Abstract
Aerated caves receive minimal light energy, yet host diverse microbial communities and the strategies allowing them to meet energy and carbon needs remain unclear. We determined the processes and mediators of primary production in aerated limestone and basalt caves through paired metagenomic and biogeochemical profiling. Four caves were sampled, including sediments and biofilms, yielding 94 metagenomes. Based on 1458 metagenome-assembled genomes, over half of microbial cells encode enzymes to use atmospheric trace gases as energy and carbon sources. The most abundant microbes are chemosynthetic primary producers, notably the gammaproteobacterial methanotrophic order Ca. Methylocavales and two uncultivated actinobacterial genera predicted to grow on atmospheric hydrogen, carbon dioxide, and carbon monoxide. Biogeochemical and isotopic measurements confirmed that these gases are rapidly consumed at rates likely sustaining a substantial fraction of the community and potentially driving primary production. Conventional chemolithoautotrophs, using ammonium and sulfide, are also enriched and active. Altogether, these results indicate that caves are unique in microbial biodiversity and the biogeochemical processes sustaining them. Consumption of atmospheric trace gases likely has a dual role in caves, providing energy for microbial survival and potentially supporting chemosynthetic growth, thereby introducing organic carbon. This process, defined as ‘aerotrophy’, operates alongside organic and inorganic inputs.
Similar content being viewed by others
Introduction
Terrestrial caves provide unique habitats for life, given that they are dark, humid, thermally insulated, and relatively isolated systems1,2. The sediments and mineral surfaces of caves harbour abundant and diverse microbiota. The nine dominant bacterial phyla from surface soils are also the most abundant microbes in caves, with most assumed to be aerobes dependent on organic carbon (i.e. organoheterotrophs)3,4,5,6. However, a long-standing question is how these microbes obtain sufficient carbon and energy to live in these environments with minimal photosynthesis7. Some caves receive extensive external inputs of organic matter (allochthonous sources), for example, due to seasonal flooding, groundwater percolation, or animal sources (e.g. bat guano8). However, these inputs are spatially and temporally variable9, and there is extensive evidence that most caves are highly nutrient-limited ecosystems10. It’s also well-recognised that chemosynthetic microbes produce organic matter (autochthonous sources) by harnessing the inorganic energy sources from reduced sulphur, nitrogen, and iron compounds present in drip water and mineral surfaces6,11,12,13. Given that these inorganic compounds generally occur in trace amounts, chemosynthetic processes are thought to play a minor role in cave microbial ecosystems, except in globally rare, deep geothermally-heated caves usually isolated from the surface13,14,15,16,17,18.
A potential alternative source of energy and carbon in typical cave ecosystems is the atmosphere itself. We have recently discovered that atmospheric molecular hydrogen (H2) and carbon monoxide (CO) are critical energy sources supporting the biodiversity of soils and waters worldwide, and enable complex ecosystems to form in oligotrophic environments such as Antarctic soils19,20,21,22,23. Bacteria use high-affinity hydrogenases and CO dehydrogenases to liberate electrons from these gases for aerobic respiration and carbon fixation via the Calvin–Benson–Bassham (CBB) cycle24. Aerobic methanotrophs, which use atmospheric methane (CH4) as a dual energy and carbon source25,26,27, have also been identified in various cave systems and mediate CH4 oxidation rates comparable to those of surface soils28,29,30,31,32,33,34,35,36,37. Considering these findings, we sought to disentangle the relative roles of atmospheric, rock-derived, and solar energy sources in supporting primary production and energy conservation in cave ecosystems. To do so, we integrated genome-resolved metagenomic profiling, in situ and ex situ biogeochemical and isotopic measurements, and thermodynamic modelling of sediment and biofilm microbial communities collected along transects from four aerated limestone and basalt caves sampled within Australia. Our results reveal that aerated caves host abundant communities of methane and hydrogen oxidisers, which continuously oxidise atmospheric trace gases to assimilate carbon and fix CO2. We predict that this process enables continuous growth, suggesting that aerated caves are potentially major systems sustained by aerotrophic primary production.
Results and discussion
Most cave microbes encode enzymes to harvest atmospheric energy sources
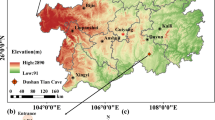
We studied four aerated caves in southeastern Australia, two limestone caves and two basalt lava tubes (Fig. 1a). In each cave, we sampled along a transect extending from the entrance to twilight, transition and deep zones. We collected a total of 94 environmental samples, including sediments, biofilm, and mineral samples, which were analysed using metagenomics as well as gas and isotopic studies to determine microbial composition, capabilities, and metabolic activity. The samples from the four caves (Fig. 1a, Supplementary Data 1) spanned a broad range of organic carbon (0.2–31.3%), pH (3.6–8.7), and moisture levels (10.8–55.8%). This suggests much spatial variability in their allochthonous supply of organic carbon and other resources, in support of previous studies7. Based on shotgun metagenomic profiling, microbial communities varied between cave sediments and biofilms, between rock types, and with cave depth (Fig. 1b–e; Supplementary Data 3a–c). Microbial abundance (av. 6.7 × 109 16S rRNA gene copies per gram of dry sediment) and richness (av. 450 taxa per sample; Chao1 based on metagenomic reads of 16S rRNA genes of sediment) decreased by 3.3-fold and 2-fold, respectively, between the cave entrance and interior sediments (Supplementary Data 2 and 3b). In line with most other sampled caves5, Actinobacteriota, Proteobacteria, Acidobacteriota, Chloroflexota and Gemmatimonadota were the most common phyla, along with Thermoproteota (predominantly Nitrosophaerales) (Fig. 1b; Supplementary Data 3c). Metagenomic assembly and binning yielded 1458 dereplicated high- and medium-quality metagenome-assembled genomes (MAGs) spanning 35 different phyla, including one unclassified (Supplementary Data 4c).
a Maps showing geographic and lithology setting of the caves studied103. Detailed cave morphology, major environment features, study sites and scale are shown, redrawn from material provided by the Victorian Speleological Association Inc. b Stacked barchart showing phylum-level community structure at the sample level, vertical-coloured bars show the environmental diversity captured according to sample types. c Boxplot of 16S rRNA gene copy number wet weight for bulk sediments and brown biofilms, with biological replicates (n = 44) grouped by site and host rock. d Boxplot of observed and estimated richness with biological replicates (n = 94) grouped by site and host rock. e Non-metric multidimensional scaling (NMDS) plot of beta diversity (Bray-Curtis), visualising differences in community structure according to host rock. Ellipses represent 95% concentration regions for each group, which summarise the dispersion of samples in NMDS space. Boxplots display the median (centre line), interquartile range (IQR; box) and whiskers extending to values within 1.5 × IQR of the box.
The energy and carbon acquisition strategies of the cave microbes are inferred by searching for 52 conserved marker genes encoding the potential of key energy and carbon acquisition processes in the MAGs and short reads (Fig. 2a; Supplementary Data 4a–c). These analyses suggested that most cave bacteria mediate aerobic respiration using both organic compounds and trace gases as substrates (Fig. 2a, b). 44% of MAGs, accounting for 54% of mapped metagenomic reads (Supplementary Data 4c), encode enzymes to consume one or more atmospheric trace gases, namely form I CO dehydrogenases for CO oxidation (25.4% genomes/74.3% community based on short reads; Supplementary Fig. 1), group 1 and 2 [NiFe]-hydrogenases for H2 oxidation (25.8%/43%; Supplementary Fig. 2), and particulate methane monooxygenases for CH4 oxidation (2.9%/5.5%; Supplementary Fig. 3). These findings suggest that most microbial cells in caves encode enzymes that could enable trace gas oxidation. A smaller proportion of cave microbes can also use rock-derived energy sources such as sulphide (13.9%/18%), thiosulfate (6.7%/7%), ammonia (4.1%/3.5%; Supplementary Fig. 4), nitrite (1.2%/3.7%; Supplementary Fig. 5), and ferrous iron (2.7%/2.2%).
a Heatmap showing the metabolic potential of the community as average gene copies per organisms for major energy and carbon acquisition pathways across limestone and volcanic caves in Australia in comparison to global samples. b Dot plot showing the metabolic potential at the genome-resolved level (MAGs). Each dot in the blue shading represents encoded metabolic function and the shading represents average genome completeness at the phylum level. Orange lollipop charts show the percentage of MAGs encoding each gene and purple lollipop charts show the relative abundance maximum for each phylum.
As expected, photosynthesis genes were abundant in sediments and biofilms at the entrance of each cave (29%) but declined by an average of 73-fold in the cave interior. Conversely, there were marked increases in cave interior compared to entrance communities for the genes enabling the oxidation of CH4 (7.3-fold), ammonium (1.5-fold), nitrite (2.3-fold), sulphide (1.4-fold), H2 (2-fold, average group 1h & 1l [NiFe]-hydrogenases) and CO (1.1-fold). Concordant patterns were observed for carbon fixation genes, with the photosynthetic cyanobacterial Type IB RuBisCO decreasing 39-fold, and the chemosynthetic, predominantly actinobacterial Type IE RuBisCO increasing 2-fold in cave interiors compared to entrances (Supplementary Data 4b, c; Fig. 2a, b; Supplementary Fig. 6). Some carbon fixation also likely occurs through primarily 4-hydroxybutyrate cycle (3.6%/2.2%), reductive tricarboxylic acid cycle (1.6%/2%), and 3-hydroxypropionate cycle (0.9%/0.7%). Altogether, these findings indicate a shift from photosynthetic to chemosynthetic primary production in cave ecosystems, driven both by atmospheric and rock-derived substrates. A driver analysis, using both random forest and linear models, revealed that key soil physicochemical components significantly influence the abundance of different community members. Trace gas oxidisers encoding genes for CH4, H2 and CO responded variably between basalt and limestone caves to a wide range of factors, primarily to electrical conductivity (EC), pH and total organic carbon (TOC). The response of ammonia, nitrite, sulphide, and iron oxidisers was more conserved to limestone caves, suggesting pH, TOC and soil water content (SWC) as dominant drivers of their abundance (Supplementary Fig 11a, d; Supplementary Data 7d).
To ensure these insights were representative of caves worldwide, we further analysed twelve previously published metagenomes representative of diverse global cave ecosystems (Fig. 2a, Supplementary Data 4a). Oxidation of ambient trace gases and, to a lesser extent, rock-derived inorganic compounds are widespread strategies in sediments and rocks of cave interiors. For example, in rock metagenomes from Monte Cristo Cave (Brazil) and in white microbial mats in Kipuka Kanohina Cave (Hawaii), almost all microbes encode high-affinity hydrogenases38. The three exceptions are photosynthetic biofilms collected from an illuminated entrance and sinkhole, as well as a cave lake likely to receive considerable organic inputs (Supplementary Data 4b).
