Abstract
Many non-small cell lung cancer (NSCLC) patients remain unresponsive to the current standard of care, which includes chemotherapy and immune checkpoint inhibitors, like anti-PD-1/PD-L1 antibodies. While interleukin (IL)-1β is known to promote lung cancer growth in humans and mice, we show here that IL-1β administration or overexpression overcomes resistance to classical chemo-immunotherapy (cisplatin/pemetrexed/anti-PD-1) in mouse lung cancer models. The antitumor effects of IL-1β rely on cancer cell-derived CXCL10 which mediates CD8 T cell recruitment at the tumor site. In lung cancer cells, Thioredoxin Interacting Protein (TXNIP) induces mitochondrial DNA (mtDNA) release in the cytosol, activating Absence in Melanoma 2 (AIM2) inflammasome, which subsequently triggers IL-1β and CXCL10 secretion, thereby reversing chemo-immunotherapy resistance. The clinical relevance of our findings is supported by the transcriptomic analysis of patient tumors, showing that high expression of IL1B, IL1R1, AIM2 and/or TXNIP is associated with better response to immunotherapy in NSCLC patients. Additionally, drug screening identifies MEK and MDM2 inhibitors as inducers of TXNIP expression capable of reversing resistance to chemo-immunotherapy. This study highlights a positive role of IL-1β in lung cancer treatment and suggests that enhancing IL-1β production at the tumor site can overcome resistance to chemo-immunotherapy.
Similar content being viewed by others
Introduction
Non-small cell lung cancer (NSCLC) is the most frequent thoracic cancer. In the absence of oncogenic addiction, the treatment of metastatic disease relies on the association of chemotherapy and immune checkpoint inhibitors (ICIs) targeting PD-1 and PD-L1. The association of anti-PD-1/PD-L1 monoclonal antibodies (mAb) with platinum-based doublet chemotherapy is now standard of care in all-comer metastatic NSCLC patients in first line, and is proposed to counter resistance to ICI monotherapies1,2. The rationale for such an association is based notably on the capacity of chemotherapy to promote immunogenic cancer cell death, which is able to stimulate an immune response against cancer cells and consequently, turn “cold” tumors (with little immune infiltration) into “hot” tumors (with greater T cell infiltration), more prone to respond to checkpoint inhibitors3,4. However, while the association of cisplatin and pemetrexed with an anti-PD-L1 antibody demonstrates efficacy, only 50% of patients gain benefit from such combotherapy, and most patients finally escape this regimen5. Finding markers to predict resistance and identifying new treatments to fight resistance to chemo-immunotherapy are unmet needs.
The IL-1 family is composed of 11 cytokines. The main members, IL-1α, IL-1β, and IL-1 receptor antagonist (IL-1RA), are the most widely studied. IL-1α and IL-1β bind the type 1 receptor (IL-1R1). After ligation, IL-1α and IL-1β induce transcription of several genes involved in inflammatory and immune pathways. In the context of cancer, both positive and negative effects on tumor growth have been described. IL-1β has different influences on cancer occurrence, cancer aggressiveness (by regulating angiogenesis and metastasis) and response to treatment, thus leading to controversial conclusions. These contrasting results could also be explained by the level of IL-1β produced, the type of producing cells, the microenvironment (immune cells or fibroblasts), the stage of cancer, and the anti-cancer treatments used6,7.
In the context of lung cancer, IL-1β is described as a pro-tumoral cytokine. Increased levels of IL-1β at the tumor site favor angiogenesis and induction of Epithelial to Mesenchymal Transition (EMT)8,9,10,11. Deficiency of Il1b in mice inhibits tumor growth of lung and other types of mouse subcutaneously (s.c.) implanted cancers12. Moreover, air pollutants cause the release of IL-1β by lung macrophages, and this process results in the proliferation of progenitor-like EGFR mutant epithelial cells2. The human CANTOS (Canakinumab Anti-Inflammatory Thrombosis Outcome Study) trial evaluated the capacity of the anti-IL-1β Ab canakinumab to prevent cardiovascular events. In the group of patients at high risk of developing lung cancer, canakinumab reduced the incidence and mortality of lung cancer13. Thus, these data raised the interesting hypothesis that inhibiting IL-1β could be a target to fight human lung cancer. However, the CANOPY program, evaluating the association of canakinumab with first-line platinum-based doublet chemotherapy and anti-PD-L1 (CANOPY-1 phase III14), second-line docetaxel (CANOPY-2 phase III15), surgical complete resection and neoadjuvant cisplatin-based chemotherapy (CANOPY-A phase III16) or neoadjuvant anti-PD-1 and surgery (CANOPY-N phase II17) did not show any clinical efficacy of IL-1β blockade.
In this work, we set out to evaluate the impact of IL-1β on the efficacy of chemo-immunotherapy in NSCLC. We observe that a high IL1B and IL1R1 gene expression is a marker of response for patients treated with chemo-immunotherapy. In mice, injection or overexpression of IL-1β at the tumor site reverses resistance to cisplatin/pemetrexed/anti-PD-1 combination in murine LLC1 and KP1.9 lung cancer models. IL-1β is essential to trigger CXLC10 production by cancer cells and for the chemo-attraction of CD8 T cells at the tumor site. The AIM2 inflammasome, which is widely expressed in murine and human lung cancer cell lines and more expressed in cancer tissues as compared to normal lung tissues, could be used to trigger IL-1β and CXCL10 production. In this way, TXNIP (Thioredoxin-Interacting Protein) overexpression leads to the release of mitochondrial DNA (mtDNA) in the cytosol, which activates AIM2 inflammasome, induces caspase-1 activation and IL-1β and CXCL10 production and reverses chemo-immunotherapy resistance. Moreover, a high score including IL1B, IL1R1, AIM2 and TXNIP expression is associated with better response to immunotherapy in NSCLC patients. Using a screening of potential TXNIP inductors, we find that trametinib (a Mitogen-activated protein kinase kinase (MEK) inhibitor) and JNJ-26854165 (a tryptamine derivative, MDM2 inhibitor) have the capacity to reverse resistance to cisplatin/pemetrexed/anti-PD1 treatment in a caspase-1 and IL-1β-dependent manner.
Results
IL-1β improves NSCLC response to chemo-immunotherapy
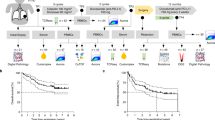
In order to evaluate the role of the IL-1R1/IL-1β pathway in the response to chemo-immunotherapy in mice, we used LLC1 (a spontaneous Kras/Nras p53 mutant cancer cell line) and KP1.9 (a lung adenocarcinoma cell line isolated from C57BL/6, KrasLSL G12D/+; p53fl/fl mice) mouse lung cancer cells. These cell lines were subcutaneously injected in mice, and the resulting mouse models were resistant to cisplatin/pemetrexed/anti-PD-1 mAb (CDDP/Pem + αPD-1) treatment (Fig. 1A). To understand the effects of IL-1β in these models, we used either recombinant IL-1β intratumor injections or overexpressing cancer cells. While recombinant IL-1β had no effect on tumor growth, its overexpression led to opposite effects with an increase in LLC1 tumor growth and a delay in KP1.9 tumor growth (Supplementary Fig. 1A). However, intra-tumor injections of recombinant IL-1β (rIL-1β) enabled chemo-immunotherapy efficacy and inhibition of tumor growth (Fig. 1B and Supplementary Fig. 1B). Similar results were observed when LLC1 cells or KP1.9 cells were genetically modified to overexpress bioactive IL-1β (Fig. 1C and Supplementary Fig. 1A). In these models with high IL-1β production in the tumors, some mice (from 6 to 44%, depending on the model) were cured by the treatment (Supplementary Fig. 1C). IL-1β had no or slight effects on anti-PD-1 or chemotherapy monotherapy efficacy and only improved CDDP/Pem + αPD-1 treatment (Supplementary Fig. 1D). The association of CDDP/Pem + αPD-1 with intratumor injections of IL-1β had also an effect on the establishment of tumor lung foci of LLC1 cells i.v. injected (Fig. 1D). To evaluate the systemic and abscopal effects of IL-1β, mice bearing two LLC1 tumors (one in each flank) were treated with chemo-immunotherapy with injections of recombinant IL-1β only in the left tumor. The effects of intra-tumor injections of recombinant IL-1β delayed the growth of both tumors, thus confirming that the immune response generated by IL-1β is systemic (Fig. 1E, left panel). This result was not observed when the experiment was performed with an LLC1 tumor (which received intra-tumor IL-1β injections) and a pancreas tumor (KPC) on the other flank (Fig. 1E, right panel).
A Control LLC1 or KP1.9 lung cancer cells were s.c. injected into C57BL/6 mice at d0. Tumor-bearing mice were treated with cisplatin (CDDP: 6 mg/kg ip), pemetrexed (Pem: 100 mg/kg ip) at d9 and anti-PD-1 mAb (10 mg/kg ip, 3 times a week from d7, for 3 weeks). Tumor size (n = 10) was monitored. B Control LLC1 or KP1.9 lung cancer cells were s.c. injected into C57BL/6 mice at d0. Recombinant IL-1β (rIL-1β - 50 µg/kg) was daily injected in the tumor from d7 to d15. Tumor-bearing mice were treated with cisplatin/pemetrexed/anti-PD-1 mAb as above. Tumor size (n = 10) was monitored. C mature IL-1β (mIL1β) overexpressing LLC1 or KP1.9 lung cancer cells were s.c. injected into C57BL/6 mice at d0. Tumor-bearing mice were treated with cisplatin/pemetrexed/anti-PD-1 mAb as above. Tumor size (n = 10) was monitored. Insets: mature IL-1β expression was analyzed by Western Blot. β-Actin was used as a loading control. Numbers indicate molecular weights in kDa. D LLC1 cells were s.c. and i.v. injected into C57BL/6 mice at d0. Recombinant IL-1β (rIL-1β - 50 µg/kg) was daily injected in the tumor from d7 to d15. Tumor-bearing mice were treated with cisplatin/pemetrexed/anti-PD-1 mAb as above. Lung tumor foci were numerated at day 15. Box and whiskers represent min to max, with the center line representing the median (n = 7). E C57BL/6 mice were s.c. injected into the left flank with LLC1 cells and also with LLC1 (left panel) or KPC (right panel) cells into the right flank. Tumor-bearing mice were treated or not by chemotherapy and anti-PD-1 mAb, as in A and recombinant IL-1β was only injected in the LLC1 tumors localized in the left flank. Tumor volumes were monitored at day 25 (n = 8) and represented as a percentage of the corresponding untreated tumor. Box and whiskers represent min to max, with the center line representing the median. F Kaplan–Meier estimation of progression-free survival of patients from cohort 1 (left panel), cohort 2 (middle panel) or the TCGA LUAD database (right panel). Patients were split into four groups with high (H) or low (L) expression of IL1B and IL1R1 genes. High expression was determined using the optimal cut-off estimated from maximally selected rank statistics. Data represent mean ± SEM (A–E). Two-way ANOVA (A–C) and unpaired t tests (D, E). n.s., not significant. Log-rank t test p-values (F). Source data are provided as a Source Data file. Panel (A) was created in part with BioRender.
Moreover, the effects of IL-1β seem restricted to lung cancer models treated with CDDP/Pem + anti-PD-1. No impact of IL-1β on tumor growth was found in the KPC pancreatic cancer model, treated with CDDP/Pem + anti-PD-1 or FOLFIRINOX (5-FU, calcium folinate, irinotecan, oxaliplatin – one of the chemotherapy regimens usually used for pancreatic cancer patients) + anti-PD-1 or in the LLC1 model treated with FOLFIRINOX + anti-PD-1 (Supplementary Fig. 2).
