Abstract
Rhabdomyosarcoma (RMS) is marked by a myogenesis differentiation blockade, and while the AKT/mTOR pathway is universally activated, its pharmacological inhibition has shown limited success. Here, we evaluate the activity of pan-AKT inhibitors Ipatasertib, ATP-competitive, and Miransertib, allosteric inhibitor, in RMS cell lines and fusion-positive/negative patient-derived xenografts (PDX). Unlike Miransertib, Ipatasertib show significant antitumor activity against a subset of RMS. Besides AKT, the other target of Ipatasertib, but not of Miransertib, is PRKG1, a cGMP-dependent protein kinase that shares the ATP binding pocket with AKT. We investigate the role of PRKG1 in PRKG1-depleted RMS cells and in xenograft models by transcriptomic approaches. PRKG1 silencing in RMS cells reduces tumor formation in xenograft models and induces a differentiated myogenic transcriptome. RMS show higher PRKG1 expression compared to any other developmental cancer, akin to fetal skeletal muscle. Importantly, PRKG1 expression in RMS correlates with mesodermal transcriptional signature and enhanced sensitivity to Ipatasertib, regardless of the fusion oncogene status. The antitumor activity of Ipatasertib is dose-dependent, reaching an effective intra-tumor concentration when administered at 25 mg/kg daily. This study unveils the role of PRKG1 in myogenesis and highlights the potential of PRKG1 as a clinical biomarker for Ipatasertib therapy in RMS.
Similar content being viewed by others
Introduction
Pediatric Rhabdomyosarcoma (RMS) is the most common pediatric soft tissue sarcoma1, a developmental tumor that shares features with arrested skeletal muscle precursor cells2,3. Despite the expression of myogenic-committed transcription factors like MYOD1 and MYOG4, RMS cells fail to fully progress through the myogenic differentiation program. Historically, RMS has been divided into two main histological subtypes, alveolar and embryonal. The presence of the PAX3/7-FOXO1 fusion translocation defines the fusion-positive (FP) RMS subtype, in contrast to fusion-negative (FN) RMS5. This molecular classification as FP or FN is a powerful prognostic indicator6,7. RMS patients with high-risk features including PAX3/7-FOXO1 translocations, age < 1 year or older than 10 years of age, primary tumor at unfavorable locations, failing to achieve local tumor control and/or progressing to metastatic disease or at relapse have a dismal prognosis with survival rates below 20%8,9,10. For the last 20 years all different chemotherapy agents in trials have failed to improve the outcome of RMS patients where 25–30% of patients continue to experience disease recurrence with dismal post-relapse survival11. Rescue therapy for relapsed RMS includes regimens associating mTOR complex inhibitors like everolimus or temsirolimus12,13. Temsirolimus was evaluated in a recently completed phase III trial for intermediate-risk FN-RMS14.
Recently, the developmental cell hierarchy has gained considerable interest as an alternative exploiting therapeutic vulnerability3,15,16,17. Multi-omic studies have described three main cellular states present in RMS samples transitioning between early mesenchymal progenitors, myoblast and terminally differentiated myocyte cells3,15,16,17. This developmental diversity, both intra- and inter-tumoral, could be exploited for therapeutic purposes. Indeed, recent studies have demonstrated that terminal myogenic differentiation can be triggered in RAS-mutant FN-RMS cells by interfering with RAS signaling18,19 and BAF complexes in FP-RMS20. Other studies also demonstrate that MEK or ERK inhibitors selectively inhibit the viability of RAS-mutant FN-RMS cells, cause myogenic differentiation and tumor growth delay, although single-target approaches did not result in durable responses18. Combining RAS/MEK/ERK and PI3K/AKT/mTOR inhibition offered an approach to overcome therapeutic resistance21 but proved toxic in clinical trials, highlighting the challenge of targeting these pathways22.
High levels of AKT phosphorylation in RMS were shown to be associated with poor overall and disease-free survival23. Mutations in the PI3K catalytic subunit are frequently associated with MYOD1 mutations24,25. FGFR4 mutations in FN-RMS and FGFR4 hyperactivation in FP-RMS24,26 and loss of PTEN in FP and FN-RMS24 are recurrently found in RMS to activate the PI3K/AKT/mTOR pathway, which is negatively associated with patient survival23. A substantial amount of literature can be found on small molecule AKT inhibitors tested in RMS cell lines27,28. The experience of AKT inhibitors in more clinically relevant models of RMS was reported by Manzella et al., where a subgroup of RMS was found particularly sensitive to AKT inhibitors, including both FP- and FN-RMS29. However, in the clinic, it is yet unknown how to identify those AKT inhibitor sensitive RMS cases and the mechanism behind this effect.
AKT is a member of the AGC kinase [protein kinase A, G, and C families (PKA, PKC, PKG)] family, which is a threonine/serine protein kinase, widely known as protein kinase B (PKB). AKT lies in the core of the PI3K/AKT/mTOR pathway, a signaling pathway frequently activated in cancer30. The development of small molecule inhibitors targeting AKT has recently gained attraction, with several compounds currently in clinical trials. These inhibitors are classified into two main groups according to the mechanism of action: ATP-competitive and allosteric inhibitors31. ATP-competitive inhibitors bind to the ATP pocket of the AKT kinase domain, while allosteric inhibitors target a different pocket located in the AKT pleckstrin-homology domain. Among the ATP-competitives, ipatasertib (GDC-0068) has been tested broadly in clinical trials32,33,34,35. Ipatasertib is a pan-AKT inhibitor, blocking its enzymatic functions and downstream signaling. Among the allosteric inhibitors, miransertib (ARQ-092) is a highly selective inhibitor of all three AKT isoforms, binding to the unphosphorylated form of AKT36. Miransertib has shown promising antitumor activity, particularly in tumors harboring mutations in the PI3K/AKT signaling pathway. Miransertib is currently in clinical development for patients with PROTEUS syndrome and PI3K-related overgrowth spectrum of diseases37.
In this work we explore different AKT inhibitors in search of improved targeted therapy strategies for patients with RMS. We use a combination of in vitro and in vivo studies to characterize the differential anti-RMS activity of ipatasertib using a collection of cell lines and primary cultures from patient-derived RMS xenografts (PDXs). We uncover the role of PRKG1 in the myogenesis blockade characteristic of RMS that explains, in part, the singular antitumor activity of ipatasertib against a subgroup of RMS, regardless of the fusion oncogene. Pharmacokinetics (PK) and pharmacodynamics (PD) of ipatasertib in RMS PDX models show effective intratumor levels achieved with clinically feasible doses and adequate animal tolerance. Furthermore, we describe the expression levels of PRKG1 in the primary tumors and PDX models correlating to ipatasertib response and suggesting that PRKG1 could be used as a clinical biomarker to predict efficacy in the clinical setting.
Results
Treatment of a pediatric RMS patient with the AKT inhibitor ipatasertib
A previously healthy 4-month-old female presented with a right cervical soft-tissue mass that invaded the spinal canal. Histological analysis and next-generation sequencing of an intra-spinal biopsy demonstrated embryonal, MYCN amplified, FN-RMS. The tumor was classified as IRSG III (para-meningeal), TNM stage 3 (T1N1M0). The patient received treatment including chemotherapy, gross total surgical resection, and proton beam radiotherapy achieving first complete remission. Three months later, craniospinal magnetic resonance imaging (MRI) demonstrated a tumor mass in the left parieto-occipital cortex and two leptomeningeal enhancing nodules in the inter-hemispheric line, suggestive of leptomeningeal (LM) metastasis. The rest of the diagnostic workup was negative, confirming an isolated central nervous system (CNS) relapse. Patient presentation, clinical course, and pathological evaluation are summarized in Fig. 1A.
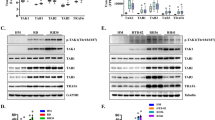
A Clinical course from initial presentation, relapse disease, and overview of sample collection. T: time in months. Created with BioRender.com B Phosphorylation and total proteins in the AKT and MEK pathways and c-PARP in E001_s cells treated with ipatasertib (AKT inhibitor) and trametinib (MEK inhibitor) for 24 h (representative of three independent experiments). Tubulin was used as loading control. C Effect of ipatasertib and trametinib in the tumoral growth of a primary RMS model in vivo. Tumor growth of E001_s subcutaneous xenografts in mice treated with 100 mg/kg per day of ipatasertib (green), 3 mg/kg per day of trametinib (red), the combination of the two drugs (purple) or vehicle control (dark gray). E001_s mice received drug administration oral gavage, daily for 15 days (with 2 days off each week). 5 animals per group with one subcutaneous tumor each. Source Data are available. D Summary of molecular, biological and clinical features of the RMS PDX used in this study. E Hematoxylin/eosin and immunohistochemistry (IHC) of rhabdomyosarcoma tumoral markers (MYOD1, MYOG and Desmin), pS6 and human nuclei in HSJD-RMS PDX samples included in this article. Black bar indicates the scale bar of 100 µm. Observe nuclear positiveness for MYOD1 and anti-human nuclei in all RMS PDX. Images representative of four independent tumor replicates.
Prior to radical resection of the relapsed tumor, the family consented to donate tissue to the HSJD tumor bank approved by Institutional Review Board, enabling the establishment of tumor spheres in vitro, HSJD-E001_s. Postoperatively, the patient completed 41.4 Gy of consolidative radiotherapy and began salvage chemotherapy with irinotecan, temozolamide and temsirolimus. Patient did well for a year, asymptomatic, with no evidence of disease.
Meanwhile, HSJD-E001_s tumor spheres were cultured and tested in vitro for their sensitivity to the AKT inhibitor ipatasertib and the MEK inhibitor trametinib. Cells were treated for 24 h with the inhibitors alone or in combination, and the AKT/mTOR and MAPK pathways analyzed by western blot (WB). Consistent with the mechanism of action, ipatasertib increased pAKT levels, while S6 phosphorylation (pS6, used here as a functional readout of AKT/mTOR pathway activity) was reduced, confirming the efficacy of AKT inhibition. In turn, trametinib decreased pERK levels, while pS6 levels remained unchanged, as expected (Fig. 1B). Analysis of cleaved PARP, a hallmark of the apoptotic effect, showed that only ipatasertib effectively induced cell death in this model, and that combination with trametinib did not further potentiate this effect (Fig. 1B).
To further investigate the effect of these inhibitors, HSJD-E001_s cells were subcutaneously implanted into immune suppressed mice. When tumors reached a volume of 100–350 mm3, mice were treated according to 4 different strategies: vehicle; 3 mg/kg of trametinib; 100 mg/kg of ipatasertib; and the combination administered 5 days per week at the maximal tolerated dose (MTD). Mice carrying E001_s xenografts received treatment for 2 weeks. As shown in Fig. 1C, trametinib did not affect tumor growth, whereas ipatasertib showed a significant anti-tumor effect when administrated alone or in combination and even achieved tumor eradication in some animals.
By the time all these in vivo experiments were completed, the patient’s MRI showed new LM lesions as well as new (and large) right posterior temporal and (small) left temporal lobe masses (Fig. 1A). A new surgical gross total resection was performed, and pathology confirmed MYCN amplified FN-RMS. The HSJD-ERMS-024 PDX model was established as an outgrowth from the second CNS tumor sample. Postoperatively, given the preclinical model studies showing activity of ipatasertib against the ERMS-E001 model, the patient began salvage therapy with compassionate use of 30 mg twice daily of ipatasertib. Rapid progression of the LM disease occurred, and patient died one month later.
To understand the failure to translate the preclinical results suggesting ipatasertib as an active drug, the lack of response observed in this patient and how to better personalize targeted therapy for patients with RMS, we conceived this study. To capture patient heterogeneity, we established new cell lines from PDXs and thereby increased the spectrum of our RMS experimental models. Figure 1D and E illustrate the molecular and histological profiles and clinical features of six primary RMS tumors and their corresponding PDXs (including ERMS-024) used in this study, with two FN and four FP cases. We generated cell lines from those PDXs and together with the commercial RH4 and RD RMS cell lines were used for in vitro testing. Unfortunately, the culture of E001_s cells could not be perpetuated, since it stopped proliferating after several passages.
The AKT inhibitors ipatasertib and miransertib show differential anti-tumor activity against RMS
Based on the observed sensitivity of E001_s to AKT inhibition, we wanted to test whether other RMS experimental models could also benefit from AKT inhibition by using the ATP-competitive inhibitor ipatasertib and the allosteric inhibitor miransertib. To this end, xenografted mice bearing the ERMS-011 model were treated by oral gavage with ipatasertib or miransertib daily at 100 mg/kg. The ipatasertib posology was five days on, followed by two days off, per week. All tumors of miransertib-treated mice grew similar to controls (Fig. 2A). In contrast, ipatasertib-treated animals showed significant tumor growth reduction at both 25 mg/kg and 100 mg/kg doses, as was the case with E001_s. Indeed, tumor volumes in the ipatasertib group reached undetectable levels at 100 mg/kg doses. As shown in Supplementary Fig. 1A, both the miransertib and ipatasertib treated animals showed stable body weight over treatment courses. To strengthen the consistency of our comparative analyses, we extended the in vivo efficacy studies to the ARMS-010 model, which confirmed the differential responses to each inhibitor (Fig. 2B).
A Antitumoral effectiveness of miransertib and ipatasertib in vivo. Plots of individual tumor size for ERMS-011 PDX in mice under ipatasertib or miransertib treatments. Highlighted in gray indicates treatment period. Source Data is available. B Antitumoral effectiveness of miransertib and ipatasertib in vivo. Tumor growth of ARMS-010 PDX in mice under ipatasertib or miransertib treatments. Mean ± sem. N = 5 miransertib, n = 5 vehicle and n = 7 mice in 100 mg/kg ipatasertib group. Source Data are available. C IHC analysis of pS6, cleaved PARP, and Ki67 in ERMS-011 (left) and ARMS-010 PDX (right) after treatment with ipatasertib or miransertib. Samples collected after 5 doses for ERMS-011 and after 10 doses for ARMS-010. Images representative of three different tumors/mice per condition. D Comparison of miransertib (purple) and ipatasertib (blue) effects in a panel of RMS primary cultures and cell lines. Cells were seeded and treated under identical conditions. Cell viability at 72 h is shown as % viable cells (mean ± SEM). IC50 values were determined using a four-parameter model with variable slope, constraining the bottom and top values to 0 and 100, respectively. ARMS-010 (n = 5), RH4 (n = 4), ARMS-006 (n = 2), ERMS-024 one representative experiment of n = 2. Source data are provided as a Source Data file. E RH4 cells were treated with 1 (+) or 10 ( + + ) μM concentrations of ipatasertib or miransertib for 24 h, pAKT (S473) and pS6 (S240/244) and their corresponding total protein levels were assessed by western blot analysis. ERMS-024, ARMS-010 and ARMS-006 were treated for 24 h with 0.5 µM of ipatasertib and miransertib, respectively. Results were reproduced in three independent experiments. F Significantly downregulated and enriched hallmark terms in RH4 cells treated for 24 h with 1 µM ipatasertib or miransertib. Enrichment was performed using pre-ranked GSEA based on limma-derived statistics (–log(p-value) × signFC). Dot size reflects the number of core enriched genes (also known as leading-edge genes); color indicates FDR-adjusted p-values. Source data provided in Supplementary Data file. G Representation of genes from the hallmark myogenesis in ipatasertib versus control. Visualization includes genes contributing to pathway-level enrichment (core enriched genes), as identified by pre-ranked GSEA using limma-derived statistics (–log(p-value) × signFC). Node color reflects limma-derived statistics. Source data provided in Supplementary Data file.