Previously uncharacterised microbes drive cave energy acquisition and primary production
We used genome-resolved metagenomics and phylogenetic analyses to resolve which microbes mediate these processes (Supplementary Data 4c; Supplementary Figs. 1–6). Most hydrogenases, CO dehydrogenases, and RuBisCOs were co-encoded by the most abundant Actinobacteriota lineages residing in the caves (primarily classes Actinomycetia, Thermoleophilia, Acidimicrobiia, and Ca. Aridivitia)21 (Fig. 3a), suggesting that they may be important primary producers in these ecosystems, although we cannot exclude the possibility of transient and dynamic heterotrophic and mixotrophic strategies. Multiple phyla nevertheless encoded each of these enzymes (14 hydrogenase-, 11 CO dehydrogenase-, and 10 RuBisCO-encoding phyla), highlighting that trace gas oxidation and chemosynthesis are ubiquitous traits (Supplementary Data 4c). These enzymes were also encoded by various uncultivated lineages, for example with CO dehydrogenases being encoded by high-quality genomes from the candidate bacterial phyla CSP1-3 and KSB1, as well as two enigmatic orders (RBG-16-68-12, UBA184) of Thermoplasmata archaea inhabiting diverse cave samples (Supplementary Fig. 1). Corroborated by the short-read analysis (Fig. 2a, Supplementary Data 4b), almost all of these belong to high-affinity clades of hydrogenases (groups 1h, 1l and 2a [NiFe]-hydrogenases21,39,40) (Fig. 2a), suggesting they adapted to atmospheric rather than elevated levels of these gases. Of the 30 most abundant microbes based on genome read mapping (Supplementary Data 4), 21 were capable of trace gas oxidation, including four putative methanotrophs (all affiliated with the USCγ/JACCXJ01 clade), whereas none mediated photosynthesis, nitrification, sulphide oxidation, or iron oxidation. The top ten most abundant microbes (comprising 20% of all mapped reads) were all from uncultivated genera from Pseudonocardiaceae and Egibacteraceae (both within class Actinomycetia), each of which co-encode RuBisCO with either CO dehydrogenase and/or uptake hydrogenases. Thus, caves appear to select for highly productive actinobacterial primary producers that grow on atmospheric energy and carbon sources. These Actinobacteriota are the most abundant lineages in the cave biofilms, whereas the methanotrophs are the single most abundant species in the cave sediments.
a Differential abundance of key taxa (MAGs n = 271), at genus and family level, between cave entrance, interior sediment, and interior biofilm samples. Box plots show the relative abundance, based on read mapping, of MAGs of hydrogenotrophs (teal), methanotrophs (red), nitrifiers (blue), sulphur oxidisers (yellow) and phototrophs (green). Pairs are denoted with asterisks showing significance. Taxonomic classification is shown at the family level and at its preceding rank if unclassified, and brackets show phylum-level affiliation. Kruskal–Wallis tests were performed per family (two-sided) with Bonferroni correction across families. For significant results, Dunn’s post-hoc tests were applied with Bonferroni correction across pairwise comparisons. */ **/ ***/ **** corresponding to adjusted p-values ≤ 0.05, ≤0.01, ≤0.001, and ≤0.0001, respectively. Exact p-values are provided in the source data. Boxplots display the median (centre line), interquartile range (box) and whiskers extending to values within 1.5 × IQR of the box. b Metabolic reconstruction of the three dominant hydrogenotrophic taxa, the candidate genera Hydrogenomurus, Hydrogenocavus and Hydrogenolapis. All encode genes consistent with trace atmospheric gas oxidation, including a group 1h [NiFe] hydrogenase and CO dehydrogenase. However, only two of the three taxa encode a Type IE RuBisCo consistent with carbon fixation via the Calvin–Benson–Bassham cycle. Hydrogenolapis are predicted to utilise organic carbon sources. c Metabolic reconstruction of the three dominant methanotrophic MAGs, Methylocella (USCα), Methyloligotrophales (USCγ), and Methylocavales. All encode particulate methane monooxygenase and a methanol dehydrogenase. Carbon fixation in Methylocella MAGs can occur via the tetrahydrofolate pathway and serine cycle, but remains unresolved in the Gammaproteobacterial MAGs, which lack key genes in the serine and RuMP pathways, and Calvin-Benson-Bassham cycle.
We comprehensively analysed the energy and carbon acquisition pathways of the three most abundant predicted hydrogenotrophs in the caves (Fig. 3b), namely the candidate genera herein named Hydrogenomurus, Hydrogenocavus and Hydrogenolapis (all etymological information in Supplementary Note 2; formerly Pseudonocardiaceae GCA-003244245, Egibacteraceae JACCXR01, and Actinomycetia JACCUZ01). These lineages were selectively enriched in distinct habitats, with Hydrogenomurus prevalent across basalt caves and constituting over half of multiple biofilm communities (up to 73%), Hydrogenocavus dominant in limestone biofilms and moonmilks (up to 54%), and Hydrogenolapis abundant in both biofilms and sediments within limestone caves (Fig. 3a; Supplementary Data 4c). All three taxa encode high-affinity group 1h [NiFe]-hydrogenases and CO dehydrogenases, consistent with the use of trace gases as an energy source. In addition, these MAGs encode a complete TCA cycle and a full suite of aerobic respiratory complexes (I-V), indicating they can conserve energy through both lithotrophic and organotrophic aerobic respiration. Both Hydrogenomurus and Hydrogenocavus also encode Type IE RuBisCO and a complete CBB cycle, indicating they are facultative chemolithoautotrophs; the lack of enzymes for oxidation of rock-derived inorganic compounds strongly suggests reductants necessary for carbon fixation are provided by H2 and CO. Their autotrophic capacity likely underlines their dominance along nutrient-limited cave mineral surfaces. Conversely, Hydrogenolapis appears to be solely reliant on organic carbon sources, potentially including peptides and mono- and disaccharides, based on the presence of various ABC transporters. Energy provided by atmospheric trace gases likely enables these microbes to allocate more organic carbon for anabolism than catabolism41. All taxa encode the pentose phosphate and Embden–Meyerhof–Parnas pathways for organic carbon catabolism, although only Hydrogenomurus MAGs encode all genes necessary for complete glycolysis.
Methanotrophs are the most enriched metabolic specialists in the cave interior, based on both genomic read mapping (Fig. 3a; Supplementary Data 4b, c) and marker gene profiles (Fig. 2a, b). Given this, we performed an in-depth analysis to resolve the evolutionary history and functional capabilities of putative cave methanotrophs encoding particulate methane monooxygenases (PmoABC), as elaborated in Supplementary Note 1. A genome tree revealed that these bacteria span the alphaproteobacterial genus Methylocella (encompassing Methylocapsa within the GTDB framework), the gammaproteobacterial order Methylococcales, and two candidate gammaproteobacterial orders herein named Methyloligotrophales and Methylocavales (formerly JACCXJ01 and CAJXQU01; etymology in Supplementary Note 2). These methanotrophs were progressively enriched from entrance and cave sediments to biofilms, with a single Methyloligotrophales MAG encompassing 10.8% of microbes in a complex limestone sediment (Fig. 3a; Supplementary Data 4c), suggesting CH4 is a primary growth substrate of cave communities. While Methylocella and Methyloligotrophales encompass the USCα and USCγ lineages of atmospheric methanotrophs25,26,42,43, Methylocavales is not known to be methanotrophic and is represented by just one previously reported genome that lacks pmo genes. Consistent with being novel methanotrophs, the cave-exclusive Methylocavales bacteria each encode complete pmoCAB operons and their PmoA protein formed a novel sister clade with Methyloligotrophales (Supplementary Fig. 3a-b). They also encode a complete set of genes to oxidise methanol (lanthanide-dependent methanol dehydrogenases), formaldehyde (tetrahydromethanopterin pathway), and formate (formate dehydrogenase) to carbon dioxide for energy conservation (Fig. 3c, Supplementary Fig. 3c). However, in common with other atmospheric gammaproteobacterial methanotrophs, including the Methyloligotrophales MAGs analysed, the carbon assimilation pathways used remain incompletely resolved (Supplementary Note 1).
Though the capacity for oxidation of rock-derived inorganic substrates was less widespread, numerous chemolithoautotrophs were nevertheless highly enriched in cave sediments and biofilms. Most notable are nitrifiers, including ammonia-oxidising archaea and bacteria, as well as nitrite-oxidising and comammox Nitrospirales (Fig. 2a, b; Supplementary Data 4b, c). Members of three archaeal families, Nitrososphaeraceae, Nitrosopumilaceae (e.g. acidophilic Nitrosotalea dominant in basalt caves), and novel clade Nitrosomiraceae (e.g. Ca. Nitrosomirus44 abundant in limestone caves) vastly outnumber ammonia-oxidising bacteria (Nitrosospira) (Fig. 3a); most encode high-affinity Amt1 ammonia transporters, carbon sequestering ABC-type bicarbonate transporters, and carbonic anhydrase consistent with their oligotrophic lifestyle45. Remarkably, urease, cyanase, and glycine cleavage system genes were also present in most MAGs, suggesting these archaea also sequester ammonia from organic substrates such as urea, cyanate, and glycine. Comammox Nitrospirales (genus Palsa-1315), known for their high-affinity for ammonia46, were also widely distributed (Supplementary Data 4b, c). These cave nitrifiers use distinct pathways to fix CO2, spanning the 4-hydroxybutyrate cycle (Nitrososphaerales), the reductive tricarboxylic acid cycle (Nitrospirales), and the CBB cycle (Nitrosospira). Altogether, these findings indicate that caves select for oligotrophic chemolithoautotrophs and that nitrifiers are likely important primary producers given their autotrophic lifestyle and enrichment. Apart from nitrification, some nine phyla were capable of sulphide oxidation, including numerous Proteobacteria and Actinobacteriota MAGs. We also reconstructed genomes of iron-oxidising Acidobacteriota and Proteobacteria (Fig. 2a, b).
