Abstract
Metabolic biomarker discovery in trace body fluids remains a significant challenge, toward molecular diagnosis and pathology studies in many diseases. Especially for eye-related diseases, such an approach based on non-invasive tear fluids remains an unsatisfied urgent need in ophthalmology. Here we construct a metabolic biomarker panel from 10 nL of tear fluids in seconds using nanoparticle-enhanced laser desorption/ionization -mass spectrometry (MS), which achieves an area under the curve of 0.923 for discriminating diabetic cataracts from alone age-related cataracts. Importantly, we integrate liquid chromatography -MS into the above analysis process to construct an integrated strategy, allowing reliable metabolite annotation by nanoliter sample volume without compromising high throughput. Further, using matched aqueous humors, we identify 1,5-anhydroglucitol as a biomarker of diabetic cataracts, revealing its protective effect against high glucose-induced lens oxidative stress and opacification, as a demonstration of the metabolic reprogramming. Our approach can be universally applied to uncover biomarkers using trace body fluid, promising next-generation metabolic reprogramming identification.
Similar content being viewed by others
Introduction
Cataract, a clouding of the lens, is the leading cause of visual impairment worldwide, affecting more than 95 million people1. With the increasing global incidence of diabetes, the efficient treatment and prevention of diabetic cataracts (DCs) are of growing importance to mitigate the heavy burden of chronic disease2,3. Clinically, cataract surgery treatment requires effective preoperative diagnosis of DCs to minimize complications4. Nevertheless, most clinical diagnoses of DC are based on patients’ self-reported diabetic history or simple preoperative one-point blood tests with a considerable underdiagnose rate of up to 60%, whereas comprehensive evaluations of diabetes are too cumbersome to apply to all cataract patients5,6,7. Also, further pathological explorations of DCs are still needed to achieve a satisfactory preventive effect8,9. In this regard, molecular biomarkers are urgently required for not only rapid and precise diagnosis but also further pathological explorations of DC. Unlike protein and RNA, metabolic biomarkers are closely linked to the disease phenotype and can be manipulated for metabolic programming10,11, making them attractive for improving DC diagnosis and pathology elucidation.
Non-invasive diagnostic technology exhibits powerful clinical utility due to its nature of being pain-free, low complication risk, and patient-friendly12,13. Metabolic biomarker screening based on non-invasive body fluids promises such a diagnostic system regardless of personal clinical experience and imaging changes regarding diseases14. Especially for DC, there is an urgent clinical need in ophthalmology for a non-invasive metabolic diagnostic approach that has not yet been fulfilled. Due to the blood-eye barrier, previous metabolomic studies focused on invasive intraocular samples, such as aqueous humors (AHs), with limited diagnostic prospects15,16. In contrast, tear fluids are uniquely non-invasive ocular body fluids and thus ideally suitable for the non-invasive diagnosis of DC. Besides, a joint analysis for metabolic biomarkers in tear fluids and intraocular samples like AHs promises a more comprehensive understanding of signature metabolic programming in DC eyes, aiding pathology elucidation. Nevertheless, the limited volume of non-stimulated tear film (single-digital microliters) is a significant challenge for discovering tear metabolic biomarkers17,18, highlighting the importance of developing appropriate metabolic analysis tools.
Mass spectrometry (MS) offers a broad metabolic profile with molecular identification capability by measuring mass to charge ratios (m/z) of ions19,20. Currently, mainstream MS technology for metabolic analysis relies on gas chromatography (GC) or liquid chromatography (LC), which requires tens to hundreds of microliter samples with long-time pretreatments and chromatographic separations21,22. However, when it comes to analyzing clinical tear fluid samples of very limited volume, traditional LC-MS and GC-MS are inadequate for metabolic detection23. In contrast, nanoparticle-enhanced laser desorption/ionization MS (NELDI-MS) can achieve direct solid-phase metabolic detection for metabolites within tens of seconds by less than 1 μL of body fluid samples23,24,25. This makes it a compelling tool for metabolic detection and biomarker discovery in trace body fluid samples, including tear fluids.
Despite its promising application for discovering metabolic biomarkers in trace body fluid samples, NELDI-MS faces challenges related to metabolite annotation. The current feature annotation strategy in most NELDI-MS-based metabolic biomarker studies is primarily based on 1-dimensional (1-D) m/z information feature matching with higher resolution MS platforms like Fourier transform ion cyclotron resonance (FT-ICR) MS and LC-MS/MS23,24,26, which may lead to annotation errors due to ion adduction variation and diverse metabolites with similar molecular weights in NELDI-MS. An improved 2-D information feature matching strategy, considering both m/z and fold change (FC), can exclude incorrect matching results25. However, this strategy’s reliance on LC-MS/MS analysis for each individual sample limits NELDI-MS’s advantages in throughput and trace body fluid detection. Overcoming these challenges is essential for realizing the full potential of NELDI-MS in identifying metabolic biomarkers in trace tear fluids.
Herein, based on metabolic analysis by the high-performance NELDI-MS platform (Fig. 1), we constructed a diagnostic biomarker panel from trace tear fluids for discriminating DCs and alone age-related cataracts (ARCs), with an area under the curve (AUC) of 0.923, a sensitivity of 85.9%, and a specificity of 82.0%. The metabolic analysis takes only a detection time of 30 s per sample with a tear fluid consumption of 10 nL, achieving a rapid, precise, and non-invasive diagnosis of DC. For metabolite annotation, we proposed a rapid 2-D information feature matching strategy based on trace samples (R2DIFMS-TS) and annotated the NELDI-MS features in the panel as biomarkers by LC-MS/MS with a tear fluid consumption of 140 nL, providing a desirable solution for discovering metabolic biomarkers in trace body fluid samples. Further, using this solution, we identified 1,5-anhydroglucitol (1,5-AG) as a new anti-cataract AH biomarker of DC, revealing the unique but closely related metabolic disorders on the ocular surface and within the eye of DC patients. 1,5-AG exhibited a protective effect against high glucose-induced lens oxidant stress and opacification, which suggests its potential key role in DC development and highlights the reliability and excellent application prospects of our solution.
The tear fluid metabolic fingerprints (TMFs) and aqueous humor metabolic fingerprints (AHMFs) were extracted from clinical samples of alone age-related cataract (ARC) and diabetic cataract (DC) patients using the high-performance nanoparticle-enhanced laser desorption/ionization mass spectrometry (NELDI-MS) within nanoliter-scale and second-scale. Features in TMFs or AHMFs were first selected by machine learning, fold changes (FCs) and p-values. Next, features selected were annotated as metabolic biomarkers using the rapid 2-D information feature matching strategy based on trace samples (R2DIFMS-TS) proposed here, supporting diagnostic application and pathologic exploration.
Results
Construction of the NELDI-MS platform for metabolic analysis
To achieve high-performance metabolic analysis, the NELDI-MS platform with ferric nanoparticles (NPs, Fig. 2a) was constructed, which featured low sample consumption and high throughput. The ferric NPs were prepared by an optimized solvothermal method, and the uniform distribution of the Fe and O elements on the NPs was characterized by the high-angle annular dark field and elemental mapping analysis (Fig. 2b). Further, the morphology and crystal structure of the NPs were respectively confirmed by scanning electron microscopy and X-ray diffraction (Supplementary Fig. 1). Using the ferric NPs as the matrix, the microarrayed NELDI-MS chip achieved automatic m/z data acquisition from up to 384 samples with 1 μL of loading volume, where the pretreatment and detection for each sample could be finished within 1 min and 30 s respectively. For detection reproducibility, the NELDI-MS showed desirable coefficients of variation (CVs) of 2.4% to 4.9% in the typical metabolite detection among eight independent tests (Fig. 2c), which was critical for reproducible and credible metabolic analysis. Importantly, the signal response for those typical metabolites was enhanced by one to three orders of magnitude by the NELDI-MS compared to the commercial organic matrix-assisted LDI-MS (MALDI-MS, Fig. 2d), supporting the metabolic analysis in trace body fluid samples.
a Illustration of the NELDI-MS platform. The lower left digital image showed the on-chip microarray of sample spots after the ferric matrix printing (Scale bar 5 mm). b High-angle annular dark field (HAADF) and elemental mapping analysis for the ferric nanoparticles (NPs) with Fe in red and O in green. The scale bar was 100 nm. c, d Detection for standard metabolites (concentrations of 1 mg mL−1) of glutamate (Glu), lysine (Lys), arginine (Arg), glucose (Glc), and mannitol (Man) using the NELDI-MS or the organic matrix-assisted LDI-MS. The data was acquired in eight independent tests. The coefficients of variation (CVs) of the NELDI-MS detection for those standard metabolites were in (c). The MS intensity comparison of those standard metabolites between the NELDI-MS (using the ferric nanoparticles, NPs) and organic matrix-assisted LDI-MS (using 2,5-dihydroxybenzoic acid, DHB, or using α-cyano-4-hydroxy-cinnamic acid, CHCA) was in (d). The data in d was expressed as means ± standard errors and p-values in the analysis of variance (ANOVA) with two-sided Tukey post-hoc tests were shown. e The distribution and median of CVs of mass to charge ratio (m/z) features extracted from the representative mixture tear fluid and AH samples by the NELDI-MS. The line segments indicated quartiles and medians. The CVs were calculated by the data from eight independent tests and the representative mixture samples were prepared by mixing six ARC and six DC samples to include all possible m/z features. f Typical MS spectra of tear fluid samples and AH samples in the ARC and DC groups at m/z of 100–400. Source data are provided as a Source data file.
To demonstrate the advantages of the NELDI-MS over other established direct ionization MS platforms, we performed a performance comparison, regarding the throughput, sensitivity, and reproducibility, between the NELDI-MS and desorption electrospray ionization MS (DESI-MS), the latter of which is a mature direct ionization MS platform for metabolic detetion27,28,29. For throughput, DESI-MS generally needs more than 2 min of detection time to scan each sample spot of 1 μL of loading volume, slower than the NELDI-MS (<30 s for detection). For sensitivity, the NELDI-MS exhibited enhanced performance with limits of detection (LODs) of equal to or less than 0.1 ng for the five typical metabolites (Supplementary Table 1), which were one to three orders of magnitude better than those of DESI-MS (LODs of 0.1 to 50 ng, Supplementary Table 1). Besides, DESI-MS showed limited reproducibility in the eight independent tests for the typical metabolites (Supplementary Fig. 2), with higher CVs (17.0% to 33.2%) than those of the NELDI-MS (2.4% to 4.9%).
