Abstract
Alzheimer’s disease, the leading cause of dementia in the elderly, is a neurodegenerative disorder that has been studied to uncover therapeutic pathways through its molecular and cellular hallmarks. Here, we present a comprehensive investigation of cellular heterogeneity from the temporal cortex region of 40 individuals, comprising healthy donors and individuals with differing AD pathology. Using single-nucleus transcriptomic analysis of 430,271 nuclei from both gray and white matter of these individuals, we identified cell type-specific subclusters in both neuronal and glial cell types with varying degrees of association with AD pathology. We extended this analysis by performing multiplexed in situ hybridization using the CARTANA platform, capturing 155 genes in 13 individuals with differing tau pathology. We not only replicated snRNA data key findings from our spatial data analysis but also identified a set of cell type-specific genes that show selective enrichment or depletion near pathological inclusions.
Similar content being viewed by others
Introduction
The two pathological hallmarks of Alzheimer’s Disease (AD) - amyloid-beta plaques and neurofibrillary tangles - show accumulation and spread that are associated with the severity of symptoms1,2,3. Although the complex pathophysiology of AD remains poorly understood, recent transcriptomic and epigenomic analyses have revealed that post-mortem human AD brain tissue exhibits downregulation of genes associated with neuronal function4,5 and upregulation of the genes involved in the innate immune response6,7. However, not many studies have looked systematically at the spatial distribution of gene expression and its relationship to neuropathology8. Thus, our overall understanding of cell type heterogeneity and compositional changes during pathological accumulation is still under-explored, hindering our ability to understand the biological processes underlying AD9.
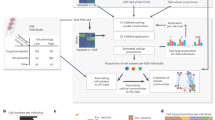
While the pathogenesis of Alzheimer’s disease has been extensively studied, the predominant focus has traditionally been on gray matter alterations, overlooking the essential role that white matter plays in neurological health. Recognizing the involvement of the temporal cortex in neurodegenerative diseases is pivotal for developing precise interventions and therapies. Here, we investigate the cellular heterogeneity in the white and gray matter of the temporal cortex (TC) region of the right hemisphere of 40 individuals taken from the Netherlands Brain Bank (NBB), comprising healthy donors and individuals at varying Braak stages. We generated 430,271 single-nucleus RNA profiles from both white and gray matter samples, which were collected separately from the midsection of the mid-temporal cortex (mid-TC) in 40 individuals. Each participant provided paired white and gray matter samples, allowing for a comprehensive analysis to identify molecular associations corresponding to varying degrees of AD pathology (Supplementary Data 1). To further expand our analysis of transcriptomic signatures within TC, we incorporated data from six previously published studies, including 888,784 total nuclei (Supplementary Data 2). This comprehensive analysis with different brain regions such as the entorhinal cortex (EC)10,11, prefrontal cortex (PFC)5,12,13,14, and superior frontal gyrus (SFG)10 allowed us to replicate a subset of the cell type associations. Finally, we explored the spatial distribution of a subset of key cell type specific genes in 13 individuals through the CARTANA targeted in situ sequencing (ISS) platform15 (Fig. 1a). Starting with a panel of 155 genes, we investigated spatial enrichment or depletion with Braak stage (validating our snRNA-seq results), as well as enrichment or depletion near specific pathological inclusions identified on the same tissue sections.
a Overview of the experimental scheme (created in BioRender. Gaur, P. (2025)). b UMAP embedding of 430,271 single-nucleus RNA profiles from the TC brain region of 40 individuals; colored by cell type. c Dot plot of the canonical markers distinguishing 8 major cell types with different levels of expression (color) and percentage of expressing nuclei (dot size) across 430,271 TC nuclei. d UMAP embedding of 430,271 single-nucleus RNA profiles from the TC brain region colored by different subclusters. e Dot plot of differentially expressed (DE) genes distinguishing neuronal cell subgroups, showing different levels of expression and percentage of expressing nuclei across different subclusters, with numbers of nuclei (n) shown in the right. f Dot plot of DE genes distinguishing glial cell subgroups, showing different levels of expression and percentage of expressing nuclei across different subclusters, with numbers of nuclei (n) shown in the right. Source data are provided as a Source Data file.
Results
Transcriptional states of CNS cell types in the temporal cortex gray and white matter of the aged human brain
We profiled tissues from the temporal cortex region to investigate the association of pathology with different sub cell types proportions. First, we profiled the transcriptome of 80 post-mortem human brain samples (40 individuals; gray and white matter) encompassing controls and individuals with varying degrees of AD pathology (Supplementary Fig. 1a, b, Supplementary Data 1). We retained 430,271 nuclei for AD trait association analysis after quality control and cluster identification. Nuclei were segregated into eight major cell types: glutamatergic neurons, GABAergic neurons, microglia, astrocytes, oligodendrocytes, oligodendrocyte precursor cells (OPCs), endothelial cells, and pericytes (Fig. 1b and Supplementary Data 3). Nuclei in these broad cell types showed specific expression of canonical marker genes such as SLC17A6 and SLC17A7 for glutamatergic neurons, GAD2 and SLC32A1 for GABAergic neurons, AQP4, FGFR3 and ALDH1L1 for astrocytes, MOG for oligodendrocytes, PDGFRA for OPCs, TMEM119, P2RY12, AIF1 and C1QA for microglia, CLDN5 for endothelial cells, and RGS5 for pericytes (Fig. 1c, e, f). Each major cell type was then subclustered independently, and subclusters were named based on a combination of top differentially expressed markers (“Methods”, Fig. 1d, e, f). We identified 20 subpopulations of glutamatergic neurons, 22 subpopulations of GABAergic neurons, 5 of astrocytes, 4 of oligodendrocytes, 3 of OPCs, 3 of endothelial cells, 6 of pericytes, and 7 of microglia (Supplementary Data 4). One microglial cluster (RIPOR2 + ) is likely enriched for monocytes, based on low expression of TMEM119, P2RY12, and high expression of LYZ (Fig. 1f).
Neuronal and glial subpopulations with lower prevalence in gray matter in late Braak stages
Our key analysis was to associate differences in subpopulation composition within each major cell type with specific AD phenotypes. We selected neurofibrillary tangle (NFT) spread as the AD phenotype of interest, for reasons described below3,12. In particular, we used the Braak staging paradigm, which assigns a stage (from 1 to 6) based on the overall spread of tau pathology in an individual, with early stages (Braak 1/2) indicating NFT accumulation primarily in the entorhinal cortex and hippocampus, and later stages (Braak 3/4/5/6) describing individuals with tau pathology in both early and late-affected (cortical) regions. Individuals at Braak stage 3 and beyond tend to show NFT accumulation in the temporal cortex, a fact we verified with staining for tau in 13 of our donors as part of our CARTANA experiments. Although amyloid plaque aggregation is the other hallmark pathology of AD, we selected tau as our proxy (through the Braak staging paradigm) variable for disease progression analysis, given that the spread of tangles in the brain is more stereotyped, and because global post-mortem tau is more strongly correlated with cognitive impairment than post-mortem amyloid-beta16. With Braak stage as our primary pathology variable for disease progression, we then examined cell subpopulations that showed coordinated changes with this trait of interest.
In gray matter (GM), we identified multiple subsets of neurons showing lower proportions in individuals at late Braak stages (Fig. 2a, b, Supplementary Data 5). Out of 5 total RORB+ glutamatergic subpopulations, two were found to be less prevalent in later Braak stages compared to earlier stages; interestingly, both of these RORB+ subpopulations express IL1RAPL2. This aligns with recent findings where RORB has been recognized as an indicative marker of selectively vulnerable excitatory neurons in the entorhinal cortex10, although we find that not all RORB+ subpopulations are equally vulnerable at late Braak stages. We also observed a glutamatergic subset with high levels of SPARCL1 was lower in the GM in donors at advanced Braak stages17. Other subpopulations with lower proportions at later Braak stages included those marked by COL5A2 (putatively in Layer 4) and GLIS3 (putatively in Layer 3). Prior microarray-based studies have found COL5A2 to be downregulated in AD18,19, and GLIS3 has been implicated in a genome-wide significant association with cerebrospinal fluid tau and phospho-tau levels in the context of a specific genetic variant20. Finally, we also observed lower proportions of GNAL+ and TLL1+ glutamatergic subpopulations at Braak stages 5/6. The glutamatergic subpopulation expressing GNAL has been reported to have an alternative splicing association with AD21, and TLL1 has been reported as one of the five genes (Tshz2, Gm12695, St3gal1, Isx and Tll1) implicated in Aβ processing and is affected in Tg-5xFAD mice treated with REMFS22.
a Differential abundance of statistically significant neuronal subpopulations with respect to AD pathology indicated by varying Braak stages in GM and WM obtained using ANCOM-BC, which applies a log-linear regression model to assess proportion differences. Two-sided Z-tests were performed, and p-values were adjusted for multiple comparisons using the Benjamini-Hochberg method. The x-axis represents the estimated log fold-change in the abundance of a subpopulation between different Braak stages, and the y-axis represents subpopulations that were found to be statistically significant after multiple testing corrections. b Box plots of all neuronal subpopulations shown in panel a, representing the distribution of nuclei in each subpopulation with respect to their concerned major cell type (GM and WM samples from n = 36 individuals). c Differential abundance of statistically significant glial subpopulations with respect to AD pathology indicated by varying Braak stages. X and Y axis representations and statistical approach are consistent with panel a. d Box plots of all glial subpopulations shown in panel c, representing the distribution of nuclei in each subpopulation with respect to their concerned major cell type (GM and WM samples from n = 36 individuals). Boxplot: center line = median; box = upper and lower quartiles; whiskers = values within ± 1.5*IQR (interquartile range); points = outliers beyond whiskers. e Differential abundance of statistically significant subpopulations with respect to sex (male). X and Y axis representations and statistical approach are consistent with panel a. Source data are provided as a Source Data file.
With respect to GABAergic neurons in GM, we found multiple subgroups with lower proportions at late Braak stages. These included parvalbumin (PVALB) neurons (Fig. 2a), which have been reported to be selectively depleted in the frontal cortex of Alzheimer’s disease mouse models23. SPARCL1+ GABAergic neurons also showed depleted proportions in GM. Notably, SPARCL1 has been reported to have altered levels in the cerebrospinal fluid (CSF) of AD patients, and it is conceivable that alterations in the regulation of SPARCL1 may play a significant role in the development of AD24,25. On the glial side, we found associations between microglial subsignatures with later Braak stages. The strongest signal was in RPL19+ microglia, which were significantly lower in intermediate and late Braak stages in GM, possibly reflecting the loss of a protective type of microglia (Fig. 2c, d).