Trace gas oxidation occurs together with nitrification and sulphide oxidation
To substantiate these findings, we performed in situ and ex situ profiling of the processes of trace gas oxidation, nitrification, and sulphide oxidation in each cave. In situ measurements of sediment-to-atmosphere fluxes revealed ambient average concentrations in limestone/basalt cave entrances of CH4 (1.5/0.65 ppmv) H2 (0.71/0.34 ppmv), and CO (0.06/0.03 ppmv), respectively (Fig. 4a; Supplementary Fig. 7). It is likely that these gases are predominantly atmospherically derived, given their concentrations are at or below global average concentrations of these gases in the lower troposphere47,48,49, though it can’t be ruled out that they are also produced through geogenic and biological processes and maintained at such concentrations through microbial consumption. On average, ambient air concentrations of all three gases decreased 4-fold from the cave entrances to the interiors of the limestone caves, suggesting microbial consumption (Fig. 4a; Supplementary Fig. 7). Equivalent trends were not observed in the basalt caves, though this is potentially due to their relatively consistent tunnel-like architecture (Fig. 1a) facilitating rapid equilibration with outside air, as opposed to decreased gas consumption. Though air exchange rates were not measured, based on in situ gas fluxes, consumption was observed across all four caves. On average, in situ CH4 fluxes increased from 1.1 nmol m−2 s−1 at the entrance to 3 nmol m−2 s−1 inside (mean of transient and equilibrium flux estimates), consistent with localised methanotrophic activity within caves (Fig. 4b; Supplementary Fig. 7, Supplementary Data 5a). In contrast, H2 fluxes were highest at the entrance and modestly declined inside the caves, averaging around 7.5 nmol m−2 s−1 (Fig. 4b; Supplementary Fig. 7, Supplementary Data 5a).
a Boxplots showing in situ atmospheric concentrations (ppm), with independent replicates (n = 81) grouped by gas, site and host rock. Dashed horizontal lines show average global concentration (2021-2022) for atmospheric H2 (0.53 ppm), CO (0.10 ppm) and methane (1.9 ppm). b Boxplots showing in situ sediment–atmosphere uptake gas fluxes (values indicate net gas consumption) with biological replicates (n = 12) grouped by gas and site. Flux phases provide upper (transient) and lower bound (equilibrium) estimates. c Boxplots showing bulk sediment and biofilm oxidation rates normalised to global average atmospheric gas and cell numbers, with biological replicates (n = 44) grouped by gas, site and host rock. d Boxplot showing the amount of power per cell derived from the oxidation of each trace gas, with biological replicates (n = 44) grouped by gas, site and host rock. Coloured bars depict the range of literature values of energy yield per cell by atmospheric methane oxidising isolates M. aurea KYG, M. gorgona MG08, M. palsarum NE2 and M. rosea SV97 (orange42), maintenance energy requirements or endogenous metabolic rates of different pure cultures (green104, yellow105, jade106) and hydrogen oxidisers in deep marine sediments (pink107). e Boxplot showing rates of nitrogen and sulphur compound metabolism, with positive values indicating accumulation and negative values showing uptake with biological replicates (n = 17) grouped by site. Nitrification is expected to result in ammonium (NH4+) consumption and nitrite (NO2−), nitrate (NO3−), whereas sulphide oxidation is expected to cause sulphate (SO42−) production. Boxplots display the median (centre line), interquartile range (box) and whiskers extending to values within 1.5 × IQR of the box.
Given the limitations in conducting flux measurements only in areas with sufficient sediment depth for flux chambers, we employed microcosm incubations with bulk sediments and biofilms extracted from cave walls to validate these observations. In these incubations, H2, CO2 and CH4 were supplied at slightly elevated levels (10 ppmv each) in otherwise ambient headspaces, in order to calculate robust first-order rate constants and atmospheric consumption rates (see “Materials and Methods”). These trace gases were consumed by the microbial communities in all samples, including biofilms and sediments, though at variable rates; most samples consumed these gases to below atmospheric concentrations, reaching observed ambient in situ ranges and/or falling below average global mixing ratios (100%, 93%, and 43% of samples for H2, CO, and CH4, respectively) (Supplementary Figs. 8 and 9, Supplementary Data 5b). H2 was consumed most quickly on average, followed by CO and CH4. CH4 oxidation rates closely matched in situ patterns, especially in limestone caves, with a 16.5-fold increase in cell-specific CH4 oxidation rates (6.89 × 10−13 – 1.13 × 10−11 nmol cell−1 s−1) occurring from entrance to deep zones (Fig. 4c; Supplementary Figs. 8 and 9, Supplementary Data 5b), in line with metagenomic observations of increased methanotrophic abundance (Fig. 2a). A driver analysis for cell-specific reaction rates revealed a significant influence of soil physicochemistry, with pH, TOC, and SWC and EC as drivers of activity for H2, CO and CH4 oxidisers in limestone caves, whereas CO oxidisers responded more strongly to the concentration of these compounds in basalt caves (Supplementary Fig. 11b, e; Supplementary Data 7d).
We also tested whether rock-derived inorganic substrates, namely ammonium and sulphide, were also used as energy sources given the metagenomic observations (Fig. 2a). The cave sediments contained varying concentrations of ammonium (1.35–40.2 mg/kg), sulphur (7.3–2468 mg/kg), and iron (6.96–2003 mg/kg) (Supplementary Data 1). Microcosm incubations showed ammonium was oxidised at variable rates across the samples and increased from the entrance to the interior of the cave (Fig. 4e; Supplementary Fig. 10; Supplementary Data 6). Nitrite and nitrate also accumulated in these incubations, consistent with stepwise nitrification processes occurring within the cave environments. Sulphide oxidation was also evident from the accumulation of the end-product sulphate (Fig. 4e; Supplementary Fig. 10; Supplementary Data 6). It should be noted that our in situ and ex situ biogeochemistry results are based on a single sampling expedition per cave, yet there can be much seasonal variability in cave hydrology and substrate availability, including trace gases; more extensive seasonal sampling would be required to understand temporal variability in biogeochemical activities.
Trace gas oxidation drives community-wide carbon and energy provision
Thermodynamic calculations based on bulk oxidation rates of H2, CO and CH4 revealed that trace gas oxidation rates inside the cave yielded an average power output of 1.6 × 10−15, 6.2 × 10−16 and 4.7 × 10−13 W per H2, CO and CH4-oxidising cell, respectively. Power per cell outputs were similar between rock types and cave depths for H2 and CO, but were higher in basalt compared to limestone, and between surface and subsurface for CH4 (Fig. 4d; Supplementary Data 5b). As we did not simultaneously measure oxygen consumption, we are unable to determine what proportion of total aerobic respiration is attributable to trace gas oxidation. These values represent theoretical estimates averaged across all cells, and it is likely that only a fraction of the community is metabolically active, meaning the per-cell power for active trace gas oxidisers may be higher. Nevertheless, the observed rates may theoretically provide sufficient power to support the growth and survival of localised populations of trace gas oxidisers in these caves, particularly at the entrance and twilight sites in basalt caves. Indeed, based on our culture-based studies of four oligotrophic bacteria that grow on trace gases alone (e.g. M. gorgona), the CH4 power output in cave greatly exceeds growth requirements and H2 power output is at the same magnitude to support growth42,50. Importantly, these rates are averaged for all cells and samples, while in reality, some microbes may co-consume these gases, notably the highly abundant Hydrogenocavus and Hydrogenomurus. Altogether, our data indicate that atmospheric trace gas consumption can theoretically support localised activity, including growth of the methanotrophs and the survival of the hydrogenotrophs, with some bacteria potentially capable of chemolithoautotrophic growth using H2 and/or CO. We note that cultivation of these microorganisms will be a necessary next step to validate their capacity to grow on atmospheric gases.
To probe the major pathways forming organic matter in caves, we quantified the fractionation signature of biomass 13C/12C in cave sediments and biofilms (Fig. 5a). Two autotrophically-grown hydrogenotrophs and three methanotrophs (including atmospheric CH4 oxidiser M. gorgona) were also analysed for their carbon fractionation as a comparison. The organic fractions in the cave samples exhibited a depletion of 13C (δ13Corganic) from −21.7 to 40.7‰, consistent with biomass derived from CBB cycle, but distinct from other carbon fixation pathways51,52 (Fig. 5a). It is likely that organic carbon from external plant-derived inputs (allochthonous inputs) is transported into cave interiors. However, despite δ¹³Corganic values overlapping those of C₃-plant detritus (a shared CBB signature), multiple lines of evidence suggest cave-dwelling primary producers are also major sources of organic matter (autochthonous inputs). First, δ13Corganic was progressively more negative from sediments in limestone caves to biofilms in basalt caves, in line with the increasing trends of bacterial rbcL and pmoA gene abundance (Figs. 2a and 5b). Second, random forest analysis reveals rbcL abundance is the best carbon assimilation gene predictor for δ13Corganic (Fig. 5b), and also the most negatively correlated with δ13Corganic among all metabolic marker genes (Spearman’s rho = −0.56, p = 2.0 × 10−6), also suggesting autochthonous inputs from the CBB cycle. Moreover, the δ13Corganic signatures are similar to those of the tested hydrogenotrophs (Fig. 5a) and, as elaborated below, we directly detected considerable hydrogenotrophic CO2 fixation through ex situ assays. CH4 assimilation, which yields biomass strongly depleted in 13C, may also contribute to the negative δ13Corganic in some cave samples (Fig. 5a). This was supported by calculating methane carbon assimilation as a function of cell-specific methane oxidation rates (Fig. 5c). 13C/12C signatures were significantly associated with both SWC and TOC (Supplementary Fig. 11c; Supplementary Data 7d).
a Boxplot showing depletion of biomass 13C stable isotope (δ13Corganic) across biological replicates of cave sediments (n = 39), biofilms (n = 50), and selected autotrophically-grown hydrogenotroph (n = 6) and methanotroph (n = 9) pure cultures. Coloured bars depict the range of literature values of δ13Corganic of biomass produced from 4-hydroxybutyrate cycle (4HB cycle; green), 3-hydroxypropionate cycle (3HP cycle; yellow), reductive tricarboxylic acid cycle (rTCA; jade), and Calvin–Benson–Bassham cycle (CBB cycle; pink). δ13Corganic for pure cultures and literature values were adjusted based on the use of atmospheric CO2 (δ13C: −8.5‰) and CH4 (δ13C: −47.2‰) as sole carbon sources. b Heatmap showing the abundance ratio of key carbon assimilation marker genes (pmoA, mcr, hbsT, aclB, rbcL) in biofilm against sediment communities (top) and random forest analysis of these genes as predictors for cave δ13Corganic values (bottom). c Boxplot showing methane carbon assimilation rate as a function of bulk sediment oxidation rates normalised to median assimilation rates commonly observed across soil ecosystems, with biological replicates (n = 39) grouped by site/type and host rock. d Boxplots showing 14C-CO2 fixation rates under three conditions: dark carbon assimilation (basal rate of CO2 incorporation under dark ambient conditions due to carbon fixation or anaplerotic processes); photosynthetic carbon fixation (CO2 fixed under light ambient conditions); and hydrogenotrophic CO2 fixation (CO2 fixed under dark H2-enriched conditions). For each cave, nine independent biological replicates were pooled, and three replicates for the entrance. Each pooled sample was analysed in technical triplicates (n = 24), yielding two independent biological duplicates for each host rock and site. Note that the rates do not represent gross carbon fixation rates due to the presence of unlabelled native inorganic carbon and internally recycled CO2 within samples. Boxplots display the median (centre line), interquartile range (box) and whiskers extending to values within 1.5 × IQR of the box.