The performance of the NELDI-MS was further evaluated in metabolic detection for ocular fluid samples. In MS data acquisition for body fluids, the tear fluid and AH samples were diluted with ultrapure water at 100 folds and 5 folds respectively following the literature procedure23,30, which could avoid interference by the salt crystallization in NELDI-MS detection (Supplementary Fig. 3). From 1 μL of diluted body fluids, corresponding to 10 nL of tear fluids or 200 nL of AHs, the NELDI-MS recorded ~120,000 m/z data points at 100–1000 Da for further m/z feature extraction by peak detection. Consistent with the result in typical metabolite detection, the NELDI-MS also exhibited satisfactory detection reproducibility for the representative mixture tear fluid and AH samples, where the median of CVs of m/z features is 12.2% and 10.7%, respectively (eight independent tests, Fig. 2e). Importantly, the typical MS spectra (Fig. 2f) demonstrated the strong detection of the NELDI-MS for small metabolites in trace tear fluids and AHs, in which over 350 apparent m/z features were detected and mostly concentrated on the low mass range of 100–400 Da. In sum, the above results demonstrated the high performance of the NELDI-MS for acquiring tear fluid metabolic fingerprints (TMFs) and AH metabolic fingerprints (AHMFs).
Diagnostic machine learning model by TMFs for DC
To support distinguishing DC patients from common ARC patients in ophthalmology clinics, the monocular tear fluids from 168 individuals were collected (200 nL per individual, ARC/DC of 82/86, main cohort) for machine learning based on TMFs and further diagnostic biomarker panel construction. The 168 individuals were randomly split into the discovery and validation cohorts with a proportion of 75% and 25%, for diagnostic machine learning model building and validation. The age and gender information were shown in Fig. 3a and Supplementary Table 2. And, other demographic characteristics, related comorbidity information, as well as diabetic characteristics were summarized in Supplementary Tables 3 and 4.
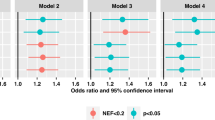
a The cohort design and distribution of age and sex for the cataract cohort, including 82 ARC patients and 86 DC patients. b The heatmap of the 168 TMFs, each of which contained 422 m/z features. c The workflow for algorithm evaluation and hyperparameter tuning in model building. d The comparison of cross-validated area under the curves (AUCs) for the four best-tuned machine learning algorithms in 5-fold cross-validation with 20 rounds, including the Logistic Regression (LR), K-Nearest Neighbor (KNN), Adaptive Boosting (AB), and Decision Tree (DT). The boxes indicated quartiles and medians. The whiskers indicated minimum and maximum values. P values in the ANOVA with two-sided Tukey post-hoc tests were shown. e, f The receiver operating characteristic (ROC) curves by the LR model for diagnosing DC in the discovery cohort (in e) and validation cohort (in f). Source data are provided as a Source data file.
Notably, except ages, no significant differences in the above clinical information existed between the ARC and DC patients (Supplementary Tables 2, 3, and 4). For ages, the two-tailed t-test showed that the ages of DC patients were significantly less than that of ARC patients in the cohort (p value < 0.01, Supplementary Table 2), which was in line with clinical observations2. To eliminate the potential extra interference by age, the stratified analysis was performed in the performance evaluation of the diagnostic model (Supplementary Table 5), which was widely applied to remove biases by covariates31,32. Based on the NELDI-MS analysis, the metabolic data set for the entire 168 tear fluid samples was established. From the original MS data, 422 apparent m/z features were extracted by the peak detection and composed the TMF of each sample. The distribution of m/z features in a total of the 168 TMFs was visually displayed in the heatmap (Fig. 3b). Of note, the TMFs from ARC and DC patients could not be separated by the principal component analysis, a commonly used unsupervised method, indicating machine learning was required to distinguish those by TMFs (Supplementary Fig. 4). In the power analysis of the pilot data (ARC/DC, 15/15), a sample number of 40 per group could achieve a predicated power of 0.84 (FDR = 0.1), demonstrating the sample size was adequate for machine learning with a sufficient confidence level (Supplementary Fig. 5).
For building the TMF-based diagnostic model, the performance of four representative machine learning algorithms with various hyperparameter combinations was evaluated by the averaged cross-validated AUC (5 folds with 20 rounds, Fig. 3c) in the discovery cohort, including the Logistic Regression (LR), K-Nearest Neighbor (KNN), Adaptive Boosting (AB), and Decision Tree (DT). Consequently, the LR algorithm with the best-tuned hyperparameters achieved significantly higher performance than the other three algorithms (p value < 0.001, Fig. 3d). Next, the diagnostic model was trained by TMFs from the discovery cohort with the best-tuned LR algorithm. For diagnosing DC, the LR model achieved high performance with an AUC of 0.956 (95% CI of 0.926 to 0.987), of which the sensitivity was 90.6% when the specificity reached 83.6% (Fig. 3e). Notably, in the validation cohort, the high diagnostic performance of the LR model was validated with the AUC of 0.946 (95% CI of 0.880–1.000), the sensitivity of 90.9%, and the specificity of 90.5%, excluding the risk of overfitting (Fig. 3f). Lastly, to exclude the potential influence of age on the LR model by the stratified analysis, the discovery and validation cohorts were stratified into two age groups by age of 70, the median age of the entire 168 individuals. Consequently, the LR model reached AUCs of 0.934 to 0.974 in all two stratified discovery and two stratified validation cohorts (Supplementary Fig. 6 and Supplementary Tables 5 and 6), demonstrating the high diagnostic performance of the LR model did not arise from the differential ages between the ARC and DC groups. Altogether, the diagnostic model based on TMFs realized a precise, rapid, and non-invasive diagnosis of DC, while constructing a high-performance biomarker panel from complex metabolic fingerprints was still necessary to support potential clinical diagnostic applications.
Feature selection for the diagnostic panel from TMFs
To further support the diagnosis of DC, m/z features in TMFs were selected to construct a diagnostic feature panel. As the process in Fig. 4a, ten key m/z features were first selected from the 422 features by the coefficients in the LR model and FWER p-values (two-tailed t-test, Bonferroni correction) in the discovery cohort. Then, by the AUC evaluation for all combinations of the ten key features, three target features, including the features of m/z 173.0, 337.2, and 348.0, were selected to construct the panel. For group differences, in the DC group, m/z 173.0 and 337.2 were significantly increased with the FCs of 1.55 and 2.51 respectively, whereas m/z 348.0 significantly decreased with the FC of 0.66 (Fig. 4b, FWER p values < 0.001, FCs were calculated by the ratio of DC to ARC).
a The workflow for selecting m/z features for the diagnostic panel. b The intensity comparison of the three target features for ARCs and DCs (ARC/DC, 61/64, discovery cohort). The boxes indicated quartiles and medians. The whiskers indicated minimum and maximum values. FWER p-values in the two-tailed t-tests (Bonferroni correction) were shown. c When using the feature panel or single features, the different ROC curves by the LR model for diagnosing DC in the discovery cohort. d The ROC curve by the feature panel-based LR model for diagnosing DC in the validation cohort. e The ROC curve by the feature panel-based LR model for diagnosing DC in the extra external cohort, which included 79 ARCs and 27 DCs. f The workflow of the R2DIFMS-TS for annotating the NELDI-MS features as metabolites. g The representative extracted ion chromatogram (XIC) partially showing the XIC features that had the matched m/z with the NELDI-MS feature of m/z 337.2. Those five XIC features were labeled with letters from a to e. Only features a and e were finalized as the matching results in the R2DIFMS-TS. The XIC was extracted from the LC-MS/MS data of the mixed tear fluid samples (three replicates with similar results). The metabolite annotations for features a and e were yielded by high-resolution m/z and MS/MS fragments. h The FCs of the target NELDI-MS feature (m/z 337.2 in the TMFs) and its matched XIC features in the R2DIFMS-TS. The FCs were calculated by the ratio of DC to ARC. The bar graph for the NELDI-MS feature represented the data in the TMFs of the discovery cohort (ARC/DC, 61/64), and that for the XIC features represented the data acquired by the three replicated LC-MS/MS analyses for the mixed tear fluid samples. Source data are provided as a Source data file.
For diagnosing DC, in the discovery cohort, the LR model by the three target features achieved an enhanced AUC of 0.923 (95% CI of 0.876 to 0.970), while those features showed limited AUCs of 0.715 to 0.870 when singly employed (Fig. 4c), showing the necessity of the combined use of the three features as a panel. Notably, the LR model by the panel could diagnose DC with a sensitivity of 85.9% and a specificity of 82.0%. In the validation cohort, the LR model by the panel also reached a consistent AUC of 0.896 (95% CI of 0.788–1.000) with a sensitivity of 86.4% and a specificity of 90.5% (Fig. 4d), thus validating the high performance of the panel.
To further evaluate the reproducibility of the feature panel’s between-group differences and diagnostic performance at a larger-scale level, we additionally collected the monocular tear fluids from 106 individuals (200 nL per individual, ARC/DC of 79/27) in this study, as an extra external cohort. These samples were detected in another lab using another MS instrument (Methods section) to evaluate the cross-lab reproducibility of our diagnostic approach. Of note, to ensure those metabolite changes are directly related to DC, this cohort excluded patients with some related comorbidities, which may lead to metabolic disorders. Its cohort information was summarized in Supplementary Tables 7, 8, and 9. Notably, in the extra external cohort, all three targeted features exhibited consistent between-group differences with those in the discovery cohort from the main cohort (Supplementary Fig. 7). In terms of diagnostic performance, the feature panel-based LR model afforded a strong AUC of 0.869 for diagnosing DC (Fig. 4e, 95% CI of 0.799 to 0.939, the sensitivity was 81.5% when the specificity reached 79.8%). These results demonstrated the reproducibility of our findings.
To investigate if the metabolic changes represented by the target features are directly linked to DC formation rather than diabetes itself, we collected the monocular tear fluids from 39 individuals without cataracts as the additional comparison groups, including 20 healthy individuals (healthy control group) and 19 diabetic patients without cataracts (diabetic control group). Its cohort information was summarized in Supplementary Tables 10, 11, and 12. Interestingly, the MS signals of the three target features of m/z 173.0, 337.2, and 348.0, did not show significant differences or apparent change trend (defined by fold changes > 1.2 or <0.83) between the healthy control and diabetic control groups (Supplementary Fig. 8). These results suggested that those metabolic changes may not be simply caused by the single factor of the presence of diabetes, and thus, those changes may be directly linked to DC formation rather than diabetes itself.
In summary, a diagnostic panel consisting of three m/z features was discovered from TMFs and featured high performance for diagnosing DC. While those features still need to be reliably annotated as metabolites.
