Abstract
The resistance to KRAS-targeted therapies, particularly due to co-occurring gene mutations, remains a significant challenge. Through a metabolite library screening, we reveal that polyamines sensitize KRAS inhibitors only in KRASMU/KEAP1WT cells but not in KRASMU/KEAP1MU cells. Transcriptome sequencing and metabolome profiling pinpoint SAT1, the key enzyme in polyamine metabolism, as essential for this divergence. In KRASMU/KEAP1WT context, treatment of KRAS inhibitors activates JNK/c-Jun pathway and SAT1 expression, while the augmented SAT1 facilitates polyamine metabolism and KRAS inhibitors-induced ferroptosis. Conversely, in KRASMU/KEAP1MU cells, activated JNK promotes the degradation of NRF2, thereby inhibiting SAT1 expression. Our results further demonstrate that polyamine supplementation enhances KRAS-targeted therapy in KRASMU/KEAP1WT resistant cells, patient-derived organoids, xenografts, and spontaneously tumorigenic mice, while KRASMU/KEAP1MU models require lentivirus or adeno-associated virus-mediated SAT1 overexpression prior to polyamine treatment, to augment ferroptosis and drug sensitivity. Our findings highlight SAT1-mediated polyamine metabolism as a promising target in precision treatments for KRAS-mutant cancers.
Similar content being viewed by others
Introduction
Kirsten rats arcomaviral oncogene homolog (KRAS) mutation is among the most prevalent oncogenic mutations in human cancers, particularly in pancreatic adenocarcinoma (95%), colorectal cancer (35%), and lung cancers (25%)1,2. Although KRAS mutations were still considered undruggable a few years ago, recent advancements in targeted therapies, such as the development of KRASG12C inhibitors like sotorasib3 and adagrasib4 have opened up promising therapeutic avenues5,6,7. However, the effectiveness of these inhibitors is often substantially compromised by the presence of gene co-mutations, especially those involving kelch-like ECH-associated protein 1 (KEAP1) and liver kinase B1 (LKB1), which necessitates the exploration of novel strategies to enhance their sensitivity8,9,10,11.
Ferroptosis, an iron-dependent cell death characterized by polyunsaturated fatty acid (PUFA) peroxidation driven by an imbalance of reactive oxygen species (ROS)12,13, has recently been highlighted for its role in overcoming tumor treatment resistance14,15. Considering KRAS mutations’ extensive and intricate regulatory effects on lipid metabolism and antioxidant responses16,17, a promising opportunity emerges to enhance the sensitivity of KRAS-targeted therapies by promoting ferroptosis, as reported before18. However, this scenario could be further complicated by gene co-mutations. KEAP1 mutations, for instance, disrupt the KEAP1-neuclear factor-erythroid 2-related factor 2 (NRF2) antioxidant signaling pathway, resulting in enhanced antioxidant responses and a potential shift in the balance toward ferroptosis induction.
Polyamines, a class of small polycationic alkylamines that includes spermine, spermidine, and putrescine, play pivotal roles in a myriad of cellular processes, such as gene regulation, cell proliferation, and cell death19,20. Spermidine/spermine N1-acetyltransferase 1 (SAT1), the critical enzyme involved in polyamine catabolism, catalyzes the conversion of spermidine and spermine into their N1-acetylated forms. The oxidation of N1-acetylated polyamines by polyamine oxidase (PAOX) generates ROS, which in turn promotes ferroptosis21,22. Notably, tumor cells, especially those harboring KRAS mutations, often rely on elevated intracellular polyamine levels to sustain their relentless proliferation23. Given this dependency, targeting SAT1-mediated polyamine metabolism has emerged as a promising therapeutic strategy in cancer treatment21,24. However, it has yet to be well explored in KRAS-targeted therapies.
In this study, through a metabolite library screening, we identified and validated that polyamines can enhance the efficacy of KRAS-targeted therapies, but only in KRAS-mutant tumors with wild-type KEAP1(KRASMU/KEAP1WT). In KRAS-mutant tumors with mutated KEAP1(KRAS MU/KEAP1MU), lentivirus or adeno-associated virus (AAV)-mediated SAT1 overexpression, combined with polyamine treatment, is required to augment ferroptosis and drug sensitivity. Further exploration in resistant cells, organoids, xenografts, and spontaneous lung cancer models demonstrated that targeting SAT1, with different strategies based on the KEAP1 mutation status, enhances KRAS-targeted therapies by modulating polyamine metabolism and inducing ferroptosis.
Results
Polyamines synergistically enhance the efficacy of KRAS inhibitors in a KEAP1-dependent manner
The reprogramming of cellular metabolism to satisfy the uncontrolled energy demands of proliferation is a major hallmark of cancer25,26, particularly in tumors harboring KRAS mutations27,28. This metabolic reprogramming modulates various oncogenic signaling pathways and presents an opportunity to exploit KRAS-associated metabolic vulnerabilities to enhance the efficacy of KRAS-targeted therapies29. By integrating a comprehensive metabolic library comprising 889 exogenous human metabolites with the KRAS-targeted inhibitors sotorasib and adagrasib (Fig. 1A), we pinpointed polyamine metabolism components—arginine, ornithine, spermidine, and spermine—as significant sensitizers of KRAS inhibitors in MIAPACA2 cells (Fig. 1B).
A Flow chart of the metabolic library screening process. Created in BioRender. Bian, Y. (2025) https://BioRender.com/3p0ref1. B Relative viability of metabolic-treated vs vehicle (DMSO)-treated MIAPACA2 cells showed the metabolic library screening results. MIAPACA2 cells were treated with KRAS G12C inhibitors sotorasib or adagrasib (MIAPACA2, sotorasib, IC50 = 40 nM; adagrasib, IC50 = 60 nM), along with metabolics contained in the library (MCE HY-L030) or vehicle (DMSO) for 72 h. The algorithm for “Relative viability” showed that “X” means a specific metabolic, “V” means vehicle (DMSO). C Relative viabilities of MIAPACA2 and H23 cells treated with gradient concentrations of spermine and KRASG12C inhibitors (sotorasib or adagrasib) for 72 h were shown. 2D surface response for cell viability and 3D surface Bliss synergy score were shown. (n = 3 independent experiments). D Relative viability of 15 pan-cancer KRASG12C mutant cell lines treated with vehicle (DMSO), spermine (100 nM), sotorasib (IC50), and combination (spermine and sotorasib) for 72 h. The cell lines including pancreatic adenocarcinoma cell line MIAPACA2; bladder urothelial carcinoma cell line UMUC3; esophageal squamous cell carcinoma cell line KYSE410; lung adenocarcinoma cell lines H23, H2122, LU99, SW1573, H2030, H1792, H358, LU65 and HCC44; ovarian epithelial tumor cell line OV56; and colorectal adenocarcinoma cell lines JVE-015 and SW1463. (n = 3 independent experiments) Data were analyzed by two-tailed Student’s t-test and were presented by mean ± SD. E Venn plot displayed two genes (KRAS and KEAP1) from the intersection of mutant genes in non-synergy effect cell lines, including H23, H2122, H2030, H1792, and HCC44. F Workflow of KRASG12C mutant PAAD and LUAD patient-derived organoids (PDOs) establishment. G The number of PAAD and LUAD organoids, accompanied by an informational schematic that includes the patients’ type of mutation and differentiation stage were shown. H, I Representative brightfield images of PAAD PDOs (from PAAD patient #1 and #4) (H) and LUAD PDOs (from LUAD patient #1 and #2) (I) treated with KRASG12C inhibitors (sotorasib, 300 nm; adagrasib, 300 nM), spermine (1000 nM) or combination for 72 h (scale bars, 100 μm). J–K Luminescence measurement showed the relative viability of PAAD PDOs (from PAAD patient #1 and #4) (J) and LUAD PDOs (from LUAD patient #1 and #2) (K) treated with KRASG12C inhibitors (sotorasib, 300 nm; adagrasib, 300 nM), spermine (1000 nM) or combination for 72 h. (n = 3 independent experiments) Data were analyzed by two-tailed Student’s t-test and were presented by mean ± SD. Source data are provided as a Source Data file.
Given the critical role of polyamine metabolism in tumor proliferation and ferroptosis, we further evaluated these sensitizing effects across diverse cell lines besides MIAPACA2. Intriguingly, synergistic effect was also observed in H1373 cells, while no synergistic interaction was detected in H23 cells when spermine or spermidine was administered alongside KRAS inhibitors (Fig. 1C, Fig. S1A, B). Similarly, assessment of arginine and ornithine sensitization levels showed no significant effects in H23 cells. Furthermore, exogenous putrescine and cadaverine supplementation supplementation exhibited moderate sensitization in MIAPACA2 cells but no synergistic effects in H23 cells, likely due to the low endogenous levels of putrescine and cadaverine. (Fig. S1C, D). Upon extending our investigation to a set of 15 KRASG12C-mutant cell lines, we observed sensitization in ten lines (demonstrating a synergy effect) and no sensitization in the remaining five (Fig. 1D, Fig. S2A).
We hypothesized that co-occurring gene mutations might determine the sensitizing effects of polyamines. Analysis of mutational profiles from the CCLE database (https://portals.broadinstitute.org/ccle) revealed that polyamine supplementation significantly potentiated KRAS inhibitors only in KRASG12C/KEAP1WT cell lines (including MIAPACA2, JVE015, KYSE410, OV56, LU99, SW1573, H358, LU65, UMUC3, SW1463 and H1373), but not in KRASG12C/KEAP1MU cell lines (such as H23, H2122, H2030, H1792, and HCC44) (Fig. 1D, E, Fig. S2A). Notably, mutations in LKB1, cyclin dependent kinase inhibitor 2 A (CDKN2A), and tumor protein p53 (TP53) were inconsequential. Additionally, we validated the polyamine-mediated sensitization effects in the newly approved KRASG12C inhibitors fulzerasib and garsoasib (Fig. S2B, C). Validation in MIAPACA2, H23, HPAC (KRASG12D/KEAP1WT), and HEC1 (KRASG12D/KEAP1MU) cells demonstrated that the sensitizing effect of polyamines extended to the pan-KRAS inhibitor BI-2493 and the KRASG12D inhibitor MRTX1133 (Fig. S2D, E).
To address potential biases arising from cell line heterogeneity and two-dimensional growth conditions, we established patient-derived organoids (PDOs) from KRASG12C pancreatic and lung tumors to validate the impact of polyamines on KRAS-targeted therapy (Fig. 1F, G, Fig. S2F). Consistent with our previous findings, spermidine sensitized KRAS inhibitors in KRASMU/KEAP1WT organoids, but not in KRASMU/KEAP1MU organoids (Fig. 1H–K, Fig. S2G-H).
To further dissect the role of KEAP1 status in polyamine-mediated sensitization, gain- and loss-of-function experiments were performed (Fig. S2I). The results showed that knocking out KEAP1 in MIAPACA2 and H1373 (KRASMU/KEAP1WT) cells abolished the sensitizing effect of polyamines, whereas reintroduction of KEAP1 into H23 (KRASMU/KEAP1MU) cells restored the sensitizing effect (Fig. 2A, Fig. S3A, B). Collectively, these results underscore the pivotal role of KEAP1 mutation status in polyamine-induced sensitization to KRAS inhibitors.