To rule out the possibility that the differential response was due to incomplete AKT inhibition by miransertib and for better characterization of the effect of ipatasertib in vivo, we performed immunohistochemical (IHC) analyses to evaluate markers of the AKT pathway, proliferation and apoptosis (Fig. 2C). pS6 levels were reduced in tumors treated with either ipatasertib or miransertib, confirming that both inhibitors effectively engage the AKT pathway in vivo. In addition, Ki67 staining was reduced in tumors treated with both inhibitors, with a more pronounced decrease in ipatasertib-treated tumors, while cleaved PARP (cPARP) levels were increased in ipatasertib-treated tumors compared to those treated with miransertib, suggesting enhanced induction of apoptosis. These findings demonstrated that, although both inhibitors reduce the AKT pathway activity in vivo, ipatasertib exerts a greater impact on tumor proliferation and apoptosis compared to miransertib.
In vitro experiments confirmed a discordant cytotoxic activity between the two AKT inhibitors (Fig. 2D). In ERMS-024 PDX-derived cells, miransertib IC50 was lower than ipatasertib (0.24 μM vs. 0.62 μM). However, in both ARMS-010 and ARMS-006 PDX-derived cells and the RH4 cell line, miransertib showed lower cytotoxicity than ipatasertib (IC50 of miransertib vs. ipatasertib in ARMS-010, 0.29 µM vs. IC50 0.098 µM; in ARMS-006, 0.43 µM vs. IC50 0.12 µM; in RH4, 0.84 μM vs. 0.45 μM). Taken together, AKT inhibitor ipatasertib, but not miransertib, displayed tumor inhibitory capacities in RMS.
To understand the differential cytotoxic effect of miransertib and ipatasertib we analyzed the status of the AKT/mTOR pathway in each of the models analyzed. Cells were treated for 24 h with either ipatasertib or miransertib, and the AKT/mTOR pathway analyzed by western blot. As observed in E001_s, and in accordance with their respective mechanisms of action, ipatasertib increased pAKT levels, while miransertib decreased them. An effective inhibition of AKT/mTOR was observed with both treatments, as pS6 was reduced while total S6 levels remained unchanged (Fig. 2E). In the RH4 RMS cells, higher doses of ipatasertib and miransertib did not further reduce S6 phosphorylation levels, suggesting that residual pS6 is AKT-independent (Fig. 2E).
To compare the transcriptomic effects of each AKT inhibitor in RMS, a whole genome expression profiling was performed from ipatasertib and miransertib treated RH4 cells with 1 µM for 24 h of each AKT inhibitor. Supervised gene expression analysis between treated and untreated cells, resulted in 1829 differentially expressed genes (DEGs) in cells treated with ipatasertib for 24 h, compared to 799 DEGs with miransertib (Supplementary Data 1). At the transcriptomic level, consistent with their described mechanism of action, the most significantly depleted genes were those from the mTORC1 signaling pathway, confirming its downregulation (Fig. 2F). Since both ipatasertib and miransertib equally inhibited the AKT/mTOR signaling pathway, the observed anti-tumor effect of ipatasertib could not be related to AKT inhibition.
Interestingly, among the subgroup of genes upregulated after treatment, the hallmark “myogenesis” was significant for ipatasertib but not in miransertib treated cells (Fig. 2F, G). Genes such as TNNT1, MYL4, MYL3 and MYH11 were found within this hallmark (Supplementary Data 1).
In an attempt to identify differentially phosphorylated targets, we performed a discovery-based phosphoproteomic profiling of cells treated for 1 h with each inhibitor. Volcano plots showing changes in phosphoprotein levels revealed that, consistent with its differential inhibitory mechanism, treatment with ipatasertib resulted in a greater number of downregulated phosphopeptides than with miransertib (Supplementary Data 1 and Supplementary Fig. 1).
Therefore, we hypothesized that, besides AKT, other kinases relevant for RMS might be concomitantly inhibited by the ATP-competitive inhibitor. Recently, the kinome profile of ipatasertib had been reported38. In the top proteins inhibited with high specificity, PRKG1 was found second to AKT (Supplementary Fig. 1C). Nanomolar concentrations of ipatasertib inhibited the cGMP-dependent protein kinase PRKG1 which belongs to the AGC kinase family and thus share the ATP binding pocket of AKT. Thus, we hypothesized that PRKG1 inhibition could explain the anti-tumor effects of ipatasertib in RMS.
Ipatasertib exerts its physiological effects via dual inhibition of AKT and PRKG1, whereas miransertib operates mainly through AKT
AKT has been crystallized bound to ipatasertib (PDB ID 4ekl) showing key hydrogen bonds in the hinge region (Glu228 and Ala230) and salt bridges at the pocket entrance (Glu234 and Glu278) (Fig. 3A). Unrestrained self-docking of ipatasertib to this AKT holo conformation reproduces the experimental binding pose (heavy atom RMSD = 0.73Å) with appreciable docking score (−11.38).
A Binding pose of ipatasertib on AKT binding site extracted from PDB ID 4ekl. B Superposition of ATP binding pocket in AKT (gray) and PRKG1 (blue) showing a different 3D conformation for the equivalent glutamic acid residue (278 in Akt, 488 in PRKG1). C Binding pose of ipatasertib docked to PRKG1 after induced fit (pink). Original PRKG1 conformation in blue. D 3D structures of AKT (PDB ID 5kcv, gray and orange) and PRKG1-Iβ (7lv3, blue). E Western blot showing PRKG1 and AKT levels in whole cell extracts upon PRKG1 knockdown (shPRKG1#1 and shPRKG1#2) and AKT knockdown (shAKT#1 and shAKT#2) in RH4 cells. Tubulin as loading control. Blots are representative of two independent biological replicates. F Cell viability curves at 72 h in shControl (shCtrl#LKO, purple), shAKT (shAKT#1, orange), and shPRKG1 (shPRKG1#2, blue) upon treatment with ipatasertib or miransertib, respectively. Mean ± SEM from independent biological replicates (n = 2 for ipatasertib, n = 3 for miransertib). Dose-response curves fitted using a four-parameter logistic model with variable slope. Source Data is available. G Western blot analysis of PRKG1 levels in protein extracts from SCR (control) and KO_PRKG1 cells in RH4. Tubulin was used as loading control. Blot is representative of four independent biological replicates. H Cell viability curves at 72 h for AKT inhibitors, showing the percentage of viable cells in control (SCR) versus PRKG1_KO cells treated with ATP-competitive inhibitors (ipatasertib and GSK690693) and allosteric inhibitors (MK-2206 and miransertib). Data represent mean ± SEM; n = 3 independent biological replicates for ipatasertib, miransertib, and MK-2206, and n = 2 for GSK. Dose-response curves were fitted using a four-parameter logistic model to calculate IC50 values (see table on the right). Source Data is available.
The ATP-binding site of PRKG1 shows a high degree of sequence and structure similarity with that of AKT. To query if ipatasertib could also bind to PRKG1 we retrieved the protein conformation from the holo structure deposited with PDB ID 7lv3. Since one of the key glutamic acid residues at the entrance of the pocket (Glu488) adopts an outwards-facing conformation, differing from the inward orientation observed for Glu278 in the AKT-ipatasertib complex (Fig. 3B), we performed induced fit docking to allow a reorganization of this side chain. The resulting docked pose of ipatasertib in PRKG1 achieved an equivalent docking score (−11.40) to that of AKT, with Glu488 sidechain rearranging to make a salt-bridge interaction, therefore preserving the binding mode (Fig. 3C).
To further validate the proposed binding of ipatasertib, we perform molecular dynamics (MD) simulations for PRKG1 and AKT, starting from the docking and co-crystallized poses, respectively. Both simulations preserved the initial binding mode during the whole simulation time (0.5 microseconds), adding evidence that the proposed pose on PRKG1 is highly plausible. Canonical hinge hydrogen bonds with Glu228/Ala230 (AKT) and Glu439/Cys441 (PRKG1) were preserved for over 91% of the time. Additionally, PRKG1 formed a specific hydrogen bond between the hydroxyl group of ipatasertib and the thiol of Cys441, maintained for 56% of the simulation. On the other end of the pocket, the salt bridges with the glutamic acids prevail during different amounts of time, with Glu234 (AKT) and Glu445 (PRKG1) reaching the more stable interactions (87.5% and 98.7% of the simulation time, respectively). From these MD simulations, we estimated the binding affinity of the ligand through MM-GBSA. Similar free energies of binding were obtained for both targets, even slightly stronger for PRKG1: −40.0 + /− 0.3 kcal/mol for AKT and −46.9 + /− 0.3 kcal/mol for PRKG1. Taken together, these results provide a robust prediction that ipatasertib can bind PRKG1 with similar pose and affinity to AKT.
Finally, we evaluated whether miransertib, an allosteric AKT inhibitor, could also bind to PRKG1. Noteworthy, the allosteric site where miransertib is bound (PDB ID 5kcv) is formed by the PH domain at the N-terminal of AKT, which is not present in PRKG1. Figure 3D shows that the allosteric pocket in the inhibited form of AKT (PDB ID 5kcv) is not formed in the inhibited form reported for PRKG1 (PDB ID 7lv3). Docking calculations of miransertib against PRKG1 reveal scores too low to further try to characterize a plausible binding complex. Lastly, Yu et al36. reported that there is no appreciable inhibition of PRKG1 by this compound.
After verifying that ipatasertib could inhibit both AKT and PRKG1, we investigated the impact of their knockdown in RMS. Western blots confirmed efficient PRKG1 and AKT silencing by two independent lentiviral short hairpin RNAs (shRNAs) in RH4 cells (Fig. 3E), and downregulation of both kinases reduced cell proliferation (Supplementary Fig. 2A and 2B).
To determine whether AKT and PRKG1 expression conferred sensitivity to ipatasertib, IC50 values following gene silencing were calculated. As shown in Fig. 3F, both AKT and PRKG1-knockdown (KD) cells were less sensitive to the inhibitor. In contrast, PRKG1-depleted cells showed the same sensitivity to miransertib as control cells, whereas the IC50 of AKT-depleted cells increased, as expected (Fig. 3F).
To further investigate the involvement of PRKG1 in the anti-tumor effect of ipatasertib, PRKG1 was deleted using CRISPR-Cas9 technology (Fig. 3G). As observed under PRKG1-depleted conditions, PRKG1-knockout (KO) cells exhibited an over tenfold increase in the IC50 for ipatasertib compared to control cells (Fig. 3H). Furthermore, Carboxyfluorescein Succinimidyl Ester (CFSE) staining confirmed that the proliferation of PRKG1-KO cells was not affected by ipatasertib (Supplementary Fig. 2C). Importantly, the IC50 value for GSK690693 (another ATP-competitive AKT inhibitor) was also higher in PRKG1-KO cells (Fig. 3H). In contrast, the IC50 of allosteric inhibitors such as miransertib and MK-2206 barely changed in PRKG1-KO cells (Fig. 3H).
Altogether, these results demonstrate that in RMS, miransertib acts specifically inhibiting AKT whereas the anti-tumor activity of ipatasertib is dependent on both AKT and PRKG1.
PRKG1 is essential for RMS growth
To study the significance of PRKG1 in RMS in vivo, we silenced PRKG1 RNA expression in the FP-RMS cell line RH4 and in the FN-RMS cell line RD using shRNAs (Figs. 3E, 4A). We subcutaneously injected NOD-SCID mice with RH4 and RD cells infected with LKO-shControl (shCTRL), shPRKG1#1 and shPRKG1#2 and monitored tumor growth. PRKG1-KD tumors showed a significant delay in tumor growth compared to controls (Fig. 4B). Thirty-two days post-injection, KD-RD derived tumors were significantly smaller than shCTRL tumors (tumor volume mean of 131 (SEM: ±19,8) and 64 (SEM: ±13,3) mm3 for shPRKG1#1 and #2, respectively, and 593 mm3 (SEM: ±77,6) for shCTRL). Similarly, tumors generated from PRKG1-KD RH4 cells showed a statistically significant delay in tumor growth compared to control. Although this effect was more pronounced with the shPRKG1#2 sequence than with shPRKG1#1 (Fig. 4B). Western blot confirmed the downregulation of PRKG1 in shPRKG1-derived tumors (Fig. 4C).
A Western blot showing PRKG1 levels in whole cell extracts upon PRKG1 knockdown and with two shRNA sequences (shPRKG1#1 and shPRKG1#2) in RD cells. Tubulin was used as loading control. Blot is representative of two independent experiments. B Quantification of tumor growth based on tumor volume measurements (mm3) in RH4 (left) and RD (right) xenografts. Error bars represent Mean ± SEM at each time point. ***p ≤ 0.001 and **p ≤ 0.01. Two-tailed Wilcoxon signed-rank test with Bonferroni correction. Comparisons between shControl groups (in black) and each knockdown (shPRKG1#1 and shPRKG1#2) groups (in purple and pink) were statistically significant (RD: p = 0.001; RH4: p = 0.0078). In RH4, n = 10 and in RD, n = 12 tumors per group are represented. Source data are provided as a Source Data file. C Western blot analysis of PRKG1 levels in protein extracts collected from two independent tumors of shControl groups and knockdown (shPRKG1#1 and shPRKG1#2) groups at endpoint in RH4 and RD. D Western blot showing levels of PRKG1 in whole cell extracts upon PRKG1 knockout in RD cells. Tubulin was used as loading control. Results reproduced in four independent biological replicates. E Quantification of tumor growth based on tumor volume measurements (mm3) in RH4 (n = 10) and RD (n = 8) xenografts. Error bars represent Mean ± SEM at each time point. ****p ≤ 0.0001 and ***p ≤ 0.001. Two-tailed Wilcoxon matched-pairs signed-rank test comparing SCR- control [black] and PRKG1-KO tumors [purple]. RD: p = 0.000061; RH4: p = 0.0005. Source data are provided as a Source Data file. F Western blot analysis of PRKG1 levels in protein extracts collected from 3 different tumors of SCR (control) and KO_PRKG1 groups at endpoint in RH4 and RD. Tubulin was used as loading control. G Representative Ki67 IHC of SCR and PRKG1_KO xenografted tumors. Images are representative of four different tumors per condition. H Boxplot depicting the quantification of Ki67-positive cells within SCR and PRKG1_KO xenografted tumors. Mann Whitney test two-tailed Center line = median; box = 25th–75th percentiles; whiskers = min–max. Each dot represents one tumor. RH4 * p < 0.05 RH4 (p = 0.0286); RD (p = 0.0286). N = 4 tumors per group were quantified. Source data are provided as a Source Data file.
To further validate these results, we then used CRISPR-Cas9 technology to knockout PRKG1 expression (Figs. 3G, 4D). Following the same xenograft tumor generation strategy, we observed that PRKG1-KO cells gave rise to significantly smaller tumors than control cells (Fig. 4E), in agreement with the results obtained by shRNA strategy. The efficiency of the PRKG1 KO in tumor samples was evaluated by western blot (Fig. 4F). Accordingly, IHC analysis of Ki67 showed lower proliferation rates in PRKG1-KO tumors (Fig. 4G, H). Altogether, these results confirm PRKG1 as a critical factor for RMS tumor growth in vivo.