In order to confirm the potential role of IL-1β in human NSCLC response to chemo-immunotherapy, RNAseq data from 2 cohorts of patients from Dijon and Montreal, treated with platinum-based doublet chemotherapy followed by anti-PD-1 or anti-PD-L1 mAb18 (Supplementary Table 1; n = 92 for Cohort 1 and n = 70 for Cohort 2) and from the TCGA (506 patients in the LUAD cohort without immunotherapy) were analyzed. Using progression-free survival (PFS) under anti-PD-1/PD-L1 mAbs and overall survival (OS) after initial anti-PD-1/PD-L1 mAbs, we observed in cohorts 1 and 2 that high IL1R1 was associated with better PFS, while IL1B was significantly associated with better PFS only in cohort 1 (Supplementary Fig. 3A). In both cohorts, patients with high IL1B and high IL1R1 expression had better PFS and OS than other patients (Fig. 1F and Supplementary Fig. 3B). In contrast, IL1B or IL1R1 genes were not associated with outcome in patients from the TCGA LUAD cohort who had not received immunotherapy (Fig. 1F and Supplementary Fig. 3A and B).
Thus, these results show a positive effect of IL-1β on chemotherapy and immunotherapy efficacy in human and mouse NSCLC.
IL-1β induces CXCL10-mediated CD8 T cell recruitment
In RNAseq data from the TCGA LUAD and our 2 cohorts of NSCLC patients, we analyzed GSEA hallmarks in responders vs non-responders, using median cohort PFS as a cut-off. We observed an enrichment of inflammatory and immune response pathways (TNFA signaling, IL6/JAK/STAT3 signaling, Allograft rejection, inflammatory response, IL2/STAT5 signaling) in NSCLC patients who received chemotherapy and immunotherapy with prolonged PFS but not in the TCGA database, where patients were not treated with such treatment (Supplementary Fig. 4A). Because chemo-immunotherapy efficacy mainly relies on CD8 T cell effector functions and abundance within the tumor3, we evaluated the Extended Immune Gene (EIG) and IFN signatures, previously described as predictive of response to immunotherapy19. We found that expression of these gene sets was increased in IL1BHigh and IL1R1High patients from NSCLC cohorts 1 and 2 and TCGA LUAD databases (Supplementary Fig. 4B). This suggests that the IL-1β/IL-1R1 pathway is correlated with IFN signatures and CD8 infiltration or CXCL10-dependent chemoattraction in the tumor. To explore this hypothesis, we examined the impact of IL-1β on CD8 T cell accumulation and activation in mouse lung cancer cells subcutaneously implanted in mice. We observed that CDDP/Pem + αPD-1 did not increase gene expression of T cell immune markers at the tumor site (Cxcl9, Cxcl10, Cd3e, Cd8a, Ifng, Gzmb) (Fig. 2A). However, in IL-1β-overexpressing tumors, the association of CDDP/Pem + αPD-1 promoted a major increase in gene expression of T cell immune markers (Fig. 2A). Cxcl10 induction started 6 h after chemo-immunotherapy treatment and was followed by increased Cd3e and Cd8a expression, starting 24 h after treatment initiation. Similar results were observed in LLC1 with intra-tumor recombinant IL-1β injections and in KP1.9-overexpressing IL-1β tumor-bearing mice treated with chemo-immunotherapy (Fig. 2B, C). The recruitment of CD8 T cells in the tumor following chemo-immunotherapy and intra-tumor recombinant IL-1β treatments was confirmed by immunohistochemistry and flow cytometry (Fig. 2D and Supplementary Fig. 5). To determine the role of CXCL10-mediated CD8 T cell recruitment at the tumor site, we depleted CD8 T cells or inhibited CXCR3 (the receptor of CXCL9, 10 and 11), using specific mAbs. We observed that both mAbs inhibited the therapeutic efficacy of CDDP/Pem + αPD-1 + recombinant IL-1β on LLC1 tumor growth (Fig. 2E). Similarly, the anti-CXCR3 mAb also reversed the antitumor effect of chemo-immunotherapy in the context of IL-1β overexpression in LLC1 cells (Supplementary Fig. 6). IL-1β can fix IL-1R1 expressed either on cancer cells or host cells. CDDP/Pem/αPD-1 was still effective at reducing tumor growth of IL-1β overexpressing KP1.9 cells in WT and Il1r1-/- mice, thus suggesting an effect of IL-1β on tumor cells (Supplementary Fig. 7). Moreover, CDDP/Pem + αPD-1 + rIL-1β therapeutic effects were inhibited when tumors were generated using LLC1 knockdown for Il1r1 (Fig. 2F, G). CXCL10 can be produced by numerous cells within the tumor, such as myeloid or cancer cells3. To evaluate the implication of an IL-1β/CXCL10-autocrine pathway in cancer cells, we used Cxcl10-deficient LLC1 cells (Fig. 2F). We observed that the effect of CDDP/Pem + αPD-1 + rIL-1β on tumor growth was completely inhibited in tumors generated with Cxcl10-deficient LLC1 cells (Fig. 2G). These results thus demonstrate an essential role of the IL-1 receptor and CXCL10 production within the cancer cells.
A Control LLC1 cells or mature IL-1β (mIL1β) overexpressing LLC1 cells were s.c. injected into C57BL/6 mice at d0. Tumor-bearing mice were treated with cisplatin (CDDP: 6 mg/kg ip), pemetrexed (Pem: 100 mg/kg ip) at d9 and anti-PD-1 mAb (10 mg/kg ip at d9 and d11). Heat map showing the expression (z-score) of indicated genes evaluated by RT-qPCR in tumors 6, 12, 24 or 72 h after chemotherapy treatment (n = 4–10). B LLC1 cells were s.c. injected into C57BL/6 mice at d0. Recombinant IL-1β (rIL-1β - 50 µg/kg) was daily injected in the tumor from d7 to d15. Tumor-bearing mice were treated with cisplatin/pemetrexed/anti-PD-1 mAb as in A. Heat map showing the expression (z-score) of indicated genes evaluated by RT-qPCR in tumors 8 days after chemotherapy treatment (n = 7). C Control KP1.9 cells or mature IL-1β (mIL1β) overexpressing KP1.9 cells were s.c. injected into C57BL/6 mice at d0. Tumor-bearing mice were treated with cisplatin/pemetrexed/anti-PD-1 mAb as in A. Heat map showing the expression (z-score) of indicated genes evaluated by RT-qPCR in tumors 8 days after chemotherapy treatment (n = 5–7). D LLC1 tumor-bearing C57BL/6 mice were treated with cisplatin/pemetrexed/anti-PD-1/rIL-1β as above. Tumors were recovered 6 and 8 days after treatment (d15 and d17), and CD8 staining was analyzed by IHC. Left panel: representative images; right panel: quantification of CD8 T cells, box and whiskers represent min to max, with center line representing the median (n = 5). E LLC1 tumor-bearing C57BL/6 mice were treated with cisplatin/pemetrexed/anti-PD-1/rIL-1β as above in the presence of anti-CD8, anti-CXCR3 mAbs or control Ig (5 mg/kg ip, 3 times a week from d9). Tumor size (n = 7) was monitored. F LLC1 knockdown for Il1r1 or Cxcl10 were generated by CRISPRCas9. Upper panel: the expression of IL-1R1 was detected by flow cytometry; lower panel: Cxcl10 expression was analyzed by RT-qPCR (n = 6). G Control, Cxcl10 (Cxcl10KD) or Il1r1 (Il1r1KD)-deficient LLC1 cells were s.c. injected into C57BL/6 mice. Tumor-bearing mice were treated with cisplatin/pemetrexed/anti-PD-1/rIL-1β as above. Tumor size was monitored (left panel: n = 10; middle panel: n = 6; right panel: n = 8). H LLC1 cells were treated in vitro with 100 ng/mL rIL-1β for 2 h and analyzed by RNAseq. Volcano plot representation of genes differentially expressed in rIL-1β-treated cells versus untreated cells. The X-axis represents the log2 fold change, and the Y-axis represents the − log10 adjusted p-values. Chemokine genes were tagged. I LLC1 cells were treated in vitro with 100 ng/mL rIL-1β for the indicated times. Cxcl10 mRNA expression was analyzed by RT-qPCR (n = 6). J Control or mIL1β overexpressing LLC1 cells were seeded in plates. After 24 h, Cxcl10 mRNA expression was analyzed by RT-qPCR (n = 12). K Scatter plot of CXCL10 mRNA expression after recombinant IL-1β treatment and IL-1R1 membrane expression (normalized MFI = IL-1R1 MFI / Ig MFI) in the indicated human lung cancer cell lines. Data represent mean ± SEM (E, G) or z-score analysis (A–C). Data represent mean ± SD of two or three independent experiments (F and I–K). Data presented as boxplots of median ± interquartile range; whiskers correspond to min and max (D). Unpaired t test comparing each condition with untreated (-) control cells. No star = not significant (A–C). Unpaired t test comparing indicated conditions (D). Unpaired t test comparing each condition with control LLC1 cells (F and J) or with 0 h (I). Two-way ANOVA (E and G). *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001, ns, not significant. P-values were estimated using a two-sided Wald test and were adjusted using the Benjamini-Hochberg method (H). Source data are provided as a Source Data file. Panels (A, B, C, E and G) were created in part with BioRender.
We evaluated the effects of IL-1β in vitro on LLC1 and KP1.9 cells, which both expressed IL-1R1 (Supplementary Fig. 8A). First, recombinant IL-1β did not impact the cytotoxic effects of CDDP/Pem on LLC1 cells (Supplementary Fig. 8B). Second, using RNAseq, we observed that Cxcl10 was the gene whose expression was most induced by IL-1β among genes coding for chemokines (Fig. 2H). This result was confirmed by QPCR and ELISA in LLC1 and KP1.9 cells (Fig. 2I and Supplementary Fig. 8C and D). Moreover, the overexpression of mature IL-1β in LLC1 and KP1.9 cells also induced CXCL10 expression and production in vitro (Fig. 2J and Supplementary Fig. 8E and F), strengthening the direct effect of IL-1β on cancer cells. IL-1β induced Cxcl10 expression mainly through NF-κB, and not the type I IFN pathway. In fact, while NF-κB inhibitors inhibited IL-1β-induced Cxcl10 expression, IFNAR blocking Ab had no effect and no Ifn gene expression induction was observed (Supplementary Fig. 8G and H).
Finally, we evaluated the capacity of IL-1β to increase CXCL10 expression in different human lung cancer cell lines with different levels of IL-1R1 expression at the cell surface (Supplementary Fig. 9). Recombinant human IL-1β was able to increase CXCL10 expression at different levels in all the cell lines tested, and the level of CXCL10 expression was correlated with IL-1R1 expression (Fig. 2K).
Altogether, these results show that the association of chemo-immunotherapy and IL-1β triggers CXCL10 production and expression by lung cancer cells, thus enabling the recruitment of CD8 T cells within the tumor and the anti-tumor effects of chemo-immunotherapy.
Caspase-1 activation and IL-1β production can be induced by the AIM2 inflammasome in lung cancer cells
Because the use of recombinant IL-1β in mice leads to weight loss (Supplementary Fig. 1B) and the treatment of cancer patients by intra-tumor injections of IL-1β seems complex, we searched for ways to induce IL-1β production by cancer cells. IL-1β maturation and production require cleavage by caspase-1, an enzyme that is activated within inflammasome complexes20. The analysis of paired normal and tumor samples from TCGA LUAD, showed that one of the inflammasome sensors whose expression is most enriched in tumors compared to normal lung tissues is AIM2 (Fig. 3A). By Western Blot analysis, AIM2 seemed to be homogenously expressed in human and mouse lung cancer cells, contrary to other inflammasome receptors, like NLRP3 or NLRP1, which were sometimes undetectable depending on the cell line (Fig. 3B). In patients treated with chemotherapy and immunotherapy, higher expression of AIM2 was associated with improved PFS and OS (statistically significant for cohort 2 but not cohort 1), while in patients from the TCGA LUAD cohort who had not received immunotherapy, AIM2 was not associated with outcome (Fig. 3C and Supplementary Fig. 10A). Higher AIM2 expression was also correlated with both EIG and IFN signature expression (Supplementary Fig. 10B). We also showed that all tumor cell lines expressed caspase-1 and ASC, thus supporting their capacity to form an active AIM2 inflammasome (Fig. 3B). To confirm this hypothesis, we evaluated the capacity of inflammasome activators to trigger caspase-1 activation and IL-1β cleavage in LLC1 cancer cells and in J774 macrophage-like cells, as a control. While AIM2 (poly(dA:dT)) or NLRP3 (ATP) activators triggered IL-1β cleavage in the macrophage J774 cell line supernatants, the AIM2 activator was the only one able to induce caspase-1 activation and IL-1β cleavage and release in LLC1 cells (Fig. 3D and Supplementary Fig. 11). Moreover, AIM2 activation was the sole signal able to trigger Cxcl10 expression in LLC1 cells (Fig. 3E). This effect was dependent on AIM2 and IL-1β, as poly(dA:dT) was not able to trigger Cxcl10 expression in LLC1 cells with knock down Aim2 or Il1b expression (Fig. 3F).