Specific neuronal and glial subclusters selectively enriched in gray matter at late Braak stages
Our analysis above highlighted multiple neuronal groups with lower proportions at late Braak stages. However, since these proportions are measured as a fraction of the total glutamatergic or GABAergic neurons, they are offset by concomitantly higher proportions of other neuronal subtypes; these latter groups are putatively less vulnerable to disease pathology and are interesting for further study. For example, we found that a few glutamatergic neuron populations from the deep layers of the cortex (L5 − 6) in GM, marked by THEMIS and POSTN, are overrepresented at later Braak stages, which suggests that this subgroup could be relatively more resistant to increasing tau burden. In addition, a unique SST+/THSD7B+ GABAergic subpopulation marked by Thyrotropin Releasing Hormone Degrading Enzyme (TRHDE) was also enriched in Braak stage 5/6 (Fig. 2a, b). Among glial cells, OPCs specific to heat shock protein were higher in GM in individuals with more advanced AD pathology. A similar pattern was observed in MGP+ pericytes and TNC+ astrocytes, whose proportions were higher in individuals at later Braak stages (Fig. 2c, d and Supplementary Data 5).
Neuronal and glial subclusters selectively enriched and depleted in white matter at late Braak stages
Although the vast majority of cells in brain white matter are oligodendrocytes, we found changes in glial subpopulations, and even some neuronal populations, associated with Braak stage in TC-associated white matter (WM). Endothelial cells marked by TSHZ2 were present at higher proportions in donors at advanced Braak stages (Fig. 2c), as were OPCs with high expression of heat-shock proteins; this latter finding resembles the same finding in GM. Finally, RPL19+ microglia and TAGLN+ pericytes were lower in individuals at intermediate and advanced Braak stages, respectively; RPL19+ microglia were one of the cell signatures showing the same trend in WM and GM (Fig. 2c and Supplementary Data 5).
Despite the sparsity of neuronal nuclei in the WM, we observed statistically significant differences in neuronal subgroups in the WM at different Braak stages. Similar to our findings in GM, PVALB+ /TMEM132C+ and PVALB+ /SPARCL1+ subpopulations were lower in WM in donors at advanced Braak stages (Fig. 2a, b and Supplementary Data 5). By contrast, RORB + /IL1RAPL2+ glutamatergic subpopulations showed a different pattern in WM, exhibiting higher proportions at intermediate Braak stages 2/4 (Fig. 2a, b and Supplementary Data 5). Finally, GABAergic neurons expressing LAMP5 - a gene previously implicated in dysfunction in Alzheimer’s disease26 and CHST9, a drug metabolism-related gene in AD27, showed lower prevalence in WM in late Braak stages (5-6) (Fig. 2a, b and Supplementary Data5). Overall, the changes in WM appear to be less distinct than those found in GM and are mostly restricted to these rarer cell populations.
Consistency of subcluster associations with Braak stage across multiple brain regions and studies
In order to assess the generalizability of our results, we integrated our snRNA-Seq data with other publicly available datasets spanning various brain regions. We incorporated entorhinal cortex10,11, prefrontal cortex5,12,13, superior frontal gyrus10 and deep white matter from prefrontal cortex14 to examine the single-nuc RNA-seq profiles (Supplementary Fig. 2a, b and Supplementary Data 2). After the removal of potential doublets from 959,237 nuclei (“Methods”), we retained a total of 888,784 nuclei for trait association analysis (Supplementary Data 2, 4). We then investigated which AD trait associations in the TC were found across multiple brain regions (Supplementary Data 6). We found that THEMIS + /POSTN+ deep layer glutamatergic neurons were consistently overrepresented in late Braak stages, suggesting that these deep layer neurons may be generally resistant to AD pathology across the cortex (Supplementary Fig. 3f, g). Conversely, PVALB+ /TMEM132C+ GABAergic neurons were lower at advanced Braak stages in all brain regions we examined (Supplementary Fig. 3g). On the glial side, TAGLN+ pericytes and RPL19+ microglia were lower in Braak stages 5/6 in multiple cortical regions, whereas HSPA1A+ OPCs were enriched in donors at these Braak stages (Supplementary Fig. 3j). To further investigate these signatures with more recent datasets, we performed trait association analyses in the middle temporal gyrus (MTG) from 84 individuals in Seattle Alzheimer’s Disease Brain Cell Atlas (SEA-AD) consortium28. We mapped our TC subclusters onto the MTG dataset using the Cell Type Mapper algorithm29 (Supplementary Data 7) and observed a high degree of similarity between the two datasets (Supplementary Fig. 3a, b). Using our TC subcluster labels transferred to MTG nuclei, we then conducted the trait association analysis on 84 individuals (Supplementary Fig. 3e, h and Supplementary Data 8) and found that the potentially resilient THEMIS+ /POSTN+ glutamatergic neurons were consistently overrepresented in late Braak stages of AD in the MTG as well (Supplementary Fig. 3e). To further validate these findings, we included an additional study of the prefrontal cortex (PFC) from the ROSMAP cohort (424 individuals)30 and compared the similarity between our TC subclusters and those identified in the larger study (Supplementary Fig. 3c, d and Supplementary Data 9). We then re-investigated all PFC data from multiple studies, ran the trait association analysis (Supplementary Fig. 3f, i and Supplementary Data 10) and again found that THEMIS+ /POSTN+ glutamatergic neurons were enriched in advanced AD pathology (Supplementary Fig. 3f). Along with this, TAGLN+ pericytes were also lower in advanced Braak stages (Supplementary Fig. 3i) in PFC as well as several other significant signatures that merit further investigation (Supplementary Fig. 3 and Supplementary Data 8, 10).
Differential expression analysis identifies additional cell type-specific genes in late stage AD pathology in the TC
Whereas cluster proportion analyses can identify differential vulnerability and resistance, disease-associated cellular signatures may be obscured when cluster boundaries are not discrete. To identify additional signatures potentially missed by cluster-based analyses, we then looked for pathological changes in gene expression levels in each broad cell type using a pseudobulk approach (“Methods”). As our experimental design included two samples per individual (GM matter from the TC + associated WM), we used a mixed-effect negative binomial model to test for changes in gene expression related to AD pathology (“Methods”). We found a few expression changes in the early stages of AD (Braak 2-4 vs Braak 0-1), with only 17 genes differentially expressed (FDR-adjusted p-value < 0.05), and all of these were in GABAergic neurons, oligodendrocytes, and OPCs (Fig. 3a and Supplementary Data 11). As our statistical model included an interaction term between Braak stage and tissue, we also tested whether any genes were differentially affected by early AD pathology in gray vs white matter for each broad cell type. We found only a few genes that were affected at early Braak stage differentially between GM and WM (10 genes, FDR-adjusted p-value < 0.05, Supplementary Data11).
a Number of differentially expressed genes per cell type when compared across Braak stages using a false discovery rate (FDR) of 5%. b Differentially expressed genes with the largest log fold change for each cell type at late Braak stage (5-6) vs early Braak stage (0-1). ‘*’ indicates statistical significance at a false discovery rate (FDR) of < 5%, based on a two-sided Wald test. c Pathway enrichment of differentially expressed genes, examining pathways specifically associated with AD genes. The color of the dot reflects NES, which refers to the Normalized Enrichment Score (NES) for Gene set enrichment analysis, whereas the size refers to the absolute value of NES. d LOEUF score (a measure of gene constraint) for differentially expressed genes in glutamatergic neurons (GM and WM samples from n = 39 individuals). p-values are derived from two-sided Wilcoxon rank-sum tests. No multiple testing correction was applied. e Pathway enrichment (10% FDR) of differentially expressed genes in pathways specifically associated with AD genes (20% FDR). The color and size of the dots are the same as in c. f Expression of NPTX2 in Glutamatergic neurons at different Braak stages. g Expression of FCER1G in Microglia at different Braak stages and in GM vs WM. h Expression of APOE in endothelial cells at different Braak stages and in GM vs WM. For (f–h), Boxplot: center line = median; box = upper and lower quartiles; whiskers = values within ± 1.5*IQR; points = individual data with outliers beyond whiskers. GM and WM samples from n = 39 individuals with varying Braak stages. Differential expression was assessed using a negative binomial mixed model (glmmTMB) fitted separately for each cell type. Two-sided Wald tests were performed, and p-values were adjusted for multiple comparisons using the Benjamini-Hochberg method. i Differentially expressed genes affected in a different manner between GM and WM that belong to pathways associated with AD genes (5% FDR, denoted by ‘*’). Source data are provided as a Source Data file.
However, we found a large number of differentially expressed genes (1230 at FDR-adjusted p-value < 0.05) in late Braak stages (Braak 5-6 vs Braak 0-1), which is not unexpected given that cortical pathologies in the TC are only observed at later Braak stages in AD. Interestingly, the majority of differentially expressed genes were specific to individual cell types (~ 90%, 1102/1230). This could be due to the genes being exclusively expressed in a particular cell type, indicating that the cellular response to AD may be cell-type-specific, or because the genes are expressed in multiple cell types but exhibit differential expression in only one (Supplementary Data11). For example, NPTX2 (Fig. 3f), a putative CSF prognostic AD biomarker31, was significantly lower in late Braak stages in glutamatergic neurons. Rerunning our negative binomial mixed model, which had the capability to test for differential expression in AD pathology in gray vs white matter by incorporating an interaction term between Braak stage and tissue, we found that APOE, a well-known AD risk gene belonging to pathways related to amyloid formation32, is differentially expressed in endothelial cells between gray and white matter cells (Fig. 3h). Upon examining the APOE levels in endothelial cells, we observed that the difference between controls and cases was more pronounced in the GM compared to the WM, though APOE levels were downregulated in both GM and WM. Specifically, the magnitude of the decrease was greater in the GM than in the WM. Although APOE expression is usually associated with microglia and astrocytes, this differential signal in endothelial cells suggests compartment (gray and white matter)-specific alterations in the vasculature in AD. We also found that 644 genes (FDR adjusted p-value < 0.05) were affected differently by late stage pathology in gray vs white matter (Fig. 3a). For example, FCER1G, a gene associated with AD genetic risk14 was affected differentially in late Braak stages in gray (lower) vs white matter (higher) (Fig. 3g), suggesting that transcriptional changes in AD can differ between different brain tissue types. ABCA7, another AD risk gene33, was significantly higher in GM microglia than in WM microglia (Supplementary Data 11). Altogether, these results indicate that gene expression changes related to AD pathology are both cell-type specific and tissue specific, suggesting that cross-tissue differential expression analysis at the cell type level will be necessary to fully capture the transcriptional changes associated with AD.
