the levels of ≈1 spot/µm2 in the basal PM with the cells expressing the non-tagged ORs at levels comparable to or higher than those of SNAPf-ORs or ORs-Halo7, because the cells expressing the non-tagged ORs should serve as the positive controls. For this purpose, the expression levels of non-tagged ORs were monitored by using cells transfected with the cDNA linking the non-tagged OR sequence to the mCherry sequence via the self-cleavable 2 A linker sequence (mCherry-2A-OR). This way, mCherry is released from the OR into the cytoplasm at the ER, and the non-tagged OR is then transported to the PM. The expression of non-tagged OR was detected by the presence of mCherry in the cytoplasm using the epi-illumination at 561 nm (sensitivities much lower than single-molecule imaging, showing the presence of rather high concentrations of mCherry). Therefore, the expression of the non-tagged OR used in this study is considered higher than that of SNAPf-ORs or ORs-Halo7.
cAMPinG1 cAMP assay
CHO-K1 cells stably expressing SNAPf-DOR were transfected with the cDNAs encoding cAMPinG169. Approximately 2 × 105 cells were mixed with ≈1 µg cAMPinG1 plasmids in 100 µL transfection buffer, and electroporation was carried out with a 4D-Nucleofector electroporation apparatus, according to the manufacturer’s instructions (4D-Nucleofector, Lonza; SF Cell Line solution and program for CHO-K1 cells). The transfected cells were seeded in glass-base dishes and cultured for 24–48 h before fluorescence microscopy observations. All microscope observations were performed at 37 °C.
For the cAMP inhibition assay, cells expressing SNAPf-DOR and MOR-Halo7, labeled with SNAP-Surface 549 and SaraFluor650T, respectively, on the basal PM were selected using TIRF microscopy with 561-nm and 642-nm excitations. Time-lapse imaging of cAMPinG1 was performed every 30 s with 100-ms 405-nm and 488-nm laser illuminations (sequential 100 ms × 2). After 5 min of baseline imaging, forskolin (Sigma-Aldrich) was added to a final concentration of 7 µM. Following 10 min of further imaging, DAMGO was added to a final concentration of 50 nM, and imaging continued for an additional 10 min. Image sequences were analyzed as previously described69.
Quantifying the internalization of SNAPf-ORs
We examined OR internalization, which was monitored by using the membrane-impermeable fluorescence quencher, Mn(III) meso-tetra(4-sulfonatophenyl)porphine (Mn3+-TSP). This quencher only suppresses fluorescence emission from the SNAP-Surface 549 dye on the SNAPf-OR on the cell surface, but not that in the cytoplasm105. Accordingly, by subtracting the signal intensity after the quencher addition from that before the addition, we evaluated the percentages of OR molecules remaining in the PM. The time courses of the decrease in the numbers of ORs remaining in the PM during 55 min (due to their internalization) were examined in both the presence and absence of 0.5 µM agonists and ≈1 µM Dpep(20-42)DM in the cytoplasm.
Systemic morphine administration to mice and examination of its antinociceptive effect
Morphine hydrochloride (Takeda Pharmaceutical) was dissolved in saline and subcutaneously administered to mice once each day (10 mg/kg) for 11 days. Antinociception was measured by the tail-flick test on days 1, 5, 8, and 11 from 0 to 120 min after morphine administration (details will be described later). Body weight was measured before each morphine injection. The control group received saline without morphine. All animal experiments were randomized and performed by a blinded researcher, who was then unblinded before statistical analysis.