A Analysis of inflammasome sensor gene expression between tumor and normal lung tissues from the TCGA LUAD database. The fold increase was calculated by dividing mRNA expression in tumors by mRNA expression in corresponding normal adjacent lung tissue for each patient (n = 58). B Western Blot analysis of indicated proteins in indicated human and mouse lung cancer cell lines. β-Actin was used as a loading control. Numbers indicate molecular weights in kDa. C Kaplan–Meier estimation of progression-free survival in patients from cohort 1 (left panel), cohort 2 (middle panel) or the TCGA LUAD database (right panel). Patients were split into two groups with high (H) or low (L) expression of the AIM2 gene. High expression was determined using the optimal cut-off estimated from maximally selected rank statistics. D and E LLC1 cells were treated either with LPS (0.5 µg/mL), the NLRP1 activator Val-boroPro (0.05 mM), the NLRP3 activator ATP (5 mM) or the AIM2 activator poly(dA:dT) (5 µg/mL). After 1 h of treatment, caspase-1 activity was analyzed using FLICA1 by flow cytometry (n = 6 independent wells), and after 2 h of treatment, the presence of IL-1β was analyzed by Western Blot in cell supernatants (D). After 18 h of treatment, Cxcl10 mRNA expression was analyzed by RT-qPCR (E, n = 9 independent wells). F LLC1 control cells or knock down for Aim2 (Aim2KD) or Il1b (Il1bKD) were treated with the AIM2 activator poly(dA:dT) (5 µg/mL) for 18 h, and Cxcl10 mRNA expression was analyzed by RT-qPCR (n = 9 independent wells). Data represent mean ± SD of three independent experiments (D–F). The data represent one experiment representative of two independent experiments (B). Unpaired t test comparing each condition with the untreated condition (-) (D, E). Unpaired t test comparing indicated conditions (F). ns, not significant. Wald test (A) and log-rank t test p-values (C). Source data are provided as a Source Data file.
Thus, these results show that lung cancer cells highly express AIM2 and that its activation induces IL-1β cleavage and release, and subsequent CXCL10 production.
TXNIP overexpression improves CDDP/Pem-induced CXCL10 expression and anti-tumor effects
AIM2 is activated by double-strand DNA (dsDNA), and in sterile conditions, the main sources of host-derived dsDNA are the nucleus and mitochondria21,22. Therefore, we searched for ways to trigger mitochondrial DNA (mtDNA) accumulation in the cytosol. Chemotherapy by inducing cell death can induce mitochondrial permeabilization and mtDNA release or can induce migration of nuclear DNA into the cytoplasm by generation of micronuclei23. We observed that CDDP/Pem triggered modest release of mtDNA (and not nuclear DNA) into the cytosol of LLC1 cells at early times (Supplementary Fig. 12A and B) but could not induce significant caspase-1 activation (Supplementary Fig. 12C). Consequently, CDDP/Pem only induced modest expression of Cxcl10 (Supplementary Fig. 12D). The release of mtDNA into the cytosol is mainly orchestrated by membrane pores, mainly generated by the oligomerization of Bax/Bak24. Thus, we evaluated the capacity of BH3-mimetics ABT-263 (navitoclax), a Bcl-2 and Bcl-XL inhibitor, and ABT-199 (venetoclax), a specific Bcl-2 inhibitor, to trigger mtDNA cytosolic relocalization in LLC1 cells. Navitoclax, but not venetoclax, was able to induce small amounts of mtDNA in the cytosol, an effect that was not enhanced in association with CDDP/Pem (Supplementary Fig. 13A). Thus, these drugs were not able to enhance the expression of Cxcl10 induced by chemotherapy (Supplementary Fig. 13B). Together, these data support that targeting BH3 could not be used in this model to trigger mtDNA release and AIM2 activation.
Another way to induce mtDNA release is to modulate mitochondrial oxidative stress. One of the main regulators of oxidative stress is TRX2, which binds either ASK1 or TXNIP. Thus, the association of TRX2 with ASK1 inhibits its phosphorylation and the consequent release of mitochondrial content into the cytosol, an effect that is inhibited by the sequestration of TRX2 by TXNIP25. In this way, TXNIP was deemed to be a suitable target to induce mtDNA release26. To explore whether TXNIP could improve AIM2 inflammasome activation in our model, we overexpressed Txnip in LLC1 cells (Fig. 4A). TXNIP overexpression induced ASK1 phosphorylation (Fig. 4A) and triggered high amounts of mtDNA release in the cytosol (Fig. 4B). TXNIP overexpression also triggered caspase-1 activation and IL-1β release by cancer cells (Fig. 4C). We also evaluated the synergy between TXNIP overexpression and CDDP/Pem on Cxcl10. While chemotherapy only slightly induced Cxcl10 expression, its association with Txnip overexpression was synergic, enhancing Cxcl10 expression (Fig. 4D). Ethidium bromide (EtBr), which induces mtDNA depletion, and ASK1 inhibitor, completely blunted Cxcl10 induction, thus demonstrating the essential role of ASK1 phosphorylation and mtDNA cytosolic release in TXNIP/CDDP/Pem-induced Cxcl10 expression (Fig. 4D). In order to investigate whether this effect was dependent on IL-1β production, we overexpressed Txnip in Il1b-deficient or control LLC1 cells and showed that Txnip overexpression did not influence Cxcl10 expression in the absence of IL-1β (Fig. 4E). Finally, while CDDP/Pem/αPD-1 had no antitumor effects on subcutaneous LLC1 tumors, the overexpression of TXNIP in LLC1 cells enabled chemo-immunotherapy to decrease tumor growth (Fig. 4F and Supplementary Fig. 14). This therapeutic effect was dependent on CXCL10, since, when TXNIP was overexpressed in Cxcl10-deficient LLC1 cells, chemo-immunotherapy was inefficient in vivo (Fig. 4F). The effect of TXNIP plus chemo-immunotherapy was also associated with induction of T cell marker gene expression in the tumors (Fig. 4G). Similarly, in human lung cancer cell lines, the overexpression of TXNIP also increased the capacity of CDDP/Pem to induce CXCL10 expression (Fig. 4H).
A Western Blot analysis of indicated proteins in LLC1 cells stably transfected with empty vector (EV) or Txnip (Txnip#). β-Actin was used as a loading control. Numbers indicate molecular weights in kDa. B Cox1, Dloop and Non-Numt mitochondrial DNA expression was analyzed by RT-qPCR in cytosolic fractions from EV or Txnip# LLC1 cells (n = 6 independent wells). C EV or Txnip# LLC1 cells were seeded in plates, and caspase-1 activity was analyzed after FLICA1 staining by flow cytometry (n = 12 independent wells) and the presence of IL-1β was analyzed in cell supernatants by Western Blot. Number indicates molecular weight in kDa. D EV or Txnip# LLC1 cells were treated or not with 3.7 µM cisplatin (CDDP) and 33 µM pemetrexed (Pem). After 18 h of treatment in the presence or absence of EtBr or ASK1 inhibitor (Selonsertib 25 µM), Cxcl10 mRNA expression was analyzed by RT-qPCR (n = 9 independent wells). E control or Il1bKD LLC1 cells transfected with empty vector (EV) or overexpressing Txnip were treated as in (D). Cxcl10 mRNA expression was analyzed by RT-qPCR after 18 hours of treatment (n = 12 independent wells). F, G C57BL/6 mice were s.c. injected either with control, Txnip overexpressing LLC1 cells (Txnip#) or Cxcl10-deficient and Txnip overexpressing LLC1 cells (Cxcl10KD/Txnip#) at d0. Tumor-bearing mice were treated with cisplatin (6 mg/kg ip), pemetrexed (100 mg/kg ip) at d9 and anti-PD-1 mAb (10 mg/kg ip, 3 times a week from d9, for 3 weeks). Tumor size (left panel: n = 7, middle panel: n = 8, right panel: n = 7) was monitored (F), and tumor biopsies (Control: n = 10, Txnip#: n = 10, Cxcl10KD: n = 4, Cxcl10KD/Txnip#: n = 6) were recovered 8 days after chemotherapy treatment to analyze indicated gene expression (z-score) by RT-qPCR (G). H H460 and H1299 human lung cancer cell lines were transfected with TXNIP (TXNIP#) or empty vector (EV). Left panel: TXNIP mRNA expression was analyzed by RT-qPCR (n = 9). Right panel: CXCL10 mRNA expression was analyzed by RT-qPCR after 18 h of treatment with 3.7 µM cisplatin (CDDP) and 33 µM pemetrexed (Pem) (H460: n = 6 independent wells; H1299: n = 12 independent wells). Data represent mean ± SEM (F) or z-score analysis (G). Data represent mean ± SD of three independent experiments (B–E and H). The data represent one experiment representative of three independent experiments (A). Unpaired t test comparing indicated conditions (B–E and H). Unpaired t test comparing CDDP/Pem/αPD-1 (+) with untreated conditions (−) for each cell type. No star = not significant (G). Two-way ANOVA (F). *p < 0.05, **p < 0.01, ***p < 0.001, ns, not significant. Source data are provided as a Source Data file. Panel (F) was created in part with BioRender.
Taken together, these results show that TXNIP overexpression enables CDDP/Pem to trigger mtDNA release and, following caspase-1 activation and IL-1β production, to induce Cxcl10 expression, CD8 T cell recruitment and tumor growth control.
Effects of TXNIP inductors on CDDP/Pem-induced CXCL10 expression and anti-tumor effects
Because TXNIP overexpression was shown to enable CDDP/Pem to induce Cxcl10 expression and reverse resistance to chemo-immunotherapy, we evaluated its impact together with the previously identified targets, i.e., IL1B, IL1R1 and AIM2, on NSCLC patients’ outcomes. Using high vs low gene score classification, we observed that high expression of the 4 genes was correlated with better PFS and OS for patients who had received chemotherapy and immunotherapy (either cohort 1 and 2) but not for patients from the TCGA LUAD, who were never treated with immunotherapy (Fig. 5A and Supplementary Fig. 15).
A Kaplan–Meier estimation of progression-free survival in patients from cohort 1 (left panel), cohort 2 (middle panel) or the TCGA LUAD database (right panel). Patients were split into five groups with high (H) or low (L) expression of 0 to 4 genes (IL1B, IL1R1, AIM2, TXNIP). High expression was determined using the optimal cut-off estimated from maximally selected rank statistics. B Public RNAseq and microarray database analysis to identify molecules able to induce TXNIP expression. Red dots correspond to compounds with a fold increase > 1.5 and a p-value < 0.05. C LLC1 cells were treated for the indicated times with increasing concentrations of the molecules identified in (B). Txnip mRNA expression was determined by RT-qPCR. D LLC1 cells were treated or not for 2 h with the indicated molecules. TXNIP expression was analyzed by Western Blot. β-Actin was used as a loading control. Numbers indicate molecular weights in kDa. Molecules; F: Trametinib (0,1 µM), H: Panobinostat (25 nM), I: SNX-2112 (0,125 µM), J: Sunitinib (1,25 µM), K: YM-155 (0,05 µM), L: JNJ-26854165 (25 nM), M: Foretinib (12,5 nM), N: TW-37 (2,5 µM). E LLC1 cells were pre-treated or not for 2 h with indicated molecules and then with 3.7 µM cisplatin (CDDP) and 33 µM pemetrexed (Pem). After 18 h of treatment, Cxcl10 mRNA expression was analyzed by RT-qPCR (n = 6 independent wells). Molecules; F: Trametinib (0,1 and 1 µM), H: Panobinostat (5 and 25 nM), I: SNX-2112 (0,025 and 0,125 µM), J: Sunitinib (0,250 and 1,25 µM), K: YM-155 (0,01 and 0,05 µM), L: JNJ-26854165 (5 and 25 nM), M: Foretinib (2,5 and 12,5 nM), N: TW-37 (0,5 and 2,5 µM). F, G, LLC1 tumor-bearing C57BL/6 mice (s.c. injection at day 0) were daily treated or not with indicated molecules from d7 to d15 (see supplementary Table 2 for dosage). Mice were also treated once i.p. or not with 6 mg/kg cisplatin (CDDP) and 100 mg/kg pemetrexed (Pem) at day 9 and three times a week with 10 mg/kg anti-PD-1 Ab (from day 9). Tumor size (n = 8) was monitored at day 30. Box and whiskers represent min to max, with the center line representing the median (F). Tumor biopsies (n = 9) were recovered 8 days after chemotherapy treatment to analyze indicated gene expression (z-score) by RT-qPCR (G). Data represent mean ± SEM (F) or z-score analysis (G). Data represent mean ± SD of three independent experiments (C and E). The data represent one experiment representative of two independent experiments (D). Unpaired t test comparing each condition with CDDP/Pem alone (E) or CDDP/Pem/αPD-1 alone (F and G). *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001, ns, not significant. For panels (G), no star means not significant. Log-rank t test p-values (A). Differential tests were two_sided and P-values were adjusted using the Benjamini-Hochberg FDR correction method (B). Source data are provided as a Source Data file. Panel (F) was created in part with BioRender.