Silencing PRKG1 causes myogenic differentiation in RMS
To decipher the molecular basis of PRKG1-dependent regulation of RMS tumor growth, we performed RNA sequencing (RNA-seq) analysis of three biological replicates of control and PRKG1-KO RH4 cells. Hierarchical clustering of the RNA-seq data showed that PRKG1 silenced and control cells separated into two distinct groups (Supplementary Fig. 3A). Analysis of DEGs using a statistical significance cut-off, showed 1460 upregulated and 568 downregulated genes in the PRKG1-KO RH4 cells (Supplementary Data 2, Fig. 5A). Upregulation of some of the top DEGs identified in the transcriptomic analysis such as MYH8 and TNNI1 was validated by RT-qPCR in RH4 and RD PRKG1-KO cells (Fig. 5B). To determine whether these transcriptional changes were translated to phenotypic changes, we stained the sarcomere protein TNNI1 (troponin I) by immunofluorescence in cultured cells (Supplementary Fig. 3B) and by immunohistochemistry in xenografts (Supplementary Fig. 3C). Both RH4 PRKG1-KO and RD PRKG1-KO tumors showed increased TNNI1 expression and cytoplasmic elongation and striation, which were particularly pronounced in PRKG1_KO RD tumors (Supplementary Fig. 3C). In contrast, the expression of MYOG, an intermediate myogenic marker, was consistently reduced in PRKG1 KO xenografts, as well as in tumor samples from mice treated with ipatasertib but not with miransertib (Supplementary Figs. 4A, B). Importantly, although there is some evidence that the RAS pathway regulates differentiation in RMS18 we did not observe significant changes in the levels of phosphorylated ERK in PRKG1-KO cells nor in ipatasertib-treated cells (Supplementary Fig. 4C). These phenotypic changes, along with the increased expression of muscle markers in PRKG1-KO cells, indicate myogenic differentiation associated with PRKG1 suppression.
A Volcano plot showing upregulated genes (red) and downregulated genes (blue) in RH4 PRKG1-KO cells vs SCR Control (n = 3 independent biological replicates). Adjusted p value < 0.05; log2(FC) > | 1 | . p-values were adjusted using the false discovery rate (FDR). Source Data is available as Supplementary Data 2. B RT-qPCR analysis of TNNI1, TTN, MYL1, MYL4, TNNT3 and MYH8 in RH4 and RD PRKG1-KO cells. GAPDH was used as a housekeeping gene. C Dot plot of significantly enriched up and down regulated terms in PRKG1 KO vs SCR cells. Over-Representation Analysis (ORA) was performed using a one-sided hypergeometric test, with p-values adjusted via the Benjamini-Hochberg (BH) method to control the false discovery rate (FDR). D Western blot showing MYOD1 and PRKG1 levels in RH4 (n = 2) and RD (n = 1) cells transfected with siPRKG1, siMYOD1 or siRNA control and analyzed 72 h later. E RT-qPCR of genes involved in muscle differentiation in RH4 (left panel) and RD (right panel) cells, transfected as above indicated. Two biological replicates are shown. Source data is provided as a Source Data file.
Functional analysis of DEGs in PRKG-KO cells, performed by over-representation analysis (ORA), revealed the enrichment in genes involved in muscle differentiation and under representation of mesenchymal development genes (Fig. 5C). The corresponding gene set enrichment analysis (GSEA) identified the hallmark “myogenesis” as positively enriched (Supplementary Fig. 5A). Accordingly, the expression of several terminal differentiation myogenic genes was significantly induced (TNNI1, MYL1, MYH8, TTN) while genes regulating the mesenchymal stem cell (MSC) phenotype (COL3A1, MEST, BMPR1A, MESD, TNS1) were downregulated. A similar approach using RNA-Seq from PRKG1-KD RH4 cells showed the overexpression of myogenic markers (MYL1, MYH8, TNNI1 and TTN) (Supplementary Fig. 5B). In addition, TNNI1 protein overexpression was also observed in PRKG1-KD RH4 and RD cells (Supplementary Fig. 5C).
These results suggested that silencing PRKG1 unblock, at least in part, the undifferentiated myogenic transcriptome associated with RMS. To identify potential mechanisms that regulate PRKG1 expression during myogenic differentiation, we explored the relationship between PRKG1 and the master transcription factor MYOD1. Publicly available data from chromatin immunoprecipitation sequencing (ChIP) of the PRKG1 genomic region revealed MYOD1 binding to the transcription start site (TSS) and to an intragenic peak close to the TSS in RH4 cells and RD cells (Supplementary Fig. 5D). This was further confirmed by ChIP followed by PCR (Supplementary Fig. 5E), suggesting that PRKG1 might be part of the MYOD1-regulated myogenic differentiation program. To further understand this regulatory loop, MYOD1 was downregulated by transient transfection of short interfering RNA (siRNA). MYOD1 silencing in RH4 and RD cells resulted in enhancement of PRKG1 expression (Fig. 5D) and, concomitantly, in downregulation of terminal myogenic differentiation markers expression (Fig. 5E). Overall, this data confirms the role of PRKG1 in myogenesis and its relevance in RMS block of differentiation.
PRKG1 is expressed in fetal tissues and relates to mesodermal cell states in RMS
The role of PRKG1 in myogenic differentiation is mostly undescribed and for RMS completely unexplored. Therefore, we first analyzed PRKG1 expression in a collection of healthy embryonic tissues and stem cells by extracting data from public databases. Skeletal muscle precursor cells showed the highest PRKG1 expression levels (Fig. 6A). Accordingly, PRKG1 was detected by IHC in muscle precursor cells of 7–8 weeks human embryo samples (Fig. 6B). Across different developmental cancers including sarcomas, neuroblastoma, and brain tumors, PRKG1 is highly and differentially expressed in RMS, with levels only comparable to fetal skeletal muscle (Fig. 6C, D). Importantly, PRKG1 levels are independent of the oncogenic fusion status, the most determinant and prognostic significant molecular feature in RMS (Fig. 6E). IHC and confocal immunofluorescence of primary RMS tumor samples revealed high PRKG1 expression in FP and FN RMS patient samples (Fig. 6F, G). After verifying protein expression of PRKG1 in RMS, we performed a quantitative assessment of the protein levels in our experimental models of RMS. Protein analysis by western blot of primary tumors and PDXs showed varied levels of PRKG1, ranging from samples with low expression to samples with high PRKG1 expression (Fig. 7A).
A PRKG1 mRNA expression from publicly available GEO databases (Affymetrix U133plus2.0 array probe PRKG1_207119_at) of human mesenchymal and embryonic stem cells (hMSC and hESC, respectively) and embryonal tissues (skeletal muscle precursors in blue), fetal cerebellum and brainstem). Each dot represents an individual sample. Sample sizes per group are as follows: Skeletal muscle precursors (n = 10), hESC (n = 17), brainstem (n = 2), hMSC (n = 73), and fetal cerebellum (n = 5). Source data is provided as a Source Data file. B IHC of PRKG1 in a 7-week human embryo, with staining reproduced in three consecutive sections. C Confocal imaging of PRKG1 (in green) distribution in skeletal fetal muscle from HSJD biobank. The plasma membrane was stained with wheat germ agglutinin (WGA; in red). In blue, nuclear counterstain with DAPI. Observe the pronounced PRKG1 localization at cytoplasm. Representative field selected from multiple fields acquired at different magnifications. D PRKG1 mRNA expression from publicly available GEO databases (Affymetrix U133plus2.0 array probe PRKG1_207119_at) of pediatric tumors including: low-grade glioma (LGG), pilocytic astrocytoma (P. Astrocytoma), high-grade glioma or glioblastoma (HGG or GBM), diffuse intrinsic pontine glioma (DIPG), ependymoma, atypical teratoid/rhabdoid tumors (ATRT), medulloblastoma, neuroblastoma, osteosarcoma, Ewing sarcoma (Ewing_S), synovial sarcoma (Synovial_S), human skeletal muscle precursor cells (hSMPs) and Rhabdomyosarcoma (in turquoise). Each dot represents a different sample. Sample sizes per group are as follows: RMS (n = 58), ATRT (n = 58), Ewing_S (n = 142), GBM (n = 48), DIPG (n = 29) P_Astrocytoma (n = 41) Osteosarcoma (n = 17), Ependymoma (n = 11), Synovial_S (n = 34), Medulloblastoma (n = 217), Neuroblastoma (n = 122), LGG Brainstem (n = 6). Source Data is available. E Dot plot representing PRKG1 mRNA levels from 58 RMS patients in GSE66533. Each dot represents an individual sample. Sample sizes are: FP-RMS (n = 33) and FN-RMS (n = 25). Data are presented as mean values +/- SD. A two-tailed Mann-Whitney test was used to assess statistical significance, ns: p = 0,189. Source Data is available. F Immunofluorescence of PRKG1 (in green) distribution in one FN and one FP RMS tumors from HSJD biobank. In blue, nuclear counterstain with DAPI. Observe the pronounced PRKG1 localization at cytoplasm. Images from three independent tumors in FP RMS and three in FN RMS. G Histo (H)-score of IHC staining of PRKG1 in n = 24 RMS tumor samples from HSJD Biobank. Both FP and FN-RMS patient samples showed high levels of PRKG1 expression. Source Data are available.
A Western blots showing levels and quantification of PRKG1 in primary tumors (patient samples) and xenografts (PDX). Blot representative blot of one biological replicate per patient sample. B Box plot representing PRKG1 mRNA levels on the bases of a cut-off point ( = 5.1875) (left panel). Here, we define the UpperCP (High PRKG1 levels, n = 5) versus LowerCP (Low PRKG1 levels; n = 6) samples. Mann Whitney test two-tailed Center line = median; box = 25th–75th percentiles; whiskers = min–max. ** indicates p = 0.0043. Source data are provided as Source Data file. C Hierarchical clustering of the supervised analysis of RMS primary tumors (T) and PDX (X) in UpperCP vs LowerCP samples (see Fig. 7B). D GSEA plots of the core enriched genes in high (UpperCP) versus low (LowerCP) PRKG1 samples. GSEA was performed using the Broad Institute tool with default settings. Statistical significance was assessed by permutation-based, two-sided testing with FDR adjustment. Differential expression was analyzed using limma; probes were considered significant at FDR < 0.05. E Illustration of DEGs included in Mesoderm development gene signature (GSEA: NES = 1.97, FDR = 0.00089), shown by logFC and limma p values. Source Data is available (F) Analysis of RNA-seq data from 139 RMS samples (StJude Cloud) classified into three myogenic clusters: Mesoderm/ Myoblast/ Myocyte myogenic clusters by Patel et al. (2022). PRKG1 expression levels significantly correlate with mesoderm gene signature. Gene expression (x-axis) is plotted against GSVA scores (y-axis), with a linear regression line fitted for each facet. GSVA scores, and the corresponding correlation coefficient (R) and p value are shown on each panel. The shaded area indicates the 95% confidence interval for the estimated fit. Pearson correlation test and linear regression with 95% CI (calculated as the predicted mean ± 1.96 times the standard error. G Violin plot of integrated single-nucleus RNA-seq data from all patients in Patel et al., showing PRKG1 expression across annotated clusters from the original publication. H UMAP representation of tumor cells from patient SJRHB012_R displaying annotated clusters. I UMAP representation of tumor cells from patient SJRHB012_R displaying PRKG1 expression. J UMAP representation of tumor cells from patient SJRHB012_R displaying MEOX2 expression.
To molecularly characterize RMS with high PRKG1 expression, we classified samples (six primary tumors and their corresponding PDX models, Fig. 1D) according to their PRKG1 expression levels (LowerCP_PRKG1 and UpperCP_PRKG1) using the Human Genome U219 Array (Affymetrix, Thermo Fisher) and then performed a supervised transcriptional analysis (Fig. 7B, C). GSEA of DEGs in the High_PRKG1 group revealed enrichment in mesoderm developmental genes (Fig. 7D) and the mTORC1 pathway (Supplementary Fig. 6A). As shown in Fig. 7E, samples with elevated PRKG1 levels showed increased expression of genes such as BMPR1A, EYA1, MEST and SIX1, all involved in early mesodermal processes. In contrast, the expression of genes involved in myogenesis were negatively correlated to this subgroup (Supplementary Fig. 6B).
According to these results, we hypothesized that PRKG1 might be playing a role in the aberrant myogenic differentiation program of RMS. Furthermore, the observed mesodermal enrichment in the High_PRKG1 RMS group suggested that it could potentially be involved in stem cell maintenance. To ascertain whether PRKG1 expression in RMS was associated with an early myogenic transcriptional program, we used the RMS Transcriptome sequencing (RNA-Seq) data from 139 patient samples collected from various public resources, primarily from the St. Jude Cloud (detailed in Supplementary Fig. 6C and Supplementary Data 3)39. The 139 bulk RNA-seq samples were split into high and low PRKG1 groups using the 25th and 75th percentiles, with 35 samples in each group DEGs of PRKG1high_Q vs PRKG1low_Q in 139 RNA-seq samples are presented in Supplementary Data 4. A Venn diagram was used to depict the overlap between DEGs of PRKG1high_Q vs PRKG1low_Q in 139 RNA-seq samples and DEGs associated with our High_PRKG1 and Low_PRKG1 groups (Supplementary Fig. 6D, Supplementary Data 5). The analysis identified 921 shared genes, which are enriched in mesodermal and stem cell transcriptional profiles (Supplementary Fig. 6D). This enrichment highlights the role of PRKG1 in regulating genes associated with mesodermal development and stem cell characteristics.
To further analyze the role of PRKG1 in myogenesis, we computed the scores for the differentiation gene set classification (mesoderm, myoblast and myocyte) reported by Patel et al.3. Correlation analysis between PRKG1 expression levels and the three myogenic-state groups revealed high PRKG1 levels strongly correlated to the mesoderm gene set (R = 0.37, p = 9.6e-06) and, to a considerably lesser extent, the gene signature specific to myocyte (R = 0.26, p = 0.0022) but not to the myoblast gene signature (Fig. 7F).
Moreover, we analyzed integrated single nucleus data from 18 patient samples provided by Patel et al3. Violin plots visualized PRKG1 expression in seven myogenesis-related clusters defined by Patel et al. using the Leiden algorithm40, representing distinct stages from early mesodermal cells to fully differentiated myocytes. The highest PRKG1 expression levels were found in the mesoderm and early myoblast clusters (Fig. 7G), highlighting the importance of PRKG1 in early phases of myogenesis and providing insights into its temporal expression patterns during muscle development. Accordingly, UMAP representation of PRKG1 expression in a tumor sample, determined by integrated single nucleus RNA sequencing data, showed its expression in mesoderm and early myoblast clusters also expressing MEOX2, a transcription factor expressed in mesoderm and critical for mammalian muscle and bone development (Fig. 7H–J). Concomitant expressions of PRKG1 and MEOX2 in the same cell clusters during muscle development further supports that high PRKG1 expression is coupled to specific myogenic states on the continuum of the myogenic differentiation trajectory.
