Abstract
Despite the well-known role of the transforming growth factor-β (TGFβ) pathway in cancer progression, therapies targeting it have largely failed in the clinic. This suggests our understanding of TGFβ’s function is incomplete. Here we show that this therapeutic failure is rooted in a fundamental paradox: while TGFβ promotes malignant traits in gastric cancer, many cancer cells remain sensitive to its potent tumor-suppressive effects. We uncover that this suppression works by impairing the cell’s energy metabolism through modulating dihydrolipoamide dehydrogenase (DLD). Therefore, broadly blocking the TGFβ pathway can inadvertently release a natural brake on tumor growth. Based on this insight, we demonstrate that co-targeting this metabolic vulnerability with an inhibitor (devimistat) alongside an anti-TGFβ agent significantly enhances therapeutic efficacy in gastric cancer models. This combination approach presents a promising strategy to overcome the limitations of current therapies.
Similar content being viewed by others
Introduction
Gastric cancer (GC) represents a significant global health challenge due to its insidious onset, late diagnosis, high heterogeneity, rapid progression, and poor response to available treatments1, and improving therapeutic outcomes in GC requires a better understanding of the malignant features that worsen as the disease progresses. In this regard, although the transforming growth factor-β (TGFβ) pathway has been associated with the high metastatic potential and poor prognosis of GC2,3, our understanding of the role of TGFβ in tumorigenesis and progression remains unclear and context-dependent, with evidence showing contradictory effects4,5. Despite this, targeted therapies against the TGFβ pathway have yet to demonstrate satisfactory response rates or survival benefits in clinical trials, and no such therapies have been approved for cancer treatment6,7. Therefore, further research is essential to address the limitations of anti-TGFβ therapy and to optimize its integration into GC treatment regimens.
Tumors develop within a complex microenvironment where various physicochemical properties influence the hallmarks of cancer8. Emerging evidence, including our findings, indicates that microenvironmental signals can affect therapeutic response by altering cancer metabolism9,10. TGFβ, a key signaling molecule in the tumor microenvironment (TME), has been implicated in regulating diverse aspects of cancer metabolism, including glucose, amino acid, and lipid metabolism through transcriptional regulation of key metabolic regulators, transporters, and enzymes (such as PGC-1α, GLUT-1, and FASN)11,12,13. However, this metabolic regulation exhibits significant heterogeneity across different cancer types and microenvironmental conditions. Importantly, the mechanistic basis of TGFβ-mediated metabolic control, particularly through post-translational modification (PTM)-regulated enzyme activity, remains poorly understood14,15. Furthermore, understanding how metabolic mechanisms contribute to the limited efficacy of anti-TGFβ therapy and whether combining metabolic interventions with this therapy could enhance treatment effectiveness remains to be explored.
In this study, we demonstrate that while the malignant characteristics mediated by the TGFβ pathway become more pronounced during gastric cancer (GC) progression and are associated with poor prognosis, many GC cells retain sensitivity to the tumor-suppressive effects of TGFβ. This dual nature of TGFβ contributes to the unintended pro-tumor effects observed with anti-TGFβ therapies. Moreover, we identified that metabolic regulation plays a critical role in mediating TGFβ‘s tumor-suppressive effects. Specifically, TGFβ induces the interaction between acetyl-CoA acetyltransferase 1 (ACAT1) and the phosphorylated SMAD3/3/4 complex, preventing its translocation into the mitochondria. This interaction downregulates the acetylation of dihydrolipoamide dehydrogenase (DLD) at lysine 430 (K430), thereby inhibiting its dimerization and integration into the pyruvate dehydrogenase (PDH) and α-ketoglutarate dehydrogenase (α-KGDH) enzyme complexes, ultimately disrupting cellular energy metabolism. Importantly, co-targeting DLD effectively mitigates the pro-proliferative side effects of anti-TGFβ therapy, thereby enhancing its efficacy in GC models. Collectively, this study identifies the ACAT1/DLD axis as a metabolism-centered effector of TGFβ‘s tumor-suppressive function and proposes a combinatorial strategy to improve therapeutic outcomes in GC.
Results
Antagonizing TGFβ signaling fails to improve therapeutic outcomes in GC models
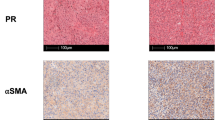
To identify the malignant features that accumulate progressively during GC onset and progression, we performed bulk RNA sequencing on 12 paired samples of GC tissues, adjacent normal gastric tissues, and peritoneal metastases (Fig. 1A). Our analysis revealed that epithelial-mesenchymal transition (EMT) was one of the most significantly enriched pathways and was progressively upregulated in the sequential samples of GC progression (Fig. 1B, Supplementary Fig. 1A), underscoring its critical role in driving GC malignancy. Given the established role of the TGFβ pathway in EMT16, we examined public data from the TCGA and GTEx databases and found that components of the canonical TGFβ pathway were upregulated in GC (Supplementary Fig. 1B). Notably, elevated expression of TGFB1 and TGFBR1 correlated with poor overall survival (Supplementary Fig. 1C). We further validated these findings through immunohistochemical (IHC) analysis of paired GC tissues, adjacent non-cancerous tissues, and peritoneal metastasis samples from the Sun Yat-sen University Cancer Center (SYSUCC; Supplementary Table 1). This analysis confirmed that TGFβ1 and TβR1 expression progressively increased with disease progression and were inversely correlated with overall survival in GC patients (Fig. 1C–E).
A Sample sources for bulk RNA sequencing. Created with BioRender.com. https://BioRender.com/z5unugi. B Top 5 pathways from GSEA, showing significant NES differences in tumor vs. normal tissue (left) and metastasis vs. primary tumor (right). Representative IHC images and score analysis for TGFβ1 (C) and TβR1 (D) in paired GC (n = 97), normal gastric (n = 97), and peritoneal metastatic tissues (n = 16) from the SYSUCC-GC cohort. Scale bar = 100 μm. Centerline, median; box limits, upper and lower quartiles; whiskers, 1.5× interquartile range; two-sided paired t-test. E Kaplan‒Meier survival analysis in SYSUCC-GC cohort stratified by IHC scores for TGFβ1 (left) and TβR1 (right). F Establishment and treatment timeline for orthotopic GC-PDX model. G Metastasis incidence in autopsies of orthotopic GC-PDX mice. H Kaplan‒Meier survival analysis of GC-PDX model mice with indicated treatments (n = 10 mice per group, two-sided log-rank test). I Representative 18FDG PET/CT images of PDX growth, showing orthotopic xenografts (orange circles) and liver metastasis (magenta circles), n = 4 mice for the control group, n = 3 for the galunisertib group. J Tumor volume and SUVmean of 18FDG uptake in 3D ROIs. SUV, standardized uptake value. Data presented as mean ± SD, two-sided Student’s t-test. K Growth of GC/CRC cell lines with or without 10 ng/mL TGFβ1, normalized to control, n = 3 independent experiments, data presented as mean ± SD, two-sided Student’s t-test. L–N HGC27 cells were injected into BALB/c-nu mice, treated with control, galunisertib, or vactosertib. Tumor images (L) with volume (M), and weight (N) analyses on day 27; n = 6 mice per group, scale bar = 1 cm. Data presented as mean ± SD, two-sided one-way ANOVA with Tukey’s multiple comparison test. O, P Representative bright-field images of GC patient-derived organoids with diameter analysis under TGFβ1 (10 ng/mL), galunisertib (20 μM), vactosertib (2 μM), or combination treatments (n = 3 independent experiments). Data are presented as mean ± SD; two-tailed one-way ANOVA with Tukey’s multiple comparison test.
To evaluate the potential of targeting TGFβ signaling in GC therapy, we established orthotopic patient-derived xenograft (PDX) models and treated them with galunisertib or vactosertib, two TβR1 inhibitors currently in clinical trials (Fig. 1F)17,18,19. Despite initial reductions in metastasis observed at autopsy, these anti-TGFβ treatments did not translate into improved survival outcomes; rather, there was a trend toward worse survival in the treatment groups (Fig. 1G, H). Additionally, 18F-fluorodeoxyglucose (18FDG) PET/CT scans revealed that while anti-TGFβ treatment reduced metastatic signals, the orthotopic tumors in these mice exhibited faster growth and increased metabolic activity (Fig. 1I, J), indicating a potential paradoxical effect of anti-TGFβ therapy in promoting tumor growth.
Although it is generally believed that cancer cells must escape TGFβ’s tumor-suppressive effects to evolve malignantly20, our findings suggest that TGFβ’s tumor-suppressive role is maintained in many GC cells. To investigate this further, we assessed the growth of various gastric and colorectal cancer cell lines in response to TGFβ1 stimulation in vitro. While most colorectal cancer cells escaped TGFβ’s tumor-suppressive effects, the majority of GC cells retained sensitivity to TGFβ-induced growth inhibition (Fig. 1K). This sensitivity may be linked to fewer mutations in TGFβ pathway components in their genomic profiles (Supplementary Fig. 1D, E). While confirming the on-target drug effect, we observed this sensitivity in both galunisertib or vactosertib-treated HGC27-derived subcutaneous tumors in vivo and in patient-derived GC organoid cultures with intact TGFβ signaling components in vitro (Fig. 1L–P, Supplementary Fig. 1F–H). In contrast, this sensitivity was absent in TGFβ signaling-deficient HCT116 tumors (Supplementary Fig. 1I–K), further confirming the requirement for an intact TGFβ pathway. These results suggest that TGFβ’s tumor-suppressive effect is preserved in a significant proportion of GC cells and that disruption of this effect by anti-TGFβ therapies may reduce their effectiveness.
TGFβ restricts GC metabolism by suppressing the TCA cycle enzymes PDH and α-KGDH
To gain a deeper understanding of the mechanisms through which TGFβ inhibits GC cell proliferation and how anti-TGFβ therapies might disrupt this effect, we investigated the role of metabolic rewiring in TGFβ-mediated growth inhibition. Given our observation of increased 18FDG uptake in orthotopic tumors treated with galunisertib and the essential role of metabolism in supporting cell growth, we aimed to explore whether TGFβ affects GC metabolism. To obtain a comprehensive assessment of TGFβ’s metabolic effects, we employed non-targeted metabolomics profiling. This analysis revealed significant alterations in the metabolic profiles of GC cells treated with TGFβ1 compared to control cells (Fig. 2A, B, Supplementary Fig. 2A, B, Supplementary Data 1). Metabolite set enrichment analysis21 indicated that the differential metabolites were predominantly centered around the TCA cycle (Fig. 2C, D). Seahorse real-time cell metabolic analyses further revealed that TGFβ1 significantly reduced the oxygen consumption rate (OCR) in GC cells, suggesting a decreased flux through the TCA cycle. Notably, this reduction was effectively counteracted by the anti-TGFβ agent galunisertib (Fig. 2E, F).