This observation encouraged us to find a way to increase TXNIP expression, with a view to improving the efficacy of chemo-immunotherapy. Therefore, we aimed to identify molecules capable of inducing TXNIP expression, using a bioinformatic strategy based on analysis of public RNAseq and microarray databases, as previously described by Gu et al.27. Using existing transcriptomic studies on drug treatment effects in the GEO database with differential expression, we searched for drugs that could induce TXNIP expression. This transcriptomic database involved 719 transcriptomic profiles and tested the effects of 235 different drugs on human cancer cell lines. Sixteen molecules able to statistically increase TXNIP expression after p-value adjustment with the Benjamini-Hochberg method (FDR correction) were selected as possible TXNIP inducers (Fig. 5B and Supplementary Table 2). Then, the capacity of these molecules to increase TXNIP expression in LLC1 cells was studied. Only half of the selected molecules were able to induce TXNIP mRNA expression in our model (Fig. 5C). The effect of these molecules on TXNIP expression was confirmed at the protein level by Western Blot (Fig. 5D). Six of these molecules were able to increase Cxcl10 expression and/or CXCL10 production in the supernatant of LLC1 cells following CDDP/Pem treatment (Fig. 5E and Supplementary Fig. 16). The potential effect of these molecules was then investigated in vivo. We found that three molecules were able to improve CDDP/Pem/αPD-1 effects on subcutaneous LLC1 tumor growth and also to increase Cd8a, Cd3e, Cxcl10, Gzmb and/or Ifng expression at the tumor site (Fig. 5F, G). We then decided to focus our interest on two of the most efficient of these molecules, namely trametinib (molecule F), a MEK inhibitor, and JNJ-26854165 (molecule L), an MDM2 inhibitor.
MEKi plus chemo-immunotherapy anti-tumor effects are mediated by the TXNIP/AIM2/caspase-1/IL-1β pathway
We first evaluated the capacity of CDDP/Pem/MEKi to activate the AIM2 inflammasome. We observed that CDDP/Pem/MEKi was able to increase caspase-1 activity in LLC1 cells, after 30 minutes of treatment, while MEKi or CDDP/Pem alone did not (Fig. 6A and Supplementary Fig. 12C). Moreover, the association of CDDP/Pem with MEKi enabled detection of IL-1β in the cell supernatant after two hours of treatment (Fig. 6A and Supplementary Fig. 17A). We also showed that CDDP/Pem/MEKi induced an increase in mtDNA content in the cytosol (Supplementary Fig. 17B) to a greater extent than CDDP/Pem alone (Supplementary Fig. 12B). Moreover, in LLC1 cells invalidated for Txnip, CDDP/Pem/MEKi was not able to induce mtDNA release into the cytosol (Fig. 6B). Using AIM2 immunoprecipitation, we detected an interaction between cytosolic mtDNA and AIM2 following CDDP/Pem/MEKi treatment (Fig. 6C). In the absence of TXNIP, CDDP/Pem/MEKi was not able to induce caspase-1 activation or IL-1β secretion (Fig. 6D). To study the role of AIM2 in the effects of CDDP/Pem/MEKi treatment, we used Aim2 knock down LLC1 cells (Fig. 6E). Invalidation of Aim2 expression inhibited caspase-1 activation and IL-1β production, following CDDP/Pem/MEKi treatment (Fig. 6E). Moreover, CDDP/Pem/MEKi was less effective at inducing Cxcl10 mRNA expression and CXCL10 protein production in Aim2 or Txnip-invalidated LLC1 cells, compared to control cells (Fig. 6F and Supplementary Fig. 18A and B).
A LLC1 cells were pre-treated for 2 h with 100 nM trametinib (MEKi) and then with 3.7 µM cisplatin (CDDP) and 33 µM pemetrexed (Pem) for the indicated times. The presence of IL-1β was analyzed in cell supernatants by Western Blot (upper panel, number indicates molecular weight in kDa.) and caspase-1 activity was assayed after FLICA1 staining by flow cytometry (lower panel, n = 6 independent wells). B LLC1 cells control or knock down for Txnip by CRISPRCas9 (TxnipKD) were treated or not with CDDP/Pem/MEKi as indicated in A. After 30 min of chemotherapy treatment, Cox1, Dloop and Non-Numt mitochondrial DNA expression was analyzed by RT-qPCR in cytosolic fractions (Co: n = 9, TxnipKD: n = 6 independent wells). Inset: Western Blot analysis of TXNIP expression in untreated cells. β-Actin was used as a loading control. Numbers indicate molecular weights in kDa. C LLC1 cells were treated or not with CDDP/Pem/MEKi as indicated in A. After 30 min of chemotherapy treatment, immunoprecipitation of AIM2 was performed, and associated DNA was purified. RT-qPCR analysis of indicated mitochondrial genes (n = 6 independent wells). Data were normalized to values obtained after immunoprecipitation with a control Ig. D Control or TxnipKD LLC1 cells were treated or not with CDDP/Pem/MEKi as indicated in A. Caspase-1 activity was assayed after FLICA1 staining by flow cytometry, after 30 min of chemotherapy treatment (upper panel, n = 6 independent wells) and the presence of IL-1β in cell supernatants was analyzed by Western Blot, after 2 h of treatment (lower panel). E Control or Aim2KD LLC1 cells were treated or not with CDDP/Pem/MEKi as indicated in A. Caspase-1 activity (upper panel, n = 9 independent wells) and IL-1β in cell supernatants (lower panel) were assayed as indicated in (D). Inset: Western Blot analysis of AIM2 in untreated cells. β-Actin was used as a loading control. Numbers indicate molecular weights in kDa. F Control or TxnipKD or Aim2KD LLC1 cells were treated or not with CDDP/Pem/MEKi as indicated in A. After 18 hours of treatment, Cxcl10 mRNA expression was analyzed by RT-qPCR (n = 9 independent wells). G Control or Casp1KD LLC1 cells were treated or not with CDDP/Pem/MEKi as indicated in A. Caspase-1 activity (upper panel, n = 6 independent wells) and IL-1β in cell supernatants (lower panel) were assayed as indicated in D. Inset: Western Blot analysis of Pro-caspase-1 in untreated cells. β-Actin was used as a loading control. Numbers indicate molecular weights in kDa. H Control or Casp1KD or Il1bKD LLC1 cells were treated or not with CDDP/Pem/MEKi as indicated in A. After 18 h of treatment, Cxcl10 mRNA expression was analyzed by RT-qPCR (n = 9 independent wells). Inset: Western Blot analysis of Pro-IL-1β in untreated cells. β-Actin was used as a loading control. Numbers indicate molecular weights in kDa. I C57BL/6 mice were s.c. injected either with control, Txnip, Aim2, Casp1 or Il1b-deficient LLC1 cells (d0). Tumor-bearing mice were treated or not p.o. daily with 1 mg/kg trametinib (MEKi) from d7 to d15. Mice were also treated or not i.p. once with 6 mg/kg cisplatin (CDDP) and 100 mg/kg pemetrexed (Pem) at day 9 and three times a week with 10 mg/kg anti-PD-1 Ab (from day 9). Tumor size was monitored (control, Aim2KD: n = 10, TxnipKD: n = 15, Casp1KD: n = 9 or Il1bKD n = 7). J H460 and H1299 human lung cancer cells were treated as in (A). The presence of IL-1β was detected in cell supernatants by Western Blot (upper panel, number indicates molecular weight in kDa) and caspase-1 activity was assayed after FLICA1 staining by flow cytometry (lower panel, n = 6 independent wells) at the indicated times. K, L H460, H1299 human lung cancer cells were pre-treated or not with EtBr, caspase-1 inhibitor (YVAD, 50 µM) or IL-1R1 inhibitor (Anakinra, 10 µg/mL) (K, H460: n = 7, H1299: n = 5 independent wells) or transiently transfected with siRNA control or specific for AIM2 (L, H460 n = 10, H1299: n = 12 independent wells) and treated as in A. After 18 hours of treatment, CXCL10 mRNA expression was analyzed by RT-qPCR. Data represent mean ± SEM (I). Data represent mean ± SD of three independent experiments (A –H and J–L). The data represent one experiment representative of two or three independent experiments (A, J). Unpaired t test comparing treated conditions with untreated conditions (0) (A, J). Unpaired t test comparing indicated conditions (B–H and K, L). Two-way ANOVA (I). ns, not significant. Source data are provided as a Source Data file. Panel I was created in part with BioRender.
The importance of mtDNA in CDDP/Pem/MEKi-mediated effects was confirmed by using EtBr (Supplementary Fig. 19A). Our laboratory have previously shown that the use of trametinib improves CDDP/Pem-mediated CXCL10 expression by cancer cells, through TLR9 activation by mtDNA28. Here, we showed that in Tlr9-deficient LLC1 cells, the expression of Aim2 was drastically decreased as compared to untreated control LLC1 cells (Supplementary Fig. 19B), thus suggesting that TLR9 may sense mtDNA to provide a “priming signal” (the first signal required for inflammasome activation6) in LLC1 cells.
Because NLRP3 can also participate in dsDNA sensing29, we investigated its implication in our model. Neither MCC950 NLRP3 inhibitor nor Nlrp3 knockdown were able to dampen CDDP/Pem/MEKi-mediated caspase-1 activation, CXCL10 expression and production in LLC1 cells (Supplementary Fig. 20A and B), suggesting that the NLRP3 inflammasome was not implicated in this context.
To demonstrate the importance of caspase-1 and IL-1β in the effects of CDDP/Pem/MEKi, we first used the YVAD caspase-1 inhibitor to reduce caspase-1 activation (Supplementary Fig. 21A). The use of this inhibitor also decreased the expression and production of CXCL10 induced by CDDP/Pem/MEKi (Supplementary Fig. 21B and C). To confirm the implication of caspase-1, we established Casp1-invalidated LLC1 cells (Fig. 6G). In these cells, CDDP/Pem/MEKi was inefficient at inducing caspase-1 activation and IL-1β maturation as compared to control cells (Fig. 6G). Moreover, we observed that CDDP/Pem/MEKi-mediated Cxcl10 mRNA expression and CXCL10 protein production in Casp1-invalidated LLC1 cells was reduced, compared to control cells (Fig. 6H and Supplementary Fig. 18C). Similarly, using LLC1 cells invalidated for Il1b (Fig. 6H), we showed that CDDP/Pem/MEKi-mediated CXCL10 expression and production were dramatically decreased as compared to control cells (Fig. 6H and Supplementary Fig. 18D). Furthermore, the knock down of Il1r1 expression in LLC1 cells or the use of anakinra (a competitor of IL-1β on IL-1R1) decreased CXCL10 mRNA expression and protein release in cancer cells treated in vitro with CDDP/Pem/MEKi (Supplementary Fig. 22A to C).