Metabolite annotation for biomarkers in the diagnostic panel
To achieve the reliable metabolite annotation for the three target NELDI-MS features in TMFs, the R2DIFMS-TS was proposed. This strategy allowed rapid obtaining necessary information supporting the 2-D information (m/z and FC) matching, through LC-MS/MS analysis for the mixed samples rather than each individual sample (Fig. 4f and Supplementary Fig. 9). Specifically, LC-MS/MS analysis for the mixed tear fluid samples (140 nL per sample mixed by group) could offer the three key information items used for feature annotating and matching, including FC, high-resolution m/z, and MS/MS fragments of metabolites, without long-time detection for each individual sample in the cohort. Importantly, the available tear fluid volume for the LC-MS/MS analysis was increased by over 60 folds to the single sample (depending on sample size) while acquisition for the three key information items was unaffected. The 2-D information of m/z and FC could serve as identifiers to match the m/z features arising from the same metabolite in the LC-MS/MS and NELDI-MS, transferring reliable metabolite annotations from LC-MS/MS to NELDI-MS features.
The specific process of the R2DIFMS-TS for annotating the three target NELDI-MS features in TMFs was illustrated by the case of m/z 337.2, the feature with the highest coefficient in the LR model. For initial matching by m/z, extracted ion chromatogram (XIC) features corresponding to m/z 337.2 were obtained from the LC-MS/MS analysis (for the mixed ARC and DC samples), according to the common ion adductions in NELDI-MS metabolic analysis33. For further matching by FC, these XIC features were filtered by the matching criteria that the FC of the XIC feature must be greater than the FC of the NELDI-MS feature, with a consistent trend, considering that the NELDI-MS features could arise from multiple metabolites. As an illustration, in the XIC shown in Fig. 4g, five XIC features from a to e had the matched m/z with this NELDI-MS feature, while only features a and e had the matched FCs (Fig. 4h, Supplementary Fig. 10, and Supplementary Table 13). By high-resolution m/z and MS/MS fragments, XIC features a and e were annotated as two isomeric metabolites, 9,10-DHOME and 12,13-DHOME, which were finalized as the biomarkers corresponding to the NELDI-MS feature of m/z 337.2. Notably, each XIC feature with the matched m/z but unmatched FC, including those not shown here, might be finalized as a matching result in the 1-D m/z information-based feature matching process, but the between-group difference of m/z 337.2 did not arise from those, making the risk of unreliable biomarker discovery. Similarly, using the R2DIFMS-TS, the NELDI-MS features of m/z 173.0 and 348.0 were finally annotated as glycerol 3-phosphate (G3P) and N-acetylneuraminic acid (NeuAc) respectively (Supplementary Figs. 10 and 11 and Supplementary Table 13). In short, the R2DIFMS-TS addressed the problem of reliable metabolite annotation in NELDI-MS-based biomarker discovery, yielding a desired solution for discovering metabolic biomarkers in trace body fluid samples. Using this solution, a high-performance diagnostic biomarker panel of DC was constructed, with a tear fluid consumption of down to 150 nL.
Exploration for DC pathogenesis based on tear and AH biomarkers
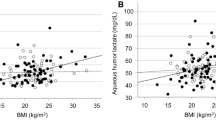
To further reveal DC pathogenesis, the joint analysis for metabolic biomarkers in tear fluids and AHs was performed, based on the 46 matched monocular AH samples collected from the patients undergoing surgery (ARC/DC of 30/16 out of the 168 participants, Fig. 5a and Supplementary Table 14). The AHMFs of the 46 AH samples were recorded by the NELDI-MS, which consisted of 383 apparent m/z features (Supplementary Fig. 12). By evaluating FWER p values (criteria was p value < 0.05, two-tailed t-test, Bonferroni correction) and FCs between the two groups (criteria was FC > 1.2, ARC/DC or DC/ARC), m/z 225.0 was identified as the target feature (Fig. 5b and Supplementary Fig. 13), which was annotated as 1,5-AG as the AH biomarker using the R2DIFMS-TS (Fig. 5c, Supplementary Fig. 14, and Supplementary Table 15). Of note, the three features of the tear biomarkers either did not show consistent differences or failed to be detected in AHs (Supplementary Table 16), indicating differences in metabolic disorders of DC between the ocular surface and within the eye. Nevertheless, in the Spearman correlation analysis, the AHMF feature of 1,5-AG significantly correlated with the TMF feature of the DHOMEs (p value < 0.05, Supplementary Fig. 15), showing the metabolic disorders on the ocular surface and within the eye were different but closely related. In terms of metabolic pathways involved, tear biomarkers were related to energy metabolism, inflammation, and oxidative stress, whereas the AH biomarker was also involved in energy metabolism and oxidative stress. In short, these results suggested there were unique but closely related metabolic disorders between the ocular surface and within the eye, which might be involved in the pathogenesis of DC.
a Illustration of discovering AH biomarkers of DC. b The volcano plots representing FWER p-values (dotted line indicated p-value < 0.05, two-tailed t-test, Bonferroni correction) and FCs (dotted lines indicated FC > 1.2, ARC/DC or DC/ARC) of the 383 features in the AHMFs. c The FCs (DC/ARC) of the AHMF feature of m/z 225.0 and its matched XIC features in the R2DIFMS-TS. This XIC feature was annotated as 1,5-anhydroglucitol (1,5-AG) by high-resolution m/z and MS/MS fragments. The bar graph represented AHMF data (ARC/DC, 30/16) and LC-MS/MS XIC data (three replicates for the mixed AH samples). d, e The expression of the three antioxidant enzymes measured by Western blotting (in d) and qPCR analysis (in e) in the different groups’ HLECs, including superoxide dismutase 1 (SOD1), glutathione peroxidase 1 (GPx1), and catalase (CAT). f The glutathione (GSH) levels of the different groups’ HLECs. g, h The 2,7-dichlorofluorescein diacetate (DCFHDA) staining to measure the ROS levels of the different groups’ HLECs (representative staining image in (g), quantified fluorescence comparison in (h). Blue (Hoechst) and green (DCFHDA) represented stained nuclei and ROS, respectively (scale bar 50 μm). i, j The comparison of lens opacity for the cultured rat lenses in the different groups. The representative lens image and its grayscale quantification data were in i (scale bar 1 mm). The quantification comparison was in (j). k, l The comparison of superoxide anion levels for the different groups’ cultured rat lenses (representative staining image in (k), quantified fluorescence comparison in (l). Blue (4’,6-diamidino-2-phenylindole, DAPI) and red (dihydroethidium, DHE) represented stained nuclei and superoxide anion, respectively (scale bar 20 μm). The data in Fig. d to h represented three biological replicates, and those in Fig. i to l represented six biological replicates. P values in the ANOVA with two-sided Tukey post-hoc tests were shown. The bar graphs in Fig. d to l were expressed as means ± standard errors. Source data are provided as a Source data file, including the three replicated uncropped Western blot scans for Fig. d.
To validate the DC biomarkers discovered here by the MS-based strategy, we performed a nuclear magnetic resonance (NMR)-based metabolic annotation validation for the tear and AH biomarkers using a mixed sample analysis framework. The AH biomarker of 1,5-AG, as well as the tear biomarkers of G3P and DHOMEs, were identified in the NMR spectra, which showed the same change trend between groups as the results of the AHMFs and TMFs (Supplementary Table 17). Nevertheless, we did not find the specific peak cluster of the tear biomarker of NeuAc, maybe due to the limited sensitivity of NMR. In summary, the NMR-based metabolic characterization validated the DC biomarkers discovered by MS, in both AH and tear fluid samples.
Notably, 1,5-AG is a blood biomarker of glycemic status approved by the FDA and was herein identified as an AH biomarker of DC34. As a metabolite in AHs, it could act directly on the lens by the AH-lens interface, which aroused our great interest in further investigating its role in DC development. For this purpose, human lens epithelial cells (HLECs) were employed to evaluate the effect of 1,5-AG on lens oxidative stress, which was one of the main causes of cataracts induced by high glucose. For safety assessment, the cell viability analysis demonstrated excellent safety of 1,5-AG when treated with HLECs at different concentrations ranging from 0.05 to 500 μM (cell viability > 85% at all concentrations, Supplementary Fig. 16). Thus, concentrations of 50 and 500 μM were employed in the activity evaluation. For activity evaluation, the detection of antioxidant enzymes, including superoxide dismutase 1 (SOD1), glutathione peroxidase 1 (GPx1), and catalase (CAT), was performed in HLECs cultured under normal glucose (Control group), high glucose (200 mM of glucose, HG group), high glucose with low 1,5-AG (200 mM of glucose with 50 μM of 1,5-AG, HG + LAG group), and high glucose with high 1,5-AG (200 mM of glucose with 500 μM of 1,5-AG, HG + HAG group). In the Western blot (Fig. 5d) and quantitative real-time PCR (qPCR) analysis (Fig. 5e, Supplementary Table 18), compared to the Control group, the levels of the three antioxidant enzymes were significantly downregulated in the HG group. Interestingly, the high 1,5-AG treatment restored the levels of all three antioxidant enzymes, indicating the protective effect of 1,5-AG against oxidative stress induced by high glucose in HLECs. In the detection of glutathione (GSH), the GSH level decreased by high glucose was also reversed after the high 1,5-AG treatment (Fig. 5f), consistent with the Western blot and qPCR results. Further, staining of HLECs with 2,7-dichlorofluorescein diacetate (DCFHDA) for direct measurement of reactive oxygen species (ROS) levels showed that the high glucose-induced increase in ROS was significantly inhibited by high 1,5-AG treatment (Fig. 5g, h, blue and green respectively represented stained nuclei and ROS). These results indicated that 1,5-AG could attenuate high glucose-induced oxidative stress in HLECs via upregulating antioxidant enzymes.
Encouraged by the above findings based on HLECs, we cultured rat lenses to obtain direct evidence for the protective effect of 1,5-AG against cataracts in the high glucose environment. As shown in Fig. 5i, j, the high glucose environment noticeably increased the opacity area of the lens, which was a manifestation of cataracts. Excitingly, treatment with high concentrations of 1,5-AG markedly inhibited the increase in opacity area, demonstrating the protective effect of 1,5-AG against high glucose-induced cataracts in actual lens tissues. In histopathological analysis, the hematoxylin and eosin staining for the lens tissues exhibited lens fibers with edema and irregular arrangement in the HG group, while the lens in HG + HAG presented a normal close and regular arrangement like the control group (Supplementary Fig. 17). Moreover, superoxide anion in lens epithelium was measured by dihydroethidium (DHE) staining to illustrate the relationship between oxidative stress and cataract changes. Compared to the Control group, the superoxide production was increased in the lens epithelium (indicated by white lines) with high glucose, while was attenuated by the high 1,5-AG treatment (Fig. 5k, l, blue and red respectively represented stained nuclei and superoxide anion). These results based on animal lenses indicated that 1,5-AG may prevent high glucose-induced cataracts via attenuating oxidative stress.