A Relative viabilities of MIAPACA2 with KEAP1 knockout (KO) and H23 cells with KEAP1 overexpression (OE) treated with gradient concentrations of spermine and sotorasib or adagrasib for 72 h. 2D surface response for cell viability and 3D surface Bliss synergy response were shown. (n = 3 independent experiments). B Schematic depiction of polyamine metabolism. Created in BioRender. Bian, Y. (2025) https://BioRender.com/3p0ref1. C The volcano plots showed differentially expressed genes (DEGs) in MIAPACA2 and H23 cells treated with KRASG12C inhibitors (sotorasib: MIAPACA2, IC50 = 40 nM; H23, IC50 = 500 nM; adagrasib: MIAPACA2, IC50 = 60 nM; H23, IC50 = 100 nM, 72 h) by RNA-seq analysis. ODC1, SAT1, SMOX, and PAOX are critical in polyamine metabolism. “DN” means downregulated in the KRAS inhibitors treatment group, and “UP” means upregulated in the KRAS inhibitors treatment group (|log2FC | > 0.4 and p < 0.05). D qRT-PCR and WB verified the mRNA and protein expression level of ODC1, SAT1, SMOX, and PAOX in MIAPACA2 and H23 cell lines treated with sotorasib or adagraisb (IC50) for 72 h. (n = 3 independent experiments) Data were analyzed by two-tailed Student’s t-test and were presented by mean ± SD. Source data are provided as a Source Data file. E Polyamine metabolites level in MIAPACA2 and H23 cells following KRAS inhibitors treatment (IC50, 72 h). (n = 3 independent experiments) Data were analyzed by two-tailed Student’s t-test and were presented by mean ± SD. Source data are provided as a Source Data file. F Polyamine metabolites level in MIAPACA2 (NC and SAT1-KO) and H23 (NC and SAT1-OE) cells. (n = 3 independent experiments) Data were analyzed by two-tailed Student’s t-test and were presented by mean ± SD. Source data are provided as a Source Data file. G Relative viabilities of MIAPACA2 cells (NC and SAT1-KO group) and H23 cells (NC and SAT1-OE group) treated with spermine (100 nM) or spermidine (100 nM), KRAS inhibitors (IC50), or combination for 72 h. (n = 3 independent experiments) Data were analyzed by two-tailed Student’s t-test and were presented by mean ± SD. Source data are provided as a Source Data file. H qRT-PCR and WB showed the expression level of SAT1 in MIAPACA2 cells (NC, KEAP1-KO, KEAP1-WT, and KEAP1-MU group) and H23 cells (NC and KEAP1-OE group). (n = 3 independent experiments) Data were analyzed by two-tailed Student’s t-test and were presented by mean ± SD. Source data are provided as a Source Data file. Source data are provided as a Source Data file.
KEAP1 WT-dependent SAT1 upregulation enhances the synergy between polyamines and KRAS inhibitors
To pinpoint the metabolic step influencing polyamine sensitization to KRAS inhibitors, we conducted RNA-seq analysis on MIAPACA2 and H23 cells treated with sotorasib and adagrasib (IC50, 72 h), and several crucial enzymes involved in polyamine metabolism, namely ornithine decarboxylase 1 (ODC1), SAT1, PAOX, and spermine oxidase (SMOX) (Fig. 2B), were examined. Our results revealed SAT1, the key enzyme in polyamine catabolism, exhibited divergent alterations in cell lines with distinct KEAP1 status: it was upregulated in MIAPACA2 and H1373 cells and downregulated in H23 cells following the treatment of KRAS inhibitors. Meanwhile, ODC1, essential for polyamine synthesis, was consistently downregulated in both cell lines after treatment (Fig. 2C, D, Fig. S3C). Metabolite analysis further revealed reduced levels of spermidine, spermine, N1-acetylspermine, and N1-acetylspermidine (spermine and spermidine are acetylated in their N1 positions) upon KRAS inhibitor administration (Fig. 2E), a finding validated in PDOs (Fig. S3D). Notably, KRAS inhibitor-treated H23 cells exhibited a more pronounced reduction in N1-acetylspermine and N1-acetylspermidine compared to MIAPACA2 cells (Fig. 2E). Coupled with their direct association with SAT1 (Fig. 2B), cell line-specific SAT1 expression divergence, stable PAOX levels, and reduced or unchanged SMOX levels (Fig. 2C, D, Fig. S3C), these data collectively suggest SAT1 as the central regulator of the differential cellular response to KRAS inhibitors.
To validate this hypothesis, we first validated that SAT1 enzymatic activity changes corresponded with its mRNA and protein level alterations (Fig. S3E), then generated SAT1 knockout (SAT1-KO) MIAPACA2 and H1373 cells, as well as SAT1 overexpressing (SAT1-OE) H23 cells (Fig. S3F). Metabolomics identified SAT1 as the key regulator of N1-acetylspermine/spermidine alterations (Fig. 2F). To eliminate the effects of ODC1 alterations, we performed exogenous polyamine supplementation to validate the critical role of SAT1. As expected, exogenous polyamine supplementation failed to enhance KRAS inhibitors sensitivity in SAT1-KO MIAPACA2 and H1373 cells, in contrast to the negative control (NC) MIAPACA2 and H1373 cells. Conversely, in SAT1-OE H23 cells, polyamine supplementation instead enhanced the efficacy of KRAS inhibitors, confirming the crutial role of SAT1 upregulation in polyamine-mediated sensitization to KRAS-targeted therapy (Fig. 2G, Fig. S3G).
Our findings thus far establish that polyamine sensitization to KRAS inhibitors is contingent upon KEAP1 mutation status, and the contrasting SAT1 expression patterns in different KEAP1 status backgrounds underlie the observed variations in sensitizing effects. This prompted us to further investigate KEAP1’s role in modulating SAT1 expression. In MIAPACA2 and H1373 cells (KRASMU/KEAP1WT), knocking out KEAP1 significantly abolished the KRAS inhibitors-induced upregulation in SAT1 expression. Conversely, in KEAP1-deficient H23 cells, restoring KEAP1 function via overexpression prevented KRAS inhibitor-mediated SAT1 downregulation (Fig. 2H Fig. S3H). These observations elucidate the inability of polyamines to sensitize KRAS inhibitors in KEAP1-KO MIAPACA2 and H1373 cells while enhancing their effect in KEAP1-OE H23 cells. Notably, in the absence of inhibitors, KEAP1 knockout significantly elevated SAT1 expression in MIAPACA2 and H1373 cells, whereas KEAP1 overexpression decreased SAT1 levels in H23 cells (Fig. 2H, Fig. S3H). Furthermore, through rescue experiments reconstituting both KEAP1-WT and KEAP1-MU, the results demonstrated that KEAP1-MU similarly significantly increased the SAT1 levels in KEAP1-WT, confirming the critical regulatory role of KEAP1 on SAT1(Fig. 2H). This indicates a negative regulatory relationship between KEAP1 and SAT1 that is reversed upon KRAS inhibitors treatment, suggesting a more complex KRAS-KEAP1-SAT1 regulatory axis awaits exploration.
SAT1-mediated polyamine catabolism enhances KRAS inhibition through ferroptosis
Previous studies have demonstrated that SAT1-regulated polyamine catabolism primarily exerts its cytotoxic effects by promoting ferroptosis, which also plays a key role in the efficacy of KRAS inhibitors18,21,22. In this study, we treated MIAPACA2 and H23 cells with a combination of ferroptosis inhibitors—ferrostatin-1 (fer-1) and deferoxamine (DFO)—alongside with the apoptosis inhibitor Z-VAD-FMK (Z-VAD) and the necrosis inhibitor necrosulfonamide (necro) for 72 h, in conjunction with KRAS inhibitors. Our findings revealed that ferroptosis inhibitors significantly attenuated the cell death induced by KRAS inhibitors (Fig. S4A). Concurrently, elevated levels of malondialdehyde (MDA) further substantiated the role of ferroptosis in the treatment of KRAS inhibitors (Fig. S4B). Furthermore, the lack of synergistic sensitization between KRAS inhibitors and ferroptosis inducers (RSL3/IKE) likely stems from mechanistic overlap in GPX4-GSH axis disruption (Fig. S4C). This redundancy underscores SAT1’s engagement of an alternative ferroptosis pathway via H2O2 overproduction, as previously demonstrated22,30. Exogenous polyamine supplementation increased H2O2 production, which was abolished by SAT1-KO and PAOX-KO (Fig. S4D–F). The detection of PAOX enzymatic activity further indicates that PAOX mediates the oxidation of N¹-acetylspermidine and N¹-acetylspermine (Fig.z S4G).These findings indicate that spermine/spermidine undergo SAT1-dependent acetylation followed by PAOX-mediated oxidation to generate H2O2, which subsequently reacts with iron via the Fenton reaction to promote ferroptosis. Therefore, we hypothesized that SAT1-mediated polyamine catabolism might augment the sensitivity of KRAS-targeted therapies by facilitating ferroptosis.
Our results revealed that in MIAPACA2 cells, polyamines potentiated the efficacy of KRAS inhibitors, an effect exclusively abrogated by ferroptosis inhibitors (Fig. 3A). Notably, several hallmarks of ferroptosis, including MDA (Fig. 3B), lipid peroxidation (Fig. 3C), and intracellular ferrous iron levels (Fig. 3D), and were markedly elevated when KRAS inhibitors were administered in combination with polyamines, as compared to KRAS inhibitors alone, which were validated in H1373 cells (Fig. S4H, I). Transmission electron microscopy analysis further disclosed more pronounced mitochondrial shrinkage and cristae disappearance in cells receiving the combination treatment (Fig. 3E, F). These observations demonstrated in MIAPACA2 cells (KRASMU/KEAP1WT), KRAS inhibitors upregulate SAT1 expression, and polyamine supplementation further increases polyamine catabolism and ferroptosis, thereby amplifying the cytotoxic effect of KRAS inhibitors and elevating drug sensitivity. Conversely, in H23 cells (KRASMU/KEAP1MU), which exhibit downregulated SAT1 after the treatment of KRAS inhibitors, polyamine supplementation failed to augment the effects of KRAS inhibitors, and ferroptosis markers remained essentially unchanged.