To investigate the role of the IL-1/IL-1R1 pathway in the therapeutic efficacy of this treatment, we evaluated the effects of anakinra on LLC1 tumor growth. We observed that anakinra impeded CDDP/Pem/MEKi + αPD-1 antitumor effects (Supplementary Fig. 22D). Anakinra also decreased tumor expression of T cell markers (Supplementary Fig. 22E). Similarly, CDDP/Pem/MEKi + αPD-1 treatment was inefficient at reducing the growth of subcutaneous LLC1 tumors invalidated for Txnip, Aim2, Il1b or Casp1 as compared with control LLC1 cells, in vivo (Fig. 6I). This effect was associated with the absence of induction of T cell markers following CDDP/Pem/MEKi treatment in Txnip or Aim2-deficient LLC1 tumor bearing mice (Supplementary Fig. 23). Finally, after cell sorting, we showed that the treatment of tumor-bearing mice by CDDP/Pem/MEKi increased IL-1β and CXCL10 production only by cancer cells and not myeloid cells (Supplementary Fig. 24A). Moreover, tumor growth experiments showed a similar therapeutic effect of CDDP/Pem/MEKi + αPD-1 in WT, Il1b-/- and Il1r1-/- mice (Supplementary Fig. 24B), thus showing that CDDP/Pem/MEKi effects are dependent on an autocrine cancer-cell derived IL-1β loop.
The effect of the MEKi was also investigated in the KP1.9 model. Trametinib was able to increase Cxcl10 expression in CDDP/Pem-treated cells (Supplementary Fig. 25A) but also to improve the therapeutic effect of the CDDP/Pem/αPD-1 association (Supplementary Fig. 25B). Finally, the capacity of CDDP/Pem/MEKi to induce IL-1β production and caspase-1 activation was further explored in human lung cancer cells. We observed that CDDP/Pem/MEKi was able to induce IL-1β production and caspase-1 activation in H460 and H1299 RAS-mutated cell lines (Fig. 6J). Moreover, the capacity of this treatment to induce CXCL10 expression in these cells was impeded by anakinra, YVAD and EtBr (Fig. 6K). The importance of AIM2 was further explored in human lung cancer cells. We observed that CDDP/Pem/MEKi-induced CXCL10 expression was inhibited in the presence of siRNA targeting AIM2 (Fig. 6L).
Altogether, these results show that CDDP/Pem/MEKi-induced TXNIP expression drives mtDNA release in the cytosol, all required to allow IL-1β and CXCL10 production and anti-tumor effects.
MDM2i plus chemo-immunotherapy has anti-tumor effects that depend on TXNIP, caspase-1, IL-1β and CXCL10
Because the association of MDM2i (compound L: JNJ-26854165) with chemo-immunotherapy also had a strong effect on tumor growth and immune recruitment in vivo (Fig. 5F and G), we also studied this compound. JNJ-26854165 is a tryptamine derivative known to promote mitochondrial damage and mitochondrial cell death in cancer cells30. First, we showed that the association MDM2i/ CDDP/Pem enhanced mtDNA release in the cytoplasm (Supplementary Fig. 26A). MDM2i was also able to induce caspase-1 activation and IL-1β production in association with CDDP/Pem (Supplementary Fig. 26B). To investigate the role of TXNIP, caspase-1 and IL-1β in MDM2i effects, the expression of Cxcl10 was measured in Txnip, Casp1 or Il1b-deficient LLC1 cells. We found that the association of MDM2i with CDDP/Pem was ineffective at inducing Cxcl10 expression in Txnip, Casp1 or Il1b-deficient cell lines, contrary to control cells (Supplementary Fig. 26C). The effects of MDM2i in association with CDDP/Pem/αPD-1 on tumor growth were dependent on TXNIP, caspase-1 and CXCL10 production by cancer cells, as the therapeutic effects of the treatment observed in control LLC1 cells were inhibited when using Cxcl10, Casp1 or Txnip-deficient LLC1 cells (Supplementary Fig. 26D). These effects were correlated with the modification of T cell marker gene expression in the tumors lacking Cxcl10 or Txnip within cancer cells (Supplementary Fig. 26E). The effect of MDM2i was also investigated in the KP1.9 model. MDM2i was able to increase Cxcl10 expression in CDDP/Pem-treated cells (Supplementary Fig. 26F) but also able to improve the therapeutic effect of the CDDP/Pem/αPD-1 (Supplementary Fig. 26G). Lastly, in human lung cancer cell lines, MDM2i improved the capacity of CDDP/Pem treatment to trigger CXCL10 expression (Supplementary Fig. 26H).
Altogether, these results show that the association of MDM2i with CDDP/Pem is able to increase TXNIP/CXCL10 expression in cancer cells and improve the anti-tumor effects of CDDP/Pem/αPD-1.
Discussion
Despite the major benefit of combining chemotherapy and immune checkpoint inhibitors, most patients with NSCLC fail to obtain long-term benefit from this treatment. This treatment strategy needs to be improved, notably by discovering better predictive biomarkers of chemo-immunotherapy efficacy, and by finding new associations to fight resistance mechanisms. To tackle these issues, we first show that induction of active IL-1β (recombinant protein, overexpression or chemical induction) in the tumor improves chemo-immunotherapy efficacy. This effect is dependent on CXCL10-mediated CD8 T cell recruitment at the tumor site.
IL-1β has been described to have ambivalent effects in the context of cancer6,7. In lung cancer, IL-1β favors angiogenesis and EMT8,9,10,11 and deficiency of Il1b in mice inhibits lung tumor growth12. A similar pro-tumor effect was also described in humans, with the CANTOS trial. In that study, patients with atherosclerosis who had had myocardial infarction and who were free of previously diagnosed cancer were enrolled and treated with the anti-IL-1β Ab canakinumab. In treated patients, the incidence and mortality of lung cancer was significantly lower compared to patients who received a placebo, thus suggesting that IL-1β has procarcinogenic effects and favors the development of NSCLC13. However, in therapeutic settings, canakinumab in either second line docetaxel (CANOPY-2), or first line platinum-based doublet chemotherapy plus anti-PD-L1 (CANOPY-1) was ineffective and did not improve the antitumoral effect of chemotherapy or chemo-immunotherapy31. These previous studies are in agreement with our results, i.e., 1) IL-1β continuous overexpression can accelerate tumor growth in the LLC1 model (but not in the KP1.9 model, and this discrepancy might be investigated, and 2) IL-1β should not be inhibited, but rather induced or added in the context of chemo-immunotherapy treatment of NSCLC to trigger therapeutic effects.
In humans, we have also shown that high expression of IL1B and IL1R1 in tumors is associated with better PFS and OS for patients under chemotherapy and immunotherapy treatments, but not among patients who were not treated with immunotherapy, thus corroborating the fact that IL-1β may have different effects depending on the context. Moreover, scoring the expression of genes related to our pathway (IL1B, IL1R1, AIM2, TXNIP) can be a useful marker, because high expression of these four genes is associated with better PFS and OS for patients treated with chemo-immunotherapy. However, due to the limited number of patients and the retrospective design of the study, additional studies are needed to validate our observations.
We demonstrate that local or systemic IL-1β could reverse resistance to CDDP/Pem plus anti-PD-1 in subcutaneous lung cancer models. In addition, IL-1β therapy can also reverse the resistance of lung tumor foci. However, this last model only mimics lung metastasis, and our experiments do not completely investigate in situ lung cancer. Thus, the local tumor microenvironment of the orthotopic lung cancer model is different from the microenvironment of sc tumors, and we cannot eliminate that it may limit the efficacy of our combination therapy.
When we compared the effects of IL-1β in association with cisplatin/pemetrexed + anti-PD-1 or FOLFIRINOX + anti-PD-1 in lung or pancreatic cancer models, a therapeutic effect was observed only when IL-1β was associated with cisplatin/pemetrexed + anti-PD-1 in the lung cancer model. This suggests that the effects of IL-1β are restricted to lung cancer models treated with cisplatin/pemetrexed and anti-PD-1. However, our investigation should be extended to other mouse cancer models, and further experiments are needed to understand why IL-1β has therapeutic effects only in some conditions.
Like IL-1β, AIM2 has been reported to favor lung cancer cell proliferation and invasion in the absence of treatment32,33. However, a pan-cancer analysis of public RNA databases previously described that AIM2-related gene expression was associated with the expression of chemokine receptors (CXCR3, CXCR6, CCL5 and CCL2) and corresponding chemokines (CXCL9, 10, 11, 16 and CCL2, 3, 4, 5) and the infiltration of CD8, CD4, NK cells and macrophages in almost all cancer types studied34. In our patient cohorts treated with chemo-immunotherapy, high expression of AIM2 was slightly correlated with a clinical benefit of immunotherapy. In addition, high AIM2 expression was associated with higher immune infiltration and IFNG signature, thus suggesting its essential role in positively regulating antitumor immune response. Because AIM2 is highly expressed in lung tumor tissues, compared to healthy tissues, and appears to be the most homogeneously expressed inflammasome sensor in human and mouse lung cancer cells, it could be considered as an interesting target for NSCLC treatment, as compared to other inflammasome sensors, such as NLRP3, which is not strongly expressed in lung cancer cells35.
One way to activate AIM2 is to induce the release of mtDNA in the cytosol. This phenomenon is regulated by the Bcl2 family members. Mitochondrial membrane pores are mainly generated by the oligomerization of Bax/Bak24. However, in our model, only the Bax/Bak mimetic navitoclax and not venetoclax induced the accumulation of low amounts of mtDNA in the cytosol. This discrepancy between the two inductors might be explained by the fact that navitoclax targets both Bcl-2 and Bcl-XL, while venetoclax targets only Bcl-2. This suggests that the consideration of such a strategy needs to strengthen the understanding of Bcl-2 member expression and functions in lung cancer models, and maybe to use inhibitors with a wider effect and targeting more Bcl-2 members. For this reason, we decided to search for a target upstream of mitochondria, and found TXNIP, which has been described to participate in mitochondrial dysfunction, mtDNA release into the cytosol and IL-1β production in human and mouse adipose tissue26,36. When overexpressed in lung cancer cells, TXNIP can trigger caspase-1 activation, IL-1β and CXCL10 production, and improve chemo-immunotherapy efficacy.
Because IL-1β has some cytotoxic effects in mice (loss of weight) and that its injection to patients is difficult to consider, we aimed at finding inductors of TXNIP expression and consequently IL-1β production. By screening several chemical components capable of increasing TXNIP expression, we discovered 8 molecules able to increase TXNIP protein expression in NCSLC cell lines. Based on these results, we focused on 2 compounds, JNJ-26854165, an MDM2 inhibitor, and trametinib, a MEK inhibitor, which triggered caspase-1 activation, IL-1β cleavage and CXCL10 production, thus leading to CD8 T cell recruitment and tumor growth delay in the presence of chemo-immunotherapy.
In a previous work, we found that to obtain an anti-tumor effect of cisplatin/pemetrexed/anti-PD-L1, the KRAS (Kirsten Rat Sarcoma 2 viral oncogene homolog)/MEK/ERK (Extracellular signal-Regulated Kinase) pathway must be inhibited. The use of the MEK inhibitor trametinib enables chemotherapy to improve expression and production of CXCL10 by cancer cells, a chemokine that triggers the recruitment of CD8 T cells, thus enabling the anti-PD-L1 to be fully efficient in association with chemotherapy28. In this previous work, we showed that mtDNA was sensed by TLR9 to allow CXCL10 expression and production by lung cancer cells. Here, we provide further insights into the mechanism of action of trametinib. Indeed, mtDNA binds not only to TLR9 but also to AIM2. In cancer cells, mitochondria are highly subject to fusion/fission and may become defective. These abnormal mitochondria are degraded by mitophagy, thus enabling the release of mtDNA in the autophagolysosome, where it activates TLR9. Importantly, TLR9 is essential to enable AIM2 expression in cancer cells, thus suggesting that mitophagy could regulate AIM2 expression in lung cancer cells. Under treatments that enhance mitochondria destabilization, high amounts of mtDNA are then released in the cytosol and sensed by AIM2, thus leading to caspase-1 activation, IL-1β and CXCL10 production. We also highlighted here the importance of IL-1β in MEKi effects. Within tumors, IL-1β has been described to be mainly produced by myeloid cells and to be implicated in tumor growth2,37,38. However, in our lung mouse models, IL-1β is produced by cancer cells and acts in an autocrine manner to trigger therapeutic effects when associated with chemo-immunotherapy.