Given the potential key role of 1,5-AG in AHs in DC development, we performed targeted detection for 1,5-AG through multiple reaction monitoring (MRM) analysis, which allowed absolute quantification of its concentrations in AHs and stronger validation regarding metabolite identification. With the ion pair determined by the 1,5-AG standard, the presence of 1,5-AG in the AH samples was further confirmed, and 1,5-AG’s concentrations in the 12 AH samples (ARC/DC of 6/6, randomly selected from all 46 AH samples) were quantified by the standard curve method. 1,5-AG exhibited a significant down-regulation (p value < 0.05, mean concentration of 3.28 μg ml−1 for ARCs and 1.28 μg ml−1 for DCs, Supplementary Fig. 18) in the DC group compared to the ARC group, consistent with the results in the R2DIFMS-TS process. Notably, concentrations of 1,5-AG and intensities of AHMF feature of m/z 225.0 showed a significantly strong correlation (p value < 0.001, Pearson correlation coefficient of 0.867, Supplementary Table 19), which demonstrated that the R2DIFMS-TS reliably identified the differential metabolite that led to the intensity change of the AHMF feature of m/z 225.0. These results offered further validation of 1,5-AG as an AH biomarker regarding DC, and highlighted the value of the strategy of R2DIFMS-TS in rapid biomarker discovery.
Discussion
As the rapid increase in the number of diabetic people worldwide, the treatment for DC has become an important concern in ophthalmology clinics. Though the technology of cataract extraction surgery has advanced, the incidence of intraoperative and postoperative complications in DC is about 30% higher when compared with ARCs35,36. And, the surgery may accelerate the progressions of diabetic retinopathy and macular edema37,38. Those extra risks highlight the pressing need for precise discrimination of DC from common ARC patients before surgery to prepare individualized therapeutic regimens. A reliable diagnosis of DC relies on comprehensive evaluations, including clinical symptoms, patient history, multiple measurements for blood glucose level, glycosylated hemoglobin level, and glucose tolerance. These laborious evaluations usually require hours to days and are sometimes limited by the availability in ophthalmology clinics, making them difficult to apply to all cataract patients6. In this case, simple preoperative blood tests or self-reported history remain the mainstay in the clinical diagnosis of DC, which may lead to unacceptable underdiagnosis5,6,7. Moreover, blood tests, including common blood glucose or glycosylated hemoglobin tests, are invasive and would cause injury to patients’ skin39.
Recently, advancements in molecular diagnosis have offered a promising tool for diagnosing DC. A recent work reported an encouragingly strong performance (AUC of 0.978) for distinguishing DC from ARC through LDI-MS analysis for AH metabolic biomarkers15. Nevertheless, AHs are collected invasively from within the eye and therefore not applicable for the preoperative diagnosis of DC, which encouraged us to develop a novel non-invasive molecular diagnosis method of DC by identifying metabolic biomarkers in tear fluids, uniquely non-invasive ocular body fluids.
Rapid and in-depth metabolic analysis for non-stimulated tear film, featuring the limited volume of single-digital microliters, is a significant challenge for conventional metabolic analysis tools like LC-MS and GC-MS, due to their well-known limitations on sample volume requirements and analysis throughput. In this respect, direct ionization MS platforms promise rapid metabolic profiling with less sample consumption, like the NELDI-MS employed here, direct infusion MS, MALDI-MS, as well as MS coupled with in situ sample extraction techniques represented by DESI-MS27,28,29,40,41,42. Compared to the NELDI-MS, direct infusion MS relying on ESI typically requires complex metabolome extraction and longer detection time (typically more than 2 min for that while less than 30 s for the NELDI-MS), especially for protein-rich or salt-rich biofluids40,41. In terms of LDI-MS, the NELDI-MS exhibited one to three orders of magnitude higher signal response for the typical metabolites compared to the conventional MALDI-MS, which is crucial for in-depth metabolic analysis. As for MS coupled with in situ sample extraction techniques, we performed a performance comparison between the NELDI-MS and DESI-MS, a commercially available direct ionization platform for metabolic analysis27,28,29. The NELDI-MS exhibited advantages in throughput (detection time of less than 30 s, while that of more than 2 min for the DESI-MS), sensitivity (one to three orders of magnitude lower LODs than the DESI-MS), and reproducibility (CVs of 2.4%–4.9%, while those of 17.0%–33.2% for the DESI-MS). These comparisons highlighted the NELDI-MS’s strengths and promising application in metabolic analysis.
Here, using trace tear fluids non-invasively collected from ARC and DC patients, we constructed a diagnostic biomarker panel of four metabolites, based on machine learning for the TMFs recorded by the high-performance NELDI-MS (tear fluid consumption of 10 nL). Excitingly, the diagnostic panel achieved a precise diagnosis of DC with an AUC of 0.923, sensitivity of 85.9%, and specificity of 82.0%. Importantly, the sample pretreatment and metabolic detection for each tear fluid sample could be finished within 1 min and 30 s respectively, supporting the rapid diagnosis of DC. Further, through validating the biomarker panel by including an extra external cohort, we demonstrated the panel-based model’s reproducibility regarding diagnostic performance. Compared to existing molecular and clinical diagnosis methods, the DC diagnosis based on the tear metabolic biomarkers covered the advantages of rapidness, precision, and non-invasive sampling, and thus is expected to apply to cataract patients in ophthalmology clinics.
Reliable metabolite annotation has been a crucial bottleneck of NELDI-MS for its application in biomarker discovery. Metabolite annotations for NELDI-MS features are generally transferred from FT-ICR-MS or LC-MS/MS through matching features arising from the same metabolite23,24,25,26. In this regard, feature matching based on 1-D information of m/z is the most mainstream strategy23,24,26. However, it may yield unreliable metabolite annotations with several or even a dozen incorrect matching results due to the complexity of biofluids and ion adduction variation. In the process of metabolite annotation for the TMF feature of m/z 337.2, we provided an illustration regarding the ambiguous or incorrect annotation risk caused by the 1-D matching of m/z. Specifically, five LC-MS/MS XIC features showed the matched m/z and might be identified as the signal of the biomarker in a 1-D matching process, while three of those were potential incorrect results. On the other hand, though 2-D information matching relying on long-time LC-MS/MS analysis for each individual sample within the cohort can yield reliable metabolite annotation25, it not only loses the advantage of analysis throughput but also fails to be applicable for trace samples. Therefore, reliable metabolite annotation is a critical limitation of NELDI-MS on biomarker discovery in trace body fluids.
In our study, we proposed the R2DIFMS-TS, which reconstructs the metabolic annotation framework by integrating NELDI-MS-derived metabolic fingerprints with LC-MS/MS data from mixed samples, utilizing m/z and fold-change matching for accurate metabolite identification. Compared to the 1-D information matching strategy, the R2DIFMS-TS effectively eliminated the potential incorrect results, like the three XIC features in Fig. 4g. Compared to long-time LC-MS/MS analysis for individual samples within the cohort, the R2DIFMS-TS allowed that the available sample volume could multiply for trace samples and the detection time did not to be increased in LC-MS/MS analysis, as the sample number increases. Using the R2DIFMS-TS, we annotated the target NELDI-MS features within the diagnostic panel as metabolites by 140 nL of tear fluid per sample, including 9,10-DHOME, 12,13-DHOME, G3P, and NeuAc, and the NMR analysis validated the MS-based metabolic characterization. Up to this point, we successfully achieved metabolic biomarker discovery for DC with a total tear fluid consumption of 150 nL per sample, through TMF acquisition by NELDI-MS and metabolite annotation by LC-MS/MS. Of note, we next successfully applied R2DIFMS-TS for biomarker discovery in AHs and identified 1,5-AG as a new anti-cataract active DC biomarker, which was further validated by NMR analysis and targeted MRM analysis, highlighting the reliability and excellent application prospect of our solution. Our strategy integrating NELDI-MS and LC-MS/MS provides a solution for rapidly discovering metabolic biomarkers in large-size trace body fluid samples with nanoliter-scale sample consumption and second-scale detection time, which has the potential to be generally used in biomarker research based on such samples. It promises a desirable tool for developing non-invasive molecular diagnosis technology and revealing disease molecular mechanisms.
In addition to their diagnostic applications, metabolic biomarkers also help to understand the pathogenesis, laying the basis for improving DC prevention. The major mechanism of DC was related to the dysregulated sorbitol pathway of glucose metabolism, non-enzymatic glycation, and oxidative stress due to sustained high blood In this regard, DC metabolic biomarkers within the eye have been reported, which are mainly related to energy metabolism, glycosylation, and oxidative stress15,16. On this basis, here, the signature metabolic patterns of DC on the ocular surface and within the eye were comprehensively revealed. Within the eye, 1,5-AG was identified as a DC biomarker in AHs, related to glucose metabolism and oxidative stress43,44. For the ocular surface, metabolic biomarkers of DC in non-stimulated tear film were first reported in the present study. Specifically, G3P is a key metabolite in the glycolysis metabolic pathway45, whereas the DHOMEs are the strong pro-inflammatory lipids as the oxidation products of linoleate46,47. It is well-known that oxidative stress and inflammation are mutually reinforcing48. And, NeuAc is related to either glucose metabolism, inflammation, and oxidative stress49,50. Interestingly, the comparison of the healthy control and diabetic control groups suggested that those tear biomarkers may not be simply caused by the single factor of the presence of diabetes, and thus, they may be directly linked to DC formation rather than diabetes itself. These findings are critical for future exploration of the pathogenic factors of DC and their metabolic influence, including genotypes, living habits, glucose control, and so on. Of note, although the different biomarkers in AHs and tear fluids suggested unique metabolic disorders on the ocular surface and within the eye, they showed consistency in the metabolic function abnormalities involved. Moreover, Spearman correlation analysis also showed close relationships between biomarkers in AHs and tear fluids. In summary, our findings suggested there were unique but closely related metabolic disorders between the ocular surface and within the eye of DC patients, which indicated dysregulated energy metabolism, aberrant inflammation, and abnormal oxidative stress. These metabolic disorders might be involved in the pathogenesis of DC.