Finally, we traced radioisotope incorporation of 14C-CO2 into biomass to ascertain the relative contributions of dark, hydrogenotrophic, and photosynthetic CO2 fixation pathways, including as a source of the δ13Corganic signatures. Whereas photosynthesis was strongly stimulated at the entrance of basalt caves, it was negligible elsewhere. Hydrogenotrophic CO2 fixation was observed in cave interiors, with two-fold and five-fold more carbon fixed in limestone and basalt caves respectively compared to dark conditions (Fig. 5d; Supplementary Data 7a). Notably, basalt biofilm and sediment microbes mediated higher hydrogenotrophic CO2 fixation activities (5.7 × 10−13 nmol 14C cell−1 min−1) compared to average rates by limestone sediment microbes (5.3 × 10−14 nmol 14C cell−1 min−1). This supports our metagenomic inferences that H2 is the predominant driver of CO2 fixation and may contribute to the observed δ13Corganic signatures in caves. Gross carbon fixation rates are probably orders of magnitude higher than measured 14CO2-fixation rates, as unlabelled native inorganic carbon, CO2 from the ambient air headspace, and internally recycled CO2 within samples invariably competed with the trace levels of added 14CO2 (400 ppm), especially in carbonate-rich limestone samples (1.7–29% carbon; Supplementary Data 1). Therefore, these rates are a conservative estimate to only reflect the potential for hydrogenotrophic CO2 fixation under elevated H2 conditions, and should not be interpreted as ambient net CO2 fixation rates. Methanotrophs also contribute to carbon acquisition from methane, as they are estimated to assimilate −3.5 × 10−7 to −1.1 × 10−12 nmol C cell−1 min−1 based on their ex situ cell-specific oxidation rates, and a theoretical 40% methane-carbon conversion efficiency53,54,55, which is further supported by their activities based on the in situ flux analysis (Fig. 5c; Supplementary Data 7b). We acknowledge that methanotrophic CO2 fixation via the CBB cycle, as well as other dark and anaplerotic processes not accounted for in this study, may contribute to CO2 fixation in caves. These experiments demonstrate that microbial energy and carbon acquisition from atmospheric substrates occur continuously, with chemosynthetic primary productivity being sustained across a range of cave surfaces.
Aerotrophy as a key process in cave ecosystems
Here, we provide extensive metagenomic and biogeochemical evidence for the consumption of atmosphere-derived trace gases in cave sediments and biofilms. This consumption may support a potentially large fraction of the microbial community. Primary production appears to be driven by highly abundant and active methanotrophs oxidising atmospheric methane. Previously undescribed lineages of actinobacterial lithoautotrophs are predicted to oxidise hydrogen to fix CO2. Together, these organisms continuously use the gases methane, hydrogen, carbon dioxide and carbon monoxide present in cave atmospheres. Cave ecosystems differ from polar desert soils, the other major type of ecosystem shown to be atmospherically powered 19,20,21, in that trace gases appear to drive continual growth rather than long-term survival in these nutrient-deprived environments. This is reflected by the abundant primary producers in cave sediments and biofilms, the rapid fluxes and activities of trace gas oxidisers, and the theoretical considerations based on thermodynamic and biogeochemical modelling. Given the relatively stable environment, humid conditions, and aeration of most caves, it is predicted that aerotrophs will continually cycle gases such as hydrogen, methane, and carbon dioxide, whether of atmospheric origin or geogenic origin (including at atmospheric concentrations), allowing continuous primary production. On this basis, we propose defining the term “aerotrophy” as “the process of growth through the use of atmospheric trace gases, as energy and carbon sources” (derived from Greek, by combining the prefix “aero” meaning “air” and the suffix “trophic”, derived from “trophē”, meaning “nourishment”), with caves as major systems often driven by ‘aerotrophic microorganisms’.
Aerotrophy nevertheless co-occurs with other energy and carbon acquisition processes in cave systems. There is likely much spatiotemporal variation in the relative contributions of atmospheric, inorganic, organic, and even solar energy sources within and between caves. We provide metagenomic and biogeochemical evidence that other chemosynthetic processes, including nitrification and sulphide oxidation, are also important in diverse caves and are likely to be dominant where rock-derived substrates are locally elevated. Moreover, while the atmosphere is most likely the primary source of trace gases in the caves sampled, some caves or microenvironments therein will have elevated inputs through geological processes (e.g. serpentinisation, radiolysis) and biological production (e.g. fermentation, nitrogen fixation, methanogenesis) that could supplement primary production30,37,56. Our analyses also do not differentiate the relative contribution of aerotrophy and other chemosynthetic carbon fixation (autochthonous inputs) and external organic matter (allochthonous inputs) to the organic carbon levels observed in the sampled caves. External inputs are likely to be substantial in certain sections of the caves, in line with the high moisture and carbon content of certain samples, and will also vary in response to seasonal hydrological dynamics. These inputs are predicted to primarily enter the cave via sinking streams and surface water that gradually seeps through soil and rock layers9. While sinking streams can provide substantial concentrations of organic carbon comparable with surface streams, water percolation tends to spread smaller amounts of organic carbon over a wider area. Depending on the geomorphology of each cave, these modes of carbon and nutrient inputs likely interact to create the varied local conditions we observed. Given the genome collection suggests most of the bacteria in these caves can use both trace gases and organic carbon to meet their resource needs, it is likely most will take advantage of external carbon whenever available across space and time, but continually sustain themselves on trace gases otherwise. Indeed, given that atmospheric trace gases are such rich and dependable energy sources, they are continuously used by the most abundant microbes in these ecosystems. Exceptions likely include those caves that have more extensive solar or guano inputs, are disconnected from the atmosphere, or are otherwise largely anoxic15. Altogether, our analyses suggest that atmospheric trace gases support a basal level of primary production and energy conservation, which in turn sustains the substantial biodiversity found in various cave ecosystems.
Methods
Site description and sampling
Samples were collected from four caves in Victoria Australia, belonging to two major lithologies, sedimentary limestone and igneous volcanic rock: 3M-3—Shades of death (S 37.40591; E 148.21114); 3M-49—Scrubby creek (S 37.44000; E 148.17000); 3H-11—Harman 1 (S 37.9075; E 141.9769); and 3H-9—Tunnel cave (S 38.05706; E 141.92000). Two limestone caves (M3 / M49) were sampled in April 2021, and two volcanic caves (H-11 / H-9) were sampled in November 2021. Samples were collected using sterile techniques along a distance transect traversing four sites consisting of the cave entrance at the surface (Entrance) and cave interior (Twilight, Transition and Deep), with the Deep sites representing the furthest point from the entrance (Fig. 1a). At each site (~25 m2), three sampling plots (~1 m2) were selected at random (Fig. 1a). At each plot, sediment-atmosphere fluxes were measured, and samples of sediments and biofilms were collected in triplicate for ex situ oxidation measurements, physicochemical analysis, and DNA extractions to perform qPCR and metagenomic sequencing. Additionally, opportunistic sampling of conspicuously hued biofilms, fungal spores, moonmilk57 and sediments were collected for metagenomic sequencing. An alternative design was used for Harman 1 cave, given the lack of bulk sediments on the cave floor and high prevalence of brown biofilms meant flux measurements could not be taken. Instead, sampling focused on ambient air measurements and the collection of sediments from three plots at the Entrance and biofilm from three plots at the Deep zone and two plots at the Transition site. All sampling occurred during daylight hours and dry weather conditions, gas samples and incubations were processed within 48 h of collection, and sediment samples for DNA extraction were initially stored at 5 °C for transport and frozen at −20 °C until DNA extraction. Sampling at Harman 1 and Tunnel cave were conducted under Parks Victoria Access Agreement AA-0000038. Sampling at Scrubby Creek cave and Shades of Death cave were privately arranged with owners at the Rimstone Cooperative Ltd.
Sediment physicochemical analysis
For sediment chemistry analysis, sediment samples from each plot were pooled to form one representative composite sample per site and sent to the Environmental Analysis Laboratory (EAL), Southern Cross University. In total, 37 separate soil physicochemical parameters were selected for analysis, based on commonly reported drivers of soil microbial composition globally. These included: Phosphorus (mg/kg P), Nitrate Nitrogen (mg/kg N), Ammonium Nitrogen (mg/kg N), Sulphur (mg/kg S), pH, EC (dS/m), Estimated Organic Matter (% OM), Exchangeable Calcium (mg/kg), Exchangeable Magnesium (mg/kg), Exchangeable Potassium (mg/kg), Exchangeable Sodium (mg/kg), Exchangeable Aluminium (mg/kg), Exchangeable Hydrogen (mg/kg), Effective Cation Exchange Capacity (ECEC) (cmol + /kg), Calcium (%), Magnesium (%), Potassium (%), Sodium - ESP (%), Aluminium (%), Hydrogen (%), Calcium/Magnesium Ratio, Zinc (mg/kg), Manganese (mg/kg), Iron (mg/kg), Copper (mg/kg), Boron (mg/kg), Silicon (mg/kg Si), Total Carbon (%), Total Nitrogen (%), Carbon/Nitrogen Ratio, Chloride Estimate (equiv. mg/kg), TOC (%), Moisture content (%), Gravel (%), Sand (%), Silt (%), Clay (%).
Community DNA extraction
Total community DNA was extracted from a total of 105 samples (94 environmental samples +11 controls) using 0.25 g of sediments and biofilms. Extractions were performed using the FastDNA Spin Kit for Soil according to the manufacturer’s instructions with an additional round of bead beating to improve DNA yields. Samples were eluted in DNase- and RNase-free UltraPure Water (ThermoFisher). Eleven sample-free negative controls were also extracted. Nucleic acid purity and yield were measured using a NanoDrop ND-1000 spectrophotometer and a Qubit Fluorometer 2.0.
Quantitative PCR
Quantitative polymerase chain reactions (qPCR) were used to estimate total bacterial and archaeal abundance. The 16S rRNA gene was amplified using the degenerate primer pair (515 F 5ʹ-GTGYCAGCMGCCGCGGTAA-3ʹ and 806 R 5ʹ-GGACTACNVGGGTWTCTAAT-3ʹ). A synthetic Escherichia coli 16S rRNA gene sequence in a pUC-like cloning vector (pMA plasmid; GeneArt, ThermoFisher Scientific) was used as a standard. PCR reactions were set up in each well of a 96-well plate using LightCycler 480 SYBR Green I Master Mix. Each sample was run in triplicate and standards in duplicate on a QuantStudio 7 Flex Instrument (Applied Biosystems). The qPCR conditions were as follows: pre-incubation at 95 °C for 3 min and 45 cycles of denaturation 95 °C for 30 s, annealing at 54 °C for 30 s, and extension at 72 °C for 24 s. 16S rRNA gene copy numbers were calculated based on a standard curve constructed by plotting average Ct values of a serial dilution of the plasmid-borne standard against their copy numbers.