A Workflow for non-targeted metabolomics profiling of HGC27 cells treated w/wo TGFβ1 for 6 h. Created with BioRender.com. https://BioRender.com/z5unugi. B PCA of metabolites from TGFβ1-treated and untreated cells (n = 1 experiment with 10 technical replicates per group). C Top 15 enriched pathways from MSEA. D Network analysis of metabolic pathways from profiling. E, F OCR in HGC27 and MKN74 cells treated w/wo TGFβ1 and/or galunisertib for 6 h using Seahorse XFe24. Rot/AA, rotenone, and antimycin A; n = 4 independent experiments, data presented as mean ± SD. Schematic of carbon flux from U-13C6-glucose (G) or U-13C5-glutamine (H) into TCA intermediates. Abundance of labeled metabolites and product-to-substrate ratios for PDH and α-KGDH reactions in HGC27 cells treated w/wo TGFβ1 for 6 h post-tracing with U-13C6-glucose (I, J) or U-13C5-glutamine (K, L); n = 5 independent experiments, data presented as mean ± SD, two-sided Student’s t-test. M–O Relative activities of PDH, CS, and α-KGDH in HGC27, MKN74, and two patient-derived organoids treated w/wo TGFβ1/galunisertib for 6 h (TGFβ1 = 10 ng/mL, galunisertib = 20 μM, unless specified); n = 3 independent experiments, data presented as mean ± SD, two-sided one-way ANOVA with Tukey’s multiple comparison test. TGFβ1 and galunisertib concentrations were 10 ng/mL and 20 μM, respectively, unless otherwise specified.
To identify the specific steps in the TCA cycle affected by TGFβ1, we conducted [U-13C6]-glucose and [U-13C5]-glutamine tracing assays to analyze the mass isotopologue distribution of TCA cycle intermediates in GC cells (Fig. 2G, H). Consistent with our previous findings, we observed a general reduction in the abundance of [U-13C6]-glucose-labeled intermediates in the TGFβ1-treated group, with the exceptions of M + 3 Pyr and M + 2 α-KG (Fig. 2I, Supplementary Fig. 2C, Supplementary Data 2). Notably, there were significant decreases in the conversion of M + 3 Pyr to M + 2 Cit and M + 2 α-KG to M + 2 Suc, as evidenced by decreased product/substrate ratios (Fig. 2J). Similarly, [U-13C5]-glutamine tracing revealed impaired TCA cycle flux, with decreased conversion of M + 5 α-KG to M + 4 Suc and M + 0 Pyr to M + 4 Cit under TGFβ1 stimulation (Fig. 2K, L, Supplementary Fig. 2D, Supplementary Data 3). Since these reactions involve three key enzymes in the rate-limiting and usually irreversible steps of the TCA cycle—pyruvate dehydrogenase (PDH), citrate synthase (CS), and α-ketoglutarate dehydrogenase (α-KGDH)—we next examined the impact of TGFβ1 on these enzymes. Enzyme activity measurements revealed that TGFβ1 significantly inhibited the activities of PDH and α-KGDH in GC cell lines and organoids, whereas the activity of CS remained unchanged (Fig. 2M–O). These findings demonstrate that TGFβ suppresses TCA cycle flux by simultaneously inhibiting the critical enzymes PDH and α-KGDH.
TGFβ inhibits PDH and α-KGDH by reducing the acetylation of the E3 enzyme DLD at K430
Next, we investigated how the simultaneous regulation of PDH and α-KGDH by TGFβ was achieved. Since both PDH and α-KGDH are organized as enzyme complexes comprising E1, E2, and E3 components22, we examined their composition. We found that the two complexes utilize distinct E1 and E2 subunits but share dihydrolipoamide dehydrogenase (DLD) as a common E3 component (Fig. 3A). We then focused on DLD and found that its knockdown significantly impaired the activities of PDH and α-KGDH, reduced the OCR, and inhibited the proliferation of GC cells in vitro (Supplementary Fig. 3A–D), highlighting the important role of DLD in maintaining TCA cycle flux and cell viability. While galunisertib effectively reversed the decline in PDH and α-KGDH activity induced by TGFβ1 and mitigated the reduced growth of wild-type (WT) GC cells, it failed to restore these phenotypes in GC cells with DLD knockdown (Supplementary Fig. 3E, F). Moreover, galunisertib promoted tumor growth in vivo only in subcutaneous tumors derived from WT GC cells, not in those with DLD knockdown (Supplementary Fig. 3G–I). These results suggest that DLD is crucial for TGFβ-mediated growth inhibition.
A Composition of PDH and α-KGDH complexes. B Co-IP analysis of lysine acetylation (Kac), succinylation (Ksucc), and phosphorylation (pSer, pThr, pTyr) of DLD in HGC27 and MKN74 cells w/wo TGFβ1 for 6 h. The samples derive from the same experiment but different gels were processed in parallel: separate gels were used for the IP detection of Kac, Ksucc, pSer, pThr, pTyr, and DLD. For the input controls, one gel was used for Vinculin and DLD, and another for pSMAD2/3 and SMAD2/3. C Co-IP analysis of DLD acetylation in HEK293T cells with DLD mutations ± TGFβ1 for 6 h. The samples derive from the same experiment but different gels were processed in parallel: separate gels were used for the IP detection of Kac and HA. For the input controls, one gel was used for Vinculin and HA, and another for pSMAD2/3 and SMAD2/3. D MS/MS spectrum of acetylated “AK (Acetyl) TNADTDGMVK”. E Sequence alignment near DLD K430 among species. F PDH and α-KGDH activity in HEK293T cells with endogenous DLD knockdown and transfected DLD variants; n = 3 independent experiments, data presented as mean ± SD, two-sided two-way ANOVA with Tukey’s multiple comparison test. G OCR of HGC27 cells with endogenous DLD knockdown and transfected DLD variants, measured by Seahorse XFe24; n = 4 independent experiments, data presented as mean ± SD. H Growth of HGC27 cells transfected with DLD variants w/wo TGFβ1, assessed by cell counts; n = 3 independent experiments, data presented as mean ± SD, two-sided two-way ANOVA with Tukey’s multiple comparison test. I Design for subcutaneous injection of HGC27 cells with DLD variants into BALB/c-nu mice, followed by control or galunisertib treatment. Subcutaneous tumor images (J) and quantification of tumor volume (K) and weight (L) in indicated treatment groups on day 27; n = 6 mice per group, data presented as mean ± SD, two-sided two-way ANOVA with Tukey’s multiple comparison test, scale bar = 1 cm. M Immunoblot of DLD-K430ac in HGC27 and MKN74 cells under the indicated treatments for 6 h. The samples derive from the same experiment but different gels were processed in parallel: one gel was used for DLD-K430ac, DLD, and Vinculin, and another for pSMAD2/3 and SMAD2/3. Representative images (N) and statistical analysis (O) of IHC staining of DLD-K430ac in paired primary GC, normal gastric tissue, and peritoneal metastasis from the SYSUCC-GC cohort, n = 97 for the normal group, n = 97 for the primary GC group, n = 16 for the peritoneal metastasis group. Scale bar = 100 μm. Centerline, median; box limits, upper and lower quartiles; whiskers, 1.5× interquartile range; two-sided paired t-test. TGFβ1 and galunisertib concentrations were 10 ng/mL and 20 μM, respectively, unless otherwise specified. Images in (B, C, M) are representative of two independent experiments. The samples are derived from the same experiment but different gels for Kac, Ksucc, pSer, pThr, pTyr, and DLD, and B, another for protein C, and another for β-actin, were processed in parallel.
To explore how TGFβ modulates DLD, we assessed the mRNA and protein levels of DLD and other components of the PDH and α-KGDH complexes and observed no significant changes following TGFβ1 treatment (Fig. 3B, Supplementary Fig. 4A, B). Given the important role of post-translational modifications in enzyme regulation23, we detected the levels of several common types of post-translational modifications on DLD with or without TGFβ1 treatment and observed a marked decrease in acetylation instead of other common modifications such as phosphorylation or succinylation (Fig. 3B).
Acetylation primarily targets lysine residues within proteins24. Using LC-MS/MS, we identified 10 lysine residues on DLD that are potential sites for acetylation (Supplementary Fig. 4C). We individually mutated each of these lysines to non-acetylatable arginine (R) residues and evaluated the acetylation level of each mutant with and without TGFβ1 treatment using a pan-acetyllysine antibody. We noticed that the acetylation level of DLD could not be further reduced by TGFβ1 when K430 was mutated, and this acetylatable lysine is evolutionarily conserved (Fig. 3C–E). To further examine K430’s role, we reintroduced shRNA-resistant WT, K430R, or K430Q (mimicking hyperacetylation) DLD proteins into GC cells with depleted endogenous DLD, using the K445R mutant as a control. The K430R mutation markedly inhibited, while the K430Q mutation significantly enhanced the enzyme activity of PDH and α-KGDH. Importantly, TGFβ1 failed to modulate enzyme activity when K430 was mutated (Fig. 3F). Cells expressing the DLDK430Q mutant exhibited increased OCR, whereas those expressing DLDK430R showed decreased OCR (Fig. 3G, Supplementary Fig. 4D). These metabolic effects translated into functional outcomes: both in vitro proliferation assays and subcutaneous tumor models showed consistent growth patterns (Fig. 3H–L, Supplementary Fig. 4E). In contrast, the K445R mutant behaved similarly to WT (Supplementary Fig. 4F, G), underscoring the specificity of DLD-K430 acetylation in TGFβ-mediated proliferation control. Notably, metabolic regulation via modulation of DLD-K430 acetylation also interferes with the well-documented cytostatic mechanism of TGFβ, which acts through transcriptional regulation of CDK inhibitors and MYC (Supplementary Fig. 4H), suggesting crosstalk between TGFβ’s metabolic and transcriptional regulatory roles.
Next, we generated an antibody specifically recognizing acetylated DLD at K430 (DLD-K430ac) and confirmed its specificity by dot blotting and IHC (Supplementary Fig. 4I, J). Using this antibody, we observed a significant reduction in DLD-K430ac levels in GC cells upon TGFβ1 stimulation, which was reversed by galunisertib (Fig. 3M). IHC analysis further demonstrated that DLD-K430ac levels were significantly lower in GC tissues compared to paired normal gastric tissues, with the lowest levels observed in peritoneal metastases (Fig. 3N, O). Notably, DLD-K430ac levels showed a modest but statistically significant negative correlation with TGFβ1 expression (Supplementary Fig. 4K). And reverse to the trend indicated by TGFβ1/TβR1, the level of DLD-K430ac was positively associated with the overall survival of GC patients (Supplementary Fig. 4L). These findings suggest that TGFβ1 inhibits PDH and α-KGDH by reducing the acetylation of the common E3 enzyme DLD at K430, and the TGFβ1-DLD-K430ac axis correlates with the clinicopathological features of GC.
Acetylation at K430 facilitates the dimerization of DLD and the assembly of PDH and α-KGDH complexes
Next, we investigated how acetylation of DLD at K430 affects the catalytic activity of PDH and α-KGDH. Notably, K430 belongs to the interface domain crucial for DLD dimerization, rather than within the substrate-binding domains (Fig. 4A). Given that PDH and α-KGDH complex assembly relies on E3 dimers interacting with an oligomeric E2 core (Fig. 4B)25,26, and considering the essential role of the interface domain in maintaining enzymatic activity27, we hypothesized that K430 acetylation might influence DLD dimer formation. This perturbation could subsequently affect the structural integrity and catalytic function of both enzyme complexes. Immunoprecipitation assays revealed that the ability of HA-tagged DLD to pull down its Myc-tagged counterpart was strongest for the hyperacetylation-mimicking K430Q mutant and weakest for the deacetylation-mimicking K430R mutant (Fig. 4C). Protein cross-linking assays further demonstrated increased dimer formation for the K430Q mutant and decreased dimerization for the K430R mutant (Fig. 4D). While TGFβ1 treatment reduced dimerization of WT DLD, this effect was not observed in the K430Q mutant (Fig. 4E). Additionally, TGFβ1 stimulation decreased the levels of endogenous dimeric DLD in HGC27 and MKN74 GC cells and this effect was reversed by galunisertib treatment (Fig. 4F). Molecular dynamics simulations suggested that the state of residue K430 affects local interactions within the DLD dimer (Supplementary Fig. 5A). Root Mean Square Deviation (RMSD) analysis revealed that the K430R dimer exhibited the greatest deviation from the reference conformation, indicating lower stability, whereas the K430Q dimer was the most stable (Fig. 4G). Furthermore, Molecular Mechanics/Generalized Born Surface Area (MM/GBSA) calculations showed that the K430Q dimer had a more negative binding energy than the K430R dimer (Fig. 4H), supporting the hypothesis that acetylation at K430 enhances DLD dimerization.