PRKG1 levels predict the anti-tumor activity of ipatasertib in RMS
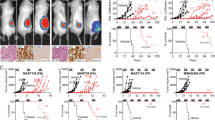
Ipatasertib efficacy was tested in vivo in six different RMS-PDXs, including FP and FN-RMS models. Briefly, at the maximum tolerated dose (MTD, 100 mg/kg/day), Ipatasertib substantially reduced tumor growth in some of the tested PDX models (Fig. 8A). According to the response, we could differentiate between two subgroups of RMS PDXs. One group including ERMS-011, ARMS-010 and ARMS-006, characterized by tumors that reduced more than 50% the tumor-initiating-volume by 2 weeks of treatment (10 doses). The second group characterized by delayed but continued growth while on ipatasertib treatment (ERMS-024, ARMS-007 and ARMS-001) (Fig. 8A, B). We explored one-quarter of the MTD dose in one FP and one FN-RMS PDX since 25 mg/kg is a feasible dose to translate into the clinic. As shown in Fig. 8C, treatment with ipatasertib showed a dose-dependent effect in the RMS models tested since 25 mg/kg slowed tumor growth rate for those models where 100 mg/kg completely abrogated the tumor. At the highest dose, the mean survival time for ARMS-010 model was extended to 71 days (mean survival time for control mice, 38 days), and to 35 days (versus 10 days of the control group) for mice carrying the aggressive ERMS-024 model (from the second CNS relapse of our index case).
A Efficacy of ipatasertib administrated at 100 mg/kg orally (PO) and daily (QD) in six RMS-PDX. Individual tumors growth curves for vehicle (black) and ipatasertib (blue). The treatment period highlighted in gray. Data came from one single experiment for each model, except for ARMS-006, in which two independent experiments were performed. ARMS-A001, n = 8; ARMS-006 n = 8; ARMS-007, n = 6; ARMS-010, n = 6; ERMS-011 n = 6; and ERMS-024, n = 8. Source Data are available. B Percentage of tumor response, evaluated after 10 doses of ipatasertib at 100 mg/kg administered PO, QD. Response was classified as: complete response (CR: tumor volume: < 100 mm3 + reduction >50%), partial response (PR: tumor volume ≥100 mm3 +reduction ≥ 50%), stable disease (SD: reduction < 50% or increase ≤ 25%) and progressive disease (PD: reduction < 50% or increase >25%). ARMS-001 (n = 12), ARMS-006 (n = 10, from two independent experiments), ARMS-010 (n = 7), ARMS-007 (n = 7), ERMS-011 (n = 9), and ERMS-024 (n = 11); n indicates number of treated tumors in each PDX. Source Data are available (C) Dose-response tumor volumes fitted to a simple linear regression comparing vehicle (dark purple) and ipatasertib-treated tumors (100 mg/kg in dark blue and 25 mg/kg in turquoise). Mean values +/- SEM. Data from vehicle and 100 mg/kg treated tumors came from previous graph. Kaplan-Meier survival curves for ARMS-010 and ERMS-024 PDXs treated with ipatasertib (100 or 25 mg/kg). The survival event was reported at 1500 mm3 tumor size. Log-rank test with Bonferroni correction with three comparisons in each PDX: vehicle vs. 100 mg/kg—p = 0.0054 (ARMS-010) and p = 0.00011 (ERMS-024); vehicle vs. 25 mg/kg—(p = 0.4704 (ARMS-010) and p = 0.0126 (ERMS-024); 100 mg/kg vs. 25 mg/kg — (p = 0.0067 (ARMS-010) and p = 0.00020 (ERMS-024). Tumors per group: ARMS-010 vehicle (n = 5), 25 mg/Kg ipatasertib (n = 7), 100 mg/Kg ipatasertib (n = 8); ERMS-024 vehicle (n = 5), 25 mg/Kg ipatasertib (n = 9), 100 mg/Kg ipatasertib (n = 12). Source Data are available (D) Individual tumor volume in ipatasertib-treated mice. Each plot represents an individual mouse with tumors in the right (TD) and left (TI) flanks. ID identification for mouse. E Scatter plot of PRKG1 mRNA levels (11740294_a_at probe) positively correlating with tumor regression after ten ipatasertib doses. Spearman R = 0.8828 correlation two tailed (p value = 0.0007). Source Data are available.
As shown in Fig. 8A, tumors responding to ipatasertib regrew upon stopping therapy. To investigate tumor escape mechanisms, ipatasertib was re-administered to the ERMS-011 mice when the tumor regrew at 150–300 mm3. As seen in Fig. 8D, re-challenge with ipatasertib reduced the tumor volume twice after two serial sequences of on/off treatment, extending the survival of the ipatasertib-treated mice to 100 days.
The PRKG1 mRNA levels in PDXs and primary tumors positively correlated to ipatasertib efficacy in vivo evaluated after 10 doses (Fig. 8E). RMS models with complete response to ipatasertib corresponded to those with higher PRKG1 levels (High_PRKG1), while RMS models with stable or progressive disease displayed low mRNA levels of PRKG1 (Low_PRKG1). This data identified PRKG1 as a potential biomarker for ipatasertib-response in RMS. The established cut-off was a useful tool to cluster both groups for subsequent analysis. None of the AKT mRNAs (AKT1, AKT2 and AKT3) were among the DEGs in the unsupervised clustering analysis (SD > 1), nor correlated to ipatasertib efficacy (Supplementary Fig. 7). These results demonstrate PRKG1 as a bona fide biomarker for ipatasertib response in RMS. Of note, PRKG1 protein levels in E001_1 (first CNS relapse) compared with ERMS-024 (second CNS relapse) models were reduced (Supplementary Fig. 8A), in agreement with a reduced sensitivity to ipatasertib (Supplementary Fig. 8B), further confirming the role of PRKG1 as biomarker for ipatasertib response. Notably, EGFR and PROM1(CD133) (mesodermal markers) expression were reduced in the second relapse samples while the expression of the differentiation marker MYH8 expression was increased (Supplementary Fig. 8C).
The differential anti-tumor efficacy of ipatasertib in the eight RMS models tested could be related to drug bioavailability. To test this hypothesis ipatasertib pharmacokinetics (PK) was studied in the ERMS-E011 model (complete remission upon ipatasertib treatment) (Fig. 9A) and the ARMS-006 model (partial response to ipatasertib) (Fig. 9C). After a single ipatasertib dose, either at 100 mg/kg or at 25 mg/kg, samples were collected at different time points and analyzed by liquid chromatography/mass spectrometry (LC/MS). As expected, the area under the curve (AUC) and the maximum observed concentration (Cmax) were higher at 100 mg/kg than at 25 mg/kg in both PDX models (Supplementary Data 6). Drug accumulation phase was longer in the ARMS-006 xenografted mice, in which the peak concentration (Tmax) in the tumor was 8 h, while in the ERMS-011 model Tmax was 1 h at 25 mg/kg dose and 3 h at 100 mg/kg dose, an earlier decay phase (Fig. 9A, C). This profile curve allowed keeping intra-tumor levels in the ARMS-006 model higher than 1 µM for both 25 and 100 mg/kg doses within 24 h post administration ensuring prolonged tumor exposure to ipatasertib. At 100 mg/kg dose, the intra-tumor concentration remained high above 1 µM in both models (Fig. 9C). In addition, plasma AUC₀–₂₄ values following both 100 mg/kg and 25 mg/kg dosing as well as tumor-to-plasma ratios and intra-tumoral concentrations at 1, 3, 8, and 24 h were similar in the ARMS-006 and ERMS-011 models (Supplementary Data 6). Therefore, these data demonstrate how ipatasertib reached enough intra-tumor concentrations to inhibit AKT signaling and to effectively slow tumor growth and indicate that the variation in treatment efficacy cannot be attributed to differences in drug exposure.
A, C Plasma and tumor concentrations of ipatasertib in RMS PDX over time (0–24 h). For reference, the value of 1 μM is indicated in all graphs. AUC (area under the curve), Cmax (maximum observed concentration) and Tmax (the time of Cmax) in plasma and tumor are shown in Supplementary Data 6 for ERMS-011 and ARMS-006 bearing mice. Pharmacodynamic changes at 3 h and 24 h after a single ipatasertib dose. Source data are provided as a Source Data file. B, D IHC for pS6 (240/244), cleaved-PARP and human nuclei in control and treated tumors in ARMS-006 and ERMS-011 at 100 mg/kg and 25 mg/kg. One representative tumor per condition is shown; IHC was performed on all tumors included in the PK analysis (images of representative fields). Pictures, 20X magnification.
Since ipatasertib inhibits both AKT and PRKG1, S6 phosphorylation levels were used as a pharmacodynamic marker. Compared to controls, the strongest reduction in pS6 signal was observed 3 h after ipatasertib treatment, and pS6 levels were restored at 24 h both in ERMS-E011 and ARMS-006 (Fig. 9B, D). Moreover, cleaved PARP levels notably increased 24 h after a single dose of ipatasertib at 100 mg/kg in tumor samples (Fig. 9B, D). Notably, pharmacodynamic analysis in the ipatasertib-resistant ARMS-007 model showed that pS6 levels were also reduced, confirming that ipatasertib effectively reaches the tumor and inhibits AKT (Supplementary Fig. 9). In summary, the differential anti-tumor response to ipatasertib is due to intrinsic differences in tumor cell biology, rather than pharmacological variability.
Taken together, 25 mg/kg as well as 100 mg/kg doses were sufficient to achieve effective ipatasertib intra-tumor concentrations and cause cell death. However, only the 100 mg/kg dose was able to fully inhibit the AKT pathway at 24 h. Our data suggests that the 25 mg/kg dose would require daily ipatasertib administration whereas the 100 mg/kg dosing may allow every other day administration since S6 inactivation persisted and the intra-tumor concentration of the active drug stayed over 1 µM for more than 24 h.
Discussion
Robust evidence supports that RMS is driven by cells blocked in their myogenic potential, failing to differentiate. Various terms describe distinct subpopulations within RMS. Studies by Patel et al.3, Wei et al.15, and Danielli et al16. detail the cellular heterogeneity and impaired differentiation that compound RMS at the single-cell level. We could confirm through a meta-analysis of these RMS single cell RNA-seq datasets that similar gene signatures are conceptually the same.
Despite intensive research, differentiation blockade has not yet provided a real therapeutic opportunity for the management of patients with high-risk RMS. In this study we unveiled the role of PRKG1 in the complex process of myogenic blockade that defines RMS tumorigenesis and provide evidence for effectively targeting a critical pathway common to all RMS with ipatasertib, an oral drug with adequate pharmacological characteristics to be tested in the clinic.
PRKG1 (cGMP-dependent protein kinase) is a protein kinase regulated by cGMP with multiple activities including smooth muscle relaxation by activation of myosin phosphatase41. In smooth muscle a crosstalk between PRKG1 and BMP signaling pathways was described42. PRKG1 regulates FOXO1A activity during myoblast cell fusion43 and in cardiomyocytes PRKG1 controls the sarcomere and cytoskeleton, including the activity of TTN and TNNI144, and phosphorylates TSC2, the GTPase regulatory subunit of mTORC145. In other physiological backgrounds, PRKG1 mediates osteoblast differentiation46.
The role of PRKG1 in cancer includes enhanced SRC activation, DNA synthesis and cell proliferation in high-grade ovarian cancer47; increased cell viability in glioma neurospheres48; and contradictory roles in apoptosis in breast and non-small cell lung carcinoma49,50. To our knowledge, the role of PRKG1 in pediatric cancers has not been described.
In the RMS literature, PRKG1 appears almost unnoticed in the supplementary list of a study analyzing the interaction between NCOR/HDAC3 and MYOD151. Moreover, PRKG1 is listed among the genes harboring PAX3-FOXO1 binding sites52 and genes related to PAX3-FOXO153. These data, together with the presence of MYOD1 at the PRKG1 transcription start site in RH4 cells, strongly suggest that PRKG1 acts as an effector of the MYOD1-regulated differentiation blockade in RMS. In this study we showed that PRKG1 expression is under the control of MYOD1 and confirmed its role in the myogenic blockade that defines RMS.
Ipatasertib, but not miransertib, potently inhibits PRKG138. Similar ATP-competitive AKT inhibitors, such as afuresertib and uprosertib, were previously described to inhibit PRKG1 as well54. Ipatasertib is an orally administered inhibitor of all three isoforms of pAKT and has been studied in solid tumors in adults both in pre-clinical and clinical trials33,34,35,55,56,57. These clinical advances in adults provide a real opportunity to test ipatasertib in the clinic for a rare tumor such as RMS.
High pAKT expression predicted sensitivity to ipatasertib in preclinical models and in clinical trials58, but not in endometrial cancer cells59. In our study we showed that expression levels of PRKG1 correlated to ipatasertib response in RMS providing a useful biomarker to be tested in clinical trials.
At clinically feasible doses, ipatasertib monotherapy blocked PDX-RMS growth in vivo and induced tumor responses in mice. However, tumors regrew after discontinuation. Re-challenging with ipatasertib inhibited regrowth in sensitive models, suggesting that a maintenance type of strategy could prolong tumor remission in patients.
Ipatasertib pharmacodynamics was demonstrated in RMS models, confirming the ability of ipatasertib to inhibit tumor growth by blocking mTOR signaling and causing cell death. The efficacy data associated with favorable biodistribution, and limited toxicity reinforces the potentials of ipatasertib for clinical development in RMS. Ipatasertib achieved intra-tumor concentrations higher than required to induce apoptosis in the models and doses tested explaining tumor regression. Importantly, the intra-tumor concentration 24 h after the lower ipatasertib dose (25 mg/kg) was yet higher than the Cmax. These data suggest that the administration schedule could be reconsidered for the 100 mg/kg dosing with the aim of reducing toxicity. Every other day regimen, instead of daily administration currently in use, might be equally effective and less toxic for RMS patients. Based on human adult data33,60,61 and body surface area–adjusted doses62, the doses of 25 and 100 mg/kg in mice (equivalent to ~120–480 mg in humans) fall within the clinical range. Notably, intra-tumoral concentrations remain elevated at 24 h despite declining plasma levels, indicating efficient tumor distribution. These findings support the translational relevance of our murine RMS models. A limitation of this study is the lack of additional late time points needed to accurately compare elimination curves and t1/2. Here, we focused on drug penetration at the target site and tumor-to-plasma concentration ratios.
In the literature, ipatasertib was administered to a 12-year-old patient with an epithelioid neoplasm with a fusion oncogene that involved AKT. This patient was reported to receive 300 mg/day and showed an objective tumor response63. This dosing is consistent with our PK data. However, in younger children this dose might turn exceedingly toxic. Based upon our data, for patients experiencing toxicity to the 100 mg/kg equivalent dosing of ipatasertib, it would be worth exploring every 48 h administration rather than reducing the dose of ipatasertib. Furthermore, clinical studies should examine the potential of 25 mg/kg dosing with PK guided schedules.
The in vivo efficacy studies of ipatasertib in the tumor model from the first CNS relapse of the index patient supported the compassionate use of ipatasertib in the context of a second aggressive LM relapse. In vivo efficacy studies were performed in the second CNS relapse model (E024) parallel with the patient’s treatment with ipatasertib. Treatment was shown to be equally ineffective in the E024 model and the patient. Several reasons explain this. First, the dose. No previous experience with ipatasertib in infants existed at that time, therefore, to reduce potential toxicities, the dose approved by the Ethics Committee informed by data from Roche was reduced five times the dose we had proposed based upon preclinical data. Second, the CNS location. Ipatasertib appears to have a reduced blood-brain barrier passage when preserved. Third, the patient had received one year of temsirolimus (plus irinotecan and temozolomide) prior to ipatasertib treatment. We showed how the PRKG1 levels decreased from first to second relapse. According to our current understanding of the role of PRKG1 as a biomarker of response to ipatasertib in RMS, we could have predicted limited efficacy by the time the patient was treated. Whether the inhibition of PRKG1 enhances the effect of cytotoxic agents against RMS as shown for pAKT inhibition remains an ongoing question currently being tested in the laboratory.