We also tested whether the LOEUF (loss of function observed/expected upper bound fraction) scores (a measure of gene constraint)34 were significantly different for our differentially expressed genes. Gene constraint refers to the degree to which evolutionary pressures restrict genetic variation within a gene, typically due to its functional importance. In other words, it reflects how much a gene can tolerate changes (mutations) without causing harm or disrupting normal biological function. Highly constrained genes are those that tolerate little to no variation because changes in these genes are likely to be deleterious and negatively impact the organism’s fitness. Interestingly, upregulated genes in late Braak stages (5-6) in glutamatergic neurons, as well as genes differentially affected between gray and white matter in glutamatergic neurons, were significantly more constrained than non-differentially expressed genes (Fig. 3d), suggesting that differentially expressed genes in glutamatergic neurons upregulate essential functions in late Braak stages and might also play an important developmental role. We speculate that these genes could be important genes for cell survival and that glutamatergic neurons upregulate these genes as a survival mechanism in an extreme environment.
Other than these, the differentially expressed genes (Braak 5-6 vs Braak 0-1) with the largest effect sizes included several interesting genes (Fig. 3b). For example, FOXJ1, a key transcription factor involved in the production of motile cilia35, was significantly upregulated in astrocytes. In addition, PVALB was significantly downregulated in inhibitory neurons, which is consistent with our proportion analysis (Fig. 2a). We also found that RELN, which inhibits tau phosphorylation36, was strongly upregulated in oligodendrocytes. We also present volcano plots depicting other significant differentially expressed genes associated with Braak stage progression (Supplementary Fig. 4a), as well as genes exhibiting tissue-specific expression patterns (Supplementary Fig. 4b and Supplementary Data11).
Pathway enrichment analysis of these differential genes (Braak 5-6 vs Braak 0-1) highlighted several associations with Alzheimer’s disease (“Methods”). For example, genes involved in the negative regulation of amyloid fibril formation were lower at late Braak stages in inhibitory neurons, while ‘phospholipid binding’ and ‘cellular response to lipids’ were higher at late Braak stages in microglia and OPCs, respectively (Fig. 3c). The genes with higher expression GM astrocytes in late stage pathology were enriched in endocytosis and kinase binding processes (Fig. 3e), while the analogous genes in GM glutamatergic neurons were enriched in the regulation of vesicle transport. Genes with higher expression in WM microglia (but not GM microglia) at later Braak stages were enriched in cellular response to lipoprotein particle stimuli, while the analogous genes in WM oligodendrocytes showed enrichment for the regulation of amyloid-beta formation. Finally, we looked for differentially expressed genes that were affected in a different manner between gray and white matter in late-stage pathology and belonged to pathways specifically associated with AD genes (Fig. 3i).
Characterizing susceptible and resilient cell signatures with high-resolution spatial transcriptomics and pathological staining
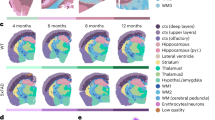
To complement and validate our analysis of dissociated nuclei from tissue, we next examined intact tissue using the CARTANA15 probe-based in situ sequencing platform with 155 genes on 13 tissue samples from our donors (Supplementary Data 12). In addition, we also performed staining on the same tissue sections to localize amyloid and tau pathological inclusions. The workflow included detection of β-amyloid and tau pathology, classification of 100 × 100 pixel (1056 μm2) ‘tiles’ based on the quantitative plaque/tangle density, segmentation of white and gray matter, labeling of cells, and quantification of expression of 155 genes. These genes were selected to identify all major cell classes in the cortical gray and temporal cortex-associated white matter, as well as a combinatorial set of genes to identify the subclusters from our single-nucleus RNA-seq analysis. Because of probe design challenges in the system, alternative markers for several subclusters had to be selected, instead of the optimal markers based on single-nucleus RNA-seq. Nevertheless, these alternative markers still show clear discrimination among the subtypes (Supplementary Fig. 5); for example, the THEMIS + /POSTN+ glutamatergic subpopulation could be marked alternatively using the combination of PPP1R1A and CC2D1B (Supplementary Fig. 5d). This ultimately generated a resource with cells assigned to their major class, tissue coordinates of every pathological inclusion, and annotation of cells as present in white or gray matter (“Methods”, Fig. 4a, b, Supplementary Figs. 5, 6 and Supplementary Data 13). Whereas the majority of cells were assigned to a ‘major class’ label (Supplementary Data 13), the sparse detection of many subcluster-specific genes meant that we were not able to definitively assign every cell to an snRNA-seq-derived subcluster. Thus, instead of calculating cell subcluster counts directly in the tissue, we modeled the expression of subtype-specific genes in tissue by fitting a linear mixed model that investigated cell type-specific association of these marker genes and pathology.
a A visual representation of the spatial distribution of cells in the specified samples (from control to progressing AD pathology), with a background grid for all samples based on cell coordinates, while different layers highlighting cells expressing specific markers (Oligodendrocytes = light blue to dark blue, RORB = green and LAMP5 = orange). GM and WM are well distinguished with plaque deposition sites (blue squares) on tissue. b Same as a except for tangles deposition instead of plaques in magenta squares. c Boxplots of the genes found to have significant associations with Braak stages in GM and WM of TC in tissue, validating snRNA trait association signatures, as determined by linear mixed-effects models using the lmerTest R package. Two-sided tests were performed, and no correction was applied for multiple comparisons. Statistical significance indicated by */** on the plots corresponds to p-values derived from the model output, not from tests performed separately on the plotted data (GM and WM samples from n = 11 individuals). ‘*’ represents p-value ≤ 0.05 and ‘**’ represents p-value ≤ 0.01. Specific p-values GM: ACTG1_Glut- 0.02, CC2D1B_Glut- 0.001, PPP1R1A_Glut- 0.02, ALDH1A1_Glut-0.011, TM2D1_Glut-0.02, ACTG1_GABA- 0.006, SCG3_GABA- 0.016, 0.002, PDCD6_GABA- 0.013, CD74_Microg- 0.012, P2RY12_Microg- 0.001, GFAP_Astro- 0.0001. Specific p-values WM: PDCD6_GABA- 0.03, ANXA7_GABA- 0.012. Boxplot: center line = median; box = upper and lower quartiles; whiskers = values within ± 1.5*IQR. Source data are provided as a Source Data file.
After localizing all amyloid-beta and tau pathological inclusions in the tissue and assigning cells to their major class, we performed two analyses to validate our snRNA-seq results and two discovery analyses based on the spatial data, as described in detail in the “Methods”:
(1) Validation of snRNA-seq-derived proportion associations with Braak stage, using a mixed effects model to assess subcluster marker gene expression differences within each major cell class. (2) Validation of snRNA-seq-derived differential expression signatures at late Braak stages, using 1056 μm2 tile-based pseudobulked differential gene expression analysis (see “Methods”) across Braak stages, and also with donors binarized into high and low pathology groups. (3) Discovery of cell type-specific genes showing preferential enrichment or depletion close to/far away from amyloid-beta plaques, tau tangles, or both pathologies, using a series of mixed effects models. (4) Discovery of cell type-specific genes affected by plaque density (instead of proximity to the closest plaque), by examining tiles (1056 μm2 area) having amyloid-beta plaques versus tiles with no plaques, with gene expression value pseudobulked per tile. This microenvironment-based analysis was only carried out for amyloid-beta plaques, because individuals with cortical tangle pathology did not have sufficient tangle-free tiles.
Validation of snRNA-seq derived associations in neuronal and glial populations with advanced Braak stage
Our first analysis using the ISH data was to identify which snRNA-seq-derived proportion differences and gene expression were also present in intact tissue. To this end, we performed two analyses: (1) differential expression of cell subcluster markers within each broad class across Braak stages, (2) global differential expression using all cells in GM and WM across Braak stages, as well as with donors binarized into low and high pathology groups.
In our cell type-specific analysis, we found that the expression of ACTG1, ALDH1A1, and TM2D1 within glutamatergic neurons was statistically significantly lower in GM tissue from donors at late Braak stages. This expression pattern is consistent with lower proportions of the SPARCL1+ (marked by ACTG1) and RORB+ /IL1RAPL2+ (marked by ALDH1A1 and TM2D1) glutamatergic clusters in the dissociated nuclei (Fig . 4c, 2a, Supplementary Fig. 5d and Supplementary Data 14). Within GABAergic neurons, we found lower expression of ACTG1 and SCG3, which mark the PVALB+ /SPARCL+ subgroup in Braak stages 5/6 in GM, consistent with our snRNA-seq findings (Fig. 4c). In addition to this, we found relatively lower expression of PDCD6 in Braak stages 5/6, consistent with our GABAergic PVALB+ /TMEM132C+ subgroup in GM. Finally, our finding on the relatively resilient THEMIS+ /POSTN+ glutamatergic population was also confirmed in our ISH data, where we found higher expression of PPP1R1A and CC2D1B within glutamatergic neurons at Braak stages 2/4 and 5/6, respectively (Figs. 4c, 5d and Supplementary Data 14).
On the glial side in GM, the ISH data recapitulated lower proportions of the RPL19+ microglial subcluster, as evidenced by lower expression of CD74 and P2RY12 in microglia in intact tissue (Fig. 4c, Supplementary Fig. 5e and Supplementary Data 14). By contrast, higher expression of GFAP at advanced Braak stages was consistent with our snRNA-seq findings regarding higher proportions of the TNC+ (GFAP+) astrocyte cluster at Braak stage 5/6 (Fig. 4c, Supplementary Fig. 5e and Supplementary Data 14).