A Schematic of human DLD domains and homodimer structure. B Assembly process of PDH/α-KGDH multienzyme complexes. C Co-IP analysis of HA-DLD and Myc-DLD interaction in HEK293T cells with indicated DLD variants. The samples derive from the same experiment but different gels were processed in parallel: separate gels were used for the IP detection of Myc and HA. For the input controls, one gel was used for Myc, and another for HA and Vinculin. IB detection of monomeric and dimeric DLD in HEK293T cells transfected with DLD variants (D) and treated ± TGFβ1 for 6 h (E); DSS, disuccinimidyl suberate. The samples derive from the same experiment but different gels were processed in parallel: one for DSS cross-linked samples, another for HA and Vinculin, and another for pSMAD2/3 and SMAD2/3. F IB detection of endogenous monomeric and dimeric DLD in HGC27 and MKN74 cells under the indicated conditions. The samples derive from the same experiment but different gels were processed in parallel: one for DSS cross-linked samples, another for DLD and Vinculin, and another for pSMAD2/3 and SMAD2/3. G RMSD analysis of DLD dimer stability across indicated variants. H MM/GBSA calculations of binding energy for DLD dimers by indicated variants (n = 50 conformations). Centerline, median; box limits, upper and lower quartiles; whiskers, 1.5x interquartile range, two-sided one-way ANOVA with Tukey’s multiple comparison test. I Co-IP of HA-DLD with E2 components (DLAT, DLST) of PDH and α-KGDH complexes in HEK293T cells transfected with indicated DLD variants. The samples derive from the same experiment but different gels were processed in parallel: for IP samples, one gel was used for DLAT and DLST, and another for HA. For the input controls, one gel was used for Vinculin and HA, and another for DLAT and DLST. Co-IP of HA-DLD with E2 components in HEK293T cells treated w/wo TGFβ1 for 6 h (J) and of endogenous DLD in HGC27 and MKN74 cells (K). The samples derive from the same experiment but different gels were processed in parallel: for IP samples, one gel was used for DLAT and DLST, and another for HA/DLD. For the input controls, one gel was used for Vinculin and HA/DLD, another for DLAT and DLST, and another for pSMAD2/3 and SMAD2/3. TGFβ1 and galunisertib concentrations were 10 ng/mL and 20 μM, respectively, unless otherwise specified. Images in (C–F, I–K) are representative of two independent experiments with similar results.
To assess the impact of K430 acetylation on the assembly of PDH and α-KGDH complexes, we examined the interactions between the E3 component (DLD) and the E2 components (DLAT in the PDH complex and DLST in the α-KGDH complex). The interaction between the hyperacetylation-mimicking K430Q mutant and DLAT or DLST was stronger compared to WT DLD, whereas the interaction was weaker for the deacetylation-mimicking K430R mutant (Fig. 4I). TGFβ1 diminished the E2–E3 interaction in cells transfected with WT DLD but did not affect the interaction in cells transfected with the K430Q mutant (Fig. 4J). This effect was corroborated by decreased interaction between endogenous E2 and E3 components in HGC27 and MKN74 GC cells upon TGFβ1 treatment (Fig. 4K). Additionally, surface plasmon resonance (SPR) assays with purified His-tagged DLD mutants ruled out the possibility that acetylation at K430 affects DLD’s affinity for substrates, as the KD values for NAD+ or FAD binding to DLD were comparable among the three variants (Supplementary Fig. 5B–D). Collectively, these findings demonstrate that TGFβ modulates the activity of PDH and α-KGDH through K430 acetylation-mediated DLD dimerization and enzyme complex assembly.
TGFβ downregulates DLD-K430 acetylation through SMAD3/4-dependent mitochondrial-nuclear translocation of ACAT1
Lysine acetylation is a dynamic and reversible modification regulated by acetyltransferases and deacetylases28. To identify enzymes involved in DLD-K430 acetylation, we performed mass spectrometry on proteins immunoprecipitated with DLD and identified two lysine acetyltransferases: ACAT1 and lysine acetyltransferase 2B (KAT2B) (Fig. 5A, Supplementary Data 4). However, only alterations in ACAT1 expression, not KAT2B, resulted in changes in DLD-K430 acetylation levels in GC cells (Fig. 5B, Supplementary Fig. 6A, B). We confirmed the direct interaction between DLD and ACAT1 through a GST pull-down assay using purified GST-DLD and His-SUMO-ACAT1 (Fig. 5C, Supplementary Fig. 6C). Additionally, in vitro acetylation assays demonstrated that ACAT1 directly catalyzes the acetylation of DLD at K430 (Fig. 5D). Immunoprecipitation, proximity ligation and immunofluorescence assays showed that, upon TGFβ1 treatment, despite unchanged ACAT1 expression levels, the interaction between DLD and ACAT1 was significantly reduced in GC cells and that this decrease could be reversed by galunisertib (Fig. 5E, Supplementary Fig. 6D–G). Notably, TGFβ1 stimulation led to a marked redistribution of ACAT1, with decreased mitochondrial localization and increased nuclear localization, as confirmed by immunofluorescence and subcellular fractionation assays (Fig. 5F, G, Supplementary Fig. 6H). To assess the necessity of mitochondrial translocation for ACAT1-mediated acetylation of DLD at K430, we generated an ACAT1 mutant lacking the mitochondrial targeting sequence (ACAT1∆MTS, Fig. 5H). Unlike cells expressing wild-type ACAT1, cells expressing this mutant exhibited lower levels of DLD-K430ac that did not respond to TGFβ1 (Fig. 5I). These results indicate that TGFβ1 downregulates DLD-K430ac by inhibiting the mitochondrial translocation of ACAT1.
A MS analysis of DLD-immunoprecipitated proteins from HGC27 cells, ranked by enriched peptide number, identified acetyltransferases, including KAT2B and ACAT1 (n = 1). B IB detection of DLD-K430ac in HGC27 cells with ACAT1 knockdown or overexpression. The samples derive from the same experiment but different gels were processed in parallel: one gel for ACAT1, one gel for DLD, and another gel for DLD-K430ac and Vinculin. C GST pull-down showing direct binding between purified GST-DLD and His-SUMO-ACAT1. The samples derive from the same experiment but different gels were processed in parallel: one for pull-down samples, one for His, and another for GST for the input samples. D IB detection of GST-DLDWT or GST-DLDK430R acetylation at K430 by His-SUMO-ACAT1 in vitro. The samples derive from the same experiment but different gels were processed in parallel: one for DLD-K430ac, one for His, and another for GST samples. E Co-IP of endogenous DLD and ACAT1 in HGC27 cells after 6 h treatment. The samples derive from the same experiment but different gels were processed in parallel: one gel for IP samples. For the input controls, one gel was used for ACAT1, DLD, and Vinculin, and another for pSMAD2/3 and SMAD2/3. F Immunofluorescence of MitoTracker and ACAT1 colocalization in HGC27 cells w/wo TGFβ1 for 6 h. Scale bar, 10 μm. G IB of ACAT1 in mitochondrial (Mito), nuclear (Nuc), and whole cell lysates (WCL) in HGC27 cells after 6 h TGFβ1 treatment. The samples derive from the same experiment but different gels were processed in parallel: one gel for ACAT1 and Lamin b1, another gel for ATP5A and Vinculin. H Schematic of ACAT1 variants and IB of ACAT1 in mitochondrial and WCL in HEK293T cells. MTS, mitochondrial targeting sequence. The samples derive from the same experiment but different gels were processed in parallel: one for Flag and Vinculin, another for ATP5A. The samples derive from the same experiment but different gels were processed in parallel: one for Flag and Vinculin, another for ATP5A. I IB of DLD-K430ac in HEK293T cells with ACAT1 depletion and vector, flag-ACAT1WT, or flag-ACAT1ΔMTS w/wo TGFβ1. The samples derive from the same experiment but different gels were processed in parallel: one for Flag and Vinculin, one for DLD-K430ac and DLD, and another for pSMAD2/3 and SMAD2/3. J IB of DLD-K430ac in HGC27 cells with SMAD2/3/4 knockdown w/wo TGFβ1. The samples derive from the same experiment but different gels were processed in parallel: one for DLD-K430ac and Vinculin, one for pSMAD2/3 and SMAD2/3, one for DLD, and another for SMAD4. K Co-IP of endogenous ACAT1 and SMAD2/3/4 in HGC27 cells w/wo 6 h TGFβ1. The samples derive from the same experiment but different gels were processed in parallel: one SMAD2/3 and Vinculin, one for SMAD4, and another for ACAT1. Immunofluorescence of ACAT1 colocalization with SMAD3 (L) or SMAD4 (M) in HGC27 cells w/wo TGFβ1 for 6 h. Scale bar, 10 μm. N, O Co-IP of Flag-ACAT1 and HA-tagged SMAD2/3 variants or SMAD4. The samples derive from the same experiment but different gels were processed in parallel: separate gels were used for the IP detection of Flag and HA. For the input controls, one gel was used for HA, and another for Flag and Vinculin. P Molecular docking showing interaction between ACAT1 tetramer and SMAD3/3/4 complex. Images in (B–D, G–K, N, O) are representative of two independent experiments with similar results. Images in (E, F, L, M) are representative of three independent experiments with similar results.
To elucidate the mechanisms driving ACAT1’s redistribution from mitochondria to the nucleus and to determine if this process is dependent on the canonical TGFβ-SMAD pathway, we performed knockdown experiments targeting SMAD proteins. Knockdown of SMAD3 or SMAD4 led to an increase in DLD-K430 acetylation, which remained unresponsive to TGFβ1 stimulation, while knockdown of SMAD2 did not affect this process (Fig. 5J, Supplementary Fig. 6I), suggesting a dependency on SMAD3/4. Cell fractionation assays revealed that ACAT1 translocation closely mirrored that of SMAD proteins, occurring rapidly within 30 min of TGFβ stimulation and peaking by 3 h (Supplementary Fig. 6J, K). Immunoprecipitation and immunofluorescence assays further indicated that TGFβ1 treatment enhanced the interaction between ACAT1 and SMAD3/4 within the nucleus (Fig. 5K–M, Supplementary Fig. 6L), implying that ACAT1 binds to phosphorylated SMAD3 and the complex formed with SMAD4. To explore this interaction in detail, we constructed HA-tagged SMAD proteins with phospho-mimicking serine-to-aspartate (SD) mutations and non-phosphorylatable serine-to-alanine (SA) mutations in SMAD2 and SMAD3 and examined their interactions with Flag-tagged ACAT1. The results revealed that ACAT1 primarily interacts with SMAD3/4, with the phospho-mimicking SMAD3S423D/S425D mutant significantly enhancing this interaction (Fig. 5N). Moreover, expression of the SMAD3S423D/S425D mutant also increased the interaction between ACAT1 and SMAD4, whereas the SMAD3S423A/S425A mutant did not (Fig. 5O), suggesting that TGFβ1-induced phosphorylation of SMAD3 facilitates the interaction between ACAT1, SMAD3, and SMAD4. Protein docking analysis further supports that ACAT1 interacts with the SMAD3/4 heterotrimer, with the interaction interface located at the junction of SMAD3 and SMAD4, and the phosphorylated S423 residue on SMAD3 being involved in this interaction (Fig. 5P). Critically, SMAD3 knockdown blocked ACAT1 nuclear translocation, resulting in its cytosolic retention, confirming that nuclear translocation of ACAT1 depends on SMAD3/4 (Supplementary Fig. 6M, N). In summary, these findings reveal that TGFβ1 downregulates DLD-K430ac through the SMAD3/4-dependent mitochondrial-nuclear translocation of ACAT1.