Intracerebroventricular administration of the Dpep(20-42)DM peptide in mice
The Dpep(20-42)DM peptide was dissolved in freshly prepared aCSF (127 mM NaCl, 1.5 mM KCl, 1.24 mM KH2PO4, 1.4 mM MgSO4, 26 mM NaHCO3, 2.4 mM CaCl2, 10 mM D-glucose). For continuous intracerebroventricular administration, an osmotic minipump (Alzet model 2002) filled with the peptide solution was implanted according to the manufacturer’s protocol and as described previously106. In order to stabilize the pump flow rate, the pumps were primed by placing them in sterile saline at room temperature overnight. The osmotic minipumps were filled with 200 µL of aCSF with or without the peptide solution (0.83 mg/mL in aCSF). The minipump was connected to a 2.5-mm-long cannula (Alzet Brain Infusion Kit 3) via a 1.5-cm-long polyvinylchloride tube. Before surgery, the mice were anesthetized with a combination anesthetic (0.75 mg/kg of medetomidine, 4.0 mg/kg of midazolam, and 5.0 mg/kg of butorphanol). The scalp was shaved, Povidone-Iodine (Isodine, Mundipharma K.K.) was applied, and an incision was made along the midline of the scalp; hemostats were then used to make a pocket under the skin between the shoulder blades. The skull was scraped clean of periosteum so that the cannula can properly adhere to the skull. A microdrill (Tamiya, Inc.) was used to create a hole approximately 1.1 mm lateral and 0.5 mm caudal to the bregma. The minipump was placed in the pocket under the skin between the shoulder blades, the cannula was inserted through the drilled hole into the lateral ventricle, and the cannula pedestal was affixed to the skull with cyanoacrylate glue and resin cement. The incision was closed with soft silk sutures and antibacterial ointment was applied to the wound. The animals were allowed to rest on a disposable heating pad until they were recovered by using 0.75 mg/kg of atipamezole and then returned to their home cages in the animal facility for 1–2 days until the measurement. Cannula placement into the lateral ventricle was verified with trypan blue (4%, 10 μl).
Morphine-induced antinociception assay
Morphine-induced antinociception was evaluated by using the tail-flick test15. Using a tail-flick apparatus (IITC Life Science), the intensity of the heat source was set at 10, which resulted in a basal tail-flick latency of 2–3 s for most of the animals. The tail-flick latency was recorded before (0 min, baseline latency) and every 30 min after morphine injection (30, 60, 90, and 120 min). The results are presented as a percentage of the maximal possible effect, %MPE, which was calculated for each mouse at each time point according to the following formula: %MPE = [(latency after drug − baseline latency)/ (10 − baseline latency)] × 100. Cutoff latency was selected at 10 s to minimize tissue damage. The area under the %MPE vs. time curve (AUC) for each treatment condition was also used to evaluate antinociception (Fig. 9h). Normal distribution of the data was verified before performing the parametric statistical analysis. Wherever appropriate, data were analyzed using two-way ANOVA, followed by Tukey’s post hoc tests or a multiple unpaired t-test with significance set at P < 0.05. All calculations were performed using the GraphPad Prism 9 software (GraphPad Software).
Software and statistical analysis
The microscope station that combined a single-molecule imaging system and a super-resolution confocal microscope was controlled by in-house LabVIEW2018-based software, and the single-molecule movie acquisitions were performed using the MCR software (Hamamatsu Photonics) for Windows. All statistical analyses for in vitro experiments were performed with Tukey’s multiple comparison test except for the analysis of the colocalization lifetime data, which was performed with the Brunner-Munzel test, using RStudio 1.2.1335 for Windows. P values less than 0.05 were considered statistically significant. The confocal images were processed and analyzed using ImageJ for Windows. Curve fitting was performed by OriginPro 2019b for Windows. The simulation study was performed using the in-house software based on MATLAB 2019a for Windows. Statistical parameters, including the numbers of movies, cells, and independent replicates as well as p values, are provided in Supplementary Data 2. We used the Chai Discovery web https://lab.chaidiscovery.com/ interface to predict the homodimer structures for the C-terminal cytoplasmic domains. The resulting highest-ranked model was visualized as cartoons highlighted with the stick-models, using PyMol 3.1 by Schrödinger. Figures and videos were edited using Photoshop CC 2021, Illustrator CC 2021 (Adobe).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Source data are provided with this paper. All data necessary to support the conclusions of this study are provided in the main text, main figures, Supplementary Information, and Source data file. Raw movies and images used for quantification are available from the corresponding authors upon request. Source data are provided with this paper.
References
Zhou, P. et al. Single-molecule methods for characterizing receptor dimers reveal metastable opioid receptor homodimers that induce functional modulation. Nat. Commun. https://doi.org/10.1038/s41467-025-64694-3 (2025).