With respect to WM cell signatures, we identified lower expression of PDCD6 and ANXA7 in PVALB+ GABAergic neurons in Braak stage 5/6, corresponding to our earlier finding of lower proportions of PVALB+ /TMEM132C+ and PVALB + /SPARCL1+ GABAergic clusters (Fig. 4c, Supplementary Fig. 5d and Supplementary Data 15). Thus, even though neuronal cells are rare in white matter, our findings for PVALB+ subgroups are consistent across the dissociated nuclei and intact tissue.
In addition to validating gene expression with each broad cell class in our ISH data, we also examined global signatures aggregated across all cell classes in both GM and WM (Supplementary Fig. 7a) at different Braak stages. At the global level, we did not find any statistically significant genes after multiple testing corrections. However, we found 29 genes with nominal significance, which is a two-fold enrichment with respect to the null. For example, we recapitulated our prior astrocyte-specific analysis and found a general increase in GFAP expression at late Braak stages 5/6. Notably, the expression of TMEM119, a presumed homeostatic microglial gene, is unexpectedly upregulated in donors with advanced pathology. This contrasts with P2RY12, another homeostatic microglial gene, which is typically downregulated in similar conditions; this discrepancy is not found in the snRNA-seq data. We examine the implications of this finding in more detail in the “Discussion”.
Finally, when we grouped donors into low and high pathology (instead of by individual Braak stage), we identified 31 genes with a nominal p-value below 0.05, which is a ~ 4-fold enrichment compared to what would be expected under the null (i.e., 155 tests*0.05 = 8 genes) (Supplementary Fig. 7b). Notably, we found a decrease in SST expression (significantly downregulated in GABAergic neurons in our single nucleus data), as well as a decrease in NEFL expression (significantly downregulated in excitatory neurons in our single nucleus data), lending further evidence to the dysregulation of these neuronal genes in AD. Thus, at both the global and broad cell class level, our in situ analysis confirmed the strongest signals from our dissociated single nucleus RNA-seq proportion and gene expression analyses with nominal statistical significance.
Spatial analysis identifies genes with altered expression near pathological inclusions
In addition to using the spatial data to confirm global and cell type-specific differences from our snRNA-seq data, we also conducted analyses that explicitly used the spatial information from CARTANA. Notably, we examined cell type-specific gene expression changes in the microenvironment of plaques and tangles in GM tissue, taking advantage of our experimental approach localizing RNA and proteinopathic inclusions in the same tissue sections. We used two approaches - a distance-based model and a tile-based model - to examine differences in tissue composition near and far from plaques and tangles. Thus, although our snRNA-seq analysis focused on tangle-based proxies for overall disease progression (Braak stage), our additional spatial analysis includes assessment of changes associated with both hallmark pathologies of AD.
For the distance-based model, we categorized every cell in the GM into one of three groups: close (< 70 μm) to plaques, intermediate distance from plaques (70 - 154 μm), and far away (> 154 μm) from plaques, and the same for close (< 98 μm), intermediate distance (98 - 262 μm) and far (> 262 μm) from the nearest tangles. The distance cutoffs were calculated empirically based on the overall distribution (see “Methods”). For each of these pathologies, we then identified statistically significant differences in cell type-specific gene expression in cells belonging to each of these three groups. In glutamatergic cells, there was higher expression of ALDH5A1 in cells at intermediate distance and far from plaques, whereas NEFL and NEFM showed lower expression in these groups (Fig. 5b and Supplementary Data 16), suggesting that higher expression of neurofilament genes in close proximity may be associated with plaque-related stress. In GABAergic cells, RELN had lower expression in the proximal cells (higher expression in the plaque-intermediate and distant group), consistent with previous findings that RELN+ interneurons are likely affected by AD pathology37,38. In microglia, expression of CD68 was higher in cells at intermediate distance from plaques, while astrocytic GFAP expression was higher in plaque-distant cells (see “Discussion” below for further interpretation of this latter finding). Finally, in endothelial and pericytes, ID3 showed higher expression in plaque-distant cells (Fig. 5b and Supplementary Data 16). For the distance-based model with tangles (Fig. 5c), we found fewer strong associations of cell type signatures with distance from pathological inclusions, due partly to the absence of tangles in the early Braak stage individuals. Notably, we found that THEMIS expression was higher in glutamatergic neurons far from tangles, as was GFAP expression in astrocytes and PLP1 expression in oligodendrocytes (Supplementary Data 17). Interestingly, we also found a weak association of NEFL and NEFM in tangle-distant cells (Supplementary Data 17), in opposition to the stronger effect seen in plaque-proximal cells. If these neurofilament genes are indeed more highly expressed in cells undergoing pathology-induced stress, it suggests that neurons close to cells with tangle aggregates may be less stressed than those in relatively tangle-free areas, consistent with the notion that neurons may aggregate tau as a protective measure against pathogenic soluble tau39.
a A cartoon representing the distance for a cell to get assigned as close, intermediate, or far from plaque/tangle depositions. (Partly created in BioRender. Gaur, P. (2025)). b Heatmap displaying the Δ (change in log-transformed expression) estimated by a linear mixed model (LMM) comparing gene expression across varying distances relative to the close-proximity from plaques in GM of TC on tissue. Each row represents a gene, grouped by cell type, while the color gradient reflects the magnitude and direction of the effect- positive values indicate enriched expression, and negative values indicate depleted expression with distance. Bold values (Δ) annotated with ‘**’ denote statistically significant differences (p-value ≤ 0.01). c Same as b, for tangles. Source data are provided as a Source Data file.
Because amyloid and tau pathology do not occur independently of each other in later Braak stage tissue, we also scrutinized the joint effect of plaque and tangles by including an interaction term in the distance-based model (Supplementary Data 18). This model highlighted several effects of interacting pathologies on cell type-specific gene expression: in glutamatergic neurons, the expression of ACTG1 was lower at an intermediate distance from plaques and when located far from tangles, and DYNC2LI1 expression was lower in cells furthest from both pathologies. Among inhibitory cells, only KIT expression varied in the interaction model, with reduced abundance at an intermediate distance from both tangles and plaques. Overall, our set of distance-based analyses highlighted an additional set of cell type-specific genes whose variability in spatial proximity to pathological inclusions is not directly observable from data obtained through tissue dissociation methods.
Finally, in addition to assessing gene expression based on distance from the closest pathological inclusion, we also analyzed the effect of local pathological density on cell types using a tile-based analysis. We divided the tissue into 100 × 100 pixel tiles and identified those that contained amyloid plaques versus those that did not. This alternative model takes into account not just the closest plaque but rather captures differences due to the overall plaque density in a given microenvironment (“Methods”). Although this analysis did not yield any significant genes after multiple testing correction, 15 genes were nominally significant (p < 0.05), corresponding to a 2-fold enrichment with respect to the null hypothesis (Supplementary Fig. 7c). Similar to our distance-based analysis, this approach also detected lower NEFM expression in glutamatergic neurons in tiles with no plaques. In microglia, we observed an increase of BIN1, a well-known AD risk gene, in plaque-containing tiles. Finally, we found a reduction of SERPINE1, a vascular and microglial specific gene involved in the inhibition of plasmin (which degrades amyloid-beta plaques), in plaque-containing tiles only in individuals with low pathology, suggesting that activation and amyloid-beta degradation may be greater in these individuals in earlier stages of the disease (Supplementary Fig. 7c).
Discussion
Our study is among the first to investigate molecular differences in the temporal cortex in controls and AD donors by employing three techniques; snRNA-Seq, ISH using the CARTANA platform, and immunohistochemical characterization of pathology. Our overall findings can be grouped into four sets of discoveries, as summarized below.
First, we found that multiple subpopulations of neurons had lower proportions in GM in donors at late Braak stages, with the exception of glutamatergic subpopulations marked by THEMIS and POSTN. THEMIS, previously described in T cells, has been identified as a marker of deep-layer neurons in humans, and periostin (POSTN) is an extracellular glycoprotein originally identified as a molecule differentially expressed in osteoblasts and fibroblasts40. Recent studies have shown that POSTN exhibits neurite outgrowth activity in cerebellar granule neurons (CGN) or dorsal root ganglion (DRG) neurons41 and neuroprotective activity42 in adult cortical neurons43. A recent study39 identified an NR4A2+/NTNG2+ deep layer (L6) glutamatergic neuronal type that harbors the lowest number of neurofibrillary tangles in AD, marking it as potentially resistant to pathology. We observed that our THEMIS+/POSTN+ subcluster has high expression of NR4A2 and NTNG2, suggesting this is a subclass of excitatory neurons that are less affected by AD pathology. With respect to GABAergic neurons, the higher proportion of TRHDE-expressing cells at later Braak stages suggests that thyrotropin-associated pathways may show differential regulation in disease. The other GABAergic neuron marker gene with altered expression in individuals with pathology is THSD7B, which has been reported to be associated with age-related cognitive decline, suggesting that this GABAergic subpopulation could also serve as a candidate for further exploration44.
Second, we found multiple glial subgroups with altered proportions in AD, including RPL19+ microglia and TNC+ astrocytes. The former are less abundant in individuals at late Braak stages, whereas the latter shows the opposite effect. However, our cross-sectional study does not provide information as to whether these signatures are protective, reactionary, or pathological. Indeed, prior work on the extracellular matrix protein TNC (Tenascin) has shown it to be at very low levels in the healthy adult mouse nervous system but upregulated in lesioned adult mouse brain45. In human brain tissue, TNC has also been shown to have high expression in EC and SFG within GFAP-high astrocytes10, suggesting that this signature may be associated with astrocytic reactivity in response to neuronal damage or stress.
Third, we found surprising differences in neuronal composition in WM, despite the overall rarity of neurons in this tissue. The RORB+/IL1RAPL2+ glutamatergic neuron subgroup is relatively more abundant in WM in intermediate Braak stages, in contrast to the lower proportions in GM at late Braak stages, suggesting either potential mislocalization of these neurons from GM to WM in disease, or differences in selective vulnerability in GM and WM. By contrast, we found that LAMP5+ /CHST9 + , PVALB +/TMEM132C+ and PVALB +/SPARCL1+ GABAergic clusters are preferentially lower in WM at late Braak stages. Thus, even though neuronal cells are rare in WM, we identified neuronal distribution changes in WM in both directions. Finally, we also observed some sex specific enriched/depleted sub-populations in TC and multiple regions as well (Fig. 2e and Supplementary Fig. 3f-j). Despite our inability to confirm the correlation of these signatures in our ISH data, we still make a note cataloging them for future study, particularly in light of their possible AD specificity.