Dual targeting of DLD and TGFβ signaling enhances treatment efficacy in GC models
Given the critical role of metabolic regulation in mediating the downstream effects of TGFβ signaling and the adverse pro-tumor growth consequences associated with reversing these metabolic effects using anti-TGFβ therapy, we aimed to enhance therapeutic efficacy through a combined approach involving metabolic intervention. Devimistat (CPI-613), a lipoate analog, serves as a potent disruptor of mitochondrial energy production by inhibiting both PDH and α-KGDH complexes29 (Fig. 6A). Although the specific binding targets of devimistat have not been definitively established, it is likely that, as a lipoate analog, devimistat competes with the dihydrolipoyl moiety of the E2 components for binding at the active site of E3, specifically DLD. Molecular docking analysis revealed that devimistat fits snugly into the pocket adjacent to the cofactor FAD on DLD, exhibiting a binding energy of −16.82 kJ/mol (Fig. 6B). Both bio-layer interferometry and SPR assays confirmed that devimistat binds to purified His-DLD with a dissociation constant (KD) of approximately 500~600 nM (Fig. 6C, D, Supplementary Fig. 7A). Moreover, devimistat effectively inhibited the activity of PDH and α-KGDH, restricted TCA cycle flux, and impeded the in vitro and in vivo growth of GC cells expressing either WT DLD or the hyperacetylation-mimicking DLDK430Q mutant, demonstrating its broad applicability (Supplementary Fig. 7B–G).
A Schematic illustrating devimistat’s pharmacological effect. B Molecular docking model showing devimistat binding to DLD. Bio-layer interferometry-derived binding curves (C) and simulated fits (D) for devimistat-DLD interactions, n = 2 independent experiments with similar results. E, F Representative images and statistical analysis of GC organoid diameters under indicated treatments. Galunisertib and devimistat concentrations were 20 μM and 200 μM, respectively. n = 3 independent experiments, data presented as mean ± SD, two-sided one-way ANOVA with Tukey’s multiple comparison test. Subcutaneous injection of HGC27 cells in BALB/c-nu mice, followed by indicated treatments; tumor images (G), volumes (H), and weights (I) on day 27 are shown. n = 6 mice per group, data presented as mean ± SD, two-sided one-way ANOVA with Tukey’s multiple comparison test, scale bar = 1 cm. J–M Experimental setup for intraperitoneal injection of luciferase-expressing HGC27 cells in BALB/c-nu mice; treatments and tumor burden monitored by bioluminescence imaging (BLI), with analysis of peritoneal bioluminescence (L) and tumor weight (M). n = 6 mice per group, data presented as mean ± SD, Kruskal–Wallis test followed by pairwise Wilcoxon test with BH adjustment. N Schematic of orthotopic GC-PDX model and treatment timeline. O Kaplan‒Meier survival analysis of orthotopic GC-PDX model mice with indicated treatments. n = 10 mice per group, two-sided log-rank test. P Working model for this study.
Combination treatment experiments indicated that devimistat counteracted the growth-promoting effects of galunisertib on GC cells and GC organoids, both in vitro and in vivo (Fig. 6E–I, Supplementary Fig. 7H, I). Given the pivotal role of TGFβ signaling in tumor metastasis, we further compared the efficacy of anti-TGFβ monotherapy with that of anti-TGFβ plus devimistat combination therapy in a peritoneal metastasis model and a lung metastasis model of GC. Consistent with previous studies30,31, the reduction in tumor burden confirmed the therapeutic efficacy of anti-TGFβ monotherapy in preventing metastasis. However, the combined treatment with galunisertib and devimistat demonstrated a remarkable antimetastatic effect that surpassed the efficacy of either drug alone (Fig. 6J–M, Supplementary Fig. 8A–E). This superior efficacy is likely due to the dual inhibition of metastatic seeding and subsequent tumor growth following seeding. Metabolic analyses further indicated that while galunisertib monotherapy enhanced PDH and α-KGDH activity in metastatic tumors, devimistat effectively counteracted this metabolic activation (Supplementary Fig. 8F, G). Additionally, the combination therapy exhibited superior efficacy in prolonging the survival of mice bearing GC PDXs compared to either monotherapy in the orthotopic model (Fig. 6N, O, Supplementary Fig. 8H, I), with no overt toxicity as evidenced by comparable body weights among groups (Supplementary Fig. 8J–L). These findings underscore the translational significance of concurrently targeting TGFβ signaling and DLD for the treatment of GC.
Discussion
The TGFβ signaling pathway is a complex and versatile regulatory mechanism in metazoans, governing processes such as cell proliferation, differentiation, morphogenesis, and tissue homeostasis, with its malfunction linked to various diseases32,33. In cancer, TGFβ is recognized for its dual role—acting as a tumor suppressor in early stages and promoting tumor progression in later stages34. It is widely believed that tumor cells must evade the tumor-suppressive effects of TGFβ to achieve malignant evolution20. Our findings reveal that many GC cells, despite having transformed malignantly, retain responsiveness to the tumor-suppressive effects of TGFβ, particularly (though not absolutely) those with intact TβR and SMAD components. This observation supports the context-dependent nature of TGFβ’s actions and aligns with previous findings35,36,37. Furthermore, it suggests that evasion of TGFβ’s cytostatic effect is a promotive but not strictly necessary step in cancer progression.
The discrepancy between theoretical expectations and the practical efficacy of anti-TGFβ therapy highlights the challenge of translating TGFβ-related research into effective treatments6. Herein, our study suggests that this challenge may partly result from suboptimal tumor burden control due to the growth-promoting effects of anti-TGFβ drugs through metabolic regulation (Fig. 6P). Interestingly, preclinical studies have predominantly shown successful outcomes in metastatic tumor models30,31, while clinical evaluations often rely on measuring lesion size according to the Response Evaluation Criteria In Solid Tumors (RECIST)38. In our study, anti-TGFβ therapy was found to effectively inhibit tumor metastasis but simultaneously abrogated the anti-proliferative effects of TGFβ. This unintended consequence is significant, particularly in cancers involving critical anatomical structures where a large tumor burden can rapidly deteriorate patients’ physical status due to complications such as obstruction39. However, caution is warranted given the high heterogeneity among cancer types—for instance, TGFβ’s tumor-suppressive role in diffuse-type gastric cancer may not apply to breast cancer models40,41. The dual nature of anti-TGFβ therapy underscores the need for combination strategies that incorporate additional therapeutic modalities. Currently, anti-TGFβ agents are being tested in combination with chemotherapy, radiotherapy, and immune checkpoint inhibitors18,42,43. In this regard, our study proposes that DLD could be a rationale-based choice for co-targeting in future clinical practice to enhance therapeutic efficacy.
Compared with the well-characterized molecular events in TGFβ-regulated processes like EMT, the metabolic impacts of TGFβ signaling on cancer cells remain ambiguous and context-dependent14,15. Our study elucidates the specific mechanisms by which TGFβ influences GC metabolism through the regulation of ACAT1’s subcellular distribution and DLD’s acetylation. This metabolic reprogramming not only alters cellular energy metabolism but also functionally intersects with TGFβ’s transcriptional regulatory network, contributing to a crucial dimension of its cytostatic effects. While ACAT1 is traditionally recognized for its role in mitochondrial metabolism, it has also been observed in non-classical subcellular locations and exhibits acetyltransferase activity with substrates such as METTL3, GNPAT, and TFEB44,45,46. Our findings reveal DLD as a substrate for ACAT1 and provide insights into the mechanism behind ACAT1’s nuclear translocation in the presence of TGFβ through interacting with SMAD3/4. This enrichment of ACAT1’s regulatory role extends beyond its metabolic functions, highlighting its involvement in both metabolic and non-metabolic processes. However, the precise mechanism driving ACAT1’s exit from mitochondria to engage with the SMAD3/4 complex remains unresolved.
PDH and α-KGDH are essential for mitochondrial bioenergetics as they act as key entry points for substrates into the TCA cycle47. Up to now, related investigations have primarily focused on the enzyme-specific E1 components of these complexes48,49,50, with relatively little attention given to the universally utilized E3 component, DLD. Historically, studies on DLD have concentrated on disease-related mutations51,52, and only a few proteomics investigations have explored its post-translational modifications, such as phosphorylation, acetylation, and succinylation53,54,55. However, these studies have not provided comprehensive insights into how specific modifications affect DLD’s structure and function. Our study offers insights by demonstrating that acetylation of DLD at K430 enhances the formation of DLD dimers and the assembly of PDH and α-KGDH enzyme complexes.
In conclusion, our present study highlighted the inherent weakness of anti-TGFβ therapy, demonstrated the central role of metabolic regulation in the tumor-suppressive aspect of TGFβ, and proposed a mechanism-based strategy to enhance the efficacy of anti-TGFβ therapy by co-inhibiting DLD.
Methods
Antibodies, reagents, and critical commercial assays
The names, sources, and catalog numbers of all antibodies, reagents, and commercial assays utilized in this study are detailed in Supplementary Data 5.
Cell culture
HEK293T (ATCC, Cat# CRL-11268), KATO III (ATCC, Cat# HTB-103), RKO (ATCC, Cat# CRL-2577), and cells were maintained in high-glucose Dulbecco’s modified Eagle’s medium (Gibco) supplemented with 10% fetal bovine serum (FBS, Gibco) and 1× Penicillin-Streptomycin (WISENT). HGC27 (SIBCB Cell library, Cat# TCHu22), NCI-N87 (ATCC, Cat# CRL-5822), DLD-1 (ATCC, Cat# CCL-221), SW480 (ATCC, Cat# CCL-228), HT-29 (ATCC, Cat# HTB-38), SNU-668 (KCLB, Cat# 00668), SNU-216 (KCLB, Cat# 00216), MKN45 (RIKEN Cell Bank, Cat# RCB1001) and MKN74 (RIKEN Cell Bank, Cat# RCB1002) cells were maintained in RPMI-1640 medium (Gibco) supplemented with 10% FBS and 1× Penicillin-Streptomycin. AGS (ATCC, Cat# CRL-1739) and LoVo (ATCC, Cat#CCL-229) cells were maintained in Ham’s F-12K medium (Gibco) supplemented with 10% FBS and 1× Penicillin-Streptomycin. All the cell lines were tested to confirm no mycoplasma contamination. All cells were incubated in 5% CO2 at 37 °C in a humidified incubator.