To conclude, our study shows an anti-tumor effect of IL-1β in lung cancer, in a chemo-immunotherapeutic context, which remains the first-line standard of treatment for lung cancer without oncogenic addiction. We also show the role of TXNIP in increasing IL-1β-mediated CXCL10 production and consequently chemo-immunotherapy efficacy, thus demonstrating the potential interest of combining TXNIP inductors with chemo-immunotherapy. These data warrant validation in clinical trials in human patients.
Methods
Cell culture
LLC1 (Lewis lung Carcinoma), J774A.1 (macrophage-like) mouse cells were obtained from the American Type Culture Collection (ATCC). KPC (PDAC) mouse cells were obtained from Ximbio. KP1.9 mouse cells were kindly provided by Prof. Alfred Zippelius (University Hospital, Basel). LLC1, J774A.1 and KPC cells were grown in DMEM with 4 g/L glucose and ultraglutamine (Dutscher) and KP1.9 in IMDM (Dutscher). All media were supplemented with 10% (vol/vol) fetal bovine serum (Dutscher) and with penicillin and streptomycin (Gibco), in an atmosphere of 95% air and 5% CO2 at 37 °C.
A427, A549, H460, H1650, H1792 and H1975 human cells were obtained from the ATCC. H322 and H358 human cells were obtained from Dr Laurence Dubrez, and H1299 and H2228 human cells from Prof. Beau Faller.
H322, H358, H460, H1299, H1650, H1792, H1975 and H2228 human lung cancer cells were cultured in RPMI with 10% (vol/vol) fetal bovine serum (Dutscher) supplemented with penicillin and streptomycin (Gibco). A549 lung carcinoma cells were cultured in DMEM with 10% (vol/vol) fetal bovine serum (Dutscher) supplemented with penicillin and streptomycin (Gibco). All cell lines were cultured at 37 °C under 5% CO2.
Stable Aim2, Casp1, Cxcl10, Il1b, Il1r1, Nlrp3, Tlr9 or Txnip-deficient LLC1 cells were generated by lentiviral-mediated expression of specific sgRNAs in two different doxycycline-inducible Cas9-expressing stable clones. A non-targeting sgRNA designated by sgRNA-irrelevant was used as a control. Pooled cell populations were used for experiments after blasticidin selection. The pCW-Cas9 lentiviral plasmid was a gift from Eric Lander & David Sabatini (Addgene # 5066139). The lenti-sgRNA blast lentiviral plasmid was a gift from Brett Stringer (Addgene #10499340). Primers designed to assess qPCR experiments are described in Supplementary Table 3.
LLC1 cells were transfected with pEF1A-mIl1b (a plasmid containing the sequence coding for mature IL-1β under the EF1A promoter (Vector Builder)) or GFP-TXNIP (a gift from Clark Distelhorst41 (Addgene plasmid # 18758; http://n2t.net/addgene:18758; RRID:Addgene_18758) using the Lipofectamine 3000 transfection reagent (Invitrogen), according to the manufacturer’s instructions. Stable TXNIP overexpressing LLC1 cells were selected 2 days after transfection with 500 µg/mL G418 (Invivogen).
Human lung cancer cells were transiently transfected with 0.5 µg of silencer select negative control siRNA (4390844, Thermofisher) or silencer select AIM2 siRNA (sc18092 and sc18093, Thermofisher) with opti-MEM medium, Lipofectamine 3000 reagent and P3000 reagent for 24 h before treatments.
The following reagents were used: Ac-YVAD-cmk, MCC950, Val-boroPro, Adenosine 5’-triphosphate disodium salt, Poly(dA:dT) / LyoVec (Invivogen), Anakinra (Sobi), recombinant mouse IL-1β (Miltenyi Biotech), Alisertib, GDC-0941, GSK1210151A, KW-2449, methylstat, MST-312, NVP-231, panobinostat, SNX-2112, sunitinib, YM-155, JNJ-26854165, foretinib, TW-37, GSK461364 and GSK436 (Coger), Trametinib, Selonsertib, Navitoclax, Venetoclax, JSH-23, Bortezomib (Selleckchem), Lipopolysaccharides (Ozyme), IFNAR-1 (Euromedex).
Mice
All animals were bred and maintained according to both the FELASA and the Animal Experimental Ethics Committee Guidelines (N° C 21 464 04 EA, University of Burgundy, France). Animals used were between 6 and 22 weeks of age. Female C57BL/6 mice (aged 6–8 weeks) were obtained from Charles River Laboratories. Il1r1-/- and Il1b-/- C57BL/6 mice42,43 were bred at the TAAM (Typage et Archivage d’Animaux Modèles – Orléans, France). All experiments were performed after permission of the local ethics committee of the University of Burgundy (agreement numbers: #53835 and #43998). The maximal tumor size (300mm2) permitted by the ethics committee was not exceeded, and mice were euthanized by cervical dislocation.
In vivo experiments
To induce subcutaneous tumor formation, 1.106 LLC1, KP1.9, KPC cells were injected subcutaneously into mice, and 7 days after tumor-cell injection (tumor size, 30 mm2), animals were treated.
For trametinib treatment, trametinib was prepared in a 0.5% HPMC 0.2% Tween80 solution. Mice received either HPMC or trametinib (1 mg/kg/day) per os, every day for 9 days. The treatment was started two days before chemotherapy.
For chemotherapy, mice received a single intraperitoneal (i.p.) injection of 6 mg per kg body weight of cisplatin (CDDP - Mylan) with 100 mg per kg of pemetrexed (Pem - Accord) or FOLFIRINOX (50 mg/kg Folinate, 25 mg/kg 5-FU, 25 mg/kg Irinotecan, 2.5 mg/kg Oxaliplatin).
For anti-PD-1, mice were injected i.p. three times a week with 10 mg/kg of anti-PD-1 Ab (BE0146 - BioXcell), starting from the beginning of chemotherapy treatment.
For some experiments, 100 mg/kg of Anakinra (Kineret - Sobi), 5 mg/kg anti-CXCR3 Ab (BE0249), 5 mg/kg anti-CD8β Ab (BE0223) or rat IgG1 isotype control (BioXcell) were injected i.p. three times a week, beginning with the chemotherapeutic treatment. In other experiments, mice received intratumoral injection of recombinant IL-1β (50 µg/kg every day for 9 days) (Miltenyi Biotec). Tumor size (Length x Width) was measured three times a week with a caliper, and tumor surface (L x W) was expressed in mm2.
For the lung metastasis model, mice were subcutaneously injected with 1.106 LLC1 (day 0). One week later, 1.106 LLC1 were intravenously injected in the tail vein to induce lung tumor formation. Some mice were treated with intratumor injections of recombinant IL-1β (from day 7 to day 15), one i.p. injection of cisplatin/pemetrexed (day 9) and i.p. injections of anti-PD-1 (from day 9, three times a week) with dosages indicated above. Lungs were collected at day 15 to evaluate the number of tumor foci.
The experiments were performed one or two times, and the total number of animals is indicated in the figure legend.
Ex vivo experiments
After dissection, tumors were mechanically and enzymatically dissociated using a mouse tumor dissociation kit, according to the manufacturer’s recommendations (130-096-730, Miltenyi Biotech). Tumor cells and myeloid cells were isolated by using, respectively, the Tumor Cell Isolation Kit (130-110-187) and CD11b MicroBeads (130-126-725) according to the manufacturer’s instructions (Miltenyi Biotech). Cells were then cultured for 4 hours at 37 °C, and supernatants were collected.
RT-qPCR
Cells or tumor biopsies were collected and lysed in Trizol reagent (Invitrogen), and total RNA was extracted. Three hundred nanograms of RNA were reverse-transcribed into cDNA using M-MLV reverse transcriptase, Random Primers and RNAseOUT inhibitor (Invitrogen). cDNA was quantified by real-time PCR using Power SYBR Green Real-time PCR kit (Applied Biosystem) on a QuantStudio 5 detection system (Applied Biosystems). Relative mRNA levels were determined using the ΔΔCt method. Values were expressed relative to Actb. Each independent experiment was performed in triplicate. The sequences of the oligonucleotides used are described in Supplementary Table 3.
Cytokine quantification
ELISA assays (R&D systems) were used to quantify mouse CXCL10 (DY466), human CXCL10 (DY266-05) and mouse IL-1β (MLB00C-1) according to the manufacturer’s information. Each independent experiment was performed in triplicate.
Flow cytometry experiments
To assess caspase-1 activation, 400,000 cells were treated and stained with FAM-FLICA Caspase-1 kit (BioRad) during the last hour of treatment at 37 °C. After trypsin detachment, cells were then stained with fixable viability dye (FVD) eFluor780 (Invitrogen) for 15 min at room temperature. After washing, cells were analyzed with a CytoFlex (Beckman Colter) and CytExpert software.
To evaluate IL1-R1 and R2 expression, cells were stained with anti-IL-1R1-PE (113505- Biolegend), anti-IL-1R2-FITC (orb466068 - Biorbyt) or corresponding control Ig (Biolegend) in the presence of FVD eFluor780 for 15 min at room temperature. After washing, cells were analyzed with a CytoFlex (Beckman Colter) and CytExpert software.
To assess cell death, 400,000 cells were treated and stained with Annexin V and DAPI (Miltenyi Biotech) according to the manufacturer’s instructions. After washing, cells were analyzed with a CytoFlex (Beckman Colter) and CytExpert software.
To study the infiltration of lymphoid and myeloid cells in the tumor tissue, tumors were collected and mechanically and enzymatically dissociated using a mouse tumor dissociation kit (130-096-730), according to the manufacturer’s instructions (Miltenyi Biotech). The tumor cell suspension (106 cells) was stained in Flow Cytometry Staining Buffer (FSB, eBioscience) with viability dye eFluor 780 and specific antibodies, for 15 min at RT in the dark, washed twice in FSB, and analyzed by flow cytometry. Flow cytometry acquisition was performed on a Fortesa cytometer (BD Biosciences). Flow Jo was used for the analysis. The following antibodies were used (Supplementary Table 4): anti-CD8a BUV805 (612898), anti-CD45 BUV395 (564279) from BD Biosciences, anti-CD11c FITC REAfinity (130-110-700), anti-Ly-6G APC REAfinity (130-120-734), anti-Ly-6C Viogreen (130-102-207), Miltenyi Biotec, anti-F4/80 Brilliant Violet 605 (123133), anti-CD3 Brilliant Violet 711 (100241), anti-CD11b Brilliant Violet 785 (101243) from Biolegend.
All the gating strategy are explained in Supplementary Fig. 27. Each independent experiment was performed in duplicate.
Cell viability assay
Cells (100,000) were seeded the day before treatment. After treatment, cells were washed with PBS and fixed with ethanol 100% for 30 min before crystal violet staining. Crystal violet was then resuspended in 50% ethanol/10% SDS, and OD was read at 575 nm with a Spark 10 M spectrophotometer (Tecan Life Sciences).
Western blotting
To detect the active form of IL-1β, cells were treated in OptiMEM without FBS. The supernatants (500 µL) were harvested by centrifugation for 5 min at 350 × g and precipitated using methanol (500 µL) and chloroform (150 µL). After centrifugation at 12,000 × g for 10 min., the aqueous phase (at the top) was discarded and 800 µL of methanol were added. Samples were centrifuged at 12,000 × g for 10 min, and the supernatants were removed. Pellets (containing proteins) were dried for 10 min at 37 °C, mixed with 20 µL of loading buffer (125 mM Tris-HCl [pH 6.8], 10% β-mercaptoethanol, 4.6% SDS, 20% glycerol, and 0.003% bromophenol blue) and incubated at 95 °C for 5 min.
To detect cellular proteins, whole-cell lysates were prepared by lysing cells in RIPA buffer (Sigma-Aldrich) in the presence of complete protease inhibitor (Roche). The protein concentration was assessed by a Lowry dosage (Biorad), and fifty µg of proteins were mixed with loading buffer.