Valentino, R. J. & Volkow, N. D. Untangling the complexity of opioid receptor function. Neuropsychopharmacology 43, 2514–2520 (2018).
Levac, B. A. R., O’Dowd, B. F. & George, S. R. Oligomerization of opioid receptors: generation of novel signaling units. Curr. Opin. Pharmacol. 2, 76–81 (2002).
Schmid, C. L. et al. Bias factor and therapeutic window correlate to predict safer opioid analgesics. Cell 171, 1165–1175.e1113 (2017).
Darcq, E. & Kieffer, B. L. Opioid receptors: drivers to addiction?. Nat. Rev. Neurosci. 19, 499–514 (2018).
Chan, H. C. S., McCarthy, D., Li, J., Palczewski, K. & Yuan, S. Designing safer analgesics via μ-opioid receptor pathways. Trends Pharmacol. Sci. 38, 1016–1037 (2017).
Cong, X. et al. Molecular insights into the biased signaling mechanism of the μ-opioid receptor. Mol. Cell 81, 4165–4175.e4166 (2021).
Ferré, S. et al. G protein–coupled receptor oligomerization revisited: functional and pharmacological perspectives. Pharmacol. Rev. 66, 413 (2014).
Pan, Z. Z. -Opposing actions of the κ-opioid receptor. Trends Pharmacol. Sci. 19, 94–98 (1998).
Gomes, I. et al. A role for heterodimerization of μ and δ opiate receptors in enhancing morphine analgesia. Proc. Natl. Acad. Sci. USA 101, 5135 (2004).
Gupta, A. et al. Increased abundance of opioid receptor heteromers after chronic morphine administration. Sci. Signal 3, ra54–ra54 (2010).
Olson, K. M. et al. Synthesis and evaluation of a novel bivalent selective antagonist for the mu-delta opioid receptor heterodimer that reduces morphine withdrawal in mice. J. Med. Chem. 61, 6075–6086 (2018).
He, S. Q. et al. Facilitation of μ-Opioid receptor activity by preventing δ-opioid receptor-mediated codegradation. Neuron 69, 120–131 (2011).
Chefer, V. I. & Shippenberg, T. S. Augmentation of morphine-induced sensitization but reduction in morphine tolerance and reward in delta-opioid receptor knockout mice. Neuropsychopharmacology 34, 887–898 (2009).
Gomes, I. et al. Identification of a μ-δ opioid receptor heteromer-biased agonist with antinociceptive activity. Proc. Natl. Acad. Sci. USA 110, 12072 (2013).
Wang, D. et al. Functional divergence of delta and mu opioid receptor organization in CNS pain circuits. Neuron 98, 90–108.e105 (2018).
Zhu, Y. et al. Retention of supraspinal delta-like analgesia and loss of morphine tolerance in delta opioid receptor knockout mice. Neuron 24, 243–252 (1999).
Egan, T. M. & North, R. A. Both mu and delta opiate receptors exist on the same neuron. Science 214, 923–924 (1981).
Wang, H. B. et al. Coexpression of δ- and μ-opioid receptors in nociceptive sensory neurons. Proc. Natl. Acad. Sci. USA 107, 13117–13122 (2010).
DiCello, J. J. et al. Mu and delta opioid receptors are coexpressed and functionally interact in the enteric nervous system of the mouse colon. Cell. Mol. Gastroenterol. Hepatol. 9, 465–483 (2020).
Erbs, E. et al. A mu–delta opioid receptor brain atlas reveals neuronal co-occurrence in subcortical networks. Brain. Struct. Funct. 220, 677–702 (2015).
Erbs, E., Faget, L., Veinante, P., Kieffer, B. L., Massotte, D. In vivo neuronal co-expression of mu and delta opioid receptors uncovers new therapeutic perspectives. Receptors Clin. Investig. 1, 210 (2014).
Pierre, F. et al. Morphine-dependent and abstinent mice are characterized by a broader distribution of the neurons co-expressing mu and delta opioid receptors. Neuropharmacology 152, 30–41 (2019).