Animal studies
Six-week-old BALB/c-nu or NOD.Cg-Prkdcscid IL2rgtm1Wjl/SzJ (NSG) mice were purchased from Vital River Laboratory Animal Technology Co., Ltd. (Beijing, China) or Gempharmatech Co., Ltd. (Nanjing, China) and housed in a specific-pathogen-free facility under standard conditions. All animal protocols were approved by the Institutional Animal Care and Use Committee of Sun Yat-sen University. For subcutaneous tumor models, 2 × 106 HGC27 cells were injected into the right flanks of BALB/c-nu mice. Galunisertib (75 mg/kg), vactosertib (40 mg/kg), devimistat (25 mg/kg), or vehicle were administered intraperitoneally every 3 days, and tumor size was monitored using a caliper. For orthotopic GC-PDX models, fresh GC tissues from patients were trimmed into 1 × 1 × 1 mm blocks and sutured to the serous surface of the glandular stomach of six-week-old NSG mice. Galunisertib (75 mg/kg) or vactosertib (40 mg/kg) was administered intraperitoneally every 3 days, and survival was monitored for analysis. For peritoneal/lung metastasis models, 2 × 106 luciferase-expressing HGC27 cells were injected into the abdominal cavities/tail veins of six-week-old BALB/c-nu mice. Galunisertib (75 mg/kg) and/or devimistat (25 mg/kg) were administered intraperitoneally every 3 days. At week 8, bioluminescence signals were measured using an IVIS Spectrum In Vivo Imaging System (PerkinElmer, Massachusetts, USA) following the intraperitoneal injection of 1.5 mg XenoLight D-Luciferin Potassium Salt (PerkinElmer) in 100 μL volume. The criteria for the experimental endpoint were defined based on the specific tumor model used. For the subcutaneous tumor model, the endpoint was reached when tumors achieved a maximum diameter of 2.0 cm or 10% of body weight. For the orthotopic GC-PDX and metastasis models, endpoints were defined as natural death or humane euthanasia, which was performed if tumor progression interfered with normal locomotion or the ability to access food and water. The sample size was determined with reference to the experimental designs reported in similar previous studies. Mice were stratified by sex and body weight at day 0 for grouping. Simple randomization was employed to assign animals to experimental groups on day 0. Treatment administrations and tumor volume measurements were performed blindly. The data analysts were blinded to the group assignments. Transplanted tumors were not to exceed a diameter of 2.0 cm or 10% of body weight as permitted by the Institutional Ethics Committee for Clinical Research and Animal Trials of the Sun Yat-sen University Cancer Center.
Human tumor samples
Clinical samples, including 12 pairs of primary GC, adjacent normal gastric, and peritoneal metastasis samples for bulk RNA sequencing, 172 primary GC tumor samples, with 97 paired adjacent normal gastric tissue samples, and 16 paired peritoneal metastasis samples for IHC analysis, and GC primary tumor samples for the establishment of orthotopic GC models and GC organoids were collected from Sun Yat-sen University Cancer Center (Guangzhou, China). Samples were collected from both male and female patients. All patients went through endoscopic examination and were histologically diagnosed with GC before the operation. Written informed consent was obtained from all patients. Patient characteristics in the SYSUCC-GC cohort are shown in Supplementary Table 1. The study was approved by the Medical Ethics Committee of Sun Yat-sen University Cancer Center in accordance with the Declaration of Helsinki.
RNA-seq analysis
After RNA extraction and library construction, sequencing was conducted on the Illumina platform, generating fastq files. Subsequently, quality control was performed using fastQC, followed by alignment to the human genome hg38 to generate a gene expression matrix. Differential gene expression analysis was carried out using DESeq2, with genes selected based on a p-value < 0.05. Enrichment analysis was then conducted using the enricher function from the ClusterProfiler package56, focusing on HALLMARK, GO_BP, REACTOME, and KEGG gene sets57, and the results were visualized using bar plots. GSEA analysis was performed within gene sets of interest after sorting all genes by log2FoldChange.
Immunohistochemistry (IHC)
For immunohistochemistry (IHC), tissue slides were first baked for 2 h at 65 °C. Deparaffinization was performed by incubating the slides twice in 100% xylene for 15 min, followed by rehydration through a graded ethanol series (100%, 95%, and 70%) and finally distilled water, with each step lasting 5 min. To block endogenous peroxidase activity, slides were treated with 3% H2O2 in methanol for 10 min. For antigen retrieval, slides were boiled for 20 min in either 10 mM sodium citrate (pH 6.0) or 0.5 mM EDTA (pH 8.0) solution, then allowed to cool at room temperature for 20 min. Following several PBS washes, the slides were blocked for 1 h using 5% BSA in PBST. Samples were then incubated overnight at 4 °C with primary antibodies (1:100–1:500 dilution). After a final set of washes, the signal was detected using the Dako REAL™ EnVision™ Detection System (Peroxidase/DAB+; Dako, Denmark). Staining scores were determined based on the intensity and proportion of positive cells.
Organoid preparation and culture
To generate gastric cancer organoids, fresh tumor tissues were transported on ice in cold DMEM/F12 (Gibco) with primocin (InvivoGen), then washed in cold PBS, minced into fragments (≤1 mm³), and digested for 1 h at 37 °C with collagenase and hyaluronidase. The dissociated cell suspension was centrifuged (300 × g, 5 min), treated for 5 min with RBC lysis buffer (eBioscience) to remove red blood cells, and filtered through a 100 µm cell strainer (Corning). After further centrifugation and resuspension, 4–5 × 10⁵ cells were seeded in 300 µL of Matrigel (Corning) into a pre-warmed 24-well plate. The Matrigel was solidified at 37 °C for 5–10 min before adding 500 µL of complete growth medium, which consisted of Advanced DMEM/F12 supplemented with 10 mM HEPES (Invitrogen), 1× Penicillin-Streptomycin (WISENT), 1× GlutaMAX (Gibco), 1× B-27 Supplement (Gibco), 1 mM N-Acetyl-L-cysteine (Sigma), 10 ng/mL EGF (Gibco), 25 ng/mL FGF-10 (Gibco), 100 ng/mL Wnt-3a (R&D), 500 ng/mL R-Spondin 1 (R&D), 100 ng/mL Noggin (Gibco), 10 µM SB202190 (Sigma), 10 µM Y-27632 (MCE), 1 µM A 83-01 (MCE), and 10 nM human Gastrin I (Sigma). For TGFβ treatment experiments, A 83-01 was omitted from this medium. Organoids were passaged every 1–2 weeks at a 1:3–1:5 dilution by dissociating them with a TrypLE-based (Gibco) harvesting solution for 5–10 min at 37 °C and re-seeding the resulting cell pellet in fresh Matrigel.
PET/CT imaging
To evaluate the therapeutic efficacy of galunisertib in orthotopic GC-PDX models, ¹⁸F-FDG PET/CT imaging was conducted following a 6-week treatment period. Prior to scanning, mice were fasted for 8 h, anesthetized with 50 mg/kg pentobarbital sodium, and administered 5 µCi/g of ¹⁸F-FDG in saline via tail vein injection. A 15-min static scan was acquired 40 m post-injection using an Albira microPET/CT Scanner (Bruker, Germany). The acquired microPET images were co-registered with microCT images and corrected for attenuation, scatter, normalization, and camera dead time. Tumor radiotracer uptake was then quantified by calculating the standardized uptake value (SUV) from three-dimensional regions of interest (ROIs).
Non-targeted metabolomics analysis
For non-targeted metabolomics, HGC27 cells were cultured for 6 h with or without 10 ng/mL TGFβ1. Metabolites were then extracted from 2 × 10⁷ cells by rapid freezing in liquid nitrogen within a 1 mL solution of acetonitrile, methanol, and water (2:2:1, v/v/v). After thawing at room temperature, the samples were homogenized with a tissue grinder and centrifuged at 16,000 × g for 10 min at 4 °C. The resulting supernatant was collected, concentrated, dried, and re-dissolved in a 1:9 (v/v) mixture of acetonitrile and a 4 ppm solution of 2-Amino-3-(2-chloro-phenyl)-propionic acid in 0.1% formic acid, then filtered through a 0.22 µm membrane prior to analysis. LC-MS analysis was performed on a Vanquish UHPLC System coupled to a Q Exactive™ Mass Spectrometer (Thermo Fisher Scientific). Chromatographic separation was achieved using an ACQUITY UPLC HSS T3 column (150 × 2.1 mm, 1.8 µm; Waters) maintained at 40 °C, with a flow rate of 0.25 mL/min and an injection volume of 2 µL. For positive ion mode (ESI+), the mobile phases were 0.1% formic acid in acetonitrile (A) and 0.1% formic acid in water (B), run on the following gradient: 0–1 min, 2% A; 1–9 min, 2–50% A; 9–12 min, 50–98% A; 12–13.5 min, 98% A; 13.5–14 min, 98-2% A; 14–20 min, 2% A. For negative ion mode (ESI−), the mobile phases were acetonitrile (C) and 5 mM ammonium formate (D), with the gradient: 0–1 min, 2% C; 1–9 min, 2–50% C; 9–12 min, 50–98% C; 12–13.5 min, 98% C; 13.5–14 min, 98-2% C; 14–17 min, 2% C. The mass spectrometer was operated with an ESI source in Full MS-ddMS² (data-dependent) mode. Key source parameters included a sheath gas pressure of 30 arb, an aux gas flow of 10 arb, a capillary temperature of 325°C, and spray voltages of +3.50 kV and −2.50 kV for positive and negative modes, respectively. Full MS scans were acquired over a range of m/z 81–1000 at 70,000 FWHM resolution, triggering up to 10 dependent MS/MS scans at 17,500 FWHM resolution using a normalized collision energy of 30% and automatic dynamic exclusion. Finally, Principal Component Analysis (PCA) was performed on the relative metabolite abundances using the ROPLS R package58, and pathway analysis was conducted via Metabolite Set Enrichment Analysis21.
Oxygen consumption rate analysis (OCR)
Mitochondrial respiratory capacity was evaluated using the XF Cell Mito Stress Test Kit (Agilent Technologies). To begin, 2 × 10⁴ cells per well were seeded into a Seahorse 24-well culture microplate and incubated overnight. In parallel, an XF24 FluxPak sensor cartridge was hydrated overnight at 37 °C in a non-CO₂ incubator using Seahorse Calibrant. The following day, the cell medium was replaced with Seahorse base medium supplemented with 1 mM pyruvate, 2 mM glutamine, and 10 mM glucose for a 1-h incubation period. The oxygen consumption rate (OCR) was then measured on an XFe24 extracellular flux analyzer (Agilent). The analysis involved the sequential injection of 1 mmol/L oligomycin A, 0.5 mmol/L FCCP, and 0.5 mmol/L of a rotenone/antimycin A mixture. After the experiment, all OCR values were normalized to the total protein concentration in each well, as determined by a Pierce BCA Protein Assay Kit (Thermo Fisher Scientific).