Fourth, our multiplexed ISH data, combined with immunostaining for pathology on the same section, allowed us to recapitulate the signatures found in our snRNA-seq AD trait association analysis as well as characterize cell type-specific gene expression differences between regions close to and far away from amyloid plaques and neurofibrillary tangles. Given differences in localization of plaques (extracellular) versus neurofibrillary tangles (primarily within neurons), it is not surprising that the pathology-proximal signatures of neurons were different in the two cases. This could reflect the selective vulnerability of neuronal classes described in the “Results” section to each of the two pathologies. By contrast, microglial signatures in plaque-proximal regions showed consistently lower expression of genes such as CD68, and astrocytes in both pathological microenvironments also showed shared signatures, including higher mean expression of GFAP in low pathology environments, a potentially counter-intuitive finding. Notably, prior studies using IHC have shown increased GFAP+ astrocyte density near plaques in the temporal neocortex46. In contrast, our study finds that overall GFAP transcript levels are higher farther from plaques. This can be reconciled if both GFAP-high and GFAP-low astrocyte densities increase near plaques, resulting in a non-unimodal expression pattern. To test this, we used kernel density estimation to identify local minima in the expression distribution and defined density-based thresholds to classify GFAP into ordered expression groups. Analyzing proportions of ‘very low’ and ‘very high’ GFAP expressers across plaque proximity (distance bins) revealed a non-unimodal distribution near plaques, aligning with prior staining data (Supplementary Fig. 7d). We further evaluated the modality of the distributions by computing Hartigan’s dip statistic, which revealed distinct differences among the three plaque distance bins. Plaque proximal (close) bin exhibited the highest median dip value (0.124), indicating a stronger deviation from unimodality and suggesting possible multimodal behavior or underlying GFAP subgroup structure. In contrast, intermediate and far bins showed lower median dip values (0.119 and 0.121, respectively) than the proximal group. These findings point to a more heterogeneous or potentially skewed GFAP expression pattern in plaque-proximal regions, as shown in Supplementary Fig. 7d.
In addition to these intriguing findings, we observe an opposing expression pattern of TMEM119 and P2RY12 in our in situ data during late pathology. Both genes, classified as homeostatic microglial genes, are typically downregulated in advanced pathology. However, in our single-nucleus RNA-seq data, we do not identify clusters with relatively higher or lower expression of TMEM119 versus P2RY12 that are differentially represented in late Braak stages (Figs. 1 and 2). Furthermore, we do not observe strong differential expression of these two genes in opposite directions (Fig. 3). While the underlying cause of this unusual pattern remains unclear, we reference some studies that have reported a similar expression trend for TMEM119. In particular, Satoh et al. (2016)47 reported elevated TMEM119 mRNA levels in the frontal cortex of Alzheimer’s patients, linking increased TMEM119 expression in microglia to tissue inflammation. Similarly, Mercurio et al. (2022)48 observed Tmem119 upregulation in frozen brain sections of traumatic brain injury (TBI) mouse models at Day 7, as determined by RT-PCR and in situ hybridization. This increase was specifically noted in the contused area and surrounding regions, coinciding with heightened microglial activation. Further contextualizing these findings, Ruan and Elyaman (2022)49 published a commentary on Mercurio et al. (2022), which provides a detailed discussion of TMEM119’s role in microglial identification and characterization, integrating insights from multiple studies. Overall, the spatial analysis, while confirming some of the key findings of our dissociated snRNA-seq analysis, also sheds light on signatures that are likely altered in proximity to AD proteinopathies. We identified coordinated changes across almost all major cell types in space, which may not be observable in studies looking at dissociated nuclei. Although we cannot establish causality from proximity-based studies such as this one, we can nonetheless refine the set of candidate signatures that are most likely to be interacting directly with pathological inclusions in AD.
Given the specific combinations of methods described here, it is important to contextualize our findings in light of some of the limitations of the experimental approaches. Notably, our use of multiplexed ISH through the CARTANA platform does suffer from sparsity, leading to noncomprehensive assignment of individual cells to broad cell classes or subtypes. Although our cell class-specific differential expression analysis did allow us to replicate a subset of our snRNA-seq findings, several of the subgroups identified through snRNA-seq were not clearly delineated in ISH, preventing the possibility of validating their proportion differences. This is an important caveat, because differential expression in the CARTANA data may result from depletion of a specific cell cluster, or it may result from a reduction in the expression of that gene within the cluster, without a change in the relative cluster abundance. Although we cannot distinguish these two possibilities from the CARTANA data alone, our differential expression analysis of the single-nucleus data does not suggest that neuronal marker gene expression is not altered within a cluster in association with Braak stages. Thus, although the inability to annotate cells specifically to subclusters in the in situ data precludes a transcriptome-wide validation of the single-nucleus signatures, the CARTANA data set does provide additional evidence that the patterns seen in the single-nucleus data are not due to dissociation artifacts. Another important limitation of the CARTANA analyses is that they are constrained to 2-D sections, which limits the ability to capture the effect of pathological inclusions above and below the plane, in 3-D. This lack of information will increase the variability in gene expression, and thus probably obscures additional signals that could be uncovered if the 3-D location of pathological inclusions was known. As spatial transcriptomics advances, integrating 3-D structure, either experimentally or computationally, will be essential to fully resolve the effect of pathology on cellular architecture and spatial gradients. In addition, our cohort size (40 individuals), while substantial, is smaller than those from recent studies on the prefrontal cortex30,37, suggesting again that there may be some false negatives masked by the heterogeneous presentation of AD and the need for larger cohort studies.
Overall, the study presented here provides a multimodal examination of GM and WM in the temporal cortex, examining both compositional changes as well as spatial differences in tissue from individuals at various Braak stages. Together with other studies looking at cell type differences in post-mortem tissue from other brain regions (primarily GM), this type of examination is necessary not only to characterize cell type-specific alterations in AD, but also to refine a set of candidate signatures to probe in targeted downstream experiments. As with prior studies, we find that multiple classes of cells in the brain show differences in individuals at early versus late stages of pathological accumulation, highlighting key compositional and spatial alterations that may ultimately lead to novel cellular signature-based therapeutics.
Methods
Nuclei isolation and library preparation for Single-nucleus RNA-Seq
Brain autopsies were performed by the Netherlands Brain Bank (NBB; Amsterdam, The Netherlands (https://www.brainbank.nl)), as approved by the local medical ethics committee of the VUmc (registration 2009.148). Donors granted informed consent for autopsy, tissue storage, and the utilization of anonymized clinical and neuropathological data for research. Tissue for this study was obtained from the middle part of the mid-temporal cortex of the right hemisphere. At autopsy, the middle temporal gyrus was isolated and divided in 5 equal parts. For this study, we used the middle (or third) part. This procedure was performed in a consistent manner for all donors. After cryosectioning (10 μm), subcortical WM was separated from the GM using a razor blade. GM and subcortical WM tissue were both processed separately for homogenization and lysis.
Nuclei were isolated from 10 μm fresh-frozen sections using the Nuclei PURE Prep Nuclei Isolation Kit (Sigma-Aldrich), with specific modifications. Regions of interest were macrodissected using a scalpel blade and lysed in Nuclei Pure Lysis Solution containing 0.1% Triton X, 1 mM DTT, and 0.4 U µl-1 SUPERase-In RNase Inhibitor (Thermo Fisher Scientific), freshly added before use. The samples were homogenized sequentially with 23-gauge and 29-gauge syringes.
Cold 1.8 M Sucrose Cushion Solution, prepared immediately before use with 1 mM DTT and 0.4 U µl-1 SUPERase-In RNase Inhibitor, was added to the suspensions, which were then filtered through a 30 μm strainer. The lysates were carefully layered on top of 1.8 M Sucrose Cushion Solution in new Eppendorf tubes and centrifuged for 45 min at 16,000 × g and 4 °C. Resulting pellets were resuspended in Nuclei Storage Buffer containing 0.4 U µl-1 SUPERase-In RNase Inhibitor and transferred to new Eppendorf tubes for centrifugation at 500 g and 4 °C for 5 min. This step was repeated once.
Finally, purified nuclei were re-suspended in Nuclei Storage Buffer with 0.4 U µl-1 SUPERaseIn RNase Inhibitor, stained using trypan blue, and counted with Countess II (Life Technologies). Approximately 12,000 cells per sample were loaded onto the 10 x Single Cell Next GEM G Chip. cDNA libraries were prepared using the Chromium Single Cell 3′ Library and Gel Bead v3 kit, following the manufacturer’s instructions. Sequencing was performed on the Illumina NovaSeq 6000 system with the NovaSeq 6000 S2 Reagent Kit version 1.5 (100 cycles), targeting a minimum sequencing depth of 30,000 reads per nucleus.
Data processing and quality control
All samples from our temporal cortex dataset, as well as from the public datasets, underwent uniform processing using Cell Ranger50 (version 3.1.0), employing the GRCh38 reference human genome and the Ensembl Homo_sapiens GRCh38.91 reference annotation, modified to include intronic reads. Nuclei were characterized as barcodes containing a minimum of 500 unique molecular identifiers (UMIs) (excluding mitochondrial RNA) and less than 5% mitochondrial RNA. For samples with over 10,000 nuclei, only the top 10,000 nuclei with the highest UMI counts were retained. Doublets were marked using scDblFinder51 (version 1.4.0) but not excluded, as none of the clusters were enriched in doublets, though we removed nuclei that were flagged as potential doublets (Supplementary Data 2). We filtered out two samples from TC region (from one individual) due to low quality (less than 500 cells with a lower amount of UMI, detected genes and higher mitochondrial reads percentage than the other samples) (Supplementary Fig. 1d). The primary QC visualization for TC region and other multiple brain region studies included in the analysis can be seen in Supplementary Fig. 2a, b.
