Abstract
Chromatin interactions play critical roles in gene regulation. The genomes of distinct maize inbreds exhibit substantial structural variation, however, F1 hybrids lead to significant hybrid vigor. The long-range genomic interactions present in different inbreds and their hybrid offspring have not been assessed in maize. Here, we perform H3K4me3/H3K27ac HiChIP in two maize inbreds and their reciprocal F1 hybrids. We identify interactions specific to inbred or hybrid genotypes. Some genes involved in hybrid-specific interactions show dominant or over-dominant expression patterns. The analysis of the inbred-specific or shared interactions between two parental inbred lines in the hybrids reveal unexpected gain or loss of interactions compared to the parents. The unexpected hybrid interactions are partially associated with unexpected gene regulation patterns. We also identify some inter-allelic interactions in hybrids, suggesting potential somatic chromosome interactions in maize. Our analyses of 3D regulatory networks provide a potential explanation for non-additive gene expression in maize hybrids.
Similar content being viewed by others
Introduction
The characterization of 3D genomes provides an opportunity to decipher the highly dynamic, spatiotemporal processes in nuclei linked to transcriptional regulation. This information bridges the genotype-related chromosome 3D architecture with phenotype-related functional performance1,2. Launched from the original dilution Hi-C3 and estrogen receptor α-centered ChIA-PET4 simultaneously in 2009, the 3D toolkit2,5 continuously expands to be more sophisticated and provide higher signal-to-noise for depicting chromatin interactions. Among the 3D toolkit of approaches, HiChIP6 or PLAC-seq7 offer an efficient and sensitive approach to interpret long-range physical contacts. While HiChIP has been used for several discoveries in mammals8,9,10,11,12, this 3D approach has only recently been applied to plants to explore the 3D nuclear architecture of polyploidy wheat13 and long-range cis-regulatory elements in maize14,15.
Structural variation (SV), including transposable element (TE)-associated insertion/deletion polymorphisms, are prevalent among different maize inbred lines16,17,18,19,20. In mammals, TEs contribute to genotype specific-interactions and are also found in the conserved interactions21,22,23. In rice, scores of genes with Presence/Absence variation (PAV)-associated line-specific interactions display higher expression, suggesting that the SVs contribute to the genotype specific interactions24,25. However, the extent to which SVs influence the 3D genome architecture has not been systematically examined in plant species. As a pivotal crop for genetic and genomic research in plants, maize plays a crucial role of both economical and biological significance as starch source, animal fodder, petroleum substitution and degradable fiber for sustainable development confronting global warming26,27. Moreover, ~85% of maize genome is composed of TEs, and maize inbred lines exhibit exceptional SVs with hundreds of megabases of non-alignable sequence between any two inbreds18,28,29,30. This provides an opportunity to investigate how SVs/TEs contribute to long-range interactions in distinct maize haplotypes.
Crosses between two maize inbreds with distinct genome content result in substantial heterosis31. Heterosis refers to the phenomenon in which the F1 offspring outperform its parents in yield, biomass, fitness, and resistance to biotic and abiotic stress32,33,34,35. While there are several hypotheses that seek to explain potential genetic mechanisms of heterosis, the specific basis of heterosis and the molecular mechanisms that confer non-additive expression patterns remain unclear. During the past two decades, genomic analyses have been conducted to explore the underlying mechanisms of heterosis36,37. Recent 3D work on plant hybrids delineates the chromatin topological landscapes in a species-specific and objective-centered way based on the Hi-C platform. In Arabidopsis, the two sub-genomes in a hybrid tend to have different chromatin compactness increases38. In Brassica napus, the hybrids consisted of more active A compartment compared to their parents39. Further, in barley, it was found that both the contacts between the homologues and the non-homologues were detected40. However, to date a higher resolution 3D chromatin interaction landscape, as well as the relationship between the allele-specific expression and the interactions has not been assessed.
In this study, we use the H3K4me3 and H3K27ac-directed HiChIP to assess the role of SVs in the context of 3D chromatin interactions in the seedlings of maize inbred lines B73 and Mo17. We find that SVs contribute to differential long-range interactions between the parents and that these differences can lead to altered interactions of distal sequences with genes in distinct haplotypes. We also probe the hybrid-specific interactions and inter-allelic interactions in the reciprocal hybrids B73 x Mo17 and Mo17 x B73 relative to the inbred parents. This reveals evidence for hybrid-specific interactions, including examples of inter-allelic interactions. Our work provides evidence into the role of SVs on long-range interaction variation in plant species and highlights the potential role of long-range interactions in explaining non-additive patterns of gene expression in hybrids.
Results
Generation of high-quality HiChIP data for B73, Mo17 and their hybrids
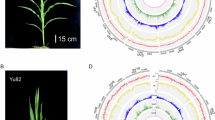
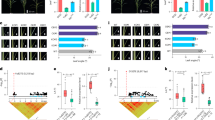
To compare long-range interactions among maize inbreds and their F1 hybrids, we performed HiChIP as well as ChIP-seq for two biological replicates of seedlings from B73, Mo17, B73 x Mo17 (BM), and Mo17 x B73 (MB). Specifically, we generated HiChIP and ChIP-seq data per sample using antibodies against H3K4me3 and H3K27ac, which mark putative proximal and distal regulatory elements, respectively15,41,42 (Fig. 1a). The two replicates of HiChIP and ChIP-seq data exhibited a high level of reproducibility (Supplementary Figs. 1 and 2). For each replicate per genotype for either H3K4me3 or H3K27ac, we generated ~13−47 million valid read pairs by aligning to the B73_AGPv4 reference genome or modified B73 reference genome that includes single-nucleotide polymorphism (SNP) derived from Mo17 substituted to enable improved mapping rates16,43 (Supplementary Table 1). For each chromatin modification per genotype, valid read pairs and peaks identified from ChIP-seq data were used to identify long-range interactions ranging from 20 Kb to 2 Mb between two anchors. We identified 3369–51,184 interactions for either replicate per genotype (Supplementary Table 1). We found the alignment of ChIP-seq data derived from Mo17 to the B73 reference genome rather than the correct genome (Mo17) resulted in a 15%−32% reduction of uniquely mapping reads (Fig. 1b); while the number of peaks that were detected in each biological replicate is reduced by 4.88%−11.47% due to alignment to the other genome rather than alignment to the proper reference genome (Fig. 1c). For the HiChIP dataset, the alignment was reduced by 40.96%−44.80%, by aligning to the other reference genome with SNP-between the correct genome and the other genome substituted (Fig. 1d). Consequently, the number of interactions detected reduced by up to 42.35% of ones detected by using the correct genome (Fig. 1e). As expected, no obvious difference between the read mapping ratio, or the number of peaks, or the interactions obtained was observed in the BM or MB hybrids based on alignments to either B73 or Mo17 reference genome (Supplementary Fig. 3).
a Overview of the datasets we generated and contents to be explored in this study. By using newly generated HiChIP, ChIP-seq, and corresponding RNA-seq data, we would detect inbred-specific, hybrid-specific, and inter-allelic interactions; and then aimed to investigate how structure variations (SVs) contribute to inbred-specific interactions, how hybrid-specific ones regulate gene expression, and are there any inter-allelic ones existed in the maize hybrids. Some interactive examples were further confirmed by 4C-seq. b, c Comparisons of the number of uniquely mapped reads (b) and the number of peaks (c) obtained, when aligning B73 or Mo17 inbred ChIP-seq data to their cognate and counterpart reference genomes, respectively. d, e Comparisons of the number of valid read pairs (d) and the number of interactions (e) we obtained when aligning HiChIP datasets of individual inbred lines to their cognate and counterpart reference genomes. B73 data was mapped to B73 and B73-SNP substituted Mo17 reference genome, and Mo17 data was mapped to Mo17 and Mo17-SNP substituted B73 reference genome. B73-ref, B73 reference genome (B73_AGPv4); Mo17-ref, Mo17 reference genome (Mo17-CAU-v1). For b-e, B73-R1 indicates the replicate 1 dataset of B73 while B73-R2 for the replicate 2 of B73 for H3K4me3 or H3K27ac; Mo17-R1 and Mo17-R2 have the same meanings. f Pipeline of long-range interaction detection using combined B73 + Mo17 reference genome. Taking B73 inbred as an example of inbred samples, the HiChIP data will be aligned to the B73_AGPv4 genome and Mo17-CAU-v1 with B73-SNP substituted, respectively. All the valid read pairs along 20 chromosomes, as well as their corresponding peaks obtained based on ChIP-seq data, will be utilized to call loops using FitHiChIP; by contrast, the pipeline for hybrids was quite similar, except that the HiChIP data being aligned to B73_AGPv4 genome and Mo17-CAU-v1, respectively. B73-ref and Mo17-ref have the same meaning as in (b).