Excitingly, 1,5-AG, a DC biomarker in AHs identified here, exhibited a protective effect against high glucose-induced lens oxidant stress and opacification. In HLECs, 1,5-AG significantly attenuated high glucose-induced oxidative stress, a major factor in cataract formation, via upregulating antioxidant enzymes. Further, in rat lenses, 1,5-AG attenuated glucose-induced lens opacity and the swelling of lens cortical fibers. These findings suggested that the low level of 1,5-AG in AHs of DC patients may exacerbate oxidative stress by high glucose, contributing to cataract development. On the other hand, 1,5-AG was also significantly related to the pro-inflammatory lipids, DHOMEs, in tear fluids, indicating it might be involved in ocular inflammation. On this basis, we targeted quantified the concentrations of 1,5-AG in AHs by MRM analysis, which are 3.28 μg ml−1 for ARCs and 1.28 μg ml−1 for DCs (mean concentrations). Given that 1,5-AG is an FDA-approved blood biomarker reflecting short-time glycemia and its close relationships to diabetic complications34,51, it is possible that a decrease of 1,5-AG might be common in diabetic patients’ AHs and plays an important role in the development of other diabetic eye diseases.
This study has limitations regarding cohort design. (1) Inclusion/exclusion criteria and clinical characterization. To represent the inherent characteristics of diabetic cataract in the real clinical population, some chronic comorbidities were not excluded in this study. And the information on non-diabetic medications (e.g., antihypertensives, statins) for those comorbidities was limited. Although no significant inter-group differences were observed, these comorbidities may still have the potential to influence metabolic profiles. Additionally, only DC patients with a history of diabetes for more than 10 years were included, which may introduce some degree of heterogeneity. Consequently, future studies are still needed to clarify the influence of comorbidities, non-diabetic medications, and diabetic duration on tear fluid composition and DC-related metabolic reprogramming. (2) Comparison group choosing. In this study, ARC patients were chosen as the comparison group to identify DC-specific metabolic biomarkers, to support the diagnosis and molecular mechanism exploration of DC. However, Nevertheless, setting broader comparison groups, including healthy individuals and diabetic patients without cataracts, would help to obtain a baseline of normal tear metabolites and further investigate if those biomarker changes are directly linked to DC formation rather than diabetes itself. In this regard, though we have included a small-sized cohort (39 individuals) here to provide some initial insights, future studies incorporating these groups with larger sample sizes are significant to further elucidate the unique metabolic signatures of DC.
In conclusion, we proposed a rapid, precise, and noninvasive diagnosis of DC through metabolic biomarker detection in trace tear fluids by the high-performance NELDI-MS. In this process, by integrating NELDI-MS and LC-MS/MS, we achieved reliably annotating m/z features as metabolites within nanoliter-scale sample consuming in NELDI-MS-based biomarker discovery, providing a promising solution for the rapid discovery of metabolic biomarkers in large-size trace body fluid samples. Further, based on DC biomarkers in tear fluids and AHs, we revealed unique but closely related metabolic disorders on the ocular surface and within the eye of DC patients, in which the low level of 1,5-AG in AHs might contribute to DC development. Our work would contribute to the noninvasive diagnosis and pathologies exploration for DC, as well as other studies on metabolic biomarker discovery based on trace body fluids.
Methods
Chemicals and reagents
The chemicals and reagents in the present study included: (1) reagents for the preparation of the ferric nanoparticles (NPs): (2) standard metabolites and conventional organic matrices for the performance evaluation of the nanoparticle-enhanced laser desorption/ionization mass spectrometry (NELDI-MS) platform; (3) reagents for the biological evaluations; (4) other reagents and chemicals. For the preparation of the ferric NPs, ferric chloride hexahydrate (97%), trisodium citrate dihydrate (99%), sodium acetate (99%), ethylene glycol (99.5%), and absolute ethanol (99.7%) were purchased from Sinopharm Chemical Reagent Beijing Co., Ltd. (Beijing, China). For the standard metabolites and conventional organic matrices, D-glucose (99.5%), L-arginine (99.5%), L-lysine (98%), L-glutamate (99%), and D-mannitol (99%) were purchased from Sigma-Aldrich (St. Louis, MO, USA). α-cyano-4-hydroxy-cinnamic acid (CHCA, 99.0%) and 2,5-dihydroxybenzoic acid (DHB, 99.0%) were also obtained from Sigma-Aldrich (St. Louis, MO, USA). For reagents for the biological evaluations, D-glucose (99.5%) was purchased from Sigma-Aldrich (St. Louis, MO, USA), and 1,5-anhydroglucitol (1,5-AG, 97%) was purchased from Adamas (Shanghai, China). For other reagents and chemicals, acetonitrile (HPLC), water (HPLC), and methanol (HPLC) were obtained from J. T. Baker (Phillipsburg, NJ, USA). Formic acid (99.5%) was ordered from Fisher Scientific (Waltham, MA, USA). Trifluoroacetic acid (99.5%) was purchased from Macklin Biochemical Co., Ltd. (Shanghai, China). Ammonium acetate (99.9%) was purchased from Sigma-Aldrich (St. Louis, MO, USA). Except for the liquid chromatography (LC) coupled MS experiment, all other aqueous solutions were prepared using ultrapure water (18.2 MΩ cm, Milli-Q, Millipore, GmbH) throughout the experiments.
Preparation and characterization of the NPs
The ferric chloride hexahydrate was first dissolved in the ethylene glycol solution, after which the trisodium citrate dihydrate and sodium acetate were added. Then, after the sonication at 25 °C for 45 min, the mixture was transferred to the autoclave for hydrothermal reaction at 200 °C for 10 h. Lastly, the particles were washed and dried for further use.
In the characterization, scanning electron microscopy images were recorded by an S4800 field emission scanning electron microscope (Hitachi, Japan), and X-ray diffraction patterns were recorded using a D8 Advanced X-Ray Diffractometer (Bruker, Germany). The high-angle annular dark field and elemental mapping images were recorded by a JEOL JEM-2100F instrument.
Clinical participants
This study was approved by the Medical Science Research Ethics Committee of Shanghai Ninth People’s Hospital (SH9H-2023-T64-2). All participants in this study provided written consent for the use of samples.
In this study, tear fluid samples were collected from the 168 participants (main cohort, which was split into the discovery and validation cohorts for machine learning) at Shanghai Ninth People’s Hospital. And, aqueous humor (AH) samples were collected from the 46 participants who underwent cataract surgery out of the 168 participants. All these participants were diagnosed as cataract patients by two experienced ophthalmologists. Of these, participants with type II diabetes were taken into the diabetic cataract (DC) group, whereas the other participants without diabetes were taken into the alone age-related cataract (ARC) group. Before being included in the cohort, all participants in the DC group had a history of diabetes for more than 10 years, and were examined by two experienced physicians. The fasting blood-glucose level was 5.7 ± 0.9 mmol L−1 for the ARC patients and 8.4 ± 3.7 mmol L−1 for the DC patients (mean ± SD). The glycosylated hemoglobin level was 5.6% ± 0.7% for the ARC patients and 8.3% ± 3.1% for the DC patients (mean ± SD). Of note, patients with one or several ocular characteristics of the following were excluded: (1) eye axis greater than 26 mm; (2) other ocular history, including but not limited to diabetic retinopathy, glaucoma, ocular trauma, prior eye surgeries, as well as recent ocular infections; (3) recent eye drops treatments. Recent diseases or treatments were defined as those within the past 12 weeks in this section. Besides the above exclusion criteria regarding ocular, the exclusion criteria also included common autoimmune diseases (including type I diabetes, systemic lupus erythematosus, autoimmune arthritis, chronic autoimmune skin diseases, and Sjögren’s syndrome), recent localized or systemic infections, and recent acute inflammatory diseases. Information regarding ages, genders, demographic characteristics, and other major comorbidities not in the exclusion criteria of the 168 participants was summarized in Supplementary Tables 2 and 3 (comorbidity information included kidney, liver, thyroid, and cardiovascular diseases, as well as cancers, whereas demographic characteristics included residency, educational status, occupational status, and body mass index). Diabetic characteristics for the DC patients (n = 86) were summarized in Supplementary Table 4 (including diabetic treatments, duration, complications, etc.).
Besides, we recruited an independent single-center temporal external cohort (extra external cohort), including 106 participants at Shanghai Ninth People’s Hospital. The diagnostic criteria of this cohort were consistent with those in the main cohort of the 168 participants. The fasting blood-glucose level was 5.3 ± 0.6 mmol L−1 for the ARC patients and 7.9 ± 2.2 mmol L−1 for the DC patients (mean ± SD). The glycosylated hemoglobin level was 5.5% ± 0.4% for the ARC patients and 8.2% ± 2.2% for the DC patients (mean ± SD). To avoid metabolic disturbance by some related comorbidities, this cohort additionally excluded patients with chronic kidney diseases, chronic liver diseases, thyroid diseases, and cancers, besides the main cohort’s exclusion criteria. Nevertheless, some chronic basic diseases were not included in this cohort’s exclusion criteria, including hypertension, hypercholesterolemia, and coronary heart disease. The 106 participants’ information regarding ages, genders, demographic characteristics, and other major comorbidities was summarized in Supplementary Tables 7 and 8, and diabetic characteristics for the DC patients (n = 27) were summarized in Supplementary Table 9.
We also included a cohort of individuals without cataracts at Shanghai Ninth People’s Hospital, including 20 healthy individuals (healthy control group) and 19 diabetic patients without cataracts (diabetic control group). Two experienced ophthalmologists examined their eyes to exclude cataracts. Before being included in the cohort, all participants in the diabetic control group had a history of diabetes for more than 10 years, and were examined by two experienced diabetologists. The fasting blood-glucose level was 5.7 ± 1.0 mmol L−1 for the healthy control group and 6.8 ± 2.3 mmol L−1 for the diabetic control group (mean ± SD). The glycosylated hemoglobin level was 5.5% ± 0.7% for the healthy control group and 6.1% ± 0.7% for the diabetic control group (mean ± SD). The exclusion criteria, except cataracts, were consistent with those in the extra external cohort of the 106 participants. The 39 participants’ information regarding ages, genders, demographic characteristics, and other major comorbidities was summarized in Supplementary Tables 10 and 11, and diabetic characteristics for the diabetic patients (n = 19) were summarized in Supplementary Table 12.
Tear fluid and AH sample harvesting
For tear fluid sample harvesting, 200 nL of tear fluid was collected from the single eyes of participants using a microcapillary tube. After the collection, each sample was immediately diluted 100-fold by ultrapure water and stored at −80 °C. For AH sample harvesting, about 100 μL of AHs were collected at the onset of cataract surgery and stored at −80 °C. All freezing samples were thawed at 4 °C before use.