Chakrabarti, S., Liu, N. J. & Gintzler, A. R. Formation of μ-/κ-opioid receptor heterodimer is sex-dependent and mediates female-specific opioid analgesia. Proc. Natl. Acad. Sci. USA 107, 20115–20119 (2010).
Jordan, B. A. & Devi, L. A. G-protein-coupled receptor heterodimerization modulates receptor function. Nature 399, 697–700 (1999).
Berg, K. A. et al. Allosteric Interactions between δ and κ opioid receptors in peripheral sensory neurons. Mol. Pharmacol. 81, 264 (2012).
Jacobs, B. A. et al. Signaling characteristics and functional regulation of delta opioid-kappa opioid receptor (DOP-KOP) heteromers in peripheral sensory neurons. Neuropharmacology 151, 208–218 (2019).
Jacobs, B. A. et al. Allosterism within δ opioid-κ opioid receptor heteromers in peripheral sensory neurons: regulation of κ opioid agonist efficacy. Mol. Pharmacol. 93, 376–386 (2018).
Waldhoer, M. et al. A heterodimer-selective agonist shows in vivo relevance of G protein-coupled receptor dimers. Proc. Natl. Acad. Sci. USA 102, 9050 (2005).
Gomes, I. et al. G protein–coupled receptor heteromers. Annu. Rev. Pharmacol. Toxicol. 56, 403–425 (2016).
Gurevich, V. V. & Gurevich, E. V. GPCRs and signal transducers: interaction stoichiometry. Trends Pharmacol. Sci. 39, 672–684 (2018).
Ferré, S., Ciruela, F., Woods, A. S., Lluis, C. & Franco, R. Functional relevance of neurotransmitter receptor heteromers in the central nervous system. Trends Neurosci. 30, 440–446 (2007).
Bellot, M. et al. Dual agonist occupancy of AT1-R-α2C-AR heterodimers results in atypical Gs-PKA signaling. Nat. Chem. Biol. 11, 271–279 (2015).
Szafran, K. et al. Potential role of G protein-coupled receptor (GPCR) heterodimerization in neuropsychiatric disorders: a focus on depression. Pharmacol. Rep. 65, 1498–1505 (2013).
Fribourg, M. et al. Decoding the signaling of a GPCR heteromeric complex reveals a unifying mechanism of action of antipsychotic drugs. Cell 147, 1011–1023 (2011).
Di Marino, D., Conflitti, P., Motta, S. & Limongelli, V. Structural basis of dimerization of chemokine receptors CCR5 and CXCR4. Nat. Commun. 14, 6439 (2023).
Ngamlertwong, N. et al. Agonist dependency of the second phase access of β-arrestin 2 to the heteromeric µ-V1b receptor. Sci. Rep. 11, 15813 (2021).
Hübner, H. et al. Structure-guided development of heterodimer-selective GPCR ligands. Nat. Commun. 7, 12298 (2016).
Callén, L. et al. Cannabinoid receptors CB1 and CB2 form functional heteromers in brain. J. Biol. Chem. 287, 20851–20865 (2012).
Verma, V., Hasbi, A., O’Dowd, B. F. & George, S. R. Dopamine D1-D2 receptor Heteromer-mediated calcium release is desensitized by D1 receptor occupancy with or without signal activation: dual functional regulation by G protein-coupled receptor kinase 2. J. Biol. Chem. 285, 35092–35103 (2010).
White, J. H. et al. Heterodimerization is required for the formation of a functional GABAB receptor. Nature 396, 679–682 (1998).
Jones, K. A. et al. GABAB receptors function as a heteromeric assembly of the subunits GABABR1 and GABABR2. Nature 396, 674–679 (1998).
Calebiro, D. et al. Single-molecule analysis of fluorescently labeled G-protein–coupled receptors reveals complexes with distinct dynamics and organization. Proc. Natl. Acad. Sci. USA 110, 743 (2013).
Dijkman, P. M. et al. Dynamic tuneable G protein-coupled receptor monomer-dimer populations. Nat. Commun. 9, 1710 (2018).
Filizola, M. & Devi, L. A. Structural biology: how opioid drugs bind to receptors. Nature 485, 314–317 (2012).