To reduce the impact of these differences in alignment to the two different reference genomes on detected interactions, we developed a strategy for read mapping and interaction identification as shown in Fig. 1f. In detail, B73 HiChIP data was separately aligned to the B73_AGPv443 and Mo17_ref_v116 genomes with SNP substituted. We then combined all the valid read pairs mapped to B73 and/or Mo17 genomes in 20 chromosomes (10 chromosomes from B73 and 10 chromosomes from Mo17) for interaction detection (Fig. 1f). A similar alignment was performed for the Mo17 dataset by using Mo17_ref_v1 and SNP-substituted B73_AGPv4 genome. Since hybrid HiChIP data included both of parental genomes, hybrid data was aligned to B73_AGPv4 genome and Mo17_ref_v1 genome separately. As a result, we obtained 10,179 to 97,983 interactions for either replicate per genotype by this strategy (Supplementary Table 1).
To further assess the interactions detected by using the combined reference genomes, we compared them with the interactions detected using a single reference genome. It showed that the interactions detected based on the combined genome includes >99% interactions detected using a single reference genome. We compared the interactions detected based on the previously published ChIA-PET method, also using B73 seedlings44. It revealed that about 51.8% H3K4me3 and 40% H3K27ac interactions detected from ChIA-PET and HiChIP overlapped, including ZmRap2.7, BX1, UB3, and PSBS1-centered loci44,45. In addition, other different interactions centered at these four loci were also detected using our dataset (Supplementary Fig. 4). For example, at the ZmRap2.7 locus we identified three previously reported H3K4me3 interactions captured by ChIA-PET44, but also detected four other interactions (Supplementary Fig. 4a). These analyses suggested that our HiChIP method can produce a robust set of long-range interactions in maize inbreds and hybrids studied.
Identification of differential and conserved long-range interactions in B73 and Mo17
Considering the tremendous structural variation between the B73 and Mo17 genomes, we hypothesized that there would be substantial variation in chromatin interactions between the two genotypes. To identify inbred-specific interactions, we used “Differential Analysis” function46 in FitHiChIP to compare interactions between B73 and Mo17 inbreds (Supplementary Fig. 5, see Methods). We identified 7613/1930 and 4937/725 (H3K4me3/H3K27ac) B73-specific and Mo17-specific interactions, respectively (Supplementary Data 1, 2, 3, and 4). There were also 9547 H3K4me3 and 3670 H3K27ac conserved interactions identified between B73 and Mo17 (Supplementary Data 5 and 6; See details in Methods). We assessed the quality of inbred-specific and conserved interactions by comparing them to an independent dataset that profiled H3K4me3 interactions in B7314. Many (60%) of B73-specific H3K4me3 interactions, and 91.2% of conserved interactions in our study, overlapped with the independent dataset, but only 4% of Mo17-specific H3K4me3 interactions were detected in the independent dataset. Further, we selected one B73-specific interaction for validation using 4C-seq (Supplementary Fig. 6). We detected a significant enrichment (74,026 reads captured) in B73, but no significant peaks (399 reads) were obtained in Mo17. Both of these results indicated the high quality of the inbred-specific interaction list we detected.
We proceeded to investigate genomic distributions of interaction anchors in inbred-specific and conserved interactions. More than 80% of both H3K4me3- and H3K27ac-based conserved interactions represented genic-genic interactions (both anchor regions overlapped with annotated protein-coding genes) (Fig. 2a). However, only <17% of them represented genic-intergenic interactions (one anchor region overlapped with annotated protein-coding genes, while the other one did not overlap with any annotated protein-coding genes) with relatively few representing intergenic-intergenic interactions (both anchor regions did not overlap with any annotated protein-coding genes). The B73- or Mo17-specific interactions exhibited higher frequency of gene-intergenic contacts compared to the conserved interactions (Fig. 2a). This suggested that the inbred-specific interactions have higher proportion of gene-intergenic interactions. On the other hand, we found that 85% or more of the genic-genic conserved interactions were syntenic gene-syntenic gene (syn-syn) (syntenic genes indicate the genes are syntenic between B73 and Mo17 genomes; there are 25,810 syntenic genes in B73 genome and 25,576 genes in Mo17 genome) interactions, while the number for inbred-specific ones is 64% ~ 74% (Fig. 2b). These syn-syn interactions enabled us to investigate the loop length variation between inbreds. We found almost 85% of conserved syn-syn interactions have >1 Kb difference in distance between two anchors of B73 and Mo17, and ~70% were greater than 10 Kb (Fig. 2c, Supplementary Data 7 and 8). By contrast, ~93% of syn-syn interactions detected from B73- or Mo17- specific ones have >1 Kb difference in distance between two anchors of B73 and Mo17 and about 75% with the difference greater than 10 Kb (Fig. 2c). Take a syn-syn (the Zm00001d012517–pmg1) interaction detected from the conserved interaction list as an example (Fig. 2d, e), a 17 Kb Presence-Absence Variations (PAV) region primarily comprised by TEs (Transposable Elements) was present within the loop region of the interaction. Overall, the results above indicated chromatin interactions could be primarily resulted from syntenic genes, and sequence variation was widely tolerated in the loop regions of both inbred-specific and conserved interactions.
a The proportion of the interactions with their anchor sequences overlapping genic or non-genic sequences was determined for conserved and genotype-specific interactions, respectively. b The proportion of the anchor sequences that overlap syntenic and non-syntenic genes was determined for genic-genic class of conserved and genotype-specific interactions, respectively. c Comparison of loop length difference among B73- and Mo17-specific, and conserved interactions in B73 and Mo17. Loop length difference was calculated based on the distance between the inside boundary of a pair of anchors from each interaction in B73 and Mo17 genome. d, e An example shows conserved long-range interactions between two highlighted regions, separately in red and green, between two pairs of corresponding syntenic genes in B73 and Mo17 genomes. Variable loop distances between two syntenic genes in B73 and Mo17 coordinates were 29 Kb and 17 Kb respectively. f Comparison of expression levels of genes targeted by B73- or Mo17-specific interactions in B73 and Mo17. Only interactions with at least one anchor overlapping with only one protein-coding genes were used for further analyzation. The boxplots depict the median (central line), interquartile range (box boundaries), and 1.5×IQR whisker ranges, with outliers shown as individual points. Wilcoxon test (two-sided) was performed to assess the difference between the two inbred lines. From left to right, the p-values and sample size are: < 2.2e−16 (n = 2718), < 2.2e−16 (n = 1708), 1.575e−14 (n = 1074), 9.251e−13 (n = 369). g Comparison of expression levels of genes targeted by conserved interactions with different loop length. All conserved interactions were divided into three categories based on their loop length, “long” indicates the interactions with their loop length in the top third (n = 9926 in H3K4me3; n = 7306 in H3K27ac), while “short” indicates ones with their loop length in the bottom third (n = 9926 in H3K4me3; n = 7304 in H3K27ac). The remaining ones belong to “middle” (n = 9928 in H3K4me3; n = 7306 in H3K27ac). Only interactions with at least one anchor overlapping with only one protein-coding genes were used, and these genes’ expression level was calculated by FPKM. FPKM, fragments per kilobase per million reads. The boxplots depict the median (central line), interquartile range (box boundaries), and 1.5×IQR whisker ranges, with outliers shown as individual points. Wilcoxon test (two-sided) was performed to assess the expression difference. From left to right, the p-values are: < 2.2e−16, 0.002, < 2.2e−16, 3.991e−6, 1.997e−10, 2.409e−5, 0.001, 6.396e−6. Source data are provided as a Source Data file.