A Relative viabilities in MIAPACA2 cells treated with spermine (100 nM) or spermidine (100 nM) combined with or without DFO (5000 nM), fer-1 (500 nM), z-VAD (500 nM), or necrosulfonamide (250 nM) under treatment with sotorasib or adagrasib. (n = 3 independent experiments) Data were analyzed by two-tailed Student’s t-test and were presented by mean ± SD. B–D MDA (B), lipid peroxidation (C), and ferrous iron (D) level of MIAPACA2 and H23 cells treated with spermine (100 nM) or spermidine (100 nM), sotorasib or adagrasib (IC50) and combination for 72 h. (n = 3 independent experiments) Data were analyzed by two-tailed Student’s t-test and were presented by mean ± SD. E Transmission electron microscopy images of MIAPACA2 and H23 cells treated with spermine (100 nM) or spermidine (100 nM), sotorasib or adagrasib (IC50) and combination for 72 h (scale bars, 5 μm). F Number of damaged mitochondria per MIAPACA2 or H23 cell with cristae disappearance and mitochondrial shrinkage. (n = 3 independent experiments) Data were analyzed by two-tailed Student’s t-test and were presented by mean ± SD. G Relative enzymatic activity of SAT1 in H23 NC cells, SAT1-OE cells with or without critical enzymatic site mutation (Tyrosine 140 replaced by phenylalanine, Y140F).(n = 3 independent experiments) Data were analyzed by two-tailed Student’s t-test and were presented by mean ± SD. H, I MDA (H), lipid peroxidation (I) level of MIAPACA2 cells with SAT1-KO and H23 cells with SAT1-OE or SAT1-Y240F treated with spermine (100 nM) or spermidine (100 nM), sotorasib or adagrasib (IC50) and combination for 72 h. (n = 3 independent experiments) Data were analyzed by two-tailed Student’s t-test and were presented by mean ± SD. J Transmission electron microscopy images of MIAPACA2 cells with SAT1-KO and H23 cells with SAT1-OE cells treated with spermine (100 nM) or spermidine (100 nM), sotorasib or adagrasib (IC50) and combination for 72 h (scale bars, 5 μm). K Number of damaged mitochondria per MIAPACA2 SAT1-KO or H23 SAT1-OE Cell with cristae disappearance and mitochondrial shrinkage. (n = 3 independent experiments) Data were analyzed by two-tailed Student’s t-test and were presented by mean ± SD. L Ferrous iron level of MIAPACA2 cells with SAT1-KO and H23 cells with SAT1-OE treated with spermine (100 nM) or spermidine (100 nM), sotorasib or adagrasib (IC50) and combination for 72 h. (n = 3 independent experiments) Data were analyzed by two-tailed Student’s t-test and were presented by mean ± SD. Source data are provided as a Source Data file.
To further validate the critical role of SAT1, we examined SAT1-KO MIAPACA2 and SAT1-OE H23 cells with or without critical enzymatic site mutation (Y140F)31 (Fig. 3G). SAT1-KO MIAPACA2 cells displayed no alterations in ferroptosis markers upon combined treatment, contrasting sharply with SAT1-OE H23 cells, which exhibited increased MDA (Fig. 3H), lipid peroxidation (Fig. 3I), mitochondrial shrinkage, cristae disappearance (Fig. 3J, K) and intracellular ferrous iron level (Fig. 3L), whereas catalytic inactivation of SAT1 (Y140F) abolished polyamine-enhanced KRAS inhibitor effects in SAT1-OE H23 cells, despite equivalent SAT1 mRNA expression. This definitively establishes SAT1’s enzymatic activity as essential for polyamine-driven ferroptosis. Prior studies suggest fetal bovine serum contains amine oxidase (BSAO) that oxidizes polyamines to ammonia and aldehyde32. However, low-dose polyamines alone showed no cytotoxicity, and combining aminoguanidine (BSAO inhibitor) with polyamines maintained ferroptosis sensitization, demonstrating SAT1-mediated effects rather than oxidative byproduct mechanisms (Fig. S4J). These findings collectively reinforce the essential role of SAT1 upregulation in promoting polyamine catabolism-induced ferroptosis and enhancing sensitivity to KRAS-targeted therapies in KRASMU/KEAP1WT tumors. In KRASMU/KEAP1MU tumors, since SAT1 expression is inhibited by the treatment of KRAS inhibitors, prior overexpression of SAT1 is required for the sensitization.
Overcoming drug resistance through SAT1-mediated polyamine metabolism and ferroptosis
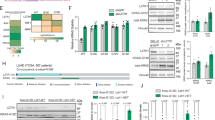
To further substantiate these findings, we generated drug-resistant cell lines by exposing them to sotorasib or adagrasib for seven months, resulting in resistant lines (denoted as MIAPACA2-Sotorasib-R, MIAPACA2-Adagrasib-R, H23-Sotorasib-R, and H23-Adagrasib-R) with IC50s over 10-fold higher than parental cells (Fig. 4A, B). In KRASG12C inhibitor-resistant MIAPACA2 cells, polyamine supplementation persisted in enhancing the potency of KRAS inhibitors. Conversely, this effect was conspicuously absent in KRAS inhibitor-resistant H23 cells (Fig. 4C). RNA-seq analysis of these resistant cells mirrored the expression trends of key enzymes observed in parental cells treated with KRASG12C inhibitors for 72 h (Fig. 4D). SAT1 was markedly upregulated in resistant MIAPACA2 cells (KRASMU/KEAP1WT) and significantly downregulated in resistant H23 cells (KRASMU/KEAP1MU) (Fig. 4E, F). Metabolomic analysis revealed decreased levels of putrescine, spermidine, and spermine in MIAPACA2 and H23 resistant cells compared to parental cells, consistent with downregulated ODC1 expression. Notably, following KRAS inhibitors treatment, H23 cells exhibited a more pronounced reduction in N1-acetylspermine and N1-acetylspermidine levels, whereas MIAPACA2 cells showed a more marked decrease in putrescine, spermidine, and spermine compared to controls, aligning with differential SAT1 expression patterns (Fig. 4G), underscoring the critical role of SAT1 in determining the impact of polyamine supplementation on KRAS inhibitors in resistant cells.
A Stepwise dose escalation of sotorasib (from 0 to 6400 nM) or adagrasib (from 0 to 2560 nM) throughout the process to generate KRASG12C inhibitors-resistant cell lines (MIAPACA2-Sotorasib-R, MIAPACA2-Adagrasib-R, H23-Sotorasib-R, and H23-Adagrasib-R). Created in BioRender. Bian, Y. (2025) https://BioRender.com/3p0ref1. B Relative viability of parental and four resistant cells incubated in increasing concentrations of sotorasib or adagrasib for 72 h (MIAPACA2-sotorasib-R, IC50 = 1020 nM; MIAPACA2-adagrasib-R, IC50 = 1420 nM; H23-sotorasib-R, IC50 > 5000 nM; H23-adagrasib-R, IC50 = 2338 nM). (n = 3 independent experiments) Data were analyzed by two-way ANOVA test. C Relative viabilities of MIAPACA2 and H23 resistant cells treated with gradient concentrations of spermine and sotorasib or adagrasib for 72 h. 2D surface response for cell viability and 3D surface Bliss synergy response were shown. (n = 3 independent experiments). D The volcano plots showed DEGs in MIAPACA2-Sotorasib-R, MIAPACA2-Adagrasib-R, H23-Sotorasib-R, and H23-Adagrasib-R cells versus parental cells by RNA-seq analysis. “DN” means downregulated in KRASG12C inhibitors resistant cells, and “UP” means upregulated in KRASG12C inhibitors resistant cells (|log2FC | > 0.4 and p < 0.05).E, F qRT-PCR (E) and WB (F) verified the expression level of ODC1, SAT1, SMOX, and PAOX in KRASG12C inhibitors-resistant MIAPACA2 and H23 cell lines. (n = 3 independent experiments) Data were analyzed by two-tailed Student’s t-test and were presented by mean ± SD. G The heatmap displayed the polyamine metabolites level in MIAPACA2 and H23 resistant cells following KRAS inhibitors treatment (IC50, 72 h). H Lipid peroxidation level of KRASG12C inhibitors-resistant MIAPACA2 and H23 cells treated with spermine (100 nM), sotorasib or adagrasib (IC50 in resistant cells), and combination for 72 h. (n = 3 independent experiments) Source data are provided as a Source Data file.
Furthermore, polyamine supplementation in resistant cells enhanced KRAS inhibitors’ efficacy by promoting ferroptosis. In resistant MIAPACA2 cells, combined treatment increased lipid peroxidation (Fig. 4H), MDA level (Fig. S5A), and intracellular ferrous iron level (Fig. S5B) compared to KRAS inhibitors monotherapy, while in H23 resistant cells, polyamine supplementation showed no such effect. These findings further support that SAT1-mediated polyamine catabolism promotes ferroptosis and sensitizes KRAS-targeted therapy in a KEAP1-dependent manner, even in drug-resistant cells, offering a potential solution to address drug resistance.
KRAS inhibitors modulate SAT1 expression through JNK/NRF2/SAT1 and JNK/c-Jun/SAT1 pathways
Given the differential regulation of SAT1 expression by KRAS inhibitors in MIAPACA2 and H23 cells, contingent on their KEAP1 mutation status, we delved into the intricate mechanisms linking KRAS, KEAP1, and SAT1. KEAP1 is primarily recognized for its role in promoting the ubiquitination and subsequent degradation of NRF233,34, therefore, the expression of NRF2 was first assessed. Our examination of NRF2 expression levels revealed a notable finding: in H23 cells (KRASMU/KEAP1MU), NRF2 expression was significantly reduced upon KRAS inhibitors treatment, a pattern that paralleled the downregulation of SAT1 (Fig. 5A). Our findings align with prior reports demonstrating NRF2-mediated regulation of SAT135,36.
A WB showed the expression level of SAT1 and the involved pathway factors, including JNK and c-Jun, NRF2, as well as KEAP1 in MIAPACA2 (NC and KEAP1-KO group) and H23 (NC and KEAP1-OE group) cells treated with vehicle (DMSO) or KRAS inhibitors (IC50) for 72 h. (n = 3 independent experiments). B WB displayed the change in the expression level of SAT1 and the involved pathway factors, including JNK and c-Jun, NRF2 and KEAP1 in MIAPACA2 (NC and KEAP1-KO group) and H23 (NC and KEAP1-OE group) cells treated with vehicle (DMSO) or KRAS inhibitors, under treatment of JNK inhibitor JNK-IN-8 (10 nM). (n = 3 independent experiments). C Genome-wide data of SAT1 from the ENCODE database showed the c-Jun and NRF2 binding peak within the promotor region of SAT1. D ChIP-qPCR experiments showed the binding intensities of c-Jun and NRF2 to the promotor region of SAT1 in MIAPACA2 and H23 cells (n = 3 independent experiments). Data were analyzed by two-tailed Student’s t-test and were presented by mean ± SD. E WB showed the results of DNA pull down assays of SAT1. F Dual-luciferase assays showed the fluorescence intensity in four groups: a wild-type control lacking any mutations, a mutant with disrupted putative NRF2 binding sites, a mutant with disrupted putative c-Jun binding sites, and a double mutant with both NRF2 and c-Jun putative binding sites disrupted in MIAPACA2 and H23 cells treated with vehicle (DMSO) or KRAS inhibitors (IC50) (n = 3 independent experiments). Data were analyzed by two-tailed Student’s t-test and were presented by mean ± SD. G The immunofluorescence images of KEAP1, p-JNK, p-c-Jun, NRF2, and SAT1 in MIAPACA2 cells (NC and KEAP1-KO group) treated with KRAS inhibitors (IC50) for 72 h (scale bars, 50 μm). H Relative fluorescence intensity of KEAP1, p-JNK, p-c-Jun, NRF2, and SAT1 in MIAPACA2 cells (NC and KEAP1-KO group) treated with KRAS inhibitors (IC50) for 72 h. (n = 3 independent experiments) Data were analyzed by two-tailed Student’s t-test and were presented by mean ± SD. Source data are provided as a Source Data file.