In conclusion, our results provide several clues on how to personalize target therapy against RMS using PRKG1 as a biomarker. The characterization of the role of PRKG1 in the pathophysiology of RMS opens a new therapeutic avenue to be readily explored in patients with RMS.
Methods
Cell lines
Rhabdomyosarcoma commercial cell lines RH4, RH30 and RD were kindly provided by Dr. Martinez-Tirado (Instituto de Investigación Biomédica de Bellvitge (IDIBELL), Barcelona). Cells were cultured in RPMI 1640 medium supplemented with 10% fetal bovine serum (FBS) (Hyclone, Lonza), 5 mM of L-glutamine, and 10,000 U/ml of penicillin-streptomycin (Gibco) and were grown in monolayer conditions. All cell cultures were maintained at 370C in a humidified atmosphere containing 5% CO2. All human cell lines used in this study were authenticated by Short Tandem Repeat (STR) profiling, performed at qGenomics.
Establishment of RMS primary cultures
Primary cultures (ARMS-006, ARMS-010 and ERMS-024) were generated from patient derived xenografts (PDX). Tumor samples were mechanically dissociated using two scalpels and then filtered with 0.45 µm Corning™ Sterile Cell Strainers (Fisher scientific). After collected for centrifugation at 400 x g for 3 min, tumors were cultured in tumor stem media (TSM) consisting of a 50:50 mixture of DMEM/F12 (Dulbecco’s Modified Eagle Medium/Nutrient Mixture F-12) and Neurobasal medium (Invitrogen), HEPES buffer solution (10 mM), 1% Sodium pyruvate MEM (100 mM), 1% MEM non-essential aminoacids (10 mM), 1% glutamax-I supplement and antibiotic and antimycotic (Ref 15240096) all of them purchased from Fisher™. In addition, the medium also contained the following factors: B-27™ Supplement minus vitamin A (Gibco™), heparin (2 μg/ml) (H3149-10KU, Sigma), H-EGF (20 ng/ml), H-FGFb (20 ng/ml), H-PDGF-AA (10 ng/ml) and H-PDGF-BB (10 ng/ml)64. All the human recombinant growth factors were purchased from PeproTech. The cultures were incubated at 37 °C, 5% CO2, and 95% humidity. Media was refreshed every 2–3 days.
A stock of cells was frozen in freezing medium (Synth-a-Freeze cryopreservation medium, ThermoFisher). Upon the establishment of each primary culture, the presence of RMS and human tissue-specific markers were analyzed. To this end, the expression of human nucleus antigen, MyoG, MyoD1, and the fusion genes when applicable, were routinely checked in primary cultures in vitro.
Clinical specimens
All human samples used to generate our RMS-PDX models came from fresh biopsies of patients at Hospital Sant Joan de Déu (HSJD). For clinical and molecular features see Fig. 1D. All patient material was collected after signed informed consent and procedures approved by the Institutional Ethics review Committee. The goal was to establish new RMS models in vitro from patient samples and from PDX. Briefly, fresh tissue specimens were obtained from surgical samples from patients at HSJD. The RMS diagnosis was confirmed by a Board-certified pathologist and molecular biologist using standardized tests including RT-PCR and FISH for fusion gene confirmation when applicable.
Pharmacological treatments
Ipatasertib (GDC-0068) was provided by Genentech (affiliated to F. Hoffmann-La Roche Ltd), and miransertib (ARQ-092-2MSD) was provided by Arqule (recently affiliated to MSD), and the use of both inhibitors were under material transfer agreements (MTA). Trametinib (GSK1120212) was purchased from Selleck Chemicals.
Structure preparation
Starting structures for docking and MD simulations were downloaded from the Protein Data Bank (www.rcsb.org): PDB ID 4ekl for AKT and 7lv3 for PRKG1. All ions, solvent molecules and ligand molecules were removed, except for the MD simulation of the AKT-ipatasertib complex. The resulting structures were prepared with the Protein Preparation Workflow from Schrödinger implemented in Maestro65: missing hydrogen atoms were added, protonation state of titratable residues at pH 7.4 was computed on the basis of their predicted pKa values, and the hydrogen-bonding network was optimized. Asn, Gln and His residues were checked for adequate side chain flipping. The final structures were subjected to restrained minimization with OPLS-AA force field, keeping heavy atoms in place and optimizing the positions of hydrogen atoms.
Docking simulations
The co-crystallized ligands were used to center the docking grid which had an inner box size of 12Åx12Åx12Å and an outer box size of 22Åx22Åx22Å, covering the ATP-binding pocket. Docking simulations were carried out using the Standard Precision (SP) protocol of Glide66. For the induced fit protocol, hydrogen bond constraints to the hinge residues (Glu439 backbone CO and Cys441 backbone NH) were applied. Briefly, a constrained minimization of the receptor is performed, followed by docking of the compound using a softened potential, refinement of the docked poses with Prime (side-chains energy minimization), and final redocking to the best receptor structures. All reported scores correspond to glide gscores.
Molecular dynamics simulations
The preprocessed protein structures (AKT-ipatasertib complex from PDB ID 4ekl and PRKG1-ipatasertib complex from docking) were immersed in a truncated octahedral box of water solvent extending at least 12 Å from any complex atom. TIP3P model was used for water molecules. The force fields used were Amber ff14SB for protein residues and gaff2 for small molecule atoms. Solvated systems were subjected first to an energy minimization procedure to adjust solvent orientation, eliminate local clashes and stereochemical inaccuracies. The following equilibration protocol consisted in 1 ns of constant volume MD, where temperature was slowly raised from 10 to 300 K, after which 10 ns of constant pressure and temperature MD was performed (1 bar, 300 K) to allow the system to reach proper density. In all MD simulations, temperature and pressure control was achieved using the Langevin thermostat and Monte Carlo barostat, respectively. Systems were simulated using periodic boundary conditions and Ewald sums (grid spacing of 1 Å) for treating long-range electrostatic interactions with a 8 Å cutoff for direct interactions. The SHAKE algorithm was used to keep bonds involving H atoms at their equilibrium length, and hydrogen mass repartition was applied allowing the employment of a 4 fs time step for the integration of Newton’s equations. Simulations were performed with the PMEMD implementation of SANDER for GPU from the Amber 20 program package. Finally, the system was subjected to 0.5 microsecond MD production run. No restraints were applied to any atom.
Cell viability studies by MTS assay: IC50 calculation
Cells were seeded in 96-well plates in a proportion of 3.000 cells/well (RH4 and RD) or 10.000 cells/well (A006_s, A010_s, and E024_s) 24 h prior to treatment (370C, 5% CO2). They were treated with each drug at 1:2 or 1:10 serial dilutions into final concentration ranges starting from 10 μM. Seventy-two hours later, 10% MTS was added and absorbance at 490 nm was read using a Tecan microplate reader. Percent viability was calculated by normalizing absorbance values to those from cells grown in media with vehicle treatment, after background subtraction. IC50 was determined with log (inhibitor) vs. response. Variable slope (four parameters) curves to determine the right value using Prism 8 Software (GraphPad).
CFSE proliferation assay
CellTrace™ CFSE Cell Proliferation Kit (Invitrogen™, C34570) was used according to the manufacturer’s instructions. RH4 SCR or PRKG1-KO cells were seeded at a density of 150,000 cells per well. After overnight attachment, cells were incubated with 5 µM CFSE (working concentration) for 20 min at 37 °C in the dark. Following incubation, cells were washed with RPMI supplemented with 10% fetal bovine serum (FBS). Thirty minutes later, cells were treated with 1 µM ipatasertib or vehicle. After 120 h of treatment, cells were detached using trypsin, washed twice, fixed with paraformaldehyde, and analyzed by flow cytometry using the ACEA NovoCyte 3000 (ACEA Bioscience Inc.). Data were processed using NovoExpress Software (ACEA Bioscience Inc.).
siRNA oligofection to knockdown PRKG1 in the RH4 cell line
Transient transfection of siRNA to knockdown PRKG1 was performed as follows. Briefly, 2.5 × 105 cells/well were seeded in 6-well dishes, and 24 h later were transfected with four different siRNAs targeting PRKG1, or with a Luciferase siRNA as a negative control. The siRNA-lipid complex was produced by diluting 25 pmol of each siRNA duplex in OPTIMEM media with 7 μL of Lipofectamine™ RNAiMAX reagent (Thermo Fisher Scientific, CA, USA) following the protocol guidelines of the manufacturer. Cells were harvested 72 h after transfection and subjected to transcriptional and proteomic analysis. Small interfering RNA (siRNA) knockdown sequences are presented in Supplementary Table 1.
Knockout of PRKG1 in the RMS cell lines using CRISPR-Cas9 technology
RH4 cells were nucleofected. The guide RNA (gRNA) sequence targeting the third exon of PRKG1 was selected using the CRISPR-Cas9 gRNA design tool (https://www.synthego.com/products/bioinformatics/crispr-design-tool). Single-guide RNA (sgRNA) were purchase from Synthego (CRISPRevolution sgRNA EZ Kit). Nucleofection was carried out using the Amaxa nucleofection system (P3 Primary Cell 4D-Nucleofector Kit, Cat#V4XP-3032) according to the manufacturer’s instructions. Briefly, for RNP formation, sgRNA:Cas9 ratio of 4.5:1 was used. 4 × 105 RH4 cells were electroporated using the CA137 nucleofection program and plated into 24 well plates. After 48 h, the cells were sorted and plated as a single cell on 96 well plates for 15–30 days to make clones. Genomic DNA (gDNA) was extracted using Cell lysis solution QIAGEN (cat#158908). Protein precipitation solution QIAGEN (cat#158912) and DNA hydration solution QIAGEN (cat#158914) following manufacturer’s instructions.
For RD cells lipofection was used. Genome knockout of PRKG1 in RMS cells was performed using a ribonucleoprotein (RNP) complex that consists of purified Cas 9 nuclease duplexed with chemically modified synthetic single guide RNA (sgRNA). For RNP formation a sgRNA ratio of Cas9 1,3:1 was used; therefore, the reaction comprised 3,9 pmoles sgRNA and 3 pmoles recombinant Cas9. In the same tube, Lipofectamine TM Cas9 Plus reagent was added and incubated for 10 min to generate Cas9/sgRNA RNP, following Synthego guidelines. For the RNP delivery into cells, LipofectamineTM CRISPRMAX was used as transfection reagent. The RNP-transfection solution was mixed with cells prior to plating, RNP transfection solution mixed together with cells were splited into 24 well plates. Each reaction was designed for a 24 well plate and 6×104 cells. After 72 h, the cells were sorted and plated as a single cell on 96 well plates for 15–30 days to make clones. Sequences of sgRNA guides used for CRISPR-Cas9 experiments are shown in Supplementary Table 2.
Individual clones were sequenced and clones with confirmed knockout were expanded for further experiments. Amplified fragments were Sanger sequenced and indel frequency was calculated using the ICE tool.
Here, we used a multi-sgRNA strategy, in which multiple sgRNAs are designed to jointly knockout PRKG1 gene. When co-transfected, the sgRNA includes a large fragment deletion, resulting in a robust knockout. We used as negative control Cas9 complexed. Also, mock conditions with parental (untransfected) cells were routinely included.
Cell culture and transduction
The lentiviral silencing constructs from MISSION® shRNA Library TRCN0000000996 and TRCN0000010033 were used for PRKG1. TRCN0000039794 and TRCN0000001612 were used for silencing AKT. The MISSION® pLKO.1-puro Empty Vector Control Plasmid DNA as control all of them from the TRC library (sigma-aldrich). 2 μg VSV-G, 3 μg pRRE, 3 μg pRSV-Rev and 4 μg shRNA plasmids with X-tremeGENE™ HP transfection reagent were transfected to 293FT cells to package all vectors. The RMS cells were transduced with 293FT supernatant containing lentivirus for two to three days. To select for stable infection cells were maintained in medium containing 2 μg/ml puromycin.
RNA extraction and quantitative real time PCR
RNA was isolated and purified with the RNA easy Mini kit Quiagen (cat# 50974104) following manufacturer instructions. For in vivo or human tissue samples, RNA was isolated with Trizol reagent (Sigma-Aldrich) (composed by a phenol solution), followed by the addition of chloroform. Finally, the RNA in the aqueous phase was precipitated by isopropanol and ethanol. RNA was quantified using Nanodrop one C (Thermo Fisher). The retrotranscription of RNA to cDNA was performed with 1 µg RNA with Applied Biosystems™ GeneAmp™ dNTP Blend (2.5 mM each), Fisher Random Hexamers (50uM), and M-MLV Reverse Transcriptase (200 u/μL) (Promega, Fitchburg, WI, USA). The quantitative analysis of gene expression was performed with Syber Green PCR Master Mix (Applied Biosystems) and specific forward and reverse primer. The qPCR was accomplished on Applied Biosystems Quant Studio 6 Real-Time PCR system (Thermo Fisher), using the ΔΔCT relative quantification method. GAPDH or TBP were used as housekeeping genes. The sequences of cDNA primers used for qPCR are presented in Supplementary Table 3.
Gene expression analysis
Microarray: High versus Low PRKG1 gene expression in HSJD-RMS tumors
To study transcriptomic differences between RMS tumors with high and low PRKG1 gene expression, RNA was summited to microarray hybridization at the Institut d’Investigacions Biomèdiques August Pi Sunyer (IDIBAPS) Genomic Service following standard procedures after the 3’IVT Pico method. Quality and integrity RNA sample were analyzed using bioanalyzer, and the PrimeView Human Genome U219 Array Plate (Thermo Fisher Scientific) was used. Data were quality controlled and normalized using the robust multichip average (RMA) algorithm using the R oligo package67,68. Standard deviation (SD) density plots were used to determine the cut-off value for gene expression unsupervised analyses (SD > 1). The median of the expression of PRKG1 probe 11740294_a_at (5.1875) was used as PRKG1 expression cut-off to classify samples into High-PRKG1 (UpperCP) and Low-PRKG1 (LowerCP). The supervised analyses were performed using the R limma package69. Probes were considered significantly differentially expressed when the adjusted false discovery rate (FDR) was < 0.05. To functionally compare High-PRKG1 samples to Low-PRKG1 samples gene we performed gene set enrichment analysis (GSEA). GSEA analysis (http://www.broadinstitute.org/gsea/index.jsp) was performed using default parameter settings.
Microarray gene expression in RMS cells treated with ipatasertib or miransertib
Samples were processed at MARGenomics IMIM’s core facility. Samples background was corrected, quantile-normalized and summarized to gene-level using the robust multi-chip average (RMA)64,67 obtaining a total of 20893 transcript clusters. The differential gene expression (DGE) analysis was assessed with an empirical Bayes moderated t-statistics model implemented in the limma package (v.3.54.2)69. P values were adjusted using the FDR70. Genes were considered differentially expressed with adj. p value < 0.05.
Pre-Ranked Gene Set Enrichment Analysis (GSEA)71 implemented in clusterProfiler (v.3.18.0)72 was used to retrieve enriched functional pathways. The ranked list of genes was generated using the –log (p.val)*signFC for each gene from the statistics obtained in the DGE analysis with limma. Functional annotation was obtained based on the enrichment of gene sets belonging to Hallmark gene set collection (v.7.2) in Molecular Signatures Database (MSigDB). All analyses were performed using R version 4.0.3.