Further, we analyzed the expression levels of genes that were located within the anchor sequences of inbred-specific interactions, and it revealed that genes involved in B73-specific interactions were significantly higher expressed in B73 than in Mo17, while the genes targeted by high-confidence Mo17-specific ones expressed higher in Mo17 than B73 (Wilcoxon test, p-value < 0.01 in both of histone modifications, Fig. 2f). Additionally, we found the longer distance between the gene pairs, the lower the expression level of the corresponding genes tends to be (Fig. 2g, Wilcoxon test, p-value < 0.01 in every comparison). These results suggested long-range interaction might directly or indirectly affect gene expression.
Structural variation contributed to the occurrence of a subset of genotype-specific long-range interactions
Since there were a large number of sequence differences between B73 and Mo17 genome16, we hypothesized that many of the interactions detected in only one genome might be the result of large structural variants between genomes. As expected, a previously reported large PAV, an ~2.91 Mb deletion16 in the Mo17 genome relative to the B73 genome, was a region with interactions in B73 data but no detected interactions in Mo17 (Fig. 3a). In contrast, an ~ 5.35 Mb identical-by-state (IBS) region30 in the two genomes exhibited similar patterns and numbers of interactions (Fig. 3b). In addition, we identified 2038 and 2157 PAV regions with the length >50 Kb on the B73 and Mo17 genomes, respectively, and compared the number of valid read pairs obtained from B73 and Mo17 inbred lines. It was found that 91.2% of the B73-presence (Mo17-absence) regions contained more B73 valid read pairs. Similarly, 95.9% of the Mo17-presence (B73-absence) regions contained more Mo17 valid pair reads (Supplementary Fig. 7). Furthermore, it was worth investigating the contribution of structural variation (SV) to anchors in inbred-specific interactions. Since a certain number of anchors (Supplementary Fig. 8) involved in inbred-specific interactions were also detected to interact with other anchors forming other non-inbred-specific interactions, we divided the anchors employed by inbred-specific interactions into two groups: specific anchors and shared anchors. Shared anchor means that the anchors were involved in inbred-specific interactions as well as other non-inbred-specific ones, while the specific anchors indicate that the anchors were only involved in inbred-specific interactions. Moreover, given that ChIP-seq peaks were considered as the key region for interactions and at least 86.2% anchors containing ChIP-seq peaks per sample, we further assessed the SVs of anchor regions by identifying the ChIP-seq peak regions. As expected, specific anchors have a higher proportion of SVs compared to shared anchors. For B73-specific interactions, about 68.5% (1414 of 2064) and 79.8% (927 of 1161) peaks of specific anchors were detected with SVs in H3K4me3 and H3K27ac, respectively (Fig. 3c). In contrast, for the shared anchors, about 38.0% (1444 of 3804) H3K4me3 and 51.5% (644 of 1250) H3K27ac were detected with SVs (Fig. 3c). Similar results were observed in Mo17-specific interactions (Fig. 3c). Further, we compared the length of SVs overlapped with peak regions in specific and shared anchors. Regarding the varied sizes of peak regions and the overlapping regions between peaks and SVs in anchors, we adopted the relative length of an SV on a peak region and found the relative length of SVs on specific anchors was obviously much larger than that on shared anchors (Fig. 3d). These results indicated that specific anchors were often located within or near SVs compared to the shared anchors.
a, b Comparison of normalized contact matrix between B73-B73 and Mo17-B73 H3K4me3 on a previously identified PAV (present in B73, but absent in Mo17) region (Chr6:22,483,501-25,398,000); Comparison of normalized contact matrix between B73-B73 and Mo17-B73 H3K4me3 on a previously identified IBS region (Chr2:124,400,000-129,750,000). c The proportion of anchors with their peak regions overlapping with SVs, taking the shared anchors as controls. Specific anchors were filtered out from B73- or Mo17-specific interactions but requiring that they were not shared in the conserved interactions, while shared anchors from inbred-specific interactions were also shared in the conserved interactions. d Distribution of the percentages of SVs accounting for the peak region of 5 Kb anchor for specific anchors and shared ones from B73- or Mo17-specific interactions. The boxplots depict the median (central line), interquartile range (box boundaries), and 1.5×IQR whisker ranges, with outliers shown as individual points. From left to right, the sample sizes are: 3804, 2064, 2817, 2084, 1250, 1161, 474, 562. e Overview of a PAV-related chromatin example. L1 is a B73-specific interaction with two anchors targeting at R1 and R2 regions, separately. As there is a large PAV (present in B73, but absent in Mo17) in the peak region of R2, another L2 interaction between R1 and R3 was detected in the Mo17 genome, and it’s Mo17-specific. R2 region overlaps Zm00001d033948, and R3 overlaps Zm00001d033951. L1 and L2 were both detected in H3K4me3 and H3K27ac. f, g Comparisons of the expression levels of Zm00001d033948 and Zm00001d033951 in B73 or Mo17. The genes, Zm00001d033948 and Zm00001d033951 was targeted by B73- and Mo17-specific interactions, respectively. FPKM, fragments per kilobase per million reads. The boxplots depict the median (central line), interquartile range (box boundaries), and 1.5×IQR whisker ranges, with outliers shown as individual points. Source data are provided as a Source Data file.
Subsequently, we assessed the impact when one of the anchor regions of a documented interaction in one inbred line was deleted in the other inbred line. By employing inbred-specific interactions and related SV genes (see Methods), we detected 11 cases (Supplementary Table 2). A detailed analysis of the PAV-related interaction case between the two inbreds was performed to monitor potential changes in interactions (Fig. 3e). In this scenario, an interaction was observed at an existing peak in B73, but this peak was absent in Mo17, resulting in no interaction in Mo17. However, another interaction was observed in Mo17 at a flanking peak near the missing one. As shown in Fig. 3e, the L1 interaction between the R1 and R2 regions was detected as specific to B73. The R1 region is located within the gene body of Zm00001d033945, distal to its transcription start site (TSS), suggesting it as a potential enhancer region. In contrast, the R2 region contains a ChIP-seq peak at the first exon and promoter region of the gene Zm00001d033948. This peak was identified as a PAV region, present in B73 but absent in Mo17. Correspondingly, the gene Zm00001d033948 displayed a high expression level in B73 but lost its expression in Mo17 (Fig. 3f). In the Mo17 genome, with the peak region of R2 lost, we detected a Mo17-specific L2 interaction between the R1 and R3 regions. The R3 region includes a ChIP-seq peak that overlaps with the TSS of Zm00001d033951 in Mo17. The expression level of Zm00001d033951 exhibited elevated expression in Mo17 relative to B73 (Fig. 3f). In other words, after R1 lost its interaction with R2, R1 retargeted another region R3, in the Mo17 genome. These examples suggested that these changes in interactions may be associated with genotype-specific expression in maize.