To delve deeper into the mechanisms underlying NRF2 downregulation in KRAS inhibitors-treated H23 cells, particularly given the mutated KEAP1 status, we explored alternative pathways reported to significantly mediate NRF2 degradation, including c-Jun N-terminal protein kinase (JNK)33,37 and nuclear factor kappa-B (NFκB)38,39,40. Through the application of JNK inhibitors (JNK-IN-8 and JNK-IN-11) and NFκB inhibitors (MYLS22 and BI-69A11), we discovered that only JNK inhibitors but not NFκB inhibitors effectively abrogated the SAT1 downregulation observed with KRAS inhibitors in H23 cells, implicating JNK in the control of the NRF2/SAT1 axis (Fig. S5C).
We also detected c-Jun, the major downstream effector of JNK41,42, and observed that both phosphorylated JNK (p-JNK) and phosphorylated c-Jun (p-c-Jun) were upregulated in both cell lines following KRAS inhibition (Fig. 5A). This upregulation hinted that JNK/c-Jun might contribute to SAT1 upregulation in MIAPACA2 cells, which exhibit low NRF2 expression under the regulation of wild-type KEAP1. Conversely, in H23 cells with high NRF2 expression due to KEAP1 mutation, JNK-mediated NRF2 degradation predominantly led to the downregulation of the NRF2/SAT1 axis and SAT1 expression (Fig. 5A). Furthermore, JNK inhibition reversed the SAT1 upregulation observed in MIAPACA2 cells while it prevented NRF2 degradation and subsequent SAT1 reduction in H23 cells treated with KRAS inhibitors, confirming the pivotal role of JNK in this regulatory network (Fig. 5B). Additionally, NRF2 knockout in H23 cells resulted in SAT1 upregulation upon KRAS inhibition, reinforcing the promotional effect of the JNK/c-Jun pathway on SAT1 expression (Fig. S5D). Consistent results were obtained in MIAPACA2 KEAP1-KO cells (which behave similarly to H23) and H23 KEAP1-OE cells (which behave similarly to MIAPACA2), providing further robust support for our proposed mechanism (Fig. 5A).
Furthermore, ChIP-Seq data obtained from the Encode database unveiled significant binding peaks for both NRF2 and c-Jun at the SAT1 promoter region, suggesting their direct role in promoting SAT1 transcription (Fig. 5C). This regulatory relationship was further validated through ChIP-qPCR and DNA pull-down assays. In MIAPACA2 cells with wild-type KEAP1, KRAS inhibitors augmented c-Jun binding to the SAT1 promoter, while NRF2 binding remained unaffected. Conversely, in H23 cells harboring KEAP1 mutation, KRAS inhibitors increased c-Jun binding but decreased NRF2 binding at the SAT1 promoter (Fig. 5D, E). Luciferase reporter assays also demonstrated that KRAS inhibitors enhanced SAT1 transcription in MIAPACA2 cells but suppressed it in H23 cells, and these effects were altered by mutations in the NRF2/c-Jun binding sites within the SAT1 promoter.
Specifically, mutating the NRF2-binding site in MIAPACA2 cells did not affect KRAS inhibitor responses, whereas it enhanced their efficacy in H23 cells. Mutating the c-Jun binding site reduced the effects of KRAS inhibitors in MIAPACA2 cells. Double mutation of both binding sites abolished the effects of KRAS inhibitors in both cell lines (Fig. 5F). Collectively, these results indicate that KRAS inhibitors and the KEAP1 mutation status influence NRF2/c-Jun binding to the SAT1 promoter, thereby modulating its transcription. Finally, immunofluorescence experiments confirmed the distinct regulatory mechanisms operating in these two cell types (Fig. 5G, H, Fig. S5E, F).
Precisely targeting SAT1-mediated polyamine metabolism to enhance KRAS inhibitors in PDOs, xenograft, and spontaneous lung cancer mouse models
Using PDOs, cell-derived xenografts, and KRASG12C spontaneous lung cancer mouse models that closely mimic the heterogeneity of human tumors and the in vivo microenvironment, we further validated the efficacy of different strategies targeting SAT1-mediated polyamine catabolism in KRAS-mutant tumors with differing KEAP1 statuses. Firstly, we treated PAAD and LUAD organoids with lentivirus to manipulate SAT1 expression (Fig. 6A). SAT1 was knocked out in organoids derived from patients with KRASMU/KEAP1WT (PAAD patient #1 and LUAD patient #1), while it was overexpressed in organoids from KRASMU/KEAP1MU patients (PAAD patient #4 and LUAD patient #2) (Fig. 6B). The results indicated that in KRASMU/KEAP1WT organoids, polyamine supplementation increased MDA levels, suggesting enhanced ferroptosis due to SAT1 upregulation, which in turn increased the efficacy of KRAS-targeted drugs. This effect was blocked by SAT1 depletion. However, in KRASMU/KEAP1MU organoids, polyamine supplementation alone did not enhance sensitivity to KRAS inhibitors. Only under SAT1 overexpression did polyamine catabolism increase significantly, promoting ferroptosis and improving the efficacy of KRAS-targeted drugs (Fig. 6C, D).
A Workflow of the establishment of PDOs with SAT1 KO or SAT OE. Created in BioRender. Bian, Y. (2025) https://BioRender.com/3p0ref1. B WB showed the effect of SAT1 KO and SAT1 OE in PAAD and LUAD PDOs. C Luminescence measurement showed the relative viability of PAAD and LUAD PDOs (NC and SAT1-KO or SAT1-OE group) treated with spermine (100 nM), sotorasib or adagrasib (300 nM), and combination for 72 h. (n = 3 independent experiments) Data were analyzed by two-tailed Student’s t-test and were presented by mean ± SD. D MDA level of PAAD and LUAD PDOs (NC and SAT1-KO or SAT1-OE group). (n = 3 independent experiments) Data were analyzed by two-tailed Student’s t-test and were presented by mean ± SD. E Schematic design of xenograft tumor assay. Created in BioRender. Bian, Y. (2025) https://BioRender.com/3p0ref1. F–H Tumor images (F), growth curves (G) (Data were analyzed by two-way ANOVA test.), and tumor weight at the end of the treatment (H) (Data were analyzed by two-tailed Student’s t-test.) of resected tumors from MIAPACA2 cells. (n = 5 mice/group, one experiment) (I–K) Tumor images (I), growth curves (J) (Data were analyzed by two-way ANOVA test.), and tumor weight at the end of the treatment (K) (Data were analyzed by two-tailed Student’s t-test.) of resected tumors from H23 cells. (n = 5 mice/group, one experiment) (L) Flow chart of spontaneous lung cancer mouse model with KRASG12C mutation. Created in BioRender. Bian, Y. (2025) https://BioRender.com/3p0ref1. M Survival analysis of spontaneous lung cancer mice. N CT scan images of lung cancer in mice models treated with vehicle, spermine, sotorasib, or combination (prior group and post-treatment group). Data were analyzed by log-rank test. Source data are provided as a Source Data file.
We further validated the precision therapy strategy by considering co-mutations in xenograft (Fig. 6E) and spontaneous lung cancer mouse models. KRAS-targeted therapy reduced endogenous spermidine and spermine in subcutaneous tumors (Fig. S5G), and exogenous polyamine supplementation modulated therapeutic efficacy. In nude mice with MIAPACA2 cell-derived tumors, combined polyamine and KRAS-targeted therapy significantly reduced tumor volume and weight compared to KRAS inhibitors monotherapy (Fig. 6F–H). In nude mice with H23 cell-derived tumors, AAV-mediated SAT1 overexpression followed by combined treatment also enhanced the efficacy of KRAS inhibitors (Fig. 6I–K). Additionally, in the KRASG12C mutant spontaneous lung cancer mouse model (KRASMU/KEAP1WT)(Fig. 6L), polyamine supplementation improved the efficacy of KRAS-targeted therapy. CT scans revealed near-complete tumor disappearance in the combined therapy group, with extended survival compared to the KRAS inhibitor monotherapy group (Fig. 6M, N). These findings confirm SAT1-mediated polyamine metabolism as a promising therapeutic target for precision treatments in KRAS-mutant cancers with different KEAP1 status.
Discussion
The emergence of KRAS-targeted therapies like sotorasib and adagrasib has provided promising avenues for treating KRAS-mutant cancers, but resistance remains a significant challenge43,44,45,46,47, while ferroptosis has recently been highlighted for its role in overcoming tumor treatment resistance26,48. Numerous studies have documented that KRAS-mutant tumors develop multiple metabolic pathways to evade ferroptosis and drive tumor progression. Specifically, Son et al. revealed that KRAS mutations upregulate glutamate oxaloacetate transaminases 1 and 2 (GOT1 and GOT2), which in turn enhance NADPH production via malic enzyme 1 (ME1)49. Separately, previous studies found that elevated levels of solute carrier family 7 member 11 (SLC7A11) in KRAS-mutant tumors protect cells from oxidative stress by boosting intracellular cysteine uptake, stimulating glutathione (GSH) synthesis and accelerating ROS detoxification17,50.
Concurrently, precisely targeting the vulnerabilities within these altered metabolic pathways to induce ferroptosis has emerged as a potential therapeutic strategy for KRAS-mutant tumors. For instance, Padanad et al. demonstrated that disrupting lipid metabolism reprogramming can promote ferroptosis in these tumors51. Similarly, another study revealed that targeting ALDH1A1, a key enzyme involved in aldehyde metabolism and retinoic acid biosynthesis, effectively enhances ferroptosis and boosts the efficacy of KRAS-targeted therapy. In this study, through high-content screening of a metabolite library, we identified and validated that SAT1-mediated polyamine catabolism enhances the effectiveness of KRAS-targeted therapies through ferroptosis, highlighting the promising role of ferroptosis in augmenting the therapeutic effects of KRAS inhibitors.
A growing amount of evidence indicates that polyamine metabolism plays an important role in regulating ferroptosis32,52,53. SAT1, as the rate-limiting enzyme in polyamine catabolism, acetylates spermidine and spermine to form their N1-acetylated counterparts54,55, which are subsequently oxidized by PAOX to produce H2O256, thereby facilitating ROS production, promoting cellular lipid peroxidation, and accelerating ferroptosis57. Due to the upregulation of polyamine levels in various cancer cell types, polyamine metabolism has emerged as a potential therapeutic target for cancer treatment, particularly through targeting SAT120,23,58,59,60. However, the efficacy of SAT1-targeting monotherapy is limited20, and there have been minimal advancements in combining it with other drugs to improve therapeutic outcomes. Also, the role of targeting SAT1-mediated polyamine metabolism in KRAS therapy and its relationship with KEAP1 status has not been previously reported. As PAOX-mediated oxidation generates H₂O₂, it critically regulates how tumor polyamine metabolism affects ferroptosis. It has been previously reported that PAOX expression is very low in some tumor cell lines30. In such PAOX-deficient contexts, acetylated polyamines undergo extracellular export, potentially compromising the ferroptosis-dependent efficacy enhancement by polyamine supplementation or SAT1 overexpression during targeted therapy. Using PAOX-KO models and activity assays, we demonstrate PAOX’s important role in H₂O₂ production, further refining the understanding of SAT1’s involvement in different condition. In this study, we found that SAT1 upregulation significantly promotes polyamine catabolism-induced ferroptosis and enhances sensitivity to KRAS-targeted therapies in KRASMU/KEAP1WT tumors, but prior overexpression of SAT1 is required for the sensitization to these therapies in KRASMU/KEAP1MU tumors.