Filizola, M., Olmea, O. & Weinstein, H. Prediction of heterodimerization interfaces of G-protein coupled receptors with a new subtractive correlated mutation method. Protein Eng. Des. Sel. 15, 881–885 (2002).
Pan, Y. X. Expression of opioid receptors in mammalian cell Lines. In: Opioid Research: Methods and Protocols (ed Pan, Z. Z.) (Humana Press, 2003).
Liu, X., Kai, M., Jin, L. & Wang, R. Computational study of the heterodimerization between μ and δ receptors. J. Comput. Aided Mol. Des. 23, 321–332 (2009).
Filizola, M. & Weinstein, H. Structural models for dimerization of G-protein coupled receptors: the opioid receptor homodimers. Pept. Sci. 66, 317–325 (2002).
Meral, D. et al. Molecular details of dimerization kinetics reveal negligible populations of transient µ-opioid receptor homodimers at physiological concentrations. Sci. Rep. 8, 7705 (2018).
Marino, K. A., Prada-Gracia, D., Provasi, D. & Filizola, M. Impact of lipid composition and receptor conformation on the spatio-temporal organization of μ-opioid receptors in a multi-component plasma membrane model. PLOS Comput. Biol. 12, e1005240 (2016).
Grasberger, B., Minton, A. P., DeLisi, C. & Metzger, H. Interaction between proteins localized in membranes. Proc. Natl. Acad. Sci. USA 83, 6258–6262 (1986).
Yekkirala, A. S., Kalyuzhny, A. E. & Portoghese, P. S. Standard opioid agonists activate heteromeric opioid receptors: evidence for morphine and [d-Ala(2)-MePhe(4)-Glyol(5)]enkephalin as selective μ-δ agonists. Acs. Chem. Neurosci. 1, 146–154 (2010).
Fagan, R. R. et al. Selective targeting of mu opioid receptors to primary cilia. Cell Rep. 43, 114164 (2024).
Décaillot, F. M., Rozenfeld, R., Gupta, A. & Devi, L. A. Cell surface targeting of mu-delta opioid receptor heterodimers by RTP4. Proc. Natl. Acad. Sci. USA 105, 16045–16050 (2008).
Bao, F. et al. Clinical opioids differentially induce co-internalization of μ- and δ-opioid receptors. Mol. Pain. 14, 1744806918769492 (2018).
Rozenfeld, R. & Devi, L. A. Receptor heterodimerization leads to a switch in signaling: β-arrestin2-mediated ERK activation by μ-δ opioid receptor heterodimers. FASEB J. 21, 2455–2465 (2007).
Milan-Lobo, L., Whistler, J. Heteromerization of the mu and delta opioid receptor produces ligand-biased antagonism and alters mu receptor trafficking. J. Pharmacol. Exp. Ther. 337, 868–875 (2011).
Grimes, J. et al. Plasma membrane preassociation drives β-arrestin coupling to receptors and activation. Cell 186, 2238–2255.e2220 (2023).
Kusumi, A., Tsunoyama, T. A., Suzuki, K. G. N., Fujiwara, T. K. & Aladag, A. Transient, nano-scale, liquid-like molecular assemblies coming of age. Curr. Opin. Cell Biol. 89, 102394 (2024).
Suzuki, K. G. N., Fujiwara, T. K., Edidin, M. & Kusumi, A. Dynamic recruitment of phospholipase Cγ at transiently immobilized GPI-anchored receptor clusters induces IP3–Ca2+ signaling: single-molecule tracking study 2. J. Cell Biol. 177, 731–742 (2007).
Suzuki, K. G. N. et al. GPI-anchored receptor clusters transiently recruit Lyn and Gα for temporary cluster immobilization and Lyn activation: single-molecule tracking study 1. J. Cell Biol. 177, 717–730 (2007).
Metcalf, M. D. et al. The δ opioid receptor agonist SNC80 selectively activates heteromeric μ-δ opioid receptors. Acs. Chem. Neurosci. 3, 505–509 (2012).
Dratz, E. A. et al. NMR structure of a receptor-bound G-protein peptide. Nature 363, 276–281 (1993).