Identification of hybrid-specific long-range interactions
F1 hybrids resulting from crosses between B73 and Mo17 exhibited substantial better-parent heterosis for yield, plant height, and many other traits31,36,47. The long-range interaction data from the reciprocal F1 hybrids were used to search for hybrid-specific interactions. For each modification per hybrid, we used the same approach to identify genotype-specific interactions by requiring the interactions detected in each hybrid but not detected in either of the two parents. Comparing each hybrid (BM or MB) with both of inbreds (B73 or Mo17), we detected five hybrid-specific H3K4me3 interactions (zero for H3K27ac) that are commonly present in BM and MB but not in the inbreds (Fig. 4a); by contrast, we didn’t detect any hybrid-specific interactions when using in silico hybrid data (negative control) in either modification (see Methods, Supplementary Fig. 9). Further analysis showed that only one expressed protein-coding gene overlap with the ChIP-seq peaks of anchors of these five hybrid-specific H3K4me3 interactions, and this gene has a dominant expression pattern (Supplementary Fig. 10). However, it is hard to draw a conclusion based on a limited number of cases. Nevertheless, we detected hundreds of to a few thousand of hybrid-preferable interactions (with FDR < 0.05 when doing comparison between any of one inbred and one hybrid) when comparing interactions between hybrids and inbreds (Fig. 4a). By assessing the expression levels of genes overlapping hybrid-specific anchors (see Methods), we found these genes exhibited significantly higher expression level in hybrids than that of inbred lines (Fig. 4b). By contrast, the genes overlapping inbred-specific anchors displayed decreased expression level in hybrids compared to that in the inbred lines (Supplementary Fig. 11, see Methods). This result further indicated that long-range interactions may contribute to the expression of hybrid-preferable expression.
a A four-way Venn diagram shows all long-range interactions found in hybrid genotypes relative to inbred parents. MB > B73, represented MB specific interactions relative to B73, and these were identified by FitHiChIP using “differential analysis” model. BM > Mo17, BM > B73, and MB > Mo17 have similar meaning as MB > B73, all referring to interactions detected in a specific hybrid relative to a certain inbred. b Comparisons of the expression levels of genes targeted by hybrid-specific interactions. The boxplots depict the median (central line), interquartile range (box boundaries), and 1.5×IQR whisker ranges, with outliers shown as individual points. Wilcoxon test (two-sided) was performed to assess the difference between the two inbred lines. The p-values and sample size are: 1.554e−11 (BM > B73 H3K4me3, n = 271), 6.459 e−8 (MB > B73 H3K4me3, n = 65), < 2.2e−16 (MB > B73 H3K27ac, n = 281), 1.105e−11 (BM > Mo17 H3K4me3, n = 337), 0.013 (MB > Mo17 H3K4me3, n = 48), 1.074 e−6 (MB>Mo17, n = 377). c The proportion of valid read pairs derived from the B73 allele was determined based on the merge of the two inbred datasets (x-axis) and in the F1 hybrid (y-axis) for the hybrid gain interactions. d Hybrid loss interactions were identified using the Fisher exact test (two-sided) with adjusted p-value < 0.05. e, f Proportion of regulatory genes that were uniquely targeted by unexpected and expected interactions. Genes following seven categories including cis only, trans only, cis+trans, cisxtrans, compensatory, conserved and ambiguous were classified following the approach reported in McManus et al.69. For simplify comparisons, we classified trans only, cis+trans and cisxtrans into “trans” regulatory patterns. Genes with conserved and ambiguous expression patterns were not considered in the analysis. Source data are provided as a Source Data file.
In order to investigate allele-specific changes in long-range interactions between inbred and hybrid lines, we searched for examples of allele-specific gain or loss of interactions in the F1 hybrids relative to the parental inbreds by using the strategy as shown in Supplementary Fig. 12. If these genotype-specific interactions were caused by cis-acting sequence variation between the two alleles, we would expect only one of the alleles to exhibit evidence for interactions in the F1 hybrid. However, if trans-acting variation influenced the variable interactions in the two parents, we would expect to potentially observe interactions for both alleles based on an allelic gain of an interaction. To document examples of allele-specific gain of long-range interactions, we focused on the subset of inbred-specific interactions that are present in only B73 or Mo17. Considering the FitHiChIP software we used under the Peak-to-All mode will not consider a region as an interaction, when both anchors of this region do not overlap any ChIP-seq peaks but have HiChIP reads, therefore, we used 6764/1814 and 3874/703 (H3K4me3/H3K27ac) B73-specific and Mo17-specific interactions for further analysis by excluding such interactions. To perform allele-specific analysis of the interactions, we only used the subset of valid read pairs overlapping SNP positions because these reads can be classified based on which parent they were derived from. A comparison of the parental difference in the number of valid read pairs in the inbreds compared to the proportion of valid read pairs from B73 in the F1 hybrids, revealed 3275 H3K4me3 and 925 H3K27ac examples of cis-acting variation such that the bias observed in the parents was maintained in the F1 as expected (Fig. 4c). However, there were also 22 H3K4me3 and 64 H3K27ac examples, in which the bias observed between the parents was not maintained in the F1; meanwhile, we did not detect any H3K4me3 or H3K27ac interactions using in silico hybrid datasets (negative control). Overall, these results reflected unexpected gains of interactions for an allele within the F1 compared to the parent and suggested potential hybrid-specific allelic interactions. The potential allele-specific loss of interactions were identified from the conserved interactions (detected both in B73 and Mo17 inbred lines) that were detected to be of significant difference in B73 or Mo17 allele of F1 hybrids (Fig. 4d). The analysis revealed that 7549 H3K4me3 and 2751 H3K27ac examples in the F1 hybrids exhibited proportions similar to the bias observed between the parents. In contrast, 41 H3K4me3 and 83 H3K27ac examples showed an unexpected level of bias in the F1 hybrids (3 H3K4me3 and 4 H3K27ac interactions detected from in silico hybrid datasets), suggesting a preferential interaction for one allele over the other (Fig. 4d). The examples of biased interactions in the F1 could result from allelic competition for factors necessary for loop formation that can occur in the F1 but not in the inbred lines.
The examples of allele-specific gain or loss of interactions in the hybrids could result in unexpected patterns of gene expression for genes located on the anchors of these interactions. The pattern of allele-specific expression was assessed for all differentially expressed genes located on the anchors of the interactions with allele-specific interaction data (Fig. 4e, f). The allele-specific expression data could be used to classify genes as having cis-variation (allelic expression bias observed in F1 was similar to the proportion predicted in the parents), trans-variation (allelic expression bias in F1 was much lower than expected based on the proportion predicted from parents) or compensatory variation (allelic expression bias in F1 was higher than predicted based on parents). The genes that showed the unexpected patterns of hybrid-specific loss of interactions were enriched for compensatory patterns. This suggested that the hybrid-specific changes in allelic interactions may result in higher levels of allele-expression bias for nearby genes.
Identification of potential long-range inter-allelic interactions in maize
The analysis of allele-specific interactions also enabled us to search for potential examples of inter-allelic interactions in maize that could reflect interactions of homologous chromosomes in non-meiotic cells (Fig. 5a). In order to capture inter-allelic interactions, we developed a strategy by calculating valid read pairs with each read including at least one SNP, which distinguished reads from B73 or Mo17 allele (see Methods). We were concerned that a subset of these putative somatic chromosome pairing interactions might reflect false-positive SNP calls or complex structural rearrangements, so we ran the same analysis process on the inbred samples and removed any interactions that had any valid pair reads for candidate B73-Mo17 inter-allelic interactions. By limiting the number of valid read pairs, a different number of allele-specific interactions were detected (Fig. 5b, c). If we focused on the subset of interactions which had >= 2 valid read pairs in both BM and MB, we obtained 377 and 422 H3K4me3/H3K27ac inter-allelic interactions, respectively (Fig. 5b, c, Supplementary Data 9 and 10). Using a much stricter criterion that requiring >= 10 valid read pairs detected in BM and MB hybrids, we obtained 7 and 3 H3K4me3/H3K27ac inter-allelic interactions.
a The inter-allelic interaction (blue lines) could be detected when an interaction was detected in a hybrid and either anchor of the valid read pairs overlapped with a different allele-specific SNP. Yellow stars indicate the SNPs between B73 and Mo17. b, c The number of valid read pairs in BM and MB was shown for all the candidate inter-allelic interactions. The dots in dark green indicates the interactions with at least one valid read pairs, while the dots in black, red, and yellow indicate the number of detected valid-pair reads be greater than 5, 3 or 4, and 2, that supported inter-allelic interactions between two alleles was determined by both reciprocal hybrids (BM and MB) for H3K4me3 (b) and H3K27ac (c). d The proportion of the anchor-sequences that overlap with syntenic or non-syntenic genes as well other types was determined for inter-allelic interactions. e Expression of genes targeted by H3K4me3 inter-allelic interactions overlapping with syntenic genes in inbred or hybrid lines. The boxplots depict the median (central line), interquartile range (box boundaries), and 1.5×IQR whisker ranges, with outliers shown as individual points (Wilcoxon test (two-sided). The p-values and sample size are: < 2.2e−16 (B73 compare to MB, n = 255), 3.714e−15 (B73 compare to BM, n = 255), < 2.2e−16 (Mo17 compare to BM, n = 239), 0.056 (Mo17 compare to MB, n = 239). f The browser view of the inter-allelic interaction on chromosome 7. The left anchor (Chr7: 20,505,000 – 20,510,000) overlapped Zm00001d019180 (from B73), and the right anchor (chr7: 21,065,000 – 21,070,000) overlapped Zm00014a006917 (from Mo17). The numbers below the black straight line represent inter-allelic valid read pairs number from HiChIP data, and the numbers below the blue curved line are presented validated reads number from 4C-seq. g, h The expression level of Zm00001d019180 (g) and Zm00014a006917 (h) in inbred or hybrid lines. The boxplots depict the median (central line), interquartile range (box boundaries), and 1.5 × IQR whisker ranges, with outliers shown as individual points. Source data are provided as a Source Data file.