Metabolic flux analysis
HGC27 cells were cultured in complete RPMI-1640 media supplemented with or without 10 ng/mL TGFβ1 for 6 h. For glucose tracing, the medium was replaced with SILAC RPMI-1640 FLEX-MEDIA (Gibco) supplemented with 200 mg/L L-arginine (Thermo), 40 mg/L L-lysine-2HCl (Thermo), 300 mg/L L-glutamine (Thermo), 10% dialyzed FBS, and 2 g/L D-glucose-13C6 (Sigma) for 2 h. For glutamine tracing, the medium was replaced with SILAC RPMI-1640 FLEX-MEDIA (Gibco) supplemented with 200 mg/L L-arginine (Thermo), 40 mg/L L-lysine-2HCl (Thermo), 2 g/L D-glucose, 10% dialyzed FBS, and 300 mg/L L-glutamine-13C5 (Sigma) for 2 h. For GC-MS analysis, cellular metabolism was quenched with liquid nitrogen, and metabolites were subsequently extracted with a prechilled (−80 °C) 80% methanol solution. The extract then underwent a two-step derivatization process. First, samples were incubated for 90 min at 37 °C with 30 µL of 20 mg/mL methoxyamine hydrochloride in pyridine. This was followed by the addition of 30 µL of N-tert-butyldimethylsilyl-N-methyl-trifluoroacetamide (MTBSTFA) with 1% tBDMS (Regis Technologies) and a second 90-min incubation at 37 °C. The derivatized samples were analyzed on an Agilent 7890 gas chromatography system coupled to an Agilent 5975 C inert MSD. A 1 µL aliquot was injected in splitless mode with a 5.5-min solvent delay. The GC oven program began at 100 °C for 2 min, then ramped to 180 °C at 10 °C/min, to 260 °C at 5 °C/min, and finally to 300 °C at 10 °C/min, with a final 8-min hold. The injector, transfer line, and electron impact ion source were set to 250 °C, 250 °C, and 230 °C, respectively. Mass spectra were acquired across a range of m/z 50–600 in full scan mode with an electron energy of 70 eV. Finally, the natural isotopic abundance in the data was corrected using IsoCor software59.
PDH, α-KGDH, CS activity assays
The activity of PDH was measured using a Pyruvate Dehydrogenase (PDH) Enzyme Activity Microplate Assay Kit (Abcam) following the manufacturer’s instructions. Briefly, the PDH enzyme complex was captured onto an antibody-coated microplate and supplemented with substrates. PDH activity was determined by following the reduction of NAD+ to NADH, coupled to the reduction of a reporter dye to yield a colored reaction product that can be monitored by measuring the absorbance at 450 nm. The activity of α-KGDH was measured using an α-Ketoglutarate Dehydrogenase (α-KGDH) Activity Assay Kit following the manufacturer’s instructions (Solarbio, Beijing, China). Briefly, cells were lysed for protein extraction. Substrates of α-KGDH were supplemented, and the absorbance at 340 nm was measured for monitoring changes in the concentration of the product NADH. The activity of the CS was measured using a Citrate Synthase (CS) Activity Assay Kit following the manufacturer’s instructions (Solarbio). Briefly, cells were lysed for protein extraction. Substrates of CS were supplemented. CS activity was determined by following the production of citrate, coupled to the change of color of a reporter dye that can be monitored by measuring the absorbance at 412 nm.
DNA construction and mutagenesis
Short hairpin RNA (shRNA) sequences of DLD, ACAT1, and KAT2B were amplified by polymerase chain reaction (PCR) and cloned into the pCLenti-U6-shRNA-CMV-Puro-WPRE vector from OBio Biotechnology (Shanghai, China). Short hairpin RNA (shRNA)-resistant (r) human rDLD-WT, rDLD-K430R, and rDLD-K430Q were made by introducing missense mutations at K430 and synonymous mutations in shRNA-targeting sites in PCR-amplified DLD and cloned into pLenti-EGFP-F2A-Hygro-CMV-MCS-WPRE vectors from OBio Biotechnology. C-terminal HA-tagged DLD-K66R, DLD-K122R, DLD-K159R, DLD-K166R, DLD-K277R, DLD-K320R, DLD-K410R, DLD-K417R, DLD-K430R, DLD-K430Q, and DLD-K445R, and Myc-tagged DLD-K430R and DLD-K430Q were made by introducing missense mutations in the corresponding lysine sites in DLD and cloned into pcDNA3.1 vector from OBio Biotechnology. PCR-amplified ACAT1, ACAT1ΔMTS, and KAT2B were cloned into pLenti-CMV-ACAT1-3×FLAG-PGK-Neo-WPRE vectors from OBio Biotechnology. For prokaryotic expression, DLD-WT, DLD-K430R, and DLD-K430Q were cloned into pET28A or pGEX vectors from Genecreate Biotechnology (Wuhan, China), and ACAT1 was cloned into the pET-SUMO vector from Genecreate Biotechnology.
RNA isolation and qPCR analysis
Total RNA was extracted using TRIzol reagent, and first-strand cDNA was subsequently synthesized with the Prime Script RT Master Mix Kit (Takara). This cDNA then served as the template for quantitative real-time PCR (qPCR), which was performed using the GoTaq qPCR Master Mix (Promega) according to the manufacturer’s protocol. All resulting gene expression data were analyzed and normalized to ACTB levels. A complete list of the primers used for this analysis can be found in Supplementary Table 2.
Viral production and transfection
Lentiviruses were produced by co-transfecting 2 μg of the constructed target plasmid with 2.8 μg packing plasmid psPAX2 and 1.56 μg envelope vector pMD2.G into HEK293T cells cultured in T25 flasks using ViaFect Transfection Reagent (Promega) according to the manufacturer’s instructions. Lentiviruses were collected 72 h after transfection. Sequences for RNA interference are listed in Supplementary Table 3. Virus transfection was performed by incubating cells with medium containing lentivirus and 8 μg/mL polybrene (OBio) for 24 h. Cells were then selected with puromycin (Sigma) or hygromycin B (Sigma) until no cells in the empty control group were alive.
Immunoprecipitation and immunoblot
Cells were harvested in lysis buffer for IB and IP (Beyotime, Shanghai, China) containing Halt Protease and Phosphatase Inhibitor Cocktail (Thermo Fisher Scientific, Waltham, MA). The protein concentrations were normalized using a BCA assay kit (Thermo Fisher Scientific). For IP assays, protein lysates were incubated with the indicated IP antibody and protein A/G-conjugated magnetic beads (MCE) or anti-HA beads (MCE) overnight at 4 °C on a rotary shaker. After 3 times of washing with lysis buffer, the bound proteins were eluted with SDS-loading buffer for immunoblot analysis. For western blotting, protein lysates were resolved by 5× loading buffer (Beyotime) and transferred to PVDF membranes with 0.45 μm pore (Millipore). All membranes were incubated with the indicated primary antibodies diluted in TBST (20 mM Tris, pH 7.5, 150 mM NaCl, 0.1% Tween-20) supplemented with 5% bovine serum albumin (BSA, Sigma–Aldrich) overnight at 4 °C. After 3 times of washing with TBST, membranes were incubated with horseradish peroxidase-conjugated secondary antibodies diluted in TBST-5% BSA for 1 h at RT. Chemiluminescence was detected by ChemiDoc MP (Bio-Rad, California, USA), following exposure to SuperSignal West Femto Maximum Sensitivity Substrate (Thermo Fisher Scientific). Quantification was performed with ImageJ60.
Mass spectrometry analysis of DLD acetylation
For mass spectrometry analysis of DLD acetylation, immunoprecipitated protein was eluted from anti-DLD antibody beads using an SDT lysis buffer (4% SDS, 100 mM DTT, 100 mM Tris-HCl, pH 8.0) with boiling and ultrasonication. The cleared protein supernatant was then reduced with 0.05 M TCEP for 1 h at 60 °C and alkylated with 55 mM MMTS for 45 min at room temperature in darkness. A two-step trypsin digestion was performed, first overnight (1:50 enzyme:protein ratio) and then for 4 h (1:100 ratio), before the peptides were vacuum dried. For the LC-MS/MS analysis, peptides were reconstituted in 0.1% formic acid (FA) and 2% acetonitrile (ACN) and loaded directly onto a reversed-phase analytical column (75 µm i.d. × 150 mm, Acclaim PepMap RSLC C18). Chromatographic separation was performed over a 50-min gradient, ramping from 5% to 90% solvent B (0.1% FA in 80% ACN) at a constant flow rate of 300 nL/min. The HPLC was coupled online to a Q Exactive™ Orbitrap mass spectrometer (Thermo Fisher Scientific) with a nanospray ionization (NSI) source. The instrument operated in a data-dependent acquisition (DDA) mode, alternating between one full MS scan (70,000 resolution) and 20 subsequent MS/MS scans. The top 20 precursor ions with an intensity threshold above 1E4 were selected for fragmentation using a normalized collision energy (NCE) of 27. The resulting fragment ions were detected in the Orbitrap at a resolution of 17,500. Key parameters included an electrospray voltage of 2.0 kV, a dynamic exclusion of 30 s, and an automatic gain control (AGC) target of 1E5 ions for MS/MS scans to prevent ion trap overfilling. During the acquisition, full MS scans were performed over a mass range of m/z 350 to 1800, with a fixed first mass of 100 m/z for fragmentation. For protein identification, the resulting raw files were searched against the Uniprot Homo sapiens database using MASCOT software. The search parameters specified trypsin as the enzyme, allowing for a maximum of one missed cleavage. Carbamidomethylation of cysteine was set as a fixed modification, while acetylation of lysine was treated as a variable modification. The mass tolerances for precursor and fragment ions were set to 20 ppm and 0.6 Da, respectively, using monoisotopic masses, and a significance threshold of 0.05 was applied to all identifications.
Protein expression, purification, and verification
pET28A-His-DLD WT, K430R, K430Q, pGEX-6P-1-GST-DLD WT, K430R or pET-SUMO-ACAT1 were transformed into BL21/DE3 competent cells. Recombinant protein expression was induced for 24 h at 30 °C using 0.5 mM IPTG, after which the cells were lysed by sonication. For His-tagged proteins, the lysate was purified using Ni-NTA resin (Genescript) for 4 h at 4 °C; the column was washed with a 20 mM imidazole buffer, and proteins were eluted with a 0–250 mM imidazole gradient. For GST-tagged proteins, cleared lysates were loaded onto a GSTrap HP column, washed with PBS, and eluted with 10 mM reduced glutathione. All proteins then underwent a second purification step via size exclusion chromatography on a Superdex 200 Increase 10/300 GL column using an AKTA FPLC System. The final purified proteins were concentrated and desalted into PBS using an Amicon Ultra-4 centrifugal filter (Millipore), and their concentrations were quantified with a BCA protein assay kit (Pierce).
Protein cross-linking assay
Protein cross-linking was achieved with the crosslinker disuccinimidyl suberate (DSS). Cells were treated with 1 mM DSS in conjugation buffer containing 0.15 M NaCl at pH 7 at room temperature for 30 min. Then the reaction mixture was quenched by adding the quenching buffer (1 M Tris-HCl, pH 7.5) to a final concentration of 50 mM Tris and incubated at room temperature for 15 min. Cross-linked proteins were prepared for further detection.