Metagenome sequencing, assembly, and binning
Metagenomic shotgun libraries were prepared using the Nextera XT DNA Sample Preparation Kit (Illumina Inc., San Diego, CA, USA) and subject to paired-end sequencing (2 × 150 bp) on an Illumina NovaSeq6000 platform at the Australian Centre for Ecogenomics (ACE), University of Queensland. Raw metagenomic sequences were subjected to quality filtering using the BBDuk function of the BBTools v38.80 (https://sourceforge.net/projects/bbmap/); contaminating adaptors (k-mer size of 23 and hamming distance of 1), PhiX sequences (k-mer size of 31 and hamming distance of 1), and bases from 3’ ends with a Phred score below 20 were trimmed. After removing resultant reads with lengths shorter than 50 bp, 93% high-quality read pairs were retained for downstream analysis. Coassemblies were conducted according to cave type (limestone vs basalt) and sample nature (dividing by biofilm, sediment, and moonmilk) were conducted using quality-controlled reads in megahit (v1.2.9) using kmers of 27, 37, 47, 57, 67, 77, 87, 97, 107, 117 and 127. In addition, assemblies for each individual sample were performed using metaSPAdes (v3.15.5) with the same set of kmers58. Contigs from both coassemblies and individual assemblies with a minimum length of 500 bp were used for binning with MetaBAT2 (v2.12.1)59, MaxBin2 (v2.2.7)60 and CONCOCT (v1.1.0)61 to reconstruct MAGs, which is implemented in the “binning” module of MetaWRAP (v1.3.2)62. Subsequently, the ‘bin refinement’ modules of MetaWRAP and DAS tool63 were used in parallel to consolidate the MAGs from the three binners into one bin set for each assembly. A final MAG set was obtained by dereplicating MAGs from all assemblies at 99% average nucleotide identity using dRep “dereplicate” (v3.4.2)13,64. The completeness and contamination of the MAGs were estimated using CheckM2 (v0.1.3)65 “predict”. Taxonomy of each MAG was assigned using GTDB-Tk “classify_wf” (v2.3.0)66 against GTDB release 214. CoverM (v0.6.1) “genome” was used to calculate the relative abundance of each MAGs within each sample
Bacterial and archaeal community profiling
To profile bacterial, archaeal, and eukaryotic community composition based on the metagenomes, quality-filtered short reads encoding 16S rRNA genes were retrieved and assigned using PhyloFlash v.3.067. The function PhyloFlash.pl was given a -taxlevel 6 flag for bacteria/archaea specifying a taxonomic assignment down to genus level. Retrieved sequences were then clustered using PhyloFlash.pl into NTUs (nearest taxonomic units), mapped to the SILVA database (release 138)68, and filtered to exclude rare hits (min. read count ≥ 10). Alpha and beta diversity were calculated using R (version 4.1.0 (2021-05-18)—“Camp Pontanezen”) and the Phyloseq package69. First, reads were normalised using the coverage-based rarefaction and extrapolation method implemented in the R package iNEXT70. Coverage was calculated for each sample using the function phyloseq_coverage, followed by rarefaction of all reads, using the function phyloseq_coverage_raref with default parameters. Observed and estimated richness was calculated using the estimate_richness function specifying with “observed” and “chao1” flags. To calculate beta diversity, sample counts were first transformed to relative abundance and clustered using a Non-metric Multidimensional Scaling ordination via the ordinate function with flags “NMDS” and “Bray” function. A stress plot was used to determine linear (R2 = 0.83) and non-metric (R2 = 0.96) fit between ordination distance and dissimilarity (Bray-Curtis). The final stress of <0.2 (0.18) indicated a good representation in reduced dimension.
Metagenomic functional analysis and metabolic reconstruction
To estimate the metabolic capability of the sediment and biofilm communities, quality-filtered and unassembled short reads were searched against custom protein databases of representative metabolic markers using DIAMOND v.0.9.31 (query cover > 80%)71. Searches were carried out using all quality-filtered unassembled reads with lengths over 120 bp. The metabolic markers searched are involved in oxidative phosphorylation (AtpA), NADH oxidation (NuoF), aerobic respiration (CoxA, CcoN, CyoA, CydA), formate oxidation (FdhA), arsenic cycling (ARO, ArsC), selenium cycling (YgfK), sulphur cycling (AsrA, FCC, Sqr, DsrA, Sor, SoxB), nitrogen cycling (AmoA, HzsA, NifH, NarG, NapA, NirS, NirK, NrfA, NosZ, NxrA, NorB), iron cycling (Cyc2, MtrB, OmcB), reductive dehalogenation (RdhA), phototrophy (PsaA, PsbA, energy-converting microbial rhodopsin (RHO)), methane cycling (McrA, MmoX, PmoA), hydrogen cycling (large subunit of NiFe-, FeFe-, and Fe-hydrogenases), isoprene oxidation (IsoA), CO oxidation (CoxL, CooS), succinate oxidation (SdhA), fumarate reduction (FrdA), and carbon fixation (RbcL, AcsB, AclB, Mcr, HbsT, HbsC)72,73,74. Results were filtered based on an identity threshold of 50%, except for RHO (40%), NuoF, group 4 NiFe-hydrogenases, MmoX, FeFe-hydrogenases, CoxL, AmoA, NxrA, RbcL (all 60%), PsaA (80%), PsbA, IsoA, AtpA, ARO, YgfK (70%), and HbsT (75%). Read counts for each gene were normalised to reads per kilobase per million (RPKM) and average gene copy per organism as previously described 21,27,75. Proteins from each MAGs were predicted intrinsically in CheckM65. Best gene hits were filtered to retain only those either at least 40 amino acids in length or with at least 80% query or 80% subject coverage3. For predicted proteins, the same thresholds were used as above except for AtpA (60%), PsbA (60%), RdhA (45%), Cyc2 (35%) and RHO (30%)23. Metabolic reconstruction for the three most dominant hydrogenotrophs and methanotrophs was performed via annotation of relevant MAGs with DRAM76, based on the KEGG database. Pathways were identified through the KEGG pathway database and literature77. No MAGs encoded a GSH-linked pathway for formaldehyde oxidation, which is consequently omitted from the metabolism figure.
Methanotroph phylogenetic and metabolic analysis
GraftM v 0.1478 was used to ascertain gene phylogeny. PmoA and MmoX packages79 were used and updated with sequences published in ref. 21,27,80,81,82,83,84,85,86. HMM and DIAMOND (hmmsearch+diamond) search methods were applied, along with a conditional E-value (c-Evalue) threshold of 1e-1026,78. Multiple sequence alignment of protein sequences was performed with MAFFT v 7.49087, trimming at minimum 20% representation was performed with TrimAl v.1.4.188. Phylogenetic trees of protein sequences were generated with IQ-TREE v. 2.2.0.389 with the ultrafast bootstrap approximation option using 1000 iterations and enabling the ModelFinder option (Best-fit model: LG + F + R6). The trees were rerooted and grouped in ARB v. 6.0.390. Visualisation of protein trees was done in iTOL v691. The genome phylogenetic tree was generated by extracting the multiple sequence alignment of high-quality MAGs (completion >90% and contamination <5%) and known methanotrophs generated by GTDB-Tk (v2.1.1)66 against GTDB release 214. The tree was constructed with IQ-TREE v. 2.2.0.389 using model WAG + G and ultrafast bootstrap approximation option using 1000 iterations. The trees were rerooted and grouped in ARB v. 6.0.335 and visualised in iTOL v691. Annotation was performed using DRAM and the KEGG database, and pathways were identified through the KEGG pathway database and literature77. No MAGs encoded a GSH-linked pathway for formaldehyde oxidation, which is consequently omitted from the metabolism figure.
In situ sediment–atmosphere gas fluxes
In situ sediment-atmosphere fluxes of H2, CO and CH4 were measured using static flux chambers. The chamber consisted of a polyvinylchloride (PVC) pipe of 20 cm height and 15 cm diameter, with a threaded access cap. The cap was fitted with a gastight O-ring, two butyl rubber septa (one for air sampling and one for a thermometer), and an axial fan on the inside to promote internal mixing. At each plot, a PVC base collar of 10 cm height and 14.8 cm diameter was inserted ~5 cm into the sediment and left to equilibrate for ~15 min prior to sampling to reduce lateral gas fluxes. Once the chamber was fitted over the collar, an ambient air measurement was taken at the base of the collar, the cap was immediately closed, and the axial fan was started, and the next measurement was taken within 30 s. Additionally, ambient measurements were taken over a 30-min period. For each measurement, 25 mL of gas was collected using a gas-tight Terumo syringe fitted with a Luer Lock and Discofix three-way stopcock. Data quality checks and measurements by gas chromatography were conducted as described below. The temperature of the chamber, ambient air, and sediment were monitored throughout. Concentrations were then converted to nmol m−3 at ambient pressure and temperature using the ideal gas law. Atmospheric flux (Jatm; nmol m2 s−1) was calculated from the initial ambient conditions (t0) and rate of change at chamber deployment, using linear regression and an exponential model for each chamber measurement as previously described27. We calculated two separate flux phases “transient flux”, which includes the t0 ambient state, and the equilibrium flux, which excludes t0 to capture steady-state conditions. This was done to obtain an upper and lower bound flux range, to better reflect the unique conditions inside each cave, including shallow sediments, unique microclimate and absence of soil matrix.
Ex situ bulk oxidation rates
Microcosms were used to determine the capacity of sediment and biofim microbial communities to oxidise H2, CO and CH4 by gas chromatography. Overall, 44 biological replicates from four caves consisting of the Entrance (n = 12), Twilight (n = 9 plots), Transition (n = 11) and Deep (n = 12) zones were used. 5 g of sediment/brown biofilm (Harman 1) samples were placed in a 120 ml serum vial and incubated at 20 °C. The ambient air headspace was amended with H2, CO and CH4 (via a mixed gas cylinder containing 0.1 % v/v H2, CO and CH4 each in N2, BOC Australia) to give starting mixing ratios of approximately 10 parts per million (ppmv) for each gas. At each time interval, 2 ml of headspace gas was sampled using a gas-tight syringe and stored in a sealed 3 ml glass exetainer flushed prior with ultra-high purity N2 (99.999% pure, BOC Australia). A VICI gas chromatograph instrument with a pulsed discharge helium ionisation detector (model TGA-6791-W-4U-2, Valco Instruments Company Inc.) and an autosampler was used to measure gas concentrations as previously described92. The instrument was calibrated against ultra-pure H2, CO and CH4 standards. Calibration mixed gas (10.20 ppmv of H2, 10.10 ppmv of CH4, 9.95 ppmv of CO in N2, Air Liquide Australia) and pressurised air (Air Liquide Australia) with known trace gas concentrations were used as internal reference standards. Pooled heat-killed sediments for each triplicate (treated at 121 °C, 15 p.s.i. for 60 min) and empty vials (duplicate) were prepared as negative controls. All measurements were quality checked for peak anomalies, injection failures or leaks and excluded from downstream analysis. Reflecting the natural variation oxidation rates, measurements were halted once oxidation approached sub-atmospheric concentrations for individual samples. First order reaction rate constants were calculated by fitting an exponential model as determined by the lowest overall Root Mean Squared Error when compared to a linear model and normalised to average global concentrations for each gas.