The detailed method information on tear fluid harvesting was below. Microcapillary tubes (30 to 32 mm of length, total volume of 3.0 μL, Hirschmann, German) were employed to harvest non-stimulated tear fluid (more than 400 nL, less than 1 μL) from the lateral canthus via capillary action. The collected volume was estimated by the length of capillary filling. Tear fluid collected in the capillary tube was expelled into a centrifuge tube using a rubber bulb, and a calibrated pipette (0.1–2.5 μL range, Eppendorf, German) was used to transfer 200 nL of tear fluid, which was then diluted in 19.8 μL of ultrapure water (100-fold dilution) and stored at –80 °C for subsequent analysis. All tear fluid samples were exclusively collected from cataract-affected eyes. For patients with bilateral cataracts, only one eye was included in the study. All tear fluid samples were collected between 9:00 and 12:00 AM to mitigate diurnal variability. Compared to well-established tear fluid collection methods (e.g., Schirmer strips, capillary tubes, or sponges), our approach has the following advantages: (1) Little patient stimulation. (2) Convenient clinical sampling. (3) Precise collection of trace non-stimulated tear fluids. Nevertheless, it has a limitation regarding cumbersome demanding post-processing steps (e.g., precise pipetting of nanoliter volumes).
LDI-MS analysis for performance evaluation and body fluid metabolic detection
The major LDI-MS experiment, including the detection for the tear fluid samples (except the samples in the extra external cohort), AH samples, and standard metabolites, was conducted in the same lab using an Autoflex time-of-flight mass spectrometry (Bruker, Germany), where a Nd:YAG solid-state SmartBeam (355 nm) was employed. The LDI-MS experiment for the tear fluid samples in the extra external cohort was conducted in another lab using an ultrafleXtreme (Bruker, Germany) with a Nd:YAG solid-state SmartBeam-II (355 nm).
For matrix preparation, the ferric NPs were dispersed in ultrapure water at 1 mg mL−1 for further use in the NELDI-MS, whereas CHCA and DHB were dissolved in 0.1% TFA solution (acetonitrile/water, 3/7, v/v) at saturation concentration and 10 mg/mL, respectively. In standard metabolite detection, each standard metabolite was prepared as an aqueous solution of different concentrations (1 mg mL−1 for reproducibility evaluation and NELDI-MS/MALDI-MS comparison, gradual concentration gradients for detection limit evaluation). In preparation for the body fluid samples, the tear fluids and AHs were respectively prepared as 100-fold dilution and 5-fold dilution with ultrapure water before LDI-MS detection, to decrease possible salt crystallization.
For LDI-MS analysis, in a typical process, 1 μL of body fluid dilution sample or standard metabolite solution was dropped and deposited on the 384-well target plate (Bruker, Germany), and then the matrix was dropped to cover the dried spot. After the target plate was dried at 25 °C, the MS spectra of the samples were acquired under the positive mode. To monitor the data acquisition and ensure data reliability, each sample was independently detected in five replicates (those replicates could be totally finished within 30 s), and standard spots containing metabolites of fixed concentrations were tested at regular intervals. The pulse laser was used with a frequency of 1000 HZ, and the number of shots was 2000 per acquisition.
DESI-MS analysis for performance evaluation
Desorption electrospray ionization MS (DESI-MS) analysis was conducted using a DESI XS (Waters, USA) source equipped with a Q-TOF mass analyzer (SELECT SERIES Cyclic IMS, Waters, USA). For sample preparation, each standard metabolite was prepared as an aqueous solution of different concentrations (1 mg mL−1 for reproducibility evaluation, gradual concentration gradients for detection limit evaluation). For DESI-MS analysis, similar to the LDI-MS analysis, in a typical process, 1 μL of standard metabolite solution was dropped and deposited on the glass slide to form a dried sample spot at 25 °C. Then, the sample spot was scanned by the DESI source to acquire MS data under the positive mode.
For parameters of the DESI-MS, the step size was 100 μm for the X-axis and 250 μm for the Y-axis, and the rate was 500 μm s−1. The DESI spray solvent was methanol/water (50/50, v/v) with a flow rate of 2 μL min−1. The mass range was set at 50–1000 Da, the capillary was set at 0.60 kV, the cone was set at 40 V, the source temperature was set at 150 °C, and the heated transfer line was set at 250 °C.
Data processing, machine learning, and feature panel construction for metabolic fingerprints
The processing for raw MS data was performed using a pyOpenMS framework-based Python script (version 3.7), including average spectrum calculation, baseline correction, peak detection, alignment, peak filtration, and median normalization. After the processing, m/z features were extracted from the raw MS data, constituting the tear fluid metabolic fingerprint (TMF) and AH metabolic fingerprint (AHMF).
For further machine learning and diagnostic feature panel construction, the 168 tear fluid samples were randomly split into the discovery and validation cohorts with a ratio of 3:1. In the following study, only data from the discovery cohort would be used for modeling algorithm tuning, model training and evaluation, as well as feature selection, while the data from the validation cohort was only used for validating the model performance.
The machine learning of metabolic fingerprints was performed with the scikit-learn scientific computing framework. The m/z feature data was scaled to 0–1 for machine learning. The candidate modeling algorithms included Logistic Regression (LR), K-Nearest Neighbor (KNN), Adaptive Boosting (AB), and Decision Tree (DT). The modeling algorithm was determined and tuned according to the averaged cross-validated area under the curves (AUCs, 5 folds, 20 rounds) in the discovery cohort. With the best-tuned algorithm, the models for TMFs were trained by data in the discovery cohort, which were validated in the validation cohort.
For feature panel construction, the p values in the two-tailed t-tests and the coefficients in the TMF-based diagnostic model were used to select the key features. Specifically, the differential features (FWER p value < 0.05, two-tailed t-test, Bonferroni correction) were ranked by the coefficient in the LR model to obtain the 10 key features. Next, the cross-validated AUCs (5 folds, 10 rounds) of all combinations of those 10 features were employed to select the target features making up the diagnostic panel. The panel-based model was also trained in the discovery cohort and validated in the validation cohort. Besides, the panel-based model was further evaluated using the data of the extra external cohort, which was acquired by another instrument in another laboratory.
LC-MS/MS analysis for the mixed samples
In the LC-MS/MS sample preparation, in order to precipitate salts and proteins, methanol/acetonitrile (50/50, v/v) solution was added to each mixed tear fluid sample and mixed AH sample to reach an organic solvent proportion of 75%. After being placed at −20 °C for 2 hours, the mixture was centrifuged for 20 min at 16,200 × g to collect the supernatant. Finally, the supernatant was dried by centrifugation and then redissolved in methanol/water (30/70, v/v) according to the specific dilution factor of body fluids (1:6 for tear fluids, 1:2 for AHs).
The LC-MS/MS metabolic analysis for the mixed tear fluid samples and AH samples was performed using an LC-20A (Shimadzu, Japan) coupled ZenoTOF 7600 (SCIEX, USA) system. RP-C18 and HILIC separation modes were performed in both positive and negative electrospray ionization modes. For the HILIC separation, the Atlantis BEH Z-HILIC column (particle size, 1.7 μm; 100 mm (length) × 2.1 mm (i.d.)) was used. The mobile phases A and B were 10 mM ammonium acetate in water and pure acetonitrile, respectively. The binary gradient was 0–18 min: 95%–40% B; 18–22 min: 40%–40% B; 22–23 min: 40%–95% B; 23–35 min: 95%–95%. For the RP-C18 separation, the ACQUITY BEH C18 (particle size, 1.7 μm; 50 mm (length) × 2.1 mm (i.d.)) was used with 0.1% formic acid in water for mobile phase A and 0.1% formic acid in acetonitrile for mobile phase B. The binary gradient was 0–6 min: 5%–30% B; 6–12 min: 30%–50% B; 12–25 min: 50%–98% B; 25–30 min: 98%–98% B; 30–30.1 min: 98%–5% B; 30.1–35 min: 5%–5% B. In both the HILIC and RP-C18 separations, the flow rate was 0.2 mL min−1, the oven temperature was 40 °C, and the injection volume was 3 μL.
The information-dependent acquisition mode was applied in the LC-MS/MS analysis. An acquisition cycle consisted of one MS experiment and a maximum of fifteen MS/MS experiments. For the MS experiment, the mass range was set at 50–1000 Da and the accumulation time was set at 0.3 s. For the MS/MS experiment, the candidate ions were selected by intensity, the mass ranges were set at 50–500 Da, the accumulation time was set at 0.05 s, the fragmentation mode was CID, and the Zeno pulsing was on. The collision energy was 35 V for positive mode and −35 V for negative mode, as well as the collision energy spread was set at 15 V.
Targeted MRM analysis for the AH samples
For the preparation of each AH sample, 60 μL of methanol/acetonitrile (50/50, v/v) solution was added to 20 μL of AH. After being placed at −20 °C for 2 h, the mixture was centrifuged for 20 min at 16,200 × g to collect the supernatant, and the supernatant was directly used for multiple reaction monitoring (MRM) analysis. For the preparation of the standard series of 1,5-AG, 10 mg of 1,5-AG standard was accurately weighed to prepare the standard solution of 1 mg mL−1 (ultrapure water), and it was then gradually diluted to the standard series at concentrations of 10, 2.5, 1, and 0.1 μg mL−1 (75% organic solvent proportion, methanol/acetonitrile of 50/50, v/v).
A Xevo TQ-XS triple quadrupole mass spectrometer (Waters, USA) was employed to carry out the MRM analysis to achieve absolute quantification of 1,5-AG in the AH samples. A HSS T3 column (particle size, 1.8 μm; 100 mm (length) × 2.1 mm (i.d.)) was used at an oven temperature of 40 °C. The gradient system was 0–2 min: 1%–15% B; 2–3 min: 15%–35% B; 3–4 min: 35%–99% B; 4–6.5 min: 99%–99% B; 6.5–6.7 min: 99%–1% B; 6.7–10 min: 1%–1% B with a flow rate of 0.2 mL min−1 (mobile phase A: 0.1% formic acid in water, mobile phase B: pure acetonitrile). The injection volume was 2 μL. For parameters of the MRM analysis, the analysis was performed with negative electrospray ionization mode, the quantitative ion pair was 163.0− > 113.0, the capillary was set at 3.0 kV, the cone was set at 30 V, the collision was set at 12 V, the desolvation gas flow was set at 1000 L h−1, the cone gas flow was set at 150 L h−1, and the collision gas flow was set at 0.12 mL min−1.