After analyzing the genomic distribution of inter-allelic interactions across the genome, we found that about 75% of them were genic-genic ones, including syn-syn and nonsyn-nonsyn types. More interestingly, about 50% of these interactions occurred as syn-syn genes between B73 and Mo17 (Fig. 5d). Further, we compared the expression levels of these genes in hybrids and inbreds and found that both hybrids exhibited significantly higher expression compared to B73 (Fig. 5e, Supplementary Fig. 13). One example of these genic-genic inter-allelic interactions was shown in Fig. 5f. Based on HiChIP dataset, 2 and 10 valid read pairs spanning B73 and Mo17 alleles were detected in BM and MB, respectively. Importantly, using an independent 4C-seq experiment, we obtained 1304 and 2223 read pairs spanning the B73 and Mo17 alleles in the BM and MB samples, respectively (Fig. 5f). These suggested that the gene Zm00001d019180 in the B73 genome interacted with its homologous gene Zm00014a006917 in the Mo17 genome. Additionally, we found that Zm00001d19180 in the B73 genome was significantly up-regulated in the MB sample compared to B73 (Fig. 5g), and Zm00014a006917 in the Mo17 genome displayed increased expression in both BM and MB compared to that in Mo17 (Fig. 5h). Overall, these results indicated that inter-allelic interactions occurred primarily between homolog genes and typically resulted in higher expression levels in the hybrids.
Discussion
The 3D chromatin architecture can shed light on the functional genome and interactions that likely influence gene expression. The maize genome has similar attributes as other crop species with high TE content, abundant structural variation among diverse inbred lines, and exhibit strong heterosis in F1 hybrids35,37,48. In this study, we interrogated the long-range chromatin interactions in two maize inbred lines, B73 and Mo17, along with their reciprocal hybrids BM and MB using HiChIP with H3K4me3 and H3K27ac antibodies in duplicates by using the B73 and Mo17 combined genome. For the two inbreds with structurally diverse genomes, we identified the differential interactions and obtained variable numbers of long-range interactions for the inbred lines. There is significant enrichment of gene-intergenic (putative promoter-putative enhancer) interactions for inbred-specific interactions, which were also observed for tissue-specific interactions in both plants and mammals44,49. We found that structural variation between the inbreds is likely responsible for the presence of about one-third of genotype-specific interactions. Interestingly, we discovered several SV-triggered interaction cases in which another interaction(s) will be tethered as the presence/absence of ChIP-seq peak(s) in given inbreds in maize, and this finding was consistent with the recently reported ERR (enhancer release and retargeting) phenomenon in mammals50. ERR suggests that when one anchor of an interaction is deleted, the remaining ‘released’ anchor may form other interactions with other sequences, potentially altering regulation for nearby genes. Thus, structural variation among inbred lines could result in gene regulation through ERR-like examples in which a distal enhancer interacts with different loci based on structural variation.
Many structural variations are due to polymorphic TE insertions. We explored the relationship between TEs and the existence of long-range interactions. Prior work postulated that chromatin interactions involving putative promoters and their gene-proximal regions, predated the TE proliferation inserted within the interactions14,51,52. The interaction distance became gradually farther with the TE proliferation, and thus the ancestral gene-proximal regulatory regions became distal regulatory elements that must interact through long-range interactions. We found many examples, such as the pmg1 locus (Fig. 1g), in which similar interactions were maintained in B73 and Mo17 even though very different sequences and transposons were present within the loop middle region of the interaction. It was possible that, with continual TE insertions, some genes located in the interacting anchors became functionally lost pseudogenes, intergenic regions, or even TEs. In supporting this, a number of interactions displayed gene-TE interactions (here TE refers to a real TE or a region with TE inserted) (Supplementary Fig. 14). During this process, some interactions could also be interrupted, and the associated genes had the basal promoter function alone.
Heterotic B73-Mo17 reciprocal hybrids include examples of unique long-range interactions relative to the parents. The hybrid-specific interactions include several examples of over-dominant expression patterns53,54,55,56. The exact molecular mechanisms that lead to over-dominant expression for specific loci are not well understood. Hybrid-specific interactions may be one of the mechanisms that allow for unexpected expression patterns in hybrids relative to the inbred parents. We also assessed allele-specific gain or loss of interactions in the hybrids, which provides evidence for potential trans-acting influences on long-range interactions. We also find that genes located near the allele-specific gain or loss of long-range interactions in hybrids are enriched for compensatory allele-specific expression patterns. Nevertheless, there are several potential limitations to be noted: firstly, only a very limited number of hybrid-specific, and allele-specific gain or loss interactions were detected. We proposed that this is largely due to the unsaturated dataset and the data imbalance in read coverage between replicates (e.g., 51,181/16,992 interactions detected in BM H3K27ac and 14,955/37,354 in MB H3K4me3). Difference between the numbers of certain types of differential interactions doesn’t reflect the difference in biology; for example, it seems there are more BM-specific interactions for H3K4me3 and more MB-specific interactions for H3K27ac based on Fig. 4a, while this difference was demonstrated to be due to the obvious variations in the dataset size (after balancing the data volume and re-detecting the BM- and MB-specific interactions, we did not observe a obviously repeatable difference in loop numbers between different type of genotypes or histone modifications (Supplementary Fig. 15, Supplementary Table 3). Overall, a nearly saturated dataset, as well as experimental validation (e.g., by employing CRISPR-Cas based system) is needed to support these initial findings in future. Besides, consisting with previous studies in mice, wheat, and barley25,40,57, we detected a small number of inter-allelic specific interactions between homologous chromosomes in the hybrids. This number could be much greater, as it was highly limited by the numbers of SNPs and valid read pairs, as well as the strategy to detect loops by requiring the distance between anchors, and so on. It is possible that some unique aspects of these chromosomal regions, such as some specific transcription factor(s) or other mechanisms, might lead to interactions for these regions.
Finally, we expect that our current findings represent only a subset of the full set of inbred-specific or hybrid-specific long-range interactions. However, our findings highlight the potential for unique interactions in hybrids relative to the inbred parents and suggest that these differences may explain some of the unexpected gene expression patterns observed in hybrids.
Methods
Plant materials and growth conditions
The seedlings of B73, Mo17, and their reciprocal crosses (B73 crossed by Mo17 (B73 as the maternal line, abbreviated as B73xMo17), and Mo17 crossed by B73 (Mo17 as the maternal line; abbreviated as Mo17xB73)) were planted in the light incubator (Ningbo Ledian Company, Model: RLD-1000I-2DZ) at 28 °C for 14 hs in light and at 22 °C for 10 h in darkness, and the humidity was 60% relative humidity. The above-ground tissues of whole plants were harvested 14 days after sowing (DAS).