Cell specific power calculations
The calculation was performed as previously described21,27, involving the determination of cell-specific power generated by the oxidation of H2, CO and CH4, measured as Gibbs energy per unit time per microbial cell. This was accomplished by factoring in the reaction rate for each gas, the Gibbs energy of the reaction, and the number of microbial cells involved. The Gibbs energy calculations considered the standard Gibbs energy, the reaction quotient, gas-phase compound activities, and sediment conditions. Measurement of reactant concentrations and reaction rates was conducted via gas chromatography. Estimates of microbial cells performing these reactions were calculated from 16S rRNA gene copy number datasets and abundances inferred from metagenomics datasets, focusing on trace gas-oxidising communities. This approach was compared to previous literature by recalculating the theoretical maximum population of trace-gas oxidising bacteria using modelled Gibbs energy and measured reaction rates.
Ammonium and sulphide oxidation
Oxic slurry experiments were undertaken to determine the oxidation rates of ammonium and sulphide of the cave sediments. Biotic and abiotic oxidation rates were distinguished using sterilised (autoclaved at 120 °C for 1 h) and non-sterilised cave sediments. Slurries containing 10 g sediment (wet weight) and 200 mL substrate amended ultrapure water (50 µM of either ammonium chloride or sodium sulphide) were prepared in 250 mL Schott bottles. The slurries were aerated for 5 min to ensure oxic conditions. The bottles containing the slurries were left uncapped but loosely covered with pre-combusted aluminium foil. The slurries were incubated in the dark and were mixed from time to time for the duration of the incubation period (up to 22 days). At each time point, 15 mL of samples were collected and filtered through 0.22 µm pore-sized filters (Sartorius Minisart syringe filter). Filtered samples were analysed for ammonium, nitrate, nitrite and sulphate concentrations using a Lachat Quickchem 8000 Flow Injection Analyzer (APHA standard method). Oxidation rates of ammonium were calculated using linear and non-linear regression of uptake over time as well as liner regression of nitrite and nitrate accumulation over time. Similarly, rates of sulphide oxidation were calculated from linear regression of sulphate increase over time.
Measurement of carbon isotope at natural abundance level (δ13C)
Stable isotope measurements were performed to probe possible pathways contributing to the formation of organic matter in caves. Approximately 1 g of sediment was dried at 60 °C for 24 h, pulverised and homogenised with a clean mortar and pestle. The pulverised sample was further dried at 60 °C overnight. Biofilm samples were first resuspended in sterile ultrapure water, pelleted by centrifugation at 13500 g for 2 min, and dried at 60 °C. Samples were weighed into 8 mm × 5 mm silver capsules, repeatedly treated with 10% hydrochloric acid (HCl) to remove inorganic carbon, and dried at 60 °C until no further effervescence was observed. All samples were analysed on an ANCA GSL2 elemental analyser interfaced to a Hydra 20-22 continuous flow isotope ratio mass spectrometer (CF-IRMS; Sercon Ltd., UK). The precision of the analysis was ±0.2‰. To ensure the accuracy of the isotopic values, internal standards (i.e. sucrose, gelatine and bream) were run concurrently with the samples. These internal standards have been calibrated against internationally recognised reference materials (i.e. USGS 40, USGS41 and IAEA C-6). In addition to δ13C, the instrumental analysis also yielded information on the percentage of organic carbon in each sample, with a quantification limit of 0.1 mg and a precision of 0.5 µg for carbon content.
Culture-based natural isotope preparation
To contextualise the δ13C of biomass originating from different carbon fixation pathways, we referred to the carbon isotopic fractionation effects observed in pure cultures. Our dataset includes previously reported literature values for autotrophic pure cultures52,93,94,95,96 and new analyses of two hydrogenotrophs (Cupriavidus necator H16, Hydrogenophaga pseudoflava 1034) and three methanotrophs (Methylocapsa gorgona MG08, Methylococcus capsulatus NCIB 11132, Methylosinus trichosporium OB3b). H. pseudoflava and C. necator grew autotrophically (CBB cycle) on H2 as follows. Pre-culture of both species was prepared by cultivating autotrophically on 20% H2 + 5% CO2 in minimal medium DSMZ 81 with an initial OD600 of 0.03 in gas-tight 120 ml serum vials sealed with rubber stoppers and aluminium caps at 30 °C with an agitation speed of 180 rpm. The main cultures were prepared by inoculating with an initial OD600 of 0.05 under identical conditions in a 1 L Schott bottle, sealed with rubber stoppers (GL 45 open-top cap) and screw caps (GL 45 cap open-top). M. gorgona, M. capsulatus, M. trichosporium were grown on CH4 as follows. NCIMB nitrate mineral salts (NMS) medium 131 was used for M. capsulatus and M. trichosporium while DSMZ medium 921 supplemented with 10 ml/L of 10x NMS salts and lanthanum (a final concentration of 1 µM) was used for M. gorgona. Exponential M. capsulatus and M. trichosporium cultures were grown in 1 L Schott bottles under 20% v/v CH4 in air at 37 °C and 30 °C, respectively, with shaking (200 rpm) for 7 days. M. gorgona cultures were prepared by inoculating 250 mL medium with 70 mL of an early stationary phase culture in 1 L glass bottles under 30% v/v CH4 in air at 20 °C without shaking. δ13C values of CO2 and CH4 were determined to be −10.6 ± 0.1‰ and −51.9 ± 1.5‰, respectively. Triplicate cultures were prepared for all strains. Following incubation, cells at late exponential or stationary phase were subject to centrifugation at 4 °C. Supernatants were discarded, and cell pellets were resuspended in 1x PBS buffer, followed by another round of centrifugation. Supernatants were discarded again, and the cell pellets were stored at −80 °C until freeze-drying. Cell pellets were lyophilised for 48 h and analysed for δ13C, using the same method as for the sediment and biofilm samples. One outlier was removed with an anomalously light signature of −67.40‰. The expected δ13C values of autotrophs reported in the literatures and current measurements when grown on atmospheric CO2 (δ13C average −8.5‰)97 and CH4 (δ13C average −47.2‰)98 were reported. A random forest model was then generated assessing the importance of key functional genes (pmoA, mcr, hbsT, aclB and rbcL) against depletion of biomass 13C stable isotope (δ13Corganic), using the randomForest function in R. Parameters were as follows: Number of variables randomly sampled as candidates at each split = 10, number of trees grown = 10,000 and sampling of cases was conducted with replacement.
14CO₂ fixation assay
A radiolabelled carbon dioxide (14CO2) incubation assay was used to measure the capacity of cave sediments/biofilms to mediate three processes: (i) dark CO2 assimilation/fixation, (ii) hydrogenotrophic CO2 fixation, and (iii) photosynthetic CO2 fixation as previously reported19,20,22,99. The sample was added with 50% v/w of sterile water containing 0.0833 μmol of NaH14CO3 (equivalent amount as 400 ppmv headspace 14CO2; Perkin Elmer, 56.6 mCi mmol−1) and the vial was immediately sealed with PTFE/silicone septum lid (Supelco, Sigma-Aldrich). The samples were then incubated for 96 h at either 20 °C under three conditions: (1) dark (covered in aluminium foil), (2) light (40 μmol photons m−2 s−1 under constant illumination), and (3) dark hydrogenotrophic condition (100 ppmv of headspace H2). For each condition and location, a technical triplicate and a heat-killed sediment control was prepared. Following incubation, each sample was treated with 2 ml of 1 M HCl to remove unfixed CO2 and the content was transferred to a 20ml scintillation vial. After overnight acidification treatment, the sample was added with an additional 1 ml of 1 M HCl and allowed to dry at 60 °C under a heat lamp. 20 ml of liquid scintillation cocktail (EcoLume™, MP Biomedical) was added to the dried sample and the signal of fixed 14C was measured on an automated liquid scintillation counter (Tri-Carb 2810 TR, Perkin Elmer) for 5 min. The instrument was regularly calibrated with 14C standards of known activity.
Phylogenetic analysis
The evolutionary relationships of maker genes, including methane oxidation (PmoA), hydrogen oxidation (NiFe), CO oxidation (CoxL), ammonia oxidation (AmoA), nitrite oxidation (NxrA) and carbon fixation (RbcL) were visualised by constructing phylogenetic trees of protein sequences. Briefly, protein sequences retrieved from the MAGs and metagenomic assembled reads by homology-based searches against a subset of reference sequences from a custom database100, and were aligned using MAFFT87 with default parameters, and subsequently trimmed using Trimal v1.2rev59 (“-gappyout”)88. The trimmed multiple sequence alignments were used to infer phylogenetic trees with the maximum likelihood method incorporating substitution model selection and 1000 bootstrap iterations implement in IQ-TREE2 v1.6.12 (“-alrt 1000 -B 1000 -m TEST”)89. Tree rendering was performed using the Python package ETE3101 combined with visualisation in iTOL91.
Methane carbon conversion efficiency
Theoretical estimation of carbon assimilation into microbial biomass was based on measured methane oxidation rates. Three seminal papers on methane carbon assimilation were used to calculate a constant conversion factor54,55,102. The percentage of methane carbon assimilation varied between 20 and 62%. By utilising the median for each range and computing the mean (40%), we determined the amount of methane carbon assimilated in units of nmol C cell−1 min−1 by multiplying the cell-specific methane oxidation rate by a constant fraction of 0.4. This measurement provides a quantifiable indicator of the role of aerobic methanotrophs in the carbon cycle. One drawback of this approach is that it does not consider variations in assimilation across different cells and variations in assimilation across different physicochemical conditions.
Data availability
The metagenomic data generated in this study have been deposited at NCBI Sequence Read Archive BioProject: PRJNA1048116: https://www.ncbi.nlm.nih.gov/bioproject/PRJNA1048116. All individual accession numbers with hyperlinks are provided in Supplementary Data 4a. Metagenome assembled genomes have been deposited in GenBank, with accession numbers and hyperlinks provided in Supplementary Data 4c. Processed metagenomic data are listed in Supplementary Data 4a–c. Raw data for all figures has been provided in the Source Data. Source data are provided with this paper.