NMR analysis for the mixed samples
In terms of the nuclear magnetic resonance (NMR) sample preparation, 20 ARC tear fluid samples and 20 DC tear fluid samples in the validation cohort (from the main cohort) were used to prepare the corresponding mixed tear fluid samples (140 nL per sample mixed by group), whereas 5 ARC AH samples and 5 DC AH samples were used to prepare the corresponding mixed AH samples (20 μL per sample mixed by group). The mixed tear fluid samples were dried by centrifugation, redissolved in 100 μL of heavy water (D2O, 99.9%, the same below), and then diluted with 300 μL of PBS D2O buffer (45 mM, pH = 7.4, the same below). The mixed AH samples were directly diluted with 300 μL of PBS D2O buffer. The diluted samples were directly used for NMR analysis after centrifugations (4 °C, 11,180 × g, 10 min). For the NMR analysis method, NMR spectra were acquired at 298 K and operated at 600.13 MHz for proton, using an Avance III spectrometer equipped with a 5 mm 1H/13C/15N/D CP TCI probe (Bruker, Germany). For each sample, a standard 1D 1H NMR spectrum was acquired by Bruker TopSpin (version 3.2), using a “noesypr1d” (Bruker library) pulse sequence with water suppression during the relaxation delay of 2.3 s. The spectral width was 12,019 Hz. The number of scans was 1024.
For NMR data analysis, an R script (version 4.4.1) based on the PepsNMR package was used. The regions of water (ppm of 4.5–5.1) were removed before processing. The signal of the alpha anomer of glucose at 5.233 ppm was used for the spectra calibration. The data was divided into discrete bins of 0.002 ppm and normalized by their median intensities. The NMR signals were assigned to the metabolites using the Human Metabolome Database (https://hmdb.ca/), and the integrals of the corresponding signal regions were used to represent the metabolite’s abundance.
Cell culture and treatment
Human lens epithelial cells (HLECs, male human origin) were purchased from BLUEFBIO (China) and were cultured in RPMI1640 cell medium (Gibco, USA) containing 10% fetal bovine serum (FBS, Gibco, USA), and 1% penicillin/streptomycin (Gibco, USA). Cells were cultured at 37 °C under 5% CO2 in a humidified atmosphere. After serum starvation for 24 h, HLECs were divided into four groups and treated accordingly for 48 h, including the normal glucose group (Control), high glucose group (HG, 200 mM of glucose), high glucose with low 1,5-AG group (HG + LAG, 50 μM of 1,5-AG), and high glucose with high 1,5-AG group (HG + HAG, 500 μM of 1,5-AG). The culture medium with different treatments was refreshed every day.
Cell viability detection
HLECs were seeded in 96-well plates with an equal volume of cell suspension and then incubated with Cell Counting Kit-8 (CCK-8, Dojindo, Japan) for 4 h. After 4 h of incubation, the optical density was measured using a microplate reader at a wavelength of 450 nm. Cell viability was expressed as a percentage normalized to the value of the control group.
Western blot analysis
Total proteins from the treated cells were harvested in RIPA lysis buffer (Thermo Scientific, USA) with a 1% protease and phosphatase inhibitor cocktail and then sonicated. Protein concentration was determined using a BCA protein assay kit (Thermo Scientific, USA). Equal amounts of lysate (50 μg) were separated on SDS–PAGE gels (Bio-Rad, USA), transferred to PVDF membranes (Millipore, USA) and then blocked in 5% BSA for 1 h. The membranes were then incubated with primary antibodies (SOD1 [1:2000, Abcam, UK], GPx1 [1:1000, Abcam, UK], CAT [1:2000, Proteintech, China] at 4 ◦C overnight and then probed with horseradish peroxidase (HRP)-conjugated secondary antibodies for 1 h at room temperature. Protein signaling was visualized using image scanning systems (Tanon, China, and e-Blot, China).
qPCR analysis
Total RNA from HLECs was prepared using TRIzol reagent. Reverse transcription of total RNA from HLECs was performed using PrimeScript RT master mix (Takara, Japan). Quantitative real-time PCR (qPCR) to measure mRNA expression levels was performed with SYBR Green PCR Mix (Applied Biosystems, USA). β-actin was utilized as an internal control to normalize mRNA expression, and relative mRNA expression is presented as the fold change compared to the control group. The sequences of all primers used are listed in Supplementary Table 18.
Detection of GSH
Cell pellets were resuspended in phosphate buffer solution (PBS), homogenized, and centrifuged at 1500 × g for 10 min. Cell supernatant was used to detect the content of glutathione (GSH) using a commercial kit from Beyotime Biotechnology (Shanghai, China) following the manufacturer’s instructions.
Detection of ROS
The 2,7-dichlorofluorescein diacetate (DCFHDA, Thermo Scientific, USA) was used as a fluorescent probe to detect intracellular reactive oxygen species (ROS) in HLECs. The DCFHDA concentration used was 10 µM. The cells were incubated with DCFHDA at 37 °C for 30 min and then washed twice with PBS. The cell nuclei were labeled with Hoechst staining solution (Biosharp, China). After that, cells were photographed using a Nikon fluorescence microscope.
Lens organ culture ex vivo
Six-week-old male Sprague-Dawley rats were treated according to the Association for Research in Vision and Ophthalmology (ARVO) guidance, and study was approved by the Institutional Animal Care Committee of the Ninth People’s Hospital, Shanghai Jiao Tong University School of Medicine. Lenses were extracted from these rats euthanized by CO2 asphyxiation followed by cervical dislocation. Whole eyes were removed immediately and placed in PBS. Lenses were then dissected from the globe and incubated in the culture medium with different treatments at a humidified atmosphere at 37 °C with 5% CO2. The culture medium with different treatments was refreshed every day. After incubation for 7 days, lenses were imaged under darkfield and brightfield microscopy. Lens opacity was measured and calculated using ImageJ software (National Institutes of Health, Maryland, USA).
Detection of superoxide anion
For rat lens epithelial cells, superoxide anions were measured using dihydroethidium (DHE, MedChemExpress, USA) probe according to the manufacturer’s instructions. Briefly, 10-μm-thick unfixed rat lens frozen sections were prepared and incubated with 10 μM DHE staining solution for 30 min at 37 °C followed by immediate observation under a fluorescence microscope. The nuclei were labeled with 4′,6-diamidino-2-phenylindole (DAPI, Invitrogen, USA) staining solution.
Histological staining
The lenses were prepared into frozen sections (10 μm thickness). The slices were fixed in formalin and washed in water, and then stained with hematoxylin and eosin (HE, Servicebio, China). Images were acquired by fluorescence microscope (Nikon).
Statistics and reproducibility
For significance testing, the two-sided Chi-square test was applied to examine the significance of differences in discrete variables (like genders and comorbidities) between the ARC participants and DC participants. The two-tailed t-test was applied for mean comparison between two groups, and the one-way analysis of variance with two-sided Tukey post-hoc test was applied for mean comparison among multiple groups. The two-tailed tests were used to examine the significance of the Spearman correlation and Pearson correlation analysis. All significance tests were conducted by GraphPad Prism 10 and Python 3.7, and the significance level was set at 0.05. The exact sample size for conducting each significance testing was noted in the figure legend.
The AUC, sensitivity, specificity, and corresponding 95% CI of machine learning models were calculated by GraphPad Prism 10 according to the predicted result by the model in Python 3.7. The principal component analysis and power analysis were performed by MetaboAnalyst 5.0. The correlation analysis was conducted using GraphPad Prism 10.
The DESI-MS data and targeted MRM analysis data were analyzed using MassLynx (version 4.2), including visualization and integration. The standard curve for MRM analysis was constructed by OriginPro 2025. The LC-MS/MS data analysis for the mixed samples was performed using SCIEX OS (version 3.4.0), including extracted ion chromatogram (XIC) peak extraction, peak integration, and metabolite identification by high-resolution MS and MS/MS.
The electron microscope experiments for the ferric NPs (including scanning electron microscopy analysis, high-angle annular dark field analysis, and elemental mapping analysis) were repeated three times independently with similar results. The three replicated uncropped Western blot scans were in Source Data. Scatter points in the bar graphs represented individual data points (if applicable).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Metabolic fingerprints by the NELDI-MS generated in this study are provided in Supplementary Data 1. The raw MS data can be freely accessed in the MassIVE database (accession code: MSV000099221, ProteomeXchange database identifier: PXD068490, https://doi.org/10.25345/C5QZ22W5B). Other data generated in this study are provided in the Supplementary Information and Source Data file. Source data are provided with this paper.
References
Liu, Y. C., Wilkins, M., Kim, T., Malyugin, B. & Mehta, J. S. Cataracts. Lancet 390, 600–612 (2017).
Ang, M. J. & Afshari, N. A. Cataract and systemic disease: a review. Clin. Exp. Ophthalmol. 49, 118–127 (2021).
Ong, K. L. et al. Global, regional, and national burden of diabetes from 1990 to 2021, with projections of prevalence to 2050: a systematic analysis for the global burden of disease study 2021. Lancet 402, 203–234 (2023).
Wong, T. Y. et al. Guidelines on diabetic eye care: the international council of ophthalmology recommendations for screening, follow-up, referral, and treatment based on resource settings. Ophthalmology 125, 1608–1622 (2018).
Wang, L. M. et al. Prevalence and treatment of diabetes in China, 2013-2018. J. Am. Med. Assoc. 326, 2498–2506 (2021).
Danaei, G. et al. Effects of diabetes definition on global surveillance of diabetes prevalence and diagnosis: a pooled analysis of 96 population-based studies with 331288 participants. Lancet Diabetes Endocrinol. 3, 624–637 (2015).
Malkani, S. & Mordes, J. P. Implications of using hemoglobin A1c for diagnosing diabetes mellitus. Am. J. Med. 124, 395–401 (2011).
Ye, W. et al. LncRNA MALAT1 regulates miR-144-3p to facilitate epithelial-mesenchymal transition of lens epithelial cells via the ROS/NRF2/Notch1/Snail pathway. Oxid. Med. Cell. Longev. 2020, 8184314 (2020).
Obrosova, I. G., Chung, S. S. M. & Kador, P. F. Diabetic cataracts: mechanisms and management. Diabetes Metab. Res. Rev. 26, 172–180 (2010).
Guijas, C., Montenegro-Burke, J. R., Warth, B., Spilker, M. E. & Siuzdak, G. Metabolomics activity screening for identifying metabolites that modulate phenotype. Nat. Biotechnol. 36, 316–320 (2018).
Qiu, S. et al. Small molecule metabolites: discovery of biomarkers and therapeutic targets. Signal Transduct. Target. Ther. 8, 132 (2023).