ChIP-seq library construction
The ChIP-seq libraries were constructed as reported44 with minor modifications. Briefly, seedlings of 14 DAS were harvested in the chamber and sequentially dual crosslinked with EGS (ethylene glycol bis (succinimidyl succinate)) and formaldehyde, followed by glycine quenching. Take 2 grams of 14-DAS seedling as starting material with 2 μg antibody of relevant histone modification for immunoprecipitation for each ChIP-seq round, and the Protein A/G beads were reduced from 40 μL to 10 μL bed volume. Enrichment checks of ΔΔCt>3 in qPCR were further utilized for library preparation and X-ten 150PE sequencing. All the FRiP values in the ChIP-seq data sets were between 0.27 ~ 0.60.
HiChIP library construction
The HiChIP libraries were constructed with H3K4me3 (Abcam, ab8580) and H3K27ac (Abcam, ab4729) antibodies for maize plants. Two grams of crosslinked seedlings were ground into fine powder with liquid nitrogen, and the nuclei were retrieved with identical buffers44. The nuclei were suspended in 500 μL 1.2xNEBuffer 3.1, put into 15 μL 10%(w/v) SDS to a final concentration of 0.4% and incubated at 62 °C for 5 min. When it cooled down to room temperature, another 50 μL 10% Triton was added, mixed gently, and incubated at 37 °C for 15 min. The enzyme DpnII was selected in our HiChIP, and 7.5 μL of 50 U/μL DpnII was used for each reaction. After 2-hour incubation on the ThermoMixer at 37 °C, inactivated the enzyme at 62 °C for 20 min. The following steps were similar to those in the original HiChIP protocol6.
Strand-specific RNA-seq library construction
The 14-DAS seedlings for B73 and Mo17 inbreds and their F1 hybrids were harvested. For each genotype, the above-ground parts of at least three plants were collected using a scissor and immediately snap freezing in liquid nitrogen. After the extraction of total RNA44, the ssRNA-seq libraries were constructed with the VAHTS Stranded mRNA-seq Library Prep Kit for Illumina V2 (Nanjing Vazyme, NR612-01) according to the manufacturer’s manual.
Reference genomes
Four reference genomes were used in this study for analysis. The B73 AGPv4 reference genome and Mo17 (CAU version1) reference genome were downloaded from MaizeGDB (https://maizegdb.org/). A total of 9,867,467 SNP data of the Mo17 genome relative to B73 AGPv4 genome was retrieved from a previous publication16. In addition, 8,817,085 SNPs in the B73 genome relative to Mo17 genome were identified by SyRI58. To generate a “simulated Mo17 reference genome”, we replaced corresponding nucleotides on SNP positions inB73 AGPv4 reference genome by Mo17 nucleotides. The “simulated B73 reference genome” was built by replacing corresponding nucleotides on SNP positions in Mo17 reference genome by B73 nucleotides.
ChIP-seq data processing
ChIP-seq raw reads were preprocessed using Trim_galore with default settings (https://www.bioinformatics.babraham.ac.uk/projects/trim_galore/). All samples were aligned with B73 or Mo17 reference genome using Bowtie259. Reads with MAPQ > 20 were retained for further analysis. For each sample, aligned reads from the two reference genomes (B73- and Mo17-reference genome) were combined and sorted using SAMtools60 and combined. ChIP-seq peaks were then identified using MACS2 callpeak function with parameters “ --shift -100 --extsize 200 --nomodel -B --SPMR -g 1.7e9 61. The Irreproducibility Discovery Rate (IDR)62 method was used to assess the repeatability between two replicates.
HiChIP data processing
For B73 samples, raw HiChIP reads were aligned to the B73 AGPv4 reference genome or “simulated Mo17 reference genome” using Bowtie259. For Mo17 samples, raw HiChIP reads were aligned against Mo17 reference genome or“simulated B73 reference genome”. For B73xMo17 and Mo17xB73 samples, raw HiChIP reads were aligned to B73 or Mo17 reference genome. Alignments with MAPQ > 5 were retained for calling valid read pairs using the HiC-Pro pipeline v2.11.363. Combined valid pairs reads from two reference genomes were then used to call Peak-to-All interaction interactions using FitHiChIP46 with 5 Kb bin size. ChIP-seq peaks produced from merged replicates in corresponding antibodies were used as anchors for interaction calling. Lower and upper distance thresholds of interactions between two segments were set as defaults to be 20 Kb and 2 Mb, respectively. Significant interactions were determined using coverage bias regression correction with FDR < 0.01.
In addition, for calculating the mapping efficiency aligning into different reference genomes in Fig. 1, B73 HiChIP data were also processed using the HiC-Pro pipeline but aligned against the Mo17 reference genome to assess alignment difference. Meanwhile, Mo17 HiChIP data were processed using the HiC-Pro pipeline but aligned against the B73 reference genome.
Structure variation detection
The B73 AGPv4 genome was mapped to the Mo17 genome using the minimap264 with default settings. To reduce computation, the alignment occurs between the corresponding chromosomes, e.g., between B73 Chr1 and Mo17 Chr1. Then, structure variations based on B73 reference genome were detected using SyRI58. Meanwhile, structure variations based on Mo17 reference genome were also identified using minimap2-SyRI pipeline.
RNA-seq data processing
Raw RNA-seq reads data were preprocessed using Trim_galore. The pre-processed reads of inbred and hybrid samples were aligned against B73 AGPv4 or Mo17 reference genome using HISAT265. Uniquely mapped reads were assigned to and counted for B73 or Mo17 gene models using HTSeq-count v0.11.266. DESeq267 was used to identify differentially expressed genes (DEGs) between genotypes in pairwise. Additionally, uniquely mapped reads were also used to measure gene expression (i.e., FPKM) by StringTie68.
Gene regulatory divergence assay
B73 and Mo17-allele specific expression was calculated in hybrid RNA-seq data using SNPs between B73 and Mo17. Differential expressed (DE) genes between genotypes were identified using DESeq2 with absolute log2 Fold Change > 1 and FDR < 0.05. Significant expression differences between B73/Mo17 in inbreds and B73/Mo17 in either of hybrids (BM or MB) were determined using Fisher exact tests with FDR < 0.05. Genes in B73 or Mo17 gene model following seven categories including cis only, trans only, cis + trans, cis x trans, compensatory, conserved, and ambiguous were classified following the approach reported in McManus et al.69. To simplify comparisons, we classified trans only, cis + trans and cis x trans into “trans” regulatory patterns.
Conserved and high-confidence differential interaction identification between genotypes
To call differential interactions, raw H3K4me3 or H3K27ac interactions in two replicates of genotype A and genotype B were processed using the FitHiChIP “DiffAnalysisHiChIP.r” function with default settings. The interactions with FDR < 0.05 were identified as specific interactions. Significant interactions (Peak-to-All) called from replicates of genotype A and replicates of genotype B, were used to detect conserved interactions between two genotypes in either H3K4me3 or H3K27ac. The overlapped interactions from replicates of genotype A and replicates of genotype B were retained, and those with FDR > 0.5 in FitHiChIP “DiffAnalysisHiChIP.r” analysis were further selected as conserved interactions.
Hybrid gain and hybrid loss interactions
To understand valid read pair divergence between inbred and hybrid, we separately tested two different types of interactions (hybrid gain interactions and hybrid loss interactions) in hybrids depending on the interaction’s status (conserved or high-confidence differential) between inbred samples. When aligning the inbred HiChIP data to the corresponding reference genome, we observed an apparent heterozygosity (>0.2) at a small subset of single-nucleotide positions (0.78% of SNPs in B73 and 0.99% of SNPs in Mo17). These are likely false-positive ‘heterozygous’ sites that are the result of mis-mapping of reads from highly similar duplicated sequences and could obscure the parental origin of interactions during the identification of hybrid losses and gains. Therefore, such positions were also filtered out and excluded from the determination of parental origin.
For hybrid loss interactions, we collected coordinates of previously identified conserved interactions between B73 and Mo17 and employed a two-sided Fisher exact test to test the ratio of B73 and Mo17 valid read pairs between inbreds and allele-specific B73 valid read pairs and allele-specific Mo17 valid read pairs in either of the hybrids. The Benjamin-Hochberg procedure70 was used for multiple testing corrections.