Molecular dynamics simulation
Molecular dynamics simulations were conducted using the Amber software61, with the dimeric structure of DLD (PDB ID: 2F5Z) serving as the initial conformation. Mutations at the K430 residue were introduced using Pymol to create K430Q and K430R variants, resulting in three distinct initial systems. Each protein system was solvated using the TIP3P explicit solvent model via the leap program62, with the system shape set to a truncated octahedron and the solvent edge positioned 15 Å from the protein structure. Neutralization was achieved by adding 8 Na+ counterions to the K430Q system and 6 Na+ counterions to the K430R and wild-type K430 systems, ensuring an overall neutral charge. Periodic boundary conditions (PBC) were applied to eliminate boundary effects. Following energy minimization, the systems were heated from 10 K to 300 K over 100 ps under constant volume conditions (NVT ensemble), and subsequently equilibrated for 500 ps under constant temperature and pressure (NPT ensemble). The SHAKE algorithm was utilized to constrain bonds involving hydrogen atoms, with a 2 fs time step for integration. Production dynamics simulations were performed with all restraints removed, using the pmemd.cuda program compiled for GPU acceleration. Trajectories were output every 2 fs and processed using the Amber traj program to calculate RMSD63 for the complex, protein αC atoms, receptor backbone, and small molecules. Visualizations were generated using Grace software. To assess binding energies, 50 random frames from the steady-state phase of the simulation were selected for Molecular Mechanics Generalized Born Surface Area (MM/GBSA) calculations64, allowing for comparative analysis across the different systems.
Bio-layer interferometry
The binding kinetics between His-tagged DLD and devimistat were analyzed by bio-layer interferometry (BLI) on an Octet Red 96 system (Fortebio). The experiment was conducted at 30 °C with 1000 rpm shaking, and data were acquired at a frequency of 2.0 Hz. Following a 10-min equilibration, His-DLD protein was immobilized onto Ni-NTA biosensors (Sartorius). A dilution series of devimistat was then introduced to the sensor tips to measure association. The resulting raw sensorgrams were aligned to the baseline, and all kinetic parameters were calculated by fitting the data to a 1:1 binding model using Octet Analysis 11.0 software.
Surface plasmon resonance (SPR) assay
Binding interactions of DLD-WT, DLD-K430R, and DLD-K430Q with NAD⁺, FAD, or devimistat were evaluated at 25 °C on a BIAcore T200 system (Cytiva) equipped with CM7 sensor chips. Data analysis was performed using the BIAcore T200 Evaluation software (v2.0, Cytiva) according to the manufacturer’s instructions. In short, the CM7 chip surface was first activated with a mixture of 200 μM 1-ethyl-3-(3-dimethylaminopropyl)carbodiimide (EDC) and 50 μM N-hydroxysuccinimide (NHS) at a flow rate of 10 μL/min for 10 min. Subsequently, 30 μL of purified protein solution (1 mg/mL, adjusted to pH 4.0 with 10 mM sodium acetate, pH 4.0) was injected at 10 μL/min for 5 min in three consecutive cycles, resulting in ~14,000 response units. Residual reactive groups were blocked with 1 M ethanolamine (10 μL/min, 10 min). A reference flow cell was treated in the same manner, except PBS adjusted to pH 4.0 was used instead of protein solution. Following immobilization, all cells were equilibrated with PBS. For NAD⁺ and FAD measurements, stock solutions (1 mM) were serially diluted in PBS and injected at 30 μL/min for 180 s per cycle, followed by surface regeneration with 10 mM glycine-HCl (pH 2.0) for 5 min at 10 μL/min. For devimistat binding assays, PBS containing 5% DMSO was used as the running buffer, and a solvent correction protocol was applied after the final injection. Sensorgrams from protein-coated flow cells were reference-subtracted, and binding kinetics were determined by global fitting of the data to a 1:1 Langmuir binding model.
Immunofluorescence analysis
HGC27 cells with indicated treatments were fixed with 4% paraformaldehyde for 15 min, permeabilized with 0.2% Triton X-100 for 10 min, and blocked with blocking buffer containing 1% BSA for 1 h at room temperature. Cells were then incubated with primary antibodies (at a 1:100 or 1:200 dilution) at 4 °C overnight. The next day, cells were washed and incubated with fluorescence dye-conjugated secondary antibodies for 1 h, and DAPI at room temperature. Immunofluorescence microscopic images of the cells were obtained using LSM880 and processed with ZEN software (ZEISS).
Proximity ligation assay (PLA)
Protein-protein interactions in HGC27 cells were investigated using the Duolink in situ Proximity Ligation Assay (PLA) kit (Sigma–Aldrich). According to the manufacturer’s protocol, fixed cells were first incubated with a pair of primary antibodies (1:200 dilution) and subsequently with secondary PLA probes. A positive signal, which is generated only when the two target antigens are within 40 nm of each other, was visualized using a 594 nm fluorescence detection kit. Cell nuclei were counterstained with DAPI, and all images were acquired on an LSM880 confocal microscope and processed with ZEN software (ZEISS).
GST pull-down assay
Purified 10 μg GST-DLD protein was incubated with purified 10 μg His-SUMO-ACAT1 with anti-GST magnetic beads (MCE) in binding buffer (20 mM Tris-Cl pH 8.0, 200 mM NaCl, 1 mM EDTA pH 8.0, 0.5% NP-40, and protease inhibitor cocktail) in a final volume of 500 μL at 4 °C overnight. The beads were then washed three times with binding buffer, and the bound proteins were eluted with SDS-loading buffer prior to immunoblot analysis.
In vitro acetylation assay
For the in vitro acetylation assay, 2 µg of purified His-SUMO-ACAT1 enzyme was incubated with 2 µg of substrate (either GST-DLD WT or K430R) in a 50 µL reaction. The reaction buffer contained 50 mM Tris-HCl (pH 8.0), 1 mM DTT, 75 mM KCl, 20 µM acetyl CoA, and a protease inhibitor cocktail. After a 1-h incubation at 30 °C, the reaction was terminated by adding SDS-PAGE sample buffer and heating the mixture at 95 °C for 5 min. The resulting acetylated proteins were then resolved and analyzed by SDS-PAGE and immunoblot.
Subcellular fractionation
Mitochondria were isolated using a Mitochondria Isolation Kit for Cultured Cells (Thermo Fisher Scientific) according to the manufacturer’s directions. Briefly, cells were collected and underwent sequential treatment with reagents A and B. Mitochondria were then collected through differential centrifugation. Nuclei were isolated using a Nuclear and Cytoplasmic Protein Extraction Kit (Beyotime) according to the manufacturer’s directions. Briefly, cells underwent sequential treatment with reagents A and B containing PMSF. Nuclei were then collected by centrifugation. Nuclear proteins were then extracted for further analysis.
Molecular docking
For protein-small molecule docking, the X-ray crystal structure of DLD (PDB ID: 1ZMC) was obtained from the Protein Data Bank. The 3D structure of devimistat was obtained from the ZINC database (ZINC ID: ZINC43208701). Molecular docking calculations were performed using Autodock465. For protein-protein docking, the X-ray crystal structure of ACAT1 tetramer (PDB ID: 2F2S) and the SMAD3/3/4 heterotrimer (PDB ID: 1U7F) were obtained from the Protein Data Bank. Protein-protein docking was performed with the HDOCK server66. The conformation with the best affinity was selected as the final docking conformation. The docking results were visualized and displayed as 3D diagrams with PyMol 2.4.
Statistical analysis
Comparisons between two groups were performed with a paired/unpaired Student’s t-test. One-way or two-way ANOVA test with Tukey multiple comparisons was used to compare continuous outcomes across multiple experimental groups. Kruskal–Wallis test with pairwise Wilcoxon test was used for a non-parametric test between multiple groups. Survival functions were estimated by Kaplan–Meier methods, and the log-rank test was used to compare the survival distributions of different groups. Pearson correlation was used to evaluate the association between two indicators. Statistical analyses were performed with R version 4.3.0 or Graphpad Prism 8.0. A p-value less than 0.05 was considered to represent a statistically significant difference. All statistical tests were two-sided. No data were excluded from the analyses.
Ethical statement
All tissues were obtained with approval from the Institutional Research Ethics Committee of SYSUCC (G2022-080-01). All animal studies were performed in accordance with a protocol approved by the Institutional Ethics Committee for Clinical Research and Animal Trials of the SYSUCC (L102012022221U).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The mass spectrometry proteomics data related to this study have been deposited in the ProteomeXchange Consortium under dataset identifier PXD060756. RNA-seq data related to this study have been deposited in the Genome Sequence Archive (GSA) under the accession number HRA009315. All the other data are available within the article and its Supplementary Information. Source data are provided with this paper.
References
Smyth, E. C., Nilsson, M., Grabsch, H. I., van Grieken, N. C. & Lordick, F. Gastric cancer. Lancet 396, 635–648 (2020).
Cristescu, R. et al. Molecular analysis of gastric cancer identifies subtypes associated with distinct clinical outcomes. Nat. Med. 21, 449–456 (2015).
Tanaka, Y. et al. Multi-omic profiling of peritoneal metastases in gastric cancer identifies molecular subtypes and therapeutic vulnerabilities. Nat. Cancer 2, 962–977 (2021).
Massagué, J. TGFβ signalling in context. Nat. Rev. Mol. Cell Biol. 13, 616–630 (2012).
David, C. J. & Massague, J. Contextual determinants of TGF beta action in development, immunity and cancer. Nat. Rev. Mol. Cell Biol. 19, 419–435 (2018).
Ciardiello, D., Elez, E., Tabernero, J. & Seoane, J. Clinical development of therapies targeting TGF beta: current knowledge and future perspectives. Ann. Oncol. 31, 1336–1349 (2020).
Peng, D., Fu, M., Wang, M., Wei, Y. & Wei, X. Targeting TGF-β signal transduction for fibrosis and cancer therapy. Mol. Cancer 21, 104 (2022).
Hanahan, D. Hallmarks of cancer: new dimensions. Cancer Discov. 12, 31–46 (2022).
Han, Y. et al. IL-1β-associated NNT acetylation orchestrates iron-sulfur cluster maintenance and cancer immunotherapy resistance. Mol. Cell 83, 1887–1902.e1888 (2023).
Zhang, Y. Y., Han, Y., Li, W. N., Xu, R. H. & Ju, H. Q. Tumor iron homeostasis and immune regulation. Trends Pharm. Sci. 45, 145–156 (2024).
Yadav, H. et al. Protection from obesity and diabetes by blockade of TGF-β/Smad3 signaling. Cell Metab. 14, 67–79 (2011).
Nilchian, A., Giotopoulou, N., Sun, W. & Fuxe, J. Different regulation of Glut1 expression and glucose uptake during the induction and chronic stages of TGFβ1-Induced EMT in breast cancer cells. Biomolecules 10, 1621 (2020).
Jiang, L. et al. Metabolic reprogramming during TGFβ1-induced epithelial-to-mesenchymal transition. Oncogene 34, 3908–3916 (2015).
Liu, H. & Chen, Y. G. The interplay between TGF-β signaling and cell metabolism. Front. Cell Dev. Biol. 10, 846723 (2022).
Shi, X. et al. TGF-β signaling in the tumor metabolic microenvironment and targeted therapies. J. Hematol. Oncol. 15, 135 (2022).
Youssef, K. K. & Nieto, M. A. Epithelial-mesenchymal transition in tissue repair and degeneration. Nat. Rev. Mol. Cell Biol. 25, 720–739 (2024).
Rodon, J. et al. First-in-human dose study of the novel transforming growth factor-β receptor I kinase inhibitor LY2157299 monohydrate in patients with advanced cancer and glioma. Clin. Cancer Res. 21, 553–560 (2015).