Integration of temporal cortex data with other cortical regions
The 80 samples from the TC region were first merged using the Seurat workflow (Supplementary Fig. 1c) without batch correction. We then used Harmony52 to integrate our TC dataset (GM and WM) with other published studies from different brain regions, including entorhinal cortex10,11, prefrontal cortex5,12,13, superior frontal gyrus10 and deep white matter from prefrontal cortex14. The matrix of filtered UMI counts (> 500 per nucleus) for each study was converted to a single cell experiment (SCE) object using the function read10xCounts from the SingleCellExperiment53 package in R (version 4.0.1). The data class conversion from sce to Seurat object and merging of all datasets were executed using conventional functions CreateSeuratObject and merge respectively from Seurat package (Version 3.2.0.)54. The merging of all datasets did not colocalize the same brain regions (Supplementary Fig. 2c), which led to the important step of integration. We then integrated all datasets using Harmony52 (version 0.1.0) (Supplementary Fig. 2d, e) within the Seurat workflow utilizing the RunHarmony function. Before performing the integration, we followed the conservative steps of normalizing, scaling the counts and running the PCA on a total of 959,237 nuclei from all the studies together using inbuilt functions from the Seurat package, and selected 30 principal components for the Harmony integration (version 0.1.0) (resolution = 0.8, ‘study’ as the integration variable). Since the Harmony algorithm iteratively corrects PCA embeddings, the downstream analyses used the Harmony embeddings instead of PCA.
We clustered the integrated datasets using the Seurat::FindClusters function at a resolution of 0.8. The FindClusters function, which leverages the Louvain algorithm for community detection, identifies clusters of cells as groups of densely connected nodes in a graph. Specifically, the function constructs a shared nearest neighbor (SNN) graph using the FindNeighbors function and then applies the Louvain algorithm to partition the graph into clusters. These pre-clusters were annotated using a canonical set of marker genes after machine learning based refinement and were split into 8 broad cell type clusters (Supplementary Fig. 2f and Supplementary Data 3) with a reasonable distribution of various cell types (Supplementary Fig. 2g) that were visualized employing a 2D UMAP55 projection executing DimPlot function from the Seurat package and a bar plot using dittoSeq::dittoBarPlot56 function respectively. After identification and clustering of broad cell types at the top level, we sub-clustered all broad cell types independently using Harmony integrated data to investigate further subtypes within each class. To subcluster, we started by subsetting the specific cell type cluster from the integrated dataset and reintegrated them using the same workflow as described above. The downstream analyses, including the trait association investigation, were performed on these subclusters generated from Harmony integrated data (Supplementary Data 4).
Machine learning based refinement of integrated clusters
We used Random Forest as a robustness method for our subclusters obtained for all major cell types. To train the model, we included the subclusters within each cell type with at least 50 cells, where 75% of the cells could be used as a training set and the remaining 25% for testing, at a minimum, with the topmost variable genes. We identified genes that drove biological heterogeneity in the data set by modeling the per-gene variance using the function modelGeneVar from the Bioconductor package scran57 (version 1.18.7). After predicting the classes on the testing set of clusters, we evaluated the accuracy of a classification with the help of a confusion matrix (Supplementary Fig. 8). Based on this post-hoc Random Forest-derived confusion matrix, we refined the clustering by merging clusters that exhibited approximately ≥ 10% mutual confusion and were spatially adjacent in the UMAP projection. This refinement aimed to improve cluster coherence by integrating both quantitative similarity and qualitative spatial context, resulting in a more robust set of clusters for the integrated dataset. It is important to note that this training was performed on the sub-clusters prior to the final assignment of identity labels.
Cross-comparison between Integrated and TC region specific clusters
After employing random forest as a robustness method for our subclusters in the integrated dataset, we clustered the TC nuclei separately as well and compared the clusters from the TC region to the integrated ones and ran another round of refinement of clusters. This led us to merge some additional clusters in the integrated dataset for each broad cell type (Supplementary Fig. 9) based on approximately 10% similarity and adjacency in a 2-D UMAP projection. The integrated clusters were annotated based on the top cluster markers present within each identity cluster. We obtained these markers using the function FindAllMarkers from the Seurat package. We only tested genes that were detected in minimum 25% cells in a subcluster and showed at least 0.25-fold difference (log-scale). The glutamatergic subclusters were annotated using layer marker genes co-expressed with other top cluster identity markers. The layer association of top gene markers for glutamatergic cell types were established using transcriptomics explorer offered by Allen brain map atlas encompassing human multiple cortical areas (Transcriptomics Explorer:: Allen Brain Atlas: Cell Types). Our multiple-layered refinement approach led us to have 888,784 (after QC and potential doublet removal) nuclei to analyze further. (Supplementary Data 2 and Supplementary Fig. 2e, f, h).
Trait association in TC region and comparison to different brain regions
We excluded samples AD001 and AD002 from further analysis (both trait association and differential expression) because they did not meet our quality control (QC) criteria (Supplementary Fig. 1d). Furthermore, we removed all non-AD dementia samples from our study examining changes in cell proportions related to AD pathology in the TC region. To explore the cluster proportion changes in early or late AD pathology in WM and GM of the TC region, we implemented ANCOM-BC58 (Analysis of Compositions of Microbiomes with Bias Correction, version 1.0.5), which is a log-linear model that determines differentially abundant taxa according to the variable of interest. In our analysis, this enables the identification of subpopulations whose compositional abundances differ significantly across AD pathology, denoted by various Braak stages. The output of ANCOM-BC includes a dataframe with beta values, which correspond to the log fold-change in the compositional abundance of a subpopulation between varying Braak stages. A positive beta value indicates that the Braak stage is associated with increased abundance of the subcluster, while a negative beta value signifies a decrease in abundance. We inspected WM and GM from TC in each cell type independently to investigate any associations of the changes in cluster proportion with AD pathology. The clusters were implemented as taxa, and the AD pathology marked by different Braak stages was used as a covariate of interest. We regressed out sex, age of death and postmortem interval. Numeric covariates, like age of death and postmortem interval, were divided by two times their standard deviations to scale the regression59. The Braak stages were classified in three discrete categories. Braak stages less than 2, ranging from 2-4, and more than 5 were designated as ‘Braak_stage 0-1’, ‘Braak_stage 2-4’, and ‘Braak_stage 5-6’ respectively.
We implemented the same strategy for the integrated cell type classes with an additional covariate of different brain regions in the model. Before fitting the log linear model, we removed any cells from the integrated dataset that had no information of PMI, age of death, sex, and Braak stage. We also excluded the Grubman et al.11 study from this part of the analysis due to variations in demographic details. This additional filtering resulted in a total of 835,413 nuclei for the trait association analysis that were derived from a total of 216 samples. The trait association analysis was performed separately for each broad cell type.
In order to compare the TC region signatures with two recent datasets from MTG (Gabitto et al.28, 2024) and PFC region (Green et al.30, 2024), we first mapped our TC sub cellular populations onto these both datasets separately. We utilized the cell_type_mapper package from Cell Type Mapper to transfer annotations of subcellular populations from our single-nuclei dataset from TC onto both regions. Initially, we generated the taxonomy file from our TC dataset using the command - ‘python3 -m cell_type_mapper.cli.precompute_stats_scrattch’. Subsequently, we performed dataset mapping with the command ‘python3 -m cell_type_mapper.cli.map_to_on_the_fly_markers’, designating our dataset as the reference and aforementioned datasets as the query, using default parameters. After retrieving the mapped labels (Supplementary Data 7, 9), we gathered the required metadata information like braak stage, age of death, sex and PMI for 84 donors in MTG region from SEA-AD consortium and 424 in PFC from ROSMAP and implemented a log linear regression model using ANCOM-BC60 (version 2.0.2) similarly as our previous trait association analysis with TC region (Supplementary Data 8, 10). Notably, we used the ‘ancombc’ function and ensured that the model remained consistent with the one applied in the TC region analysis.
Differential expression analysis
We performed our differential expression analyses using a ‘pseudo-bulk’ approach, that is, we summed the total number of UMI for each sample and each cell type, resulting in a single count value for each gene in each sample and each cell type. We only retained samples (including non-AD related dementia samples) which had at least 10 nuclei from a specific cell type, and subpopulations that were detected in at least 20 samples. We also only tested genes with a median count per million (CPM) of at least 1 and expressed in at least 20 samples. Furthermore, we removed genes with low counts using the ‘filterByExpr’ function from the edgeR61 R package. We then used the glmmTMB62 R package to fit a negative binomial mixed model to the data for each cell type (family = nbinom2). We used the following models: counts (raw) ~ Braak_stage (or diagnosis) * tissue (gray/white) + age + sex + pmi + (1|individual) (random effect as each individual has a sample from gray/white matter), effectively testing for an effect of Braak_stage (or diagnosis) in our cohort, as well as allowing brain region-specific effects. As an offset, we used the log of the TMM normalized library size63 minus the log of 1 million. The following variables were categorized prior to model fitting as this improved the convergence of our statistical model: Braak_stage (0-1, 2-4, 5-6), age (< 60, 60-80, > 80), pmi (< 6, 6-8, > 8).
Enrichment of differentially expressed genes
First, we identified genetically enriched AD pathways by using MAGMA64. Briefly, we obtained gene-level genetic associations using GWAS summary statistics from a recent AD GWAS65 (using all SNPs within a window starting from 35 kb upstream of each gene to 10 kb downstream). We then used MAGMA to test whether any pathways from the GO_BP, GO_CC, GO_MF, Hallmark, KEGG and REACTOME databases were enriched in AD genetic risk. Multiple testing correction was performed using the Benjamini-Hochberg procedure. Finally, we then tested whether the differentially expressed genes were enriched in any AD genetically associated pathways using the fgsea66 R package.
In situ sequencing
Immunostaining of human samples
Based on snRNA results and previously published studies, we chose 155 genes (Supplementary Data 12) to analyze the spatial arrangement of cell types on the tissue sections of 13 individuals comprising different stages of AD pathology as well as controls. We used CARTANA to generate and sequence clonally amplified barcode sequences, deploying gene-specific probes ligation onto tissue sections.