Gleixner, J. et al. Illuminating neuropeptide Y Y4 receptor binding: fluorescent cyclic peptides with subnanomolar binding affinity as novel molecular tools. ACS Pharmacol. Transl. Sci. 7, 1142–1168 (2024).
Yokoyama, T., Kato, N. & Yamada, N. Development of a high-throughput bioassay to screen melatonin receptor agonists using human melatonin receptor expressing CHO cells. Neurosci. Lett. 344, 45–48 (2003).
Han, Y., Moreira, I. S., Urizar, E., Weinstein, H. & Javitch, J. A. Allosteric communication between protomers of dopamine class A GPCR dimers modulates activation. Nat. Chem. Biol. 5, 688–695 (2009).
Conklin, B. R., Farfel, Z., Lustig, K. D., Julius, D. & Bourne, H. R. Substitution of three amino acids switches receptor specificity of Gqα to that of Giα. Nature 363, 274–276 (1993).
Yokoyama, T. et al. A multicolor suite for deciphering population coding of calcium and cAMP in vivo. Nat. Methods 21, 897–907 (2024).
Kabli, N., Nguyen, T., Balboni, G., O’Dowd, B. F. & George, S. R. Antidepressant-like and anxiolytic-like effects following activation of the μ-δ opioid receptor heteromer in the nucleus accumbens. Mol. Psychiatry 19, 986–994 (2014).
Fujita, W., Gomes, I. & Devi, L. A. Revolution in GPCR signalling: opioid receptor heteromers as novel therapeutic targets: IUPHAR review 10. Br. J. Pharmacol. 171, 4155–4176 (2014).
Keresztes, A. et al. Antagonism of the mu-delta opioid receptor heterodimer enhances opioid antinociception by activating Src and calcium/calmodulin-dependent protein kinase II signaling. Pain 163, 146–158 (2022).
Möller, J. et al. Single-molecule analysis reveals agonist-specific dimer formation of µ-opioid receptors. Nat. Chem. Biol. 16, 946–954 (2020).
Asher, W. B. et al. Single-molecule FRET imaging of GPCR dimers in living cells. Nat. Methods 18, 397–405 (2021).
Cechova, K. et al. Kappa but not delta or mu opioid receptors form homodimers at low membrane densities. Cell. Mol. Life Sci. 78, 7557–7568 (2021).
Drakopoulos, A. et al. Investigation of inactive-state κ opioid receptor homodimerization via single-molecule microscopy using new antagonistic fluorescent probes. J. Med. Chem. 63, 3596–3609 (2020).
Baiula, M. Monitoring opioid receptor interaction in living cells by bioluminescence resonance energy transfer (BRET). Methods Mol. Biol. 2201, 35–43 (2021).
Cvejic, S. & Devi, L. A. Dimerization of the delta opioid receptor: implication for a role in receptor internalization. J. Biol. Chem. 272, 26959–26964 (1997).
Gomes, I., Filipovska, J., Jordan, B. A. & Devi, L. A. Oligomerization of opioid receptors. Methods 27, 358–365 (2002).
He, L., Fong, J., von Zastrow, M. & Whistler, J. L. Regulation of opioid receptor trafficking and morphine tolerance by receptor oligomerization. Cell 108, 271–282 (2002).
Kasai, R. S., Fujiwara, T. K., Kusumi, A. Metastable GPCR dimers trigger the basal signal by recruiting G-proteins. bioRxiv, 2020.2002.2010.929588 (2020).
Johnston, J. M. et al. Making structural sense of dimerization interfaces of delta opioid receptor homodimers. Biochemistry 50, 1682–1690 (2011).
Manglik, A. et al. Crystal structure of the µ-opioid receptor bound to a morphinan antagonist. Nature 485, 321–326 (2012).
De Oliveira, P. A. et al. Preferential Gs protein coupling of the galanin Gal(1) receptor in the µ-opioid-Gal(1) receptor heterotetramer. Pharmacol. Res. 182, 106322 (2022).
Kaneko, S. et al. Structural and dynamic insights into the activation of the μ-opioid receptor by an allosteric modulator. Nat. Commun. 15, 3544 (2024).