Sigrist, R. M. S., Liau, J., El Kaffas, A., Chammas, M. C. & Willmann, J. K. Ultrasound elastography: review of techniques and clinical applications. Theranostics 7, 1303–1329 (2017).
Shaukat, A. & Levin, T. R. Current and future colorectal cancer screening strategies. Nat. Rev. Gastroenterol. Hepatol. 19, 521–531 (2022).
Mani, V. et al. Electrochemical sensors targeting salivary biomarkers: a comprehensive review. Trac Trends Anal. Chem. 135, 116164 (2021).
Yang, C. J. et al. Precise detection of cataracts with specific high-risk factors by layered binary co-ionizers assisted aqueous humor metabolic analysis. Adv. Sci. 9, 2105905 (2022).
Pietrowska, K. et al. An exploratory LC-MS-based metabolomics study reveals differences in aqueous humor composition between diabetic and non-diabetic patients with cataract. Electrophoresis 39, 1233–1240 (2018).
Davies, N. M. Biopharmaceutical considerations in topical ocular drug delivery. Clin. Exp. Pharmacol. Physiol. 27, 558–562 (2000).
Sarwat, S. et al. Feasibility of silicon quantum dots as a biomarker for the bioimaging of tear film. Nanomaterials 12, 1965 (2022).
Wishart, D. S. Emerging applications of metabolomics in drug discovery and precision medicine. Nat. Rev. Drug Discov. 15, 473–484 (2016).
Liu, X. J. & Locasale, J. W. Metabolomics: a primer. Trends Biochem. Sci. 42, 274–284 (2017).
Dunn, W. B. et al. Procedures for large-scale metabolic profiling of serum and plasma using gas chromatography and liquid chromatography coupled to mass spectrometry. Nat. Protoc. 6, 1060–1083 (2011).
Theodoridis, G., Gika, H. G. & Wilson, I. D. LC-MS-based methodology for global metabolite profiling in metabonomics/metabolomics. Trac Trends Anal. Chem. 27, 251–260 (2008).
Wu, J. et al. Glaucoma characterization by machine learning of tear metabolic fingerprinting. Small Methods 6, 2200264 (2022).
Huang, Y. D. et al. Diagnosis and prognosis of breast cancer by high-performance serum metabolic fingerprints. Proc. Natl. Acad. Sci. USA 119, e2122245119 (2022).
Li, S. X. et al. Serum metabolic fingerprints characterize systemic lupus erythematosus. Adv. Sci. 11, 2304610 (2024).
Cao, J. et al. Deciphering the metabolic heterogeneity of hematopoietic stem cells with single-cell resolution. Cell Metab. 36, 209–221.e6 (2024).
Siebenhaar, M., Küllmer, K., Fernandes, N. M. D., Hüllen, V. & Hopf, C. Personalized monitoring of therapeutic salicylic acid in dried blood spots using a three-layer setup and desorption electrospray ionization mass spectrometry. Anal. Bioanal. Chem. 407, 7229–7238 (2015).
Soudah, T., Zoabi, A. & Margulis, K. Desorption electrospray ionization mass spectrometry imaging in discovery and development of novel therapies. Mass Spectrom. Rev. 42, 751–778 (2023).
Takáts, Z., Wiseman, J. M. & Cooks, R. G. Ambient mass spectrometry using desorption electrospray ionization (DESI): instrumentation, mechanisms and applications in forensics, chemistry, and biology. J. Mass Spectrom. 40, 1261–1275 (2005).
Liu, W. S. et al. Monitoring retinoblastoma by machine learning of aqueous humor metabolic fingerprinting. Small Methods 6, 2101220 (2022).
Kohli-Lynch, C. N. et al. Cost-effectiveness of low-density lipoprotein cholesterol level-guided statin treatment in patients with borderline cardiovascular risk. JAMA Cardiol. 4, 969–977 (2019).
Tatonetti, N. P., Ye, P. P., Daneshjou, R. & Altman, R. B. Data-driven prediction of drug effects and interactions. Sci. Transl. Med. 4, 125ra31 (2012).
Sun, S. Y., Liu, W. S., Yang, J., Wang, H. & Qian, K. Nanoparticle-assisted cation adduction and fragmentation of small metabolites. Angew. Chem. Int. Ed. 60, 11310–11317 (2021).
Halama, A. et al. Measurement of 1,5-anhydroglucitol in blood and saliva: From non-targeted metabolomics to biochemical assay. J. Transl. Med. 14, 140 (2016).
Pinarci, E. Y. et al. Anterior segment complications after phacovitrectomy in diabetic and nondiabetic patients. Eur. J. Ophthalmol. 23, 223–229 (2013).
Greenberg, P. B. et al. Prevalence and predictors of ocular complications associated with cataract surgery in United States veterans. Ophthalmology 118, 507–514 (2011).
Haddad, N. M. N., Sun, J. K., Abujaber, S., Schlossman, D. K. & Silva, P. S. Cataract surgery and its complications in diabetic patients. Semin. Ophthalmol. 29, 329–337 (2014).
Shah, A. S. & Chen, S. H. Cataract surgery and diabetes. Curr. Opin. Ophthalmol. 21, 4–9 (2010).
Moreno-Oyervides, A. et al. Early, non-invasive sensing of sustained hyperglycemia in mice using millimeter-wave spectroscopy. Sensors 19, 3347 (2019).
Fuhrer, T., Heer, D., Begemann, B. & Zamboni, N. High-throughput, accurate mass metabolome profiling of cellular extracts by flow injection - time-of-flight mass spectrometry. Anal. Chem. 83, 7074–7080 (2011).
Lin, L. et al. Direct infusion mass spectrometry or liquid chromatography mass spectrometry for human metabonomics? A serum metabonomic study of kidney cancer. Analyst 135, 2970–2978 (2010).
Lai, X. P. et al. Combining MALDI-MS with machine learning for metabolomic characterization of lung cancer patient sera. Anal. Methods 14, 499–507 (2022).
Tanabe, K. et al. Supplemental feeding of 1,5-anhydro-d-glucitol prevents the onset and development of diabetes through the suppression of oxidative stress in KKAy mice. Food Sci. Technol. Res. 29, 413–421 (2023).
Buse, J. B., Freeman, J. L. R., Edelman, S. V., Jovanovic, L. & McGill, J. B. Serum 1,5-anhydroglucitol (GlycoMark™): a short-term glycemic marker. Diabetes Technol. Ther. 5, 355–363 (2003).
Luo, Z. L. et al. Angiotensin II induces podocyte metabolic reprogramming from glycolysis to glycerol-3-phosphate biosynthesis. Cell. Signal. 99, 110443 (2022).
Askari, A. A. et al. Basal and inducible anti-inflammatory epoxygenase activity in endothelial cells. Biochem. Biophys. Res. Commun. 446, 633–637 (2014).
Edwards, L. M. et al. Metabolomics reveals increased isoleukotoxin diol (12,13-DHOME) in human plasma after acute intralipid infusion. J. Lipid Res. 53, 1979–1986 (2012).
Jianbing, H. et al. The effect of allograft inflammatory factor-1 on inflammation, oxidative stress, and autophagy via miR-34a/ATG4B pathway in diabetic kidney disease. Oxid. Med. Cell. Longev. 2022, 1668000 (2022).
Yida, Z. et al. High fat diet-induced inflammation and oxidative stress are attenuated by N-acetylneuraminic acid in rats. J. Biomed. Sci. 22, 96 (2015).
Kohnz, R. A. et al. Protein sialylation regulates a gene expression signature that promotes breast cancer cell pathogenicity. ACS Chem. Biol. 11, 2131–2139 (2016).
Kim, W. J. & Park, C. Y. 1,5-anhydroglucitol in diabetes mellitus. Endocrine 43, 33–40 (2013).
Acknowledgements
We would like to express our gratitude to the patients and healthy volunteers who made this work possible. We’d like to thank Tian Xu, Ye Tian (undergraduate students from Shanghai Jiao Tong University School of Medicine) for their valuable assistance in animal handling and laboratory experiments during this study. We’d like to thank Dr. Bona Dai (Instrumental Analysis Center of Shanghai Jiao Tong University) for assistance with the NMR analyses. This work was supported by the National Key Research and Development Program of China (Projects 2022YFC2502801 (J. Wan) and 2024YFE0102200 (J. Wan)), the National Natural Science Foundation of China (Projects 22474040 (J. Wan), 82371102 (Y. Su), 82371054 (T. Guo), 82201137 (C. Yan)), Shanghai Key Clinical Specialty, Shanghai Eye Disease Research Center (2022ZZ01003, X. Fan), the Key Program of Shanghai Science and Technology Commission (23JC1402400, X. Fan). This work was supported by the Shanghai Frontiers Science Center of Molecule Intelligent Syntheses (J. Wan). This work was sponsored by the foundation of Shanghai Key Laboratory of Thoracic Tumor Biotherapy (No. 2025SZ1006, J. Wan). We also acknowledge SciDraw.io and its contributors (Vivek Kumar, John Chilton, and Alexander Bates) for providing freely available open resources (CC-BY 4.0), which inspired some of the scientific drawings in this work.
Author information
Authors and Affiliations
Contributions
J. Wan, Y. Su, X. Fan and T. Guo conceived the study and acquired the financial support. Z. Qi, Y. Zhao, S. Li, YT. Wang, Y. Lin and J. Hou contributed analytical method development and data analysis. M. Wang, C. Yan, and W. Zhuang collected clinical samples and analyzed clinical data. Z. Qi, M. Wang, C. Yan, and W. Zhuang contributed cell and animal evaluation. Z. Qi, YH. Wang, X. Chen, and W. Shu contributed synthesis, characterization, and performance testing for the matrix material. All authors joined in the critical discussion and edited the manuscript. Z. Qi, M. Wang and C. Yan contributed equally to this work.
Corresponding authors
Ethics declarations
Competing interests
Z. Qi and J. Wan have patents for both the analytical technology and the use of the technology to detect bio-samples in preparation, and not yet filed. The other authors declare no competing interests. Received: ((will be filled in by the editorial staff)) Revised: ((will be filled in by the editorial staff)) Published online: ((will be filled in by the editorial staff)).
Peer review
Peer review information
Nature Communications thanks Rosa Fernandes, Akos Vertes, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Qi, Z., Wang, M., Yan, C. et al. A mass spectrometry-based strategy allows signature metabolite identification in tear fluid from people with diabetic cataracts. Nat Commun 16, 10246 (2025). https://doi.org/10.1038/s41467-025-65082-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65082-7