Innate resistance to KRAS-targeted drugs is often due to co-mutations, which can be classified into three distinct subtypes: KRAS-LKB1/KEAP1 (KL), KRAS-CDKN2A (KC), and KRAS-TP53 (KP)61. Clinical studies have shown that the presence of co-mutations, particularly KEAP1 co-mutations, significantly correlates with drug resistance, lower objective response rates, and poorer clinical outcomes to single-agent targeted therapies62,63,64,65, highlighting the crucial to develop more precise therapeutic strategies in co-mutated patients. KEAP1 ubiquitinates and degrades NRF233,66, whereas NRF2 exerts a vital antagonistic role in ferroptosis67,68. Previous studies reported that NRF2 regulates SAT1 transcription35,36, our study further demonstrates that NRF2 significantly promotes SAT1 expression, polyamine metabolism, and subsequent ferroptosis, suggesting a multifaceted role of NRF2 in ferroptosis. Additionally, when both JNK and NFκB have been implicated in NRF2 degradation37,39,40,69, our results indicate that JNK, but not NFκB, is markedly activated by KRAS inhibitors. We found that JNK not only promotes SAT1 expression through c-Jun but also exerts a complex regulation on SAT1 expression, polyamine metabolism, and ferroptosis by facilitating NRF2 degradation. Although JNK pathway activation in response to KRAS inhibitors has been previously reported70,71, our findings provide unique insights into the intricate role of JNK in the context of co-mutations, particularly alterations in KEAP1. These observations emphasize the importance of precision therapy tailored to specific co-mutations and underscore the potential significance of our findings for developing targeted therapies.
PDOs, as three-dimensional, ex vivo cultures of viable tissues, meticulously mimic the native extracellular niche observed in vivo, providing an invaluable tool for biological research and preclinical studies, including drug sensitivity tests, disease modeling, and personalized medicine72,73. In this research, we constructed patient-derived organoids from KRASMU/KEAP1WT and KRASMU/KEAP1MU samples, validated the pivotal role of SAT1-mediated polyamine metabolism in KRAS mutant patients with different KEAP1 statuses, and elucidated the mechanism underlying the efficacy of targeted drugs via ferroptosis regulation. Furthermore, our findings were further validated in mouse models of lung spontaneous tumors, which provide an unparalleled approximation of tumor characteristics, including heterogeneity, invasiveness, metastatic potential, and the immune microenvironment of human tumors74,75. In our study, we leveraged the Cre/LoxP system to establish spontaneous lung cancer models in C57BL/6 mice harboring the KRASG12C mutation. This genetically engineered approach closely mimics the spontaneous onset of lung cancer in humans. This model allowed us to investigate the therapeutic potential of polyamine in combination with KRAS inhibitors. Recent advances highlight the health-promoting effects of polyamines. Clinical studies demonstrate their potential for improving cognitive function, neuroprotection, and anti-aging therapies76,77. Furthermore, dietary polyamine supplementation is associated with reduced mortality from cardiovascular disease and cancer78, with spermidine intake linked to prolonged survival in early-stage cancer patients78. Studies also indicate that polyamine supplementation enhances the efficacy of radiotherapy and chemotherapy in cancer treatment through inducing ferroptosis22. Our findings revealed that polyamine treatment significantly augmented the efficacy of KRAS inhibitors, leading to improved tumor control and prolonged survival. These observations underscore the potential clinical utility of polyamine-based therapeutic strategies in KRAS-mutant cancers. In summary, our results revealed that in KRASMU/KEAP1 WT cells, treatment of KRAS inhibitors activates the JNK/c-Jun pathway and SAT1 expression, and polyamine supplementation enhances the efficacy of KRAS-targeted therapy. However, in KRASMU/KEAP1MU cells, it promotes the degradation of NRF2 and inhibits SAT1 expression, necessitating SAT1 overexpression combined with polyamine treatment to augment ferroptosis and drug sensitivity (Fig. 7). Our findings underscore the significance of co-mutations and highlight SAT1-mediated polyamine metabolism as a promising therapeutic target in precision treatments for KRAS-mutant cancers with different KEAP1 statuses.
A Mechanism diagram for KRAS inhibitors regulate SAT1 expression through JNK/c-Jun or JNK/NRF2 pathway and mediated polyamine and ferroptosis in KRAS-mutant cancer with differential KEAP1 status. Created in BioRender. Bian, Y. (2025) https://BioRender.com/5ji2693.
Methods
This study complied with all ethical regulations of Zhongshan Hospital Fudan University for research involving patient samples and animal models, following review and approval by its Institutional Review Board and Institutional Animal Care and Use Committee.
Cell culture and compounds
Lung adenocarcinoma (LUAD) cell lines H2122, LU99, LU65, and HCC44 were procured from Beyotime (Zhejiang, China), ChuanQiu Biotechnology (Shanghai, China), Meisen Cell (Zhejiang, China) and Qiansi Biotechnology (Shanghai, China), respectively. Colorectal adenocarcinoma (COADREAD) cell line JVE-015 was acquired from DSMZ (Braunschweig, Germany), and SW1573 was obtained from Meisen Cell. KYSE410, an esophageal squamous cell carcinoma (ESCC) cell line, was obtained from Meisen Cell. Ovarian epithelial tumor cell line OV56 was obtained from Mlbio company (Shanghai, China). Endometrial Carcinoma (UCEC) cell line HEC1 was acquired from Immocell Biotechnology (Xiamen, China). Additional cell lines included pancreatic adenocarcinoma (PAAD) cell lines MIAPACA2 and HPAC, LUAD cell lines H23, H2030, H1792, H358 and H1373, COADREAD cell lines SW1463, and bladder urothelial carcinoma (BLCA) cell line UMUC3 all from the cell bank of the Chinese Academy of Sciences. The cells were maintained in high-glucose DMEM (Hyclone, UT, USA) supplemented with 10% fetal bovine serum (FBS, Every Green, Hangzhou, Zhejiang, China), 0.1 mg/ml streptomycin, and 100 U/ml penicillin (Sangon Biotech, Shanghai, China) in a humidified incubator with 95% air and 5% CO2.
The following compounds were obtained from TargetMol (Shanghai, China): KRASG12C inhibitors sotorasib and adagrasib, KRASG12D inhibitor MRTX1133, RSL3, imidazole ketone erastin (IKE), ferrostatin-1 (fer-1), Z-VAD-FMK (Z-VAD), deferoxamine (DFO), necrosulfonamide (necro), JNK inhibitors JNK-IN-8 and JNK-IN-11, NFkB inhibitors MYLS22 and BI-69A11, spermine and spermidine. Pan-KRAS inhibitor BI-2493 and aminoguanidine were sourced from MedChemExpress (Shanghai, China). KRASG12C inhibitors fulzerasib and garsoasib were from Selleck chemicals (Houston, USA).
The sotorasib- and adagrasib-resistant cell lines were established through continuous exposure to the respective inhibitors for seven months, followed by IC50 value measurements. Four resistant cell lines with more than a 10-fold increase in IC50 compared to parental cells were obtained: MIAPACA2-Sotorasib-R, MIAPACA2-Adagrasib-R, H23-Sotorasib-R, and H23-Adagrasib-R.
Screening of a metabolite library
A collection of 889 endogenous human metabolites obtained from MedChemExpress, was utilized for this study. MIAPACA2 cells were seeded into 96-well plates at a density of 2000 cells per well. Following a 72 h period, the cells underwent incubation with specific metabolites or DMSO at a concentration of 10 μM and KRAS inhibitors sotorasib (IC50 = 40 nM) or adagrasib (IC50 = 60 nM). Luminescent cell viability assessments were conducted, and the resultant data were recorded.
Cell viability assays
MIPACA2 cells were seeded in 96-well plates at a density of 2000 cells per well and allowed to adhere and proliferate for 24 h. (The density of H23 cells is 4000 cells per well). Cells were treated with various doses of KRAS-targeted inhibitors and spermine or spermidine, as well as RSL3, IKE, fer-1, Z-VAD, DFO, and necro for 72 h. Cell viability was assessed using the Cell Counting Kit-8 (TargetMol), with optical density (OD) measured using a microplate reader. Determination of IC50 values was achieved by employing a sigmoidal dose-response curve with a nonlinear regression model, utilizing GraphPad Prism 9 software (GraphPad Software, California, USA). For the analysis of combinatorial effects, the SynergyFinder tool79 (accessible at https://synergyfinder.org/) was leveraged.
Patient-derived organoids (PDOs) culture and drug sensitivity assays
With the approval of the ethics committee (B2022-180R), tumor tissues were harvested from patients diagnosed with PAAD and LUAD who underwent surgical resection at Zhongshan Hospital, Fudan University. Written informed consent was obtained from all subjects. In accordance with the approved protocol, no compensation was provided to participants as the review board waived this requirement. Patients’ characteristics are indicated in Fig. 1G. Analysis includes patients from both sexes with an age range of 45–72 years. The tissues were minced, thoroughly washed, and subjected to enzymatic digestion using PDO culture media kits (Absin, Shanghai, China). After centrifugation, the resulting pellet was blended with matrigel (YEASEN, Shanghai, China) in a proportion of 25:1. This mixture was then dispensed into 24-well plates and allowed to solidify at 37 °C for 10 min, leading to the formation of PDOs. Subsequently, these PDOs were preserved in 4% paraformaldehyde and processed for HE staining to facilitate histological examination. To manipulate the genetic expression of SAT1 in PDOs, CRISPR/Cas-9 technology was employed for knockout purposes, while lentiviral vectors were utilized to achieve overexpression. For the assessment of drug sensitivity, PDOs were incubated with various therapeutic agents in 96-well plates for a duration of 72 h. Cell viability was then quantified using the CellTiter-Lumi Luminescent 3D Assay Kit (Beyotime).
Transfection of lentiviruses
Lentiviral vectors targeting specific genes for knockout (KO) and overexpression (OE) were designed and constructed by Genechem Technology (Shanghai, China). These vectors included those for KEAP1-specific sgRNA (designated as KEAP1-KO), SAT1-KO, PAOX-KO and NRF2-KO, along with corresponding overexpression constructs. Lentivirus vectors mediate the overexpression of KEAP1 with of without KEAP1 mutant were constructed based on the KEAP1-KO cell lines. Lentivirus vectors mediate the overexpression of SAT with or without mutated Y to F at 140 residue of SAT were constructed (SAT-OE and SAT1-Y193F) were used to infect the H23 cell lines. These vectors were used to infect cell lines, and stable cell lines were selected using 2.5 μM puromycin (Beyotime) for 48 h. The sequences of the sgRNAs used to guide the CRISPR/Cas9 system in targeting these genes are detailed in Supplementary Table 1.