RNA sequencing and gene set enrichment analysis
Raw sequencing reads in the fastq files were mapped with STAR (v.2.7.8)73 Gencode release 41 based on the GRCh38.p13 reference genome and the corresponding GTF file. The table of counts was obtained with featureCounts function in the package subread (v.2.0.3)74. The DGE analysis was assessed with voom and limma in the limma package (v.3.54.2)69.
Genes having more than 10 reads in at least 3 samples were kept. Raw library size differences between samples were treated with the weighted “trimmed mean method” TMM75 implemented in the edgeR package version 3.40.276. The normalized counts were used for the unsupervised analysis. For the DGE analysis, read counts were converted to log2-counts-per-million (logCPM) and the mean-variance relationship was modeled with precision weights using voom approach in limma package. Linear models were fitted using the limma package, and empirical Bayes moderated two-sided t-tests were applied to assess differential expression. P values were adjusted for multiple testing using the Benjamini–Hochberg false discovery rate (FDR)70. Genes were considered differentially expressed with adjusted. p value < 0.05 and |log2(FC)| > 1.
Functional enrichment was performed with functions implemented in the clusterProfiler package (v.4.2.2). GSEA71 was used with the list of all genes ranked by -log(p.val)*signFC from the statistics obtained in the DGE analysis with limma. Universal over-representation analysis (ORA) of DE genes (adjusted p value < 0.05) was also used to obtain enriched functional pathways. Functional annotations were obtained from Hallmark (v.2023.2)77 and differentiation-related pathways from both GO.BP (v.2023.2)78 and CGP (v.2023.2). All analyses were performed using R software version 4.2.1.
PRKG1 expression in pediatric databases, microarray platform
Thirty public GO_datasets for pediatric tumor transcriptomes, generated with the platform U133 plus2.0, Affymetrix were identified, in collaboration with FSJD genomic group. A dataset of 783 pediatric tumors plus mesenchymal (hMSC), embryonic stem cells (hESC), and human Skeletal Muscle Precursor cells (hSMPs), was generated. Tumors represented include medulloblastoma (MB), osteosarcoma (OS), neuroblastoma, Ewing Sarcoma (Ewing_S), synovial sarcoma (Synovial_S), diffuse intrinsic pontine glioma (DIPG), low-grade glioma (LGG), high-grade glioma or glioblastoma (HGG or GBM), ependymoma, pilocytic astrocytoma (P_Astrocytoma) and Rhabdomyosarcoma. Importantly, 58 RMS samples are included. Gene expression data normalization was performed using RMA algorithm included in the oligo R-package (R/Bioconductor). Quality control was done using oligo and limma R-packages (R/Bioconductor). There were three probes associated with PRKG1 gene. For our analysis, the PRKG1 207119_at, 211380_s_at, 228396_at probes were analyzed. Datasets included in this study have been previously reported and were obtained from the NCBI Gene Expression Omnibus (GEO) data repository. Reference Accession Numbers GSE67851, GSE70678, GSE44971, GSE26576, GSE74195, GSE13828, GSE7896, GSE8884, GSE9440, GSE9510, GSE17679, GSE34620, GSE37371, GSE34824, GSE36245, GSE26576, GSE10327, GSE37418, GSE49243, GSE67851, GSE74195, GSE10315, GSE13604, GSE6460, GSE7637, GSE9520, GSE9451, GSE9593, GSE16254, GSE73537, GSE14827, GSE44971, GSE26576, GSE66533, GSE20196 and GSE44227. All microarray analysis was performed by Dr. Soledad Gómez (HSJD DAI-Omics unit).
PRKG1 expression in Rhabdomyosarcoma patient databases, RNAseq platform
RNA-Seq raw counts of Rhabdomyosarcoma human samples were obtained from the St. Jude Cloud platform (https://www.stjude.cloud)39. It includes a compendium of datasets, mainly from Childhood Solid Tumor Network (CSTN), Pediatric Cancer Genome Project (PCGP), Real-time Clinical Genomics (RTCG). After removal of duplicate samples, this public cohort comprises 139 samples.
We kept genes having mean CPM > 1. Raw library size differences between samples were treated with the weighted “trimmed mean method” TMM implemented in the edgeR package (v3.40.2). The normalized counts per million (log2CPM) were used for the unsupervised analysis, PCA and clusters. We used Combat from sva package (v3.44.0)79 to correct the batch effect detected due to library preparation protocols. Batch variable was defined as the combination of “attr_library_selection_protocol” and “attr_tissue_preservative” variables into three categories: PolyA Fresh/Frozen (n = 45), total FFPE (n = 38) and Total Fresh/Frozen (n = 69). We preserved in the model the biological variables: “sample type” and “sj_associated_diagnoses_disease_code_curated”. Combat corrected expression data was then used for all downstream analysis.
We calculated gene signature scores for each sample with GSVA (v1.44.2)80 of gene sets related to myogenesis obtained from previous publications (Patel et al., Dev Cell 2022; Danielli et al., Sci Adv 2023; De Micheli et al., Cell Rep 2020; Oprescu et al., iScience 2020; Yohe et al., Sci Transl Med 2018) (Supplementary Data 7). GSVA scores and PRKG1 gene expression (log2CPM) were correlated using Pearson correlation. An unsupervised hierarchical clustering (pheatmap function, euclidean distance, method complete) of GSVA scores and samples showed clustering of biologically similar genesets.
Differential expression analysis was performed between samples with high vs low PRKG1 expression (median log2cpm PRKG1 expression was used as cut-off). An empirical Bayes moderated t-statistics model (limma package v3.52.2)69 was built to detect differentially expressed genes between the studied conditions. Correction for multiple comparisons was performed using FDR. Genes were considered differentially expressed with FDR < 0.05.
Single-nucleus RNA-seq analysis of Rhabdomyosarcoma patient database
Processed single-nucleus RNA-seq Seurat files for individual and integrated patient data were kindly provided by the authors of the original publication (Patel et al. 2022)3. The individual patient data was filtered to discard stromal cells, retaining only tumor cells based on the annotations from the original publication. Tumor cells were re-clustered using Seurat (v5.1.0)81 The re-clustering involved the following steps: setting the default assay to “SCT”, running RunPCA using the variable features of the dataset, identifying neighbors FindNeighbors based on the first 10 principal components, finding clusters FindClusters with a resolution of 0.5, and performing Uniform Manifold Approximation and Projection (UMAP) RunUMAP using the first 10 principal components. All single-nucleus analyses were performed in R (v4.3.1).
Chromatin immunoprecipitation
Cells were treated with 1% formaldehyde for 10 min for cross-linking reaction. A previous incubation with gold-fixative (ref. C01019027, Diagenode) was done for PRKG1 immunoprecipitation to assure fixation. Cross-linking was stopped by adding 500 μl glycine (1.25 M). Cells were resuspended in lysis buffer [0.1% SDS, 0.15 M NaCl, 1% Triton X-100, 1 mM EDTA, 20 mM tris (pH 8), and protease inhibitors (1 mg/ml)] and sonicated with Bioruptor Pico (Diagenode) for 10 cycles until chromatin was sheared to an average fragment length of 200 bp. After centrifugation, a small fraction of eluted chromatin was measured with Qubit. Starting with 30 μg of sample, immunoprecipitation for each antibody was performed overnight; 50 μl of Dynabeads Protein A (Invitrogen) was then added and incubated for 2 h at 4 °C under rotation. Immunoprecipitates were washed with different buffers and the obtained pellets were eluted with 120 μl of a solution of 1% SDS and 0.1 M NaHCO3. Eluted pellets were decrosslinked for 5 h at 65 °C and purified on 50 μl of tris-EDTA buffer with the QIAquick PCR Purification Kit (Qiagen). Differences in the DNA content at each binding region were determined by RT-qPCR as previously described. The reported data represent RT-qPCR values normalized to input DNA and are expressed as percentage (%) of bound/input signal. Primers used for ChIP-qPCR were design according to the peaks of using UCSC genome browser and Primer Three Plus. Primers used for CHIPqPCR are presented in Supplementary Table 4.
Clustering of PRKG1 peaks with epigenomic marks
For visualization of ChIP-seq data from public databases, the UCSC genome browser was used to generate the screenshots presented in this work. Publicly available ChIP-seq tracks from the accession number GSE215202.
Protein extraction
Cultured cells were scraped and collected by centrifugation. For whole cell protein extracts, cells were lysed with Lysis buffer 6 (Biotechne#: 895561) to preserve stability of phosphoproteins.
For protein isolation from tissues, tumor samples were homogenized in 0.5% Igepal, 0.5% sodium deoxycholate 0.1% sodium dodecyl sulfate, 50 mM TRIS-HCl (pH7.5) and 150 mM NaCl with stainless-steel beads (0.2 mm diameter) in the bullet blender at 12000 rpm for 2–5 min until a homogeneous mixture was obtained.
Phosphoproteomic analysis
Two biological replicates of each sample were included. Samples were lysed using BeatBox with 2% SDS, 100 mM DTT, 100 mM Tris/HCl. Digestion was performed following FASP standard protocol with LysC/Trypsin (1 µg/µL, Promega). Digested peptides were cleaned up with MonoSpin C-18 (type S) spin columns and phosphopeptides enriched in an immobilized metal affinity chromatography (MagReSyn® Zr-IMAC HP Zirconium ion (Zr4 + ) functional magnetic microparticles, Resyn biosciences).
Peptide and phosphopeptide samples were dried and reconstituted in 3 % ACN and 1% FA, then loaded onto the EVOTIPs (EV-2013 EVOSEP) following the manufacturer’s instructions. The samples were injected into a nanoLC-MS system: EVOSEP One (EVOSEP, Odense, Denmark) chromatograph connected to an Orbitrap Eclipse Tribrid mass spectrometer (ThermoFisher Scientific, San Jose, CA) via a nanoEasy Spray Source interface with a stainless-steel emitter (EV-1086 EVOSEP). The Evosep One method was 15 SPD (88-minute gradient), and the flow rate 0.22 µL/min with a Dr Maisch C18 AQ, 1.5 µm beads (EV-1137 EVOSEP) analytical column (oven set at 40 °C). The mass spectrometer was operated in DIA mode using 33 variable precursor isolation windows (14.5 to 105.5 m/z) with 0.5 m/z overlap. Full MS1 scans were acquired in the Orbitrap with a scan range of 350 - 1200 m/z and a resolution of 120,000 (at 200 m/z). Automatic gain control (AGC) was set to a target of 1 × 106 and a maximum injection time of 56 ms. MS2 spectra were acquired in the orbitrap at a resolution of 30,000 (at 200 m/z). A higher energy collision induced dissociation (HCD) method (28% NCE) was applied with an AGC target of 5 × 104 and a maximum injection time of 55 ms. Orbitrap Eclipse Tune Application 4.2.431 and Xcalibur version 4.7.69.37 were used to operate the instrument and to acquire data.
Western blotting
Protein extracts were quantified with Bradford Reagent (BioRad), and about 35–50ug of protein extracts were boiled for 3 min at 95 °C prior and loaded in 8–12% polyacrylamide gels for electrophoresis. Proteins were then transferred into nitrocellulose membranes (GE Healthcare Life Science). Membranes were blocked in 3% BSA or 5% milk powder solved in TBS-T (Tris-buffered saline, 0.1% Tween 20) for 1 h. In general, membranes were incubated overnight at 40C with the primary antibodies and 1 h at room temperature with fluorescently labeled secondary antibodies. Fluorescence was detected using LI-COR Odyssey Classic Infrared Imaging System (LI-COR Inc.). Relative levels of protein expression were measured by Image J software.
Fluorescent westerns were imaged using the Odyssey Infrared Imaging System (LI-COR Biosciences). Secondary antibodies were IRDye 680LT donkey anti-rabbit (925-68023), IRDye 800CW donkey anti-mouse (926-32212), IRDye 800 CW goat anti-rabbit IgG (926-32211), and IRDye 680RD goat anti-mouse IgG (926-68070). For chemiluminescent detection, secondary HRP-conjugated antibodies were obtained from Dako and Immobilon Western Chemiluminescent HRP substrate for detection. All the antibodies used in this work are presented in Supplementary Table 5.
Immunofluorescence (IF) on primary samples
RMS samples were embedded in OCT and placed into slides for fixation with 4% PFA for 10 min. RMS adherent cells were grown over a slide and fixed with 4% formaldehyde for 15 min at 37 ºC. Before permeabilization, cells were labeled with Wheat Germ Agglutinin conjugated to Alexa Fluor® 647 at a concentration of 5.0 μg/ml. After 10 min of incubation at room temperature, cells were washed and permeabilized with 0.1% Triton X-100. Then, cells were incubated with blocking solution (3% BSA) for 30 min. PRKG1 was used as primary antibody for 1 h. TNNI1 as primary antibody was incubated ON at 4 ºC. Cells were washed and labeled with a secondary antibody, Alexa Fluor® 488 goat anti-rabbit (1:400). Samples were incubated with secondary labeled antibodies for 1 h at room temperature.
Immunohistochemistry (IHC)
First PDX engraftments as well as successive mouse-to-mouse transplantation tumors were characterized by immunohistochemical detection of MyoD1 (1:20, M3512; DAKO Agilent Technologies, Santa Clara, CA, USA) and anti-human nuclei antigen (1:200, MAB4383; Merck Millipore, Burlington, MA, USA), and counterstained with hematoxylin and eosin. Pharmacodynamic analyses were performed by immunohistochemistry detection of pS6 (Ser240/244) (1:1000) and Cleaved-PARP (1:50, #5625S, Cell signaling). Briefly, tumor samples were fixed in 10% formalin and embedded in paraffin. Paraffin block samples were cut into sections of 3 μm with the microtome and deparaffined at 60 ºC for 45 min. The rehydration and then xylol and ethanol standard baths protocol was followed by the antigen retrieval was heat-induced in sodium citrate (pH 6.0) buffer and subsequently, slides were inhibited endogenous peroxidase action with peroxide de hydrogen y 0.1% azide sodic, following manufacturer’s instructions. The staining was carried out using the corresponding primary antibodies. The DAB (Polymer) Kit (Buffer + Chromogen) Novocastra (Leica Biosystems) was used Slides were finally cover slipped with dibutyl phthalate polystyrene xylene (DPX) and dried at room temperature.
In vivo experiments
Ethics and animal experimentation
All in vivo studies were performed according to the Institutional Animal Research Ethics Committee and European guidelines (EU Directive 2010/63/EU) on the principle of the 3 R. That is to Replace, Reduce and Refine the use of animals for scientific purposes. Work with mice adhered to the European regulations and was approved by Comitè Ètic d’Experimentació Animal (CEEA) de la Universitat de Barcelona (animal protocol number 293/19_10964). Mice were housed under specific pathogen-free (SPF) conditions, with food and water provided ad libitum. Tumor burden was monitored three or more times per week. Animals were euthanized when the tumor volume exceeded 1500 mm3 or if body weight loss was more than 20% of the starting weight. In a limited number of cases, tumor volume slightly exceeded the 1,500 mm³ limit between scheduled measurements due to the rapid exponential growth phase characteristic of these models. Upon detection, animals were euthanized according to humane endpoint criteria, and all procedures were conducted under continuous veterinary supervision. For all xenograft experiments, NOD SCID (NOD.CB17-Prkdc<scid > /J; ENVIGO) mice (mainly female) aged 4–6 weeks at the time of injection were used. The number of animals per group (n) is indicated in the figure legends and is consistently between 5 and 10 animals per group. Sex was not considered in the study design or analysis. This decision was based on previous studies showing no sex-dependent differences in tumor growth in this model.