Samples were separated by sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) with a Biorad Kit (1610180, 1610183, 1610185) and electroblotted to a nitrocellulose membrane (Amersham, GE Healthcare) with a Transfer Buffer from Biorad (10026938). After incubation for 1 hour at RT with 5% nonfat milk in phosphate-buffered saline (PBS)–0.1% Tween-20 or 5% Bovine Serum Albumin (BSA) in Tris Buffered Saline (TBS) 0.1% Tween-20, membranes were incubated overnight with the primary antibody diluted in PBS-milk-Tween or TBS-BSA-Tween, washed, incubated with the secondary antibody for 1 h at RT, and washed again before analysis with a chemiluminescence detection kit (Amersham, GE Healthcare) thanks to the Chemidoc device (Biorad). The following mouse mAbs were used (Supplementary Table 4): anti–β-actin (A1978) from Sigma-Aldrich, anti-mouse anti-caspase-1 (AG-20B-0044) from Adipogen and anti-COX IV (11967S) from Cell Signaling. The following rabbit pAbs were used (Supplementary Table 4): anti-AIM2 (Ab93015) from Abcam, anti-ASC (AG-25B-0006-C100) from Adipogen, anti-NLRP1 (NB100-56148ss) and anti-NLRP3 (NBP1-77080) from Novus, anti-TXNIP (40-3700) from Invitrogen, anti-cleaved-IL-1beta (Asp117) (63124S), anti-ASK1 (3762S) and anti-phospho-ASK1 (3764S) and anti-α-Tubulin (2144S) from cell signaling Technology. We also used rat pAb anti-IL-1β (MAB-4011) from R&D Systems. Secondary Abs HRP-conjugated polyclonal anti-mouse, anti-rabbit or anti-rat immunoglobulins (Jackson ImmunoResearch) were also used.
Immunoprecipitation
After washing with cold PBS, 20 million cells per sample were lysed for 10 minutes on ice and then for one hour at − 80 °C in lysis buffer (Hepes pH7 10 mM, KCl 100 mM, MgCl2 5 mM, EDTA 25 mM, DTT 2 mM, NP40 0.5%, protease inhibitor). The cell lysate was then recovered after centrifugation at 10,000 × g for 15 min. Immunoprecipitations were carried out using the Dynabeads Protein G Immunoprecipitation kit (10007D) from Invitrogen, according to the manufacturer’s recommendations. For immunoprecipitation, 1 µg of anti-AIM2 primary antibody (Abcam - Ab93015) was added to the beads. Immunoprecipitations with control Ig were performed in the same way as for the beads. Elution was done without denaturation in order to extract DNA and analyze mitochondrial DNA gene expression. Each independent experiment was performed in duplicate.
Cell fractionation
After washing with cold PBS, 15 million cells per sample were lysed for 30 minutes on ice in hypotonic lysis buffer (Sucrose 250 mM, Hepes pH7 20 mM, KCl 10 mM, MgCl2 1.5 mM, EDTA 1 mM, DTT 1 mM, protease inhibitor). The cells were then manually lysed with a syringe and a 26 G needle to reach approximately 50% dead cells. Two centrifugations at 750 g for 10 minutes were carried out before the supernatant was centrifuged at 10,000 × g for 30 min. The supernatant constitutes the cytosolic fraction, and the pellet, the mitochondrial fraction. The pellet was recovered in 50 µL of hypotonic lysis buffer. Sixty µg of protein per fraction were analyzed by Western Blotting to check the purity of the fractions. The expression of COX IV (mitochondria) and α-tubulin (cytosol) were analyzed. Each independent experiment was performed in duplicate.
DNA Extraction
DNA from cytosolic fractions was extracted with QIAamp DNA Micro Kit from Qiagen (56304) according to the manufacturer’s recommendations. The expression of the genes, CoxI, Dloop and Non-Numt, representative of the mitochondrial DNA or Tert, representative of nuclear DNA, was then analyzed by real-time PCR using Power SYBR Green Real-time PCR kit (Applied Biosystem) on a QuantStudio 5 detection system (Applied Biosystems).
Mitochondrial DNA (mtDNA) depletion
LLC1 cells were cultured in the presence of 150 ng/mL ethidium bromide (Sigma Aldrich) for 3 days before experiments.
Immunohistochemistry
Tumors were collected, fixed in PFA, and embedded in paraffin by the pathology laboratory. Four-micron slices were cut from formalin-fixed paraffin-embedded tumor samples. The tissues embedded in paraffin were cut on a Leica rotary microtome (RM2145). Four µm thick sections were deposited on a glass slide, deparaffinized and stained using a PT link (Agilent) and an Autostainer 48 (Agilent). Briefly, slides were deparaffinized using a pH9 buffer for 50 min at 95 °C. After cooling, slides were washed in wash buffer (Agilent) twice for 5 min. Peroxydase blocking was performed by incubating samples in S2023 solution (Agilent) for 5 min. Blocking was performed with Normal Goat Serum (MP-7451-50 kit, Vector Laboratories). Then anti-mouse CD8 (HS-361 00, Sysy Ab - Supplementary Table 4) antibody was added for 30 minutes at RT. Immpress HRP Goat polymers (MP-7451-50 kit Vector Laboratories) were added for 15 minutes at RT after two washing steps. DAB (SM803) was then added to the samples for 2 min. After two new washing steps, slides were finally incubated with hematoxylin (Enzo) for 20 min and permanently mounted using a Leica automated coverslipper. Slides were imaged with a Mantra Quantitative Pathology Workstation and analyzed using inForm Cell Analysis. CD8 and DAB staining was extracted. A mean of four or five areas for each slide (depending on tumor area) was calculated, and one representative image was chosen for each tumor group.
RNAseq on LLC1 cells
RNA Sequencing was performed by the “plateforme de génotypage et séquençage” (Institut du Cerveau, Paris) using NovaSeq X plus 1.5B reagent kit on a NovaSeq sequencer. Differential expression analysis was performed with the DESeq2 R package44. Untreated LLC1 cells or LLC1 cells treated with recombinant IL-1β (100 g/mL for 2 hours) were compared to each other. Raw p-values associated with each gene were adjusted using Bonferroni correction, as appropriate. For each comparison, genes with an adjusted p-value < 0.10 were selected as differential.
GEO/NCBI database analysis
Using existing transcriptomic studies on drug treatment effects in the GEO database, we searched for drugs that could induce TXNIP expression. More precisely, we used a transcriptomic database involving 719 transcriptomic profiles based on microarray or RNAseq studies, collected as previously described by Gu et al.27. In this way, the effect of 235 different drugs on human cancer cell lines was tested. For this purpose, the differential effect of TXNIP was tested using limma45 and DESeq244, respectively for microarray and RNA-seq. Sixteen molecules that were able to statistically increase TXNIP expression after p-value adjustment with the Benjamini-Hochberg46 method (FDR correction) were selected as possible TXNIP inducers (Supplementary Table 2).
Statistical analyses of in vivo and in vitro studies
In vitro results are shown as means ± standard deviations (SD.), in vivo results are shown as means ± standard error of the mean (SEM.) and comparisons of datasets were performed using GraphPad Prism 8. The following statistical tests were used: for tumor growth, two-way ANOVA; for in vitro experiments, nonparametric unpaired t test with FDR adjustment according to the Benjamini-Hochberg method. All P-values were two-tailed. All experiments were reproduced at least three times, and all attempts at replication were successful, with self-consistent results. The investigators were not blinded to allocation during the experiments or outcome assessment. No statistical method was used to predetermine the sample size. In vitro, the experiments were not randomized. In vivo, mice were randomized before treatments.
Biostatistical analysis on patient samples
Study population
We used two retrospective cohorts of patients with NSCLC receiving treatment with anti-programmed death 1 (PD-1/PD-L1) checkpoint inhibitors between 2014 and 2020 (Supplementary Table 1)18. Cohort 1 was composed of 92 patients treated in the Georges François Leclerc Cancer center, or the University Hospital of Dijon or the Hospital of Montréal; and cohort 2 was composed of 70 patients treated in the Georges François Leclerc Cancer center. We also used 506 patients with NSCLC from The Cancer Genome Atlas (TCGA), this cohort is hereafter named TCGA LUAD.
Ethics for cohorts 1 and 2
Only patients from whom informed consent was obtained and recorded in the medical chart were included in this retrospective study. This study was approved by the CNIL (French national commission for data privacy) and the local ethics committee, and was performed in accordance with the principles of Good Clinical Practice and the Declaration of Helsinki and European legislation. For RNAseq analysis, this study falls within the scope of the biological collection authorization registered under the numbers AC-2014-2260 (France) and #18.085 (Canada).
RNAseq data
For both cohorts, tumors were collected at diagnosis.
For cohort 1, total RNA was extracted from formalin-fixed paraffin-embedded (FFPE) tumor slices (5 × 5 µm) using the Maxwell 16 LEV RNA FFPE Purification kit (Promega) following the manufacturer’s instructions. Libraries were prepared from 12 µl of total RNA with the TruSeq Stranded Total RNA using Ribo-Zero (Illumina) following the manufacturer’s instructions. Once qualified, paired-end libraries were sequenced using 2 × 75 bp output on a NextSeq 500 device (Illumina).
For cohort 2, 10 ng of total RNA or 40 ng from FFPE were fragmented, and 3’-end of mRNA were captured using an RT primer poly(T) containing a UMI (Unique Molecular Identifier). Illumina adapters were added during this step by template switching. Fragments were then amplified with two PCR to complete Illumina adapters and add indexing sequences. Libraries were sequenced on the NovaSeq 6000 platform.
Abundance of transcripts was quantified using the Kallisto program47. This program is based on pseudo alignment for rapidly determining the compatibility of reads with targets, without the need for alignment. The Kallisto transcript index used as reference was built from merged human cDNA and ncDNA files from the GRCh37 assembly ENSEMBL. Gene-level count matrices were then created with the DESeq2 library44. Low-count genes were pre-filtered by removing genes with too few reads.
In the case of cohort 2, specific processing was used before applying the Kallisto program to take the UMI (Unique Molecular Identifier) into account. After a trimming step using cutadapt48, UMI sequences were retrieved using UMItools49. Processed reads were then aligned to the reference genome using STAR50, and the resulting aligned reads were deduplicated using UMItools again.
For the TCGA LUAD cohort, RSEM were downloaded on the GDC Data Portal (https://portal.gdc.cancer.gov/), and raw counts were downloaded using the R package TCGAbiolinks51 by selecting STAR counts. RSEM and counts were then normalized using the “median of ratios” method52. Variance stabilizing transformation was next applied on RSEM.
In each cohort, two transcriptomic signatures were computed as a metagene by taking the mean expression of corresponding genes: the Extended Immune gene signature (EIG) and Interferon gamma (IFNG)18.
In the TCGA LUAD cohort, 58 patients had paired normal and tumor samples, these patients were used to perform a ratio of tumor to normal gene expression.
Statistical analysis
Expression levels of genes of interest (IL1B, IL1R1, AIM2 and TXNIP) were transformed as dichotomous variables using the methodology of Lausen et al via the maxstat R library53.
The expression of transcriptomic signatures was compared using the Wilcoxon test according to dichotomized gene expression of genes of interest.
In cohorts 1 and 2, Progression-Free-Survival (PFS) was calculated from the date of first immunotherapy administration to recurrence or death from any cause, and evaluated at 6 months. Patients who were alive with no progression at 6 months were censored. Overall Survival (OS) was calculated from the date of first immunotherapy administration until death from any cause, and was censored at 12 months. In the TCGA LUAD cohort, PFS and OS were censored at 60 months54.
For each cohort, patients were considered responders when their PFS was greater than the cohort median survival.
Survival analysis was performed using the survival R library. The prognostic value of the different variables was tested using univariate Cox regression for PFS and OS. Proportional hazards assumptions were tested based on Schoenfeld residuals. Survival probabilities were estimated using the Kaplan-Meier method, and survival curves were compared using the log-rank test.
Differential gene expression analysis was performed using the DESeq2 R package44 according to responder status or normal and tumor samples. Gene Set Enrichment Analysis (GSEA)55 was performed on resulting differential genes according to responder status using Hallmarks of cancer gene sets from the Broad Institute and the R package “fgsea”56.