Western Blot (WB) experiments
Cellular proteins were extracted by lysing the cells in a chilled RIPA buffer (Beyotime), supplemented with a mixture of protease and phosphatase inhibitors (TargetMol). This lysis process was carried out for 10 min on ice. The protein concentration was quantified using a BCA assay kit (YEASEN).SDS-PAGE (YAMAY Biotech, Shanghai, China) was performed to separate the proteins based on their molecular weight. The separated proteins were transferred onto PVDF membranes (Millipore, Billerica, MA, USA). The membranes were blocked with 5% non-fat milk for 1 h. Following this, the membranes were incubated overnight at 4 °C with specific primary antibodies, diluted per the manufacturer’s instructions. After thorough washing with TBST solution to remove any unbound antibodies, the membranes were further incubated for 1 h with an HRP-conjugated secondary antibody (diluted 1:2500, Beyotime). The Moon Chemiluminescence Kit (Beyotime) was utilized to visualize the protein bands. A detailed list of all primary antibodies employed in this study, including their sources and dilutions, is provided in Supplementary Table 2.
RNA sequencing (RNA-seq) and differential gene analysis
RNA-seq was conducted by oebiotech (Shanghai, China), adhering to previously described protocols56. Briefly, total RNA was extracted using TRIzol reagent (Tiangen, Beijing, China) from MIAPACA2 and H23 cells treated with DMSO or sotorasib (MIAPACA2 IC50 = 40 nM, H23 IC50 = 500 nM) and adagrasib (MIAPACA2 IC50 = 60 nM, H23 IC50 = 100 nM) for 72 h. Also, total RNA was extracted from parental cells and four resistant cells (MIAPACA2-Sotorasib-R, MIAPACA2-Adagrasib-R, H23-Sotorasib-R, and H23-Adagrasib-R). As previously reported, RNA-seq was performed using the Illumina HiSeq platform (Illumina, USA)80. Differentially expressed genes (DEGs) were analyzed and visualized using the limma (two-sided testing, applying Benjamini-Hochberg correction for multiple comparisons) and ggplot2 packages in R software, applying criteria of p-value < 0.05 and |log2 fold change | > 0.5.
RNA isolation and quantitative real-time PCR (qRT-PCR)
The TRIzol reagent (Tiangen) was employed for RNA extraction, adhering strictly to the manufacturer’s recommended protocol. RNA was reverse transcribed into complementary DNA (cDNA) utilizing the Hifair® II First-strand cDNA Synthesis Kit (YEASEN). The expression levels of mRNA were quantified by quantitative real-time PCR (qRT-PCR) analysis. This was achieved using the Hieff® qPCR SYBR Green Master Mix (YEASEN) on an ABI QuantStudio 5 real-time PCR instrument (Thermo Fisher). Each sample was amplified in three replicates. The relative expression of mRNA was then normalized to the β-actin using the 2-ΔΔCT method. The primers utilized in this study were acquired from Sangon Biotech (Shanghai, China). A detailed list of these primers is provided in Supplementary Table 1.
Metabolomics analysis
Following KRASG12C inhibitors administration (sotorasib and adagrasib, IC50, 72 h), cellular and organoid samples were washed three times with ice-cold PBS to remove residual media and inhibitors, followed by metabolic quenching using 1 mL of a methanol-acetonitrile-water mixture in a 2:2:1 ratio. For subcutaneous tumor tissues, specimens were rapidly dissected within 5 min post-euthanasia, rinsed thoroughly with PBS to eliminate blood contamination, and flash-frozen in liquid nitrogen to preserve metabolic integrity. All sample types, including cells, organoids, and tumors, underwent three cycles of rapid freezing in liquid nitrogen for 1 minute, thawing at 4 °C, and vigorous vortex-mixing for 30 seconds to ensure complete cell lysis and metabolite extraction. Subsequently, samples were sonicated in an ice-water bath for 10 min to disrupt cellular debris and incubated at −40 °C for 1 h to precipitate proteins. After centrifugation at 13,800 g for 15 min at 4 °C, supernatants were collected, dried under a gentle stream of nitrogen gas to concentrate metabolites, and reconstituted in a suitable extraction solvent for a secondary sonication and centrifugation step under identical conditions. Metabolite separation and detection were performed using a Vanquish Ultra-High-Performance Liquid Chromatography system coupled to an Orbitrap Exploris 120 mass spectrometer (Thermo Fisher Scientific). Raw mass spectrometry data were converted to mzXML format using ProteoWizard software and cross-referenced with the BiotreeDB version 3.0 database for precise metabolite annotation. All detection and analytical procedures were performed by Biotree Biomedical Technology Co., Ltd (Shanghai, China) and oebiotech (Shanghai, China).
Measurement of SAT1 and PAOX enzymatic activity
Enzyme activity of SAT1 was determined by continuously monitoring the production of CoASH through its reaction with 4,4′-dithiopyridine (DTDP), which generates the 4-thiopyridone chromophore absorbing at 324 nm (ε = 19,800 M⁻¹ cm⁻¹)31. The assay mixture (1 mL final volume) contained 50 mM Tris-HCl (pH 7.5), 0.2 mM DTDP, and specified concentrations of substrates or inhibitors. Reactions were initiated by adding the enzyme, and absorbance changes were recorded at 24 °C for 1–2 min using a UVIKON XL spectrophotometer. Activity values were derived from the linear slope of absorbance versus time, with one unit defined as the amount of enzyme producing 1 μmol of CoASH per minute under the assay conditions. Enzyme activity of PAOX was detected by CheKine™ Polyamine Oxidase (PAOX) Activity Assay Kit (Abbkine, Wuhan, China) following the instructions.
Intracellular H2O2 level measurement with H2DCFDA
Intracellular H2O2 levels were quantified using the H2DCFDA-based reactive oxygen species detection kit (Beyotime) following the manufacturer’s guidelines. Cells were cultured in six-well plates and subjected to specified treatments. Prior to analysis, cells were loaded with 10 μM H2DCFDA in serum-free medium for 20 min under standard culture conditions (37 °C, 5% CO2). Unbound probes were removed by rinsing three times with serum-free medium. Fluorescence intensity was determined using a microplate reader with excitation and emission wavelengths set at 488 nm and 525 nm, respectively.
Malondialdehyde (MDA) and ferrous ion measurement
MDA level was detected by the MDA Assay Kit (Beyotime). The level of intracellular ferrous ions was determined using an Cell Ferrous Iron Colorimetric Assay Kit (Elabscience, Wuhan, China). The procedure was executed in strict adherence to the manufacturer’s guidelines. Briefly, the harvested cells were lysed and diluted to achieve a final Fe²⁺ concentration range of 0.4–35 μmol/L. A 96-well plate was configured with three experimental groups: standard wells (loaded with 80 μL of serially diluted Fe²⁺ standard solution at 100 μmol/L initial concentration), test wells (80 μL of diluted lysate), and blank control wells (80 μL of assay buffer). All wells, except the blank controls, received 80 μL of chromogenic agent, followed by thorough mixing and 10 min incubation at 37 °C protected from light. Absorbance at 593 nm was measured using a microplate reader. The standard curve was generated using the linear regression equation y = ax + b, where y represents the blank-corrected OD value, x denotes the Fe²⁺ concentration, a is the slope, and b is the intercept. Intracellular Fe²⁺ content was calculated as nmol per 10⁶ cells using the formula:
Fe²⁺ content (nmol/106)= [(Sample OD – Blank OD) – b] / a × Dilution Factor / Cell Count (per 10⁶). The relative intracellular Fe²⁺ levels were quantified as the ratio of drug-treated groups to vehicle (DMSO) controls. The quantification of malondialdehyde (MDA) similarly relies on a standardized calibration curve. A microplate reader set to an absorbance wavelength of 593 nm was utilized to acquire the optical density readings.
Lipid peroxidation measurement
Cells cultured in 12-well plates were pretreated with various agents and subsequently rinsed with PBS (Beyotime). For the assessment of lipid peroxidation level, the cells were then incubated with 4 μM BODIPY 581/591 C11 dye (Thermo Fisher Scientific) at 37 °C for 30 min. This fluorescent probe undergoes a shift in emission from red to green when oxidized by lipid-ROS81. Following incubation, the cells were washed three times with PBS and resuspended. Lipid peroxidation levels were then measured with the FITC channel of an Accuri 6 cytometer (BD Biosciences, San Diego, USA) and analyzed through FlowJo software (TreeStar Inc, OR, USA).
Transmission electron microscopy
Cells were initially seeded in 6 cm dishes and subjected to various treatments. The cells were fixed with 2.5% glutaraldehyde solution to maintain their structure and subsequently washed three times using 0.1 mol/L phosphate buffer (pH 7.4). Post-fixation was carried out with a phosphate buffer solution containing 1% osmic acid, followed by another three washes. The cells then underwent dehydration and were embedded in resin, after which they were cured in an oven at 60 °C for 48 h. Ultrathin sections of the samples were stained with a combination of uranyl acetate and lead citrate, allowed to dry overnight, and finally observed and photographed using a transmission electron microscope (Hitachi, Tokyo, Japan).
Chromatin Immunoprecipitation (ChIP) assays
ChIP assays were performed using the SimpleChIP® Plus Enzymatic Chromatin IP Kit (Cell Signaling Technology, USA), following the manufacturer’s guidelines. Briefly, cells were crosslinked with formaldehyde to stabilize DNA-protein interactions. Chromatin was then digested with micrococcal nuclease to produce DNA/protein fragments ranging from 150 to 900 bp. Immunoprecipitation was carried out with either control IgG or an anti-c-Jun antibody (1:50 dilution, cell signaling technology) and an anti-NRF2 antibody (1:50 dilution, Abways). Protein G magnetic beads were used to capture protein-DNA complexes, and the crosslinks were reversed. The isolated DNA fragments were purified using spin columns and quantified by qRT-PCR with specific primer pairs: BS-1 (F: TCTCTGTTACCCTTCCTGCCCTA, R: GGCTAGACCTCATGGATGACCAG), BS-2 (F: TGAAGAATGTACCTCGGGCAGAA, R: ATTAGACATCAGGTCGCCTCTGG), BS-3 (F: GTGCCTGGGAGAGGAAACGAT, R: AACCACCTCCTCCTACTGTTCAA).
Dual-luciferase reporter assays
Four distinct firefly luciferase reporter plasmids were constructed, each incorporating the transcriptional start region (TSR) of SAT1 with specific modifications: a wild-type control lacking any mutations, a mutant with disrupted putative NRF2 binding sites, a mutant with disrupted putative c-Jun binding sites, and a double mutant with both NRF2 and c-Jun putative binding sites disrupted. The high-probability binding sites were predicted based on consensus motifs form JASPAR database (https://jaspar.genereg.net/). These plasmids, accompanied by a Renilla luciferase normalization plasmid, were individually transfected into MIAPACA2 and H23 cells using Lipo8000 (Beyotime) reagent, which were subsequently treated with KRAS inhibitors (IC50, 72 h). Luciferase activity was measured using a Luciferase Reporter Gene Assay kit (Beyotime) according to the manufacturer’s protocol. Sequences used are available in the Supplemental Table 3.