NOD-SCID mice were subcutaneously implanted in both flanks with 3 × 3 mm3 fresh PDX tumor. PDXs HSJD- ARMS-001, ARMS-006, ARMS-007, ARMS-010, ERMS-011 and ERMS-024 PDX models were generated from primary tumor biopsies. The specific number of animals in each experimental group is indicated in the graphs. Tumor volume was calculated as follows: (longer measurement x (shorter measurement) ^2) / 2.
Antitumoral activity of ipatasertib (GDC-0068)
Ipatasertib was diluted in water and freshly prepared every day before administration. When tumor volumes reached 150–350 mm3, mice were randomized into different groups (5–10 animals per group) and divided into four different groups: saline (control), ipatasertib (100 mg/kg), ipatasertib (50 mg/kg) and ipatasertib (25 mg/kg) when applicable. Ipatasertib treatments or vehicle control were orally administrated five times per week with 2 days off each week, during four consecutive weeks, (dx5) x4.
The regimen of 100 mg/kg of ipatasertib group was the Maximum Tolerated Dose (MTD). Note that doses of 25 or 50 mg/kg, below the MTD, were only included in graphs of dose-dependent effect.
Survival curves of mice bearing HSJD-ARMS-010 and ERMS-024 PDX, treated with 100 mg/kg or 25 mg/kg of ipatasertib were calculated by Kaplan-Meier Method. Survival event was reported at 1500 mm3 tumor size. Log-rank statistics with Bonferroni correction test was used to compare statistically significant between treatment groups, performed with GraphPad Prism 8 software.
Antitumoral activity of miransertib (ARQ-092.2MSA)
To test miransertib (ARQ-092) efficacy on tumoral reduction, in vivo experiments were accomplished following the given instructions: a solution of 10 mg/ml of ARQ-092.2MSA was prepared in 0.01 M Phosphoric Acid and freshly prepared every day before administration. The compound amount weigh was corrected by a Potency Correction Factor (Free Base content) of 0.649. To get a dose of 100 mg/kg, 200 µl were administrated for a mouse of 20 g weigh by oral gavage. This volume was corrected by the animal weight. Tumor volume criteria were the same as for ipatasertib treated mice.
Here we present data from an HSJD-ERMS-011 model. When tumor volumes reached 150–450 mm3, mice were randomized into five groups (“n” range between 4–9 animals per group): (i) Only water (vehicle), (ii)0.01 M Phosphoric Acid (vehicle), (iii) miransertib solved in 0.01 M Phosphoric Acid and (iv) two groups of ipatasertib at high and low doses solved in water. The treatment was started in a regimen of 5 days on and 2 days-off during 2 weeks for all groups to compare toxicities and responses. However, due to the high toxicities seen in ARQ-092 after 10 doses, we switched the administration protocol to 3 days off between cycles of ARQ-0.92-phosphoric acid to avoid this high toxicity.
Antitumoral activity of trametinib
E001_s cells were suspended in 1:1 TSM - Matrigel (Corning® Basement Membrane Matrix, Cultek) and these suspensions were inoculated in the flank of the mouse. When tumor volumes reached 150–350 mm3, mice were divided into four different groups: vehicle (control); ipatasertib (100 mg/kg); ipatasertib (100 mg/kg) plus trametinib (3 mg/Kg) and trametinib (3 mg/kg). Ipatasertib was administrated at 100 mg/kg diluted in water and freshly prepared every day before administration. Trametinib was administrated as suspension in vehicle (0.5% hydroxypropylmethylcellulose/0.2% Tween 80/5% sucrose). It was prepared weekly and stored at 4 °C until use. In the ipatasertib + trametinib group, independent drug administration was separated by at least 1 h.
Pharmacokinetic analysis: ipatasertib methodology detection
To investigate the (PK) of ipatasertib, we used female nod-scid mice bearing bilateral subcutaneous HSJD-ARMS-006 and HSJD-ERMS-011 PDX. Mice received ipatasertib oral administration, at a dose of 100 mg/kg or 25 mg/kg, when tumor reached >250mm3. Blood and tumor samples were collected at 0, 1, 3, 8, and 24 h after a single dose of ipatasertib. Blood samples (approximately 800 μL) were collected from each animal at the scheduled sample collection time by terminal cardiac puncture into tubes containing Heparin as an anticoagulant and centrifuged at 1500–2000g to isolate plasma. In each time condition, 4 animals were included for each dose tested, therefore, 8 tumors and 4 plasma samples were analyzed for each condition and time. A total of 32 animals were included for the ERMS-011 PK and 25 animals for the PDX HSJD-ARMS-006 PK. Plasma and tumor tissues were snap-frozen in liquid nitrogen and stored at −80 °C.
The concentration of ipatasertib in each plasma ant tumor sample was determined by a internally validated Liquid Chromatography with tandem mass spectrometry (LC-MS-MS) assay in the Department of Pharmacology, Therapeutics and Toxicology, Universitat Autònoma de Barcelona (UAB). For PK studies of a small molecule, such as ipatasertib, using an LC-MS/MS instrument provides much higher sensitivity and specify than UV detectors commonly used on HPLC units. Chromatography was performed using Zorbax Eclipse Plus C18 (2.1 × 50, 1.8 µm) (Agilent Technologies) with a mobile phase of 0.1% formic acid in acetonitrile and with a flow rate of 0.4 ml/min. Lincomycin was used as internal standard. The drug and the internal standard were extracted by liquid–liquid extraction and analyzed by mass spectrometry The Autosampler model Agilent 1260 Infinity together with the detector model Agilent 6420 mass spectrometer were used for the analysis.
Phosphoproteomic data analysis
Database searches
The spectrum files were analyzed in DirectDIA mode using the Spectronaut software from Biognosys (v. 19.0.240604.62635)82, searching against the reviewed Uniprot database for Homo sapiens (SwissProt Human, released on 2024/01) and common contaminants83. Oxidation in methionine, acetylation, methionine excision in protein N-terminal and phosphorylation in STY (only for phosphopeptide samples) were set as variable modifications. Carbamidomethylation in cysteine was set as a fixed modification. The maximum number of tryptic missed cleavages and the maximum number of variable modifications allowed were set to 2 and 5, respectively. The m/z range for the precursors and the fragment ions were set to 350—1200 and 200—3000, respectively. The remaining Spectronaut parameters were kept at their default values. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE84 partner repository with the dataset identifier PXD064262.
Statistical analysis
We preprocessed, visualized and statistically analyzed Spectronaut’s search output at the precursor and p-sites levels using the R programming language85. For proteome analysis, we dropped precursors with elution group Q-value or protein group Q-value above 0.01 before using the package diann-rpackage86 to generate log2-transformed PG-level MaxLFQ quantifications87. For the phosphoproteome analysis, we used the p-site level table from Spectronaut. Then, we perform an equivalent analysis in parallel for proteome and phosphoproteome. We dropped contaminant protein groups and normalized datasets using the cyclic LOESS algorithm implementation included in the limma R package69,88. For protein groups we requested at least two different quantifying peptides per protein group. For protein groups and p-sites we demand full observations in at least one group and impute the missing values with normally distributed random numbers with mean μimp and standard deviation σimp, where μimp = μdata - 1.8 σdata and σimp = 0.3 σdata (μdata and σdata are mean and the standard deviation of the original data). We performed the subsequent differential abundance analysis at the protein group and p-site level leveraging the limma R package68. Comparison between groups was done (ipa vs Vehicle and Mira vs Vehicle) to find out changes. For each comparison, we calculated estimated log2 fold changes (logFC) from the mean of each replica. Finally, we applied standard cutoffs for the fold change ( | FC | > 1.5) and the p value (p < 0.05) to define significant protein groups and p-sites.
Statistical analysis and reproducibility
Statistical analyses were performed with R 4.0.3 and GraphPad Prism 8.0. Variable slope (four parameters) curves were used to determine the IC50 value using Prism 8 Software (GraphPad). Student’s t test and Mann-Whitney t test (for nonnormally distributed data) were used for nonpaired comparisons of two groups. Wilcoxon signed rank test was used for non-normally distributed data for paired comparison. Survivals were calculated using the Kaplan-Meier method, and curves were compared using the log-rank test, with Bonferroni correction. A P value < 0.05 was considered significant.
Western blot experiments were independently repeated at least three times with similar results. Representative blots are shown. For xenograft experiments, all collected tumors were analyzed by immunohistochemistry (IHQ) for the indicated protein, and the image shown corresponds to a representative tumor displaying the observed staining pattern. Results were reproducible across biological replicates. The number of tumors per condition (n) is indicated in the graphs and figure legends.
Ethics statement
The Hospital Sant Joan de Deu IRB Medical Council approved an informed consent for all excedent of clinical and biological material to be used and collected for research purposes (Forms 1608/09). This consent underwent ethical review and was approved following the Spanish National Biomedical Law from 2007 (article 69.6 Law 14/2007). All legal guardians sign the informed consent at Sant Joan de Déu Hospital (SJD, Barcelona, Spain) for all the samples stored at the institutional biobank. Furthermore, an extra written informed consent to publish identifiable data from the case report was obtained from the patient’s legal guardian. Embryos included in this study were derived from spontaneous abortions and were collected by the institutional Tissue Bank under the above-mentioned informed consent. No compensation is involved in tissue donations from participants of the HSJD Tissue Bank.
Work with mice adhered to the European regulations and was approved by Comitè Ètic d’Experimentació Animal (CEEA) de la Universitat de Barcelona (animal protocol number 293/19_10964).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The expression data generated for this study have been deposited in the GEO database (NCBI Gene Expression Omnibus) under accession code GSE268193 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE268193) which contains three subseries: The RNA-seq-based profiling of the PRKG1-depleted RMS cells is available from GSE268190, (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi? acc=GSE268190). Expression profiling by array (Clariom_S_Human) from Rhabdomyosarcoma human cells treated with ipatasertib or miransertib is available from GSE268191. (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE 268191) Expression profiling by array (Human Genome U219 Array Plate) of pediatric rhabdomyosarcoma tumors and their corresponding subcutaneous patient derived xenograft is available from GSE268192 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE268192). The RNAseq publicly available data used in this study are available in the St. Jude Cloud database (https://www.stjude.cloud). The dataset used to obtain the myogenesis gene set was derived from previously published studies: Patel et al.3, Danielli et al.16, De Micheli et al.89, Oprescu et al.90, Yohe et al.18. The LC-MS data have been deposited in the ProteomeXchange Consortium under the accession code PXD064262 (https://www.ebi.ac.uk/pride/archive/projects/PXD064262). The publicly available ChIP-seq tracks used are available in the accession number GSE215202 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE215202). The remaining data are available within the Article, Supplementary Information or Source Data file. The raw numbers for charts and graphs are available in the Source Data file. Source data are provided with this paper.
References
Skapek, S. X. et al. Rhabdomyosarcoma. Nat. Rev. Dis. Prim. 5, 1 (2019).
Hettmer, S. & Wagers, A. J. Muscling in: uncovering the origins of rhabdomyosarcoma. Nat. Med 16, 171–173 (2010).
Patel, A. G. et al. The myogenesis program drives clonal selection and drug resistance in rhabdomyosarcoma. Dev. Cell 57, 1226–1240.e1228 (2022).
Sebire, N. J. & Malone, M. Myogenin and MyoD1 expression in paediatric rhabdomyosarcomas. J. Clin. Pathol. 56, 412–416 (2003).
Davicioni, E. et al. Identification of a PAX-FKHR gene expression signature that defines molecular classes and determines the prognosis of alveolar rhabdomyosarcomas. Cancer Res 66, 6936–6946 (2006).
Williamson, D. et al. Fusion gene-negative alveolar rhabdomyosarcoma is clinically and molecularly indistinguishable from embryonal rhabdomyosarcoma. J. Clin. Oncol. 28, 2151–2158 (2010).
Missiaglia, E. et al. PAX3/FOXO1 fusion gene status is the key prognostic molecular marker in rhabdomyosarcoma and significantly improves current risk stratification. J. Clin. Oncol. 30, 1670–1677 (2012).
Oberlin, O. et al. Prognostic factors in metastatic rhabdomyosarcomas: results of a pooled analysis from United States and European cooperative groups. J. Clin. Oncol. 26, 2384–2389 (2008).
Joshi, D. et al. Sarcoma committee of the children’s oncology, age is an independent prognostic factor in rhabdomyosarcoma: a report from the soft tissue sarcoma committee of the children’s oncology group. Pediatr. blood cancer 42, 64–73 (2004).
Shern, J. F. et al. Genomic classification and clinical outcome in rhabdomyosarcoma: a report from an international consortium. J. Clin. Oncol. 39, 2859–2871 (2021).
Hawkins, D. S. et al. Addition of vincristine and irinotecan to vincristine, dactinomycin, and cyclophosphamide does not improve outcome for intermediate-risk rhabdomyosarcoma: a report from the Children’s Oncology Group. J. Clin. Oncol. 36, 2770–2777 (2018).
Mascarenhas, L. et al. Randomized phase II trial of bevacizumab or temsirolimus in combination with chemotherapy for first relapse rhabdomyosarcoma: a report from the Children’s Oncology Group. J. Clin. Oncol. 37, 2866–2874 (2019).
Geoerger, B. et al. Phase II trial of temsirolimus in children with high-grade glioma, neuroblastoma and rhabdomyosarcoma. Eur. J. Cancer 48, 253–262 (2012).
Gupta A. A. et al. Addition of temsirolimus to chemotherapy in children, adolescents, and young adults with intermediate-risk rhabdomyosarcoma (ARST1431): a randomised, open-label, phase 3 trial from the Children’s Oncology Group. Lancet Oncol. 7, 912–921 (2024).
Wei, Y. et al. Single-cell analysis and functional characterization uncover the stem cell hierarchies and developmental origins of rhabdomyosarcoma. Nat. Cancer 3, 961–975 (2022).
Danielli, S. G. et al. Single-cell profiling of alveolar rhabdomyosarcoma reveals RAS pathway inhibitors as cell-fate hijackers with therapeutic relevance. Sci. Adv. 9, eade9238 (2023).
DeMartino, J. et al. Single-cell transcriptomics reveals immune suppression and cell states predictive of patient outcomes in rhabdomyosarcoma. Nat. Commun. 14, 3074 (2023).
Yohe, M.E. et al. MEK inhibition induces MYOG and remodels super-enhancers in RAS-driven rhabdomyosarcoma. Sci Transl Med 10, eaan4470 (2018).
Pomella, S. et al. Interaction between SNAI2 and MYOD enhances oncogenesis and suppresses differentiation in fusion negative rhabdomyosarcoma. Nat. Commun. 12, 192 (2021).
Laubscher, D. et al. BAF complexes drive proliferation and block myogenic differentiation in fusion-positive rhabdomyosarcoma. Nat. Commun. 12, 6924 (2021).
Renshaw, J. et al. Dual blockade of the PI3K/AKT/mTOR (AZD8055) and RAS/MEK/ERK (AZD6244) pathways synergistically inhibits rhabdomyosarcoma cell growth in vitro and in vivo. Clin. Cancer Res 19, 5940–5951 (2013).
Shimizu, T. et al. The clinical effect of the dual-targeting strategy involving PI3K/AKT/mTOR and RAS/MEK/ERK pathways in patients with advanced cancer. Clin. Cancer Res 18, 2316–2325 (2012).
Petricoin, E. F. 3rd et al. Liotta, Phosphoprotein pathway mapping: Akt/mammalian target of rapamycin activation is negatively associated with childhood rhabdomyosarcoma survival. Cancer Res 67, 3431–3440 (2007).