Mahoney, J. P. & Sunahara, R. K. Mechanistic insights into GPCR–G protein interactions. Curr. Opin. Struct. Biol. 41, 247–254 (2016).
Whorton, M. R. et al. A monomeric G protein-coupled receptor isolated in a high-density lipoprotein particle efficiently activates its G protein. Proc. Natl. Acad. Sci. USA 104, 7682–7687 (2007).
Nitsche, J. F. et al. Genetic dissociation of opiate tolerance and physical dependence in delta-opioid receptor-1 and preproenkephalin knock-out mice. J. Neurosci. 22, 10906–10913 (2002).
Xie, W. Y. et al. Disruption of Cdk5-associated phosphorylation of residue threonine-161 of the δ-opioid receptor: impaired receptor function and attenuated morphine antinociceptive tolerance. J. Neurosci. 29, 3551 (2009).
Gaborit, M. & Massotte, D. Therapeutic potential of opioid receptor heteromers in chronic pain and associated comorbidities. Br. J. Pharmacol. 180, 994–1013 (2023).
Matsoukas, M. T. et al. Design of small non-peptidic ligands that alter heteromerization between cannabinoid CB(1) and serotonin 5HT(2A) receptors. J. Med. Chem. 68, 261–269 (2025).
Gallo, M. et al. Orally active peptide vector allows using cannabis to fight pain while avoiding side effects. J. Med. Chem. 64, 6937–6948 (2021).
Schulz, R., Wehmeyer, A. & Schulz, K. Opioid receptor types selectively cointernalize with G protein-coupled receptor kinases 2 and 3. J. Pharmacol. Exp. Ther. 300, 376 (2002).
Fukuda, K., Kato, S., Mori, K., Nishi, M. & Takeshima, H. Primary structures and expression from cDNAs of rat opioid receptor δ-and μ-subtypes. FEBS Lett. 327, 311–314 (1993).
Shaner, N. C. et al. Improved monomeric red, orange and yellow fluorescent proteins derived from Discosoma sp. red fluorescent protein. Nat. Biotechnol. 22, 1567–1572 (2004).
Lindberg, F. P., Gresham, H. D., Schwarz, E. & Brown, E. J. Molecular cloning of integrin-associated protein: an immunoglobulin family member with multiple membrane-spanning domains implicated in alpha v beta 3-dependent ligand binding. J. Cell. Biol. 123, 485–496 (1993).
James, J. R., Oliveira, M. I., Carmo, A. M., Iaboni, A. & Davis, S. J. A rigorous experimental framework for detecting protein oligomerization using bioluminescence resonance energy transfer. Nat. Methods 3, 1001–1006 (2006).
Yamashiro, S. et al. Substrate specificity of beta 1,4-N-acetylgalactosaminyltransferase in vitro and in cDNA-transfected cells. GM2/GD2 synthase efficiently generates asialo-GM2 in certain cells. J. Biol. Chem. 270, 6149–6155 (1995).
Ando, R. et al. StayGold variants for molecular fusion and membrane-targeting applications. Nat. Methods 21, 648–656 (2024).
Koyama-Honda, I. et al. Fluorescence imaging for monitoring the colocalization of two single molecules in living cells. Biophys. J. 88, 2126–2136 (2005).
Murakoshi, H. et al. Single-molecule imaging analysis of Ras activation in living cells. Proc. Natl. Acad. Sci. USA 101, 7317 (2004).
Fujiwara, T. K. et al. Confined diffusion of transmembrane proteins and lipids induced by the same actin meshwork lining the plasma membrane. Mol. Biol. Cell. 27, 1101–1119 (2016).
Kasai, R. S. et al. Full characterization of GPCR monomer-dimer dynamic equilibrium by single molecule imaging. J. Cell. Biol. 192, 463–480 (2011).
Stone, M. B. & Veatch, S. L. Steady-state cross-correlations for live two-colour super-resolution localization data sets. Nat. Commun. 6, 7347 (2015).
Keppler, A., Pick, H., Arrivoli, C., Vogel, H. & Johnsson, K. Labeling of fusion proteins with synthetic fluorophores in live cells. Proc. Natl. Acad. Sci. USA. 101, 9955–9959 (2004).