References
Chen, Z. et al. The World Karst Aquifer Mapping project: concept, mapping procedure and map of Europe. Hydrogeol. J. 25, 771–785 (2017).
Palmer, A. N. Origin and morphology of limestone caves. Geol. Soc. Am. Bull. 103, 1–21 (1991).
Joseph, S. J., Hugenholtz, P., Sangwan, P., Osborne, C. A. & Janssen, P. H. Laboratory cultivation of widespread and previously uncultured soil bacteria. Appl. Environ. Microbiol. 69, 7210–7215 (2003).
De Mandal, S., Chatterjee, R. & Kumar, N. S. Dominant bacterial phyla in caves and their predicted functional roles in C and N cycle. BMC Microbiol. 17, 1–9 (2017).
Zhu, H. Z. et al. Bacteria and metabolic potential in karst caves revealed by intensive bacterial cultivation and genome assembly. Appl. Environ. Microbiol. 87, 1–17 (2021).
Zhu, H. Z. et al. Diversity, distribution and co-occurrence patterns of bacterial communities in a karst cave system. Front. Microbiol. 10, 1726 (2019).
Culver, D. C. & Pipan, T. Sources of energy in subterranean environments. The Biology of Caves and Other Subterranean Habitats 23–39 (Oxford University Press, 2009).
Tanalgo, K. C., Oliveira, H. F. M. & Hughes, A. C. Mapping global conservation priorities and habitat vulnerabilities for cave-dwelling bats in a changing world. Sci. Total Environ. 843, 156909 (2022).
Simon, K. S., Pipan, T. & Culver, D. C. A conceptual model of the flow and distribution of organic carbon in caves. J. Cave Karst Stud. 69, 279–284 (2007).
Huppop, K. How do cave animals cope with the food scarcity in caves? Ecosystems of the World 159–188 (2000).
Holmes, A. J. et al. Phylogenetic structure of unusual aquatic microbial formations in Nullarbor caves, Australia. Environ. Microbiol. 3, 256–264 (2001).
Ortiz, M. et al. Making a living while starving in the dark: metagenomic insights into the energy dynamics of a carbonate cave. ISME J. 8, 478–491 (2014).
Nicolosi, G. et al. Sulfidic Habitats in the Gypsum Karst System of Monte Conca (Italy) Host a Chemoautotrophically Supported Invertebrate Community. Int. J. Environ. Res. Public Health 19, 2671 (2022).
Sarbu, S. M., Kane, T. C. & Kinkle, B. K. A chemoautotrophically based cave ecosystem. Science 272, 1953–1954 (1996).
Chen, Y. et al. Life without light: microbial diversity and evidence of sulfur- and ammonium-based chemolithotrophy in Movile Cave. ISME J. 3, 1093–1104 (2009).
Kumaresan, D. et al. Aerobic proteobacterial methylotrophs in Movile Cave: genomic and metagenomic analyses. Microbiome 6, 1 (2018).
Read, K. J. H., Melim, L. A., Winter, A. S. & Northup, D. E. Bacterial diversity in vadose cave pools: evidence for isolated ecosystems. J. Cave Karst Stud. 83, 163–188 (2021).
Spanning et al. Methanotrophy by a mycobacterium species that dominates a cave microbial ecosystem. Nat. Microbiol 7, 2089–2100 (2022).
Ji, M. et al. Atmospheric trace gases support primary production in Antarctic desert surface soil. Nature 552, 400–403 (2017).
Bay, S. K. et al. Chemosynthetic and photosynthetic bacteria contribute differentially to primary production across a steep desert aridity gradient. ISME J. 15, 3339–3356 (2021).
Ortiz, M. et al. Multiple energy sources and metabolic strategies sustain microbial diversity in Antarctic desert soils. Proc. Natl. Acad. Sci. USA 118, e2025322118 (2021).
Ray, A. E. et al. Atmospheric chemosynthesis is phylogenetically and geographically widespread and contributes significantly to carbon fixation throughout cold deserts. ISME J. 16, 2547–2560 (2022).
Lappan, R. et al. Molecular hydrogen in seawater supports growth of diverse marine bacteria. Nat. Microbiol. 8, 581–595 (2023).
Greening, C. & Grinter, R. Microbial oxidation of atmospheric trace gases. Nat. Rev. Microbiol. 20, 513–528 (2022).
Dunfield, P. F., Knief, C. & Lipski, A. Diversity and activity of methanotrophic bacteria. Am. Soc. Microbiol. 69, 6703–6714 (2003).
Knief, C. Diversity and habitat preferences of cultivated and uncultivated aerobic methanotrophic bacteria evaluated based on pmoA as molecular marker. Front. Microbiol. 6, 1346 (2015).
Bay, S. K. et al. Trace gas oxidizers are widespread and active members of soil microbial communities. Nat. Microbiol. 6, 246–256 (2021).
Mattey, D. P. et al. Methane in underground air in Gibraltar karst. Earth Planet Sci. Lett. 374, 71–80 (2013).
Fernandez-Cortes, A. et al. Subterranean atmospheres may act as daily methane sinks. Nat. Commun. 6, 1–11 (2015).
McDonough, L. K. et al. Spatial variability of cave-air carbon dioxide and methane concentrations and isotopic compositions in a semi-arid karst environment. Environ. Earth Sci. 75, 700 (2016).
Webster, K. D., Mirza, A., Deli, J. M., Sauer, P. E. & Schimmelmann, A. Consumption of atmospheric methane in a limestone cave in Indiana. USA Chem. Geol. 443, 1–9 (2016).
Waring, C. L. et al. Seasonal total methane depletion in limestone caves. Sci. Rep. 7, 1–12 (2017).
Nguyễn-Thuỳ, D. et al. Subterranean microbial oxidation of atmospheric methane in cavernous tropical karst. Chem. Geol. 466, 229–238 (2017).
Cheng, X.-Y. et al. USC γ dominated community composition and cooccurrence network of methanotrophs and bacteria in subterranean Karst Caves. Microbiol. Spectr. 9, e0082021 (2021).
Cheng, X. et al. Niche differentiation of atmospheric methane-oxidizing bacteria and their community assembly in subsurface karst caves. Environ. Microbiol. Rep. 14, 886–896 (2022).
Zhao, R., Wang, H., Cheng, X., Yun, Y. & Qiu, X. Upland soil cluster γ dominates the methanotroph communities in the karst Heshang cave. FEMS Microbiol. Ecol. 94, 1–13 (2018).
Allenby, A. et al. Occurrence of methane-oxidizing bacteria and methanogenic archaea in earth’s cave systems—a metagenomic analysis. Front. Ecol. Evol. 10, 909865–909865 (2022).
Bendia, A. G. et al. Metagenome-assembled genomes from Monte Cristo Cave (Diamantina, Brazil) reveal prokaryotic lineages as functional models for life on Mars. Astrobiology 22, 293–312 (2022).
Greening, C., Berney, M., Hards, K., Cook, G. M. & Conrad, R. A soil actinobacterium scavenges atmospheric H2 using two membrane-associated, oxygen-dependent [NiFe] hydrogenases. Proc. Natl. Acad. Sci. USA 111, 4257–4261 (2014).
Grinter, R. et al. Structural basis for bacterial energy extraction from atmospheric hydrogen. Nature 615, 541–547 (2023).
Carini, P. Hazardous gases sustain microbes underfoot. Nat. Microbiol. 6, 145–146 (2021).
Tveit, A. T. et al. Simultaneous oxidation of atmospheric methane, carbon monoxide and hydrogen for bacterial growth. Microorganisms 9, 1–12 (2021).
Tveit, A. T. et al. Widespread soil bacterium that oxidizes atmospheric methane. Proc. Natl. Acad. Sci. USA 116, 8515–8524 (2019).
Zheng, Y. et al. Novel order-level lineage of ammonia-oxidizing archaea widespread in marine and terrestrial environments. ISME J. 18, wrad002 (2024).
Offre, P., Kerou, M., Spang, A. & Schleper, C. Variability of the transporter gene complement in ammonia-oxidizing archaea. Trends Microbiol. 22, 665–675 (2014).
Kits, K. D. et al. Kinetic analysis of a complete nitrifier reveals an oligotrophic lifestyle. Nature 549, 269–272 (2017).
Constant, P., Poissant, L. & Villemur, R. Tropospheric H2 budget and the response of its soil uptake under the changing environment. Sci. Total Environ. 407, 1809–1823 (2009).
Conrad, R. The global methane cycle: recent advances in understanding the microbial processes involved. Environ. Microbiol. Rep. 1, 285–292 (2009).
Novelli, P. C., Masarie, K. A., Tans, P. P. & Lang, P. M. Recent changes in atmospheric carbon monoxide. Science 263, 1587–1590 (1994).
Schmider, T. et al. Physiological basis for atmospheric methane oxidation and methanotrophic growth on air. Nat. Commun. 15, 4151 (2024).
Berg, I. A. et al. Autotrophic carbon fixation in archaea. Nat. Rev. Microbiol. 8, 447–460 (2010).
Garcia, A. K., Cavanaugh, C. M. & Kacar, B. The curious consistency of carbon biosignatures over billions of years of Earth-life coevolution. ISME J. 15, 2183–2194 (2021).
Roslev, P., Iversen, N. & Henriksen, K. Oxidation and assimilation of atmospheric methane by soil methane oxidizers. Appl. Environ. Microbiol. 63, 874–80 (1997).
Roslev, P. & Iversen, N. Radioactive fingerprinting of microorganisms that oxidize atmospheric methane in different soils. Appl. Environ. Microbiol. 65, 4064–4070 (1999).
Holmes, A. J. et al. Characterization of methanotrophic bacterial populations in soils showing atmospheric methane uptake. Appl. Environ. Microbiol. 65, 3312–3318 (1999).
Schimmelmann, A., Fernandez-Cortes, A., Cuezva, S., Streil, T. & Lennon, J. T. Radiolysis via radioactivity is not responsible for rapid methane oxidation in subterranean air. PLoS ONE 13, e0206506 (2018).
Park, S. et al. Microbial diversity in moonmilk of Baeg-Nyong Cave, Korean CZO. Front. Microbiol. 11, 1–13 (2020).
Nurk, S., Meleshko, D., Korobeynikov, A. & Pevzner, P. A. MetaSPAdes: a new versatile metagenomic assembler. Genome Res. 27, 824–834 (2017).
Kang, D. D. et al. MetaBAT 2: an adaptive binning algorithm for robust and efficient genome reconstruction from metagenome assemblies. PeerJ 2019, e7359 (2019).