Neuropathologically characterized human brain blocks from controls and Alzheimer’s patients obtained from Netherland Brain Bank were cryosectioned into 10 µm sections, onto SuperFrost Plus glass slides (Thermo Fisher) and stored at - 80 °C. In situ sequencing was performed using the reagents supplied by CARTANA (PN1110-02, HS Library Preparation Kit; PN3110-02 - In situ sequencing kit) and following the recommended protocol for human fresh frozen samples, with autofluorescence quenching. Briefly, the sections underwent fixation, permeabilization, probes hybridization and ligation, rolling circle amplification, and seven sequential cycles of fluorescence labeling and image acquisition at 20x magnification on an ECLIPSE Ti2 inverted microscope (Nikon) (Supplementary Fig. 5a). The probes used belonged to a panel of 155 genes, designed for this study by CARTANA, based on their state-of-the-art guidelines.
After completing in situ sequencing, the same sections underwent immunostaining of amyloid plaques and phosphorylated Tau. Briefly, after removing the coverslips, in situ sequencing probes were detached through sequential washings with formamide. Afterwards, blocking solution (SP5030-250, Vector Labs) was added for 20 min before performing an incubation with mouse anti-Phospho-Tau (Ser202, Thr205) (MN1020, ThermoFisher) for two hours, followed by one hour incubation with donkey anti-mouse IgG [H + L]/ AF 550 (SA5-10167, Invitrogen) and one hour incubation with mouse anti-Beta-Amyloid MOAB-2/AF 750 (NBP2-13075AF750, Novus Biologicals). Finally, sections were incubated for 30 seconds in 1x TrueBlack (23007, Biotium) to reduce autofluorescence and for 5 min in DAPI (62248, ThermoFisher) for nuclei detection. Sections were then mounted using SlowFade™ Gold Antifade Mountant (S36940, ThermoFisher), and imaging was carried out using a 20x magnification on an ECLIPSE Ti2 inverted microscope (Supplementary Fig. 6a).
Gene expression
For each sample, we obtained from in situ sequencing a reference DAPI image and 6 sequencing images (one for each barcode position) that were aligned and combined using ISSanalysis software (Quantified Biology) in order to generate a map of the spatial coordinates of each cell (Supplementary Data 13) and each target RNA molecule on the section. Afterwards, for each tissue section, it was determined the total count of spots for each target gene and this value was normalized on the tissue area (Supplementary Data 19). To compare the gene expression of different samples while removing any bias due to technical factors, the expression of each gene was normalized also on the total number of identified molecules per sample (Supplementary Data 20).
Gene enrichment in plaques proximity
First Halo v3.1 (Indica Lab) was used to build a classifier able to automatically recognize beta-amyloid plaques from the immunostaining images (Supplementary Fig. 5b). Plaques coordinates (Supplementary Data 21) were then exported and uploaded on ISSanaysis software, where they were used to identify genes, whose expression was enriched in proximity of plaques. In order to do so, a grid was applied to each tissue section and for each tile was determined the presence/absence of a plaque and the presence of any of the in situ sequencing target genes (Supplementary Fig. 5c). Similarly, to what was done previously, for each sample it was determined the normalized count of reads for each gene in the areas with and without plaques.
In situ cell-type classification
We used the following strategy to label cells in our CARTANA ISS experiment. First, we aggregated all counts for each gene and each broad cell type (across samples) from our temporal cortex single nuclei experiment. Second, we normalized the broad cell type pseudo-bulk expression data so that each broad cell type had a total of 10k reads (CP10k). We then computed a specificity metric for each gene by dividing the CP10k from each broad cell type by the total CP10k of all broad cell types. We then identified broad cell-type marker genes as genes with a CP10k of at least 0.01 and a specificity metric greater than 0.5 (indicating that more than 50% of the expression of the gene across cell types is quantified in the given cell type). We then used the subset of marker genes that were also quantified in our CARTANA ISH experiment to label cell types. For each CARTANA cell, we summed the total number of CARTANA counts across all markers from the different broad cell types (i.e., Astrocytes, Oligodendrocytes, etc.). Then, for each cell, we computed a z-score of the counts in marker genes from the different cell types, which were then transformed into a one-sided p-value using the pnorm R function. Finally, cell type labels (i.e., Astrocytes, Oligodendrocytes, etc.) were given to each individual cell from the tissue section if the p-value was below 0.05 (indicating that the sum of count for a marker of a specific cell type was significantly higher than the counts of marker genes for the other cell types).
Pathology-induced gene expression variability in the spatial environment
We modeled the expression of 155 genes within each broad cell type for WM and GM of the TC region separately to test whether there’s a significant association between the expression of 155 CARTANA genes and AD phenotypic trait (Braak stage). We fit a linear mixed-effects model using lmerTest67 R package to investigate this relationship of gene expression with different Braak stages while taking individual variation as a random effect for each major cell type. Similar to our snRNA trait association analysis, we categorized the Braak stages into three groups; Braak stage 0-1, 2-4 and 5-6 and modeled the association for every cell type in WM and GM of TC (Fig. 4c).
Microenvironment-induced gene expression variability in the spatial environment
We modeled the expression of 155 genes based on the microenvironment of plaque and tangles depositions. Based on the spatial cell coordinates for all samples (except for AD006) along with those of the plaques and tangles, we were able to represent the potential location of pathology on tissue via a 2-D projection (Fig. 4a, b). Detailed information on cell coordinates for all samples and samples used in plaques and tangles distance based analyses is available in Supplementary Data 12, 13, 21, 22. For the distance based model, we computed the Euclidean distance of each cell from the center of each plaque location for every sample and considered the minimum distance to the closest plaque for each cell to consider the microenvironment. We then categorized (using ntile function from dplyr68 R package) and calculated these distances based on an area of 0.1056 µm² per pixel into three groups: close, intermediate, and far. The same strategy was applied to compute the tangles’ microenvironment. To associate the cell location from the plaques on the tissue we distributed the distances into three bins marking each cell either being close (bin1), intermediate (bin2) or far away (bin3) from the plaques or tangles (Fig. 5a). We then fit the linear mixed-effects model to investigate this relationship while including distance bin as the fixed effect and regressing out the variation coming from individuals.
To analyze the global signature in the in situ transcriptomics data (refer to Supplementary Fig 7 and Supplementary Data 23), we performed the following steps: (1) data summation and normalization: For each brain section, we summed the counts for each gene. These counts were then normalized to a total of 6000 counts per section, which is close to the median count of 5649 per section. The normalized counts were subsequently log-transformed. (2) Statistical analysis by Braak stage: We tested for differences in normalized gene counts across different Braak stage categories (Braak 0-1, Braak 2-4, and Braak 5-6) using the Wilcoxon rank-sum test. To account for multiple comparisons, we applied the Benjamini-Hochberg procedure to adjust the p-values. (3) Comparison between AD cases and controls: Similarly, we used the Wilcoxon rank-sum test to identify significant differences in gene expression between Alzheimer’s disease (AD) cases and control samples. (4) Plaque analysis: For the plaque analysis, we categorized image tiles into those containing plaques and those that were plaque-free (Supplementary Data 24). We summed the counts for each gene within each sample and plaque category, normalizing these counts to 20,000 counts per sample and plaque category. This was followed by a log + 1 transformation. To test for differences in gene expression between plaque-containing and plaque-free tiles, we used a paired Wilcoxon rank-sum test and corrected for multiple comparisons using the Benjamini-Hochberg procedure.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Single-nuclei RNA-seq data for the Temporal cortex datasets have been deposited at the European Genome-phenome Archive, which is hosted by the European Bioinformatics Institute and the Center for Genomic Regulation, under accession ID- EGAD00001009166. Ensembl Homo_sapiens GRCh38.91 reference annotation: http://ftp.ensembl.org/pub/release-91/gtf/homo_sapiens/. The Supplementary Figures are available in Supplementary Information. Source data are provided in this paper.
Code availability
The code used to perform the analysis described in this study is available at https://github.com/Gaur-et-al.
References
Mucke, L. Alzheimer’s disease. Nature 461, 895–897 (2009).
Dolgin, E. Alzheimer’s disease is getting easier to spot. Nature 559, S10–S12 (2018).
Mostafavi, S. et al. A molecular network of the aging human brain provides insights into the pathology and cognitive decline of Alzheimer’s disease. Nat. Neurosci. 21, 811–819 (2018).
Williams, J. B., Cao, Q. & Yan, Z. Transcriptomic analysis of human brains with Alzheimer’s disease reveals the altered expression of synaptic genes linked to cognitive deficits. Brain Commun. 3, https://doi.org/10.1093/braincomms/fcab123 (2021).
Mathys, H. et al. Single-cell transcriptomic analysis of Alzheimer’s disease. Nature 570, 332–337 (2019).
Heneka, M. T. et al. Neuroinflammation in Alzheimer’s disease. Lancet Neurol. 14, 388–405 (2015).
Obulesu, M. & Jhansilakshmi, M. Neuroinflammation in Alzheimer’s disease: an understanding of physiology and pathology. Int. J. Neurosci. 124, 227–235 (2013).
Miyoshi, E. et al.Spatial and single-nucleus transcriptomic analysis of genetic and sporadic forms of Alzheimer’s disease.https://doi.org/10.1101/2023.07.24.550282 (2023).
Luquez, T. et al. Cell type-specific changes identified by single-cell transcriptomics in Alzheimer’s disease. Genome Med. 14, https://doi.org/10.1186/s13073-022-01136-5 (2022).
Leng, K. et al. Molecular characterization of selectively vulnerable neurons in Alzheimer’s disease. Nat. Neurosci. 24, 276–287 (2021).
Grubman, A. et al. A single-cell atlas of entorhinal cortex from individuals with Alzheimer’s disease reveals cell-type-specific gene expression regulation. Nat. Neurosci. 22, 2087–2097 (2019).
Cain, A. et al. Multi-cellular communities are perturbed in the aging human brain and Alzheimer’s disease. https://doi.org/10.1101/2020.12.22.424084 (2020).
Zhou, Y. et al. Human and mouse single-nucleus transcriptomics reveal TREM2-dependent and TREM2-independent cellular responses in Alzheimer’s disease. Nat. Med. 26, 131–142 (2020).
Bryois, J. et al. Cell-type-specific cis-eQTLs in eight human brain cell types identify novel risk genes for psychiatric and neurological disorders. Nat. Neurosci. 25, 1104–1112 (2022).
Application Note: Mapping brain cell types with CARTANA in situ sequencing on the Nikon Ti2-E microscope. Nature https://www.nature.com/articles/d42473-019-00264-8 (2019).