DNA pull-down assays
DNA pull-down assays were performed as previously reported82. DNA probes corresponding to the SAT1 promoter region were synthesized by Genechem Technology and amplified using TaKaRa LA Taq® with GC Buffer (Takara Bio, Kyoto, Japan). Following amplification, probes were purified using the SanPrep Column DNA Gel Extraction Kit (Sangon Biotech). Labeled probes were incubated with streptavidin-conjugated magnetic beads (CST) for 30 min at room temperature with gentle agitation. Unbound DNA was removed through three washes with BS/THES buffer. Lysates of cell nuclei were extracted using a Nuclear and Cytoplasmic Protein Extraction Kit (Beyotime) and then added to the beads and incubated overnight at 4 °C with gentle agitation. Non-specifically bound proteins were washed away, and the eluted proteins were analyzed using the WB experiment.
Immunofluorescence staining
Cells were fixed with 4% paraformaldehyde for 15 min at room temperature and permeabilized with 0.5% Triton X-100 for 20 min. Non-specific binding was blocked with 1% bovine serum albumin (BSA) for 1 h, followed by overnight incubation at 4 °C with primary antibodies against KEAP1(1:100, Absin), p-JNK(1:100, Abways), p-c-Jun(1:100, Cell signaling technology), NRF2(1:100, Abways) and SAT1(1:100, Abcam). After three washes with PBST, cells were incubated with secondary antibodies conjugated to fluorescent dyes for 1 h at room temperature. Following additional PBST washes, cells were mounted with a DAPI-containing medium (Beyotime) and imaged using an Olympus IX71 microscope.
Xenograft experiments
Our xenograft studies were conducted to comply with the ethical standards established by the Animal Ethics Committee of Zhongshan Hospital, Fudan University (Approval No. B2023-182R). The tumor burden never exceeded the maximum allowable size of 15 mm, in accordance with the Chinese national guideline RB/T 173-2018 on humane endpoint assessment. Euthanasia was performed by carbon dioxide inhalation whenever humane endpoints were reached. These criteria included a ≥ 20% reduction in baseline body weight or a tumor volume exceeding 1,500 mm³. Tumor volumes were determined as follows: subcutaneous tumors were measured by caliper using the formula 0.5 × length × width², while spontaneous lung tumors were approximated from the largest axial CT slice using the formula 0.5 × (maximum diameter)³. The nude mice were maintained in a sterile laminar flow environment. Mice were housed under controlled conditions (21–23 °C, 45% humidity) with free access to water and a standard chow diet (LabDiet, #5053). Six-week-old male BALB/c nude mice were subcutaneously inoculated with 2 × 106 MIAPACA2 cells, genetically modified to express either NC (negative control) or SAT1-KO). Other mice were injected with H23 cells and were randomly assigned to two groups: AAV-NC (1 × 1010 vg/ml) and AAV-SAT1 (1×1010 vg/ml). Upon reaching a tumor volume of roughly 150 mm3(14 days after injection), the mice were randomly assigned to four treatment cohorts: vehicle control, sotorasib (30 mg/kg), spermine (200 mg/kg), and sotorasib plus spermine. These treatments were administered orally once daily for 2 weeks, as illustrated in Fig. 6E. Tumor dimensions were measured at biweekly intervals. Following the completion of the 2-week treatment period, all mice were humanely euthanized.
Spontaneous lung cancer mice model
Male C57BL/6 mice, aged six weeks, harboring a Cre-LoxP-mediated G12C mutant KRAS (Shanghai Model Organisms Center, Shanghai, China) were intranasally administered 5 × 1010 copies of Scgb1a1-Cre adeno-associated virus (AAV) (GENECHEM) to activate oncogenic KRASG12C expression and induce lung adenocarcinoma development. Twelve weeks post-AAV administration, mice received daily oral gavage of vehicle control or sotorasib (30 mg/kg) or spermine (200 mg/kg) or sotorasib plus spermine for four weeks. Tumor progression was monitored via computed tomography (CT) scans. Survival analysis of mice was conducted.
Statistics & reproducibility
All data were analyzed using GraphPad Prism 9 and R (version 4.1.2). Reads were aligned to genes using TopHat (v.2.0.13) and HISAT2 (v.2.1.0), with raw data normalized to fragments per kilobase of exon per million mapped reads (FPKM) for subsequent analysis. All experiments were conducted a minimum of three times, and the findings are presented as mean ± standard deviation. Continuous variables were compared using Student’s t-test or two-way ANOVA as appropriate, with p < 0.05 considered statistically significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
We used JASPAR network resource (https://jaspar.genereg.net/) to predict the binding site of NRF2 and c-Jun. We used ChIP-Seq data in the following datasets: ENCFF245VLB, ENCFF528PWS, ENCFF815NVB, ENCFF213CMB, ENCFF354ZTB, ENCFF392PIY, ENCFF690PPG, and ENCFF796HRU from ENCODE database (https://www.encodeproject.org/). Raw RNA-seq data were deposited in GSA-Human database at HRA012845. Source data are provided with this paper. The raw metabolomics data reported in this paper have been deposited in the OMIX, China National Center for Bioinformation, Chinese Academy of Sciences (https://ngdc.cncb.ac.cn/omix/release/OMIX012140). Source data are provided with this paper.
References
Kim, D., Xue, J. Y. & Lito, P. Targeting KRAS(G12C): From Inhibitory Mechanism to Modulation of Antitumor Effects in Patients. Cell 183, 850–859 (2020).
Li, S., Balmain, A. & Counter, C. M. A model for RAS mutation patterns in cancers: finding the sweet spot. Nat. Rev. Cancer 18, 767–777 (2018).
Hong, D. S. et al. KRAS(G12C) Inhibition with Sotorasib in Advanced Solid Tumors. N. Engl. J. Med 383, 1207–1217 (2020).
Hallin, J. et al. The KRAS(G12C) Inhibitor MRTX849 Provides Insight toward Therapeutic Susceptibility of KRAS-Mutant Cancers in Mouse Models and Patients. Cancer Discov. 10, 54–71 (2020).
Strickler, J. H. et al. Sotorasib in KRAS p.G12C-Mutated Advanced Pancreatic Cancer. N. Engl. J. Med 388, 33–43 (2023).
Yaeger, R. et al. Adagrasib with or without Cetuximab in Colorectal Cancer with Mutated KRAS G12C. N. Engl. J. Med 388, 44–54 (2023).
Cox, A. D., Fesik, S. W., Kimmelman, A. C., Luo, J. & Der, C. J. Drugging the undruggable RAS: Mission possible?. Nat. Rev. Drug Discov. 13, 828–851 (2014).
Zhu, C. et al. Targeting KRAS mutant cancers: from druggable therapy to drug resistance. Mol. Cancer 21, 159 (2022).
Akhave, N. S., Biter, A. B. & Hong, D. S. Mechanisms of Resistance to KRAS(G12C)-Targeted Therapy. Cancer Discov. 11, 1345–1352 (2021).
Punekar, S. R., Velcheti, V., Neel, B. G. & Wong, K. K. The current state of the art and future trends in RAS-targeted cancer therapies. Nat. Rev. Clin. Oncol. 19, 637–655 (2022).
Dilly, J. et al. Mechanisms of Resistance to Oncogenic KRAS Inhibition in Pancreatic Cancer. Cancer Discov. 14, 2135–2161 (2024).
Dixon, S. J. et al. Ferroptosis: an iron-dependent form of nonapoptotic cell death. Cell 149, 1060–1072 (2012).
Stockwell, B. R. et al. Ferroptosis: A Regulated Cell Death Nexus Linking Metabolism, Redox Biology, and Disease. Cell 171, 273–285 (2017).
Lei, G., Zhuang, L. & Gan, B. Targeting ferroptosis as a vulnerability in cancer. Nat. Rev. Cancer 22, 381–396 (2022).
Conche, C. et al. Combining ferroptosis induction with MDSC blockade renders primary tumours and metastases in liver sensitive to immune checkpoint blockade. Gut 72, 1774–1782 (2023).
Mukhopadhyay, S., Vander Heiden, M. G. & McCormick, F. The Metabolic Landscape of RAS-Driven Cancers from biology to therapy. Nat. Cancer 2, 271–283 (2021).
Hu, K. et al. Suppression of the SLC7A11/glutathione axis causes synthetic lethality in KRAS-mutant lung adenocarcinoma. J. Clin. Invest 130, 1752–1766 (2020).
Bian, Y. et al. Targeting ALDH1A1 to enhance the efficacy of KRAS-targeted therapy through ferroptosis. Redox Biol. 77, 103361 (2024).
Holbert, C. E., Cullen, M. T., Casero, R. A. Jr & Stewart, T. M. Polyamines in cancer: integrating organismal metabolism and antitumour immunity. Nat. Rev. Cancer 22, 467–480 (2022).
Casero, R. A. Jr, Murray Stewart, T. & Pegg, A. E. Polyamine metabolism and cancer: treatments, challenges and opportunities. Nat. Rev. Cancer 18, 681–695 (2018).
Han, X. et al. Activation of polyamine catabolism promotes glutamine metabolism and creates a targetable vulnerability in lung cancer. Proc. Natl Acad. Sci. USA 121, e2319429121 (2024).
Bi, G. et al. Polyamine-mediated ferroptosis amplification acts as a targetable vulnerability in cancer. Nat. Commun. 15, 2461 (2024).
Novita Sari, I. et al. Metabolism and function of polyamines in cancer progression. Cancer Lett. 519, 91–104 (2021).
Casero, R. A. Jr & Woster, P. M. Recent advances in the development of polyamine analogues as antitumor agents. J. Med Chem. 52, 4551–4573 (2009).
Hanahan, D. Hallmarks of Cancer: New Dimensions. Cancer Discov. 12, 31–46 (2022).
Jiang, X. et al. Cellular metabolism: A key player in cancer ferroptosis. Cancer Commun. (Lond.) 44, 185–204 (2024).
Mayers, J. R. et al. Tissue of origin dictates branched-chain amino acid metabolism in mutant Kras-driven cancers. Science 353, 1161–1165 (2016).
Yuneva, M. O. et al. The metabolic profile of tumors depends on both the responsible genetic lesion and tissue type. Cell Metab. 15, 157–170 (2012).
Kelly, M. R. et al. Combined Proteomic and Genetic Interaction Mapping Reveals New RAS Effector Pathways and Susceptibilities. Cancer Discov. 10, 1950–1967 (2020).
Ivanova, O. N. et al. Polyamine Catabolism Revisited: Acetylpolyamine Oxidase Plays a Minor Role due to Low Expression. Cells 13, 1134 (2024).
Hegde, S. S., Chandler, J., Vetting, M. W., Yu, M. & Blanchard, J. S. Mechanistic and structural analysis of human spermidine/spermine N1-acetyltransferase. Biochemistry 46, 7187–7195 (2007).
Murray Stewart, T., Dunston, T. T., Woster, P. M. & Casero, R. A. Jr.: Polyamine catabolism and oxidative damage. J. Biol. Chem. 293, 18736–18745 (2018).
Baird, L. & Yamamoto, M. The Molecular Mechanisms Regulating the KEAP1-NRF2 Pathway. Mol. Cell Biol. 40, e00099-20 (2020).
Jaramillo, M. C. & Zhang, D. D. The emerging role of the Nrf2-Keap1 signaling pathway in cancer. Genes Dev. 27, 2179–2191 (2013).