Wu, Y.-W., Simmons, B. A. & Singer, S. W. MaxBin 2.0: an automated binning algorithm to recover genomes from multiple metagenomic datasets. Bioinformatics 32, 605–607 (2015).
Alneberg, J. et al. Binning metagenomic contigs by coverage and composition. Nat. Methods 11, 1144–1146 (2014).
Uritskiy, G. V., DiRuggiero, J. & Taylor, J. MetaWRAP—a flexible pipeline for genome-resolved metagenomic data analysis. Microbiome 6, 158 (2018).
Sieber, C. M. K. et al. Recovery of genomes from metagenomes via a dereplication, aggregation and scoring strategy. Nat. Microbiol. 3, 836–843 (2018).
Olm, M. R., Brown, C. T., Brooks, B. & Banfield, J. F. dRep: a tool for fast and accurate genomic comparisons that enables improved genome recovery from metagenomes through de-replication. ISME J. 11, 2864–2868 (2017).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015).
Chaumeil, P. A., Mussig, A. J., Hugenholtz, P. & Parks, D. H. GTDB-Tk v2: memory friendly classification with the genome taxonomy database. Bioinformatics 38, 5315–5316 (2022).
Gruber-Vodicka, H. R., Seah, B. K. B. & Pruesse, E. PhyloFlash: rapid small-subunit rRNA profiling and targeted assembly from metagenomes. mSystems 5, e00920–20 (2020).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2012).
McMurdie, P. J. & Holmes, S. Phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8, e61217 (2013).
Chao, A. et al. Measuring temporal change in alpha diversity: a framework integrating taxonomic, phylogenetic and functional diversity and the iNEXT.3D standardization. Methods Ecol. Evol. 12, 1926–1940 (2021).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 12, 59–60 (2015).
Greening, C. et al. Diverse hydrogen production and consumption pathways influence methane production in ruminants. ISME J. 13, 2617–2632 (2019).
Søndergaard, D., Pedersen, C. N. S. & Greening, C. HydDB: a web tool for hydrogenase classification and analysis. Sci. Rep. 6, 34212 (2016).
Cordero, P. R. F. et al. Atmospheric carbon monoxide oxidation is a widespread mechanism supporting microbial survival. ISME J. 13, 2868–2881 (2019).
Martinez-Perez, C. et al. Phylogenetically and functionally diverse microorganisms reside under the Ross Ice Shelf. Nat. Commun. 13, 117 (2022).
Shaffer, M. et al. DRAM for distilling microbial metabolism to automate the curation of microbiome function. Nucleic Acids Res. 48, 8883–8900 (2020).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Boyd, J. A., Woodcroft, B. J. & Tyson, G. W. GraftM: a tool for scalable, phylogenetically informed classification of genes within metagenomes. Nucleic Acids Res. 46, e59–e59 (2018).
Singleton, C. M. et al. Methanotrophy across a natural permafrost thaw environment. ISME J. 12, 2544–2558 (2018).
Putkinen, A. et al. New insight to the role of microbes in the methane exchange in trees: evidence from metagenomic sequencing. N. Phytol. 231, 524–536 (2021).
Vorobev, A. V. et al. Methyloferula stellata gen. nov., sp. nov., an acidophilic, obligately methanotrophic bacterium that possesses only a soluble methane monooxygenase. Int J. Syst. Evol. Microbiol. 61, 2456–2463 (2011).
Rgen Heyer, J., Galchenko, V. F., Dunfield, P. F. & Dunfield, P. Molecular phylogeny of type II methane-oxidizing bacteria isolated from various environments. Microbiology 148, 2831–2846 (2002).
Theisen, A. R. et al. Regulation of methane oxidation in the facultative methanotroph Methylocella silvestris BL2. Mol. Microbiol. 58, 682–692 (2005).
Dedysh, S. N. et al. Methylocella tundrae sp. nov., a novel methanotrophic bacterium from acidic tundra peatlands. Int J. Syst. Evol. Microbiol. 54, 151–156 (2004).
Edwards, C. R. et al. Draft genome sequence of uncultured upland soil cluster Gammaproteobacteria gives molecular insights into high-affinity methanotrophy. Genome Announc. 5, 10–1128 (2017).
Murphy, C. L. et al. Genomic analysis of the yet-uncultured Binatota reveals broad methylotrophic, alkane-degradation, and pigment production capacities. mBio 12, 10–1128 (2021).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Capella-Gutiérrez, S., Silla-Martínez, J. M. & Gabaldón, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Minh, B. Q. et al. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Ludwig, W. et al. ARB: a software environment for sequence data. Nucleic Acids Res. 32, 1363–1371 (2004).
Letunic, I. & Bork, P. Interactive Tree of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 49, W293–W296 (2021).
Islam, Z. F. et al. Two Chloroflexi classes independently evolved the ability to persist on atmospheric hydrogen and carbon monoxide. ISME J. 13, 1801–1813 (2019).
Ruby, E. G., Jannasch, H. W. & Deuser, W. G. Fractionation of stable carbon isotopes during chemoautotrophic growth of sulfur-oxidizing bacteria. Appl Environ. Microbiol. 53, 1940–1943 (1987).
Thomas, P. J. et al. Isotope discrimination by form IC RubisCO from Ralstonia eutropha and Rhodobacter sphaeroides, metabolically versatile members of ‘Proteobacteria’from aquatic and soil habitats. Environ. Microbiol. 21, 72–80 (2019).
Preuß, A., Schauder, R., Fuchs, G. & Stichler, W. Carbon isotope fractionation by autotrophic bacteria with three different C02 fixation pathways. Z. f.ür. Naturforsch. C. 44, 397–402 (1989).
House, C. H., Schopf, J. W. & Stetter, K. O. Carbon isotopic fractionation by Archaeans and other thermophilic prokaryotes. Org. Geochem 34, 345–356 (2003).
Graven, H. et al. Compiled records of carbon isotopes in atmospheric CO2 for historical simulations in CMIP6. Geosci. Model Dev. 10, 4405–4417 (2017).
Nisbet, E. G. et al. Rising atmospheric methane: 2007–2014 growth and isotopic shift. Glob. Biogeochem. Cycles 30, 1356–1370 (2016).
Jordaan, K. et al. Hydrogen-oxidizing bacteria are abundant in desert soils and strongly stimulated by hydration. mSystems 5, e01131–20 (2020).
Leung, P. M. & Greening, C. Greening lab metabolic marker gene databases. https://doi.org/10.26180/c.5230745.
Huerta-Cepas, J., Serra, F. & Bork, P. ETE 3: reconstruction, analysis, and visualization of phylogenomic data. Mol. Biol. Evol. 33, 1635–1638 (2016).
Roslev, P., Iversen, N. & Henriksen, K. Oxidation and assimilation of atmospheric methane by soil methane oxidizers. Appl Environ. Microbiol. 63, 874–880 (1997).
Raymond, O. L., Liu, S., Gallagher, R., Zhang, W. & Highet, L. M. Surface Geology of Australia 1:1 million scale dataset 2012 edition. Commonwealth of Australia (Geoscience Australia). https://doi.org/10.26186/5c636e559cbe1 (2012).
Tijhuis, L., Van Loosdrecht, M. C. M. & Heijnen, J. J. A thermodynamically based correlation for maintenance gibbs energy requirements in aerobic and anaerobic chemotrophic growth. Biotechnol. Bioeng. 42, 509–519 (1993).
LaRowe, D. E. & Amend, J. P. Power limits for microbial life. Front. Microbiol. 6, 718 (2015).
DeLong, J. P., Okie, J. G., Moses, M. E., Sibly, R. M. & Brown, J. H. Shifts in metabolic scaling, production, and efficiency across major evolutionary transitions of life. Proc. Natl. Acad. Sci. USA 107, 12941–12945 (2010).
Bradley, J. A. et al. Widespread energy limitation to life in global subseafloor sediments. Sci. Adv. 6, eaba0697 (2020).
Acknowledgements
This research was carried out as part of the Australian Research Council (ARC) SRIEAS grant Securing Antarctica’s Environmental Future (SR200100005; awarded to S.L.C., C.G., M.A.M.) and a Monash University Faculty of Science Strategic Uplift Seed Grant (awarded to S.K.B., P.C., W.W.W.). S.K.B. and R.L. are supported by an ARC Discovery Early Career Research award (DE230101346; DE230100542). C.G. is supported by an NHMRC EL2 Fellowship (APP1178715) and ARC Future Fellowship (FT240100502). C.M.S. is supported by a Novo Nordisk Foundation Postdoctoral Fellowship grant (NNF20OC0065005). K.S.K. is supported by the Dark Science project (Villum Foundation). G.N. and P.M.L. acknowledge the Early Career Postdoctoral Fellowship (ECPF23-8566329039 & ECPF23-1113137961) awarded by Faculty of Medicine, Nursing and Health Science (FMNHS) at Monash University. A.T.T. and O.S. were supported by the Research Council of Norway (projects Living on Air, 315129 and Harvesting Energy from Air, 347122). Z.F. is supported by Monash University-China Scholarship Council Joint Scholarship (CSC202308240008). This study used the MASSIVE M3 and MonARCH supercomputing infrastructure. We are grateful of Tess Hutchinson and Michaela Wawryk for technical support.
Author information
Authors and Affiliations
Contributions
S.K.B., C.G. and S.L.C. conceived this study. S.K.B. planned and led field work, designed and led experiments and analyzed data. Different authors were responsible for performing fieldwork (S.K.B., R.L., T.F., P.M.L., S.W. and N.W.), field logistics (S.K.B., S.W., T.F. and N.W.), DNA extraction and qPCR (S.K.B., R.L. and T.J.), gas chromatography (S.K.B.), nitrification and sulphide oxidation assays (W.W.W., V.E. and P.M.L.C.), pure culture preparation (N.A., S.J., O.S. and A.T.), carbon stable isotope measurement (W.W.W., P.M.L., Z.F., V.E. and P.M.L.C.), 14C radioisotope tracing (S.K.B.), spatial analysis and mapping (D.C., T.F., S.W. and M.A.M.), metagenomic community and metabolic analysis (S.K.B., G.N., C.G. and P.M.L.), MAG construction and annotation (G.N.), and metabolic reconstruction (S.H., K.S.K., N.K., M.K, C.M.S., S.K.B., G.N., C.G. and P.M.L.). S.L.C. and C.G. provided most resourcing, supervision and funding. S.K.B., C.G., S.L.C. and P.M.L. wrote the paper with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Tillmann Lueders, and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Bay, S.K., Ni, G., Lappan, R. et al. Microbial aerotrophy enables continuous primary production in diverse cave ecosystems. Nat Commun 16, 10295 (2025). https://doi.org/10.1038/s41467-025-65209-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65209-w
This article is cited by
-
Metabolically flexible microorganisms rapidly establish glacial foreland ecosystems
Nature Communications (2025)