Bennett, D. A., Wilson, R. S., Boyle, P. A., Buchman, A. S. & Schneider, J. A. Relation of neuropathology to cognition in persons without cognitive impairment. Ann. Neurol. 72, 599–609 (2012).
Muratore, C. R. et al. Cell-type dependent Alzheimer’s dsease phenotypes: Probing the biology of selective neuronal vulnerability. Stem Cell Rep. 9, 1868–1884 (2017).
Kong, W. et al. Independent component analysis of Alzheimer’s DNA microarray gene expression data. Mol. Neurodegener. 4, https://doi.org/10.1186/1750-1326-4-5 (2009).
Squillario, M. & Barla, A. A computational procedure for functional characterization of potential marker genes from molecular data: Alzheimer’s as a case study. BMC Med. Genom. 4, 55 (2011).
Cruchaga, C. et al. GWAS of Cerebrospinal fluid tau levels identifies risk variants for Alzheimer’s disease. Neuron 78, 256–268 (2013).
Lai, M. K. P., Esiri, M. M. & Tan, M. G. K. Genome-wide profiling of alternative splicing in Alzheimer’s disease. Genom. Data 2, 290–292 (2014).
Jeong, Y. et al. 1950 MHz Electromagnetic fields ameliorate Aβ pathology in Alzheimer’s disease mice. Curr. Alzheimer Res. 12, 481–492 (2015).
Ali, F., Baringer, S. L., Neal, A., Choi, E. Y. & Kwan, A. C. Parvalbumin-positive neuron loss and amyloid-β deposits in the frontal cortex of Alzheimer’s disease-related mice. J. Alzheimer’s Dis. 72, 1323–1339 (2019).
Yin, G. N., Lee, H. W., Cho, J.-Y. & Suk, K. Neuronal pentraxin receptor in cerebrospinal fluid as a potential biomarker for neurodegenerative diseases. Brain Res. 1265, 158–170 (2009).
Richens, J. L. et al. Practical detection of a definitive biomarker panel for Alzheimer’s disease; comparisons between matched plasma and cerebrospinal fluid. Int. J. Mol. Epidemiol. Genet. 5, 53–70 (2014).
Deng, Y. et al. Loss of LAMP5 interneurons drives neuronal network dysfunction in Alzheimer’s disease. Acta Neuropathol. 144, 637–650 (2022).
Cacabelos, R., Cacabelos, P. & Torrellas, C. Personalized medicine of Alzheimer’s disease. in Handbook of Pharmacogenomics and Stratified Medicine 563–615 (Elsevier, 2014).
Gabitto, M. I. et al. Integrated multimodal cell atlas of Alzheimer’s disease. Nat. Neurosci. 27, 2366–2383 (2024).
AllenInstitute. GitHub - AllenInstitute/cell_type_mapper: Repository for storing prototype functionality implementations for the BKP. GitHub https://github.com/AllenInstitute/cell_type_mapper.
Green, G. S. et al. Cellular communities reveal trajectories of brain ageing and Alzheimer’s disease. Nature 633, 634–645 (2024).
Libiger, O. et al. Identification of NPTX2 as a prognostic biomarker of Alzheimer’s disease through a longitudinal CSF proteomics study in ADNI subjects. Alzheimer’s Dementia 16, e047605 (2020).
Namba, Y., Tomonaga, M., Kawasaki, H., Otomo, E. & Ikeda, K. Apolipoprotein E immunoreactivity in cerebral amyloid deposits and neurofibrillary tangles in Alzheimer’s disease and kuru plaque amyloid in Creutzfeldt-Jakob disease. Brain Res. 541, 163–166 (1991).
Steinberg, S. et al. Loss-of-function variants in ABCA7 confer risk of Alzheimer’s disease. Nat. Genet. 47, 445–447 (2015).
Karczewski, K. J. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581, 434–443 (2020).
Yu, X., Ng, C. P., Habacher, H. & Roy, S. Foxj1 transcription factors are master regulators of the motile ciliogenic program. Nat. Genet. 40, 1445–1453 (2008).
Deutsch, S. I., Rosse, R. B. & Deutsch, L. H. Faulty regulation of tau phosphorylation by the reelin signal transduction pathway is a potential mechanism of pathogenesis and therapeutic target in Alzheimer’s disease. Eur. Neuropsychopharmacol. 16, 547–551 (2006).
Mathys, H. et al. Single-cell atlas reveals correlates of high cognitive function, dementia, and resilience to Alzheimer’s disease pathology. Cell 186, 4365–4385 (2023).
Kocherhans, S. et al. Reduced reelin expression accelerates amyloid- plaque formation and tau pathology in transgenic Alzheimer’s disease Mice. J. Neurosci. 30, 9228–9240 (2010).
Otero-Garcia, M. et al. Molecular signatures underlying neurofibrillary tangle susceptibility in Alzheimer’s disease. Neuron 110, 2929–2948 (2022).
Horiuchi, K. et al. Identification and characterization of a novel protein, periostin, with restricted expression to periosteum and periodontal ligament and increased expression by transforming growth factor β. J. Bone Miner. Res. 14, 1239–1249 (1999).
Shih, C.-H., Lacagnina, M., Leuer-Bisciotti, K. & Pröschel, C. Astroglial-derived periostin promotes axonal regeneration after spinal cord injury. J. Neurosci. 34, 2438–2443 (2014).
Shimamura, M. et al. Role ofcentral nervous system periostin in cerebral Ischemia. Stroke 43, 1108–1114 (2012).
Matsunaga, E. et al. Periostin, a neurite outgrowth-promoting factor, is expressed at high levels in the primate cerebral cortex. Dev. Growth Differ. 57, 200–208 (2015).
De Jager, P. L. et al. A genome-wide scan for common variants affecting the rate of age-related cognitive decline. Neurobiol. Aging 33, 1017.e1–1017.e15 (2012).
Laywell, E. D. et al. Enhanced expression of the developmentally regulated extracellular matrix molecule tenascin following adult brain injury. Proc. Natl. Acad. Sci. USA 89, 2634–2638 (1992).
Serrano-Pozo, A. et al. Differential relationships of reactive astrocytes and microglia to fibrillar amyloid deposits in Alzheimer disease. J. Neuropathol. Exp. Neurol. 72, 462–471 (2013).
Satoh, J. et al. TMEM119 marks a subset of microglia in the human brain. Neuropathology 36, 39–49 (2015).
Mercurio, D. et al. Protein expression of the microglial marker Tmem119 decreases in association with morphological changes and location in a mouse model of traumatic brain injury. Front. Cell. Neurosci.16, https://doi.org/10.3389/fncel.2022.820127 (2022).
Ruan, C. & Elyaman, W. A new understanding of TMEM119 as a marker of microglia. Front. Cell. Neurosci. 16, https://doi.org/10.3389/fncel.2022.902372 (2022).
Zheng, G. X. Y. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Germain, P.-L., Lun, A., Macnair, W. & Robinson, M. D. Doublet identification in single-cell sequencing data using scDblFinder. F1000Res. 10, 979 (2021).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Amezquita, R. A. et al. Orchestrating single-cell analysis with Bioconductor. Nat. Methods 17, 137–145 (2019).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
McInnes, L., Healy, J., Saul, N. & Großberger, L. UMAP: Uniform Manifold Approximation and Projection. J. Open Source Softw. 3, 861 (2018).
dittoSeq. Bioconductor https://bioconductor.org/packages/dittoSeq (2020).
Lun, A. T. L., McCarthy, D. J. & Marioni, J. C. A step-by-step workflow for low-level analysis of single-cell RNA-seq data with Bioconductor. F1000Res. 5, 2122 (2016).
Lin, H. & Peddada, S. D. Analysis of compositions of microbiomes with bias correction. Nat. Commun. 11, 3514 (2020).
Gelman, A. Scaling regression inputs by dividing by two standard deviations. Stat. Med. 27, 2865–2873 (2007).
Lin, H. & Peddada, S. D. Multigroup analysis of compositions of microbiomes with covariate adjustments and repeated measures. Nat. Methods 21, 83–91 (2024).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2009).
Brooks, M. et al. glmmTMB Balances speed and flexibility among packages for zeroinflated generalized linear mixed modeling. R. J. 9, 378 (2017).
Robinson, M. D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 11, https://doi.org/10.1186/gb-2010-11-3-r25 (2010).
de Leeuw, C. A., Mooij, J. M., Heskes, T. & Posthuma, D. MAGMA: Generalized Gene-Set Analysis of GWAS Data. PLOS Comput. Biol. 11, e1004219 (2015).
Wightman, D. P. et al. A genome-wide association study with 1,126,563 individuals identifies new risk loci for Alzheimer’s disease. Nat. Genet. 53, 1276–1282 (2021).
Korotkevich, G. et al. Fast gene set enrichment analysis. https://doi.org/10.1101/060012 (2016).
Kuznetsova, A., Brockhoff, P. B. & Christensen, R. H. B. lmerTest Package: Tests in Linear Mixed Effects Models. J. Stat. Softw. 82, 1–26 (2017).
Wickham, H., François, R., Henry, L., Müller, K. & Vaughan, D. dplyr: A Grammar of Data Manipulation. R package version 1.1.4, https://dplyr.tidyverse.org (2025).
Acknowledgements
We extend our gratitude to the generous individuals who donated their brains to further research at the Netherlands Brain Bank. This work was supported in part by NIH grants R01AG066831 (VM) and U19AG074862 (VM).
Author information
Authors and Affiliations
Contributions
V.M. and D.M. acquired funding, and V.M. and D.M. jointly designed and supervised the study. P.G. performed the majority of the data analysis, generated plots, and wrote most of the manuscript with input from all authors. J.B. performed additional data analysis, figure generation, and manuscript writing. D.C. conducted scRNA-seq and CARTANA experiments. L.F. and J.H. contributed to the experiments and image analysis. A.Y. assisted with manuscript revision. All authors reviewed and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
D.M. was a full-time employee of F. Hoffmann-La Roche Ltd when this study was done. D.M. is now an employee of Biogen. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Vivek Swarup and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Gaur, P., Bryois, J., Calini, D. et al. Single-nucleus and spatial transcriptomic profiling of human temporal cortex and white matter reveals key associations with AD pathology. Nat Commun 16, 10395 (2025). https://doi.org/10.1038/s41467-025-65350-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65350-6