For hybrid gain interaction, we used inbred-specific interactions as background. In our aforementioned analysis, inbred-specific interactions were identified using a Peak-to-All mode. Consequently, some of the specific interactions detected between B73 and Mo17 may result from the absence of ChIP-seq peaks rather than the actual absence of HiChIP reads. To avoid the confounding effects of these discrepancies on the identification of hybrid gains, we excluded such differential interactions from subsequent analysis.
Any interactions with adjusted p-value less than 0.05 and a difference in the proportion of reads derived from B73 and Mo17 between inbred lines and hybrids greater than 0.3 were considered as unexpected cases and were classified as hybrid gain or hybrid loss interactions.
Inter-allelic interaction identifications in F1 HiChIP data
We inspected the number of covered inter-allelic valid read pairs between B73 allele and Mo17 allele using SNPs between B73 and Mo17. First, left end (R1) and right end (R2) reads of HiChIP data from PE sequencing were mapped to B73 or Mo17 reference genome separately using bowtie2. Unmapped reads were remapped to genome after removing the ligation motif. Second, B73 or Mo17-allele specific reads were filtered. “B73-allele read” means the read can be mapped to B73-ref without SNPs or Indels, while it can’t be mapped to Mo17-ref or can be mapped to Mo17-ref with SNPs or Indels. The same principle applies for “Mo17-allele read”. The candidate inter-allelic pairing reads need to satisfy: R1 belonging to “B73-allele read” and R2 belonging to “Mo17-allele read”, or R2 belonging to “B73-allele read” and R1 belonging to “Mo17-allele read”. Third, inter-allelic interactions were identified. Only unique mapped sole-source reads were kept for interaction detection. Candidate inter-allelic pairing reads were assigned to 5 kb genome region, and total number of pairing inter-allelic reads were calculated in 5 kb region. The criterion of inter-allelic interaction in hybrid lines was: 1) having no inter-allelic pairing reads in inbred lines; 2) having >=2 inter-allelic pairing reads in BM and MB.
Construction of in silico hybrid datasets
We constructed in silico hybrid datasets by sampling equal amounts of data from each parent (B73 and Mo17). Specifically, we randomly selected 130 Mb of paired-end reads from each parent and combined them to create an in silico hybrid dataset with a total of 260 Mb paired-end reads, matching the largest dataset size among our hybrid HiChIP data. We generated four in silico hybrid datasets: in silico hybrid H3K4me3 rep1, and in silico hybrid H3K4me3 rep2, in silico hybrid H3K27ac rep1, in silico hybrid H3K27ac rep2.
When we utilized in silico hybrid datasets as a negative control for identifying hybrid-specific and allele-specific interactions, we also applied the FitHiChIP “DiffAnalysisHiChIP.r” function pipeline as used for the actual hybrid dataset. In the identification of in silico hybrid losses, conserved interactions between B73 and Mo17 were used as the background. However, since the in silico hybrid data were generated by randomly sampling reads from the inbred HiChIP datasets, an imbalance in the number of reads from B73 and Mo17 could occur even within conserved interaction regions. Such imbalances may result in false-positive hybrid losses. To address this, we first removed regions exhibiting read imbalance prior to the identification of in silico hybrid losses. Specifically, regions were excluded if the absolute difference between the proportion of B73 reads in the in silico dataset and that in the original inbred HiChIP data exceeded 0.3.
4C-seq library construction
For 4C-seq library construction, we used the enzyme combination of DpnII-NlaIII and followed the detailed steps as published44. When the 4 C template was prepared, we performed two-round PCR instead of using ultralong primers in one-round PCR. We performed the first round PCR of 16 cycles, followed by 0.8 x beads purification to 20 ul ddH2O. We took 5 μL for the second PCR with 14 cycles, checked the products on 1.5% agarose gel, and purified the remaining with 0.8 x beads to 20 μL ddH2O for Nova 2 × 150 bp sequencing. The principles for primer design should be highlighted here. For a satisfactory reverse PCR, the products amplified with a pair of 20 nt specific primers of the viewpoint were firstly amplified with 18-cycle PCR and would be regarded usable if the majority of PCR product was smaller than 700 bp in size, no over-focused undigested or re-ligated band, and a relatively equivalently amplified smear. When designing the first round PCR primers after accessing these short specific primers, two sequences of 5’-ACACTCTTTCCCTACACGACGCTCTTCCGATCT-3’ and 5’-GTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT-3’ were flanked 5’ on the specific primers as forward and reverse respectively. When designing the second round PCR primers, a primer pair of 5’-AATGATACGGCGACCACCGAGATCTACACTCTTTCCCTACACGACGCTCTTCCGATCT-3’ and 5’-CAAGCAGAAGACGGCATACGAGATNNNNNNGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT-3’ were used and the six N bases depicted the index for sample pooling. 4 C data was processed using the 4C-ker pipeline71.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data generated in this study have been deposited in the NCBI database under BioProject accession PRJNA825068. Additionally, the data from this study can be viewed interactively on the UCSC genome browser [https://genome.ucsc.edu/s/LiangLab/B73_Mo17_hybrid]. Source data are provided with this paper.
Code availability
The code used in this study is available at Github [https://github.com/Maxuxu-IBCAS/Maize-HiChIP].
References
Grubert, F. et al. Genetic control of chromatin states in humans involves local and distal chromosomal interactions. Cell 162, 1051–1065 (2015).
Tang, Z. et al. CTCF-mediated human 3D genome architecture reveals chromatin topology for transcription. Cell 163, 1611–1627 (2015).
Lieberman-aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–294 (2009).
Fullwood, M. J. et al. An oestrogen-receptor-α-bound human chromatin interactome. Nature 462, 58–64 (2009).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Mumbach, M. R. et al. HiChIP: Efficient and sensitive analysis of protein-directed genome architecture. Nat. Methods 13, 919–922 (2016).
Fang, R. et al. Mapping of long-range chromatin interactions by proximity ligation-assisted ChIP-seq. Cell Res 26, 1345–1348 (2016).
Mumbach, M. R. et al. Enhancer connectome in primary human cells identifies target genes of disease-associated DNA elements. Nat. Genet. 49, 1602–1612 (2017).
Huang, H. et al. CTCF mediates dosage- and sequence-context-dependent transcriptional insulation by forming local chromatin domains. Nat. Genet. 53, 1064–1074 (2021).
Rowley, M. J. et al. Evolutionarily conserved principles predict 3D chromatin organization. Mol. Cell 67, 837–852.e7 (2017).
Petrovic, J. et al. Oncogenic notch promotes long-range regulatory interactions within hyperconnected 3D Cliques. Mol. Cell 73, 1174–1190.e12 (2019).
Weintraub, A. S. et al. YY1 Is a Structural regulator of enhancer-promoter loops. Cell 171, 1573–1588.e28 (2017).
Concia, L. et al. Wheat chromatin architecture is organized in genome territories and transcription factories. Genome Biol. 21, 104–124 (2020).
Ricci, W. A. et al. Widespread long-range cis-regulatory elements in the maize genome. Nat. Plants 5, 1237–1249 (2019).
Xu, G. et al. Evolutionary and functional genomics of DNA methylation in maize domestication and improvement. Nat. Commun. 11, 5539–5551 (2020).
Sun, S. et al. Extensive intraspecific gene order and gene structural variations between Mo17 and other maize genomes. Nat. Genet. 50, 1289–1295 (2018).
Lai, J. et al. Genome-wide patterns of genetic variation among elite maize inbred lines. Nat. Genet. 42, 1027–1030 (2010).
Hufford, M. B. et al. De novo assembly, annotation, and comparative analysis of 26 diverse maize genomes. Science 373, 655–662 (2021).
Haberer, G. et al. European maize genomes highlight intraspecies variation in repeat and gene content. Nat. Genet. 52, 950–957 (2020).
Yang, N. et al. Genome assembly of a tropical maize inbred line provides insights into structural variation and crop improvement. Nat. Genet. 51, 1052–1059 (2019).