Wang, Y., Devereux, W., Stewart, T. M. & Casero, R. A. Jr.: Cloning and characterization of human polyamine-modulated factor-1, a transcriptional cofactor that regulates the transcription of the spermidine/spermine N(1)-acetyltransferase gene. J. Biol. Chem. 274, 22095–22101 (1999).
Wang, Y., Xiao, L., Thiagalingam, A., Nelkin, B. D. & Casero, R. A. Jr.: The identification of a cis-element and a trans-acting factor involved in the response to polyamines and polyamine analogues in the regulation of the human spermidine/spermine N1-acetyltransferase gene transcription. J. Biol. Chem. 273, 34623–34630 (1998).
Chen, Y. et al. c-Jun NH(2) -Terminal Protein Kinase Phosphorylates the Nrf2-ECH Homology 6 Domain of Nuclear Factor Erythroid 2-Related Factor 2 and Downregulates Cytoprotective Genes in Acetaminophen-Induced Liver Injury in Mice. Hepatology 71, 1787–1801 (2020).
Yu, M. et al. Nuclear factor p65 interacts with Keap1 to repress the Nrf2-ARE pathway. Cell Signal 23, 883–892 (2011).
Jiang, Z. Y. et al. Insight into the intermolecular recognition mechanism between Keap1 and IKKβ combining homology modelling, protein-protein docking, molecular dynamics simulations and virtual alanine mutation. PLoS One 8, e75076 (2013).
Narkiewicz, J. et al. Changes in mRNA and protein levels of human HtrA1, HtrA2 and HtrA3 in ovarian cancer. Clin. Biochem 41, 561–569 (2008).
Santarpia, L., Lippman, S. M. & El-Naggar, A. K. Targeting the MAPK-RAS-RAF signaling pathway in cancer therapy. Expert Opin. Ther. Targets 16, 103–119 (2012).
Miller, M. A. et al. Reduced Proteolytic Shedding of Receptor Tyrosine Kinases Is a Post-Translational Mechanism of Kinase Inhibitor Resistance. Cancer Discov. 6, 382–399 (2016).
Awad, M. M. et al. Acquired Resistance to KRAS(G12C) Inhibition in Cancer. N. Engl. J. Med 384, 2382–2393 (2021).
Mechanisms of KRAS Inhibitor Resistance Revealed. Cancer Discov. 11, 1311–1312 (2021).
Zhao, Y. et al. Diverse alterations associated with resistance to KRAS(G12C) inhibition. Nature 599, 679–683 (2021).
Huang, K., Wang, Y. & Roberts, T. M. ERK-dependent protein phosphorylation in KRAS-mutant cancer: a mix of the expected and surprising. Trends Biochem. Sci. 50, 6–8 (2024).
Loi, P. et al. Epigenetic and Oncogenic Inhibitors Cooperatively Drive Differentiation and Kill KRAS-Mutant Colorectal Cancers. Cancer Discov. 14, 2430–2449 (2024).
Hadian, K. & Stockwell, B. R. The therapeutic potential of targeting regulated non-apoptotic cell death. Nat. Rev. Drug Discov. 22, 723–742 (2023).
Son, J. et al. Glutamine supports pancreatic cancer growth through a KRAS-regulated metabolic pathway. Nature 496, 101–105 (2013).
Lim, J. K. M. et al. Cystine/glutamate antiporter xCT (SLC7A11) facilitates oncogenic RAS transformation by preserving intracellular redox balance. Proc. Natl Acad. Sci. USA 116, 9433–9442 (2019).
Padanad, M. S. et al. Fatty Acid Oxidation Mediated by Acyl-CoA Synthetase Long Chain 3 Is Required for Mutant KRAS Lung Tumorigenesis. Cell Rep. 16, 1614–1628 (2016).
Yang, J., Dai, X., Xu, H., Tang, Q. & Bi, F. Regulation of Ferroptosis by Amino Acid Metabolism in Cancer. Int J. Biol. Sci. 18, 1695–1705 (2022).
Liang, M. et al. l-Arginine induces antioxidant response to prevent oxidative stress via stimulation of glutathione synthesis and activation of Nrf2 pathway. Food Chem. Toxicol. 115, 315–328 (2018).
Casero, R. A. Jr. & Pegg, A. E. Spermidine/spermine N1-acetyltransferase-the turning point in polyamine metabolism. Faseb j. 7, 653–661 (1993).
Pegg, A. E. Spermidine/spermine-N(1)-acetyltransferase: a key metabolic regulator. Am. J. Physiol. Endocrinol. Metab. 294, E995–E1010 (2008).
Bi, G. et al. Retinol Saturase Mediates Retinoid Metabolism to Impair a Ferroptosis Defense System in Cancer Cells. Cancer Res 83, 2387–2404 (2023).
Ou, Y., Wang, S. J., Li, D., Chu, B. & Gu, W. Activation of SAT1 engages polyamine metabolism with p53-mediated ferroptotic responses. Proc. Natl Acad. Sci. USA 113, E6806–e6812 (2016).
Affronti, H. C. et al. Pharmacological polyamine catabolism upregulation with methionine salvage pathway inhibition as an effective prostate cancer therapy. Nat. Commun. 11, 52 (2020).
Murray-Stewart, T. et al. Jr.: Curcumin mediates polyamine metabolism and sensitizes gastrointestinal cancer cells to antitumor polyamine-targeted therapies. PLoS One 13, e0202677 (2018).
Casero, R. A. Jr. & Marton, L. J. Targeting polyamine metabolism and function in cancer and other hyperproliferative diseases. Nat. Rev. Drug Discov. 6, 373–390 (2007).
Skoulidis, F. & Heymach, J. V. Co-occurring genomic alterations in non-small-cell lung cancer biology and therapy. Nat. Rev. Cancer 19, 495–509 (2019).
Arbour, K. C. et al. Effects of Co-occurring Genomic Alterations on Outcomes in Patients with KRAS-Mutant Non-Small Cell Lung Cancer. Clin. Cancer Res 24, 334–340 (2018).
Skoulidis, F. et al. Sotorasib for Lung Cancers with KRAS p.G12C Mutation. N. Engl. J. Med 384, 2371–2381 (2021).
Jänne, P. A. et al. Adagrasib in Non-Small-Cell Lung Cancer Harboring a KRAS(G12C) Mutation. N. Engl. J. Med 387, 120–131 (2022).
Skoulidis, F. et al. CTLA4 blockade abrogates KEAP1/STK11-related resistance to PD-(L)1 inhibitors. Nature 635, 462–471 (2024).
Pillai, R., Hayashi, M., Zavitsanou, A. M. & Papagiannakopoulos, T. NRF2: KEAPing Tumors Protected. Cancer Discov. 12, 625–643 (2022).
Sun, X. et al. Activation of the p62-Keap1-NRF2 pathway protects against ferroptosis in hepatocellular carcinoma cells. Hepatology 63, 173–184 (2016).
Dodson, M., Castro-Portuguez, R. & Zhang, D. D. NRF2 plays a critical role in mitigating lipid peroxidation and ferroptosis. Redox Biol. 23, 101107 (2019).
Sivandzade, F., Prasad, S., Bhalerao, A. & Cucullo, L. NRF2 and NF-қB interplay in cerebrovascular and neurodegenerative disorders: Molecular mechanisms and possible therapeutic approaches. Redox Biol. 21, 101059 (2019).
Jansen, R. A. et al. Small-molecule inhibition of MAP2K4 is synergistic with RAS inhibitors in KRAS-mutant cancers. Proc. Natl Acad. Sci. USA 121, e2319492121 (2024).
Fallahi-Sichani, M. et al. Systematic analysis of BRAF(V600E) melanomas reveals a role for JNK/c-Jun pathway in adaptive resistance to drug-induced apoptosis. Mol. Syst. Biol. 11, 797 (2015).
Wang, H. M. et al. Using patient-derived organoids to predict locally advanced or metastatic lung cancer tumor response: A real-world study. Cell Rep. Med 4, 100911 (2023).
Driehuis, E., Kretzschmar, K. & Clevers, H. Establishment of patient-derived cancer organoids for drug-screening applications. Nat. Protoc. 15, 3380–3409 (2020).
Johnson, L. et al. Somatic activation of the K-ras oncogene causes early onset lung cancer in mice. Nature 410, 1111–1116 (2001).
Canon, J. et al. The clinical KRAS(G12C) inhibitor AMG 510 drives anti-tumour immunity. Nature 575, 217–223 (2019).
Hirano, R., Shirasawa, H. & Kurihara S. Health-Promoting Effects of Dietary Polyamines. Med. Sci. (Basel) 9, 8 (2021).
Arthur, R., Jamwal, S. & Kumar, P. A review on polyamines as promising next-generation neuroprotective and anti-aging therapy. Eur. J. Pharm. 978, 176804 (2024).
Fan, J., Feng, Z. & Chen, N. Spermidine as a target for cancer therapy. Pharm. Res 159, 104943 (2020).
Zheng, S. et al. SynergyFinder Plus: Toward Better Interpretation and Annotation of Drug Combination Screening Datasets. Genomics Proteom. Bioinforma. 20, 587–596 (2022).
Bian, Y. et al. Integrative genome-scale analysis of immune infiltration in esophageal carcinoma. Int Immunopharmacol. 93, 107371 (2021).
Murphy, M. P. et al. Guidelines for measuring reactive oxygen species and oxidative damage in cells and in vivo. Nat. Metab. 4, 651–662 (2022).
Liang, J. et al. MAFF confers vulnerability to cisplatin-based and ionizing radiation treatments by modulating ferroptosis and cell cycle progression in lung adenocarcinoma. Drug Resist Updat 73, 101057 (2024).
Acknowledgements
This work was supported by grants from The National Natural Science Foundation of China (Nos. 82473184, to C.Z., 82403100, to G.B.), the Natural Science Foundation of Shanghai (24ZR1409900, to G.B.), the China Postdoctoral Science Foundation (No. 2024M760555, to G.B.), and the Shanghai Sailing Program (Nos. 24YF2731100, to Q.S., 24YF2704100, to J.L.).
Author information
Authors and Affiliations
Contributions
(Ⅰ) Cheng Zhan and Wei Jiang contributed conceptualization and design. (Ⅱ) Wei Jiang, Qun Wang and Hong Fan contributed administrative support. (III) Yunyi Bian, Guangyao Shan and Guoshu Bi contributed experiments. (IV) Yunyi Bian, Zhijie Xu, Yuanliang Yan, Huan Zhang and Jiaqi Liang contributed collection and assembly of data. (V) Yunyi Bian, Wei Guo, Qihai Sui, Yanjun Yi, Haochun Shi, and Tao Lu contributed data analysis and interpretation. (VI) Yunyi Bian and Guangyao Shan contributed manuscript writing. (VII) All authors final contributed approval of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Jiyeon Kim and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Bian, Y., Shan, G., Bi, G. et al. Targeting polyamine metabolism and ferroptosis enhances the efficacy of KRAS-targeted therapy depending on KEAP1 status. Nat Commun 16, 9923 (2025). https://doi.org/10.1038/s41467-025-65441-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-65441-4