Yamazaki, T. et al. Galunisertib plus neoadjuvant chemoradiotherapy in patients with locally advanced rectal cancer: a single-arm, phase 2 trial. Lancet Oncol. 23, 1189–1200 (2022).
Kim, B. G., Malek, E., Choi, S. H., Ignatz-Hoover, J. J. & Driscoll, J. J. Novel therapies emerging in oncology to target the TGF-β pathway. J. Hematol. Oncol. 14, 55 (2021).
Massagué, J. TGFbeta in Cancer. Cell 134, 215–230 (2008).
Xia, J. & Wishart, D. S. MSEA: a web-based tool to identify biologically meaningful patterns in quantitative metabolomic data. Nucleic Acids Res. 38, W71–W77 (2010).
Mathias, R. A. et al. Sirtuin 4 is a lipoamidase regulating pyruvate dehydrogenase complex activity. Cell 159, 1615–1625 (2014).
Li, W., Li, F., Zhang, X., Lin, H.-K. & Xu, C. Insights into the post-translational modification and its emerging role in shaping the tumor microenvironment. Signal. Transduct. Target. Ther. 6, 422 (2021).
Narita, T., Weinert, B. T. & Choudhary, C. Functions and mechanisms of non-histone protein acetylation. Nat. Rev. Mol. Cell Biol. 20, 156–174 (2019).
Patel, M. S., Nemeria, N. S., Furey, W. & Jordan, F. The pyruvate dehydrogenase complexes: structure-based function and regulation. J. Biol. Chem. 289, 16615–16623 (2014).
Sheu, K. F. & Blass, J. P. The alpha-ketoglutarate dehydrogenase complex. Ann. N. Y. Acad. Sci. 893, 61–78 (1999).
Brautigam, C. A., Chuang, J. L., Tomchick, D. R., Machius, M. & Chuang, D. T. Crystal structure of human dihydrolipoamide dehydrogenase: NAD+/NADH binding and the structural basis of disease-causing mutations. J. Mol. Biol. 350, 543–552 (2005).
Shvedunova, M. & Akhtar, A. Modulation of cellular processes by histone and non-histone protein acetylation. Nat. Rev. Mol. Cell Biol. 23, 329–349 (2022).
Stuart, S. D. et al. A strategically designed small molecule attacks alpha-ketoglutarate dehydrogenase in tumor cells through a redox process. Cancer Metab. 2, 4 (2014).
Xia, X. et al. Neutrophil extracellular traps promote metastasis in gastric cancer patients with postoperative abdominal infectious complications. Nat. Commun. 13, 1017 (2022).
Xie, L. et al. Gastric cancer-derived LBP promotes liver metastasis by driving intrahepatic fibrotic pre-metastatic niche formation. J. Exp. Clin. Cancer Res. 42, 258 (2023).
Deng, Z. et al. TGF-β signaling in health, disease, and therapeutics. Signal. Transduct. Target Ther. 9, 61 (2024).
Massagué, J. & Sheppard, D. TGF-β signaling in health and disease. Cell 186, 4007–4037 (2023).
Colak, S. & Ten Dijke, P. Targeting TGF-β signaling in cancer. Trends Cancer 3, 56–71 (2017).
Takahashi, K. et al. TGF-β generates a population of cancer cells residing in G1 phase with high motility and metastatic potential via KRTAP2-3. Cell Rep. 40, 111411 (2022).
Wang, Z. et al. Involucrasin B inhibits the proliferation of Caco-2 cells by regulating the TGFβ/SMAD2-3-4 pathway. Molecules 29, 686 (2024).
Li, Q. S. & Zheng, P. S. ESRRB inhibits the TGFβ signaling pathway to drive cell proliferation in cervical cancer. Cancer Res. 83, 3095–3114 (2023).
Eisenhauer, E. A. et al. New response evaluation criteria in solid tumours: revised RECIST guideline (version 1.1). Eur. J. Cancer 45, 228–247 (2009).
Chen, X. J. et al. Palliative gastrectomy versus gastrojejunostomy for advanced gastric cancer with outlet obstruction: a propensity score matching analysis. BMC Cancer 21, 188 (2021).
Komuro, A. et al. Diffuse-type gastric carcinoma: progression, angiogenesis, and transforming growth factor beta signaling. J. Natl. Cancer Inst. 101, 592–604 (2009).
Yang, Y. A. et al. Lifetime exposure to a soluble TGF-beta antagonist protects mice against metastasis without adverse side effects. J. Clin. Invest. 109, 1607–1615 (2002).
Liu, S., Ren, J. & Ten Dijke, P. Targeting TGFβ signal transduction for cancer therapy. Signal. Transduct. Target Ther. 6, 8 (2021).
Lan, Y. et al. Simultaneous targeting of TGF-β/PD-L1 synergizes with radiotherapy by reprogramming the tumor microenvironment to overcome immune evasion. Cancer Cell 39, 1388–1403.e1310 (2021).
Gu, L. et al. Stabilization of FASN by ACAT1-mediated GNPAT acetylation promotes lipid metabolism and hepatocarcinogenesis. Oncogene 39, 2437–2449 (2020).
Zhang, G. et al. ACAT1-mediated METTL3 acetylation inhibits cell migration and invasion in triple negative breast cancer. Genes Immun. 24, 99–107 (2023).
Zhang, J. et al. Importance of TFEB acetylation in control of its transcriptional activity and lysosomal function in response to histone deacetylase inhibitors. Autophagy 14, 1043–1059 (2018).
Chalifoux, O., Faerman, B. & Mailloux, R. J. Mitochondrial hydrogen peroxide production by pyruvate dehydrogenase and α-ketoglutarate dehydrogenase in oxidative eustress and oxidative distress. J. Biol. Chem. 299, 105399 (2023).
Stacpoole, P. W. Therapeutic targeting of the pyruvate dehydrogenase complex/pyruvate dehydrogenase kinase (PDC/PDK) axis in cancer. J. Natl. Cancer Inst. 109, djx071 (2017).
Mailloux, R. J., Craig Ayre, D. & Christian, S. L. Induction of mitochondrial reactive oxygen species production by GSH mediated S-glutathionylation of 2-oxoglutarate dehydrogenase. Redox Biol. 8, 285–297 (2016).
East, M. P., Laitinen, T. & Asquith, C. R. M. BCKDK: an emerging kinase target for metabolic diseases and cancer. Nat. Rev. Drug Discov. 20, 498 (2021).
Odievre, M. H. et al. A novel mutation in the dihydrolipoamide dehydrogenase E3 subunit gene (DLD) resulting in an atypical form of alpha-ketoglutarate dehydrogenase deficiency. Hum. Mutat. 25, 323–324 (2005.
Campanholi, D. R. R. et al. Molecular basis of various forms of maple syrup urine disease in Chilean patients. Mol. Genet. Genom. Med. 9, e1616 (2021).
Huttlin, E. L. et al. A tissue-specific atlas of mouse protein phosphorylation and expression. Cell 143, 1174–1189 (2010).
Rardin, M. J. et al. Label-free quantitative proteomics of the lysine acetylome in mitochondria identifies substrates of SIRT3 in metabolic pathways. Proc. Natl. Acad. Sci. USA 110, 6601–6606 (2013).
Park, J. et al. SIRT5-mediated lysine desuccinylation impacts diverse metabolic pathways. Mol. Cell 50, 919–930 (2013).
Wu, T. et al. clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innovation 2, 100141 (2021).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Thévenot, E. A., Roux, A., Xu, Y., Ezan, E. & Junot, C. Analysis of the human adult urinary metabolome variations with age, body mass index, and gender by implementing a comprehensive workflow for univariate and OPLS statistical analyses. J. Proteome Res. 14, 3322–3335 (2015).
Millard, P. et al. IsoCor: isotope correction for high-resolution MS labeling experiments. Bioinformatics 35, 4484–4487 (2019).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Pearlman, D. A. et al. AMBER, a package of computer programs for applying molecular mechanics, normal mode analysis, molecular dynamics and free energy calculations to simulate the structural and energetic properties of molecules. Comput. Phys. Commun. 91, 1–41 (1995).
Zhang, Z., Yan, K., Liu, X. & Liu, J. A leap-frog algorithm-based efficient unified thermostat scheme for molecular dynamics. Chin. Sci. Bull. 63, 3467–3483 (2018).
Coutsias, E. A., Seok, C. & Dill, K. A. Using quaternions to calculate RMSD. J. Comput. Chem. 25, 1849–1857 (2004).
Genheden, S. & Ryde, U. The MM/PBSA and MM/GBSA methods to estimate ligand-binding affinities. Expert Opin. Drug Discov. 10, 449–461 (2015).
Morris, G. M. et al. AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J. Comput. Chem. 30, 2785–2791 (2009).
Yan, Y., Tao, H., He, J. & Huang, S. Y. The HDOCK server for integrated protein-protein docking. Nat. Protoc. 15, 1829–1852 (2020).
Acknowledgements
This work was supported by National Key Research and Development Program of China (2022YFA1105300), National Natural Science Foundation of China (82273241, 82102921, 82203822), China Postdoctoral Science Foundation (2023M744049), Guangdong Basic and Applied Basic Research Foundation (2023B1515040030, 2023A1515010547, 2021A1515111001), Sanming Project of Medicine in Shenzhen (SZSM202211017), Chih Kuang Scholarship for Outstanding Young Physician-Scientists of Sun Yat-sen University Cancer Center (CKS-SYSUCC-2023001). Some of the figures in this study were generated with BioRender. We sincerely thank Prof. Ye-Guang Chen for his invaluable guidance and insightful advice, which greatly contributed to this work. We are also grateful to the doctors and patients at SYSUCC for their contributions to tissue collection. Additionally, we appreciate the staff of the SYSUCC Animal Center for their dedicated care of the experimental animals.
Author information
Authors and Affiliations
Contributions
H.-Q.J. and R.-H.X. conceptualized the study. Y.-Q.P., Y.H., Z.-Y.Q., T.-T.W., H.C., K.L., Y.-Q.Z., Q.-N.W., and W.C. collected the data. X.-Y.Z., Z.-R.C., Y.L., Y.-Y.Z., H.-X.W., and T.T. analyzed and interpreted the data. Y.-Q.P., Z.-Y.Q., X.-Y.Z., and Y.-Q.Z. performed statistical analysis. H.-X.W., T.T., Y.C., Z.-L.Z., Z.-X.L., H.-L.P., and X.-Y.G. contributed to the discussion and data interpretation. Y.-Q.P., Y.H., Z.-Y.Q., X.-Y.Z., and H.-Q.J. wrote the original draft of the manuscript. R.-H.X., H.-L.P., and X.-Y.G. reviewed and edited the manuscript. H.-Q.J. supervised and supervised and accepted full responsibility for the work as the guarantor. All authors reviewed the manuscript and approved the final version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Keiji Miyazawa, Laurent Le Cam, Ming Sun, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Pan, YQ., Han, Y., Qian, ZY. et al. A preserved TGFβ cytostatic response through DLD-mediated metabolic modulation undermines anti-TGFβ therapy in gastric cancer. Nat Commun 16, 10016 (2025). https://doi.org/10.1038/s41467-025-64997-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-64997-5
This article is cited by
-
Triple targeting of STING, TGF-β, and PD-L1 boosts CXCL16–CXCR6 signaling for potent antitumor response
Nature Communications (2026)