Choudhary, M. N. et al. Co-opted transposons help perpetuate conserved higher-order chromosomal structures. Genome Biol. 21, 16–30 (2020).
Choudhary, M. N., Quaid, K., Xing, X., Schmidt, H. & Wang, T. Widespread contribution of transposable elements to the rewiring of mammalian 3D genomes. Nat. Commun. 14, 634–646 (2023).
Diehl, A. G., Ouyang, N. & Boyle, A. P. Transposable elements contribute to cell and species-specific chromatin looping and gene regulation in mammalian genomes. Nat. Commun. 11, 1796–1814 (2020).
Zhao, L. et al. Chromatin loops associated with active genes and heterochromatin shape rice genome architecture for transcriptional regulation. Nat. Commun. 10, 3640–3653 (2019).
Yuan, J. et al. Open chromatin interaction maps reveal functional regulatory elements and chromatin architecture variations during wheat evolution. Genome Biol. 23, 34–55 (2022).
Schnable, P. S. et al. The B73 Maize genome: complexity, diversity, and dynamics. Science 326, 1112–1115 (2009).
Walbot, V. 10 reasons to be tantalized by the B73 maize genome. PLOS Genet 5, e1000723 (2009).
Messing, J. et al. Sequence composition and genome organization of maize. Proc. Natl Acad. Sci. Usa. 101, 14349–14354 (2004).
Buckler, E. S., Gaut, B. S. & McMullen, M. D. Molecular and functional diversity of maize. Curr. Opin. Plant Biol. 9, 172–176 (2006).
Anderson, S. N. et al. Transposable elements contribute to dynamic genome content in maize. Plant J. 100, 1052–1065 (2019).
Flint-Garcia, S. A., Buckler, E. S., Tiffin, P., Ersoz, E. & Springer, N. M. Heterosis is prevalent for multiple traits in diverse maize germplasm. PLoS One 4, e7433 (2009).
Hochholdinger, F. & Hoecker, N. Towards the molecular basis of heterosis. Trends Plant Sci. 12, 427–432 (2007).
Birchler, J. A., Auger, D. L. & Riddle, N. C. In search of the molecular basis of heterosis. Plant Cell 15, 2236–2239 (2003).
Birchler, J. A., Yao, H., Chudalayandi, S., Vaiman, D. & Veitia, R. A. Heterosis. Plant Cell 22, 2105–2112 (2010).
Chen, Z. J. Genomic and epigenetic insights into the molecular bases of heterosis. Nat. Rev. Genet. 14, 471–482 (2013).
He, G. et al. Conservation and divergence of transcriptomic and epigenomic variation in maize hybrids. Genome Biol. 14, R57–R72 (2013).
Hofmann, N. R. A global view of hybrid vigor: DNA methylation, small RNAs, and gene expression. Plant Cell 24, 841–841 (2012).
Zhu, W. et al. Altered chromatin compaction and histone methylation drive non-additive gene expression in an interspecific Arabidopsis hybrid. Genome Biol. 18, 1–16 (2017).
Hu, Y. et al. Comparison of dynamic 3D chromatin architecture uncovers heterosis for leaf size in Brassica napus. J. Adv. Res. 42, 289–301 (2022).
Mascher, M. et al. A chromosome conformation capture ordered sequence of the barley genome. Nature 544, 427–433 (2017).
Lin, Y., Zhao, H., Kotlarz, M. & Jiang, J. Enhancer-mediated reporter gene expression in Arabidopsis thaliana: a forward genetic screen. Plant J. 106, 661–671 (2021).
Zhu, B., Zhang, W., Zhang, T., Liu, B. & Jiang, J. Genome-wide prediction and validation of intergenic enhancers in Arabidopsis using open chromatin signatures. Plant Cell 27, 2415–2426 (2015).
Jiao, Y. et al. Improved maize reference genome with single-molecule technologies. Nature 546, 524–527 (2017).
Li, E. et al. Long-range interactions between proximal and distal regulatory regions in maize. Nat. Commun. 10, 2633–2647 (2019).
Peng, Y. et al. Chromatin interaction maps reveal genetic regulation for quantitative traits in maize. Nat. Commun. 10, 2632–2643 (2019).
Bhattacharyya, S., Chandra, V., Vijayanand, P. & Ay, F. Identification of significant chromatin contacts from HiChIP data by FitHiChIP. Nat. Commun. 10, 4221–4235 (2019).
Swanson-Wagner, R. A. et al. All possible modes of gene action are observed in a global comparison of gene expression in a maize F1 hybrid and its inbred parents. Proc. Natl Acad. Sci. Usa. 103, 6805–6810 (2006).
Schnable, P. S. & Springer, N. M. Progress toward understanding heterosis in crop plants. Annu. Rev. Plant Biol. 64, 71–88 (2013).
Li, G. et al. Extensive promoter-centered chromatin interactions provide a topological basis for transcription regulation. Cell 148, 84–98 (2012).
Oh, S. et al. Enhancer release and retargeting activates disease-susceptibility genes. Nature 595, 735–740 (2021).
Lu, Z. et al. The prevalence, evolution and chromatin signatures of plant regulatory elements. Nat. Plants 5, 1250–1259 (2019).
Wang, M. et al. Comparative genome analyses highlight transposon-mediated genome expansion and the evolutionary architecture of 3D genomic folding in cotton. Mol. Biol. Evol. 38, 3621–3636 (2021).
Xiao, Y. et al. The genetic mechanism of heterosis utilization in maize improvement. Genome Biol. 22, 148–177 (2021).
Paschold, A. et al. Complementation contributes to transcriptome complexity in maize (Zea mays L.) hybrids relative to their inbred parents. Genome Res 22, 2445–2454 (2012).
Li, L. et al. Dominance, overdominance and epistasis condition the heterosis in two heterotic rice hybrids. Genetics 180, 1725–1742 (2008).
Huang, X. et al. Genomic analysis of hybrid rice varieties reveals numerous superior alleles that contribute to heterosis. Nat. Commun. 6, 6258–6267 (2015).
Han, Z. et al. Diploid genome architecture revealed by multi-omic data of hybrid mice. Genome Res 30, 1097–1106 (2020).
Goel, M., Sun, H., Jiao, W.-B. & Schneeberger, K. SyRI: finding genomic rearrangements and local sequence differences from whole-genome assemblies. Genome Biol. 20, 277–290 (2019).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Li, Q., Brown, J. B., Huang, H. & Bickel, P. J. Measuring reproducibility of high-throughput experiments. Ann. Appl. Stat. 5, 1752–1779 (2011).
Servant, N. et al. HiC-Pro: An optimized and flexible pipeline for Hi-C data processing. Genome Biol. 16, 259–270 (2015).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Anders, S., Pyl, P. T. & Huber, W. HTSeq-a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550–571 (2014).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
McManus, C. J. et al. Regulatory divergence in Drosophila revealed by mRNA-seq. Genome Res 20, 816–825 (2010).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. 57, 289–300 (1995).
Raviram, R. et al. 4C-ker: A method to reproducibly identify genome-wide interactions captured by 4C-Seq experiments. PLoS Comput. Biol. 12, e1004780 (2016).
Acknowledgements
This work was funded by National Natural Science Foundation of China (32422063) to M.Z., and the National Science Foundation (NSF) Plant Genome Research Program (IOS-1934384) to N.M.S.
Author information
Authors and Affiliations
Contributions
M.Z. and H.L. designed the project. H.L. performed all the experiments, including the preparation of HiChIP, ChIP-seq, and 4C-seq libraries. X.M., Z.K.L., S.J., and W.D. performed the computational analysis. X.M., Z.K.L., Z.R.L, and others produced and improved the figures. H.L., Z.K.L., N.M.S., X.M., and M.Z. wrote and revised the manuscript. All the authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Bao Liu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, H., Ma, X., Jiang, S. et al. Conservation and variability of long-range interactions in structurally diverse maize genomes. Nat Commun 16, 10105 (2025). https://doi.org/10.1038/s41467-025-65065-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65065-8