Shern, J. F. et al. Comprehensive genomic analysis of rhabdomyosarcoma reveals a landscape of alterations affecting a common genetic axis in fusion-positive and fusion-negative tumors. Cancer Discov. 4, 216–231 (2014).
Kohsaka, S. et al. A recurrent neomorphic mutation in MYOD1 defines a clinically aggressive subset of embryonal rhabdomyosarcoma associated with PI3K-AKT pathway mutations. Nat. Genet 46, 595–600 (2014).
McKinnon T. et al. Functional screening of FGFR4-driven tumorigenesis identifies PI3K/mTOR inhibition as a therapeutic strategy in rhabdomyosarcoma. Oncogene. 20, 2630–2644 (2018).
Cen, L. et al. PDK-1/AKT pathway as a novel therapeutic target in rhabdomyosarcoma cells using OSU-03012 compound. Br. J. Cancer 97, 785–791 (2007).
Codenotti, S. et al. Hyperactive Akt1 signaling increases tumor progression and DNA repair in embryonal rhabdomyosarcoma RD line and confers susceptibility to glycolysis and mevalonate pathway inhibitors. Cells 11, 2859 (2022).
Manzella, G. et al. Phenotypic profiling with a living biobank of primary rhabdomyosarcoma unravels disease heterogeneity and AKT sensitivity. Nat. Commun. 11, 4629 (2020).
Fruman, D. A. & Rommel, C. PI3K and cancer: lessons, challenges and opportunities. Nat. Rev. Drug Discov. 13, 140–156 (2014).
Savill, K. M. Z. et al. Distinct resistance mechanisms arise to allosteric vs. ATP-competitive AKT inhibitors. Nat. Commun. 13, 2057 (2022).
Schmid, P. et al. First-Line Ipatasertib, Atezolizumab, and Taxane Triplet for Metastatic Triple-Negative Breast Cancer: Clinical and Biomarker Results. Clin. Cancer Res 30, 767–778 (2024).
Saura, C. et al. I Study of the ATP-Competitive AKT Inhibitor Ipatasertib Demonstrates Robust and Safe Targeting of AKT in Patients with Solid Tumors. Cancer Discov. 7, 102–113 (2017).
Oliveira, M. et al. FAIRLANE, a double-blind placebo-controlled randomized phase II trial of neoadjuvant ipatasertib plus paclitaxel for early triple-negative breast cancer. Ann. Oncol. 30, 1289–1297 (2019).
Shapiro, G. I. et al. A phase Ib open-label dose escalation study of the safety, pharmacokinetics, and pharmacodynamics of cobimetinib (GDC-0973) and ipatasertib (GDC-0068) in patients with locally advanced or metastatic solid tumors. Invest N. Drugs 39, 163–174 (2021).
Yu, Y. et al. Targeting AKT1-E17K and the PI3K/AKT Pathway with an Allosteric AKT Inhibitor, ARQ 092. PLoS One 10, e0140479 (2015).
Keppler-Noreuil, K. M. et al. Pharmacodynamic Study of Miransertib in Individuals with Proteus Syndrome. Am. J. Hum. Genet 104, 484–491 (2019).
You, I. et al. Discovery of an AKT Degrader with Prolonged Inhibition of Downstream Signaling. Cell Chem. Biol. 27, 66–73.e67 (2020).
McLeod, Gout et al. Jude Cloud-a Pediatric Cancer Genomic Data Sharing Ecosystem. Cancer Discov 11, 1082–1099 (2021).
Traag, V. A., Waltman, L. & van Eck, N. J. From Louvain to Leiden: guaranteeing well-connected communities. Sci. Rep. 9, 5233 (2019).
Dippold, R. P. & Fisher, S. A. Myosin phosphatase isoforms as determinants of smooth muscle contractile function and calcium sensitivity of force production. Microcirculation 21, 239–248 (2014).
Schwappacher, R. et al. Novel crosstalk to BMP signalling: cGMP-dependent kinase I modulates BMP receptor and Smad activity. EMBO J. 28, 1537–1550 (2009).
Bois, P. R., Brochard, V. F., Salin-Cantegrel, A. V., Cleveland, J. L. & Grosveld, G. C. FoxO1a-cyclic GMP-dependent kinase I interactions orchestrate myoblast fusion. Mol. Cell Biol. 25, 7645–7656 (2005).
Adler, J., Kuret, A., Langst, N. & Lukowski, R. Targets of cGMP/cGKI in Cardiac Myocytes. J. Cardiovasc Pharm. 75, 494–507 (2020).
Ranek, M. J. et al. PKG1-modified TSC2 regulates mTORC1 activity to counter adverse cardiac stress. Nature 566, 264–269 (2019).
Schall, N. et al. Protein kinase G1 regulates bone regeneration and rescues diabetic fracture healing. JCI Insight 5, e135355 (2020).
Leung, E. L., Wong, J. C., Johlfs, M., Tsang, B. K. & Fiscus, R. R. Protein kinase G type Ialpha activity in human ovarian cancer cells significantly contributes to enhanced Src activation and DNA synthesis/cell proliferation. Mol. Cancer Res 8, 578–591 (2010).
Wilson, T. J., Zamler, D. B., Doherty, R., Castro, M. G. & Lowenstein, P. R. Reversibility of glioma stem cells’ phenotypes explains their complex in vitro and in vivo behavior: Discovery of a novel neurosphere-specific enzyme, cGMP-dependent protein kinase 1, using the genomic landscape of human glioma stem cells as a discovery tool. Oncotarget 7, 63020–63041 (2016).
Fallahian, F., Karami-Tehrani, F., Salami, S. & Aghaei, M. Cyclic GMP induced apoptosis via protein kinase G in oestrogen receptor-positive and -negative breast cancer cell lines. FEBS J. 278, 3360–3369 (2011).
Wong, J. C., Bathina, M. & Fiscus, R. R. Cyclic GMP/protein kinase G type-Ialpha (PKG-Ialpha) signaling pathway promotes CREB phosphorylation and maintains higher c-IAP1, livin, survivin, and Mcl-1 expression and the inhibition of PKG-Ialpha kinase activity synergizes with cisplatin in non-small cell lung cancer cells. J. Cell Biochem 113, 3587–3598 (2012).
Phelps, M. P., Bailey, J. N., Vleeshouwer-Neumann, T. & Chen, E. Y. CRISPR screen identifies the NCOR/HDAC3 complex as a major suppressor of differentiation in rhabdomyosarcoma. Proc. Natl. Acad. Sci. USA 113, 15090–15095 (2016).
Cao, L. et al. Genome-wide identification of PAX3-FKHR binding sites in rhabdomyosarcoma reveals candidate target genes important for development and cancer. Cancer Res 70, 6497–6508 (2010).
Gryder, B. E. et al. Myogenic super enhancers and confers BET bromodomain vulnerability. Cancer Discov. 7, 884–899 (2017).
Dumble, M. et al. Discovery of novel AKT inhibitors with enhanced anti-tumor effects in combination with the MEK inhibitor. PLoS One 9, e100880 (2014).
Isakoff, S. J. et al. Antitumor activity of ipatasertib combined with chemotherapy: results from a phase Ib study in solid tumors. Ann. Oncol. 31, 626–633 (2020).
Kim, S. B. et al. investigators, Ipatasertib plus paclitaxel versus placebo plus paclitaxel as first-line therapy for metastatic triple-negative breast cancer (LOTUS): a multicentre, randomised, double-blind, placebo-controlled, phase 2 trial. Lancet Oncol. 18, 1360–1372 (2017).
Dent, R. et al. investigators, Final results of the double-blind placebo-controlled randomized phase 2 LOTUS trial of first-line ipatasertib plus paclitaxel for inoperable locally advanced/metastatic triple-negative breast cancer. Breast Cancer Res Treat. 189, 377–386 (2021).
Shi, Z. et al. Functional mapping of AKT signaling and biomarkers of response from the FAIRLANE trial of neoadjuvant ipatasertib plus paclitaxel for triple-negative breast cancer. Clin. Cancer Res 28, 993–1003 (2022).
O'Donnell, J. et al. Ipatasertib exhibits anti-tumorigenic effects and enhances sensitivity to paclitaxel in endometrial cancer in vitro and in vivo. Int J Oncol 63, 103 (2023).
Lin, J. et al. Targeting activated Akt with GDC-0068, a novel selective Akt inhibitor that is efficacious in multiple tumor models. Clin. Cancer Res 19, 1760–1772 (2013).
Yoshida, K. et al. Population pharmacokinetics of ipatasertib and its metabolite in cancer patients. J. Clin. Pharmacol. 61, 1579–1591 (2021).
Nair, A. B. & Jacob, S. A simple practice guide for dose conversion between animals and human. J. Basic Clin. Pharm. 7, 27–31 (2016).
Slotkin, E. K. et al. Patient-driven discovery, therapeutic targeting, and post-clinical validation of a novel AKT1 fusion-driven cancer. Cancer Discov. 9, 605–616 (2019).
Meel, M. H. et al. Culture methods of diffuse intrinsic pontine glioma cells determine response to targeted therapies. Exp. Cell Res. 360, 397–403 (2017).
Schrödinger Release 2024-4: Maestro, Schrödinger (LLC, New York, NY, 2025).
Friesner, R. A. et al. Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J. Med. Chem. 47, 1739–1749 (2004).
Irizarry R. A., Ooi S. L., Wu Z. & Boeke J. D. Use of mixture models in a microarray-based screening procedure for detecting differentially represented yeast mutants. Stat. Appl. Genet. Mol. Biol. 2, 1 (2003).
Irizarry, R. A. et al. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4, 249–264 (2003).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B 57, 289–300 (1995).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. ClusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Dobin, Davis, A. “STAR: Ultrafast Universal RNA-Seq Aligner.” Bioinformatics (Oxford, England) 29,15–21. (2013).
Liao, Y., Smyth, G.K. & Shi, W. “featurecounts: an efficient general purpose program for assigning sequence reads to genomic features.”. Bioinforma. (Oxf., Engl.) 30, 923–930 (2014).
Robinson, M. D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 11, R25 (2010).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. “edgeR: a bioconductor package for differential expression analysis of digital gene expression data”. Bioinforma. (Oxf., Engl.) 26, 139–140 (2012).
Liberzon, A. The molecular signatures database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Köhler, S., Gargano, M. & Matentzoglu, N. “The human phenotype ontology in 2021.”. Nucleic Acids Res. 49, D687–D692 (2021).
Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E. & Storey, J. D. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28, 882–883 (2012).
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-Seq data. BMC Bioinforma. 14, 7 (2013).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2024).
Software. In: Biognosys. https://biognosys.com/software/spectronaut/ (2021).
Frankenfield, A. M., Ni, J., Ahmed, M. & Hao, L. Protein contaminants matter: building universal protein contaminant libraries for DDA and DIA proteomics. J. Proteome Res. 21, 2104–2113 (2022).
Perez-Riverol, Y. et al. The PRIDE database at 20 years: 2025 update. Nucleic Acids Res. 53, D543–D553 (2025).
The R Project for Statistical Computing. https://www.R-project.org/ (2023).
GitHub - vdemichev/diann-rpackage: report processing and protein quantification for MS-based proteomics. In: GitHub https://github.com/vdemichev/diann-rpackage (2023).
Cox, J. et al. Accurate proteome-wide label-free quantification by delayed normalization and maximal peptide ratio extraction, termed MaxLFQ. Mol. Cell Proteom. 13, 2513–2526 (2014).
Bolstad, B. M., Irizarry, R. A., Astrand, M. & Speed, T. P. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19, 185–193 (2003).
De Micheli, A. J. et al. Single‑cell analysis of the muscle stem cell hierarchy identifies heterotypic communication signals involved in skeletal muscle regeneration. Cell Rep. 30, 3583–3595 (2020).
Oprescu, S. N., Yue, F., Qiu, J., Brito, L. F. & Kuang, S. Temporal dynamics and heterogeneity of cell populations during skeletal muscle regeneration. iScience 23, 100993 (2020).
Acknowledgements
The authors thank the support of the parents and families of children with rhabdomyosarcoma, especially the family of the index case who provided all consents to specifically describe the clinical course of the disease; and the Asociación contra el Rabdomiosarcoma, Cancer del Desarrollo. We are grateful to the Band of Parents at Hospital Sant Joan de Déu for supporting the overall research activities of the Cancer Pediatrics Group (IRSJD PCCB). The authors thank the Xarxa de Bancs de Tumors de Catalunya (XBTC; sponsored by Pla Director d’Oncologia de Catalunya), Dr. Martinez-Tirado for providing the commercial cell lines used in this study and Dr. Anand Patel for providing the Seurat objects of single-nucleus RNA-seq data from the RMS sample included in this study. This work has been awarded the 37th Schweisguth Prize from the International Society of Pediatric Oncology (SIOP), Hawaii, October 2024. Funding: This Project was funded by “Association against Rhabdomyosarcoma from HSJD” (Asociación Contra el Rabdomiosarcoma, Cancer del Desarrollo). (https://curarelrabdo.org/) PT ackowledges funding from PFIS predoctoral fellowship from the Instituto de Salud Carlos III (ISCIII) [grant number FI21/00047]. AA acknowledges funding from the European Research Council [grant number ERC-StG 101076506] and the Ramón y Cajal fellowship [grant number RYC2019-027738-I]. This project was supported by the PID2020-118241RA-I00 (RHABDOSEQ) project, and an FPI PhD fellowship contract for ACX [grant number PRE2021-098532], both funded by the Spanish Ministry of Science and Innovation (MCIN). This work has also been supported by the XXV FERO award (Fundación FERO) granted to AA. IHM acknowledges funding from Instituto de Salud Carlos III (ISCIII, co-funded by the European Union; PI22/00364) and from La Marató TV3 Foundation (201928).
Author information
Authors and Affiliations
Contributions
E.P.: Conceptualization, Methodology, Validation, Formal analysis, Investigation, Writing- Original draft preparation. P.T.: Methodology, Investigation. E.A.: Methodology, Investigation. S.G.G.: Software, Formal analysis, Computing Resources S.M.L.: Methodology, Investigation. A.C.X.: Software, Investigation. P.B.M.: Software, Formal analysis, Computing Resources. J.P.B.: Software, Formal analysis, Computing Resources. J.A.P.: Software, Formal analysis, Computing Resources. S.S.O.: Software, Formal analysis, Computing Resources. L.D.: Software, Formal analysis, Computing Resources. M.G.: Software, Formal analysis, Computing Resources. I.F.C.: Software, Formal analysis, Computing Resources. A.O.: Methodology, Investigation. M.V.: Computing Resources. Q.R.: Software, Formal analysis, Computing Resources. L.G.-G.: Conceptualization, Visualization. E.R.: Investigation, Validation. C.R.: Validation. G.R.: Conceptualization, A.M.C.: Resources. A.A.: Conceptualization, Computing Resources. I.H.M.: Conceptualization, Resources, Writing-Original Draft, Visualization, Supervision, Project administration. J.M.: Conceptualization, Resources, Writing-Original Draft, Supervision, Project administration, Funding acquisition.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Prada, E., Táboas, P., Andrades, E. et al. PRKG1 hinders myogenic differentiation and predicts response to AKT inhibitor ipatasertib in Rhabdomyosarcoma. Nat Commun 16, 9816 (2025). https://doi.org/10.1038/s41467-025-64783-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-64783-3