P-values ≤ 0.05 were considered statistically significant. When appropriate, P-values were adjusted using Benjamini-Hochberg FDR correction46, and adjusted p-values < 0.05 were considered significant.
Statistical analyses were performed using R software version 4.1.3 (http://www.R-project.org/), and graphs were drawn using GraphPad Prism version 8.0.2.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
RNA-Seq data were reused and previously deposited into the Gene Expression Omnibus database under accession numbers GSE190264, GSE190265, GSE190266. All other data are available in the article and its Supplementary files or from the corresponding author upon request. Source data are provided in this paper.
References
Gandhi, L. et al. Pembrolizumab plus chemotherapy in metastatic non-small-cell lung cancer. N. Engl. J. Med. 378, 2078–2092 (2018).
Hill, W. et al. Lung adenocarcinoma promotion by air pollutants. Nature 616, 159–167 (2023).
Roussot, N., Ghiringhelli, F. & Rébé, C. Tumor immunogenic cell death as a mediator of intratumor CD8 T-cell recruitment. Cells 11, 3672 (2022).
Ghiringhelli, F. & Rébé, C. Using immunogenic cell death to improve anticancer efficacy of immune checkpoint inhibitors: From basic science to clinical application. Immunological Rev. 321, 335–349 (2024).
Frederickson, A. M. et al. Pembrolizumab plus chemotherapy for first-line treatment of metastatic nonsquamous non-small-cell lung cancer: a network meta-analysis. Immunotherapy 11, 407–428 (2019).
Rébé, C. & Ghiringhelli, F. Interleukin-1β and Cancer. Cancers 12, E1791 (2020).
Perrichet, A., Ghiringhelli, F. & Rébé, C. Understanding inflammasomes and PD-1/PD-L1 crosstalk to improve cancer treatment efficiency. Cancers 12, E3550 (2020).
Krelin, Y. et al. Interleukin-1β–driven inflammation promotes the development and invasiveness of chemical carcinogen–induced tumors. Cancer Res. 67, 1062–1071 (2007).
Saijo, Y. et al. Proinflammatory cytokine IL-1β promotes tumor growth of Lewis Lung Carcinoma by induction of angiogenic factors: In vivo analysis of tumor-stromal interaction. J. Immunol. 169, 469–475 (2002).
Nakao, S. et al. Infiltration of COX-2–expressing macrophages is a prerequisite for IL-1β–induced neovascularization and tumor growth. J. Clin. Invest. 115, 2979–2991 (2005).
Huang, J. et al. Targeting the IL-1β/EHD1/TUBB3 axis overcomes resistance to EGFR-TKI in NSCLC. Oncogene 39, 1739–1755 (2020).
Tian, T. et al. IL1α Antagonizes IL1β and promotes adaptive immune rejection of malignant tumors. Cancer Immunol. Res 8, 660–671 (2020).
Ridker, P. M. et al. Effect of interleukin-1β inhibition with canakinumab on incident lung cancer in patients with atherosclerosis: exploratory results from a randomised, double-blind, placebo-controlled trial. Lancet 390, 1833–1842 (2017).
Tan, D. S. W. et al. Canakinumab versus placebo in combination with first-line pembrolizumab plus chemotherapy for advanced non–small-cell lung cancer: Results From the CANOPY-1 Trial. JCO 42, 192–204 (2024).
Paz-Ares, L. et al. Canakinumab in combination with docetaxel compared with docetaxel alone for the treatment of advanced non-small cell lung cancer following platinum-based doublet chemotherapy and immunotherapy (CANOPY-2): A multicenter, randomized, double-blind, phase 3 trial. Lung Cancer 189, 107451 (2024).
Garon, E. B. et al. Canakinumab as adjuvant therapy in patients with completely resected non–small-cell lung cancer: Results from the CANOPY-A double-blind, randomized clinical trial. JCO 42, 180–191 (2024).
Mok, T. S. K. et al. LBA4 CANOPY-N: A phase II study of canakinumab (CAN) or pembrolizumab (PEM), alone or in combination, as neoadjuvant therapy in patients (pts) with resectable stage Ib–IIIa non-small cell lung cancer (NSCLC). Ann. Oncol. 33, S1547–S1548 (2022).
Lecuelle, J. et al. MER4 endogenous retrovirus correlated with better efficacy of anti-PD1/PD-L1 therapy in non-small cell lung cancer. J. Immunother. Cancer 10, e004241 (2022).
Ayers, M. et al. IFN-γ–related mRNA profile predicts clinical response to PD-1 blockade. J. Clin. Invest. 127, 2930–2940 (2017).
Evavold, C. L. & Kagan, J. C. Inflammasomes: Threat-assessment organelles of the innate immune system. Immunity 51, 609–624 (2019).
Dang, E. V., McDonald, J. G., Russell, D. W. & Cyster, J. G. Oxysterol restraint of cholesterol synthesis prevents AIM2 inflammasome activation. Cell 171, 1057–1071.e11 (2017).
Rathinam, V. A. K. & Fitzgerald, K. A. Inflammasome complexes: Emerging mechanisms and effector functions. Cell 165, 792–800 (2016).
Miller, K. N. et al. Cytoplasmic DNA: sources, sensing, and role in aging and disease. Cell 184, 5506–5526 (2021).
McArthur, K. et al. BAK/BAX macropores facilitate mitochondrial herniation and mtDNA efflux during apoptosis. Science 359, eaao6047 (2018).
Choi, E.-H. & Park, S.-J. TXNIP: A key protein in the cellular stress response pathway and a potential therapeutic target. Exp. Mol. Med. 55, 1348–1356 (2023).
Bronner, D. N. et al. Endoplasmic reticulum stress activates the inflammasome via NLRP3- and Caspase-2-driven mitochondrial damage. Immunity 43, 451–462 (2015).
Gu, S. S. et al. Therapeutically increasing MHC-I expression potentiates immune checkpoint blockade. Cancer Discov. 11, 1524–1541 (2021).
Limagne, E. et al. MEK inhibition overcomes chemoimmunotherapy resistance by inducing CXCL10 in cancer cells. Cancer Cell 40, 136–152 (2022).
Shimada, K. et al. Oxidized mitochondrial DNA activates the NLRP3 inflammasome during apoptosis. Immunity 36, 401–414 (2012).
Li, Z. et al. Dual effect of tryptamine on prostate cancer cell growth regulation: A pilot study. Int. J. Mol. Sci.23, 11087 (2022).
Lythgoe, M. P. & Prasad, V. Repositioning canakinumab for non-small cell lung cancer—important lessons for drug repurposing in oncology. Br. J. Cancer 127, 785–787 (2022).
Qi, M. et al. AIM2 promotes the development of non-small cell lung cancer by modulating mitochondrial dynamics. Oncogene 39, 2707–2723 (2020).
Zhang, M. et al. AIM2 promotes non-small-cell lung cancer cell growth through inflammasome-dependent pathway. J. Cell. Physiol. 234, 20161–20173 (2019).
Qin, Y. et al. Pan-cancer analysis of AIM2 inflammasomes with potential implications for immunotherapy in human cancer: A bulk omics research and single cell sequencing validation. Front. Immunol.13, https://doi.org/10.3389/fimmu.2022.998266 (2022).
Bodnar-Wachtel, M. et al. Inflammasome-independent NLRP3 function enforces ATM activity in response to genotoxic stress. Life Sci. Alliance6, https://doi.org/10.26508/lsa.202201494 (2023).
Koenen, T. B. et al. Hyperglycemia activates caspase-1 and TXNIP-mediated IL-1β transcription in human adipose tissue. Diabetes 60, 517–524 (2011).
Park, M. D. et al. Hematopoietic aging promotes cancer by fueling IL-1α–driven emergency myelopoiesis. Science 386, eadn0327 (2024).
Caronni, N. et al. IL-1β+ macrophages fuel pathogenic inflammation in pancreatic cancer. Nature 623, 415–422 (2023).
Wang, T., Wei, J. J., Sabatini, D. M. & Lander, E. S. Genetic screens in human cells using the CRISPR-Cas9 system. Science 343, 80–84 (2014).
Stringer, B. W. et al. A reference collection of patient-derived cell line and xenograft models of proneural, classical and mesenchymal glioblastoma. Sci. Rep. 9, 4902 (2019).
Wang, Z. et al. Thioredoxin-interacting protein (txnip) is a glucocorticoid-regulated primary response gene involved in mediating glucocorticoid-induced apoptosis. Oncogene 25, 1903–1913 (2006).
Labow, M. et al. Absence of IL-1 signaling and reduced inflammatory response in IL-1 type I receptor-deficient mice. J. Immunol. 159, 2452–2461 (1997).
Horai, R. et al. Production of mice deficient in genes for interleukin (IL)-1α, IL-1β, IL-1α/β, and IL-1 receptor antagonist shows that IL-1β is crucial in turpentine-induced fever development and glucocorticoid secretion. J. Exp. Med. 187, 1463–1475 (1998).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B 57, 289–300 (1995).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527 (2016).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet. J. 17, 10–12 (2011).
Smith, T., Heger, A. & Sudbery, I. UMI-tools: modeling sequencing errors in Unique Molecular Identifiers to improve quantification accuracy. Genome Res. 27, 491–499 (2017).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics29, 15–21 (2013).
Colaprico, A. et al. TCGAbiolinks: an R/Bioconductor package for integrative analysis of TCGA data. Nucleic Acids Res. 44, e71 (2016).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Hothorn, T. & Lausen, B. On the exact distribution of maximally selected rank statistics. Comput. Stat. Data Anal. 43, 121–137 (2003).
Liu, J. et al. An integrated TCGA pan-cancer clinical data resource to drive high-quality survival outcome analytics. Cell 173, 400–416 (2018).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Korotkevich, G. et al. Fast gene set enrichment analysis. 060012 Preprint at https://doi.org/10.1101/060012 (2021).
Acknowledgements
This study was supported by grants from the ‘Institut National du Cancer’ (INCa) PLBIO2020-104 (F.G.) and PLBIO23-124-2023-163 (F.G. and C.R.), the ‘Ligue Nationale Contre le Cancer’ (F.G.) and by a French Government grant managed by the French National Research Agency under the program ‘Investissements d’Avenir’, reference ANR-11-LABX-0021 (Lipstic Labex). FG’s team is ‘Equipe labellisée Ligue Nationale Contre le Cancer’. A.P. is a recipient of grants from the ‘Région Bourgogne-Franche-Comté’ and the ‘Association pour la Recherche sur le Cancer’ (ARC). We acknowledge F. Ecarnot, PhD, for helping with the preparation of the paper and correction of the English. We thank the animal housing facility at the University of Burgundy and the Imaflow platform (Dijon, France). The authors would like to thank Integragen (Genopole Campus 1, Evry) and R. Boidot and S. Chevrier (Georges-François Leclerc Center) for the generation of sequencing data. The mouse within Figs. 1, 2, 4, 5 and 6 was created in BioRender (Limagne, E. (2025) https://BioRender.com/6q83rg6).
Author information
Authors and Affiliations
Contributions
A.P. contributed to the design of the study, analyzed data and edited the manuscript. A.P. performed or contributed to all experiments, with help from M.T., H.B., P.J., P.R., F.Ga., N.R., X.Y., T.V. and L.N. J.L. performed all bioinformatics analyses, under the supervision of C.T. R.A. and A.A. developed the CRISPRCas9 cells. A.I. and D.R. performed IHC experiments, with help from V.D. and T.H. for computational analysis. A.Z. provided resources. F.Gh. and B.R. conducted the trials on NSCLC. E.L. and F.C. provided key scientific inputs. C.R. prepared figures. C.R. and F.Gh. conceptualized and coordinated the study, acquired funding, analyzed the data and wrote the paper. All authors read and edited the manuscript and approved the final version of the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Damya Laoui, who co-reviewed with Sofie Deschoemaeker and Lize Allonsius; Renato Ostuni and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Perrichet, A., Lecuelle, J., Limagne, E. et al. Cancer cell-derived IL-1β reverses chemo-immunotherapy resistance in non-small cell lung cancer. Nat Commun 16, 10244 (2025). https://doi.org/10.1038/s41467-025-64839-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-64839-4