Hagg, T. Intracerebral infusion of neurotrophic factors. In: Neuroprotection Methods and Protocols (ed Borsello, T.) (Humana Press, 2007).
Zhou, P. et al. Single-molecule methods for characterizing receptor dimers reveal metastable opioid receptor homodimers that induce functional modulation, Zenodo, https://doi.org/10.5281/zenodo.17160485 (2025).
Acknowledgements
We thank Profs. H. Takeshima of Kyoto University, R. Schülz of the University of Münich, and R. Y. Tsien of the University of California San Diego, for their kind gifts of cDNAs encoding rat KOR and DOR, that encoding rat MOR, and that encoding mCherry, respectively. We also thank Ms. Irina Meshcheryakova of OIST for technical help in preparing the cDNAs and Dr. Michael C. Roy of OIST for technical help in testing the peptide stability. We are particularly grateful for the encouragement of Prof. Yuki Sugiura of the Center for Cancer Immunotherapy and Immunobiology, Kyoto University Graduate School of Medicine, for this work. This work was supported in part by Japan Society for the Promotion of Science (JSPS) Grants-in-Aid for Scientific Research (Kiban A to A.K. [21H04772], Kiban S to A.K. [16H06386], Kiban B to T.K.F. [16H04775, 20H02585], H.U. [21H03024], and M.S. [JP23H02782], Kiban C to R.S.K. [17K07333] and W.F. [21K07275, 24K10036], Wakate B to R.S.K. [26870292], Wakate to T.A.T. [21K15058] and T.Y. [24K18240]) and Challenging Exploratory Research to T.K.F. [18K19001] and A.K. [22K19334]), a Grant-in-Aid from the Ministry of Education, Culture, Sports, Science and Technology of Japan (MEXT) for Transformative Research Areas (A) to T.K.F. (21H05252), Transformative Research Areas (B) to M.S. (JP24H00861), a JST grant ACT-X to T.A.T. (JPMJAX211B) and T.Y. (JPMJAX211K), an AMED grant Brain/MIND 2.0 to M.S. (JP24wm0625119), NSTC grant (R.O.C., Taiwan) to H.U. (112-2320-B-016-003), and OIST Proof of Concept Program Technology Pioneer Fellowship to P.Z. (R11_61). WPI-iCeMS of Kyoto University is supported by the World Premiere Research Center Initiative (WPI) of MEXT.
Author information
Authors and Affiliations
Contributions
P.Z., R.S.K., and A.K. conceived and formulated the project. A.K., T.A.T., T.K.F., and P.Z. developed the simultaneous dual-color single-molecule tracking station. P.Z. and A.K. designed biological experiments, and P.Z., with help from T.A.T., performed virtually all of the microscopic experiments. T.K.F., T.A.T., and A.K. developed and improved the in-house image analysis software and instrument control software, and discussed single-molecule imaging data. S.P. developed the theory for evaluating the dimer lifetimes from the single-molecule experimental data and assisted with its application to experimental results. T.A.T. and P.Z. analyzed the PCCF histograms and obtained dimer dissociation constants. W.F. and H.N., with help from H.U., performed mouse experiments. P.Z., with help from M.S., T.Y., and T.A.T. conducted a cAMPinG1-based cAMP assay. P.Z., A.K., and T.A.T. wrote the manuscript, and all authors discussed the results and participated in the manuscript revisions.
Corresponding authors
Ethics declarations
Competing interests
P.Z. and A.K. are inventors on a patent application (patent pending) for DpepDMs, MpepMDs and peptides from EL3 domains of ORs described in this study. The remaining authors declare no competing interest.
Peer review
Peer review information
Nature Communications thanks Tijana Jovanović-Talisman who co-reviewed with Carina Lima; Vittorio Limongelli who co-reviewed with Stefano Raniolo; Sergi Ferre and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhou, P., Kasai, R.S., Fujita, W. et al. Single-molecule characterization of opioid receptor heterodimers reveals soluble µ-δ dimer blocker peptide alleviates morphine tolerance. Nat Commun 16, 9859 (2025). https://doi.org/10.1038/s41467-025-64695-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-64695-2